code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
from matplotlib import pyplot
from matplotlib import pyplot as plt
import seaborn as sns
from tqdm import tqdm
from scipy.stats import ttest_ind
from numpy import median
import pickle
# +
# Make dictionaries for converting between sequence IDs
# Need list of proteins for each family
enzyme_order_df = pd.read_excel("data/initial_enzymes_1.xlsx")
sp_df = pd.read_excel("data/sp_prot_translations.xls")
# Map true protein ids
enzyme_order_df['prot_seq_trunc'] = enzyme_order_df['Protein-met-sigp'].str[:80]
sp_df['prot_seq_trunc'] = sp_df['prot_seq'].str[1:81]
merge_df = enzyme_order_df.merge(sp_df, how='inner', on='prot_seq_trunc')
# Get shuffled enzyme ID list back out
enzIDkeydf = merge_df[['enzyme_id','enzyme_type','seqID', 'new_id']]
seq2enzyme = pd.Series(enzIDkeydf.enzyme_id.values, index= enzIDkeydf.seqID).to_dict()
seq2family = pd.Series(enzIDkeydf.enzyme_type.values, index= enzIDkeydf.seqID).to_dict()
seq2newid = pd.Series(enzIDkeydf.new_id.values, index= enzIDkeydf.seqID).to_dict()
newid2seq = pd.Series(enzIDkeydf.seqID.values, index= enzIDkeydf.new_id).to_dict()
# -
# Read in preprocessed data
df = pd.read_csv('data/preprocessed.csv')
df['run_index'] = [i for i in range(len(df))]
df = df.drop(columns=['Unnamed: 0'])
# +
# Make data tidy
tidy_df = df.melt(id_vars=['correct', 'protein_id', 'sp_id', 'run_label', 'true_prot_id', 'prot_correct', 'sp_correct', 'family', 'run_id', 'run_index'],
var_name='assay', value_name='assay_value')
tidy_df.rename(columns={'true_prot_id':'seqID'}, inplace=True)
tidy_df = tidy_df.sort_values(by = 'run_id', ascending = True).dropna()
# Rescreen Column remapping
remap_dict = {'amylase_2_10x': 'amylase_10x',
'amylase_1_10x': 'amylase_10x',
'positive_amy_10x': 'amylase_10x',
'positive_xyl_50x': 'xylanase_50x',
'xylanase_1_50x': 'xylanase_50x',
}
tidy_df['family'].replace(remap_dict, inplace=True)
# +
# Create new 'correct' column based on results from pairwise alignments, which found more SPs
def combine_cols(row):
if row["prot_correct"] == True and row["sp_correct"] == True:
return True
else:
return False
tidy_df['old_correct'] = tidy_df['correct'].values
tidy_df['correct'] = tidy_df.apply(lambda x: combine_cols(x), axis=1)
# -
rescreen_fams = [fam for fam in list(set(tidy_df['family'].values)) if fam[-2:] == '0x']
all_fams = [fam for fam in list(set(tidy_df['family'].values))]
reg_fams = [fam for fam in all_fams if fam not in rescreen_fams]
print(all_fams)
print(rescreen_fams)
print(reg_fams)
# Remove rescreens
tidy_df = tidy_df[tidy_df['run_id'] != '26_15-2']
tidy_df = tidy_df[tidy_df['family'].isin(reg_fams)]
set(tidy_df['family'].values)
# +
# Need list of proteins for each family
enzyme_order_df = pd.read_excel("data/initial_enzymes_1.xlsx")
sp_df = pd.read_excel("data/sp_prot_translations.xls")
# Map true protein ids
enzyme_order_df['prot_seq_trunc'] = enzyme_order_df['Protein-met-sigp'].str[:80]
sp_df['prot_seq_trunc'] = sp_df['prot_seq'].str[1:81]
merge_df = enzyme_order_df.merge(sp_df, how='inner', on='prot_seq_trunc')
merge_df[['enzyme_id','enzyme_type','seqID', 'new_id']]
# Get shuffled enzyme ID list back out
enzIDkeydf = merge_df[['enzyme_id','enzyme_type','seqID', 'new_id']]
# -
tidy_df.head(2)
prot_fams = ['amylase', 'lipase', 'protease', 'xylanase']
# ## Visualize negative controls
#
# These constructs either did not have a SP or enzymatic sequence post Golden Gate Assembly.
for fam in prot_fams:
enzkey = enzIDkeydf[enzIDkeydf['enzyme_type'] == fam]
fam_seqIDs = list(set(enzkey['seqID'].values)) # Shuffled seqIDs
# Entries to search for within df
search_fams = [fam, 'positive', 'negative'] # also search positive/negative libraries
# Pick points that are in the correct family and are the correct seqIDs
fam_df = tidy_df[(tidy_df['family'].isin(search_fams)) & tidy_df['seqID'].isin(fam_seqIDs)].copy()
# Take incorrect constructs as negative controls
false_construct_df = fam_df[fam_df['correct'] == False]
random_construct_df = fam_df[fam_df['family'] == 'negative']
neg_df = pd.concat([false_construct_df, random_construct_df])
# neg_df = neg_df.assign(run_id='neg_controls')
neg_values = neg_df['assay_value'].values
neg_avg = np.average(neg_values)
try:
fig, ax = pyplot.subplots(figsize=(6,3))
chart = sns.swarmplot(ax=ax, x="run_id", y="assay_value", data=neg_df)
chart.set_title(fam + " controls")
chart.set_xticklabels(
chart.get_xticklabels(),
rotation=90,
horizontalalignment='right',
fontweight='light',
fontsize='large'
)
chart.xaxis.label.set_visible(False)
except ValueError:
print(f"{fam} cannot be plotted")
tidy_df[tidy_df['seqID']=='seq19']
# ## Now looking at all variants
# #### Get proteins that have some level of function (by seqID)
# Defined as having a mean activity higher than that of incorrect constructs, and also have a p-value < 0.01 as compared to incorrect constructs. This does include separate into individual runs.
# +
### Get proteins that have some level of function (by seqID)
p_cutoff = 0.05
effect_size = 2
alive_proteins = set() #seqIDs that have some function on some construct.
significant_count, total_count = 0,0
for fam in prot_fams:
enzkey = enzIDkeydf[enzIDkeydf['enzyme_type'] == fam]
fam_seqIDs = list(set(enzkey['seqID'].values)) # Shuffled seqIDs
# Entries to search for within df
search_fams = [fam, 'positive', 'negative'] # also search positive/negative libraries
# Pick points that are in the correct family and are the correct seqIDs
fam_df = tidy_df[(tidy_df['family'].isin(search_fams)) & tidy_df['seqID'].isin(fam_seqIDs)].copy()
# Take incorrect constructs as negative controls
false_construct_df = fam_df[fam_df['correct'] == False]
neg_values = false_construct_df['assay_value'].values
neg_avg = np.average(neg_values)
max_diff = np.max(neg_values) - neg_avg
cutoff = neg_avg + effect_size * max_diff
for run_id in set(fam_df['run_id'].values):
_df = fam_df[fam_df['run_id'] == run_id]
curr_values = _df['assay_value'].values
curr_prot = _df.iloc[0]['seqID']
p = ttest_ind(curr_values, neg_values, equal_var=False).pvalue
if np.average(curr_values) > cutoff and p < p_cutoff:
if curr_prot not in alive_proteins:
print(curr_prot, p, neg_avg, np.average(curr_values))
alive_proteins.add(curr_prot)
alive_proteins = list(alive_proteins)
alive_proteins.sort()
print(len(alive_proteins), alive_proteins)
# +
# Update with new plot_ids
seq2newid2 = {k[3:]:v for k,v in seq2newid.items()}
seq2newid.update({
'003':'AprE', '021':'LipB', '066':'YbdG', '093':'YkvV', '144':'YvcE', '193':'YcnJ'
})
seq2newid2.update({
'003':'AprE', '021':'LipB', '066':'YbdG', '093':'YkvV', '144':'YvcE', '193':'YcnJ'
})
pickle.dump(seq2newid2, open('data/final_plot_ids_2.p','wb'))
seq2newid2 = pickle.load(open("data/final_plot_ids_2.p", "rb"))
sp_plot_id = dict()
for i in range(41):
for j in range(4):
key = 'sps' + str(i+<KEY>)
value = i*4 + j + 1
sp_plot_id.update({key:f"Gen_{value:03d}"})
sp_plot_id
with open('data/final_sp_plot_dict.p','wb') as f:
pickle.dump(sp_plot_id, f)
with open('data/final_sp_plot_dict.p','rb') as f:
sp_plot_id = pickle.load(f)
def get_new_id(row, include_prot=False):
if row['run_id'] == 'false_construct':
return 'incorrect construct'
else:
old_id = row['run_id'].split('_')
if len(old_id) == 3: # then it's a control
prot, sp, _ = old_id
if old_id[-1] == 'neg':
new_id = 'random_' + sp
elif old_id[-1] == 'pos':
try:
new_id = seq2newid2[sp]
except KeyError:
new_id = 'incorrect construct pos'
# print(old_id)
else:
raise ImplementationError
elif len(old_id) == 2:
# prot, sp = old_id
# sp_ref, sp_num = sp.split('-')
# new_id = seq2newid2[sp_ref] + '-' + sp_num
new_id = sp_plot_id[row['sp_id']]
else:
raise ImplementationError
if include_prot:
new_id = seq2newid2[prot] + '_' + new_id
return new_id
tidy_df['plot_id'] = df.apply(lambda row: get_new_id(row), axis=1)
tidy_df[(tidy_df['plot_id'] == 'incorrect_construct_pos') & (tidy_df['correct']==True)]
# +
p_cutoff = 0.05
effect_size = 2.0
significant_count, total_count = 0,0
all_func_df = pd.DataFrame()
# all_nonfunc_df = pd.DataFrame()
all_func_constructs = []
all_nonfunc_constructs = []
run_ix_classification = dict()
for fam in prot_fams:
enzkey = enzIDkeydf[enzIDkeydf['enzyme_type'] == fam]
# Union with functional prots!!!
fam_seqIDs = list(set(enzkey['seqID'].values).intersection(set(alive_proteins)))
# Entries to search for within df
search_fams = [fam, 'positive', 'negative'] # also search positive/negative libraries
# Pick points that are in the correct family and are the correct seqIDs
fam_seqIDs.sort()
# print(set(enzkey['seqID'].values))
# print(fam_seqIDs)
fam_df = tidy_df[(tidy_df['family'].isin(search_fams)) & tidy_df['seqID'].isin(fam_seqIDs)] # .copy()
fam_df = fam_df[fam_df['correct'] == True]
fam_run_ids = list(set(fam_df['run_id'].values))
# Take incorrect constructs as negative controls, also get cutoff for p-value
all_in_family_df = tidy_df[(tidy_df['family'].isin(search_fams)) &
(tidy_df['seqID'].isin(list(set(enzkey['seqID'].values))))].copy()
false_construct_df = all_in_family_df[all_in_family_df['correct']==False].copy()
false_construct_df['run_id'] = 'false_construct'
neg_values = false_construct_df['assay_value'].values
neg_avg = np.average(neg_values)
max_diff = np.max(neg_values) - neg_avg
cutoff = neg_avg + (effect_size*max_diff)
func_constructs = set()
# nonfunc_constructs = set()
for run_ix in set(fam_df['run_index'].values):
_df = fam_df[fam_df['run_index'] == run_ix]
# Classify as functioning or not
curr_values = _df['assay_value'].values
p = ttest_ind(curr_values, neg_values, equal_var=True).pvalue
# Store both run_id for plot categories, and run_ix for subcategory hues
if np.average(curr_values) > cutoff and p < p_cutoff:
func_constructs.add(_df.iloc[0]['run_id']) # Add to functional constructs
run_ix_classification.update({run_ix:'Func Replicate'})
assert _df.iloc[0]['correct']==True, 'need to reconsider correct constructs'
else:
run_ix_classification.update({run_ix:'NonFunc Replicate'})
# Get nonfunctional constructs
nonfunc_constructs = [seqid for seqid in fam_df['run_id'] if seqid not in func_constructs]
all_func_constructs = all_func_constructs + list(func_constructs)
all_nonfunc_constructs = all_nonfunc_constructs + list(nonfunc_constructs)
# Separate plot for each enzyme
for i, seqid in enumerate(fam_seqIDs):
# Get functional constructs
correct_df = tidy_df[(tidy_df['correct']==True) & (tidy_df['seqID'] == seqid)].copy()
func_df = correct_df[correct_df['run_id'].isin(func_constructs)].copy()
# Get nonfunctional constructs
nonfunc_df = correct_df[correct_df['run_id'].isin(nonfunc_constructs)].copy()
# Save to all_func_df for downstream visualization/stats
func_df['func'] = True
all_func_df = all_func_df.append(func_df, ignore_index=True, sort=False)
nonfunc_df['func'] = False
all_func_df = all_func_df.append(nonfunc_df, ignore_index=True, sort=False)
# Add hues to appropriate portions for plotting
func_df['hue_class'] = func_df['run_index'].map(run_ix_classification) # functional has two
false_construct_df['hue_class'] = 'Incorrect Construct'
nonfunc_df['hue_class'] = 'NonFunc Construct'
plot_df = pd.concat([func_df, false_construct_df, nonfunc_df], sort=False)
plot_df['new_plot_id'] = plot_df.apply(lambda row: get_new_id(row), axis=1)
# if 'positive' in plot_df['family'].values:
# Plot
sns.set(font='helvetica', context='notebook', font_scale=1, style='ticks')
f = plt.figure(figsize=(8,7))
color_palette_dict = {'Func Replicate':sns.xkcd_rgb["green"],
'NonFunc Replicate':sns.xkcd_rgb["nice blue"],
'Incorrect Construct':sns.xkcd_rgb["black"],
'NonFunc Construct':sns.xkcd_rgb["steel grey"]}
chart = sns.swarmplot(x="new_plot_id", y="assay_value", hue='hue_class', data=plot_df, palette=color_palette_dict, size=6) #
title = seq2family[seqid].capitalize() + ' ' + seq2enzyme[seqid]
title = seq2newid[func_df.iloc[0]['seqID']]
if title[:3] == 'Pro':
title = 'Protease ' + title[4:]
elif title[:3] == 'Xyl':
title = 'Xylanase ' + title[4:]
elif title[:3] == 'Lip':
title = 'Lipase ' + title[4:]
elif title[:3] == 'Amy':
title = 'Amylase ' + title[4:]
# title += " : func | pos || negs || nonfunc"
plt.xlabel('Signal Peptide', fontsize=14)
plt.ylabel('Assay Value', fontsize=14)
plt.title(title, fontsize=16)
# chart.set_title(title)
chart.set_xticklabels(
chart.get_xticklabels(),
rotation=90,
horizontalalignment='right',
fontweight='light',
fontsize='medium')
plt.tight_layout()
plt.legend().set_title('Classification')
# plt.legend().set_visible(False)
display(chart)
plt.savefig('data/figs/' + seq2family[seqid] + '_' + seq2enzyme[seqid] + '.svg')
pickle.dump(run_ix_classification, open('data/200225_runix_classification.p', 'wb'))
# -
# Save functionality classification to csv
filename = 'data/func_class_p'+str(p_cutoff) + '_effectsize'+str(effect_size) +'.csv'
all_func_df.to_csv(filename)
on_func_df = all_func_df[all_func_df['func'] == True]
func_gen_sps = list(set(on_func_df[~on_func_df['family'].isin(['positive', 'negative'])]['sp_id'].values))
func_gen_sps.sort()
len(func_gen_sps)
off_func_df = all_func_df[all_func_df['func'] == False]
nonfunc_gen_sps = list(set(off_func_df[~off_func_df['family'].isin(['positive', 'negative'])]['sp_id'].values))
nonfunc_gen_sps.sort()
len(nonfunc_gen_sps)
# +
sp_df = pd.read_excel("data/sp_prot_translations.xls")
spid_to_seq = dict()
for sp in func_gen_sps+nonfunc_gen_sps:
seqID = 'seq' + sp[3:-2]
version = sp[-1]
if version == '1':
seq = sp_df[sp_df['seqID']==seqID].iloc[0]['75'][:-6]
elif version == '2':
seq = sp_df[sp_df['seqID']==seqID].iloc[0]['90'][:-6]
elif version == '3':
seq = sp_df[sp_df['seqID']==seqID].iloc[0]['95'][:-6]
elif version == '4':
seq = sp_df[sp_df['seqID']==seqID].iloc[0]['99'][:-6]
spid_to_seq.update({sp:seq})
func_sps = pd.DataFrame({'spid':func_gen_sps, 'seq':[spid_to_seq[s] for s in func_gen_sps]})
# -
func_sps.to_csv('outputs/functional_gen_sps_200129.csv')
# ### Check sequence distribution
# #### for only nonfunctional and sometimes or always functional generated SPs
func_sps
off_func_df = all_func_df[all_func_df['func'] == False]
nonfunc_gen_sps = list(set(off_func_df[~off_func_df['family'].isin(['positive', 'negative'])]['sp_id'].values))
nonfunc_gen_sps.sort()
len(nonfunc_gen_sps)
overlap = 0
nonfunc = 0
only_nonfunc_gen_sps = []
for sp in nonfunc_gen_sps:
if sp in func_gen_sps:
overlap += 1
else:
nonfunc += 1
only_nonfunc_gen_sps.append(sp)
print(overlap, nonfunc)
len(func_gen_sps)
assert len(set(func_gen_sps + only_nonfunc_gen_sps)) == \
len(set(func_gen_sps)) + len(set(only_nonfunc_gen_sps)), 'error, check for ONLY nonfunc'
func_data = [[sp, spid_to_seq[sp], True] for sp in func_gen_sps]
nonfunc_data = [[sp, spid_to_seq[sp], False] for sp in only_nonfunc_gen_sps]
classification_data = func_data+nonfunc_data
class_df = pd.DataFrame(classification_data, columns = ['spid', 'seq', 'any_functional'])
class_df
class_df.to_csv('outputs/any_functionality_classification.csv')
only_nonfunc_gen_sps
# +
# t = [spid_to_seq[s] for s in nonfunc_gen_sps]
# t.sort()
# for i in t:
# print(i)
# -
| experimental_validation/2a_negative_control_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.feature_extraction import DictVectorizer
from sklearn.tree import DecisionTreeClassifier
# #### 加载数据
# 加载数据
train_data = pd.read_csv('./train.csv')
test_data= pd.read_csv('./test.csv')
# ### 准备阶段
# #### 数据探索
train_data.info()
train_data.describe()
# 使用 describe(include=[‘O’]) 查看字符串类型(非数字)的整体情况
train_data.describe(include=['O'])
train_data.head()
train_data.tail()
# #### 数据清洗
# 使用平均年龄来填充年龄中的 nan 值
train_data['Age'].fillna(train_data['Age'].mean(), inplace=True)
test_data['Age'].fillna(test_data['Age'].mean(), inplace=True)
# 使用票价的均值填充票价中的 nan 值
train_data['Fare'].fillna(train_data['Fare'].mean(), inplace=True)
test_data['Fare'].fillna(test_data['Fare'].mean(), inplace=True)
train_data['Embarked'].value_counts()
# 使用登录最多的港口来填充登录港口的 nan 值
train_data['Embarked'].fillna('S', inplace=True)
test_data['Embarked'].fillna('S', inplace=True)
# #### 特征选择
# 特征选择是分类器的关键。特征选择不同,得到的分类器也不同。那么我们该选择哪些特征做生存的预测呢?
#
# 通过数据探索我们发现,PassengerId 为乘客编号,对分类没有作用,可以放弃;Name 为乘客姓名,对分类没有作用,可以放弃;Cabin 字段缺失值太多,可以放弃;Ticket 字段为船票号码,杂乱无章且无规律,可以放弃。其余的字段包括:Pclass、Sex、Age、SibSp、Parch 和 Fare,这些属性分别表示了乘客的船票等级、性别、年龄、亲戚数量以及船票价格,可能会和乘客的生存预测分类有关系。具体是什么关系,我们可以交给分类器来处理。
#
# 因此我们先将 Pclass、Sex、Age 等这些其余的字段作特征,放到特征向量 features 里。
features = ['Pclass', 'Sex', 'Age', 'SibSp', 'Parch', 'Fare', 'Embarked']
train_features = train_data[features]
train_labels = train_data['Survived']
test_features = test_data[features]
# #### 特征工程-数据标准化
# DictVectorizer 类,用它将可以处理符号化的对象,将符号转成数字 0/1 进行表示。
devc = DictVectorizer(sparse=False)
train_features = devc.fit_transform(train_features.to_dict(orient='record'))
devc.feature_names_
# ### 分类阶段
# #### 决策树模型
# 构造 ID3 决策树
clf = DecisionTreeClassifier(criterion='entropy')
# 决策树训练
clf.fit(train_features, train_labels)
# #### 模型评估&预测
test_features = devc.transform(test_features.to_dict(orient='record'))
# 决策树预测
pred_labels = clf.predict(test_features)
# 得到决策树准确率
acc_decision_tree = clf.score(train_features, train_labels)
print('score 准确率为 %.4f' % acc_decision_tree)
test_data.head()
# ##### 使用K折交叉验证 统计决策树准确率
import numpy as np
from sklearn.model_selection import cross_val_score
'cross_val_score准确率为 %.4f' % np.mean(
cross_val_score(clf, train_features, train_labels, cv=10))
# #### 使用AdaBoost 模型预测
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=200)
ada.fit(train_features, train_labels)
'ada cross_val_score准确率为 %.4f' % np.mean(
cross_val_score(ada, train_features, train_labels, cv=10))
| 35-AdaBoost(下):如何使用AdaBoost对房价进行预测?/titanic_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
# %matplotlib inline
from nltk.sentiment.vader import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
from newsapi import NewsApiClient
import nltk
api_key = os.getenv("NEWS_API_KEY")
newsapi = NewsApiClient(api_key=api_key)
# Fetch the Corona Virus news articles
coronavirus_headlines = newsapi.get_everything(q='coronavirus', language='en')
#
print(f"Total Articles about CoronaVirus: {coronavirus_headlines['totalResults']}")
# +
# Create Sentiment Dataframe
corona_virus_sentiments = []
for article in coronavirus_headlines['articles']:
try:
text = article['content']
date = article['publishedAt']
sentiment = analyzer.polarity_scores(text)
compound = sentiment['compound']
pos = sentiment['pos']
neu = sentiment['neu']
neg = sentiment['neg']
corona_virus_sentiments.append({
'Text': text,
'Compound': compound,
'Positive': pos,
'Negative': neg,
'Neutral': neu
})
except AttributeError:
pass
corona_df = pd.DataFrame(corona_virus_sentiments)
# Reorder Columns
columns = ['Compound', 'Negative', 'Neutral', 'Positive', 'Text']
corona_df = corona_df[columns]
corona_df.head()
# -
#Describe Cornoa df
corona_df.describe()
# Tokenize Ariticles
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer, PorterStemmer
from string import punctuation
import re
# +
# Addtion of stop words
sw_addon = {'u', 'f', 'ha'}
# Complete the tokenizer function
def tokenizer(text):
"""Tokenizes text."""
# Remove the stop words
sw = set(stopwords.words('english'))
# Remove the punctuation
regex = re.compile("[^a-zA-Z ]")
re_clean = regex.sub('', text)
# Tokenize clean version of words
lemmatizer = WordNetLemmatizer()
words = word_tokenize(re_clean)
# Lemmatize Words into root words
lem = [lemmatizer.lemmatize(word) for word in words]
# Convert the words to lowercase
tokens = [word.lower() for word in lem if word.lower() not in sw.union(sw_addon)]
return tokens
#tokens
# +
# Create a new tokens column for bitcoin
coronavrus_tokens = corona_df.copy()
tokenized = []
for i in coronavrus_tokens['Text']:
tokenized_articles = tokenizer(i)
tokenized.append({'tokens': tokenized_articles})
coronavrus_tokens_df = pd.DataFrame(tokenized)
coronavrus_tokens['Tokens'] = coronavrus_tokens_df
coronavrus_tokens.head()
# -
# # NGRAMS AND FREQUENCY ANALYSIS
from collections import Counter
from nltk import ngrams
# Generate the Corona N-grams where n=2
coronavirus_words = []
for text in coronavrus_tokens['Tokens']:
for word in text:
coronavirus_words.append(word)
corona_count = Counter(ngrams(coronavirus_words, n=2), ascending=True)
corona_count.most_common(20)
# Use the token_count function to generate the top 10 words from Corona Virus
tokens = [coronavrus_tokens]
def token_count(tokens, N=10):
"""Returns the top N tokens from the frequency count"""
return Counter(tokens).most_common(N)
# Get the top 10 words for Corona Virus
token_count(coronavirus_words)
# # Generate A WordCloud
#imports
from wordcloud import WordCloud
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = [20.0, 10.0]
# +
#Set the function
def process_text(corpus):
# Combine all articles in corpus into one large string
big_string = ' '.join(corpus)
return big_string
corona_word_cloud = process_text(coronavirus_words)
wc = WordCloud(colormap="RdYlBu").generate(corona_word_cloud)
plt.imshow(wc)
wc = WordCloud(width=1200, height=800, max_words = 50).generate(corona_word_cloud)
plt.title("Corona Virus Word Cloud", fontsize=40)
plt.imshow(wc)
# -
| Corona_Virus.ipynb |
# ---
# title: "pandas Time Series Basics"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "pandas time series basics."
# type: technical_note
# draft: false
# aliases:
# - /python/pandas_time_series_basics.html
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import modules
from datetime import datetime
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as pyplot
# ### Create a dataframe
data = {'date': ['2014-05-01 18:47:05.069722', '2014-05-01 18:47:05.119994', '2014-05-02 18:47:05.178768', '2014-05-02 18:47:05.230071', '2014-05-02 18:47:05.230071', '2014-05-02 18:47:05.280592', '2014-05-03 18:47:05.332662', '2014-05-03 18:47:05.385109', '2014-05-04 18:47:05.436523', '2014-05-04 18:47:05.486877'],
'battle_deaths': [34, 25, 26, 15, 15, 14, 26, 25, 62, 41]}
df = pd.DataFrame(data, columns = ['date', 'battle_deaths'])
print(df)
# ### Convert `df['date']` from string to datetime
df['date'] = pd.to_datetime(df['date'])
# ### Set `df['date']` as the index and delete the column
df.index = df['date']
del df['date']
df
# ### View all observations that occured in 2014
df['2014']
# ### View all observations that occured in May 2014
df['2014-05']
# ### Observations after May 3rd, 2014
df[datetime(2014, 5, 3):]
# ### Observations between May 3rd and May 4th
df['5/3/2014':'5/4/2014']
# ### Truncation observations after May 2nd 2014
df.truncate(after='5/3/2014')
# ### Observations of May 2014
df['5-2014']
# ### Count the number of observations per timestamp
df.groupby(level=0).count()
# ###
# ### Mean value of battle_deaths per day
df.resample('D').mean()
# ### Total value of battle_deaths per day
df.resample('D').sum()
# ### Plot of the total battle deaths per day
df.resample('D').sum().plot()
| docs/python/data_wrangling/pandas_time_series_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:research]
# language: python
# name: conda-env-research-py
# ---
# ## Redo the figures to compare the three characteristic functions
# %run -i startup2.py
bg_mpl_file = "~/PycharmProjects/bg-mpl-stylesheets/bg_mpl_stylesheet/bg_mpl_stylesheet"
plt.style.use(bg_mpl_file)
# ## Load the data
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
file_info = pd.read_csv("data_info.csv", index_col=0)
file_info
# + jupyter={"source_hidden": true} tags=[]
profile = io.load_profile(file_info["gr"]["JBNP33L"], {"qdamp":0.0313, "qbroad": 0.0131})
# -
# ## Fit the model using different characteristic functions
# ### Spherical bronze + Spherical anatase
# + jupyter={"source_hidden": true} tags=[]
model = create_model_spherical()
model.set_profile(profile)
model.load("results2/JBNP33L.txt")
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
model.set_order(["B_scale", "B_f_psize", "A_scale", "A_f_psize"])
model.set_xrange(1.6, 50., 0.1)
model.optimize()
model.set_xrange(1.6, 50., 0.01)
model.optimize()
model.set_order("all")
model.optimize()
model.plot()
# + jupyter={"source_hidden": true} tags=[]
model.update()
# + jupyter={"source_hidden": true} tags=[]
model.save_all("results3", "JBNP33L_two_spherical")
# + jupyter={"source_hidden": true} tags=[]
del model
# -
# ### Core bronze + Shell anatase
# + jupyter={"source_hidden": true} tags=[]
model = create_model_core_shell()
model.set_profile(profile)
model.load("results2/JBNP33L.txt")
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
model.set_order(["B_scale", "A_f_radius", "A_scale", "A_f_thickness"])
model.set_xrange(1.6, 50., 0.1)
model.optimize()
model.set_xrange(1.6, 50., 0.01)
model.optimize()
model.set_order("all")
model.optimize()
model.plot()
# + jupyter={"source_hidden": true} tags=[]
model.update()
# + jupyter={"source_hidden": true} tags=[]
model.save_all("results3", "JBNP33L_core_shell")
# + jupyter={"source_hidden": true} tags=[]
del model
# -
# ## Calculate the partial PDF of lognormal + spherical
# + jupyter={"source_hidden": true} tags=[]
model = create_model_with_ligand()
model.load("results3/JBNP33L.txt")
model.set_profile(profile)
model.set_xrange(0.01, 50.0, 0.01)
# + jupyter={"source_hidden": true} tags=[]
model.set_equation("B_f * B")
model.eval()
arr_b = model.export_fits()
# + jupyter={"source_hidden": true} tags=[]
model.set_equation("A_f * A")
model.eval()
arr_a = model.export_fits()
# + jupyter={"source_hidden": true} tags=[]
model.set_equation("L")
model.eval()
arr_l = model.export_fits()
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
arr = xr.concat([arr_b, arr_a, arr_l], dim="phase").assign_coords({"phase": ["bronze", "anatase", "ligand"]}).drop_dims(["xobs"]).drop_vars(["y"])["ycalc"]
arr
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
arr.plot(hue="phase");
# + jupyter={"source_hidden": true} tags=[]
arr.to_netcdf("summary/partial_pdfs_lognormal_spherical")
# -
# ## Summarize the results
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
ds = load_and_concat("results3", "JBNP33L*_fits.nc")
ds = ds.sortby("files").sel({"dim_0": [2, 0, 1]}).assign({"shape": (["dim_0"], ["spherical and spherical", "core and shell", "lognormal and spherical"])})
ds
# + jupyter={"source_hidden": true} tags=[]
ds.to_netcdf("summary/fits_different_shapes.nc")
# + jupyter={"source_hidden": true} tags=[]
del ds
# -
# ## Visualize the results
ds = xr.load_dataset("summary/fits_different_shapes.nc")
ds
partial = xr.load_dataarray("summary/partial_pdfs_lognormal_spherical")
partial
# +
def visualize_fits(axes, start=30, end=50, margin=1):
ds2 = ds.sel({"x": slice(start, end), "xobs": slice(start, end)})
n = len(axes)
for i in range(n):
md.plot_fits(ds2.sel(dim_0=i), offset=-0.02, ax=axes[i])
axes[i].set_xlim(start - margin, end + margin)
letters = "abc"
legends = ds2.shape.values
for i in range(n):
axes[i].set_title("")
axes[i].set_title("({})".format(letters[i]), x=0.05, y=0.9)
axes[i].annotate(legends[i], (0.45, 0.9), xycoords='axes fraction')
use_parentheses(axes[i])
return
def plot_partial_pdfs(ax):
texts=partial["phase"].values
ones = np.ones_like(partial.x)
offsets = np.stack([0 * ones, -0.85 * ones, -1.1 * ones])
ycalc = partial + offsets
ycalc.attrs = partial.attrs
ycalc.plot.line(hue="phase", ax=ax, add_legend=False)
coords = [(40, 0.2), (40, -0.75), (40, -1.03)]
colors = get_colors()
for i in range(3):
ax.annotate(texts[i], coords[i], color=colors[i])
ax.set_title("(d)", x=0.05, y=0.9)
use_parentheses(ax)
return
# fig, axes = plt.subplots(2, 2, figsize=(4 * 1.4 * 2, 3 * 1.4 * 2), gridspec_kw={"wspace": 0.0, "hspace": 0.0})
fig = plt.figure(figsize=(4 * 2 * 1.4, 3 * 2 * 1.4))
grids = mpl.gridspec.GridSpec(
2, 2,
wspace=0, hspace=0
)
axes = [None] * 4
axes[0] = fig.add_subplot(grids[0, 0])
axes[1] = fig.add_subplot(grids[0, 1])
axes[2] = fig.add_subplot(grids[1, 0])
axes[3] = fig.add_subplot(grids[1, 1])
# plot on axes
visualize_fits(axes[:3])
plot_partial_pdfs(axes[3])
# disable ticks and set ylim
axes[0].set_xlabel("")
axes[0].set_xticklabels([])
axes[1].set_xlabel("")
axes[1].set_xticklabels([])
axes[1].set_ylabel("")
axes[1].set_yticklabels([])
axes[3].set_ylabel("")
axes[3].set_yticklabels([])
ylim = (-0.19, 0.13)
for i in range(3):
axes[i].set_ylim(*ylim)
# show
plt.show()
# -
paper_exporter("compare_shapes.pdf", fig)
| analysis7.ipynb |
% -*- coding: utf-8 -*-
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% + [markdown] slideshow={"slide_type": "slide"}
%
% [**Dr. <NAME>**](mailto:<EMAIL>), _Lecturer in Biomedical Engineering_
%
% National University of Ireland Galway.
%
% ---
% + [markdown] slideshow={"slide_type": "skip"}
% \newpage
% + [markdown] slideshow={"slide_type": "slide"}
% # Learning objectives
% At the end of this lecture you should be able to:
%
% * Derive the finite element equations for thermal problems on 1D linear elastic trusses
% * Solve basic thermal problems for linear elastic trusses
%
% + [markdown] slideshow={"slide_type": "slide"}
% # The truss element (mechanical loads)
% 
%
% * Behaviour Hooke's law for a bar:
% $$\begin{Bmatrix} \sigma \end{Bmatrix}=E \begin{Bmatrix}\epsilon\end{Bmatrix}$$
% $$\begin{Bmatrix} f_{1} \\ f_{2} \end{Bmatrix}=\frac{AE}{L}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}\begin{Bmatrix} u_1 \\ u_2\end{Bmatrix}$$
%
% * Truss (or bar) is a 1D element but has a **cross-sectional area** $A$
%
% * **strain** a relative metric for deformation:
% $$\begin{Bmatrix} \varepsilon \end{Bmatrix}= \begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$$
%
% + [markdown] slideshow={"slide_type": "slide"}
% # The truss element (thermal loads)
%
% ## Adding termal strain
% 
%
% Define a total strain $\varepsilon_{tot}$ and split into a thermal $\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}$ and mechanical (elastic) strain $\begin{Bmatrix}\varepsilon_{e}\end{Bmatrix}$:
% $$\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}=\begin{Bmatrix}\varepsilon_e\end{Bmatrix}+\begin{Bmatrix}\varepsilon_0\end{Bmatrix}$$
% $$\begin{Bmatrix}\varepsilon_e\end{Bmatrix}=\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}-\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}$$
% and
% $$\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}=\alpha \Delta T \rightarrow \begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}=\frac{\begin{Bmatrix}\sigma\end{Bmatrix}}{E}+\alpha \Delta T=\begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$$
% $\alpha$: Thermal expansion coefficient
% $\Delta T$: Temperature change
% + [markdown] slideshow={"slide_type": "slide"}
% ## Developing the finite element equations
% Using the variational principle, the potential energy of a system can be expressed as:
% $$\Pi=\Lambda-W$$
% $\Pi$ : Potential energy of the system
% $\Lambda$ : The sum of internal strain energy
% $W$ : Work done by external forces
%
% + [markdown] slideshow={"slide_type": "slide"}
% Recall that the strain energy is derived through integration over the volume but this time features $\begin{Bmatrix}\varepsilon_e\end{Bmatrix}$ instead of $\begin{Bmatrix}\varepsilon\end{Bmatrix}$:
% $$\Lambda=\int_V \frac{\begin{Bmatrix}\sigma\end{Bmatrix} \begin{Bmatrix}\varepsilon_e\end{Bmatrix}}{2} dV$$
% Which using
% $$\begin{Bmatrix} \sigma \end{Bmatrix}=E \begin{Bmatrix}\varepsilon_e\end{Bmatrix}$$
% Leads to
% $$\Lambda=\int_V \frac{E\begin{Bmatrix}\varepsilon_e\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_e\end{Bmatrix}}{2} dV$$
% + [markdown] slideshow={"slide_type": "slide"}
% $$\Lambda=\int_V \frac{E\begin{Bmatrix}\varepsilon_e\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_e\end{Bmatrix}}{2} dV$$
%
% First we will split this equation up and use $\begin{Bmatrix}\varepsilon_e\end{Bmatrix}=\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}-\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}$ to note that:
% $$\frac{\varepsilon_{e}^2}{2}=\frac{(\varepsilon_{tot}-\varepsilon_{0})^2}{2}=\frac{\varepsilon_{tot}^2}{2}-\varepsilon_{0}\varepsilon_{tot}+\frac{\varepsilon_{0}^2}{2}$$
% Or for matrices:
% $$\frac{\begin{Bmatrix}\varepsilon_{e}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{e}\end{Bmatrix}}{2} =\frac{\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}}{2}-\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}+\frac{\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}}{2}$$
% Leading to:
% $$\Lambda=\int_V \frac{E\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}}{2} dV-\int_V E\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV + \int_V \frac{E\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}}{2} dV=a-b+c$$
%
%
% + [markdown] slideshow={"slide_type": "slide"}
% Using $\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}=\begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$, and $\big(\begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}\big)^{\top}=\begin{Bmatrix} u \end{Bmatrix}^{\top}\begin{bmatrix} B \end{bmatrix}^{\top}$ we can write:
%
% $$a=\int_V \frac{E\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}}{2} dV=\int_V \frac{E}{2}\begin{Bmatrix} u \end{Bmatrix}^{\top}\begin{bmatrix} B \end{bmatrix}^{\top} \begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix} dV$$
% $$ = \frac{1}{2} \begin{Bmatrix} u \end{Bmatrix}^{\top} \int_V E\begin{bmatrix} B \end{bmatrix}^{\top} \begin{bmatrix} B \end{bmatrix} dV \begin{Bmatrix} u \end{Bmatrix} $$
% $$ = \frac{1}{2}\begin{Bmatrix} u \end{Bmatrix}^{\top} \begin{bmatrix} K \end{bmatrix} \begin{Bmatrix} u \end{Bmatrix} $$
%
% + [markdown] slideshow={"slide_type": "slide"}
% Using $\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}=\begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$, and $\big(\begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}\big)^{\top}=\begin{Bmatrix} u \end{Bmatrix}^{\top}\begin{bmatrix} B \end{bmatrix}^{\top}$ we can write:
%
% $$b=\int_V E\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV=\int_V E \begin{Bmatrix} u \end{Bmatrix}^{\top}\begin{bmatrix} B \end{bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV$$
% $$=\begin{Bmatrix} u \end{Bmatrix}^{\top} \int_V \begin{bmatrix} B \end{bmatrix}^{\top} E \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV$$
% + [markdown] slideshow={"slide_type": "slide"}
% Leading to:
% $$\Lambda=a-b+c=\frac{1}{2}\begin{Bmatrix} u \end{Bmatrix}^{\top} \begin{bmatrix} K \end{bmatrix} \begin{Bmatrix} u \end{Bmatrix}-\begin{Bmatrix} u \end{Bmatrix}^{\top}\int_V \begin{bmatrix} B \end{bmatrix}^{\top} E \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV + \int_V \frac{E\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}}{2} dV$$
%
% Next recall solving for:
% $$\frac{\partial\Pi}{\partial \begin{Bmatrix} u \end{Bmatrix}}=0$$
% With:
% $$\partial\Pi=\Lambda-W$$
% $$W=\begin{Bmatrix} u \end{Bmatrix}^\top\begin{Bmatrix} F \end{Bmatrix}$$
% + [markdown] slideshow={"slide_type": "slide"}
% Leading to:
% $$\Lambda=a-b+c=\frac{1}{2}\begin{Bmatrix} u \end{Bmatrix}^{\top} \begin{bmatrix} K \end{bmatrix} \begin{Bmatrix} u \end{Bmatrix}-\begin{Bmatrix} u \end{Bmatrix}^{\top}\int_V \begin{bmatrix} B \end{bmatrix}^{\top} E \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV + \int_V \frac{E\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}^{\top} \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}}{2} dV$$
%
% Next recall solving for:
% $$\frac{\partial\Pi}{\partial \begin{Bmatrix} u \end{Bmatrix}}=0=\begin{bmatrix} K \end{bmatrix} \begin{Bmatrix} u \end{Bmatrix}- \int_V \begin{bmatrix} B \end{bmatrix}^{\top} E \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV - \begin{Bmatrix} F \end{Bmatrix}$$
% With:
% $$\Pi=\Lambda-W$$
% $$W=\begin{Bmatrix} u \end{Bmatrix}^\top \begin{Bmatrix} F \end{Bmatrix}$$
% + [markdown] slideshow={"slide_type": "slide"}
% Rearranging $\frac{\partial\Pi}{\partial \begin{Bmatrix} u \end{Bmatrix}}$ gives:
%
% $$\begin{bmatrix} K \end{bmatrix} \begin{Bmatrix} u \end{Bmatrix} = \int_V \begin{bmatrix} B \end{bmatrix}^{\top} E \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV + \begin{Bmatrix} F \end{Bmatrix}$$
% Next define $\begin{Bmatrix} F \end{Bmatrix}^{thermal}$:
% $$\begin{Bmatrix} F \end{Bmatrix}^{thermal} = \int_V \begin{bmatrix} B \end{bmatrix}^{\top} E \begin{Bmatrix}\varepsilon_{0}\end{Bmatrix} dV$$
%
% + [markdown] slideshow={"slide_type": "slide"}
% Using $\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}=\alpha \Delta T$, $\begin{bmatrix} B \end{bmatrix}^\top =\begin{bmatrix} -\frac{1}{L} \\ \frac{1}{L}\end{bmatrix}$,and $dV=AdX$ we can rewrite $\begin{Bmatrix} F \end{Bmatrix}^{thermal}$:
%
% $$\begin{Bmatrix} F \end{Bmatrix}^{thermal}=\int_0^L \begin{bmatrix} -\frac{1}{L} \\ \frac{1}{L}\end{bmatrix} E \alpha \Delta T A dx $$
%
% $$= \begin{bmatrix} -\frac{1}{L} \\ \frac{1}{L}\end{bmatrix} E \alpha \Delta T A \int_0^L dx = \begin{bmatrix} -\frac{1}{L} \\ \frac{1}{L}\end{bmatrix} E \alpha \Delta T A L $$
%
% $$\begin{Bmatrix} F \end{Bmatrix}^{thermal}=E \alpha \Delta T A \begin{bmatrix} -1 \\ 1 \end{bmatrix} $$
%
%
% + [markdown] slideshow={"slide_type": "slide"}
% # The truss element (mechanical loads)
% 
%
% * Behaviour Hooke's law for a bar:
% $$\begin{Bmatrix} \sigma \end{Bmatrix}=E \begin{Bmatrix}\epsilon\end{Bmatrix}$$
% $$\begin{Bmatrix} f_{1} \\ f_{2} \end{Bmatrix}=\frac{AE}{L}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}\begin{Bmatrix} u_1 \\ u_2\end{Bmatrix}$$
%
% * Truss (or bar) is a 1D element but has a **cross-sectional area** $A$
%
% * **strain** a relative metric for deformation:
% $$\begin{Bmatrix} \varepsilon \end{Bmatrix}= \begin{bmatrix} B \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$$
%
% + [markdown] slideshow={"slide_type": "slide"}
% # The truss element (thermal loads)
%
% ## Adding termal strain
% 
%
% $$\begin{Bmatrix} f_{1} \\ f_{2} \end{Bmatrix}+E \alpha \Delta T A \begin{bmatrix} -1 \\ 1 \end{bmatrix} =\frac{AE}{L}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}\begin{Bmatrix} u_1 \\ u_2\end{Bmatrix}$$
%
% Define a total strain $\varepsilon_{tot}$ and split into a thermal $\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}$ and mechanical (elastic) strain $\begin{Bmatrix}\varepsilon_{e}\end{Bmatrix}$:
% $$\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}=\begin{Bmatrix}\varepsilon_e\end{Bmatrix}+\begin{Bmatrix}\varepsilon_0\end{Bmatrix}$$
%
% $\alpha$: Thermal expansion coefficient
% $\Delta T$: Temperature change
% + [markdown] slideshow={"slide_type": "slide"}
% # Example 1: Thermal loads in a contrained set of trusses
% A 2 truss structure subjected to temperature variations
%
% 
%
% $$\begin{Bmatrix} f_{1} \\ f_{2} \end{Bmatrix}+E \alpha \Delta T A \begin{bmatrix} -1 \\ 1 \end{bmatrix} =\frac{AE}{L}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}\begin{Bmatrix} u_1 \\ u_2\end{Bmatrix}$$
%
% $E_1=2000 N/m^2$, $A_1=1 m^2$, $L_1=10 m$, $\alpha_1=2\cdot10^{-6} / ^{\circ}C$
% $E_2=2000 N/m^2$, $A_2=0.5 m^2$, $L_2=8 m$, $\alpha_2=2\cdot10^{-6} / ^{\circ}C$
% $\Delta T$: Temperature change
%
% Given that the neutral (zero force) temperature is 18 °C, determine the nodal displacements and forces at night and during the day.
% + [markdown] slideshow={"slide_type": "slide"}
% ## Compute thermal forces
%
% ### Day situation
% During the day $\Delta T=\Delta T_{day}=25-18=7$:
%
% $$\begin{Bmatrix} f_1^{(1)} \\ f_2^{(1)} \end{Bmatrix}^{thermal} = E_1 \alpha_1 \Delta T_{day} A_1 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = 2000 \cdot (2\cdot10^{-6}) \cdot 7 \cdot 1 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{Bmatrix} -0.028 \\ 0.028 \end{Bmatrix}$$
%
% $$\begin{Bmatrix} f_2^{(2)} \\ f_3^{(2)} \end{Bmatrix}^{thermal} = E_2 \alpha_2 \Delta T_{day} A_2 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = 2000 \cdot (2\cdot10^{-6}) \cdot 7 \cdot 0.5 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{Bmatrix} -0.014 \\ 0.014 \end{Bmatrix}$$
%
% Superposition:
% $$\begin{Bmatrix} F \end{Bmatrix}^{thermal} = \begin{Bmatrix} -0.028 \\ 0.028-0.014 \\ 0.014 \end{Bmatrix}= \begin{Bmatrix} -0.028 \\ 0.014 \\ 0.014 \end{Bmatrix}$$
%
% + [markdown] slideshow={"slide_type": "slide"}
% ### Night situation
% During the night $\Delta T=\Delta T_{night}=-2-18=-20$:
%
% $$\begin{Bmatrix} f_1^{(1)} \\ f_2^{(1)} \end{Bmatrix}^{thermal} = E_1 \alpha_1 \Delta T_{day} A_1 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = 2000 \cdot (2\cdot10^{-6}) \cdot -20 \cdot 1 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{Bmatrix} 0.08 \\ -0.08 \end{Bmatrix}$$
%
% $$\begin{Bmatrix} f_2^{(2)} \\ f_3^{(2)} \end{Bmatrix}^{thermal} = E_2 \alpha_2 \Delta T_{day} A_2 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = 2000 \cdot (2\cdot10^{-6}) \cdot -20 \cdot 0.5 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{Bmatrix} 0.04 \\ -0.04 \end{Bmatrix}$$
%
% Superposition:
% $$\begin{Bmatrix} F \end{Bmatrix}^{thermal} = \begin{Bmatrix} 0.08 \\ -0.08+0.04 \\ -0.04 \end{Bmatrix}= \begin{Bmatrix} 0.08 \\ -0.04 \\ -0.04 \end{Bmatrix}$$
% + slideshow={"slide_type": "skip"}
E1=2000; A1=1; L1=10; a1=2e-6;
E2=2000; A2=0.5; L2=8; a2=2e-6;
T_neutral=18;
T_day=25;
T_night=-2;
dT_day=T_day-T_neutral;
dT_night=T_night-T_neutral
f12_day = E1*a1*dT_day*A1*[-1;1]
f23_day = E2*a2*dT_day*A2*[-1;1]
f12_night = E1*a1*dT_night*A1*[-1;1]
f23_night = E2*a2*dT_night*A2*[-1;1]
% + [markdown] slideshow={"slide_type": "slide"}
% ## Set up element stiffness matrices
%
% $$ \begin{bmatrix} K^{(1)} \end{bmatrix}=\frac{A_1 E_1}{L_1}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\frac{1 \cdot 2000}{10}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\begin{bmatrix} 200 & -200 \\ -200 & 200\end{bmatrix}$$
%
% $$ \begin{bmatrix} K^{(2)} \end{bmatrix}=\frac{A_2 E_2}{L_2}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\frac{0.5 \cdot 2000}{8}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\begin{bmatrix} 125 & -125 \\ -125 & 125\end{bmatrix}$$
%
%
% + slideshow={"slide_type": "slide"}
% Numerical check
k1=(A1*E1)./L1;
k2=(A2*E2)./L2;
Q=[1 -1; -1 1];
K1=k1*Q
K2=k2*Q
% + [markdown] slideshow={"slide_type": "slide"}
% ## Use superposition to assemble total stiffness matrix
% $$\begin{bmatrix} K \end{bmatrix}=\underbrace{\begin{bmatrix} 200 & -200 & 0 \\ -200 & 200 & 0 \\ 0 & 0 & 0\end{bmatrix}}_{\begin{bmatrix} K^{(1)} \end{bmatrix}}+\underbrace{\begin{bmatrix} 0 & 0 & 0 \\ 0 & 125 & -125 \\ 0 & -125 & 125\end{bmatrix}}_{\begin{bmatrix} K^{(2)} \end{bmatrix}}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}$$
%
% + slideshow={"slide_type": "slide"}
% Numerical check
K=zeros(3,3);
K(1:2,1:2)=K(1:2,1:2)+K1;
K(2:3,2:3)=K(2:3,2:3)+K2
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solving for unknown forces and displacements during day
%
% $$\begin{Bmatrix} F \end{Bmatrix}+\begin{Bmatrix} F \end{Bmatrix}^{thermal}=\begin{bmatrix} K \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$$
%
% $$\begin{Bmatrix} F_1 \\ 0 \\ F_3 \end{Bmatrix}+\begin{Bmatrix} f_1^{(1)} \\ f_2^{(1)}+f_2^{(2)} \\ f_3^{(2)} \end{Bmatrix}^{thermal}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ 0 \end{Bmatrix}$$
%
% $$\begin{Bmatrix} F_1 \\ 0 \\ F_3 \end{Bmatrix}+\begin{Bmatrix} -0.028 \\ 0.014 \\ 0.014 \end{Bmatrix}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ 0 \end{Bmatrix}$$
%
% $$\rightarrow 0.014=325*u_2 \rightarrow u_2=\frac{0.014}{325}\approx 4.308\cdot10^{-5}$$
%
% + slideshow={"slide_type": "slide"}
% Numerical check
F_thermal_day=zeros(3,1);
F_thermal_day(1:2)=F_thermal_day(1:2)+f12_day;
F_thermal_day(2:3)=F_thermal_day(2:3)+f23_day
F2=0+F_thermal_day(2)
u2=F2./K(2,2)
% + [markdown] slideshow={"slide_type": "slide"}
% $$\begin{Bmatrix} F_1 \\ 0 \\ F_3 \end{Bmatrix}+\begin{Bmatrix} -0.028 \\ 0.014 \\ 0.014 \end{Bmatrix}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ 0 \end{Bmatrix}$$
%
% $$\rightarrow F_1-0.028=-200*u_2 \rightarrow F_1=-200 \cdot \frac{0.014}{325}+0.028 \approx 0.01939$$
% $$\rightarrow F_3+0.014=-125*u_2 \rightarrow F_3=-125 \cdot \frac{0.014}{325}-0.014 \approx -0.01939$$
%
% + slideshow={"slide_type": "slide"}
u=[0 u2 0]' %Displacements
F_total=K*u %Total forces
F=F_total-F_thermal_day %Direct forces
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solving for unknown forces and displacements during night
%
% $$\begin{Bmatrix} F \end{Bmatrix}+\begin{Bmatrix} F \end{Bmatrix}^{thermal}=\begin{bmatrix} K \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$$
%
% $$\begin{Bmatrix} F_1 \\ 0 \\ F_3 \end{Bmatrix}+\begin{Bmatrix} f_1^{(1)} \\ f_2^{(1)}+f_2^{(2)} \\ f_3^{(2)} \end{Bmatrix}^{thermal}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ 0 \end{Bmatrix}$$
%
% $$\begin{Bmatrix} F_1 \\ 0 \\ F_3 \end{Bmatrix}+\begin{Bmatrix} 0.08 \\ -0.04 \\ -0.04 \end{Bmatrix}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ 0 \end{Bmatrix}$$
%
% $$\rightarrow -0.04=325*u_2 \rightarrow u_2=\frac{-0.04}{325}\approx -12.31\cdot10^{-5}$$
%
% + slideshow={"slide_type": "slide"}
% Numerical check
F_thermal_night=zeros(3,1);
F_thermal_night(1:2)=F_thermal_night(1:2)+f12_night
F_thermal_night(2:3)=F_thermal_night(2:3)+f23_night
F2=0+F_thermal_night(2)
u2=F2./K(2,2)
% + [markdown] slideshow={"slide_type": "slide"}
% $$\begin{Bmatrix} F_1 \\ 0 \\ F_3 \end{Bmatrix}+\begin{Bmatrix} 0.08 \\ -0.04 \\ -0.04 \end{Bmatrix}=\begin{bmatrix} 200 & -200 & 0 \\ -200 & 325 & -125 \\ 0 & -125 & 125\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ 0 \end{Bmatrix}$$
%
% $$\rightarrow F_1+0.08=-200*u_2 \rightarrow F_1=-200 \cdot \frac{-0.04}{325}-0.08 \approx -0.05539$$
% $$\rightarrow F_3-0.04=-125*u_2 \rightarrow F_1=-125 \cdot \frac{-0.04}{325}+0.04 \approx 0.05539$$
%
% + slideshow={"slide_type": "slide"}
u=[0 u2 0]' %Displacements
F_total=K*u %Total forces
F=F_total-F_thermal_night %Direct forces
% + [markdown] slideshow={"slide_type": "slide"}
% # Question 2 thermal loads in a partially constrained set of trusses
% Consider the two truss system of Figure Q2. Both trusses are linear elastic. The first node is constrained from moving. Node 3 is subjected to a direct nodal force of $P=20$ N (in the x-direction). Furthermore, the system is heated, from an initial temperature of 18° Celsius, to 100° Celsius. The truss parameters for all elements are given in Table Q2.
%
% | | Element 1 | Element 2 |
% | ----------- | ----------- | ----------- |
% | Youngs modulus | 100 MPa | 100 MPa |
% | Area | 50 $mm^2$ | 25 $mm^2$ |
% | Length | 250 mm | 200 mm |
% | Thermal exp. coeff. | 2x$10^{-4}$/° Celsius | 2x$10^{-4}$/° Celsius |
%
% __(a)__ The increase in temperature causes thermal expansion and forces to develop in the truss system. Compute the element thermal force arrays as well as the global thermal force array.
%
% __(b)__ Derive the element stiffness matrices as well as the global stiffness matrix.
%
% __(c)__ Formulate the finite element equations for this system and use the finite element method to compute the nodal displacements and the direct nodal forces.
%
% __(d)__ Use the truss shape functions to compute the displacement in truss 1 at x=125 mm, assuming node 1 is at x=0 mm.
%
% __(e)__ Use the B-matrix to compute the element strains.
%
% 
%
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solution (a)
%
% Compute the temperature change:
% $$\Delta T=100-18=82$$
%
% Compute thermal force contributions:
% $$\begin{Bmatrix} f_1^{(1)} \\ f_2^{(1)} \end{Bmatrix}^{thermal} = E_1 \alpha_1 \Delta T A_1 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = 100 \cdot (2\cdot10^{-4}) \cdot 82 \cdot 50 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{Bmatrix} -82 \\ 82 \end{Bmatrix}$$
%
% $$\begin{Bmatrix} f_2^{(2)} \\ f_3^{(2)} \end{Bmatrix}^{thermal} = E_2 \alpha_2 \Delta T A_2 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = 100 \cdot (2\cdot10^{-4}) \cdot 82 \cdot 25 \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{Bmatrix} -41 \\ 41 \end{Bmatrix}$$
%
% Use superposition to get total thermal force contribution:
% $$\begin{Bmatrix} F \end{Bmatrix}^{thermal} = \begin{Bmatrix} -82 \\ 82-41 \\ 41 \end{Bmatrix}= \begin{Bmatrix} -82 \\ 41 \\ 41 \end{Bmatrix}$$
%
% + slideshow={"slide_type": "slide"}
% Numerical check
E1=100; A1=50; L1=250; a1=2e-4;
E2=100; A2=25; L2=200; a2=2e-4;
T_neutral=18; T=100; dT=T-T_neutral;
f12 = E1*a1*dT*A1*[-1;1]
f23 = E2*a2*dT*A2*[-1;1]
F_th=zeros(3,1);
F_th(1:2)=f12+F_th(1:2);
F_th(2:3)=f23+F_th(2:3)
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solution (b)
%
% Set up element stiffness matrices
%
% $$ \begin{bmatrix} K^{(1)} \end{bmatrix}=\frac{A_1 E_1}{L_1}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\frac{50 \cdot 100}{250}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\begin{bmatrix} 20 & -20 \\ -20 & 20\end{bmatrix}$$
%
% $$ \begin{bmatrix} K^{(2)} \end{bmatrix}=\frac{A_2 E_2}{L_2}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\frac{25 \cdot 100}{200}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}=\begin{bmatrix} 12.5 & -12.5 \\ -12.5 & 12.5\end{bmatrix}$$
%
% Use superposition to assemble total stiffness matrix
% $$\begin{bmatrix} K \end{bmatrix}=\underbrace{\begin{bmatrix} 20 & -20 & 0 \\ -20 & 20 & 0 \\ 0 & 0 & 0\end{bmatrix}}_{\begin{bmatrix} K^{(1)} \end{bmatrix}}+\underbrace{\begin{bmatrix} 0 & 0 & 0 \\ 0 & 12.5 & -12.5 \\ 0 & -12.5 & 12.5\end{bmatrix}}_{\begin{bmatrix} K^{(2)} \end{bmatrix}}=\begin{bmatrix} 20 & -20 & 0 \\ -20 & 32.5 & -12.5 \\ 0 & -12.5 & 12.5\end{bmatrix}$$
% + slideshow={"slide_type": "slide"}
% Numerical check
K1=(A1*E1)/L1*[1 -1; -1 1]
K2=(A2*E2)/L2*[1 -1; -1 1]
K=zeros(3,3);
K(1:2,1:2)=K(1:2,1:2)+K1;
K(2:3,2:3)=K(2:3,2:3)+K2
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solution (c)
%
% The total system of equations is written as:
%
% $$\begin{Bmatrix} F \end{Bmatrix}+\begin{Bmatrix} F \end{Bmatrix}^{thermal}=\begin{bmatrix} K \end{bmatrix}\begin{Bmatrix} u \end{Bmatrix}$$
%
% Leading to:
%
% $$\begin{Bmatrix} F_1 \\ 0 \\ 20 \end{Bmatrix}+\begin{Bmatrix} -82 \\ 41 \\ 41 \end{Bmatrix}=\begin{bmatrix} 20 & -20 & 0 \\ -20 & 32.5 & -12.5 \\ 0 & -12.5 & 12.5\end{bmatrix}\begin{Bmatrix} 0 \\ u_2 \\ u_3 \end{Bmatrix}$$
%
% Providing the following equations:
% $$ \rightarrow 32.5 \cdot u_2-12.5 \cdot u_3 =41 $$
% $$ \rightarrow -12.5 \cdot u_2+12.5 \cdot u_3 =61 $$
%
% + [markdown] slideshow={"slide_type": "slide"}
% Leading to:
% $$ \rightarrow 20 \cdot u_2=102 $$
% $$ \rightarrow u_2=\frac{102}{20}=\frac{51}{10}=5.1 $$
%
% And therefore:
% $$ \rightarrow 32.5 \cdot u_2-12.5 \cdot u_3 =41 $$
% $$ \rightarrow 32.5 \cdot 5.1-12.5 \cdot u_3 =41 $$
% $$ \rightarrow -12.5 \cdot u_3 =41-165.75=-124.75 $$
% $$ \rightarrow u_3 =\frac{-124.75}{-12.5}=9.98 $$
% + [markdown] slideshow={"slide_type": "slide"}
% Leading to:
% $$\begin{Bmatrix} F_1 \\ 0 \\ 20 \end{Bmatrix}+\begin{Bmatrix} -82 \\ 41 \\ 41 \end{Bmatrix}=\begin{bmatrix} 20 & -20 & 0 \\ -20 & 32.5 & -12.5 \\ 0 & -12.5 & 12.5\end{bmatrix}\begin{Bmatrix} 0 \\ 5.1 \\ 9.98 \end{Bmatrix}=\begin{Bmatrix} -102 \\ 41 \\ 61 \end{Bmatrix}$$
%
% And therefore $F_1=-20$ since:
% $$\begin{Bmatrix} F_1 \\ 0 \\ 20 \end{Bmatrix}+\begin{Bmatrix} -82 \\ 41 \\ 41 \end{Bmatrix}=\begin{Bmatrix} -102 \\ 41 \\ 61 \end{Bmatrix}=\begin{Bmatrix} -20 \\ 0 \\ 20 \end{Bmatrix}+\begin{Bmatrix} -82 \\ 41 \\ 41 \end{Bmatrix}$$
% + slideshow={"slide_type": "slide"}
% Numerical check
F2=0; %Force in middle node is zero
F3=20; %The applied force to node 3
u2=((F2+F_th(2))+(F3+F_th(3)))/(K(2,2)+K(3,2)) %Displacement u2
u3=((F2+F_th(2))-(K(2,2).*u2))./K(2,3) %Displacement u3
u=[0 u2 u3]' %Displacements
F_tot=K*u %Compute total force
F1=F_tot(1,1)-F_th(1,1) %get F1
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solution (d)
%
% The coordinate $x=125$ mm is in element 1 (in fact it is in the middle). The truss shape functions for this element can be expressed as:
% $$\begin{bmatrix} \mathbf{N}^{(1)} \end{bmatrix}=\begin{bmatrix} 1-\frac{x}{L_1} & \frac{x}{L_1}\end{bmatrix}=\begin{bmatrix} 1-\frac{125}{250} & \frac{125}{250}\end{bmatrix}$$
% Shape function based displacement interpolation is written as:
% $$u(x)=\begin{bmatrix} \mathbf{N} \end{bmatrix} \begin{Bmatrix} \mathbf{u} \end{Bmatrix}=\begin{bmatrix} 0.5 & 0.5 \end{bmatrix} \begin{Bmatrix} 0 \\ 5.1 \end{Bmatrix}=2.55$$
% + slideshow={"slide_type": "slide"}
% Numerical check
x=125; %X-coordinate for interpolation
N=[1-x/L1 x/L1] %Shape function metrics
u %Displacement vector
ux=N*u([1 2]) %Interpolated displacement
% + [markdown] slideshow={"slide_type": "slide"}
% ## Solution (e)
%
% The $\begin{bmatrix} \mathbf{B} \end{bmatrix}$ matrix is expressed as:
%
% $$\begin{bmatrix} \mathbf{B} \end{bmatrix}=\begin{bmatrix} -\frac{1}{L} & \frac{1}{L} \end{bmatrix} $$
%
% and the strain can be computed from;
% $$\varepsilon^{(i)}=\begin{bmatrix} B \end{bmatrix}^{(i)} \begin{Bmatrix} \mathbf{u} \end{Bmatrix}^{(i)}$$
%
% For element 1 we have:
% $$\varepsilon^{(1)}=\begin{bmatrix} -\frac{1}{250} & \frac{1}{250} \end{bmatrix} \begin{Bmatrix} 0 \\ 5.1 \end{Bmatrix}=0.0204$$
%
% For element 2 we have:
% $$\varepsilon^{(2)}=\begin{bmatrix} -\frac{1}{200} & \frac{1}{200} \end{bmatrix} \begin{Bmatrix} 5.1 \\ 9.98\end{Bmatrix}=0.0244$$
% + slideshow={"slide_type": "slide"}
% Numerical check
B1=[-1/L1 1/L1];
u_e1=u([1 2]);
eps_1=B1*u_e1
B2=[-1/L2 1/L2];
u_e2=u([2 3]);
eps_2=B2*u_e2
% + [markdown] slideshow={"slide_type": "slide"}
% # Summary
%
% * To allow for thermal strains the total strain $\begin{Bmatrix}\varepsilon\end{Bmatrix}$ is decomposed into the elastic strain $\begin{Bmatrix}\varepsilon_e\end{Bmatrix}$ and the termal strain $\begin{Bmatrix}\varepsilon_0\end{Bmatrix}$:
% $$\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}=\begin{Bmatrix}\varepsilon_e\end{Bmatrix}+\begin{Bmatrix}\varepsilon_0\end{Bmatrix}$$
% $$\begin{Bmatrix}\varepsilon_e\end{Bmatrix}=\begin{Bmatrix}\varepsilon_{tot}\end{Bmatrix}-\begin{Bmatrix}\varepsilon_{0}\end{Bmatrix}$$
%
%
% * The truss equations featuring thermal effects were developed as:
% $$\begin{Bmatrix} f_{1} \\ f_{2} \end{Bmatrix}+E \alpha \Delta T A \begin{bmatrix} -1 \\ 1 \end{bmatrix} =\frac{AE}{L}\begin{bmatrix} 1 & -1 \\ -1 & 1\end{bmatrix}\begin{Bmatrix} u_1 \\ u_2\end{Bmatrix}$$
%
% With:
%
% $\alpha$: Thermal expansion coefficient
% $\Delta T$: Temperature change
%
% + [markdown] slideshow={"slide_type": "skip"}
% \newpage
% + [markdown] slideshow={"slide_type": "skip"}
% **About this document**
%
% This document was created using a [Jupyter notebook](https://jupyter.org/) which allows for the presentation of theory and equations, as well as live (running code) numerical implementations.
%
% This Jupyter notebook is available [open source](https://github.com/Kevin-Mattheus-Moerman/NUIG_BME_402_6101) and features the [Octave](https://www.gnu.org/software/octave/index) programming language (an open source alternative to MATLAB). If you are interested in running this Jupyter notebook yourself [download and install Octave](https://www.gnu.org/software/octave/download) and install [the Jupyter environment](https://jupyter.org/install). Once both Jupyter and Octave are installed follow [these instructions](https://github.com/calysto/octave_kernel) to configure the use of Octave with Jupyter notebooks.
%
% To run Jupyter call `jupyter notebook` from your Terminal/Command Prompt.
| notebooks/nb7_thermal_problems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import sys
import h5py
from tqdm import tqdm
import numpy as np
import torch
# import BART
os.environ['TOOLBOX_PATH'] = "/home/svangurp/scratch/samuel/bart-0.6.00/"
sys.path.append('/home/svangurp/scratch/samuel/bart-0.6.00/python/')
import bart
import fastmri.data.transforms as T
from fastmri.data.subsample import create_mask_for_mask_type
from fastmri import tensor_to_complex_np
import matplotlib.pyplot as plt
from skimage.metrics import peak_signal_noise_ratio, structural_similarity
# +
def save_zero_filled(data_dir, out_dir, which_challenge):
reconstructions = {}
for fname in tqdm(list(data_dir.glob("*.h5"))):
with h5py.File(fname, "r") as hf:
et_root = etree.fromstring(hf["ismrmrd_header"][()])
masked_kspace = transforms.to_tensor(hf["kspace"][()])
# extract target image width, height from ismrmrd header
enc = ["encoding", "encodedSpace", "matrixSize"]
crop_size = (
int(et_query(et_root, enc + ["x"])),
int(et_query(et_root, enc + ["y"])),
)
# inverse Fourier Transform to get zero filled solution
image = fastmri.ifft2c(masked_kspace)
# check for FLAIR 203
if image.shape[-2] < crop_size[1]:
crop_size = (image.shape[-2], image.shape[-2])
# crop input image
image = transforms.complex_center_crop(image, crop_size)
# absolute value
image = fastmri.complex_abs(image)
# apply Root-Sum-of-Squares if multicoil data
if which_challenge == "multicoil":
image = fastmri.rss(image, dim=1)
reconstructions[fname.name] = image
fastmri.save_reconstructions(reconstructions, out_dir)
# -
fname = '/scratch/svangurp/samuel/data/knee/train/file1000002.h5'
data = h5py.File(fname, 'r')
kspace = data["kspace"][()]
| Samuel_notebooks/.ipynb_checkpoints/zero_filled_whole_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This script is used to do some analysis using RGC data in the paper (Baden, 2016, Nature).
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import FormatStrFormatter
import cv2
import glob
import h5py
from skimage.morphology import disk
from scipy.stats import pearsonr
from scipy.ndimage import gaussian_filter
import scipy.io
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# for plot figures
plt.rcParams['svg.fonttype'] = 'none'
def adjust_spines(ax, spines):
for loc, spine in ax.spines.items():
if loc in spines:
spine.set_position(('outward', 2))
else:
spine.set_color('none')
if 'left' in spines:
ax.yaxis.set_ticks_position('left')
else:
ax.yaxis.set_ticks([])
if 'bottom' in spines:
ax.xaxis.set_ticks_position('bottom')
else:
ax.xaxis.set_ticks([])
#load data
data_name = "data/BadenEtAl_RGCs_2016_v1.mat"
rgcdata=scipy.io.loadmat(data_name)
print (rgcdata.keys())
# ### Analysis
#
# group index 1-9 and 31 and 32: OFF
#
# group index 15-30: ON
#
# rf_qi>0.2, rf_size>50 and <450
#
# then use the corresponding onoff index
group_idx=rgcdata['group_idx'].flatten()#group number
rf_qi=rgcdata['rf_qi'].flatten()
rf_size=rgcdata['rf_size'].flatten()#RF diamter (2 sd of gaussian)
cell_oo_idx=rgcdata['cell_oo_idx'].flatten()#onoff index
print ('shape of group_idx:{}'.format(group_idx.shape))
print ('shape of rf_qi:{}'.format(rf_qi.shape))
print ('shape of rf_size:{}'.format(rf_size.shape))
print ('shape of cell_oo_idx:{}'.format(cell_oo_idx.shape))
# +
oncells_group_idx=[]
oncells_size=[]
offcells_group_idx=[]
offcells_size=[]
onoffcells=[]#on:1, off:-1 and RF size
onoffcells_group=[]#group index and RF size
onoffcells_ooi=[]#onoff index and RF size for all cells
oncells_ooi =[]#onoff index and RF size for ON cells
offcells_ooi =[]#onoff index and RF size for OFF cells
on_group_idx=np.arange(15,31)
on_group_idx=list(on_group_idx)
off_group_idx=np.arange(1,10)
off_group_idx=list(off_group_idx)
off_group_idx.append(31)
off_group_idx.append(32)
for ii in range(len(rf_size)):
if rf_size[ii]<450 and rf_size[ii]>50 and rf_qi[ii]>0.2:
if group_idx[ii] in on_group_idx:
oncells_group_idx.append(group_idx[ii])
oncells_size.append(rf_size[ii])
onoffcells.append(np.array([1,rf_size[ii]]))
onoffcells_group.append(np.array([group_idx[ii],rf_size[ii]]))
onoffcells_ooi.append(np.array([cell_oo_idx[ii],rf_size[ii]]))
oncells_ooi.append(np.array([cell_oo_idx[ii],rf_size[ii]]))
if group_idx[ii] in off_group_idx:
offcells_group_idx.append(group_idx[ii])
offcells_size.append(rf_size[ii])
onoffcells.append(np.array([-1,rf_size[ii]]))
onoffcells_group.append(np.array([group_idx[ii],rf_size[ii]]))
onoffcells_ooi.append(np.array([cell_oo_idx[ii],rf_size[ii]]))
offcells_ooi.append(np.array([cell_oo_idx[ii],rf_size[ii]]))
onoffcells=np.array(onoffcells)
onoffcells_group=np.array(onoffcells_group)
oncells_size=np.array(oncells_size)
offcells_size=np.array(offcells_size)
onoffcells_ooi=np.array(onoffcells_ooi)
oncells_ooi =np.array(oncells_ooi)
offcells_ooi =np.array(offcells_ooi)
print ('shape of onoffcells:{}'.format(onoffcells.shape))
print ('shape of onoffcells:{}'.format(onoffcells.shape))
print ('shape of oncells_size:{}'.format(oncells_size.shape))
print ('shape of offcells_size:{}'.format(offcells_size.shape))
print ('shape of onoffcells_ooi:{}'.format(onoffcells_ooi.shape))
# +
#plot onoff index histogram
xmax=1.0
xmin=-1.0
log_flag=False
weights = np.ones_like(onoffcells_ooi[:,0])/float(len(onoffcells_ooi[:,0]))
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
axes.hist(onoffcells_ooi[:,0], bins=16,color='gray',range=[xmin,xmax],\
weights=weights,log=log_flag)
axes.set_yticks([0,0.1])
axes.set_xlabel('Onoff index', fontsize=16)
axes.set_ylabel('Probability', fontsize=16)
adjust_spines(axes, ['left', 'bottom'])
plt.tight_layout()
# -
#plot onoff index histogram for on cells
xmax=1.0
xmin=-1.0
log_flag=False #log scale histogram if it is True
weights = np.ones_like(oncells_ooi[:,0])/float(len(oncells_ooi[:,0]))
print (len(oncells_ooi))
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
axes.hist(oncells_ooi[:,0], bins=16,color='gray',range=[xmin,xmax],\
weights=weights,log=log_flag)
axes.set_yticks([0,0.1])
axes.set_xlabel('Onoff index', fontsize=16)
axes.set_ylabel('Probability', fontsize=16)
adjust_spines(axes, ['left', 'bottom'])
plt.tight_layout()
#plot onoff index histogram for off cells
xmax=1.0
xmin=-1.0
log_flag=False #log scale histogram if it is True
weights = np.ones_like(offcells_ooi[:,0])/float(len(offcells_ooi[:,0]))
print (len(offcells_ooi))
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
axes.hist(offcells_ooi[:,0], bins=16,color='gray',range=[xmin,xmax],\
weights=weights,log=log_flag)
axes.set_yticks([0,0.1])
axes.set_xlabel('Onoff index', fontsize=16)
axes.set_ylabel('Probability', fontsize=16)
adjust_spines(axes, ['left', 'bottom'])
plt.tight_layout()
# +
#plot 2d histogram of onoff index and rf size
xmax=450
xmin=30
ymax=1.0
ymin=-1.0
weights = np.ones_like(onoffcells_ooi[:,1])/float(len(onoffcells_ooi[:,1]))
rfsize_ooidx=np.zeros((64,64))
rfsize_ooidx,_,_=np.histogram2d(onoffcells_ooi[:,1], \
onoffcells_ooi[:,0], \
bins=64,range=[[xmin,xmax],[ymin,ymax]],weights=weights)
rfsize_ooidx=np.log10(rfsize_ooidx+1e-6)
rfsize_ooidx=rfsize_ooidx.T
H_max=-2
H_min=-6
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
im=axes.imshow(rfsize_ooidx, interpolation='nearest', origin='low',cmap='jet',vmin=H_min, vmax=H_max)
labels = ['2','6','10','14']
axes.set_xticks([4.5,22.5,40.5,58.5]) # 0: 30um, 63:450um,
axes.set_xticklabels(labels)
labels = [float("{0:.1f}".format(ymin)),float("{0:.1f}".format(ymax))]
axes.set_yticks([0,63])
axes.set_yticklabels(labels)
axes.axhline(y=31.5,color='k',linewidth=2)
axes.set_xlabel('RF size', fontsize=16)
axes.set_ylabel('Onoff index', fontsize=16)
#
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([0.85, 0.3, 0.03, 0.5])
ticks=[-2,-4,-6]
fig.colorbar(im, cax=cbar_ax,ticks=ticks,format="%d")
handles, labels = axes.get_legend_handles_labels()
lgd = axes.legend(handles, labels, loc='center left', bbox_to_anchor=(1, 0.5))
# +
#plot 2d histogram of onoff index and rf size, ON cells
xmax=450
xmin=30 # 50
ymax=1.0
ymin=-1.0
weights = np.ones_like(oncells_ooi[:,1])/float(len(oncells_ooi[:,1]))
rfsize_ooidx=np.zeros((64,64))
rfsize_ooidx,_,_=np.histogram2d(oncells_ooi[:,1], \
oncells_ooi[:,0], \
bins=64,range=[[xmin,xmax],[ymin,ymax]],weights=weights)
rfsize_ooidx=np.log10(rfsize_ooidx+1e-6)
rfsize_ooidx=rfsize_ooidx.T
H_max=-2
H_min=-6
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
im=axes.imshow(rfsize_ooidx, interpolation='nearest', origin='low',cmap='jet',vmin=H_min, vmax=H_max)
labels = ['2','6','10','14']
axes.set_xticks([4.5,22.5,40.5,58.5])
axes.set_xticklabels(labels)
labels = [float("{0:.1f}".format(ymin)),float("{0:.1f}".format(ymax))]
axes.set_yticks([0,63])
axes.set_yticklabels(labels)
axes.axhline(y=31.5,color='white',linestyle='--')
axes.set_xlabel('RF size', fontsize=16)
axes.set_ylabel('Onoff index', fontsize=16)
#
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([0.85, 0.3, 0.03, 0.5])
ticks=[-2,-4,-6]
fig.colorbar(im, cax=cbar_ax,ticks=ticks,format="%d")
handles, labels = axes.get_legend_handles_labels()
lgd = axes.legend(handles, labels, loc='center left', bbox_to_anchor=(1, 0.5))
# +
#plot 2d histogram of onoff index and rf size, OFF cells
xmax=450
xmin=30
ymax=1.0
ymin=-1.0
weights = np.ones_like(offcells_ooi[:,1])/float(len(offcells_ooi[:,1]))
rfsize_ooidx=np.zeros((64,64))
rfsize_ooidx,_,_=np.histogram2d(offcells_ooi[:,1], \
offcells_ooi[:,0], \
bins=64,range=[[xmin,xmax],[ymin,ymax]],weights=weights)
rfsize_ooidx=np.log10(rfsize_ooidx+1e-6)
rfsize_ooidx=rfsize_ooidx.T
H_max=-2
H_min=-6
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
im=axes.imshow(rfsize_ooidx, interpolation='nearest', origin='low',cmap='jet',vmin=H_min, vmax=H_max)
labels = ['2','6','10','14']
axes.set_xticks([4.5,22.5,40.5,58.5])
axes.set_xticklabels(labels)
labels = [float("{0:.1f}".format(ymin)),float("{0:.1f}".format(ymax))]
axes.set_yticks([0,63])
axes.set_yticklabels(labels)
axes.axhline(y=31.5,color='white',linestyle='--')
axes.set_xlabel('RF size', fontsize=16)
axes.set_ylabel('Onoff index', fontsize=16)
#
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([0.85, 0.3, 0.03, 0.5])
ticks=[-2,-4,-6]
fig.colorbar(im, cax=cbar_ax,ticks=ticks,format="%d")
handles, labels = axes.get_legend_handles_labels()
lgd = axes.legend(handles, labels, loc='center left', bbox_to_anchor=(1, 0.5))
# -
#plot OFF/ON as RF size
#sort cells as RF size
temp=np.argsort(onoffcells_ooi[:,1])
onoffcells_ooi_sorted=onoffcells_ooi[temp][:2376]
onoffcells_sorted =onoffcells[temp][:2376]#on:1, off:-1 and RF size
num=6
delta=int(len(onoffcells_ooi_sorted)/num)
OFFtoONs=np.zeros(num)
rfsizegroups=np.zeros(num,np.int32)
for ii in range(num):
temp1=onoffcells_ooi_sorted[ii*delta:ii*delta+delta,0]#onoff index
temp2=onoffcells_ooi_sorted[ii*delta:ii*delta+delta,1]#rf size
OFFtoONs[ii]=len(np.where(temp1<0)[0])/len(np.where(temp1>0)[0])
rfsizegroups[ii]=int(np.median(temp2))
#
fig, ax = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
ax.plot(rfsizegroups,OFFtoONs,'o',color='k',linestyle='-')
ax.set_xticks([6*30,8*30,10*30])
ax.set_yticks([0,1,2])
ax.set_xlabel('RF size (um)', fontsize=16)
ax.set_ylabel('OFF/ON', fontsize=16)
adjust_spines(ax, ['left', 'bottom'])
plt.tight_layout()
#boxplot of onoff index, also for bootstrapping and statistical test
Onoff_boxplot=[] #OOi index
Onoff_polarity_boxplot=[] #polarity
for ii in range(num):
Onoff_boxplot.append(onoffcells_ooi_sorted[ii*delta:ii*delta+delta,0])
Onoff_polarity_boxplot.append(onoffcells_sorted[ii*delta:ii*delta+delta,0])
Onoff_boxplot_array=np.array(Onoff_boxplot) #OOi index
Onoff_polarity_boxplot_array=np.array(Onoff_polarity_boxplot) #polarity, on:1, off:-1
print (Onoff_boxplot_array.shape)
print (Onoff_polarity_boxplot_array.shape)
#bootstrapping
#apply bootstrapping to estimate standard deviation (error)
#statistics can be offratios, median, mean
#for offratios, be careful with the threshold
#data: for statistics offratios, median, mean: numpy array with shape (sample_size,1)
#num_exp: number of experiments, with replacement
def bootstrap(statistics,data,num_exp=10000,seed=66):
if statistics == 'offratios':
def func(x): return len(x[np.where(x<0)])/len(x[np.where(x>0)]) #threshold is 0, may be different
elif statistics == 'median':
def func(x): return np.median(x)
elif statistics == 'mean':
def func(x): return np.mean(x)
sta_boot=np.zeros((num_exp))
num_data=len(data)
for ii in range(num_exp):
np.random.seed(seed+ii)
tempind=np.random.choice(num_data,num_data,replace=True)
sta_boot[ii]=func(data[tempind])
return np.percentile(sta_boot,2.5),np.percentile(sta_boot,97.5)
#plot OFF/ON as RF size with bootstrapping
OFFtoONlowqs=np.zeros(len(Onoff_boxplot_array)) #lower_quartile
OFFtoONhigqs=np.zeros(len(Onoff_boxplot_array)) #upper_quartile
for ii in range(len(OFFtoONlowqs)):
temp=Onoff_boxplot_array[ii]
tempmax=0
low_perc,high_perc=bootstrap('offratios',temp,num_exp=10000,seed=66)
OFFtoONlowqs[ii] = OFFtoONs[ii]-low_perc
OFFtoONhigqs[ii] =-OFFtoONs[ii]+high_perc
#
#plot
fig, ax = plt.subplots(nrows=1, ncols=1,figsize=(2,3))
ax.errorbar(rfsizegroups,OFFtoONs,yerr=(OFFtoONlowqs,OFFtoONhigqs),marker='o',color='k',linestyle='-',capsize=4)
ax.set_xticks([6*30,8*30,10*30])
ax.set_yticks([0,1,2])
ax.set_xlabel('RF size (um)', fontsize=16)
ax.set_ylabel('OFF/ON', fontsize=16)
adjust_spines(ax, ['left', 'bottom'])
plt.tight_layout()
#get the median of onoff index
Onoffmedians=np.zeros(num)
Onoffmeans=np.zeros(num)
rfsizegroups=np.zeros(num,np.int32)
for ii in range(num):
temp1=onoffcells_ooi_sorted[ii*delta:ii*delta+delta,0]#onoff index
temp2=onoffcells_ooi_sorted[ii*delta:ii*delta+delta,1]#rf size
Onoffmedians[ii]=np.median(temp1)
Onoffmeans[ii] =np.mean(temp1)
rfsizegroups[ii]=int(np.median(temp2))
#
fig, ax = plt.subplots(nrows=1, ncols=1,figsize=(3,3))
ax.plot(rfsizegroups,Onoffmedians,'o',color='k',linestyle='-')
ax.set_xlabel('RF size (um)', fontsize=16)
ax.set_ylabel('Onoff median', fontsize=16)
adjust_spines(ax, ['left', 'bottom'])
handles, labels = ax.get_legend_handles_labels()
lgd = ax.legend(handles, labels, loc='center left', frameon=False,bbox_to_anchor=(1, 0.5))
#plot median of onoff index with bootstrapping
Onofflowqs=np.zeros(len(Onoff_boxplot_array)) #lower_quartile
Onoffhigqs=np.zeros(len(Onoff_boxplot_array)) #upper_quartile
for ii in range(len(Onofflowqs)):
temp=Onoff_boxplot_array[ii]
low_perc,high_perc=bootstrap('median',temp,num_exp=10000,seed=66)
Onofflowqs[ii] = Onoffmedians[ii]-low_perc
Onoffhigqs[ii] =-Onoffmedians[ii]+high_perc
#
fig, ax = plt.subplots(nrows=1, ncols=1,figsize=(2,3))
ax.errorbar(rfsizegroups,Onoffmedians,yerr=(Onofflowqs,Onoffhigqs),marker='o',color='k',linestyle='-',capsize=4)
ax.axhline(y=0,color='k',linestyle='--')
ax.set_xticks([6*30,8*30,10*30])
ax.set_yticks([0.2,0,-0.2,-0.4])
ax.set_xlabel('RF (um)', fontsize=16)
ax.set_ylabel('Onoff median', fontsize=16)
adjust_spines(ax, ['left', 'bottom'])
handles, labels = ax.get_legend_handles_labels()
lgd = ax.legend(handles, labels, loc='center left',frameon=False, bbox_to_anchor=(1, 0.5))
#plot median of onoff index with bootstrapping, ON and OFF separately
#ON
Onofflowqs_ON =np.zeros(len(Onoff_boxplot_array)) #lower_quartile
Onoffhigqs_ON =np.zeros(len(Onoff_boxplot_array)) #upper_quartile
Onoffmedians_ON=np.zeros(len(Onoff_boxplot_array))
for ii in range(len(Onofflowqs_ON)):
temp=Onoff_boxplot_array[ii]
temp=temp[np.where(Onoff_polarity_boxplot_array[ii]>0)] # ON
Onoffmedians_ON[ii]=np.median(temp)
low_perc,high_perc=bootstrap('median',temp,num_exp=10000,seed=66)
Onofflowqs_ON[ii] = Onoffmedians_ON[ii]-low_perc
Onoffhigqs_ON[ii] =-Onoffmedians_ON[ii]+high_perc
#OFF
Onofflowqs_OFF =np.zeros(len(Onoff_boxplot_array)) #lower_quartile
Onoffhigqs_OFF =np.zeros(len(Onoff_boxplot_array)) #upper_quartile
Onoffmedians_OFF=np.zeros(len(Onoff_boxplot_array))
for ii in range(len(Onofflowqs_OFF)):
temp=Onoff_boxplot_array[ii]
temp=temp[np.where(Onoff_polarity_boxplot_array[ii]<0)] # OFF
Onoffmedians_OFF[ii]=np.median(temp)
low_perc,high_perc=bootstrap('median',temp,num_exp=10000,seed=66)
Onofflowqs_OFF[ii] = Onoffmedians_OFF[ii]-low_perc
Onoffhigqs_OFF[ii] =-Onoffmedians_OFF[ii]+high_perc
#
fig, ax = plt.subplots(nrows=1, ncols=1,figsize=(2,3))
ax.errorbar(rfsizegroups,Onoffmedians_ON,yerr=(Onofflowqs_ON,Onoffhigqs_ON),\
marker='o',color='k',linestyle='-',capsize=4,label='ON')
ax.errorbar(rfsizegroups,Onoffmedians_OFF,yerr=(Onofflowqs_OFF,Onoffhigqs_OFF),\
marker='o',color='k',linestyle='--',capsize=4,label='OFF')
ax.set_xticks([6*30,8*30,10*30])
ax.set_yticks([-0.8,-0.4,0,0.4])
ax.set_xlabel('RF (um)', fontsize=16)
ax.set_ylabel('Onoff median', fontsize=16)
adjust_spines(ax, ['left', 'bottom'])
handles, labels = ax.get_legend_handles_labels()
lgd = ax.legend(handles, labels, loc='center left',frameon=False, bbox_to_anchor=(1, 0.5))
# ### Statistical test
# #### permutation test
#permutation test using monte-carlo method
def perm_test(xs, ys, nmc, randomseed):
n, k = len(xs), 0
diff = np.abs(np.mean(xs) - np.mean(ys))
zs = np.concatenate([xs, ys])
for j in range(nmc):
np.random.seed(randomseed+j)
np.random.shuffle(zs)
k += diff < np.abs(np.mean(zs[:n]) - np.mean(zs[n:]))
return k / nmc
perm_res=[]
for ii in np.arange(len(Onoff_boxplot)):
for jj in np.arange(ii+1,len(Onoff_boxplot)):
temp=perm_test(Onoff_boxplot[ii], Onoff_boxplot[jj], 10000, ii*jj+jj+100)
perm_res.append(temp)
print(perm_res)
| code/RGC_Baden_2016_Nature_reanalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('thesis')
# language: python
# name: python3
# ---
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tqdm
import json
import glob
from numpyencoder import NumpyEncoder
import re
import open3d
import shutil
PATH="Dataset"
METADATA= glob.glob(f"{PATH}/**/**.json")
METADATA
folder_name = [i.split("/")[1] for i in glob.glob(f"{PATH}/**/")]
existing_metadata = [i.split("/")[1] for i in METADATA]
for n_folder in folder_name:
if n_folder not in existing_metadata:
print(n_folder)
# +
def save_meta_data(file_name,dict_file):
with open(f'{file_name}.json', 'w') as fp:
json.dump(dict_file, fp, indent=4,cls=NumpyEncoder)
def read_meta_data (path):
with open(path, 'r') as j:
contents = json.loads(j.read())
return contents
def del_data(files_name,folder_name="Dataset"):
try:
for d in files_name:
os.remove(f"{folder_name}/{d}")
except:
os.remove(f"{folder_name}/{files_name}")
# -
b_data = [i.split("/")[1] for i in METADATA]
for b_d in b_data :
buff = glob.glob(f"{PATH}/{b_d}/**.txt")
data = np.array([i.split("/") for i in buff if "building" in i or "buidling" in i])
new= data[:,-1].copy()
new = [i.split("_") for i in new]
for i,_ in enumerate(new):
if len(new[i])==2:
new[i][1] = new[i][1].split(".")
new[i][1][0]= new[i][1][0].zfill(3)
new[i][1]= ".".join(new[i][1])
else:
new[i][1]=new[i][1].zfill(3)
new[i]= "_".join(new[i])
index = np.where(data[:,-1]!=new)[0]
if index.size != 0:
old_data= data[:,-1]
METADATA_buff = read_meta_data(f'{PATH}/{b_d}/meta_data.json')
OLD_METADATA_buff = METADATA_buff.copy()
for i in index:
try:
data_edited= METADATA_buff.pop(old_data[i][:-4])
except:
continue
list_new_data=new[i].split("/")
METADATA_buff[list_new_data[-1][:-4]]={
"path":f"{b_d}/{new[i]}",
"jumlah_point":data_edited["jumlah_point"]
}
print(index)
save_meta_data(f'{PATH}/{b_d}/meta_data',METADATA_buff)
for idx in index:
os.rename(f"{PATH}/{b_d}/{old_data[idx]}",f"{PATH}/{b_d}/{new[idx]}")
# # print()
# # print(old_data)
# METADATA= glob.glob(f"{PATH}/**/**.json")
# +
def get_list_per_instace(str_name ,file_list,Area,dict_file):
result_dict = {}
id_ns = np.unique(np.array([b.split("_")[1] for b in dict_file],dtype="int16"))
zf = 3 #if Area == "T_316000_233500_NE_T_316000_233500_SW" or Area=="T_315500_234500_SE" else 2
for idn in id_ns :
r = re.compile(f"{str_name}_{str(idn).zfill(zf)}")
result_dict[f"{str_name}_{str(idn).zfill(zf)}"] = list(filter(r.match, file_list))
return result_dict
for i,_ in tqdm.tqdm(enumerate(METADATA),total=len(METADATA)):
Area = METADATA[i].split("/")[1]
md = read_meta_data(METADATA[i])
file_names = md.keys()
building,ground,undefined,vegetation=[],[],[],[]
label = set([i.split("_")[0] for i in file_names])
for fn in file_names:
if 'building' in fn.lower() or "buidling" in fn.lower():
building.append(fn)
if 'ground' in fn.lower():
ground.append(fn)
if 'undefined' in fn.lower() or "undifined" in fn.lower() or "undefined" in fn.lower() or 'Undefined' in fn.lower():
undefined.append(fn)
if 'vegetation'in fn.lower():
vegetation.append(fn)
dict_area={
"building":get_list_per_instace(str_name = "building",file_list = building,dict_file=building,Area=Area),
"vegetation":vegetation,
"ground":ground,
"undefined":undefined
}
save_meta_data(f"map_ins_{Area}",dict_area)
def maping_meta_data(class_name,map_data,meta_data):
inst_id = {}
try:
for count,key in enumerate(map_data[class_name]):
try:
k_split = key.split("_")
if len(k_split)==3:
_,sub_class,key_id = k_split
else :
sub_class,key_id[:-4] = k_split
except:
sub_class,key_id=key,0
inst_id[str(count).zfill(3)]={
"total_point": meta_data[key]["jumlah_point"],
"path": meta_data[key]["path"],
"sub_class": sub_class,
"id": str(int(key_id)-1).zfill(3)
}
except Exception as e:
print(class_name)
print(meta_data[class_name])
inst_id["000"]={
"total_point": meta_data[class_name]["jumlah_point"],
"path": meta_data[class_name]["path"],
"sub_class": "000",
"id": "000"
}
return inst_id
def map_to_inst(map_data_json,meta_data_json):
map_data = read_meta_data(map_data_json)
meta_data= read_meta_data(meta_data_json)
for k in map_data.keys():
if k == "building":
inst_id_building = {}
list_build_id= list(map_data[k].keys())
for id_b in list_build_id:
id_ins = str(int(id_b.split("_")[-1])-1).zfill(3)
total_point = sum([meta_data[i]["jumlah_point"] for i in map_data[list(map_data.keys())[0]][id_b]])
list_all_txt = [meta_data[i]["path"] for i in map_data[list(map_data.keys())[0]][id_b]]
inst_id_building[id_ins]={
"total_point":total_point,
"path":list_all_txt
}
elif k=="vegetation" :
inst_id_vegetation = maping_meta_data("vegetation",map_data,meta_data)
elif k=="ground" :
inst_id_ground = maping_meta_data("ground",map_data,meta_data)
else :
inst_id_undefined = maping_meta_data("undefined",map_data,meta_data)
return [inst_id_building,inst_id_vegetation,inst_id_ground,inst_id_undefined]
for i,_ in enumerate(METADATA):
area=METADATA[i].split("/")[1]
ins = map_to_inst(f"map_ins_{area}.json",METADATA[i])
save_meta_data(f"map_ins_{area}",{"building":ins[0],"vegetation":ins[1],"ground":ins[2],'undefined':ins[3]})
# -
MAPTOINST_META_LIST= glob.glob("map_**")
print(MAPTOINST_META_LIST)
MAPTOINST_META_LIST= glob.glob("map_**")
def get_combining_building_data(map_data_json = MAPTOINST_META_LIST,buff_folder="Dataset") :
for i,meta_name in tqdm.tqdm(enumerate(map_data_json),total=len(map_data_json)):
map_data = read_meta_data(map_data_json[i])["building"]
buff_folder = f"all_data_building/{map_data_json[i][8:-5]}"
os.makedirs(buff_folder,exist_ok=True)
map_data_id= list(map_data.keys())
for mdi in map_data_id:
list_data_txt = map_data[mdi]["path"]
# Start from init = 0 and append until last
all_data = np.loadtxt(f"Dataset/{list_data_txt[0]}")[:,:6]
for idx in range(1,len(list_data_txt)):
try:
buff = np.loadtxt(f"Dataset/{list_data_txt[idx]}")[:,:6]
all_data = np.append(all_data,buff,axis=0)
except :
buff = np.loadtxt(f"Dataset/{list_data_txt[idx]}")[:6]
all_data = np.append(all_data,[buff],axis=0)
# del_data(list_data_txt)
if "T_316000_233500_NE_T_316000_233500_SW" in meta_name:
data_building_index = np.where(all_data[:,0]<318000)
data_bulding_sub_building_index = np.where(all_data[:,0]>318000)
data_building = all_data[data_building_index]
data_bulding_sub_building = all_data[data_bulding_sub_building_index]
data_bulding_sub_building[:,0] = data_bulding_sub_building[:,0]-2000
data_bulding_sub_building[:,1] = data_bulding_sub_building[:,1]-3619
data_bulding_sub_building[:,2] = data_bulding_sub_building[:,2]+37.4
all_data = np.append(data_building,data_bulding_sub_building,axis=0)
save_txt_loc = f"{buff_folder}/building_{mdi}.txt"
map_data[mdi]["path"] = save_txt_loc
np.savetxt(save_txt_loc,all_data,fmt='%.8f %.8f %.8f %d %d %d')
# break
old_map = read_meta_data(map_data_json[i])
old_map["building"]= map_data
save_meta_data(f"{map_data_json[i][8:-5]}",old_map)
os.remove(f"{map_data_json[i]}")
get_combining_building_data()
| Data Clean and Extraction/meta_dataekstraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="dKIcnfoBYoC5" outputId="011f1477-684f-4e6e-ce5b-7df4d555a4de" colab={"base_uri": "https://localhost:8080/"}
# %load_ext autoreload
# %autoreload 2
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from skimage import io
matplotlib.rcParams['figure.figsize'] = (13,7)
# + id="kyOG3phoY_Kj" outputId="b946b164-564d-4bb0-9829-73a685410e2e" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="vw8ioO3OZD8k"
# !ln -s /root /content
# + id="xdl2DVe8ZJiJ"
import sys
sys.path.append('/content/gdrive/MyDrive/ColabNotebooks/fastaiPart2/course-v3/nbs/dl2')
# + id="eFk3EERbZSQQ" outputId="d814924e-8623-4159-b1ea-f8a61168a987" colab={"base_uri": "https://localhost:8080/"}
# !git clone https://github.com/eladrich/pix2vertex.pytorch.git
# + id="B7hg2Kv8Z3zi"
# %cd pix2vertex.pytorch
# !python setup.py install
# + id="g_dO1ecYZCmn"
import pix2vertex as p2v
# + id="4mQJqptZZOOE"
# + [markdown] id="2m8DzntcYoDI"
# ## Initializations
# + id="cUw1wxkPbD_D"
#dlib and facial landmarks: cf: https://www.pyimagesearch.com/2017/04/03/facial-landmarks-dlib-opencv-python/
# + id="v9K38v4lYoDK" outputId="780ca5c0-9810-44f3-be36-1fb130b94fc0" colab={"base_uri": "https://localhost:8080/"}
detector = p2v.Detector()
reconstructor = p2v.Reconstructor(detector=detector)
# + id="kDM8a0J1YoDK"
im_path = 'examples/sample.jpg' # im_path can be a URL as well!
img = io.imread(im_path)
fig = plt.figure()
plt.imshow(img)
plt.show()
# + [markdown] id="1Fk446iGYoDL"
# ## Inference
# + id="m_Ow011nYoDM"
img_crop = detector.detect_and_crop(img)
fig = plt.figure()
plt.imshow(img_crop)
plt.show()
# + id="Hq5yvvPOYoDM"
net_res = reconstructor.run_net(img_crop)
p2v.vis_net_result(img_crop,net_res)
final_res = reconstructor.post_process(net_res)
# + [markdown] id="zEv5j337YoDN"
# ## Interactive Visualizations
# + id="rMWiL1_ZYoDN"
plot = p2v.vis_depth_interactive(final_res['Z_surface'])
# + id="rS8j1rhuYoDO"
plot = p2v.vis_pcloud_interactive(final_res,img_crop)
# + id="gfO_un2UYoDO"
# Fallback matplotlib visualization
p2v.vis_depth_matplotlib(img_crop,final_res['Z_surface'])
# + [markdown] id="w3D0GaWBYoDP"
# ## Saving Result
# + id="Jx6CFgdOYoDP"
p2v.save2stl(final_res['Z_surface'],'res.stl')
# + [markdown] id="99ZRIS-7YoDQ"
# Create link to make accessible from notebook
# + id="2Ue63TmRYoDQ"
from IPython.display import FileLink
FileLink('res.stl')
# + id="cnoZ8PBBYoDT"
| reconstruct_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # `MaxwellVacuum`: Solving Maxwell Equations in Vacuum with NRPy+ and the Einstein Toolkit
#
# ## Authors: <NAME>, <NAME>, & <NAME>
#
# ## This notebook generates `MaxwellVacuum`, an [Einstein Toolkit](https://einsteintoolkit.org) thorn for solving Maxwell's equations in vacuum, in flat spacetime and Cartesian coordinates. This thorn is highly optimized for modern CPU architectures, featuring SIMD intrinsics and OpenMP support.
#
# ### Formatting improvements courtesy Brandon Clark
#
# [comment]: <> (Abstract: TODO)
#
# **Notebook Status:** <font color='green'><b> Validated</b></font>
#
# **Validation Notes:** As demonstrated [in the plot below](#code_validation), the numerical errors converge to zero at the expected rate, and we observe similar qualitatitve behavior of of the error nodes for Systems I (ADM - like) and II (BSSN - like). Also validated against the NRPy+ [Tutorial-Start_to_Finish-Solving_Maxwells_Equations_in_Vacuum-Cartesian](Tutorial-Start_to_Finish-Solving_Maxwells_Equations_in_Vacuum-Cartesian.ipynb) notebook.
#
# ### NRPy+ Source Code for this module:
# * [Maxwell/VacuumMaxwell_Flat_Evol_Cartesian.py](../edit/Maxwell/VacuumMaxwell_Flat_Evol_Cartesian.py) [\[**tutorial**\]](Tutorial-VacuumMaxwell_formulation_Cartesian.ipynb)
#
# ## Introduction:
# This tutorial notebook constructs an Einstein Toolkit (ETK) thorn (module) that will set up expressions for the right-hand sides of Maxwell's equations as described in [Knapp, Walker, & Baumgarte (2002)](https://arxiv.org/abs/gr-qc/0201051) and the [NRPy+ `Tutorial-VacuumMaxwell_formulation_Cartesian` Jupyter notebook](Tutorial-VacuumMaxwell_formulation_Cartesian.ipynb), both for System I:
#
# \begin{align}
# \partial_t A^i &= -E^i - \partial^i \varphi, \\
# \partial_t E^i &= \partial^i \partial_j A^j - \partial_j \partial^j A^i, \\
# \partial_t \varphi &= -\partial_i A^i,
# \end{align}
#
# with associated constraint
#
# $$
# \partial_i E^i = 0,
# $$
#
# and for System II:
#
# \begin{align}
# \partial_t A^i &= -E^i - \partial^i \varphi, \\
# \partial_t E^i &= \partial^i \Gamma - \partial_j \partial^j A^i, \\
# \partial_t \Gamma &= - \partial_i \partial^i \varphi, \\
# \partial_t \varphi &= -\Gamma,
# \end{align}
#
# with associated constraints
#
# \begin{align}
# \Gamma - \partial_i A^i &= 0, \\
# \partial_i E^i &= 0.
# \end{align}
#
# This thorn is largely based on and should function similarly to the [`BaikalETK`](Tutorial-BaikalETK.ipynb) thorns, which solve Einstein's equations of general relativity in Cartesian coordinates, in the BSSN formalism. Further, we generate the C kernels for a number of finite difference orders, so that users are free to choose the finite-differencing order at runtime.
#
# When interfaced properly with the ETK, this module will propagate the initial data for $E^i$, $A^i$, and $\varphi$ defined in [`MaxwellVacuumID`](Tutorial-ETK_thorn-MaxwellVacuumID.ipynb) forward in time by integrating the equations for $\partial_t E^i$, $\partial_t A^i$ and $\partial_t \varphi$ subject to spatial boundary conditions. The time evolution itself is handled by the `MoL` (Method of Lines) thorn in the `CactusNumerical` arrangement, and the boundary conditions by the `Boundary` thorn in the `CactusBase` arrangement. Specifically, we implement ETK's `NewRad` boundary condition driver, i.e. a radiation (Sommerfeld) boundary condition.
#
# Similar to the [`MaxwellVacuumID`](Tutorial-ETK_thorn-MaxwellVacuumID.ipynb) module, we will construct the MaxwellVacuum module in two steps.
#
# 1. Call on NRPy+ to convert the SymPy expressions for the evolution equations into C-code kernels.
# 1. Write the C code drivers and linkages to the Einstein Toolkit infrastructure (i.e., the .ccl files) to complete this Einstein Toolkit module.
#
# The above steps will be performed in this notebook by first outputting *Python* code into a module, and then loading the Python module to generate the C code *in parallel* (using the [`multiprocessing`](https://docs.python.org/3/library/multiprocessing.html) built-in Python module).
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Initialize needed Python/NRPy+ modules
# 1. [Step 2](#mwev): NRPy+-generated C code kernels for Maxwell spacetime solve
# 1. [Step 2.a](#mwevrhs): Maxwell RHS expressions
# 1. [Step 2.b](#constraints): Divergence Constraint
# 1. [Step 2.c](#driver_call_codegen_funcs): Given `WhichPart` parameter choice, generate algorithm for calling corresponding function within `MaxwellVacuum_C_kernels_codegen_onepart()` to generate C code kernel
# 1. [Step 2.e](#kernel_codegen): Generate C code kernels for `MaxwellVacuum`
# 1. [Step 2.e.i](#feature_choice): Set compile-time and runtime parameters for `MaxwellVacuum`
# 1. [Step 2.e.ii](#parallel_codegen): Generate all C-code kernels for `MaxwellVacuum`, in parallel if possible
# 1. [Step 3](#cclfiles): CCL files - Define how this module interacts and interfaces with the wider Einstein Toolkit infrastructure
# 1. [Step 3.a](#paramccl): `param.ccl`: specify free parameters within `MaxwellVacuum`
# 1. [Step 3.b](#interfaceccl): `interface.ccl`: define needed gridfunctions; provide keywords denoting what this thorn provides and what it should inherit from other thorns
# 1. [Step 3.c](#scheduleccl): `schedule.ccl`:schedule all functions used within `MaxwellVacuum`, specify data dependencies within said functions, and allocate memory for gridfunctions
# 1. [Step 4](#cdrivers): C driver functions for ETK registration & NRPy+ generated kernels
# 1. [Step 4.a](#etkfunctions): Needed ETK functions: Banner, Symmetry registration, Parameter sanity check, Method of Lines (`MoL`) registration, Boundary condition
# 1. [Step 4.b](#mwevrhss) Evaluate Maxwell right-hand-sides (RHSs)
# 1. [Step 4.c](#diagnostics): Diagnostics: Computing the divergence constraint
#
# 1. [Step 4.d](#outcdrivers): Output all C driver functions needed for `MaxwellVacuum`
# 1. [Step 4.e](#makecodedefn): `make.code.defn`: List of all C driver functions needed to compile `MaxwellVacuum`
# 1. [Step 5](#code_validation): Code Validation
# 1. [Step 5.a](#convergence): Error Convergence
# 1. [Step 5.b](#errornodes): Behavior of Error Nodes
# 1. [Step 6](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize needed Python/NRPy+ modules \[Back to [top](#toc)\]
#
# $$\label{initializenrpy}$$
# +
MaxwellVacuumdir = "MaxwellNRPy_py_dir"
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
import os, sys # Standard Python modules for multiplatform OS-level functions
cmd.mkdir(os.path.join(MaxwellVacuumdir))
# Write an empty __init__.py file in this directory so that Python2 can load modules from it.
with open(os.path.join(MaxwellVacuumdir, "__init__.py"), "w") as file:
pass
# +
# %%writefile $MaxwellVacuumdir/MaxwellVacuum_C_kernels_codegen.py
# Step 1: Import needed core NRPy+ modules
from outputC import lhrh # NRPy+: Core C code output module
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import loop as lp # NRPy+: Generate C code loops
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import os, sys # Standard Python modules for multiplatform OS-level functions
import time # Standard Python module; useful for benchmarking
import Maxwell.VacuumMaxwell_Flat_Evol_Cartesian as rhs
def MaxwellVacuum_C_kernels_codegen_onepart(params=
"WhichPart=Maxwell_RHSs,ThornName=MaxwellVacuum,FD_order=4"):
# Set default parameters
WhichPart = "Maxwell_RHSs"
ThornName = "MaxwellVacuum"
FD_order = 4
import re
if params != "":
params2 = re.sub("^,","",params)
params = params2.strip()
splitstring = re.split("=|,", params)
# if len(splitstring) % 2 != 0:
# print("outputC: Invalid params string: "+params)
# sys.exit(1)
parnm = []
value = []
for i in range(int(len(splitstring)/2)):
parnm.append(splitstring[2*i])
value.append(splitstring[2*i+1])
for i in range(len(parnm)):
parnm.append(splitstring[2*i])
value.append(splitstring[2*i+1])
for i in range(len(parnm)):
if parnm[i] == "WhichPart":
WhichPart = value[i]
elif parnm[i] == "ThornName":
ThornName = value[i]
elif parnm[i] == "FD_order":
FD_order = int(value[i])
else:
print("MaxwellVacuum Error: Could not parse input param: "+parnm[i])
sys.exit(1)
# Set output directory for C kernels
outdir = os.path.join(ThornName, "src") # Main C code output directory
# Set spatial dimension (must be 3 for Maxwell)
par.set_parval_from_str("grid::DIM",3)
# Step 2: Set some core parameters, including CoordSystem MoL timestepping algorithm,
# FD order, floating point precision, and CFL factor:
# Choices are: Spherical, SinhSpherical, SinhSphericalv2, Cylindrical, SinhCylindrical,
# SymTP, SinhSymTP
# NOTE: Only CoordSystem == Cartesian makes sense here; new
# boundary conditions are needed within the ETK for
# Spherical, etc. coordinates.
CoordSystem = "Cartesian"
par.set_parval_from_str("reference_metric::CoordSystem",CoordSystem)
rfm.reference_metric() # Create ReU, ReDD needed for rescaling B-L initial data, generating Maxwell RHSs, etc.
# Set the gridfunction memory access type to ETK-like, so that finite_difference
# knows how to read and write gridfunctions from/to memory.
par.set_parval_from_str("grid::GridFuncMemAccess","ETK")
# -
# <a id='mwev'></a>
#
# # Step 2: NRPy+-generated C code kernels for solving \[Back to [top](#toc)\]
# $$\label{mwev}$$
#
# <a id='mwevrhs'></a>
#
# ## Step 2.a: Maxwell RHS expressions \[Back to [top](#toc)\]
# $$\label{mwevrhs}$$
#
# `MaxwellVacuum` implements Maxwell's equations in flat spacetime and in Cartesian coordinates, which is fully documented within NRPy+ ([start here](Tutorial-VacuumMaxwell_formulation_Cartesian.ipynb)).
#
# Here, we simply call the [Maxwell/VacuumMaxwell_Flat_Evol_Cartesian.py](../edit/Maxwell/VacuumMaxwell_Flat_Evol_Cartesian.py); [\[**tutorial**\]](Tutorial-VacuumMaxwell_formulation_Cartesian.ipynb) NRPy+ Python module to generate the symbolic expressions and then output the finite-difference C code form of the equations using NRPy+'s [finite_difference](finite_difference.py) ([**tutorial**](Tutorial-Finite_Difference_Derivatives.ipynb)) C code generation module.
#
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_kernels_codegen.py
def Maxwell_RHSs__generate_symbolic_expressions_SystemI():
######################################
# START: GENERATE SYMBOLIC EXPRESSIONS
print("Generating symbolic expressions for Maxwell RHSs SystemI...\n")
start = time.time()
# Set which system to use, which are defined in Maxwell/InitialData.py
par.initialize_param(par.glb_param("char", "Maxwell.InitialData","System_to_use","System_I"))
par.set_parval_from_str("reference_metric::enable_rfm_precompute","True")
par.set_parval_from_str("reference_metric::rfm_precompute_Ccode_outdir",os.path.join(outdir,"rfm_files/"))
rhs.VacuumMaxwellRHSs()
end = time.time()
print("(BENCH) Finished Maxwell RHS symbolic expressions in "+str(end-start)+" seconds.\n")
# END: GENERATE SYMBOLIC EXPRESSIONS
######################################
# Step 2: Register new gridfunctions so they can be written to by NRPy.
# System I:
AIU = ixp.register_gridfunctions_for_single_rank1("EVOL","AIU")
EIU = ixp.register_gridfunctions_for_single_rank1("EVOL","EIU")
psiI = gri.register_gridfunctions("EVOL","psiI")
C_SystemI = rhs.C
Maxwell_RHSs_SymbExpressions_SystemI = [\
lhrh(lhs=gri.gfaccess("rhs_gfs","AIU0"),rhs=rhs.ArhsU[0]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","AIU1"),rhs=rhs.ArhsU[1]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","AIU2"),rhs=rhs.ArhsU[2]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","EIU0"),rhs=rhs.ErhsU[0]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","EIU1"),rhs=rhs.ErhsU[1]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","EIU2"),rhs=rhs.ErhsU[2]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","psi"),rhs=rhs.psi_rhs)]
# Avoid re-registering GFs later
del gri.glb_gridfcs_list[:8]
return [Maxwell_RHSs_SymbExpressions_SystemI, C_SystemI]
def Maxwell_RHSs__generate_symbolic_expressions_SystemII():
######################################
# START: GENERATE SYMBOLIC EXPRESSIONS
print("Generating symbolic expressions for Maxwell RHSs SystemII...\n")
start = time.time()
# Set which system to use, which are defined in Maxwell/VacuumMaxwell_Flat_Cartesian_ID.py
par.set_parval_from_str("Maxwell.InitialData::System_to_use","System_II")
# Avoid re-registering GFs
gri.glb_gridfcs_list = []
rhs.VacuumMaxwellRHSs()
end = time.time()
print("(BENCH) Finished Maxwell RHS symbolic expressions in "+str(end-start)+" seconds.\n")
# END: GENERATE SYMBOLIC EXPRESSIONS
######################################
# Register gridfunctions so they can be written to by NRPy.
# System II:
AIIU = ixp.register_gridfunctions_for_single_rank1("EVOL","AIIU")
EIIU = ixp.register_gridfunctions_for_single_rank1("EVOL","EIIU")
psiII = gri.register_gridfunctions("EVOL","psiII")
GammaII = gri.register_gridfunctions("EVOL","GammaII")
C_SystemII = rhs.C
G_SystemII = rhs.G
Maxwell_RHSs_SymbExpressions_SystemII = [\
lhrh(lhs=gri.gfaccess("rhs_gfs","AIIU0"),rhs=rhs.ArhsU[0]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","AIIU1"),rhs=rhs.ArhsU[1]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","AIIU2"),rhs=rhs.ArhsU[2]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","EIIU0"),rhs=rhs.ErhsU[0]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","EIIU1"),rhs=rhs.ErhsU[1]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","EIIU2"),rhs=rhs.ErhsU[2]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","psi"),rhs=rhs.psi_rhs),\
lhrh(lhs=gri.gfaccess("rhs_gfs","Gamma"),rhs=rhs.Gamma_rhs)]
return [Maxwell_RHSs_SymbExpressions_SystemII, C_SystemII, G_SystemII]
def Maxwell_RHSs__generate_Ccode(all_RHSs_exprs_list_SystemI, all_RHSs_exprs_list_SystemII):
Maxwell_RHSs_SymbExpressions_SystemI = all_RHSs_exprs_list_SystemI
Maxwell_RHSs_SymbExpressions_SystemII = all_RHSs_exprs_list_SystemII
print("Generating C code for Maxwell RHSs (FD_order="+str(FD_order)+") in "+par.parval_from_str("reference_metric::CoordSystem")+" coordinates.")
start = time.time()
# Store original finite-differencing order:
FD_order_orig = par.parval_from_str("finite_difference::FD_CENTDERIVS_ORDER")
# Set new finite-differencing order:
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order)
Maxwell_RHSs_string_SystemI = fin.FD_outputC("returnstring",Maxwell_RHSs_SymbExpressions_SystemI,
params="outCverbose=False,enable_SIMD=True").replace("AU","AIU")\
.replace("EU","EIU")\
.replace("psi","psiI")
Maxwell_RHSs_string_SystemII = fin.FD_outputC("returnstring",Maxwell_RHSs_SymbExpressions_SystemII,
params="outCverbose=False,enable_SIMD=True").replace("AU","AIIU")\
.replace("EU","EIIU")\
.replace("psi","psiII")\
.replace("Gamma","GammaII")
filename = "Maxwell_RHSs"+"_FD_order_"+str(FD_order)+".h"
with open(os.path.join(outdir,filename), "w") as file:
file.write(lp.loop(["i2","i1","i0"],["cctk_nghostzones[2]","cctk_nghostzones[1]","cctk_nghostzones[0]"],
["cctk_lsh[2]-cctk_nghostzones[2]","cctk_lsh[1]-cctk_nghostzones[1]","cctk_lsh[0]-cctk_nghostzones[0]"],
["1","1","SIMD_width"],
["#pragma omp parallel for",
"#include \"rfm_files/rfm_struct__SIMD_outer_read2.h\"",
r""" #include "rfm_files/rfm_struct__SIMD_outer_read1.h"
#if (defined __INTEL_COMPILER && __INTEL_COMPILER_BUILD_DATE >= 20180804)
#pragma ivdep // Forces Intel compiler (if Intel compiler used) to ignore certain SIMD vector dependencies
#pragma vector always // Forces Intel compiler (if Intel compiler used) to vectorize
#endif"""],"",
"#include \"rfm_files/rfm_struct__SIMD_inner_read0.h\"\n"+Maxwell_RHSs_string_SystemI+Maxwell_RHSs_string_SystemII))
# Restore original finite-differencing order:
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order_orig)
end = time.time()
print("(BENCH) Finished Maxwell_RHS C codegen (FD_order="+str(FD_order)+") in " + str(end - start) + " seconds.\n")
# <a id='constraints'></a>
#
# ## Step 2.b: Constraints \[Back to [top](#toc)\]
# $$\label{constraints}$$
#
# Finally output the C code for evaluating the constraints, described in [this tutorial](Tutorial-VacuumMaxwell_Cartesian_RHSs.ipynb). In the absence of numerical error, the constraints should evaluate to zero, but due to numerical (typically truncation and roundoff) error they do not. We will therefore measure constraint violations to gauge the accuracy of our simulation, and ultimately determine whether errors are dominated by numerical finite differencing (truncation) error as expected.
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_kernels_codegen.py
def Maxwell_constraints__generate_symbolic_expressions_and_C_code(C_SystemI,
C_SystemII,
G_SystemII):
DivEI = gri.register_gridfunctions("AUX", "DivEI")
DivEII = gri.register_gridfunctions("AUX", "DivEII")
GII = gri.register_gridfunctions("AUX", "GII")
# Store original finite-differencing order:
FD_order_orig = par.parval_from_str("finite_difference::FD_CENTDERIVS_ORDER")
# Set new finite-differencing order:
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order)
start = time.time()
print("Generating optimized C code for constraints.")
Constraints_string_SystemI = fin.FD_outputC("returnstring",
[lhrh(lhs=gri.gfaccess("aux_gfs", "DivEI"), rhs=C_SystemI)],
params="outCverbose=False").replace("AU","AIU")\
.replace("EU","EIU")\
.replace("psi","psiI")
Constraints_string_SystemII = fin.FD_outputC("returnstring",
[lhrh(lhs=gri.gfaccess("aux_gfs", "DivEII"), rhs=C_SystemII),
lhrh(lhs=gri.gfaccess("aux_gfs", "GII"), rhs=G_SystemII)],
params="outCverbose=False").replace("AU","AIIU")\
.replace("EU","EIIU")\
.replace("psi","psiII")\
.replace("Gamma","GammaII")
with open(os.path.join(outdir,"Maxwell_constraints"+"_FD_order_"+str(FD_order)+".h"), "w") as file:
file.write(lp.loop(["i2","i1","i0"],["cctk_nghostzones[2]","cctk_nghostzones[1]","cctk_nghostzones[0]"],
["cctk_lsh[2]-cctk_nghostzones[2]","cctk_lsh[1]-cctk_nghostzones[1]","cctk_lsh[0]-cctk_nghostzones[0]"],
["1","1","1"],["#pragma omp parallel for","",""], "",Constraints_string_SystemI+Constraints_string_SystemII))
# Restore original finite-differencing order:
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order_orig)
end = time.time()
print("(BENCH) Finished constraint C codegen (FD_order="+str(FD_order)+") in " + str(end - start) + " seconds.\n")
# <a id='driver_call_codegen_funcs'></a>
#
# ## Step 2.d: Given `WhichPart` parameter choice, generate algorithm for calling corresponding function within `MaxwellVacuum_C_kernels_codegen_onepart()` to generate C code kernel \[Back to [top](#toc)\]
# $$\label{driver_call_codegen_funcs}$$
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_kernels_codegen.py
SystemI_exprs = Maxwell_RHSs__generate_symbolic_expressions_SystemI()
SystemII_exprs = Maxwell_RHSs__generate_symbolic_expressions_SystemII()
if WhichPart=="Maxwell_RHSs":
Maxwell_RHSs__generate_Ccode(SystemI_exprs[0], SystemII_exprs[0])
elif WhichPart=="Maxwell_constraints":
Maxwell_constraints__generate_symbolic_expressions_and_C_code(SystemI_exprs[1],
SystemII_exprs[1],
SystemII_exprs[2])
else:
print("Error: WhichPart = "+WhichPart+" is not recognized.")
sys.exit(1)
# Avoid re-registering GFs
del gri.glb_gridfcs_list[7:17]
# Store all NRPy+ environment variables to an output string so NRPy+ environment from within this subprocess can be easily restored
import pickle
# https://www.pythonforthelab.com/blog/storing-binary-data-and-serializing/
outstr = []
outstr.append(pickle.dumps(len(gri.glb_gridfcs_list)))
for lst in gri.glb_gridfcs_list:
outstr.append(pickle.dumps(lst.gftype))
outstr.append(pickle.dumps(lst.name))
outstr.append(pickle.dumps(lst.rank))
outstr.append(pickle.dumps(lst.DIM))
outstr.append(pickle.dumps(len(par.glb_params_list)))
for lst in par.glb_params_list:
outstr.append(pickle.dumps(lst.type))
outstr.append(pickle.dumps(lst.module))
outstr.append(pickle.dumps(lst.parname))
outstr.append(pickle.dumps(lst.defaultval))
outstr.append(pickle.dumps(len(par.glb_Cparams_list)))
for lst in par.glb_Cparams_list:
outstr.append(pickle.dumps(lst.type))
outstr.append(pickle.dumps(lst.module))
outstr.append(pickle.dumps(lst.parname))
outstr.append(pickle.dumps(lst.defaultval))
return outstr
# <a id='kernel_codegen'></a>
#
# ## Step 2.c: Generate C code kernels for `MaxwellVacuum` \[Back to [top](#toc)\]
# $$\label{kernel_codegen}$$
#
# Here we generate the C code kernels (i.e., the C-code representation of the equations needed) for `MaxwellVacuum`.
#
# <a id='feature_choice'></a>
#
# ### Step 2.c.i: Set compile-time and runtime parameters for `MaxwellVacuum` \[Back to [top](#toc)\]
# $$\label{feature_choice}$$
#
# NRPy+ is a code generation package that is designed to offer maximum flexibility *at the time of C code generation*. As a result, although NRPy+ can in principle output an infinite variety of C code kernels for solving systems of partial differential equations, generally free parameters in each kernel steerable at *runtime* are restricted to simple scalars. This leads to more optimized kernels, but at the expense of flexibility in generating multiple kernels (e.g. one per finite-differencing order). Reducing the number of kernels and adding more flexibility at runtime will be a focus of future work.
#
# For now, `MaxwellVacuum` supports the following runtime options:
#
# * `MaxwellVacuum`: Evolution of the Maxwell's equations.
# * Finite differencing of orders 2, 4, 6, and 8 via runtime parameter `FD_order`
#
# Next we set up the default parameter lists for `MaxwellVacuum_C_kernels_codegen_onepart()` for the `MaxwellVacuum` thorn. We set these parameter lists as strings to make parallelizing the code generation far easier (easier to pass a list of strings than a list of function arguments to Python's `multiprocessing.Pool()`).
# +
# Step 2.e.i: Set compile-time and runtime parameters for Maxwell and MaxwellVacuum
# Runtime parameters for
# MaxwellVacuum: FD_orders = [2,4,6,8];
paramslist = []
FD_orders = [2,4,6,8]
WhichParamSet = 0
ThornName = "MaxwellVacuum"
for WhichPart in ["Maxwell_RHSs","Maxwell_constraints"]:
for FD_order in FD_orders:
paramstr = "WhichPart="+WhichPart+","
paramstr+= "ThornName="+ThornName+","
paramstr+= "FD_order="+str(FD_order)+","
paramslist.append(paramstr)
WhichParamSet = WhichParamSet + 1
paramslist.sort() # Sort the list alphabetically.
# -
# <a id='parallel_codegen'></a>
#
# ### Step 2.e.ii: Generate all C-code kernels for `MaxwellVacuum`, in parallel if possible \[Back to [top](#toc)\]
# $$\label{parallel_codegen}$$
# +
nrpy_dir_path = os.path.join(".")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
# Create all output directories if they do not yet exist
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
for ThornName in ["MaxwellVacuum"]:
outrootdir = ThornName
cmd.mkdir(os.path.join(outrootdir))
outdir = os.path.join(outrootdir,"src") # Main C code output directory
# Copy SIMD/SIMD_intrinsics.h to $outdir/SIMD/SIMD_intrinsics.h, replacing
# the line "#define REAL_SIMD_ARRAY REAL" with "#define REAL_SIMD_ARRAY CCTK_REAL"
# (since REAL is undefined in the ETK, but CCTK_REAL takes its place)
cmd.mkdir(os.path.join(outdir,"SIMD"))
import fileinput
f = fileinput.input(os.path.join(nrpy_dir_path,"SIMD","SIMD_intrinsics.h"))
with open(os.path.join(outdir,"SIMD","SIMD_intrinsics.h"),"w") as outfile:
for line in f:
outfile.write(line.replace("#define REAL_SIMD_ARRAY REAL", "#define REAL_SIMD_ARRAY CCTK_REAL"))
# Create directory for rfm_files output
cmd.mkdir(os.path.join(outdir,"rfm_files"))
# Start parallel C code generation (codegen)
# NRPyEnvVars stores the NRPy+ environment from all the subprocesses in the following
# parallel codegen
NRPyEnvVars = []
import time # Standard Python module for benchmarking
import logging
start = time.time()
if __name__ == "__main__":
# try:
# if os.name == 'nt':
# # Windows & Jupyter multiprocessing do not mix, so we run in serial on Windows.
# # Here's why: https://stackoverflow.com/questions/45719956/python-multiprocessing-in-jupyter-on-windows-attributeerror-cant-get-attribut
# raise Exception("Parallel codegen currently not available in Windows")
# # Step 3.d.ii: Import the multiprocessing module.
# import multiprocessing
# print("***************************************")
# print("Starting parallel C kernel codegen...")
# print("***************************************")
# # Step 3.d.iii: Define master function for parallelization.
# # Note that lambdifying this doesn't work in Python 3
# def master_func(i):
# import MaxwellNRPy_py_dir.MaxwellVacuum_C_kernels_codegen as MCk
# return MCk.MaxwellVacuum_C_kernels_codegen_onepart(params=paramslist[i])
# # Step 3.d.iv: Evaluate list of functions in parallel if possible;
# # otherwise fallback to serial evaluation:
# pool = multiprocessing.Pool() #processes=len(paramslist))
# NRPyEnvVars.append(pool.map(master_func,range(len(paramslist))))
# pool.terminate()
# pool.join()
# except:
# logging.exception("Ignore this warning/backtrace if on a system in which serial codegen is necessary:")
print("***************************************")
print("Starting serial C kernel codegen...")
print("***************************************")
# Steps 3.d.ii-iv, alternate: As fallback, evaluate functions in serial.
# This will happen on Android and Windows systems
import MaxwellNRPy_py_dir.MaxwellVacuum_C_kernels_codegen as MCk
import grid as gri
for param in paramslist:
gri.glb_gridfcs_list = []
MCk.MaxwellVacuum_C_kernels_codegen_onepart(params=param)
NRPyEnvVars = [] # Reset NRPyEnvVars in case multiprocessing wrote to it and failed.
# # Steps 3.d.ii-iv, alternate: As fallback, evaluate functions in serial.
# # This will happen on Android and Windows systems
print("(BENCH) Finished C kernel codegen for MaxwellVacuum in "+str(time.time()-start)+" seconds.\n")
# -
# %tb
# <a id='cclfiles'></a>
#
# # Step 3: ETK `ccl` file generation \[Back to [top](#toc)\]
# $$\label{cclfiles}$$
#
# The Einstein Toolkit (ETK) ccl files contain runtime parameters (`param.ccl`), registered gridfunctions (`interface.ccl`), and function scheduling (`schedule.ccl`). As parameters and gridfunctions are registered with NRPy+ when the C-code kernels are generated, and this generation occurs on separate processes in parallel, we store the entire NRPy+ environment for *each* process. This results in a tremendous amount of duplication, which is sorted out next. Once all duplicated environment variables (e.g., registered gridfunctions) are removed, we replace the current NRPy+ environment with the new one, by setting `gri.glb_gridfcs_list[],par.glb_params_list[],par.glb_Cparams_list[]`.
# +
# Store all NRPy+ environment variables to file so NRPy+ environment from within this subprocess can be easily restored
import pickle # Standard Python module for converting arbitrary data structures to a uniform format.
import grid as gri # NRPy+: Functions having to do with numerical grids
import NRPy_param_funcs as par # NRPy+: Parameter interface
if len(NRPyEnvVars) > 0:
# https://www.pythonforthelab.com/blog/storing-binary-data-and-serializing/
grfcs_list = []
param_list = []
Cparm_list = []
for WhichParamSet in NRPyEnvVars[0]:
# gridfunctions
i=0
# print("Length of WhichParamSet:",str(len(WhichParamSet)))
num_elements = pickle.loads(WhichParamSet[i]); i+=1
for lst in range(num_elements):
grfcs_list.append(gri.glb_gridfc(gftype=pickle.loads(WhichParamSet[i+0]),
name =pickle.loads(WhichParamSet[i+1]),
rank =pickle.loads(WhichParamSet[i+2]),
DIM =pickle.loads(WhichParamSet[i+3]))) ; i+=4
# parameters
num_elements = pickle.loads(WhichParamSet[i]); i+=1
for lst in range(num_elements):
param_list.append(par.glb_param(type =pickle.loads(WhichParamSet[i+0]),
module =pickle.loads(WhichParamSet[i+1]),
parname =pickle.loads(WhichParamSet[i+2]),
defaultval=pickle.loads(WhichParamSet[i+3]))) ; i+=4
# Cparameters
num_elements = pickle.loads(WhichParamSet[i]); i+=1
for lst in range(num_elements):
Cparm_list.append(par.glb_Cparam(type =pickle.loads(WhichParamSet[i+0]),
module =pickle.loads(WhichParamSet[i+1]),
parname =pickle.loads(WhichParamSet[i+2]),
defaultval=pickle.loads(WhichParamSet[i+3]))) ; i+=4
grfcs_list_uniq = []
for gf_ntuple_stored in grfcs_list:
found_gf = False
for gf_ntuple_new in grfcs_list_uniq:
if gf_ntuple_new == gf_ntuple_stored:
found_gf = True
if found_gf == False:
grfcs_list_uniq.append(gf_ntuple_stored)
param_list_uniq = []
for pr_ntuple_stored in param_list:
found_pr = False
for pr_ntuple_new in param_list_uniq:
if pr_ntuple_new == pr_ntuple_stored:
found_pr = True
if found_pr == False:
param_list_uniq.append(pr_ntuple_stored)
# Set glb_paramsvals_list:
# Step 1: Reset all paramsvals to their defaults
par.glb_paramsvals_list = []
for parm in param_list_uniq:
par.glb_paramsvals_list.append(parm.defaultval)
Cparm_list_uniq = []
for Cp_ntuple_stored in Cparm_list:
found_Cp = False
for Cp_ntuple_new in Cparm_list_uniq:
if Cp_ntuple_new == Cp_ntuple_stored:
found_Cp = True
if found_Cp == False:
Cparm_list_uniq.append(Cp_ntuple_stored)
gri.glb_gridfcs_list = []
par.glb_params_list = []
par.glb_Cparams_list = []
gri.glb_gridfcs_list = grfcs_list_uniq
par.glb_params_list = param_list_uniq
par.glb_Cparams_list = Cparm_list_uniq
# -
# <a id='paramccl'></a>
#
# ## Step 3.a: `param.ccl`: specify free parameters within `MaxwellVacuum` \[Back to [top](#toc)\]
# $$\label{paramccl}$$
#
# All parameters necessary to evolve the right-hand side (RHS) expressions of Maxwell's equations are registered within NRPy+; we use this information to automatically generate `param.ccl`. NRPy+ also specifies default values for each parameter.
#
# More information on `param.ccl` syntax can be found in the [official Einstein Toolkit documentation](http://einsteintoolkit.org/usersguide/UsersGuide.html#x1-184000D2.3).
# +
# %%writefile $MaxwellVacuumdir/MaxwellVacuum_ETK_ccl_files_codegen.py
# Step 1: Import needed core NRPy+ modules
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import os, sys # Standard Python modules for multiplatform OS-level functions
def keep_param__return_type(paramtuple):
keep_param = True # We'll not set some parameters in param.ccl;
# e.g., those that should be #define'd like M_PI.
typestring = ""
# Separate thorns within the ETK take care of grid/coordinate parameters;
# thus we ignore NRPy+ grid/coordinate parameters:
if paramtuple.module == "grid" or paramtuple.module == "reference_metric":
keep_param = False
partype = paramtuple.type
if partype == "bool":
typestring += "BOOLEAN "
elif partype == "REAL":
if paramtuple.defaultval != 1e300: # 1e300 is a magic value indicating that the C parameter should be mutable
typestring += "CCTK_REAL "
else:
keep_param = False
elif partype == "int":
typestring += "CCTK_INT "
elif partype == "#define":
keep_param = False
elif partype == "char":
# FIXME: char/string parameter types should in principle be supported
print("Error: parameter "+paramtuple.module+"::"+paramtuple.parname+
" has unsupported type: \""+ paramtuple.type + "\"")
sys.exit(1)
else:
print("Error: parameter "+paramtuple.module+"::"+paramtuple.parname+
" has unsupported type: \""+ paramtuple.type + "\"")
sys.exit(1)
return keep_param, typestring
def output_param_ccl(ThornName="MaxwellVacuum"):
with open(os.path.join(ThornName,"param.ccl"), "w") as file:
file.write("""
# This param.ccl file was automatically generated by NRPy+.
# You are advised against modifying it directly; instead
# modify the Python code that generates it.
shares: MethodOfLines
#EXTENDS CCTK_KEYWORD evolution_method "evolution_method"
#{
# "MaxwellVacuum" :: ""
#}
restricted:
CCTK_INT FD_order "Finite-differencing order"
{\n""")
FDorders = []
for _root, _dirs, files in os.walk(os.path.join(ThornName,"src")): # _root,_dirs unused.
for Cfilename in files:
if (".h" in Cfilename) and ("RHSs" in Cfilename) and ("intrinsics" not in Cfilename):
array = Cfilename.replace(".","_").split("_")
FDorders.append(int(array[-2]))
FDorders.sort()
for order in FDorders:
file.write(" "+str(order)+":"+str(order)+" :: \"finite-differencing order = "+str(order)+"\"\n")
FDorder_default = 4
if FDorder_default not in FDorders:
print("WARNING: 4th-order FD kernel was not output!?! Changing default FD order to "+str(FDorders[0]))
FDorder_default = FDorders[0]
file.write("} "+str(FDorder_default)+ "\n\n") # choose 4th order by default, consistent with ML_Maxwell
paramccl_str = ""
for i in range(len(par.glb_Cparams_list)):
# keep_param is a boolean indicating whether we should accept or reject
# the parameter. singleparstring will contain the string indicating
# the variable type.
keep_param, singleparstring = keep_param__return_type(par.glb_Cparams_list[i])
if keep_param:
parname = par.glb_Cparams_list[i].parname
partype = par.glb_Cparams_list[i].type
singleparstring += parname + " \""+ parname +" (see NRPy+ for parameter definition)\"\n"
singleparstring += "{\n"
if partype != "bool":
singleparstring += " *:* :: \"All values accepted. NRPy+ does not restrict the allowed ranges of parameters yet.\"\n"
singleparstring += "} "+str(par.glb_Cparams_list[i].defaultval)+"\n\n"
paramccl_str += singleparstring
file.write(paramccl_str)
# -
# <a id='interfaceccl'></a>
#
# ## Step 3.b: `interface.ccl`: define needed gridfunctions; provide keywords denoting what this thorn provides and what it should inherit from other thorns \[Back to [top](#toc)\]
# $$\label{interfaceccl}$$
#
# `interface.ccl` declares all gridfunctions and determines how `MaxwellVacuum` interacts with other Einstein Toolkit thorns.
#
# The [official Einstein Toolkit (Cactus) documentation](http://einsteintoolkit.org/usersguide/UsersGuide.html#x1-179000D2.2) defines what must/should be included in an `interface.ccl` file.
# +
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_ETK_ccl_files_codegen.py
# First construct lists of the basic gridfunctions used in NRPy+.
# Each type will be its own group in MaxwellVacuum.
evol_gfs_list = []
auxevol_gfs_list = []
aux_gfs_list = []
for i in range(len(gri.glb_gridfcs_list)):
if gri.glb_gridfcs_list[i].gftype == "EVOL":
evol_gfs_list.append( gri.glb_gridfcs_list[i].name+"GF")
if gri.glb_gridfcs_list[i].gftype == "AUX":
aux_gfs_list.append( gri.glb_gridfcs_list[i].name+"GF")
if gri.glb_gridfcs_list[i].gftype == "AUXEVOL":
auxevol_gfs_list.append(gri.glb_gridfcs_list[i].name+"GF")
# NRPy+'s finite-difference code generator assumes gridfunctions
# are alphabetized; not sorting may result in unnecessary
# cache misses.
evol_gfs_list.sort()
aux_gfs_list.sort()
auxevol_gfs_list.sort()
rhs_list = []
for gf in evol_gfs_list:
rhs_list.append(gf.replace("GF","")+"_rhsGF")
def output_interface_ccl(ThornName="MaxwellVacuum"):
outstr = """
# This interface.ccl file was automatically generated by NRPy+.
# You are advised against modifying it directly; instead
# modify the Python code that generates it.
# With "implements", we give our thorn its unique name.
implements: MaxwellVacuum
# By "inheriting" other thorns, we tell the Toolkit that we
# will rely on variables/function that exist within those
# functions.
inherits: Boundary grid MethodofLines\n"""
outstr += """
# Needed functions and #include's:
USES INCLUDE: Symmetry.h
USES INCLUDE: Boundary.h
# Needed Method of Lines function
CCTK_INT FUNCTION MoLRegisterEvolvedGroup(CCTK_INT IN EvolvedIndex, \
CCTK_INT IN RHSIndex)
REQUIRES FUNCTION MoLRegisterEvolvedGroup
# Needed Boundary Conditions function
CCTK_INT FUNCTION GetBoundarySpecification(CCTK_INT IN size, CCTK_INT OUT ARRAY nboundaryzones, CCTK_INT OUT ARRAY is_internal, CCTK_INT OUT ARRAY is_staggered, CCTK_INT OUT ARRAY shiftout)
USES FUNCTION GetBoundarySpecification
CCTK_INT FUNCTION SymmetryTableHandleForGrid(CCTK_POINTER_TO_CONST IN cctkGH)
USES FUNCTION SymmetryTableHandleForGrid
CCTK_INT FUNCTION Boundary_SelectVarForBC(CCTK_POINTER_TO_CONST IN GH, CCTK_INT IN faces, CCTK_INT IN boundary_width, CCTK_INT IN table_handle, CCTK_STRING IN var_name, CCTK_STRING IN bc_name)
USES FUNCTION Boundary_SelectVarForBC
# Needed for EinsteinEvolve/NewRad outer boundary condition driver:
CCTK_INT FUNCTION \\
NewRad_Apply \\
(CCTK_POINTER_TO_CONST IN cctkGH, \\
CCTK_REAL ARRAY IN var, \\
CCTK_REAL ARRAY INOUT rhs, \\
CCTK_REAL IN var0, \\
CCTK_REAL IN v0, \\
CCTK_INT IN radpower)
REQUIRES FUNCTION NewRad_Apply
# Tell the Toolkit that we want all gridfunctions
# to be visible to other thorns by using
# the keyword "public". Note that declaring these
# gridfunctions *does not* allocate memory for them;
# that is done by the schedule.ccl file.
public:
"""
# Next we declare gridfunctions based on their corresponding gridfunction groups as registered within NRPy+
def output_list_of_gfs(gfs_list,description="User did not provide description"):
gfsstr = " "
for i in range(len(gfs_list)):
gfsstr += gfs_list[i]
if i != len(gfs_list)-1:
gfsstr += "," # This is a comma-separated list of gridfunctions
else:
gfsstr += "\n} \""+description+"\"\n\n"
return gfsstr
# First EVOL type:
outstr += "CCTK_REAL evol_variables type = GF Timelevels=3\n{\n"
outstr += output_list_of_gfs(evol_gfs_list,"Maxwell evolved gridfunctions")
# Second EVOL right-hand-sides
outstr += "CCTK_REAL evol_variables_rhs type = GF Timelevels=1 TAGS=\'InterpNumTimelevels=1 prolongation=\"none\"\'\n{\n"
outstr += output_list_of_gfs(rhs_list,"right-hand-side storage for Maxwell evolved gridfunctions")
# Then AUX type:
outstr += "CCTK_REAL aux_variables type = GF Timelevels=3\n{\n"
outstr += output_list_of_gfs(aux_gfs_list,"Auxiliary gridfunctions for Maxwell diagnostics")
# Finally, AUXEVOL type:
# outstr += "CCTK_REAL auxevol_variables type = GF Timelevels=1 TAGS=\'InterpNumTimelevels=1 prolongation=\"none\"\'\n{\n"
# outstr += output_list_of_gfs(auxevol_gfs_list,"Auxiliary gridfunctions needed for evaluating the Maxwell RHSs")
with open(os.path.join(ThornName,"interface.ccl"), "w") as file:
file.write(outstr.replace("MaxwellVacuum",ThornName))
# -
# <a id='scheduleccl'></a>
#
# ## Step 3.c: `schedule.ccl`: schedule all functions used within `MaxwellVacuum`, specify data dependencies within said functions, and allocate memory for gridfunctions \[Back to [top](#toc)\]
# $$\label{scheduleccl}$$
#
# Official documentation on constructing ETK `schedule.ccl` files is found [here](http://einsteintoolkit.org/usersguide/UsersGuide.html#x1-187000D2.4).
# +
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_ETK_ccl_files_codegen.py
def output_schedule_ccl(ThornName="MaxwellVacuum"):
outstr = """
# This schedule.ccl file was automatically generated by NRPy+.
# You are advised against modifying it directly; instead
# modify the Python code that generates it.
# Next allocate storage for all 3 gridfunction groups used in MaxwellVacuum
STORAGE: evol_variables[3] # Evolution variables
STORAGE: evol_variables_rhs[1] # Variables storing right-hand-sides
STORAGE: aux_variables[3] # Diagnostics variables
# The following scheduler is based on Lean/LeanMaxwellMoL/schedule.ccl
schedule MaxwellVacuum_Banner at STARTUP
{
LANG: C
OPTIONS: meta
} "Output ASCII art banner"
schedule MaxwellVacuum_Symmetry_registration at BASEGRID
{
LANG: C
OPTIONS: Global
} "Register symmetries, the CartGrid3D way."
schedule MaxwellVacuum_zero_rhss at BASEGRID after MaxwellVacuum_Symmetry_registration
{
LANG: C
} "Idea from Lean: set all rhs functions to zero to prevent spurious nans"
# MoL: registration
schedule MaxwellVacuum_MoL_registration in MoL_Register
{
LANG: C
OPTIONS: META
} "Register variables for MoL"
# MoL: compute RHSs, etc
schedule MaxwellVacuum_RHSs in MoL_CalcRHS as MaxwellVacuum_RHS
{
LANG: C
} "MoL: Evaluate Maxwell RHSs"
schedule MaxwellVacuum_NewRad in MoL_CalcRHS after MaxwellVacuum_RHS
{
LANG: C
} "NewRad boundary conditions, scheduled right after RHS eval."
schedule MaxwellVacuum_BoundaryConditions_evolved_gfs in MoL_PostStep
{
LANG: C
OPTIONS: LEVEL
SYNC: evol_variables
} "Apply boundary conditions and perform AMR+interprocessor synchronization"
schedule GROUP ApplyBCs as MaxwellVacuum_ApplyBCs in MoL_PostStep after MaxwellVacuum_BoundaryConditions_evolved_gfs
{
} "Group for applying boundary conditions"
# Compute divergence and Gamma constraints
schedule MaxwellVacuum_constraints in MoL_PseudoEvolution
{
LANG: C
OPTIONS: Local
} "Compute Maxwell (divergence and Gamma) constraints"
"""
with open(os.path.join(ThornName,"schedule.ccl"), "w") as file:
file.write(outstr.replace("MaxwellVacuum",ThornName))
# +
import MaxwellNRPy_py_dir.MaxwellVacuum_ETK_ccl_files_codegen as cclgen
ThornName="MaxwellVacuum"
cclgen.output_param_ccl(ThornName)
cclgen.output_interface_ccl(ThornName)
cclgen.output_schedule_ccl(ThornName)
# -
# <a id='cdrivers'></a>
#
# # Step 4: C driver functions for ETK registration & NRPy+ generated kernels \[Back to [top](#toc)\]
# $$\label{cdrivers}$$
#
# Now that we have constructed the basic C code kernels and the needed Einstein Toolkit `ccl` files, we next write the driver functions for registering `MaxwellVacuum` within the Toolkit and the C code kernels. Each of these driver functions will be called directly from the thorn's [`schedule.ccl`](#scheduleccl) in the ETK.
#
# <a id='etkfunctions'></a>
# ## Step 4.a: Needed ETK functions: Banner, Symmetry registration, Parameter sanity check, Method of Lines (`MoL`) registration, Boundary condition \[Back to [top](#toc)\]
# $$\label{etkfunctions}$$
#
# ### To-do: Parameter sanity check function. E.g., error should be thrown if `cctk_nghostzones[]` is set too small for the chosen finite-differencing order within NRPy+.
# +
# %%writefile $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
# Step 1: Import needed core NRPy+ and Python modules
from outputC import lhrh # NRPy+: Core C code output module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import finite_difference as fin # NRPy+: Finite difference C code generation module
import grid as gri # NRPy+: Functions having to do with numerical grids
import loop as lp # NRPy+: Generate C code loops
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import os, sys # Standard Python modules for multiplatform OS-level functions
# We need the function keep_param__return_type() from this module:
import MaxwellNRPy_py_dir.MaxwellVacuum_ETK_ccl_files_codegen as ccl
make_code_defn_list = []
def append_to_make_code_defn_list(filename):
if filename not in make_code_defn_list:
make_code_defn_list.append(filename)
return filename
# +
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
def driver_C_codes(Csrcdict, ThornName,
rhs_list,evol_gfs_list,aux_gfs_list,auxevol_gfs_list):
# First the ETK banner code, proudly showing the NRPy+ banner
import NRPy_logo as logo
outstr = """
#include <stdio.h>
void MaxwellVacuum_Banner()
{
"""
logostr = logo.print_logo(print_to_stdout=False)
outstr += "printf(\"MaxwellVacuum: another Einstein Toolkit thorn generated by\\n\");\n"
for line in logostr.splitlines():
outstr += " printf(\""+line+"\\n\");\n"
outstr += "}\n"
# Finally add C code string to dictionaries (Python dictionaries are immutable)
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("Banner.c")] = outstr.replace("MaxwellVacuum",ThornName)
# -
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
# Next MaxwellVacuum_Symmetry_registration(): Register symmetries
full_gfs_list = []
full_gfs_list.extend(evol_gfs_list)
full_gfs_list.extend(auxevol_gfs_list)
full_gfs_list.extend(aux_gfs_list)
outstr = """
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
#include "Symmetry.h"
void MaxwellVacuum_Symmetry_registration(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
// Stores gridfunction parity across x=0, y=0, and z=0 planes, respectively
int sym[3];
// Next register parities for each gridfunction based on its name
// (to ensure this algorithm is robust, gridfunctions with integers
// in their base names are forbidden in NRPy+).
"""
outstr += ""
for gfname in full_gfs_list:
gfname_without_GFsuffix = gfname[:-2]
outstr += """
// Default to scalar symmetry:
sym[0] = 1; sym[1] = 1; sym[2] = 1;
// Now modify sym[0], sym[1], and/or sym[2] as needed
// to account for gridfunction parity across
// x=0, y=0, and/or z=0 planes, respectively
"""
# If gridfunction name does not end in a digit, by NRPy+ syntax, it must be a scalar
if gfname_without_GFsuffix[len(gfname_without_GFsuffix) - 1].isdigit() == False:
outstr += " // (this gridfunction is a scalar -- no need to change default sym[]'s!)\n"
elif len(gfname_without_GFsuffix) > 2:
# Rank-1 indexed expression (e.g., vector)
if gfname_without_GFsuffix[len(gfname_without_GFsuffix) - 2].isdigit() == False:
if int(gfname_without_GFsuffix[-1]) > 2:
print("Error: Found invalid gridfunction name: "+gfname)
sys.exit(1)
symidx = gfname_without_GFsuffix[-1]
if int(symidx) < 3: outstr += " sym[" + symidx + "] = -1;\n"
# Rank-2 indexed expression
elif gfname_without_GFsuffix[len(gfname_without_GFsuffix) - 2].isdigit() == True:
if len(gfname_without_GFsuffix) > 3 and gfname_without_GFsuffix[len(gfname_without_GFsuffix) - 3].isdigit() == True:
print("Error: Found a Rank-3 or above gridfunction: "+gfname+", which is at the moment unsupported.")
print("It should be easy to support this if desired.")
sys.exit(1)
symidx0 = gfname_without_GFsuffix[-2]
if int(symidx0) >= 0: outstr += " sym[" + symidx0 + "] *= -1;\n"
symidx1 = gfname_without_GFsuffix[-1]
if int(symidx1) >= 0: outstr += " sym[" + symidx1 + "] *= -1;\n"
else:
print("Don't know how you got this far with a gridfunction named "+gfname+", but I'll take no more of this nonsense.")
print(" Please follow best-practices and rename your gridfunction to be more descriptive")
sys.exit(1)
outstr += " SetCartSymVN(cctkGH, sym, \"MaxwellVacuum::" + gfname + "\");\n"
outstr += "}\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("Symmetry_registration_oldCartGrid3D.c")] = \
outstr.replace("MaxwellVacuum",ThornName)
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
# Next set RHSs to zero
outstr = """
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
#include "Symmetry.h"
void MaxwellVacuum_zero_rhss(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
"""
set_rhss_to_zero = ""
for gf in rhs_list:
set_rhss_to_zero += gf+"[CCTK_GFINDEX3D(cctkGH,i0,i1,i2)] = 0.0;\n"
outstr += lp.loop(["i2","i1","i0"],["0", "0", "0"],
["cctk_lsh[2]","cctk_lsh[1]","cctk_lsh[0]"],
["1","1","1"],
["#pragma omp parallel for","","",],"",set_rhss_to_zero)
outstr += "}\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("zero_rhss.c")] = outstr.replace("MaxwellVacuum",ThornName)
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
# Next registration with the Method of Lines thorn
outstr = """
//--------------------------------------------------------------------------
// Register with the Method of Lines time stepper
// (MoL thorn, found in arrangements/CactusBase/MoL)
// MoL documentation located in arrangements/CactusBase/MoL/doc
//--------------------------------------------------------------------------
#include <stdio.h>
#include "cctk.h"
#include "cctk_Parameters.h"
#include "cctk_Arguments.h"
#include "Symmetry.h"
void MaxwellVacuum_MoL_registration(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
CCTK_INT ierr = 0, group, rhs;
// Register evolution & RHS gridfunction groups with MoL, so it knows
group = CCTK_GroupIndex("MaxwellVacuum::evol_variables");
rhs = CCTK_GroupIndex("MaxwellVacuum::evol_variables_rhs");
ierr += MoLRegisterEvolvedGroup(group, rhs);
if (ierr) CCTK_ERROR("Problems registering with MoL");
}
"""
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("MoL_registration.c")] = outstr.replace("MaxwellVacuum",ThornName)
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
# Next register with the boundary conditions thorns.
# PART 1: Set BC type to "none" for all variables
# Since we choose NewRad boundary conditions, we must register all
# gridfunctions to have boundary type "none". This is because
# NewRad is seen by the rest of the Toolkit as a modification to the
# RHSs.
# This code is based on Kranc's McLachlan/ML_Maxwell/src/Boundaries.cc code.
outstr = """
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
#include "cctk_Faces.h"
#include "util_Table.h"
#include "Symmetry.h"
// Set `none` boundary conditions on Maxwell RHSs, as these are set via NewRad.
void MaxwellVacuum_BoundaryConditions_evolved_gfs(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
CCTK_INT ierr CCTK_ATTRIBUTE_UNUSED = 0;
"""
for gf in evol_gfs_list:
outstr += """
ierr = Boundary_SelectVarForBC(cctkGH, CCTK_ALL_FACES, 1, -1, "MaxwellVacuum::"""+gf+"""", "none");
if (ierr < 0) CCTK_ERROR("Failed to register BC for MaxwellVacuum::"""+gf+"""!");
"""
outstr += """
}
// Set `none` boundary conditions on Maxwell constraints
void MaxwellVacuum_BoundaryConditions_aux_gfs(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
CCTK_INT ierr CCTK_ATTRIBUTE_UNUSED = 0;
"""
for gf in aux_gfs_list:
outstr += """
ierr = Boundary_SelectVarForBC(cctkGH, CCTK_ALL_FACES, cctk_nghostzones[0], -1, "MaxwellVacuum::"""+gf+"""", "none");
if (ierr < 0) CCTK_ERROR("Failed to register BC for MaxwellVacuum::"""+gf+"""!");
"""
outstr += "}\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("BoundaryConditions.c")] = outstr.replace("MaxwellVacuum",ThornName)
# PART 2: Set C code for calling NewRad BCs
# As explained in lean_public/LeanMaxwellMoL/src/calc_mwev_rhs.F90,
# the function NewRad_Apply takes the following arguments:
# NewRad_Apply(cctkGH, var, rhs, var0, v0, radpower),
# which implement the boundary condition:
# var = var_at_infinite_r + u(r-var_char_speed*t)/r^var_radpower
# Obviously for var_radpower>0, var_at_infinite_r is the value of
# the variable at r->infinity. var_char_speed is the propagation
# speed at the outer boundary, and var_radpower is the radial
# falloff rate.
outstr = """
#include <math.h>
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
void MaxwellVacuum_NewRad(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
"""
for gf in evol_gfs_list:
var_at_infinite_r = "0.0"
var_char_speed = "1.0"
var_radpower = "3.0"
outstr += " NewRad_Apply(cctkGH, "+gf+", "+gf.replace("GF","")+"_rhsGF, "+var_at_infinite_r+", "+var_char_speed+", "+var_radpower+");\n"
outstr += "}\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("BoundaryCondition_NewRad.c")] = outstr.replace("MaxwellVacuum",ThornName)
# <a id='mwevrhss'></a>
#
# ## Step 4.b: Evaluate Maxwell right-hand-sides (RHSs) \[Back to [top](#toc)\]
# $$\label{mwevrhss}$$
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
###########################
###########################
# Maxwell_RHSs
###########################
common_includes = """
#include <math.h>
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
#include "SIMD/SIMD_intrinsics.h"
"""
common_preloop = """
DECLARE_CCTK_ARGUMENTS;
const CCTK_REAL NOSIMDinvdx0 = 1.0/CCTK_DELTA_SPACE(0);
const REAL_SIMD_ARRAY invdx0 = ConstSIMD(NOSIMDinvdx0);
const CCTK_REAL NOSIMDinvdx1 = 1.0/CCTK_DELTA_SPACE(1);
const REAL_SIMD_ARRAY invdx1 = ConstSIMD(NOSIMDinvdx1);
const CCTK_REAL NOSIMDinvdx2 = 1.0/CCTK_DELTA_SPACE(2);
const REAL_SIMD_ARRAY invdx2 = ConstSIMD(NOSIMDinvdx2);
"""
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
path = os.path.join(ThornName,"src")
MaxwellVacuum_src_filelist = []
for _root, _dirs, files in os.walk(path): # _root, _dirs unused.
for filename in files:
MaxwellVacuum_src_filelist.append(filename)
MaxwellVacuum_src_filelist.sort() # Sort the list in place.
Maxwell_FD_orders_output = []
for filename in MaxwellVacuum_src_filelist:
if "Maxwell_RHSs_" in filename:
array = filename.replace(".","_").split("_")
FDorder = int(array[-2])
if FDorder not in Maxwell_FD_orders_output:
Maxwell_FD_orders_output.append(FDorder)
Maxwell_FD_orders_output.sort()
###########################
# Output Maxwell RHSs driver function
outstr = common_includes
for filename in MaxwellVacuum_src_filelist:
if ("Maxwell_RHSs_" in filename) and (".h" in filename):
outstr += """extern void """ + ThornName+"_"+filename.replace(".h", "(CCTK_ARGUMENTS);") + "\n"
outstr += """
void MaxwellVacuum_RHSs(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
const CCTK_INT *FD_order = CCTK_ParameterGet("FD_order","MaxwellVacuum",NULL);
"""
for filename in MaxwellVacuum_src_filelist:
if ("Maxwell_RHSs_" in filename) and (".h" in filename):
array = filename.replace(".", "_").split("_")
outstr += " if(*FD_order == " + str(array[-2]) + ") {\n"
outstr += " " + ThornName+"_"+filename.replace(".h", "(CCTK_PASS_CTOC);") + "\n"
outstr += " }\n"
outstr += "} // END FUNCTION\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("Maxwell_RHSs.c")] = outstr.replace("MaxwellVacuum", ThornName)
def SIMD_declare_C_params():
SIMD_declare_C_params_str = ""
for i in range(len(par.glb_Cparams_list)):
# keep_param is a boolean indicating whether we should accept or reject
# the parameter. singleparstring will contain the string indicating
# the variable type.
keep_param, singleparstring = ccl.keep_param__return_type(par.glb_Cparams_list[i])
if (keep_param) and ("CCTK_REAL" in singleparstring):
parname = par.glb_Cparams_list[i].parname
SIMD_declare_C_params_str += " const "+singleparstring + "*NOSIMD"+parname+\
" = CCTK_ParameterGet(\""+parname+"\",\"MaxwellVacuum\",NULL);\n"
SIMD_declare_C_params_str += " const REAL_SIMD_ARRAY "+parname+" = ConstSIMD(*NOSIMD"+parname+");\n"
return SIMD_declare_C_params_str
# Create functions for the largest C kernels (Maxwell RHSs and Ricci) and output
# the .h files to .c files with function wrappers; delete original .h files
path = os.path.join(ThornName, "src")
for filename in MaxwellVacuum_src_filelist:
if ("Maxwell_RHSs_" in filename) and (".h" in filename):
outstr = common_includes + "void MaxwellVacuum_"+filename.replace(".h","")+"(CCTK_ARGUMENTS) {\n"
outstr += common_preloop+SIMD_declare_C_params()
with open(os.path.join(path,filename), "r") as currfile:
outstr += currfile.read()
# Now that we've inserted the contents of the kernel into this file,
# we delete the file containing the kernel
os.remove(os.path.join(path,filename))
outstr += "} // END FUNCTION\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list(filename.replace(".h",".c"))] = outstr.replace("MaxwellVacuum",ThornName)
# <a id='diagnostics'></a>
#
# ## Step 4.c: Diagnostics: Computing the divergence constraint \[Back to [top](#toc)\]
# $$\label{diagnostics}$$
#
# The divergence constraint is a useful diagnostics of a calculation's health. Here we construct the driver function.
# %%writefile -a $MaxwellVacuumdir/MaxwellVacuum_C_drivers_codegen.py
# Next, the driver for computing the Maxwell Hamiltonian & momentum constraints
outstr = """
#include <math.h>
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
void MaxwellVacuum_constraints(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
const CCTK_REAL invdx0 = 1.0/CCTK_DELTA_SPACE(0);
const CCTK_REAL invdx1 = 1.0/CCTK_DELTA_SPACE(1);
const CCTK_REAL invdx2 = 1.0/CCTK_DELTA_SPACE(2);
"""
for filename in MaxwellVacuum_src_filelist:
if "Maxwell_constraints_" in filename:
array = filename.replace(".","_").split("_")
outstr += " if(FD_order == "+str(array[-2])+") {\n"
outstr += " #include \""+filename+"\"\n"
outstr += " }\n"
outstr += "}\n"
# Add C code string to dictionary (Python dictionaries are immutable)
Csrcdict[append_to_make_code_defn_list("driver_constraints.c")] = outstr.replace("MaxwellVacuum",ThornName)
# <a id='outcdrivers'></a>
#
# ## Step 4.d: Output all C driver functions needed for `MaxwellVacuum` \[Back to [top](#toc)\]
# $$\label{outcdrivers}$$
#
# First we call the above functions (output above to the `MaxwellVacuum_validate.MaxwellVacuum_C_drivers_codegen` Python module) to store all needed driver C files to a Python dictionary, then we simply outputs the dictionary to the appropriate files.
# +
import MaxwellNRPy_py_dir.MaxwellVacuum_C_drivers_codegen as driver
# The following Python dictionaries consist of a key, which is the filename
# in the thorn's src/ directory (e.g., "driver_Maxwell_constraints.c"),
# and a value, which is the corresponding source code, stored as a
# Python string.
Vac_Csrcdict = {}
Reg_Csrcdict = {}
# We'll need lists of gridfunctions for these driver functions
evol_gfs_list = cclgen.evol_gfs_list
aux_gfs_list = cclgen.aux_gfs_list
auxevol_gfs_list = cclgen.auxevol_gfs_list
# Generate driver codes for MaxwellVacuum thorn (i.e., populate the Vac_Csrcdict dictionary)
driver.driver_C_codes(Vac_Csrcdict, "MaxwellVacuum",
cclgen.rhs_list,cclgen.evol_gfs_list,cclgen.aux_gfs_list,
cclgen.auxevol_gfs_list)
# Next we output the contents of the Reg_Csrcdict and
# Vac_Csrcdict dictionaries to files in the respective
# thorns' directories.
for key,val in Vac_Csrcdict.items():
with open(os.path.join("MaxwellVacuum","src",key),"w") as file:
file.write(val)
# -
# <a id='makecodedefn'></a>
#
# ## Step 4.e: `make.code.defn`: List of all C driver functions needed to compile `MaxwellVacuum` \[Back to [top](#toc)\]
# $$\label{makecodedefn}$$
#
# When constructing each C code driver function above, we called the `append_to_make_code_defn_list()` function, which built a list of each C code driver file. We'll now add each of those files to the `make.code.defn` file, used by the Einstein Toolkit's build system.
# +
# Finally output the thorns' make.code.defn files, consisting of
# a list of all C codes in the above dictionaries. This is
# part of the ETK build system so that these files are output.
def output_make_code_defn(dictionary, ThornName):
with open(os.path.join(ThornName,"src","make.code.defn"), "w") as file:
file.write("""
# Main make.code.defn file for thorn """+ThornName+"""
# Source files in this directory
SRCS =""")
filestring = ""
list_of_C_driver_files = list(dictionary.keys())
for i in range(len(list_of_C_driver_files)):
filestring += " "+list_of_C_driver_files[i]
if i != len(list_of_C_driver_files)-1:
filestring += " \\\n"
else:
filestring += "\n"
file.write(filestring)
output_make_code_defn(Vac_Csrcdict,"MaxwellVacuum")
# -
# <a id='code_validation'></a>
#
# # Step 5: Code validation \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# Here we will show plots demonstrating good error convergence and proper behavior of error nodes in the systems.
# <a id='convergence'></a>
#
# ## Step 5.a: Error Convergence \[Back to [top](#toc)\]
# $$\label{convergence}$$
#
# **Code tests adopting fourth-order finite differencing, coupled to 2nd order Iterative Crank-Nicholson method-of-lines for time integration**
#
# Inside the directory *`MaxwellVacuum/example_parfiles/`* are the files used for this convergence test:
# **maxwell_toroidaldipole-0.125_OB4.par & maxwell_toroidaldipole-0.0625_OB4.par** : ETK parameter files needed for performing the tests. These parameter files set up a toroidal dipole field propagating in a 3D numerical grid that extends from -4. to +4. along the x-, y-, and z-axes (in units of $c=1$). The parameter files are identical, except the latter has grid resolution that is twice as high as the former (so the errors should drop in the higher resolution case by a factor of $2^2$, since we adopt fourth-order finite differencing coupled to 2nd order Iterative Crank-Nicholson time integration.)
#
# **Second-order code validation test results:**
#
# The plot below shows the discrepancy between numerical and exact solutions to x-components of system I $\vec{E}$ and $\vec{A}$ at two different resolutions, at t = 2.0 (to not have errors at the boundary propagate too far inward): dashed is low resolution ($\Delta x_{\rm low}=0.125$) and solid is high resolution ($\Delta x_{\rm high}=0.0625$). Since this test adopts **fourth**-order finite differencing for spatial derivatives and **second**-order Iterative Crank-Nicholson timestepping, we would expect this error to drop by a factor of approximately $(\Delta x_{\rm low}/\Delta x_{\rm high})^2 = (0.125/0.0625)^2 = 2^2=4$ when going from low to high resolution, and after rescaling the error in the low-resolution case, we see that indeed it overlaps the high-resolution result quite nicely, confirming second-order convergence. We note that we also observe convergence for all other evolved variables (in both systems) with a nonzero exact solutions.
from IPython.display import Image
Image("MaxwellVacuum/example_parfiles/Ex-convergence.png", width=500, height=500)
Image("MaxwellVacuum/example_parfiles/Ax-convergence.png", width=500, height=500)
# <a id='errornodes'></a>
#
# ## Step 5.b: Behavior of Error Nodes \[Back to [top](#toc)\]
# $$\label{errornodes}$$
#
# Because System I is weakly hyperbolic (see [Tutorial-VacuumMaxwell_formulation_Cartesian](Tutorial-VacuumMaxwell_formulation_Cartesian.ipynb) for more discussion), zero speed error nodes of the constraint violation sit on our numerical grid, adding to the errors of our evolution variables. In contrast, System II is strongly hyperbolic, and the error nodes propagate away at the speed of light, leading to more stable evolution of the evolution variables. The plot below demostrates the qualitative behavior for both systems.
#
# Contrast these plots to Figure 1 in [Knapp, <NAME> (2002)](https://arxiv.org/abs/gr-qc/0201051); we observe excellent qualitative agreement.
Image("MaxwellVacuum/example_parfiles/constraintviolation.png", width=500, height=500)
# <a id='latex_pdf_output'></a>
#
# # Step 6: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ETK_thorn-MaxwellVacuum.pdf](Tutorial-ETK_thorn-MaxwellVacuum.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ETK_thorn-MaxwellVacuum")
| Tutorial-ETK_thorn-MaxwellVacuum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Tuples
tup_cntry_1 = ('India','SriLanka','Bangladesh','Australia','China','Nepal')
tup_cntry_2 = ('USA','Russia','Canada','UK','WestIndies','Iraq','SouthAfrica')
tup_cntry_1[0] = 'Indonasia'
#Indexing
tup_cntry_1[1:4]
# + tags=[]
## Adding the tuples
tup_cntry = tup_cntry_1 +tup_cntry_2
print(tup_cntry)
# -
# ## Tuples are immutable
tup_cntry_mod[8] = 'New York'
| Day_5/Tuple_Datatype.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import GRU, Dense
from keras.layers import LSTM
from keras import callbacks
from keras import optimizers
import pandas as pd
import tensorflow as tf
import numpy as np
df = pd.read_csv('data/international-airline-passengers.csv', index_col='Month')
print(df.head())
df.plot()
# # Data cleansing
columns_to_keep = ['Passengers']
df = df[columns_to_keep]
df['Passengers'] = df['Passengers'].apply(lambda x: x*1000)
df.index.names = ['Month']
df.sort_index(inplace=True)
print('Total rows: {}'.format(len(df)))
df.head()
df.describe()
df.plot()
# Null values?
df.isnull().sum()
null_columns=df.columns[df.isnull().any()]
df[null_columns].isnull().sum()
print(df[df.isnull().any(axis=1)][null_columns].head())
df.dropna(inplace=True)
df.isnull().sum()
df.hist(bins=10)
len(df[df['Passengers'] == 0])
# # Scaled data
print('Min', np.min(df))
print('Max', np.max(df))
# We can then extract the NumPy array from the dataframe and convert the integer values to floating point values, which are more suitable for modeling with a neural network.
dataset = df.astype('float32')
# LSTMs are sensitive to the scale of the input data, specifically when the sigmoid (default) or tanh activation functions are used. It can be a good practice to rescale the data to the range of 0-to-1, also called normalizing. We can easily normalize the dataset using the MinMaxScaler preprocessing class from the scikit-learn library.
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(dataset)
print('Min', np.min(scaled))
print('Max', np.max(scaled))
print(scaled[:10])
# # Create the RNN
# A simple method that we can use is to split the ordered dataset into train and test datasets. The code below calculates the index of the split point and separates the data into the training datasets with 70% of the observations that we can use to train our model, leaving the remaining 30% for testing the model.
# Split into train and test sets
train_size = int(len(scaled) * 0.70)
test_size = len(scaled - train_size)
train, test = scaled[0:train_size, :], scaled[train_size: len(scaled), :]
print('train: {}\ntest: {}'.format(len(train), len(test)))
# convert an array of values into a dataset matrix
# The function takes two arguments: the dataset, which is a NumPy array that we want to convert into a dataset, and the look_back, which is the number of previous time steps to use as input variables to predict the next time period — in this case defaulted to 1.
# This default will create a dataset where X is the energy quantity at a given time (t) and Y is the qty of energy at the next time (t + 1).
def create_dataset(dataset, look_back=1):
print(len(dataset), look_back)
dataX, dataY = [], []
for i in range(len(dataset)-look_back-1):
a = dataset[i:(i+look_back), 0]
print(i)
print('X {} to {}'.format(i, i+look_back))
print(a)
print('Y {}'.format(i + look_back))
print(dataset[i + look_back, 0])
dataset[i + look_back, 0]
dataX.append(a)
dataY.append(dataset[i + look_back, 0])
return np.array(dataX), np.array(dataY)
# reshape into X=t and Y=t+1
look_back = 1
X_train, y_train = create_dataset(train, look_back)
X_test, y_test = create_dataset(test, look_back)
# The LSTM network expects the input data (X) to be provided with a specific array structure in the form of: [samples, time steps, features].
#
#
# reshape input to be [samples, time steps, features]
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 1))
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
print(X_train.shape)
print(X_test.shape)
# The network has a visible layer with 1 input, a hidden layer with 4 LSTM blocks or neurons, and an output layer that makes a single value prediction. The default sigmoid activation function is used for the LSTM blocks. The network is trained for 100 epochs and a batch size of 1 is used.
#
#
# create and fit the LSTM network
batch_size = 1
model = Sequential()
model.add(LSTM(4, batch_input_shape=(batch_size, look_back, 1), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X_train, y_train, epochs=100, batch_size=batch_size, verbose=2, shuffle=True)
# Note that we invert the predictions before calculating error scores to ensure that performance is reported in the same units as the original data (thousands of passengers per month).
#
# Make preditions
# +
import math
from sklearn.metrics import mean_squared_error
trainPredict = model.predict(X_train, batch_size=batch_size)
model.reset_states()
testPredict = model.predict(X_test, batch_size=batch_size)
# invert predictions
trainPredict = scaler.inverse_transform(trainPredict)
y_train = scaler.inverse_transform([y_train])
testPredict = scaler.inverse_transform(testPredict)
y_test = scaler.inverse_transform([y_test])
# calculate root mean squared error
trainScore = math.sqrt(mean_squared_error(y_train[0], trainPredict[:,0]))
print('Train Score: %.2f RMSE' % (trainScore))
testScore = math.sqrt(mean_squared_error(y_test[0], testPredict[:,0]))
print('Test Score: %.2f RMSE' % (testScore))
# -
# Because of how the dataset was prepared, we must shift the predictions so that they align on the x-axis with the original dataset. Once prepared, the data is plotted, showing the original dataset in blue, the predictions for the training dataset in green, and the predictions on the unseen test dataset in red.
#
#
# shift train predictions for plotting
trainPredictPlot = np.empty_like(scaled)
trainPredictPlot[:, :] = np.nan
trainPredictPlot[look_back:len(trainPredict)+look_back, :] = trainPredict
# shift test predictions for plotting
testPredictPlot = np.empty_like(scaled)
testPredictPlot[:, :] = np.nan
testPredictPlot[len(trainPredict)+(look_back*2)+1:len(scaled)-1, :] = testPredict
# plot baseline and predictions
plt.figure(figsize=(20,10))
plt.plot(scaler.inverse_transform(scaled))
plt.plot(trainPredictPlot)
plt.plot(testPredictPlot)
plt.show()
| .ipynb_checkpoints/timeseries-prediction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TimeDistributed testing
from numpy import array
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import TimeDistributed
# ## One-to-One LSTM for Sequence Prediction
# prepare sequence
length = 5
seq = array([i/float(length) for i in range(length)])
X = seq.reshape(length, 1, 1)
y = seq.reshape(length, 1)
# define LSTM configuration
n_neurons = length
n_batch = length
n_epoch = 500
# create LSTM
model = Sequential()
model.add(LSTM(n_neurons, input_shape=(1, 1)))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
print(model.summary())
# train LSTM
model.fit(X, y, epochs=n_epoch, batch_size=n_batch, verbose=2)
# evaluate
result = model.predict(X, batch_size=n_batch, verbose=0)
print(X.shape, result.shape)
# print(X)
# print(result)
for value in result:
print('%.1f' % value)
# ## Many-to-One LSTM for Sequence Prediction (without TimeDistributed)
# prepare sequence
length = 5
seq = array([i/float(length) for i in range(length)])
X = seq.reshape(1, length, 1)
y = seq.reshape(1, length)
# define LSTM configuration
n_neurons = length
n_batch = 1
n_epoch = 500
# create LSTM
model = Sequential()
model.add(LSTM(n_neurons, input_shape=(length, 1)))
model.add(Dense(length))
model.compile(loss='mean_squared_error', optimizer='adam')
print(model.summary())
# train LSTM
model.fit(X, y, epochs=n_epoch, batch_size=n_batch, verbose=2)
# evaluate
result = model.predict(X, batch_size=n_batch, verbose=0)
print(X.shape, result.shape)
print(X)
print(result)
for value in result[0,:]:
print('%.1f' % value)
# ## Many-to-Many LSTM for Sequence Prediction (with TimeDistributed)
# prepare sequence
length = 19
seq = array([i/float(length) for i in range(length)])
X = seq.reshape(1, length, 1)
y = seq.reshape(1, length, 1)
# define LSTM configuration
n_neurons = 5
n_batch = 1
n_epoch = 500
# create LSTM
model = Sequential()
model.add(LSTM(n_neurons, input_shape=(length, 1), return_sequences=True))
model.add(TimeDistributed(Dense(1)))
model.compile(loss='mean_squared_error', optimizer='adam')
print(model.summary())
# ## Many-to-Many LSTM for Sequence Prediction (without TimeDistributed)
# prepare sequence
length = 19
seq = array([i/float(length) for i in range(length)])
X = seq.reshape(1, length, 1)
y = seq.reshape(1, length, 1)
# define LSTM configuration
n_neurons = 5
n_batch = 1
n_epoch = 500
# create LSTM
model = Sequential()
model.add(LSTM(n_neurons, input_shape=(length, 1), return_sequences=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
print(model.summary())
# train LSTM
model.fit(X, y, epochs=n_epoch, batch_size=n_batch, verbose=1)
# evaluate
result = model.predict(X, batch_size=n_batch, verbose=0)
print(X.shape, result.shape)
for x, value in zip(X[0, :, 0], result[0, :, 0]):
print('{:>.2f} {:>.3f}'.format(x, value))
# train LSTM
model.fit(X, y, epochs=n_epoch, batch_size=n_batch, verbose=1)
# evaluate
result = model.predict(X, batch_size=n_batch, verbose=0)
print(X.shape, result.shape)
for x, value in zip(X[0, :, 0], result[0, :, 0]):
print('{:>.2f} {:>.3f}'.format(x, value))
# +
import pandas as pd
# create a differenced series
def difference(dataset, interval=1):
diff = list()
for i in range(interval, len(dataset)):
value = dataset[i] - dataset[i - interval]
diff.append(value)
return pd.Series(diff)
# invert differenced value
def inverse_difference(history, yhat, interval=1):
return yhat + history[-interval]
# load dataset
def parser(x):
return pd.datetime.strptime('190'+x, '%Y-%m')
series = pd.read_csv('shampoo-sales.csv', header=0, parse_dates=[0], index_col=0, squeeze=True, date_parser=parser)
print(series.head())
# transform to be stationary
differenced = difference(series, 1)
print(differenced.head())
# invert transform
inverted = list()
for i in range(len(differenced)):
value = inverse_difference(series, differenced[i], len(series)-i)
inverted.append(value)
inverted = pd.Series(inverted)
print(inverted.head())
# +
batch_size = 1 # this example is too small to use anything larger than 1.
window_size = 7 # This is the size after unrolling.
n_features = 1 # Assume each red square represents a single number.
n_layers = 3 # These are the middle 3 horizontal layers.
n_neurons = 2 # One neuron for the green block(s) and one for the yellow.
n_outputs = 7 # because it's many to many.
seq = array([i/float(length) for i in range(length)])
X = seq.reshape(1, length, 1)
y = seq.reshape(1, length)
x = Input(batch_shape=(batch_size, window_size, n_features)) # bottom row of red blocks.
h = LSTM(n_neurons, return_sequences=True)(x) # hidden layer 1
h = LSTM(n_neurons, return_sequences=True)(h) # hidden layer 2
h = LSTM(n_neurons, return_sequences=True)(h) # hidden layer 3
z = TimeDistributed(Dense(n_outputs, activation='softmax'))(h) # top row of blue blocks
model = Model(inputs=[x],outputs=[z])
model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
model.summary()
# -
# # Stateful Test
# +
from pandas import DataFrame
from pandas import Series
from pandas import concat
from pandas import read_csv
from pandas import datetime
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from math import sqrt
import matplotlib
import numpy
from numpy import concatenate
# date-time parsing function for loading the dataset
def parser(x):
return datetime.strptime('190'+x, '%Y-%m')
# frame a sequence as a supervised learning problem
def timeseries_to_supervised(data, lag=1):
df = DataFrame(data)
columns = [df.shift(i) for i in range(1, lag+1)]
columns.append(df)
df = concat(columns, axis=1)
return df
# create a differenced series
def difference(dataset, interval=1):
diff = list()
for i in range(interval, len(dataset)):
value = dataset[i] - dataset[i - interval]
diff.append(value)
return Series(diff)
# invert differenced value
def inverse_difference(history, yhat, interval=1):
return yhat + history[-interval]
# scale train and test data to [-1, 1]
def scale(train, test):
# fit scaler
scaler = MinMaxScaler(feature_range=(-1, 1))
scaler = scaler.fit(train)
# transform train
train = train.reshape(train.shape[0], train.shape[1])
train_scaled = scaler.transform(train)
# transform test
test = test.reshape(test.shape[0], test.shape[1])
test_scaled = scaler.transform(test)
return scaler, train_scaled, test_scaled
# inverse scaling for a forecasted value
def invert_scale(scaler, X, yhat):
new_row = [x for x in X] + [yhat]
array = numpy.array(new_row)
array = array.reshape(1, len(array))
inverted = scaler.inverse_transform(array)
return inverted[0, -1]
# fit an LSTM network to training data
def fit_lstm(train, batch_size, nb_epoch, neurons):
X, y = train[:, 0:-1], train[:, -1]
X = X.reshape(X.shape[0], 1, X.shape[1])
model = Sequential()
model.add(LSTM(neurons, batch_input_shape=(batch_size, X.shape[1], X.shape[2]), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
for i in range(nb_epoch):
model.fit(X, y, epochs=1, batch_size=batch_size, verbose=0, shuffle=False)
model.reset_states()
return model
# make a one-step forecast
def forecast_lstm(model, batch_size, X):
X = X.reshape(1, 1, len(X))
yhat = model.predict(X, batch_size=batch_size)
return yhat[0,0]
# run a repeated experiment
def experiment(repeats, series):
# transform data to be stationary
raw_values = series.values
diff_values = difference(raw_values, 1)
# transform data to be supervised learning
supervised = timeseries_to_supervised(diff_values, 1)
supervised_values = supervised.values[1:,:]
# split data into train and test-sets
train, test = supervised_values[0:-12, :], supervised_values[-12:, :]
# transform the scale of the data
scaler, train_scaled, test_scaled = scale(train, test)
# run experiment
error_scores = list()
for r in range(repeats):
# fit the base model
lstm_model = fit_lstm(train_scaled, 1, 1000, 1)
# forecast test dataset
predictions = list()
for i in range(len(test_scaled)):
# predict
X, y = test_scaled[i, 0:-1], test_scaled[i, -1]
yhat = forecast_lstm(lstm_model, 1, X)
# invert scaling
yhat = invert_scale(scaler, X, yhat)
# invert differencing
yhat = inverse_difference(raw_values, yhat, len(test_scaled)+1-i)
# store forecast
predictions.append(yhat)
# report performance
rmse = sqrt(mean_squared_error(raw_values[-12:], predictions))
print('%d) Test RMSE: %.3f' % (r+1, rmse))
error_scores.append(rmse)
return error_scores
# execute the experiment
def run():
# load dataset
series = read_csv('shampoo-sales.csv', header=0, parse_dates=[0], index_col=0, squeeze=True, date_parser=parser)
# experiment
repeats = 10
results = DataFrame()
# run experiment
results['results'] = experiment(repeats, series)
# summarize results
print(results.describe())
# save results
results.to_csv('experiment_stateful.csv', index=False)
# entry point
run()
# -
# fit an LSTM network to training data. stateful=True, shuffle=False
def fit_lstm(train, batch_size, nb_epoch, neurons):
X, y = train[:, 0:-1], train[:, -1]
X = X.reshape(X.shape[0], 1, X.shape[1])
model = Sequential()
model.add(LSTM(neurons, batch_input_shape=(batch_size, X.shape[1], X.shape[2]), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
for i in range(nb_epoch):
model.fit(X, y, epochs=1, batch_size=batch_size, verbose=0, shuffle=False)
model.reset_states()
return model
# fit an LSTM network to training data. stateful=False, shuffle=False
def fit_lstm(train, batch_size, nb_epoch, neurons):
X, y = train[:, 0:-1], train[:, -1]
X = X.reshape(X.shape[0], 1, X.shape[1])
model = Sequential()
model.add(LSTM(neurons, batch_input_shape=(batch_size, X.shape[1], X.shape[2]), stateful=False))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X, y, epochs=nb_epoch, batch_size=batch_size, verbose=0, shuffle=False)
return model
# fit an LSTM network to training data. stateful=False, shuffle=True
def fit_lstm(train, batch_size, nb_epoch, neurons):
X, y = train[:, 0:-1], train[:, -1]
X = X.reshape(X.shape[0], 1, X.shape[1])
model = Sequential()
model.add(LSTM(neurons, batch_input_shape=(batch_size, X.shape[1], X.shape[2]), stateful=False))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X, y, epochs=nb_epoch, batch_size=batch_size, verbose=0, shuffle=True)
return model
from pandas import DataFrame
from pandas import read_csv
from matplotlib import pyplot
# load results into a dataframe
filenames = ['experiment_stateful.csv', 'experiment_stateful2.csv']
results = DataFrame()
for name in filenames:
results[name[11:-4]] = read_csv(name, header=0)
# describe all results
print(results.describe())
# box and whisker plot
results.boxplot()
pyplot.show()
from pandas import DataFrame
from pandas import read_csv
from matplotlib import pyplot
# load results into a dataframe
filenames = ['experiment_stateful.csv', 'experiment_stateless.csv', 'experiment_stateless_shuffle.csv']
results = DataFrame()
for name in filenames:
results[name[11:-4]] = read_csv(name, header=0)
# describe all results
print(results.describe())
# box and whisker plot
results.boxplot()
pyplot.show()
| rnn_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MendozaJessaMae/OOP-1-1/blob/main/OOP_Concepts_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nRgJOeoYPIIA"
# Application 1
#
# 1. Create a Python program that displays the name of three students (Student 1, Student 2, Student 3) and their term grades
# 2. Create a class name Person and attributes - std1, std2, std3, pre,mid,fin
# 3. Compute the average of each term using Grade() method
# 4. Information about student's grades must be hidden from others
# + colab={"base_uri": "https://localhost:8080/"} id="CRm4PaBbPTSy" outputId="f165679b-8cfb-4fb4-8487-602634623495"
class Person:
def __init__(self):
self.name = input("\nEnter your name:")
def Grade_input(self):
self.prelim = float(input("Enter Prelim Grades:"))
self.midterm = float(input("Enter Midterm Grades:"))
self.finals = float(input("Enter Finals Grades:"))
def Grade(self):
print("Average:",((self.prelim * .3) + (self.midterm * .3) + (self.finals * .4)))
class Student(Person):
def student_info(self):
total = self.prelim + self.midterm + self.finals
average = final//3
list = []
list.append(self.name)
list.append(total)
print(list)
S1 = Person()
S1.Grade_input()
S1.Grade()
S2 = Person()
S2.Grade_input()
S2.Grade()
S3 = Person()
S3.Grade_input()
S3.Grade()
| OOP_Concepts_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Develop Kubeflow Pipeline in Notebook
#
# This notebook shows how to build, compile and run a Kubeflow pipeline using Kubeflow SDK in Jupyter notebooks.
#
# This notebook will make sure you understand
# - Develop a Kubeflow pipeline with the KFP SDK
# - Submit Kubeflow pipelines using the KFP SDK
# - Monitor Kubeflow pipeline running status
#
# For KFP SDK details, please check documentaion [SDK Overview](https://www.kubeflow.org/docs/pipelines/sdk/sdk-overview/)
# ## Import kfp sdk
import kfp
import kfp.dsl as dsl
from kfp import compiler
# ## Develop your Kubeflow Pipeline
# ### Create pipeline component from a python function
def add_op(a, b):
return dsl.ContainerOp(
name='Add Operation',
image='python:3.6.5',
command=['sh', '-c'],
arguments=['python -c "print($0+$1)" | tee $2', str(a), str(b), '/tmp/output'],
file_outputs={
'data': '/tmp/output',
}
)
# ### Build a pipeline using the component
@dsl.pipeline(
name='Calculation pipeline',
description='A toy pipeline that performs arithmetic calculations.'
)
def calc_pipeline(
a=0,
b=7,
c=17,
):
#Passing pipeline parameter and a constant value as operation arguments
add_task = add_op(a, 4) #Returns a dsl.ContainerOp class instance.
#You can create explicit dependency between the tasks using xyz_task.after(abc_task)
add_2_task = add_op(a, b)
add_3_task = add_op(add_task.output, add_2_task.output)
# ### Compile and run the pipeline
#
# Create an Experiment in the Pipeline System
pipeline_func = calc_pipeline
pipeline_filename = pipeline_func.__name__ + '.pipeline.zip'
compiler.Compiler().compile(pipeline_func, pipeline_filename)
# ### Create an Experiment in the Pipeline System
#
# Pipeline system requires an "Experiment" to group pipeline runs. You can create a new experiment, or call client.list_experiments() to get existing ones.
#
# You can also use `default` Experiment to host your pipeline
# Get or create an experiment and submit a pipeline run
EXPERIMENT_NAME='add_operation'
client = kfp.Client()
experiment = client.create_experiment(EXPERIMENT_NAME)
# ### Submit the pipeline for execution
# +
# Specify pipeline argument values
arguments = {'a': 7, 'b': 8}
# Submit a pipeline run
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id, run_name, pipeline_filename, arguments)
# This link leads to the run information page.
# Note: There is a bug in JupyterLab that modifies the URL and makes the link stop working
| notebooks/01_Jupyter_Notebook/01_03_Notebook_Development_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (XPython)
# language: python
# name: xpython
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Monte Carlo Simulation and Linear Algebra
# + [markdown] slideshow={"slide_type": "-"} tags=["remove-cell"]
# **CS1302 Introduction to Computer Programming**
# ___
# + slideshow={"slide_type": "fragment"} tags=["remove-cell"]
# %reload_ext mytutor
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Monte Carlo simulation
# + [markdown] slideshow={"slide_type": "subslide"}
# **What is Monte Carlo simulation?**
# + [markdown] slideshow={"slide_type": "fragment"}
# > The name Monte Carlo refers to the [Monte Carlo Casino in Monaco](https://en.wikipedia.org/wiki/Monte_Carlo_Casino) where Ulam's uncle would borrow money from relatives to gamble.
# + [markdown] slideshow={"slide_type": "fragment"}
# It would be nice to simulate the casino, so Ulam's uncle did not need to borrow money to go.
# Actually...,
# - Monte Carlo is the code name of the secret project for creating the [hydrogen bomb](https://en.wikipedia.org/wiki/Monte_Carlo_method).
# - [Ulam](https://en.wikipedia.org/wiki/Stanislaw_Ulam) worked with [<NAME>](https://en.wikipedia.org/wiki/John_von_Neumann) to program the first electronic computer ENIAC to simulate a computational model of a thermonuclear reaction.
#
# (See also [The Beginning of the Monte Carlo Method](https://permalink.lanl.gov/object/tr?what=info:lanl-repo/lareport/LA-UR-88-9067) for a more detailed account.)
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to compute the value of $\pi$**?
# + [markdown] slideshow={"slide_type": "fragment"}
# An application of Monte Carlo simulation is in approximating $\pi$ using
# the [Buffon's needle](https://en.wikipedia.org/wiki/Buffon%27s_needle_problem).
# There is [a program](https://www.khanacademy.org/computer-programming/pi-by-buffons-needle/6695500989890560) written in javascript to do this.
# + [markdown] slideshow={"slide_type": "fragment"}
# The javascript program a bit long to digest, so we will use an alternative simulation that is easier to understand/program.
# + [markdown] slideshow={"slide_type": "subslide"}
# If we uniformly randomly pick a point in a square. What is the chance it is in the inscribed circle, i.e., the biggest circle inside the square?
# + [markdown] slideshow={"slide_type": "-"}
# The chance is the area of the circle divided by the area of the square. Suppose the square has length $\ell$, then the chance is
#
# $$ \frac{\pi (\ell /2)^2}{ (\ell)^2 } = \frac{\pi}4 $$
# independent of the length $\ell$.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Complete the following function to return an approximation of $\pi$ as follows:
# 1. Simulate the random process of picking a point from a square repeatedly `n` times by
# generating the $x$ and $y$ coordinates uniformly randomly from a unit interval $[0,1)$.
# 2. Compute the fraction of times the point is in the first quadrant of the inscribed circle as shown in the figure below.
# 3. Return $4$ times the fraction as the approximation.
# <p><a href="https://commons.wikimedia.org/wiki/File:Pi_30K.gif#/media/File:Pi_30K.gif"><img src="https://upload.wikimedia.org/wikipedia/commons/8/84/Pi_30K.gif" alt="Pi 30K.gif"></a></p>
# + nbgrader={"grade": false, "grade_id": "approximate_pi", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
import random, math
def approximate_pi(n):
### BEGIN SOLUTION
return 4*len([True for i in range(n)
if random.random()**2 + random.random()**2 < 1])/n
### END SOLUTION
print(f'Approximate: {approximate_pi(int(1e7))}\nGround truth: {math.pi}')
# + [markdown] slideshow={"slide_type": "subslide"}
# **How accurate is the approximation?**
# + [markdown] slideshow={"slide_type": "fragment"}
# The following uses a powerful library `numpy` for computing to return a [$95\%$-confidence interval](http://onlinestatbook.com/2/estimation/mean.html#:~:text=To%20compute%20the%2095%25%20confidence,be%20between%20the%20cutoff%20points.).
# + slideshow={"slide_type": "-"}
import numpy as np
def np_approximate_pi(n):
in_circle = (np.random.random((n,2))**2).sum(axis=-1) < 1
mean = 4 * in_circle.mean()
std = 4 * in_circle.std() / n**0.5
return np.array([mean - 2*std, mean + 2*std])
interval = np_approximate_pi(int(1e7))
print(f'''95%-confidence interval: {interval}
Estimate: {interval.mean():.4f} ± {(interval[1]-interval[0])/2:.4f}
Ground truth: {math.pi}''')
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that the computation done using `numpy` is over $5$ times faster despite the additional computation of the standard deviation.
# + [markdown] slideshow={"slide_type": "subslide"}
# There are faster methods to approximate $\pi$ such as the [Chudnovsky_algorithm](https://en.wikipedia.org/wiki/Chudnovsky_algorithm), but Monte-Carlo method is still useful in more complicated situations.
# E.g., see the Monte Carlo simulation of a [real-life situation](https://www.youtube.com/watch?v=-fCVxTTAtFQ) in playing basketball:
# > "When down by three and left with only 30 seconds is it better to attempt a hard 3-point shot or an easy 2-point shot and get another possession?" --<NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Linear Algebra
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to solve a linear equation?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Given the following linear equation in variable $x$ with real-valued coefficient $a$ and $b$,
#
# $$ a x = b,$$
# what is the value of $x$ that satisfies the equation?
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Complete the following function to return either the unique solution of $x$ or `None` if a unique solution does not exist.
# + nbgrader={"grade": false, "grade_id": "solve_linear_equation", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def solve_linear_equation(a,b):
### BEGIN SOLUTION
return b/a if a != 0 else None
### END SOLUTION
import ipywidgets as widgets
@widgets.interact(a=(0,5,1),b=(0,5,1))
def linear_equation_solver(a=2, b=3):
print(f'''linear equation: {a}x = {b}
solution: x = {solve_linear_equation(a,b)}''')
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to solve multiple linear equations?**
# + [markdown] slideshow={"slide_type": "fragment"}
# In the general case, we have a system of $m$ linear equations and $n$ variables:
#
# $$ \begin{aligned}
# a_{00} x_0 + a_{01} x_1 + \dots + a_{0(n-1)} x_{n-1} &= b_0\\
# a_{10} x_0 + a_{11} x_1 + \dots + a_{1(n-1)} x_{n-1} &= b_1\\
# \vdots\kern2em &= \vdots\\
# a_{(m-1)0} x_0 + a_{(m-1)1} x_1 + \dots + a_{(m-1)(n-1)} x_{n-1} &= b_{m-1}\\
# \end{aligned}
# $$
# where
# - $x_j$ for $j\in \{0,\dots,n-1\}$ are the variables, and
# - $a_{ij}$ and $b_j$ for $i\in \{0,\dots,m-1\}$ and $j\in \{0,\dots,n-1\}$ are the coefficients.
#
# A fundamental problem in linear algebra is to compute the unique solution to the system if it exists.
# + [markdown] slideshow={"slide_type": "fragment"}
# We will consider the simpler 2-by-2 system with 2 variables and 2 equations:
#
# $$ \begin{aligned}
# a_{00} x_0 + a_{01} x_1 &= b_0\\
# a_{10} x_0 + a_{11} x_1 &= b_1.
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# To get an idea of the solution, suppose
#
# $$a_{00}=a_{11}=1, a_{01} = a_{10} = 0.$$
# The system of equations become
#
# $$ \begin{aligned}
# x_0 \hphantom{+ x_1} &= b_0\\
# \hphantom{x_0 +} x_1 &= b_1,
# \end{aligned}
# $$
# which gives the solution directly.
# + [markdown] slideshow={"slide_type": "fragment"}
# What about $a_{00}=a_{11}=2$ instead?
#
# $$ \begin{aligned}
# 2x_0 \hphantom{+ x_1} &= b_0\\
# \hphantom{2x_0 +} 2x_1 &= b_1,
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# To obtain the solution, we simply divide both equations by 2:
#
# $$ \begin{aligned}
# x_0 \hphantom{+ x_1} &= \frac{b_0}2\\
# \hphantom{x_0 +} x_1 &= \frac{b_1}2.
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# What if $a_{01}=2$ instead?
#
# $$ \begin{aligned}
# 2x_0 + 2x_1 &= b_0\\
# \hphantom{2x_0 +} 2x_1 &= b_1\\
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# The second equation gives the solution of $x_1$, and we can use the solution in the first equation to solve for $x_0$. More precisely:
# - Subtract the second equation from the first one:
#
# $$ \begin{aligned}
# 2x_0 \hphantom{+2x_1} &= b_0 - b_1\\
# \hphantom{2x_0 +} 2x_1 &= b_1\\
# \end{aligned}
# $$
# - Divide both equation by 2:
#
# $$ \begin{aligned}
# x_0 \hphantom{+ x_1} &= \frac{b_0 - b_1}2\\
# \hphantom{x_0 +} x_1 &= \frac{b_1}2\\
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# The above operations are called *row operations* in linear algebra: each row is an equation.
# A system of linear equations can be solved by the linear operations of
# 1. multiplying an equation by a constant, and
# 2. subtracting one equation from another.
# + [markdown] slideshow={"slide_type": "fragment"}
# How to write a program to solve a general 2-by-2 system? We will use the `numpy` library.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Creating `numpy` arrays
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to store the coefficients?**
# + [markdown] slideshow={"slide_type": "fragment"}
# In linear algebra, a system of equations such as
#
# $$ \begin{aligned}
# a_{00} x_0 + a_{01} x_1 &= b_0\\
# a_{10} x_0 + a_{11} x_1 &= b_1
# \end{aligned}
# $$
# is written concisely in *matrix* form as $ \mathbf{A} \mathbf{x} = \mathbf{b} $:
#
# $$\overbrace{\begin{bmatrix}
# a_{00} & a_{01}\\
# a_{10} & a_{11}
# \end{bmatrix}}^{\mathbf{A}}
# \overbrace{
# \begin{bmatrix}
# x_0\\
# x_1
# \end{bmatrix}}
# ^{\mathbf{x}}
# = \overbrace{\begin{bmatrix}
# b_0\\
# b_1
# \end{bmatrix}}^{\mathbf{b}},
# $$
# where
# $ \mathbf{A} \mathbf{x}$ is the *matrix multiplication*
#
# $$ \mathbf{A} \mathbf{x} = \begin{bmatrix}
# a_{00} x_0 + a_{01} x_1\\
# a_{10} x_0 + a_{11} x_1
# \end{bmatrix}.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# We say that $\mathbf{A}$ is a [*matrix*](https://en.wikipedia.org/wiki/Matrix_(mathematics)) and its dimension/shape is $2$-by-$2$:
# - The first dimension/axis has size $2$. We also say that the matrix has $2$ rows.
# - The second dimension/axis has size $2$. We also say that the matrix has $2$ columns.
# $\mathbf{x}$ and $\mathbf{b}$ are called column vectors, which are matrices with one column.
# + [markdown] slideshow={"slide_type": "subslide"}
# Consider the example
# $$ \begin{aligned}
# 2x_0 + 2x_1 &= 1\\
# \hphantom{2x_0 +} 2x_1 &= 1,
# \end{aligned}$$
# or in matrix form with
# $$ \begin{aligned}
# \mathbf{A}&=\begin{bmatrix}
# a_{00} & a_{01} \\
# a_{10} & a_{11}
# \end{bmatrix}
# = \begin{bmatrix}
# 2 & 2 \\
# 0 & 2
# \end{bmatrix}\\
# \mathbf{b}&=\begin{bmatrix}
# b_0\\
# b_1
# \end{bmatrix} = \begin{bmatrix}
# 1\\
# 1
# \end{bmatrix}\end{aligned}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# Instead of using `list` to store the matrix, we will use a `numpy` array.
# + slideshow={"slide_type": "-"}
A = np.array([[2.,2],[0,2]])
b = np.array([1.,1])
A, b
# + [markdown] slideshow={"slide_type": "subslide"}
# Compared to `list`, `numpy` array is often more efficient and has more useful attributes.
# + slideshow={"slide_type": "-"}
array_attributes = set(attr for attr in dir(np.array([])) if attr[0]!='_')
list_attributes = set(attr for attr in dir(list) if attr[0]!='_')
print('\nCommon attributes:\n',*(array_attributes & list_attributes))
print('\nArray-specific attributes:\n', *(array_attributes - list_attributes))
print('\nList-specific attributes:\n',*(list_attributes - array_attributes))
# + [markdown] slideshow={"slide_type": "subslide"}
# The following attributes give the dimension/shape, number of dimensions, size, and datatype.
# + slideshow={"slide_type": "-"}
for array in A, b:
print(f'''{array}
shape: {array.shape}
ndim: {array.ndim}
size: {array.size}
dtype: {array.dtype}
''')
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that the function `len` only returns the size of the first dimension:
# + slideshow={"slide_type": "-"}
assert A.shape[0] == len(A)
len(A)
# + [markdown] slideshow={"slide_type": "subslide"}
# Unlike `list`, every `numpy` array has a data type. For efficient computation/storage, numpy implements different data types with different storage sizes:
# * integer: `int8`, `int16`, `int32`, `int64`, `uint8`, ...
# * float: `float16`, `float32`, `float64`, ...
# * complex: `complex64`, `complex128`, ...
# * boolean: `bool8`
# * Unicode: `string`
# * Object: `object`
# + [markdown] slideshow={"slide_type": "fragment"}
# E.g., `int64` is the 64-bit integer. Unlike `int`, `int64` has a range.
# + slideshow={"slide_type": "-"}
# np.int64?
print(f'range: {np.int64(-2**63)} to {np.int64(2**63-1)}')
np.int64(2**63) # overflow error
# + [markdown] slideshow={"slide_type": "subslide"}
# We can use the `astype` method to convert the data type:
# + slideshow={"slide_type": "-"}
A_int64 = A.astype(int) # converts to int64 by default
A_float32 = A.astype(np.float32) # converts to float32
for array in A_int64, A_float32:
print(array, array.dtype)
# + [markdown] slideshow={"slide_type": "fragment"}
# We have to be careful about assigning items of different types to an array.
# + slideshow={"slide_type": "-"}
A_int64[0,0] = 1
print(A_int64)
A_int64[0,0] = 0.5
print(A_int64) # intended assignment fails
np.array([int(1), float(1)]) # will be all floating point numbers
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Create a heterogeneous numpy array to store both integer and strings:
# ```Python
# [0, 1, 2, 'a', 'b', 'c']
# ```
# *Hint:* There is an numpy data type called `object`.
# + nbgrader={"grade": false, "grade_id": "hetero", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# np.object?
### BEGIN SOLUTION
heterogeneous_np_array = np.array([*range(3),*'abc'],dtype=object)
### END SOLUTION
heterogeneous_np_array
# + [markdown] slideshow={"slide_type": "fragment"}
# Be careful when creating arrays of `tuple`/`list`:
# + slideshow={"slide_type": "-"}
for array in (np.array([(1,2),[3,4,5]],dtype=object),
np.array([(1,2),[3,4]],dtype=object)):
print(array, '\nshape:', array.shape, 'length:', len(array), 'size:', array.size)
# + [markdown] slideshow={"slide_type": "subslide"}
# `numpy` provides many functions to create an array:
# + slideshow={"slide_type": "-"}
# np.zeros?
np.zeros(0), np.zeros(1), np.zeros((2,3,4)) # Dimension can be higher than 2
# + slideshow={"slide_type": "-"}
# np.ones?
np.ones(0, dtype=int), np.ones((2,3,4), dtype=int) # initialize values to int 1
# +
# np.eye?
np.eye(0), np.eye(1), np.eye(2), np.eye(3) # identity matrices
# +
# np.diag?
np.diag(range(1)), np.diag(range(2)), np.diag(np.ones(3),k=1) # diagonal matrices
# + slideshow={"slide_type": "-"}
# np.empty?
np.empty(0), np.empty((2,3,4), dtype=int) # create array faster without initialization
# + [markdown] slideshow={"slide_type": "subslide"}
# `numpy` also provides functions to build an array using rules.
# + slideshow={"slide_type": "-"}
# np.arange?
np.arange(5), np.arange(4,5), np.arange(4.5,5.5,0.5) # like range but allow non-integer parameters
# + slideshow={"slide_type": "-"}
# np.linspace?
np.linspace(4,5), np.linspace(4,5,11), np.linspace(4,5,11) # can specify number of points instead of step
# + slideshow={"slide_type": "-"}
# np.fromfunction?
np.fromfunction(lambda i, j: i * j, (3,4)) # can initialize using a function
# + [markdown] slideshow={"slide_type": "subslide"}
# We can also reshape an array using the `reshape` method/function:
# + slideshow={"slide_type": "-"}
array = np.arange(2*3*4)
# array.reshape?
(array,
array.reshape(2,3,4), # last axis index changes fastest
array.reshape(2,3,-1), # size of last axis calculated automatically
array.reshape((2,3,4), order='F')) # first axis index changes fastest
# + [markdown] slideshow={"slide_type": "fragment"}
# `flatten` is a special case of reshaping an array to one dimension.
# (Indeed, `flatten` returns a copy of the array but `reshape` returns a dynamic view whenever possible.)
# + slideshow={"slide_type": "-"}
array = np.arange(2*3*4).reshape(2,3,4)
array, array.flatten(), array.reshape(-1), array.flatten(order='F')
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Correct the following function to print every element of an array line-by-line.
# ```Python
# def print_array_entries_line_by_line(array):
# for i in array:
# print(i)
# ```
# + nbgrader={"grade": false, "grade_id": "flatten", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"} tags=["remove-output"]
def print_array_entries_line_by_line(array):
### BEGIN SOLUTION
for i in array.flatten():
print(i)
### END SOLUTION
print_array_entries_line_by_line(np.arange(2*3*4).reshape(2,3,4))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Operating on `numpy` arrays
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to verify the solution of a system of linear equations?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Before solving the system of linear equations, let us try to verify a solution to the equations:
#
# $$ \begin{aligned}
# 2x_0 + 2x_1 &= 1\\
# \hphantom{2x_0 +} 2x_1 &= 1
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# `numpy` provides the function `matmul` and the operator `@` for matrix multiplication.
# + slideshow={"slide_type": "-"}
print(np.matmul(A,np.array([0,0])) == b)
print(A @ np.array([0,0.5]) == b)
# + [markdown] slideshow={"slide_type": "subslide"}
# Note that the comparison on `numpy` arrays returns a boolean array instead of a boolean value, unlike the comparison operations on lists.
# + [markdown] slideshow={"slide_type": "fragment"}
# To check whether all items are true, we use the `all` method.
# + slideshow={"slide_type": "-"}
print((np.matmul(A,np.array([0,0])) == b).all())
print((A @ np.array([0,0.5]) == b).all())
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to concatenate arrays?**
# + [markdown] slideshow={"slide_type": "-"}
# We will operate on an augmented matrix of the coefficients:
#
# $$ \begin{aligned} \mathbf{C} &= \begin{bmatrix}
# \mathbf{A} & \mathbf{b}
# \end{bmatrix}\\
# &= \begin{bmatrix}
# a_{00} & a_{01} & b_0 \\
# a_{10} & a_{11} & b_1
# \end{bmatrix}
# \end{aligned}
# $$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# `numpy` provides functions to create block matrices:
# + slideshow={"slide_type": "-"}
# np.block?
C = np.block([A,b.reshape(-1,1)]) # reshape to ensure same ndim
C
# + [markdown] slideshow={"slide_type": "fragment"}
# To stack an array along different axes:
# + slideshow={"slide_type": "-"}
array = np.arange(1*2*3).reshape(1,2,3)
for concat_array in [array,
np.hstack((array,array)), # stack along the first axis
np.vstack((array,array)), # second axis
np.concatenate((array,array), axis=-1), # last axis
np.stack((array,array), axis=0)]: # new axis
print(concat_array, '\nshape:', concat_array.shape)
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to perform arithmetic operations on a `numpy` array?**
# + [markdown] slideshow={"slide_type": "fragment"}
# To divide all the coefficients by $2$, we can simply write:
# + slideshow={"slide_type": "-"}
D = C / 2
D
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that the above does not work for `list`.
# + slideshow={"slide_type": "-"}
C.tolist() / 2 # deep convert to list
# + [markdown] slideshow={"slide_type": "subslide"}
# Arithmetic operations on `numpy` arrays apply if the arrays have compatible dimensions. Two dimensions are compatible when
# - they are equal, except for
# - components equal to 1.
# + [markdown] slideshow={"slide_type": "fragment"}
# `numpy` uses [broadcasting rules](https://numpy.org/doc/stable/user/basics.broadcasting.html#general-broadcasting-rules) to stretch the axis of size 1 up to match the corresponding axis in other arrays.
# `C / 2` is a example where the second operand $2$ is broadcasted to a $2$-by-$2$ matrix before the elementwise division. Another example is as follows.
# + slideshow={"slide_type": "-"}
three_by_one = np.arange(3).reshape(3,1)
one_by_four = np.arange(4).reshape(1,4)
print(f'''
{three_by_one}
*
{one_by_four}
==
{three_by_one * one_by_four}
''')
# + [markdown] slideshow={"slide_type": "subslide"}
# Next, to subtract the second row of the coefficients from the first row:
# + slideshow={"slide_type": "-"}
D[0,:] = D[0,:] - D[1,:]
D
# + [markdown] slideshow={"slide_type": "fragment"}
# Notice the use of commas to index different dimensions instead of using multiple brackets:
# + slideshow={"slide_type": "-"}
assert (D[0][:] == D[0,:]).all()
# + [markdown] slideshow={"slide_type": "subslide"}
# Using this indexing technique, it is easy extract the last column as the solution to the system of linear equations:
# + slideshow={"slide_type": "-"}
x = D[:,-1]
x
# + [markdown] slideshow={"slide_type": "fragment"}
# This gives the desired solution $x_0=0$ and $x_1=0.5$ for
#
# $$ \begin{aligned}
# 2x_0 + 2x_1 &= 1\\
# \hphantom{2x_0 +} 2x_1 &= 1\\
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# `numpy` provides many [convenient ways](https://numpy.org/doc/stable/reference/arrays.indexing.html#advanced-indexing) to index an array.
# + slideshow={"slide_type": "-"}
B = np.arange(2*3).reshape(2,3)
B, B[(0,1),(2,0)] # selecting the corners using integer array
# + slideshow={"slide_type": "-"}
B = np.arange(2*3*4).reshape(2,3,4)
B, B[0], B[0,(1,2)], B[0,(1,2),(2,3)], B[:,(1,2),(2,3)] # pay attention to the last two cases
# + slideshow={"slide_type": "-"}
assert (B[...,-1] == B[:,:,-1]).all()
B[...,-1] # ... expands to selecting all elements of all previous dimensions
# + slideshow={"slide_type": "-"}
B[B>5] # indexing using boolean array
# + [markdown] slideshow={"slide_type": "subslide"}
# Finally, the following function solves a system of 2 linear equations with 2 variables.
# + slideshow={"slide_type": "-"}
def solve_2_by_2_system(A,b):
'''Returns the unique solution of the linear system, if exists,
else returns None.'''
C = np.hstack((A,b.reshape(-1,1)))
if C[0,0] == 0: C = C[(1,0),:]
if C[0,0] == 0: return None
C[0,:] = C[0,:] / C[0,0]
C[1,:] = C[1,:] - C[0,:] * C[1,0]
if C[1,1] == 0: return None
C[1,:] = C[1,:] / C[1,1]
C[0,:] = C[0,:] - C[1,:] * C[0,1]
return C[:,-1]
# + slideshow={"slide_type": "-"}
# tests
for A in (np.eye(2),
np.ones((2,2)),
np.stack((np.ones(2),np.zeros(2))),
np.stack((np.ones(2),np.zeros(2)),axis=1)):
print(f'A={A}\nb={b}\nx={solve_2_by_2_system(A,b)}\n')
# -
# ### Universal functions
# + [markdown] slideshow={"slide_type": "subslide"}
# Why does the first line of code below return two arrays but the second code return only one array? Shouldn't the first line of code return the following?
# ```Python
# array([[(0,1), (0,2), (0,3)],
# [(1,1), (1,2), (1,3)]])
# ```
# + slideshow={"slide_type": "-"}
print(np.fromfunction(lambda i,j:(i,j), (2,3), dtype=int))
print(np.fromfunction(lambda i,j:(i*j), (2,3), dtype=int))
# + [markdown] slideshow={"slide_type": "-"}
# From the documentation, `fromfunction` applies the given function to the two arrays as arguments.
# - The first line of code returns a tuple of the arrays.
# - The second line of code multiplies the two arrays to give one array, according to how multiplication works for numpy arrays.
# + [markdown] slideshow={"slide_type": "subslide"}
# Indeed, `numpy` implements [universal/vectorized functions/operators](https://numpy.org/doc/stable/reference/ufuncs.html) that take arrays as arguments and perform operations with appropriate broadcasting rules. The following is an example that uses the universal function `np.sin`:
# + slideshow={"slide_type": "-"}
import matplotlib.pyplot as plt
@widgets.interact(a=(0,5,1),b=(-1,1,0.1))
def plot_sin(a=1,b=0):
x = np.linspace(0,2*math.pi)
plt.plot(x,np.sin(a*x+b*math.pi)) # np.sin, *, + are universal functions
plt.title(r'$\sin(ax+b\pi)$')
plt.xlabel(r'$x$ (radian)')
# + [markdown] slideshow={"slide_type": "subslide"}
# In addition to making the code shorter, universal functions are both efficient and flexible. (Recall the Monte Carlo simulation to approximate $\pi$.)
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Explain how the Monte Carlo simulation work using universal functions:
# ```Python
# def np_approximate_pi(n):
# in_circle = (np.random.random((n,2))**2).sum(axis=-1) < 1
# mean = 4 * in_circle.mean()
# std = 4 * in_circle.std() / n**0.5
# return np.array([mean - 2*std, mean + 2*std])
# ```
# + [markdown] nbgrader={"grade": true, "grade_id": "universal", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false}
# - `random.random` generates a numpy array for $n$ points in the unit square randomly.
# - `sum` sums up the element along the last axis to give the squared distance.
# - `<` returns the boolean array indicating whether each point is in the first quadrant of the inscribed circle.
# - `mean` and `std` returns the mean and standard deviation of the boolean array with True and False interpreted as 1 and 0 respectively.
| source/Lecture9/Monte Carlo Simulation and Linear Algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Taylor problem 3.23
#
# last revised: 04-Jan-2020 by <NAME> [<EMAIL>]
#
# **This notebook is almost ready to go, except that the initial conditions and $\Delta v$ are different from the problem statement and there is no statement to print the figure. Fix these and you're done!**
# This is a conservation of momentum problem, which in the end lets us determine the trajectories of the two masses before and after the explosion. How should we visualize that the center-of-mass of the pieces continues to follow the original parabolic path?
# Plan:
# 1. Plot the original trajectory, also continued past the explosion time.
# 2. Plot the two trajectories after the explosion.
# 3. For some specified times of the latter two trajectories, connect the points and indicate the center of mass.
#
# The implementation here could certainly be improved! Please make suggestions (and develop improved versions).
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# First define some functions we think we will need. The formulas are based on our paper-and-pencil work.
# The trajectory starting from $t=0$ is:
#
# $
# \begin{align}
# x(t) &= x_0 + v_{x0} t \\
# y(t) &= y_0 + v_{y0} t - \frac{1}{2} g t^2
# \end{align}
# $
def trajectory(x0, y0, vx0, vy0, t_pts, g=9.8):
"""Calculate the x(t) and y(t) trajectories for an array of times,
which must start with t=0.
"""
return x0 + vx0*t_pts, y0 + vy0*t_pts - g*t_pts**2/2.
# The velocity at the final time $t_f$ is:
#
# $
# \begin{align}
# v_{x}(t) &= v_{x0} \\
# v_{y}(t) &= v_{y0} - g t_f
# \end{align}
# $
def final_velocity(vx0, vy0, t_pts, g=9.8):
"""Calculate the vx(t) and vy(t) at the end of an array of times t_pts"""
return vx0, vy0 - g*t_pts[-1] # -1 gives the last element
# The center of mass of two particles at $(x_1, y_1)$ and $(x_2, y_2)$ is:
#
# $
# \begin{align}
# x_{cm} &= \frac{1}{2}(x_1 + x_2) \\
# y_{cm} &= \frac{1}{2}(y_1 + y_2)
# \end{align}
# $
def com_position(x1, y1, x2, y2):
"""Find the center-of-mass (com) position given two positions (x,y)."""
return (x1 + x2)/2., (y1 + y2)/2.
# **1. Calculate and plot the original trajectory up to the explosion.**
# +
# initial conditions
x0_before, y0_before = [0., 0.] # put the origin at the starting point
vx0_before, vy0_before = [6., 3.] # given in the problem statement
g = 1. # as recommended
# Array of times to calculate the trajectory up to the explosion at t=4
t_pts_before = np.array([0., 1., 2., 3., 4.])
x_before, y_before = trajectory(x0_before, y0_before,
vx0_before, vy0_before,
t_pts_before, g)
# +
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(x_before, y_before, 'ro-')
ax.set_xlabel('x')
ax.set_ylabel('y')
# -
# Does it make sense so far? Note that we could use more intermediate points to make a more correct curve (rather than the piecewise straight lines) but this is fine at least for a first pass.
# **2. Calculate and plot the two trajectories after the explosion.**
#
# For the second part of the trajectory, we reset our clock to $t=0$ because that is how our trajectory function is constructed. We'll need initial positions and velocities of the pieces just after the explosion. These are the final position of the combined piece before the explosion and the final velocity plus and minus $\Delta \mathbf{v}$. We are told $\Delta \mathbf{v}$. We have to figure out the final velocity before the explosion.
# +
delta_v = np.array([2., 1.]) # change in velociy of one piece
# reset time to 0 for calculating trajectories
t_pts_after = np.array([0., 1., 2., 3., 4., 5.])
# Also could have used np.arange(0.,6.,1.)
x0_after = x_before[-1] # -1 here means the last element of the array
y0_after = y_before[-1]
vxcm0_after, vycm0_after = final_velocity(vx0_before, vy0_before,
t_pts_before, g)
# The _1 and _2 refer to the two pieces after the explosinon
vx0_after_1 = vxcm0_after + delta_v[0]
vy0_after_1 = vycm0_after + delta_v[1]
vx0_after_2 = vxcm0_after - delta_v[0]
vy0_after_2 = vycm0_after - delta_v[1]
# Given the initial conditions after the explosion, we calculate trajectories
x_after_1, y_after_1 = trajectory(x0_after, y0_after,
vx0_after_1, vy0_after_1,
t_pts_after, g)
x_after_2, y_after_2 = trajectory(x0_after, y0_after,
vx0_after_2, vy0_after_2,
t_pts_after, g)
# This is the center-of-mass trajectory
xcm_after, ycm_after = trajectory(x0_after, y0_after,
vxcm0_after, vycm0_after,
t_pts_after, g)
# These are calculated points of the center-of-mass
xcm_pts, ycm_pts = com_position(x_after_1, y_after_1, x_after_2, y_after_2)
# -
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(x_before, y_before, 'ro-', label='before explosion')
ax.plot(x_after_1, y_after_1, 'go-', label='piece 1 after')
ax.plot(x_after_2, y_after_2, 'bo-', label='piece 2 after')
ax.plot(xcm_after, ycm_after, 'r--', label='original trajectory')
ax.plot(xcm_pts, ycm_pts, 'o', color='black', label='center-of-mass of 1 and 2')
for i in range(len(t_pts_after)):
ax.plot([x_after_1[i], x_after_2[i]],
[y_after_1[i], y_after_2[i]],
'k--'
)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.legend();
| 2020_week_1/Taylor_problem_3.23.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scrape zomato data ✨ FOR RESTAUTANTS IN MUMBAI
# +
##Zomato
# #Importing packages
# import pandas as pd
# from selenium import webdriver
# from selenium.webdriver.chrome.options import Options
# from bs4 import BeautifulSoup as soup
# from time import sleep
# import random
# import csv
# driver = webdriver.Chrome("C:\\Users\\PRIYANSH\\Desktop\\Zomato\\chromedriver.exe")
# driver.get('https://developers.zomato.com/api/v2.1/categories')
# from urllib.request import urlopen as ureq
# from bs4 import BeautifulSoup
# import requests
# header = {
# "User-agent": 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_11_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/61.0.3163.100 Safari/537.36',
# "Accept": "application/json",
# "user_key": '<KEY>'
# }
# url = 'https://developers.zomato.com/api/v2.1/categories'
# page = requests.get(url)
# html = BeautifulSoup(page.content,'html.parser')
# html
# -
# # Scrape 150 pages each page contains 15 restaurant
# +
#Importing packages
import pandas as pd
from selenium import webdriver
from selenium.webdriver.chrome.options import Options
from bs4 import BeautifulSoup as soup
from time import sleep
import random
import csv
'''options = Options()
options.add_argument("--headless")
options.add_argument('--no-sandbox')
options.add_argument('--disable-gpu')
options.add_argument('start-maximized')
options.add_argument('disable-infobars')
options.add_argument("--disable-extensions")'''
CHROMEDRIVER_PATH = "C:\\Users\\PRIYANSH\\Desktop\\Zomato\\chromedriver.exe"
browser = webdriver.Chrome(CHROMEDRIVER_PATH)
def get_no_page():
try:
_page_no = browser.find_element_by_xpath(
'//*[@id="search-results-container"]/div[2]/div[1]/div[1]/div/b[2]').text
return(int(_page_no))
except:
return("cant access number of page results")
def generate_random():
#It returns a random value between 3 and 5. That number indicates the seconds to be wait
rand = random.randint(3, 5)
return rand
#Restautants or outlets in Mumbai
def extract_datasets(link):
browser.get(link)
sleep(generate_random())
contents = soup(browser.page_source, "html.parser")
al = contents.find_all('a' ,{'class': 'result-title hover_feedback zred bold ln24 fontsize0'})
return al
url = 'https://www.zomato.com/mumbai/restaurants?'
browser.get(url)
print("Total Number of Pages for this search %s" %(get_no_page()))
# get = extract_datasets(url)
pages =150 # <how many pages to scrape
get = []
for n in range(1,pages+1):
page_links = "page=%s" % (n)
web_url = url+page_links
get.append(extract_datasets(web_url))
sleep(generate_random())
browser.quit()
print(" Out of these Pages i've scraped only %s pages : " %(len(get)))
# -
names = []
urls = []
for j in range(len(get)):
lin = get[j]
for i in range(len(lin)):
#print(i , lin[i].text.strip() , lin[i].get('href'))
names.append(lin[i].text.strip())
urls.append( lin[i].get('href'))
# # sTORE DATA IN .CSV FILE
import pandas as pd
df = pd.DataFrame()
df['Restaurant_Name'] = names
df['URLs'] = urls
df.to_csv('restaurant.csv' , index = False)
data = pd.read_csv('restaurant.csv')
print(data.shape)
data.head()
# +
# #Importing packages
# import pandas as pd
# from selenium import webdriver
# from selenium.webdriver.chrome.options import Options
# from bs4 import BeautifulSoup as soup
# from time import sleep
# import random
# import csv
# '''options = Options()
# options.add_argument("--headless")
# options.add_argument('--no-sandbox')
# options.add_argument('--disable-gpu')
# options.add_argument('start-maximized')
# options.add_argument('disable-infobars')
# options.add_argument("--disable-extensions")'''
# CHROMEDRIVER_PATH = "C:\\Users\\PRIYANSH\\Desktop\\Zomato\\chromedriver.exe"
# browser = webdriver.Chrome(CHROMEDRIVER_PATH)
# def extract_datasets(link):
# browser.get(link)
# contents = soup(browser.page_source, "html.parser")
# al = contents.find_all('img')
# return al
# url ='https://www.zomato.com/mumbai/cafe-safar-oshiwara' + '/reviews'
# browser.get(url)
# get = extract_datasets(web_url)
# browser.quit()
# print(len(get))
# +
# for i in range(len(get)):
# print( i , get[i].get('alt'))
# -
# # Ultimate
# +
#Importing packages
import pandas as pd
from selenium import webdriver
from selenium.webdriver.chrome.options import Options
from bs4 import BeautifulSoup as soup
from time import sleep
import random
import csv
import re
headers = {'User-Agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_11_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/77.0.3865.120 Safari/537.36'}
'''options = Options()
options.add_argument("--headless")
options.add_argument('--no-sandbox')
options.add_argument('--disable-gpu')
options.add_argument('start-maximized')
options.add_argument('disable-infobars')
options.add_argument("--disable-extensions")'''
CHROMEDRIVER_PATH = "C:\\Users\\PRIYANSH\\Desktop\\Zomato\\chromedriver.exe"
cll = []
cnn = []
cfl =[]
cre = []
crr = []
rnn = []
rrr = []
rll = []
time = []
browser = webdriver.Chrome(CHROMEDRIVER_PATH)
for lnk in range(1 , 19):
link = 'https://www.zomato.com/mumbai/restaurants?page={}'.format(lnk)
browser.get(link)
contents = soup(browser.page_source, "html.parser")
search_list = contents.find_all("div", {'id': 'orig-search-list'})
list_content = search_list[0].find_all("div", {'class': 'content'})
print("Page_Number : " , lnk)
for i in range(0,len(list_content)): #len(list_content)
review = []
follower = []
users = []
cl = []
Time = []
cr = []
rr=[]
rn =[]
rl=[]
res_name = list_content[i].find("a", {'data-result-type': 'ResCard_Name'})
res_ratings = list_content[i].find("div", {'data-variation': 'mini inverted'})
if ratings is None:
continue
res_url = res_name.get('href')
response_url = requests.get(res_url, headers=headers)
content_url = response_url.content
soup_url = BeautifulSoup(content_url, "html.parser")
prof = soup_url.find_all('span' , {'class': "grey-text fontsize5 nowrap"})
merch_name = soup_url.find_all("div", {'class': 'header nowrap ui left'})
times = soup_url.find_all('time') #'a' , {'class': 'grey-text'}
#followers and reviews
for h in range(len(prof)):
review.append(' '.join(prof[h].text.strip().split()[:2]))
follower.append(' '.join(prof[h].text.strip().split()[3:]))
#user details
for h in range(len(merch_name)):
users.append(merch_name[h].text.strip())
cl.append(merch_name[h].a.get('href'))
#time
for h in range(len(times)):
Time.append(times[h].get('datetime'))
merch_ratings = soup_url.find_all("div", {'class': re.compile(r'ttupper fs12px left bold zdhl2 tooltip*')})
for k in range(len(merch_name)):
cr.append(merch_ratings[k].get('aria-label').split()[1])
rn.append(res_name.text.strip() )
rr.append(res_ratings.text.strip())
rl.append(res_url)
cll+=cl
cnn+=users
cfl+=follower
cre+=review
crr+=cr
rnn+=rn
rrr+= rr
rll+=rl
time+=Time
print(len(cll), len(cnn) , len(cfl) , len(cre) , len(crr) , len(rnn) , len(rrr) , len(rll) , len(time))
browser.quit()
# -
len(cll)
import pandas as pd
out = pd.DataFrame()
out['RESTAURANT_NAME'] = rnn
out['RESTAURANT_RATING'] = rrr
out['CUSTOMER_NAME']= cnn
out['CUSTOMER_RATING'] = crr
out['REVIEWS_BY_USER'] = cre
out['FOLLOWERS_OF_USER'] =cfl
out['DELIVERY_TIME'] = time
out['USER_PROFILE_LINK'] = cll
out['RESTAURANT_LINK'] = rll
print(out.shape)
out.head()
out = out[out['RESTAURANT_RATING']!='NEW']
print(out.shape)
out['RESTAURANT_RATING'] = out['RESTAURANT_RATING'].astype('float')
out['CUSTOMER_RATING'] = out['CUSTOMER_RATING'].astype('float')
out.info()
out.to_csv('output.csv' , index = False)
# # Modification
import pandas as pd
data = pd.read_csv('output.csv')
print(data.shape)
data.head()
data['DELIVERY_TIME'] = data['DELIVERY_TIME'].apply(lambda x: x.split()[0])
data['DATE'] = data['DELIVERY_TIME'].apply(lambda x: int(x.split('-')[2]))
data['MONTH'] = data['DELIVERY_TIME'].apply(lambda x: int(x.split('-')[1]))
data['YEAR'] = data['DELIVERY_TIME'].apply(lambda x:int( x.split('-')[0]))
print(data.shape)
data.head()
data['YEAR'].value_counts()
data['MONTH'].value_counts()
# # Now i'm gonna write some SQL queries to get data from above table ....here
# # Write a SQL query to find the number of Zomato users
#
SELECT COUNT(*)
FROM table;
# # Write a SQL query to find the list of Zomato users who made more than 10 orders in a particular month
#first of all perform grouping on users by GROUPBY
SELECT *
FROM table
WHERE MONTH=10
GROUP BY CUSTOMER_NAME;
# +
#IT WILL SHOW ALLL THE USERS WHO HAVE ORDERS IN OCTOBER MONTH
#AFTER THAT WE WILL CALCULATE THE TOTAL NUMBER OF USERS WHO ARE MORE THAN 10 TIMES
# -
# # Write a SQL query to find the list of Zomato users who order food from the same restaurants more than 3 times in a week
# let's take November (9) month and take date 1-7
SELECT *
FROM table
WHERE MONTH=9 AND (DATE=>1 AND DATE<=7)
GROUP BY CUSTOMER_NAME;
# HAVING COUNT()
# +
#AGAIN WE UNIQUE USERS HERE IN A WEEK ,WE WILL COUNT TO CHECK WITHER IT IS HAVING 3 ORDERS IN A WEEK OR NOT
# -
# # ///@<NAME>
# # +++++++++++++++++++++++++++++++++++++++++
| Zomato_scraping_for_mumbai/zomato_scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="YQGhuaNsRLO1"
# ##LunarLanderContinuous-v2 by OpenAI Gym
# **Importing Modules**
# + id="Q0OLWrL_RQ6y"
# !pip3 install box2d-py
import gym
import pickle
import tensorflow as tf
import tensorflow.compat.v1 as tf1
tf1.disable_v2_behavior()
from tensorflow import keras
from keras import layers, initializers, regularizers
import numpy as np
import threading
from functools import reduce
import time
import os
from collections import deque
import matplotlib.pyplot as plt
# + [markdown] id="e1LXGgThRWRy"
# **Initial prep work**
# + id="mXiDeGCiRf-J"
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
checkpoint_path = "./models_checkpoints"
try:
os.mkdir(checkpoint_path)
except FileExistsError:
pass
class StateTrasitionRecorder:
def __init__(self):
self.recorder_memory = deque()
def save_state_transition(self, transition):
self.recorder_memory.append(transition)
def flush_recorder_memory(self):
self.recorder_memory = deque()
class RolloutBuffer(StateTrasitionRecorder):
def __init__(self, policy_net_args):
super().__init__()
self.rollout_memory = deque()
self.gamma = policy_net_args["gamma"]
def save_rollout(self, episode):
complete_episode = self.compute_total_rewards(episode, self.gamma)
self.rollout_memory.append(complete_episode)
self.flush_recorder_memory()
def compute_total_rewards(self, episode_transitions, gamma):
states, actions, rewards, nex_states, dones = zip(*episode_transitions)
Q_s_a = []
for i in range(len(rewards)):
Q_i = 0
for j in range(i, len(rewards)):
Q_i += rewards[j] * self.gamma ** (j - i)
Q_s_a.append(Q_i)
episode = deque(zip(states, actions, rewards,
nex_states, dones, Q_s_a))
return(episode)
def unroll_state_transitions(self):
states = ()
actions = ()
next_states = ()
rewards = ()
dones = ()
Q_sa = ()
for episode in self.rollout_memory:
ep_states, ep_actions, ep_next_states, ep_rewards, ep_dones, ep_Q_s_a = zip(
*episode)
states += ep_states
actions += ep_actions
next_states += ep_next_states
rewards += ep_rewards
dones += ep_dones
Q_sa += ep_Q_s_a
states = np.asarray(states)
actions = np.asarray(actions)
next_states = np.asarray(next_states)
rewards = np.asarray(rewards)
dones = np.asarray(dones, dtype=int)
Q_sa = np.asarray(Q_sa).reshape(-1, 1)
return states, actions, next_states, rewards, dones, Q_sa
def flush_rollout_memory(self):
self.rollout_memory = deque()
def build_networks(network_name, num_Hlayers, activations_Hlayers, Hlayer_sizes, n_output_units, output_layer_activation, regularization_constant, network_type, input_features,):
assert(num_Hlayers == (len(activations_Hlayers)) and num_Hlayers ==
len(Hlayer_sizes))
with tf1.variable_scope(network_type):
network = tf1.layers.Dense(Hlayer_sizes[0], activation=activations_Hlayers[0], kernel_initializer=tf.initializers.glorot_normal(),
kernel_regularizer=tf.keras.regularizers.L2(l2=regularization_constant), name="Layer_1")(input_features)
for layer in range(1, num_Hlayers):
network = tf1.layers.Dense(units=Hlayer_sizes[layer], kernel_initializer=tf.initializers.glorot_normal(), kernel_regularizer=tf.keras.regularizers.L2(l2=regularization_constant), activation=activations_Hlayers[layer], name=(
"Layer_" + str(layer + 1)))(network)
if network_type == "Actor":
mu = tf1.layers.Dense(units=n_output_units, kernel_initializer=tf.initializers.glorot_normal(),
kernel_regularizer=tf.keras.regularizers.L2(l2=regularization_constant), activation=output_layer_activation, name="mu")(network)
covariance = tf1.layers.Dense(
units=n_output_units, kernel_initializer=tf.initializers.glorot_normal(), kernel_regularizer=tf.keras.regularizers.L2(l2=regularization_constant), activation=tf.nn.softplus, name="covariance")(network)
params = tf1.get_collection(tf1.GraphKeys.TRAINABLE_VARIABLES, network_name + "/" + network_type)
return mu, covariance, params
else:
critic = tf1.layers.Dense(units=n_output_units, kernel_initializer=tf.initializers.glorot_normal(
), activation=output_layer_activation, kernel_regularizer=tf.keras.regularizers.L2(l2=regularization_constant), name="V")(network)
params = tf1.get_collection(tf1.GraphKeys.TRAINABLE_VARIABLES, network_name + "/" + network_type)
return critic, params
class ComputationGraph:
def __init__(self, name, policy_network_args, value_function_network_args):
super().__init__(policy_network_args)
with tf1.variable_scope(name):
self.actor_optimizer = policy_network_args["optimizer"]
self.critic_optimizer = value_function_network_args["optimizer"]
self.st_placeholder = tf1.placeholder(dtype=tf.float32, shape=[
None, policy_network_args["state_space_size"]], name="State")
self.rewards_placeholder = tf1.placeholder(
tf.float32, shape=[None, 1], name="rewards")
self.actions_placeholder = tf1.placeholder(
tf.float32, shape=[None, policy_network_args["action_space_size"]], name="actions")
self.dones_placeholder = tf1.placeholder(
tf.float32, shape=[None, 1], name="dones")
self.Qsa_placeholder = tf1.placeholder(
dtype=tf.float32, shape=[None, 1], name="Q_sa")
self.mu, self.covariance, self.actor_params = build_networks(name, policy_network_args["num_Hlayers"], policy_network_args["activations_Hlayers"], policy_network_args[
"Hlayer_sizes"], policy_network_args["n_output_units"], policy_network_args["output_layer_activation"], policy_network_args["regularization_constant"], "Actor", self.st_placeholder)
self.critic, self.critic_params = build_networks(name, value_function_network_args["num_Hlayers"], value_function_network_args["activations_Hlayers"], value_function_network_args[
"Hlayer_sizes"], value_function_network_args["n_output_units"], value_function_network_args["output_layer_activation"], policy_network_args["regularization_constant"], "Critic", self.st_placeholder)
with tf1.variable_scope("Train_value_function_estimator"):
self.value_function_net_cost = tf.losses.mean_squared_error(
self.Qsa_placeholder, self.critic) + tf1.losses.get_regularization_loss(scope=name + "/" + "Critic")
tf1.summary.scalar("Critic_Cost", self.value_function_net_cost)
if name == "Global_Agent":
for variable in self.actor_params:
var_name = "Actor_" + variable.name.replace("kernel:0", "w").replace("bias:0", "b")
tf.summary.histogram(var_name, variable)
for variable in self.critic_params:
var_name = "Critic_" + variable.name.replace("kernel:0", "w").replace("bias:0", "b")
tf.summary.histogram(var_name, variable)
with tf1.variable_scope("Train_policy_network"):
self.advantage_funtion = tf.math.subtract(
self.Qsa_placeholder, self.critic)
self.probability_density_func = tf1.distributions.Normal(
self.mu, self.covariance)
self.log_prob_a = self.probability_density_func.log_prob(
self.actions_placeholder)
auxiliary = tf.multiply(
self.log_prob_a, self.advantage_funtion)
entropy = self.probability_density_func.entropy()
self.auxiliary = policy_network_args["Entropy"] * \
entropy + auxiliary
self.policy_net_cost = tf.reduce_sum(-self.auxiliary) + tf1.losses.get_regularization_loss(scope=name + "/" + "Actor")
self.summary_policy_cost = tf.summary.scalar("Policy_Cost", self.policy_net_cost)
with tf.name_scope("choose_a"):
self.action = tf1.clip_by_value(self.probability_density_func.sample(
1), policy_network_args["action_space_lower_bound"], policy_network_args["action_space_upper_bound"])
with tf.name_scope("get_grad"):
self.actor_grads = tf1.gradients(self.policy_net_cost, self.actor_params)
self.critic_grads = tf1.gradients(self.value_function_net_cost, self.critic_params)
for act_grad, critic_grad in zip(self.actor_grads, self.critic_grads):
var_name_actor = "Actor_" + act_grad.name.replace("Addn", "w")
var_name_critic = "Critic_" + critic_grad.name.replace("Addn", "w")
tf.summary.histogram(var_name_actor, act_grad)
tf.summary.histogram(var_name_critic, critic_grad)
self.summaries = tf1.summary.merge_all()
# + [markdown] id="oLy8Xy-bRxvb"
# **Training the Model**
# + id="fKTk5Zt7R63o"
class RLAgent(ComputationGraph, RolloutBuffer):
def __init__(self, name, policy_network_args, value_function_network_args, session, summary_writer, Global_Agent=None):
super().__init__(name, policy_network_args, value_function_network_args)
self.current_num_epi = 0
self.env = gym.make('LunarLanderContinuous-v2')
self.total_number_episodes = policy_network_args["total_number_episodes"]
self.num_episodes_before_update = policy_network_args["number_of_episodes_before_update"]
self.Global_Agent = Global_Agent
self.ep_rewards = []
self.frequency_printing_statistics = policy_network_args["frequency_of_printing_statistics"]
self.episodes_back = policy_network_args["episodes_back"]
self.rendering_frequency = policy_network_args["frequency_of_rendering_episode"]
self.max_steps = policy_network_args["max_steps_per_episode"]
self.summary_writer = summary_writer
self.name = name
self.session = session
if Global_Agent is not None:
with tf.name_scope(name):
with tf.name_scope('sync'):
with tf.name_scope('pull_from_global'):
self.pull_actor_params_op = [local_params.assign(
global_params) for local_params, global_params in zip(self.actor_params, Global_Agent.actor_params)]
self.pull_critic_params_op = [local_params.assign(
global_params) for local_params, global_params in zip(self.critic_params, Global_Agent.critic_params)]
with tf.name_scope("push_to_global"):
self.push_actor_params_op = self.actor_optimizer.apply_gradients(zip(self.actor_grads, self.Global_Agent.actor_params))
self.push_critic_params_op = self.critic_optimizer.apply_gradients(zip(self.critic_grads, Global_Agent.critic_params))
def update_Global_Agent(self, feed_dict):
_, _, = self.session.run([self.push_actor_params_op,
self.push_critic_params_op], feed_dict)
def save_summary(self, feed_dict):
summary = self.session.run(self.Global_Agent.summaries, feed_dict)
self.summary_writer.add_summary(summary, self.Global_Agent.current_num_epi)
def pull_from_global(self):
self.session.run([self.pull_actor_params_op,
self.pull_critic_params_op])
def take_action(self, state):
state = state.reshape(-1, 8)
action = self.session.run([self.action], feed_dict={
self.st_placeholder: state})
return action[0].reshape(2,)
def collect_rollouts(self, n_rolls, max_steps, render=False):
for i in range(n_rolls):
n_steps = 0
state = self.env.reset()
done = False
sum_rewards = 0
while not done and n_steps <= max_steps:
if render:
self.env.render()
action = self.take_action(state)
next_state, reward, done, info = self.env.step(action)
if not done and n_steps == max_steps:
state_feed = next_state.reshape(-1, 8)
reward = reward + float(self.session.run([self.critic], feed_dict={self.st_placeholder: state_feed})[0])
self.save_state_transition(
[state, action, reward, next_state, done])
sum_rewards += reward
state = next_state
n_steps += 1
if self.name == "Global_Agent":
print(f"Episode Reward: {sum_rewards}")
self.ep_rewards.append(sum_rewards)
self.save_rollout(self.recorder_memory)
def training_loop(self):
"""Runs episodes in a loop and performs steps of gradient descent after every episode"""
while not coord.should_stop() and self.Global_Agent.current_num_epi <= self.total_number_episodes:
self.collect_rollouts(
self.num_episodes_before_update, self.max_steps, render=False)
states, actions, next_states, rewards, dones, Q_sa = self.unroll_state_transitions()
feed_dict = {self.st_placeholder: states,
self.actions_placeholder: actions,
self.Qsa_placeholder: Q_sa}
self.update_Global_Agent(feed_dict)
self.Global_Agent.current_num_epi += self.num_episodes_before_update
feed_dict_global_summary = {self.Global_Agent.st_placeholder: states,
self.Global_Agent.actions_placeholder: actions,
self.Global_Agent.Qsa_placeholder: Q_sa}
self.save_summary(feed_dict_global_summary)
self.flush_rollout_memory()
self.pull_from_global()
if self.Global_Agent.current_num_epi % self.frequency_printing_statistics == 0:
average_reward = self.Global_Agent.compute_average_rewards(self.episodes_back)
print(
f"Global ep number {self.Global_Agent.current_num_epi}: Reward = {average_reward}")
class Global_Agent(RLAgent):
def __init__(self, name, policy_network_args, value_function_network_args, session, summary_writer, child_agents=[]):
super().__init__(name, policy_network_args, value_function_network_args, session, summary_writer)
self.child_agents = child_agents
self.num_childs = len(child_agents)
def compute_average_rewards(self, episodes_back):
"""Computes the average reward of each child agent going n episodes back, and returnes the average of those average rewards"""
reward = 0
for agent in self.child_agents:
agent_average_reward = reduce(
lambda x, y: x + y, agent.ep_rewards[-episodes_back:]) / episodes_back
reward += agent_average_reward
reward /= self.num_childs
return reward
# + [markdown] id="5j_ph4KMS6xc"
# **Testing the model**
# + id="AWN9xhEVS9Q6"
if __name__ == "__main__":
env = gym.make('LunarLanderContinuous-v2')
action_space_upper_bound = env.action_space.high
action_space_lower_bound = env.action_space.low
subdir = time.strftime("%Y%m%d-%H%M%S", time.localtime())
logdir = "./summary/" + subdir
writer = tf.summary.create_file_writer(logdir)
sess = tf1.Session()
policy_net_args = {"num_Hlayers": 2,
"activations_Hlayers": ["relu", "relu"],
"Hlayer_sizes": [100, 100],
"n_output_units": 2,
"output_layer_activation": tf.nn.tanh,
"state_space_size": 8,
"action_space_size": 2,
"Entropy": 0.01,
"action_space_upper_bound": action_space_upper_bound,
"action_space_lower_bound": action_space_lower_bound,
"optimizer": tf1.train.RMSPropOptimizer(0.0001),
"total_number_episodes": 5000,
"number_of_episodes_before_update": 1,
"frequency_of_printing_statistics": 100,
"frequency_of_rendering_episode": 1000,
"number_child_agents": 8,
"episodes_back": 20,
"gamma": 0.99,
"regularization_constant": 0.01,
"max_steps_per_episode": 2000
}
valuefunction_net_args = {"num_Hlayers": 2,
"activations_Hlayers": ["relu", "relu"],
"Hlayer_sizes": [100, 64],
"n_output_units": 1,
"output_layer_activation": "linear",
"state_space_size": 8,
"action_space_size": 2,
"optimizer": tf1.train.RMSPropOptimizer(0.01),
"regularization_constant": 0.01}
global_agent = Global_Agent("Global_Agent", policy_net_args, valuefunction_net_args, sess, writer)
child_agents = []
for i in range(policy_net_args["number_child_agents"]):
i_name = f"ChildAgent_{i}"
child_agents.append(RLAgent(i_name, policy_net_args, valuefunction_net_args, sess, writer, global_agent))
global_agent.child_agents = child_agents
global_agent.num_childs = len(child_agents)
saver = tf1.train.Saver()
coord = tf.train.Coordinator()
if len(os.listdir(checkpoint_path)) == 0:
sess.run(tf1.global_variables_initializer())
else:
saver.restore(sess, checkpoint_path + "/variables.ckpt")
child_agents_threads = []
subdir = time.strftime("%Y%m%d-%H%M%S", time.localtime())
logdir = "./summary/" + subdir
writer = tf1.summary.FileWriter(logdir)
writer.add_graph(sess.graph)
for child_agent in child_agents:
def job(): return child_agent.training_loop()
t = threading.Thread(target=job)
t.start()
child_agents_threads.append(t)
coord.join(child_agents_threads)
saver.save(sess, checkpoint_path + "/variables.ckpt")
for i in range(1):
global_agent.collect_rollouts(10, 2000, render=True)
global_agent.collect_rollouts(90, 2000)
rewards = global_agent.ep_rewards
average = sum(rewards)/len(rewards)
average = [average] * 100
fig, ax = plt.subplots()
ax.plot(rewards, label="Episode Reward")
ax.plot(average, label="Average")
ax.legend(loc="best")
plt.show()
| Lunar-Lander-Continuos-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="XIelfH4uoLOQ"
# # Working with Animations in Coldtype
# + id="vQqKKBu6fSZh" executionInfo={"status": "ok", "timestamp": 1632773263364, "user_tz": 420, "elapsed": 17083, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}}
# #!pip install coldtype[notebook]
# !pip install -q "coldtype[notebook] @ git+https://github.com/goodhertz/coldtype"
from coldtype.notebook import *
# + [markdown] id="CwcUbsLlm-dr"
# ## A circle moving
# + colab={"base_uri": "https://localhost:8080/", "height": 291} id="A8_pGQ6vfe7j" executionInfo={"status": "ok", "timestamp": 1632773266471, "user_tz": 420, "elapsed": 155, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="90d0fcd7-f7f8-4c5d-b6f0-6703423d0f21"
@animation((540, 540), timeline=60, bg=1)
def circle(f):
return PS([
P(f.a.r).f(1),
(P().oval(f.a.r.inset(120)
.offset(f.e("eeio", 1, rng=(-f.a.r.w/2, f.a.r.w/2)), 0))
.f(hsl(0.75)))])
# + [markdown] id="jvmWCCXPnOgZ"
# Here's a line of code to render and compile the entire animation as an h264.
#
# (If you're using the local viewer app, this is the equivalent of hitting the `a` key, adding a line like `release = circle.export("h264")`, and then hitting the `r` key to trigger that release action.)
# + colab={"base_uri": "https://localhost:8080/", "height": 291, "referenced_widgets": ["250a266549d54878acaeef5ff3de8aa3", "7ffca08a3dc04f849845892b22aeeca0", "90f8775d27354f6caa379d82e8e7fc4b", "cb1311f995c64122bdfb5eb1947571d1", "fef5545a8ff14645add1323553aa64bb", "b3e4c3bed9e34ce2a7d756525526ee9e", "c5e8d4a763ff4effb469a372b06a6a0b", "a54eac3fb47c4b00b3aeba6c3902d889", "ba65a43dea984b5ea5e819ad5eab807f", "c8b819ac83ca4cad886ffb550f88d7a9", "ed5a1cd0c27e4878acb5be76679f4867"]} id="lPp0_ZK_iAvQ" executionInfo={"status": "ok", "timestamp": 1632773277690, "user_tz": 420, "elapsed": 3373, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="224f6f14-b0ef-4d45-ca81-0ad93cbe6b10"
circle.render().show()
# + [markdown] id="ckjOCzI3np1A"
# ## A letter flying
# + colab={"base_uri": "https://localhost:8080/", "height": 323, "referenced_widgets": ["329a9a6e231b45cd92ec049096eb7ef6", "fc8c4c749e38405bbf72b8cfddb3e2ed", "00a2133e7f3f43639f0f6192da6c6c3f", "0322ecd81ee741d5aa53a04af7e4bc4e", "<KEY>", "8ce2c8a222be45a1a0a7826bd8f22c2d", "99f4b6e457b24d7984b4e36ec453fe14"]} id="CH7Y341Ffr_M" executionInfo={"status": "ok", "timestamp": 1632762351590, "user_tz": 420, "elapsed": 155, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="db5c97ba-712c-44c6-fbb8-e7ad322cb913"
@animation((540, 540), bg=0, timeline=24)
def flying(f):
return PS([
(P().rect(f.a.r)
.f(hsl(f.e("qeio")))),
(StSt("A", Font.MutatorSans(),
50, wght=0.2)
.align(f.a.r)
.scale(f.e("eei", rng=(1, 91)))
.rotate(f.e("qeio", rng=(0, 360)))
.f(1))])
# + colab={"base_uri": "https://localhost:8080/", "height": 291, "referenced_widgets": ["0d08e0465d124739a176ffb9f5a86044", "4b68ce5f47124027ac51fcd0242a174b", "fa0bf8519f3a419aacfe7b267a673a6b", "c939edc5f7af48e689656d4913d4e3df", "ac6ab1ebe5934c6a826b0154f3436cb2", "<KEY>", "1b8fec3ffc51407aae48fa39ff4e3c7a", "643f224452c24a94a6e729421be0523b", "c4a1f8afb2e644af861edc4c4a347a31", "bf457392fd5c48369f80edbf9d52a511", "cb91d86b175e42e182e8dd51d38c54cb"]} id="MzHTn4-_mdMu" executionInfo={"status": "ok", "timestamp": 1632762356138, "user_tz": 420, "elapsed": 1492, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="b6d5b27f-8cdb-4b31-f189-b7f3bac58e47"
flying.render().show()
# + [markdown] id="yl-jQtUqqiGU"
# ## Simple variation
# + colab={"base_uri": "https://localhost:8080/", "height": 323, "referenced_widgets": ["b77161fd1cda499199c84c330ab90f8d", "6badf339543d4dd09e848ad69118117e", "f908cfac39f84cd0b541410de36adabd", "27c6a70b32b2421ead387fa3187a3435", "<KEY>", "9eeec7da73954271ac23f41efe91859e", "7d69bdf998514afea835976cc71986b6"]} id="KUH3FPI0qHbN" executionInfo={"status": "ok", "timestamp": 1632762361448, "user_tz": 420, "elapsed": 234, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="b67fff5d-8a27-4982-eecf-211be5b7a743"
@animation((1080, 540), timeline=50, bg=hsl(0.4))
def vari(f):
return (StSt("COLDTYPE",
Font.ColdtypeObviously(), 200,
wdth=f.e("eeio", 1))
.align(f.a.r)
.f(1))
# + colab={"base_uri": "https://localhost:8080/", "height": 291, "referenced_widgets": ["f58dc4524959479aa9e799326f4803a8", "80d5b1f4ba544570813d0c4dd82bd7b2", "9225ebe5ca4742efbb169fedecf69474", "2398aed8840842d5ab0bb0be6206e791", "<KEY>", "02d4bb8bb974471ea12ad7668e66afe0", "251842ec57a448179ea51379544a0913", "3f0687a6f8264127bcef94a7a3d8f3eb", "4d2ed4a806964063b29c774b102ebd66", "12b22a76950a45bd99a687e2ea709054", "f6caa35df9ca4eb89378d3e5e4ad48c3"]} id="5XHCUK5oqT73" executionInfo={"status": "ok", "timestamp": 1632762381977, "user_tz": 420, "elapsed": 4803, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="5ea83205-167b-4132-996f-a6b31e6391b6"
vari.render().show(loops=2)
# + [markdown] id="QvTOQcXTp8Dh"
# ## A variable wave
# + colab={"base_uri": "https://localhost:8080/", "height": 323, "referenced_widgets": ["af27b28020ae4a37aa9f6bcfdc85f957", "<KEY>", "fd719fb1e0384b379b9c3cc4f799f5b3", "864c1dbea83946ddb5e4e1aaf97fd009", "495597551d1646fd954f78d860232fee", "49024b8ab4b047e7a864e202f6a2042e", "008f25a7062a4b4ebe9d6b260a33de81"]} id="4E1F649RoK4C" executionInfo={"status": "ok", "timestamp": 1632762385693, "user_tz": 420, "elapsed": 179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="98c7b11c-c221-452f-dc67-2f8247daf5e3"
@animation((1080, 540), timeline=50, bg=hsl(0.6))
def wave(f):
return (Glyphwise("COLDTYPE", lambda g:
Style(Font.ColdtypeObviously(), 200,
wdth=f.adj(-g.i*3).e("seio", 1)))
.align(f.a.r)
.f(1))
# + colab={"base_uri": "https://localhost:8080/", "height": 291, "referenced_widgets": ["ef064f3c3fc84c64989be832dc740514", "d41ebe4c76a64f1390db765dfd612066", "4edde883b9ed4b44b130fe6dd1ef88ed", "88aa233ebca94cfe95f8e8617fa0f29d", "28c671df67464418a8eae170d40a9778", "04724f3bf2be4758a3e7441f9cfadee2", "51c5e3aca04d4daeba29fb62207fbef7", "da15add7039a4e50875d3ae77c0a6a9b", "<KEY>", "1f7c6ce2d16644d9b4e7a1f03a27d8ff", "de2bd46b173b4d1e87b0039e067cad7e"]} id="CyAQ8StPoUm2" executionInfo={"status": "ok", "timestamp": 1632762404086, "user_tz": 420, "elapsed": 5574, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg9X2Vci7VOjUJPYvBOav8xi6FVNGs-VYJlhcUP=s64", "userId": "14730280017638687953"}} outputId="48846f91-dad3-489d-ddea-0a7ae31a5ff3"
wave.render().show(loops=2)
| docs/notebooks/tutorials/animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ### R Squared (R^2)学习
#
# #### 1.MSE,RMSE,MAE的问题
#
# 这三种衡量指标的缺点就是:对于不同的量纲,不容易评判模型的好换
#
# 为了解决上述问题,引入了R方来评测线性回归法的好坏,也是sklearn用来评测线性回归模型好坏的指标
#
#
# #### 2.R方
#
# R方公式:
# [公式](img/R方公式.png)
#
# 分子含义:使用我们的模型预测产生的错误
#
# 父母含义:使用y=y_mean模型预测产生的错误, y=y_mean模型可以理解为最基础的模型(Baseline Model)
#
# var(y): 方差,variance
#
# 根据公式,可以知道以下几点:
# - 衡量指标在0-1之间,容易判断模型好坏,越接近于1,模型越好
# - 值等于0,即分子等于分母,表示我们的模型和最基础模型一样,预测情况属于最坏情况
# - 值等于1,即分子等于0,表示我们的模型预测结果最好
# - 值小于0,即分子大于分母,表示我们的模型预测错误比最基础模型预测错误还多,说明我们的模型很差,可以认为数据不存在任何线性关系
# + pycharm={"name": "#%%\n", "is_executing": false}
import numpy as np
from sklearn import datasets
from sklearn.model_selection import train_test_split
# 构造数据
boston = datasets.load_boston()
x = boston.data[:,5]
y = boston.target
x = x[y < 50]
y = y[y < 50]
x_train, x_test, y_train, y_test = train_test_split(x, y)
# + pycharm={"name": "#%%\n", "is_executing": false}
from SimpleLinearRegression import SimpleLinearRegression
from accuracy_score.metrics import mean_squared_error
# 训练模型
estimator = SimpleLinearRegression()
estimator.fit(x_train, y_train)
print('estimator.a_ = ',estimator.a_)
print('estimator.b_ = ',estimator.b_)
# 预测
y_predict = estimator.predict(x_test)
# 计算MSE
mse = mean_squared_error(y_test, y_predict)
# 计算方差
var = np.var(y_test)
# r2
r2 = 1 - mse/var
r2
# + [markdown] pycharm={"name": "#%% md\n"}
# ### 封装我们自己的r2_score
# 代码[这里](accuracy_score/metrics.py)
# + pycharm={"name": "#%%\n", "is_executing": false}
from accuracy_score.metrics import r2_score
result = r2_score(y_test, y_predict)
result
# + [markdown] pycharm={"name": "#%% md\n"}
# ### 使用sklean的r2_score
# + pycharm={"name": "#%%\n", "is_executing": false}
from sklearn.metrics import r2_score
result = r2_score(y_test, y_predict)
result
# + [markdown] pycharm={"name": "#%% md\n"}
# 我们自己封装的R2计算的结果和sklearn封装的R2计算结果基本一致
#
# 0.5都不到,绝大多数是数据的问题,毕竟我们只使用了波士顿房价数据的某一个特征(RM)来测试
| ml/04-Linear-Regression/05-R-Squared/05-R-Squared.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Multiple Linear Regression Introduction
#
# In this notebook (and following quizzes), you will be creating a few simple linear regression models, as well as a multiple linear regression model, to predict home value.
#
# Let's get started by importing the necessary libraries and reading in the data you will be using.
# +
import numpy as np
import pandas as pd
import statsmodels.api as sm;
df = pd.read_csv('./house_prices.csv')
df.head()
# -
# `1.` Using statsmodels, fit three individual simple linear regression models to predict price. You should have a model that uses **area**, another using **bedrooms**, and a final one using **bathrooms**. You will also want to use an intercept in each of your three models.
#
# Use the results from each of your models to answer the first two quiz questions below.
df['intercept'] = 1
lm = sm.OLS(df['price'], df[['intercept','area']])
results = lm.fit()
results.summary()
df['intercept'] = 1
lm = sm.OLS(df['price'], df[['intercept','bathrooms']])
results = lm.fit()
results.summary()
df['intercept'] = 1
lm = sm.OLS(df['price'], df[['intercept','bedrooms']])
results = lm.fit()
results.summary()
# `2.` Now that you have looked at the results from the simple linear regression models, let's try a multiple linear regression model using all three of these variables at the same time. You will still want an intercept in this model.
mlr = sm.OLS(df['price'], df[['intercept','area','bathrooms','bedrooms']])
results_mlr = mlr.fit()
results_mlr.summary()
X = df[['intercept','bathrooms','bedrooms','area']]
Y = df['price']
np.dot(np.dot(np.linalg.inv(np.dot(X.transpose(),X)),X.transpose()),Y)
# `3.` Along with using the **area**, **bedrooms**, and **bathrooms** you might also want to use **style** to predict the price. Try adding this to your multiple linear regression model. What happens? Use the final quiz below to provide your answer.
mlr2 = sm.OLS(df['price'], df[['intercept', 'area', 'bedrooms', 'bathrooms', 'style']])
results_mlr2 = mlr2.fit()
results_mlr2.summary()
| House Price/Multiple Linear Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata_df = pd.read_csv(mouse_metadata_path)
study_results_df = pd.read_csv(study_results_path)
# Combine the data into a single dataset
merged_df = pd.merge(mouse_metadata_df,study_results_df, on="Mouse ID", how = "left")
# Display the data table for preview
#merged_df.head()
#mouse_metadata_df.count()
#study_results_df.count()
#merged_df.count()
#test = mouse_metadata_df["Mouse ID"].nunique(dropna=True)
#test
# -
# Check the number of mice.
total_mice = merged_df["Mouse ID"].nunique(dropna=True)
total_mice
# Find any duplicate rows with the same Mouse ID's and Timepoints.
find_duplicates = merged_df.duplicated(subset=["Mouse ID", "Timepoint"],keep = False)
find_duplicates
#cleaned_df = merged_df.drop_duplicates(subset=["Mouse ID", "Timepoint"])
# Drop any duplicate rows
merged_cleaned_df = merged_df.drop_duplicates(subset=["Mouse ID", "Timepoint"])
merged_cleaned_df.head()
# Recheck the number of mice
total_mice = merged_cleaned_df["Mouse ID"].nunique(dropna=True)
total_mice
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance,
# standard deviation, and SEM of the tumor volume for each regimen.
#summary_statistics_test = merged_cleaned_df.groupby('Drug Regimen').describe()
#summary_statistics_test
summary_statistics = merged_cleaned_df.groupby('Drug Regimen').agg({'Tumor Volume (mm3)':['mean',
'median',
'var',
'std',
'sem'
]})
summary_statistics
# -
# ## Bar and Pie Charts
# +
# Generate a bar plot showing the total number of datapoints for each drug regimen using pandas.
# There should be a single bar per regimen
bar_plot = merged_cleaned_df.groupby(["Drug Regimen"]).count()["Mouse ID"]
#bar_plot
bar_plot.plot(kind="bar", color="pink", stacked=True, figsize=(10,5))
plt.title("Drug Treatment Regimen",)
plt.ylabel("Number of Mice Treated by Drug Regimen")
plt.xlabel("Drug Regimen Used")
plt.show()
# -
# Generate identical bar plot using pyplot instead of pandas.
# Getting datapoints count by examining previus datatapoint variable
bar_plot
data_points = [230, 178, 178, 188, 186, 181, 161, 228, 181, 182]
drug_names = ["Capomulin", "Ceftamin", "Infubinol", "Ketapril", "Naftisol","Placebo", "Propriva", "Ramicane", "Stelasyn", "Zoniferol"]
plt.figure(figsize=(10,5))
plt.bar(drug_names,data_points, color="pink", width=0.5)
plt.xticks(rotation='vertical')
plt.title("Drug Treatment Regimen",)
plt.ylabel("Number of Mice Treated by Drug Regimen")
plt.xlabel("Drug Regimen Used")
plt.show()
# Generate a pie plot showing the distribution of female versus male mice using pandas
gender_count = merged_cleaned_df.groupby('Sex')['Mouse ID'].count()
gender_count.head()
colors = ['pink', 'skyblue']
explode = (0.1,0)
gender_count.plot(kind="pie", autopct='%1.1f%%', colors=colors, shadow=True, explode=explode, startangle=90)
plt.title("Male vs Female Mice Distribution")
plt.axis("off")
plt.show()
# +
# Generate identical pie plot using pyplot
labels = ["Female", "Male"]
#sizes = [15, 30, 45, 10]
explode = (0.1, 0)
colors = ['pink', 'skyblue']
plt.pie(gender_count, labels=labels, autopct='%1.1f%%', colors=colors, shadow=True, explode=explode, startangle=90)
plt.title("Male vs Female Mice Distribution")
plt.show()
# -
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
# HINT: Not all mice lived until timepoint 45
# Start by getting the last (greatest) timepoint for each mouse
each_mouse = merged_cleaned_df.groupby("Mouse ID").max()
max_timepoint = each_mouse.reset_index()
#max_timepoint
max_timepoint_df = max_timepoint[['Mouse ID','Timepoint']]
max_timepoint_df.head()
# -
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
tumor_volume_df = max_timepoint_df.merge(merged_cleaned_df, on = ['Mouse ID', 'Timepoint'], how="left")
tumor_volume_df.head()
# Calculate the quartiles and IQR and quantitatively determine if there are any potential outliers across all four treatment regimens.
# +
# Calculate quartiles, IQR, and identify potential outliers for each regimen.
##############################################################################
# Put treatments into a list for for loop (and later for plot labels)
treatment_drugs = ['Capomulin', 'Ramicane', 'Infubinol', 'Ceftamin']
# Create empty list to fill with tumor vol data (for plotting)
tumor_vol_data = []
# Calculate the IQR and quantitatively determine if there are any potential outliers.
#######################################################################################
# Locate the rows which contain mice on each drug and get the tumor volumes
# this was done inside the loop. Below is code commented out to check before the loop.
# mouse_by_drug_group_df = tumor_volume_df.groupby("Drug Regimen")
# mouse_by_drug_group_df
# -
# Determine outliers using upper and lower bounds
for i in treatment_drugs:
volume_drug = tumor_volume_df.loc[tumor_volume_df["Drug Regimen"] == i]["Tumor Volume (mm3)"]
quartiles = volume_drug.quantile([.25,.5,.75])
lower_q = round(quartiles[0.25],2)
upper_q = round(quartiles[0.75],2)
iqr = upper_q-lower_q
lower_bound = round(lower_q - (1.5*iqr),2)
upper_bound = round(upper_q + (1.5*iqr),2)
tumor_vol_data.append(volume_drug)
print(f"-----------------------")
print(f" The lower quartile of {i} is : {lower_q}")
print(f" The upper quartile of {i}: {upper_q}")
print(f" The interquartile range of {i}: {iqr}")
print(f" These values below {lower_bound} could be outliers.")
print(f" These values above {upper_bound} could be outliers.")
print(f"-----------------------")
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
# There should be a single chart with four box plots inside it.
# using "tumor_vol_data" referenced above for x-axis tickers
marker = dict(markerfacecolor='g', marker='D')
fig1, ax1 = plt.subplots()
ax1.set_title('Tumor Volume for Promising Drugs')
ax1.set_ylabel('Tumor Volume (mm3)')
bp = ax1.boxplot(tumor_vol_data,flierprops=marker)
ax1.set_xticklabels(treatment_drugs)
plt.savefig('../Images/drug_treatment_quartiles_charts.png')
plt.show()
# -
# ## Line and Scatter Plots
# Generate a line plot of tumor volume vs. time point for a single mouse
# treated with Capomulin
mouse_by_capomulin = merged_cleaned_df.loc[merged_cleaned_df["Drug Regimen"] == "Capomulin"]
mouse_by_capomulin
single_mouse_id = "s185"
single_mouse_capomulin = mouse_by_capomulin.loc[mouse_by_capomulin["Mouse ID"] == single_mouse_id]
single_mouse_capomulin
# +
x_axis = single_mouse_capomulin["Timepoint"]
y_axis = single_mouse_capomulin["Tumor Volume (mm3)"]
plt.plot(x_axis,y_axis)
plt.xlabel("Timepoint")
plt.ylabel("Tumor Volume (mm3)")
plt.title("Capomulin time series for Mouse S185")
plt.savefig('../Images/capomulin_series_mouse_S185.png')
plt.show()
# -
# Generate a scatter plot of average tumor volume vs. mouse weight
# for all mice in the Capomulin regimen
single_drug = merged_cleaned_df[merged_cleaned_df["Drug Regimen"]=="Capomulin"]
capomulin_regimen_analysis = single_drug.groupby("Mouse ID")
weight = capomulin_regimen_analysis["Weight (g)"].min()
avg_tumor_volume = capomulin_regimen_analysis["Tumor Volume (mm3)"].mean()
#weight
#avg_tumor_volume
plt.scatter(weight,avg_tumor_volume)
plt.xlabel("Weight(g) per Mouse")
plt.ylabel("Average Tumor Volume (mm3) per Mouse")
plt.title("Capomulin Regimen")
plt.savefig('../Images/tumor_volume_vs_weight.png')
plt.show
# ## Correlation and Regression
# +
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
#merged_cleaned_df.head(10)
(slope, intercept, rvalue, pvalue, stderr) = st.linregress(weight, avg_tumor_volume)
regression_values = weight * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(weight,avg_tumor_volume)
plt.plot(weight,regression_values,"r-")
plt.xlabel("Weight (g) per Mouse")
plt.ylabel("Average Tumor Volume (mm3) per Mouse")
plt.title("Capomulin Regimen")
print(f"The Correlation Coefficient is: {round(st.pearsonr(weight,avg_tumor_volume)[0],2)}")
print(f"The linear regression equation is: {line_eq}")
plt.savefig('../Images/correlation_regression.png')
plt.show()
# -
# # Observations and Insights
#
# 1. Datapoints for all the drugs were around the same amount, providing quality data for analysis.
# 2. Mouse gender distribution was almost excatly equal.
# 3. Two drugs (Capomulin, Ramicane) were the most effective on reducing the tumor size.
#
| Pymaceuticals/pymaceuticals_main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
import networkx as nx
from custom import custom_funcs as cf
from networkx.algorithms import bipartite
from circos import CircosPlot
import numpy as np
import matplotlib.pyplot as plt
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction
#
# Bipartite graphs are graphs that have two (bi-) partitions (-partite) of nodes. Nodes within each partition are not allowed to be connected to one another; rather, they can only be connected to nodes in the other partition.
#
# Bipartite graphs can be useful for modelling relations between two sets of entities. We will explore the construction and analysis of bipartite graphs here.
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's load a [crime data](http://konect.uni-koblenz.de/networks/moreno_crime) bipartite graph and quickly explore it.
#
# > This bipartite network contains persons who appeared in at least one crime case as either a suspect, a victim, a witness or both a suspect and victim at the same time. A left node represents a person and a right node represents a crime. An edge between two nodes shows that the left node was involved in the crime represented by the right node.
# + slideshow={"slide_type": "subslide"}
G = cf.load_crime_network()
G.edges(data=True)[0:5]
# + slideshow={"slide_type": "slide"}
G.nodes(data=True)[0:10]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Projections
#
# Bipartite graphs can be projected down to one of the projections. For example, we can generate a person-person graph from the person-crime graph, by declaring that two nodes that share a crime node are in fact joined by an edge.
# -
# 
# ### Exercise
#
# Find the bipartite projection function in the NetworkX `bipartite` module [docs](https://networkx.github.io/documentation/networkx-1.10/reference/algorithms.bipartite.html), and use it to obtain the `unipartite` projection of the bipartite graph.
# + slideshow={"slide_type": "subslide"}
person_nodes =
pG =
pG.nodes(data=True)[0:5]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Try visualizing the person-person crime network by using a Circos plot. Ensure that the nodes are grouped by gender and then by number of connections.
# + slideshow={"slide_type": "subslide"}
nodes = sorted(____, key=lambda x: (____________, ___________))
edges = pG.edges()
edgeprops = dict(alpha=0.1)
node_cmap = {0:'blue', 1:'red'}
nodecolor = [__________________ for n in nodes]
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111)
c = CircosPlot(nodes, edges, radius=10, ax=ax, fig=fig, edgeprops=edgeprops, nodecolor=nodecolor)
c.draw()
c.fig.savefig('images/crime-person.png', dpi=300)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Use a similar logic to extract crime links.
# + slideshow={"slide_type": "subslide"}
crime_nodes = _________
cG = _____________ # cG stands for "crime graph"
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Can you plot how the crimes are connected, using a Circos plot? Try ordering it by number of connections.
# + slideshow={"slide_type": "subslide"}
nodes = sorted(___________, key=lambda x: __________)
edges = cG.edges()
edgeprops = dict(alpha=0.1)
nodecolor = plt.cm.viridis(np.arange(len(nodes)) / len(nodes))
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111)
c = CircosPlot(nodes, edges, radius=10, ax=ax, fig=fig, edgeprops=edgeprops, nodecolor=nodecolor)
c.draw()
plt.savefig('images/crime-crime.png', dpi=300)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# NetworkX also implements centrality measures for bipartite graphs, which allows you to obtain their metrics without first converting to a particular projection. This is useful for exploratory data analysis.
#
# Try the following challenges, referring to the [API documentation](https://networkx.github.io/documentation/networkx-1.9/reference/algorithms.bipartite.html) to help you:
#
# 1. Which crimes have the most number of people involved?
# 1. Which people are involved in the most number of crimes?
# + slideshow={"slide_type": "subslide"}
# Degree Centrality
bpdc = _______________________
sorted(___________, key=lambda x: ___, reverse=True)
| 7. Bipartite Graphs (Student).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# ML_in_Finance-Backpropagation
# Author: <NAME>
# Version: 1.0 (24.7.2019)
# License: MIT
# Email: <EMAIL>
# Notes: tested on Mac OS X with Python 3.6.9 with the following packages
# numpy=1.18.1, keras=2.3.1, tensorflow=2.0.0
# Citation: Please cite the following reference if this notebook is used for research purposes:
# <NAME>., <NAME>. and <NAME>, Machine Learning in Finance: From Theory to Practice, Springer Graduate textbook Series, 2020.
# This notebook is courtesy of <NAME>, Imperial College (<EMAIL>) and demonstrates backpropagation.
# -
# # Overview
# This notebooks demonstrate the back-propagation algorithm in detail and compares the results of an implementation with tensorflow. See Section 5.1 of Chapter 4 for further details.
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
from numpy.linalg import norm
import copy
import os
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
def relu(x):
return x*(np.sign(x)+1.)/2.
# + _uuid="ab7671c8b804e06579cfab4d74d8c6fec680eb0a"
def sigmoid(x):
return 1./(1.+np.exp(-x))
# + _uuid="9644cbf438ea83c5b6797eea3d2d8d61d32c673d"
def softmax(x):
return np.exp(x)/sum(np.exp(x))
# + _uuid="6aba0cf2b87a1a1f8c040f3f4df19a8d752501da"
def mynorm(Z):
return np.sqrt(np.mean(Z**2))
# + [markdown] _uuid="ba0305a6cb5cd493c395c105bf2b7f01442893b3"
# Let us consider a feed-forward architecture with an input layer, $L-1$ hidden layers and one output layer, with $K$ units in the output layer. As a result, we have $L$ sets of weights and biases $(W^{(\ell)}, \mathbf{b}^{(\ell)})$ for $\ell=1,\dots, L$, corresponding to the layer inputs $Z^{(\ell-1)}$ and outputs $Z^{(\ell)}$ for $\ell=1,\dots, L$. Recall that each layer is an activation of a semi-affine transformation, $I^{(\ell)}(Z^{(\ell-1)}):=W^{(L)}Z^{(\ell-1)}+ b^{(L)}$. The corresponding activation functions are denoted as $\sigma^{(\ell)}$. The activation function for the output layer is a softmax function, $\sigma_s(x)$.
#
# Here we use the cross-entropy as the loss function, which is defined as
# $$ \mathcal{L}:= -\sum_{k=1}^{K}Y_{k}\log \hat{Y}_{k}.$$
#
# The relationship between the layers, for $\ell\in\{1,\dots, L\}$ are:
#
# \begin{align*}
# \hat{Y} (X) & = Z^{(L)}=\sigma_s(I^{(L)}) \in [0,1]^{K},\\
# Z^{(\ell)} & = \sigma^{(\ell)} \left ( I^{(\ell)} \right ), ~\ell=1,\dots,L-1,\\
# Z^{(0)} & = X.\\
# \end{align*}
#
#
# The update rules for the weights and biases are
# \begin{align*}
# \Delta W^{(\ell)} &= - \gamma \nabla_{W^{(\ell)}}\mathcal{L},\\
# \Delta \mathbf{b}^{(\ell)} &= - \gamma \nabla_{\mathbf{b}^{(\ell)}}\mathcal{L}.
# \end{align*}
# We now begin the Back-Propagation.
#
# For the gradient of $\mathcal{L}$ w.r.t. $W^{(L)}$ we have
# \begin{align*}
# \frac{\partial \mathcal{L}}{\partial w_{ij}^{(L)}} &= \sum_{k=1}^{K}\frac{\partial \mathcal{L}}{\partial Z_{k}^{(L)}} \frac{\partial Z_{k}^{(L)}}{\partial w_{ij}^{(L)}}\\
# %%%%%%%%%%%
# &= \sum_{k=1}^{K}\frac{\partial \mathcal{L}}{\partial Z_{k}^{(L)}} \sum_{m=1}^{K}\frac{\partial Z_{k}^{(L)}}{\partial I_{m}^{(L)}} \frac{\partial I_{m}^{(L)}}{\partial w_{ij}^{(L)}}
# \end{align*}
# But
# \begin{align*}
# \frac{\partial \mathcal{L}}{\partial Z_{k}^{(L)}} &= -\frac{Y_{k}}{Z_{k}^{(L)}}\\
# %%%%%%%%%%%%%%
# \frac{\partial Z_{k}^{(L)}}{\partial I_{m}^{(L)}} &= \frac{\partial}{\partial I_{m}^{(L)}}[\sigma(I^{(L)})]_{k}\\
# %%%%%%%%%%%%%%
# &= \frac{\partial}{\partial I_{m}^{(L)}} \frac{\exp[I_{k}^{(L)}]}{\sum_{n=1}^{K}\exp[I_{n}^{(L)}]}\\
# %%%%%%%%%%%%%%
# &= \begin{cases}
# -\frac{\exp[I_{k}^{(L)}]}{\sum_{n=1}^{K}\exp[I_{n}^{(L)}]} \frac{\exp[I_{m}^{(L)}]}{\sum_{n=1}^{K}\exp[I_{n}^{(L)}]} & \text{if } k \neq m \\
# \frac{\exp[I_{k}^{(L)}]}{\sum_{n=1}^{K}\exp[I_{n}^{(L)}]} - \frac{\exp[I_{k}^{(L)}]}
# {\sum_{n=1}^{K}\exp[I_{n}^{(L)}]} \frac{\exp[I_{m}^{(L)}]}{\sum_{n=1}^{K}\exp[I_{n}^{(L)}]}
# & \text{otherwise}
# \end{cases}\\
# %%%%%%%%%%%%%%
# &= \begin{cases}
# -\sigma_{k}\sigma_{m}& \text{if } k \neq m \\
# \sigma_k(1 - \sigma_m) & \text{otherwise}
# \end{cases}\\
# %%%%%%%%%%%%%%
# &= \sigma_k(\delta_{km} - \sigma_m) \quad \text{where} \, \delta_{km} \, \text{is the Kronecker's Delta}\\
# %%%%%%%%%%%%%%
# \frac{\partial I_{m}^{(L)}}{\partial w_{ij}^{(L)}} &= \delta_{mi}Z_{j}^{(L-1)}\\
# %%%%%%%%%%%%%%
# \implies \frac{\partial \mathcal{L}}{\partial w_{ij}^{(L)}} &= -\sum_{k=1}^{K}\frac{Y_{k}}{Z_{k}^{(L)}}
# \sum_{m=1}^{K} Z_{m}^{(L)}(\delta_{km} - Z_{m}^{(L)}) \delta_{mi}Z_{j}^{(L-1)}\\
# %%%%%%%%%%%%%%
# &= -Z_{j}^{(L-1)} \sum_{k=1}^{K}Y_{k} (\delta_{ki} - Z_{i}^{(L)}) \\
# %%%%%%%%%%%%%%
# &= Z_{j}^{(L-1)} (Z_{i}^{(L)}-Y_{i})
# \end{align*}
# Where we have used the fact that $\sum_{k=1}^{K}Y_{k}=1$ in the last equality.
#
# Similarly for $\mathbf{b}^{(L)}$, we have
# \begin{align*}
# \frac{\partial \mathcal{L}}{\partial b_{i}^{(L)}} &= \sum_{k=1}^{K}\frac{\partial \mathcal{L}}{\partial Z_{k}^{(L)}} \sum_{m=1}^{K}\frac{\partial Z_{k}^{(L)}}{\partial I_{m}^{(L)}} \frac{\partial I_{m}^{(L)}}{\partial b_{i}^{(L)}}\\
# %%%%%%%%%%%%%%%%%%
# &= Z_{i}^{(L)}-Y_{i}
# \end{align*}
# It follows that
# \begin{align*}
# \nabla_{\mathbf{b}^{(L)}}\mathcal{L} &= Z^{(L)}-Y\\
# \nabla_{W^{(L)}}\mathcal{L} &= \nabla_{\mathbf{b}^{(L)}}\mathcal{L} \otimes {Z^{(L-1)}}
# \end{align*}
# Where $\otimes$ denotes the outer product.
# + [markdown] _uuid="baff8933f9f9ab7421c873f385975555d58efcec"
# For the gradient of $\mathcal{L}$ w.r.t. $W^{(L-1)}$ we have
# \begin{align*}
# \frac{\partial \mathcal{L}}{\partial w_{ij}^{(L-1)}} &= \sum_{k=1}^{K}\frac{\partial L}{\partial Z_{k}^{(L)}} \frac{\partial Z_{k}^{(L)}}{\partial w_{ij}^{(L-1)}}\\
# %%%%%%%%%%%
# &= \sum_{k=1}^{K}\frac{\partial \mathcal{L}}{\partial Z_{k}^{(L)}} \sum_{m=1}^{K}\frac{\partial Z_{k}^{(L)}}{\partial I_{m}^{(L)}} \sum_{n=1}^{n^{(L-1)}} \frac{\partial I_{m}^{(L)}}{\partial Z_{n}^{(L-1)}} \sum_{p=1}^{n^{(L-1)}} \frac{\partial Z_{n}^{(L-1)}}{\partial I_{p}^{(L-1)}} \frac{\partial I_{p}^{(L-1)}}{\partial w_{ij}^{(L-1)}}
# \end{align*}
# If we assume that $\sigma^{(\ell)}(x)=\text{sigmoid}(x), ~\ell \in \{1,\dots, L-1\}$ then
# \begin{align*}
# \frac{\partial I_{m}^{(L)}}{\partial Z_{n}^{(L-1)}} &= w_{mn}^{(L)}\\
# %%%%%%%%%%%%%%%%%
# \frac{\partial Z_{n}^{(L-1)}}{\partial I_{p}^{(L-1)}} &= \frac{\partial}{\partial I_{p}^{(L-1)}}\bigg(\frac{1}{1+\exp(-I_{n}^{(L-1)})}\bigg)\\
# %%%%%%%%%%%%%%%%%
# &= \frac{1}{1+\exp(-I_{n}^{(L-1)})} \frac{\exp(-I_{n}^{(L-1)})}{1+\exp(-I_{n}^{(L-1)})} \, \delta_{np} \\
# %%%%%%%%%%%%%%%%%
# &= Z_{n}^{(L-1)} (1-Z_{n}^{(L-1)}) \, \delta_{np} = \sigma^{(L-1)}_n(1-\sigma^{(L-1)}_n)\delta_{np} \\
# %%%%%%%%%%%%%%%%%
# \frac{\partial I_{p}^{(L-1)}}{\partial w_{ij}^{(L-1)}} &= \delta_{pi} Z_{j}^{(L-2)} \\
# %%%%%%%%%%%%%%%%%
# \implies \frac{\partial L}{\partial w_{ij}^{(L)}} &= -\sum_{k=1}^{K}\frac{Y_{k}}{Z_{k}^{(L)}} \sum_{m=1}^{K}Z_{k}^{(L)}(\delta_{km} - Z_{m}^{(L)}) \sum_{n=1}^{n^{(L-1)}} w_{mn}^{(L)} \sum_{p=1}^{n^{(L-1)}} Z_{n}^{(L-1)} (1-Z_{n}^{(L-1)}) \, \delta_{np} \delta_{pi} Z_{j}^{(L-2)} \\
# %%%%%%%%%%%%%%%%%
# &= -\sum_{k=1}^{K}Y_{k} \sum_{m=1}^{K}(\delta_{km} - Z_{m}^{(L)}) \sum_{n=1}^{n^{(L-1)}} w_{mn}^{(L)} Z_{n}^{(L-1)} (1-Z_{n}^{(L-1)}) \, \delta_{ni} Z_{j}^{(L-2)} \\
# %%%%%%%%%%%%%%%%%
# &= -\sum_{k=1}^{K}Y_{k} \sum_{m=1}^{K}(\delta_{km} - Z_{m}^{(L)}) w_{mi}^{(L)} Z_{i}^{(L-2)} (1-Z_{i}^{(L-1)}) Z_{j}^{(L-2)} \\
# %%%%%%%%%%%%%%%%%
# &= -Z_{j}^{(L-2)}Z_{i}^{(L-1)}(1-Z_{i}^{(L-1)}) \sum_{m=1}^{K} w_{mi}^{(L)} \sum_{k=1}^{K}(\delta_{km}Y_{k} - Z_{m}^{(L)}Y_{k}) \\
# %%%%%%%%%%%%%%%%%
# &= Z_{j}^{(L-2)}Z_{i}^{(L-1)} (1-Z_{i}^{(L-1)}) \sum_{m=1}^{K} w_{mi}^{(L)} (Z_{m}^{(L)} - Y_{m}) \\
# %%%%%%%%%%%%%%%%%
# &= Z_{j}^{(L-2)}Z_{i}^{(L-1)} (1-Z_{i}^{(L-1)}) (Z^{(L)} - Y)^{T} \mathbf{w}_{,i}^{(L)} \\
# \end{align*}
#
# Similarly we have
#
# $$ \frac{\partial \mathcal{L}}{\partial b_{i}^{(L-1)}} = Z_{i}^{(L-1)} (1-Z_{i}^{(L-1)}) (Z^{(L)} - Y)^{T} \mathbf{w}_{,i}^{(L)}. $$
# It follows that we can define the following recursion relation for the loss gradient:
#
# \begin{align*}
# \nabla_{b^{(L-1)}}\mathcal{L} &= Z^{(L-1)} \circ (\mathbf{1}-Z^{(L-1)}) \circ
# ({W^{(L)}}^{T} \nabla_{b^{(L)}}\mathcal{L}) \\
# \nabla_{W^{(L-1)}}\mathcal{L} &= \nabla_{b^{(L-1)}}\mathcal{L} \otimes Z^{(L-2)}\\
# & = Z^{(L-1)} \circ (\mathbf{1}-Z^{(L-1)}) \circ
# ({W^{(L)}}^{T} \nabla_{W^{(L)}}\mathcal{L})
# \end{align*}
#
# Where $\circ$ denotes the Hadamard Product (elementwise multiplication). This recursion relation generalizes for all layers. To see this, let the back-propagation error $\delta^{(\ell)}:=\nabla_{b^{(\ell)}}\mathcal{L}$, and since
#
# \begin{align*}
# \left[\frac{\partial \sigma^{(\ell)}}{\partial I^{(\ell)}}\right]_{ij}&=\frac{\partial \sigma_i^{(\ell)}}{\partial I_j^{(\ell)}}\\
# &=\sigma_i^{(\ell)}(1-\sigma_i^{(\ell)})\delta_{ij}\\
# \end{align*}
#
# Equivalently in matrix-vector form
# $$\nabla_{I^{(\ell)}} \sigma^{(\ell)}=\text{diag}(\sigma^{(\ell)} \circ (\mathbf{1}-\sigma^{(\ell)})).$$
#
# We can write, in general, for any choice of activation function for the hidden layer,
#
# $$ \delta^{(\ell)}=(\nabla_{I^{(\ell)}} \sigma^{(\ell)})(W^{(\ell+1)})^T\delta^{(\ell+1)}.$$
# and
#
# $$\nabla_{W^{(\ell)}}\mathcal{L} = \delta^{(\ell)} \otimes Z^{(\ell-1)}.$$
# -
# ## Backpropagation Example
# Here we define a three layer feed-forward network as in the Example given in Chapter 4.
# On each pass through the training loop, the training input is fed forward through the network to calculate the loss, then the gradient of the loss with respect to each of the weights is calculated and the weights updated for the next pass through the loop.
#
# Below, we will compare this backpropagation implementation's results with Keras
# + _uuid="d5e7c8262ea14a5f1e3678534442f0c6292486f2"
def myANN(Y, Xtrain, Xpred, W01, W02, W03, b01, b02, b03):
# Initialization of Weights and Biases
W1 = copy.copy(W01)
W2 = copy.copy(W02)
W3 = copy.copy(W03)
b1 = copy.copy(b01)
b2 = copy.copy(b02)
b3 = copy.copy(b03)
# Initialize ad hoc variables
k = 1
change = 999
# Begin the training loop
while (change > 0.001 and k < 201):
print("Iteration", k)
## Begin Feedforward (assume learning rate is one)
# Hidden Layer 1
Z1 = sigmoid(W1 @ Xtrain + b1)
# Hidden Layer 2
Z2 = sigmoid(W2 @ Z1 + b2)
# Output Layer
Yhat = softmax(W3 @ Z2 + b3)
# Find cross-entropy loss
loss = -Y @ np.log(Yhat)
print("Current Loss:",loss)
## Find gradient of loss with respect to the weights
# Output Later
dLdb3 = Yhat - Y
dLdW3 = np.outer(dLdb3, Z2)
# Hidden Layer 2
dLdb2 = (W3.T @ (dLdb3)) * Z2 * (1-Z2)
dLdW2 = np.outer(dLdb2,Z1)
# Hidden Layer 1
dLdb1 = (W2.T @ (dLdb2)) * Z1 * (1-Z1)
dLdW1 = np.outer(dLdb1, Xtrain)
## Update Weights by Back Propagation
# Output Layer
b3 -= dLdb3 # (learning rate is one)
W3 -= dLdW3
# Hidden Layer 2
b2 -= dLdb2
W2 -= dLdW2
# Hidden Layer 1
b1 -= dLdb1
W1 -= dLdW1
change = norm(dLdb1)+norm(dLdb2)+norm(dLdb3)+norm(dLdW1)+norm(dLdW2)+norm(dLdW3)
k += 1
Z1pred = W1 @ Xpred + b1
Z2pred = W2 @ sigmoid(Z1pred) + b2
Z3pred = W3 @ sigmoid(Z2pred) + b3
Ypred = softmax(Z3pred)
print("")
print("Summary")
print("Target Y \n", Y)
print("Fitted Ytrain \n", Yhat)
print("Xpred\n", Xpred)
print("Fitted Ypred \n", Ypred)
print("Weight Matrix 1 \n", W1)
print("Bias Vector 1 \n", b1)
print("Weight Matrix 2 \n", W2)
print("Bias Vector 2 \n", b2)
print("Weight Matrix 3 \n", W3)
print("Bias Vector 3 \n", b3)
# -
# Defining the initial weights of the network prior to training:
# + _uuid="5c41fd636007d1448a7a3d9e8481a417e771faec"
W0_1 = np.array([[0.1,0.3,0.7], [0.9,0.4,0.4]])
b_1 = np.array([1.,1.])
W0_2 = np.array([[0.4,0.3], [0.7,0.2]])
b_2 = np.array([1.,1.])
W0_3 = np.array([[0.5,0.6], [0.6,0.7], [0.3,0.2]])
b_3 = np.array([1.,1.,1.])
# -
# Defining the training input and the desired output of the model:
X_train = np.array([0.1,0.7,0.3])
YY = np.array([1.,0.,0.])
X_pred = X_train
# Finally we can run the input through the model, updating the weights on each iteration.
# + _uuid="9807df2b46c07bcd1f58c6e19e75b47294911283"
myANN(YY, X_train, X_pred, W0_1, W0_2, W0_3, b_1, b_2, b_3)
# -
# ## Implementing the Model with Keras
# To recreate our example neural network with Keras, we must first import the components of the package necessary to build our model. Keras provides a rich suite of model architectures, layers, activation functions and other building blocks for creating deep learning models.
#
# As we are creating a typical sequential network with three densely connected layers, we simply need the classes for the `Sequential` model and the `Dense` layer.
#
# For training, we will instantiate a stochastic gradient descent optimiser, setting its learning rate to 1 as above.
# + _uuid="3452568af4e2b0eef09b05faf226e7ca100d4340"
from keras.models import Sequential
from keras.layers import Dense
from keras.optimizers import SGD
# -
# Creating the model, we define its architecture by adding the layers in the order they are applied to the inputs, specifying their activation functions, and initialising their weights to the values defined above. Note that the weights are transposed:
# Create the model
model = Sequential()
# Add the first hidden layer
model.add(Dense(2, input_dim=3, activation='sigmoid', weights = [W0_1.T, b_1]))
# Second hidden layer
model.add(Dense(2, activation='sigmoid', weights = [W0_2.T, b_2]))
# Output layer
model.add(Dense(3, activation='softmax', weights = [W0_3.T, b_3]))
# Now we can compile the model. This is where the learning strategy for the model is specified.
#
# We provide Keras with the objective function (`loss`) to minimise during training, and the optimisation algorithm to apply in order to do so.
# + _uuid="c3022e1a7b78fbe0ca1bf116e3e96882b4f2a7c9"
sgd = SGD(lr=1)
model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['categorical_crossentropy'])
model.get_weights()
# -
# Finally, we can fit the compiled model to our training input and output, each formatted as column vectors.
# + _uuid="67897067fabc2f208b360a454d5a524940d34f2f"
model.fit(X_train.reshape((1,3)), YY.reshape((1, 3)), epochs=200, batch_size=1)
# -
# With the trained model, we can get its predicted output for an input instance.
# + _uuid="1dc231803e034841a955e31c2ce2e3d68b9eced1"
model.predict(X_pred.reshape((1, 3)))
# + [markdown] _uuid="889e04a3f4b08b56ba9dbc0a5771186bd40c23ce"
# Now we can get the weights of the trained model. Note that by convention Keras' representation of the weights and activations are transposed compared to our example above.
# + _uuid="8a0592357283bc20cd7cd47d40132b49a2cbb160"
model.get_weights()
| Chapter4-NNs/ML_in_Finance-Backpropagation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Logistic Regression
# ## classification with binary response variable
train=pd.read_csv("trainT.csv")
test=pd.read_csv("testT.csv")
train.head(6)
df=train[['Survived','Pclass','Sex','Age','Fare']]
df.head(5)
#change male to 1 and female to 0
df["Sex"] = df["Sex"].apply(lambda sex:1 if sex=="male" else 0)
df.head(5) #sex as ctegorical varible
#handle missing values of age
df["Age"] = df["Age"].fillna(df["Age"].median())
df["Fare"] = df["Fare"].fillna(df["Fare"].median())
df.isnull().sum()
# +
X_train = df.drop("Survived", axis=1) #predictors from training
Y_train = df["Survived"]
# -
logreg = LogisticRegression() #define the logistic regression model
logreg.fit(X_train, Y_train)
# +
## lets look at the test data
# -
test.head()
X_test=test[['Pclass','Sex','Age','Fare']]
X_test["Sex"] = X_test["Sex"].apply(lambda sex:1 if sex=="male" else 0)
X_test.isnull().sum()
X_test["Age"] = X_test["Age"].fillna(X_test["Age"].median())
X_test=X_test.dropna()
X_test.isnull().sum()
X_test.head(5)
Y_pred = logreg.predict(X_test)
acc_log = round(logreg.score(X_train, Y_train) * 100, 2) #accuracy of the model
acc_log
X_train.head(3)
Y_train.head(3)
g=X_train[['Sex']]
logreg = LogisticRegression() #define the logistic regression model
logreg.fit(g, Y_train)
gT=X_test[['Sex']]
gT.head(4)
preds = logreg.predict_proba(gT)
preds = pd.DataFrame(preds)
preds.columns = ["Death_prob", "Survival_prob"]
# Generate table of predictions vs Sex
pd.crosstab(train["Sex"], preds.ix[:, "Survival_prob"]) #predict the probability of survival according to gender
# # Test for Classification Accuracy
train=pd.read_csv("trainT.csv")
df=train[['Survived','Pclass','Sex','Age','Fare']]
#change male to 1 and female to 0
df["Sex"] = df["Sex"].apply(lambda sex:1 if sex=="male" else 0)
#handle missing values of age
df["Age"] = df["Age"].fillna(df["Age"].median())
df["Fare"] = df["Fare"].fillna(df["Fare"].median()) #more immune to outliers
df.head(7)
X= df.drop("Survived", axis=1)
Y = df["Survived"]
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = .3, random_state=25) #30% hold out for testing
logreg = LogisticRegression() #define the logistic regression model
logreg.fit(X_train, y_train)
Y_pred = logreg.predict(X_test) #predict the response variable based on predictors in the test set
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, Y_pred)
confusion_matrix
from sklearn.metrics import accuracy_score
accuracy_score(y_test, Y_pred) #compare with the actual y values, y_test (hold outs) with predicted y
from sklearn.metrics import cohen_kappa_score
cohen_kappa_score(y_test, Y_pred) #Scores above .8 are generally considered good agreement;
from sklearn.metrics import classification_report
report = classification_report(y_test, Y_pred)
print(report)
| section 6_part 1/Accuracy_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <div class="alert alert-info">
#
# # PSY 4016-G-H20
# # Cueillette et traitement des données en neurosciences cognitives
# ## Chapitre 3: Modification de données
# <p>
# <li> Modification des données
# <li> Combinaison des données
# <li> Les opérateurs
# <ul> <li>opérateurs de comparaison
# <li>opérateurs d'aggregation
# <li>opérateurs arithmétiques (incluses dans NumPy)
# <li>opérateurs trigonométrique; exponents, logarithms
# <li>opérateurs sur les valuers manquantes</ul>
# <li> Pandas groupby
# <div class="alert alert-info">
# <h3>Modifications de données</h3></div>
# <div class="alert alert-warning">
# <h3>modification des valeurs des chaînes</h3>
# </div>
a = "bonjour"
a
# +
a[3] = "T"
# la chaîne est un objet immuable
# -
a.index("j")
# +
# remplacement des données dans la chaîne
a.replace("njo","nTo")
# +
# chaîne -> liste
a.split("n")
# -
# <div class="alert alert-warning">
# <h3>modification des valeurs d'une liste</h3>
# </div>
ls1 = [3, 2, 1, 0]
# +
# ajout de données
ls1.append('text-ver2')
ls1
# +
# changement de données
ls1[1] = 7
print(ls1)
# +
# faire des copies
ls2 = ls1.copy()
# -
ls1.remove(3)
# +
# ls1
# +
ls1 = [3, 2, 1, 0]
ls1.pop(2)
ls1
# +
ls1 = [3, 2, 1, 0]
ls1.sort()
ls1
# -
ls1.reverse()
ls1
# <div class="alert alert-warning">
# <h3>modification des valeurs d'un tuple</h3>
# </div>
tup1 = (3, 2, 1, 0)
tup1[1] = 7
# +
# faire des copies
tup2 = tup1
print(tup2)
# +
tup1.remove(3)
# Les tuples ne peuvent être changés, une fois définis
# Les tuples sont habituellement plus rapides que les listes
# -
# <div class="alert alert-warning">
# <h3>modification des valeurs d'un dictionnaire</h3>
# </div>
# +
# ajout des valeurs
d1 = {
'cle1':'variable',
'cle2':[],
'cle3': [1,2,3],
'cle4': ('a', 'b',)
}
d1['autre'] = 'valeur2'
d1
# +
# changements des valeurs:
d1['cle1'] = 'var 2'
d1['cle1']
# +
# donnee la valeur de la clé SI elle existe
var = d1.get('cle3')
var
# +
# supprimer une clé du dictionnaire SI elle existe
d1.pop('cle4')
d1
# +
d1 = {
'cle1':'variable',
'cle2':[],
'cle3': [1,2,3],
'cle4': ('a', 'b',)
}
del d1['cle4']
d1
# +
# # supprimer tous les élements du dictionnaire
d1.clear()
d1
# -
print(len(d1), len(d1['cle1']))
# +
# faire des copies
d3 = d1
# -
d1
d3
d3['kqyky'] = 4.0
d3
d1
# +
# créer une copie superficielle
d3 = d1.copy()
# +
# créer une copie complète
d3 = d1.deepcopy()
# +
import copy
d3 = copy.deepcopy(d1)
# Il faut créer une copie indépendante
# Sinon, les modifications apportées au dictionnaire copié
# vont également être envoyées à la version d'origine
# +
# plus d'info sur la différence entre copy et deepcopy
https://thispointer.com/python-how-to-copy-a-dictionary-shallow-copy-vs-deep-copy/
https://stackabuse.com/deep-vs-shallow-copies-in-python/#:~:text=Deep%20copy%20stores%20copies%20of%20an%20object's%20values,,implementation,%20I'd%20like%20you%20to%20imagine%20this%20scenario.
http://tech.quebecblogue.com/python-tips-copie-superficielle-copie-profonde-dans-python3/
https://www.ipgirl.com/675/quelle-est-la-difference-entre-une-copie-profonde-et-une-copie-superficielle.html
# -
# <div class="alert alert-warning">
# <h3>modification des valeurs d'un pandas DataFrame</h3>
# <li>loc (location)
# <li>iloc (index location)
# <li>at
# </div>
# + slideshow={"slide_type": "fragment"}
import pandas
# -
fichier = ""
df_qc.ETABLISSEMENT[1]
# +
# pour montrer la colonne
df_qc['ETABLISSEMENT']
# +
# semblable à la précédente; loc = location
df_qc.loc[:, 'ETABLISSEMENT']
# -
df_qc.iloc[1:] # affiche toutes les lignes de 1 :; iloc = index-location
df_qc.iloc[0:, [1,2]] # affiche les lignes de 0 à : et les colonnes 1 et 2
df_qc.at[0,'ETABLISSEMENT'] # permet l'accès aux cellules
#index 0 et col 'str'
col = df_qc.columns[1] #
df_qc[col][:3]
df_qc.ETABLISSEMENT
df_qc.loc[1, 'ETABLISSEMENT'] = 'POLY'
df_qc.at[2, 'ETABLISSEMENT'] = 'Sherbrooke'
df_qc.iloc[4, 1] = 'ULaval'
df_qc.ETABLISSEMENT
df_qc2 = df_qc
df_qc.ETABLISSEMENT.head(3)
df_qc2.ETABLISSEMENT.head(3)
df_qc2.at[1, 'ETABLISSEMENT'] = 'poly_new'
df_qc2.at[1, 'ETABLISSEMENT']
df_qc.at[1, 'ETABLISSEMENT']
# +
# faire des copies
df_qc = pandas.read_csv(fichier)
df_qc2 = df_qc.copy(deep=True)
df_qc2.at[1, 'ETABLISSEMENT'] = 'poly_new'
df_qc2.ETABLISSEMENT[1]
df_qc.ETABLISSEMENT[1]
# -
import copy
df_qc2 = copy.deepcopy(df_qc)
df_qc2.at[1, 'ETABLISSEMENT'] = 'poly_new'
df_qc3 = pandas.read_csv(fichier, usecols = ['ETABLISSEMENT', 'COUT_TOTAL'])
df_qc3.columns
df_qc4 = df_qc[["ETABLISSEMENT", "ETABLISSEMENT_PARTENAIRE",
'ANNEE_FINANCIERE', 'PROGRAMME', 'VOLET', 'RESPONSABLE_DU_PROJET',
'TITRE', 'RESEAU', 'MONTANT_MEES', 'MONTANT_MSSS', 'MONTANT_MEI',
'MONTANT_TOTAL_QUEBEC', 'COUT_TOTAL']]
df_qc4.columns
# +
# df.department.tolist() permet d'extraire le contenu d'une col en une liste
ls_df = df.department.tolist()
#ls_df
# -
ls_df = list(df.items())
ls_df[1]
# +
# création d'une nouvelle colonne appelée «autre»
df_qc2['autre'] = 'valeur2'
#df_qc2['autre'][:5]
# -
df_qc
# +
# modification d'index -- prend une col et la définit en tant qu"index
df_qc.set_index('ETABLISSEMENT')
# -
df_qc
df_qc.set_index('ETABLISSEMENT', inplace = True)
## pour faire le changeent, on peut utiliser inplace ou simplement stocker le changement dans un nouveau jeu de donées
df_qc3 = df_qc.set_index('ETABLISSEMENT')
df_qc
df_qc.index[1]
df_qc.at[1, 'COUT_TOTAL']
#contenu dans la ligne avec le nb 1 et la col "cout total"
df_qc.index
df_qc.at['École Polytechnique de Montréal', 'COUT_TOTAL']
#
df_qc.to_excel('chemin/tmp2.xls')
df_qc.to_csv('chemin/tmp2.csv')
# <div class="alert alert-warning">
# <h3>modification des valeurs d'un tableau Numpy</h3>
# <li> np.reshape
# <li> np.newaxis
# <li> copy
# <li> changer les éléments
# </div>
# + slideshow={"slide_type": "fragment"}
import numpy as np
# -
x1
x1_2D = x1.reshape(0,1)
x1_2D
x1_2D2 = np.reshape(x1, (3, -1)) #-1 décide automatiquement le nombre de cols
x1_2D2
# +
# np.newaxis pour convertir un tableau 1D
# en un vecteur de ligne ou colonne
x1[:,np.newaxis]
# +
# changer de plus grandes tables
mon_tab = np.array(((6, 12, 93, 2),
(5, 26, 78, 90),
(3, 12, 16, 22),
(5, 3, 1, 16)))
mon_tab
# -
mon_tab.shape
mon_tab_remodel = np.reshape(mon_tab, (8,2))
m3 = mon_tab[:, np.newaxis]
m3
m3.shape
mon_tab
np.reshape(mon_tab, (8,3))
np.reshape(mon_tab, (8,-1))
x2
x2_v1 = x2[1:,2:].copy()
x2_v1
x2_v2 = x2[:2, :2].copy()
x2_v2
x2[0,0] = 12
x2
x2[0,1] = 3.141
x2 # ? ou est la nouvelle valeur ?
# <div class="alert alert-warning">
# <b>REMARQUE ! </b>
# <ul> contrairement aux listes, les tableaux numpy ont un type fixe, i.e., float in int = int</div>
# <div class="alert alert-warning">
# <b>trier les tableaux numpy</b>
# <li> np.sort
# <li> np.argsort</div>
a = np.array([2,5,3,7,5,3])
a
#np.sort(a)
a.sort()
a
np.argsort(a) #affiche les indices des éléments triés
# <div class="alert alert-info">
# <h3>Combinaison des données</h3></div>
# <div class="alert alert-warning">
# <h3>combiner les chaînes</h3>
# </div>
var2 = 12
var2
var1+var2
text3, c, text4 = 'montreal', 4, '.ca'
text3
text3, c, text4
print(c)
print(text3+c)
print(text3+str(c))
# +
# string() commands:
# len()
# strip() # Les espaces de début sont supprimés
# strip(char) # retourne une copie de la chaîne dans laquelle tous les caractères ont été supprimés du DÉBUT et de la FIN de la chaîne
# rstrip(char) # retourne une copie de la chaîne dans laquelle tous les caractères ont été supprimés de la FIN de la chaîne
# replace(char)
# split(char, char)
# find(char)
# isdigit()
# -
s = 'bonjour'
# len(s)
#s.find(j)
s.rfind('jo')
#s.rstrip('ur')
#s.replace('b','c')
# all string methods:
https://docs.python.org/2/library/string.html
https://docs.python.org/3/library/stdtypes.html
https://www.programiz.com/python-programming/methods/string
# +
a = "aBDnnc"
a.islower()
# +
# ATTENTION avec strip(), rstrip()
# si la commande comprend plusieurs caractères,
# les caractères du milieu seront traités comme une commande de recherche
# ex2 = "texts.xlsx"
# ex2.strip(".xlsx")
# ex2.rstrip(".xlsx")
# ex3 = "textn.fini"
# ex3.strip(".fini")
# ex3.rstrip(".fini")
# ex4 = "textt.fiti"
# ex4.strip(".fiti")
# ex4.rstrip(".fiti")
# strip() supprime les caractères du début et de la fin
# rstrip() supprime le caractères de la fin.
# -
# <div class="alert alert-warning">
# <h3>combiner les entieres et les flottants</h3>
# </div>
i = 30
ii = i+7
ii
f = 3.0
f
ff = f+8
ff
fff = ff+ii
fff
f/i
ff/f
fi = float(i)
fi
iff = int(f)
iff
type(i)
type(f)
text3
type(text3)
# +
# integer methods:
https://docs.python.org/3/library/stdtypes.html
# +
a = 7676
a.bit_length()
# -
m = 8.0
m.is_integer()
# <div class="alert alert-warning">
# <h3>combiner les listes</h3>
# </div>
# +
# concatenation:
ls3 = ls1+ls1[2:]
ls3
# -
tup3 = tup1+tup1[-2:]
tup3
tup4 = ls1+tup1
tup4
# +
# commandes python utilisables sur les listes
help(ls1)
len(ls) # taille de la liste ls
sorted(ls) # enregistrera une liste triée
# commande spécifique pour les listes:
ls.sort # affichera une liste triée mais ne l'enregistrera pas
ls.append # ajout d’un element à la fin de la liste ls
ls.reverse # inverser la liste ls
ls.index # rechercher un élément dans la liste ls
ls.remove # retirer un element de la liste ls
ls.pop # retirer le dernier element de la liste ls
# -
ls1_sorted = sorted(ls1[:-3])
ls1_sorted
a = [5, 2, 3, 1, 4]
a.sort()
a
ls_fruits = ["pomme",'banane','orange','poire','raisins']
fruits = pandas.DataFrame(ls_fruits)
fruits
fruits.values
fruits.to_csv("C:/Users/USER/Desktop/tmp.csv")
# +
# fruits.to_excel("C:/Users/USER/Desktop/tmp.csv")
# -
# <center><img src="img/Tableau_comparaison_liste_tuple.png"></center>
# <div class="alert alert-warning">
# <h3>combiner les dictionnaires</h3>
# </div>
# +
# La concaténation des dictionnaires peut comporter des erreurs
# Vérifiez la version de python.
# -
x = {'a': 1, 'b': 2}
y = {'b': 3, 'c': 4}
# +
# python >3.9
z = x | y
z
# ATTENTION!
# la valeur des clés avec le même nom
# prendra TOUJOURS la valeur du deuxième dictionnaire
# +
# python >3.5
z = {**x, **y}
z
# +
# la fusion peut être effectuée en incluant de nouvelles clés:
z = {**x, 'foo': 1, 'bar': 2, **y}
z
# +
# python 2
z = x.copy()
z.update(y)
z
# +
# pour une réponse plus complète:
# https://stackoverflow.com/questions/38987/how-do-i-merge-two-dictionaries-in-a-single-expression-take-union-of-dictionari
# -
# <div class="alert alert-warning">
# <h3>combiner les pandas.DataFrame</h3>
# </div>
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <ul> <h5>Concat/ Append</h5>
# <li> Concat donne la flexibilité de rejoindre les jeux de données en fonction de l'axe (toutes les lignes ou toutes les colonnes)
# <p><p> pandas.concat(objs, axis=0, join='outer', join_axes=None, ignore_index=False, keys=None, levels=None, names=None, verify_integrity=False, sort=None, copy=True)<p>
# <li> Append est le cas spécifique (axe = 0, join = 'outer') de concat. il a été écrit avant l'écriture de la fonction concat. il peut être supprimé dans les versions ultérieures.
# <p><p> pandas.DataFrame.append(other, ignore_index=False, verify_integrity=False, sort=None)</ul>
# <p>
# <ul> <h5>Merge/ Join</h5>
# <li> Merge est basée sur une colonne particulière de chacun des deux cadres de données, ces colonnes sont des variables comme «left_on», «right_on», «on»
# <p><p> pandas.DataFrame.merge(right, how='inner', on=None, left_on=None, right_on=None, left_index=False, right_index=False, sort=False, suffixes=('_x', '_y'), copy=True, indicator=False, validate=None)<p>
# <li> Join est basée sur les index et sur la façon dont les variable se combinent =['left','right','inner','outer']. Utilisez le join pour la jointure index sur index (par défaut) et colonne (s) sur index. Si vous vous joignez uniquement à l'index, vous souhaiterez peut-être utiliser join pour vous éviter de taper.
# <p><p> pandas.DataFrame.join(self, other, on=None, how='left', lsuffix='', rsuffix='', sort=False)</ul></div>
# -
# <div class="alert alert-success">
# <li> Créons deux cadres de données à partir de deux dictionnaires
# </div>
df_A = pandas.DataFrame({'a':5, 'b':2, 'c':1}, index = [0])
df_B = pandas.DataFrame({'a':3, 'b':4, 'c':2}, index = [0])
df_C = pandas.DataFrame({'a':7, 'b':2, 'c':8}, index = [0])
df_A.append(df_B)
df_Concat = pandas.concat([df_A, df_B], axis = 0) # , axis=0 is default - pour les colonnes
df_Concat
df_Concat = pandas.concat([df_A, df_B], axis = 1) # axis=1 = pour l'index
df_Concat
pandas.concat([df_A, df_B], keys=['rouge','bleue'], axis=0)
df_office1 = pandas.DataFrame({'name':['Alana','Pierre','Corinne','Marianne'],
'group':['Bank','Science','Resources','Plannification']})
df_office2 = pandas.DataFrame({'name':['Alana','Pierre','Corinne','Marianne'],
'date':['2004','2008','2006','2012']})
pandas.concat([df_office1, df_office1])
pandas.concat([df_office1, df_office1])
pandas.concat([df_office1, df_office2], axis=1)
df_office1.merge(df_office2)
df_office1
df_office2
df_office1.join(df_office2.set_index('name'), on='name')
# <div class="alert alert-warning">
# <h3>combiner les numpy.array</h3>
# </div>
# <div class="alert alert-warning">
# <ul> <b>Concatenation </b></div>
# +
# concaténation de tableaux
tab1 = np.array(((1, 2),
(3, 4)))
tab2 = np.array(((5, 6),
(7, 8)))
tab3 = np.array(((9, 10),
(11, 12)))
# -
tab1.shape
tab1
concat1 = np.concatenate((tab1, tab2, tab3))
concat1
concat1.shape
# +
# concaténation de tableaux, different types
a = np.array([1,2,3])
b = np.array([4,5,6])
c = np.array([7,8,9])
print(a, b,c)
# -
# <center><img src="img/numpy_vstack_hstack.png"><center>
np.vstack([a,b,c]) #concaténation vertical
chk1 = np.vstack([a,b,c]) #concaténation vertical
print(chk1.shape, chk1.ndim)
np.hstack([a,b,c]) #concaténation horizontale
chk_v = np.hstack([a,b,c]) #concaténation vertical
print(chk_v.shape, chk_v.ndim)
np.c_[a,b,c] #concaténation vertical, Version 2
np.r_[a,b,c] #concaténation horizontale, Version 2
# <div class="alert alert-warning">
# <b>Division de tableaux </b>
# </div>
# <div class="alert alert-success">
# <li> np.split
# <li> np.vsplit
# <li> np.hsplit</div>
tab_concat = np.array(((1, 2),
(3, 4),
(5, 6),
(7, 8),
(9, 10),
(11, 12)))
tab_concat
tab_concat.shape, tab_concat.ndim
# <center><img src="img/numpy_split.png"><center>
np.split(tab_concat, 3)
# <center><img src="img/numpy_vsplit.png"><center>
# +
# numpy.vsplit(ary, indices_or_sections)
np.vsplit(tab_concat, [3])
# -
u1, u2 = np.vsplit(tab_concat, [3])
u1
tab_concat
[ 1, 2],[ 3, 4],
# <center><img src="img/numpy_hsplit.png"><center>
u1, u2 = np.hsplit(tab_concat, [1])
u2
# <div class="alert alert-info">
# <h2>Les Operateurs</h2>
# <ul> <li>opérateurs de comparaison
# <li>opérateurs d'aggregation
# <li>opérateurs arithmétiques (incluses dans NumPy)
# <li>opérateurs trigonométrique; exponents, logarithms
# <li>opérateurs sur les valuers manquantes</ul></div>
# + active=""
# # types de data
#
# bool_ True/False
# int_ entier par défaut
# int8 Octet
# int16/int32/int64 entier
# float_ flotte64/ flotte32/ flotte16
# complex64/complex128 nombres complexes de deux flottants 32/64 octet
# -
# <div class="alert alert-warning">
# <h3>opérateurs de comparaison</h3></div>
# + active=""
# == Égal à
# != Inégal
# < Moins que
# > Plus grand que
# <= Inférieur ou égal à
# >= Égal ou supérieur à
#
# Opérateurs logiques
#
# and
# or
# not
#
# Opérateurs d'identité
# is
# is not
#
# Opérateurs d'adhésion
# in
# not in
#
# Opérateurs au niveau du bit
# & AND
# | OR
# +
import pandas
fichier = r'C:\Users\Dylan\Downloads\proj_infra_rechr_approu_20210423.csv'
df_qc = pandas.read_csv(fichier)
df_qc = pandas.read_csv(fichier, usecols = ['REGION_ADMINISTRATIVE', 'ETABLISSEMENT'])
# -
df_qc.columns
# +
# utiliser les méthodes de Bool pour examiner les
# clés / indices et les valeurs - pour vérifier si la valeur est présente
'ETABLISSEMENT' in df_qc
# -
'etab' in df_qc
df_qc.head
df_qc.col.tolist()
df_qc[(df_qc["COUT_TOTAL"]>1400000) & (df_qc["COUT_TOTAL"]<5500000)]
df_qc[df_qc.ETABLISSEMENT == 'Université de Montréal']
df_qc[
(df_qc.ETABLISSEMENT == 'Université de Montréal') &
(df_qc.COUT_TOTAL < 100000)]
a = np.random.randint(10, size=8)
a
a<4
a!=3
# <div class="alert alert-warning">
# <h3>opérateurs d'aggregation</h3></div>
# + active=""
# python:
#
# min(a) = minimum
# max(a) = maximum
#
# pandas:
# min() = minimum
# max() = maximum
# std() = écart type
# var() = variance
# count() = Nombre total d'éléments
# first() = premier élément
# last() = dernier élément
# mean() = moyenne
# median() = médiane
# mad() = écart absolu moyen
# prod() = produit de tous les articles
# sum() = somme de tous les éléments
#
#
# numpy: (operate more quickly)
# np.min
# np.max
# np.sum
# np.prod
# np.mean
# np.std
#
# np.var = variance
# np.argmin = index of min value
# np.argmax = index of max value
# np.median = compute median
# np.percentile = compute rank
# np.any = if any True
# np.all = if all True
# -
import seaborn as sns
#pip3 install seaborn
# +
# lecture à partir de jeux de données de modules:
planets = sns.load_dataset('planets')
#titanic = sns.load_dataset('titanic')
#iris = sns.load_dataset('iris')
planets.head()
# -
planets.head(10)
planets.columns
# +
# planets.head(3)
# -
planets.count()
planets.max()
planets.method.count(), len(planets.method)
planets.mass.max()
planets.orbital_period.mad()
ls = planets.method.tolist()
# #ls
planets.method.value_counts()
# +
# planets.dropna().describe()
# -
planets['number'].mean()
planets.number.mean()
x1 = planets.number
np.any(x1 > 5)
planets.number.mean()
np.sum(a>5)
import numpy as np
np.sum(b ==3, axis=1) #nombre de valeurs = 3 dans chaque ligne
np.all(b<8) # toutes les valeurs sont-elles inférieures à 8?
# <div class="alert alert-warning">
# <h3>opérateurs arithmétiques</h3></div>
# <div class="alert alert-warning">
# <ul> <b>opérateurs arithmétiques - pandas</b></div>
# + slideshow={"slide_type": "subslide"} active=""
# pandas:
#
# add() = +
# sub(), substract() = -
# mul(), multiply() = *
# truediv(), div(), divide() = /
# floordir() = //
# mod() = %
# pow() = **
# + active=""
# numpy:
#
# np.add + Addition (3 + 6 = 9)
# np.subtract - Subtraction (6 – 3 = 3)
# np.negative - Negation ( -3)
# np.multiply * Multiplication (3 * 3 = 9)
# np.divide / Division (6/3 = 2; 3/2 = 1)
# np.floor_divide // Floor division (6//3 = 2; 3//2 = 1)
# np.power ** Exponentiation (3**3 = 27; 2**3 = 8)
# np.mod % Modulus/remainder(9%3=0; 9%4 = 1)
# -
df_qc['COUT/3'] = df_qc.COUT_TOTAL/3
#df_qc['COUT/3'].tail()
df_qc['COUT/3'].head(3)
type(df_qc['COUT/3'][0])
# +
# df_qc.columns
# -
df_qc['soustraction'] = df_qc['COUT_TOTAL'].subtract(df_qc['MONTANT_TOTAL_QUEBEC'])
df_qc['soustraction'].head()
# <div class="alert alert-danger">
# <h3>Exercise</h3>
# <li> Comment vérifier si le code de soustraction est correct ?
# </div>
df_qc.COUT_TOTAL.head()
df_qc.MONTANT_TOTAL_QUEBEC.head()
df_qc['soustraction'].head()
# +
# vérification:
val1 = df_qc['COUT_TOTAL'][1]
val2 = df_qc['MONTANT_TOTAL_QUEBEC'][1]
finale = val1-val2
val3 = df_qc['soustraction'][1]
finale == val3
(df_qc.COUT_TOTAL[1])-(df_qc.MONTANT_TOTAL_QUEBEC[1]) == df_qc.soustraction[1]
# -
df_test1 = df_qc.loc[df_qc.COUT_TOTAL>20000000, ['REGION_ADMINISTRATIVE', 'ANNEE_FINANCIERE']].head()
df_test1
df_test1.T
# +
# df_qc[df_qc.COUT_TOTAL > 200000]
# +
# df_qc.loc[df_qc.COUT_TOTAL>20000000, ['REGION_ADMINISTRATIVE', 'ANNEE_FINANCIERE']].head()
# -
df_qc[(df_qc.COUT_TOTAL>1400000) & (df_qc.COUT_TOTAL<5500000)]
# <div class="alert alert-warning">
# <ul> <b>opérateurs arithmétiques - NumPy</b></div>
np.add + Addition (3 + 6 = 9)
np.subtract - Subtraction (6 – 3 = 3)
np.negative - Negation ( -3)
np.multiply * Multiplication (3 * 3 = 9)
np.divide / Division (6/3 = 2; 3/2 = 1)
np.floor_divide // Floor division (6//3 = 2; 3//2 = 1)
np.power ** Exponentiation (3**3 = 27; 2**3 = 8)
np.mod % Modulus/remainder(9%3=0; 9%4 = 1)
# +
# opérations arithmétiques
arr1 = np.random.randint(5, size=4)
arr2 = np.random.randint(5, size=4)
arr1, arr2
# -
# Ajouter des tableaux ensemble
arr1 + arr2
# Multiplication matricielle
arr1 * arr2
-arr1
arr1**arr2
import numpy as np
np.add(arr1, 2)
np.negative(arr1)
np.multiply(arr1, 3)
abs(arr1) #python command
np.absolute(arr1) #np command
np.add.reduce(arr1) # renvoie l'addition de tous les éléments
np.multiply.reduce(a) #produire de multiplier tous les éléments
np.add.accumulate(a) #afficher toutes les valeurs intermédiaires
np.multiply.outer(a,a) #créer une table de multiplication
# <div class="alert alert-warning">
# <h3>opérateurs trigonométrique; exponents, logarithms</h3></div>
exmpl = np.linspace(0, 4, 4)
exmpl
# +
sin, cos, tan, arcsin, arccos, arctan
np.sin(exmpl)
np.cos(exmpl)
np.tan(exmpl)
exp, exp2, power, log, log2, log10
np.exp(exmpl)
np.exp2(exmpl)
np.log(exmpl)
np.log2(exmpl)
np.power(exmpl, 2)
# -
np.power(exmpl, 2)
np.sin(exmpl)
# <div class="alert alert-warning">
# <h3>opérateurs sur les valuers manquantes</h3></div>
# <div class="alert alert-success">
# <li>pandas marque les données manquantes avec NaN (pas un nombre) ou aucun (méthode du masque)
# <li>NaN / None - ne sont pas disponibles pour tous les types de données
# <li>-9999 (valeur sentinelle)
# <li>la valeur sentinelle réduit la plage de valeurs valides pouvant être représentées et peut nécessiter une logique supplémentaire dans l'arithmétique CPU et GPU</div>
# +
# quand quelque chose manque, pandas fusionne la cellule avec NaN:
A = {'a':1, 'b':2}
B = {'b':3, 'c':4}
df_na = pandas.DataFrame([A, B])
# -
df_na
type(df_na['a'][0])
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <li>isnull() = générer un masque booléen indiquant les valeurs manquantes
# <li>notnull() = opposé à isnull()
# <li>dropna() = renvoie une version filtrée des données, les NaN sont supprimés
# <li>fillna() = renvoie une copie des données avec les valeurs manquantes renseignées ou imputées</div>
# -
# df_na.isnull()
df_na.isna()
df_na.notnull()
# df_na.notna()
df_na.dropna() # supprime les colonnes où au moins un élément est manquant
df_na['a'].isnull().values.any()
# df_na.a.isnull().values.any()
# +
#df_na['a']
# -
df_na['a'].fillna(0)
df_na['a'].fillna(3)
# +
#df_na.fillna(0)
# +
# plus d'informations ici:
https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.dropna.html
# -
# <div class="alert alert-info">
# <h3>Groupby: pandas.DataFrame.groupby</h3></div>
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <li> DataFrame.groupby(by=None, axis=0, level=None, as_index=True, sort=True, group_keys=True, squeeze=False, observed=False, **kwargs)
#
# <p>
#
# groupby s'effectue en 3 étapes:
#
# <li> <b>fractionnement</b> pour scinder et grouper un Cadre de données en fonction de la valeur de la clé spécifiée
# <li> <b>application</b> pour calculer une fonction (agrégat, transformation, filtrage, au sein des groupes individuels)
# <li> <b>combinaison</b> fusionne les résultats dans un tableau de sortie.
# -
planets.columns
planets.method.tolist()
planets.to_excel(r"C:\Users\Dylan\Downloads\tmp3.xls")
planets.groupby('method').mean()
planets.orbital_period[planets.method=='Astrometry'].mean()
planets.distance[planets.method=='Astrometry'].mean()
planets.groupby('method').groups.keys()
planets.groupby(['method','year']).first()
planets.groupby('method')['year'].describe()
# +
#dispatch:
planets.groupby('method')['year'].describe().unstack()
# -
# python commands:
# help(i)
sorted()
# <div class="alert alert-danger">
# <b>les commandes et codes à apprendre par cœur </b></div>
# + active=""
# STRINGS:
# ''.replace()
# ''.strip() ! Attention
# ''.rstrip() ! Attention
# ''.split()
# ''.find()
# ''.rfind()
# ''.upper()
# ''.lower()
# ''.isdigit()
# ''.isupper()
# ''.islower()
# ''.endswith()
# ''.startswith()
# ''.count()
# ''.join()
# ''.zfill()
# https://docs.python.org/3/library/stdtypes.html
# https://www.programiz.com/python-programming/methods/string
#
# LIST:
# ls.remove()
# ls.append()
# ls.pop
# ls.sort()
# ls.reverse
# ls+ls
# https://docs.python.org/3/library/stdtypes.html
#
# TUPLE:
# tup+tup
# https://docs.python.org/3/library/stdtypes.html
#
# DICTIONARY:
# d.copy()
# d.deepcopy()
# d.get(cle)
# d.pop()
# del d[cle]
# d.clear()
#
# COPY
# copy.deepcopy(d)
#
# PANDAS:
# df.loc, df.iloc, df.at
# df.copy(deep=True)
# df.set_index(NAME, inplace=True)
# df.describe()
# COMBINAISON:
# df_A.append(df_B)
# pandas.concat([df_A, df_B], keys=['rouge','bleue'], axis=0, sort=True)
# df_A.merge(df_B)
# df_A.join(df_B.set_index('name'), on='name')
# GROUPBY:
# df.groupby('COL').mean()
# df.distance[df.COL=='VAL'].mean()
# df.groupby('COL')['COL2'].describe().unstack()
# df.groupby(['COL' , decade])['COL1'].sum().unstack().fillna(0)
# NUMPY:
# np.reshape
# np.newaxis
# np.sort
# np.argsort
# np.concatenate
# np.vstack
# np.hstack
# np.split
# np.vsplit
# np.hsplit
#
#
# OPÉRATEURS:
# == != < > <= >=
# and or not
# is, is not
# in, not in
# & |
# astype()
# PANDAS:
# df[df.COL > int]
# df.loc[df.COL > int, ['COL1', 'COL2']].head()
# dfc[(dfc.COL>int) & (df.COL<int]
# count, first, last, mean, median, min, max, sum, std, var, mad, prod
# mean(), add(), sub(), substract(), mul(), multiply(), div(), divide()
# df.COL.mean()
# df['COL'] = df_qc['COL1'].subtract(df_qc['COL2'])
# NAN:
# isnull, notnull, dropna, fillna
# isnull().values.any()
# NUMPY:
# np.min
# np.max
# np.sum
# np.mean
# np.std
# np.any
# np.all
# np.percentile
# np.add
# np.subtract
# np.negative
# np.multiply
# np.divide
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-danger">
# <h3>Exercises</h3>
# </div>
# -
# <div class="alert alert-success">
# <li> Comment ajouter une variable à un tuple ?
# <li> Triez et affichez la liste
# <li> Ajoutez l’élément 12 à la liste et affichez la liste
# <li> Enlevez l’élément 38 et affichez la liste
# <li>convertir un tableau 1D en tableau 2D avec 2 lignes
# </ul>
# </div>
# <div class="alert alert-success">
# <li> Comment concaténer 2 listes ?
# <li> Comment concaténer un tuple et une liste ?
# <li> Quelles sont les commandes habituelles utilisées pour travailler avec une liste ?
# <li> Quelles sont les commandes pour travailler avec une chaîne ?
# </ul>
# </div>
# +
# Trouvez l'erreur
list1 = ['1','2','3', 4, '5', 6, 'c']
list2 = ['a','b','c','d']
list3 = ('e','f','g','h')
print(list1+list1['3'])
#print(list1[3]+list2)
#print(list1+list3)
# -
planets.head()
decade = 10 * (planets['year'] // 10)
decade
# <div class="alert alert-success">
# <li> Décrivez les étapes du code 'CODE GROUPBY EXERCISE'
# <li> Qu'est-ce que la fonction fillna() apportera au code final?
# </div>
# +
# CODE GROUPBY EXERCISE
decade = decade.astype(str) + 's'
decade.name = 'decade'
planets.groupby(['method' , decade])['number'].sum().unstack().fillna(0)
# + slideshow={"slide_type": "subslide"}
# Créez un cadre de données de pandas
# à partir des éléments ci-dessous:
mydict = dict(zip(mylist, myarr))
# +
# Combinez ls1 et dict1 pour former une base de données
ls1 = list('abcedfghijklmnopqrstuvwxyz')
dict1 = {i: [str(i)+'_'+str(i*2)] for i in range(1,len(ls1))}
# -
np.concatenate
# Créez un tableau numpy 3 × 3 de tous les True
import numpy as np
np.ones((2, 3), dtype=bool)
# obtenir les positions de 2 élèments dans un tableau numpy, où les éléments A et B correspondent
# joindre deux tableaux verticalement
# +
# réponses pour la quéstion dans chapitre 2
# version 1
d1 = {'Subj_ID':['001'],'score': [16], 'group': [2], 'condition': ['cognition']}
df_d = pandas.DataFrame(d1)
# version 2
d1 = {'Subj_ID':'001','score': 16, 'group': 2, 'condition': 'cognition'}
df_d = pandas.DataFrame(d1, index=[0])
| ch_cours/ch03_modifDonnee_combin_opers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot
matplotlib.style.use('fivethirtyeight')
# %matplotlib inline
def lkup(x):
if x == 's':
return 1000000000
elif x == 'ms':
return 1000000
elif x == 'µs':
return 1000
else:
return 1
# -
def fibb1(x):
if x == 0:
return 0
if x == 1:
return 1
return fibb1(x-1) + fibb1(x-2)
def fibb2(x):
x -= 1
x1 = 0
x2 = 1
for i in range(x):
x1 += x2
x1, x2 = x2, x1
return x2
# +
#for i in range(1, 40):
# %timeit fibb1(i)
# -
resStr = """198 ns ± 8.62 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
544 ns ± 12 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
976 ns ± 16.5 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.83 µs ± 34.7 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
3.12 µs ± 89.3 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
4.99 µs ± 276 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
8.28 µs ± 378 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
13.2 µs ± 82.3 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
21.6 µs ± 1.42 µs per loop (mean ± std. dev. of 7 runs, 10000 loops each)
34.8 µs ± 949 ns per loop (mean ± std. dev. of 7 runs, 10000 loops each)
57 µs ± 1.5 µs per loop (mean ± std. dev. of 7 runs, 10000 loops each)
92.1 µs ± 1.32 µs per loop (mean ± std. dev. of 7 runs, 10000 loops each)
160 µs ± 4.95 µs per loop (mean ± std. dev. of 7 runs, 10000 loops each)
245 µs ± 7.9 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
420 µs ± 31 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
680 µs ± 41.9 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
1.07 ms ± 24.6 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
1.7 ms ± 19.1 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
2.77 ms ± 147 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
4.5 ms ± 111 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
7.21 ms ± 181 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
13.9 ms ± 2.94 ms per loop (mean ± std. dev. of 7 runs, 100 loops each)
19 ms ± 696 µs per loop (mean ± std. dev. of 7 runs, 10 loops each)
29.9 ms ± 974 µs per loop (mean ± std. dev. of 7 runs, 10 loops each)
49.8 ms ± 2.43 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
80 ms ± 2.09 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
129 ms ± 1.88 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
211 ms ± 8.61 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
366 ms ± 42.5 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
561 ms ± 9.79 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
895 ms ± 21.7 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
1.46 s ± 59.4 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
2.44 s ± 118 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
3.72 s ± 48.2 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
6.51 s ± 391 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
10.3 s ± 164 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
15.9 s ± 332 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
25.5 s ± 182 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
41.5 s ± 301 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)"""
fibb1Data = [(float(x[0])*lkup(x[1]), float(x[3])*lkup(x[4])) for x in [y.split(' ') for y in resStr.split('\n')]]
# +
#for i in range(1, 40):
# %timeit fibb2(i)
# -
resStr = """470 ns ± 18.1 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
651 ns ± 33.4 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
704 ns ± 30.2 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
789 ns ± 35.2 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
838 ns ± 13.7 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
908 ns ± 26.3 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
941 ns ± 13.6 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.05 µs ± 28.3 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.14 µs ± 44.9 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.3 µs ± 68.2 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.27 µs ± 26.5 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.32 µs ± 35.6 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.37 µs ± 23 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.47 µs ± 19.6 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.58 µs ± 21.5 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.74 µs ± 85.4 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.74 µs ± 24 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.83 µs ± 34.2 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
1.92 µs ± 25.7 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
2.01 µs ± 37.6 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
2.1 µs ± 32.9 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
2.15 µs ± 98.9 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.26 µs ± 97.9 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.33 µs ± 85.3 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.4 µs ± 120 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.54 µs ± 87 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.65 µs ± 86.2 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.81 µs ± 145 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.1 µs ± 219 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.12 µs ± 269 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
2.95 µs ± 47.8 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.1 µs ± 139 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.23 µs ± 124 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.2 µs ± 138 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.28 µs ± 48.9 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.5 µs ± 72.6 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.51 µs ± 144 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
3.95 µs ± 438 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
4.05 µs ± 229 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)"""
fibb2Data = [(float(x[0])*lkup(x[1]), float(x[3])*lkup(x[4])) for x in [y.split(' ') for y in resStr.split('\n')]]
pd.DataFrame([x[0] for x in fibb1Data]).plot()
pd.DataFrame([x[0] for x in fibb2Data]).plot()
| WhyAlgorithmsMatter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Hands-on: Training and deploying Question Answering with BERT
# -
# Pre-trained language representations have been shown to improve many downstream NLP tasks such as question answering, and natural language inference. Devlin, Jacob, et al proposed BERT [1] (Bidirectional Encoder Representations from Transformers), which fine-tunes deep bidirectional representations on a wide range of tasks with minimal task-specific parameters, and obtained state- of-the-art results.
#
# In this tutorial, we will focus on adapting the BERT model for the question answering task on the SQuAD dataset. Specifically, we will:
#
# - understand how to pre-process the SQuAD dataset to leverage the learnt representation in BERT,
# - adapt the BERT model to the question answering task, and
# - load a trained model to perform inference on the SQuAD dataset
# ## Sagemaker configuration
#
# This notebook requires mxnet-cu101 >= 1.6.0b20191102, gluonnlp >= 0.8.1
# We can create a sagemaker notebook instance with the lifecycle configuration file: sagemaker-lifecycle.config
# +
# One time script
# # !bash sagemaker-lifecycle.config
# -
# !pip list | grep mxnet
# !pip list | grep gluonnlp
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Load MXNet and GluonNLP
#
# We first import the libraries:
# +
import argparse, collections, time, logging
import json
import os
import io
import copy
import random
import warnings
import numpy as np
import gluonnlp as nlp
import mxnet as mx
import bert
import qa_utils
from gluonnlp.data import SQuAD
from bert.model.qa import BertForQALoss, BertForQA
from bert.data.qa import SQuADTransform, preprocess_dataset
from bert.bert_qa_evaluate import get_F1_EM, predict, PredResult
# Hyperparameters
parser = argparse.ArgumentParser('BERT finetuning')
parser.add_argument('--epochs', type=int, default=3)
parser.add_argument('--batch_size', default=32)
parser.add_argument('--num_epochs', default=1)
parser.add_argument('--lr', default=5e-5)
args = parser.parse_args([])
epochs = args.epochs
batch_size = args.batch_size
num_epochs = args.num_epochs
lr = args.lr
# output_dir = args.output_dir
# if not os.path.exists(output_dir):
# os.mkdir(output_dir)
# test_batch_size = args.test_batch_size
# optimizer = args.optimizer
# accumulate = args.accumulate
# warmup_ratio = args.warmup_ratio
# log_interval = args.log_interval
# max_seq_length = args.max_seq_length
# doc_stride = args.doc_stride
# max_query_length = args.max_query_length
# n_best_size = args.n_best_size
# + [markdown] slideshow={"slide_type": "slide"}
# ## Inspect the SQuAD Dataset
# + [markdown] slideshow={"slide_type": "skip"}
# Then we take a look at the Stanford Question Answering Dataset (SQuAD). The dataset can be downloaded using the `nlp.data.SQuAD` API. In this tutorial, we create a small dataset with 3 samples from the SQuAD dataset for demonstration purpose.
#
# The question answering task on the SQuAD dataset is setup the following way. For each sample in the dataset, a context is provided. The context is usually a long paragraph which contains lots of information. Then a question asked based on the context. The goal is to find the text span in the context that answers the question in the sample.
# + slideshow={"slide_type": "fragment"}
full_data = nlp.data.SQuAD(segment='dev', version='1.1')
# loading a subset of the dev set of SQuAD
num_target_samples = 3
target_samples = [full_data[i] for i in range(num_target_samples)]
dataset = mx.gluon.data.SimpleDataset(target_samples)
print('Number of samples in the created dataset subsampled from SQuAD = %d'%len(dataset))
# -
target_samples[0]
# As we can see, the above example is the structure of the SQuAD dataset. Here, the question index is 2, the context index is 3, the answer index is 4, and the answer position index is 5.
# + slideshow={"slide_type": "subslide"}
question_idx = 2
context_idx = 3
answer_idx = 4
answer_pos_idx = 5
# + [markdown] slideshow={"slide_type": "skip"}
# Let's take a look at a sample from the dataset. In this sample, the question is about the location of the game, with a description about the Super Bowl 50 game as the context. Note that three different answer spans are correct for this question, and they start from index 403, 355 and 355 in the context respectively.
# + slideshow={"slide_type": "subslide"}
sample = dataset[2]
print('\nContext:\n')
print(sample[context_idx])
print("\nQuestion")
print(sample[question_idx])
print("\nCorrect Answer Spans")
print(sample[answer_idx])
print("\nAnswer Span Start Indices:")
print(sample[answer_pos_idx])
# + [markdown] slideshow={"slide_type": "skip"}
# ## Data Pre-processing for QA with BERT
#
# Recall that during BERT pre-training, it takes a sentence pair as the input, separated by the 'SEP' special token. For SQuAD, we can feed the context-question pair as the sentence pair input. To use BERT to predict the starting and ending span of the answer, we can add a classification layer for each token in the context texts, to predict if a token is the start or the end of the answer span.
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "skip"}
# In the next few code blocks, we will work on pre-processing the samples in the SQuAD dataset in the desired format with these special separators.
#
# -
# ### Get Pre-trained BERT Model
# + [markdown] slideshow={"slide_type": "skip"}
# First, let's use the *get_model* API in GluonNLP to get the model definition for BERT, and the vocabulary used for the BERT model. Note that we discard the pooler and classifier layers used for the next sentence prediction task, as well as the decoder layers for the masked language model task during the BERT pre-training phase. These layers are not useful for predicting the starting and ending indices of the answer span.
#
# The list of pre-trained BERT models available in GluonNLP can be found [here](http://gluon-nlp.mxnet.io/model_zoo/bert/index.html).
# + slideshow={"slide_type": "fragment"}
bert_model, vocab = nlp.model.get_model('bert_12_768_12',
dataset_name='book_corpus_wiki_en_uncased',
use_classifier=False,
use_decoder=False,
use_pooler=False,
pretrained=False)
with open('vocab.json', 'w') as f:
f.write(vocab.to_json())
# + [markdown] slideshow={"slide_type": "skip"}
# Note that there are several special tokens in the vocabulary for BERT. In particular, the `[SEP]` token is used for separating the sentence pairs, and the `[CLS]` token is added at the beginning of the sentence pairs. They will be used to pre-process the SQuAD dataset later.
# + slideshow={"slide_type": "fragment"}
print(vocab)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Tokenization
# + [markdown] slideshow={"slide_type": "skip"}
# The second step is to process the samples using the same tokenizer used for BERT, which is provided as the `BERTTokenizer` API in GluonNLP. Note that instead of word level and character level representation, BERT uses subwords to represent a word, separated `##`.
#
# In the following example, the word `suspending` is tokenized as two subwords (`suspend` and `##ing`), and `numerals` is tokenized as three subwords (`nu`, `##meral`, `##s`).
# + slideshow={"slide_type": "subslide"}
tokenizer = nlp.data.BERTTokenizer(vocab=vocab, lower=True)
tokenizer("as well as temporarily suspending the tradition of naming each Super Bowl game with Roman numerals")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Sentence Pair Composition
#
# With the tokenizer inplace, we are ready to process the question-context texts and compose sentence pairs. The functionality is available via the `SQuADTransform` API.
# + slideshow={"slide_type": "fragment"}
transform = bert.data.qa.SQuADTransform(tokenizer, is_pad=False, is_training=False, do_lookup=False)
dev_data_transform, _ = preprocess_dataset(dataset, transform)
logging.info('The number of examples after preprocessing:{}'.format(len(dev_data_transform)))
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's take a look at the sample after the transformation:
# + slideshow={"slide_type": "fragment"}
sample = dev_data_transform[2]
print('\nsegment type: \n' + str(sample[2]))
print('\ntext length: ' + str(sample[3]))
print('\nsentence pair: \n' + str(sample[1]))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Vocabulary Lookup
#
# Finally, we convert the transformed texts to subword indices, which are used to contructor NDArrays as the inputs to the model.
# + slideshow={"slide_type": "fragment"}
def vocab_lookup(example_id, subwords, type_ids, length, start, end):
indices = vocab[subwords]
return example_id, indices, type_ids, length, start, end
dev_data_transform = dev_data_transform.transform(vocab_lookup, lazy=False)
print(dev_data_transform[2][1])
# + [markdown] slideshow={"slide_type": "slide"}
# ## Model Definition
#
# After the data is processed, we can define the model that uses the representation produced by BERT for predicting the starting and ending positions of the answer span.
# + [markdown] slideshow={"slide_type": "slide"}
# We download a BERT model trained on the SQuAD dataset, prepare the dataloader.
# + slideshow={"slide_type": "fragment"}
net = BertForQA(bert_model)
ctx = mx.gpu(0)
ckpt = qa_utils.download_qa_ckpt()
net.load_parameters(ckpt, ctx=ctx)
batch_size = 1
dev_dataloader = mx.gluon.data.DataLoader(
dev_data_transform, batch_size=batch_size, shuffle=False)
# +
all_results = collections.defaultdict(list)
total_num = 0
for data in dev_dataloader:
example_ids, inputs, token_types, valid_length, _, _ = data
total_num += len(inputs)
batch_size = inputs.shape[0]
output = net(inputs.astype('float32').as_in_context(ctx),
token_types.astype('float32').as_in_context(ctx),
valid_length.astype('float32').as_in_context(ctx))
pred_start, pred_end = mx.nd.split(output, axis=2, num_outputs=2)
example_ids = example_ids.asnumpy().tolist()
pred_start = pred_start.reshape(batch_size, -1).asnumpy()
pred_end = pred_end.reshape(batch_size, -1).asnumpy()
for example_id, start, end in zip(example_ids, pred_start, pred_end):
all_results[example_id].append(PredResult(start=start, end=end))
# -
qa_utils.predict(dataset, all_results, vocab)
# ### Let's Train the Model
#
# Now we can put all the pieces together, and start fine-tuning the model with a few epochs.
# +
# net = BertForQA(bert=bert_model)
# nlp.utils.load_parameters(net, pretrained_bert_parameters, ctx=ctx,
# ignore_extra=True, cast_dtype=True)
net.span_classifier.initialize(init=mx.init.Normal(0.02), ctx=ctx)
net.hybridize(static_alloc=True)
loss_function = BertForQALoss()
loss_function.hybridize(static_alloc=True)
# -
# ## Deploy on SageMaker
#
# 1. Preparing functions for inference
# 2. Saving the model parameters
# 3. Building a docker container with dependencies installed
# 4. Launching a serving end-point with SageMaker SDK
# ### 1. Preparing functions for inference
#
# Two functions:
# 1. model_fn() to load model parameters
# 2. transform_fn() to run model inference given an input
# +
# %%writefile code/serve.py
import collections, json, logging, warnings
import multiprocessing as mp
from functools import partial
import gluonnlp as nlp
import mxnet as mx
from mxnet.gluon import Block, nn
import bert
from bert.data.qa import preprocess_dataset, SQuADTransform
import bert_qa_evaluate
class BertForQA(Block):
"""Model for SQuAD task with BERT.
The model feeds token ids and token type ids into BERT to get the
pooled BERT sequence representation, then apply a Dense layer for QA task.
Parameters
----------
bert: BERTModel
Bidirectional encoder with transformer.
prefix : str or None
See document of `mx.gluon.Block`.
params : ParameterDict or None
See document of `mx.gluon.Block`.
"""
def __init__(self, bert, prefix=None, params=None):
super(BertForQA, self).__init__(prefix=prefix, params=params)
self.bert = bert
with self.name_scope():
self.span_classifier = nn.Dense(units=2, flatten=False)
def forward(self, inputs, token_types, valid_length=None): # pylint: disable=arguments-differ
"""Generate the unnormalized score for the given the input sequences.
Parameters
----------
inputs : NDArray, shape (batch_size, seq_length)
Input words for the sequences.
token_types : NDArray, shape (batch_size, seq_length)
Token types for the sequences, used to indicate whether the word belongs to the
first sentence or the second one.
valid_length : NDArray or None, shape (batch_size,)
Valid length of the sequence. This is used to mask the padded tokens.
Returns
-------
outputs : NDArray
Shape (batch_size, seq_length, 2)
"""
bert_output = self.bert(inputs, token_types, valid_length)
output = self.span_classifier(bert_output)
return output
def get_all_results(net, vocab, squadTransform, test_dataset, ctx = mx.cpu()):
all_results = collections.defaultdict(list)
def _vocab_lookup(example_id, subwords, type_ids, length, start, end):
indices = vocab[subwords]
return example_id, indices, type_ids, length, start, end
dev_data_transform, _ = preprocess_dataset(test_dataset, squadTransform)
dev_data_transform = dev_data_transform.transform(_vocab_lookup, lazy=False)
dev_dataloader = mx.gluon.data.DataLoader(dev_data_transform, batch_size=1, shuffle=False)
for data in dev_dataloader:
example_ids, inputs, token_types, valid_length, _, _ = data
batch_size = inputs.shape[0]
output = net(inputs.astype('float32').as_in_context(ctx),
token_types.astype('float32').as_in_context(ctx),
valid_length.astype('float32').as_in_context(ctx))
pred_start, pred_end = mx.nd.split(output, axis=2, num_outputs=2)
example_ids = example_ids.asnumpy().tolist()
pred_start = pred_start.reshape(batch_size, -1).asnumpy()
pred_end = pred_end.reshape(batch_size, -1).asnumpy()
for example_id, start, end in zip(example_ids, pred_start, pred_end):
all_results[example_id].append(bert_qa_evaluate.PredResult(start=start, end=end))
return(all_results)
def _test_example_transform(test_examples):
"""
Change test examples to a format like SQUAD data.
Parameters
----------
test_examples: a list of (question, context) tuple.
Example: [('Which NFL team represented the AFC at Super Bowl 50?',
'Super Bowl 50 was an American football game ......),
('Where did Super Bowl 50 take place?',,
'Super Bowl 50 was ......),
......]
Returns
----------
test_examples_tuples : a list of SQUAD tuples
"""
test_examples_tuples = []
i = 0
for test in test_examples:
question, context = test[0], test[1] # test.split(" [CONTEXT] ")
tup = (i, "", question, context, [], [])
test_examples_tuples.append(tup)
i += 1
return(test_examples_tuples)
def model_fn(model_dir = "", params_path = "bert_qa-7eb11865.params"):
"""
Load the gluon model. Called once when hosting service starts.
:param: model_dir The directory where model files are stored.
:return: a Gluon model, and the vocabulary
"""
bert_model, vocab = nlp.model.get_model('bert_12_768_12',
dataset_name='book_corpus_wiki_en_uncased',
use_classifier=False,
use_decoder=False,
use_pooler=False,
pretrained=False)
net = BertForQA(bert_model)
if len(model_dir) > 0:
params_path = model_dir + "/" +params_path
net.load_parameters(params_path, ctx=mx.cpu())
tokenizer = nlp.data.BERTTokenizer(vocab, lower=True)
transform = SQuADTransform(tokenizer, is_pad=False, is_training=False, do_lookup=False)
return net, vocab, transform
def transform_fn(model, input_data, input_content_type=None, output_content_type=None):
"""
Transform a request using the Gluon model. Called once per request.
:param model: The Gluon model and the vocab
:param dataset: The request payload
Example:
## (example_id, [question, content], ques_cont_token_types, valid_length, _, _)
(2,
'56be4db0acb8001400a502ee',
'Where did Super Bowl 50 take place?',
'Super Bowl 50 was an American football game to determine the champion of the National
Football League (NFL) for the 2015 season. The American Football Conference (AFC)
champion Denver Broncos defeated the National Football Conference (NFC) champion
Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played
on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara,
California. As this was the 50th Super Bowl, the league emphasized the "golden
anniversary" with various gold-themed initiatives, as well as temporarily suspending
the tradition of naming each Super Bowl game with Roman numerals (under which the
game would have been known as "Super Bowl L"), so that the logo could prominently
feature the Arabic numerals 50.',
['Santa Clara, California', "Levi's Stadium", "Levi's Stadium
in the San Francisco Bay Area at Santa Clara, California."],
[403, 355, 355])
:param input_content_type: The request content type, assume json
:param output_content_type: The (desired) response content type, assume json
:return: response payload and content type.
"""
net, vocab, squadTransform = model
data = json.loads(input_data)
test_examples_tuples = _test_example_transform(data)
test_dataset = mx.gluon.data.SimpleDataset(test_examples_tuples) # [tup]
all_results = get_all_results(net, vocab, squadTransform, test_dataset, ctx=mx.cpu())
all_predictions = collections.defaultdict(list) # collections.OrderedDict()
data_transform = test_dataset.transform(squadTransform._transform)
for features in data_transform:
f_id = features[0].example_id
results = all_results[f_id]
prediction, nbest = bert_qa_evaluate.predict(
features=features,
results=results,
tokenizer=nlp.data.BERTBasicTokenizer(vocab))
nbest_prediction = []
for i in range(3):
nbest_prediction.append('%.2f%% \t %s'%(nbest[i][1] * 100, nbest[i][0]))
all_predictions[f_id] = nbest_prediction
response_body = json.dumps(all_predictions)
return response_body, output_content_type
# -
# ### 2. Saving the model parameters
# +
## save parameters, model definition and vocabulary in a zip file
# output_dir = "model_outputs"
# if not os.path.exists(output_dir):
# os.mkdir(output_dir)
with open('vocab.json', 'w') as f:
f.write(vocab.to_json())
import tarfile
with tarfile.open("model.tar.gz", "w:gz") as tar:
# tar.add("Dockerfile")
tar.add("code/serve.py")
tar.add("bert/data/qa.py")
tar.add("bert_qa_evaluate.py")
tar.add("bert_qa-7eb11865.params")
tar.add("vocab.json")
# +
## test
my_test_example_0 = ('Which NFL team represented the AFC at Super Bowl 50?',
'Super Bowl 50 was an American football game to determine the champion of the National Football League (NFL) for the 2015 season. The American Football Conference (AFC) champion Denver Broncos defeated the National Football Conference (NFC) champion Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara, California. As this was the 50th Super Bowl, the league emphasized the "golden anniversary" with various gold-themed initiatives, as well as temporarily suspending the tradition of naming each Super Bowl game with Roman numerals (under which the game would have been known as "Super Bowl L"), so that the logo could prominently feature the Arabic numerals 50.')
my_test_example_1 = ('Where did Super Bowl 50 take place?',
'Super Bowl 50 was an American football game to determine the champion of the National Football League (NFL) for the 2015 season. The American Football Conference (AFC) champion Denver Broncos defeated the National Football Conference (NFC) champion Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara, California. As this was the 50th Super Bowl, the league emphasized the "golden anniversary" with various gold-themed initiatives, as well as temporarily suspending the tradition of naming each Super Bowl game with Roman numerals (under which the game would have been known as "Super Bowl L"), so that the logo could prominently feature the Arabic numerals 50.')
my_test_example_concat_0 = " [CONTEXT] ".join(my_test_example_0)
my_test_example_concat_1 = " [CONTEXT] ".join(my_test_example_1)
my_test_examples = (my_test_example_0, my_test_example_1)
## prepare test examples
# with open('my_test_examples.json', 'w') as f:
# json.dump(my_test_examples, f)
# mymodel = model_fn()
# transform_fn(mymodel, my_test_examples)
# -
# ### 3. Building a docker container with dependencies installed
#
# Let's prepare a docker container with all the dependencies required for model inference. Here we build a docker container based on the SageMaker MXNet inference container, and you can find the list of all available inference containers at https://docs.aws.amazon.com/sagemaker/latest/dg/pre-built-containers-frameworks-deep-learning.html
#
# Here we use local mode for demonstration purpose. To deploy on actual instances, you need to login into AWS elastic container registry (ECR) service, and push the container to ECR.
#
# ```
# docker build -t $YOUR_EDR_DOCKER_TAG . -f Dockerfile
# $(aws ecr get-login --no-include-email --region $YOUR_REGION)
# docker push $YOUR_EDR_DOCKER_TAG
# ```
# +
# %%writefile Dockerfile
ARG REGION
FROM 763104351884.dkr.ecr.$REGION.amazonaws.com/mxnet-inference:1.4.1-gpu-py3
RUN pip install --upgrade --user --pre 'mxnet-mkl' 'https://github.com/dmlc/gluon-nlp/tarball/v0.9.x'
RUN pip list | grep mxnet
COPY *.py /opt/ml/model/code/
COPY bert/data/qa.py /opt/ml/model/code/bert/data/
COPY bert/bert_qa_evaluate.py /opt/ml/model/code/bert/
# -
# !export REGION=$(wget -qO- http://169.254.169.254/latest/meta-data/placement/availability-zone) &&\
# docker build --no-cache --build-arg REGION=${REGION::-1} -t my-docker:inference . -f Dockerfile
# ### 4. Launching a serving end-point with SageMaker SDK
#
# We create a MXNet model which can be deployed later, by specifying the docker image, and entry point for the inference code. If serve.py does not work, use dummy_hosting_module.py for debugging purpose.
import sagemaker
from sagemaker.mxnet.model import MXNetModel
sagemaker_model = MXNetModel(model_data='file:///home/ec2-user/SageMaker/ako2020-bert/tutorial/model.tar.gz',
image='my-docker:inference', # docker images
role=sagemaker.get_execution_role(),
py_version='py3', # python version
entry_point='serve.py',
source_dir='.')
# We use 'local' mode to test our deployment code, where the inference happens on the current instance.
# If you are ready to deploy the model on a new instance, change the `instance_type` argument to values such as `ml.c4.xlarge`.
#
# Here we use 'local' mode for testing, for real instances use c5.2xlarge, p2.xlarge, etc.
predictor = sagemaker_model.deploy(initial_instance_count=1, instance_type='local')
output = predictor.predict(my_test_examples)
print('\nPrediction output: {}\n\n'.format(output))
# ### Clean Up
#
# Remove the endpoint after we are done.
predictor.delete_endpoint()
# ### Resources
#
# Amazon SageMaker Developer Guide
# https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-dg.pdf
| nlp/resources/temp/question_answering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from PIL import Image
from functools import reduce, partial
from multiprocessing import Pool
from matplotlib import pyplot as mp
from mpmath import mp
from mpmath import pi as Pi
from mpmath import sin as Sin
from mpmath import asin as Asin
from mpmath import sqrt as Sqrt
import matplotlib.image as mpimg
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# <NAME>, in his paper "Real numbers, data science and chaos: How to fit any dataset with a single parameter" shows how any dataset of any modality (time-series, images, sound...) can be approximated by a well-behaved (continuous, differentiable...) scalar function with a single real-valued parameter. The paper can be found here.
# All the code and concepts in this Jupyter notebook are based on the work of <NAME>.
print('https://arxiv.org/pdf/1904.12320.pdf')
# The equation is the following:
#
# $$ f_{\alpha \left ( X \right )}=sin^{^{2}}(2^{x\tau }arcsin\sqrt{\alpha })\$$
color = './images/color.png'
b_and_w = './images/blackandwhite.png'
image_color = mpimg.imread(color)
image_bw = mpimg.imread(b_and_w)
image_pil = Image.open(b_and_w)
# +
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(15, 7))
axes[0].imshow(image_color)
axes[1].imshow(image_bw)
fig.tight_layout()
# -
image_color
image_bw
image = image_pil.convert('L').point(lambda x : 0 if x > 100 else 255).convert('1')
img = np.rot90(np.asarray(image), k=3)
width, height = img.shape
possiblePositions = list(zip(*np.where(img)))
# +
sampleAndScale = lambda gr: gr['y'].sample().values[0] / height
data = pd.DataFrame(possiblePositions, columns=['x', 'y']).groupby('x').apply(sampleAndScale)
data = data.reindex(np.arange(0, width), fill_value=-10)
data = data[data > 0]
numbPoints = len(data)
data.index = range(numbPoints)
# -
phiInv = lambda z: np.arcsin(np.sqrt(z)) / (2.0 * np.pi)
decimalToBinary_phiInv = lambda z: decimalToBinary(phiInv(z))
phi = lambda theta: Sin(theta * Pi * 2.0) ** 2
dyadicMap = lambda x : (2 * x) % 1
tau = 12
def decimalToBinary(decimalInitial, targetBinaryPrecision = tau):
return reduce(lambda acc, _: [dyadicMap(acc[0]), acc[1] + ('0' if acc[0] < 0.5 else '1')],
range(targetBinaryPrecision),
[decimalInitial, ''])[1]
def binaryToDecimal(binaryInitial):
return reduce(lambda acc, val: acc + int(val[1]) / mp.power(2, (val[0] + 1)),
enumerate(binaryInitial),
mp.mpf(0.0))
def binaryReducer(val):
return int(val[1]) / mp.power(2, (val[0] + 1))
def binaryToDecFaster(binaryInitial):
with Pool(8) as p:
tt = p.map(binaryReducer, enumerate(binaryInitial))
res = mp.mpf(0)
for _ in tt:
res += _
return res
def dyadicDecoder(decimalInitial, k):
return (2 ** (k * tau) * decimalInitial) % 1
def logisticDecoder(decimalInitial, k):
return float(Sin(2 ** (k * tau) * Asin(Sqrt(decimalInitial))) ** 2)
def findInitialCondition(trainData):
conjugateInitial_binary = ''.join(map(decimalToBinary_phiInv, trainData))
necessaryPrecision = len(conjugateInitial_binary)
assert tau * len(trainData) == necessaryPrecision
# data is passed through sequentially so no need to worry
# plus, all samples have the same size anyway
# to be safe, these global settings should be handled more carefully with context managers
mp.prec = necessaryPrecision
print('significance = %d bits ; %d digits (base-10) ; ratio = %.3f\n' % (mp.prec, mp.dps, mp.prec / mp.dps))
# conjugateInitial = binaryToDecimal(conjugateInitial_binary)
conjugateInitial = binaryToDecFaster(conjugateInitial_binary)
decimalInitial = phi(conjugateInitial)
return decimalInitial
def generateData(decimalInitial, howManyPoints):
p_logisticDecoder = partial(logisticDecoder, decimalInitial)
with Pool(8) as p:
decodedValues = p.map(p_logisticDecoder, range(howManyPoints))
return decodedValues
decimalInitial = findInitialCondition(data)
decodedValues = generateData(decimalInitial, len(data))
decimalInitial
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(10, 3))
axes[0].imshow(image_color)
axes[0].set_title('Color', fontsize=20)
axes[1].imshow(image_bw)
axes[1].set_title('Black and White Contour', fontsize=20)
axes[2].scatter(range(len(decodedValues)), decodedValues, color='black', s=2)
axes[2].set_title('One parameter fit', fontsize=20)
fig.tight_layout()
| Single Parameter Fit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sayakpaul/Handwriting-Recognizer-in-Keras/blob/main/Initial_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="D6B83kI1VDpQ"
# ## Reference
# - Keras example on [Captcha OCR](https://keras.io/examples/vision/captcha_ocr/)
# + [markdown] id="BOJ3n-LZbxls"
# ## Dataset collection
# + id="w1MdZdnrJq5l"
# !wget -q https://github.com/sayakpaul/Handwriting-Recognizer-in-Keras/releases/download/v1.0.0/IAM_Words.zip
# !unzip -qq IAM_Words.zip
# + id="jo2n2mBxL3N6"
# !mkdir data
# !mkdir data/words
# !tar -xf IAM_Words/words.tgz -C data/words
# !mv IAM_Words/words.txt data
# + id="OwwnL7cwL8q0" colab={"base_uri": "https://localhost:8080/"} outputId="42be5c86-2b9e-4026-80c6-86a37a638ded"
# !head -20 data/words.txt
# + [markdown] id="T5Br7NfLbz_E"
# ## Imports
# + id="JdA5uc6ATAWx" colab={"base_uri": "https://localhost:8080/"} outputId="c8be0648-3d44-43af-835c-589dcc1a335e"
# !pip install -q -U imgaug
# + id="AzAn4MGOMGyP"
from imutils import paths
from tqdm.notebook import tqdm
from itertools import groupby
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import imgaug
import cv2
import os
np.random.seed(42)
tf.random.set_seed(42)
# + [markdown] id="eFUUdA4Ib1q4"
# ## Dataset preparation
# + id="5nw0wiDWssNI"
BASE_IMAGE_PATH = os.path.join("data", "words")
# + id="M0OnjVA6MQTu" colab={"base_uri": "https://localhost:8080/"} outputId="fa95e066-14e3-4abb-d1c9-f70f43d1ec6a"
# Image path: part1-part2-part3 --> part1/part1-part2/part1-part2-part3.png
# The above format DOES NOT include the base path which is "data/words" in
# this case.
all_images = list(paths.list_images(BASE_IMAGE_PATH))
all_images[:5]
# + id="bWcMSzy9NCdp" colab={"base_uri": "https://localhost:8080/", "height": 83, "referenced_widgets": ["863bfd50e00a43909f7e36186bcb0e45", "3d1abccd6bf04921a2f2d5a06e485efe", "3cc7c624c1ca4d958b926e040bc91761", "c2a3f91652e443c59e180ae7028a5490", "0053f7f84f754596963b7544b8080d06", "<KEY>", "062bdcf2881b4f54a8750c8c30190554", "6433d6803ba84d37bfd4abe39c9c1a9a"]} outputId="0ba72633-3d43-4e12-930b-d95c8c208209"
words_list = []
words = open('data/words.txt', 'r').readlines()
for line in tqdm(words):
if line[0]=='#':
continue
if line.split(" ")[1]!="err": # We don't need to deal with errored entries.
words_list.append(line)
len(words_list)
# + id="2MkkUDt6SCr5"
np.random.shuffle(words_list)
# + [markdown] id="Q_1JJr-joiMb"
# ### Prepare the splits (90:5:5)
# + id="OlWJ8FP2Rpht" colab={"base_uri": "https://localhost:8080/"} outputId="4611f0eb-c891-45d9-c691-fbe5715edf4a"
split_idx = int(0.9 * len(words_list))
train_samples = words_list[:split_idx]
test_samples = words_list[split_idx:]
val_split_idx = int(0.5 * len(test_samples))
validation_samples = test_samples[:val_split_idx]
test_samples = test_samples[val_split_idx:]
assert len(words_list) == len(train_samples) + len(validation_samples) + len(test_samples)
print(f"Total training samples: {len(train_samples)}")
print(f"Total validation samples: {len(validation_samples)}")
print(f"Total test samples: {len(test_samples)}")
# + id="c9oBlfUkNGVz" colab={"base_uri": "https://localhost:8080/"} outputId="990d7f0b-98ec-49ee-dcd2-f782f1c7c941"
# Since the labels start appearing after eighth index we use that
# to retrieve the grounth-truth labels. Remember indexing starts from
# zero in Python.
start_idx = 8
train_words = [line.split(' ')[start_idx:][0].strip() for line in train_samples]
max_label_len = max([len(str(text)) for text in train_words])
print(f"Maximum label length: {max_label_len}")
padding_method = "post"
padding_token = 99
tokenizer = tf.keras.preprocessing.text.Tokenizer(filters="\n", char_level=True)
tokenizer.fit_on_texts(train_words)
tokenized_words = tokenizer.texts_to_sequences(train_words)
padded_train_words = tf.keras.preprocessing.sequence.pad_sequences(tokenized_words,
maxlen=max_label_len,
padding=padding_method,
value=padding_token)
# Maximum sequence length is 4, hence a word is padded to that length.
print(f"Integer representation of a word: {padded_train_words[0]}")
# + id="6aBAK36wos4G" colab={"base_uri": "https://localhost:8080/"} outputId="655dfad9-e7d4-4695-87f8-94bdc83d737c"
# Unique characters.
len(tokenizer.word_index)
# + id="nU9pG8p8nRd_"
# View some word index mappings.
def process_word(word):
processed_word = []
for i in word:
if i != 99:
processed_word.append(tokenizer.index_word[i])
return "".join(processed_word)
def view_sample_mappings(sample_padded_words):
for t in sample_padded_words:
word = process_word(t)
print (f"{t.tolist()}----> {word}")
# + id="V5Z3bGvk7n9e" colab={"base_uri": "https://localhost:8080/"} outputId="aa9f6af3-f929-466a-e548-ca7ea28d86e2"
view_sample_mappings(padded_train_words[:15])
# + id="FhOEwqXcpMPH"
valid_words = [line.split(' ')[start_idx:][0].strip() for line in validation_samples]
tokenized_valid_words = tokenizer.texts_to_sequences(valid_words)
padded_valid_words = tf.keras.preprocessing.sequence.pad_sequences(tokenized_valid_words,
maxlen=max_label_len,
padding=padding_method,
value=padding_token)
# + id="f1d8444YpgQa"
test_words = [line.split(' ')[start_idx:][0].strip() for line in test_samples]
tokenized_test_words = tokenizer.texts_to_sequences(test_words)
padded_test_words = tf.keras.preprocessing.sequence.pad_sequences(tokenized_test_words,
maxlen=max_label_len,
padding=padding_method,
value=padding_token)
# + id="RROe32g7GD6k"
# Credit: https://github.com/githubharald/SimpleHTR/blob/master/src/SamplePreprocessor.py
def distortion_free_resize(image, img_size):
# Target size and current image size.
(wt, ht) = img_size
(h, w) = image.shape
# Compute the individual resolution scales and take
# the maximum between them.
fx = w / wt
fy = h / ht
f = max(fx, fy)
# Compute the new image size such that the aspect ratio is respected.
new_size = (max(min(wt, int(w / f)), 1), max(min(ht, int(h / f)), 1))
# First, resize the image to this newly computed size. Then
# copy its pixels over appropriately to another blank image
# having the target size.
image = cv2.resize(image, new_size)
target = np.ones([ht, wt]) * 255
target[0:new_size[1], 0:new_size[0]] = image
# Tranpose to (w, h) format.
image = cv2.transpose(target)
return image
# + id="f5XfPrBuQdGd"
IMG_WIDTH = 128
IMG_HEIGHT = 32
def process_image(img_path, img_size=(IMG_WIDTH, IMG_HEIGHT)):
# Read image in grayscale mode.
image = cv2.imread(img_path, 0)
# Scale pixel values to [0, 1].
image = image.astype("float32")/255
# Resize image.
image = distortion_free_resize(image, img_size)
# Add channel otherwise Conv2D won't be compatible.
image = np.expand_dims(image, axis=-1)
return image
# + id="HBVTcwqBZMnX"
def prepare_images(samples):
images = np.zeros(shape=(len(samples), IMG_WIDTH, IMG_HEIGHT, 1))
for (i, file_line) in enumerate(tqdm(samples)):
line_split = file_line.strip()
line_split = line_split.split(" ")
# Each line split will have this format for the corresponding image:
# part1/part1-part2/part1-part2-part3.png
image_name = line_split[0]
partI = image_name.split("-")[0]
partII = image_name.split("-")[1]
img_path = os.path.join(BASE_IMAGE_PATH, partI,
partI + "-" + partII,
image_name + ".png"
)
if os.path.getsize(img_path):
preprocessed_image = process_image(img_path)
images[i] = preprocessed_image
return images
# + id="E806c2cVa2mI" colab={"base_uri": "https://localhost:8080/", "height": 164, "referenced_widgets": ["dbf74a73d9824a288c69d6d1e0f3ab96", "e5ca3abd1b634507aa115c823e8cc660", "d7b83d9b73dd42c9ac876217de6181ba", "1783481b39a14cda842025ea2aba6be7", "361567a72c9e4750adc15c850186b313", "e1ece62e03cc4c5a901e51a42bc60d2b", "87a354d4275b47cda0f09a224bce94e2", "<KEY>", "d359280a815b445aa70af200057fb4a0", "d5a78f3ea43a42a9814f1270ae7bdac5", "<KEY>", "<KEY>", "<KEY>", "e4083ace92034ceb8967aa20c78fd545", "1638930591bf4a1bb280dd4039aa755b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "5fef5c3fa26c4e748a2f67e57d2b2d76", "<KEY>", "d8c9ff19908a488098f52232afefbf83", "e7a3d9a582534898a53130abd42ca2ac", "a8e7018577444f0fa90f0de6a17f616f"]} outputId="6fea8a9c-feee-4b3f-b47d-bf88f1dfc2d0"
train_images = prepare_images(train_samples)
validation_images = prepare_images(validation_samples)
test_images = prepare_images(test_samples)
# + id="9cv3QeQ-SGJS"
# Construct TensorFlow datasets.
BATCH_SIZE = 64
AUTOTUNE = tf.data.AUTOTUNE
augmenter = imgaug.augmenters.Sequential([
imgaug.augmenters.GammaContrast(gamma=(0.25, 3.0)),
imgaug.augmenters.Sometimes(
0.3,
imgaug.augmenters.GaussianBlur(sigma=(0, 0.5))
)
])
def augment(images):
return augmenter(images=images.numpy())
def make_dicts(images, labels):
return {"images": images, "labels": labels}
def make_datasets(images, labels, training=True):
dataset = tf.data.Dataset.from_tensor_slices((images, labels))
if training:
dataset = dataset.shuffle(BATCH_SIZE * 25)
dataset = dataset.batch(BATCH_SIZE)
if training:
dataset = dataset.map(
lambda x, y: (tf.py_function(augment, [x], [tf.float32])[0], y),
num_parallel_calls=AUTOTUNE
)
dataset = dataset.map(make_dicts).prefetch(AUTOTUNE)
return dataset
train_dataset = make_datasets(train_images, padded_train_words)
validation_dataset = make_datasets(validation_images, padded_valid_words, False)
test_dataset = make_datasets(test_images, padded_test_words, False)
# + id="doV5CtiBVgMQ"
def plot_samples(images, labels):
_, ax = plt.subplots(4, 4, figsize=(12, 12))
for i in range(16):
img = (images[i] * 255).numpy().astype("uint8")
label = process_word(labels[i].numpy())
ax[i // 4, i % 4].imshow(img.squeeze(-1).T, cmap="gray")
ax[i // 4, i % 4].set_title(label)
ax[i // 4, i % 4].axis("off")
plt.show()
# + id="SvslZTljVtQ6" colab={"base_uri": "https://localhost:8080/", "height": 594} outputId="c28c0e90-e091-4396-d4b1-48e697d9f978"
batch = next(iter(train_dataset))
plot_samples(batch["images"], batch["labels"])
# + [markdown] id="7KQ8ueDJb46h"
# ## Model building
# + id="QZRjUkjyWUXl" colab={"base_uri": "https://localhost:8080/"} outputId="38ae65d6-9ddf-4493-fe28-dc163090ea02"
class CTCLayer( tf.keras.layers.Layer):
def __init__(self, name=None):
super().__init__(name=name)
self.loss_fn = tf.keras.backend.ctc_batch_cost
def call(self, y_true, y_pred):
batch_len = tf.cast(tf.shape(y_true)[0], dtype="int64")
input_length = tf.cast(tf.shape(y_pred)[1], dtype="int64")
label_length = tf.cast(tf.shape(y_true)[1], dtype="int64")
input_length = input_length * tf.ones(shape=(batch_len, 1), dtype="int64")
label_length = label_length * tf.ones(shape=(batch_len, 1), dtype="int64")
loss = self.loss_fn(y_true, y_pred, input_length, label_length)
self.add_loss(loss)
# At test time, just return the computed predictions
return y_pred
def build_model():
# Inputs to the model
input_img = tf.keras.layers.Input(
shape=(IMG_WIDTH, IMG_HEIGHT, 1), name="images")
labels = tf.keras.layers.Input(name="labels", shape=(None,))
# First conv block
x = tf.keras.layers.Conv2D(
32,
(3, 3),
activation="relu",
kernel_initializer="he_normal",
padding="same",
name="Conv1",
)(input_img)
x = tf.keras.layers.MaxPooling2D((2, 2), name="pool1")(x)
# Second conv block
x = tf.keras.layers.Conv2D(
64,
(3, 3),
activation="relu",
kernel_initializer="he_normal",
padding="same",
name="Conv2",
)(x)
x = tf.keras.layers.MaxPooling2D((2, 2), name="pool2")(x)
# We have used two max pool with pool size and strides 2.
# Hence, downsampled feature maps are 4x smaller. The number of
# filters in the last layer is 64. Reshape accordingly before
# passing the output to the RNN part of the model
new_shape = ((IMG_WIDTH // 4), (IMG_HEIGHT // 4) * 64)
x = tf.keras.layers.Reshape(target_shape=new_shape, name="reshape")(x)
x = tf.keras.layers.Dense(64, activation="relu", name="dense1")(x)
x = tf.keras.layers.Dropout(0.2)(x)
# RNNs
x = tf.keras.layers.Bidirectional( tf.keras.layers.LSTM(128, return_sequences=True, dropout=0.25))(x)
x = tf.keras.layers.Bidirectional( tf.keras.layers.LSTM(64, return_sequences=True, dropout=0.25))(x)
# Output layer (the tokenizer is char-level)
# +2 recommendation came from here - https://github.com/MaybeShewill-CV/CRNN_Tensorflow/issues/69#issuecomment-383992527
x = tf.keras.layers.Dense(len(tokenizer.word_index) + 2, activation="softmax", name="dense2")(x)
# Add CTC layer for calculating CTC loss at each step
output = CTCLayer(name="ctc_loss")(labels, x)
# Define the model
model = tf.keras.models.Model(
inputs=[input_img, labels], outputs=output, name="handwriting_recognizer"
)
# Optimizer
opt = tf.keras.optimizers.Adam()
# Compile the model and return
model.compile(optimizer=opt)
return model
# Get the model
model = build_model()
model.summary()
# + [markdown] id="GUCgn5jJb7hP"
# ## Model training and inference
# + id="zUwk56nCWxxQ" colab={"base_uri": "https://localhost:8080/"} outputId="e4ac1ead-2c12-4dda-b908-3a4591a6af15"
epochs = 100
early_stopping_patience = 10
early_stopping = tf.keras.callbacks.EarlyStopping(
monitor="val_loss", patience=early_stopping_patience, restore_best_weights=True
)
# Train the model
model = build_model()
history = model.fit(
train_dataset,
validation_data=validation_dataset,
epochs=epochs,
callbacks=[early_stopping],
)
# + id="aakOMS1c4c1I" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="ed5fe7dd-ccc7-41bd-bd4f-f75d0a79296c"
plt.plot(history.history["loss"], label="train_loss")
plt.plot(history.history["val_loss"], label="val_loss")
plt.xlabel("Epochs")
plt.ylabel("CTC Loss")
plt.title("Model's Loss over Epochs")
plt.legend()
plt.show()
# + id="-yyn2luJQ1OU" colab={"base_uri": "https://localhost:8080/"} outputId="179b2264-fbe1-431b-8030-720f726a022e"
# Get the prediction model by extracting layers till the output layer.
prediction_model = tf.keras.models.Model(
model.get_layer(name="images").input, model.get_layer(name="dense2").output
)
prediction_model.summary()
# + id="0n-ZPN5sRLui"
# Reference: https://github.com/rajesh-bhat/spark-ai-summit-2020-text-extraction/blob/master/CRNN_CTC_wandb.ipynb
def ctc_decoder(predictions):
'''
input: given batch of predictions from text rec model
output: return lists of raw extracted text
'''
text_list = []
pred_indcies = np.argmax(predictions, axis=2)
for i in range(pred_indcies.shape[0]):
ans = ""
## merge repeats
merged_list = [k for k,_ in groupby(pred_indcies[i])]
## remove blanks
for p in merged_list:
if p != len(tokenizer.index_word) + 1:
ans += tokenizer.index_word[int(p)]
text_list.append(ans)
return text_list
# + id="JI5nbx2xRm7R" colab={"base_uri": "https://localhost:8080/", "height": 594} outputId="c2332c59-b7fd-4a9d-ec96-8e06912dbd91"
# Let's check results on some test samples.
for batch in test_dataset.take(1):
batch_images = batch["images"]
batch_labels = batch["labels"]
preds = prediction_model.predict(batch_images)
pred_texts = ctc_decoder(preds)
fig , ax = plt.subplots(4, 4, figsize=(12, 12))
for i in range(16):
img = (batch_images[i, :, :, 0] * 255).numpy().astype(np.uint8)
title = f"Prediction: {pred_texts[i]}"
ax[i // 4, i % 4].imshow(img.T, cmap="gray")
ax[i // 4, i % 4].set_title(title)
ax[i // 4, i % 4].axis("off")
plt.show()
| Initial_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
#
# * y와 한 개 이상의 독립 변수 (또는 설명 변수) X와의 선형 상관 관계를 모델링하는 회귀분석 기법이다. 한 개의 설명 변수에 기반한 경우에는 단순 선형 회귀, 둘 이상의 설명 변수에 기반한 경우에는 다중 선형 회귀라고 한다. [참고: 위키피디아](https://ko.wikipedia.org/wiki/선형_회귀)
#
# $$y_{\textrm{pred}} = \boldsymbol{W}^{\top}\boldsymbol{x} + b$$
#
# * $\boldsymbol{x} = [x_{1}, x_{2}, \cdots, x_{d}]$
# * $\boldsymbol{W} = [w_{1}, w_{2}, \cdots, w_{d}]$
# * Loss function: $\mathcal{L} = \sum^{N} (y_{\textrm{pred}} - y)^{2}$
# ## Training Pseudo Code
#
# ```python
# for epoch in max_epochs: # 1 epoch: 모든 데이터(N)를 한번 학습 시켰을 때
# for step in num_batches: # num_batches = int(data_size / batch_size)
# 1. sampling mini-batches with batch_size
# 1-1. data augmentation (필요하면)
# 2. calculate the logits # logits = f(x)
# 3. calculate the loss # loss = loss(logits, labels)
# 4. calculate the gradient with respect to weights
# 5. update weights
# ```
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import time
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
sess_config = tf.ConfigProto(gpu_options=tf.GPUOptions(allow_growth=True))
os.environ["CUDA_VISIBLE_DEVICES"]="0"
# -
# ## Phase 1. Build a model
# ### Make data
# +
np.random.seed(219)
_a = 3
_b = -3
N = 200
data_x = np.random.uniform(low=0, high=5, size=N)
data_y = _a * data_x + _b + np.random.normal(0, 2, N)
plt.plot(data_x, data_y, 'ro')
plt.axhline(0, color='black', lw=1)
plt.axvline(0, color='black', lw=1)
plt.show()
# -
# ### Create placeholders for inputs and labels
# x: inputs
x = tf.placeholder(tf.float32, name='x')
# y: labels
y = tf.placeholder(tf.float32, name='y')
# ### Create weight and bias
# +
tf.set_random_seed(219)
# create Variables
W = tf.get_variable(name="weights", shape=[], initializer=tf.random_normal_initializer())
b = tf.get_variable(name="bias", shape=[], initializer=tf.random_uniform_initializer())
# -
# ### Build a model: $y = Wx + b$
y_pred = W * x + b
# ### Define loss function
loss = tf.square(y_pred - y, name="loss")
# ### Create a optimizer
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.001).minimize(loss)
# ## Phase 2. Train a model
# ### Train a model
with tf.Session(config=sess_config) as sess:
# Initialize all variables
sess.run(tf.global_variables_initializer())
writer = tf.summary.FileWriter('graphs/01.1.linear.regression', sess.graph)
writer.close()
# train the model
max_epochs = 100
loss_history = []
start_time = time.time()
for epoch in range(max_epochs):
total_loss = 0.0
shuffle_index = np.random.permutation(len(data_x))
for i in shuffle_index:
x_ = data_x[i]
y_ = data_y[i]
_, loss_ = sess.run([train_op, loss],
feed_dict={x: x_,
y: y_})
total_loss += loss_
total_loss /= len(data_x)
loss_history.append(total_loss)
if (epoch+1) % 10 == 0:
print('Epoch %d: total_loss: %f' % (epoch+1, total_loss))
print('training done!')
print('elapsed time {} sec'.format(time.time() - start_time))
W_, b_ = sess.run([W, b])
# ### Print the results: W and b
#
# * 정답 W = 3, b = -3
print(W_, b_)
# ### Plot the loss funtion
plt.plot(loss_history, label='loss_history')
plt.legend()
plt.show()
# ### Plot the results
plt.plot(data_x, data_y, 'ro', label='Real data')
plt.plot(data_x, W_ * data_x + b_, lw=5, label='model')
plt.axhline(0, color='black', lw=1)
plt.axvline(0, color='black', lw=1)
plt.legend()
plt.show()
| tf.version.1/02.regression/01.1.linear.regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://github.com/luisguiserrano/hmm/blob/master/Simple%20HMM.ipynb
# s stands for Sunny
# r stands for Rainy
# h stands for Happy
# g stands for Grumpy
from numpy import random
# +
# Transition Probabilities
p_ss = 0.8 #prob of Sunny day after Sunny day
p_sr = 0.2 #prob of Rainy day after a Sunny day
p_rs = 0.4 #prob of Rainy day after Sunny day
p_rr = 0.6 #prob of Rainy day after Rainy day
# Initial Probabilities
p_s = 2/3 #prob of Sunny day
p_r = 1/3 #prob of Rainy day
# Emission Probabilities
p_sh = 0.8 #prob of being Happy given Sunny
p_sg = 0.2 #prob of being Grumpy given Sunny
p_rh = 0.4 #prob of being Happy given Rainy
p_rg = 0.6 #prob of being Grumpy given Rainy
# +
moods = ['H', 'H', 'G', 'G', 'G', 'H']
probabilities = []
weather = []
if moods[0] == 'H':
probabilities.append((p_s*p_sh, p_r*p_rh))
else:
probabilities.append((p_s*p_sg, p_r*p_rg))
for i in range(1,len(moods)):
yesterday_sunny, yesterday_rainy = probabilities[-1]
if moods[i] == 'H':
today_sunny = max(yesterday_sunny*p_ss*p_sh, yesterday_rainy*p_rs*p_sh)
today_rainy = max(yesterday_sunny*p_sr*p_rh, yesterday_rainy*p_rr*p_rh)
probabilities.append((today_sunny, today_rainy))
else:
today_sunny = max(yesterday_sunny*p_ss*p_sg, yesterday_rainy*p_rs*p_sg)
today_rainy = max(yesterday_sunny*p_sr*p_rg, yesterday_rainy*p_rr*p_rg)
probabilities.append((today_sunny, today_rainy))
for p in probabilities:
if p[0] > p[1]:
weather.append('S')
else:
weather.append('R')
weather
# -
#rounding off elements in a tuple
rounded_probs = [ ( round ( a [ 0 ], 3 ), round ( a [ 1 ], 3 )) for a in probabilities ]
rounded_probs
probabilities
| bayes/Old_code/Untitled2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#replacing a char
import numpy as np
a="hello"
print(np.char.replace(a,'o','l'))
# +
#replacing a string
import numpy as np
print(np.char.replace("Hatred to all","all","None"))
# -
| String_operations/Replace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SSNR validation
# ---
# Validate SSNR method on synthetic images.
# +
from pathlib import Path
from tqdm.notebook import tqdm
import numpy as np
import pandas as pd
from skimage import io
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
# -
#Local imports
import sys
# Add code directory to path
code_dir = '../code/'
if code_dir not in sys.path:
sys.path.insert(1, code_dir)
# Import ssnr code
import ssnr
import noise
# Suppress warnings
import warnings
warnings.simplefilter('ignore', RuntimeWarning)
# ## White noise
# ---
# ### Generate noisy images
# +
# Parameters
SNR_set = 1
# Generate image
image = noise.generate_image(512, 512)
# Add noise
noisy_1 = image + noise.white_noise(image.var() / SNR_set, image.shape)
noisy_2 = image + noise.white_noise(image.var() / SNR_set, image.shape)
# Plot images
fig, axes = plt.subplots(ncols=3, figsize=(12, 4))
axes[0].imshow(image)
axes[1].imshow(noisy_1)
axes[2].imshow(noisy_2);
# -
# ### SSNR
# +
# Direct SSNR calculation
SSNR_data = ssnr.SSNR_ring([noisy_1, noisy_2])
# Plot SSNR data
fig, ax = plt.subplots(figsize=(8, 3))
ax.loglog(SSNR_data, label='SSNR')
ax.set_xlabel('Spatial frequency [-]')
ax.set_ylabel('SSNR [-]')
ax.legend()
ax.grid(ls=':')
# -
# ### SNR calculations
# +
# From JOY
SNR_JOY = (ssnr.SNR_JOY(noisy_1) +
ssnr.SNR_JOY(noisy_2)) / 2
# From SSNR
SNR_SSNR = ssnr.SSNR_full([noisy_1, noisy_2])
# Output
out = f"""\
True SNR........ {SNR_set:.10f}
SNR from JOY.... {SNR_JOY:.10f}
SNR from SSNR... {SNR_SSNR:.10f}
"""
print(out)
# -
# ## Batch of images with white noise
# ---
# +
# Set up DataFrame
df = pd.DataFrame(columns=['SNR', 'SNR SSNR', 'SNR JOY'])
df['SNR'] = np.geomspace(1e-4, 1e4, 30)
for i, row in tqdm(df.iterrows(),
total=len(df)):
SNR_set = row['SNR']
# Generate noisy images
image = noise.generate_image(512, 512)
noisy_1 = image + noise.white_noise(image.var()/SNR_set, image.shape)
noisy_2 = image + noise.white_noise(image.var()/SNR_set, image.shape)
# (S)SNR calculations
df.loc[i, 'SNR SSNR'] = ssnr.SSNR_full([noisy_1, noisy_2])
df.loc[i, 'SNR JOY'] = (ssnr.SNR_JOY(noisy_1) +
ssnr.SNR_JOY(noisy_2)) / 2
# Preview
df[::5]
# -
# Plot SNR calculations
fig, ax = plt.subplots(figsize=(7, 5))
ax.plot(df['SNR'], df['SNR'], 'k--')
for s in ['SNR SSNR', 'SNR JOY']:
ax.plot(df['SNR'], df[s], 'o-', mfc='none', label=s)
# Aesthetics
ax.set_xlabel('SNR set [-]')
ax.set_ylabel('SNR measured [-]')
ax.set_xscale('log')
ax.set_yscale('log')
ax.legend()
ax.grid(ls=':')
# ## Batch of images with non-white noise
# ---
# Shows that SSNR method is more robust to correlated noise than Joy method.
# +
# Set up DataFrame
df = pd.DataFrame(columns=['SNR', 'SNR SSNR', 'SNR JOY'])
df['SNR'] = np.geomspace(1e-4, 1e4, 30)
for i, row in tqdm(df.iterrows(),
total=len(df)):
SNR_set = row['SNR']
# Generate noisy images
image = noise.generate_image(512, 512)
noisy_1 = image + noise.nonwhite_noise(image.var()/SNR_set, image.shape)
noisy_2 = image + noise.nonwhite_noise(image.var()/SNR_set, image.shape)
# (S)SNR calculations
df.loc[i, 'SNR SSNR'] = ssnr.SSNR_full([noisy_1, noisy_2])
df.loc[i, 'SNR JOY'] = (ssnr.SNR_JOY(noisy_1) +
ssnr.SNR_JOY(noisy_2)) / 2
# Preview
df[::5]
# -
# Plot SNR calculations
fig, ax = plt.subplots(figsize=(7, 5))
ax.plot(df['SNR'], df['SNR'], 'k--')
for s in ['SNR SSNR', 'SNR JOY']:
ax.plot(df['SNR'], df[s], 'o-', mfc='none', label=s)
# Aesthetics
ax.set_xlabel('SNR set [-]')
ax.set_ylabel('SNR measured [-]')
ax.set_xscale('log')
ax.set_yscale('log')
ax.legend()
ax.grid(ls=':')
# ## Images with fewer features
# ---
# Shows that SSNR method is also robust to images with fewer features.
# +
# Set up DataFrame
df = pd.DataFrame(columns=['SNR', 'SNR SSNR', 'SNR JOY'])
df['SNR'] = np.geomspace(1e-4, 1e4, 30)
for i, row in tqdm(df.iterrows(),
total=len(df)):
SNR_set = row['SNR']
# Generate noisy images
image = noise.generate_image(512, 512, N_features=10)
noisy_1 = image + noise.white_noise(image.var()/SNR_set, image.shape)
noisy_2 = image + noise.white_noise(image.var()/SNR_set, image.shape)
# (S)SNR calculations
df.loc[i, 'SNR SSNR'] = ssnr.SSNR_full([noisy_1, noisy_2])
df.loc[i, 'SNR JOY'] = (ssnr.SNR_JOY(noisy_1) +
ssnr.SNR_JOY(noisy_2)) / 2
# Preview
df[::5]
# -
# Plot SNR calculations
fig, axes = plt.subplots(ncols=2, figsize=(12, 5),
gridspec_kw={'width_ratios': [5, 7]})
axes[0].imshow(noisy_1)
axes[1].plot(df['SNR'], df['SNR'], 'k--')
for s in ['SNR SSNR', 'SNR JOY']:
axes[1].plot(df['SNR'], df[s], 'o-', mfc='none', label=s)
# Aesthetics
axes[1].set_xlabel('SNR set [-]')
axes[1].set_ylabel('SNR measured [-]')
axes[1].set_xscale('log')
axes[1].set_yscale('log')
axes[1].legend()
axes[1].grid(ls=':')
| notebooks/SSNR_validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data visualization
# ### The gender gap in college degrees
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
women_degrees = pd.read_csv('percent-bachelors-degrees-women-usa.csv')
cb_dark_blue = (0/255,107/255,164/255)
cb_orange = (255/255, 128/255, 14/255)
stem_cats = ['Engineering', 'Computer Science', 'Psychology', 'Biology', 'Physical Sciences', 'Math and Statistics']
fig = plt.figure(figsize=(18, 3))
for sp in range(0,6):
ax = fig.add_subplot(1,6,sp+1)
ax.plot(women_degrees['Year'], women_degrees[stem_cats[sp]], c=cb_dark_blue, label='Women', linewidth=3)
ax.plot(women_degrees['Year'], 100-women_degrees[stem_cats[sp]], c=cb_orange, label='Men', linewidth=3)
ax.spines["right"].set_visible(False)
ax.spines["left"].set_visible(False)
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
ax.set_xlim(1968, 2011)
ax.set_ylim(0,100)
ax.set_title(stem_cats[sp])
ax.tick_params(bottom="off", top="off", left="off", right="off")
if sp == 0:
ax.text(2005, 87, 'Men')
ax.text(2002, 8, 'Women')
elif sp == 5:
ax.text(2005, 62, 'Men')
ax.text(2001, 35, 'Women')
plt.show()
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
women_degrees = pd.read_csv('percent-bachelors-degrees-women-usa.csv')
cb_dark_blue = (0/255,107/255,164/255)
cb_orange = (255/255, 128/255, 14/255)
stem_cats = ['Engineering', 'Computer Science', 'Psychology', 'Biology', 'Physical Sciences', 'Math and Statistics']
fig = plt.figure(figsize=(18, 3))
for sp in range(0,6):
ax = fig.add_subplot(1,6,sp+1)
ax.plot(women_degrees['Year'], women_degrees[stem_cats[sp]], c=cb_dark_blue, label='Women', linewidth=3)
ax.plot(women_degrees['Year'], 100-women_degrees[stem_cats[sp]], c=cb_orange, label='Men', linewidth=3)
ax.spines["right"].set_visible(False)
ax.spines["left"].set_visible(False)
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
ax.set_xlim(1968, 2011)
ax.set_ylim(0,100)
ax.set_title(stem_cats[sp])
ax.tick_params(bottom="off", top="off", left="off", right="off")
if sp == 0:
ax.text(2005, 87, 'Men')
ax.text(2002, 8, 'Women')
elif sp == 5:
ax.text(2005, 62, 'Men')
ax.text(2001, 35, 'Women')
plt.show()
# -
stem_cats = ['Psychology', 'Biology', 'Math and Statistics', 'Physical Sciences', 'Computer Science', 'Engineering']
lib_arts_cats = ['Foreign Languages', 'English', 'Communications and Journalism', 'Art and Performance', 'Social Sciences and History']
other_cats = ['Health Professions', 'Public Administration', 'Education', 'Agriculture','Business', 'Architecture']
# +
fig = plt.figure(figsize=(18, 18))
for sp in range(0,6):
ax = fig.add_subplot(6,3,sp*3+1)
ax.plot(women_degrees['Year'], women_degrees[stem_cats[sp]], c=cb_dark_blue, label='Women', linewidth=3)
ax.plot(women_degrees['Year'], 100-women_degrees[stem_cats[sp]], c=cb_orange, label='Men', linewidth=3)
ax.spines["right"].set_visible(False)
ax.spines["left"].set_visible(False)
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
ax.set_xlim(1968, 2011)
ax.set_ylim(0,100)
ax.set_title(stem_cats[sp])
ax.set_yticks([0,100])
ax.tick_params(bottom="off", top="off", left="off", right="off",
labelbottom='off')
ax.axhline(50, c=(171/255, 171/255, 171/255), alpha=0.3)
if sp == 0:
ax.text(2005, 87, 'Men')
ax.text(2002, 8, 'Women')
elif sp == 5:
ax.text(2005, 75, 'Men')
ax.text(2001, 25, 'Women')
ax.tick_params(labelbottom='on')
for sp in range(0,5):
ax = fig.add_subplot(6,3,sp*3+2)
ax.plot(women_degrees['Year'], women_degrees[lib_arts_cats[sp]], c=cb_dark_blue, label='Women', linewidth=3)
ax.plot(women_degrees['Year'], 100-women_degrees[lib_arts_cats[sp]], c=cb_orange, label='Men', linewidth=3)
ax.spines["right"].set_visible(False)
ax.spines["left"].set_visible(False)
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
ax.set_xlim(1968, 2011)
ax.set_ylim(0,100)
ax.set_title(stem_cats[sp])
ax.set_yticks([0,100])
ax.tick_params(bottom="off", top="off", left="off", right="off",
labelbottom='off')
ax.axhline(50, c=(171/255, 171/255, 171/255), alpha=0.3)
if sp == 0:
ax.text(2005, 87, 'Men')
ax.text(2002, 8, 'Women')
elif sp==4:
ax.tick_params(labelbottom='on')
for sp in range(0,6):
ax = fig.add_subplot(6,3,sp*3+3)
ax.plot(women_degrees['Year'], women_degrees[other_cats[sp]], c=cb_dark_blue, label='Women', linewidth=3)
ax.plot(women_degrees['Year'], 100-women_degrees[other_cats[sp]], c=cb_orange, label='Men', linewidth=3)
ax.spines["right"].set_visible(False)
ax.spines["left"].set_visible(False)
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
ax.set_xlim(1968, 2011)
ax.set_ylim(0,100)
ax.set_title(stem_cats[sp])
ax.set_yticks([0,100])
ax.tick_params(bottom="off", top="off", left="off", right="off",
labelbottom='off')
ax.axhline(50, c=(171/255, 171/255, 171/255), alpha=0.3)
if sp == 0:
ax.text(2005, 87, 'Men')
ax.text(2002, 8, 'Women')
elif sp == 5:
ax.text(2005, 62, 'Men')
ax.text(2001, 35, 'Women')
ax.tick_params(labelbottom='on')
plt.show()
plt.savefig('biology_degrees.png')
# -
| assets/projects/DQ_Visualization_2/Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sqlite3
con = sqlite3.connect('todo.db') # Warning: This file is created in the current directory
con.execute("CREATE TABLE todo (id INTEGER PRIMARY KEY, task char(100) NOT NULL, status bool NOT NULL)")
con.execute("INSERT INTO todo (task,status) VALUES ('Read A-byte-of-python to get a good introduction into Python',0)")
con.execute("INSERT INTO todo (task,status) VALUES ('Visit the Python website',1)")
con.execute("INSERT INTO todo (task,status) VALUES ('Test various editors for and check the syntax highlighting',1)")
con.execute("INSERT INTO todo (task,status) VALUES ('Choose your favorite WSGI-Framework',0)")
con.commit()
# +
import sqlite3
from bottle import route, run, debug, template, request, static_file, error
@route('/todo')
def todo_list():
conn = sqlite3.connect('todo.db')
c = conn.cursor()
c.execute("SELECT id, task FROM todo WHERE status LIKE '1'")
result = c.fetchall()
c.close()
output = template('make_table', rows=result)
return output
@route('/new', method='GET')
def new_item():
if request.GET.save:
new = request.GET.task.strip()
conn = sqlite3.connect('todo.db')
c = conn.cursor()
c.execute("INSERT INTO todo (task,status) VALUES (?,?)", (new,1))
new_id = c.lastrowid
conn.commit()
c.close()
return '<p>The new task was inserted into the database, the ID is %s</p>' % new_id
else:
return template('new_task.tpl')
@route('/item<item:re:[0-9]+>')
def show_item(item):
conn = sqlite3.connect('todo.db')
c = conn.cursor()
c.execute("SELECT task FROM todo WHERE id LIKE ?", (item,))
result = c.fetchall()
c.close()
if not result:
return 'This item number does not exist!'
else:
return 'Task: %s' % result[0]
@route('/json<json:re:[0-9]+>')
def show_json(json):
conn = sqlite3.connect('todo.db')
c = conn.cursor()
c.execute("SELECT task FROM todo WHERE id LIKE ?", (json,))
result = c.fetchall()
c.close()
if not result:
return {'task': 'This item number does not exist!'}
else:
return {'task': result[0]}
@route('/edit/<no:int>', method='GET')
def edit_item(no):
if request.GET.save:
edit = request.GET.task.strip()
status = request.GET.status.strip()
if status == 'open':
status = 1
else:
status = 0
conn = sqlite3.connect('todo.db')
c = conn.cursor()
c.execute("UPDATE todo SET task = ?, status = ? WHERE id LIKE ?", (edit, status, no))
conn.commit()
return '<p>The item number %s was successfully updated</p>' % no
else:
conn = sqlite3.connect('todo.db')
c = conn.cursor()
c.execute("SELECT task FROM todo WHERE id LIKE ?", (str(no)))
cur_data = c.fetchone()
return template('edit_task', old=cur_data, no=no)
@error(403)
def mistake403(code):
return 'The parameter you passed has the wrong format!'
@error(404)
def mistake404(code):
return 'Sorry, this page does not exist!'
@route('/help')
def help():
return static_file('help.html', root='/home/sai/restapi/')
debug(True)
run()
# -
| Todolist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Denoising Autoencoder
#
# Sticking with the MNIST dataset, let's add noise to our data and see if we can define and train an autoencoder to _de_-noise the images.
#
# <img src='notebook_ims/autoencoder_denoise.png' width=70%/>
#
# Let's get started by importing our libraries and getting the dataset.
# +
import torch
import numpy as np
from torchvision import datasets
import torchvision.transforms as transforms
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# load the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# Create training and test dataloaders
num_workers = 0
# how many samples per batch to load
batch_size = 20
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size, num_workers=num_workers)
# -
# ### Visualize the Data
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# get one image from the batch
img = np.squeeze(images[0])
fig = plt.figure(figsize = (5,5))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
# -
# ---
# # Denoising
#
# As I've mentioned before, autoencoders like the ones you've built so far aren't too useful in practive. However, they can be used to denoise images quite successfully just by training the network on noisy images. We can create the noisy images ourselves by adding Gaussian noise to the training images, then clipping the values to be between 0 and 1.
#
# >**We'll use noisy images as input and the original, clean images as targets.**
#
# Below is an example of some of the noisy images I generated and the associated, denoised images.
#
# <img src='notebook_ims/denoising.png' />
#
#
# Since this is a harder problem for the network, we'll want to use _deeper_ convolutional layers here; layers with more feature maps. You might also consider adding additional layers. I suggest starting with a depth of 32 for the convolutional layers in the encoder, and the same depths going backward through the decoder.
#
# #### TODO: Build the network for the denoising autoencoder. Add deeper and/or additional layers compared to the model above.
# +
import torch.nn as nn
import torch.nn.functional as F
# define the NN architecture
class ConvDenoiser(nn.Module):
def __init__(self):
super(ConvDenoiser, self).__init__()
## encoder layers ##
self.conv1 = nn.Conv2d(1, 16, 3, padding=1)
self.conv2 = nn.Conv2d(16, 4, 3, padding=1)
# Average pooling for smoothing with a kernel of size 2 and stride 2
# self.pool = nn.AvgPool2d(2, 2)
self.pool = nn.MaxPool2d(2, 2)
## decoder layers ##
## a kernel of 2 and a stride of 2 will increase the spatial dims by 2. Stride is applied on the output not input,
## so every pixel in the input is convolved with the kernel.
self.t_conv1 = nn.ConvTranspose2d(4, 16, 2, stride=2)
self.t_conv2 = nn.ConvTranspose2d(16, 1, 2, stride=2)
def forward(self, x):
## encode ##
x = F.relu(self.conv1(x))
x = self.pool(x)
x = F.relu(self.conv2(x))
x = self.pool(x)
x = F.relu(self.t_conv1(x))
x = F.sigmoid(self.t_conv2(x))
## decode ##
return x
# initialize the NN
model = ConvDenoiser()
print(model)
# -
# ---
# ## Training
#
# We are only concerned with the training images, which we can get from the `train_loader`.
#
# >In this case, we are actually **adding some noise** to these images and we'll feed these `noisy_imgs` to our model. The model will produce reconstructed images based on the noisy input. But, we want it to produce _normal_ un-noisy images, and so, when we calculate the loss, we will still compare the reconstructed outputs to the original images!
#
# Because we're comparing pixel values in input and output images, it will be best to use a loss that is meant for a regression task. Regression is all about comparing quantities rather than probabilistic values. So, in this case, I'll use `MSELoss`. And compare output images and input images as follows:
# ```
# loss = criterion(outputs, images)
# ```
# +
# specify loss function
criterion = nn.MSELoss()
# specify loss function
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
# +
# number of epochs to train the model
n_epochs = 20
# for adding noise to images
noise_factor=0.5
for epoch in range(1, n_epochs+1):
# monitor training loss
train_loss = 0.0
###################
# train the model #
###################
for data in train_loader:
# _ stands in for labels, here
# no need to flatten images
images, _ = data
## add random noise to the input images
noisy_imgs = images + noise_factor * torch.randn(*images.shape)
# Clip the images to be between 0 and 1
noisy_imgs = np.clip(noisy_imgs, 0., 1.)
# clear the gradients of all optimized variables
optimizer.zero_grad()
## forward pass: compute predicted outputs by passing *noisy* images to the model
outputs = model(noisy_imgs)
# calculate the loss
# the "target" is still the original, not-noisy images
loss = criterion(outputs, images)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*images.size(0)
# print avg training statistics
train_loss = train_loss/len(train_loader)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(
epoch,
train_loss
))
# -
# ## Checking out the results
#
# Here I'm adding noise to the test images and passing them through the autoencoder. It does a suprising great job of removing the noise, even though it's sometimes difficult to tell what the original number is.
# +
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# add noise to the test images
noisy_imgs = images + noise_factor * torch.randn(*images.shape)
noisy_imgs = np.clip(noisy_imgs, 0., 1.)
# get sample outputs
output = model(noisy_imgs)
# prep images for display
noisy_imgs = noisy_imgs.numpy()
# output is resized into a batch of images
output = output.view(batch_size, 1, 28, 28)
# use detach when it's an output that requires_grad
output = output.detach().numpy()
# plot the first ten input images and then reconstructed images
fig, axes = plt.subplots(nrows=2, ncols=10, sharex=True, sharey=True, figsize=(25,4))
# input images on top row, reconstructions on bottom
for noisy_imgs, row in zip([noisy_imgs, output], axes):
for img, ax in zip(noisy_imgs, row):
ax.imshow(np.squeeze(img), cmap='gray')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# -
| autoencoder/denoising-autoencoder/Denoising_Autoencoder_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from statsmodels.tsa.seasonal import seasonal_decompose
import matplotlib.pyplot as plt
import scipy.stats as st
import seaborn as sns
import pandas as pd
from scipy.stats import norm, uniform, expon, t, probplot
import scipy.stats as st
from scipy.integrate import quad
from sympy.solvers import solve
from sympy import Symbol
import numpy as np
from pandas import Series, DataFrame
from statsmodels.graphics.tsaplots import plot_acf
from statsmodels.tsa.stattools import acf
from pandas.plotting import lag_plot
# + pycharm={"name": "#%%\n"}
w = DataFrame(np.random.normal(size=1000))
MA = DataFrame(w.rolling(window=3).mean()).dropna()
plot_acf(MA, lags=12, c="C1")
plt.vlines(x=2.1, ymin=0, ymax=1/3, color="red", linestyle='--', label="Geschaetzt")
plt.vlines(x=1.1, ymin=0, ymax=2/3, color="red", linestyle='--')
plt.vlines(x=0.1, ymin=0, ymax=1, color="red", linestyle='--')
plt.legend()
# -
df = pd.read_table("wave.dat")
df.head()
df.plot()
plt.xlabel("Time")
plt.ylabel("Wave height (mm)")
plt.show()
df.loc[0:59,:].plot()
plt.xlabel("Time")
plt.ylabel("Wave height (mm)")
plt.show()
from statsmodels.tsa.stattools import acf, plot_acf
acf(df["waveht"])[1]
lag_plot(df,1)
plt.xlabel("x_t")
plt.ylabel("x_(t+k)")
plt.show()
acf(df["waveht"])[[2,3,5,10]]
plt.subplot(221)
lag_plot(df,2)
plt.xlabel("x_t")
plt.ylabel("x_(t+k)")
plt.subplot(222)
lag_plot(df,3)
plt.xlabel("x_t")
plt.ylabel("x_(t+k)")
plt.subplot(223)
lag_plot(df,5)
plt.xlabel("x_t")
plt.ylabel("x_(t+k)")
plt.subplot(224)
lag_plot(df,10)
plt.xlabel("x_t")
plt.ylabel("x_(t+k)")
plt.show()
plot_acf(df["waveht"])
AirP = pd.read_csv("AirPassengers.csv")
AirP.head()
AirP["TravelDate"] = pd.to_datetime(AirP["TravelDate"])
AirP.set_index("TravelDate", inplace = True)
AirP.head()
plot_acf(AirP["Passengers"])
from statsmodels.tsa.seasonal import seasonal_decompose
remainder = seasonal_decompose(AirP["Passengers"], model="multiplicative").resid[6:138]
remainder.plot()
plot_acf(remainder, lags = 21)
print(AirP["Passengers"][6:138].std())
trend = seasonal_decompose(AirP["Passengers"], model="multiplicative").trend[6:138]
print((AirP["Passengers"][6:138]-trend).std())
decomp = seasonal_decompose(AirP["Passengers"])
seasonal_decompose(AirP["Passengers"], model="multiplicative").plot()
print((decomp.observed - decomp.trend)[6:138].std())
print(seasonal_decompose(AirP["Passengers"]).resid[6:138].std())
], model="multiplicative")
| Lernphase/SW12/Skript.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="8H1CGTFa1Lo-"
# # Bilibili视频网站排行榜评分的线性回归分析
#
# Bilibili(哔哩哔哩)是当下最热门的视频播放网站,人们在该网站对视频进行浏览,会根据自己喜好对视频进行点赞、投币、收藏、分享、回复等操作,Bilibili同时还有一个排行榜机制,记录了近期比较热门的视频,并给出了一个评分,排行榜的排序规则即根据该评分,本文将根据这些排行榜视频的各项数据,进行线性回归分析,结合实际情况找出对评分影响最大的数据。
# + [markdown] id="o_lnHqzn3IxU"
# Bilibili提供了获取排行榜数据的API,因此我们可以利用该API获取实时的排行榜数据,获取的数据为json格式,经过解析,已将数据保存至Bilibili.csv文件中。
# + [markdown] id="KWqxbjtv3aK1"
# 本文的思路即为多元线性回归,在统计学中,线性回归(linear regression)是利用称为线性回归方程的最小二乘函数对一个或多个自变量和因变量之间关系进行建模的一种回归分析。这种函数是一个或多个称为回归系数的模型参数的线性组合。只有一个自变量的情况称为简单回归,大于一个自变量情况的叫做多元回归(multivariable linear regression)。
#
# 完成了对各个自变量权重的计算,找出了对评分影响权重最高的因素,并得到了验证。
#
# 在哔哩哔哩官网首页处获取排行榜信息的API,得到为
#
# https://api.bilibili.com/x/web-interface/ranking/v2?rid=0
#
# 请求参数为
# - rid 分类,0代表全部分类
#
# 返回格式为json格式,json中的data字段为视频信息数组。
#
# 为了防止请求次数过多增加服务器压力,导致本机IP禁止访问哔哩哔哩,先将请求的response保存到data.txt文件中。
#
# 使用python的json包进行解析
# ```json
# {
# "aid": 375696064,
# "videos": 1,
# "tid": 17,
# "tname": "单机游戏",
# "copyright": 1,
# "pic": "http://i0.hdslb.com/bfs/archive/85005893d4c4959ff096d6fb061040e223842bcb.jpg",
# "title": "史上最骚魔法师!(第二集)",
# "pubdate": 1621566911,
# "ctime": 1621566912,
# "desc": "本期请到了Warma参与配音!鼓掌!!!!!\n游戏:Darkside Detective\n第一集:BV1M64y1m7gA\n各位如果看得开心,希望三连支持一下!",
# "state": 0,
# "duration": 658,
# "mission_id": 24025,
# "rights": {
# "bp": 0,
# "elec": 0,
# "download": 0,
# "movie": 0,
# "pay": 0,
# "hd5": 0,
# "no_reprint": 1,
# "autoplay": 1,
# "ugc_pay": 0,
# "is_cooperation": 0,
# "ugc_pay_preview": 0,
# "no_background": 0
# },
# "owner": {
# "mid": 546195,
# "name": "老番茄",
# "face": "http://i0.hdslb.com/bfs/face/bc5ca101313d4db223c395d64779e76eb3482d60.jpg"
# },
# "stat": {
# "aid": 375696064,
# "view": 1149043,
# "danmaku": 7300,
# "reply": 3278,
# "favorite": 37490,
# "coin": 98319,
# "share": 1780,
# "now_rank": 0,
# "his_rank": 1,
# "like": 210211,
# "dislike": 0
# },
# "dynamic": "用魔法击败魔法",
# "cid": 341808079,
# "dimension": {
# "width": 1920,
# "height": 1080,
# "rotate": 0
# },
# "short_link": "https://b23.tv/BV1jo4y117Vf",
# "short_link_v2": "https://b23.tv/BV1jo4y117Vf",
# "bvid": "BV1jo4y117Vf",
# "score": 2446535
# }
# ```
#
# 通过解析json格式文件并整理得到bilibili.csv
# + id="0-WWEKsN0TRn"
# 包引用
import pandas as pd # csv文件读写分析
import numpy as np # 线性代数
import matplotlib.pyplot as plt # 制图
# + id="XP6dF1FT2eGk"
data = pd.read_csv('./bilibili.csv', index_col='title')
# 评分
score = data['score']
# 播放量
view = data['view']
# 投币
coins = data['coins']
# 收藏
favor = data['favorite']
# 评论
reply = data['reply']
# 点赞
like = data['like']
# 弹幕数量
danmu = data['danmu']
# 分享次数
share = data['share']
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="U3B9JJPm3AW-" outputId="4be1666c-e32f-4ba4-f573-272cfafec53b"
data
# + [markdown] id="dIMw7FdF4Qyq"
# 部分数据展示
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="DOqr1mg_2sy9" outputId="d0f8ca82-f792-4b5c-e0a3-ebf0c5adcde6"
score
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="JvuOU5MD2wJl" outputId="38a2bcc8-7351-47ad-e520-f9d2cef99a22"
coins
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="anr_qZhs4Us3" outputId="fa6ed6b2-dd39-43cc-a704-2e573a42a8ee"
favor
# + id="2yjzHzqH4t5q"
x = [i for i in range(len(score))]
# 获取自变量矩阵
X = data.iloc[:,3:10].values
# 获取因变量矩阵
Y = data.iloc[:,2].values.reshape(-1,1)
# 生成设计矩阵
om = np.ones(X.shape[0]).reshape(-1,1)
X = np.hstack((X, om))
# + colab={"base_uri": "https://localhost:8080/", "height": 293} id="jCEPuU-kA6z4" outputId="788e905d-c33d-4b93-adfe-3e7cb2518db2"
# 绘制图像,按照排名递减绘制,即分数从高到低
plt.xlabel('rank')
plt.ylabel('score')
plt.plot(x, score, 'o')
# -
# 排名-总分图,总分与rank正相关。
labels = ['score']
plt.ylabel('score')
plt.boxplot(score, labels = labels, showmeans = True)
# 由箱线图可以得到score数据中,均值大致为120000
#
# 中位数大致为900000
#
# 使用numpy自带库函数检验
np.mean(score)
np.median(score)
np.var(score)
np.std(score)
plt.xlabel('rank')
plt.ylabel('favorite')
plt.plot(x, favor, 'o')
# 根据排名-收藏图,可以发现,收藏对于排名即总分,有一定的正相关。
plt.xlabel('favorite')
labels = ['favorite']
plt.ylabel('favorite')
plt.boxplot(favor, labels = labels, showmeans = True)
# 由箱线图可知,收藏数据集中在[6000, 20000]内,中位数大致为10000,均值为16000
np.mean(favor)
np.median(favor)
np.var(favor)
np.std(favor)
plt.xlabel('rank')
plt.ylabel('view')
plt.plot(x, view, 'o')
# 通过排名-播放量图,可以得出播放量与排名在一定程度上为正相关。
plt.xlabel('view')
labels = ['view']
plt.ylabel('view')
plt.boxplot(view, labels = labels, showmeans = True)
# 由箱线图可知,播放量数据集中在[600000, 800000]内,中位数大致为600000,均值大致为800000
np.mean(view)
np.median(view)
np.var(view)
np.std(view)
plt.xlabel('rank')
plt.ylabel('coins')
plt.plot(x, coins, 'o')
# 通过排名-投币数图,可以得出排名与投币数在一定程度上为正相关。
plt.xlabel('coins')
labels = ['coins']
plt.ylabel('coins')
plt.boxplot(coins, labels = labels, showmeans = True)
# 由箱线图可知,投币数据集中在[15000, 30000]内,中位数大致为17000,均值大致为30000
np.mean(coins)
np.median(coins)
# 方差
np.var(coins)
# 标准差
np.std(coins)
plt.xlabel('rank')
plt.ylabel('reply')
plt.plot(x, reply, 'o')
# 通过排名-评论数图,可以得出排名与评论数相关性较低。
plt.xlabel('reply')
labels = ['reply']
plt.ylabel('reply')
plt.boxplot(reply, labels = labels, showmeans = True)
# 由箱线图可知,评论数据集中在[2500, 4000]内,中位数大致为2500,均值大致为3000
np.mean(reply)
np.median(reply)
# 方差
np.var(reply)
# 标准差
np.std(reply)
plt.xlabel('rank')
plt.ylabel('like')
plt.plot(x, like, 'o')
# 通过排名-点赞数图,可以得出排名与点赞数在一定程度上为正相关。
plt.xlabel('like')
labels = ['like']
plt.ylabel('like')
plt.boxplot(like, labels = labels, showmeans = True)
# 由箱线图可知,点赞数据集中在[50000, 100000]内,中位数大致为60000,均值大致为80000
np.mean(like)
np.median(like)
# 方差
np.var(like)
# 标准差
np.std(like)
plt.xlabel('rank')
plt.ylabel('danmu')
plt.plot(x, danmu, 'o')
# 通过排名-弹幕数图,可以得出排名与弹幕数相关性较低。
plt.xlabel('danmu')
labels = ['danmu']
plt.ylabel('danmu')
plt.boxplot(danmu, labels = labels, showmeans = True)
# 由箱线图可知,弹幕数据集中在[2500, 5000]内,中位数大致为2500,均值大致为5000
np.median(danmu)
np.mean(danmu)
# 方差
np.var(danmu)
# 标准差
np.std(danmu)
plt.xlabel('rank')
plt.ylabel('share')
plt.plot(x, share, 'o')
# 通过排名-分享数图,可以得出排名与分享数相关性较低。
plt.xlabel('share')
labels = ['share']
plt.ylabel('share')
plt.boxplot(share, labels = labels, showmeans = True)
# 由箱线图可知,分享数据集中在[1000, 6000]内,中位数大致为3000,均值大致为6000
np.median(share)
np.mean(share)
# 方差
np.var(share)
# 标准差
np.std(share)
# + id="MFN2sGkq4oXS"
# 计算系数矩阵w-hat
w_hat = np.dot(np.dot(np.linalg.inv(np.dot(X.T, X)), X.T), Y)
# 获取b矩阵
b = w_hat[-1]
# 获取w系数矩阵
w = w_hat[:-1]
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="LpMVbf795zRI" outputId="159d7e74-963c-4911-f790-9545b9cd62d0"
w_hat
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="gk-n-xIX51rd" outputId="cc7e7a62-cda2-4665-c246-aa7e8305fdfb"
b
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="Wbcg4FZx53qA" outputId="a09bf2c8-9002-45b8-d120-cc479f37e6ae"
w
# + [markdown] id="Q5qsxHNL6GLj"
# 这里发现w系数矩阵中出现了负值,经检查发现该项目对应的自变量为like和reply,即视频的点赞次数和评论数,这与实际情况是不符的,猜测是因为该自变量对结果的影响过小,导致预测出现了偏差,同时发现弹幕数量对视频评分的影响过大,远超于其他参数,显然与实际生活不符,并且作为对视频质量的估计,弹幕数也的确不能作为一个重要的参数。
#
# 因此我们对弹幕数进行剔除重新进行拟合。
# + id="z1B0AHzjARNr"
data = pd.read_csv('./bilibili.csv', index_col='title')
# + id="g97zAppY7fxY"
# 获取自变量矩阵,剔除弹幕数
X = data.iloc[:,[3, 4, 5, 6, 8, 9]].values
# 获取因变量矩阵
Y = data.iloc[:,2].values.reshape(-1,1)
# 生成设计矩阵
om = np.ones(X.shape[0]).reshape(-1,1)
X = np.hstack((X, om))
# + id="tfKa7U8xAsVb"
# 计算系数矩阵w-hat
w_hat = np.dot(np.dot(np.linalg.inv(np.dot(X.T, X)), X.T), Y)
# 获取b矩阵
b = w_hat[-1]
# 获取w系数矩阵
w = w_hat[:-1]
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="ivbMfPD0Atvv" outputId="5c938e68-9146-4a4b-b5b3-1152ac3a0dad"
b
# -
w
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 0} id="EpIPOoXGAvNc" outputId="568e5e1c-c9cf-42a4-e0c8-31c89b709026"
# 观察发现此时各项参数对于评分的影响已经较为接近正常水平。但是share,视频分享数对于评分的影响仍为负值,依然根据上述猜测,share对于总分的影响过小,导致拟合的结果差,因此再次剔除share字段。
# -
# 获取自变量矩阵,剔除弹幕数,分享数
X = data.iloc[:,[3, 4, 5, 6, 9]].values
# 获取因变量矩阵
Y = data.iloc[:,2].values.reshape(-1,1)
# 生成设计矩阵
om = np.ones(X.shape[0]).reshape(-1,1)
X = np.hstack((X, om))
# 计算系数矩阵w-hat
w_hat = np.dot(np.dot(np.linalg.inv(np.dot(X.T, X)), X.T), Y)
# 获取b矩阵
b = w_hat[-1]
# 获取w系数矩阵
w = w_hat[:-1]
# + id="h4eod0IAGQR2"
X_test = data.iloc[:, [3, 4, 5, 6, 9]].values
Y_predict = np.dot(X_test, w) + b
# + id="NiQFS0ktHWgx"
dic = abs(Y-Y_predict)
# + id="x_lRAKrFGu2R"
sum = 0
for i in range(len(Y)):
sum += abs(Y[i][0] - Y_predict[i][0])
# + [markdown] id="4Foiw_eFHQbA"
# 误差的平均值
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="J2FhNDTLG3QR" outputId="ce8d9726-50e8-41b3-bfca-3113faea5867"
sum/len(Y)
# -
#计算平均平方误差啊
mse = 0
for i in range(len(Y)):
mse += (Y[i][0] - Y_predict[i][0])**2
mse /= len(Y)
mse
len(Y)
# 计算平均平方误差平方根
rmse = np.sqrt(mse)
rmse
# 计算R^2决定系数
# r^2 = 1 - rss/tss
rss = 0
for i in range(len(Y)):
rss += (Y[i][0] - Y_predict[i][0])**2
rss
mse*len(Y)
# 计算tss
tss = 0
y_ = 0 # average y
for i in range(len(Y)):
y_ += Y[i][0]
y_ = y_ / len(Y)
for i in range(len(Y)):
tss += (Y[i][0] - y_)**2
R2 = 1 - rss/tss
R2
# 通过对R^2决定系数的计算,可以发现模型拟合的效果较为良好。
# + [markdown] id="UGKXzBl_JYQP"
# 观察预测值与实际值的差距,发现拟合的趋势基本吻合,但仍有较大的误差
# + colab={"base_uri": "https://localhost:8080/", "height": 293} id="rQgl3TThHfoK" outputId="8fce0288-1c34-4e17-cc89-e90e8c39eb71"
x = [i for i in range(0, len(Y))]
plt.plot(x, Y_predict, 'o')
plt.plot(x, Y)
# -
plt.plot(x, Y_predict, 'o')
plt.plot(x, Y, 'o')
# 从图象中可以看出,预测值的趋势与实际值较为接近。
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="MRK9KP79Ili3" outputId="a572ad9e-bc21-4a52-811f-6e1c1680fad7"
y = [sum/len(Y) for i in range(len(x))]
plt.plot(x, dic, 'o')
plt.plot(x, y)
# -
# 分析得出,该线性模型的拟合误差较为集中,集中在[0, 181348.81788162683]即平均误差之间。
# + [markdown] id="DAir1zraKjC8"
# 根据权重分析各个数据对总分的影响,可以发现,收藏对于视频总评分的影响占比最高,其次是硬币数,播放数,说明哔哩哔哩对与视频的质量高低评判有一定的综合考量,播放量属于可以由视频制作人通过其他方式刷取,但是收藏量是由用户对于视频质量的高低做出的决定,因而更具代表性,更有说服力。反观实际观看体验中,有些视频制作人会以视频收藏满几万后,更新新一期视频,因为该项对视频的收益影响最大,更能给视频制作人带来实际收益。
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="pvMz4BEAKhkN" outputId="9ef87302-50cd-4761-8261-fabee5c1b138"
w
# + [markdown] id="Yv9O3qQELbEh"
# 本次线性回归的拟合可以较为准确的判断出各各数据对于视频评分的影响,但是对于视频评分的预测误差较大,但是仍能较准确的判断出视频评分所影响的视频排名,认为有如下原因:
# - 数据量过少,导致拟合程度不足
# - 评分规则非线形模型,应该更换模型
# - 还有其他未考虑的因素
#
# 如果要更准确的对视频评分进行预测,应该综合分析数据的特性,或者采用机器学习等更优秀的手段来进行求解和预测,线性回归仍有一定的局限性。
| lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Review of descriptive statistics
# ------
#
# Lets review some descriptive statistics.
# #### average / mean
#
# This is simple: represent the mean value.
#
# $$\mu = \frac{1}{N}\sum_i^N a_i$$
# +
import numpy as np
a = [3, 5, 9]
np.mean(a)
assert np.mean(a) == sum(a)/len(a)
print(np.mean(a))
# -
np.square(3)
# +
mean = 5.666666666666667
np.sqrt((np.square(3 - mean) + np.square(5 - mean) + np.square(9 - mean))/3)
# -
# #### standard deviation:
#
# * Discrete random variable:
#
# $$\sigma = \sqrt{\frac{1}{N} \sum_i^N (x_i - \mu)^2}$$
#
np.std(a)
# Standard deviations represents the dispersion. Low standard deviation indicates that the values are close to the mean value.
#
# <!-- <img src="figs/std.png" style="width: 350px;"> -->
#
# <a title="<NAME>, CC BY 2.5 <https://creativecommons.org/licenses/by/2.5>, via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Standard_deviation_diagram.svg"><img width="256" alt="Standard deviation diagram" src="https://upload.wikimedia.org/wikipedia/commons/thumb/8/8c/Standard_deviation_diagram.svg/256px-Standard_deviation_diagram.svg.png"></a>
#
#
# 1 standard deviation width represents 68.2% of all the data distribution. We will see more details in the histogram review and how its relates to histogram width-height and gaussian. The variance is equal to standard deviation squared sometimes descibed as $\sigma^2$.
a = [73, 68, 70, 50, 80, 90]
np.mean(a)
np.std(a)
np.mean(a) - 1.96*np.std(a)/np.sqrt(len(a)), np.mean(a) + 1.96*np.std(a)/np.sqrt(len(a)) # confidence interval 95CL%
# From the above mass sample: 68% of human mass is between 59.63kg to 84.03kg
# #### Mode
#
# The mode is the value that appears most often in a set of data values.
# +
import pandas as pd
a = [1, 1, 1, 3, 4, 5, 6]
pd.Series(a).value_counts()
# -
# #### Median
#
# Median is the value separating the higher half from the lower half of a data sample, a population, or a probability distribution.
a = [10, 20, 30, 15, 50, 60, 70]
sorted(a)
np.median(a), np.mean(a)
np.median(a)
# <a title="Cmglee, CC BY-SA 3.0 <https://creativecommons.org/licenses/by-sa/3.0>, via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Visualisation_mode_median_mean.svg"><img width="256" alt="Visualisation mode median mean" src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/33/Visualisation_mode_median_mean.svg/256px-Visualisation_mode_median_mean.svg.png"></a>
# #### Root Mean square (RMS)
#
# Root Mean Square (RMS or rms) is defined as the square root of the mean square (the arithmetic mean of the squares of a set of numbers)
#
#
# $$
# x_{RMS} = \sqrt{\frac{1}{N}\sum_i^N x_i^2} = \sqrt{\frac{1}{N}(x_1^2 + x_2^2 \cdots x_N^2)}
# $$
#
#
# RMS of the pairwise differences of the two data sets can serve as a measure how far on average the error is from 0 (predicted vs measured values).
def rms(a):
return np.sqrt(np.square(a).sum()/len(a))
rms(a)
# #### Examples
# +
import pandas as pd
from statadict import parse_stata_dict
stata_dict = parse_stata_dict('../../ThinkStats2/code/2002FemPreg.dct')
df = pd.read_fwf('../../ThinkStats2/code/2002FemPreg.dat',
names=stata_dict.names,
colspecs=stata_dict.colspecs)
# -
df[:3]
# Some feature description from ThinkStats2 book:
#
# * `caseid` is the integer ID of the respondent.
#
# * `prglength` is the integer duration of the pregnancy in weeks.
#
# * `outcome` is an integer code for the outcome of the pregnancy. The code 1 indicates a live birth.
#
# * `pregordr` is a pregnancy serial number; for example, the code for a respondent’s first pregnancy is 1, for the second pregnancy is 2, and so on.
#
# * `birthord` is a serial number for live births; the code for a respondent’s first child is 1, and so on. For outcomes other than live births, this field is blank.
#
# * `birthwgt_lb` and `birthwgt_oz` contain the pounds and ounces parts of the birth weight of the baby.
#
# * `agepreg` is the mother’s age at the end of the pregnancy.
#
# * `finalwgt` is the statistical weight associated with the respondent. It is a floating-point value that indicates the number of people in the U.S. population this respondent represents.
#
# +
import sys
sys.path.append('/Users/<EMAIL>/Documents/git/ThinkStats2/code')
from nsfg import CleanFemPreg
# perform some cleanup
CleanFemPreg(df)
# +
POUND_TO_KG = 0.453592
df['totalwgt_kg'] = df['totalwgt_lb']*POUND_TO_KG
# -
print('mean:', df['totalwgt_lb'].mean())
print('std:', df['totalwgt_lb'].std())
print('median:', df['totalwgt_lb'].median())
print('rms:', rms(df['totalwgt_lb'].dropna().values))
print('mode:', df['totalwgt_lb'].mode().values[0])
print()
print('-'*40)
print()
print('mean:', df['totalwgt_kg'].mean())
print('std:', df['totalwgt_kg'].std())
print('median:', df['totalwgt_kg'].median())
print('rms:', rms(df['totalwgt_kg'].dropna().values))
print('mode:', df['totalwgt_kg'].mode().values[0])
# ```
#
# value label Total
# 1 LIVE BIRTH 9148
# 2 INDUCED ABORTION 1862
# 3 STILLBIRTH 120
# 4 MISCARRIAGE 1921
# 5 ECTOPIC PREGNANCY 190
# 6 CURRENT PREGNANCY 352
#
# ```
df.outcome.value_counts().sort_index()
df[df['caseid'] == 10229].sort_values('datend')['outcome']
# # Correlations:
#
#
# #### Covariance:
# Covariance is a measure of the joint variability of two random variables.
#
#
# <a title="Cmglee, CC BY-SA 4.0 <https://creativecommons.org/licenses/by-sa/4.0>, via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Covariance_trends.svg"><img width="128" alt="Covariance trends" src="https://upload.wikimedia.org/wikipedia/commons/thumb/a/a0/Covariance_trends.svg/128px-Covariance_trends.svg.png"></a>
#
# **Formal definition:**
#
# $$
# cov(X,Y) = E[(X - E[X])(Y - E[y])]
# $$
#
#
# [from wiki]: For two jointly distributed real-valued random variables {\displaystyle X}X and {\displaystyle Y}Y with finite second moments, the covariance is defined as the expected value (or mean) of the product of their deviations from their individual expected values
#
#
#
# If the (real) random variable pair $(X,Y)$ can take on the values $(x_i, y_i)$ for $i = (1,...,N)$ with equal probabilities $p_i = 1/N$ then the covariance can be equivalently written in terms of the means $E(X)$ and $E(Y)$ as
#
# $$
# cov(X,Y) = \frac{1}{N}\sum_i^N(x_i - E(Y))(y_i - E(Y))
# $$
#
#
# Note: $cov(X,X) = \sigma^2$
X = np.array([1, 2, 3, 4, 5])
Y = 0.3*X
# Y = np.square(X)
X
Y
from matplotlib import pyplot as plt
plt.scatter(X,Y)
plt.xlabel('X')
plt.ylabel('Y')
np.cov(X,Y, bias=True)
# +
# cov(X*X) cov(X*Y)
# cov(Y*X) cov(Y*Y)
# -
X
np.var(X)
np.sqrt(np.var(X))
np.std(X)
# #### Pearson correlation coefficient
#
#
#
# [cite: wiki] It is the ratio between the covariance of two variables and the product of their standard deviations; thus it is essentially a normalised measurement of the covariance, such that the result always has a value between −1 and 1.
#
#
# $$
# \rho_{X,Y} = \frac{cov(X,Y)}{\sigma_X\sigma_Y}
# $$
#
#
# It is a measure of linear correlation between two sets of data.
#
#
# <a title="Kiatdd, CC BY-SA 3.0 <https://creativecommons.org/licenses/by-sa/3.0>, via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Correlation_coefficient.png"><img width="512" alt="Correlation coefficient" src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/34/Correlation_coefficient.png/512px-Correlation_coefficient.png"></a>
#
#
#
# <a title="DenisBoigelot, original uploader was Imagecreator, CC0, via Wikimedia Commons" href="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d4/Correlation_examples2.svg/2560px-Correlation_examples2.svg.png">
# <img width="512" alt="Correlation examples2" src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d4/Correlation_examples2.svg/256px-Correlation_examples2.svg.png"></a>
#
#
# <a href="https://commons.wikimedia.org/wiki/File:Correlation_examples2.svg">DenisBoigelot, original uploader was Imagecreator</a>, CC0, via Wikimedia Commons
#
#
# Take a note: correlation is not causality, see <a href="https://www.tylervigen.com/spurious-correlations">here</a>
# +
X
Y = np.sqrt(X)
# -
X
Y
np.corrcoef(X,Y)
# # Probabilities
a = ['A', 'A', 'B', 'B','B','B','B']
p_theory = 2/7
p_theory
import random
random.choice(a)
# +
probabilities = []
for i in range(100):
values = []
for j in range(10):
extracted = random.choice(a)
if extracted == 'A':
values.append('correct')
else:
values.append('fail')
num = len([i for i in values if i == 'correct'])
den = len(values)
probabilities.append(num/den)
# -
len(probabilities)
plt.hist(probabilities, bins=50, range=(0,1))
# * standard error:
#
# $$
# E_{e} = \frac{\sigma}{\sqrt{N}}
# $$
np.mean(probabilities), np.std(probabilities), np.std(probabilities)/np.sqrt(len(probabilities))
# +
# 0.285732 +/- 0.000141
# +
# import matplotlib.lines as mlines
plt.errorbar([0,1], [0.285732, 2/7], yerr=[0.000141, 0], xerr=None, marker='o', linestyle='')
# plt.plot(-0.04, 2/7, 0.04, 2/7, color='k')
# mlines.Line2D([-0.04,0.04], [2/7,2/7])
# -
2/7
np.mean(probabilities) - np.std(probabilities), np.mean(probabilities) + np.std(probabilities)
# $$
# P = N_A/ N_T
# $$
2/7 + 5/7
0.2857142857142857 + 0.7142857142857143
2/7*5/7
# # Distributions
#
# ## Histograms
a = [5, 6, 3, 4, 4, 4, 6, 6]
_ = plt.hist(a, bins=10, range=(0,10))
plt.grid()
(10 - 0)/10
df['agepreg'].min(), df['agepreg'].max()
df['agepreg'].hist(bins=100, range=(df['agepreg'].min(), df['agepreg'].max()))
df['agepreg'].mean()
df['agepreg'].median()
df['agepreg'].mode()
df['agepreg'].std()
df['agepreg'].mean() - 2*df['agepreg'].std(), df['agepreg'].mean() + 2*df['agepreg'].std()
# # Hypothesis tests
#
# Lets first revisit what statistical tests are reference [here](http://www.stats.ox.ac.uk/~filippi/Teaching/psychology_humanscience_2015/lecture8.pdf)
#
#
from numpy import random
from matplotlib import lines as mlines
obs = random.normal(0, 1, 1000)
# +
hull_hypothesis = 0.05
fig = plt.figure()
ax = fig.add_subplot(111)
_ = ax.hist(obs, range=(-5,5), bins=100, alpha=0.7)
ax.add_line(mlines.Line2D([0,0], [0, 50], color='r'))
# largura a meia altura ~ \sigma
# -
from scipy import stats
stats.ttest_1samp(obs, hull_hypothesis)
# * pvalues < 5% alternative hypothesis cannot be excluded
# * pvalue > 5% null hypothesis cannot be excluded
# - hull hypotheses: first babies have average birth lenght of 40 weeks;
# - alternative hypothesis: first babies have average birth higher from 40 weeks;
# +
first = (df['pregordr'] == 1) & (df['outcome'] == 1)
other = (df['pregordr'] != 1) & (df['outcome'] == 1)
# observe pandas have histogram plotting
plt.figure(figsize=(8,6))
df[other]['prglngth'].hist(bins=100, label='other', density=False)
df[first]['prglngth'].hist(bins=100, label='first', density=False)
plt.xlabel('preg. length [weeks]')
plt.legend()
# -
df[first]['prglngth'].mean(), df[first]['prglngth'].mean()
df[other]['prglngth'].mean(), df[other]['prglngth'].std()
err_first = df[first]['prglngth'].std()/np.sqrt(len(df[first]['prglngth']))
err_other = df[other]['prglngth'].std()/np.sqrt(len(df[other]['prglngth']))
df[other].shape[0], df[first].shape[0]
err_first, err_other
plt.errorbar([0,1], [df[first]['prglngth'].mean(), df[other]['prglngth'].mean()], yerr=[err_first, err_other], xerr=None, marker='o', linestyle='')
# +
import seaborn as sns
plt.figure(figsize=(8,6))
sns.kdeplot(data=df[other], x="prglngth", label='other')
sns.kdeplot(data=df[first], x="prglngth", label='first')
plt.xlabel('preg. length [weeks]')
plt.legend()
# -
import scipy
scipy.stats.ttest_1samp(df[other]['prglngth'].values, 40)
scipy.stats.ttest_1samp(df[first]['prglngth'].values, 40)
# - $H_0$ first babies have equal pregnance length that of others babies
# - $H_1$ first babies have different pregnance length that of others babies
stats.ttest_ind(df[first]['prglngth'].values, df[other]['prglngth'].values)
# - the null hypothesis cannot be rejected.
# - H_0 first babies does not have higher pregnance lenght that of other babies;
# - H_1 first babies have higher pregnance length that of other babies;
# Please read as ref [this](https://machinelearningmastery.com/statistical-hypothesis-tests-in-python-cheat-sheet/)
# ## Gaussian distribution
#
#
# If $X$ is Normally distributed with mean $\mu$ and standard deviation $\sigma$, we
# write
#
# $$
# X \sim N(\mu, \sigma^2)
# $$
#
#
# The Probability Density Function (PDF) of the Normal distribution is given by:
# $$
# g(x) = \frac{1}{\sqrt{2\pi}\sigma}e^{\frac{(x-\mu)^2}{2\sigma^2}}
# $$
#
# $z$ is the chosen values.
#
# $$
# P(Z<z) = \int_{-\inf}^{z} g(x) = \int_{-\inf}^{z} \frac{1}{\sqrt{2\pi}\sigma}e^{\frac{(x-\mu)^2}{2\sigma^2}}
# $$
#
# Numerically
# +
from matplotlib import lines as mlines
values = np.random.normal(0, 1, 100000)
fig = plt.figure()
ax = fig.add_subplot(111)
z = 1
ax.add_line(mlines.Line2D([z,z], [0, 0.5], color='r'))
hist_values, bins, _ = ax.hist(values, range=(-5,5), bins=200, density=True, alpha=0.6)
# -
widths = np.diff(bins[bins<z])
widths = list(widths) + [widths[0]]
area = sum(np.array(widths)*hist_values[:bins[bins<z].shape[0]])
area
# +
from matplotlib import lines as mlines
values = np.random.normal(70, 10, 100000)
fig = plt.figure()
ax = fig.add_subplot(111)
hist_values, bins, _ = ax.hist(values, range=(0,140), bins=200, density=True, alpha=0.6)
ax.add_line(mlines.Line2D([70,70], [0, ax.get_ylim()[1]], color='r'))
ax.add_line(mlines.Line2D([70-10,70-10], [0, ax.get_ylim()[1]], color='b'))
ax.add_line(mlines.Line2D([70+10,70+10], [0, ax.get_ylim()[1]], color='b'))
# -
values.mean(), values.std()
# normalize the distribution to make it centered in zero and standard deviation = 1
values = (values-values.mean())/values.std()
values
fig = plt.figure()
ax = fig.add_subplot(111)
hist_values, bins, _ = ax.hist(values, range=(-5,5), bins=200, density=True, alpha=0.6)
ax.add_line(mlines.Line2D([0,0], [0, ax.get_ylim()[1]], color='r'))
ax.add_line(mlines.Line2D([0-1,0-1], [0, ax.get_ylim()[1]], color='b'))
ax.add_line(mlines.Line2D([0+1,0+1], [0, ax.get_ylim()[1]], color='b'))
# ## Refresh:
#
#
# * standard deviation;
# * standard error: $\frac{\sigma}{\sqrt{N}}$
values = np.random.normal(0,1,10000)
values
values.std()/np.sqrt(len(values))
# +
z = 1.96
fig = plt.figure()
ax = fig.add_subplot(111)
hist_values, bins, _ = ax.hist(values, range=(-5,5), bins=200, density=True, alpha=0.6)
ax.add_line(mlines.Line2D([0,0], [0, ax.get_ylim()[1]], color='r'))
ax.add_line(mlines.Line2D([0-z,0-z], [0, ax.get_ylim()[1]], color='b'))
ax.add_line(mlines.Line2D([0+z,0+z], [0, ax.get_ylim()[1]], color='b'))
# +
z = 1.96
mu = 0
standard_error = values.std()/np.sqrt(values.shape[0])
fig = plt.figure()
ax = fig.add_subplot(111)
hist_values, bins, _ = ax.hist(values, range=(-5,5), bins=200, density=True, alpha=0.6)
ax.add_line(mlines.Line2D([0,0], [0, ax.get_ylim()[1]], color='r'))
ax.add_line(mlines.Line2D([mu-z*standard_error,mu-z*standard_error], [0, ax.get_ylim()[1]], color='b'))
ax.add_line(mlines.Line2D([mu+z*standard_error,mu+z*standard_error], [0, ax.get_ylim()[1]], color='b'))
# -
values.mean() - z*standard_error, values.mean() + z*standard_error
# +
means = list()
for i in range(10000):
values_ = np.random.normal(0, 1, 1000)
means.append(values_.mean())
# +
z = 1.96
mu = 0
standard_error = values.std()/np.sqrt(values.shape[0])
fig = plt.figure()
ax = fig.add_subplot(111)
hist_values, bins, _ = ax.hist(means, range=(-0.5,0.5), bins=200, density=True, alpha=0.6)
ax.add_line(mlines.Line2D([0,0], [0, ax.get_ylim()[1]], color='r'))
ax.add_line(mlines.Line2D([mu-z*standard_error,mu-z*standard_error], [0, ax.get_ylim()[1]], color='b'))
ax.add_line(mlines.Line2D([mu+z*standard_error,mu+z*standard_error], [0, ax.get_ylim()[1]], color='b'))
# -
pressao = np.random.normal(14, 5, 10000)
pressao[:10].mean()
pressao.mean() - 1.96*pressao.std()/np.sqrt(len(pressao)), pressao.mean() + 1.96*pressao.std()/np.sqrt(len(pressao))
import scipy
scipy.stats.ttest_1samp(df[other]['prglngth'].values, 40)
mean = df[other]['prglngth'].mean()
standard_error = df[other]['prglngth'].std()/np.sqrt(df[other]['prglngth'].shape[0])
mean, standard_error
# 95% confidence interval for the expected value
mean - 1.96*standard_error, mean + 1.96*standard_error
scipy.stats.ttest_1samp(df[other]['prglngth'].values, 38.47672347896686)
scipy.stats.ttest_1samp(df[other]['prglngth'].values, 38.61670212656948)
df[first]['prglngth'].var(), df[other]['prglngth'].var()
stats.ttest_ind(df[first]['prglngth'].values, df[other]['prglngth'].values, equal_var=False)
plt.hist(df[first]['prglngth'].values, bins=1000, range=(0, 50), alpha=0.5, label='first', density=True, cumulative=True, histtype='step')
plt.hist(df[other]['prglngth'].values, bins=1000, range=(0, 50), alpha=0.5, label='other', density=True, cumulative=True, histtype='step')
stats.ks_2samp(df[first]['prglngth'].values, df[other]['prglngth'].values)
stats.shapiro(df[first]['prglngth'].values)
stats.shapiro(df[other]['prglngth'].values)
| statistics/rw_statistics_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Pandas - Basic (1)**
#
# #### Pandas란
# - NumPy를 기반으로 개발된 데이터 분석을 위한 쉽고 성능 좋은 오픈소스 python 라이브러리
# - 많은 사람들에게 익숙한 행x열로 이뤄진 테이블 형태로 데이터를 다룰 수 있게 한다.
# - R과 비슷하게 데이터를 다룰 수 있게 되는데, R에 비해서는 속도가 훨씬 빠르다.
# - 공식문서: http://pandas.pydata.org
#
#
# #### Pandas의 data type
# 1. Series
# - 하나의 변수에 대한 데이터가 모인 1차원 데이터타입
# - index, value로 이루어짐
# 2. DataFrame
# - 여러 변수에 대한 series 여러 개가 모인 2차원 데이터타입, '표'라고 생각하면 된다.
# - index, column, value로 이루어짐
# - 각 column은 series로 구성됨
# - DataFrame은 여러개의 Series가 column을 구성하는 모양으로 만들어짐
#
# #### 다룰 내용
# Series와 DataFrame을 다루는 기초에 관해 두 편으로 나누어 정리해보자. 우선 여기서는 아래의 내용을 정리한다.
#
# 1. Series
# - Series 만들기
# - Series 데이터 보기
# - indexing과 slicing
# - Series 데이터 연산 및 Series 다루기
#
#
# 2. DataFrame
# - Create
# - Insert
# - append
# - concat
# ## **0. 설치와 import**
# - 설치: `$ pip install pandas`
# - import: numpy와 함께, alias를 `pd`로 import하는 것이 컨벤션
import numpy as np
import pandas as pd
# ## **1. Series**
# - index와 value로 되어 있는 데이터 형태 (index는 각 샘플에 붙는 번호라고 생각)
# - series에는 하나의 데이터타입만 사용 가능 (하나의 column은 같은 데이터타입을 가짐)
# - 한 변수에 대한 데이터 값들이라고 생각하면 된다.
# + [markdown] toc-hr-collapsed=true
# ### 1.1 Series 만들기
# -
# ##### Series 생성 방법
# - `pd.Series(data)`: data로는 list, array, dict. 등이 들어감
# - index: default로 0부터 1씩 증가하는 숫자로 생성됨
# 0에서 9까지의 숫자를 랜덤하게 5개 발생시켜 Series를 만들기
data = pd.Series(np.random.randint(10, size=5))
data
# ##### index 설정하기
# - index parameter로 list 데이터를 넘기면 인덱스를 설정함
# - 데이터 개수와 인덱수의 개수를 맞추어서 넣어야 함
data = pd.Series(np.random.randint(10, size=5), index=['A','B','C','D','E'])
data
# Series의 이름과 index의 이름을 붙여 줄수 있음
data.name = "rand_num"
data.index.name = "idx"
data
# + [markdown] toc-hr-collapsed=true
# ### 1.2 Series 데이터 보기
# -
# #### (1) `index`, `values`, `items`
# `index`, `values`: index와 values(데이터) 호출
print(data.index)
print(data.values)
# items를 이용해 index, val를 반복문에서 사용가능
for idx, val in data.items():
print(idx, val)
[(idx, val) for idx, val in data.items()]
# #### (2) index 속성으로 데이터 호출
# - 문자열 인덱스만 사용 가능
data.A, data.C
# 아래처럼 숫자를 str으로 index에 넣어줘도 그 index로는 호출할 수 없음
data_tmp = pd.Series(np.random.randint(10, size=5), index=['1','2','3','4','5'])
data.1
# ### 1.3 indexing과 slicing
# indexing
print(data[0])
print(data["B"])
# index로 여러개의 데이터 출력
data[["B","C","E"]]
# offset으로 slicing
data[1:3]
# step = -1: 역순으로 호출
data[::-1]
# ### 1.4 Series 데이터 연산 및 Series 다루기
# 연산: broadcasting으로 계산됨
data * 10
# series 끼리 계산 - index가 mapping 되어 연산
print("data: \n{}\n".format(data))
data2 = pd.Series({"D":7, "E":5, "F":9})
print("data2: \n{}\n".format(data2))
result = data + data2
print("data + data2: \n{}".format(result))
# 비교 연산 가능
data > 3
# 필터링: 3이상 되는 데이터 출력
data[data > 3] # []안의 값이 True인 값만 출력됨
# null 데이터(NaN) 제거
print(result.notnull(), "\n") # notnull() 함수: True/False를 return함
result = result[result.notnull()] # notnull이 True인 경우만 필터링함
print(result, "\n")
# ## **2. Dataframe**
# - series(index, value), column으로 이루어진 데이터 타입
# - table 모양으로 구성 (row와 column이 있음)
# + [markdown] toc-hr-collapsed=true
# ### 2.1 Create
# - `pd.DataFrame(data)`
# -
# #### (1) 빈 컬럼 생성 후 list 데이터 추가
df = pd.DataFrame(columns=["Email", "Name"])
df
df["Name"] = ["fcamp", "dss"]
df["Email"] = ["<EMAIL>", "<EMAIL>"]
df
# 하나의 컬럼 = Series
df["Name"]
# #### (2) Dict 데이터로 생성
name = ["fcamp", "dss"]
email = ["<EMAIL>", "<EMAIL>"]
dic = {"Name":name, "Email":email}
df = pd.DataFrame(dic) # dic을 DataFrame에 넣어줌
df
# #### (3) index 추가하기
# - index parameter에 index명 list를 넣어줌
# - index를 날짜로도 쓸 수 있음
# - index 값은 중복이 되지 않음 (unique 값만 올 수 있음)
index_list = ["first", "second"]
data = {"Email": ["<EMAIL>", "<EMAIL>"], "Name": ["fcamp", "dss"]}
df = pd.DataFrame(data, index=index_list)
df
# 데이터프레임에 대한 인덱스, 컬럼, 값 데이터(행렬로 가져옴) 가져오기
print("index: {}\n".format(df.index))
print("column: {}\n".format(df.columns))
print("values: \n{}".format(df.values))
# + [markdown] toc-hr-collapsed=true
# ### 2.2 Insert
# + [markdown] toc-hr-collapsed=true
# #### (1) Insert rows
# -
# ##### dataframe의 row와 column에 접근하기
# - row는 column과 달리 접근 method가 있음
# - row 접근: `.loc[]` (for location)
# - column 접근: `[column 이름]` - column명은 숫자도 가능 (Series의 index와 달리)
data = {"Email": ["<EMAIL>", "<EMAIL>"], "Name": ["fcamp", "dss"]}
df = pd.DataFrame(data)
df
# row 접근
df.loc[1] # series 형태로 나옴
# column 접근
df["Email"]
# ##### dataframe row에 데이터 넣기
# 특정 row 지정해서 데이터 넣기 (dict 형태로)
df.loc[2] = {"Email":"<EMAIL>", "Name":"data"}
df
# 맨 마지막 행에 자동으로 데이터 넣기
df.loc[len(df)] = {"Email":"<EMAIL>", "Name":"science"}
df
# + [markdown] toc-hr-collapsed=true
# #### (2) Insert columns
# -
# 컬럼 추가
df["Address"] = ""
df
# 컬럼 데이터 추가: row수를 맞춰서 추가 or 하나의 데이터를 넣어주면 broadcasting
df["Address"] = ["Seoul", "Busan", "Jeju", "Deagu"]
df
# ##### apply: 함수 사용해서 column 데이터 넣기
# +
# name 데이터의 문자열 길이를 세서 name(count) 형태로 출력되는 새로운 column 생성
def name_count(name):
return "{}({})".format(name, len(name))
name_count("hyeshin")
# -
# "Name_count"라는 column을 생성, "Name" column에 name_count 함수를 apply해서 데이터를 넣음
df["Name_count"] = df["Name"].apply(name_count)
df
# lambda 사용해서 column 데이터 넣기
df["Address_Count"] = df["Address"].apply(lambda addr: "{}({})".format(addr, len(addr)))
df
# + [markdown] toc-hr-collapsed=true
# ### 2.3 append
# - 두 개의 DataFrame을 이어 붙일 수 있음
# - 데이터를 붙이는 여러 방법(e.g. `concat`) 중 한 가지
# +
# 사람 이름과 나이가 들어간 데이터를 만들기
import random, string
def get_name():
names = ['Adam', 'Alan', 'Alex', 'Alvin', 'Andrew', 'Anthony', 'Arnold', 'Jim', 'Baldy','Peter']
return random.choice(names)
def get_age(start=20, end=40):
return random.randint(start, end) # random 모듈의 randint는 end가 포함됨
def make_data(rows=10): # default로 데이터 10개를 설정함
datas = []
for _ in range(rows): # '_"를 쓰는 이유는 이름 중복 피하려고(컨벤션)
data = {"Age":get_age(), "Name":get_name()}
datas.append(data)
return datas
# +
# 참고 random module의 randint는 numpy의 random.randint와 달리 end 값이 포함
import random, string
r = random.randint(0, 10)
print("random module randint: {}".format(r))
na = np.random.randint(0, 10, size=(3, 3))
print("Numpy random.randint: \n{}".format(na))
# -
make_data() # 중복되는 이름도 나올 수 있음
data1 = make_data()
df1 = pd.DataFrame(data1) # dict.를 요소로 가진 list로도 DataFrame을 만들 수 있음
df1
data2 = make_data()
df2 = pd.DataFrame(data2)
df2
# append: df1에 df2 데이터를 추가하기
df3 = df1.append(df2)
df3
# 새로운 인덱스 생성
df3.reset_index() # 원래 인덱스가 그대로 남게 됨 (drop=True: 기존 index 사라짐)
# ##### reset_index의 옵션
# - `inplace=True`: 수정된 데이터가 해당 변수에 바로 적용
# - `inplace=True`를 사용하지 않으면 결과 데이터를 받아서 저장해야 함
df3.reset_index(drop=True, inplace=True)
df3
# append 하면서 바로 인덱스를 새로 생성하기
df1.append(df2, ignore_index=True) # 기존의 index가 무시되고 새로 생성
# + [markdown] toc-hr-collapsed=true
# ### 2.4 concat
# - `pd.concat([*DataFrame])`
# -
# #### (1) concat rows
# df1 + df2 (rows로 합치기)하고 reset_index를 이용하여 index를 재정렬
df3 = pd.concat([df1, df2]).reset_index(drop=True)
df3
# #### (2) concat columns
# 인자로 axis=1을 설정하여 df1과 df2 데이터의 컬럼을 합쳐줌
df4 = pd.concat([df3, df1], axis=1) # join='outer'가 default
df4
# df3는 20개였고 df1은 10개 → 결과 20개 (10개는 NaN으로 나타남)
df4 = pd.concat([df3, df1], axis=1, join='inner')
df4
# df3는 20개였고 df1은 10개 → 결과 10개 (양 쪽에 모두 있는 index만 나타남)
# #### 참고자료
# - 패스트캠퍼스, ⟪데이터사이언스스쿨 8기⟫ 수업자료
| python_24_pandas_basic_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3.8 (venv)
# language: python
# name: py38-venv
# ---
# # Parallax effect
#
# This tutorial present how the volumetric absorption of photons can be taken into account to explain the peak shift at large incidence angle and quantify parallax the effet.
#
# This effect apparently disturbes the calibration at very large scattering angle and is likely to jeopardize PDF measurements.
#
# 
#
# Let's consider the sensor of a detector and assume it is a thick slab of material. This sensor is characterized by its photon absorption, but since it is not that obvious to find the numerical values, we will consider the **thickness** and the **efficiency** of the sensor (measured normaly). The efficiency depends on the incident photon energy which is assumed monochromatic.
#
# The volumic absorption follows a first order (exponential) decay, assymmetric, which is the cause of the apparent shift of peaks in the calibration procedure. This tutorial tries to quantify the effect.
#
# For most of the tutorial, the Perkin-Elmer flat panel detector is considered, it has a dopped CsI sintillator of 200µm which has an apparent efficiency of 80% at 35keV. The pixel size is 100µm.
#
# The incoming beam is considered with a Gaussian shape, but circular and square signals should be considered as well. We will assume the FWHM of the beam is 1mm and will tune this parameter. It correspond in first approximation to the sample size.
# %matplotlib nbagg
import time
start_time = time.perf_counter()
import numpy
from matplotlib.pyplot import subplots
import numexpr
import scipy.integrate, scipy.signal
from math import sin, cos, pi, log, sqrt
EPS = numpy.finfo("float64").eps
# +
class Decay:
OVER = 64 # Default oversampling value
def __init__(self, thickness, efficiency):
"""Class to simulate the parallax effect
:param thickness: thickness of the sensible layer, in meter or mm, µm...
:param efficiency: efficiency for the sensor material between 0 and 1
"""
self.thickness = float(thickness)
self.efficiency = float(efficiency)
self.lam = - log(1.0-efficiency)/thickness
self.formula = numexpr.NumExpr("where(x<0, 0.0, l*exp(-l*x))")
def __call__(self, x):
"Calculate the absorption at position x"
return self.formula(self.lam, x)
def integrate(self, x):
"""Integrate between 0 and x
:param x: length of the path, same unit as thickness
"""
return scipy.integrate.quad(s, 0.0, x)
def test(self):
"""Validate the formula for lambda
sum(decay) between 0 and thickness is efficiency"""
value, error = self.integrate(self.thickness)
assert abs(value - self.efficiency) < error
def absorption(self, angle, over=None):
"""Calculate the absorption along the path for a beam inclined with the given angle
:param angle:
:return position (along the detector), absorption (normalized)
"""
over = over or self.OVER
angle_r = numpy.deg2rad(angle)
length = self.thickness/cos(angle_r)
step = length/over
pos = numpy.linspace(0, length, over)
decay = self.__call__(pos)
decay /= decay.sum() # Normalize the decay
pos *= sin(angle_r) # rotate the decay to have it in the detector plan:
return pos, decay
def gaussian(self, width, over=None):
"""Model the beam as a gaussian
:param width: FWHM of the gaussian curve
:param over: oversampling factor, take that many points to describe the peak
:return: position, intensity (normalized)
"""
over = over or self.OVER
if width<EPS:
print("Warning, width too small")
width = EPS
step = width / over
sigma = width/(2.0*sqrt(2.0*log(2.0)))
nsteps = 2*int(3*sigma/step+1) + 1
pos = (numpy.arange(nsteps) - nsteps//2) * step
peak = numexpr.evaluate("exp(-pos**2/(2*(sigma**2)))")
peak /= peak.sum()
return pos, peak
def square(self, width, over=None):
"""Model the beam as a square signal
:param width: width of the signal
:param over: oversampling factor, take that many points to describe the peak
:return: position, intensity (normalized)
"""
over = over or self.OVER
if width<EPS:
print("Warning, width too small")
width = EPS
step = width / over
nsteps = 2*int(2*width/step+1) + 1
pos = (numpy.arange(nsteps) - nsteps//2) * step
peak = numexpr.evaluate("where(abs(pos)<=width/2, 1.0, 0.0)")
peak /= peak.sum()
return pos, peak
def circle(self, width, over=None):
"""Model the beam as a circular signal
:param width: Diameter of the beam
:param over: oversampling factor, take that many points to describe the peak
:return: position, intensity (normalized)
"""
over = over or self.OVER
if width<EPS:
print("Warning, width too small")
width = EPS
step = width / over
nsteps = 2*int(width/step+2) + 1
pos = (numpy.arange(nsteps) - nsteps//2) * step
peak = numexpr.evaluate("where(abs(pos)<=width/2, sqrt(1.0-(2.0*pos/width)**2), 0.0)")
peak /= peak.sum()
return pos, peak
def convolve(self, width, angle, beam="gaussian", over=None):
"""Calculate the line profile convoluted with parallax effect
:param width: FWHM of the peak, same unit as thickness
:param angle: incidence angle in degrees
:param over: oversampling factor for numerical integration
:return: position, intensity(position)
"""
over = over or self.OVER
angle_r = numpy.deg2rad(angle)
pos_dec, decay = self.absorption(angle, over)
peakf = self.__getattribute__(beam)
pos_peak, peak = peakf(width/cos(angle_r), over=over)
#Interpolate grids ...
pos_min = min(pos_dec[0], pos_peak[0])
pos_max = max(pos_dec[-1], pos_peak[-1])
step = min((pos_dec[-1] - pos_dec[0])/(pos_dec.shape[0]-1),
(pos_peak[-1] - pos_peak[0])/(pos_dec.shape[0]-1))
if step<EPS:
step = max((pos_dec[-1] - pos_dec[0])/(pos_dec.shape[0]-1),
(pos_peak[-1] - pos_peak[0])/(pos_dec.shape[0]-1))
nsteps_2 = int(max(-pos_min, pos_max)/step + 0.5)
pos = (numpy.arange(2*nsteps_2+1) - nsteps_2) * step
big_decay = numpy.interp(pos, pos_dec, decay, left=0.0, right=0.0)
dsum = big_decay.sum()
if dsum == 0:
big_decay[numpy.argmin(abs(pos))] = 1.0
else:
big_decay /= dsum
big_peak = numpy.interp(pos, pos_peak, peak, left=0.0, right=0.0)
return pos, scipy.signal.convolve(big_peak, big_decay, "same")
def plot_displacement(self, width, angle, beam="gaussian", ax=None):
"""Plot the displacement of the peak depending on the FWHM and the incidence angle"""
if ax is None:
fig,ax = subplots()
ax.set_xlabel("Radial displacement on the detector (mm)")
c = self.absorption(angle)
ax.plot(*c, label="Absorption")
peakf = self.__getattribute__(beam)
c = peakf(width)
ax.plot(*c, label=f"peak w={width} mm")
c = peakf(width/cos(angle*pi/180))
ax.plot(*c, label=f"peak w={width} mm, inclined")
c = self.convolve(width, angle, beam=beam)
ax.plot(*c, label="Convolution")
idx = numpy.argmax(c[1])
maxi = self.measure_displacement(width, angle, beam=beam)
ax.annotate(f"$\delta r$={maxi:.3f}", (maxi, c[1][idx]),
xycoords='data',
xytext=(0.8, 0.5), textcoords='axes fraction',
arrowprops=dict(facecolor='black', shrink=0.05),
horizontalalignment='right', verticalalignment='top',)
ax.set_title(f"Profile {beam}, width: {width}mm, angle: {angle}°")
ax.legend()
return ax
def measure_displacement(self, width, angle, beam="gaussian", over=None):
"""Measures the displacement of the peak due to parallax effect"""
over = over or self.OVER
x,y = self.convolve(width, angle, beam=beam, over=over)
ymax = y.max()
idx_max = numpy.where(y==ymax)[0]
if len(idx_max)>1:
return x[idx_max].mean()
idx = idx_max[0]
if idx>1 or idx<len(y)-1:
#Second order tailor expension
f_prime = 0.5*(y[idx+1]-y[idx-1])
f_sec = (y[idx+1]+y[idx-1]-2*y[idx])
if f_sec == 0:
print('f" is null')
return x[idx]
delta = -f_prime/f_sec
if abs(delta)>1:
print("Too large displacement")
return x[idx]
step = (x[-1]-x[0])/(len(x)-1)
return x[idx] + delta*step
return x[idx]
# s = Decay(0.2, 0.8) #200µm, 80% efficiency
s = Decay(0.45, 0.35) #450µm, 35% efficiency 450µ Si @ 20keV
s.test()
# -
fig, ax = subplots(3, 3, figsize=(15,15))
for id0, shape in enumerate(("gaussian", "circle", "square")):
for id1, w in enumerate((0.1, 0.4, 1.0)):
p=s.plot_displacement(w, 60, beam=shape, ax=ax[id0, id1])
# +
width = 1
angles = numpy.arange(90)
displ_g = [s.measure_displacement(width, a, beam="gaussian", over=1024) for a in angles]
displ_c = [s.measure_displacement(width, a, beam="circle", over=1024) for a in angles]
displ_s = [s.measure_displacement(width, a, beam="square", over=1024) for a in angles]
fig,ax = subplots()
ax.plot(angles, displ_g, label="Gaussian profile")
ax.plot(angles, displ_c, label="Circular profile")
# ax.plot(angles, displ_s, label="Square profile")
ax.set_title("450µm Si @ 20keV, 35% efficiency")
# ax.set_title("200µm CsI @ 35keV, 80% efficiency")
fig.suptitle(f"Displacement of beam ({width} mm) due to parallax")
ax.set_ylabel("Displacement (mm)")
ax.set_xlabel("Incidence angle (°)")
ax.plot(angles, numpy.sin(numpy.deg2rad(angles))/s.lam, label=r"Theory $sin(\alpha)/\mu$")
ax.legend()
# fig.savefig("Perkin.png")
fig.savefig("Pilatus.png")
pass
# -
# ## Conclusion
#
# The parallax effect induces a displacement of the ring, the numerical value is almost proportional to the scattering angle for all useable incidence angles (<60°) and represents at maximum one pixel (100µm).
#
# This tutorial can directly be re-run for photon counting detectors like the Pilatus detector manufactured by Dectris, with the efficiency curves provided by the manufacturer:
# 
# 
print(f"Execution time: {time.perf_counter()-start_time:.3f}s")
1/s.lam
| doc/source/usage/tutorial/ThickDetector/Parallax_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import lightgbm as lgbm
from sklearn.model_selection import KFold
from sklearn.metrics import f1_score, roc_auc_score
from tqdm.auto import tqdm
import warnings
warnings.filterwarnings('ignore')
# -
train = pd.read_csv('input/train.csv')
test = pd.read_csv('input/test.csv')
sample_submit = pd.read_csv('input/sample_submit.csv')
train.head(10)
train.info()
# +
train['main_active_loan_ratio'] = train['main_account_active_loan_no']/train['main_account_loan_no']
train['main_active_loan_ratio'] = train['main_active_loan_ratio'].fillna(0)
train['active_overdue_loan_ratio'] = train['main_account_active_loan_no']/train['main_account_loan_no']
train['active_overdue_loan_ratio'] = train['main_active_loan_ratio'].fillna(0)
# -
train['active_overdue_loan_ratio']
# histogram
plt.style.use('seaborn')
# sns.set_style("white")
# sns.set_color_codes(palette='deep')
f, ax = plt.subplots(figsize=(8, 7))
#Check the new distribution
plt.hist(train['loan_to_asset_ratio'],bins=100, color="b");
# ax.xaxis.grid(False)
ax.set(ylabel="")
ax.set(xlabel="SalePrice")
ax.set(title="SalePrice distribution")
# sns.despine(trim=True, left=True)
plt.show()
# Stats
from scipy.stats import skew, norm
from scipy.special import boxcox1p
from scipy.stats import boxcox_normmax
# Skew and kurt
print("Skewness: %f" % train['loan_to_asset_ratio'].skew())
print("Kurtosis: %f" % train['loan_to_asset_ratio'].kurt())
f, ax = plt.subplots(figsize=(8, 7))
#Check the new distribution
plt.scatter(train['loan_to_asset_ratio'],train['loan_default'],color="b");
ax.xaxis.grid(False)
ax.set(ylabel="")
ax.set(xlabel="SalePrice")
ax.set(title="SalePrice distribution")
# sns.despine(trim=True, left=True)
plt.show()
| additional_resources/additional_datasets/car_loan/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # MAT281
#
# ## Aplicaciones de la Matemática en la Ingeniería
# + [markdown] slideshow={"slide_type": "slide"}
# ## ¿Qué contenido aprenderemos?
# * Conocimiento básico sobre datos tabulares.
# * Etiqueta para la tabla.
# * Pivotear una tabla.
# * Despivotear una tabla.
# + [markdown] slideshow={"slide_type": "slide"}
# ## ¿Porqué aprenderemos eso?
# + [markdown] slideshow={"slide_type": "fragment"}
# **1**. Resulta ser la tarea más habitual en la manipulación de datos. 25% del tiempo, mínimo.
# + [markdown] slideshow={"slide_type": "fragment"}
# **2**. Visto en twitter:
#
# * Me: *So, first question, what's the oposite of pivoting a table?*
# * Interviewee: *Ehhh, melting a table?*
# * Me: *Hired!*
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
#
# Los datos tabulares, como su nombre lo indica, son aquellos que han sido representarse mediante una tabla que relaciona sus atributos y valores.
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
#
# ¿Existen datos que no pueden representarse mediante una datos tabulares?
# + [markdown] slideshow={"slide_type": "fragment"}
# No lo sé.
# -
# En general, diría que los datos siempre pueden almacenarse como relaciones entre tablas, pero donde el problema que emerge para algunos problemas más complejos es el de la eficiencia en la búsqueda o almacenamiento de la información.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
#
# Típicamente, existen ciertas columnas que son las columnas "identificadoras" y para las cuales *debiera* existir una única fila en la tabla.
#
# Por otra parte, existen columnas de datos, para las cuales pueden agregarse más información.
#
#
#
# |semestre | rut_estudiante|curso|prueba|nota|
# |-----------|----|--------|----|----|
# |2018-1S|15000000-6|mat281|Certamen_1|100|
# |2016-1S|15000000-6|mat281|Certamen_1|100|
# |2018-1S|18000000-9|mat281|Certamen_1|100|
# |2018-1S|15000000-6|fis120|Certamen_2|100|
#
# ¿Qué columnas son identificadoras en el ejemplo anterior? ¿Cuáles son los valores asociados?
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
#
# Consideremos el siguiente ejemplo:
#
# |semestre | rut_estudiante|curso|prueba|nota|
# |-----------|----|--------|----|----|
# |2018-1S|15000000-6|mat281|Certamen_1|100|
# |2016-1S|15000000-6|mat281|Certamen_1|100|
# |2018-1S|18000000-9|mat281|Certamen_1|100|
# |2018-1S|15000000-6|fis120|Certamen_2|100|
# |2018-1S|15000000-6|mat281|Certamen_1|10|
# |2016-1S|15000000-6|mat281|Certamen_1|-10|
#
# ¿La tabla está correcta?
# + [markdown] slideshow={"slide_type": "fragment"}
# En el ejemplo anterior existen conflictos de información entre las distintas filas de la tabla, que es necesario resolver en el procesamiento de la información.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
#
# Consideremos el siguiente ejemplo:
#
# |semestre | nombre_estudiante|curso|prueba|nota|
# |-----------|----|--------|----|----|
# |2018-1S|<NAME>|mat281|Certamen_1|100|
# |2016-1S|<NAME> |mat281|Certamen_1|100|
# |2018-1S|<NAME> |mat281|Certamen_1|100|
# |2018-1S|<NAME>. |fis120|Certamen_2|100|
#
# ¿La tabla está correcta?
# + [markdown] slideshow={"slide_type": "fragment"}
# En el ejemplo anterior una de las columnas que debe funcionar como identificador del estudiante no está normalizada y no permite una identificación única del estudiante.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
# ### wide format versus long format
#
# Por ejemplo, el conjunto de datos [Zoo Data Set](http://archive.ics.uci.edu/ml/datasets/zoo) presenta las características de diversos animales, de los cuales presentamos las primeras 5 columnas.
#
# |animal_name|hair|feathers|eggs|milk|
# |-----------|----|--------|----|----|
# |antelope|1|0|0|1|
# |bear|1|0|0|1|
# |buffalo|1|0|0|1|
# |catfish|0|0|1|0|
#
# La tabla así presentada se encuentra en "wide format", es decir, donde los valores se extienden a través de las columnas.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
# ### Wide format versus Long format
#
# Sería posible representar el mismo contenido anterior en "long format", es decir, donde los mismos valores se indicaran a través de las filas:
#
# |animal_name|characteristic|value|
# |-----------|----|--------|
# |antelope|hair |1|
# |antelope|feathers|0|
# |antelope|eggs|0|
# |antelope|milk|1|
# |...|...|...|...|..|
# |catfish|hair |0|
# |catfish|feathers|0|
# |catfish|eggs|1|
# |catfish|milk|0|
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Datos tabulares
# ### Wide format versus Long format
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Etiqueta Datos tabulares
#
# Antes de pasar a la tabulación de los datos, hablaremos de un tema pocas veces mencionado. ¿Cómo debemos dar formato a las tablas que compartimos? ¿Existe una *etiqueta* de datos tabulares?
#
# El formato y diseño de una tabla es crucial: si se hace bien, los datos son fáciles de revisar y comparar. Si se hace mal, dificulta y oscurece el entendimiento de la información.
#
# Al igual que con las visualizaciones, la finalidad de un buen formato de tablas es que la información pueda ser consumida de manera correcta y rápida por una persona externa, sin posibilidad de malinterpretaciones.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Etiqueta de Datos tabulares
# Las reglas básicas a considerar:
# 1. La fuente de todos los números debe ser uniforme y permitir comparaciones.
# 2. Las columnas deben tener separador de miles y, dentro de lo posible, la misma cantidad de decimales. Mientras menos, mejor.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Etiqueta de Datos tabulares
#
# 3. Mencionar las unidades en el nombre de la columna cuando sea relevante.
# 1. Datos numéricos deben estar alineados a la derecha.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Etiqueta de Datos tabulares
#
# 5. Datos de texto deben estar alineados a la izquierda.
# 1. Los títulos de las columnas deben estar alineados con sus datos.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Etiqueta de Datos tabulares
#
# 7. Usar el color de manera apropiada y frugalmente.
# 1. Entregar valores de agrupación según sea necesario.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Datos tabulares
# Ejemplo mal formato:
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Datos tabulares
# Ejemplo un poco mejor formato:
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Datos tabulares
# Ejemplo formato final:
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Pivoteando una tabla
#
# El pivoteo de una tabla corresponde al paso de una tabla desde el "long format" al "wide format". Típicamente esto se realiza para poder comparar los valores que se obtienen para algún registro en particular, o para utilizar algunas herramientas de visualización básica que requieren dicho formato.
# -
import pandas as pd
import os
df = pd.read_csv(os.path.join("data","terremotos.csv"), sep=",")
df["Pais"] = df["Pais"].str.strip()
df.head()
# + slideshow={"slide_type": "slide"}
df.describe(include="all")
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Pivoteando una tabla
#
# La tabla anterior no tiene bien definidas sus columnas de identificadores, puesto que en un año podría existir más de un terremoto por año:
# -
df.groupby(["Año","Pais"]).count()
# + [markdown] slideshow={"slide_type": "slide"}
# Si la tabla tiene un registro único por columnas identificadoras, pivotear la tabla no tiene problemas.
#
# Si existe más de un regisro por columnas identificadoras, es necesario usar una función de agrupación para seleccionar un valor representativo.
#
# Veamos que dice la documentación:
# +
# df.pivot_table?
# + [markdown] slideshow={"slide_type": "slide"}
# Intentémoslo. ¿Que tal pivotear para contar el número de terremotos por año?
# -
df.pivot_table(index="Año", values="Pais", aggfunc=pd.DataFrame.count)#.T
# + [markdown] slideshow={"slide_type": "slide"}
# ¿Cuál fue la mayor magnitud en cada año?
# -
df.pivot_table(index="Año", values="Magnitud", aggfunc=pd.np.max)#.T
# + [markdown] slideshow={"slide_type": "slide"}
# ¿Cuál fue la mayor magnitud en cada año, en cada país?
# -
df.pivot_table(index=["Año", "Pais"], values="Magnitud", aggfunc=pd.np.max)
# + [markdown] slideshow={"slide_type": "slide"}
# ¿Cómo fueron los terremotos año a año en cada país?
# -
df.pivot_table(index="Pais", columns="Año", values="Magnitud", aggfunc=pd.np.max)
df.pivot_table(index="Pais", columns="Año", values="Magnitud", aggfunc=pd.np.max, fill_value="")
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Pivoteando una tabla
#
# En general, pivotear una tabla no es particularmente complicado pero requiere saber bien cuál es la pregunta que se desea responder, puesto que como vimos, a partir de una misma tabla en formato "long" es posible generar varias tablas de agregación distintas.
#
# Y como dice el dicho, la práctica hace al maestro.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Despivoteando una tabla
#
# **Disclaimer**: No conozco una mejor traducción. En inglés tampoco se han puesto de acuerdo. He visto: *melt*, *un-pivot* y *reverse-pivot*, entre otros. Hasta ahora nadie me ha corregido con una mejor traducción.
#
# Despivotear una tabla consiste en pasar del "wide format" al "long format".
#
# El proceso de "des-pivotear" típicamente se realiza para poder agregar nuevas columnas a la tabla, o ponerla en un formato que permita un análisis apropiado con herramientas de visualización más avanzadas.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Despivoteando una tabla
#
# Típicamente existen 2 opciones:
# 1. El valor indicado para la columna es único, y sólo se requiere definir correctamente las columnas.
# 2. El valor indicado por la columna no es único o requiere un procesamiento adicional, y se requiere una iteración más profunda.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Despivoteando una tabla
# ### 4.1 Valor único: definir las columnas necesarias
# -
import pandas as pd
columns = ["sala","Lu-8:00","Lu-9:00","Lu-10:00","Ma-8:00","Ma-9:00","Ma-10:00"]
data = [
["C201","mat1","mat1", "","","",""],
["C202", "", "", "","mat1","mat1", ""],
["C203","fis1","fis1","fis1","fis1","fis1","fis1"],
]
df = pd.DataFrame(data=data, columns=columns)
# + slideshow={"slide_type": "slide"}
df
# + [markdown] slideshow={"slide_type": "slide"}
# La documentación de melt es bastante explícita aunque no muy amigable:
# +
# df.melt?
# + [markdown] slideshow={"slide_type": "slide"}
# Intentémoslo:
# -
df
df.melt(id_vars=["sala"]) # columnas identificadoras
# + slideshow={"slide_type": "slide"}
df.melt(id_vars=["sala"], # columnas identificadoras
var_name="dia-hora", # nueva variable
value_name="curso") # nombre de la columna para el valor de nueva variable
# + slideshow={"slide_type": "slide"}
df.melt(id_vars=["sala"], # columnas identificadoras
value_vars=["Lu-8:00","Lu-9:00","Lu-10:00"], # columnas a considerar
var_name="dia-hora", # nueva variable
value_name="curso") # nombre de la columna para el valor de nueva variable
# + [markdown] slideshow={"slide_type": "slide"}
# Después de despivotear puede ser necesario un poco de limpieza para evitar los valores vacíos.
# + slideshow={"slide_type": "-"}
# Despivotear el dataframe, renombrando columna indexadora y de valor
df_melt = df.melt(id_vars=["sala"], var_name="dia-hora", value_name="curso")
# Eliminar filas sin contenido y ordenarlas por nombre de sala y dia-hora.
df_melt[df_melt.curso!=""].sort_values(["sala","dia-hora"])
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Despivoteando una tabla
# ### 4.1 Valor único: definir las columnas necesarias
# Intentémoslo con un ejemplo más complejo
# -
import pandas as pd
columns = ["sala","dia","08:00","09:00","10:00"]
data = [
["C201","Lu", "mat1","mat1", ""],
["C201","Ma", "","",""],
["C202","Lu", "","",""],
["C202","Ma", "mat1","mat1", ""],
["C203","Lu", "fis1","fis1","fis1"],
["C203","Ma", "fis1","fis1","fis1"],
]
df = pd.DataFrame(data=data, columns=columns)
# + slideshow={"slide_type": "slide"}
df
# + slideshow={"slide_type": "slide"}
# Despivotear incorrectamente la tabla
df.melt(id_vars=["sala"], var_name="hora", value_name="curso")
# + slideshow={"slide_type": "slide"}
# Despivotear correctamente la tabla
df.melt(id_vars=["sala", "dia"], var_name="hora", value_name="curso")
# + slideshow={"slide_type": "slide"}
# Despivotear correctamente la tabla
df_melt = df.melt(id_vars=["sala", "dia"], var_name="hora", value_name="curso")
df_melt[df_melt.curso!=""].sort_values(["sala","dia","hora"])
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Despivoteando una tabla
# ### 4.1 Relaciones no únicas
# Consideremos el siguiente ejemplo:
# -
import pandas as pd
columns = ["sala","curso","Lu","Ma","hora"]
data = [
["C201","mat1","X","","8:00-10:00"],
["C202","mat1","","X","8:00-10:00"],
["C203","fis1","X","X","8:00-11:00"],
]
df = pd.DataFrame(data=data, columns=columns)
# + slideshow={"slide_type": "slide"}
df
# + [markdown] slideshow={"slide_type": "slide"}
# ¿Cómo podríamos des-pivotear la tabla anterior?
# + [markdown] slideshow={"slide_type": "fragment"}
# No existe un camino único. Dependiendo de la complejidad de la operación podrían haber soluciones más faciles.
# + [markdown] slideshow={"slide_type": "slide"}
# #### Idea 1:
# Despivotear manualmente y generar un nuevo dataframe.
# * **Ventajas**: Si se puede es una solución directa y rápida.
# * **Desventaja**: requiere programación explícita de la tarea, no es reutilizable.
# + slideshow={"slide_type": "slide"}
# Obtener el día lunes
df_Lu = df.loc[df.Lu=="X", ["sala","curso","hora"]]
df_Lu["dia"] = "Lu"
df_Lu
# + slideshow={"slide_type": "slide"}
# Obtener el día martes
df_Ma = df.loc[df.Ma=="X", ["sala","curso","hora"]]
df_Ma["dia"] = "Ma"
df_Ma
# -
# Juntar
pd.concat([df_Lu,df_Ma])
# + [markdown] slideshow={"slide_type": "slide"}
# #### Idea 2:
# Iterar sobre las filas y generar contenido para un nuevo dataframe.
# * **Ventajas**: En general, fácil de codificar.
# * **Desventaja**: puede ser lento, es ineficiente.
# -
# Forma de iterar sobre cada fila del dataframe
for i, row in df.iterrows():
# Procesar cada fila
print(row.sala, row.curso, row.Lu, row.Ma, row.hora)
# + slideshow={"slide_type": "slide"}
my_columns = ["sala","curso","dia","hora"]
my_data = []
for i, df_row in df.iterrows():
# Procesar cada fila
if df_row.Lu=="X":
my_row = [df_row.sala, df_row.curso, "Lu", df_row.hora]
my_data.append(my_row)
if df_row.Ma=="X":
my_row = [df_row.sala, df_row.curso, "Ma", df_row.hora]
my_data.append(my_row)
new_df = pd.DataFrame(data=my_data, columns=my_columns)
new_df
# + slideshow={"slide_type": "slide"}
my_columns = ["sala","curso","dia","hora"]
my_data = []
for i, df_row in df.iterrows():
# Procesar cada fila
for col_aux in ["Lu","Ma"]:
if df_row[col_aux]=="X":
my_row = [df_row.sala, df_row.curso, col_aux, df_row.hora]
my_data.append(my_row)
new_df = pd.DataFrame(data=my_data, columns=my_columns)
new_df
# + [markdown] slideshow={"slide_type": "slide"}
# ## Referencias:
#
# * https://medium.com/mission-log/design-better-data-tables-430a30a00d8c
# * https://medium.com/@enricobergamini/creating-non-numeric-pivot-tables-with-python-pandas-7aa9dfd788a7
# * https://nikgrozev.com/2015/07/01/reshaping-in-pandas-pivot-pivot-table-stack-and-unstack-explained-with-pictures/
| m02_analisis_de_datos/02_tabulando_datos/02_tabulando_datos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature: Out-Of-Fold Predictions from a CNN (+Magic Inputs)
# In addition to the convolutional architecture, we'll append some of the leaky features to the intermediate feature layer.
# <img src="assets/cnn-with-magic.png" alt="Network Architecture" style="height: 1200px;" />
# ## Imports
# This utility package imports `numpy`, `pandas`, `matplotlib` and a helper `kg` module into the root namespace.
from pygoose import *
import gc
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import *
from keras import backend as K
from keras.models import Model, Sequential
from keras.layers import *
from keras.callbacks import EarlyStopping, ModelCheckpoint
# ## Config
# Automatically discover the paths to various data folders and compose the project structure.
project = kg.Project.discover()
# Identifier for storing these features on disk and referring to them later.
feature_list_id = 'oofp_nn_cnn_with_magic'
# Make subsequent NN runs reproducible.
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
# ## Read data
# Word embedding lookup matrix.
embedding_matrix = kg.io.load(project.aux_dir + 'fasttext_vocab_embedding_matrix.pickle')
# Padded sequences of word indices for every question.
X_train_q1 = kg.io.load(project.preprocessed_data_dir + 'sequences_q1_fasttext_train.pickle')
X_train_q2 = kg.io.load(project.preprocessed_data_dir + 'sequences_q2_fasttext_train.pickle')
X_test_q1 = kg.io.load(project.preprocessed_data_dir + 'sequences_q1_fasttext_test.pickle')
X_test_q2 = kg.io.load(project.preprocessed_data_dir + 'sequences_q2_fasttext_test.pickle')
y_train = kg.io.load(project.features_dir + 'y_train.pickle')
# Magic features.
magic_feature_lists = [
'magic_frequencies',
'magic_cooccurrence_matrix',
]
X_train_magic, X_test_magic, _ = project.load_feature_lists(magic_feature_lists)
X_train_magic = X_train_magic.values
X_test_magic = X_test_magic.values
scaler = StandardScaler()
scaler.fit(np.vstack([X_train_magic, X_test_magic]))
X_train_magic = scaler.transform(X_train_magic)
X_test_magic = scaler.transform(X_test_magic)
# Word embedding properties.
EMBEDDING_DIM = embedding_matrix.shape[-1]
VOCAB_LENGTH = embedding_matrix.shape[0]
MAX_SEQUENCE_LENGTH = X_train_q1.shape[-1]
print(EMBEDDING_DIM, VOCAB_LENGTH, MAX_SEQUENCE_LENGTH)
# ## Define models
init_weights = initializers.TruncatedNormal(mean=0.0, stddev=0.05, seed=2)
init_bias = 'zeros'
def create_embedding_block():
input_seq = Input(shape=(MAX_SEQUENCE_LENGTH, ), dtype='int32')
embedding_seq = Embedding(
VOCAB_LENGTH,
EMBEDDING_DIM,
weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
trainable=False,
)(input_seq)
output_seq = embedding_seq
return input_seq, output_seq
def create_model_question_conv_branch(input_seq, params):
conv_1 = Conv1D(
params['num_conv_filters'],
kernel_size=params['conv_kernel_size'],
padding='same',
)(input_seq)
bn_1 = BatchNormalization()(conv_1)
relu_1 = Activation('relu')(bn_1)
dropout_1 = Dropout(params['conv_dropout_rate'])(relu_1)
conv_2 = Conv1D(
params['num_conv_filters'],
kernel_size=params['conv_kernel_size'],
padding='same',
)(dropout_1)
bn_2 = BatchNormalization()(conv_2)
relu_2 = Activation('relu')(bn_2)
dropout_2 = Dropout(params['conv_dropout_rate'])(relu_2)
flatten = Flatten()(dropout_2)
output = flatten
return output
def create_model_question_timedist_max_branch(input_seq, params):
timedist = TimeDistributed(Dense(EMBEDDING_DIM))(input_seq)
bn = BatchNormalization()(timedist)
relu = Activation('relu')(bn)
dropout = Dropout(params['timedist_dropout_rate'])(relu)
lambda_max = Lambda(
lambda x: K.max(x, axis=1),
output_shape=(EMBEDDING_DIM, )
)(dropout)
output = lambda_max
return output
def create_dense_block(input_layer, num_units, dropout_rate):
dense = Dense(
num_units,
kernel_initializer=init_weights,
bias_initializer=init_bias,
)(input_layer)
bn = BatchNormalization()(dense)
relu = Activation('relu')(bn)
dropout = Dropout(dropout_rate)(relu)
output = dropout
return output
def create_model(params):
input_q1, emb_q1 = create_embedding_block()
input_q2, emb_q2 = create_embedding_block()
# Feature extractors.
conv_q1_output = create_model_question_conv_branch(emb_q1, params)
conv_q2_output = create_model_question_conv_branch(emb_q2, params)
timedist_q1_output = create_model_question_timedist_max_branch(emb_q1, params)
timedist_q2_output = create_model_question_timedist_max_branch(emb_q2, params)
# Mid-level transforms.
conv_merged = concatenate([conv_q1_output, conv_q2_output])
conv_dense_1 = create_dense_block(conv_merged, params['num_dense_1'], params['dense_dropout_rate'])
conv_dense_2 = create_dense_block(conv_dense_1, params['num_dense_2'], params['dense_dropout_rate'])
td_merged = concatenate([timedist_q1_output, timedist_q2_output])
td_dense_1 = create_dense_block(td_merged, params['num_dense_1'], params['dense_dropout_rate'])
td_dense_2 = create_dense_block(td_dense_1, params['num_dense_2'], params['dense_dropout_rate'])
# Magic features.
magic_input = Input(shape=(X_train_magic.shape[-1], ))
# Main dense block.
merged_main = concatenate([conv_dense_2, td_dense_2, magic_input])
dense_main_1 = create_dense_block(merged_main, params['num_dense_1'], params['dense_dropout_rate'])
dense_main_2 = create_dense_block(dense_main_1, params['num_dense_2'], params['dense_dropout_rate'])
dense_main_3 = create_dense_block(dense_main_2, params['num_dense_3'], params['dense_dropout_rate'])
output = Dense(
1,
kernel_initializer=init_weights,
bias_initializer=init_bias,
activation='sigmoid',
)(dense_main_3)
model = Model(
inputs=[input_q1, input_q2, magic_input],
outputs=output,
)
model.compile(
loss='binary_crossentropy',
optimizer='nadam',
metrics=['accuracy']
)
return model
def predict(model, X_q1, X_q2, X_magic):
"""
Mirror the pairs, compute two separate predictions, and average them.
"""
y1 = model.predict([X_q1, X_q2, X_magic], batch_size=1024, verbose=1).reshape(-1)
y2 = model.predict([X_q2, X_q1, X_magic], batch_size=1024, verbose=1).reshape(-1)
return (y1 + y2) / 2
# ## Partition the data
NUM_FOLDS = 5
kfold = StratifiedKFold(
n_splits=NUM_FOLDS,
shuffle=True,
random_state=RANDOM_SEED
)
# Create placeholders for out-of-fold predictions.
y_train_oofp = np.zeros_like(y_train, dtype='float64')
y_test_oofp = np.zeros((len(X_test_q1), NUM_FOLDS))
# ## Define hyperparameters
BATCH_SIZE = 2048
MAX_EPOCHS = 200
model_params = {
'num_conv_filters': 32,
'num_dense_1': 256,
'num_dense_2': 128,
'num_dense_3': 100,
'conv_kernel_size': 3,
'conv_dropout_rate': 0.25,
'timedist_dropout_rate': 0.25,
'dense_dropout_rate': 0.25,
}
# The path where the best weights of the current model will be saved.
model_checkpoint_path = project.temp_dir + 'fold-checkpoint-' + feature_list_id + '.h5'
# ## Fit the folds and compute out-of-fold predictions
# +
# %%time
# Iterate through folds.
for fold_num, (ix_train, ix_val) in enumerate(kfold.split(X_train_q1, y_train)):
# Augment the training set by mirroring the pairs.
X_fold_train_q1 = np.vstack([X_train_q1[ix_train], X_train_q2[ix_train]])
X_fold_train_q2 = np.vstack([X_train_q2[ix_train], X_train_q1[ix_train]])
X_fold_train_magic = np.vstack([X_train_magic[ix_train], X_train_magic[ix_train]])
X_fold_val_q1 = np.vstack([X_train_q1[ix_val], X_train_q2[ix_val]])
X_fold_val_q2 = np.vstack([X_train_q2[ix_val], X_train_q1[ix_val]])
X_fold_val_magic = np.vstack([X_train_magic[ix_val], X_train_magic[ix_val]])
# Ground truth should also be "mirrored".
y_fold_train = np.concatenate([y_train[ix_train], y_train[ix_train]])
y_fold_val = np.concatenate([y_train[ix_val], y_train[ix_val]])
print()
print(f'Fitting fold {fold_num + 1} of {kfold.n_splits}')
print()
# Compile a new model.
model = create_model(model_params)
# Train.
model.fit(
[X_fold_train_q1, X_fold_train_q2, X_fold_train_magic], y_fold_train,
validation_data=([X_fold_val_q1, X_fold_val_q2, X_fold_val_magic], y_fold_val),
batch_size=BATCH_SIZE,
epochs=MAX_EPOCHS,
verbose=1,
callbacks=[
# Stop training when the validation loss stops improving.
EarlyStopping(
monitor='val_loss',
min_delta=0.001,
patience=3,
verbose=1,
mode='auto',
),
# Save the weights of the best epoch.
ModelCheckpoint(
model_checkpoint_path,
monitor='val_loss',
save_best_only=True,
verbose=2,
),
],
)
# Restore the best epoch.
model.load_weights(model_checkpoint_path)
# Compute out-of-fold predictions.
y_train_oofp[ix_val] = predict(model, X_train_q1[ix_val], X_train_q2[ix_val], X_train_magic[ix_val])
y_test_oofp[:, fold_num] = predict(model, X_test_q1, X_test_q2, X_test_magic)
# Clear GPU memory.
K.clear_session()
del X_fold_train_q1
del X_fold_train_q2
del X_fold_val_q1
del X_fold_val_q2
del model
gc.collect()
# -
cv_score = log_loss(y_train, y_train_oofp)
print('CV score:', cv_score)
# ## Save features
feature_names = [feature_list_id]
features_train = y_train_oofp.reshape((-1, 1))
features_test = np.mean(y_test_oofp, axis=1).reshape((-1, 1))
project.save_features(features_train, features_test, feature_names, feature_list_id)
# ## Explore
pd.DataFrame(features_test).plot.hist()
| notebooks/feature-oofp-nn-cnn-with-magic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.1 64-bit
# language: python
# name: python3
# ---
# Common methodology for recognizing digits in images without R-CNN:
# 1. Localize object
# 2. Extract the object
# 4. OCR on object
#
# OpenCV image processing reference: https://docs.opencv.org/3.4/d2/d96/tutorial_py_table_of_contents_imgproc.html
# +
import image_processing_util as ipu
from imutils.perspective import four_point_transform
import cv2 as cv
import imutils
import matplotlib.pyplot as plt
import os
import pytesseract
pytesseract.pytesseract.tesseract_cmd = 'C:/Program Files/Tesseract-OCR/tesseract.exe'
image = cv.imread("hall1.jpg")
image = imutils.resize(image, height=1000)
plt.imshow(image)
# -
#Step 1: Localize object
edges = ipu.get_edges(image)
candidate_list = ipu.get_quadrilaterals(edges)
plt.imshow(edges)
#Step 2: Extract object
candidates = len(candidate_list)
if candidates>10:#So the out folder doesn't blow up
candidates = 10
first_candidate_warped = four_point_transform(image, candidate_list[0].reshape(4, 2))
for i in range(candidates):
warped = four_point_transform(image, candidate_list[i].reshape(4, 2))
plt.imshow(warped)
abs_path = os.path.abspath("out")
plt.savefig(os.path.join(abs_path, str(i)+'.jpg'))
#Step 3: OCR
text = pytesseract.image_to_string(warped)
print(str(i) + ".jpg contents: " + text.replace("\n"," "))
plt.imshow(first_candidate_warped)
| numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:modern]
# language: python
# name: conda-env-modern-py
# ---
# +
# %reload_ext autoreload
# %autoreload 2
import os
import sys
import tensorflow as tf
import _pickle as pkl
from utils import *
from FCNN import FCNN
from sklearn.linear_model import LinearRegression, Ridge, LogisticRegression
# -
# ## Read Datasets
# LFW+
dataset_address = './LFWA+/lfw'
dataset = get_dataset(dataset_address)
lfw_raw, label_list = get_image_paths_and_labels(dataset)
with open("./dataset_description.pkl","rb") as foo:
dc = pkl.load(foo)
lfw_list = []
for im in dc['image_list']:
lfw_list.append(os.path.join(dataset_address, im))
lfw_attributes = (dc['attributes']>0).astype(int)
lfw_labels = (lfw_attributes[:,0]>0).astype(int)
lfw_latent_vars = dc['latent_vars'] ##Latent representation of the image in the VAE trained on CelebA
lfw_data = ((read_images(lfw_list) - 127.5)/128, lfw_labels, lfw_latent_vars)
sex = lfw_attributes[:,0]
skin = lfw_attributes[:,3]
dictionary = dc
permuted_idxs = np.random.permutation(np.arange(len(lfw_data[0])))
idxs_val, idxs_test = permuted_idxs[:6263], permuted_idxs[6263:]
x_val, y_val, a_val = lfw_data[0][idxs_val], lfw_data[1][idxs_val], lfw_attributes[:,[3,38,40]][idxs_val]
x_test, y_test, a_test = lfw_data[0][idxs_test], lfw_data[1][idxs_test], lfw_attributes[:,[3,38,40]][idxs_test]
# ## Load trained network
tf.reset_default_graph()
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=1.0)
gender_ckpt = "./model-20180428-135113_male.ckpt-111"
net = FCNN()
sess = tf.Session(config=tf.ConfigProto(gpu_options=gpu_options, log_device_placement=False))
sess.run(tf.global_variables_initializer())
sess.run(tf.local_variables_initializer())
net.load_model(sess, gender_ckpt, initialize = False)
# ## Network & VAE Representation
prelogits_val = np.zeros((len(x_val),128))
batch_size = 24
for i in range(int(np.ceil(len(x_val)/batch_size))):
prelogits_val[i*batch_size:i*batch_size+batch_size] = sess.run(
net.prelogits ,feed_dict={net.in_ph:x_val[i*batch_size:i*batch_size+batch_size]})
lts_val = dictionary['latent_vars'][idxs_val] #VAE representation
prelogits_test = np.zeros((len(x_test),128))
for i in range(int(np.ceil(len(x_test)/batch_size))):
prelogits_test[i*batch_size:i*batch_size+batch_size] = sess.run(
net.prelogits, feed_dict={net.in_ph:x_test[i*batch_size:i*batch_size+batch_size]})
lts_test = dictionary['latent_vars'][idxs_test] #VAE representation
# ## Black-box audting
def sess_run(result, x, l, sess):
num = x.shape[0]
num_batch = np.ceil(num/200).astype(int)
output = np.zeros(num)
for batch in range(num_batch):
output[batch*200:(batch+1)*200] = sess.run(result, feed_dict={net.in_ph:x[batch*200:(batch+1)*200],
latent_ph:l[batch*200:(batch+1)*200]})
return output
control = tf.cast(tf.greater(net.output[:,1],net.output[:,0]), tf.float32)
noharm = [control, 1 - control, control + 1 - control]
latent_val, latent_test = lts_val, lts_test
dim = latent_val.shape[-1]
latent_ph = tf.placeholder(tf.float32, shape=(None, dim), name="latent_var")
logits = net.output[:,1] - net.output[:,0]
max_T = 100
thresh = 1e-4 # Hyper-parameter
temp_harm = []
for s in noharm:
temp_harm.append(sess_run(s, x_val, latent_val, sess))
def res(p, y):
return y * ((p>=0.1)/(p + 1e-20) + (p<0.1) * (20 - 100 * p)) +\
(1-y) * ((p < 0.9)/(1 - p + 1e-20) + (p>=0.9) * (100 * p - 80))
plt.plot(np.arange(0, 1.0, 0.001), res(np.arange(0., 1, 0.001), 0))
plt.plot(np.arange(0, 1.0, 0.001), res(np.arange(0., 1, 0.001), 1))
plt.legend(['0', '1'])
plt.xlabel('P')
plt.ylabel('Residual')
best_epoch, best_acc = -1,0
(idxs1, idxs2, _), _ = split_data(np.arange(len(idxs_val)), ratio=[0.7,0.3,0.])
coeffs = []
for t in range(max_T):
control = tf.cast(tf.greater(net.output[:,1], net.output[:,0]), tf.float32)
noharm = [control, 1 - control, control + 1 - control]
probs_heldout = sess_run(tf.nn.sigmoid(logits), x_val[idxs2], latent_val[idxs2], sess)
heldout_loss = np.mean(-y_val[idxs2] * np.log(probs_heldout + 1e-20) - (1-y_val[idxs2]) * np.log(1-probs_heldout + 1e-20))
heldout_acc = np.mean((probs_heldout>0.5)==y_val[idxs2])
probs = sess_run(tf.nn.sigmoid(logits), x_val, latent_val ,sess)
val_loss = np.mean(-y_val * np.log(probs + 1e-20) - (1 - y_val) * np.log(1 - probs + 1e-20))
val_acc = np.mean((probs > 0.5) == y_val)
if heldout_acc > best_acc:
best_epoch = t
best_acc = heldout_acc
best_logits = logits
delta = res(probs,y_val)
residual = probs - y_val
for i, s in enumerate(noharm):
temp_s = sess_run(noharm[i], x_val[idxs1], latent_val[idxs1], sess)
temp_s_heldout = sess_run(noharm[i], x_val[idxs2], latent_val[idxs2], sess)
samples1 = np.where(temp_s == 1)[0]
samples2 = np.where(temp_s_heldout == 1)[0]
clf = Ridge(alpha=1)
clf.fit(latent_val[idxs1][samples1],delta[idxs1][samples1])
clf_prediction = clf.predict(latent_val[idxs2][samples2])
corr = np.mean(clf_prediction * residual[idxs2][samples2])
print(t, i, corr)
if corr > 1e-4:
coeffs.append(clf.coef_)
h = (tf.matmul(latent_ph, tf.constant(np.expand_dims(clf.coef_,-1),
dtype=tf.float32))[:,0] + clf.intercept_)
logits -= .1 * h * s
break
if i==2:
break
# ## Result
probs = sess_run(net.output[:,1] - net.output[:,0], x_test, latent_test, sess)
groups = ['all', 'F', 'M', 'B', 'N', 'BF', 'BM', 'NF', 'NM']
errs = []
idxs = np.where((skin[idxs_test]>-1) * (sex[idxs_test]>-10))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]>-1) * (sex[idxs_test]==0))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]>-1) * (sex[idxs_test]==1))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==1) * (sex[idxs_test]>-10))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==0) * (sex[idxs_test]>-10))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==1) * (sex[idxs_test]==0))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==1) * (sex[idxs_test]==1))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==0) * (sex[idxs_test]==0))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==0) * (sex[idxs_test]==1))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
output = ''
for group, err in zip(groups, errs):
output += group + ': ' + str(round(err, 1)) + ' & '
print('Original: ', output)
probs = sess_run(tf.nn.sigmoid(best_logits), x_test, latent_test, sess)
groups = ['all', 'F', 'M', 'B', 'N', 'BF', 'BM', 'NF', 'NM']
errs = []
idxs = np.where((skin[idxs_test]>-1) * (sex[idxs_test]>-10))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]>-1) * (sex[idxs_test]==0))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]>-1) * (sex[idxs_test]==1))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==1) * (sex[idxs_test]>-10))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==0) * (sex[idxs_test]>-10))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==1) * (sex[idxs_test]==0))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==1) * (sex[idxs_test]==1))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==0) * (sex[idxs_test]==0))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
idxs = np.where((skin[idxs_test]==0) * (sex[idxs_test]==1))[0]
errs.append(100 * np.mean((probs[idxs]>0.5)!=y_test[idxs]))
output = ''
for group, err in zip(groups, errs):
output += group + ': ' + str(round(err, 1)) + ' & '
print('MultiAccuracy Boost: ', output)
| Run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
filepath = '../data/processed/howstat/fall_of_wickets/'
file = 'fow_2400.csv'
df = pd.read_csv(filepath+file, index_col=0, parse_dates=[2], infer_datetime_format=True)
df.head(10)
df.groupby(['MatchId','MatchInnings'])['Wicket', 'Runs'].max()
# Let's say a batting collapse is defined as losing at least 3 wickets for at most 30 runs.
# We want to know for each MatchId-MatchInnings whether that innings is a batting collapse.
#
# Say we have FoW:
# 1-20
# 2-40
# 3-41
#
# This is a batting collapse.
# One approach:
# If we know we have only 5 wickets lost:
# Check each group of 3 wickets. Groups will be: 5-2, 4-1, 3-0.
# Convert the Runs into an array, with index=0 wickets.
#
#
# Test cases for groups of 3:
# Wickets 1,2,3 is not a collapse: 105 runs
# Wickets 4,5,6 is not a collapse: 94 runs
# Wickets 6,7,8 is a collapse: 10 runs
# Wickets 7,8,9 is a collapse: 3 runs
# Wickets 8,9,10 is not a collapse: 38 runs
#
# Test cases for groups of 2:
#
l_runs = list(df[df.MatchInnings==2].Runs)
print(l_runs)
l_runs.insert(0,0)
l_runs
l_test = [1,2,3]
run_difference = l_runs[l_test[-1]] - l_runs[l_test[0]]
print(run_difference)
# +
from itertools import tee, islice
def nwise(iterable, n):
iters = tee(iterable, n)
for i, it in enumerate(iters):
next(islice(it, i, i), None)
return zip(*iters)
list(nwise(l_runs, n=2))
# -
l_runs
l_runs
n = 3
for i in range(n,len(l_runs)):
print(f'Start: {i-n}')
print(f'End: {i}')
if i-n == 0:
run_difference = l_runs[i]
else:
run_difference = l_runs[i] - l_runs[i-n-1]
print(f'Run difference: {run_difference}')
l_positions_involved = [x for x in range(i-n+1, i+1)]
print(f'Positions involved: {l_positions_involved}')
print('\n')
def containsCollapse(df):
l_runs = list(df.Runs)
l_runs.insert(0,0)
n = 3
n_collapses = 0
d_collapses = {}
for i in range(n,len(l_runs)):
if i-n == 0:
run_difference = l_runs[i]
else:
run_difference = l_runs[i] - l_runs[i-n-1]
if run_difference <= 30:
n_collapses += 1
l_positions_involved = [x for x in range(i-n+1, i+1)]
d_collapses[n_collapses] = l_positions_involved
return n_collapses, d_collapses
# for each collapse, want to store:
# - number of runs lost
# - positions involved
# - (later on) batsman involved
# - (later on) bowlers involved
df.groupby(['MatchId','MatchInnings']).apply(containsCollapse)
df
| notebooks/ag-format-collapses-1.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
# import plotly.plotly as py
import matplotlib.pyplot as plt
from matplotlib import pyplot
import plotly.graph_objs as go
init_notebook_mode(connected=True)
time_series_df=pd.read_csv('../Data/main/Maritius_AOI_20200701_0731_full.csv')
time_series_df.head()
time_series_df.sort_values(by=['timestamp'], inplace=True)
time_series_df.head()
print(time_series_df.loc[time_series_df['timestamp'] == '2020-07'])
time_sorted_df = time_series_df.sort_values(by=['timestamp'], inplace=True)
time_series_df.head()
time_series_df.head()
time_series_df
time_series_df.info()
time_series_df['timestamp'] = pd.to_datetime(time_series_df['timestamp'])
time_series_df.head()
time_series_df.info()
time_series_df.head()
plt.plot( time_series_df['timestamp'], time_series_df['speed'])
time_series_df.drop(["call_sign", "flag" ,"draught" , "ship_and_cargo_type", "length", "width","eta" , "destination", "status", "maneuver", "accuracy" ,"collection_type" ,'mmsi_label'], axis=1, inplace=True)
time_series_df.info()
time_series_df.head()
time_series_df.drop(['created_at','imo', 'name'], axis=1, inplace=True)
time_series_df.head()
time_series_df = time_series_df[time_series_df['speed'].notna()]
time_series_df.head()
time_series_df.info()
plt.plot( time_series_df['timestamp'], time_series_df['speed'])
plt.plot( time_series_df['timestamp'], time_series_df['speed'])
plt.gcf().autofmt_xdate()
plt.show()
time_series_df = time_series_df.reset_index(drop=True)
time_series_df.head()
len(time_series_df[(time_series_df['heading']==0)])
plt.plot( time_series_df['timestamp'], time_series_df['heading'])
plt.gcf().autofmt_xdate()
plt.show()
plt.plot( time_series_df['timestamp'], time_series_df['course'])
plt.gcf().autofmt_xdate()
plt.show()
print(time_series_df.loc[[18000]])
time_series_df.head()
time_series_df.info()
time_series_df.info()
time_series_df.drop(time_series_df[time_series_df['speed'] == 0].index, inplace = True)
time_series_df.mmsi.unique()
mv_value = 372711000
param = 'heading'
len(time_series_df[time_series_df["mmsi"]==mv_value])
len(time_series_df[time_series_df["speed"]==0])
mv_data = time_series_df[time_series_df['mmsi']==mv_value]
mv_data.head()
plt.plot( mv_data['timestamp'], mv_data['speed'])
plt.gcf().autofmt_xdate()
plt.show()
plt.plot( mv_data['timestamp'], mv_data['course'])
plt.gcf().autofmt_xdate()
plt.show()
# mv_data.reset_index(inplace=True)
mv_data = mv_data.drop(['mmsi','msg_type','latitude', 'longitude'], axis=1)
mv_data.head()
mv_data = mv_data[mv_data['speed'].notna()]
mv_data.info()
mv_data.head()
# +
import warnings
mv_data = mv_data.set_index(['timestamp'])
mv_data.index = pd.to_datetime(mv_data.index, unit='s')
names=mv_data.columns
# Resample the entire dataset by daily average
rollmean = mv_data.resample(rule='D').mean()
rollstd = mv_data.resample(rule='D').std()
# Plot time series for each sensor with its mean and standard deviation
for name in names:
_ = plt.figure(figsize=(18,3))
_ = plt.plot(mv_data[name], color='blue', label='Original')
_ = plt.plot(rollmean[name], color='red', label='Rolling Mean')
_ = plt.plot(rollstd[name], color='black', label='Rolling Std' )
_ = plt.legend(loc='best')
_ = plt.title(name)
plt.show()
# -
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.pipeline import make_pipeline
df2 = mv_data
names=df2.columns
x = mv_data[names]
scaler = StandardScaler()
pca = PCA()
pipeline = make_pipeline(scaler, pca)
pipeline.fit(x)
features = range(pca.n_components_)
_ = plt.figure(figsize=(15, 5))
_ = plt.bar(features, pca.explained_variance_)
_ = plt.xlabel('PCA feature')
_ = plt.ylabel('Variance')
_ = plt.xticks(features)
_ = plt.title("Importance of the Principal Components based on inertia")
plt.show()
pca = PCA(n_components=2)
principalComponents = pca.fit_transform(x)
principalDf = pd.DataFrame(data = principalComponents, columns = ['pc1', 'pc2'])
mv_data['pc1']=pd.Series(principalDf['pc1'].values, index=mv_data.index)
mv_data['pc2']=pd.Series(principalDf['pc2'].values, index=mv_data.index)
from statsmodels.tsa.stattools import adfuller
# Run Augmented Dickey Fuller Test
result = adfuller(principalDf['pc1'])
# Print p-value
print(result[1])
# Compute change in daily mean
pca1 = principalDf['pc1'].pct_change()
# Compute autocorrelation
autocorrelation = pca1.dropna().autocorr()
print('Autocorrelation is: ', autocorrelation)
from statsmodels.graphics.tsaplots import plot_acf
plot_acf(pca1.dropna(), lags=20, alpha=0.05)
# Compute change in daily mean
pca2 = principalDf['pc2'].pct_change()
# Compute autocorrelation
autocorrelation = pca2.autocorr()
print('Autocorrelation is: ', autocorrelation)
# +
from statsmodels.graphics.tsaplots import plot_acf
plot_acf(pca2.dropna(), lags=20, alpha=0.05)
# +
# outlier_lower = Q1 - (1.5*IQR)
# outlier_upper = Q3 + (1.5*IQR)
# Calculate outlier bounds for pc1
q1_pc1, q3_pc1 = mv_data['pc1'].quantile([0.25, 0.75])
iqr_pc1 = q3_pc1 - q1_pc1
lower_pc1 = q1_pc1 - (1.5*iqr_pc1)
upper_pc1 = q3_pc1 + (1.5*iqr_pc1)
# Calculate outlier bounds for pc2
q1_pc2, q3_pc2 = mv_data['pc2'].quantile([0.25, 0.75])
iqr_pc2 = q3_pc2 - q1_pc2
lower_pc2 = q1_pc2 - (1.5*iqr_pc2)
upper_pc2 = q3_pc2 + (1.5*iqr_pc2)
# -
lower_pc1, upper_pc1
lower_pc2, upper_pc2
mv_data['anomaly_pc1'] = ((mv_data['pc1']>upper_pc1) | (mv_data['pc1']<lower_pc1)).astype('int')
mv_data['anomaly_pc2'] = ((mv_data['pc2']>upper_pc2) | (mv_data['pc2']<lower_pc2)).astype('int')
mv_data['anomaly_pc1'].value_counts()
mv_data['anomaly_pc2'].value_counts()
outliers_pc1 = mv_data.loc[(mv_data['pc1']>upper_pc1) | (mv_data['pc1']<lower_pc1), 'pc1']
outliers_pc2 = mv_data.loc[(mv_data['pc2']>upper_pc2) | (mv_data['pc2']<lower_pc2), 'pc2']
len(outliers_pc1)/len(mv_data)
len(outliers_pc2)/len(mv_data)
mv_data.head()
a = mv_data[mv_data['anomaly_pc1'] == 1] #anomaly
b = mv_data[mv_data['anomaly_pc2'] == 1] #anomaly
plt.figure(figsize=(18,6))
plt.plot(mv_data[param], color='blue', label='Normal')
plt.plot(a[param], linestyle='none', marker='X', color='red', markersize=12, label='Anomaly1')
plt.plot(b[param], linestyle='none', marker='X', color='green', markersize=12, label='Anomaly2')
plt.xlabel('Date and Time')
plt.ylabel(param)
plt.title(param +' Anomalies')
plt.legend(loc='best')
plt.show();
data1 = a
data2 = b
a.head()
from sklearn.cluster import KMeans
# I will start k-means clustering with k=2 as I already know that there are 3 classes of "NORMAL" vs
# "NOT NORMAL" which are combination of BROKEN" and"RECOVERING"
kmeans = KMeans(n_clusters=2, random_state=42)
kmeans.fit(principalDf.values)
labels = kmeans.predict(principalDf.values)
unique_elements, counts_elements = np.unique(labels, return_counts=True)
clusters = np.asarray((unique_elements, counts_elements))
plt.figure(figsize = (9, 7))
plt.bar(clusters[0], clusters[1], tick_label=clusters[0])
plt.xlabel('Clusters')
plt.ylabel('Number of points')
plt.title('Number of points in each cluster')
plt.show()
plt.figure(figsize=(9,7))
plt.scatter(principalDf['pc1'], principalDf['pc2'], c=labels)
plt.xlabel('pc1')
plt.ylabel('pc2')
plt.title('K-means of clustering')
plt.show()
def getDistanceByPoint(data, model):
""" Function that calculates the distance between a point and centroid of a cluster,
returns the distances in pandas series"""
distance = []
for i in range(0,len(data)):
Xa = np.array(data.loc[i])
Xb = model.cluster_centers_[model.labels_[i]-1]
distance.append(np.linalg.norm(Xa-Xb))
return pd.Series(distance, index=data.index)
outliers_fraction = 0.05
# get the distance between each point and its nearest centroid. The biggest distances are considered as anomaly
distance = getDistanceByPoint(principalDf, kmeans)
# number of observations that equate to the 13% of the entire data set
number_of_outliers = int(outliers_fraction*len(distance))
# Take the minimum of the largest 13% of the distances as the threshold
threshold = distance.nlargest(number_of_outliers).min()
# anomaly1 contain the anomaly result of the above method Cluster (0:normal, 1:anomaly)
principalDf['anomaly1'] = (distance >= threshold).astype(int)
principalDf.head()
principalDf['anomaly1'].value_counts()
# principalDf['anomaly2'].value_counts()
mv_data['anomaly1'] = pd.Series(principalDf['anomaly1'].values, index=mv_data.index)
a = mv_data[mv_data['anomaly1'] == 1] #anomaly
# b = mv_data[mv_data['anomaly2'] == 1] #anomaly
plt.figure(figsize=(18,6))
plt.plot(mv_data[param], color='blue', label='Normal')
plt.plot(a[param], linestyle='none', marker='X', color='red', markersize=12, label='Anomaly')
plt.xlabel('Date and Time')
plt.ylabel(param)
plt.title('Anomalies')
plt.legend(loc='best')
plt.show();
data3 = a
# data4 = b
# Import IsolationForest
from sklearn.ensemble import IsolationForest
# Assume that 13% of the entire data set are anomalies
outliers_fraction = 0.05
model = IsolationForest(contamination=outliers_fraction)
model.fit(principalDf.values)
principalDf['anomaly2'] = pd.Series(model.predict(principalDf.values))
# visualization
mv_data['anomaly2'] = pd.Series(principalDf['anomaly2'].values, index=mv_data.index)
a = mv_data.loc[mv_data['anomaly2'] == -1] #anomaly
plt.figure(figsize=(18,6))
plt.plot(mv_data[param], color='blue', label='Normal')
plt.plot(a[param], linestyle='none', marker='X', color='red', markersize=12, label='Anomaly')
plt.xlabel('Date and Time')
plt.ylabel(param +'Reading')
plt.title('Anomalies')
plt.legend(loc='best')
plt.show();
data4 = a
mv_data['anomaly2'].value_counts()
a.head()
# frames = [data1, data2, data3, data4]
# common = pd.merge(data1, data2 ,how='inner', on="timestamp")
# common = pd.merge(common, data3 ,how='inner', on="timestamp")
# common = pd.merge(common, data4 ,how='inner', on="timestamp")
# +
# common.info()
# common.head()
# -
data1.head()
# +
data2.head()
# -
data3.head()
data4.head()
def intersection(lst1, lst2,lst3,lst4):
lst5 = [value for value in lst2 if value in lst1]
lst6 = [value for value in lst3 if value in lst5]
lst7 = [value for value in lst4 if value in lst6]
return lst7
time_common = intersection(data1.index.unique() , data2.index.unique() , data3.index.unique() , data4.index.unique() )
time_df = pd.DataFrame(columns = mv_data.columns, index = time_common)
for time in time_common:
time_df.loc[time] = mv_data.loc[time]
plt.figure(figsize=(18,6))
plt.plot(mv_data[param], color='blue', label='Normal')
plt.plot(time_df[param], linestyle='none', marker='X', color='red', markersize=12, label='Anomaly')
plt.xlabel('Date and Time')
plt.ylabel(param +'Reading')
plt.title('Anomalies')
plt.legend(loc='best')
plt.show();
| cluster model/Oil Spill Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import os
import scipy
# plt.style.use('fivethirtyeight')
sns.set_style("whitegrid")
sns.set_context("notebook")
DATA_PATH = '../data/'
df = pd.read_csv(os.path.join(DATA_PATH,'creditcard.csv'))
df.head()
# ## Preprocessing
# Some basic checks: are there NAN's or duplicate rows?
# * NaN's : There are not
# * Duplicates: There are, so we erase them, this would disturb the metrics (biasing them towards too optimistic values)
print("Total number of NaN's:",df.isna().sum().sum())
print("Number of duplicated rows:",df.duplicated().sum())
df = df[df.duplicated()==False]
df.reset_index(inplace=True,drop=True)
# As expected, we are working with a highly unbalance dataset, the mean of Class is 0.001667,
#
# which means that only 0.17% of the entries correspond to Class 1, Fraud.
df.describe()
# ## Feature engineering: Time and Amount
# We check that the Time variable correspond to seconds (the database indicates that it correspond to two days)
print('Total number of days:',df.Time.max()/60/60/24)
# We can perform some feature engineering based on the Time variable
df['TimeScaled'] = df.Time/60/60/24/2
df['TimeSin'] = np.sin(2*np.pi*df.Time/60/60/24)
df['TimeCos'] = np.cos(2*np.pi*df.Time/60/60/24)
df.drop(columns='Time',inplace=True)
# Some basic statistics for each variable in the dataframe.
# It easily observed that all V's variables have zero mean and order 1 standard deviation (and they are sorted by it), they come from a PCA in which the variables where scaled before the PCA.
# There are entries with Amount = 0. What's is the meaning of this? Transactions with no money interchange? Is that really a Fraud? Is it interesting to detect them? Those are questions that we cannot answer here, but should be investigated in case of a real world problem.
#
# We can see that in this subgroup of the data, there is an over-representation of class 1 (FRAUD).
print('Probability of each one of the classes in the whole dataset')
for i, prob in enumerate(df.Class.value_counts(normalize=True)):
print('Class {}: {:.2f} %'.format(i,prob*100))
print('Probability of each one of the classes in the entries with Amount = 0')
for i, prob in enumerate(df[df.Amount==0].Class.value_counts(normalize=True)):
print('Class {}: {:.2f} %'.format(i,prob*100))
# The Amount variable is too disperse, so it is better to work with it in logarithm scale, and then rescale it.
#
# This does not matter for Decision Tree based methods. Exercise: Why?
plt.figure(figsize=(10,6))
df['AmountLog'] = np.log10(1.+df.Amount)
plt.subplot(121)
sns.distplot(df.Amount,bins=200)
plt.xlim((0,1000))
plt.subplot(122)
sns.distplot(df.AmountLog)
# df.drop(columns='Amount',inplace=True)
plt.show()
scipy.stats.boxcox(1+df.Amount,lmbda=None,alpha=0.05)
df['AmountBC']= points
df.drop(columns=['Amount','AmountLog'],inplace=True)
plt.figure(figsize=(10,6))
# df['AmountLog'] = np.log10(1.+df.Amount)
plt.subplot(121)
points, lamb = scipy.stats.boxcox(1+df.Amount,lmbda=None,)
sns.distplot(points-1,axlabel='BoxCox:'+str(lamb))
plt.subplot(122)
sns.distplot(df.AmountLog)
# df.drop(columns='Amount',inplace=True)
plt.show()
# Now, we save a copy of the cleaned dataframe, in order to preserve the preprocessing.
df.describe()
df.to_csv(os.path.join(DATA_PATH,'df_clean.csv'))
# ## Exploration
# ### One dimensional histograms
# Let us explore the Time variable, can we see any pattern?
bins = np.linspace(0,1,24)
plt.figure(figsize=(10,6))
plt.subplot(121)
sns.distplot(df.TimeScaled,bins=bins,label='All',color='red')
plt.legend()
plt.subplot(122)
sns.distplot(df[df.Class==0].TimeScaled,bins=bins,kde=False,norm_hist=True,label='Normal')
sns.distplot(df[df.Class==1].TimeScaled,bins=bins,kde=False,norm_hist=True,label='Fraud')
plt.legend()
plt.show()
# We can explore the histograms for all the variables, since there are around 30 of them.
for variable in df.columns:
plt.figure(figsize=(6,6))
bins = np.linspace(df[variable].min(),df[variable].max(),50)
sns.distplot(df[df.Class==0][variable],bins=bins,kde=False,norm_hist=True,label='Normal',axlabel=variable)
sns.distplot(df[df.Class==1][variable],bins=bins,kde=False,norm_hist=True,label='Fraud',axlabel=variable)
plt.legend()
plt.show()
# ### Pairwise scatterplots
# A really good way of getting intuition is through pairplots, i.e., scatter plots using two variables. In this way we can check if some variables are useful to disentangle the entries by Class.
#
# In this case, since there are 28+2 features, there would be 900/2 plots to check the pairwise relations.
# We first downsample Class 0 (normal) to obtain clearer plots
df_small = pd.merge( df[df.Class==1],df[df.Class==0].sample(n=10000),how='outer')
# We cannot plot all the variables, there are too many
variables_to_show = ['V4','V14','V17','V3']
sns.pairplot(df_small,vars=variables_to_show,
hue='Class',kind='scatter',markers="o",
plot_kws=dict(s=6, edgecolor=None, linewidth=0.01,alpha=0.5))
plt.show()
# Same pairwise scatterplot with all the data, we visualize it easily giving some transparency to the most populated class, and also using smaller markers for it.
plt.figure(figsize=(5,5))
x_var = 'V16'
y_var = 'V9'
sns.scatterplot(data=df[df.Class==0],x=x_var, y=y_var,s=5,edgecolor=None,alpha=0.3)
sns.scatterplot(data=df[df.Class==1],x=x_var, y=y_var,color='orange',s=10,edgecolor='w')
plt.show()
# ### Correlations
# It is also easy to see correlations among variables, but it is not very useful in this case.
sns.heatmap(df.corr(),vmin=-1,vmax=1)
sns.heatmap(df[df.Class==0].corr(),vmin=-1,vmax=1)
sns.heatmap(df[df.Class==1].corr(),vmin=-1,vmax=1)
| notebooks/1_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pymatgen
import ase.db
import numpy as np
import collections as coll
# !wget https://cmr.fysik.dtu.dk/_downloads/mp_gllbsc.db
con = ase.db.connect('mp_gllbsc.db')
from mpcontribs.io.archieml.mpfile import MPFile
s = """
title: GLLB-SC Bandgaps
references.AEM: Advanced Energy Materials, Juli 22, 2014
references.PRA: <NAME>, <NAME>, <NAME>, <NAME>, Phys. Rev. A 1995, 51, 1944.
references.PRB: <NAME>, <NAME>, <NAME>, <NAME>, Phys. Rev. B 2010, 82, 115106.
contributor : Technical University of Denmark
authors: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>
explanation: Bandgaps calculated using GLLB-SC potential by Gritsenko, <NAME>, <NAME>, and Baerends (GLLB), adapted by Kuisma et al to account for solids (-SC). The Kohn-Sham gap most basically refers to the energy difference between the bottom of the conduction band and the top of the valence band. The Quasi-Particle gap takes into account the derivative discontinuity of the chemical potential and gives the bandgap as the difference between ionization potenital and electron affinity.
"""
mpfile = MPFile.from_string(s)
count = 0
x=y=z=w= []
for row in con.select('mpid'):
d = coll.OrderedDict([])
d['Kohn-Sham_Bandgap'] = coll.OrderedDict([])
d['Derivative_Discontinuity'] = coll.OrderedDict([])
d['Quasi-Particle_Bandgap'] = coll.OrderedDict([])
count = count + 1
mpid = 'mp-' + str(row.mpid)
d['Kohn-Sham_Bandgap']['Indirect'] = row.gllbsc_ind_gap - row.gllbsc_disc
d['Kohn-Sham_Bandgap']['Direct'] = row.gllbsc_dir_gap - row.gllbsc_disc
d['Derivative_Discontinuity'] = row.gllbsc_disc
d['Quasi-Particle_Bandgap']['Indirect'] = row.gllbsc_ind_gap
d['Quasi-Particle_Bandgap']['Direct'] = row.gllbsc_dir_gap
mpfile.add_hierarchical_data(d, identifier=mpid)
if count == 10:
break
# +
import plotly
import plotly.graph_objs as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
plotly.offline.init_notebook_mode(connected = True)
x1 = [row.mpid for row in con.select('mpid')]
y1 = [row.gllbsc_disc for row in con.select('mpid')]
y2 = [row.gllbsc_ind_gap for row in con.select('mpid')]
y3 = [row.gllbsc_dir_gap for row in con.select('mpid')]
trace_1 = go.Bar(x = x1[0:10], y = y1[0:10], name = 'derivative_discontinuity')
trace_2 = go.Bar(x = x1[0:10], y = y2[0:10], name = 'quasi-particle_bandgap(indirect)')
trace_3 = go.Bar(x = x1[0:10], y = y3[0:10], name = 'quasi-particle_bandgap(direct)')
data = [trace_1, trace_2, trace_3]
layout = go.Layout(barmode='stack', xaxis= dict(title= 'mp-id', type= 'category'), yaxis= dict(title= 'Energy (eV)'))
fig = go.Figure(data=data, layout=layout)
stacked_bar = iplot(fig, filename='stacked-bar')
| mpcontribs-users/mpcontribs/users/dtu/dtu_mpfile_build.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:rmg_env]
# language: python
# name: conda-env-rmg_env-py
# ---
# +
import os
from IPython.display import display, Image
from rmgpy.tools.uncertainty import Uncertainty, process_local_results
from rmgpy.tools.canteraModel import get_rmg_species_from_user_species
from rmgpy.species import Species
# -
# # First Order Local Uncertainty Analysis for Chemical Reaction Systems
#
# This IPython notebook performs first order local uncertainty analysis for a chemical reaction system
# using a RMG-generated model.
# ## Step 1: Define mechanism files and simulation settings
# +
# This is a small ethane pyrolysis model
# Must use annotated chemkin file
chemkin_file = 'data/parse_source/chem_annotated.inp'
dict_file = 'data/parse_source/species_dictionary.txt'
# Initialize the Uncertainty class instance and load the model
uncertainty = Uncertainty(output_directory='./temp/uncertainty')
uncertainty.load_model(chemkin_file, dict_file)
# Map the species to the objects within the Uncertainty class
ethane = Species().from_smiles('CC')
C2H4 = Species().from_smiles('C=C')
mapping = get_rmg_species_from_user_species([ethane, C2H4], uncertainty.species_list)
# Define the reaction conditions
initial_mole_fractions = {mapping[ethane]: 1.0}
T = (1300, 'K')
P = (1, 'bar')
termination_time = (0.5, 'ms')
sensitive_species=[mapping[ethane], mapping[C2H4]]
# -
# ## Step 2: Run sensitivity analysis
#
# Local uncertainty analysis uses the results from a first-order sensitivity analysis. This analysis is done using RMG's native solver.
# Perform the sensitivity analysis
uncertainty.sensitivity_analysis(initial_mole_fractions, sensitive_species, T, P, termination_time, number=5, fileformat='.png')
# Show the sensitivity plots
for species in sensitive_species:
print('{}: Reaction Sensitivities'.format(species))
index = species.index
display(Image(filename=os.path.join(uncertainty.output_directory,'solver','sensitivity_1_SPC_{}_reactions.png'.format(index))))
print('{}: Thermo Sensitivities'.format(species))
display(Image(filename=os.path.join(uncertainty.output_directory,'solver','sensitivity_1_SPC_{}_thermo.png'.format(index))))
# ## Step 3: Uncertainty assignment and propagation of uncorrelated parameters
#
# If we want to run local uncertainty analysis, we must assign all the uncertainties using the `Uncertainty` class' `assignParameterUncertainties` function. `ThermoParameterUncertainty` and `KineticParameterUncertainty` classes may be customized and passed into this function if non-default constants for constructing the uncertainties are desired. This must be done after the parameter sources are properly extracted from the model.
#
# ### Thermo Uncertainty
#
# Each species is assigned a uniform uncertainty distribution in free energy:
#
# $$G \in [G_{min},G_{max}]$$
#
# We will propogate the standard deviation in free energy, which for a uniform distribution is defined as follows:
#
# $$\Delta G = \frac{1}{\sqrt{12}}(G_{max} - G_{min})$$
#
# Several parameters are used to formulate $\Delta G$. These are $\Delta G_\mathrm{library}$, $\Delta G_\mathrm{QM}$, $\Delta G_\mathrm{GAV}$, and $\Delta _\mathrm{group}$.
#
# $$\Delta G = \delta_\mathrm{library} \Delta G_\mathrm{library} + \delta_\mathrm{QM} \Delta G_\mathrm{QM} + \delta_\mathrm{GAV} \left( \Delta G_\mathrm{GAV} + \sum_{\mathrm{group}\; j} d_{j} \Delta G_{\mathrm{group},j} \right)$$
#
# where $\delta$ is the Kronecker delta function which equals one if the species thermochemistry parameter contains the particular source type and $d_{j}$ is the degeneracy (number of appearances) of the thermo group used to construct the species thermochemistry in the group additivity method.
#
# ### Kinetics Uncertainty
#
# Each reaction is assigned a uniform uncertainty distribution in the overall $\ln k$, or $\ln A$:
#
# $$\ln k \in [\ln(k_{min}),\ln(k_{max})]$$
#
# Again, we use the standard deviation of this distribution:
#
# $$\Delta \ln(k) = \frac{1}{\sqrt{12}}(\ln k_{max} - \ln k_{min})$$
#
# The parameters used to formulate $\Delta \ln k$ are $\Delta \ln k_\mathrm{library}$, $\Delta \ln k_\mathrm{training}$, $\Delta \ln k_\mathrm{pdep}$, $\Delta \ln k_\mathrm{family}$, $\Delta \ln k_\mathrm{non-exact}$, and $\Delta \ln k_\mathrm{rule}$.
#
# For library, training, and pdep reactions, the kinetic uncertainty is assigned according to their uncertainty type. For kinetics estimated using RMG's rate rules, the following formula is used to calculate the uncertainty:
#
# $$\Delta \ln k_\mathrm{rate\; rules} = \Delta\ln k_\mathrm{family} + \log_{10}(N+1) \left(\Delta\ln k_\mathrm{non-exact}\right) + \sum_{\mathrm{rule}\; i} w_i \Delta \ln k_{\mathrm{rule},i}$$
#
# where N is the total number of rate rules used and $w_{i}$ is the weight of the rate rule in the averaging scheme for that kinetics estimate.
# NOTE: You must load the database with the same settings which were used to generate the model.
# This includes any thermo or kinetics libraries which were used.
uncertainty.load_database(
thermo_libraries=['DFT_QCI_thermo', 'primaryThermoLibrary'],
kinetics_families='default',
reaction_libraries=[],
)
uncertainty.extract_sources_from_model()
uncertainty.assign_parameter_uncertainties()
# The first order local uncertainty, or variance $(d\ln c_i)^2$, for the concentration of species $i$ is defined as:
#
# $$(\Delta \ln c_i)^2 = \sum_{\mathrm{reactions}\; m} \left(\frac{\partial\ln c_i}{\partial\ln k_m}\right)^2 (\Delta \ln k_m)^2 + \sum_{\mathrm{species}\; n} \left(\frac{\partial\ln c_i}{\partial G_n}\right)^2(\Delta G_n)^2$$
#
# We have previously performed the sensitivity analysis. Now we perform the local uncertainty analysis and apply the formula above using the parameter uncertainties and plot the results. This first analysis considers the parameters to be independent. In other words, even when multiple species thermochemistries depend on a single thermo group or multiple reaction rate coefficients depend on a particular rate rule, each value is considered independent of each other. This typically results in a much larger uncertainty value than in reality due to cancellation error.
result = uncertainty.local_analysis(sensitive_species, correlated=False, number=5, fileformat='.png')
print(process_local_results(result, sensitive_species, number=5)[1])
# Show the uncertainty plots
for species in sensitive_species:
print('{}: Thermo Uncertainty Contributions'.format(species))
display(Image(filename=os.path.join(uncertainty.output_directory, 'uncorrelated', 'thermoLocalUncertainty_{}.png'.format(species.to_chemkin()))))
print('{}: Reaction Uncertainty Contributions'.format(species))
display(Image(filename=os.path.join(uncertainty.output_directory, 'uncorrelated', 'kineticsLocalUncertainty_{}.png'.format(species.to_chemkin()))))
# ## Step 4: Uncertainty assignment and propagation of correlated parameters
#
# A more accurate picture of the uncertainty in mechanism estimated using groups and rate rules requires accounting of the correlated errors resulting from using the same groups in multiple parameters. This requires us to track the original sources: the groups and the rate rules, which constitute each parameter. These errors may cancel in the final uncertainty calculation. Note, however, that the error stemming from the estimation method itself do not cancel.
#
# For thermochemistry, the error terms described previously are $\Delta G_\mathrm{library}$, $\Delta G_\mathrm{QM}$, $\Delta G_\mathrm{GAV}$, and $\Delta _\mathrm{group}$. Of these, $\Delta G_\mathrm{GAV}$ is an uncorrelated residual error, whereas the other terms are correlated. The set of correlated and uncorrelated parameters can be thought of instead as a set of independent parameters, $\Delta G_{ind,w}$.
#
# For kinetics, the error terms described perviously are $\Delta \ln k_\mathrm{library}$, $\Delta \ln k_\mathrm{training}$, $\Delta \ln k_\mathrm{pdep}$, $\Delta \ln k_\mathrm{family}$, $\Delta \ln k_\mathrm{non-exact}$, and $\Delta \ln k_\mathrm{rule}$. Of these, $\Delta \ln k_\mathrm{family}$ and $\Delta \ln k_\mathrm{non-exact}$ are uncorrelated error terms resulting from the method of estimation. Again, we consider the set of correlated and uncorrelated parameters as the set of independent parameters, $\Delta\ln k_{ind,v}$.
#
# The first order local uncertainty, or variance $(d\ln c_i)^2$, for the concentration of species $i$ becomes:
#
# $$(\Delta \ln c_i)^2 = \sum_v \left(\frac{\partial\ln c_i}{\partial\ln k_{ind,v}}\right)^2 \left(\Delta\ln k_{ind,v}\right)^2 + \sum_w \left(\frac{\partial\ln c_i}{\partial G_{ind,w}}\right)^2 \left(\Delta G_{ind,w}\right)^2$$
#
# where the differential terms can be computed as:
#
# $$\frac{\partial\ln c_i}{\partial\ln k_{ind,v}} = \sum_m \frac{\partial\ln c_i}{\partial\ln k_m} \frac{\partial\ln k_m}{\partial\ln k_{ind,v}}$$
#
# $$\frac{\partial\ln c_i}{\partial G_{ind,w}} = \sum_n \frac{\partial\ln c_i}{\partial G_n} \frac{\partial G_n}{\partial G_{ind,w}}$$
#
uncertainty.assign_parameter_uncertainties(correlated=True)
result = uncertainty.local_analysis(sensitive_species, correlated=True, number=10, fileformat='.png')
print(process_local_results(result, sensitive_species, number=5)[1])
# Show the uncertainty plots
for species in sensitive_species:
print('{}: Thermo Uncertainty Contributions'.format(species))
display(Image(filename=os.path.join(uncertainty.output_directory, 'correlated', 'thermoLocalUncertainty_{}.png'.format(species.to_chemkin()))))
print('{}: Reaction Uncertainty Contributions'.format(species))
display(Image(filename=os.path.join(uncertainty.output_directory, 'correlated', 'kineticsLocalUncertainty_{}.png'.format(species.to_chemkin()))))
| ipython/local_uncertainty.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Plot the number of projects that ask for funding over time in both the npm community and in the larger GitHub community.
import pandas as pd
import numpy as np
from datetime import datetime
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
# %matplotlib inline
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
df_all = pd.read_csv('../../data/asking_group_npm_gh.csv')
df_npm = pd.read_csv('../../data/asking_group_npm.csv')
# Get number of projects
print('All:', df_all['slug'].nunique())
print('Npm:', df_npm['slug'].nunique())
# Get number of projects with multiple services
def get_num_multiple(df):
services = ['tip4commit', 'salt', 'flattr', 'patreon', 'paypal', 'bountysource',
'tidelift', 'opencollective', 'liberapay', 'kickstarter']
num_multiple = 0
for index, row in df.iterrows():
has_one = 0
for service in services:
if row[service] == 1:
if has_one == 1:
num_multiple += 1
break
else:
has_one = 1
return (num_multiple, num_multiple/df['slug'].nunique())
get_num_multiple(df_all)
get_num_multiple(df_npm)
# Plot historic figure
def get_counts(df):
services = ['tip4commit', 'salt', 'flattr', 'patreon', 'paypal', 'bountysource',
'tidelift', 'opencollective', 'liberapay', 'kickstarter']
others = ['bountysource', 'issuehunt', 'kickstarter', 'salt', 'otechie', 'tip4commit', 'tidelift']
counts = {}
for service in services:
count = df[service].sum()
if service in others:
if 'other' in counts:
counts['other'] += count
else:
counts['other'] = count
else:
counts[service] = count
df_counts = pd.DataFrame(columns=['service', 'count'])
for service in counts:
df_counts = df_counts.append({'service':service, 'count':counts[service]}, ignore_index=True)
df_counts = df_counts.sort_values(by='count', ascending=False)
return df_counts
def convert_datetime(x):
if not isinstance(x, str):
return x
# Ignore time zone info
if 'T' in x and 'Z' in x:
x = x.split('T')[0]
try:
temp = datetime.strptime(x, '%Y-%m-%d %H:%M:%S')
except:
try:
temp = datetime.strptime(x, '%Y-%m-%d')
except:
try:
temp = datetime.strptime(x, '%m/%d/%Y')
except:
try:
temp = datetime.strptime(x, '%m/%d/%Y %H:%M:%S')
except:
temp = datetime.strptime(x, '%m/%d/%Y %H:%M')
return datetime(temp.year, temp.month, 1)
def get_time_data(service, df):
df_data = pd.DataFrame(columns=['date', 'count', 'service'])
if service == 'other':
df_sub = df[(df['bountysource']==1)|(df['salt']==1)|(df['tip4commit']==1)|(df['tidelift']==1)].copy()
# Merge other_date columns
df_sub['other_date'] = df_sub['bountysource_date'].combine_first(df_sub['salt_date'])
df_sub['other_date'] = df_sub['other_date'].combine_first(df_sub['tip4commit_date'])
df_sub['other_date'] = df_sub['other_date'].combine_first(df_sub['tidelift_date'])
else:
df_sub = df[df[service]==1].copy()
df_sub['date'] = df_sub[service+'_date'].apply(convert_datetime)
grouped = df_sub.groupby('date')
count = 0
for date, df_date in grouped:
num_add = df_date['slug'].nunique()
if num_add < 0:
print('num_project should not be negative')
break
count += num_add
df_data = df_data.append({'date':date, 'count':count, 'service':service}, ignore_index=True)
return df_data
def thousands(x, pos):
return '%1iK' % (x*1e-3)
def plot_time_data(df, formaty=True):
df_time1 = pd.DataFrame(columns=['date', 'count', 'service'])
services = ['paypal', 'patreon', 'flattr', 'liberapay', 'opencollective', 'other']
for service in services:
df_time1 = df_time1.append(get_time_data(service, df), ignore_index=True)
markers = {'paypal':'o', 'patreon':'^', 'opencollective':'', 'flattr':'|', 'liberapay':'', 'other':''}
linestyles = {'paypal':'solid', 'patreon':'solid', 'opencollective':'solid', 'flattr':'solid', 'liberapay':'dashed', 'other':'dotted'}
hues = {'paypal':'#5D7EA7', 'patreon':'#EFC6D1', 'flattr':'#85BB65', 'liberapay':'#C58293',
'opencollective':'#9FCDCD', 'other':'#3E3E3C'}
plt.figure(figsize=(10, 5))
for service, df_service in df_time1.groupby('service', sort=False):
plt.plot(df_service['date'].values, df_service['count'].values, label=service, linewidth=2.5,
marker=markers[service], markersize=7, color=hues[service], linestyle=linestyles[service], markevery=3)
if formaty:
formatter = FuncFormatter(thousands)
plt.gca().yaxis.set_major_formatter(formatter)
plt.legend(loc='upper left', prop={'size': 20}, frameon=False)
plt.ylabel('Count', fontsize=22, labelpad=15)
plt.xlim([convert_datetime('2008-01-01'), convert_datetime('2019-05-23')])
plt.rc('xtick',labelsize=22)
plt.rc('ytick',labelsize=22)
sns.despine()
return plt
# Plot for all
plt = plot_time_data(df_all)
# Plot for npm
plt = plot_time_data(df_npm, formaty=False)
| src/jupyter_notebooks/funding_adoption_over_time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Compiling Chemical Reaction Networks
# __Overview:__ The main utility of BioCRNpyler is to compile complex CRNs from simple specifications. BioCRNpyler compilation is summarized in the below figure:
#
# 
#
# Mixtures represent context and contain Components and default Mechanisms. Components (biochemical parts) call Mechanisms which they inherit from Mixture or override internally. Mechanisms are reaction schemas (abstract functions that produce CRN Species and Reactions) that represent different biochemical processes. Mechanisms find reaction parameters they need from a parameter file. More details on parameter loading can be seen in the parameter ipython notebook.
#
# In this notebook, a number of basic BioCRNpyler models are compiled involving enzymes and molecular binding. This notebook highlights the structure of BioCRNpyler and how to control compilation. Other notebooks highlight the available Mixtures, Components, and Mechanisms through detailed examples.
# # Example 1: Enzymes & Default Mechanisms
#
# In this example, we examine two different mechanisms for modeling an Enzyme $E$ which converts a substrate $S$ into a product $P$ in an in-vitro context (no dilution of any species):
#
# In this context, Enzyme is a Component which looks for a 'catalysis' Mechanism. Two different Mixtures will be compared with different default 'catalysis' Mechanisms.
#
# 1. Baisc Catalysis: $E + S \xrightarrow{k_{cat}} E + P$
# 2. Michalis Menten Catalysis: $E + S \underset{k_u}{\overset{k_b}{\rightleftharpoons}}E:S \xrightarrow{k_{cat}} E + P$
#
# By default, the Enzyme Component will inherit the Mechanism in its Mixture.
# +
# %matplotlib inline
from biocrnpyler import *
#We will use default parameter names
default_parameters = {"kb":100, "ku":10, "kcat":1.}
#Mixture 1 (M1) will contain an Enzyme E1
E = Enzyme("E", substrate = "S", product = "P")
#Choose a catalysis mechanism by commenting out one of them.
mech_cat = BasicCatalysis()
#mech_cat = MichalisMenten()
#place that mechanism in a dictionary: "catalysis":mech_cat
default_mechanisms = {mech_cat.mechanism_type:mech_cat}
#Create a mixture.
#Components is a list of Components in the mixture
#parameters is a dictionary of parameters. Can also accept parameter_file.
#default_mechanisms = dict sets the default_mechanisms in the Mixture
M = Mixture("Catalysis Mixture", components = [E], parameters = default_parameters, default_mechanisms = default_mechanisms)
print("repr(Mixture) gives a printout of what is in a mixture and what it's Mechanisms are:\n", repr(M),"\n")
#Compile the CRN with Mixture.compile_crn
CRN = M.compile_crn()
#CRNs can be printed in two ways
print("String representation of a CRN shows the string names of all species:\n",CRN, "\n\n")
print("Pretty_print representation of a CRN has formatting options and is easier to read:\n",
CRN.pretty_print(show_rates = True, show_attributes = True, show_materials = True))
# -
# # Example 2: Combining Multiple Enzymes into a Pathway Using Default Parameters for All Enzymes
# Next, we will combine 3 enzymes together into a pathway:
# 1. $E_1$ converts $A$ to $B$
# 2. $E_2$ converts $C$ to $D$
# 3. $E_3$ is a MultiEnzyme which converts $B$ and $D$ to $F$
#
# MultiEnzyme is a more general Enzyme Component which can have multiple substrates and products. Notice how all the Enzymes use the same default catalysis Mechanism which can be changed by changing the what is passed into default_mechanisms in the Mixture.
#
# Finally, we show how specific parameters can be used for each enzyme or default parameters shared between all Components.
# Default parameters can be given in a dictionary using the name of the parameter as a keyword:
#
# "parameter name" : value
#
# +
#These simple parameters can be used by default for all enzymes
default_parameters = {"kb":100, "ku":10, "kcat":1.}
E1 = Enzyme("E1", substrate = "A", product = "B")
E2 = Enzyme("E2", substrate = "C", product = "D")
E3 = MultiEnzyme("E3", substrates = ["B", "D"], products = ["F"])
#creeate a catalysis mechanism.
mech_cat = BasicCatalysis()
#mech_cat = MichalisMenten()
#place that mechanism in a dictionary: "catalysis":mech_cat
default_mechanisms = {mech_cat.mechanism_type:mech_cat}
#Create a mixture.
#Components is a list of Components in the mixture
#parameters is a dictionary of parameters. Can also accept parameter_file.
#default_mechanisms = dict sets the default_mechanisms in the Mixture
M = Mixture("Default Param Pathway", components = [E1, E2, E3], parameters = default_parameters, default_mechanisms = default_mechanisms)
print("repr(Mixture) gives a printout of what is in a mixture and what it's Mechanisms are:\n", repr(M),"\n")
#Compile the CRN with Mixture.compile_crn
CRN = M.compile_crn()
print(CRN.pretty_print(show_rates = True, show_attributes = True, show_materials = True))
# -
# # Example 3: A Pathway of Enzymes with Specific Parameters for Each Enzyme
# In the previous example, the same default parameters used by all the Enzymes. It is also easy to have specific parameters for each enzyme. Specific parameters can be given in a dictionary or parameter file using a variety of keys (see the Parameters notebook for details). The simplest, however, is:
#
# ("mechanism name/type", "component name", "parameter name") : value
#
# Notice that by switching the catalysis Mechanism, different parameters are used.
# +
#Or specific parameters can be made which give different rates for each enzyme
#The first row of parameters is used by BasicCatalysis
#The second-fourth row of parameters is used my MichalisMenten catalysis
specific_parameters = {("basic_catalysis", "E1", "kcat"):100, ("basic_catalysis", "E2", "kcat"):200, ("basic_catalysis", "E3", "kcat"):300,
("michalis_menten", "E1", "kb"):111, ("michalis_menten", "E1", "ku"):11, ("michalis_menten", "E1", "kcat"): 1.11,
("michalis_menten", "E2", "kb"):222, ("michalis_menten", "E2", "ku"):22, ("michalis_menten", "E2", "kcat"): 2.22,
("michalis_menten", "E3", "kb"):333, ("michalis_menten", "E3", "ku"):33, ("michalis_menten", "E3", "kcat"): 3.33}
E1 = Enzyme("E1", substrate = "A", product = "B")
E2 = Enzyme("E2", substrate = "C", product = "D")
E3 = MultiEnzyme("E3", substrates = ["B", "D"], products = ["F"])
#choose a catalysis mechanism.
#mech_cat = BasicCatalysis()
mech_cat = MichalisMenten()
#place that mechanism in a dictionary: "catalysis":mech_cat
default_mechanisms = {mech_cat.mechanism_type:mech_cat}
#To change the parameters, pass in a different dictionary using the parameter keyword
M = Mixture("Catalysis Mixture", components = [E1, E2, E3], parameters = specific_parameters, default_mechanisms = default_mechanisms)
CRN = M.compile_crn()
print("Using Specific Parameters for all reactions:\n", CRN.pretty_print())
# -
# # Example 4: Adding a Custom Mechanism to a Component
# Notice that in the above CRN, the enzymatic process is irreversible. However, many biochemists would argue that irreversibility is actually an approximation. In reality, the products of the reaction can bind to the enzyme and, if the chemical potential is high enough, cause the reverse reaction to occur. In many cases, it might be desirable to to only include the reverse reaction for some of the enzymatic reactions being in the Model. In BioCRNpyler, this can be done easily by adding a custom Mechanism to an individual Component which will override the default Mechanism provided by the Mixture.
#
# This is illustrated on the pathway built above, where here Enzyme 3 will be given a new catalysis mechanism called MichalisMentenReversible: $E + S \rightleftarrows E:S \rightleftarrows E:P \rightleftarrows E + P$
# +
#The MichalisMentenReversible has different parameter names: kb1, ku1, kb2, ku2, and kcat_rev
default_parameters = {"kb":100, "ku":10, "kcat":1., "kb1":111, "kb2":22, "ku1":11, "ku2":22, "kcat_rev":.001}
#Enzymes 1 and 2 are the same as above
E1 = Enzyme("E1", substrate = "A", product = "B")
E2 = Enzyme("E2", substrate = "C", product = "D")
#Create a dictionary of custom mechanisms to pass into E3 with the mechanisms keyword
mm_reversible = MichalisMentenReversible()
custom_mechanisms = {mm_reversible.mechanism_type:mm_reversible}
E3 = MultiEnzyme("E3", substrates = ["B", "D"], products = ["F"], mechanisms = custom_mechanisms)
#choose a catalysis mechanism.
#mech_cat = BasicCatalysis()
mech_cat = MichalisMenten()
#place that mechanism in a dictionary: "catalysis":mech_cat
default_mechanisms = {mech_cat.mechanism_type:mech_cat}
#Make the Mixture
M = Mixture("Catalysis Mixture", components = [E1, E2, E3], parameters = default_parameters, default_mechanisms = default_mechanisms)
#Compile the CRN
CRN = M.compile_crn()
print("Notice that there are additional reactions involving E3:\n", CRN.pretty_print())
# -
# # Example 5: Modelling an Enzyme which is also a ChemicalComplex (Binding)
#
# In this example, we will consider the followingbiochemical process: the tetramerization of the biosynthesis protein Inosine-5′monophosphate dehydrogenase (IMPDH) which is a homeotetramer that catalyzes the following reaction important in guanine biosynthesis:
#
# $\textrm{inosine-5'-phosphate} + \textrm{NAD}^+ + H_2O \rightleftharpoons \textrm{xanthosine-5'-phosphate} + \textrm{NADH} + H^+$
#
# The ChemicalComplex Component will represent the assembled protein Complex IMPDH formed by 4 identifical monomer subunits. ChemicalComplex is effectively a wrapper around a ComplexSpecies in order to provide binding reactions. The catalysis of the above reaction will be accomplished by simultaneously making IMPDH an Enzyme.
# +
#Use default parameters for simplicity
default_parameters = {"kb":100, "ku":10, "kcat":1., "kb1":111, "kb2":22, "ku1":11, "ku2":22, "kcat_rev":.001}
#Create a single species to represent the monomer.
monomer = Species("IMPDH", material_type = "subunit")
#A ChemicalComplex takes a list of species and allows them all to bind together.
C = ChemicalComplex(species = [monomer]*4, name = "IMPDH")
#Here, we set the internal species of MultiEnzyme (enzyme) to the the ComplexSpecies stored inside ChemicalComplex.
#This allows the same formal CRN species to be represented by multiple components and therefore participate in more reactions.
E = MultiEnzyme(enzyme = C.get_species(), substrates = ["isosine5phosphate", "NAD", "H2O"], products = ["xanthosine5phosphate", "NADH", "H"])
#choose a catalysis mechanism.
#mech_cat = BasicCatalysis()
mech_cat = MichalisMenten()
#mech_cat = MichalisMentenReversible()
#create a binding mechanism
mech_bind = One_Step_Binding() #All species bind together in a single step
#place that mechanism in a dictionary: {"catalysis":mech_cat, "binding":mech_bind}
default_mechanisms = {mech_cat.mechanism_type:mech_cat, mech_bind.mechanism_type:mech_bind}
#Make the Mixture
M = Mixture("Catalysis Mixture", components = [E1, E2, E3], parameters = default_parameters, default_mechanisms = default_mechanisms)
#Compile the CRN
CRN = M.compile_crn()
print(CRN.pretty_print())
# -
#
| examples/2. Compiling CRNs with Enzymes Catalysis and Binding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1e5jJxWTsI5n" colab_type="text"
# The objective of this project is to **deploy a Flask app** that uses a model trained with the Fast.ai v2 library following an example (bear detector) in the upcoming book **"Deep Learning for Coders with fastai and PyTorch: AI Applications Without a PhD" by <NAME> and <NAME>.**
#
# The most important part of the project is **testing a deployment process that combines a Flask app, the Gunicorn server, the Nginx server and a custom domain name with an SSL certificate**, all installed on a dedicated server.
#
# You can access the repo with all the code for this project plus detailed step by step explanations of how to deploy it, here:
# **[Repo with code and details](https://github.com/javismiles/bear-detector-flask-deploy)**
#
# Project put together by <NAME> (ideami.com)
# + id="JNZETAP2s2c3" colab_type="code" colab={}
#These lines are for google colab only, if you are on your own server, check the requirements.txt file at the repo.
# #!pip install fastai2
# #!pip install nbdev
# #!pip install azure
# #!pip install azure-cognitiveservices-vision-customvision
# #!pip install azure-cognitiveservices-vision-computervision
# #!pip install azure-cognitiveservices-search-websearch
# #!pip install azure-cognitiveservices-search-imagesearch
# #!unzip bearstotal.zip
# + id="Jfcl0AP0KpON" colab_type="code" colab={}
from utils import *
from fastai2.vision.widgets import *
from fastai2.imports import *
# + id="8Iav4fOlKVYp" colab_type="code" colab={}
# Categories of bears
bear_types = 'grizzly','black','teddy'
# Reference to the folder that holds the dataset
path = Path('bears')
# + id="VnUsC9FkKVYr" colab_type="code" outputId="a783405f-fa9a-40e3-e206-f1a0bd2ca0b2" colab={"base_uri": "https://localhost:8080/", "height": 55}
# Get a reference to the images
fns = get_image_files(path)
fns
# + id="DRbhPsr4KVYu" colab_type="code" outputId="41fefcd9-4990-4fe8-e7a6-76b86793efb9" colab={}
# Check for images with problems (corrupted, etc)
failed = verify_images(fns)
failed
# + id="JOSs_uAjKVYx" colab_type="code" colab={}
# You may delete images with issues with this command
#failed.map(Path.unlink);
# + id="U-9qdHAeKVYz" colab_type="code" colab={}
# Create a datablock structure, composed of the independente variable (an image block), and the dependent variable (a category).
# The datablock will use the get_image_files function to get items, it will split randomly (but from the same seed always) the images using
# 30% of them for the validation set. It will assign the independent variable, the label by checking the parent of the image, the name of
# its parent folder. And it will resize each image to be 128x128 pixels
bears = DataBlock(
blocks=(ImageBlock, CategoryBlock),
get_items=get_image_files,
splitter=RandomSplitter(valid_pct=0.3, seed=42),
get_y=parent_label,
item_tfms=Resize(128))
# + id="s-HkvIeGKVY2" colab_type="code" colab={}
# Create a dataloader from the reference to the path of the dataset.
dls = bears.dataloaders(path)
# + id="kdXTbg5vKVY4" colab_type="code" outputId="9dfb8e41-7b76-4ad1-e18c-929408421b65" colab={"base_uri": "https://localhost:8080/", "height": 193}
# Show 4 pictures of the validation set of the dataloader
dls.valid.show_batch(max_n=4, rows=1)
# + id="Oxmo3hGGKVY6" colab_type="code" colab={}
# Update the datablock, process each image, applying a random cropping process; also set the option of applying augmentation to
#batches of images; update the dataloader
bears = bears.new(
item_tfms=RandomResizedCrop(224, min_scale=0.5),
batch_tfms=aug_transforms())
dls = bears.dataloaders(path)
# + id="db99DEQFKVY9" colab_type="code" outputId="fdea9ec8-524b-4fa8-fb35-84359e1e350b" colab={"base_uri": "https://localhost:8080/", "height": 302, "referenced_widgets": ["eaffdeb2b74f423ea3a57562801c5155", "bec4fae3d46e4df7a7fce04eb1dccd0d", "5a258cb1ff3c4d1282e85ede33a20b0d", "0067a397c09d4a05800661ac4db70c07", "a433640c4042418b98213bc2d81e97a3", "8418f1481ba8446eb354740802f08159", "<KEY>", "0be5fd8f97df4ffeb2dd335e86505037"]}
# Create a cnn learner, passing it the dataloader, a resnet arquitecture reference, and setting the metric to evaluate to be the
# error rate
learn = cnn_learner(dls, resnet18, metrics=error_rate)
# We are doing transfer learning, to use fine_tune to fine tune the resnet during 4 epochs.
learn.fine_tune(4)
# + id="D60YWiRfKVZA" colab_type="code" outputId="7b681bd8-acf9-48ea-9282-8ad897c41721" colab={"base_uri": "https://localhost:8080/", "height": 310}
#Plot the confusion matrix to analyze the results
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix()
# + id="eK2hyxSFN0YB" colab_type="code" outputId="d4b178b3-2a21-418d-8243-b05049c36d42" colab={"base_uri": "https://localhost:8080/", "height": 241}
#Plot the top losses to analyze what kinds of images had issues
interp.plot_top_losses(5, rows=1)
# + id="W4nBsBpXN3N-" colab_type="code" colab={}
#clean the dataset optionally to polish the training (ImageClassifierCleaner may work well in google colab)
#cleaner = ImageClassifierCleaner(learn)
#cleaner
#for idx in cleaner.delete(): cleaner.fns[idx].unlink()
#for idx,cat in cleaner.change(): shutil.move(cleaner.fns[idx], path/cat)
# + id="6ugoZpGJN_rq" colab_type="code" colab={}
#This line will create a file (export.pkl), which you can then put in the "model" folder of the project to be used by the Flask app.
learn.export()
| resources/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Project Data Science Rossmann Store Sales</font>
#
# ### Dados coletados em: https://www.kaggle.com/c/rossmann-store-sales
# + [markdown] heading_collapsed=true
# ## <font color='red'>Problem</font>
#
# #### Rossmann opera mais de 3.000 drogarias em 7 países europeus. Atualmente, os gerentes de loja da Rossmann têm a tarefa de prever suas vendas diárias com até seis semanas de antecedência. As vendas da loja são influenciadas por muitos fatores, incluindo promoções, competição, feriados escolares e estaduais, sazonalidade e localidade. Com milhares de gerentes individuais prevendo vendas com base em suas circunstâncias únicas, a precisão dos resultados pode variar bastante.
#
# #### Você recebe dados históricos de vendas de 1.115 lojas Rossmann. A tarefa é prever a coluna "Vendas" para o conjunto de teste. Observe que algumas lojas no conjunto de dados foram temporariamente fechadas para reforma.
#
# #### Arquivos:
#
# **train.csv -** dados históricos incluindo vendas
#
# **test.csv -** dados históricos excluindo vendas
#
# **store.csv -** informações complementares sobre as lojas
#
# #### Campos de dados
#
# ##### A maioria dos campos é autoexplicativa. A seguir estão as descrições para aqueles que não são.
#
# **Id -** um Id que representa um (Store, Date) duple dentro do conjunto de teste
#
# **Store -** um ID único para cada loja
#
# **Sales -** o volume de negócios em qualquer dia (isto é o que você está prevendo)
#
# **Customers -** o número de clientes em um determinado dia
#
# **Open -** um indicador para saber se a loja estava aberta: 0 = fechada, 1 = aberta
#
# **StateHoliday -** indica um feriado estadual. Normalmente todas as lojas, com poucas exceções, fecham nos feriados estaduais. Observe que todas as escolas fecham nos feriados e finais de semana. a = feriado, b = feriado da Páscoa, c = Natal, 0 = Nenhum
#
# **SchoolHoliday -** indica se (loja, data) foi afetado pelo fechamento de escolas públicas
#
# **StoreType -** diferencia entre 4 modelos de loja diferentes: a, b, c, d
#
# **Assortment -** descreve um nível de sortimento: a = básico, b = extra, c = estendido
#
# **CompetitionDistance -** distância em metros até a loja concorrente mais próxima
#
# **CompetitionOpenSince [month / year] -** fornece o ano e mês aproximados em que o concorrente mais próximo foi aberto
#
# **Promo -** indica se uma loja está fazendo uma promoção naquele dia
#
# **Promo2 -** Promo2 é uma promoção contínua e consecutiva para algumas lojas: 0 = loja não participa, 1 = loja participa
#
# **Promo2Since [year / week] -** descreve o ano e a semana em que a loja começou a participar da Promo2
#
# **PromoInterval -** descreve os intervalos consecutivos em que a Promo2 é iniciada, nomeando os meses em que a promoção é reiniciada. Por exemplo, "fevereiro, maio, agosto, novembro" significa que cada rodada começa em fevereiro, maio, agosto, novembro de qualquer ano para aquela loja
# -
# # 0.0 IMPORTS
# +
import numpy as np
import pandas as pd
import seaborn as sns
import inflection
import datetime
import math
from matplotlib import pyplot as plt
from IPython.display import Image
# -
# ## 0.1. Loading data
df_sales_raw = pd.read_csv('data/train.csv', low_memory=False)
df_stores_raw = pd.read_csv('data/store.csv', low_memory=False)
# + code_folding=[0]
# Juntar os Data Frames aplicando o merge
df_raw = pd.merge(df_sales_raw, df_stores_raw, how='left', on='Store')
# + [markdown] heading_collapsed=true
# # 1.0. DESCREPTION DATA - PASSO 1
# + hidden=true
df1 = df_raw.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## 1.1. Rename columns
# + code_folding=[0] hidden=true
cols_old = ['Store', 'DayOfWeek', 'Date', 'Sales', 'Customers', 'Open', 'Promo',
'StateHoliday', 'SchoolHoliday', 'StoreType', 'Assortment',
'CompetitionDistance', 'CompetitionOpenSinceMonth',
'CompetitionOpenSinceYear', 'Promo2', 'Promo2SinceWeek',
'Promo2SinceYear', 'PromoInterval']
# snakecase = lambda x: inflection.underscore(x)
cols_new = list(map(lambda x: inflection.underscore(x), cols_old))
df1.columns = cols_new
# + [markdown] heading_collapsed=true hidden=true
# ## 1.2. Data dimensions
# + code_folding=[0] hidden=true
# Imprimindo o número de linhas
print('Number of Rows: {}'.format(df1.shape[0]))
# Imprimindo o número de colunas
print('Number of Cols: {}'.format(df1.shape[1]))
# + [markdown] heading_collapsed=true hidden=true
# ## 1.3. Data types
# + hidden=true
df1.dtypes
# + hidden=true
df1['date'] = pd.to_datetime(df1['date'])
# + [markdown] heading_collapsed=true hidden=true
# ## 1.4. Check NA
# + hidden=true
df1.isna().sum()
# + [markdown] heading_collapsed=true hidden=true
# ## 1.5. Fillout NA
# + code_folding=[0] hidden=true
# competition_distance 2642
df1['competition_distance'] = df1['competition_distance'].apply(lambda x: 200000.0 if math.isnan(x) else x)
# competition_open_since_month 323348
df1['competition_open_since_month'] = df1[['date', 'competition_open_since_month']].apply(lambda x: x['date'].month if math.isnan(x['competition_open_since_month']) else x['competition_open_since_month'], axis=1)
# competition_open_since_year 323348
df1['competition_open_since_year'] = df1[['date', 'competition_open_since_year']].apply(lambda x: x['date'].year if math.isnan(x['competition_open_since_year']) else x['competition_open_since_year'], axis=1)
# promo2_since_week 508031
df1['promo2_since_week'] = df1[['date', 'promo2_since_week']].apply(lambda x: x['date'].week if math.isnan(x['promo2_since_week']) else x['promo2_since_week'], axis=1)
# promo2_since_year 508031
df1['promo2_since_year'] = df1[['date', 'promo2_since_year']].apply(lambda x: x['date'].year if math.isnan(x['promo2_since_year']) else x['promo2_since_year'], axis=1)
# promo_interval 508031
df1['promo_interval'].fillna(0, inplace=True)
month_map = {1: 'Jan', 2: 'Feb', 3: 'Mar', 4: 'Apr', 5: 'May', 6: 'Jun', 7: 'Jul', 8: 'Aug', 9: 'Sep', 10: 'Oct', 11: 'Nov', 12: 'Dec'}
df1['month_map'] = df1['date'].dt.month.map(month_map)
df1['is_promo'] = df1[['promo_interval', 'month_map']].apply(lambda x: 0 if x['promo_interval'] == 0 else 1 if x['month_map'] in x['promo_interval'].split(',') else 0, axis=1)
# + [markdown] heading_collapsed=true hidden=true
# ## 1.6. Change types
# + hidden=true
df1['competition_open_since_month'] = df1['competition_open_since_month'].astype(int)
df1['competition_open_since_year'] = df1['competition_open_since_year'].astype(int)
df1['promo2_since_week'] = df1['promo2_since_week'].astype(int)
df1['promo2_since_year'] = df1['promo2_since_year'].astype(int)
# + [markdown] heading_collapsed=true hidden=true
# ## 1.7. Descriptive stastistical
# + hidden=true
num_attributes = df1.select_dtypes(include = ['int64', 'int32', 'float64'])
cat_attributes = df1.select_dtypes(exclude = ['int64', 'int32', 'float64', 'datetime64[ns]'])
# + [markdown] heading_collapsed=true hidden=true
# ### 1.7.1. Numerical attibutes
# + hidden=true
# + hidden=true
# Cntral - mean, median
ct1 = pd.DataFrame(num_attributes.apply(np.mean)).T
ct2 = pd.DataFrame(num_attributes.apply(np.median)).T
# Dispersion - min, max, range, std, skews, kurtoss
d1 = pd.DataFrame(num_attributes.apply(min)).T
d2 = pd.DataFrame(num_attributes.apply(max)).T
d3 = pd.DataFrame(num_attributes.apply(lambda x: x.max() - x.min())).T
d4 = pd.DataFrame(num_attributes.apply(np.std)).T
d5 = pd.DataFrame(num_attributes.apply(lambda x: x.skew())).T
d6 = pd.DataFrame(num_attributes.apply(lambda x: x.kurtosis())).T
# Concatenate
m = pd.concat([d1, d2, d3, ct1, ct2, d4, d5, d6]).T.reset_index()
# Renames columns
m.columns = ['attributes', 'min', 'max', 'range', 'mean', 'median', 'std', 'skwes', 'kurtosis']
# + hidden=true
m
# + hidden=true
# Vamos entender um pouco esses resultados
# Como temos uma skew positiva, em sales teremos um deslocamento do gráfico para a esquerda.
sns.distplot(df1['sales'])
# + hidden=true
# Como temos uma skew positiva, em sales teremos um deslocamento do gráfico para a esquerda. A kurtosis é positiva e possui
# um valor muito grande, logo teremos um pico muito grande tambem. ('curva de poison')
sns.distplot(df1['competition_distance'])
# + [markdown] heading_collapsed=true hidden=true
# ### 1.7.1. Categorical attibutes
# + hidden=true
cat_attributes.apply(lambda x: x.unique().shape[0])
# + hidden=true
# Gerando um gráfico boxplot
sns.boxplot(x = 'state_holiday', y = 'sales', data = df1)
# + hidden=true
# Filtrando
aux1 = df1[(df1['state_holiday'] != '0') & df1['sales'] > 0]
sns.boxplot(x = 'state_holiday', y = 'sales', data = aux1)
# + hidden=true
aux1 = df1[(df1['state_holiday'] != '0') & df1['sales'] > 0]
plt.subplot(1, 5, 1)
sns.boxplot(x = 'state_holiday', y = 'sales', data = aux1)
plt.subplot(1, 5, 3)
sns.boxplot(x = 'store_type', y = 'sales', data = aux1)
plt.subplot(1, 5, 5)
sns.boxplot(x = 'assortment', y = 'sales', data = aux1)
# -
# # 2.0. FEATURE ENGINEERING - PASSO 2
# + [markdown] heading_collapsed=true
# ## 2.1. Mapa menta de hipóteses
# + hidden=true hide_input=true
Image('img/DAILY_STORE_SALES.png')
# + [markdown] heading_collapsed=true
# ## 2.2. Criação das hipóteses
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2.1. Hipóteses Loja
# + [markdown] hidden=true
# **1.** Lojas com maior número de funcionários deveriam vender mais.
#
# **2.** Lojas com maior estoque deveriam vender mais.
#
# **3.** Lojas com maior porte deveriam vender mais.
#
# **4.** Lojas com maior sortimento deveriam vender mais.
#
# **5.** Lojas com competidores mais próximos deveriam vender menos.
#
# **6.** Lojas com competidores a mais tempo deveriam vender mais.
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2.2. Hipóteses Produtos
# + [markdown] hidden=true
# **1.** Lojas que investem mais em marketing deveriam vender mais.
#
# **2.** Lojas que expõe mais seu produto nas vetrines deveriam vender mais.
#
# **3.** Lojas que têm preços menores deveriam vender mais.
#
# **4.** Lojas preços menores por mais tempo deveriam vender mais.
#
# **5.** Lojas com promoções ativas por mais tempo deveriam vender mais.
#
# **6.** Lojas com mais dias de promoção deveriam vender mais.
#
# **7.** Lojas com promoções consecutivas deveriam vender mais.
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2.1. Hipóteses Tempo (sazonalidade)
# + [markdown] hidden=true
# **1.** Lojas com mais feriados deveriam vender mais.
#
# **2.** Lojas abertas durante o feriado de Natal deveriam vender mais.
#
# **3.** Lojas deveriam vender mais ao longo dos anos.
#
# **4.** Lojas deveriam vender mais no segundo semestre do ano.
#
# **5.** Lojas deveriam vender mais depois do dia 10 de cada mês.
#
# **6.** Lojas deveriam vender menos aos finais de semana.
#
# **7.** Lojas deveriam vender menos durante os feriados esolares.
# -
# ## 2.3. priorização - Lista final de hipóteses
# **1.** Lojas com maior sortimento deveriam vender mais.
#
# **2.** Lojas com competidores mais próximos deveriam vender menos.
#
# **3.** Lojas com competidores a mais tempo deveriam vender mais.
#
# **4.** Lojas com promoções ativas por mais tempo deveriam vender mais.
#
# **5.** Lojas com mais dias de promoção deveriam vender mais.
#
# **6.** Lojas com promoções consecutivas deveriam vender mais.
#
# **7.** Lojas abertas durante o feriado de Natal deveriam vender mais.
#
# **8.** Lojas deveriam vender mais ao longo dos anos.
#
# **9.** Lojas deveriam vender mais no segundo semestre do ano.
#
# **10.** Lojas deveriam vender mais depois do dia 10 de cada mês.
#
# **11.** Lojas deveriam vender menos aos finais de semana.
#
# **12.** Lojas deveriam vender menos durante os feriados esolares.
# + [markdown] heading_collapsed=true
# ## 2.4. Feature engineering
# + hidden=true
df2 = df1.copy()
# + hidden=true
# Derivar as variáveis
# year
df2['year'] = df2['date'].dt.year
# month
df2['month'] = df2['date'].dt.month
# day
df2['day'] = df2['date'].dt.day
# week of year
df2['week_of_year'] = df2['date'].dt.weekofyear
# year week
df2['year_week'] = df2['date'].dt.strftime('%Y-%W')
# competition since
df2['competition_since'] = df2.apply(lambda x: datetime.datetime(year=x['competition_open_since_year'],month=x['competition_open_since_month'], day=1 ), axis=1)
df2['competition_time_month'] = ((df2['date'] - df2['competition_since'])/30).apply(lambda x: x.days).astype(int)
# promo since
df2['promo_since'] = df2['promo2_since_year'].astype(str) + '-' + df2['promo2_since_week'].astype(str)
df2['promo_since'] = df2['promo_since'].apply(lambda x: datetime.datetime.strptime(x + '-1', '%Y-%W-%w')
- datetime.timedelta(days=7))
df2['promo_time_week'] = ((df2['date'] - df2['promo_since'])/7).apply(lambda x: x.days).astype(int)
# assortment
df2['assortment'] = df2['assortment'].apply(lambda x: 'basic' if x == 'a' else 'extra' if x == 'b' else 'extended' if x == 'c' else 'no_classification')
# state holiday
df2['state_holiday'] = df2['state_holiday'].apply(lambda x: 'public_holiday' if x == 'a' else 'easter_holiday' if x == 'b' else 'christmas' if x == 'c' else 'regular_day')
# + hidden=true hide_input=true
df2.sample(10).T
# + [markdown] heading_collapsed=true
# # 3.0. FILTERING VARIABLES - PASSO 3
# + hidden=true hide_input=false
df3 = df2.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## 3.1. Linear filtering
# + hidden=true hide_input=false
df3 = df3[(df3['open'] != 0) & (df3['sales'] > 0)]
# + [markdown] heading_collapsed=true hidden=true
# ## 3.2. Column selection
# + hidden=true
cols_drop = ['customers', 'open', 'promo_interval', 'month_map']
df3 = df3.drop(cols_drop, axis=1)
# + hidden=true
df3.columns
# + [markdown] hide_input=false
# # 4.0. EXPLORATORY DATA ANALYSIS - PASSO 4
# -
df4 =df3.copy()
# + [markdown] hide_input=false
# ## 4.1. Univariate analysis
# + [markdown] heading_collapsed=true hide_input=false
# ### 4.1.1. Response variable
# + hidden=true hide_input=false
f, axes = plt.subplots(1, 1, figsize=(10, 8), sharex=False)
sns.distplot(df4['sales']);
# + [markdown] heading_collapsed=true
# ### 4.1.2. Numerical variable
# + hidden=true
num_attributes.describe()
# + hidden=true
num_attributes.hist(bins=25, figsize=(18, 13));
# -
# ### 4.1.1. Categoical variable
# + hide_input=false
plt.figure(figsize = (15, 11))
# state_holiday
plt.subplot(3, 2, 1)
a = df4[df4['state_holiday'] != 'regular_day']
sns.countplot(a['state_holiday']);
plt.subplot(3, 2, 2)
sns.kdeplot(df4[df4['state_holiday'] == 'public_holiday']['sales'], label='public_holiday', shade=True);
sns.kdeplot(df4[df4['state_holiday'] == 'easter_holiday']['sales'], label='easter_holiday', shade=True);
sns.kdeplot(df4[df4['state_holiday'] == 'christmas']['sales'], label='christmas', shade=True);
# store_type
plt.subplot(3, 2, 3)
sns.countplot(df4['store_type']);
plt.subplot(3, 2, 4)
sns.kdeplot(df4[df4['store_type'] == 'a']['sales'], label='a', shade=True);
sns.kdeplot(df4[df4['store_type'] == 'b']['sales'], label='b', shade=True);
sns.kdeplot(df4[df4['store_type'] == 'c']['sales'], label='c', shade=True);
sns.kdeplot(df4[df4['store_type'] == 'd']['sales'], label='d', shade=True);
# assortment
plt.subplot(3, 2, 5)
sns.countplot(df4['assortment']);
plt.subplot(3, 2, 6)
sns.kdeplot(df4[df4['assortment'] == 'basic']['sales'], label='basic', shade=True);
sns.kdeplot(df4[df4['assortment'] == 'extended']['sales'], label='extended', shade=True);
sns.kdeplot(df4[df4['assortment'] == 'extra']['sales'], label='extra', shade=True);
# + [markdown] hide_input=false
# ## 4.2. Bivariate analysis
# + [markdown] heading_collapsed=true
# ### H1. Lojas com maior sortimento deveriam vender mais.
# **FALSO** Lojas com MAIOR SORTIMENTO vendem MENOS.
# + hidden=true hide_input=true
aux1 = df4[['assortment', 'sales']].groupby('assortment').sum().reset_index()
sns.barplot(x='assortment', y='sales', data=aux1);
aux2 = df4[['year_week', 'assortment', 'sales']].groupby(['year_week', 'assortment']).sum().reset_index()
aux2.pivot(index='year_week', columns='assortment', values='sales').plot()
aux3 = aux2[aux2['assortment'] == 'extra']
aux3.pivot(index='year_week', columns='assortment', values='sales').plot()
# + [markdown] heading_collapsed=true
# ### H2. Lojas com competidores mais próximos vendem menos.
#
# **FALSO** Lojas com COMPETIDORES MAIS PRÓXIMOS vendem MAIS.
# + hidden=true hide_input=true
plt.figure(figsize = (15, 11))
# Gráfico de pontos scatter
aux1 = df4[['competition_distance', 'sales']].groupby('competition_distance').sum().reset_index()
plt.subplot(1, 3, 1)
sns.scatterplot(x='competition_distance', y='sales', data=aux1);
plt.xticks(rotation=90)
# Criando grupos de 1000 em 1000 de competition_distance que vai de 0 a 20000
bins=list(np.arange(0, 20000, 1000))
# Criando nova coluna com a informação do grupo a qual os valores de competition_distance pertence
aux1['competition_distance_binned'] = pd.cut(aux1['competition_distance'], bins=bins)
# Soma dos valores das vendas de cada grupo
aux2 = aux1[['competition_distance_binned', 'sales']].groupby('competition_distance_binned').sum().reset_index()
# Gráfico de barras
plt.subplot(1, 3, 2)
sns.barplot(x='competition_distance_binned', y='sales', data=aux2);
plt.xticks(rotation=90)
# Gráfico de força (correlação)
plt.subplot(1, 3, 3)
sns.heatmap(aux1.corr(method='pearson'), annot=True);
# + [markdown] heading_collapsed=true
# ### H3. Lojas com competidores à mais tempo deveriam vender mais.
#
# **FALSO** Lojas com COMPETIDORES À MAIS TEMPO vendem MENOS.
# + hidden=true hide_input=true
plt.figure(figsize = (17, 9))
aux1 = df4[['competition_time_month', 'sales']].groupby('competition_time_month').sum().reset_index()
plt.subplot(1, 3, 1)
aux2 = aux1[(aux1['competition_time_month'] < 120) & (aux1['competition_time_month'] != 0)]
sns.barplot(x='competition_time_month', y='sales', data=aux2);
plt.xticks(rotation=90, fontsize=12);
plt.grid()
plt.subplot(1, 3, 2)
sns.regplot(x='competition_time_month', y='sales', data=aux2);
plt.grid(color='k', linestyle='--', linewidth=1)
plt.subplot(1, 3, 3)
sns.heatmap(aux1.corr(method='pearson'), annot=True);
# + [markdown] hide_input=false
# ## 4.3. Multivariate analysis
# -
| my_project_rossmann_store_sales_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python_3.6
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import mglearn
from sklearn.linear_model import Ridge
from sklearn.model_selection import train_test_split
# %matplotlib inline
# +
X, y = mglearn.datasets.load_extended_boston()
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
ridge = Ridge().fit(X_train, y_train)
print("Training set score: {:.2f}".format(ridge.score(X_train, y_train)))
print("Test set score_ {:.2f}".format(ridge.score(X_test, y_test)))
# -
# The training set score is lower than the one gotten from LinearRegression (0.95), but the test set score is greater.
#
# In Linear regression we were overfitting our data, but Ridge is a more restricted model, so we are less likely to overfit.
#
# A less complex model means worse performance on the training set, but better generalization.
# How much importance the model places on simplicity versus training set performance can be specified by the user, using the alpha parameter. In the previous example, we used the default parameter alpha=1.0. The optimum setting of alpha depends on the particular dataset we are using. Increasing alpha forces coefficients to move more toward zero, which decreases training set performance but might help generalization.
ridge10 = Ridge(alpha=10).fit(X_train, y_train)
print("Training set score: {:.2f}".format(ridge10.score(X_train, y_train)))
print("Test set score: {:.2f}".format(ridge10.score(X_test, y_test)))
# Decreasing alpha allows the coefficients to be less restricted, meaning we move right to the correct value. For very small values of alpha, coefficients are barely restricted at all, and we end up with a model that resembles LinearRegression
ridge01 = Ridge(alpha=0.1).fit(X_train, y_train)
print("Training set score: {:.2f}".format(ridge01.score(X_train,y_train)))
print("Test set score: {:.2f}".format((ridge01.score(X_test, y_test))))
# Here, alpha=0.1 seems to be working well. We could try decreasing alpha even more to improve generalization. For now, notice how the parameter alpha corresponds to the model complexity as shown in Figure 2-1.
# We can also get a more qualitative insight into how the alpha parameter changes the model by inspecting the coef_ attribute of models with different values of alpha. A higher alpha means a more restricted model, so we expect the entries of coef_ to have smaller magnitude for a high value of alpha than for a low value of alpha.
# For comparison purposes we add the Linear Regression
from sklearn.linear_model import LinearRegression
lr = LinearRegression().fit(X_train, y_train)
plt.plot(ridge.coef_, 's', label="Ridge alpha=1")
plt.plot(ridge10.coef_, '^', label="Ridge alpha=10")
plt.plot(ridge01.coef_, 'v', label="Ridge alpha=.1")
plt.plot(lr.coef_, 'o', label="Linear Regression")
plt.xlabel("Coefficient index")
plt.ylabel("Coefficient magnitude")
plt.hlines(0, 0, len(lr.coef_))
plt.ylim(-25, 25)
plt.legend()
# Another way to understand the influence of regularization is to fix a value of alpha but vary the amount of training data available. For Figure below, we subsampled the Boston Housing dataset and evaluated LinearRegression and Ridge(alpha=1) on subsets of increasing size (plots that show model performance as a function of dataset size are called learning curves):
mglearn.plots.plot_ridge_n_samples()
| Source/Chapter02/LinearModels/Regression/ridgeRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Let's import some libraries first...
# +
import pandas
from pandas.plotting import scatter_matrix
from sklearn import datasets
from sklearn import model_selection
from sklearn import linear_model
# models
from sklearn.ensemble import RandomForestRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.preprocessing import StandardScaler, PolynomialFeatures
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_squared_error, r2_score
import matplotlib.pyplot as plt
# -
# Load and examine dataset...
dataset = pandas.read_csv("data/ccp-consumer-lending-half-year.csv")
print (dataset)
# +
array = dataset.values
num_data_points = 3
for i in range(0, len(array) - (num_data_points - 1)):
begin = i
end = i + (num_data_points - 1)
X = array[begin:end+1,2:4] # data = avg_gross_loan_book, net_lending
Y = array[begin:end+1,1] # result = NPAT
model = LinearRegression()
model.fit(X, Y) # train model
print("Period %s to %s: " % (array[begin,0], array[end,0]), end =" ")
print("p = %sbr + %sl + %s" % (model.coef_[0], model.coef_[1], model.intercept_))
# -
# Where
# ```
# p = Net profit before tax (NPBT).
# r = Reporting period. Full year = 2, half year = 1.
# b = Average gross loan book.
# l = Net lending for the period.
# ```
# ## FY19 Predictions
#
# Assumptions:
#
# * Average gross loan book will be $196m.
#
# * Net lending will be $50m, on the upper range of the forecast. Quoting a high number here will actually reduce EBIT.
#
# +
b = 196000
l = 50000
r = 2
p = model.coef_[0] * b * r + model.coef_[1] * l + model.intercept_
print("EBIT = %s" % p)
print("NPAT = %s" % (p * 0.7))
| notebooks/ccp/CreditCorpConsumerLending-WayOffWithLinear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building a NER System
#
# A simple approach to building an NER system is to maintain a large collection of person/organization/location names that are the most relevant to our company (e.g., names of all clients, cities in their addresses, etc.); this is typically referred to as a gazetteer.
#
# Rule-based NER, which can be based on a compiled list of patterns based on word tokens and POS tags.
#
# Train an ML model, which can predict the named entities in unseen text.
# - Normal classifier: classify text word by word
# - Sequence classifier: looking at the context in which it's being used. (For NER models)
#
# Conditional Random Fields (CRFs), one of the popular sequence classifier training algorithms.
#
# Typical training data for NER follow BIO notation:
# - B indicates the beginning of an entity
# - I inside an entity, indicates when entities comprise more than one word
# - O other, indicates non-entities.
#
# Example: 'Peter' gets tagged as a B-PER
# # NER Training
import nltk
nltk.download('averaged_perceptron_tagger')
from nltk.tag import pos_tag
from sklearn_crfsuite import CRF, metrics
from sklearn.metrics import make_scorer,confusion_matrix
from pprint import pprint
from sklearn.metrics import f1_score,classification_report
from sklearn.pipeline import Pipeline
import string
# loading data
"""
Load the training/testing data.
input: conll format data, but with only 2 tab separated colums - words and NEtags.
output: A list where each item is 2 lists. sentence as a list of tokens, NER tags as a list for each token.
"""
def load__data_conll(file_path):
myoutput,words,tags = [],[],[]
fh = open(file_path)
for line in fh:
line = line.strip()
if "\t" not in line:
#Sentence ended.
myoutput.append([words,tags])
words,tags = [],[]
else:
word, tag = line.split("\t")
words.append(word)
tags.append(tag)
fh.close()
return myoutput
"""
Get features for all words in the sentence
Features:
- word context: a window of 2 words on either side of the current word, and current word.
- POS context: a window of 2 POS tags on either side of the current word, and current tag.
input: sentence as a list of tokens.
output: list of dictionaries. each dict represents features for that word.
"""
def sent2feats(sentence):
feats = []
sen_tags = pos_tag(sentence) #This format is specific to this POS tagger!
for i in range(0,len(sentence)):
word = sentence[i]
wordfeats = {}
#word features: word, prev 2 words, next 2 words in the sentence.
wordfeats['word'] = word
if i == 0:
wordfeats["prevWord"] = wordfeats["prevSecondWord"] = "<S>"
elif i==1:
wordfeats["prevWord"] = sentence[0]
wordfeats["prevSecondWord"] = "</S>"
else:
wordfeats["prevWord"] = sentence[i-1]
wordfeats["prevSecondWord"] = sentence[i-2]
#next two words as features
if i == len(sentence)-2:
wordfeats["nextWord"] = sentence[i+1]
wordfeats["nextNextWord"] = "</S>"
elif i==len(sentence)-1:
wordfeats["nextWord"] = "</S>"
wordfeats["nextNextWord"] = "</S>"
else:
wordfeats["nextWord"] = sentence[i+1]
wordfeats["nextNextWord"] = sentence[i+2]
#POS tag features: current tag, previous and next 2 tags.
wordfeats['tag'] = sen_tags[i][1]
if i == 0:
wordfeats["prevTag"] = wordfeats["prevSecondTag"] = "<S>"
elif i == 1:
wordfeats["prevTag"] = sen_tags[0][1]
wordfeats["prevSecondTag"] = "</S>"
else:
wordfeats["prevTag"] = sen_tags[i - 1][1]
wordfeats["prevSecondTag"] = sen_tags[i - 2][1]
# next two words as features
if i == len(sentence) - 2:
wordfeats["nextTag"] = sen_tags[i + 1][1]
wordfeats["nextNextTag"] = "</S>"
elif i == len(sentence) - 1:
wordfeats["nextTag"] = "</S>"
wordfeats["nextNextTag"] = "</S>"
else:
wordfeats["nextTag"] = sen_tags[i + 1][1]
wordfeats["nextNextTag"] = sen_tags[i + 2][1]
#That is it! You can add whatever you want!
feats.append(wordfeats)
return feats
# Extracting features
#Extract features from the conll data, after loading it.
def get_feats_conll(conll_data):
feats = []
labels = []
for sentence in conll_data:
feats.append(sent2feats(sentence[0]))
labels.append(sentence[1])
return feats, labels
# training a model
#Train a sequence model
def train_seq(X_train,Y_train,X_dev,Y_dev):
# crf = CRF(algorithm='lbfgs', c1=0.1, c2=0.1, max_iterations=50, all_possible_states=True)
crf = CRF(algorithm='lbfgs', c1=0.1, c2=10, max_iterations=50)#, all_possible_states=True)
#Just to fit on training data
crf.fit(X_train, Y_train)
labels = list(crf.classes_)
#testing:
y_pred = crf.predict(X_dev)
sorted_labels = sorted(labels, key=lambda name: (name[1:], name[0]))
print(metrics.flat_f1_score(Y_dev, y_pred,average='weighted', labels=labels))
print(metrics.flat_classification_report(Y_dev, y_pred, labels=sorted_labels, digits=3))
#print(metrics.sequence_accuracy_score(Y_dev, y_pred))
get_confusion_matrix(Y_dev, y_pred,labels=sorted_labels)
# +
def print_cm(cm, labels):
print("\n")
"""pretty print for confusion matrixes"""
columnwidth = max([len(x) for x in labels] + [5]) # 5 is value length
empty_cell = " " * columnwidth
# Print header
print(" " + empty_cell, end=" ")
for label in labels:
print("%{0}s".format(columnwidth) % label, end=" ")
print()
# Print rows
for i, label1 in enumerate(labels):
print(" %{0}s".format(columnwidth) % label1, end=" ")
sum = 0
for j in range(len(labels)):
cell = "%{0}.0f".format(columnwidth) % cm[i, j]
sum = sum + int(cell)
print(cell, end=" ")
print(sum) #Prints the total number of instances per cat at the end.
#python-crfsuite does not have a confusion matrix function,
#so writing it using sklearn's confusion matrix and print_cm from github
def get_confusion_matrix(y_true,y_pred,labels):
trues,preds = [], []
for yseq_true, yseq_pred in zip(y_true, y_pred):
trues.extend(yseq_true)
preds.extend(yseq_pred)
print_cm(confusion_matrix(trues,preds,labels),labels)
# +
train_path = 'Data/conlldata/train.txt'
test_path = 'Data/conlldata/test.txt'
conll_train = load__data_conll(train_path)
conll_dev = load__data_conll(test_path)
print("Training a Sequence classification model with CRF")
feats, labels = get_feats_conll(conll_train)
devfeats, devlabels = get_feats_conll(conll_dev)
train_seq(feats, labels, devfeats, devlabels)
print("Done with sequence model")
# -
conll_train[0]
feats[0]
# In real-world scenarios, using the trained model by itself won’t be sufficient, as the data keeps changing and new entities keep getting added, and there will also be some domain-specific entities or patterns that were not seen in generic training datasets. Hence, most NER systems deployed in real-world scenarios use a combination of ML models, gazetteers, and some pattern matching–based heuristics to improve their performance
# # NER Using an Existing Library (Spacy)
# +
# Google Colab
# https://colab.research.google.com/drive/1z1hHpd8emVHUhth5hnp-fj0UXLsmWlPZ?usp=sharing
# -
# # NER Using Active Learning
#
# The best approach to NER when we want customized solutions but don’t want to train everything from scratch is to start with an off-the-shelf product and either augment it with customized heuristics for our problem domain (using tools such as RegexNER or EntityRuler) and/or use active learning using tools like Prodigy.
#
# TIPS: Start with a pre-trained NER model and enhance it with heuristics, active learning, or both.
# +
# Google Colab NER BERT
# https://colab.research.google.com/drive/1z1hHpd8emVHUhth5hnp-fj0UXLsmWlPZ?usp=sharing
# -
# # Practical Advice
# - NER is very sensitive to the format of its input. It’s more accurate with well-formatted plain text than with, say, a PDF document from which plain text needs to be extracted first. One approach is to do custom post-processing of PDFs to extract blobs of text, then run NER on the blobs.
# - NER is also very sensitive to the accuracy of the prior steps in its processing pipeline: sentence splitting, tokenization, and POS tagging. So, some amount of pre-processing may be necessary before passing a piece of text into an NER model to extract entities.
#
# TIPS: If you’re working with documents, such as reports, etc., pre-process them to extract text blobs, then run NER on them.
| 05_Information Extraction/03_Named Entity Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("../datos/tce/tipos-cambio-minorista-cotizaciones-panel.csv")
df[df.entity == "bullmarket"].groupby(["entity", "coin", "hour", "channel"]).count().sort_values("indice_tiempo", ascending=False)[["value"]]
| notebooks/intersecciones-vacias-tce.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SkillFactory
# ## Введение в ML, введение в sklearn
# В этом задании мы с вами рассмотрим данные с конкурса [Задача предсказания отклика клиентов ОТП Банка](http://www.machinelearning.ru/wiki/index.php?title=%D0%97%D0%B0%D0%B4%D0%B0%D1%87%D0%B0_%D0%BF%D1%80%D0%B5%D0%B4%D1%81%D0%BA%D0%B0%D0%B7%D0%B0%D0%BD%D0%B8%D1%8F_%D0%BE%D1%82%D0%BA%D0%BB%D0%B8%D0%BA%D0%B0_%D0%BA%D0%BB%D0%B8%D0%B5%D0%BD%D1%82%D0%BE%D0%B2_%D0%9E%D0%A2%D0%9F_%D0%91%D0%B0%D0%BD%D0%BA%D0%B0_%28%D0%BA%D0%BE%D0%BD%D0%BA%D1%83%D1%80%D1%81%29)
# Думаю, так удобнее :-)
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
plt.rcParams['figure.figsize'] = (12,5)
# -
# ### Грузим данные
# Считаем описание данных
# +
df_descr = pd.read_csv('data/otp_description.csv', sep='\t', encoding='utf8')
# -
df_descr
# Считаем обучающую выборки и тестовую (которую мы как бы не видим)
df_train = pd.read_csv('data/otp_train.csv', sep='\t', encoding='utf8')
df_train.shape
df_test = pd.read_csv('data/otp_test.csv', sep='\t', encoding='utf8')
df_test.shape
df_train.head()
# ## Объединим две выборки
#
# Так как пока мы пока не умеем работать sklearn Pipeline, то для того, чтобы после предобработки столбцы в двух выборках находились на своих местах.
#
# Для того, чтобы в дальнейшем отделить их введем новый столбец "sample"
df_train.loc[:, 'sample'] = 'train'
df_test.loc[:, 'sample'] = 'test'
df = df_test.append(df_train).reset_index(drop=True)
df.shape
# ### Чуть-чуть посмотрим на данные
# Посмотрим типы данных и их заполняемость
df.info()
# Видим, что часть данных - object, скорее всего стоки.
#
#
# Давайте выведем эти значения для каждого столбца
for i in df_train.columns: # перебираем все столбцы
if str(df_train[i].dtype) == 'object': # если тип столбца - object
print('='*10)
print(i) # выводим название столбца
print(set(df_train[i])) # выводим все его значения (но делаем set - чтоб значения не повторялись)
print('\n') # выводим пустую строку
# Mожно заметить что некоторые переменные, которые обозначены как строки (например PERSONAL_INCOME) на самом деле числа, но по какой-то причине были распознаны как строки
#
# Причина же что использовалась запятая для разделения не целой части числа..
# Перекодировать их можно например так:
df['PERSONAL_INCOME'].map(lambda x: x.replace(',', '.')).astype('float')
# Такой эффект наблюдается в столбцах `PERSONAL_INCOME`, `CREDIT`, `FST_PAYMENT`, `LOAN_AVG_DLQ_AMT`, `LOAN_MAX_DLQ_AMT`
# ### Теперь ваше небольшое исследование
# #### Задание 1. Есть ли пропуски в данных? Что с ними сделать?
#
# (единственного верного ответа нет - аргументируйте)
# #### Ответ:
#
# Из результатов выше, мы знаем, что есть пропущенные значения. Посчитаем число пропусков в данных.
nan_vals_count = df.isnull().sum()
nan_vals_count.iloc[nan_vals_count.nonzero()[0]]
def show_data(dataframe, columns=None):
for i in df_train.columns if columns is None else columns: # перебираем столбцы
print('='*10)
print(i) # выводим название столбца
print(set(df_train[i])) # выводим все его значения (но делаем set - чтоб значения не повторялись)
print('\n') # выводим пустую строку
columns_with_na = ['GEN_INDUSTRY', 'GEN_TITLE', 'ORG_TP_STATE', 'ORG_TP_FCAPITAL', 'JOB_DIR', 'TP_PROVINCE', 'REGION_NM', 'WORK_TIME']
show_data(df, columns_with_na)
# #### Задание 2. Есть ли категориальные признаки? Что с ними делать?
# Есть и их нужно закодировать.
# #### Задание 3. Фунция предобработки
# Напишите функцию, которая бы
#
# * Удаляло идентификатор `AGREEMENT_RK`
# * Избавлялась от проблем с '.' и ',' в стобцах PERSONAL_INCOME, CREDIT, FST_PAYMENT, LOAN_AVG_DLQ_AMT, LOAN_MAX_DLQ_AMT
# * Что-то делала с пропусками
# * Кодировала категориальные признаки
#
# В результате, ваш датафрейм должен содержать только числа и не содержать пропусков!
def preproc_data(df_input):
df_output = df_input.copy()
## Your Code Here
df_output = df_output.drop(['AGREEMENT_RK'], axis=1)
columns_to_fix = ['PERSONAL_INCOME', 'CREDIT', 'FST_PAYMENT', 'LOAN_AVG_DLQ_AMT', 'LOAN_MAX_DLQ_AMT']
for column in columns_to_fix:
df_output[column] = df_output[column].map(lambda x: x.replace(',', '.')).astype('float')
family_income_list = ['до 5000 руб.', 'от 5000 до 10000 руб.', 'от 10000 до 20000 руб.', 'от 20000 до 50000 руб.', 'свыше 50000 руб.']
for idx, val in enumerate(family_income_list):
df_output['FAMILY_INCOME'].replace(to_replace = val, value="{}".format(idx), inplace=True)
df_output['FAMILY_INCOME'] = df_output.FAMILY_INCOME.astype('int64')
df_output.WORK_TIME[df_output.WORK_TIME.isnull()] = df_output.WORK_TIME.mean()
df_output = pd.get_dummies(df_output, columns=['EDUCATION', 'MARITAL_STATUS'])
df_output = df_output.drop(['GEN_INDUSTRY', 'GEN_TITLE', 'ORG_TP_STATE', 'ORG_TP_FCAPITAL', 'JOB_DIR', 'TP_PROVINCE', 'REGION_NM',
'REG_ADDRESS_PROVINCE', 'FACT_ADDRESS_PROVINCE', 'POSTAL_ADDRESS_PROVINCE', 'PREVIOUS_CARD_NUM_UTILIZED'], axis=1)
return df_output
# +
df_preproc = df.pipe(preproc_data)
df_train_preproc = df_preproc.query('sample == "train"').drop(['sample'], axis=1)
df_test_preproc = df_preproc.query('sample == "test"').drop(['sample'], axis=1)
# -
# #### Задание 4. Отделите целевую переменную и остальные признаки
#
# Должно получится:
# * 2 матрицы: X и X_test
# * 2 вектора: y и y_test
# +
y = df_train_preproc['TARGET']
X = df_train_preproc.drop('TARGET', axis=1)
y_test = df_test_preproc['TARGET']
X_test = df_test_preproc.drop('TARGET', axis=1)
# -
# #### Задание 5. Обучение и оценка качества разных моделей
# +
from sklearn.model_selection import train_test_split
# test_size=0.3, random_state=42
## Your Code Here
target = df_preproc['TARGET']
X = df_preproc.drop(['sample', 'TARGET'], axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, target, test_size=0.3, random_state=42)
# +
# Попробовать следующие "черные ящики": интерфейс одинаковый
# fit,
# predict,
# predict_proba
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
## Your Code Here
dtc = DecisionTreeClassifier()
dtc.fit(X_train, y_train)
predict_dtc = dtc.predict(X_test)
predict_proba_dtc = dtc.predict_proba(X_test)
rfc = RandomForestClassifier()
rfc.fit(X_train, y_train)
predict_rfc = rfc.predict(X_test)
predict_proba_rfc = rfc.predict(X_test)
lr = LogisticRegression()
lr.fit(X_train, y_train)
predict_lr = lr.predict(X_test)
predict_proba_lr = lr.predict_proba(X_test)
# +
# Посчитать метрики стандартные
# accuracy, precision, recall
from sklearn.metrics import accuracy_score, precision_score, recall_score
## Your Code Here
print('DecisionTreeClassifier:')
print('precision', precision_score(y_test, predict_dtc))
print('recall', recall_score(y_test, predict_dtc))
print('accuracy', accuracy_score(y_test, predict_dtc))
print('='*10)
print('RandomForestClassifier')
print('precision: {}'.format(precision_score(y_test, predict_rfc)))
print('recall: {}'.format(recall_score(y_test, predict_rfc)))
print('accuracy: {}'.format(accuracy_score(y_test, predict_rfc)))
print('='*10)
print('LogisticRegression')
print('precision: {}'.format(precision_score(y_test, predict_lr)))
print('recall: {}'.format(recall_score(y_test, predict_lr)))
print('accuracy: {}'.format(accuracy_score(y_test, predict_lr)))
print('='*10)
# +
# Визуалищировать эти метрики всех моделей на одном графике (чтоб визуально посмотреть)
## Your Code Here
from matplotlib import pyplot as plt
# %matplotlib inline
fig, ax = plt.subplots(figsize=(15,10))
precision1 = precision_score(y_test, predict_dtc)
precision2 = precision_score(y_test, predict_rfc)
precision3 = precision_score(y_test, predict_lr)
recall1 = recall_score(y_test, predict_dtc)
recall2 = recall_score(y_test, predict_rfc)
recall3 = recall_score(y_test, predict_lr)
accuracy1 = accuracy_score(y_test, predict_dtc)
accuracy2 = accuracy_score(y_test, predict_rfc)
accuracy3 = accuracy_score(y_test, predict_lr)
fd = pd.DataFrame([[precision1, precision2, precision3],[recall1, recall2, recall3], [accuracy1, accuracy2, accuracy3]],
index=['precision', 'recall', 'accuracy'], columns=pd.Index(['DecisionTreeClassifier','RandomForestClassifier','LogisticRegression'])).plot(kind='bar', ax=ax)
# +
from sklearn.metrics import roc_curve, auc
plt.figure(figsize=(10, 7))
fpr, tpr, thresholds = roc_curve(y_test, predict_proba_lr[:,1])
roc_auc = auc(fpr, tpr)
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('LR', roc_auc))
fpr2, tpr2, thresholds2 = roc_curve(y_test, predict_proba_dtc[:,1])
roc_auc2 = auc(fpr2, tpr2)
plt.plot(fpr2, tpr2, label='%s ROC (area = %0.2f)' % ('DTC', roc_auc2))
fpr3, tpr3, thresholds3 = roc_curve(y_test, predict_proba_rfc)
roc_auc3 = auc(fpr3, tpr3)
plt.plot(fpr3, tpr3, label='%s ROC (area = %0.2f)' % ('RFC', roc_auc3))
plt.plot([0, 1], [0, 1])
plt.ylabel('tpr')
plt.xlabel('fpr')
plt.grid(True)
plt.title('ROC curve')
plt.xlim((-0.01, 1.01))
plt.ylim((-0.01, 1.01))
plt.legend(loc=0, fontsize='small')
# +
from sklearn.cross_validation import cross_val_score
from sklearn.model_selection import StratifiedKFold
# Сделать k-fold (10 фолдов) кросс-валидацию каждой модели
# И посчитать средний roc_auc
skf = StratifiedKFold(n_splits=10, shuffle=True, random_state=123)
## Your Code Here
print('dtc: ', cross_val_score(dtc, X_train, y_train, scoring='roc_auc', cv=10))
print('rfc: ', cross_val_score(rfc, X_train, y_train, scoring='roc_auc', cv=10))
print('lr: ', cross_val_score(lr, X_train, y_train, scoring='roc_auc', cv=10))
print('dtc- mean: ', np.mean(cross_val_score(dtc, X_train, y_train, scoring='roc_auc', cv=10)))
print('rfc- mean: ', np.mean(cross_val_score(rfc, X_train, y_train, scoring='roc_auc', cv=10)))
print('lr- mean: ', np.mean(cross_val_score(lr, X_train, y_train, scoring='roc_auc', cv=10)))
# +
# Взять лучшую модель и сделать predict (с вероятностями (!!!)) для test выборки
## Your Code Here
predict_lr_test = lr.predict_proba(X_test)
print(predict_lr_test)
# -
# Померить roc_auc на тесте
print('lr: ', cross_val_score(lr, X_test, y_test, scoring='roc_auc', cv=10))
| hw_02/ml-intro-hw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # VISST data viewer
#
# This code views data from VISST netcdf files created by <NAME>.
# Load required libraries
from netCDF4 import Dataset
import numpy as np
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
import datetime
import math
import glob
# %matplotlib inline
# +
year = 2006
day = 1
month = 1
hour = 4
minute = 0
second = 0
year_str = "%04d" % year
day_str = "%02d" % day
month_str = "%02d" % month
data_path = '/home/rjackson/data/visst/'
print('Looking for files with format' + data_path + 'twpvisstgrid*' + year_str + month_str + day_str + '*.cdf')
data_list = glob.glob(data_path + 'twpvisstgrid*' + year_str + month_str + day_str + '*.cdf')
data_list
# -
# # NetCDF parameters in VISST file:
#
# latitude = latitude of pixel<br />
# longtiude = longtidue of pixel<br />
# time = time in seconds after midnight<br />
# cloud_percentage = Cloud percentage<br />
# optical_depth_linear = Linear average of optical depth<br />
# optical_depth_log = Log average of optical depth<br />
# optical_depth_linear_sd = Linear standard deviation of optical depth<br />
# optical_depth_log_sd = Log standard deviation of optical depth<br />
# ir_emit = IR emissivity average<br />
# it_emit_sd = IR emissiviey standard deviation<br />
# particle_size = Average particle size<br />
# particle_size_sd = Particle size standard deviation<br />
# water_path = Water path average<br />
# water_path_sd = Water path standard deviation<br />
# cloud_height_top = Cloud height top average<br />
# cloud_height_top_sd = Cloud height top standard deviation<br />
# cloud_height_base = Cloud base height average<br />
# cloud_height_base_sd = Cloud base height sd<br />
# cloud_pressure_top_sd = Cloud pressure top standard deviation<br />
# cloud_pressure_center_average = Cloud pressure center average<br />
# cloud_pressure_center_sd = Cloud pressure center std dev.<br />
# cloud_pressure_base = Cloud base pressure<br />
# ir_temperature = Infrared temperature average<br />
# broadband_shortwave_albedo = Broadband shortwave albedo average<br />
# broadband_longwave_flux = Broadband longwave flux<br />
# surface_net_shortwave_flux = Surface net shortwave flux average<br />
# surface_net_longwave_flux = Surface net longwave flux average<br />
# surface_down_shortwave_flux = Shortwave downward flux average<br />
# surface_down_longwave_flux = Longwave downward flux average<br />
#
# clearsky_ir_temperature = long_name: Infrared clear sky temperature average<br />
# clearsky_vis_reflectance = Visible clear sky reflectance average<br />
# cloud_temperature = Cloud temperature average<br />
# cloud_temperature_sd = Cloud temperature standard deviation<br />
# visible_reflectance = Visible reflectance average<br />
# cloud_percentage_level = Cloud percentage at 3 levels<br />
# cloud_temperature_top_level = Cloud temperature top average at 3 levels<br />
# cloud_temperature_center_level = Cloud temperature center average at 3 levels<br />
# cloud_temperature_base_level = Cloud temperature base average at 3 levels<br />
# cloud_pressure_top_level = Cloud pressure top average at 3 levels<br />
# cloud_pressure_center_level = Cloud pressure center average at 3 levels<br />
# cloud_pressure_base_level = Cloud pressure base average at 3 levels<br />
# optical_depth_linear_level = Optical depth linear average at 3 levels<br />
# optical_depth_log_level = Optical depth log average at 3 levels<br />
# cloud_height_top_level = Cloud height top average at 3 levels<br />
# cloud_height_center_level = Cloud height center average at 3 levels<br />
# cloud_height_base_level = Cloud height base average at 3 levels<br />
#
# The data frequency is once an hour. Since different satellites were used from 2003 to 2014 in the VISST database,<br /> different parameters at different sampling frequencies may be available. <br />
#
# Dimensions for 3D variables are time by lat by lon by (level or cloud type)
#
# +
cdf_data = Dataset(data_list[0], mode='r')
# Load lat, lon, and time parameters
Latitude = cdf_data.variables['latitude']
Longitude = cdf_data.variables['longitude']
Time = cdf_data.variables['time']
# Put lat and lon into 2D grid
Lon, Lat = np.meshgrid(Longitude, Latitude)
# Load brightness temperature
IRBrightness = cdf_data.variables['ir_temperature']
# -
# Since time is in seconds after midnight, we need to use datetime to convert it to HH:MM:SS for the plot
# +
# Find the index in the file that is closest to the timestamp you specify
def find_closest_timestep_index(scan_hour, scan_minute, scan_second=0):
time_stamp = datetime.datetime(year,
month,
day,
scan_hour,
scan_minute,
scan_second)
indicies = np.arange(1, len(Time))
# Set the initial value for minimum difference between scan time and input time to be unrealistically large (> 1 day)
min_delta_time = datetime.timedelta(40)
for i in indicies:
hours = math.floor(Time[i]/3600)
minutes = math.floor((Time[i] - hours*3600)/60)
seconds = Time[i] - hours*3600 - minutes*60;
temp = datetime.datetime(year,
month,
day,
int(hours),
int(minutes),
int(seconds))
delta_time = abs(temp - time_stamp)
if(delta_time < min_delta_time):
min_delta_time = delta_time
lowest_index = i
return lowest_index
# Convert seconds to midnight to a string format
def seconds_to_midnight_to_string(time_secs_after_midnight):
hours = math.floor(time_secs_after_midnight/3600)
minutes = math.floor((time_secs_after_midnight - hours*3600)/60)
temp = datetime.time(int(hours), int(minutes), )
return temp.strftime('%H:%M:%S')
def dms_to_decimal(deg, minutes, seconds):
return deg+minutes/60+seconds/3600
# -
IRBrightness
# Plot the satellite image over Darwin. Right now, the grid is centered in a 500 km by 500 km over the TWP site in Darwin.
#
# TWP Darwin Location: 12° 25' 28.56" S, 130° 53' 29.75" E
# +
# Set up projection
Darwin_Lat = dms_to_decimal(-12,25,28.56)
Darwin_Lon = dms_to_decimal(130,53,29.75)
m = Basemap(width=2000000, height=2000000,
resolution='l', projection='stere', \
lat_0=Darwin_Lat,
lon_0=Darwin_Lon)
xi, yi = m(Lon, Lat)
darwin_x, darwin_y = m(Darwin_Lon, Darwin_Lat)
# Plot data
figure = plt.figure(figsize=(8,6))
index = find_closest_timestep_index(hour, minute, second)
data = np.squeeze(IRBrightness[index,:,:,0])
colors = m.pcolor(xi, yi, data, cmap='gray', vmin=170, vmax=300)
plt.text(darwin_x, darwin_y, 'Darwin', fontweight='bold', color='blue')
plt.plot(darwin_x*0.985, darwin_y*1.01, 'b.')
m.drawparallels(np.arange(-80., 81., 10.), labels=[1,0,0,0], fontsize=10)
m.drawmeridians(np.arange(-180., 181., 10.), labels=[0,0,0,1], fontsize=10)
m.drawcoastlines()
m.drawcountries()
m.colorbar()
plt.title('IR Brightness Temperature '
+ year_str
+ '-'
+ month_str
+ '-'
+ day_str
+ ' '
+ seconds_to_midnight_to_string(Time[index]))
# -
| notebooks/VISST data viewer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.4 ('Baduk')
# language: python
# name: python3
# ---
# ### Model v1
#
# This version of modeling is with no features. This notebook tries several different models with same dataset.
# +
import pandas as pd
games_df = pd.read_csv("../data/processed/data.csv")
games_df
# +
labeled_df = games_df.drop(['win_name', 'lose_name', 'date'], axis=1)
labeled_df = labeled_df.rename(columns={'win_color': 'player_color', 'lose_color': 'opponent_color', 'win_rank': 'player_rank', 'lose_rank': 'opponent_rank'})
labeled_df['target'] = 1
labeled_df
# +
reversed_df = labeled_df.copy(deep=True)
reversed_df = reversed_df.rename(columns={'player_color':'opponent_color', 'opponent_color':'player_color', 'player_rank':'opponent_rank', 'opponent_rank':'player_rank'})
reversed_df['target'] = 0
reversed_df
# +
processed_df = pd.concat([labeled_df,reversed_df], ignore_index=True)
processed_df = processed_df[(~processed_df['player_rank'].isin(['Ama.', 'Insei', 'NR'])) & (~processed_df['opponent_rank'].isin(['Ama.', 'Insei', 'NR']))]
processed_df['player_rank'] = processed_df['player_rank'].apply(lambda x: int(x[0]))
processed_df['opponent_rank'] = processed_df['opponent_rank'].apply(lambda x: int(x[0]))
processed_df['player_color'] = processed_df['player_color'].apply(lambda x: 1 if x == 'b' else 0)
processed_df['opponent_color'] = processed_df['opponent_color'].apply(lambda x: 1 if x == 'w' else 0)
processed_df
# +
from sklearn.model_selection import train_test_split
X, y = processed_df[['player_color', 'opponent_color', 'player_rank', 'opponent_rank', 'komi']], processed_df['target']
X_train, X_test, y_train, y_test = train_test_split(X, y)
# -
# ### Random Forest Classification
# +
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier()
clf.fit(X_train, y_train)
clf.score(X_test, y_test)
# -
# We only get 55% accuracy. This makes sense though. We only have player and opponent's rank plus the komi, to make predictions based off of. I won't really try further with different models.
| models/v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
############## PLEASE RUN THIS CELL FIRST! ###################
# import everything and define a test runner function
from importlib import reload
from helper import run
import ecc, helper, tx, script
# -
# Signing Example
from ecc import G, N
from helper import hash256
secret = 1800555555518005555555
z = int.from_bytes(hash256(b'ECDSA is awesome!'), 'big')
k = 12345
r = (k*G).x.num
s = (z+r*secret) * pow(k, N-2, N) % N
print(hex(z), hex(r), hex(s))
print(secret*G)
# Verification Example
from ecc import S256Point, G, N
z = 0xbc62d4b80d9e36da29c16c5d4d9f11731f36052c72401a76c23c0fb5a9b74423
r = 0x37206a0610995c58074999cb9767b87af4c4978db68c06e8e6e81d282047a7c6
s = 0x8ca63759c1157ebeaec0d03cecca119fc9a75bf8e6d0fa65c841c8e2738cdaec
point = S256Point(0x04519fac3d910ca7e7138f7013706f619fa8f033e6ec6e09370ea38cee6a7574,
0x82b51eab8c27c66e26c858a079bcdf4f1ada34cec420cafc7eac1a42216fb6c4)
u = z * pow(s, N-2, N) % N
v = r * pow(s, N-2, N) % N
print((u*G + v*point).x.num == r)
# ### Exercise 1
# Which sigs are valid?
#
# ```
# P = (887387e452b8eacc4acfde10d9aaf7f6d9a0f975aabb10d006e4da568744d06c,
# 61de6d95231cd89026e286df3b6ae4a894a3378e393e93a0f45b666329a0ae34)
# z, r, s = ec208baa0fc1c19f708a9ca96fdeff3ac3f230bb4a7ba4aede4942ad003c0f60,
# ac8d1c87e51d0d441be8b3dd5b05c8795b48875dffe00b7ffcfac23010d3a395,
# 68342ceff8935ededd102dd876ffd6ba72d6a427a3edb13d26eb0781cb423c4
# z, r, s = 7c076ff316692a3d7eb3c3bb0f8b1488cf72e1afcd929e29307032997a838a3d,
# eff69ef2b1bd93a66ed5219add4fb51e11a840f404876325a1e8ffe0529a2c,
# c7207fee197d27c618aea621406f6bf5ef6fca38681d82b2f06fddbdce6feab6
# ```
#
# +
# Exercise 1
from ecc import S256Point, G, N
px = 0x887387e452b8eacc4acfde10d9aaf7f6d9a0f975aabb10d006e4da568744d06c
py = 0x61de6d95231cd89026e286df3b6ae4a894a3378e393e93a0f45b666329a0ae34
signatures = (
# (z, r, s)
(0xec208baa0fc1c19f708a9ca96fdeff3ac3f230bb4a7ba4aede4942ad003c0f60,
0xac8d1c87e51d0d441be8b3dd5b05c8795b48875dffe00b7ffcfac23010d3a395,
0x68342ceff8935ededd102dd876ffd6ba72d6a427a3edb13d26eb0781cb423c4),
(0x7c076ff316692a3d7eb3c3bb0f8b1488cf72e1afcd929e29307032997a838a3d,
0xeff69ef2b1bd93a66ed5219add4fb51e11a840f404876325a1e8ffe0529a2c,
0xc7207fee197d27c618aea621406f6bf5ef6fca38681d82b2f06fddbdce6feab6),
)
# initialize the public point
# use: S256Point(x-coordinate, y-coordinate)
point = S256Point(px, py)
# iterate over signatures
for z, r, s in signatures:
# u = z / s, v = r / s
u = z * pow(s, N-2, N) % N
v = r * pow(s, N-2, N) % N
# finally, uG+vP should have the x-coordinate equal to r
print((u*G+v*point).x.num == r)
# -
# ### Exercise 2
#
#
#
#
# #### Make [this test](/edit/session3/ecc.py) pass: `ecc.py:S256Test:test_verify`
# +
# Exercise 2
reload(ecc)
run(ecc.S256Test('test_verify'))
# -
# ### Exercise 3
#
#
#
#
# #### Make [this test](/edit/session3/ecc.py) pass: `ecc.py:PrivateKeyTest:test_sign`
# +
# Exercise 3
reload(ecc)
run(ecc.PrivateKeyTest('test_sign'))
# -
# ### Exercise 4
# Verify the DER signature for the hash of "ECDSA is awesome!" for the given SEC pubkey
#
# `z = int.from_bytes(hash256('ECDSA is awesome!'), 'big')`
#
# Public Key in SEC Format:
# 0204519fac3d910ca7e7138f7013706f619fa8f033e6ec6e09370ea38cee6a7574
#
# Signature in DER Format: 304402201f62993ee03fca342fcb45929993fa6ee885e00ddad8de154f268d98f083991402201e1ca12ad140c04e0e022c38f7ce31da426b8009d02832f0b44f39a6b178b7a1
#
# +
# Exercise 4
from ecc import S256Point, Signature
from helper import hash256
der = bytes.fromhex('304402201f62993ee03fca342fcb45929993fa6ee885e00ddad8de154f268d98f083991402201e1ca12ad140c04e0e022c38f7ce31da426b8009d02832f0b44f39a6b178b7a1')
sec = bytes.fromhex('0204519fac3d910ca7e7138f7013706f619fa8f033e6ec6e09370ea38cee6a7574')
# message is the hash256 of the message "ECDSA is awesome!"
z = int.from_bytes(hash256(b'ECDSA is awesome!'), 'big')
# parse the der format to get the signature
sig = Signature.parse(der)
# parse the sec format to get the public key
point = S256Point.parse(sec)
# use the verify method on S256Point to validate the signature
print(point.verify(z, sig))
# -
# ### Exercise 5
#
#
#
#
# #### Make [this test](/edit/session3/tx.py) pass: `tx.py:TxTest:test_parse_version`
# +
# Exercise 5
reload(tx)
run(tx.TxTest('test_parse_version'))
# -
# ### Exercise 6
#
#
#
#
# #### Make [this test](/edit/session3/tx.py) pass: `tx.py:TxTest:test_parse_inputs`
# +
# Exercise 6
reload(tx)
run(tx.TxTest('test_parse_inputs'))
# -
# ### Exercise 7
#
#
#
#
# #### Make [this test](/edit/session3/tx.py) pass: `tx.py:TxTest:test_parse_outputs`
# +
# Exercise 7
reload(tx)
run(tx.TxTest('test_parse_outputs'))
# -
# ### Exercise 8
#
#
#
#
# #### Make [this test](/edit/session3/tx.py) pass: `tx.py:TxTest:test_parse_locktime`
# +
# Exercise 8
reload(tx)
run(tx.TxTest('test_parse_locktime'))
# -
# ### Exercise 9
# What is the scriptSig from the second input in this tx? What is the scriptPubKey and amount of the first output in this tx? What is the amount for the second output?
#
# ```
# 010000000456919960ac691763688d3d3bcea9ad6ecaf875df5339e148a1fc61c6ed7a069e010000006a47304402204585bcdef85e6b1c6af5c2669d4830ff86e42dd205c0e089bc2a821657e951c002201024a10366077f87d6bce1f7100ad8cfa8a064b39d4e8fe4ea13a7b71aa8180f012102f0da57e85eec2934a82a585ea337ce2f4998b50ae699dd79f5880e253dafafb7feffffffeb8f51f4038dc17e6313cf831d4f02281c2a468bde0fafd37f1bf882729e7fd3000000006a47304402207899531a52d59a6de200179928ca900254a36b8dff8bb75f5f5d71b1cdc26125022008b422690b8461cb52c3cc30330b23d574351872b7c361e9aae3649071c1a7160121035d5c93d9ac96881f19ba1f686f15f009ded7c62efe85a872e6a19b43c15a2937feffffff567bf40595119d1bb8a3037c356efd56170b64cbcc160fb028fa10704b45d775000000006a47304402204c7c7818424c7f7911da6cddc59655a70af1cb5eaf17c69dadbfc74ffa0b662f02207599e08bc8023693ad4e9527dc42c34210f7a7d1d1ddfc8492b654a11e7620a0012102158b46fbdff65d0172b7989aec8850aa0dae49abfb84c81ae6e5b251a58ace5cfeffffffd63a5e6c16e620f86f375925b21cabaf736c779f88fd04dcad51d26690f7f345010000006a47304402200633ea0d3314bea0d95b3cd8dadb2ef79ea8331ffe1e61f762c0f6daea0fabde022029f23b3e9c30f080446150b23852028751635dcee2be669c2a1686a4b5edf304012103ffd6f4a67e94aba353a00882e563ff2722eb4cff0ad6006e86ee20dfe7520d55feffffff0251430f00000000001976a914ab0c0b2e98b1ab6dbf67d4750b0a56244948a87988ac005a6202000000001976a9143c82d7df364eb6c75be8c80df2b3eda8db57397088ac46430600
# ```
#
# +
# Exercise 9
from io import BytesIO
from tx import Tx
hex_transaction = '010000000456919960ac691763688d3d3bcea9ad6ecaf875df5339e148a1fc61c6ed7a069e010000006a47304402204585bcdef85e6b1c6af5c2669d4830ff86e42dd205c0e089bc2a821657e951c002201024a10366077f87d6bce1f7100ad8cfa8a064b39d4e8fe4ea13a7b71aa8180f012102f0da57e85eec2934a82a585ea337ce2f4998b50ae699dd79f5880e253dafafb7feffffffeb8f51f4038dc17e6313cf831d4f02281c2a468bde0fafd37f1bf882729e7fd3000000006a47304402207899531a52d59a6de200179928ca900254a36b8dff8bb75f5f5d71b1cdc26125022008b422690b8461cb52c3cc30330b23d574351872b7c361e9aae3649071c1a7160121035d5c93d9ac96881f19ba1f686f15f009ded7c62efe85a872e6a19b43c15a2937feffffff567bf40595119d1bb8a3037c356efd56170b64cbcc160fb028fa10704b45d775000000006a47304402204c7c7818424c7f7911da6cddc59655a70af1cb5eaf17c69dadbfc74ffa0b662f02207599e08bc8023693ad4e9527dc42c34210f7a7d1d1ddfc8492b654a11e7620a0012102158b46fbdff65d0172b7989aec8850aa0dae49abfb84c81ae6e5b251a58ace5cfeffffffd63a5e6c16e620f86f375925b21cabaf736c779f88fd04dcad51d26690f7f345010000006a47304402200633ea0d3314bea0d95b3cd8dadb2ef79ea8331ffe1e61f762c0f6daea0fabde022029f23b3e9c30f080446150b23852028751635dcee2be669c2a1686a4b5edf304012103ffd6f4a67e94aba353a00882e563ff2722eb4cff0ad6006e86ee20dfe7520d55feffffff0251430f00000000001976a914ab0c0b2e98b1ab6dbf67d4750b0a56244948a87988ac005a6202000000001976a9143c82d7df364eb6c75be8c80df2b3eda8db57397088ac46430600'
# bytes.fromhex to get the binary representation
bin_transaction = bytes.fromhex(hex_transaction)
# create a stream using BytesIO()
stream = BytesIO(bin_transaction)
# Tx.parse() the stream
tx_obj = Tx.parse(stream)
# print tx's second input's scriptSig
print(tx_obj.tx_ins[1].script_sig)
# print tx's first output's scriptPubKey
print(tx_obj.tx_outs[0].script_pubkey)
# print tx's second output's amount
print(tx_obj.tx_outs[1].amount)
| session3/complete/session3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Heat Map
# +
# 导入beakerx库及其他所需库文件
from beakerx import *
from beakerx.object import beakerx
import numpy as np
import random
# +
data = np.random.randint(0,10,30).reshape(6,5)
print(data)
HeatMap(
title = "BeakerX/Heat Map Demo 热力图演示 #1",
xLabel = "X轴",
yLabel = "Y轴",
data = data,
legendPosition = LegendPosition(position = LegendPosition.Position.BOTTOM),
# color = GradientColor.WHITE_BLUE, # BROWN_RED_YELLOW / GREEN_YELLOW_WHITE / WHITE_BLUE
color = GradientColor([Color(255,255,0), Color(255,32,0)]),
initWidth = 540,
initHeight = 360
)
# -
# # Category Plot
# +
data = np.random.randint(1,99,12).reshape(3, 4)
print(data)
cp1 = CategoryPlot(
title = "BeakerX/Category Plot Demo 分类图形演示 #1",
xLabel = "X轴",
yLabel = "Y轴",
categoryNames = ['系列%s' % x for x in range(1, len(data[0])+1)],
categoryMargin= 0.5,
orientation= PlotOrientationType.VERTICAL # PlotOrientationType.HORIZONTAL
)
# 加入分类柱状图
cp1.add(
CategoryBars(
value = data,
# base = [(-1+x)*10 for x in range(len(data))],
seriesNames= ['%s号货品' % x for x in range(1, len(data)+1)],
showItemLabel = True,
labelPosition= LabelPositionType.VALUE_OUTSIDE,
outlineColor= [Color.black, Color.red],
# centerSeries= False,
width = 0.5
)
)
# +
data = np.random.randint(1,99,8).reshape(2, 4)
base = [(-1+x)*10 for x in range(len(data))]
print(data)
print(base)
cp2 = CategoryPlot(
title = "BeakerX/Category Plot Demo 分类图形演示 #2",
xLabel = "X轴",
yLabel = "支出范围"
)
cp2.add(YAxis(label= "第二Y轴", upperMargin= 1))
# 加入杆图
cp2.add(
CategoryStems(
value = data,
base = base,
color = [Color(192,192,255), Color(255,192,192)],
seriesNames = ['支出范围%s' % x for x in range(1, len(data)+1)],
showItemLabel = True,
width = 30
)
)
# 加入线段(对应第二Y轴)
cp2.add(
CategoryLines(
value = data,
yAxis = "第二Y轴",
style = [StrokeType.DASH, StrokeType.DOT],
seriesNames = ['第二Y轴曲线%s' % x for x in range(1, len(data)+1)],
width = 3
)
)
# 加入数据点(对应第二Y轴)
cp2.add(
CategoryPoints(
value = data,
yAxis = "第二Y轴",
seriesNames = ['第二Y轴数据点%s' % x for x in range(1, len(data)+1)],
size = 15
)
)
# +
data = np.random.randint(1,99,12).reshape(3,4)
base = [(-1+x)*20 for x in range(len(data))]
print(data)
print(base)
cp3 = CategoryPlot(
title = "BeakerX/Category Plot Demo 分类图形演示 #2",
xLabel = "X轴",
yLabel = "支出范围"
)
cp3.add(
CategoryArea(
value = data,
base = base,
color = [Color(192,192,0), Color(0,192,192,192), Color(255,128,128,192)]
)
)
for i in range(len(base)):
cp3.add(
ConstantLine(
y = base[i],
style = [StrokeType.DOT, StrokeType.DASH, StrokeType.DASHDOT][i%3] ,
color = Color.gray,
showLabel = True
)
)
cp3
| beakerx_samples/beakerx_python_heatmap&categoryplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# name: python3
# ---
# ## 1.0.2. String Manipulation
# ## Learning Objectives
#
#
# * [Immutability](#Immutability)
# * [Slicing](#Slicing)
# * [Operators and methods](#Operators)
# * [Iteration](#Iteration)
#
# <a id='Immutability'></a>
# ## Immutability
# Let's remember!
# * Strings are a **sequence** of case sensitive characters
# * They are a type of object.
# * We compare strings with ==, >, < etc.
#
# We can create strings with the use of single, double or triple quotes! They all provide the same functionality, except the triple quotes enable us to write multi-line strings.
# remember a string is in quotation marks !
# Using single quotations can be problematic, here, the apostrophe ends the string early
print('What's the problem here?')
# we can use double quotations
print("What's the problem here?")
# or backslash if we have single and double quotes
print("What\'s the \"problem\" here?")
# Triple quotes for multi-line strings
print('''What's the
problem here? ''')
# Before we move one, let's look at **indexing**:
# * Since strings are a sequence of characters, we might want to get what character is at a certain position. We're going to tell Python, I want to know the character, at this certain position or at this certain index, inside my sting.
# * Square brackets used to perform **indexing** into a string to get the value at a certain index/position
#
# <img src="images/index.png" style="display:block; margin-left:auto; margin-right:auto; width:50%"/>
# +
s = "abc"
print(s[0]) # evaluates to "a"
print(s[1]) # evaluates to "b"
print(s[2]) # evaluates to "c"
# -
print(s[-1]) # evaluates to "c"
print(s[-2]) # evaluates to "b"
print(s[-3]) # evaluates to "a"
# Now we get to the idea of **immutability**. String objects are **immutable**, whichs means that a value (character) from a string cannot be updated. We can verify this by trying to update a part of the string which will led us to an error.
# Can not reassign
example = "Hello world!"
print(type(example))
example[0] = "M"
# +
# One possible solution is to create a new string object with necessary modifications:
example_1 = "Hello world!"
example_2 = "M"+ example_1[1:] # this an example of slicing, which we are about to see
print("example_1 = ", example_1, "\nexample_2 = ", example_2)
# -
s = "hello"
s[0] = 'y' # gives an error
s = 'y'+s[1:len(s)] # is allowed, s bound to new object
print(s)
# <br>
# <img src="images/bound.png" style="display:block; margin-left:auto; margin-right:auto; width:50%"/> <br>
#
# _Source on page 7:_ ([Click here](https://ocw.mit.edu/courses/electrical-engineering-and-computer-science/6-0001-introduction-to-computer-science-and-programming-in-python-fall-2016/lecture-slides-code/MIT6_0001F16_Lec3.pdf))<br>
# <a id='Slicing'></a>
# ## Slicing
# Sometimes, we might want to get a **substring**. So we want to:
# * start at the first character and go halfway into the string or
# * take a few characters in between or
# * skip every other letter or something like that in our string.
#
# In order to do these more complicated operation, we use **slicing**. In our daily life, let's think of a birthday cake. You want only a slice of it and basically get a slice.
# <br>
# <img src="images/cake.jpg" style="display:block; margin-left:auto; margin-right:auto; width:40%"/> <br>
# _Source:_ ([Click here](https://www.superhealthykids.com/wp-content/uploads/2020/05/VanillaStrawberryCake2-745x496.jpg))<br>
# * We can slice strings using: **[start:stop:step]**
# * If give two numbers: **[start:stop]**, step=1 by default
# * You can also omit numbers and leave just colons
#
# This notation here should seem a little bit familiar, because we saw it last lecture when we did it with range.
# We had a start, stop, and a step. The notation was a little bit different, because, in range, we had open-close parentheses and commas in between. But except for that, this sort of works the same.
#
# <img src="images/substr.png" style="display: block; margin-left:auto; margin-right:auto; width:40%"/>
#
# range(start, stop, step)
x = range(1, 6)
for n in x:
print(n)
# +
s = "abcdefgh"
print(s[3:6]) # evaluates to "def", same as s[3:6:1]
print(s[3:6:1]) # evaluates to "def", same as above
print(s[3:6:2]) # evaluates to "df"
print(s[::]) # evaluates to "abcdefgh", same as s[0:len(s):1]
print(s[:]) # evaluates to "abcdefgh", same as s[0:len(s)] step=1 by default
print(s[::-1]) # evaluates to "hgfedbca", same as s[-1:-(len(s)+1):-1]
print(s[4:1:-2]) # evaluates to "ec"
x = range(1)
print(x)
# -
# **Your Turn!**
#
# What does the code below print? Guess and run the cell to see the answer!
s = "6.00 is 6.0001 and 6.0002"
new_str = ""
new_str += s[-1]
new_str += s[0]
new_str += s[4::30]
new_str += s[13:10:-1]
print(new_str)
# <a id='Operators'></a>
# ## Operators and methods
# We can do more than just concatenate two strings together or do these little tests on them. Today we're going to start introducing the idea of a **function** or a procedure. We'll focus on function next lecture but for now, you can think of a **function** as sort of a procedure that does something for you.
#
# We'll look at some functions in this notebook and you can find the rest at:
# https://docs.python.org/2/library/stdtypes.html#string-methods
#
# <br>
# <img src="images/function.png" style="display:block; margin-left:auto; margin-right:auto; width:30%"/> <br>
#
# [Source](https://medium.com/@kuanzhasulan/python-functions-352137dd4d84)<br>
#
# As seen above, a function basically takes in an input and gives an output. It does some calculations or operations with the input and we get an output in return. Let's think about a snack machine. Here are the arguments according to the diagram shown above:
# * **Input:** putting money, pushing the button
# * **Function:** the machine gets what you wanted and a specific item drops into the output slot
# * **Output:** a specific item
# <br>
# <img src="images/snack.jpg" style="display:block; margin-left:auto; margin-right:auto; width:20%"/> <br>
#
# [Source for the machine](https://www.candymachines.com/Seaga-Infinity-INF5S-Snack-Vending-Machine-P2641.aspx)<br>
#
#
# ### len()
# * len() is a function used to retrieve the **length** of the string in the parentheses. That's going to tell you how many characters are in the string. These characters are going to be letters, digits, special characters, spaces, and so on. So it's just going to count how many characters are in a string.
s = "abc" # define a string variable called 's'
len(s) # evaluates to 3
# ### upper()
# .upper() method makes all caps (not inplace)
print(s.upper())
print(s)
# +
# if we want to change x itself, we must reassign it
print(s)
s = s.upper()
print(s)
# -
# ### lower()
# .lower() method makes all lowercase (not inplace)
print(s.lower())
# ### split()
# The split method breaks up a string at the specified __separator__ and returns a list of strings. A separator is just a commonly occuring value in your object. If your string was a piece of paper, the separators are the dotted lines where you should cut across. When dealing with strings, the recurring characters are referred to as __delimiters__. If no argument is given, it assumes a whitespace, ' ', to be the delimiter. This will __split a sentence into a list of words__. You can set other delimiters if you would like.
# +
x = "Hello World"
# split() method splits on space as default or desired separator
print(x.split())
# we gave a specific seperator "o"
print(x.split("o"))
# +
grocery = 'Milk, Chicken, Bread'
# splits at space as default
print(grocery.split())
# splits at comma+space ', '
print(grocery.split(', '))
# Splitting at ':'
print(grocery.split(':'))
# -
# <img src="images/split.png" align="center"/>
# ### .format
# * The .format method (A method is a function associated with an object) is a way of inserting something into a string.
# * This can be other strings or a variable taken from elsewhere in your code.
# * The syntax used is detailed below.
# * When using .format to add in a float to a string, we can specify the width and precision of the decimal.
# default prints in order
print("The {} {} {}".format("fox", "brown", "quick"))
# can index
print("The {2} {1} {0}".format("fox", "brown", "quick"))
# can use variable keys for readability
print("The {q} {b} {f}".format(f="fox", b="brown", q="quick"))
# +
# create long decimal
result = 100/777
print(result, end = "\n\n")
# use value:width.precisionf for formatting
# width is minimum length of string, padded with whitespace if necessary
# precision is decimal places
print("The result was {:1.3f}".format(result))
print("The result was {r:1.3f}".format(r=result))
print("The result was {r:1.7f}".format(r=result))
print("The result was {r:8.3f}".format(r=result))
# -
# <a id='Iteration'></a>
# ## Iteration
# We can apply for loops, very easily, to write very nice, readable code when dealing with strings.
#
# * for LOOPS RECAP
#
# for loops have a **loop variable** that iterates over a set of values:
# <br><img src="images/for.png" style="display:block; margin-left:auto; margin-right:auto; width:50%"/> <br>
#
# * **range** is a way to iterate over numbers, but a for loop variable can **iterate over any set of values**, not just numbers!
#
# These two code snippets below do the same thing but the bottom one is more __"pythonic”__:
# +
s = "abcdefghi"
for index in range(len(s)):
if s[index] == 'i' or s[index] == 'u':
print("There is an i or u")
for char in s:
if char == 'i' or char == 'u':
print("There is an i or u")
# -
# ### Note: Brief description of iterables in Python
# <br> The output of the range() function and strings are referred to as __iterables__. An iterable is any Python object capable of returning its members one at a time, permitting it to be iterated over in a loop. Most of the objects in Python are iterable. All sequences like strings, lists, tuples and dictionaries are iterable. Strings have some characters as its members and we can iterate its members in a loop. [Click here for more information about iterables](https://www.pythonlikeyoumeanit.com/Module2_EssentialsOfPython/Iterables.html#:~:text=Definition%3A,over%20in%20a%20for%2Dloop)
# +
string = 'London'
for member in string:
print(member)
# -
# <br>Are you confused? Now, let's give an example from daily life. Suppose, a group of 5 boys are standing in a line. You're pointing at the first boy and ask him about his name. Then, he replied. After that, you ask the next boy and so on. The below picture will illustrate the thing.
# <br>
# <img src="images/iterator.jpg" style="display:block; margin-left:auto; margin-right:auto; width:50%"/> <br>
#
# ([Source](https://medium.com/@kuanzhasulan/python-functions-352137dd4d84))<br>
# In this case, you are the **Iterator**!!!! Obviously, the group of boys is the **iterable** element.
#
# ---
# #### CODE EXAMPLE: ROBOT CHEERLEADERS
#
# <br>
# <img src="images/robotcheer.gif" style="display:block; margin-left:auto; margin-right:auto; width:50%"/><br>
#
# _Source:_ ([Click here](https://www.programmersought.com/images/613/1e4c176c84dfdfc4a5eaa207fda3a05d.gif))<br>
# +
an_letters = "aefhilmnorsxAEFHILMNORSX"
word = input("I will cheer for you! Enter a word: ")
times = int(input("Enthusiasm level (1-10): "))
i = 0
while i < len(word):
char = word[i]
if char in an_letters:
print("Give me an " + char + "! " + char)
else:
print("Give me a " + char + "! " + char)
i += 1
print("What does that spell?")
for i in range(times):
print(word, "!!!")
# -
# <br>
# <img src="images/robot.png" style="display:block; margin-left:auto; margin-right:auto; width:50%"/>
#
# _Source on page 10:_ ([Click here](https://ocw.mit.edu/courses/electrical-engineering-and-computer-science/6-0001-introduction-to-computer-science-and-programming-in-python-fall-2016/lecture-slides-code/MIT6_0001F16_Lec3.pdf))<br>
#
# **Your Turn!**
#
# Can you change the code example "ROBOT CHEERLEADERS" according to the image above? Simply add for loop!
# +
## Your code is here!
# -
# **Your Turn!**
#
# How many times will the code below print "common letter"? Guess and run the cell to see the answer!
s1 = "this make sense"
s2 = "i get this now!"
if len(s1) == len(s2):
for char1 in s1:
for char2 in s2:
if char1 == char2:
print("common letter")
break
# ### Summary
#
# * Strings are immutable – cannot be modified
# * We can slice strings using [start:stop:step]
# * We can do many operations with strings such as len(), indexing, upper(), lower(), .format etc.
# * We can apply for loops, very easily, to write very nice, readable code when dealing with strings.
# # Challenges
#
# ### Question 1:
# Write a Python program that:
#
# - Defines the first string with variable name string1 with an arbitrary string value
# - Defines the second string with variable name string2 with an arbitrary string value
# - Swaps the first two characters of string1 and string2
# - Joins the two edited strings with a space in between and print the result
#
# __Example__:
# - string1 = 'abc'
# - string2 = 'xyz'
# - Expected Result : 'xyc abz'
# ### Question 2:
# Write a Python program that:
#
# - Defines a variable named string with an arbitrary string value
# - Defines a variable named upper that is the same as 'string', but uppercase
# - Defines a variable named lower that is the same as 'string', but lowercase
# - Prints upper to console
# - Prints lower to console
#
# __Example:__
# - string = 'RestaRt'
# - Expected Result: 'RESTART'
# 'restart'
# ### Question 3:
# Write a Python program to get an input string from a given string where all occurrences of its first char have been changed to '$', __except the first char itself__.
#
# Write a Python program that:
#
# - Defines a variable named string with an arbitrary string value
# - Define another variable named output_string that is the same as the 'string' variable, but will all occurences of its first character (except) the first character itself are changed to '$'
# - Prints output_string to console
#
# __Example__:
#
# - string ='restart'
# - Expected Result : 'resta$t'
#
#
#
# Attention: We didn't change the first character 'r' here. We only changed the next one.
#
# __Example__:
#
# - string = 'pencil'
# - Expected Result : 'pencil'
#
# Attention: We didn't change the string. Because the first char is 'p' and we only have one 'p' in our input. We don't have any next one. So we couldn't change any char in 'pencil'.
| English/3. String Manipulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
# +
df = pd.read_csv('BlackFriday.csv')
del df['Product_Category_2']
del df['Product_Category_3']
# -
df.head()
sns.countplot(x = df['Stay_In_Current_City_Years'])
df.groupby('Stay_In_Current_City_Years').size().plot(kind = 'pie', autopct = "%.1f")
sns.countplot(x = 'Stay_In_Current_City_Years', hue = 'Gender', data = df)
sns.countplot(x = 'Stay_In_Current_City_Years', hue = 'Marital_Status', data = df)
sns.countplot(x = 'Stay_In_Current_City_Years', hue = 'City_Category', data = df)
sns.countplot(x = 'City_Category', hue = 'Age',data = df)
df.groupby('Stay_In_Current_City_Years').sum()['Purchase'].plot(kind = 'bar')
df.groupby('Stay_In_Current_City_Years').mean()['Purchase'].plot(kind = 'bar')
sns.countplot(x = df['Occupation'])
df.groupby('Occupation').size().sort_values().plot(kind = 'bar')
df.groupby('Occupation').sum()['Purchase'].sort_values().plot(kind = 'bar')
df.groupby('Occupation').mean()['Purchase'].sort_values().plot(kind = 'bar')
sns.countplot(x = 'Occupation', hue = 'Marital_Status', data = df)
sns.countplot(x = 'Occupation', hue = 'Gender', data = df)
df.groupby('Occupation').nunique()['Product_ID'].plot(kind = 'bar')
df.groupby('Occupation').nunique()['Product_ID'].sort_values().plot(kind = 'bar')
df.groupby('Product_Category_1').size().plot(kind = 'bar')
df.groupby('Product_Category_1').size().sort_values().plot(kind = 'bar')
df.groupby('Product_Category_1').sum()['Purchase'].sort_values().plot(kind = 'bar')
df.groupby('Product_Category_1').mean()['Purchase'].sort_values().plot(kind = 'bar')
df.groupby('Product_ID').sum()['Purchase'].nlargest(10).sort_values().plot(kind = 'bar')
df.groupby('Product_ID').size().nlargest(10).sort_values().plot(kind = 'bar')
df.groupby('Product_ID').mean()['Purchase'].nlargest(10).sort_values().plot(kind = 'bar')
sns.countplot(x = 'Product_Category_1', hue = 'Gender', data = df)
sns.countplot(x = 'Product_Category_1', hue = 'Marital_Status', data = df)
| 6. Occupation and Products Analysis/Black Friday - Occupation and Products Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cryptographie - principes de base
# ## 0- Principes
# Le but de la cryptographie est de cacher un texte donné de tel manière que seul son destinataire légitime puisse le lire.
# Les manières de cacher ce texte sont multiples. On s'intéresse ici à une transformation du texte pour le rendre illisible.
# En pratique pour nous, le texte à cacher est une chaîne de caractères s. On cherche donc à calculer une chaîne de caractère tirée de s de telle manière que notre destinataire (avec qui on peut éventuellement partager un secret, la <b>clé</b>), et avec un peu de chance seul lui, puisse déchiffrer ce texte.
# ### Cadre
# On se limite à des textes contenant uniquement des caractères en minuscule et des espaces, pour simplifier les raisonnements. Dans le texte de référence utilisé, tous les autres caractères seront remplacés par des espaces.
# ### Texte à cacher
# On fixe pour tout le cours un texte à crypter : les histoires extraordinaires d'<NAME>, traduites par <NAME> et disponible librement sur le site du projet Gutenberg ici : http://www.gutenberg.org/ebooks/20761
# Le fichier texte, légèrement nettoyé (majuscules enlevé, préface enlevée) est dans le dossier contenant ce notebook et peut donc être ouvert directement par Python :
with open('HistoiresExtraordinaires.txt') as f:
texte = f.read()
texte = texte.replace('\r', '')
print(texte[:2500]) #on regarde si le texte a bien été importé en affichant le début
# ## 1- <NAME>
# ### 1.1 - Description
# Le principe est assez simple :
# <ul>
# <li>on donne un numéro à chaque lettre de l'alphabet</li>
# <li>la clé de chiffrage est un nombre entre 1 et le nombre de lettre - 1</li>
# <li>la chaîne cryptée est obtenue en décalant tous les caractères de clé dans l'alphabet, étant entendu que l'on revient au début à la fin de la liste...</li>
# </ul>
# +
LETTRES = 'abcdefghijklmnopqrstuvwxyz '
def indice(lettre):
"""
Retourne l'indice de la lettre donnée dans la chaîne LETTRES
indice('a') retourne 0, indice('z') retourne 25
indice(' ') retourne 26, indice(n'importe quoi d'autre) retourne 26
"""
if 97<=ord(lettre)<=122:
return ord(lettre)-ord("a")
else:
return 26
# -
def cesar(chaine, cle):
#s est notre chaine codée qu'on construit au fur et à mesure
s = ''
n = len(LETTRES)
for i in range(len(chaine)):
#on décale la lettre chaine[i] de clé. On fait % n pour revenir à 0 si on dépasse 26
s = s + LETTRES[(indice(chaine[i]) + cle) % n]
return s
cesar(texte, 14)[:500]
# ### 1.2 - Retrouver le texte d'origine
# Le principe pour notre correspondant est assez simple, il suffit de décaler dans l'autre sens.
#on crypte
messageCode=cesar('hello world',3)
print(messageCode)
#on décrypte
messageDecode=cesar(messageCode,-3)
print(messageDecode)
#modulo 27, faire +14 ou -13 revient au même
cesar(cesar('hello world', 13), 14)
#bien sûr, avec notre texte qui contient des caractères autres que des minuscules
#on perd un peu d'information qui se transforme en espace
cesar(cesar(texte, 4), -4)[:500]
# ### 1.3 - Cryptanalyse
# Il est très facile de retrouver la clé secrète du chiffrement de césar si on connait l'alphabet utilisé.
# Ainsi un espion, le gouvernement, ou toute autre personne satisfaisant votre sens de la théorie du complot, peut retrouver le message en clair.
# La méthode est simple : force brute. On essaie toutes les clés possible et on regarde à la main les messages obtenus. il y a tellement peu de clés possible que la cryptanalyse est quasiment immédiate.
# +
import random
#on choisit une clé aléatoire qu'on n'affiche pas pour la démonstration
cle_choisie = random.randint(1, len(LETTRES) - 1)
message = 'ceci est un message un tout petit peu plus long'
crypte = cesar(message, cle_choisie)
for i in range(1, len(LETTRES)):
print('clé : ', i, ' message décodé : ' + cesar(crypte, -i))
# -
# Pas trop difficile de deviner ce qu'avait choisi python comme clé...
# ## 2 - Chiffrement par permutation
# ### 2.1 - Principe
# Au lieu de bêtement décaler chaque lettre, on choisit une bijection de l'alphabet dans lui même, c'est à dire que chaque lettre est transformée en une autre.
# En pratique une bijection est représentée par une chaîne de caractère ou une liste possédant les même lettres que LETTRES mais dans le désordre
def permute(chaine, bijection):
s = ''
n = len(LETTRES)
for c in chaine:
s = s + bijection[indice(c)]
return s
#ou on peut faire une permutation très simple en inversant simplement les "o" et les "e"
bij1='abcdofghijklmnepqrstuvwxyz '
print(permute('hello world', bij1))
#ou une permutation plus complexe car complètement aléatoire
bij = list(LETTRES)
random.shuffle(bij)
print(permute('hello world', bij))
# ### 2.2 - Déchiffrement
# Comment retrouver le message d'origine connaissant le message chiffré et la permutation ? Facile, il suffit de calculer la réciproque, c'est à dire trouver la position du caractère courant dans bijection.
def decode_permute(chiffre, bijection):
s = ''
n = len(LETTRES)
for c in chiffre:
s = s + LETTRES[bijection.index(c)]
return s
#on crypte
messageCode=permute('hello world', bij)
print(messageCode)
#on décrypte
messageDecode=decode_permute(messageCode,bij)
print(messageDecode)
# Cette fois le nombre de clé est plutôt vaste : le "a" peut se transformer en 26 lettres différentes, le "b" en 25 (puisque ça ne peut pas être la même que "a"), le "c" en 24, etc. Cela fait donc $26 \times 25 \times 24 \times ... \times 2 \times 1 = 26! \approx 3 \times 10^{29}$ possibilités de clés. Si on pouvait lire 1 000 000 de textes à la seconde (pour vérifier si c'est le bon), il faudrait quand même un million de milliard d'années pour arriver au bout...
#
# On ne peut donc pas attaquer ce genre de code par de la force brute.
# ### 2.3 - Cryptanalyse
# Idée : la probabilité d'apparition des lettres dans un texte en français n'est pas du tout uniforme. On doit pouvoir identifier les voyelles et les consones courantes assez rapidement. On peut aussi utiliser le fait que certaines lettres sont souvent doublées en français, comme le l ou le t ou le p ou encore que de nombreux mots se terminent par s.
def frequence(chaine):
"""
Calcule le nombre d'occurences de chaque lettre dans la chaine donnée.
"""
#on crée un liste de compteurs initialisés à 0
occ = [ 0 for c in LETTRES]
#on incrémente le compteur de la lettre rencontrée dans la chaine
for c in chaine:
i = indice(c)
occ[i] = occ[i] + 1
#on modifie les compteurs en transformant les effectifs en fréquences
total = max(len(chaine), 1)
for i in range(len(LETTRES)):
occ[i] = 100 * occ[i] / total
#on affiche les résultats triés par ordre de fréquence en affichant la fréquence et la lettre
resultat = [(occ[i], LETTRES[i]) for i in range(len(LETTRES))]
resultat.sort()
return resultat
#on suppose que le texte est codée par une bijection secrète IL FAUT FAIRE COMME SI ON NE CONNAISSAIT PAS CETTE BIJECTION!!!
bij=['u', 'm', 'k', 'i', 'q', 'y', 'l', 'c', 'p', 'a', 'j', ' ', 't', 'n', 'x', 'f', 'r', 'g', 'v', 'o', 'e', 's', 'h', 'w', 'z', 'd', 'b']
messageCode = permute(texte, bij)
frequence(messageCode)
# Il suffit maintenant d'aller sur [wikipedia](https://fr.wikipedia.org/wiki/Fr%C3%A9quence_d%27apparition_des_lettres_en_fran%C3%A7ais#Fr%C3%A9quence_des_caract%C3%A8res_dans_le_corpus_de_Wikip%C3%A9dia_en_fran%C3%A7ais) ou tout autre site pour trouver une table référence des fréquence de lettres dans les textes en français.
# On peut deviner plusieurs choses :
# <ul>
# <li>l'espace correspond au 21,14%</li>
# <li>le e doit correspondre à la lettre la plus fréquente</li>
# </ul>
# Pour trouver des chiffres plus proches des tables données, il faut considérer des textes sans espaces. Allons-y
messageCode2 = messageCode.replace('b', '')
frequence(messageCode2)
# Les chiffres obtenus sont proches de ceux de la table de référence. Essayons de décrypter.
#
# On va créer une fonction qui permute deux lettres données :
def echange(chaine, c1, c2):
s = ''
for c in chaine:
#si on rencontre c1, on remplace par c2
if c == c1:
s += c2
#si on rencontre c2, on remplace par c2
elif c == c2:
s += c1
#sinon on garde le même caractère
else:
s += c
return s
# +
c2 = echange(messageCode, 'b', ' ');
c2 = echange(c2, 'q', 'e')
c2 = echange(c2, 'v', 'a')
c2 = echange(c2, 'u', 'i')
c2 = echange(c2, 'o', 's')
c2 = echange(c2, 'n', 'n')
c2 = echange(c2, 'p', 'r')
c2 = echange(c2, 'g', 't')
c2[:5000]
# -
# Beaucoup de mots se terminent par a, ce sont sûrement des s. On voit aussi le 2eme mot avec un double b, on parie sur un double l à la place :
# +
c2 = echange(c2, 'a', 's')
c2 = echange(c2, 'b', 'l')
c2[0:5000]
# -
# Prochaine étape, utiliser les espaces de manière plus systématiques, pour réutiliser l'idée précédente. Par exemple, trouver les mots de longueur 1 et 2.
# +
mots = c2.split()
long_1 = [m for m in mots if len(m)== 1]
mots_1 = set() #ensemble : ne peut pas contenir deux fois le même élément
for m in long_1:
mots_1.add((m, long_1.count(m)))
print(mots_1)
long_2 = [m for m in mots if len(m) == 2]
mots_2 = set() #ensemble : ne peut pas contenir deux fois le même élément
for m in long_2:
mots_2.add((m, long_2.count(m)))
print(mots_2)
# -
# Le but est de trouver a, l, d qui sont très commun en mots d'une lettre comme en mots de deux ("l'", "d'", "la", "le"...)
#
# On observe que le mot "ue" apparait 5249 fois, comme on est assez sûr du e et sans doute du l, on peut parier que le u se transforme en d.
#
# De même, on on observe que "le" et "li" apparaissent environ 3000 fois, donc on parie que ce sont les mots le et la. On va donc inverser le i et le a.
c3 = echange(c2, 'u', 'd')
c3 = echange(c2, 'i', 'a')
c3[0:5000]
# On voit "elles sxni" qui ressemblent à "elles sont".
# On va tenter les permutations x<->o et i<->t.
c3 = echange(c3, 'x', 'o')
c3 = echange(c3, "i", "t")
c3[:5000]
c3 = echange(c3, 'z', 'y')
c3 = echange(c3, "i", "r")
c3[:5000]
c3 = echange(c3, 'c', 'h')
c3 = echange(c3, "x", "w")
c3 = echange(c3, "u", "d")
c3[:5000]
# et ainsi de suite...
# La décodage allant, on commence à comprendre le sens et pouvoir déchiffrer plus facilement.
#
# Pour faciliter la lecture, il faudrait afficher différemment les lettres déjà trouvées. On y est presque. Le reste est à finir en exercice.
# ## 3- Vigénère
# ### 3.1 - Principe
# La principale faiblesse du codage par substitution est que tous les "e" deviennnent une seule et même lettre dans le message codé et qu'une analyse statistique permet d'identifier par quoi elle a été substituée.
# Le code de Vigénère apporte une soution à cette faiblesse car il introduit l'idée d'une substitution polyalphabétique, c'est à dire qu'une lettre donnée ne sera pas toujours transformée en la même lettre.
#
# Au lieu d'utiliser un décalage uniforme sur tout le texte comme dans le code de César, on choisit un mot clé qui sert à indiquer de combien on se décale.
# <ul>
# <li>On transforme le mot clé en liste de chiffres, qui seront nos décalages (a->+0, b->+1, c->+2,...), par exemple, si la clé est bac, alors la liste des décalages est de (1,0,2) (a->+0, b->+1, c->+2,...)</li>
# <li>la première lettre du texte est décalée grâce au premier chiffre de la clé</li>
# <li>la deuxième lettre du texte est décalée grâce au deuxième chiffre de la clé</li>
# <li>on poursuit ainsi, en reprennant la clé depuis le début quand on l'a épuisée</li>
# </ul>
#
# La vidéo suivante, à regarder, explique le cryptage par substitution et de Vigénère : https://youtu.be/PIw_nuWsOFU
def vigenere(texte, mot_cle):
liste_cle = [indice(c) for c in mot_cle]
s = ''
long_cle = len(liste_cle)
long_alph = len(LETTRES)
for i in range(len(texte)):
s = s + LETTRES[(indice(texte[i]) + liste_cle[i % long_cle]) % long_alph]
return s
vigenere('ceci est un test', 'motcle')
def dechiffre_vigenere(texte, mot_cle):
liste_cle = [-indice(c) for c in mot_cle]
s = ''
long_cle = len(liste_cle)
long_alph = len(LETTRES)
for i in range(len(texte)):
s = s + LETTRES[(indice(texte[i]) + liste_cle[i % long_cle]) % long_alph]
return s
dechiffre_vigenere('osvkkidgswydeskv', 'motcle')
# ### 3.2 - Cryptanalyse
# Elle est beaucoup plus délicate que la précédente. L'idée est de trouver d'abord la longueur de la clé en procédant à des mesures d'incidences. Ensuite, on peut procéder à une analyse de fréquence pour déterminer chaque décalage.
# La page Bibmath consacrée à la cryptanalyse de Vigénère détaille assez clairement ce procédé : http://www.bibmath.net/crypto/index.php?action=affiche&quoi=poly/viganalyse
#
# On va regarder un exemple pour montrer le procédé :
#on code notre texte avec une clé de 5 caractères. ON SUPPOSE QU'ON NE CONNAIT PAS CETTE CLÉ
cle= "alien"
messageCode=vigenere(texte, cle)
messageCode=messageCode[:5000] #on se restreint à 5000 caractères pour ne pas surcharger le processeur
print(messageCode)
# On commence par lister tous les trigrammes existant dans ce texte et on les met dans un dicionnaire. La valeur associée dans le dictionnaire correspond à l'index de sa position dans le texte.
# +
#on crée le dictionnaire vide
dictTrigrammes={}
#on parcourt le message codé au complet jusqu'à l'avant dernier caractère
for i in range(len(messageCode)-2):
if messageCode[i:i+3] not in dictTrigrammes: #si le trigramme n'a jamais été rencontré
dictTrigrammes[messageCode[i:i+3]]=[i] #on crée une liste des positions de ce trigramme initialisée à [i]
else:
dictTrigrammes[messageCode[i:i+3]].append(i) #si le trigramme a déjà été rencontré, on ajoute sa nouvelle position
print(dictTrigrammes)
# -
# On veut connaitre l'écart qu'il y a entre les trigrammes identiques. On crée donc le dictionnaire dictTrigrammesEcart qui recense l'écart entre chacun des trigrammes.
dictTrigrammesEcart=dict()
for (key,value) in dictTrigrammes.items():
dictTrigrammesEcart[key]=[value[i+1]-value[i] for i in range(len(value)-1)]
print(dictTrigrammesEcart)
# L'idée en regardant ce dictionnaire est de dire qu'un trigramme réapparait quand ce sont les trois même lettres qui ont été codées par les mêmes lettres de la clé. Cela veut donc dire que les écarts trouvés entre les trigrammes sont des multiples de la longueur de la clé.
#
# Ici par exemple, presque tous les écarts semblent être des multiples de 5, ce qui nous amène à poser l'hypothèse que la clé est de longueur 5.
# Tous les 5 caractères, le décalage est donc le même. Il faut donc trouver quel décalage a été fait sur les caractères en position 0,5,10,15... puis de trouver quel décalage a été fait sur les caractères en position 1,6,11,16..., puis ceux en position 2,7,12,17 et ainsi de suite.
# Pour cela on va créer une fonction <code>sousTexte(texte,longueur,pos)</code> (à faire en exercice)
def sousTexte(texte,longueur,pos):
#renvoit les caractères de texte situés aux positions pos, pos+longueur, pos+longueur*2, pos+longueur*3...
#par exemple sousTexte("abcdefghijkl",2,0) renvoit "acegik" (on commence à 0 et on saute de 2 en 2)
#par exemple sousTexte("abcdefghijkl",2,1) renvoit "bdfhjl" (on commence à 1 et on saute de 2 en 2)
#par exemple sousTexte("abcdefghijkl",3,1) renvoit "behk" (on commence à 1 et on saute de 3 en 3)
# On peut désormais analyser les fréquences des sous-textes. L'analyse des fréquences permettra de deviner le décalage.
#
# Par exemple, si mon sous-texte contient une très grande fréquence de "g", alors je peux supposer que la lettre "e" a été décalée en "g" et que le décalage est donc de +2. Le caractère de la clé à cette position est donc c (a->+0, b->+1, c->+2,...)
sousTexte0=sousTexte(messageCode,5,0) #on récupère le sous-texte composé des caractères 0,5,10,15...
print(frequence(sousTexte)) #on observe quelle lettre est la plus fréquente, c'était sûrement un "e" dans le message en clair
# Le reste de la cryptanalyse de Vigénère est à terminer en exercice. Vous allez donc implémenter la fonction <code>sousTexte()</code> et utiliser les analyses de fréquences pour en déduire la clé utilisée.
| cryptographie-principes-de-base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import h5py
dataPath = "data-15.07.08/7.2/000012.SZtick.mat"
f = h5py.File(dataPath,'r+')
# + pycharm={"name": "#%%\n"}
f['r1'].keys()
# + pycharm={"name": "#%%\n"}
import numpy as np
import pandas as pd
keys = ['AccTurover', 'AccVolume', 'AskAvPrice', 'AskPrice', 'AskVolume', 'BSFlag', 'BidAvPrice', 'BidPrice', 'BidVolume', 'Code', 'CurDelta', 'Date', 'Downs', 'High', 'HoldLines', 'Index', 'Interest', 'Low', 'MatchItems', 'Open', 'Position', 'PreClose', 'PrePosition', 'PreSettle', 'Price', 'Settle', 'Stocks', 'Time', 'TotalAskVolume', 'TotalBidVolume', 'TradeFlag', 'Turover', 'Ups', 'Volume', 'WindCode']
data = []
for key in keys:
data.append(f['r1'][key])
# + pycharm={"name": "#%%\n"}
BidPrice = np.array(f['r1']['BidPrice'])
AskPrice = np.array(f['r1']['AskPrice'])
BidVolume = np.array(f['r1']['BidVolume'])
AskVolume = np.array(f['r1']['AskVolume'])
miu_n = BidVolume.astype(np.int) - AskVolume.astype(np.int)
miu_n
# + pycharm={"name": "#%%\n"}
miu_n_tmp = miu_n[0:1,:]
miu_n_final = np.sum(miu_n_tmp,axis=0)
# + pycharm={"name": "#%%\n"}
miu_n_final
# + pycharm={"name": "#%%\n"}
S_n = BidPrice[0] + AskPrice[0]
S_n = S_n / 2
for i in range(S_n.shape[0]):
if i+1 < S_n.shape[0]:
S_n[i] = S_n[i+1] - S_n[i]
indexes = []
for i in range(S_n.shape[0]):
if 1000 > S_n[i] > -1000:
indexes.append(i)
# + pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
fig = plt.figure()
plt.scatter(S_n[indexes],miu_n_final[indexes])
plt.xlabel('S_n')
plt.ylabel('miu_n')
# + pycharm={"name": "#%%\n"}
miu_n_final
# + pycharm={"name": "#%%\n"}
| data_K.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Módulo 2: Scraping con Selenium
# ## LATAM Airlines
# <a href="https://www.latam.com/es_ar/"><img src="https://i.pinimg.com/originals/dd/52/74/dd5274702d1382d696caeb6e0f6980c5.png" width="420"></img></a>
# <br>
#
# Vamos a scrapear el sitio de Latam para averiguar datos de vuelos en funcion el origen y destino, fecha y cabina. La información que esperamos obtener de cada vuelo es:
# - Precio(s) disponibles
# - Horas de salida y llegada (duración)
# - Información de las escalas
#
# ¡Empecemos!
url = 'https://www.latam.com/es_ar/apps/personas/booking?fecha1_dia=20&fecha1_anomes=2019-12&auAvailability=1&ida_vuelta=ida&vuelos_origen=Buenos%20Aires&from_city1=BUE&vuelos_destino=Madrid&to_city1=MAD&flex=1&vuelos_fecha_salida_ddmmaaaa=20/12/2019&cabina=Y&nadults=1&nchildren=0&ninfants=0&cod_promo='
from selenium import webdriver
options = webdriver.ChromeOptions()
options.add_argument('--incognito')
driver = webdriver.Chrome(executable_path='../../chromedriver', options=options)
driver.get(url)
#Usaremos el Xpath para obtener la lista de vuelos
vuelos = driver.find_elements_by_xpath('//li[@class="flight"]')
vuelo = vuelos[0]
# Obtenemos la información de la hora de salida, llegada y duración del vuelo
# Hora de salida
vuelo.find_element_by_xpath('.//div[@class="departure"]/time').get_attribute('datetime')
# Hora de llegada
vuelo.find_element_by_xpath('.//div[@class="arrival"]/time').get_attribute('datetime')
# Duración del vuelo
vuelo.find_element_by_xpath('.//span[@class="duration"]/time').get_attribute('datetime')
boton_escalas = vuelo.find_element_by_xpath('.//div[@class="flight-summary-stops-description"]/button')
boton_escalas
boton_escalas.click()
segmentos = vuelo.find_elements_by_xpath('//div[@class="segments-graph"]/div[@class="segments-graph-segment"]')
segmentos
escalas = len(segmentos) - 1 #0 escalas si es un vuelo directo
segmento = segmentos[0]
# Origen
segmento.find_element_by_xpath('.//div[@class="departure"]/span[@class="ground-point-name"]').text
# Hora de salida
segmento.find_element_by_xpath('.//div[@class="departure"]/time').get_attribute('datetime')
# Destino
segmento.find_element_by_xpath('.//div[@class="arrival"]/span[@class="ground-point-name"]').text
# Hora de llegada
segmento.find_element_by_xpath('.//div[@class="arrival"]/time').get_attribute('datetime')
# Duración del vuelo
segmento.find_element_by_xpath('.//span[@class="duration flight-schedule-duration"]/time').get_attribute('datetime')
# Numero del vuelo
segmento.find_element_by_xpath('.//span[@class="equipment-airline-number"]').text
# Modelo de avion
segmento.find_element_by_xpath('.//span[@class="equipment-airline-material"]').text
# Duracion de la escala
segmento.find_element_by_xpath('.//div[@class="stop connection"]//p[@class="stop-wait-time"]//time').get_attribute('datetime')
vuelo.find_element_by_xpath('//div[@class="modal-dialog"]//button[@class="close"]').click()
vuelo.click()
tarifas = vuelo.find_elements_by_xpath('.//div[@class="fares-table-container"]//tfoot//td[contains(@class, "fare-")]')
precios = []
for tarifa in tarifas:
nombre = tarifa.find_element_by_xpath('.//label').get_attribute('for')
moneda = tarifa.find_element_by_xpath('.//span[@class="price"]/span[@class="currency-symbol"]').text
valor = tarifa.find_element_by_xpath('.//span[@class="price"]/span[@class="value"]').text
dict_tarifa={nombre:{'moneda':moneda, 'valor':valor}}
precios.append(dict_tarifa)
print(dict_tarifa)
def obtener_tiempos(vuelo):
# Hora de salida
salida = vuelo.find_element_by_xpath('.//div[@class="departure"]/time').get_attribute('datetime')
# Hora de llegada
llegada = vuelo.find_element_by_xpath('.//div[@class="arrival"]/time').get_attribute('datetime')
# Duracion
duracion = vuelo.find_element_by_xpath('.//span[@class="duration"]/time').get_attribute('datetime')
return {'hora_salida': salida, 'hora_llegada': llegada, 'duracion': duracion}
def obtener_precios(vuelo):
tarifas = vuelo.find_elements_by_xpath(
'.//div[@class="fares-table-container"]//tfoot//td[contains(@class, "fare-")]')
precios = []
for tarifa in tarifas:
nombre = tarifa.find_element_by_xpath('.//label').get_attribute('for')
moneda = tarifa.find_element_by_xpath('.//span[@class="price"]/span[@class="currency-symbol"]').text
valor = tarifa.find_element_by_xpath('.//span[@class="price"]/span[@class="value"]').text
dict_tarifa={nombre:{'moneda':moneda, 'valor':valor}}
precios.append(dict_tarifa)
return precios
def obtener_datos_escalas(vuelo):
segmentos = vuelo.find_elements_by_xpath('//div[@class="segments-graph"]/div[@class="segments-graph-segment"]')
info_escalas = []
for segmento in segmentos:
# Origen
origen = segmento.find_element_by_xpath(
'.//div[@class="departure"]/span[@class="ground-point-name"]').text
# Hora de salida
dep_time = segmento.find_element_by_xpath(
'.//div[@class="departure"]/time').get_attribute('datetime')
# Destino
destino = segmento.find_element_by_xpath(
'.//div[@class="arrival"]/span[@class="ground-point-name"]').text
# Hora de llegada
arr_time = segmento.find_element_by_xpath(
'.//div[@class="arrival"]/time').get_attribute('datetime')
# Duración del vuelo
duracion_vuelo = segmento.find_element_by_xpath(
'.//span[@class="duration flight-schedule-duration"]/time').get_attribute('datetime')
# Numero del vuelo
numero_vuelo = segmento.find_element_by_xpath(
'.//span[@class="equipment-airline-number"]').text
# Modelo de avion
modelo_avion = segmento.find_element_by_xpath(
'.//span[@class="equipment-airline-material"]').text
# Duracion de la escala
if segmento != segmentos[-1]:
duracion_escala = segmento.find_element_by_xpath(
'.//div[@class="stop connection"]//p[@class="stop-wait-time"]//time').get_attribute('datetime')
else:
duracion_escala = ''
# Armo un diccionario para almacenar los datos
data_dict={'origen': origen,
'dep_time': dep_time,
'destino': destino,
'arr_time': arr_time,
'duracion_vuelo': duracion_vuelo,
'numero_vuelo': numero_vuelo,
'modelo_avion': modelo_avion,
'duracion_escala': duracion_escala}
info_escalas.append(data_dict)
return info_escalas
def obtener_info(driver):
vuelos = driver.find_elements_by_xpath('//li[@class="flight"]')
print(f'Se encontraron {len(vuelos)} vuelos.')
print('Iniciando scraping...')
info = []
for vuelo in vuelos:
# Obtenemos los tiempos generales del vuelo
tiempos = obtener_tiempos(vuelo)
# Clickeamos el botón de escalas para ver los detalles
vuelo.find_element_by_xpath('.//div[@class="flight-summary-stops-description"]/button').click()
escalas = obtener_datos_escalas(vuelo)
# Cerramos el pop-up con los detalles
vuelo.find_element_by_xpath('//div[@class="modal-dialog"]//button[@class="close"]').click()
# Clickeamos el vuelo para ver los precios
vuelo.click()
precios = obtener_precios(vuelo)
# Cerramos los precios del vuelo
vuelo.click()
info.append({'precios':precios, 'tiempos':tiempos , 'escalas': escalas})
return info
# Ahora podemos cargar la página con el driver y pasárselo a esta función
driver = webdriver.Chrome(executable_path='../../chromedriver', options=options)
driver.get(url)
obtener_info(driver)
# Se encontraron 0 vuelos porque la página no terminó de cargar
# Lo más simple que podemos hacer es agregar una demora fija lo suficientemente grande para asegurarnos que la página terminó de cargar.
import time
options = webdriver.ChromeOptions()
options.add_argument('--incognito')
driver = webdriver.Chrome(executable_path='../../chromedriver', options=options)
driver.get(url)
time.sleep(10)
vuelos = driver.find_elements_by_xpath('//li[@class="flight"]')
vuelos
driver.close()
# Esto funciona pero no es muy eficiente. Lo mejor sería esperar a que la página termine de cargar y luego recuperar los elementos.
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.by import By
from selenium.common.exceptions import TimeoutException
options = webdriver.ChromeOptions()
options.add_argument('--incognito')
driver = webdriver.Chrome(executable_path='../../chromedriver', options=options)
driver.get(url)
delay = 10
try:
vuelo = WebDriverWait(driver, delay).until(EC.presence_of_element_located((By.XPATH, '//li[@class="flight"]')))
print("La página terminó de cargar")
info_vuelos = obtener_info(driver)
except TimeoutException:
print("La página tardó demasiado en cargar")
driver.close()
info_vuelos
# ## Clase 6
# Ya tenemos el bloque de código que nos permite obtener información de los vuelos a partir de una URL. ¿Pero cómo consturimos la URL?
print(url)
url_base = 'https://www.latam.com/es_ar/apps/personas/booking?'
params = url.strip(url_base).split('&')
params
fecha = '10/12/2019'
fecha = time.strptime(fecha, '%d/%m/%Y')
fecha
def armar_url(url_base, fecha, origen, destino, cabina):
url = url_base
url+=f'&fecha1_dia={fecha.tm_mday}'
url+=f'&fecha1_anomes={fecha.tm_year}-{fecha.tm_mon}'
url+=f'&auAvailability=1'
url+=f'&ida_vuelta=ida'
url+=f'&from_city1={origen}'
url+=f'&to_city1={destino}'
url+=f'&vuelos_fecha_salida_ddmmaaaa={time.strftime("%d/%m/%Y",fecha)}'
url+=f'&cabina={cabina}'
url+=f'&nadults=1'
return url
url = armar_url(url_base,fecha, 'MAD', 'BUE', 'Y')
url
urls = [armar_url(url_base, fecha, 'BUE', destino, 'Y') for destino in ['MAD', 'RMA', 'PAR']]
urls
def scrape_latam(urls):
options = webdriver.ChromeOptions()
options.add_argument('--incognito')
driver = webdriver.Chrome(executable_path='../../chromedriver', options=options)
delay = 10
# Si es un string único, lo convierto en lista
if type(urls) == str:
urls = [urls]
print(urls)
info = []
for url in urls:
print('Scraping URL:',url)
driver.get(url)
try:
vuelo = WebDriverWait(driver, delay).until(EC.presence_of_element_located((By.XPATH, '//li[@class="flight"]')))
print("La página terminó de cargar o no se encontraron vuelos disponibles")
info.append(obtener_info(driver))
except TimeoutException:
print("La página tardó demasiado en cargar")
driver.close()
return info
scrape_latam(urls)
| NoteBooks/Curso de WebScraping/Unificado/web-scraping-master/Clases_old/Módulo 3_ Scraping con Selenium/M3C6. Ejecutando el scraper - Script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + language="html"
#
# <link href="https://fonts.googleapis.com/css?family=Lora:400,700|Montserrat:300" rel="stylesheet">
#
# <link href="https://fonts.googleapis.com/css2?family=Crimson+Pro&family=Literata&display=swap" rel="stylesheet">
# <style>
#
#
# #ipython_notebook::before{
# content:"Machine Learning";
# color: white;
# font-weight: bold;
# text-transform: uppercase;
# font-family: 'Lora',serif;
# font-size:16pt;
# margin-bottom:10px;
#
# }
# body > #header {
# background: maroon;
# opacity: 0.7;
# }
#
#
# .navbar-default .navbar-nav > li > a, #kernel_indicator {
# color: white;
# transition: all 0.25s;
# font-size:12pt;
# font-family: sans;
# /*font-weight:bold;*/
# }
# .navbar-default {
# padding-left:100px;
# background: none;
# border: none;
# }
#
#
# body > menubar-container {
# background-color: wheat;
# }
# #ipython_notebook img{
# display:block;
#
# background: url("") no-repeat;
# background-size: contain;
#
# padding-left: 500px;
# padding-right: 100px;
#
# -moz-box-sizing: border-box;
# box-sizing: border-box;
# }
#
#
#
# body {
# #font-family: 'Literata', serif;
# font-family:'Lora', san-serif;
# text-align: justify;
# font-weight: 400;
# font-size: 13pt;
# }
#
# iframe{
# width:100%;
# min-height:600px;
# }
#
# h1, h2, h3, h4, h5, h6 {
# # font-family: 'Montserrat', sans-serif;
# font-family:'Lora', serif;
# font-weight: 200;
# text-transform: uppercase;
# color: #EC7063 ;
# }
#
# h2 {
# color: #000080;
# }
#
# .checkpoint_status, .autosave_status {
# color:wheat;
# }
#
# #notebook_name {
# font-weight: 1000;
# font-size:20pt;
# text-variant:uppercase;
# color: wheat;
# margin-right:10px;
# }
# #notebook_name:hover {
# background-color: salmon;
# }
#
#
# .dataframe { /* dataframe atau table */
# background: white;
# box-shadow: 0px 1px 2px #bbb;
# }
# .dataframe thead th, .dataframe tbody td {
# text-align: center;
# padding: 1em;
# }
#
# .checkpoint_status, .autosave_status {
# color:wheat;
# }
#
# .output {
# align-items: center;
# }
#
# div.cell {
# transition: all 0.25s;
# border: none;
# position: relative;
# top: 0;
# }
# div.cell.selected, div.cell.selected.jupyter-soft-selected {
# border: none;
# background: transparent;
# box-shadow: 0 6px 18px #aaa;
# z-index: 10;
# top: -10px;
# }
# .CodeMirror pre, .CodeMirror-dialog, .CodeMirror-dialog .CodeMirror-search-field, .terminal-app .terminal {
# font-family: 'Source Code Pro Medium' , serif;
# font-weight: 500;
# font-size: 13pt;
# }
#
#
#
# </style>
# -
# ### IMPORTING THE NECESSARY LIBRARIES
# +
# Starting with the standard imports
import numpy as np
import pandas as pd
import pandas_profiling
# Preprocessing data
from sklearn.model_selection import train_test_split # data-splitter
from sklearn.preprocessing import StandardScaler # data-normalization
from sklearn.preprocessing import PolynomialFeatures # for polynomials
from sklearn.preprocessing import PowerTransformer # for power-transformations
from sklearn.pipeline import make_pipeline # for pipelines
np.random.seed (42) # for reproducible results
#
# Modeling and Metrics
#
# --For Regressor
from sklearn.dummy import DummyRegressor # baseline regressor (null-hypothesis)
from sklearn.linear_model import LinearRegression # linear regression
from sklearn.linear_model import ( Ridge,
Lasso,
ElasticNet,
RidgeCV,
LassoCV,
ElasticNetCV) # regularized regressions with CV
from sklearn.metrics import mean_squared_error, r2_score # model-metrics
from sklearn.ensemble import RandomForestRegressor
#
# For Classifiers
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.dummy import (DummyClassifier, DummyRegressor)
#
# For clusterers
from scipy import stats, integrate
import sklearn.cluster as cluster
from sklearn.cluster import (DBSCAN, KMeans)
from sklearn import metrics
from sklearn.datasets import make_blobs
# Yellowbrick
from yellowbrick.features import FeatureImportances
from yellowbrick.classifier import ConfusionMatrix, ClassificationReport, ROCAUC
from matplotlib import colors
import seaborn as sns
list_of_cmaps=['Blues','Greens','Reds','Purples'] # some colors to use
# Now the Graphical libraries imports and settings
# %matplotlib inline
import matplotlib.pyplot as plt # for plotting
import seaborn as sns # nicer looking plots
import altair as alt # for interactive plots
from matplotlib import colors # for web-color specs
pd.set_option('plotting.backend', 'matplotlib') # pandas_bokeh, plotly, etc
plt.rcParams[ 'figure.figsize' ] = '20,10' # landscape format figures
plt.rcParams[ 'legend.fontsize' ] = 13 # legend font size
plt.rcParams[ 'axes.labelsize' ] = 13 # axis label font size
plt.rcParams['figure.dpi'] = 144 # high-dpi monitors support
plt.style.use ('ggplot') # emulate ggplot style
# For latex-quality, i.e., publication quality legends and labels on graphs.
# Warning: you must have installed LaTeX on your system.
from matplotlib import rc
rc ('font', family='serif')
rc ('text', usetex=True) # Enable it selectively
rc ('font', size=16)
# For youtube video display
from IPython.display import YouTubeVideo
import warnings
warnings.filterwarnings ('ignore') # suppress warning
# -
def sv_table_styles():
th_props = [
('font-size', '11pt'),
('font-family', 'sans'),
('text-align', 'center'),
('font-weight', '300'),
('color', 'cornsilk'),
('background-color', 'salmon')
]
# Set CSS properties for td elements in dataframe
td_props = [
('font-size', '10px'),
#('color', 'cornsilk'),
('font-weight', 'normal')
]
# Currently, could not make this work!
first_col_props = [
('background-color', 'cornsilk'),
('color', 'black'),
('font-weight', '300'),
]
# Set table styles
styles = [
dict(selector="th", props=th_props),
dict(selector="td", props=td_props),
dict(selection="tr td:first-child()", props=first_col_props)
]
return styles
#
# Rotate Pandas dataframe column headers.
# Taken from:
# https://stackoverflow.com/questions/46715736/rotating-the-column-name-for-a-panda-dataframe
#
def format_vertical_headers(df):
"""Display a dataframe with vertical column headers"""
styles = [
dict(selector="th", props=[('width', '40px')]),
dict(selector="th.col_heading",
props=[("writing-mode", "vertical-rl"),
('transform', 'rotateZ(180deg)'),
('height', '160px'),
('vertical-align', 'top')])]
return (df.fillna('').style.set_table_styles(styles))
| notebook/common-imports.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: urf-8 -*-
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
import requests
from bs4 import BeautifulSoup
import re
# +
# 크롤링할 사이트 주소를 정의합니다.
source_url = "https://namu.wiki/RecentChanges"
# 사이트의 html 구조에 기반하여 크롤링을 수행합니다.
req = requests.get(source_url)
html = req.content
soup = BeautifulSoup(html, 'lxml')
contents_table = soup.find(name="table")
table_body = contents_table.find(name="tbody")
table_rows = table_body.find_all(name="tr")
# a태그의 href 속성을 리스트로 추출하여, 크롤링 할 페이지 리스트를 생성합니다.
page_url_base = "https://namu.wiki"
page_urls = []
for index in range(0, len(table_rows)):
first_td = table_rows[index].find_all('td')[0]
td_url = first_td.find_all('a')
if len(td_url) > 0:
page_url = page_url_base + td_url[0].get('href')
if 'png' not in page_url:
page_urls.append(page_url)
# 중복 url을 제거합니다.
page_urls = list(set(page_urls))
for page in page_urls[:5]:
print(page)
# +
# 크롤링한 데이터를 데이터 프레임으로 만들기 위해 준비합니다.
columns = ['title', 'category', 'content_text']
df = pd.DataFrame(columns=columns)
# 각 페이지별 '제목', '카테고리', '본문' 정보를 데이터 프레임으로 만듭니다.
for page_url in page_urls:
# 사이트의 html 구조에 기반하여 크롤링을 수행합니다.
req = requests.get(page_url)
html = req.content
soup = BeautifulSoup(html, 'lxml')
contents_table = soup.find(name="article")
title = contents_table.find_all('h1')[0]
# 카테고리 정보가 없는 경우를 확인합니다.
if len(contents_table.find_all('ul')) > 0:
category = contents_table.find_all('ul')[0]
else:
category = None
content_paragraphs = contents_table.find_all(name="div", attrs={"class":"wiki-paragraph"})
content_corpus_list = []
# 페이지 내 제목 정보에서 개행 문자를 제거한 뒤 추출합니다.
# 만약 없는 경우, 빈 문자열로 대체합니다.
if title is not None:
row_title = title.text.replace("\n"," ")
else:
row_title = ""
# 페이지 내 본문 정보에서 개행 문자를 제거한 뒤 추출합니다.
# 만약 없는 경우, 빈 문자열로 대체합니다.
if content_paragraphs is not None:
for paragraphs in content_paragraphs:
if paragraphs is not None:
# replace("찾을값", "바꿀값", [바꿀횟수])
content_corpus_list.append(paragraphs.text.replace("\n", " "))
else:
content_corpus_list.append("")
else:
content_corpus_list.append("")
# 페이지 내 카테고리정보에서 "분류"라는 단어와 개행 문자를 제거한 뒤 추출합니다.
# 만약 없는 경우, 빈 문자열로 대체합니다.
if category is not None:
row_category = category.text.replace("\n", " ")
else:
row_category = ""
# 모든 정보를 하나의 데이터 프레임에 저장합니다.
# "문자".join() 리스트에 특정 구분자를 추가하여 문자열로 변환함
row = [row_title, row_category, "".join(content_corpus_list)]
series = pd.Series(row, index = df.columns)
df = df.append(series, ignore_index=True)
# -
# 데이터 프레임을 출력합니다.
df.head(5)
# +
# 텍스트 정제 함수: 한글 이외의 문자는 전부 제거합니다.
def text_cleaning(text):
hangul = re.compile('[^ ㄱ-ㅣ가-힣]+') # 한글의 정규 표현식을 나타냅니다.
result = hangul.sub('', text)
return result
print(text_cleaning(df['content_text'][0]))
# -
# 각 피처마다 데이터 전처리를 적용합니다.
df['title'] = df['title'].apply(lambda x : text_cleaning(x))
df['category'] = df['category'].apply(lambda x : text_cleaning(x))
df['content_text'] = df['content_text'].apply(lambda x : text_cleaning(x))
df.head(5)
# 각 피처마다 말뭉치를 생성합니다.
title_corpus = "".join(df['title'].tolist())
category_corpus = "".join(df['category'].tolist())
content_corpus = "".join(df['content_text'].tolist())
print(title_corpus)
# !pip install konlpy==0.5.1
# !pip install jpype1
# !pip install Jpype1-py3
# +
from konlpy.tag import Okt
from collections import Counter
# konply의 형태소 분석기로 명사 단위의 키워드를 추출합니다.
nouns_tagger = Okt()
nouns = nouns_tagger.nouns(content_corpus)
count = Counter(nouns)
# -
count
# 한 글자 키워드를 제거합니다.
remove_char_counter = Counter({x : count[x] for x in count if len(x) > 1})
print(remove_char_counter)
# +
# 한국어 약식 불용어 사전 예시 파일을 적용합니다.
korean_stopwords_path = "../data/korean_stopwords.txt"
# 텍스트 파일을 오픈합니다.
with open(korean_stopwords_path, encoding='utf8') as f:
# readlines()로 파일을 읽으면 한 줄, 한 줄이 각각 리스트의 원소로 들어갑니다.
stopwords = f.readlines()
stopwords = [x.strip() for x in stopwords]
print(stopwords[:10])
# +
# 나무위키 페이지에 맞는 불용어를 추가합니다.
namu_wiki_stopwords = ['상위', '문서', '내용', '누설', '아래', '해당', '설명', '표기', '추가', '모든', '사용', '매우', '가장',
'줄거리', '요소', '상황', '편집', '틀', '경우', '때문', '모습', '정도', '이후', '사실', '생각', '인물',
'이름', '년월']
for stopword in namu_wiki_stopwords:
stopwords.append(stopword)
# 키워드 데이터에서 불용어를 제거합니다.
remove_char_counter = Counter({x : remove_char_counter[x] for x in count if x not in stopwords})
print(remove_char_counter)
# -
import random
import pytagcloud
import webbrowser
# +
# 가장 출현 빈도수가 높은 40개의 단어를 선정합니다.
ranked_tags = remove_char_counter.most_common(40)
# pytagcloud로 출력할 40개의 단어를 입력합니다. 단어 출력의 최대 크기는80으로 제한합니다.
taglist = pytagcloud.make_tags(ranked_tags, maxsize=80)
# pytagcloud 이미지를 생성합니다. 폰트는 나눔 고딕을 사용합니다.
pytagcloud.create_tag_image(taglist, 'wordcloud.jpg', size=(900,600),
fontname='NanumGothic', rectangular=False)
# 생성한 이미지를 주피터 노트북상에서 출력합니다.
from IPython.display import Image
Image(filename='wordcloud.jpg')
# -
| chapter2/01-namu-wiki-text-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Lab 2: Classification
#
# ### Machine Learning 1, September 2016
#
# * The lab exercises should be made in groups of two people.
# * The deadline is October 9th (Sunday) 23:59.
# * Assignment should be sent to your teaching assistant. The subject line of your email should be "lab\#\_lastname1\_lastname2\_lastname3".
# * Put your and your teammates' names in the body of the email.
# * Attach the .IPYNB (IPython Notebook) file containing your code and answers. Naming of the file follows the same rule as the subject line. For example, if the subject line is "lab01\_Kingma\_Hu", the attached file should be "lab01\_Kingma\_Hu.ipynb". Only use underscores ("\_") to connect names, otherwise the files cannot be parsed.
#
# Notes on implementation:
#
# * For this notebook you need to answer a few theory questions, add them in the Markdown cell's below the question. Note: you can use Latex-style code in here.
# * Focus on Part 1 the first week, and Part 2 the second week!
# * You should write your code and answers below the questions in this IPython Notebook.
# * Among the first lines of your notebook should be "%pylab inline". This imports all required modules, and your plots will appear inline.
# * If you have questions outside of the labs, post them on blackboard or email me.
# * NOTE: Make sure we can run your notebook / scripts!
#
# $\newcommand{\bx}{\mathbf{x}}$
# $\newcommand{\bw}{\mathbf{w}}$
# $\newcommand{\bt}{\mathbf{t}}$
# $\newcommand{\by}{\mathbf{y}}$
# $\newcommand{\bm}{\mathbf{m}}$
# $\newcommand{\bb}{\mathbf{b}}$
# $\newcommand{\bS}{\mathbf{S}}$
# $\newcommand{\ba}{\mathbf{a}}$
# $\newcommand{\bz}{\mathbf{z}}$
# $\newcommand{\bv}{\mathbf{v}}$
# $\newcommand{\bq}{\mathbf{q}}$
# $\newcommand{\bp}{\mathbf{p}}$
# $\newcommand{\bh}{\mathbf{h}}$
# $\newcommand{\bI}{\mathbf{I}}$
# $\newcommand{\bX}{\mathbf{X}}$
# $\newcommand{\bT}{\mathbf{T}}$
# $\newcommand{\bPhi}{\mathbf{\Phi}}$
# $\newcommand{\bW}{\mathbf{W}}$
# $\newcommand{\bV}{\mathbf{V}}$
# $\newcommand{\bA}{\mathbf{A}}$
# $\newcommand{\bj}{\mathbf{j}}$
# %matplotlib inline
# %pylab inline
import gzip, cPickle
import numpy as np
# # Part 1. Multiclass logistic regression
#
# Scenario: you have a friend with one big problem: she's completely blind. You decided to help her: she has a special smartphone for blind people, and you are going to develop a mobile phone app that can do _machine vision_ using the mobile camera: converting a picture (from the camera) to the meaning of the image. You decide to start with an app that can read handwritten digits, i.e. convert an image of handwritten digits to text (e.g. it would enable her to read precious handwritten phone numbers).
#
# A key building block for such an app would be a function `predict_digit(x)` that returns the digit class of an image patch $\bx$. Since hand-coding this function is highly non-trivial, you decide to solve this problem using machine learning, such that the internal parameters of this function are automatically learned using machine learning techniques.
#
# The dataset you're going to use for this is the MNIST handwritten digits dataset (`http://yann.lecun.com/exdb/mnist/`). You can load the data from `mnist.pkl.gz` we provided, using:
# +
def load_mnist():
f = gzip.open('mnist.pkl.gz', 'rb')
data = cPickle.load(f)
f.close()
return data
(x_train, t_train), (x_valid, t_valid), (x_test, t_test) = load_mnist()
# -
# The tuples represent train, validation and test sets. The first element (`x_train`, `x_valid`, `x_test`) of each tuple is a $N \times M$ matrix, where $N$ is the number of datapoints and $M = 28^2 = 784$ is the dimensionality of the data. The second element (`t_train`, `t_valid`, `t_test`) of each tuple is the corresponding $N$-dimensional vector of integers, containing the true class labels.
#
# Here's a visualisation of the first 8 digits of the trainingset:
# +
def plot_digits(data, numcols, shape=(28,28)):
numdigits = data.shape[0]
numrows = int(numdigits/numcols)
for i in range(numdigits):
plt.subplot(numrows, numcols, i+1)
plt.axis('off')
plt.imshow(data[i].reshape(shape), interpolation='nearest', cmap='Greys')
plt.show()
plot_digits(x_train[0:8], numcols=4)
# -
# In _multiclass_ logistic regression, the conditional probability of class label $j$ given the image $\bx$ for some datapoint is given by:
#
# $ \log p(t = j \;|\; \bx, \bb, \bW) = \log q_j - \log Z$
#
# where $\log q_j = \bw_j^T \bx + b_j$ (the log of the unnormalized probability of the class $j$), and $Z = \sum_k q_k$ is the normalizing factor. $\bw_j$ is the $j$-th column of $\bW$ (a matrix of size $784 \times 10$) corresponding to the class label, $b_j$ is the $j$-th element of $\bb$.
#
# Given an input image, the multiclass logistic regression model first computes the intermediate vector $\log \bq$ (of size $10 \times 1$), using $\log q_j = \bw_j^T \bx + b_j$, containing the unnormalized log-probabilities per class.
#
# The unnormalized probabilities are then normalized by $Z$ such that $\sum_j p_j = \sum_j \exp(\log p_j) = 1$. This is done by $\log p_j = \log q_j - \log Z$ where $Z = \sum_j \exp(\log q_j)$. This is known as the _softmax_ transformation, and is also used as a last layer of many classifcation neural network models, to ensure that the output of the network is a normalized distribution, regardless of the values of second-to-last layer ($\log \bq$)
#
# Warning: when computing $\log Z$, you are likely to encounter numerical problems. Save yourself countless hours of debugging and learn the [log-sum-exp trick](https://hips.seas.harvard.edu/blog/2013/01/09/computing-log-sum-exp/ "Title").
#
# The network's output $\log \bp$ of size $10 \times 1$ then contains the conditional log-probabilities $\log p(t = j \;|\; \bx, \bb, \bW)$ for each digit class $j$. In summary, the computations are done in this order:
#
# $\bx \rightarrow \log \bq \rightarrow Z \rightarrow \log \bp$
#
# Given some dataset with $N$ independent, identically distributed datapoints, the log-likelihood is given by:
#
# $ \mathcal{L}(\bb, \bW) = \sum_{n=1}^N \mathcal{L}^{(n)}$
#
# where we use $\mathcal{L}^{(n)}$ to denote the partial log-likelihood evaluated over a single datapoint. It is important to see that the log-probability of the class label $t^{(n)}$ given the image, is given by the $t^{(n)}$-th element of the network's output $\log \bp$, denoted by $\log p_{t^{(n)}}$:
#
# $\mathcal{L}^{(n)} = \log p(t = t^{(n)} \;|\; \bx = \bx^{(n)}, \bb, \bW) = \log p_{t^{(n)}} = \log q_{t^{(n)}} - \log Z^{(n)}$
#
# where $\bx^{(n)}$ and $t^{(n)}$ are the input (image) and class label (integer) of the $n$-th datapoint, and $Z^{(n)}$ is the normalizing constant for the distribution over $t^{(n)}$.
#
# ## 1.1 Gradient-based stochastic optimization
#
# ### 1.1.1 Derive gradient equations (20 points)
#
# Derive the equations for computing the (first) partial derivatives of the log-likelihood w.r.t. all the parameters, evaluated at a _single_ datapoint $n$.
#
# You should start deriving the equations for $\frac{\partial \mathcal{L}^{(n)}}{\partial \log q_j}$ for each $j$. For clarity, we'll use the shorthand $\delta^q_j = \frac{\partial \mathcal{L}^{(n)}}{\partial \log q_j}$.
#
# For $j = t^{(n)}$:
# $
# \delta^q_j
# = \frac{\partial \mathcal{L}^{(n)}}{\partial \log p_j}
# \frac{\partial \log p_j}{\partial \log q_j}
# + \frac{\partial \mathcal{L}^{(n)}}{\partial \log Z}
# \frac{\partial \log Z}{\partial Z}
# \frac{\partial Z}{\partial \log q_j}
# = 1 \cdot 1 - \frac{\partial \log Z}{\partial Z}
# \frac{\partial Z}{\partial \log q_j}
# = 1 - \frac{\partial \log Z}{\partial Z}
# \frac{\partial Z}{\partial \log q_j}
# $
#
# For $j \neq t^{(n)}$:
# $
# \delta^q_j
# = \frac{\partial \mathcal{L}^{(n)}}{\partial \log Z}
# \frac{\partial \log Z}{\partial Z}
# \frac{\partial Z}{\partial \log q_j}
# = - \frac{\partial \log Z}{\partial Z}
# \frac{\partial Z}{\partial \log q_j}
# $
#
# Complete the above derivations for $\delta^q_j$ by furtherly developing $\frac{\partial \log Z}{\partial Z}$ and $\frac{\partial Z}{\partial \log q_j}$. Both are quite simple. For these it doesn't matter whether $j = t^{(n)}$ or not.
#
# Given your equations for computing the gradients $\delta^q_j$ it should be quite straightforward to derive the equations for the gradients of the parameters of the model, $\frac{\partial \mathcal{L}^{(n)}}{\partial W_{ij}}$ and $\frac{\partial \mathcal{L}^{(n)}}{\partial b_j}$. The gradients for the biases $\bb$ are given by:
#
# $
# \frac{\partial \mathcal{L}^{(n)}}{\partial b_j}
# = \frac{\partial \mathcal{L}^{(n)}}{\partial \log q_j}
# \frac{\partial \log q_j}{\partial b_j}
# = \delta^q_j
# \cdot 1
# = \delta^q_j
# $
#
# The equation above gives the derivative of $\mathcal{L}^{(n)}$ w.r.t. a single element of $\bb$, so the vector $\nabla_\bb \mathcal{L}^{(n)}$ with all derivatives of $\mathcal{L}^{(n)}$ w.r.t. the bias parameters $\bb$ is:
#
# $
# \nabla_\bb \mathcal{L}^{(n)} = \mathbf{\delta}^q
# $
#
# where $\mathbf{\delta}^q$ denotes the vector of size $10 \times 1$ with elements $\mathbf{\delta}_j^q$.
#
# The (not fully developed) equation for computing the derivative of $\mathcal{L}^{(n)}$ w.r.t. a single element $W_{ij}$ of $\bW$ is:
#
# $
# \frac{\partial \mathcal{L}^{(n)}}{\partial W_{ij}} =
# \frac{\partial \mathcal{L}^{(n)}}{\partial \log q_j}
# \frac{\partial \log q_j}{\partial W_{ij}}
# = \mathbf{\delta}_j^q
# \frac{\partial \log q_j}{\partial W_{ij}}
# $
#
# What is $\frac{\partial \log q_j}{\partial W_{ij}}$? Complete the equation above.
#
# If you want, you can give the resulting equation in vector format ($\nabla_{\bw_j} \mathcal{L}^{(n)} = ...$), like we did for $\nabla_\bb \mathcal{L}^{(n)}$
# #### Answer:
#
# ---
#
# First, $\frac{\partial \log Z}{\partial Z} = \frac{1}{Z}$.
#
# Recall $Z = \sum_j \exp(\log q_j)$. Then, $\frac{\partial Z}{\partial \log q_j} = \exp(\log q_j) = q_j$.
#
# Finally, $\delta^q_j = \mathbb{I}_{j,t^{(n)}} - \frac{\partial \log Z}{\partial Z}
# \frac{\partial Z}{\partial \log q_j} = \mathbb{I}_{j,t^{(n)}} - \frac{q_j}{Z}$. Where $\mathbb{I}$ is the identity matrix.
#
# ---
#
# $\frac{\partial \log q_j}{\partial \bW_j} = \frac{\partial \bw_j^T \bx +\bb_j}{\partial \bW_j} = \bx \rightarrow \frac{\partial \log q_j}{\partial \bW_{ij}} = \bx_i$
#
# $\nabla_{\bw_j} \mathcal{L}^{(n)} = \mathbf{\delta}_j^q \bx \rightarrow \nabla_{\bW} \mathcal{L}^{(n)} = \bx {\mathbf{\delta}^q}^T$
#
# ---
# ### 1.1.2 Implement gradient computations (10 points)
#
# Implement the gradient calculations you derived in the previous question. Write a function `logreg_gradient(x, t, w, b)` that returns the gradients $\nabla_{\bw_j} \mathcal{L}^{(n)}$ (for each $j$) and $\nabla_{\bb} \mathcal{L}^{(n)}$, i.e. the first partial derivatives of the log-likelihood w.r.t. the parameters $\bW$ and $\bb$, evaluated at a single datapoint (`x`, `t`).
# The computation will contain roughly the following intermediate variables:
#
# $
# \log \bq \rightarrow Z \rightarrow \log \bp\,,\, \mathbf{\delta}^q
# $
#
# followed by computation of the gradient vectors $\nabla_{\bw_j} \mathcal{L}^{(n)}$ (contained in a $784 \times 10$ matrix) and $\nabla_{\bb} \mathcal{L}^{(n)}$ (a $10 \times 1$ vector).
#
def logreg_gradient(x,t,W,b):
lq = dot(W.T,x) + b
Z = sum(np.exp(lq))
delta_q = - exp(lq)/Z
delta_q[t] += 1
grad_b = delta_q
grad_W = dot(x,delta_q.T)
return grad_W, grad_b
#
# ### 1.1.3 Stochastic gradient descent (10 points)
#
# Write a function `sgd_iter(x_train, t_train, w, b)` that performs one iteration of stochastic gradient descent (SGD), and returns the new weights. It should go through the trainingset once in randomized order, call `logreg_gradient(x, t, w, b)` for each datapoint to get the gradients, and update the parameters using a small learning rate (e.g. `1E-4`). Note that in this case we're maximizing the likelihood function, so we should actually performing gradient ___ascent___... For more information about SGD, see Bishop 5.2.4 or an online source (i.e. https://en.wikipedia.org/wiki/Stochastic_gradient_descent)
def sgd_iter(x_train, t_train, W, b):
N, M = shape(x_train)
perm = np.random.permutation(N)
eta = 0.0001
for ix in perm:
x = reshape(x_train[ix],(M,-1))
t = t_train[ix]
grad_W, grad_b = logreg_gradient(x,t,W,b)
W += eta * grad_W
b += eta * grad_b
return W, b
# ## 1.2. Train
#
# ### 1.2.1 Train (10 points)
# Perform a handful of training iterations through the trainingset. Plot (in one graph) the conditional log-probability of the trainingset and validation set after each iteration.
#
# +
def logreg(x,t,W,b):
lq = dot(W.T,x) + b
Z = sum(np.exp(lq))
a = max(lq)
lZ = a + log(sum(np.exp(lq - ones(shape(lq))*a)))
lp = lq - ones(shape(lq))*log(Z)
lpt = lp[t]
return lpt, argmax(lp)
def pred_iter(x_set, t_set, W, b):
N_set, M = shape(x_set)
L = 0
misclass = 0
for ix in range(N_set):
x = reshape(x_set[ix],(M,-1))
t = t_set[ix]
lpt, lpix = logreg(x,t,W,b)
L += lpt
misclass += lpix != t
return L, misclass*1.0/N_set
N_tr, M = shape(x_train)
N_vl, M = shape(x_test)
K = 10
max_iter = 10
W = np.zeros((M,K))
b = np.zeros((K, 1))
L_tr = []
E_tr = []
L_vl = []
E_vl = []
L, err = pred_iter(x_train, t_train, W, b)
L_tr.append(L)
E_tr.append(100*err)
L, err = pred_iter(x_valid, t_valid, W, b)
L_vl.append(L)
E_vl.append(100*err)
for it in range(max_iter):
#To visualize weights after each iteration uncomment the following line
#plot_digits(W.T, numcols=10)
W, b = sgd_iter(x_train, t_train, W, b)
L, err = pred_iter(x_train, t_train, W, b)
L_tr.append(L)
E_tr.append(100*err)
L, err = pred_iter(x_valid, t_valid, W, b)
L_vl.append(L)
E_vl.append(100*err)
plot(range(max_iter + 1), E_tr, label='Training')
plot(range(max_iter + 1), E_vl, label='Validation')
xlabel('Iteration')
ylabel('Error (%)')
legend(loc=1,ncol=1)
show()
plot(range(max_iter + 1), L_tr, label='Training')
plot(range(max_iter + 1), L_vl, label='Validation')
xlabel('Iteration')
ylabel('Log-Likelihood')
legend(loc=4,ncol=1)
show()
# -
# ### 1.2.2 Visualize weights (10 points)
# Visualize the resulting parameters $\bW$ after a few iterations through the training set, by treating each column of $\bW$ as an image. If you want, you can use or edit the `plot_digits(...)` above.
# Final weights
plot_digits(W.T, numcols=5)
# ### 1.2.3. Visualize the 8 hardest and 8 easiest digits (10 points)
# Visualize the 8 digits in the validation set with the highest probability of the true class label under the model.
# Also plot the 8 digits that were assigned the lowest probability.
# Ask yourself if these results make sense.
# +
P = []
for ix in range(N_vl):
x = reshape(x_valid[ix],(M,-1))
t = t_valid[ix]
lpt, lpix = logreg(x,t,W,b)
P.append(lpt[0])
P = np.array(P)
ix_hard = P.argsort()[:8]
ix_easy = P.argsort()[-8:]
print ('Hardest 8 Examples')
plot_digits(x_valid[ix_hard], numcols=4)
print ('Easiest 8 Examples')
plot_digits(x_valid[ix_easy], numcols=4)
# -
# # Part 2. Multilayer perceptron
#
#
# You discover that the predictions by the logistic regression classifier are not good enough for your application: the model is too simple. You want to increase the accuracy of your predictions by using a better model. For this purpose, you're going to use a multilayer perceptron (MLP), a simple kind of neural network. The perceptron wil have a single hidden layer $\bh$ with $L$ elements. The parameters of the model are $\bV$ (connections between input $\bx$ and hidden layer $\bh$), $\ba$ (the biases/intercepts of $\bh$), $\bW$ (connections between $\bh$ and $\log q$) and $\bb$ (the biases/intercepts of $\log q$.
#
# The conditional probability of the class label $j$ is given by:
#
# $\log p(t = j \;|\; \bx, \bb, \bW) = \log q_j - \log Z$
#
# where $q_j$ are again the unnormalized probabilities per class, and $Z = \sum_j q_j$ is again the probability normalizing factor. Each $q_j$ is computed using:
#
# $\log q_j = \bw_j^T \bh + b_j$
#
# where $\bh$ is a $L \times 1$ vector with the hidden layer activations (of a hidden layer with size $L$), and $\bw_j$ is the $j$-th column of $\bW$ (a $L \times 10$ matrix). Each element of the hidden layer is computed from the input vector $\bx$ using:
#
# $h_j = \sigma(\bv_j^T \bx + a_j)$
#
# where $\bv_j$ is the $j$-th column of $\bV$ (a $784 \times L$ matrix), $a_j$ is the $j$-th element of $\ba$, and $\sigma(.)$ is the so-called sigmoid activation function, defined by:
#
# $\sigma(x) = \frac{1}{1 + \exp(-x)}$
#
# Note that this model is almost equal to the multiclass logistic regression model, but with an extra 'hidden layer' $\bh$. The activations of this hidden layer can be viewed as features computed from the input, where the feature transformation ($\bV$ and $\ba$) is learned.
#
# ## 2.1 Derive gradient equations (20 points)
#
# State (shortly) why $\nabla_{\bb} \mathcal{L}^{(n)}$ is equal to the earlier (multiclass logistic regression) case, and why $\nabla_{\bw_j} \mathcal{L}^{(n)}$ is almost equal to the earlier case.
#
# Like in multiclass logistic regression, you should use intermediate variables $\mathbf{\delta}_j^q$. In addition, you should use intermediate variables $\mathbf{\delta}_j^h = \frac{\partial \mathcal{L}^{(n)}}{\partial h_j}$.
#
# Given an input image, roughly the following intermediate variables should be computed:
#
# $
# \log \bq \rightarrow Z \rightarrow \log \bp \rightarrow \mathbf{\delta}^q \rightarrow \mathbf{\delta}^h
# $
#
# where $\mathbf{\delta}_j^h = \frac{\partial \mathcal{L}^{(n)}}{\partial \bh_j}$.
#
# Give the equations for computing $\mathbf{\delta}^h$, and for computing the derivatives of $\mathcal{L}^{(n)}$ w.r.t. $\bW$, $\bb$, $\bV$ and $\ba$.
#
# You can use the convenient fact that $\frac{\partial}{\partial x} \sigma(x) = \sigma(x) (1 - \sigma(x))$.
# #### Answer:
#
# We are given the following matrices (vectors) with their corresponding dimensions: $\bW_{L\times K}$, $\bV_{M \times L}$, $\bx_{M\times 1}$, $\bh_{L\times 1}$, $\bb_{K\times 1}$, $\ba_{L\times 1}$. The notation $\bA_{i\cdot}$ and $\bA_{\cdot j }$ represent the $i$-th row and $j$-th column of the matrix $\bA$, respectively. Let $\bj$ denote a $L$ dimensional column vector of ones and $\bA \odot \mathbf{B}$ the Hadamard product between $\bA$ and $\mathbf{B}$ of dimensions $m\times n$.
#
# In this case, the model is summarized by the following equations:
# * $\bh = \sigma( \bV^T \bx + \ba)$
# * $\log \bq = \bW^T \bh + \bb$
# * $\log q_k = \bW_{\cdot k}^T \bh + \bb_k$
# * $ Z = \sum_k q_k = \sum_k \log(\exp(q_k))$
#
# ---
#
# Note that $\nabla_{\bb} \mathcal{L}^{(n)} = \frac{\partial \mathcal{L}^{(n)}}{\partial \log q_j} \frac{\partial \log q_j}{\partial b_j} = \mathbf{\delta}_j^q \cdot 1 = \mathbf{\delta}_j^q$. Therefore, it is the same as in the previous case.
#
# $$\nabla_{\bb} \mathcal{L}^{(n)} = \mathbf{\delta}^q = \mathbb{I}_{\cdot t^{(n)}} - \frac{1}{Z} \bq$$
#
# Besides, $\nabla_{\bW_{\cdot j}} \mathcal{L}^{(n)} = \frac{\partial \mathcal{L}^{(n)}}{\partial \log q_j} \frac{\partial \log q_j}{\partial \bW_{\cdot j}} = \mathbf{\delta}_j^q \frac{\partial \bW_{\cdot j}^T \bh + b_j}{\partial \bW_{\cdot j}} = \mathbf{\delta}_j^q \bh$. Thus, this is almost the same as the previous case, replacing $\bx$ by the new input of the logistic regression $\bh$.
#
# $$\nabla_{\bW} \mathcal{L}^{(n)} = \bh{\mathbf{\delta}^q}^T$$
#
# ---
#
# $ \mathbf{\delta}^h = \nabla_{\bh} \mathcal{L}^{(n)} = \frac{\partial \log q_{t^{(n)}}}{\partial \bh} - \frac{\partial \log Z}{\partial \bh} = \frac{\partial \bW_{\cdot{t^{(n)}}}^T \bh + \bb_{t^{(n)}} }{\partial \bh} - \frac{\partial \log \sum_k exp(\bW_{\cdot k }^T \bh + \bb_k)}{\partial \bh} = \bW_{\cdot{t^{(n)}}} - \frac{1}{\sum_k exp(\bW_{\cdot k }^T \bh + \bb_k)} \frac{\partial \sum_k exp(\bW_{\cdot k}^T \bh + \bb_k)}{\partial \bh}$
# $ \hspace{5mm} = \bW_{\cdot{t^{(n)}}} - \frac{1}{\sum_k exp(\bW_{\cdot k }^T \bh + \bb_k)} \sum_k exp(\bW_{\cdot k }^T \bh + \bb_k) \bW_{\cdot k} = \bW_{\cdot{t^{(n)}}} - \sum_k \frac{q_k}{Z} \bW_{\cdot k} = \sum_k (\mathbb{I}_{k,t^{(n)}} - \frac{q_k}{Z}) \bW_{\cdot k} = \bW (\mathbb{I}_{\cdot t^{(n)}} - \frac{1}{Z} \bq)$
#
# $$ \mathbf{\delta}^h = \bW \mathbf{\delta}^q$$
#
# ---
#
# Note that $\frac { \partial \bh_l}{ \partial \bV_{ ij } } = \bh_l (1-\bh_l) \frac{\partial \bV_{\cdot l}^T \bx + \ba_l}{\partial \bV_{ij}} = \bh_l (1-\bh_l) \frac{\partial \sum_k \bV_{kl} \bx_k + \ba_l}{\partial \bV_{ij}} = \begin{cases} \bh_l (1-\bh_l) \bx_k \hspace{5mm} \text{if} \hspace{2mm} j=l \hspace{2mm} \text{and} \hspace{2mm} k=i\\ 0 \hspace{5mm} \text{else}\end{cases} $
#
# $\frac{\partial \mathcal{L}^{(n)}}{\partial \bV_{ij}} = \left( \frac{\partial \mathcal{L}^{(n)}}{\partial \bh} \right)^T \frac{\partial \bh}{\partial \bV_{ij}} = {\mathbf{\delta}^h}^T \begin{bmatrix} \frac { \partial \bh_{ 1 } }{ \partial \bV_{ ij } } \\ \vdots \\ \frac { \partial \bh_L}{ \partial \bV_{ ij } } \end{bmatrix}= \sum_l \mathbf{\delta}_l^h \frac { \partial \bh_l}{ \partial \bV_{ ij } } = \mathbf{\delta}_j^h \bh_j (1-\bh_j) \bx_i$
#
#
# $$\nabla_{\bV} \mathcal{L}^{(n)} = \bx (\mathbf{\delta}^h \odot \bh \odot (\bj - \bh))^T$$
#
# ---
#
# Note that $\frac { \partial \bh_l}{ \partial \ba_{ j } } = \bh_l (1-\bh_l) \frac{\partial \bV_{\cdot l}^T \bx + \ba_l}{\partial \ba_j} = \begin{cases} \bh_l (1-\bh_l) \hspace{5mm} \text{if} \hspace{2mm} j=l\\ 0 \hspace{5mm} \text{if} \hspace{2mm} j \ne l\end{cases}$
#
# $\frac{\partial \mathcal{L}^{(n)}}{\partial \ba_j} = \left( \frac{\partial \mathcal{L}^{(n)}}{\partial \bh} \right)^T \frac{\partial \bh}{\partial \ba_j} = {\mathbf{\delta}^h}^T \begin{bmatrix} \frac { \partial \bh_{ 1 } }{ \partial \ba_{ j } } \\ \vdots \\ \frac { \partial \bh_L}{ \partial \ba_{ j } } \end{bmatrix}= \sum_l \mathbf{\delta}_l^h \frac { \partial \bh_l}{ \partial \ba_{ j } } = \mathbf{\delta}_j^h \bh_j (1-\bh_j)$
#
# $$\nabla_{\ba} \mathcal{L}^{(n)} = \mathbf{\delta}^h \odot \bh \odot (\bj - \bh) $$
#
# ---
# ## 2.2 MAP optimization (10 points)
#
# You derived equations for finding the _maximum likelihood_ solution of the parameters. Explain, in a few sentences, how you could extend this approach so that it optimizes towards a _maximum a posteriori_ (MAP) solution of the parameters, with a Gaussian prior on the parameters.
# #### Answer:
#
# Recall that $posterior = \frac{likelihood \times prior}{evidence} \rightarrow \log posterior = \log likelihood + \log prior - \log evidence.$
#
# Therefore, $\theta_{MAP} = \text{argmax}_{\theta} \log posterior = \text{argmax}_{\theta} \log likelihood + \log prior$. In this case it would be equivalent to finding the parameters that maximize a function $\mathcal{L} - \lambda \mathcal{R}$, where $\mathcal{L}$ corresponds to the log-likelihood (as before) and $-\lambda \mathcal{R}$ corresponds to the regularization term for a positive hyperparameter $\lambda$ associated with the parameters of the prior.
# ## 2.3. Implement and train a MLP (15 points)
#
# Implement a MLP model with a single hidden layer, and code to train the model.
# +
from scipy.special import expit
def mlp_gradient(x,t,W,b,V,a):
h = expit(np.dot(V.T,x) + a)
lq = np.dot(W.T,h) + b
Z = sum(np.exp(lq))
delta_q = - np.exp(lq)/Z
delta_q[t] += 1
delta_h = dot(W,delta_q)
grad_b = delta_q
grad_W = dot(h,delta_q.T)
grad_a = delta_h * h * (ones(shape(h)) - h)
grad_V = dot(x,grad_a.T)
return grad_W, grad_b, grad_V, grad_a
def mlp_sgd_iter(x_train, t_train, W, b, V, a, eta):
N, M = shape(x_train)
perm = np.random.permutation(N)
for ix in perm:
x = reshape(x_train[ix],(M,-1))
t = t_train[ix]
grad_W, grad_b, grad_V, grad_a = mlp_gradient(x, t, W, b, V, a)
W += eta * grad_W
b += eta * grad_b
V += eta * grad_V
a += eta * grad_a
return W, b, V, a
# +
def mlp_class(x,t,W,b,V,a):
h = expit(np.dot(V.T,x) + a)
lq = np.dot(W.T,h) + b
Z = sum(np.exp(lq))
offset = max(lq)
lZ = offset + log(sum(np.exp(lq - ones(shape(lq))*offset)))
lp = lq - ones(shape(lq))*log(Z)
return argmax(lp)
def mlp_pred_iter(x_set, t_set, W, b, V, a):
N_set, M = shape(x_set)
misclass = 0
for ix in range(N_set):
x = reshape(x_set[ix],(M,-1))
t = t_set[ix]
misclass += mlp_class(x,t,W,b,V,a) != t
return misclass*1.0/N_set
N_tr, M = shape(x_train)
N_vl, M = shape(x_test)
K = 10
L = 200
max_iter = 15
W_mlp = np.random.normal(0,0.2,(L, K))
b_mlp = np.random.normal(0,0.2,(K, 1))
V_mlp = 2*np.random.random_sample((M, L)) - 1
a_mlp = 2*np.random.random_sample((L, 1)) - 1
E_tr = []
for it in range(max_iter):
eta = 0.1
err = mlp_pred_iter(x_train, t_train, W_mlp, b_mlp, V_mlp, a_mlp)
print ('Iter: ' + str(it) + ' Error: '+ str(100*err) + '%')
E_tr.append(err)
W_mlp, b_mlp, V_mlp, a_mlp = mlp_sgd_iter(x_train, t_train, W_mlp, b_mlp, V_mlp, a_mlp, eta)
plot(range(1,max_iter+1), E_tr, label='Training')
xlabel('Iteration')
ylabel('Error')
legend(loc=1)
show()
# -
# ### 2.3.1. Less than 250 misclassifications on the test set (10 bonus points)
#
# You receive an additional 10 bonus points if you manage to train a model with very high accuracy: at most 2.5% misclasified digits on the test set. Note that the test set contains 10000 digits, so you model should misclassify at most 250 digits. This should be achievable with a MLP model with one hidden layer. See results of various models at : `http://yann.lecun.com/exdb/mnist/index.html`. To reach such a low accuracy, you probably need to have a very high $L$ (many hidden units), probably $L > 200$, and apply a strong Gaussian prior on the weights. In this case you are allowed to use the validation set for training.
# You are allowed to add additional layers, and use convolutional networks, although that is probably not required to reach 2.5% misclassifications.
err = mlp_pred_iter(x_test, t_test, W_mlp, b_mlp, V_mlp, a_mlp)
print ('Test error: '+ str(100*err)+'%')
| ml1_labs/lab2/.ipynb_checkpoints/lab02_mul_gallegoposada_menyah-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Recognize named entities on Twitter with LSTMs
#
# In this assignment, you will use a recurrent neural network to solve Named Entity Recognition (NER) problem. NER is a common task in natural language processing systems. It serves for extraction such entities from the text as persons, organizations, locations, etc. In this task you will experiment to recognize named entities from Twitter.
#
# For example, we want to extract persons' and organizations' names from the text. Than for the input text:
#
# <NAME> works for Google Brain
#
# a NER model needs to provide the following sequence of tags:
#
# B-PER I-PER O O B-ORG I-ORG
#
# Where *B-* and *I-* prefixes stand for the beginning and inside of the entity, while *O* stands for out of tag or no tag. Markup with the prefix scheme is called *BIO markup*. This markup is introduced for distinguishing of consequent entities with similar types.
#
# A solution of the task will be based on neural networks, particularly, on Bi-Directional Long Short-Term Memory Networks (Bi-LSTMs).
#
# ### Libraries
#
# For this task you will need the following libraries:
# - [Tensorflow](https://www.tensorflow.org) — an open-source software library for Machine Intelligence.
# - [Numpy](http://www.numpy.org) — a package for scientific computing.
#
# If you have never worked with Tensorflow, you would probably need to read some tutorials during your work on this assignment, e.g. [this one](https://www.tensorflow.org/tutorials/recurrent) could be a good starting point.
# + [markdown] deletable=true editable=true
# ### Data
#
# The following cell will download all data required for this assignment into the folder `week2/data`.
# + deletable=true editable=true
import sys
sys.path.append("..")
from common.download_utils import download_week2_resources
download_week2_resources()
# + [markdown] deletable=true editable=true
# ### Load the Twitter Named Entity Recognition corpus
#
# We will work with a corpus, which contains twits with NE tags. Every line of a file contains a pair of a token (word/punctuation symbol) and a tag, separated by a whitespace. Different tweets are separated by an empty line.
#
# The function *read_data* reads a corpus from the *file_path* and returns two lists: one with tokens and one with the corresponding tags. You need to complete this function by adding a code, which will replace a user's nickname to `<USR>` token and any URL to `<URL>` token. You could think that a URL and a nickname are just strings which start with *http://* or *https://* in case of URLs and a *@* symbol for nicknames.
# + deletable=true editable=true
def read_data(file_path):
tokens = []
tags = []
tweet_tokens = []
tweet_tags = []
for line in open(file_path, encoding='utf-8'):
line = line.strip()
if not line:
if tweet_tokens:
tokens.append(tweet_tokens)
tags.append(tweet_tags)
tweet_tokens = []
tweet_tags = []
else:
token, tag = line.split()
# Replace all urls with <URL> token
# Replace all users with <USR> token
token = '<URL>' if token.startswith("http") or token.startswith("https") else token
token = '<USR>' if token.startswith("@") else token
tag = '<URL>' if tag.startswith("http") or tag.startswith("https") else tag
tag = '<USR>' if tag.startswith("@") else tag
tweet_tokens.append(token)
tweet_tags.append(tag)
return tokens, tags
# + [markdown] deletable=true editable=true
# And now we can load three separate parts of the dataset:
# - *train* data for training the model;
# - *validation* data for evaluation and hyperparameters tuning;
# - *test* data for final evaluation of the model.
# + deletable=true editable=true
train_tokens, train_tags = read_data('data/train.txt')
validation_tokens, validation_tags = read_data('data/validation.txt')
test_tokens, test_tags = read_data('data/test.txt')
# + [markdown] deletable=true editable=true
# You should always understand what kind of data you deal with. For this purpose, you can print the data running the following cell:
# + deletable=true editable=true
for i in range(3):
for token, tag in zip(train_tokens[i], train_tags[i]):
print('%s\t%s' % (token, tag))
print()
# + [markdown] deletable=true editable=true
# ### Prepare dictionaries
#
# To train a neural network, we will use two mappings:
# - {token}$\to${token id}: address the row in embeddings matrix for the current token;
# - {tag}$\to${tag id}: one-hot ground truth probability distribution vectors for computing the loss at the output of the network.
#
# Now you need to implement the function *build_dict* which will return {token or tag}$\to${index} and vice versa.
# + deletable=true editable=true
from collections import defaultdict
# + deletable=true editable=true
def build_dict(tokens_or_tags, special_tokens):
"""
tokens_or_tags: a list of lists of tokens or tags
special_tokens: some special tokens
"""
# Create a dictionary with default value 0
tok2idx = defaultdict(lambda: 0)
idx2tok = []
index = 0
# Create mappings from tokens to indices and vice versa
# Add special tokens to dictionaries
# The first special token must have index 0
for s in special_tokens:
tok2idx[s] = index
idx2tok.insert(index, s)
index += 1
for li in tokens_or_tags:
for t in li:
if t not in tok2idx:
tok2idx[t] = index
idx2tok.insert(index, t)
index += 1
return tok2idx, idx2tok
# + [markdown] deletable=true editable=true
# After implementing the function *build_dict* you can make dictionaries for tokens and tags. Special tokens in our case will be:
# - `<UNK>` token for out of vocabulary tokens;
# - `<PAD>` token for padding sentence to the same length when we create batches of sentences.
# + deletable=true editable=true
special_tokens = ['<UNK>', '<PAD>']
special_tags = ['O']
# Create dictionaries
token2idx, idx2token = build_dict(train_tokens + validation_tokens, special_tokens)
tag2idx, idx2tag = build_dict(train_tags, special_tags)
print (idx2token[0])
print (token2idx['<UNK>'])
# + [markdown] deletable=true editable=true
# The next additional functions will help you to create the mapping between tokens and ids for a sentence.
# + deletable=true editable=true
def words2idxs(tokens_list):
return [token2idx[word] for word in tokens_list]
def tags2idxs(tags_list):
return [tag2idx[tag] for tag in tags_list]
def idxs2words(idxs):
return [idx2token[idx] for idx in idxs]
def idxs2tags(idxs):
return [idx2tag[idx] for idx in idxs]
# + [markdown] deletable=true editable=true
# ### Generate batches
#
# Neural Networks are usually trained with batches. It means that weight updates of the network are based on several sequences at every single time. The tricky part is that all sequences within a batch need to have the same length. So we will pad them with a special `<PAD>` token. It is also a good practice to provide RNN with sequence lengths, so it can skip computations for padding parts. We provide the batching function *batches_generator* readily available for you to save time.
# + deletable=true editable=true
def batches_generator(batch_size, tokens, tags,
shuffle=True, allow_smaller_last_batch=True):
"""Generates padded batches of tokens and tags."""
n_samples = len(tokens)
if shuffle:
order = np.random.permutation(n_samples)
else:
order = np.arange(n_samples)
n_batches = n_samples // batch_size
if allow_smaller_last_batch and n_samples % batch_size:
n_batches += 1
for k in range(n_batches):
batch_start = k * batch_size
batch_end = min((k + 1) * batch_size, n_samples)
current_batch_size = batch_end - batch_start
x_list = []
y_list = []
max_len_token = 0
for idx in order[batch_start: batch_end]:
x_list.append(words2idxs(tokens[idx]))
y_list.append(tags2idxs(tags[idx]))
max_len_token = max(max_len_token, len(tags[idx]))
# Fill in the data into numpy nd-arrays filled with padding indices.
x = np.ones([current_batch_size, max_len_token], dtype=np.int32) * token2idx['<PAD>']
y = np.ones([current_batch_size, max_len_token], dtype=np.int32) * tag2idx['O']
lengths = np.zeros(current_batch_size, dtype=np.int32)
for n in range(current_batch_size):
utt_len = len(x_list[n])
x[n, :utt_len] = x_list[n]
lengths[n] = utt_len
y[n, :utt_len] = y_list[n]
yield x, y, lengths
# + [markdown] deletable=true editable=true
# ## Build a recurrent neural network
#
# This is the most important part of the assignment. Here we will specify the network architecture based on TensorFlow building blocks. It's fun and easy as a lego constructor! We will create an LSTM network which will produce probability distribution over tags for each token in a sentence. To take into account both right and left contexts of the token, we will use Bi-Directional LSTM (Bi-LSTM). Dense layer will be used on top to perform tag classification.
# + deletable=true editable=true
import tensorflow as tf
import numpy as np
# + deletable=true editable=true
class BiLSTMModel():
pass
# + [markdown] deletable=true editable=true
# First, we need to create [placeholders](https://www.tensorflow.org/versions/r0.12/api_docs/python/io_ops/placeholders) to specify what data we are going to feed into the network during the execution time. For this task we will need the following placeholders:
# - *input_batch* — sequences of words (the shape equals to [batch_size, sequence_len]);
# - *ground_truth_tags* — sequences of tags (the shape equals to [batch_size, sequence_len]);
# - *lengths* — lengths of not padded sequences (the shape equals to [batch_size]);
# - *dropout_ph* — dropout keep probability; this placeholder has a predefined value 1;
# - *learning_rate_ph* — learning rate; we need this placeholder because we want to change the value during training.
#
# It could be noticed that we use *None* in the shapes in the declaration, which means that data of any size can be feeded.
#
# You need to complete the function *declare_placeholders*.
# + deletable=true editable=true
def declare_placeholders(self):
"""Specifies placeholders for the model."""
# Placeholders for input and ground truth output.
self.input_batch = tf.placeholder(dtype=tf.int32, shape=[None, None], name='input_batch')
self.ground_truth_tags = tf.placeholder(dtype=tf.int32, shape = [None, None], name='ground_truth_tags')
# Placeholder for lengths of the sequences.
self.lengths = tf.placeholder(dtype=tf.int32, shape=[None], name='lengths')
# Placeholder for a dropout keep probability. If we don't feed
# a value for this placeholder, it will be equal to 1.0.
self.dropout_ph = tf.placeholder_with_default(tf.cast(1.0, tf.float64), shape=[], name='dropout_ph')
# Placeholder for a learning rate (float32).
self.learning_rate_ph = tf.placeholder(dtype=tf.float32, shape=[], name='learning_rate_ph')
# + deletable=true editable=true
BiLSTMModel.__declare_placeholders = classmethod(declare_placeholders)
# + [markdown] deletable=true editable=true
# Now, let us specify the layers of the neural network. First, we need to perform some preparatory steps:
#
# - Create embeddings matrix with [tf.Variable](https://www.tensorflow.org/api_docs/python/tf/Variable). Specify its name (*embeddings_matrix*), type (*tf.float32*), and initialize with random values.
# - Create forward and backward LSTM cells. TensorFlow provides a number of [RNN cells](https://www.tensorflow.org/api_guides/python/contrib.rnn#Core_RNN_Cells_for_use_with_TensorFlow_s_core_RNN_methods) ready for you. We suggest that you use *BasicLSTMCell*, but you can also experiment with other types, e.g. GRU cells. [This](http://colah.github.io/posts/2015-08-Understanding-LSTMs/) blogpost could be interesting if you want to learn more about the differences.
# - Wrap your cells with [DropoutWrapper](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn/DropoutWrapper). Dropout is an important regularization technique for neural networks. Specify all keep probabilities using the dropout placeholder that we created before.
#
# After that, you can build the computation graph that transforms an input_batch:
#
# - [Look up](https://www.tensorflow.org/api_docs/python/tf/nn/embedding_lookup) embeddings for an *input_batch* in the prepared *embedding_matrix*.
# - Pass the embeddings through [Bidirectional Dynamic RNN](https://www.tensorflow.org/api_docs/python/tf/nn/bidirectional_dynamic_rnn) with the specified forward and backward cells. Use the lengths placeholder here to avoid computations for padding tokens inside the RNN.
# - Create a dense layer on top. Its output will be used directly in loss function.
#
# Fill in the code below. In case you need to debug something, the easiest way is to check that tensor shapes of each step match the expected ones.
#
# + deletable=true editable=true
def build_layers(self, vocabulary_size, embedding_dim, n_hidden_rnn, n_tags):
"""Specifies bi-LSTM architecture and computes logits for inputs."""
batch_size = self.input_batch.shape[0]
# Create embedding variable (tf.Variable).
initial_embedding_matrix = np.random.randn(vocabulary_size, embedding_dim) / np.sqrt(embedding_dim)
embedding_matrix_variable = tf.Variable(initial_value=initial_embedding_matrix, \
name="embedding_matrix_variable")
# Create RNN cells (for example, tf.nn.rnn_cell.BasicLSTMCell) with n_hidden_rnn number of units
# and dropout (tf.nn.rnn_cell.DropoutWrapper), initializing all *_keep_prob with dropout placeholder.
forward_cell = tf.nn.rnn_cell.DropoutWrapper(tf.nn.rnn_cell.BasicLSTMCell(n_hidden_rnn), \
input_keep_prob=self.dropout_ph, \
output_keep_prob=self.dropout_ph, \
state_keep_prob=self.dropout_ph)
backward_cell = tf.nn.rnn_cell.DropoutWrapper(tf.nn.rnn_cell.BasicLSTMCell(n_hidden_rnn), \
input_keep_prob=self.dropout_ph, \
output_keep_prob=self.dropout_ph, \
state_keep_prob=self.dropout_ph)
# Look up embeddings for self.input_batch (tf.nn.embedding_lookup).
# Shape: [batch_size, sequence_len, embedding_dim].
embeddings = tf.nn.embedding_lookup(params=embedding_matrix_variable, \
ids=self.input_batch)
# Pass them through Bidirectional Dynamic RNN (tf.nn.bidirectional_dynamic_rnn).
# Shape: [batch_size, sequence_len, 2 * n_hidden_rnn].
(rnn_output_fw, rnn_output_bw), _ = tf.nn.bidirectional_dynamic_rnn( \
forward_cell, \
backward_cell, \
embeddings, \
sequence_length=self.lengths, \
dtype=tf.float64)
rnn_output = tf.concat([rnn_output_fw, rnn_output_bw], axis=2)
# Dense layer on top.
# Shape: [batch_size, sequence_len, n_tags].
self.logits = tf.layers.dense(rnn_output, n_tags, activation=None)
# + deletable=true editable=true
BiLSTMModel.__build_layers = classmethod(build_layers)
# + [markdown] deletable=true editable=true
# To compute the actual predictions of the neural network, you need to apply [softmax](https://www.tensorflow.org/api_docs/python/tf/nn/softmax) to the last layer and find the most probable tags with [argmax](https://www.tensorflow.org/api_docs/python/tf/argmax).
# + deletable=true editable=true
def compute_predictions(self):
"""Transforms logits to probabilities and finds the most probable tags."""
# Create softmax (tf.nn.softmax) function
softmax_output = tf.nn.softmax(logits=self.logits)
# Use argmax (tf.argmax) to get the most probable tags
self.predictions = tf.argmax(softmax_output, axis=-1)
# + deletable=true editable=true
BiLSTMModel.__compute_predictions = classmethod(compute_predictions)
# + [markdown] deletable=true editable=true
# During training we do not need predictions of the network, but we need a loss function. We will use [cross-entropy loss](http://ml-cheatsheet.readthedocs.io/en/latest/loss_functions.html#cross-entropy), efficiently implemented in TF as
# [cross entropy with logits](https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits). Note that it should be applied to logits of the model (not to softmax probabilities!). Also note, that we do not want to take into account loss terms coming from `<PAD>` tokens. So we need to mask them out, before computing [mean](https://www.tensorflow.org/api_docs/python/tf/reduce_mean).
# + deletable=true editable=true
def compute_loss(self, n_tags, PAD_index):
"""Computes masked cross-entopy loss with logits."""
# Create cross entropy function function (tf.nn.softmax_cross_entropy_with_logits)
ground_truth_tags_one_hot = tf.one_hot(self.ground_truth_tags, n_tags)
loss_tensor = tf.nn.softmax_cross_entropy_with_logits(labels=ground_truth_tags_one_hot, \
logits=self.logits)
# Create loss function which doesn't operate with <PAD> tokens (tf.reduce_mean)
mask = tf.cast(tf.not_equal(loss_tensor, PAD_index), tf.float64)
# mask = tf.not_equal(loss_tensor, PAD_index)
self.loss = tf.reduce_mean(input_tensor= tf.multiply(loss_tensor, mask))
# + deletable=true editable=true
BiLSTMModel.__compute_loss = classmethod(compute_loss)
# + [markdown] deletable=true editable=true
# The last thing to specify is how we want to optimize the loss.
# We suggest that you use [Adam](https://www.tensorflow.org/api_docs/python/tf/train/AdamOptimizer) optimizer with a learning rate from the corresponding placeholder.
# You will also need to apply [clipping](https://www.tensorflow.org/versions/r0.12/api_docs/python/train/gradient_clipping) to eliminate exploding gradients. It can be easily done with [clip_by_norm](https://www.tensorflow.org/api_docs/python/tf/clip_by_norm) function.
# + deletable=true editable=true
def perform_optimization(self):
"""Specifies the optimizer and train_op for the model."""
# Create an optimizer (tf.train.AdamOptimizer)
self.optimizer = tf.train.AdamOptimizer(learning_rate=self.learning_rate_ph)
self.grads_and_vars = self.optimizer.compute_gradients(self.loss)
# Gradient clipping (tf.clip_by_norm) for self.grads_and_vars
clip_norm = 1.0
self.grads_and_vars = [(tf.clip_by_norm(grad, clip_norm), var) for grad, var in self.grads_and_vars]
self.train_op = self.optimizer.apply_gradients(self.grads_and_vars)
# + deletable=true editable=true
BiLSTMModel.__perform_optimization = classmethod(perform_optimization)
# + [markdown] deletable=true editable=true
# Congratulations! You have specified all the parts of your network. You may have noticed, that we didn't deal with any real data yet, so what you have written is just recipes on how the network should function.
# Now we will put them to the constructor of our Bi-LSTM class to use it in the next section.
# + deletable=true editable=true
def init_model(self, vocabulary_size, n_tags, embedding_dim, n_hidden_rnn, PAD_index):
self.__declare_placeholders()
self.__build_layers(vocabulary_size, embedding_dim, n_hidden_rnn, n_tags)
self.__compute_predictions()
self.__compute_loss(n_tags, PAD_index)
self.__perform_optimization()
# + deletable=true editable=true
BiLSTMModel.__init__ = classmethod(init_model)
# + [markdown] deletable=true editable=true
# ## Train the network and predict tags
# + [markdown] deletable=true editable=true
# [Session.run](https://www.tensorflow.org/api_docs/python/tf/Session#run) is a point which initiates computations in the graph that we have defined. To train the network, we need to compute *self.train_op*, which was declared in *perform_optimization*. To predict tags, we just need to compute *self.predictions*. Anyway, we need to feed actual data through the placeholders that we defined before.
# + deletable=true editable=true
def train_on_batch(self, session, x_batch, y_batch, lengths, learning_rate, dropout_keep_probability):
feed_dict = {self.input_batch: x_batch,
self.ground_truth_tags: y_batch,
self.learning_rate_ph: learning_rate,
self.dropout_ph: dropout_keep_probability,
self.lengths: lengths}
session.run(self.train_op, feed_dict=feed_dict)
# + deletable=true editable=true
BiLSTMModel.train_on_batch = classmethod(train_on_batch)
# + [markdown] deletable=true editable=true
# Implement the function *predict_for_batch* by initializing *feed_dict* with input *x_batch* and *lengths* and running the *session* for *self.predictions*.
# + deletable=true editable=true
def predict_for_batch(self, session, x_batch, lengths):
feed_dict = {
self.input_batch: x_batch,
self.lengths: lengths
}
predictions = session.run(self.predictions, feed_dict=feed_dict)
return predictions
# + deletable=true editable=true
BiLSTMModel.predict_for_batch = classmethod(predict_for_batch)
# + [markdown] deletable=true editable=true
# We finished with necessary methods of our BiLSTMModel model and almost ready to start experimenting.
#
# ### Evaluation
# To simplify the evaluation process we provide two functions for you:
# - *predict_tags*: uses a model to get predictions and transforms indices to tokens and tags;
# - *eval_conll*: calculates precision, recall and F1 for the results.
# + deletable=true editable=true
from evaluation import precision_recall_f1
# + deletable=true editable=true
def predict_tags(model, session, token_idxs_batch, lengths):
"""Performs predictions and transforms indices to tokens and tags."""
tag_idxs_batch = model.predict_for_batch(session, token_idxs_batch, lengths)
tags_batch, tokens_batch = [], []
for tag_idxs, token_idxs in zip(tag_idxs_batch, token_idxs_batch):
tags, tokens = [], []
for tag_idx, token_idx in zip(tag_idxs, token_idxs):
if token_idx != token2idx['<PAD>']:
tags.append(idx2tag[tag_idx])
tokens.append(idx2token[token_idx])
tags_batch.append(tags)
tokens_batch.append(tokens)
return tags_batch, tokens_batch
def eval_conll(model, session, tokens, tags, short_report=True):
"""Computes NER quality measures using CONLL shared task script."""
y_true, y_pred = [], []
for x_batch, y_batch, lengths in batches_generator(1, tokens, tags):
tags_batch, tokens_batch = predict_tags(model, session, x_batch, lengths)
if len(x_batch[0]) != len(tags_batch[0]):
raise Exception("Incorrect length of prediction for the input, "
"expected length: %i, got: %i" % (len(x_batch[0]), len(tags_batch[0])))
ground_truth_tags = [idx2tag[tag_idx] for tag_idx in y_batch[0]]
# We extend every prediction and ground truth sequence with 'O' tag
# to indicate a possible end of entity.
y_true.extend(ground_truth_tags + ['O'])
y_pred.extend(tags_batch[0] + ['O'])
results = precision_recall_f1(y_true, y_pred, print_results=True, short_report=short_report)
return results
# + [markdown] deletable=true editable=true
# ## Run your experiment
# + [markdown] deletable=true editable=true
# Create *BiLSTMModel* model with the following parameters:
# - *vocabulary_size* — number of tokens;
# - *n_tags* — number of tags;
# - *embedding_dim* — dimension of embeddings, recommended value: 200;
# - *n_hidden_rnn* — size of hidden layers for RNN, recommended value: 200;
# - *PAD_index* — an index of the padding token (`<PAD>`).
#
# Set hyperparameters. You might want to start with the following recommended values:
# - *batch_size*: 32;
# - 4 epochs;
# - starting value of *learning_rate*: 0.005
# - *learning_rate_decay*: a square root of 2;
# - *dropout_keep_probability*: try several values: 0.1, 0.5, 0.9.
#
# However, feel free to conduct more experiments to tune hyperparameters and earn extra points for the assignment.
# + deletable=true editable=true
tf.reset_default_graph()
model = BiLSTMModel( \
vocabulary_size=len(token2idx), \
n_tags=len(tag2idx), \
embedding_dim=200, \
n_hidden_rnn=300, \
PAD_index=token2idx['<PAD>'])
batch_size = 32
n_epochs = 4
learning_rate = 0.005
learning_rate_decay = np.sqrt(2.0)
dropout_keep_probability = float(0.9)
# + deletable=true editable=true
sess = tf.Session()
sess.run(tf.global_variables_initializer())
print('Start training... \n')
for epoch in range(n_epochs):
# For each epoch evaluate the model on train and validation data
print('-' * 20 + ' Epoch {} '.format(epoch+1) + 'of {} '.format(n_epochs) + '-' * 20)
print('Train data evaluation:')
eval_conll(model, sess, train_tokens, train_tags, short_report=True)
print('Validation data evaluation:')
eval_conll(model, sess, validation_tokens, validation_tags, short_report=True)
# Train the model
print ('Learning Rate: ', learning_rate)
for x_batch, y_batch, lengths in batches_generator(batch_size, train_tokens, train_tags):
model.train_on_batch(sess, x_batch, y_batch, lengths, learning_rate, dropout_keep_probability)
# Decaying the learning rate
learning_rate = learning_rate / learning_rate_decay
print('...training finished.')
# + deletable=true editable=true
tf.reset_default_graph()
model = BiLSTMModel( \
vocabulary_size=len(token2idx), \
n_tags=len(tag2idx), \
embedding_dim=200, \
n_hidden_rnn=300, \
PAD_index=token2idx['<PAD>'])
batch_size = 32
n_epochs = 10
learning_rate = 0.01
learning_rate_decay = np.sqrt(2.0)
dropout_keep_probability = float(0.9)
# + [markdown] deletable=true editable=true
# Finally, we are ready to run the training!
# + deletable=true editable=true
sess = tf.Session()
sess.run(tf.global_variables_initializer())
print('Start training... \n')
for epoch in range(n_epochs):
# For each epoch evaluate the model on train and validation data
print('-' * 20 + ' Epoch {} '.format(epoch+1) + 'of {} '.format(n_epochs) + '-' * 20)
print('Train data evaluation:')
eval_conll(model, sess, train_tokens, train_tags, short_report=True)
print('Validation data evaluation:')
eval_conll(model, sess, validation_tokens, validation_tags, short_report=True)
# Train the model
print ('Learning Rate: ', learning_rate)
for x_batch, y_batch, lengths in batches_generator(batch_size, train_tokens, train_tags):
model.train_on_batch(sess, x_batch, y_batch, lengths, learning_rate, dropout_keep_probability)
# Decaying the learning rate
learning_rate = learning_rate / learning_rate_decay
print('...training finished.')
# + [markdown] deletable=true editable=true
# Now let us see full quality reports for the final model on train, validation, and test sets. To give you a hint whether you have implemented everything correctly, you might expect F-score about 40% on the validation set.
#
# **The output of the cell below (as well as the output of all the other cells) should be present in the notebook for peer2peer review!**
# + deletable=true editable=true
print('-' * 20 + ' Train set quality: ' + '-' * 20)
train_results = eval_conll(model, sess, train_tokens, train_tags, short_report=False)
print('-' * 20 + ' Validation set quality: ' + '-' * 20)
validation_results = eval_conll(model, sess, validation_tokens, validation_tags, short_report=False)
print('-' * 20 + ' Test set quality: ' + '-' * 20)
test_results = eval_conll(model, sess, test_tokens, test_tags, short_report=False)
# + [markdown] deletable=true editable=true
# ### Conclusions
#
# Could we say that our model is state of the art and the results are acceptable for the task? Definately, we can say so. Nowadays, Bi-LSTM is one of the state of the art approaches for solving NER problem and it outperforms other classical methods. Despite the fact that we used small training corpora (in comparison with usual sizes of corpora in Deep Learning), our results are quite good. In addition, in this task there are many possible named entities and for some of them we have only several dozens of trainig examples, which is definately small. However, the implemented model outperforms classical CRFs for this task. Even better results could be obtained by some combinations of several types of methods, e.g. see [this](https://arxiv.org/abs/1603.01354) paper if you are interested.
| LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Meta Data Exploration
#
# <NAME> -- <EMAIL>
#
# This notebook gives an overview of the buildings in this study
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import os
# %matplotlib inline
repos_path = "/Users/Clayton/temporal-features-for-buildings/"
meta = pd.read_csv(os.path.join(repos_path,"data/raw/meta_open.csv"), index_col='uid', parse_dates=["datastart","dataend"], dayfirst=True)
meta.info()
sns.set_style("whitegrid")
# # Create Bar Charts
#
# Use: http://stanford.edu/~mwaskom/software/seaborn/examples/horizontal_barplot.html
meta.info()#[['industry','timezone']].head()
meta.head()
df = pd.pivot_table(meta.reset_index(), index='timezone', columns='subindustry', values='uid', aggfunc='count')
df
# +
# crashes = sns.load_dataset("car_crashes").sort_values("total", ascending=False)
location = pd.DataFrame(meta.timezone.value_counts()).reset_index()
location
# -
def createbarchart(df, column, ylabel, color, filelabel):
# Initialize the matplotlib figure
#sns.set_context('poster', font_scale=1)
location = pd.DataFrame(df[column].value_counts()).reset_index()
f, ax = plt.subplots(figsize=(9, 5))
sns.set_color_codes("pastel")
sns.barplot(x=column, y="index", data=location[:15], color=color)
# Add a legend and informative axis label
ax.legend(ncol=2, loc="lower right", frameon=True)
ax.set(ylabel=ylabel,
xlabel="Number of Buildings")
sns.despine(left=True, bottom=True)
plt.subplots_adjust(left=0.3)
plt.tight_layout()
plt.savefig(os.path.join(repos_path,"reports/figures/metadataoverview/"+filelabel+".png"))
location[:15]
meta["string_starttime"] = meta.datastart.apply(lambda x: str(x.date()))
sns.set_context("paper", font_scale=2)
createbarchart(meta, "string_starttime", "Data Starting Time", "m","starttimesbar")
#sns.set_style("white")
createbarchart(meta, "timezone", "Time Zones", "b","timezonesbar")
createbarchart(meta, "industry", "Industry", "r", "bar_industry")
createbarchart(meta, "subindustry", "Sub-Industry", "g","bar_subindustry")
createbarchart(meta, "primaryspaceusage", "Primary Use", "y", "bar_primaryspaceuse")
# # Create 4 bar charts in a set of panels
import numpy as np
# +
# Simple data to display in various forms
x = np.linspace(0, 2 * np.pi, 400)
y = np.sin(x ** 2)
# row and column sharing
f, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, sharex='col', sharey='row')
ax1.plot(x, y)
ax1.set_title('Sharing x per column, y per row')
ax2.scatter(x, y)
ax3.scatter(x, 2 * y ** 2 - 1, color='r')
ax4.plot(x, 2 * y ** 2 - 1, color='r')
# -
sns.set_color_codes("pastel")
sns.set_context(font_scale=1)
import matplotlib.gridspec as gridspec
# +
plt.figure(figsize=(18,10))
gs = gridspec.GridSpec(2, 8)
#f, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2,2, figsize=(20, 9))#
# ax = plt.subplot2grid((2,2),(0, 0))
location = pd.DataFrame(meta["timezone"].value_counts()).reset_index()
ax1 = plt.subplot(gs[0,0:4])
ax1 = sns.barplot(x="timezone", y="index", data=location[:15], color="b")
ax1.legend(ncol=2, loc="lower right", frameon=True)
ax1.set(ylabel="Time Zones", xlabel="Number of Buildings")
location = pd.DataFrame(meta["industry"].value_counts()).reset_index()
ax2 = plt.subplot(gs[0,4:])
ax2 = sns.barplot(x="industry", y="index", data=location[:15], color="g")
ax2.legend(ncol=2, loc="lower right", frameon=True)
ax2.set(ylabel="Industry", xlabel="Number of Buildings")
location = pd.DataFrame(meta["subindustry"].value_counts()).reset_index()
ax3 = plt.subplot(gs[1,0:4])
ax3 = sns.barplot(x="subindustry", y="index", data=location[:15], color="r")
ax3.legend(ncol=2, loc="lower right", frameon=True)
ax3.set(ylabel="Sub-Industry", xlabel="Number of Buildings")
location = pd.DataFrame(meta["primaryspaceusage"].value_counts()).reset_index()
ax4 = plt.subplot(gs[1,4:])
ax4 = sns.barplot(x="primaryspaceusage", y="index", data=location[:15], color="y")
ax4.legend(ncol=2, loc="lower right", frameon=True)
ax4.set(ylabel="Primary Use", xlabel="Number of Buildings")
sns.despine(left=True, bottom=True)
plt.subplots_adjust(left=0.3)
plt.tight_layout()
plt.savefig(os.path.join(repos_path,"reports/figures/metadataoverview/allbars.pdf"))
# -
| notebooks/00_Meta Data Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SYS 611: Flip Flop (Discrete Time Simulation)
#
# <NAME> <<EMAIL>>
#
# This example shows how to construct a discrete time simulation of a simple flip flop (a digital component with one bit of memory). The flip flop system has one binary state variable (q), one binary input variable (x), and one binary output variable (y). The output function is defined by the logical operation y(t)=q(t). The state transition function stores the current input value in the state variable q(t+1)=x(t). The expected behavior is to output the input variables but offset by one time step.
#
# ## Dependencies
#
# This example is compatible with Python 2 environments through use of the `__future__` library function. Additionally, this example uses the `matplotlib.pyplot` library for plotting.
# +
# import the python3 behavior for importing, division, and printing in python2
from __future__ import absolute_import, division, print_function
# import the matplotlib pyplot package and refer to it as `plt`
# see http://matplotlib.org/api/pyplot_api.html for documentation
import matplotlib.pyplot as plt
# -
# ## Transition and Output Functions
#
# Define functions for the state transition function (delta) and output function (lambda). Note that `lambda` is a reserved word in Python (for lambda functions), so both Greek letters are prefixed by an underscore.
# +
# define the state transition function
def _delta(q, x):
# note that xor is equivalent to != in Python
return x
# define the output function (a Moore machine only allows the output to be a function of the state)
def _lambda(q):
return q
# -
# ## Input, State, and Output Trajectories
#
# Define lists to store the input trajectory and store the computed output and state trajectories (initialize with zero value).
# +
# define the input trajectory
x = [1,1,0,0,1,0,0,0,1]
# define the output and state trajectories (zero initial value)
y = [0,0,0,0,0,0,0,0,0]
q = [0,0,0,0,0,0,0,0,0,0]
# -
# ## Discrete Time Simulation Logic
#
# The discrete event simulation should start by setting the initial time and state. Then, enter a loop to:
# 1. Compute the current output
# 2. Compute the next state
# 3. Update the time
# +
# initialize the simulation
t = 0
q[0] = 0
# execute the simulation
while t <= 8:
# record output value
y[t] = _lambda(q[t])
# record state update
q[t+1] = _delta(q[t], x[t])
# advance time
t += 1
# -
# ## Visualize Outcomes
#
# Use bar plots in `matplotlib` to plot the input, state, and output trajectories.
plt.figure()
# create three subplots that all share a common x-axis
f, (ax1, ax2, ax3) = plt.subplots(3, sharex=True)
# plot the input trajectory on the first subplot
ax1.bar(range(9), x, color='k')
ax1.set_ylabel('Input ($x$)')
# plot the state trajectory on the second subplot (up to second-to-last value)
ax2.bar(range(9), q[:-1], color='k')
ax2.set_ylabel('State ($q$)')
# plot the output trajectory on the third subplot
ax3.bar(range(9), y, color='k')
ax3.set_ylabel('Output ($y$)')
# add labels and display
plt.xlabel('Time (ticks)')
plt.suptitle('Flip Flop Model')
plt.show()
| week7/FlipFlop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="KkwXnw9OfHZl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4b4d6113-cf8d-485f-aebd-dfea3c4879bd"
# %tensorflow_version 1.x
#Suppress warnings which keep poping up
import warnings
warnings.filterwarnings("ignore")
from keras import backend as K
import time
import matplotlib.pyplot as plt
import numpy as np
% matplotlib inline
np.random.seed(2017)
from keras.models import Sequential
from keras.layers.convolutional import Convolution2D, MaxPooling2D, SeparableConvolution2D
from keras.layers import Activation, Flatten, Dense, Dropout, AveragePooling2D
from keras.layers.normalization import BatchNormalization
from keras.utils import np_utils
# + id="NHpnoCHZfO8g" colab_type="code" colab={}
from keras.datasets import cifar10
(train_features, train_labels), (test_features, test_labels) = cifar10.load_data()
num_train, img_channels, img_rows, img_cols = train_features.shape
num_test, _, _, _ = test_features.shape
num_classes = len(np.unique(train_labels))
# + id="yTRtppAZUZ8a" colab_type="code" outputId="1bb83115-1632-4c64-ea6d-ca39e51b8643" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(train_features.shape)
# + id="14HyBUXdfS6G" colab_type="code" outputId="cb3f1945-b5ca-405e-f535-e6c5f4ef1268" colab={"base_uri": "https://localhost:8080/", "height": 213}
class_names = ['airplane','automobile','bird','cat','deer',
'dog','frog','horse','ship','truck']
fig = plt.figure(figsize=(8,3))
for i in range(num_classes):
ax = fig.add_subplot(2, 5, 1 + i, xticks=[], yticks=[])
idx = np.where(train_labels[:]==i)[0]
features_idx = train_features[idx,::]
img_num = np.random.randint(features_idx.shape[0])
im = features_idx[img_num]
ax.set_title(class_names[i])
plt.imshow(im)
plt.show()
# + id="bmfsk76-fadV" colab_type="code" colab={}
def plot_model_history(model_history):
fig, axs = plt.subplots(1,2,figsize=(15,5))
# summarize history for accuracy
axs[0].plot(range(1,len(model_history.history['acc'])+1),model_history.history['acc'])
axs[0].plot(range(1,len(model_history.history['val_acc'])+1),model_history.history['val_acc'])
axs[0].set_title('Model Accuracy')
axs[0].set_ylabel('Accuracy')
axs[0].set_xlabel('Epoch')
axs[0].set_xticks(np.arange(1,len(model_history.history['acc'])+1),len(model_history.history['acc'])/10)
axs[0].legend(['train', 'val'], loc='best')
# summarize history for loss
axs[1].plot(range(1,len(model_history.history['loss'])+1),model_history.history['loss'])
axs[1].plot(range(1,len(model_history.history['val_loss'])+1),model_history.history['val_loss'])
axs[1].set_title('Model Loss')
axs[1].set_ylabel('Loss')
axs[1].set_xlabel('Epoch')
axs[1].set_xticks(np.arange(1,len(model_history.history['loss'])+1),len(model_history.history['loss'])/10)
axs[1].legend(['train', 'val'], loc='best')
plt.show()
# + id="YJMT4rjgfdZz" colab_type="code" colab={}
def accuracy(test_x, test_y, model):
result = model.predict(test_x)
predicted_class = np.argmax(result, axis=1)
true_class = np.argmax(test_y, axis=1)
num_correct = np.sum(predicted_class == true_class)
accuracy = float(num_correct)/result.shape[0]
return (accuracy * 100)
# + id="T5c5nDvxm6zR" colab_type="code" colab={}
train_features = train_features.astype('float32')/255
test_features = test_features.astype('float32')/255
# convert class labels to binary class labels
train_labels = np_utils.to_categorical(train_labels, num_classes)
test_labels = np_utils.to_categorical(test_labels, num_classes)
# + id="cSOb2lkJfhVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="1ab35e42-9b58-4e9b-925d-7e32941aa169"
# Define the model
model = Sequential()
model.add(Convolution2D(48, 3, 3, border_mode='same', input_shape=(32, 32, 3)))
model.add(Activation('relu'))
model.add(Convolution2D(48, 3, 3))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Convolution2D(96, 3, 3, border_mode='same'))
model.add(Activation('relu'))
model.add(Convolution2D(96, 3, 3))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Convolution2D(192, 3, 3, border_mode='same'))
model.add(Activation('relu'))
model.add(Convolution2D(192, 3, 3))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
# Compile the model
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# + id="wvVs5tC8mR6Q" colab_type="code" outputId="f02bffc4-1c07-4c7a-c883-c86f0679d3d5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.summary()
# + id="-SRp2N_-kA6S" colab_type="code" colab={}
from keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(zoom_range=0.0,
horizontal_flip=False)
# + id="5oK_V7WyfsXX" colab_type="code" outputId="5a2ad01e-0be5-43ce-e5af-c5ddd6b4ac1d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# train the model
start = time.time()
# Train the model
model_info = model.fit_generator(datagen.flow(train_features, train_labels, batch_size = 128),
samples_per_epoch = train_features.shape[0], nb_epoch = 50,
validation_data = (test_features, test_labels), verbose=1)
end = time.time()
print ("Model took %0.2f seconds to train"%(end - start))
# plot model history
plot_model_history(model_info)
# compute test accuracy
print ("Accuracy on test data is: %0.2f"%accuracy(test_features, test_labels, model))
# + id="PUR0XaLPidbZ" colab_type="code" colab={}
# Define the model
my_model = Sequential()
my_model.add(SeparableConvolution2D(48, 3, 3, border_mode='same', input_shape=(32, 32, 3), activation='relu')) # 32*32*96
my_model.add(BatchNormalization())
my_model.add(Dropout(0.1))
my_model.add(SeparableConvolution2D(96, 3, 3, border_mode='valid', activation='relu')) # 30*30*96
my_model.add(BatchNormalization())
my_model.add(Dropout(0.1))
my_model.add(MaxPooling2D(pool_size=(2, 2))) # 15*15*96
my_model.add(Dropout(0.1))
my_model.add(SeparableConvolution2D(192, 3, 3, border_mode='same', activation='relu')) # 15*15*192
my_model.add(BatchNormalization())
my_model.add(Dropout(0.1))
my_model.add(SeparableConvolution2D(96, 3, 3, border_mode='valid', activation='relu')) # 13*13*192
my_model.add(BatchNormalization())
my_model.add(Dropout(0.1))
my_model.add(MaxPooling2D(pool_size=(2, 2))) # 6*6*192
my_model.add(Dropout(0.1))
my_model.add(SeparableConvolution2D(96, 3, 3, border_mode='same', activation='relu')) # 6*6*96
my_model.add(BatchNormalization())
my_model.add(Dropout(0.1))
my_model.add(SeparableConvolution2D(96, 3, 3, border_mode='valid', activation='relu')) # 4*4*48
my_model.add(BatchNormalization())
my_model.add(Dropout(0.1))
# my_model.add(SeparableConvolution2D(48, 3, 3, border_mode='valid', activation='relu')) # 4*4*48
# my_model.add(BatchNormalization())
# my_model.add(Dropout(0.1))
# my_model.add(MaxPooling2D(pool_size=(2, 2))) # 2*2*96
# my_model.add(Dropout(0.1))
# my_model.add(SeparableConvolution2D(48, 1, 1, border_mode='valid', activation='relu')) # 4*4*48
# my_model.add(BatchNormalization())
# my_model.add(Dropout(0.1))
# my_model.add(SeparableConvolution2D(10, 4, 4, border_mode='valid', activation='relu')) # 1*1*10
# my_model.add(BatchNormalization())
# my_model.add(Dropout(0.1))
my_model.add(AveragePooling2D())
my_model.add(Flatten())
my_model.add(Dense(num_classes, activation='softmax'))
# my_model.add(Activation('softmax'))
# my_model.add(Dense(256))
# my_model.add(Activation('relu'))
# my_model.add(Dense(num_classes, activation='softmax'))
# Compile the model
my_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# + id="uOuVQ8figWEv" colab_type="code" outputId="39081933-5db1-4e19-f358-31821b3619a1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
my_model.summary()
# + id="uMRTFsWBgHTL" colab_type="code" outputId="02812809-a944-4335-823f-a6db191278d2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# train the model
my_start = time.time()
# Train the model
my_model_info = my_model.fit_generator(datagen.flow(train_features, train_labels, batch_size = 128),
samples_per_epoch = train_features.shape[0], nb_epoch = 50,
validation_data = (test_features, test_labels), verbose=1)
my_end = time.time()
print ("Model took %0.2f seconds to train"%(my_end - my_start))
# plot model history
plot_model_history(my_model_info)
# compute test accuracy
print ("Accuracy on test data is: %0.2f"%accuracy(test_features, test_labels, my_model))
| session3/assign_3_70k_81.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import os
import warnings
import sys
warnings.filterwarnings('ignore')
from tensorflow import logging
logging.set_verbosity(logging.ERROR)
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import numpy as np
import tensorflow as tf
from tqdm import tqdm_notebook
import matplotlib.pyplot as plt
# +
sys.path.append('..')
from pydens import Solver, NumpySampler, cart_prod
from pydens import add_tokens
add_tokens()
# -
# <tr>
# <td> <img src="params_1.png" alt="Drawing" style="width: 500px; height: 550px"/> </td>
# <td> <img src="params_2.png" alt="Drawing" style="width: 500px; height: 200px"/> </td>
# </tr>
# +
m = (0.1 + 0.3) / 2
g = 9.81
k_o = (0.1 + 10) / 2
mu_o = (0.5 + 1) / 2
b_o = (1.1 + 1.5) / 2
rho_o = (750 + 950) / 2
coeff_o = k_o / (mu_o*b_o)
Q_o = 10 #(10 + 500) / 2
k_w = (0.1 + 10) / 2
mu_w = (0.2 + 12) / 2
b_w = (1.0 + 1.02) / 2
rho_w = (980 + 1040) / 2
coeff_w = k_w / (mu_w*b_w)
Q_w = 10 #(10 + 500) / 2
theta = (0.01 + 0.03) / 2
a = (-0.25 - 0.05) / 2
c = (-2 - 4) / 2
# -
# <img src="equations.png" alt="Drawing" style="width: 400px; height: 150px"/>
# +
p_ow = lambda s_w: (2*theta / np.sqrt(k_o/m)) / (a*(1 - s_w)/(1 + c*s_w))
eq_1 = lambda p_w, s_w, x, y, z, t: div(coeff_o*D(p_w + p_ow(s_w)) - np.array([0, 0, rho_o*g, 0]), length=3) - \
((m / b_o)*D(1 - s_w, t) + Q_o)
eq_2 = lambda p_w, s_w, x, y, z, t: div(coeff_w*D(p_w) - np.array([0, 0, rho_w*g, 0]), length=3) - \
((m / b_w)*D(s_w, t) + Q_w)
# -
initial_condition = [[200], [0.7]]
# +
pde = {'n_dims': 4, 'n_funs': 2, 'n_eqns': 2,
'form': [eq_1,
eq_2],
'initial_condition': initial_condition,
'time_multiplier': 'sigmoid'}
# Put it all together in model-config
config = {
'pde': pde,
# 'body': {'layout': 'fa fa fa RRfa+ fa fa+',
# 'units': [16, 32, 64, 64, 128, 64],
# 'activation': tf.nn.tanh},
# 'head': {'layout': 'fa f', 'units': [32, 1],
# 'activation': tf.nn.tanh},
'decay': {'name': 'invtime', 'learning_rate':0.05,
'decay_steps': 100, 'decay_rate': 0.05},
'track': {'p_w': lambda p_w, s_w, x, y, z, t: p_w,
'p_o': lambda p_w, s_w, x, y, z, t: p_w + p_ow(s_w),
's_w': lambda p_w, s_w, x, y, z, t: s_w,
's_o': lambda p_w, s_w, x, y, z, t: 1 - s_w},
}
# Uniform sampling scheme
s = NumpySampler('u', dim=4)
# -
# train the network on batches of 100 points
dg = Solver(config, layer_size=35)
dg.fit(batch_size=500, sampler=s, n_iters=1500, bar='notebook')
dg.fit(batch_size=1000, sampler=s, n_iters=10000, bar='notebook')
# Plot loss
plt.plot(dg.loss[50:])
plt.xlabel('iteration number', fontdict={'fontsize': 15})
plt.ylabel('Loss', fontdict={'fontsize': 15})
plt.show()
# +
# Plot one vertical trace
n_el = 100
pts_ = np.linspace(0, 1, n_el).reshape((n_el, 1))
pts = np.asarray([[0.5, 0.5, point, 0.01] for point in pts_])
approxs_p_w, approxs_p_o, approxs_s_w, approxs_s_o = dg.solve(pts, ['p_w', 'p_o', 's_w', 's_o'])
# approxs_s_w = dg.solve(pts, 's_w')
# approxs_p_o = dg.solve(pts, 'p_o')
# approxs_s_o = dg.solve(pts, 's_o')
fig, ax = plt.subplots(figsize=(6, 4))
plt.plot(pts_, approxs_p_w.reshape(n_el, ), 'y', label='p_w approximation')
plt.plot(pts_, approxs_p_o.reshape(n_el, ), 'r', label='p_o approximation')
plt.legend(loc='auto', shadow=True, ncol=2)
fig.show()
fig, ax = plt.subplots(figsize=(6, 4))
plt.plot(pts_, approxs_s_w.reshape(n_el, ), 'y', label='s_w approximation')
plt.plot(pts_, approxs_s_o.reshape(n_el, ), 'r', label='s_o approximation')
plt.legend(loc='auto', shadow=True, ncol=2)
fig.show()
# -
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import numpy as np
from ipywidgets import interact
import ipyvolume
# +
# # %matplotlib notebook
x = np.arange(0, 1, 0.1)
y = np.arange(0, 1, 0.1)
z = np.arange(0, 1, 0.1)
xx, yy, zz = np.meshgrid(x, y, z)
stacked = np.vstack([xx.flatten(), yy.flatten(), zz.flatten()]).T
def plt_solution(time):
pts = np.hstack([stacked, time * np.ones((stacked.shape[0], 1))])
# approxs_p_w = dg.solve(pts)[:, 0]
approxs = np.array(dg.solve(pts, 's_o')).reshape(stacked.shape[0])
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
img = ax.scatter(xx.flatten(), yy.flatten(), zz.flatten(), c=approxs)
fig.colorbar(img)
print(np.mean(approxs))
plt.show()
# plt.xlabel(r'$t$', fontdict={'fontsize': 14})
# plt.title(r'$\epsilon={}$'.format(e))
# plt.legend()
plt.show()
# -
_ = interact(plt_solution, time=(0, 1, 0.1))
# +
# # %matplotlib notebook
x = np.arange(0, 1, 0.1)
y = np.arange(0, 1, 0.1)
xx, yy = np.meshgrid(x, y)
stacked = np.vstack([xx.flatten(), yy.flatten()]).T
def plt_solution(time):
z = 0.5
print(time)
pts = np.hstack([stacked,
z * np.ones((stacked.shape[0], 1)),
time * np.ones((stacked.shape[0], 1))])
print(pts[:, -1])
approxs = np.array(dg.solve(pts, 's_w')).reshape(stacked.shape[0])
fig = plt.figure()
ax = fig.add_subplot(111)
img = ax.imshow(approxs.reshape(10, 10))
fig.colorbar(img)
print(np.mean(approxs))
plt.show()
# plt.xlabel(r'$t$', fontdict={'fontsize': 14})
# plt.title(r'$\epsilon={}$'.format(e))
# plt.legend()
plt.show()
# -
_ = interact(plt_solution, time=(0, 1, 0.1))
pts
| examples/Reservoir modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="BcMGrh0snfeS"
# [](https://colab.research.google.com/github/mravanba/comp551-notebooks/blob/master/MLE.ipynb)
#
# # Maximum Likelihood
# + id="p3fMYNQmnfeU"
import numpy as np
# #%matplotlib notebook
# %matplotlib inline
import matplotlib.pyplot as plt
from IPython.core.debugger import set_trace
np.random.seed(1234)
# + [markdown] id="Z4NFL1AELW_Q"
# Lets assume that a thumbtack has a probability $0 < \theta^* < 1$ of _heads_ .
# We use a **Bernoulli** distribution [$\mathrm{Bernoulli}(x | \theta) = \theta^{x} (1-\theta)^{1-x}$] to model this
# + colab={"base_uri": "https://localhost:8080/"} id="LJ07A94knfea" outputId="5128917b-782a-4f68-a830-326d2acf19dc"
#Function to compute parametric probability mass function
#If you pass arrays or broadcastable arrays it computes the elementwise bernoulli probability mass
Bernoulli = lambda theta,x: theta**x * (1-theta)**(1-x)
theta_star = .4
Bernoulli(theta_star, 0)
# + [markdown] id="LiYxzoBxnfeh"
# The **likelihood** function for this dataset is
# $$L(\theta) = \prod_{n=1}^{10} \mathrm{Bernoulli}(\theta, x^{(n)})$$
# Note that this is a function of $\theta$, it is not a probability distribution, so it doesn't sum to 1.
# Next we plot this function for $n$ randomly generated samples using $\theta^*$.
# + colab={"base_uri": "https://localhost:8080/", "height": 343} id="i6zoMYmrnfei" outputId="405f57ae-34ab-45c8-8fb1-5d7f3e5d471c"
n = 10 #number of random samples you want to consider
xn = np.random.rand(n) < theta_star #Generates n element boolean array where elements are True with probability theta_star and otherwise False
xn = xn.astype(int) #to change the boolean array to intergers [0:False, 1:True]
print("observation {}".format(xn))
#Function to compute the log likelihood
#Note that you can either pass this function a scalar(always broadcastable) theta or a broadcastable(in data axis) theta to get likelihood value or values
#Also note that we added an extra dimension in xn to broadcast it along theta dimension
L = lambda theta: np.prod(Bernoulli(theta, xn[None,:]), axis=-1)
#we generate 100 evenly placed values of theta from 0 to 1
theta_vals = np.linspace(0,1,100)[:, None] #Note that we made the array broadcastable by adding an extra dimension for data
plt.plot(theta_vals, L(theta_vals), '-')
plt.xlabel(r"$\theta$")
plt.ylabel(r"Likelihood $L(\theta)$")
#plt.title(r"likelihood function Bernoulli("+str(xn.astype(int))+r"$|\theta)$")
plt.show()
# + [markdown] id="XbXIwS4Qnfek"
# **maximum likelihood** method for learning a probabilistic model picks $\theta$ that maximizes $L(\theta)$ for a given set of observations $\mathcal{D} = \{x^{(1)}, \ldots, x^{(N)}\}$.
# For _Bernoulli_ dist. max. likelihood estimate is the $\widehat{\theta} = \frac{1}{N} \sum_n x^{(n)}$, corresponding to the peak of the likelihood plot. The value of the likelihood shrinks exponentially as we increase the number of observations $N$ -- try `N=100`.
# A more numerically stable value with the same _maximum_ is the **log-likelihood** $\ell(\theta) = \log(L(\theta))$. It is also customary to minimize the _negative log-likelihood_ (NLL).
# Let's plot NLL for different $N$ -- as we increase our data-points the ML estimate often gets better.
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="gOsr7Tgcnfel" outputId="1a8ab572-361e-4cc1-81ce-6300e4ad571a"
#Generates 2^12 element boolean array where elements are True with probability theta_star and otherwise False
xn_max = np.random.rand(2**12) < theta_star
for r in range(1,6):
n = 4**r #number of data samples for r-th iteration
xn = xn_max[:n] #slice them from the total samples generated
#Function to compute the log likelihood (Implementation exactly similar to the likelihood function)
ll = lambda theta: np.sum(np.log(Bernoulli(theta, xn[None,:])), axis=-1)
theta_vals = np.linspace(.01,.99,100)[:, None]
ll_vals = -ll(theta_vals)
#Plot the log likelihood values
plt.plot(theta_vals, ll_vals, label="n="+str(n))
max_ind = np.argmin(ll_vals) #Stores the theta corresponding to minimum log likelihood
plt.plot(theta_vals[max_ind], ll_vals[max_ind], '*')
#to get the horizontal line for theta
plt.plot([theta_star,theta_star], [0,ll_vals.max()], '--', label=r"$\theta^*$")
plt.xlabel(r"$\theta$")
plt.ylabel(r"Negative Log-Likelihood $-\ell(\theta)$")
plt.yscale("log")
plt.title("ML solution with increasing data")
plt.legend()
plt.show()
# -
| MLE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py39)
# language: python
# name: py39
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import datetime as dt
from salishsea_tools import evaltools as et, viz_tools
import os
import datetime as dt
import gsw
import matplotlib.gridspec as gridspec
import matplotlib as mpl
import matplotlib.dates as mdates
import cmocean as cmo
import scipy.interpolate as sinterp
import cmocean
import json
import f90nml
from collections import OrderedDict
from scipy.stats.mstats import gmean
fs=16
mpl.rc('xtick', labelsize=fs)
mpl.rc('ytick', labelsize=fs)
mpl.rc('legend', fontsize=fs)
mpl.rc('axes', titlesize=fs)
mpl.rc('axes', labelsize=fs)
mpl.rc('figure', titlesize=fs)
mpl.rc('font', size=fs)
mpl.rc('font', family='sans-serif', weight='normal', style='normal')
import warnings
#warnings.filterwarnings('ignore')
from IPython.display import Markdown, display
# %matplotlib inline
# -
PSdf=pd.read_excel('/ocean/ksuchy/MOAD/observe/PugetSoundBiomassDF.xlsx',engine='openpyxl')
PSdf
SoGdf=pd.read_excel('/ocean/ksuchy/MOAD/observe/SoGBiomassDF.xlsx',engine='openpyxl')
SoGdf
SoGdf.keys()
SoGdf['Copepods']=(SoGdf['Calanoids']+SoGdf['Cyclopoids']+SoGdf['Poecilostomatoids']+SoGdf['Harpacticoids'])
SoGdf.rename(columns={'Decapods':'Crabs','Aphragmophora':'Chaetognaths','Thecosomata':'Gastropods'},inplace=True)
SoGdf.keys()
SoGdf.drop(labels=['Station','Twilight', 'Net_Type', 'Mesh_Size(um)','Bottom Depth(m)', 'Diplostraca', 'Thecostraca', 'Amphipoda',
'Euphausiacea', 'Calanoida', 'Cyclopoida',
'Poecilostomatoida', 'Halocyprida', 'Copelata','Decapoda',
'Leptothecate', 'Siphonophorae', 'Trachylina', 'Cydippida',
'Pholadomyoida', 'Neotaenioglossa','Aciculata',
'Canalipalpata', 'Osmeriformes', 'Perciformes', 'Beroida', 'Teuthida',
'Gymnosomata', 'Isopoda', 'Siphonostomatoida', 'Anthoathecatae',
'Scorpaeniformes', 'Phragmophora', 'Clupeiformes', 'Ophiurida',
'Gadiformes', 'Semaeostomeae', 'Cumacea', 'Echinoida', 'Harpacticoida',
'Pleuronectiformes', 'Tricladida', 'Myodocopida', 'Phaeogromia',
'Noctilucales', 'Octopoda', 'Actiniaria', 'Foraminiferida',
'Monstrilloida', 'Oligotrichida', 'Mysida', 'Acariformes',
'Lophogastrida', 'Ophidiiformes', 'Thalassocalycida', 'Doliolida',
'Lepadomorpha', 'Cephalaspidea', 'Sygnathiformes','Calanoids',
'Cyclopoids', 'Poecilostomatoids', 'Harpacticoids','Month','OtherGroups'],axis=1,inplace=True)
SoGdf.keys()
#rename columns
PSdf.keys()
# +
#rename columns
#
# -
PSdf.drop(labels=['Site Name','Station','Sub Basin', 'Sample Year', 'Sample Month',
'Sample Time','Mesh Size', 'Diameter (cm)','Z','ACARTIA HUDSONICA', 'ACARTIA LONGIREMIS', 'AETIDEUS',
'CALANUS MARSHALLAE', 'CALANUS PACIFICUS', 'CENTROPAGES ABDOMINALIS',
'DITRICHOCORYCAEUS ANGLICUS', 'EPILABIDOCERA AMPHITRITES', 'EUCALANUS',
'EUCALANUS BUNGII', 'METRIDIA PACIFICA', 'MICROCALANUS',
'NEOCALANUS PLUMCHRUS', 'OITHONA SIMILIS', 'PARACALANUS',
'PARAEUCHAETA ELONGATA', 'PSEUDOCALANUS', 'PSEUDOCALANUS Lg',
'PSEUDOCALANUS MIMUS', 'PSEUDOCALANUS MOULTONI',
'PSEUDOCALANUS NEWMANI', 'PSEUDOCALANUS Sm', 'CYPHOCARIS CHALLENGERI',
'CALLIOPIUS PACIFICUS', 'GAMMARIDEA', 'THEMISTO PACIFICA',
'EUPHAUSIA PACIFICA', 'THYSANOESSA RASCHII', 'GASTROPODA',
'CLIONE LIMACINA', 'LIMACINA HELICINA', 'CHAETOGNATHA',
'FABIA SUBQUADRATA', 'CANCER PRODUCTUS', 'CANCRIDAE', 'Cancridae Lg',
'Cancridae Sm', 'GLEBOCARCINUS OREGONENSIS', 'LOPHOPANOPEUS BELLUS',
'METACARCINUS GRACILIS', 'METACARCINUS MAGISTER', 'PINNIXA',
'PINNOTHERES', 'PUGETTIA', 'OIKOPLEURA'],axis=1,inplace=True)
PSdf.keys()
PSdf.rename(columns={'Basin':'region_name','Sample Date':'Date'},inplace=True)
SoGdf.keys()
PSdf.keys()
SalishSeadf=pd.concat([PSdf, SoGdf], ignore_index=True)
SalishSeadf
#create new columns
SalishSeadf['Month']=[ii.month for ii in SalishSeadf['dtUTC']]
import netCDF4 as nc
fdict={'ptrc_T':1,'grid_T':1}
start_date = dt.datetime(2012,1,1)
end_date = dt.datetime(2016,12,31)
flen=1 # number of days per model output file. always 1 for 201905 and 201812 model runs
namfmt='nowcast' # for 201905 and 201812 model runs, this should always be 'nowcast'
# filemap is dictionary of the form variableName: fileType, where variableName is the name
# of the variable you want to extract and fileType designates the type of
# model output file it can be found in (usually ptrc_T for biology, grid_T for temperature and
# salinity)
filemap={'microzooplankton':'ptrc_T','mesozooplankton':'ptrc_T'}
# fdict is a dictionary mappy file type to its time resolution. Here, 1 means hourly output
# (1h file) and 24 means daily output (1d file). In certain runs, multiple time resolutions
# are available
fdict={'ptrc_T':1,'grid_T':1}
PATH= '/results2/SalishSea/nowcast-green.201905/'
data=et.matchData(SalishSeadf,filemap,fdict,start_date,end_date,'nowcast',PATH,1,quiet=False,method='vertNet');
data
data['mod_total']=data['mod_microzooplankton']+data['mod_mesozooplankton']
# define log transform function with slight shift to accommodate zero values
def logt(x):
return np.log10(x+.001)
# define inverse log transform with same shift
def logt_inv(y):
return 10**y-.001
# +
#Create columns for Log10 transformations
data['L10Total']=logt(data['Total'])
data['L10Copepods']=logt(data['Copepods'])
data['L10Amphipods']=logt(data['Amphipods'])
data['L10Euphausiids']=logt(data['Euphausiids'])
data['L10Gastropods']=logt(data['Gastropods'])
data['L10Chaetognaths']=logt(data['Chaetognaths'])
data['L10Crabs']=logt(data['Crabs'])
data['L10Larvaceans']=logt(data['Larvaceans'])
data['L10MainGroups']=logt(data['MainGroups'])
#Convert model values to mg C m-3 by muliplying value * C:N of 5.7 * molecular weight of C
data['L10mod_microzooplankton']=logt(data['mod_microzooplankton']*5.7*12)
data['L10mod_mesozooplankton']=logt(data['mod_mesozooplankton']*5.7*12)
data['L10mod_total']=logt(data['mod_total']*5.7*12)
# -
cm1=cmocean.cm.thermal
with nc.Dataset('/data/eolson/results/MEOPAR/NEMO-forcing-new/grid/bathymetry_201702.nc') as bathy:
bathylon=np.copy(bathy.variables['nav_lon'][:,:])
bathylat=np.copy(bathy.variables['nav_lat'][:,:])
bathyZ=np.copy(bathy.variables['Bathymetry'][:,:])
data['broadregions']=np.nan
# +
#data.loc[(data.region_name=='Northern Strait of Georgia')|(data.region_name=='Central Strait of Georgia')|(data.region_name=='Southern Strait of Georgia'),['broadregions']]='SoG'
data.loc[(data.region_name=='Northern Strait of Georgia')|(data.region_name=='Central Strait of Georgia'),['broadregions']]='Main SoG'
data.loc[(data.region_name=='Southern Strait of Georgia'),['broadregions']]='Southern SoG'
data.loc[(data.region_name=='Nearshore-Central East')|(data.region_name=='Nearshore-North East')|(data.region_name=='Nearshore North West'),['broadregions']]='Nearshore'
data.loc[(data.region_name=='Baynes Sound'),['broadregions']]='Baynes'
data.loc[(data.region_name=='Tidal Mixed')|(data.region_name=='Gulf Islands')|(data.region_name=='San Juan Islands')|(data.region_name=='San Juan'),['broadregions']]='Mixed'
data.loc[(data.region_name=='Juan de Fuca'),['broadregions']]='JdeF'
data.loc[(data.region_name=='Whidbey Basin')|(data.region_name=='Bellingham Bay')|(data.region_name=='Admiralty Inlet'),['broadregions']]='North Sound'
data.loc[(data.region_name=='Central Basin'),['broadregions']]='Central Sound'
data.loc[(data.region_name=='South Sound')|(data.region_name=='Hood Canal'),['broadregions']]='South Sound'
#data.loc[(data.region_name=='Hood Canal'),['broadregions']]='Hood Canal'
# +
#data.loc[data['broadregions']=='SoG']
# -
fig, ax = plt.subplots(1,1,figsize = (8,8))
with nc.Dataset('/ocean/ksuchy/MOAD/NEMO-forcing/grid/bathymetry_201702.nc') as grid:
viz_tools.plot_coastline(ax, grid, coords = 'map',isobath=.1)
colors=('black','orange','firebrick','mediumspringgreen','fuchsia','royalblue','green','darkviolet',
'lime','lightblue','darkgoldenrod','darkorange','deepskyblue','teal','darkgreen','darkblue','slateblue','purple')
datreg=dict()
for ind, iregion in enumerate(data.broadregions.unique()):
datreg[iregion] = data.loc[data.broadregions==iregion]
ax.plot(datreg[iregion]['Lon'], datreg[iregion]['Lat'],'.',
color = colors[ind], label=iregion)
ax.set_ylim(47, 51)
ax.xaxis.set_tick_params(labelsize=14)
ax.yaxis.set_tick_params(labelsize=14)
ax.legend(bbox_to_anchor=[1,.6,0,0])
ax.set_xlim(-126, -121);
ax.set_title('Salish Sea Observation Locations');
ax.legend(bbox_to_anchor=(1.05, 1), loc='upper left',frameon=False,markerscale=3.,fontsize=11)
def byRegion(ax,obsvar,modvar,lims):
SS=[]
for ind, iregion in enumerate(data.broadregions.unique()):
#ax.plot(datreg[iregion]['Lon'], datreg[iregion]['Lat'],'.',
#color = colors[ind], label=iregion)
SS0=et.varvarPlot(ax,datreg[iregion],obsvar,modvar,
cols=(colors[ind],),lname=iregion)
SS.append(SS0)
l=ax.legend(handles=[ip[0][0] for ip in SS])
ax.set_xlabel('Observations')
ax.set_ylabel('Model')
ax.plot(lims,lims,'k-',alpha=.5)
ax.set_xlim(lims)
ax.set_ylim(lims)
ax.set_aspect(1)
return SS,l
data['Month']=[ii.month for ii in data['dtUTC']]
JF=data.loc[(data.Month==1)|(data.Month==2)]
MAM=data.loc[(data.Month==3)|(data.Month==4)|(data.Month==5)]
JJA=data.loc[(data.Month==6)|(data.Month==7)|(data.Month==8)]
SOND=data.loc[(data.Month==9)|(data.Month==10)|(data.Month==11)|(data.Month==12)]
def bySeason(ax,obsvar,modvar,lims):
for axi in ax:
axi.plot(lims,lims,'k-')
axi.set_xlim(lims)
axi.set_ylim(lims)
axi.set_aspect(1)
axi.set_xlabel('Obs')
axi.set_ylabel('Model')
SS=et.varvarPlot(ax[0],JF,obsvar,modvar,cols=('crimson','darkturquoise','navy'))
ax[0].set_title('Winter')
SS=et.varvarPlot(ax[1],MAM,obsvar,modvar,cols=('crimson','darkturquoise','navy'))
ax[1].set_title('Spring')
SS=et.varvarPlot(ax[2],JJA,obsvar,modvar,cols=('crimson','darkturquoise','navy'))
ax[2].set_title('Summer')
SS=et.varvarPlot(ax[3],SOND,obsvar,modvar,cols=('crimson','darkturquoise','navy'))
ax[3].set_title('Autumn')
return
data.region_name.unique()
# +
#data.loc[(data.region_name=='Northern Strait of Georgia')|(data.region_name=='Central Strait of Georgia')|(data.region_name=='Southern Strait of Georgia'),['broadregions']]='SoG'
MainSoG=data.loc[(data.region_name=='Northern Strait of Georgia')|(data.region_name=='Central Strait of Georgia')]
SouthernSoG=data.loc[(data.region_name=='Southern Strait of Georgia')]
Nearshore=data.loc[(data.region_name=='Nearshore-Central East')|(data.region_name=='Nearshore-North East')|(data.region_name=='Nearshore North West')]
Baynes=data.loc[(data.region_name=='Baynes Sound')]
Mixed=data.loc[(data.region_name=='Tidal Mixed')|(data.region_name=='Gulf Islands')|(data.region_name=='San Juan Islands')|(data.region_name=='San Juan')]
JdeF=data.loc[(data.region_name=='Juan de Fuca')]
NorthSound=data.loc[(data.region_name=='Whidbey Basin')|(data.region_name=='Bellingham Bay')|(data.region_name=='Admiralty Inlet')]
CentralSound=data.loc[(data.region_name=='Central Basin')]
SouthSound=data.loc[(data.region_name=='South Sound')|(data.region_name=='Hood Canal')]
#data.loc[(data.region_name=='Hood Canal'),['broadregions']]='Hood Canal'
# -
monthlymeanMainSoG=MainSoG.groupby(['Month']).mean()
monthlymeanSouthernSoG=SouthernSoG.groupby(['Month']).mean()
monthlymeanNearshore=Nearshore.groupby(['Month']).mean()
monthlymeanBaynes=Baynes.groupby(['Month']).mean()
monthlymeanMixed=Mixed.groupby(['Month']).mean()
monthlymeanJdeF=JdeF.groupby(['Month']).mean()
monthlymeanSouthSound=SouthSound.groupby(['Month']).mean()
monthlymeanNorthSound=NorthSound.groupby(['Month']).mean()
monthlymeanCentralSound=CentralSound.groupby(['Month']).mean()
fig, ax = plt.subplots(1,1,figsize = (16,9))
PS,l=byRegion(ax,'L10Total','L10mod_total',(-0.5,3.5))
ax.set_title('Salish Sea Total Biomass (mg C m-3)',fontsize=14)
ax.legend(bbox_to_anchor=(1.1, 1.05),frameon=False,markerscale=2.5)
#fig.savefig('PugetSoundTotalBiomassModvsObs.jpg',bbox_inches='tight')
fig, ax = plt.subplots(1,4,figsize = (16,3.3))
bySeason(ax,'L10Total','L10mod_total',(-3,3))
'black','orange','firebrick','mediumspringgreen','fuchsia','royalblue','green','darkviolet',
'lime'
# +
fig,ax=plt.subplots(1,1,figsize=(12,2.5))
ax.plot(logt_inv(monthlymeanMainSoG['L10Total']),'--',color='k',label='Main SoG')
ax.plot(logt_inv(monthlymeanSouthernSoG['L10Total']),'--',color='lime',label='Southern SoG')
ax.plot(logt_inv(monthlymeanNearshore['L10Total']),'-',color='firebrick',label='Nearshore')
ax.plot(logt_inv(monthlymeanBaynes['L10Total']),'--',color='darkviolet',label='Baynes')
ax.plot(logt_inv(monthlymeanMixed['L10Total']),'--',color='orange',label='Mixed')
ax.plot(logt_inv(monthlymeanJdeF['L10Total']),'--',color='mediumspringgreen',label='JdeF')
ax.plot(logt_inv(monthlymeanNorthSound['L10Total']),'-',color='royalblue',label='North Sound')
ax.plot(logt_inv(monthlymeanSouthSound['L10Total']),'-',color='fuchsia',label='South Sound')
ax.plot(logt_inv(monthlymeanCentralSound['L10Total']),'-',color='green',label='Central Sound')
ax.set_ylim(0,250)
ax.set_xlim(0,12)
ax.set_title('Observation Zooplankton Seasonal Cycle')
ax.set_ylabel('Mean Biomass (mg C m-3)',fontsize=10)
ax.legend(fontsize=10,frameon=False,bbox_to_anchor=(1., 1.05))
fig,ax=plt.subplots(1,1,figsize=(12,2.5))
ax.plot(logt_inv(monthlymeanMainSoG['L10mod_total']),'--',color='k',label='Main SoG')
ax.plot(logt_inv(monthlymeanSouthernSoG['L10mod_total']),'--',color='lime',label='Southern SoG')
ax.plot(logt_inv(monthlymeanNearshore['L10mod_total']),'-',color='firebrick',label='Nearshore')
ax.plot(logt_inv(monthlymeanBaynes['L10mod_total']),'--',color='darkviolet',label='Baynes')
ax.plot(logt_inv(monthlymeanMixed['L10mod_total']),'--',color='orange',label='Mixed')
ax.plot(logt_inv(monthlymeanJdeF['L10mod_total']),'--',color='mediumspringgreen',label='JdeF')
ax.plot(logt_inv(monthlymeanNorthSound['L10mod_total']),'-',color='royalblue',label='North Sound')
ax.plot(logt_inv(monthlymeanSouthSound['L10mod_total']),'-',color='fuchsia',label='South Sound')
ax.plot(logt_inv(monthlymeanCentralSound['L10mod_total']),'-',color='green',label='Central Sound')
ax.set_ylim(0,250)
ax.set_xlim(0,12)
ax.set_title('Model Zooplankton Seasonal Cycle')
ax.set_ylabel('Mean Biomass (mg C m-3)',fontsize=10)
ax.legend(fontsize=10,frameon=False,bbox_to_anchor=(1., 1.05))
# -
| notebooks/SalishSeaZoopBiomass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Risk minimization for loan investments
#
# ## The Data
#
# We will be using a subset of the LendingClub DataSet obtained from Kaggle: https://www.kaggle.com/wordsforthewise/lending-club
#
# LendingClub is a US peer-to-peer lending company, headquartered in San Francisco, California.[3] It was the first peer-to-peer lender to register its offerings as securities with the Securities and Exchange Commission (SEC), and to offer loan trading on a secondary market. LendingClub is the world's largest peer-to-peer lending platform.
#
# ### Our Goal
#
# Given historical data on loans given out with information on whether or not the borrower defaulted (charge-off), can we build a model thatcan predict wether or nor a borrower will pay back their loan? This way in the future when we get a new potential customer we can assess whether or not they are likely to pay back the loan. Keep in mind classification metrics when evaluating the performance of your model!
#
# The "loan_status" column contains our label.
#
# ### Data Overview
# ----
# -----
# There are many LendingClub data sets on Kaggle. Here is the information on this particular data set:
#
# <table border="1" class="dataframe">
# <thead>
# <tr style="text-align: right;">
# <th></th>
# <th>LoanStatNew</th>
# <th>Description</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <th>0</th>
# <td>loan_amnt</td>
# <td>The listed amount of the loan applied for by the borrower. If at some point in time, the credit department reduces the loan amount, then it will be reflected in this value.</td>
# </tr>
# <tr>
# <th>1</th>
# <td>term</td>
# <td>The number of payments on the loan. Values are in months and can be either 36 or 60.</td>
# </tr>
# <tr>
# <th>2</th>
# <td>int_rate</td>
# <td>Interest Rate on the loan</td>
# </tr>
# <tr>
# <th>3</th>
# <td>installment</td>
# <td>The monthly payment owed by the borrower if the loan originates.</td>
# </tr>
# <tr>
# <th>4</th>
# <td>grade</td>
# <td>LC assigned loan grade</td>
# </tr>
# <tr>
# <th>5</th>
# <td>sub_grade</td>
# <td>LC assigned loan subgrade</td>
# </tr>
# <tr>
# <th>6</th>
# <td>emp_title</td>
# <td>The job title supplied by the Borrower when applying for the loan.*</td>
# </tr>
# <tr>
# <th>7</th>
# <td>emp_length</td>
# <td>Employment length in years. Possible values are between 0 and 10 where 0 means less than one year and 10 means ten or more years.</td>
# </tr>
# <tr>
# <th>8</th>
# <td>home_ownership</td>
# <td>The home ownership status provided by the borrower during registration or obtained from the credit report. Our values are: RENT, OWN, MORTGAGE, OTHER</td>
# </tr>
# <tr>
# <th>9</th>
# <td>annual_inc</td>
# <td>The self-reported annual income provided by the borrower during registration.</td>
# </tr>
# <tr>
# <th>10</th>
# <td>verification_status</td>
# <td>Indicates if income was verified by LC, not verified, or if the income source was verified</td>
# </tr>
# <tr>
# <th>11</th>
# <td>issue_d</td>
# <td>The month which the loan was funded</td>
# </tr>
# <tr>
# <th>12</th>
# <td>loan_status</td>
# <td>Current status of the loan</td>
# </tr>
# <tr>
# <th>13</th>
# <td>purpose</td>
# <td>A category provided by the borrower for the loan request.</td>
# </tr>
# <tr>
# <th>14</th>
# <td>title</td>
# <td>The loan title provided by the borrower</td>
# </tr>
# <tr>
# <th>15</th>
# <td>zip_code</td>
# <td>The first 3 numbers of the zip code provided by the borrower in the loan application.</td>
# </tr>
# <tr>
# <th>16</th>
# <td>addr_state</td>
# <td>The state provided by the borrower in the loan application</td>
# </tr>
# <tr>
# <th>17</th>
# <td>dti</td>
# <td>A ratio calculated using the borrower’s total monthly debt payments on the total debt obligations, excluding mortgage and the requested LC loan, divided by the borrower’s self-reported monthly income.</td>
# </tr>
# <tr>
# <th>18</th>
# <td>earliest_cr_line</td>
# <td>The month the borrower's earliest reported credit line was opened</td>
# </tr>
# <tr>
# <th>19</th>
# <td>open_acc</td>
# <td>The number of open credit lines in the borrower's credit file.</td>
# </tr>
# <tr>
# <th>20</th>
# <td>pub_rec</td>
# <td>Number of derogatory public records</td>
# </tr>
# <tr>
# <th>21</th>
# <td>revol_bal</td>
# <td>Total credit revolving balance</td>
# </tr>
# <tr>
# <th>22</th>
# <td>revol_util</td>
# <td>Revolving line utilization rate, or the amount of credit the borrower is using relative to all available revolving credit.</td>
# </tr>
# <tr>
# <th>23</th>
# <td>total_acc</td>
# <td>The total number of credit lines currently in the borrower's credit file</td>
# </tr>
# <tr>
# <th>24</th>
# <td>initial_list_status</td>
# <td>The initial listing status of the loan. Possible values are – W, F</td>
# </tr>
# <tr>
# <th>25</th>
# <td>application_type</td>
# <td>Indicates whether the loan is an individual application or a joint application with two co-borrowers</td>
# </tr>
# <tr>
# <th>26</th>
# <td>mort_acc</td>
# <td>Number of mortgage accounts.</td>
# </tr>
# <tr>
# <th>27</th>
# <td>pub_rec_bankruptcies</td>
# <td>Number of public record bankruptcies</td>
# </tr>
# </tbody>
# </table>
#
# ---
# ----
# ## Starter Code
#
import pandas as pd
data_info = pd.read_csv('../DATA/lending_club_info.csv',index_col='LoanStatNew')
print(data_info.loc['revol_util']['Description'])
def feat_info(col_name):
print(data_info.loc[col_name]['Description'])
feat_info('mort_acc')
# ## Loading the data and other imports
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# might be needed depending on your version of Jupyter
# %matplotlib inline
# -
df = pd.read_csv('../DATA/lending_club_loan_two.csv')
df.info()
#
# ------
#
# # Section 1: Exploratory Data Analysis
#
# **OVERALL GOAL: Get an understanding for which variables are important, view summary statistics, and visualize the data**
#
#
# ----
# **Since we will be attempting to predict loan_status, thus creating a countplot.**
sns.countplot(x='loan_status',data=df)
# **Creating a histogram of the loan_amnt column.**
plt.figure(figsize=(12,4))
sns.distplot(df['loan_amnt'],kde=False,bins=40)
plt.xlim(0,45000)
# **Let's explore correlation between the continuous feature variables using .corr() method.**
df.corr()
# **Visualize this using a heatmap.**
plt.figure(figsize=(12,7))
sns.heatmap(df.corr(),annot=True,cmap='viridis')
plt.ylim(10, 0)
# **We should have noticed almost perfect correlation with the "installment" feature. Explore this feature further.**
feat_info('installment')
feat_info('loan_amnt')
sns.scatterplot(x='installment',y='loan_amnt',data=df,)
# **Creating a boxplot showing the relationship between the loan_status and the Loan Amount.**
sns.boxplot(x='loan_status',y='loan_amnt',data=df)
# **Calculating the summary statistics for the loan amount, grouped by the loan_status.**
df.groupby('loan_status')['loan_amnt'].describe()
# **Let's explore the Grade and SubGrade columns that LendingClub attributes to the loans and exploring unique possible grades and subgrades**
sorted(df['grade'].unique())
sorted(df['sub_grade'].unique())
# **Creating a countplot per grade. Set the hue to the loan_status label.**
sns.countplot(x='grade',data=df,hue='loan_status')
# **Displaying a count plot per subgrade.**
plt.figure(figsize=(12,4))
subgrade_order = sorted(df['sub_grade'].unique())
sns.countplot(x='sub_grade',data=df,order = subgrade_order,palette='coolwarm' )
plt.figure(figsize=(12,4))
subgrade_order = sorted(df['sub_grade'].unique())
sns.countplot(x='sub_grade',data=df,order = subgrade_order,palette='coolwarm' ,hue='loan_status')
# **It looks like F and G subgrades don't get paid back that often. Isloateing those and recreating the countplot just for those subgrades.**
# +
f_and_g = df[(df['grade']=='G') | (df['grade']=='F')]
plt.figure(figsize=(12,4))
subgrade_order = sorted(f_and_g['sub_grade'].unique())
sns.countplot(x='sub_grade',data=f_and_g,order = subgrade_order,hue='loan_status')
# -
# **Creating a new column called 'load_repaid' which will contain a 1 if the loan status was "Fully Paid" and a 0 if it was "Charged Off".**
df['loan_status'].unique()
df['loan_repaid'] = df['loan_status'].map({'Fully Paid':1,'Charged Off':0})
df[['loan_repaid','loan_status']]
# **Creating a bar plot showing the correlation of the numeric features to the new loan_repaid column.
df.corr()['loan_repaid'].sort_values().drop('loan_repaid').plot(kind='bar')
# ---
# ---
# # Section 2: Data PreProcessing
#
# **Removing and filling any missing data and Converting categorical string features to dummy variables.**
#
#
df.head()
# # Missing Data
#
# **Let's explore this missing data columns. We use a variety of factors to decide whether or not they would be useful, to see if we should keep, discard, or fill in the missing data.**
# **Length of the dataframe**
len(df)
# **Creating a Series that displays the total count of missing values per column.**
df.isnull().sum()
# **Converting this Series to be in term of percentage of the total DataFrame**
100* df.isnull().sum()/len(df)
# **Let's examine emp_title and emp_length to see whether it will be okay to drop them. Print out their feature information using the feat_info() function from the top of this notebook.**
feat_info('emp_title')
print('\n')
feat_info('emp_length')
# **Unique employment job titles**
df['emp_title'].nunique()
df['emp_title'].value_counts()
# **Realistically there are too many unique job titles to try to convert this to a dummy variable feature. Let's remove that emp_title column.**
df = df.drop('emp_title',axis=1)
# **Creating a count plot of the emp_length feature column and sorting the order of the values.**
sorted(df['emp_length'].dropna().unique())
emp_length_order = [ '< 1 year',
'1 year',
'2 years',
'3 years',
'4 years',
'5 years',
'6 years',
'7 years',
'8 years',
'9 years',
'10+ years']
# +
plt.figure(figsize=(12,4))
sns.countplot(x='emp_length',data=df,order=emp_length_order)
# -
# **Ploting out the countplot with a hue separating Fully Paid vs Charged Off**
plt.figure(figsize=(12,4))
sns.countplot(x='emp_length',data=df,order=emp_length_order,hue='loan_status')
# **This still doesn't really inform us if there is a strong relationship between employment length and being charged off, what we want is the percentage of charge offs per category. Essentially informing us what percent of people per employment category didn't pay back their loan. There are a multitude of ways to create this Series. Once you've created it, see if visualize it with a bar plot.**
emp_co = df[df['loan_status']=="Charged Off"].groupby("emp_length").count()['loan_status']
emp_fp = df[df['loan_status']=="Fully Paid"].groupby("emp_length").count()['loan_status']
emp_len = emp_co/emp_fp
emp_len
emp_len.plot(kind='bar')
# **Charging off rates are extremely similar across all employment lengths and droping the emp_length column.**
df = df.drop('emp_length',axis=1)
# **Revisiting the DataFrame to see what feature columns still have missing data.**
df.isnull().sum()
# **Review the title column vs the purpose column. Is this repeated information?**
df['purpose'].head(10)
df['title'].head(10)
# **The title column is simply a string subcategory/description of the purpose column and droping the title column.**
df = df.drop('title',axis=1)
#
# **Finding out what the mort_acc feature represents**
feat_info('mort_acc')
# **Createing a value_counts of the mort_acc column.**
df['mort_acc'].value_counts()
# **There are many ways we could deal with this missing data. We could attempt to build a simple model to fill it in, such as a linear model, we could just fill it in based on the mean of the other columns, or you could even bin the columns into categories and then set NaN as its own category. There is no 100% correct approach! Let's review the other columsn to see which most highly correlates to mort_acc**
print("Correlation with the mort_acc column")
df.corr()['mort_acc'].sort_values()
# **Looks like the total_acc feature correlates with the mort_acc , this makes sense! We will group the dataframe by the total_acc and calculate the mean value for the mort_acc per total_acc entry.**
print("Mean of mort_acc column per total_acc")
df.groupby('total_acc').mean()['mort_acc']
# **Let's fill in the missing mort_acc values based on their total_acc value. If the mort_acc is missing, then we will fill in that missing value with the mean value corresponding to its total_acc value from the Series we created above. This involves using an .apply() method with two columns.**
total_acc_avg = df.groupby('total_acc').mean()['mort_acc']
total_acc_avg[2.0]
def fill_mort_acc(total_acc,mort_acc):
'''
Accepts the total_acc and mort_acc values for the row.
Checks if the mort_acc is NaN , if so, it returns the avg mort_acc value
for the corresponding total_acc value for that row.
total_acc_avg here should be a Series or dictionary containing the mapping of the
groupby averages of mort_acc per total_acc values.
'''
if np.isnan(mort_acc):
return total_acc_avg[total_acc]
else:
return mort_acc
df['mort_acc'] = df.apply(lambda x: fill_mort_acc(x['total_acc'], x['mort_acc']), axis=1)
df.isnull().sum()
# **revol_util and the pub_rec_bankruptcies have missing data points, but they account for less than 0.5% of the total data and removeing the rows that are missing those values in those columns with dropna().**
df = df.dropna()
df.isnull().sum()
# ## Categorical Variables and Dummy Variables
#
# **We're done working with the missing data! Now we just need to deal with the string values due to the categorical columns.**
#
# **List all the columns that are currently non-numeric.**
df.select_dtypes(['object']).columns
# ---
# **Let's now go through all the string features to see what we should do with them.**
#
# ---
#
#
# ### term feature
#
# **Converting the term feature into either a 36 or 60 integer numeric data type using .apply() or .map().**
df['term'].value_counts()
# Or just use .map()
df['term'] = df['term'].apply(lambda term: int(term[:3]))
# ### grade feature
#
# **We already know grade is part of sub_grade, so just drop the grade feature.**
df = df.drop('grade',axis=1)
# **Converting the subgrade into dummy variables. Then concatenate these new columns to the original dataframe. And drop the original subgrade column and to add drop_first=True to your get_dummies call.**
subgrade_dummies = pd.get_dummies(df['sub_grade'],drop_first=True)
df = pd.concat([df.drop('sub_grade',axis=1),subgrade_dummies],axis=1)
df.columns
df.select_dtypes(['object']).columns
# ### verification_status, application_type,initial_list_status,purpose
# **Converting these columns: ['verification_status', 'application_type','initial_list_status','purpose'] into dummy variables and concatenate them with the original dataframe. And setting drop_first=True and to drop the original columns.**
dummies = pd.get_dummies(df[['verification_status', 'application_type','initial_list_status','purpose' ]],drop_first=True)
df = df.drop(['verification_status', 'application_type','initial_list_status','purpose'],axis=1)
df = pd.concat([df,dummies],axis=1)
# ### home_ownership
# **Review the value_counts for the home_ownership column.**
df['home_ownership'].value_counts()
# **Convert these to dummy variables, but replace NONE and ANY with OTHER, so that we end up with just 4 categories, MORTGAGE, RENT, OWN, OTHER. Then concatenating them with the original dataframe and setting drop_first=True and to drop the original columns.**
# +
df['home_ownership']=df['home_ownership'].replace(['NONE', 'ANY'], 'OTHER')
dummies = pd.get_dummies(df['home_ownership'],drop_first=True)
df = df.drop('home_ownership',axis=1)
df = pd.concat([df,dummies],axis=1)
# -
# ### address
# **Let's feature engineer a zip code column from the address in the data set and Create a column called 'zip_code' that extracts the zip code from the address column.**
df['zip_code'] = df['address'].apply(lambda address:address[-5:])
# **Now make this zip_code column into dummy variables using pandas. Concatenate the result and drop the original zip_code column along with dropping the address column.**
dummies = pd.get_dummies(df['zip_code'],drop_first=True)
df = df.drop(['zip_code','address'],axis=1)
df = pd.concat([df,dummies],axis=1)
# ### issue_d
#
# **This would be data leakage, we wouldn't know beforehand whether or not a loan would be issued when using our model, so in theory we wouldn't have an issue_date, drop this feature.**
df = df.drop('issue_d',axis=1)
# ### earliest_cr_line
# **This appears to be a historical time stamp feature. So, Extracting the year from this feature using a .apply function, then convert it to a numeric feature. And Setting this new data to a feature column called 'earliest_cr_year', then drop the earliest_cr_line feature.**
df['earliest_cr_year'] = df['earliest_cr_line'].apply(lambda date:int(date[-4:]))
df = df.drop('earliest_cr_line',axis=1)
df.select_dtypes(['object']).columns
# ## Train Test Split
# **Importing train_test_split from sklearn.**
from sklearn.model_selection import train_test_split
# **droping the load_status column we created earlier, since its a duplicate of the loan_repaid column. We'll use the loan_repaid column since its already in 0s and 1s.**
df = df.drop('loan_status',axis=1)
# **Set X and y variables to the .values of the features and label.**
X = df.drop('loan_repaid',axis=1).values
y = df['loan_repaid'].values
# ----
# ----
#
# ## Grabbing a Sample for Training Time
#
# ### Using .sample() to grab a sample of the 490k+ entries to save time on training.
#
# ----
# ----
# df = df.sample(frac=0.1,random_state=101)
print(len(df))
# **Performing a train/test split with test_size=0.2 and a random_state of 101.**
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=101)
# ## Normalizing the Data
#
# **Using a MinMaxScaler to normalize the feature data X_train and X_test.**
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# # Creating the Model
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation,Dropout
from tensorflow.keras.constraints import max_norm
# **Building a sequential model to will be trained on the data. We have unlimited options here, but here is what the solution uses: a model that goes 78 --> 39 --> 19--> 1 output neuron.**
model = Sequential()
# +
model = Sequential()
# input layer
model.add(Dense(78, activation='relu'))
model.add(Dropout(0.2))
# hidden layer
model.add(Dense(39, activation='relu'))
model.add(Dropout(0.2))
# hidden layer
model.add(Dense(19, activation='relu'))
model.add(Dropout(0.2))
# output layer
model.add(Dense(units=1,activation='sigmoid'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer='adam')
# -
# **Fitting the model to the training data for at least 25 epochs. Also adding in the validation data for later plotting.**
model.fit(x=X_train,
y=y_train,
epochs=25,
batch_size=256,
validation_data=(X_test, y_test),
)
# **Saving the model.**
from tensorflow.keras.models import load_model
model.save('full_data_project_model.h5')
# # Section 3: Evaluating Model Performance.
#
# **Plotting out the validation loss versus the training loss.**
losses = pd.DataFrame(model.history.history)
losses[['loss','val_loss']].plot()
# **Creating predictions from the X_test set and display a classification report and confusion matrix for the X_test set.**
from sklearn.metrics import classification_report,confusion_matrix
predictions = model.predict_classes(X_test)
print(classification_report(y_test,predictions))
confusion_matrix(y_test,predictions)
# **Given the customer below, would you offer this person a loan?**
# +
import random
random.seed(101)
random_ind = random.randint(0,len(df))
new_customer = df.drop('loan_repaid',axis=1).iloc[random_ind]
new_customer
# -
model.predict_classes(new_customer.values.reshape(1,78))
# **Now checking, did this person actually end up paying back their loan?**
df.iloc[random_ind]['loan_repaid']
| Risk minimization for loan investments .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="UvYDZmH_xHPG"
# # Generative Adversarial Networks
#
# This code is based on [_Generative Adversarial Networks (2014)_](https://arxiv.org/abs/1406.2661) paper by <NAME>, <NAME>, et all.
#
# 
# -
# ## Environment Setup
# %load_ext pycodestyle_magic
# %flake8_on
# + colab={} colab_type="code" id="ShCO79wAxHPP"
import torch
# device configuration
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print("using %s" % (device))
# + [markdown] colab_type="text" id="cu6vT-jDxHPT"
# ## Prepare Dataset
# + colab={} colab_type="code" id="gU5JFzeGxHPU"
from torchvision import datasets, transforms
# setup MNIST dataset
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize([0.5], [0.5])])
mnist_dataset = datasets.MNIST(
root="./datasets/mnist", train=True, transform=transform, download=True)
mnist_width = mnist_dataset.train_data.size(1)
mnist_height = mnist_dataset.train_data.size(2)
mnist_shape = [1, mnist_width, mnist_height]
# data loader (input pipeline)
data_loader = torch.utils.data.DataLoader(
dataset=mnist_dataset, batch_size=100, shuffle=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 551} colab_type="code" executionInfo={"elapsed": 2014, "status": "ok", "timestamp": 1572647285437, "user": {"displayName": "<NAME> 1306365266", "photoUrl": "", "userId": "17835414831372489696"}, "user_tz": -420} id="Vn1wgnIkxHPX" outputId="eed50711-1a19-419d-ce5b-f8d0dc323ac6"
import matplotlib.pyplot as plt
examples = enumerate(data_loader)
_, (example_data, _) = next(examples)
# plot example data
for i in range(6):
plt.subplot(2, 3, i+1)
plt.tight_layout()
plt.imshow(example_data[i][0], cmap='gray', interpolation='none')
plt.title("Example %d" % i)
plt.xticks([])
plt.yticks([])
plt.show()
# + [markdown] colab_type="text" id="UJ_q5BflxHPc"
# ## Build Models
#
# 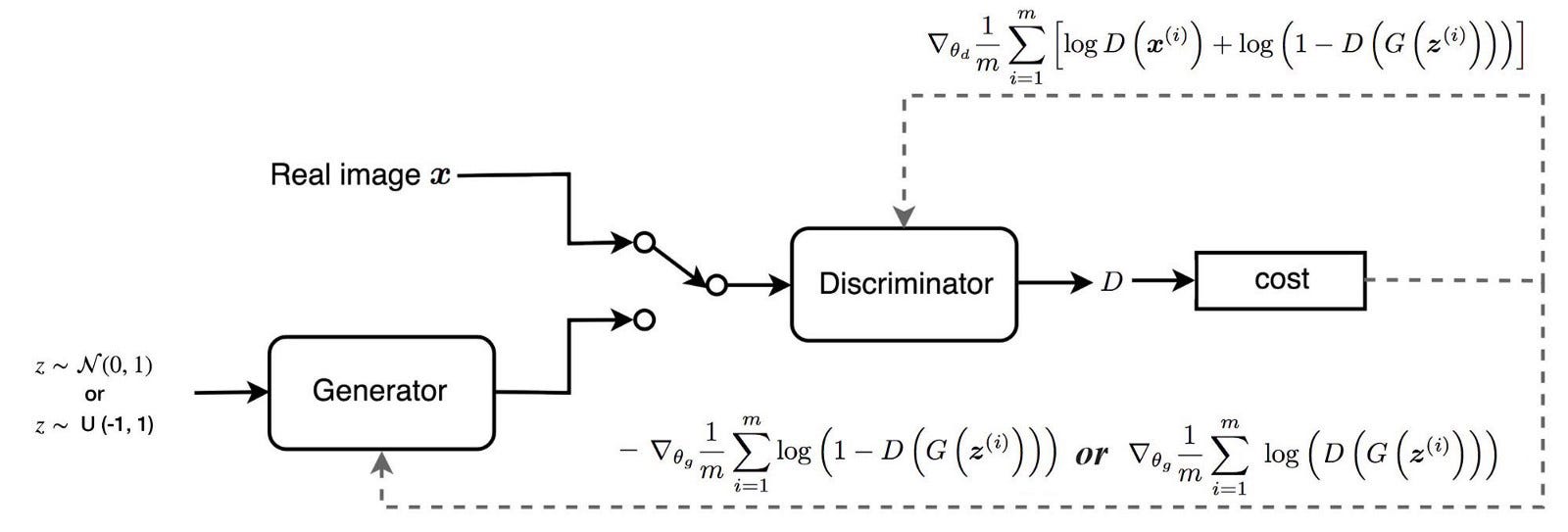
# source: https://medium.com/@jonathan_hui/gan-whats-generative-adversarial-networks-and-its-application-f39ed278ef09
# +
import torch.nn as nn
def nn_block(in_feat, out_feat, normalize=False):
layers = [nn.Linear(in_feat, out_feat)]
if normalize:
layers.append(nn.BatchNorm1d(out_feat, 0.8))
layers.append(nn.LeakyReLU(0.2, inplace=True))
return layers
# + colab={} colab_type="code" id="NddMdZ_mxHPd"
# generator model
class Generator(nn.Module):
def __init__(self, in_dim, out_dim):
super(Generator, self).__init__()
self.model = nn.Sequential(
*nn_block(in_dim, 128),
*nn_block(128, 256, normalize=True),
*nn_block(256, 512, normalize=True),
*nn_block(512, 1024, normalize=True),
nn.Linear(1024, out_dim),
nn.Tanh())
def forward(self, noise):
output = self.model(noise)
image = output.view(output.size(0), *mnist_shape)
return image
# -
# discriminator model
class Discriminator(nn.Module):
def __init__(self, in_dim):
super(Discriminator, self).__init__()
self.model = nn.Sequential(
*nn_block(in_dim, 512),
*nn_block(512, 256),
nn.Linear(256, 1),
nn.Sigmoid())
def forward(self, image):
img_flat = image.view(image.size(0), -1)
validity = self.model(img_flat)
return validity
# + colab={"base_uri": "https://localhost:8080/", "height": 72} colab_type="code" executionInfo={"elapsed": 4316, "status": "ok", "timestamp": 1572647287772, "user": {"displayName": "<NAME> 1306365266", "photoUrl": "", "userId": "17835414831372489696"}, "user_tz": -420} id="jNo6QGSAxHPg" outputId="3e926661-bc38-47fb-e5f7-9dbfa41fc825"
noise_dim = 100
mnist_dim = mnist_width * mnist_height
# build model
generator = Generator(in_dim=noise_dim, out_dim=mnist_dim).to(device)
discriminator = Discriminator(in_dim=mnist_dim).to(device)
print(generator, discriminator)
# + [markdown] colab_type="text" id="pw6SuIw-xHPm"
# ## Train Process
#
# ### Discriminator Update
#
# 
#
# ### Generator Update
#
# #### Before:
#
# 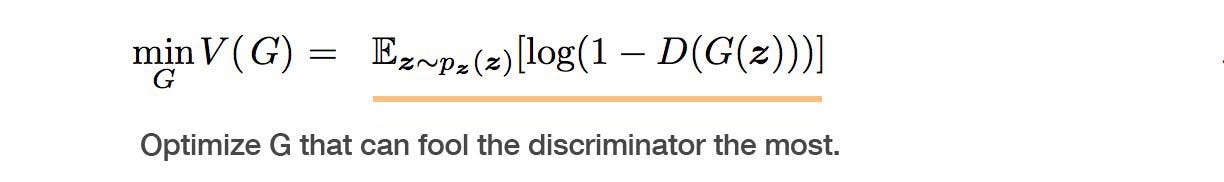
#
# #### Because Generator diminished gradient:
#
# In practice, equation 1 may not provide sufficient gradient for G to learn well. Early in learning, when G is poor,D can reject samples with high confidence because they are clearly different fromthe training data. In this case, log(1−D(G(z))) saturates. Rather than training G to minimize log(1−D(G(z))) we can train G to maximize logD(G(z)). This objective function results in thesame fixed point of the dynamics of G and D but provides much stronger gradients early in learning. (GAN Paper)
#
# 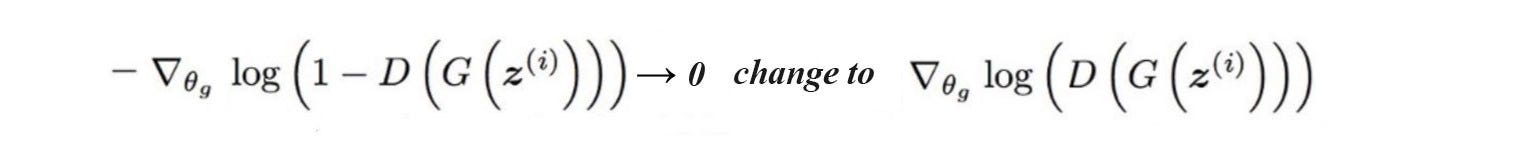
#
# 
# + colab={} colab_type="code" id="v0jzHUZ7xHPn"
# loss criterion
criterion = nn.BCELoss().to(device)
# optimizers
g_optimizer = torch.optim.Adam(
generator.parameters(), lr=0.0002, betas=(0.5, 0.999))
d_optimizer = torch.optim.Adam(
discriminator.parameters(), lr=0.0002, betas=(0.5, 0.999))
if torch.cuda.is_available():
FloatTensor = torch.cuda.FloatTensor
else:
FloatTensor = torch.FloatTensor
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 64298, "status": "error", "timestamp": 1572646592735, "user": {"displayName": "<NAME> 1306365266", "photoUrl": "", "userId": "17835414831372489696"}, "user_tz": -420} id="NX2eNdar64Xs" outputId="2f8fbc72-9810-47e4-d071-0b66d98a4729" tags=[]
from torch.autograd import Variable
from torchvision.utils import save_image
import numpy as np
import time
import os
start = time.time()
g_losses = []
d_losses = []
# train process
epochs = 200
for epoch in range(epochs):
g_loss_sum = 0
d_loss_sum = 0
for i, (imgs, _) in enumerate(data_loader):
batch_size = imgs.size(0)
# adversarial ground truths
valid = Variable(FloatTensor(batch_size, 1).fill_(1.0),
requires_grad=False)
fake = Variable(FloatTensor(batch_size, 1).fill_(0.0),
requires_grad=False)
# configure input
real_imgs = Variable(imgs.type(FloatTensor))
# -----------------
# Train Generator
# -----------------
g_optimizer.zero_grad()
# sample noise as generator input
noise = Variable(FloatTensor(
np.random.normal(0, 1, (batch_size, noise_dim))))
# generate a batch of images
gen_imgs = generator(noise)
# loss measures generator's ability to fool the discriminator
# g_loss = criterion(discriminator(gen_imgs), fake) # Normal MinMax
g_loss = criterion(discriminator(gen_imgs), valid) # Non Saturated
g_loss_sum += g_loss.item()
g_loss.backward()
g_optimizer.step()
# ---------------------
# Train Discriminator
# ---------------------
d_optimizer.zero_grad()
# loss measures discriminator's ability to classify real
# from generated samples
real_loss = criterion(discriminator(real_imgs), valid)
fake_loss = criterion(discriminator(gen_imgs.detach()), fake)
d_loss = (real_loss + fake_loss) / 2
d_loss_sum += d_loss.item()
d_loss.backward()
d_optimizer.step()
elapsed = time.time() - start
print("\r[Elapsed %02d:%02d]" % (elapsed // 60, elapsed % 60), end='')
print("\r[Epoch %d/%d] [Batch %d/%d]"
% (epoch+1, epochs, i+1, len(data_loader)), end='')
print(" [G loss: %f] [D loss: %f]"
% (g_loss.item(), d_loss.item()), end='')
g_losses.append(g_loss_sum / len(data_loader))
d_losses.append(d_loss_sum / len(data_loader))
if (epoch + 1) % 10 == 0:
# sample noise as generator input
noise = Variable(FloatTensor(
np.random.normal(0, 1, (5 * 5, noise_dim))))
# generate a batch of sample images
gen_imgs = generator(noise)
# save sample images
os.makedirs("./samples/gan", exist_ok=True)
save_image(gen_imgs, "./samples/gan/sample_%03d.png" % (epoch + 1),
nrow=5, normalize=True)
# save result models
os.makedirs("./models/gan", exist_ok=True)
torch.save(generator, "./models/gan/generator.pt")
torch.save(discriminator, "./models/gan/discriminator.pt")
# -
# ## Train Result
# plot losses
plt.plot(g_losses, label="generator loss")
plt.plot(d_losses, label="discriminator loss")
plt.legend()
plt.show()
# +
# sample noise as generator input
noise = Variable(FloatTensor(np.random.normal(0, 1, (6, noise_dim))))
# generate a batch of sample images
gen_imgs = generator(noise)
# plot sample images
imgs = gen_imgs.cpu().detach().numpy()
for i in range(6):
plt.subplot(2, 3, i+1)
plt.tight_layout()
plt.imshow(imgs[i][0], cmap='gray', interpolation='none')
plt.title("Sample %d" % i)
plt.xticks([])
plt.yticks([])
plt.show()
| 1 - Fundamental of GANs/GAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2A.soft - Git depuis le notebook
#
# [Git](http://fr.wikipedia.org/wiki/Git) est un logiciel de suivi de source décentralisé qui permet de travailler à plusieurs sur les mêmes fichiers. Aujourd'hui, on ne crée plus de logiciels sans ce type d'outil qui permet de garder l'historique des modifications apportées à un programme. [Git](http://fr.wikipedia.org/wiki/Git) a supplanté tous les autres logiciels du même type.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
import os
root = os.path.abspath('.')
# On vérifie qu'il fonctionne :
# !git
# Si cela ne fonctionne pas, cela signifie que soit [git](http://git-scm.com/) n'est pas installé, soit le notebook ne trouve pas le chemin où il est installé. Dans le premier cas, il suffit de l'installer puis de relancer le serveur de notebook pour prendre en compte les modifications. Dans le second cas, on peut s'en tirer avec une astuce du style :
# !set PATH=%PATH%;chemin_git
# Dans son utilisation la plus simple, *git* permet de synchroniser un ensemble de fichiers entre plusieurs ordinateurs et utilisateurs :
from pyquickhelper.helpgen import NbImage
NbImage("gits.png")
# Par la suite, il faudra remplacer ``SERVEURGIT`` par le nom du serveur *git* (le serveur de l'école, github, bitbucket, ...).
# ## Création d'un répertoire
#
# On crée un projet afin de pouvoir y ajouter des fichiers et de suivre leur modifications. Cette opération ne devra être réalisé qu'une seule fois. On crée ce projet dans le répertoire : ``c:\temp\premier_projet``. On crée ce répertoire puis on change le répertoire courant pour celui-ci :
import os
folder = r"premier_projet3"
if not os.path.exists(folder):
os.mkdir(folder)
# On initialise le répertoire :
# !git init premier_projet3
# ## Premiers fichiers
# On ajoute un premier fichier ``README.md`` :
# +
# %%file premier_projet3/README.md
Premier Projet
==============
* quelques listes
# -
# On copie ce notebook également :
# !copy git_notebook.ipynb premier_projet3
# Git est beaucoup plus facile à utiliser si on se place dans le repértoire du projet :
import os
os.chdir("premier_projet3")
# Il faut préciser à *git* les fichiers qu'on souhaite ajouter où dont on souhaite enregister les modifications :
# !git add README.md
# !git add git_notebook.ipynb
# On enregistre ces modifications avec un commentaire :
# !git commit -m "premier fichier et premier notebook"
# Ces modifications n'ont d'incidence que sur la copie en locale et n'ont pas encore été propagées jusqu'au serveur. On le fait en exécutant les deux instructions suivantes. La première précise dans quel répertoire du serveur *git* on souhaite ajouter les modifications. Il suffit en principe de ne le faire qu'une fois.
# !git remote add origin https://dupre@SERVEURGIT/dupre/premier_projet3.git
# Si vous vous êtes trompé, vous pouvez revenir en arrière avec :
# !git remote remove origin
# La seconde propage les modifications :
# !git push -u origin master
# Si vous avez l'erreur précédente, c'est parce que le répertoire du projet n'a pas été créé sur le serveur git. Je vous conseille de le faire directement sur le serveur lui-même (petit icône + en haut à droite sur un serveur [GitLab](https://about.gitlab.com/)) :
from pyquickhelper.helpgen import NbImage
NbImage(os.path.join(root, "gitsp.png"))
# On recommence, de préférence dans la fenêtre de ligne de commande car sinon il faudra taper le mot de passe depuis la fenêtre du serveur de notebook et ce n'est pas toujours évident de ne pas se tromper.
# !git push -u origin master
# Les fichiers devrait être sur le serveur. Pour enregistrer d'autres modifications faite dans le répertoire local, il suffit de recommencer à partir de l'instruction ``git add``.
# ## Cloner un répertoire
#
# Pour récupérer les fichiers chez soi. Il faut cloner le répertoire, par exemple dans un autre répertoire local.
import os
folder = r"projet3_copy"
if not os.path.exists(folder):
os.mkdir(folder)
# Puis taper la commande dans ce répertoire (de préférence depuis la ligne de commande s'il y a besoin de s'authentifier).
# !git clone https://SERVEURGIT/dupre/premier_projet3.git/
import os
os.listdir(r"projet3_copy")
# Pour apporter les modifications sur le serveur *git*, il faut retourner aux instructions ``git add`` vues précédemment.
# ## Mettre à jour un répertoire local
#
# Lorqu'on met à jour le serveur distant *git* depuis une copie locale, il est préférable de mettre à jour les autres copies locales avant de pouvoir les modifier. Il vaut mieux exécuter la commande depuis une ligne de commande s'il faut s'authentifier.
# !git pull https://SERVEURGIT/dupre/premier_projet3.git/
# ## Branches
#
# Le scénario classique pour travailler à plusieurs. On suppose qu'il existe un *repository* remote appelé *origin*. C'est le cas le plus classique quand on clone un répertoire de sources.
#
# * On met à jour la branche principal : ``git pull`` (local = remote)
# * On crée une branche local : ``git checkout -b name/branch``
# * On fait sa modification.
# * On commit. ``git add`` + ``git commit -m "..."``
# * On met à jour le remote en créant aussi une branche là-bas : ``git push -u origin name/branch``.
# * On envoie une [pull request](https://help.github.com/articles/about-pull-requests/).
# * Le détenteur de la branche principale accepte ou vous demande des modifications.
# * On détruit la branche une fois que les modifications sont faites ``git branch -d name/branch``.
#
# Si la branche ``master`` est mise à jour pendant que vous travaillez à la vôtre, ``git pull origin master`` rappatriera les modifications.
# ## Pour aller plus loin
# On peut faire beaucoup de choses avec *git* comme créer des branches, des tags, revenir à en arrière... La documentation sur git est assez pléthorique sur internet [tutoriels sur git](http://sixrevisions.com/resources/git-tutorials-beginners/) et il existe des [client git](http://git-scm.com/download/gui/linux). [TortoiseGit](https://code.google.com/p/tortoisegit/) est l'un des plus simples. Ces outils permettent de se servir de *git* sans ligne de commande, juste avec la souris et l'exporateur de fichiers.
| _doc/notebooks/2a/git_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/TashreefMuhammad/University_Miscellaneous_Codes/blob/main/Matrix_Factorization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="aMAJAFV0UX4H"
# # <font color = 'skyblue'>CSE 4238 : Soft Computing Lab</font>
#
# + [markdown] id="BSb79YW9gv5o"
# ## <font color = 'navyblue'>Assignment 01: Matrix Factorization</font>
#
#
# + [markdown] id="KswXlDazgxqL"
# ### Download Data
# Download the Data from drive
# + id="jgEAXzoADX-F"
# Install gdown
#======== For Kaggle and / or Colab
# ! pip install gdown
# Download index_id.csv from the url =>
# https://drive.google.com/file/d/1ENbB5srTy8_y5qDvOVNwHjAXgPW6BJiZ/view?usp=sharing
# !gdown --id 1ENbB5srTy8_y5qDvOVNwHjAXgPW6BJiZ
# # Download Matrix_Factorization_Assignment.csv from the url =>
# https://drive.google.com/file/d/1ENbB5srTy8_y5qDvOVNwHjAXgPW6BJiZ/view?usp=sharing
# !gdown --id 1lzG<KEY>
# + [markdown] id="343KUt9vgmmM"
# ### Import Libraries
#
# Import required libraries
# + id="WxASYOp4FDvS"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import mean_squared_error
from sklearn.metrics.pairwise import cosine_similarity
# + [markdown] id="igWhRHYRg2Cx"
# ### Extracting Data to DataFrame
#
# Retirieving data from CSV and saving them in DataFrame
# + id="5CK6JwORFGHp" outputId="b8281601-e5a1-4771-a437-fd25102e7afc"
indexDataFrame = pd.read_csv('index_id.csv')
# infoArray = indexDataFrame[indexDataFrame['Student Id'] == 170104014].values
# print(infoArray)
ind_id = indexDataFrame[indexDataFrame['Student Id'] == 170104014].values[0][2]
print(ind_id)
data = pd.read_csv('Matrix_Factorization_Assignment.csv')
data
# + [markdown] id="aPSxT9Dbv2EW"
# ### Preparing Data for Use
#
# * Copy All Data into another DataFrame
# * Remove the Index Column from the new DataFrame
# * Find Rows and Columns that should be removed and then drop those [Rows](https://www.geeksforgeeks.org/how-to-drop-rows-in-pandas-dataframe-by-index-labels/) and [Columns](https://www.geeksforgeeks.org/how-to-drop-one-or-multiple-columns-in-pandas-dataframe/)
# + id="3lphXv_LyeM3" outputId="37efbadf-0150-4ed3-a0e7-dbe21dc03873"
my_data = data.copy()
# print(my_data.columns)
# my_data = my_data.drop(['Index'], axis = 1)
# print(my_data.columns)
# del data
# print(len(my_data['Index'].unique()))
dropList = [my_data['Index'][i] for i in range(0, my_data.shape[0]) if int(my_data['Index'][i]) % ind_id == 0]
dropRowList = [my_data.index[i] for i in range(0, my_data.shape[0]) if int(my_data['Index'][i]) % ind_id == 0]
my_data.drop(dropRowList, inplace = True)
# print(len(dropList))
# print(dropList)
# print(sorted(dropList))
i = 0
while i < 45000:
i *= 261
if i not in dropList:
print(i, 'not found')
i += 1
print(min(dropList))
print(max(dropList))
my_data = my_data.drop(['Index'], axis = 1)
dropColList = [i for i in range(1, my_data.shape[1]) if i % ind_id == 0]
my_data.drop(my_data.columns[dropColList], axis = 1, inplace = True)
print(sum(my_data.isna().sum()))
my_data
# + [markdown] id="B84Qp5lZxmuz"
# Given index_id = 261
#
# Real Number of Rows = 45000
#
# So, Found Number of Rows that should be Removed $= 169$
#
#
# Real Number of Columns = 5000 <font color = 'red'>(Excluding Index Column) </font>
#
# So, Number of Columns that should be Removed $= \lfloor \frac{5000}{261} \rfloor = 19$
#
#
# Hence, after removing rows and column, the final DataFrame size should be:
#
# $(45000 - 169) \times (5000 - 19) \boldsymbol{\Rightarrow} 44831 \times 4981$
# + [markdown] id="cUGYUt0rhMO6"
# ### Getting NumPy Array
#
# Converting data to **NumPy** array for future processing
# + id="3m_cwUa22ON6" outputId="af30556b-dc8b-4b57-ca0c-6d2d16ac5c3b"
# tmp_data = np.array(my_data)
# print(tmp_data)
# del tmp_data
my_data = my_data.replace(np.nan, -1)
my_data = np.array(my_data, dtype = np.float32)
print(my_data)
print(sum(sum(my_data)))
my_data += 1.
print(my_data)
print(sum(sum(my_data)))
# + [markdown] id="TPlNP4MnhX0z"
# ### Define Useful Methods
#
# Define useful methods for the algorithm
# + id="wxD1OieVGYGj"
def updateMatrix(givenData, staticData, regulizer, K):
partA = (staticData.T).dot(staticData) + regulizer * np.eye(k)
partB = givenData.dot(staticData)
return partB.dot(np.linalg.inv(partA))
def lossCalc(trueData, predData):
indUse = np.nonzero(trueData)
# print(type(indUse))
errorLib = mean_squared_error(trueData[indUse], predData[indUse])
# ==== Comparison returned 7.083543050729694e-23 difference
# ==== between library value and calculated value. So, only
# ==== library value is taken for calculation
# errorCus = 0
# num = 0
# for i, j in nonZeroInd:
# errorCus += (trueData[i][j] - predData[i][j]) ** 2
# num += 1
# for i in range(trueData.shape[0]):
# for j in range(trueData.shape[1]):
# if trueData[i][j] != 0.:
# errorCus += (trueData[i][j] - predData[i][j]) ** 2
# num += 1
# errorCus /= num
# if errorLib != errorCus:
# print((errorLib - errorCus) ** 2)
return errorLib
# return errorCus
# + [markdown] id="ytki4Zlxhgnh"
# ### Matrix Factorization
#
# The main coding module for **Matrix Factorization**
# + id="4bSBncOyVEJ0" outputId="b5c72974-22d2-4e92-df0d-4393c9f97091"
N, M = my_data.shape
errorVals = []
loopK = [100, 1000, 2000, 2500]
for k in loopK:
print('Running on K = ' + str(k))
errorVal = []
V = np.zeros((M, k), dtype = np.float32)
np.random.seed(14)
U = np.random.random((N, k)).astype(dtype = np.float32)
U /= sum(sum(U))
# print(U)
# print(sum(sum(U)))
lambdaU = (0.00015 + 0.0001 * (ind_id % 8))
lambdaV = (0.00025 - 0.0001 * (ind_id % 7))
for _ in range(151):
# print(U.shape, V.shape)
V = updateMatrix(my_data.T, U, lambdaV, k)
# print(U.shape, V.shape)
U = updateMatrix(my_data, V, lambdaU, k)
# print(U.shape, V.shape)
errorVal.append(lossCalc(my_data, U.dot(V.T)))
plt.plot(errorVal, label = 'For the Value of K = ' + str(k))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
errorVals.append(errorVal)
for i, eVal in enumerate(errorVals):
plt.plot(eVal, label = 'K = ' + str(loopK[i]))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
# + id="oiomXe6mTAkc" outputId="0ec7643f-b6a9-48fc-b8b8-4d1be257b6c6"
N, M = my_data.shape
loopK = [3000, 4000, 5000]
for k in loopK:
print('Running on K = ' + str(k))
errorVal = []
V = np.zeros((M, k), dtype = np.float32)
np.random.seed(14)
U = np.random.random((N, k)).astype(dtype = np.float32)
U /= sum(sum(U))
# print(U)
# print(sum(sum(U)))
lambdaU = (0.00015 + 0.0001 * (ind_id % 8))
lambdaV = (0.00025 - 0.0001 * (ind_id % 7))
for _ in range(150):
# print(U.shape, V.shape)
V = updateMatrix(my_data.T, U, lambdaV, k)
# print(U.shape, V.shape)
U = updateMatrix(my_data, V, lambdaU, k)
# print(U.shape, V.shape)
loss = lossCalc(my_data, U.dot(V.T))
errorVal.append(loss)
if loss < 0.001:
print('Loss has reached a very small value, breaking training')
break
plt.plot(errorVal, label = 'For the Value of K = ' + str(k))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
errorVals.append(errorVal)
for i, eVal in enumerate(errorVals):
plt.plot(eVal, label = 'K = ' + str(loopK[i]))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
# + id="lKAEqViHTAkd" outputId="403eca7e-1add-4e34-9193-037f8089a45c"
N, M = my_data.shape
loopK = [5000]
for k in loopK:
print('Running on K = ' + str(k))
errorVal = []
V = np.zeros((M, k), dtype = np.float32)
np.random.seed(14)
U = np.random.random((N, k)).astype(dtype = np.float32)
U /= sum(sum(U))
# print(U)
# print(sum(sum(U)))
lambdaU = (0.00015 + 0.0001 * (ind_id % 8))
lambdaV = (0.00025 - 0.0001 * (ind_id % 7))
for _ in range(150):
# print(U.shape, V.shape)
V = updateMatrix(my_data.T, U, lambdaV, k)
# print(U.shape, V.shape)
U = updateMatrix(my_data, V, lambdaU, k)
# print(U.shape, V.shape)
loss = lossCalc(my_data, U.dot(V.T))
errorVal.append(loss)
if loss < 0.001:
print('Loss has reached a very small value, breaking training')
plt.plot(errorVal, label = 'For the Value of K = ' + str(k))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
errorVals.append(errorVal)
for i, eVal in enumerate(errorVals):
plt.plot(eVal, label = 'K = ' + str(loopK[i]))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
# + id="38boRVQ4TAke" outputId="e498af4e-e8a2-4623-e0<PASSWORD>"
loopK = [100, 1000, 2000, 2500, 3000, 4000, 5000]
for i, eVal in enumerate(errorVals):
plt.plot(eVal, label = 'K = ' + str(loopK[i]))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
loopK = [100, 1000, 2000, 2500, 3000, 4000, 5000]
for i, eVal in enumerate(errorVals):
if i < 3:
continue
plt.plot(eVal, label = 'K = ' + str(loopK[i]))
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
plt.show()
# + id="aLoXK9XRTAkh"
u_df = pd.DataFrame(U)
v_df = pd.DataFrame(V)
u_df.to_csv('U_Values.csv', index = False)
v_df.to_csv('V_Values.csv', index = False)
# + id="aCtGL4ZQTAkh"
for i, eVal in enumerate(errorVals):
with open(str(loopK[i]) + '.txt', 'w') as fle:
fle.write(str(eVal))
# + id="4BKV8CbMTAkh" outputId="df3735e1-be50-4987-8501-5216560c5012"
# del data
# del u_df
# del v_df
similarity_u = cosine_similarity(U)
similarity_v = cosine_similarity(V)
print(similarity_u.shape)
print(similarity_u)
print(similarity_v.shape)
print(similarity_v)
similarity_u_df = pd.DataFrame(similarity_u)
similarity_v_df = pd.DataFrame(similarity_v)
similarity_u_df.to_csv('Similarity_U.csv', index = False)
similarity_v_df.to_csv('Similarity_V.csv', index = False)
# + id="p8OpJ_h2TAki"
# cosine_sim_u = pd.DataFrame(columns = ['U -> 1', 'U -> 2', 'Cosine Similarity'])
# for i in range(U.shape[0]):
# for j in range(i + 1, U.shape[0]):
# sim = cosine_similarity([U[i, :]], [U[j, :]])
# val = [i, j, sim]
# cosine_sim_u.loc[len(cosine_sim_u)] = val
# # print(type(sim))
# # print(sim.shape)
# cosine_sim_u.to_csv('Cosine Similarity U.csv', index = False)
# + id="jLzUrRCPTAkp" outputId="b71e4682-1ac9-4463-b09a-cc08852f1ff1"
cosine_sim_v = pd.DataFrame(columns = ['V -> 1', 'V -> 2', 'Cosine Similarity'])
for i in range(V.shape[0]):
for j in range(i + 1, V.shape[0]):
sim = cosine_similarity([V[i, :]], [V[j, :]])
val = [i, j, sim]
cosine_sim_v.loc[len(cosine_sim_v)] = val
# print(type(sim))
# print(sim.shape)
cosine_sim_v.to_csv('Cosine Similarity V.csv', index = False)
# + id="r5Bgnv5XTAkq"
cosine_sim_v.to_csv('Cosine Similarity V.csv', index = False)
# + id="yHgMm0mETAkq"
U = pd.read_csv('U_Values.csv')
V = pd.read_csv('V_Values.csv')
U = np.array(U)
V = np.array(V)
# + id="3b4fBo3TTAkr"
# cnt = 0
# val = U.dot(V.T)
# print(val.shape)
# for i in range(my_data.shape[0]):
# for j in range(my_data.shape[1]):
# if my_data[i][j] == 0:
# print(val[i][j])
# if val[i][j] >= 1.0:
# print('Review of Movie ', j , 'by user 11071 is', val[0][j] - 1)
# cnt += 1
# if cnt == 5:
# break
# print("End Row")
# + [markdown] id="9bAAI9_p1NHX"
# <br><br>
# <center><font color = 'yellow'>$\LARGE{\text{The END}}$</font><center>
| Soft Computing/Matrix_Factorization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# <a id='introduction'></a>
# # Introduction
#
# The first example showcases the use of the XAITK-Saliency API to generate visual saliency maps using models from the [scikit-learn](https://scikit-learn.org/stable/) library trained on the MNIST dataset.
# The MNIST dataset contains grayscale images of handwritten digits (0-9), which are normalized and centered in the frame.
# It was developed to evaluate the performance of models classifying individual handwritten digits.
#
# [This example](https://scikit-learn.org/stable/auto_examples/linear_model/plot_sparse_logistic_regression_mnist.html) from scikit-learn's website uses their `LogisticRegression` class to achieve fairly high accuracy on the MNIST dataset with very short training time.
# As is shown by this example, it is easy to visualize the decision boundaries of each class by simply plotting their respective model's coefficients in the same dimensions as the input image.
#
# We use the XAITK-Saliency high-level API to mimic this visualization by creating saliency maps for several images from the dataset and averaging them to create a global decision-boundary representation for each class.
# This approach achieves comparable results to those shown in scikit-learn's example while requiring zero knowledge of the intrinsic properties of the model used.
#
# We do the same while using the `MLPClassifier` class also from the scikit-learn library.
# Our model is taken from [another example](https://scikit-learn.org/stable/auto_examples/neural_networks/plot_mnist_filters.html#sphx-glr-auto-examples-neural-networks-plot-mnist-filters-py) on their website.
# The example shows a visualization of the MLP's weights as images to gain insight on the learning behavior of the model.
# These images, however, only show generic patterns that the model has learned and therefore the behavior of each class is still obscure.
#
# Using our approach provides per-class saliency representations and also gives some insight on the different learning behaviors portrayed by both these models.
#
# ### Table of Contents
# * [MNIST Dataset Example](#MNIST-Dataset-Example-mnist)
# * [Set Up Environment](#Set-Up-Environment-mnist)
# * [Downloading the Dataset](#Download-the-Dataset-mnist)
# * [The "Application"](#The-Application-mnist)
# * [Logistic Regression Example](#Logistic-Regression-Example-mnist)
# * [Fitting the Model](#Fitting-the-Model-logistic-mnist)
# * [Black-box Classifier](#Black-box-Classifier-logistic-mnist)
# * [Heatmap Generation](#Heatmap-Generation-mnist)
# * [Calling the Application](#Calling-the-Application-logistic-mnist)
# * [MLP Example](#MLP-Example-mnist)
# * [Fitting the Model](#Fitting-the-Model-mlp-mnist)
# * [Black-box Classifier](#Black-box-Classifier-mlp-mnist)
# * [Calling the Application](#Calling-the-Application-mlp-mnist)
#
# <br>
#
# To run this notebook in Colab, use the link below:
#
# [](https://colab.research.google.com/github/XAITK/xaitk-saliency/blob/master/examples/MNIST_scikit_saliency.ipynb)
# + [markdown] tags=[]
# # MNIST Dataset Example <a name="MNIST-Dataset-Example-mnist"></a>
# ## Set Up Environment <a name="Set-Up-Environment-mnist"></a>
# -
# !pip install -qU pip
# !pip install -q xaitk-saliency
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# ## Downloading the Dataset <a name="Download-the-Dataset-mnist"></a>
#
# The MNIST dataset consists of 70,000 28x28 grayscale images of handwritten numbers.
# Each image is stored as a column vector, resulting in a (70000,784) shape for the entire dataset.
# +
from sklearn.datasets import fetch_openml
import matplotlib.pyplot as plt
import numpy as np
import os
cwd = os.getcwd()
data_dir = cwd + '/scikit-learn-example'
# Load data from https://www.openml.org/d/554
X, y = fetch_openml('mnist_784', version=1, return_X_y=True, as_frame=False, data_home=data_dir)
X = X/X.max()
# Find examples of each class
ref_inds = []
for i in range(10):
ref_inds.append(np.where(np.int64(y) == i)[0][0])
ref_imgs = X[ref_inds]
# Plot examples
plt.figure(figsize=(15,5))
for i in range(10):
plt.subplot(1, 10, i + 1)
plt.imshow(ref_imgs[i].reshape(28, 28), 'gray')
plt.axis('off')
# + [markdown] tags=[]
# ## The "Application" <a name="The-Application-mnist"></a>
#
# Our "application" will accept a set of images, a black-box image classifier, and a saliency generator and will generate saliency maps for each image provided.
# The saliency maps from the first image in the set will then be plotted to give an idea of the model's behavior on a single sample.
#
# Additionally, because all digits in the MNIST dataset are centered in the frame, we can average all the heatmaps generated for each respective class to produce a decision boundary visualization.
# The application will do this and plot the resulting averaged heatmaps for each digit class.
# This should compare to what is shown in the [first example](https://scikit-learn.org/stable/auto_examples/linear_model/plot_sparse_logistic_regression_mnist.html) discussed in the [introduction](#introduction).
# +
from smqtk_classifier import ClassifyImage
from xaitk_saliency import GenerateImageClassifierBlackboxSaliency
def app(
images: np.ndarray,
image_classifier: ClassifyImage,
saliency_generator: GenerateImageClassifierBlackboxSaliency
):
# Generate saliency maps
sal_maps_set = []
for img in images:
ref_image = img.reshape(28, 28)
sal_maps = saliency_generator(ref_image, image_classifier)
sal_maps_set.append(sal_maps)
num_classes = sal_maps_set[0].shape[0]
# Plot first image in set with saliency maps
plt.figure(figsize=(10,5))
plt.suptitle("Heatmaps for First Image", fontsize=16)
num_cols = np.ceil(num_classes/2).astype(int) + 1
plt.subplot(2, num_cols, 1)
plt.imshow(images[0].reshape(28,28), cmap='gray')
plt.xticks(())
plt.yticks(())
for c in range(num_cols - 1):
plt.subplot(2, num_cols, c + 2)
plt.imshow(sal_maps_set[0][c], cmap=plt.cm.RdBu, vmin=-1, vmax=1)
plt.xticks(())
plt.yticks(())
plt.xlabel(f"Class {c}")
for c in range(num_classes - num_cols + 1, num_classes):
plt.subplot(2, num_cols, c + 3)
plt.imshow(sal_maps_set[0][c], cmap=plt.cm.RdBu, vmin=-1, vmax=1)
plt.xticks(())
plt.yticks(())
plt.xlabel(f"Class {c}")
# Average heatmaps for each respective class
global_maps = np.sum(sal_maps_set, axis=0) / len(images)
# Plot average maps
plt.figure(figsize=(10, 5))
plt.suptitle("Average Heatmaps from All Images", fontsize=16)
for c in range(num_classes):
vcap = np.absolute(global_maps[i]).max()
plt.subplot(2, num_cols-1, c + 1)
plt.imshow(global_maps[c], cmap=plt.cm.RdBu, vmin=-vcap, vmax=vcap)
plt.xticks(())
plt.yticks(())
plt.xlabel(f"Class {c}")
# -
# # Logistic Regression Example <a name="Logistic-Regression-Example-mnist"></a>
# ## Fitting the Model <a name="Fitting-the-Model-logistic-mnist"></a>
#
# We take the same `LogisticRegression` object used in the scikit-learn example and fit it to a subset of the dataset.
# Here, an L2 penalty and a larger training set is used to yield slightly better results than those shown in the example.
# The same visualization of the coefficients is shown with these new parameters.
# +
import time
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
t0 = time.time()
# Split data into test and train sets
train_samples = 20000
X_train, X_test, y_train, y_test = train_test_split(
X, y, train_size=train_samples, test_size=10000, random_state=0)
# Define model, lower C value gives higher regulation
clf = LogisticRegression(
C=50. / train_samples, penalty='l2', solver='saga', tol=0.1, random_state=0
)
# Fit model
clf.fit(X_train, y_train)
# Score model
score = clf.score(X_test, y_test)
print("Test score with L2 penalty: %.4f" % score)
# Visualize coefficients
coef = clf.coef_.copy()
max_val = np.abs(coef).max()
plt.figure(figsize=(10, 5))
for i in range(10):
p = plt.subplot(2, 5, i + 1)
p.imshow(coef[i].reshape(28, 28), cmap=plt.cm.RdBu,
vmin=-max_val, vmax=max_val)
p.set_xticks(())
p.set_yticks(())
p.set_xlabel('Class %i' % i)
plt.suptitle('Classification vector for...')
run_time = time.time() - t0
print('Example run in %.3f s' % run_time)
plt.show()
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# ## Black-box Classifier <a name="Black-box-Classifier-logistic-mnist"></a>
#
# Here we wrap our `LogisticRegression` object in [SMQTK-Classifier's](https://smqtk-classifier.readthedocs.io/en/stable/classifier_interfaces.html#classifyimage) `ClassifyImage` class to comply with the API's interface.
# +
class MNISTClassifierLog (ClassifyImage):
def get_labels(self):
return list(range(10))
def classify_images(self, image_iter):
# Yes, "images" in this example case are really 1-dim (28*28=784).
# MLP input needs a (n_samples, n_features) matrix input.
images = np.asarray(list(image_iter)) # may fail because input is not consistent in shape
images = images.reshape(-1, 28*28) # may fail because input was not the correct shape
return (
{idx: p_i for idx, p_i in zip(range(10), p)}
for p in clf.decision_function(images)
)
# Required for implementation
def get_config(self):
return {}
image_classifier_log = MNISTClassifierLog()
# -
# ## Heatmap Generation <a name="Heatmap-Generation-mnist"></a>
#
# We create an instance of `SlidingWindowStack`, an implementation of the `GenerateImageClassifierBalckboxSaliency` interface, to carry out our image perturbation and heatmap generation.
# +
from xaitk_saliency.impls.gen_image_classifier_blackbox_sal.slidingwindow import SlidingWindowStack
gen_slidingWindow = SlidingWindowStack(
window_size=(2, 2),
stride=(1, 1),
threads=4
)
# -
# ## Calling the Application <a name="Calling-the-Application-logistic-mnist"></a>
#
# Finally, we call the application using the first 20 images in the MNIST dataset.
# Here the blue is showing positive saliency while the red is showing negative saliency.
#
# Even with a small set of images, the general shape of the digits is visible.
# Using a larger set of images should improve the visualized decision boundaries, but scales the computation time linearly.
#
# These results are largely expected when linear models like logistic regression are used.
# Each pixel from the image corresponds to a single coefficient in each of the respective classes' regressions.
# Occluding a pixel in the image should affect the output of the model, and therefore the resulting saliency maps, proportional to the value of the corresponding coefficients.
# We therefore expect the saliency maps to largely match the pattern of the learned coefficients.
app(
X[0:20],
image_classifier_log,
gen_slidingWindow
)
# # MLP Example <a name="MLP-Example-mnist"></a>
# ## Fitting the Model <a name="Fitting-the-Model-mlp-mnist"></a>
#
# Following the second [example](https://scikit-learn.org/stable/auto_examples/neural_networks/plot_mnist_filters.html#sphx-glr-auto-examples-neural-networks-plot-mnist-filters-py) from scikit-learn, we training an `MLPClassifier` on the MNIST dataset using the same hyperparameters.
#
# To shorten training time, the MLP has only one hidden layer with 50 nodes, and is only trained for 10 iterations, meaning the model does not converge.
# +
import warnings
from sklearn.exceptions import ConvergenceWarning
from sklearn.neural_network import MLPClassifier
# use the traditional train/test split
X_train, X_test = X[:60000], X[60000:]
y_train, y_test = y[:60000], y[60000:]
mlp = MLPClassifier(hidden_layer_sizes=(50,), max_iter=10, alpha=1e-4,
solver='sgd', verbose=10, random_state=1,
learning_rate_init=.1)
# this example won't converge because of CI's time constraints, so we catch the
# warning and ignore it here
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=ConvergenceWarning,
module="sklearn")
mlp.fit(X_train, y_train)
print("Training set score: %f" % mlp.score(X_train, y_train))
print("Test set score: %f" % mlp.score(X_test, y_test))
# -
# ## Black-box Classifier <a name="Black-box-Classifier-mlp-mnist"></a>
#
# We wrap our `MLPClassifier` object in [SMQTK-Classifier's](https://smqtk-classifier.readthedocs.io/en/stable/classifier_interfaces.html#classifyimage) `ClassifyImage` class to comply with the API's interface.
# +
class MNISTClassifierMLP (ClassifyImage):
def get_labels(self):
return list(range(10))
def classify_images(self, image_iter):
# Yes, "images" in this example case are really 1-dim (28*28=784).
# MLP input needs a (n_samples, n_features) matrix input.
images = np.asarray(list(image_iter)) # may fail because input is not consistent in shape
images = images.reshape(-1, 28*28) # may fail because input was not the correct shape
return (
{idx: p_i for idx, p_i in zip(range(10), p)}
for p in mlp.predict_proba(images)
)
# Required for implementation
def get_config(self):
return {}
image_classifier_mlp = MNISTClassifierMLP()
# -
# ## Calling the Application <a name="Calling-the-Application-mlp-mnist"></a>
#
# We call our application again using the same image set and saliency generator, but this time using our MLP classifier.
#
# The results show mostly negative saliency, suggesting that the MLP model has learned where the pixels are *absent* for each class more than where they are *present*.
app(
X[0:20],
image_classifier_mlp,
gen_slidingWindow
)
| examples/MNIST_scikit_saliency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pierredumontel/plane-classification/blob/main/notebooks/train_classification_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="14pFhpzH8dOf"
# Données tabulaires (reg logisitique -> pas approprié pour vecteur de coordonnées d'images)
# Pytorch (Facebook) vs tensorflow (Google) (pour les NN complexes)
#
# Pytorch + facile a utilisé en termes de structure de réseaux
#
# Keras : sur couche des 2 autres
#
# pep8
# + colab={"base_uri": "https://localhost:8080/"} id="-uJ8uTyQeOrs" outputId="391aab69-ecd2-4fb7-edf7-837a538143e5"
# !curl -O https://www.robots.ox.ac.uk/~vgg/data/fgvc-aircraft/archives/fgvc-aircraft-2013b.tar.gz
# !tar zxf fgvc-aircraft-2013b.tar.gz
# !mv fgvc-aircraft-2013b/ dataset
# + id="QnpVWmm3bXH9"
import keras
# + id="pfpp_mvybZIL" outputId="b8506fca-b15c-4f84-bc6d-aad1581a5f7c" colab={"base_uri": "https://localhost:8080/", "height": 35}
keras.__version__
# + [markdown] id="D_JkBFqTqUFb"
# # Imports
# + id="KhR21y5DePbH"
import pathlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from sklearn.model_selection import train_test_split
#from keras.utils import to_categorical
from keras.models import Sequential, load_model
from keras.layers import Conv2D, MaxPool2D, Dense, Flatten, Dropout
from PIL import Image
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ZDo7Zx0kekcP" outputId="e5027fc8-9dd4-4aae-e617-d969daee98fe"
pd.__version__
# + id="hNzZ_OVWeVkE"
DATA_DIR = pathlib.Path('dataset/data')
# + [markdown] id="-FPKx20IlC2T"
# ### Make assumptions about number of spaces (here=2)
# + colab={"base_uri": "https://localhost:8080/"} id="wzQuyfYGeYsf" outputId="9b6da9e9-119e-4c33-ce86-49c26771e547"
manufacturer_df = pd.read_csv(DATA_DIR / 'images_manufacturer_train.txt',sep=' ',
names=['image_id','m1','m2'],
usecols=['image_id','m1','m2'],#usecols for v1.4 compatibility
dtype={'image_id':str}) #ids are not int but string
manufacturer_df['manufacturer']=manufacturer_df['m1']+' '+manufacturer_df['m2']
manufacturer_df['manufacturer'].unique()
# + id="l7uHWg2DobuI"
# ! grep ',' dataset/data/images_manufacturer_train.txt
# + colab={"base_uri": "https://localhost:8080/"} id="4XxBqOT0o5F3" outputId="5804d4c7-cb27-48e1-812c-ea452317f12f"
#Recherche le caractère T dans le fichier et n'affiche que trois lignes (head -3)
# ! grep 'T' dataset/data/images_manufacturer_train.txt | head -3
# + colab={"base_uri": "https://localhost:8080/"} id="7vx44TBNpSyn" outputId="967ad121-a28a-4be6-f2da-12909f88d782"
#wc : Compte le nombre d'élements (-l : ligne, -c : caractère, -w : word)
# ! grep 'T' dataset/data/images_manufacturer_train.txt | wc
# + colab={"base_uri": "https://localhost:8080/"} id="kvsBg2wzqKAS" outputId="e37e1567-b142-45f3-e1dc-471909c950bc"
# !cut -f 1 -d' ' dataset/data/images_manufacturer_train.txt | head
# + colab={"base_uri": "https://localhost:8080/"} id="9uZKuFqglhV3" outputId="f9c61cad-acf1-4d9d-999e-8dc0bdf60fa6"
manufacturer_df = pd.read_csv(DATA_DIR / 'images_manufacturer_train.txt',sep='\t',
names=['all'],
dtype={'all':str}) #ids are not int but string
#La fonction split() découpe sur une chaîne de caractères
manufacturer_df['image_id'] = manufacturer_df['all'].apply(lambda x:x.split(' ')[0])
#La fonction '<car>'.join(liste) concatène les éléments de liste en utillisant le séparateur <car>
manufacturer_df['manufacturer']=manufacturer_df['all'].apply(lambda x:' '.join(x.split(' ')[1:]))
manufacturer_df['manufacturer'].unique()
# + [markdown] id="1UARRuQxh_B0"
# Verify data
# + colab={"base_uri": "https://localhost:8080/"} id="Lpv4gx_8ewt_" outputId="5291200c-771d-4e5c-f3e5-66301273eb51"
manufacturer_df['manufacturer'].value_counts(dropna=False)
# + [markdown] id="63Ze9CM9iC6f"
# Verify missing data
# + colab={"base_uri": "https://localhost:8080/"} id="m50meUezhkRD" outputId="0dda1de8-8843-43b7-b5e0-667f2500cc23"
manufacturer_df.isna().sum()
# + id="4bpDNnYmiIaI"
assert manufacturer_df['image_id'].isna().sum()==0,"Missing value in image_id"
assert manufacturer_df['manufacturer'].isna().sum()==0,"Missing value in manufacturer"
# + id="KCfnCd1liiJ4" outputId="5666d21e-eda0-4103-af0c-13f59c3ee31e" colab={"base_uri": "https://localhost:8080/", "height": 206}
manufacturer_df.head()
# + id="HQjt0nJv_RAx"
manufacturer_df['path'] = manufacturer_df['image_id'].apply(lambda x: pathlib.Path('dataset/data/images') / (x + '.jpg'))
# + id="sWPCupQaAd3q" outputId="6ca8d8fd-814c-4b1d-c40a-6d0533a7aadc" colab={"base_uri": "https://localhost:8080/", "height": 215}
manufacturer_df.head()
# + id="tjSmZ04TAgPI"
def build_image_database(path):
"""Build a pandas dataframe with target class and access path to images.
Parameters
----------
path (Path) : path pattern to read csv file containing images information.
Returns
-------
pandas dataframe, including target class and path to image.
"""
manufacturer_df = pd.read_csv(path,sep='\t',
names=['all'],
dtype={'all':str}) #ids are not int but string
#La fonction split() découpe sur une chaîne de caractères
manufacturer_df['image_id'] = manufacturer_df['all'].apply(lambda x:x.split(' ')[0])
#La fonction '<car>'.join(liste) concatène les éléments de liste en utillisant le séparateur <car>
manufacturer_df['manufacturer']=manufacturer_df['all'].apply(lambda x:' '.join(x.split(' ')[1:]))
#La colonne path contient le chemin d'accès à l'image
manufacturer_df['path'] = manufacturer_df['image_id'].apply(lambda x: pathlib.Path('dataset/data/images') / (x + '.jpg'))
return manufacturer_df
# + id="roWjI9HUBoYA" outputId="451247b8-2912-4300-f26b-09ab0a538564" colab={"base_uri": "https://localhost:8080/", "height": 424}
build_image_database(DATA_DIR / 'images_manufacturer_train.txt')
# + id="iRVXnzirEPqo"
# mieux que rien
def prt():
""" je fais ceci dans ma fonction
"""
return
# + id="KeFfg7JPE9TF"
def build_image_database(path,target):
"""Build a pandas dataframe with target class and access path to images.
Parameters
----------
path (Path) : path pattern to read csv file containing images information.
target (str) : name of the target column
Returns
-------
pandas dataframe, including target class and path to image.
"""
_df = pd.read_csv(path,sep='\t',
names=['all'],
dtype={'all':str}
) #ids are not int but string
#La fonction split() découpe sur une chaîne de caractères
_df['image_id'] = _df['all'].apply(lambda x:x.split(' ')[0])
#La fonction '<car>'.join(liste) concatène les éléments de liste en utillisant le séparateur <car>
_df[target]= _df['all'].apply(lambda x:' '.join(x.split(' ')[1:]))
#La colonne path contient le chemin d'accès à l'image
_df['path'] = _df['image_id'].apply(lambda x: pathlib.Path('dataset/data/images') / (x + '.jpg'))
return _df.drop(columns=['all'])
# + id="Mn0JTiasGwCB" outputId="bc2ce50b-d263-4bd7-bf84-cf0976c9ec65" colab={"base_uri": "https://localhost:8080/", "height": 112}
build_image_database(DATA_DIR / 'images_manufacturer_train.txt','manufacturer').head(2)
# + id="t5VByDsuHIYl" outputId="96644af8-6e4b-4c65-dfa6-f0c7705d9ba7" colab={"base_uri": "https://localhost:8080/", "height": 112}
build_image_database(DATA_DIR / 'images_manufacturer_train.txt','family').head(2)
# + id="EUzAgNU-Hvvm" outputId="838fd898-867e-4510-e353-22b368771282" colab={"base_uri": "https://localhost:8080/", "height": 112}
build_image_database(DATA_DIR / 'images_manufacturer_train.txt','variant').head(2)
# + id="YwrlIlroILrd"
def build_image_database(path, target) :
"""Build a pandas dataframe with target class and access path to images.
Parameters
----------
path (Path) : path pattern to read csv file containing images information.
target (str) : name of the target column.
Returns
-------
A pandas dataframe, including target class and path to image.
"""
_df =pd.read_csv(path, sep='\t'
, names=['all']
, dtype ={'all':str} # id are string not int
)
#split donne une chaine de caractères
_df['image_id'] = _df['all'].apply(lambda x: x.split(' ')[0])
#'<car>'.join(liste) concatène les éléments de la liste avec le séparateur '<car>'
_df[target] = _df['all'].apply(lambda x: ' '.join(x.split(' ')[1:]))
# La colonne path contient le chemin d'accès à l'image
_df['path'] = _df['image_id'].apply(lambda x: pathlib.Path('dataset/data/images') / (x +'.jpg'))
return _df.drop(columns=['all'])
# + id="ec-Zqr1XI1DP" outputId="7090213f-5ae3-45d2-e588-6711cd10f736" colab={"base_uri": "https://localhost:8080/", "height": 112}
build_image_database(DATA_DIR / 'images_manufacturer_train.txt','variant').head(2)
# + [markdown] id="rfge2o2VJMln"
# baselib : module qui viennent systématiquement avec Python,
#
# import os
# import pathlib
# import datetime
# + id="K_kyi1VVI2lX" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="05e3e956-b6ac-47b0-a85e-3ba5c7c91b3a"
manufacturer_df = build_image_database(DATA_DIR / 'images_manufacturer_train.txt','manufacturer')
manufacturer_df.head()
# + id="oDBtzEsoKwXi"
#On récupère un chemin d'accès
manufacturer_df.head(1)['path'].values[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="4kQ_MTs7Lk-d" outputId="1f348c92-6916-44ea-b8a4-c91daa188c5d"
plt.imshow(plt.imread(manufacturer_df.head(1)['path'].values[0]))
# + id="ih84oYsdMrrX"
#La fonction :
# -prend un df (argument)
# -prend une ligne (row : argument)
# -prend une colonne (target : argument)
# -elle affiche la classe (la valeur de target) et l'image associée, pour la ligne en argument
def show_image(df,row, target):
"""Show an image from an image database, with the associated class.
Parameters
----------
df (pd.DataFrame) : images definition dataframe
row (int) : row index in df of image to be displayed
target (str) :name of the target column
Returns
-------
None
"""
assert target in df.columns, "Missing target columns in dataframe"
assert 'path' in df.columns, 'Missing image path in dataframe'
print(df.iloc[row,][target])
plt.imshow(plt.imread(df.iloc[row,]['path']))
return
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="H1a_AigkOyTT" outputId="2d128b00-390a-4a05-e297-0ab83a56118c"
show_image(manufacturer_df,42,'manufacturer')
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="lJTvnCDNO7ef" outputId="b187b3b3-e5e6-494f-a25c-ec7422346250"
show_image(build_image_database(DATA_DIR / 'images_manufacturer_train.txt','manufacturer'),12,'manufacturer')
# + id="MQDG2rzEialY" outputId="4bd6201e-8996-4e07-dc1f-168b21b9ed19" colab={"base_uri": "https://localhost:8080/"}
manufacturer_df.shape
# + id="zqS7Yf0XjR2S" outputId="dbc83ee9-f395-4d6c-9549-f1c89a7c2d0a" colab={"base_uri": "https://localhost:8080/"}
plt.imread(manufacturer_df.head(1)['path'].values[0])
# + id="7pi4sSi5janR" outputId="0b6eef43-037d-444a-8d7d-9e43c4ea0a0f" colab={"base_uri": "https://localhost:8080/"}
manufacturer_df.head(10)['path'].apply(lambda p: plt.imread(p).shape)
# + id="oOnBYJ10m08e"
manufacturer_df['image_shape'] = manufacturer_df['path'].apply(lambda p:plt.imread(p).shape)
# + id="GnoXL_SvnBev" outputId="ccb634fe-57cb-4cc4-ee8d-3eb7b2651634" colab={"base_uri": "https://localhost:8080/"}
manufacturer_df['image_shape'].apply(lambda x:x[1]).value_counts()
# + id="OK3weE5pn7Du"
IMAGE_WIDTH = 128
IMAGE_HEIGHT = IMAGE_WIDTH
# + id="gY2qEaI8oTQB"
images_list = []
def load_resize_image(path,width, height):
"""Load an image and resize it to the target size
Parameters
----------
path (Path):access path to image file
height (int):resize image to this height
width (int):resize image to this width
Returns
-------
np.array containing resized image
"""
return np.array(Image.open(path).resize((width, height)))
# + id="q9xr1XNMqiod"
manufacturer_df['resized_image'] = manufacturer_df.apply(lambda r: load_resize_image(r['path'], IMAGE_WIDTH, IMAGE_HEIGHT), axis=1)
# + id="8VhtQeXlrsRu" outputId="2f317b4e-eb4d-4d88-9232-dcc7489531a8" colab={"base_uri": "https://localhost:8080/", "height": 285}
plt.imshow(manufacturer_df.iloc[42,]['resized_image'])
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="dKeFq3suRqcn" outputId="37f1e804-4a30-485a-89b6-5f387c8b6d0c"
#Building the model
model = Sequential()
model.add(Conv2D(filters=32, kernel_size=(5,5), activation='relu', input_shape=manufacturer_df.shape[1:]))
model.add(Conv2D(filters=32, kernel_size=(5,5), activation='relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.25))
model.add(Conv2D(filters=64, kernel_size=(3, 3), activation='relu'))
model.add(Conv2D(filters=64, kernel_size=(3, 3), activation='relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.25))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(43, activation='softmax'))
#Compilation of the model
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="mikuOHmInnEJ"
| notebooks/train_classification_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# # Image Classification with Vision Transformer
#
# **Author:** [<NAME>](https://www.linkedin.com/in/khalid-salama-24403144/)<br>
# **Date created:** 2021/01/18<br>
# **Last modified:** 2021/01/18<br>
# **Description:** Implementing the Vision Transformer (ViT) model for image classification.
# + [markdown] colab_type="text"
# ## Introduction
#
# This example implements the [Vision Transformer (ViT)](https://arxiv.org/abs/2010.11929)
# model by <NAME> et al. for image classification,
# and demonstrates it on the CIFAR-100 dataset.
# The ViT model applies the Transformer architecture with self-attention to sequences of
# image patches, without using convolution layers.
#
# This example requires TensorFlow 2.4 or higher, as well as
# [TensorFlow Addons](https://www.tensorflow.org/addons/overview),
# which can be installed using the following command:
#
# ```python
# pip install -U tensorflow-addons
# ```
# + [markdown] colab_type="text"
# ## Setup
# + colab_type="code"
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import tensorflow_addons as tfa
# + [markdown] colab_type="text"
# ## Prepare the data
# + colab_type="code"
num_classes = 100
input_shape = (32, 32, 3)
(x_train, y_train), (x_test, y_test) = keras.datasets.cifar100.load_data()
print(f"x_train shape: {x_train.shape} - y_train shape: {y_train.shape}")
print(f"x_test shape: {x_test.shape} - y_test shape: {y_test.shape}")
# + [markdown] colab_type="text"
# ## Configure the hyperparameters
# + colab_type="code"
learning_rate = 0.001
weight_decay = 0.0001
batch_size = 256
num_epochs = 100
image_size = 72 # We'll resize input images to this size
patch_size = 6 # Size of the patches to be extract from the input images
num_patches = (image_size // patch_size) ** 2
projection_dim = 64
num_heads = 4
transformer_units = [
projection_dim * 2,
projection_dim,
] # Size of the transformer layers
transformer_layers = 8
mlp_head_units = [2048, 1024] # Size of the dense layers of the final classifier
# + [markdown] colab_type="text"
# ## Use data augmentation
# + colab_type="code"
data_augmentation = keras.Sequential(
[
layers.experimental.preprocessing.Normalization(),
layers.experimental.preprocessing.Resizing(image_size, image_size),
layers.experimental.preprocessing.RandomFlip("horizontal"),
layers.experimental.preprocessing.RandomRotation(factor=0.02),
layers.experimental.preprocessing.RandomZoom(
height_factor=0.2, width_factor=0.2
),
],
name="data_augmentation",
)
# Compute the mean and the variance of the training data for normalization.
data_augmentation.layers[0].adapt(x_train)
# + [markdown] colab_type="text"
# ## Implement multilayer perceptron (MLP)
# + colab_type="code"
def mlp(x, hidden_units, dropout_rate):
for units in hidden_units:
x = layers.Dense(units, activation=tf.nn.gelu)(x)
x = layers.Dropout(dropout_rate)(x)
return x
# + [markdown] colab_type="text"
# ## Implement patch creation as a layer
# + colab_type="code"
class Patches(layers.Layer):
def __init__(self, patch_size):
super(Patches, self).__init__()
self.patch_size = patch_size
def call(self, images):
batch_size = tf.shape(images)[0]
patches = tf.image.extract_patches(
images=images,
sizes=[1, self.patch_size, self.patch_size, 1],
strides=[1, self.patch_size, self.patch_size, 1],
rates=[1, 1, 1, 1],
padding="VALID",
)
patch_dims = patches.shape[-1]
patches = tf.reshape(patches, [batch_size, -1, patch_dims])
return patches
# + [markdown] colab_type="text"
# Let's display patches for a sample image
# + colab_type="code"
import matplotlib.pyplot as plt
plt.figure(figsize=(4, 4))
image = x_train[np.random.choice(range(x_train.shape[0]))]
plt.imshow(image.astype("uint8"))
plt.axis("off")
resized_image = tf.image.resize(
tf.convert_to_tensor([image]), size=(image_size, image_size)
)
patches = Patches(patch_size)(resized_image)
print(f"Image size: {image_size} X {image_size}")
print(f"Patch size: {patch_size} X {patch_size}")
print(f"Patches per image: {patches.shape[1]}")
print(f"Elements per patch: {patches.shape[-1]}")
n = int(np.sqrt(patches.shape[1]))
plt.figure(figsize=(4, 4))
for i, patch in enumerate(patches[0]):
ax = plt.subplot(n, n, i + 1)
patch_img = tf.reshape(patch, (patch_size, patch_size, 3))
plt.imshow(patch_img.numpy().astype("uint8"))
plt.axis("off")
# + [markdown] colab_type="text"
# ## Implement the patch encoding layer
#
# The `PatchEncoder` layer will linearly transform a patch by projecting it into a
# vector of size `projection_dim`. In addition, it adds a learnable position
# embedding to the projected vector.
# + colab_type="code"
class PatchEncoder(layers.Layer):
def __init__(self, num_patches, projection_dim):
super(PatchEncoder, self).__init__()
self.num_patches = num_patches
self.projection = layers.Dense(units=projection_dim)
self.position_embedding = layers.Embedding(
input_dim=num_patches, output_dim=projection_dim
)
def call(self, patch):
positions = tf.range(start=0, limit=self.num_patches, delta=1)
encoded = self.projection(patch) + self.position_embedding(positions)
return encoded
# + [markdown] colab_type="text"
# ## Build the ViT model
#
# The ViT model consists of multiple Transformer blocks,
# which use the `layers.MultiHeadAttention` layer as a self-attention mechanism
# applied to the sequence of patches. The Transformer blocks produce a
# `[batch_size, num_patches, projection_dim]` tensor, which is processed via an
# classifier head with softmax to produce the final class probabilities output.
#
# Unlike the technique described in the [paper](https://arxiv.org/abs/2010.11929),
# which prepends a learnable embedding to the sequence of encoded patches to serve
# as the image representation, all the outputs of the final Transformer block are
# reshaped with `layers.Flatten()` and used as the image
# representation input to the classifier head.
# Note that the `layers.GlobalAveragePooling1D` layer
# could also be used instead to aggregate the outputs of the Transformer block,
# especially when the number of patches and the projection dimensions are large.
# + colab_type="code"
def create_vit_classifier():
inputs = layers.Input(shape=input_shape)
# Augment data.
augmented = data_augmentation(inputs)
# Create patches.
patches = Patches(patch_size)(augmented)
# Encode patches.
encoded_patches = PatchEncoder(num_patches, projection_dim)(patches)
# Create multiple layers of the Transformer block.
for _ in range(transformer_layers):
# Layer normalization 1.
x1 = layers.LayerNormalization(epsilon=1e-6)(encoded_patches)
# Create a multi-head attention layer.
attention_output = layers.MultiHeadAttention(
num_heads=num_heads, key_dim=projection_dim, dropout=0.1
)(x1, x1)
# Skip connection 1.
x2 = layers.Add()([attention_output, encoded_patches])
# Layer normalization 2.
x3 = layers.LayerNormalization(epsilon=1e-6)(x2)
# MLP.
x3 = mlp(x3, hidden_units=transformer_units, dropout_rate=0.1)
# Skip connection 2.
encoded_patches = layers.Add()([x3, x2])
# Create a [batch_size, projection_dim] tensor.
representation = layers.LayerNormalization(epsilon=1e-6)(encoded_patches)
representation = layers.Flatten()(representation)
representation = layers.Dropout(0.5)(representation)
# Add MLP.
features = mlp(representation, hidden_units=mlp_head_units, dropout_rate=0.5)
# Classify outputs.
logits = layers.Dense(num_classes)(features)
# Create the Keras model.
model = keras.Model(inputs=inputs, outputs=logits)
return model
# + [markdown] colab_type="text"
# ## Compile, train, and evaluate the mode
# + colab_type="code"
def run_experiment(model):
optimizer = tfa.optimizers.AdamW(
learning_rate=learning_rate, weight_decay=weight_decay
)
model.compile(
optimizer=optimizer,
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[
keras.metrics.SparseCategoricalAccuracy(name="accuracy"),
keras.metrics.SparseTopKCategoricalAccuracy(5, name="top-5-accuracy"),
],
)
checkpoint_filepath = "/tmp/checkpoint"
checkpoint_callback = keras.callbacks.ModelCheckpoint(
checkpoint_filepath,
monitor="val_accuracy",
save_best_only=True,
save_weights_only=True,
)
history = model.fit(
x=x_train,
y=y_train,
batch_size=batch_size,
epochs=num_epochs,
validation_split=0.1,
callbacks=[checkpoint_callback],
)
model.load_weights(checkpoint_filepath)
_, accuracy, top_5_accuracy = model.evaluate(x_test, y_test)
print(f"Test accuracy: {round(accuracy * 100, 2)}%")
print(f"Test top 5 accuracy: {round(top_5_accuracy * 100, 2)}%")
return history
vit_classifier = create_vit_classifier()
history = run_experiment(vit_classifier)
# + [markdown] colab_type="text"
# After 100 epochs, the ViT model achieves around 55% accuracy and
# 82% top-5 accuracy on the test data. These are not competitive results on the CIFAR-100 dataset,
# as a ResNet50V2 trained from scratch on the same data can achieve 67% accuracy.
#
# Note that the state of the art results reported in the
# [paper](https://arxiv.org/abs/2010.11929) are achieved by pre-training the ViT model using
# the JFT-300M dataset, then fine-tuning it on the target dataset. To improve the model quality
# without pre-training, you can try to train the model for more epochs, use a larger number of
# Transformer layers, resize the input images, change the patch size, or increase the projection dimensions.
# Besides, as mentioned in the paper, the quality of the model is affected not only by architecture choices,
# but also by parameters such as the learning rate schedule, optimizer, weight decay, etc.
# In practice, it's recommended to fine-tune a ViT model
# that was pre-trained using a large, high-resolution dataset.
| examples/vision/ipynb/image_classification_with_vision_transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mujoco150
# language: python
# name: mujoco150
# ---
from matplotlib.pylab import plt
import numpy as np
# +
def K_kernel(x):
#x = np.linalg.norm(x)
K = -1 / (np.exp(x/0.1) + 2 + np.exp(-x/0.1))
return K
def K_kernel2(x):
#x = np.linalg.norm(x)
x = np.clip(x, -10, 10)
K = -1 / (np.exp(x / 0.2) + np.exp(-x / 0.2))
return K
# +
c_v1 = 60*0.01
c_v2 = 1
x = np.linspace(0,1, 100)
# -
y = -c_v1 * K_kernel(c_v2 * (x))
# +
plt.plot(x,y)
#@#plt.ylim([0,2])
# +
def K_kernel3(x):
x = np.linalg.norm(x)
K = -1 / (np.exp(x/0.1) + 2 + np.exp(-x/0.1))
return K
def reward7(velocity_base, v_commdand):
v_e = np.concatenate((velocity_base[:2], velocity_base[-1:])) - v_commdand # x, y, yaw
vxy = v_commdand[:2]
wyaw = v_commdand[2]
dt =0.01
#reward calculate
kc = 1
c_w = 150* dt
c_v1 = 200 * dt
c_v2 = 1
lin_vel_cost = c_v1 * K_kernel3(c_v2 * (velocity_base[:2] - vxy))
ang_vel_cost = c_w * K_kernel3(velocity_base[-1] -wyaw )
return -lin_vel_cost, -ang_vel_cost
# -
v_b = np.array([-0.0001462446,-0.0053383102,-0.0456031509,0.0720960647,0.0809975155,0.0101319291])
v_command = np.array([-0.3,0,0])
# +
lin_reward , ang_reward = reward7(v_b, v_command)
print(lin_reward, ang_reward)
# -
# +
import gym
import numpy as np
from my_envs.mujoco import *
from my_envs.base.command_generator import command_generator
from matplotlib.pylab import plt
from evaluate.plot_results import *
# +
env = gym.make('CellrobotEnvCPG-v0') # Swimmer2-v2 SpaceInvaders-v0 CellrobotEnv-v0
#env = gym.wrappers.Monitor(env, 'tmp/tmp.mp4', force=True)
print('state: ', env.observation_space)
print('action: ', env.action_space)
q_dim = 1
command = command_generator(10000, 0.01, 2, vx_range=(-0.2, 0.2), vy_range = (0,0), wyaw_range = (0,0))
reward_fun = 1
obs = env.reset(command = command, reward_fun_choice = reward_fun)
print('test')
max_step = 1000
# while True:
# env.render()
v_e = []
c_command = []
xyz = []
rewards = []
action = np.ones(39)* ( 1)
for i in range(max_step):
# env.render()
#action = env.action_space.sample()
next_obs, reward, done, infos = env.step(action)
obs = next_obs
v_e.append(infos['velocity_base'])
c_command.append(infos['commands'])
xyz.append(infos['obs'][:3])
rewards.append(infos['rewards'])
#env.render(mode='rgb_array')#mode='rgb_array'
env.close()
dt =0.01
v_e = np.array(v_e)
c_command = np.array(c_command)
xyz = np.array(xyz)
# -
plot_velocity_curve(v_e, c_command, max_step, dt = 0.01, figsize=(8,6))
plot_position_time(xyz, max_step, dt = 0.01, figsize=(8,6))
plot_traj_xy(xyz, max_step, dt = 0.01, figsize=(8,6))
rewards
# +
rewards = np.array(rewards)
#forward_reward, ctrl_cost, contact_cost, survive_reward
plt.plot(rewards[:,0])
# +
forward_reward = np.abs(rewards[:,1]/rewards[:,0]) * np.sign(rewards[:,1])
ctrl_cost = np.abs(rewards[:,2]/rewards[:,0]) * np.sign(rewards[:,2])
contact_cost = np.abs(rewards[:,3]/rewards[:,0]) * np.sign(rewards[:,3])
survive_reward = np.abs(rewards[:,4]/rewards[:,0]) * np.sign(rewards[:,4])
plt.figure(figsize=(18,6))
plt.plot(forward_reward, label='for')
plt.plot(ctrl_cost, label='ct')
plt.plot(contact_cost, label='con')
plt.plot(survive_reward, label='sur')
plt.grid()
plt.legend()
#plt.bar(range(forward_reward.shape[0]), forward_reward)
# +
import os
os.chdir('/home/drl/PycharmProjects/rl_baselines/my_baselines')
import gym
import numpy as np
from my_envs.mujoco import *
from my_envs.base.command_generator import command_generator
from matplotlib.pylab import plt
from evaluate.plot_results import *
def plot_fitness_t(reward_fun):
env = gym.make('CellrobotEnvCPG-v0') # Swimmer2-v2 SpaceInvaders-v0 CellrobotEnv-v0
print('state: ', env.observation_space)
print('action: ', env.action_space)
command = command_generator(10000, 0.01, 2, vx_range=(-0.2, 0.2), vy_range = (0,0), wyaw_range = (0,0))
obs = env.reset( command, reward_fun)
max_step = 1000
v_e = []
c_command = []
xyz = []
rewards = []
action = np.ones(39)* ( 1)
for i in range(max_step):
# env.render()
#action = env.action_space.sample()
next_obs, reward, done, infos = env.step(action)
obs = next_obs
v_e.append(infos['velocity_base'])
c_command.append(infos['commands'])
xyz.append(infos['obs'][:3])
rewards.append(infos['rewards'])
#env.render(mode='rgb_array')#mode='rgb_array'
env.close()
dt =0.01
v_e = np.array(v_e)
c_command = np.array(c_command)
xyz = np.array(xyz)
plt.figure(figsize=(18,6))
rewards_duty = []
for i in range(rewards.shape[1]-1):
rewards_duty.append(np.abs(rewards[:,1+i]/rewards[:,0]) * np.sign(rewards[:,1+i]))
plt.plot(rewards_duty[i], label=str(i))
plt.grid()
plt.legend()
plt.show()
# -
plot_fitness_t(1)
# +
env = gym.make('CellrobotEnvCPG-v0') # Swimmer2-v2 SpaceInvaders-v0 CellrobotEnv-v0
print('state: ', env.observation_space)
print('action: ', env.action_space)
command = command_generator(10000, 0.01, 2, vx_range=(-0.2, 0.2), vy_range = (0,0), wyaw_range = (0,0))
reward_fun =1
obs = env.reset( command, reward_fun)
max_step = 1000
v_e = []
c_command = []
xyz = []
rewards = []
action = np.ones(39)* ( 1)
for i in range(max_step):
# env.render()
#action = env.action_space.sample()
next_obs, reward, done, infos = env.step(action)
obs = next_obs
v_e.append(infos['velocity_base'])
c_command.append(infos['commands'])
xyz.append(infos['obs'][:3])
rewards.append(infos['rewards'])
#env.render(mode='rgb_array')#mode='rgb_array'
env.close()
dt =0.01
v_e = np.array(v_e)
c_command = np.array(c_command)
xyz = np.array(xyz)
plt.figure(figsize=(18,6))
rewards_duty = []
for i in range(rewards.shape[1]-1):
rewards_duty.append(np.abs(rewards[:,1+i]/rewards[:,0]) * np.sign(rewards[:,1+i]))
plt.plot(rewards_duty[i], label=str(i))
plt.grid()
plt.legend()
plt.show()
# -
| test/test_fitness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import altair as alt
import altair_latimes as lat
alt.themes.register('latimes', lat.theme)
alt.themes.enable('latimes')
alt.data_transformers.enable('json')
pitches = pd.read_csv("./input/curveballs.csv")
pitches.info()
pitches.release_speed.describe()
pitches.release_spin_rate.describe()
pitcher_stats = pitches.groupby("player_name").agg(dict(
player_name="size",
release_speed="median",
release_spin_rate="median"
)).rename(columns={"player_name": "pitch_count"})
qualified_pitchers = pitcher_stats[pitcher_stats.pitch_count > 50].reset_index()
qualified_pitchers.sort_values("release_speed", ascending=False).head(20)
qualified_pitchers.sort_values("release_spin_rate", ascending=False).head(20)
pitches['player_name'] = pitches.player_name.apply(
lambda x: x if x == '<NAME>' else 'Everybody else'
)
analysis_set = pitches[(pitches.release_speed > 65) & (pitches.release_spin_rate > 1500)]
# +
dots1 = alt.Chart(analysis_set[analysis_set.player_name != '<NAME>']).mark_circle(opacity=1, clip=True).encode(
x=alt.X(
"release_speed:O",
title="Velocity (MPH)",
axis=alt.Axis(values=[65, 70, 75, 80, 85, 90])
),
y=alt.Y(
"release_spin_rate:Q",
title="Spin rate (RPM)",
scale=alt.Scale(zero=False, domain=[1500, 3600])
),
color=alt.Color(
"player_name:N",
title="Player",
scale=alt.Scale(
domain=['<NAME>', 'Everybody else'],
range=[lat.palette['highlight'], lat.palette['default']]
)
)
)
dots2 = alt.Chart(analysis_set[analysis_set.player_name == '<NAME>']).mark_circle(
opacity=1,
stroke="black",
strokeWidth=0.25,
clip=True
).encode(
x=alt.X(
"release_speed:O",
title="Velocity (MPH)",
axis=alt.Axis(values=[65, 70, 75, 80, 85, 90])
),
y=alt.Y(
"release_spin_rate:Q",
title="Spin rate (RPM)",
scale=alt.Scale(zero=False, domain=[1500, 3600])
),
color=alt.Color(
"player_name:N",
title="Player",
scale=alt.Scale(
domain=['Blake Snell', 'Everybody else'],
range=[lat.palette['highlight'], lat.palette['default']]
)
)
)
(dots1 + dots2).properties(
title="Every curveball in the StatCast era",
width=800,
)
# -
analysis_set[analysis_set.player_name == '<NAME>'].release_speed.describe()
analysis_set[analysis_set.player_name == '<NAME>'].release_spin_rate.describe()
| snell-pitches.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from keras.models import Sequential
from keras.layers import Dense, Activation
model = Sequential()
model.add(Dense(64, activation='relu', input_dim=50)) # input shape of 50
model.add(Dense(28, activation='relu'))
model.add(Dense(10, activation='softmax'))
# -
# ## Keras Fundamentals for Deep Learning
# +
## 1. Sequential Model
from keras.models import Sequential
from keras.layers import Dense, Activation, Conv2D, MaxPooling2D, Flatten, Dropout
model = Sequential()
## 2. Convolutional Layer
input_shape = (320, 320, 3)
model.add(Conv2D(48, (3, 3), activation='relu', input_shape=input_shape))
model.add(Conv2D(48, (3, 3), activation='relu'))
## 3. MaxPooling Layer
model.add(MaxPooling2D(pool_size=(2, 2)))
## 4. Dense Layer : adding a Fully Connected Layer with just specifying output size
model.add(Dense(256, activation='relu'))
## 5. Deopout Layer
model.add(Dropout(0.5))
## Compiling, Training, and Evaluation
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X_train, Y_train, batch_size=32, epoch=10, validation_data=(x_val, y_val))
score = model.evaluate(x_test, y_test, batch_size=32)
# -
# ## Let's try using simple linear regression
# +
import keras
from keras.models import Sequential
from keras.layers import Dense, Activation
import numpy as np
import matplotlib.pyplot as plt
x = data = np.linspace(1, 2, 200)
y = x * 4 + np.random.randn(*x.shape) * 0.3
model = Sequential()
model.add(Dense(1, input_dim=1, activation='linear'))
model.compile(optimizer='sgd', loss='mse', metrics=['mse'])
weights = model.layers[0].get_weights()
w_init = weights[0][0][0]
b_init = weights[1][0]
print('Linear regression model is initialized with weights w: %.2f, b: %.2f' % (w_init, b_init))
model.fit(x, y, batch_size=1, epochs=30, shuffle=False)
weights = model.layers[0].get_weights()
w_final = weights[0][0][0]
b_final = weights[1][0]
print('Linear regression model is trained to have weight w: %.2f, b: %.2f' % (w_final, b_final))
predict = model.predict(data)
plt.figure(figsize=(12, 10))
plt.plot(data, predict, 'b', data, y, 'k.')
plt.show()
# +
import tensorflow as tf
from tensorflow.keras import layers
print(tf.VERSION)
print(tf.keras.__version__)
# -
| keras/deeplearning_example/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Import all the necessary libraries here
import numpy as np
import pandas as pd
import seaborn as sns
# # Footwear
# The given dataset contains the profits generated(in %) by all the suppliers of a footwear company in 4 major cities of India - Delhi, Mumbai, Jaipur and Hyderabad. The company wants to invest more money in the city that is showing the most promise. Analyse the dataset and answer the following questions.
#loading data
df=pd.read_csv("Footwear_v2.csv")
df.head()
# +
#we note that there are no null values, and the values are treated as objects and not floats, we will have to clean the
# '%' sign at the end of all and change it to float
#we will write a function to do this like the last session
def clean(string):
clean="".join(filter(lambda x: x!='%', string))
return float(clean)
# you can also use replace
# def clean(val):
# return float(val.replace("%",""))
#
# -
# we also see the supplier column has few 'S' as upper case and few lowercase
#lets clean that too
def supply_cleaner(string):
return string.lower()
#clean the df
df['Supplier']=df['Supplier'].apply(supply_cleaner)
df['Mumbai']=df['Mumbai'].apply(clean)
df['Delhi']=df['Delhi'].apply(clean)
df['Jaipur']=df['Jaipur'].apply(clean)
df['Hyderabad']=df['Hyderabad'].apply(clean)
# ## 1. Average
# Q1)The absolute difference in the average profit percentages of Delhi and Mumbai comes out to be approximately ____
#
# - a) 1.67
#
# - b) 1.57
#
# - **c) 1.77**
#
# - d) 1.47
#
#Solution Q1
#We simply call desctibe on df
df.describe()
6.324-4.555
# ## 2. Box Plots
#
# Plot a box plot to analyse the spread of profits for each of the cities. Which city has the highest profit value at the upper fence in the box plot?
#
#
#
# - a) Delhi
#
# - b) Mumbai
#
# - **c) Hyderabad**
#
# - d) Jaipur
#
# +
sub_df=df[['Delhi', 'Mumbai', 'Jaipur', 'Hyderabad']]
sub_df.boxplot()
# -
# # Crypto Currencies
# The following datasets contain the prices of some popular cryptocurrencies such as bitcoin, litecoin, ethereum, monero, neo, quantum and ripple.Now, you would like to know how the prices of these currencies vary with each other.
# The cryptocurrencies and the corresponding columns in the dataset are as follows:
#
#
# - bitcoin (Close_btc)
# - litecoin(Close_ltc)
# - ethereum(Close_et)
# - monero(Close_mon)
# - neo(Close_neo)
#loading data
df=pd.read_csv("crypto.csv")
# ## 1. Correct Statements
# Q1)Create a pair plot with all these columns and choose the correct statements from the given ones:
#
# I)There is a good trend between litecoin and monero, one increases as the other
#
# II)There is a weak trend between bitcoin and neo.
#
#
# - **a)I**
#
# - b)II
#
# - c)Both I and II
#
# - d)None of the above.
#
#Your code here
sns.pairplot(df)
# As you can see the corelation between Close_ltc and Close_mon is positive and a good trend
#
# And Close_btc and Close_neo also show a strong trend positive line
# ## 2. Heatmap
# Q2)As mentioned earlier, Heat Maps are predominantly utilised for analysing Correlation Matrix. A high positive correlation (values near 1) means a good positive trend - if one increases, then the other also increases. A negative correlation on the other hand(values near -1) indicate good negative trend - if one increases, then the other decreases. A value near 0 indicates no correlation, as in one variable doesn’t affect the other.
# ##### Correlation Matrix
# Here, you can create a correlation matrix of the closing prices by passing the **df.corr()** function, (where df is the dataframe's name) and storing it in a variable. The code will be as follows
#
# df2 = df.corr()
#
#
# After that use this variable df2 to plot a heatmap and choose the correct option.
#
# Check out this link for creating a correlation matrix:https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.corr.html
# Create a correlation matrix for all the prices and then plot a heatmap to analyse the trends. Which of the following options is/are correct?
#
# - **a)Ethereum and Quantum have high correlation**
#
# - b)Neo and Bitcoin have pretty low correlation
#
# - **c)Ethereum has similar correlation with litecoin and neo**
#
df2=df.corr()
sns.heatmap(df2,cmap="Greens", annot=True)
# Close_et and Close_qt have high corr (0.79)
#
# Close_neo and Close_btc have a high corr (0.73)
#
# Close_et and Close_ltc have corr of (0.49)
#
# Close_et and Close_neo have a corr of (0.48)
| footware_and_crypto_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies
import requests
import json
import requests
import pandas as pd
import numpy as np
from config import api_key
# +
from pprint import pprint
pprint(...)
# -
url = "https://api.sportsdata.io/v3/nba/scores/json/Players?"
query_url = f"{url}key={api_key}"
response = requests.get(query_url)
response_json = response.json()
pprint(json.dumps(response_json,indent=4, sort_keys=True))
response_json[0]
players=pd.DataFrame(response_json)
players.to_pickle('players.pkl')
players.to_csv('players.csv', index=False)
players = pd.read_pickle('players.pkl')
players.head()
players.columns
use_col = ['Team', 'PositionCategory','Position', 'FirstName', 'LastName', 'Height', 'Weight', 'BirthDate', 'BirthCity', 'BirthState',
'BirthCountry', 'College', 'Salary', 'PhotoUrl', 'Experience']
healthinfo_df = players[use_col].copy()
healthinfo_df
healthinfo_df['BMI'] = np.round((healthinfo_df['Weight']/healthinfo_df['Height']**2) * 703, decimals = 2)
healthinfo_df.head()
healthinfo_df['PLAYER_NAME'] = healthinfo_df['FirstName'] + ' ' + healthinfo_df['LastName']
# +
# Percentage of players
with open('./_players_all_data.json') as json_file:
data = json.load(json_file)
stats = pd.read_json(data)
stats.head()
# -
stats.columns
def agg_stats(rows):
num_season = rows['SEASON_ID'].nunique()
num_games = rows['GP'].sum()
min_game = (rows['MIN']/rows['GP']).mean()
age = rows['PLAYER_AGE'].max()
return pd.Series({'age': age, 'num_season': num_season, 'num_games': num_games, 'min_game': min_game})
player_stats = stats.groupby('PLAYER_NAME').apply(agg_stats).reset_index()
player_stats.head()
combined = healthinfo_df.merge(player_stats, on='PLAYER_NAME')
combined.corr()
combined.columns
combined
import seaborn as sns
import matplotlib.pyplot as plt
sns.boxenplot(y='Position', x='BMI', data=combined)
sns.scatterplot(y='min_game', x='Experience', hue = 'BMI', data=combined)
combined.groupby('BirthState')['BMI'].mean().sort_values().plot(kind='bar', figsize=(17,5))
sns.scatterplot(y='Salary', x='BMI', data=combined, alpha=0.4)
plt.yscale('log')
sns.regplot(x='num_games', y='BMI', data=combined)
import plotly.express as px
fig.write_html('salary_bmi_player_name_scatter.html')
fig = px.scatter(combined, y='Salary', x='BMI', hover_data=['PLAYER_NAME'])
fig.show()
fig.write_html('min_game_experience_bmi_scatter.html')
fig = px.scatter(combined, y='min_game', x='Experience',color='BMI', hover_data=['PLAYER_NAME'])
fig.show()
fig.write_html('Position_BMI_box.html')
px.box(combined, y='Position', x='BMI', color='Position')
fig.write_html('PositionCategory_BMI_violinbox.html')
px.violin(combined, y='PositionCategory', x='BMI', color='PositionCategory', box=True)
| group_files/nazia/notebooks/sportsdata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Questionário 61 (Q61)
#
#
# Orientações:
#
# - Registre suas respostas no questionário de mesmo nome no SIGAA.
# - O tempo de registro das respostas no questionário será de 10 minutos. Portanto, resolva primeiro as questões e depois registre-as.
# - Haverá apenas 1 (uma) tentativa de resposta.
# - Submeta seu arquivo-fonte (utilizado para resolver as questões) em formato _.ipynb_ pelo SIGAA anexando-o à Tarefa denominada "Envio de arquivo" correspondente ao questionário.
#
# *Nota:* o arquivo-fonte será utilizado apenas como prova de execução da tarefa. Nenhuma avaliação será feita quanto ao estilo de programação.
#
# <hr>
import sympy as sym
from sympy import Symbol, pprint
import numpy as np
import matplotlib.pyplot as plt
# **Questão 1.** Observe a figura abaixo e julgue os itens a seguir.
#
# ```{figure} ../figs/q/q61.png
# ---
# width: 300px
# name: convex
# ---
# ```
#
# i) existe uma função convexa entre as quatro plotadas.
#
# ii) uma entre as funções plotadas possui convexidade parcial.
#
# iii) duas entre as funções plotadas não são convexas.
#
# Assinale a alternativa correta.
#
# A. São corretos i) e ii), apenas.
#
# B. Apenas i) é correto.
#
# C. São corretos i) e iii), apenas.
#
# <NAME>.
# +
plt.figure(figsize=(14,4))
plt.subplot(141)
x1 = np.linspace(-10, 10, 100)
plt.plot(np.sin(x1),c='r')
plt.xticks([]); plt.yticks([]);
plt.title('(a)')
plt.subplot(142)
x2 = np.linspace(-2, 2, 100)
plt.plot(x2, np.exp(x2)*10*np.sin(6*x2))
plt.xticks([]); plt.yticks([]);
plt.title('(b)')
plt.subplot(143)
x3 = np.arange(-100, 100, 1)
plt.plot(x3, x3**2, c='orange')
plt.xticks([]); plt.yticks([]);
plt.title('(c)')
plt.subplot(144)
x4 = np.arange(-100, 0, 1)
plt.plot(x4, x4**3,c='m')
plt.xticks([]); plt.yticks([]);
plt.title('(d)')
plt.show()
# -
# <hr>
#
# ## Gabarito
# Alternativa **A**
# <hr>
#
# **Questão 2.** A função a seguir simula a curva do _potencial de ação_ de uma membrana:
#
# $$P(x) = \dfrac{1.0}{(x - 0.5)^2 + 0.01} - \dfrac{1.0}{(x - 0.8)^2 + 0.04} - 70.$$
#
# Use computação simbólica para calcular uma aproximação para $P'(x=0)$ e assinale a alternativa correta.
#
# A. -67.62
#
#
# B. 0.25
#
#
# C. 11.33
#
#
# D. 0.00
#
# Nota: Use `sympy.subs(x,x0)`.
x1 = np.random.normal(0,1,10)
#Gráfico da função
plt.plot((1.0/(x1-0.5)**2)-1.0/((x1-0.8)**2 +0.04) - 70 , label='$P(x)$', c='g');
plt.legend()
plt.title('Potencial de ação de uma membrana')
plt.show()
#plt.savefig("../figs/q/q61-2.png'")
# <hr>
#
# ## Gabarito
# Alternativa **C**
x = sym.symbols('x')
p = (1.0/((x-0.5)**2+0.01))-1.0/((x-0.8)**2 +0.04) - 70
p
dp = sym.diff(p,x)
dp.subs(x,0)
# <hr>
# **Questão 3.** Considere a função
#
# $$f(x) = - \dfrac{1}{e^x \text{sen}(6x)},$$
#
# definida no domínio $[-0.5,-0.1]$. Assinale a alternativa correta:
#
# A. $f(x)$ não é convexa e $f'(x) = -\frac{e^{x}}{\text{sen}{\left(6 x \right)}} + \frac{6 e^{- x} \cos{\left(6 x \right)}}{\text{sen}^{2}{\left(6 x \right)}}$
#
# B. $f(x)$ é convexa e $f'(x) = \frac{e^{- x}}{\text{sen}{\left(6 x \right)}} + \frac{6 e^{- x} \cos{\left(6 x \right)}}{\text{sen}^{2}{\left(6 x \right)}}$
#
# C. $f(x)$ não é convexa e $f'(x) = \frac{e^{x}}{\text{sen}{\left(6 x \right)}} + \frac{6 e^{- x} \cos{\left(6 x \right)}}{\text{sen}^{2}{\left(6 x \right)}}$
#
# D. $f(x)$ é convexa e $f'(x) = -\frac{e^{- x}}{\text{sen}{\left(6 x \right)}} + \frac{6 e^{- x} \cos{\left(6 x \right)}}{\text{sen}^{2}{\left(6 x \right)}}$
# <hr>
#
# ## Gabarito
# Alternativa **B**.
# +
# domínio
a,b = -0.5,-0.1
x = sym.symbols('x')
c = 6
# função e valores
f = -1/(sym.exp(x)*sym.sin(c*x))
df = f.diff(x)
dom = np.linspace(a,b)
plt.plot(dom, -1/(np.exp(dom)*np.sin(c*dom)));
# -
| _build/jupyter_execute/todo/Q61-gab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Нарисовать маски изображений
# Этот ноутбук генерирует папку с файлами изображений, на которых изобрежены маски кораблей. Это шаг необходим для последующей передачи изображений для тюнинга масок под прямоугольники.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from PIL import Image
# +
def rle_decode(mask_rle, shape=(768, 768)):
'''
mask_rle: run-length as string formated (start length)
shape: (height,width) of array to return
Returns numpy array, 1 - mask, 0 - background
'''
s = mask_rle.split()
starts, lengths = [np.asarray(x, dtype=int) for x in (s[0:][::2], s[1:][::2])]
starts -= 1
ends = starts + lengths
img = np.zeros(shape[0]*shape[1], dtype=np.uint8)
for lo, hi in zip(starts, ends):
img[lo:hi] = 1
return img.reshape(shape).T # Needed to align to RLE direction
def masks_as_image(in_mask_list):
# Take the individual ship masks and create a single mask array for all ships
all_masks = np.zeros((768, 768), dtype = np.int16)
#if isinstance(in_mask_list, list):
for mask in in_mask_list:
if isinstance(mask, str):
all_masks += rle_decode(mask)
return np.expand_dims(all_masks, -1)
# +
## непустые маски изображений
test_images = pd.read_csv('F:\\Downloads\\all\\Submissions\\ResNet_port_v3(5).csv')
test = pd.DataFrame(columns=['ImageId', 'EncodedPixels'])
for c_img in test_images.ImageId:
if isinstance(test_images[test_images.ImageId == c_img].values[0][1], str):
test = pd.concat([test, test_images[test_images.ImageId == c_img]])
# -
test.shape
print(test.ImageId.unique().shape)
for c_img in test.ImageId.unique():
img = test[test.ImageId == c_img].EncodedPixels
img_data = masks_as_image(img)
img_data = img_data[:,:,0].astype('uint8')*255
#img_data = img_data.astype('uint8')
img = Image.fromarray(img_data)
img.save('F:\\Downloads\\all\\final_sub_masks\\simple_list_resnet_e100\\{}'.format(c_img))
test.head()
# +
def multi_rle_encode(img):
labels = label(img[:, :, 0])
return [rle_encode(labels==k) for k in np.unique(labels[labels>0])]
def rle_encode(img):
'''
img: numpy array, 1 - mask, 0 - background
Returns run length as string formated
'''
pixels = img.T.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
# +
from skimage.morphology import label
sub_final = pd.read_csv('F:\\Downloads\\all\\Submissions\\final\\sub_test_02.csv')
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (10, 5))
rle_0 = sub_final.query('ImageId=="54d955de6.jpg"')['EncodedPixels']
img_0 = masks_as_image(rle_0)
ax1.imshow(img_0[:, :, 0])
ax1.set_title('Image$_0$')
rle_1 = multi_rle_encode(img_0)
img_1 = masks_as_image(rle_1)
ax2.imshow(img_1[:, :, 0])
ax2.set_title('Image$_1$')
# -
np.sum(img_0[0:,:1,:0])
911/1600
| src/Make masks to tune.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multigrid examples
# %matplotlib inline
import matplotlib.pyplot as plt
# +
from __future__ import print_function
import numpy as np
import mesh.boundary as bnd
import mesh.patch as patch
import multigrid.MG as MG
# -
# ## Constant-coefficent Poisson equation
# We want to solve
#
# $$\phi_{xx} + \phi_{yy} = -2[(1-6x^2)y^2(1-y^2) + (1-6y^2)x^2(1-x^2)]$$
#
# on
#
# $$[0,1]\times [0,1]$$
#
# with homogeneous Dirichlet boundary conditions (this example comes from "A Multigrid Tutorial").
#
# This has the analytic solution
# $$u(x,y) = (x^2 - x^4)(y^4 - y^2)$$
# We start by setting up a multigrid object--this needs to know the number of zones our problem is defined on
nx = ny = 256
mg = MG.CellCenterMG2d(nx, ny,
xl_BC_type="dirichlet", xr_BC_type="dirichlet",
yl_BC_type="dirichlet", yr_BC_type="dirichlet", verbose=1)
# Next, we initialize the RHS. To make life easier, the `CellCenterMG2d` object has the coordinates of the solution grid (including ghost cells) as `mg.x2d` and `mg.y2d` (these are two-dimensional arrays).
# +
def rhs(x, y):
return -2.0*((1.0-6.0*x**2)*y**2*(1.0-y**2) + (1.0-6.0*y**2)*x**2*(1.0-x**2))
mg.init_RHS(rhs(mg.x2d, mg.y2d))
# -
# The last setup step is to initialize the solution--this is the starting point for the solve. Usually we just want to start with all zeros, so we use the `init_zeros()` method
mg.init_zeros()
# we can now solve -- there are actually two different techniques we can do here. We can just do pure smoothing on the solution grid using `mg.smooth(mg.nlevels-1, N)`, where `N` is the number of smoothing iterations. To get the solution `N` will need to be large and this will take a long time.
#
# Multigrid accelerates the smoothing. We can do a V-cycle multigrid solution using `mg.solve()`
mg.solve()
# We can access the solution on the finest grid using `get_solution()`
phi = mg.get_solution()
# + tags=["nbval-ignore-output"]
plt.imshow(np.transpose(phi.v()), origin="lower")
# -
# we can also get the gradient of the solution
gx, gy = mg.get_solution_gradient()
# + tags=["nbval-ignore-output"]
plt.subplot(121)
plt.imshow(np.transpose(gx.v()), origin="lower")
plt.subplot(122)
plt.imshow(np.transpose(gy.v()), origin="lower")
# -
# ## General linear elliptic equation
# The `GeneralMG2d` class implements support for a general elliptic equation of the form:
# $$\alpha \phi + \nabla \cdot (\beta \nabla \phi) + \gamma \cdot \nabla \phi = f$$
#
# with inhomogeneous boundary condtions.
#
# It subclasses the `CellCenterMG2d` class, and the basic interface is the same
# We will solve the above with
# \begin{align}
# \alpha &= 10 \\
# \beta &= xy + 1 \\
# \gamma &= \hat{x} + \hat{y}
# \end{align}
# and
# \begin{align}
# f = &-\frac{\pi}{2}(x + 1)\sin\left(\frac{\pi y}{2}\right) \cos\left(\frac{\pi x}{2}\right ) \\
# &-\frac{\pi}{2}(y + 1)\sin\left(\frac{\pi x}{2}\right) \cos\left(\frac{\pi y}{2}\right ) \\
# &+\left(\frac{-\pi^2 (xy+1)}{2} + 10\right) \cos\left(\frac{\pi x}{2}\right) \cos\left(\frac{\pi y}{2}\right)
# \end{align}
# on $[0, 1] \times [0,1]$ with boundary conditions:
# \begin{align}
# \phi(x=0) &= \cos(\pi y/2) \\
# \phi(x=1) &= 0 \\
# \phi(y=0) &= \cos(\pi x/2) \\
# \phi(y=1) &= 0
# \end{align}
#
# This has the exact solution:
# $$\phi = \cos(\pi x/2) \cos(\pi y/2)$$
import multigrid.general_MG as gMG
# For reference, we'll define a function providing the analytic solution
def true(x,y):
return np.cos(np.pi*x/2.0)*np.cos(np.pi*y/2.0)
# Now the coefficents--note that since $\gamma$ is a vector, we have a different function for each component
# +
def alpha(x,y):
return 10.0*np.ones_like(x)
def beta(x,y):
return x*y + 1.0
def gamma_x(x,y):
return np.ones_like(x)
def gamma_y(x,y):
return np.ones_like(x)
# -
# and the righthand side function
def f(x,y):
return -0.5*np.pi*(x + 1.0)*np.sin(np.pi*y/2.0)*np.cos(np.pi*x/2.0) - \
0.5*np.pi*(y + 1.0)*np.sin(np.pi*x/2.0)*np.cos(np.pi*y/2.0) + \
(-np.pi**2*(x*y+1.0)/2.0 + 10.0)*np.cos(np.pi*x/2.0)*np.cos(np.pi*y/2.0)
# Our inhomogeneous boundary conditions require a function that can be evaluated on the boundary to give the value
# +
def xl_func(y):
return np.cos(np.pi*y/2.0)
def yl_func(x):
return np.cos(np.pi*x/2.0)
# -
# Now we can setup our grid object and the coefficients, which are stored as a `CellCenter2d` object. Note, the coefficients do not need to have the same boundary conditions as $\phi$ (and for real problems, they may not). The one that matters the most is $\beta$, since that will need to be averaged to the edges of the domain, so the boundary conditions on the coefficients are important.
#
# Here we use Neumann boundary conditions
# +
import mesh.patch as patch
nx = ny = 128
g = patch.Grid2d(nx, ny, ng=1)
d = patch.CellCenterData2d(g)
bc_c = bnd.BC(xlb="neumann", xrb="neumann",
ylb="neumann", yrb="neumann")
d.register_var("alpha", bc_c)
d.register_var("beta", bc_c)
d.register_var("gamma_x", bc_c)
d.register_var("gamma_y", bc_c)
d.create()
a = d.get_var("alpha")
a[:,:] = alpha(g.x2d, g.y2d)
b = d.get_var("beta")
b[:,:] = beta(g.x2d, g.y2d)
gx = d.get_var("gamma_x")
gx[:,:] = gamma_x(g.x2d, g.y2d)
gy = d.get_var("gamma_y")
gy[:,:] = gamma_y(g.x2d, g.y2d)
# -
# Now we can setup the multigrid object
a = gMG.GeneralMG2d(nx, ny,
xl_BC_type="dirichlet", yl_BC_type="dirichlet",
xr_BC_type="dirichlet", yr_BC_type="dirichlet",
xl_BC=xl_func,
yl_BC=yl_func,
coeffs=d,
verbose=1, vis=0, true_function=true)
# just as before, we specify the righthand side and initialize the solution
a.init_zeros()
a.init_RHS(f(a.x2d, a.y2d))
# and we can solve it
a.solve(rtol=1.e-10)
# We can compare to the true solution
v = a.get_solution()
b = true(a.x2d, a.y2d)
e = v - b
# The norm of the error is
e.norm()
| multigrid/multigrid-examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import yake
from collections import defaultdict
import os
import matplotlib.pyplot as plt
import numpy as np
import yake
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from collections import Counter
from wordcloud import WordCloud
import os
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from IPython.display import Image
from collections import defaultdict
import sys
import spacy
from spacy.lang.fr.examples import sentences
import sys
from textblob import Blobber
from textblob_fr import PatternTagger, PatternAnalyzer
import collections
import os
import string
import sys
import pandas as pd
from nltk import word_tokenize
from nltk.corpus import stopwords
from pprint import pprint
from sklearn.cluster import KMeans
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.decomposition import PCA
import numpy as np
import matplotlib.pyplot as plt
from scipy.spatial.distance import cosine
import sys
from gensim.models.phrases import Phrases, Phraser
from gensim.models import Word2Vec
import nltk
from nltk.tokenize import wordpunct_tokenize
from unidecode import unidecode
# -
nlp = spacy.load('fr_core_news_md')
# + tags=[]
import nltk
nltk.download('punkt')
# -
data_path = "../data/txt/"
from collections import Counter
from wordcloud import WordCloud
import os
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from IPython.display import Image
# ## Choisir une décennie
# + tags=[]
DECADE = '1940'
# -
# ## Charger tous les fichiers de la décennie et en créer une liste de textes
# + tags=[]
#data_path = "../data/txt/"
#files = [f for f in sorted(os.listdir(data_path)) if f"_{DECADE[:-1]}" in f]
#print(files)
# Lister les fichiers de cette année
data_path = '../data'
txt_path = '../data/txt'
txts = [f for f in os.listdir(txt_path) if os.path.isfile(os.path.join(txt_path, f)) and f"_{DECADE[:-1]}" in f]
txts
# + tags=[]
len(txts)
# + tags=[]
# Stocker le contenu de ces fichiers dans une liste
content_list = []
for txt in txts:
with open(os.path.join(txt_path, txt), 'r') as f:
content_list.append(f.read())
# + tags=[]
# Imprimer les 200 premiers caractères du contenu du premier fichier
txts[0][:200]
# -
# Ecrire tout le contenu dans un fichier temporaire
temp_path = '../data/tmp'
if not os.path.exists(temp_path):
os.mkdir(temp_path)
with open(os.path.join(temp_path, f'{DECADE}.txt'), 'w') as f:
f.write(' '.join(content_list))
# + tags=[]
import os
import textract
import openpyxl
# + tags=[]
# Imprimer le contenu du fichier et constater les "déchets"
with open(os.path.join(temp_path, f'{DECADE}.txt'), 'r') as f:
before = f.read()
before[:500]
# -
# ## Stopwords
sw = stopwords.words("french")
sw += ["les", "plus", "cette", "fait", "faire", "être", "deux", "comme", "dont", "tout",
"ils", "bien", "sans", "peut", "tous", "après", "ainsi", "donc", "cet", "sous",
"celle", "entre", "encore", "toutes", "pendant", "moins", "dire", "cela", "non",
"faut", "trois", "aussi", "dit", "avoir", "doit", "contre", "depuis", "autres",
"van", "het", "autre", "jusqu"]
sw = set(sw)
print(f"{len(sw)} stopwords:\n {sorted(sw)}")
# + [markdown] tags=[]
# ## Clean text
# -
def clean_text(DECADE, folder=None):
if folder is None:
input_path = f"{DECADE}.txt"
output_path = f"{DECADE}_clean.txt"
else:
input_path = f"{folder}/{DECADE}.txt"
output_path = f"{folder}/{DECADE}_clean.txt"
output = open(output_path, "w", encoding='utf-8')
with open(input_path, encoding='utf-8') as f:
text = f.read()
words = nltk.wordpunct_tokenize(text)
kept = [w.lower() for w in words if len(w) > 2 and w.isalpha() and w.lower() not in sw]
kept_string = " ".join(kept)
output.write(kept_string)
return f'Output has been written in {output_path}!'
clean_text(DECADE, folder=temp_path)
# + [markdown] tags=[]
# ## Counting
# + tags=[]
## word's total number 1840
# !wc ../data/tmp/1940_clean.txt
# -
# ## Tokeniser
# +
# Récupération du contenu du fichier
txt_path = '../data/tmp/1940_clean.txt'
limit = 10**8
with open(txt_path) as f:
text = f.read()[:limit]
# + tags=[]
# Tokenization
words = nltk.wordpunct_tokenize(text)
print(f"{len(words)} words found")
# -
# Eliminer les stopwords et les termes non alphabétiques
kept = [w.lower() for w in words if len(w) > 2 and w.isalpha() and w.lower() not in sw]
voc = set(kept)
print(f"{len(kept)} words kept ({len(voc)} different word forms)")
# ## Finally couting related words
txt_path = '../data/tmp/1940_clean.txt'
# + tags=[]
word = 'culture'
with open(txt_path) as f:
occurrence_culture = f.read().count(word)
print(f"\n Le mot 'culture' apparaître {occurrence_culture} fois dans le corpus\n")
word = 'théâtre'
with open(txt_path) as f:
occurrence_théâtre = f.read().count(word)
print(f"\n Le mot 'cultures' apparaître {occurrence_théâtre} fois dans le corpus\n")
word = 'theatre'
with open(txt_path) as f:
occurrence_theatre = f.read().count(word)
print(f"\n Le mot 'cultures' apparaître {occurrence_theatre} fois dans le corpus\n")
word = 'théâtres'
with open(txt_path) as f:
occurrence__theatres = f.read().count(word)
print(f"\n Le mot 'théâtres' apparaître {occurrence__theatres} fois dans le corpus\n")
word = 'Théâtre'
with open(txt_path) as f:
occurrence_theatre = f.read().count(word)
print(f"\n Le mot 'Théâtre' apparaître {occurrence_theatre} fois dans le corpus\n")
word = 'Théâtres'
with open(txt_path) as f:
occurrence_theatre = f.read().count(word)
print(f"\n Le mot 'Théâtres' apparaître {occurrence_theatre} fois dans le corpus\n")
word = 'museum'
with open(txt_path) as f:
occurrence_museum = f.read().count(word)
print(f"\n Le mot 'cultures' apparaître {occurrence_museum} fois dans le corpus\n")
word = 'Museum'
with open(txt_path) as f:
occurrence_museum = f.read().count(word)
print(f"\n Le mot 'Museum' apparaître {occurrence_museum} fois dans le corpus\n")
word = 'museums'
with open(txt_path) as f:
occurrence_museums = f.read().count(word)
print(f"\n Le mot 'museums' apparaître {occurrence_museums} fois dans le corpus\n")
word = 'Museums'
with open(txt_path) as f:
occurrence_museums = f.read().count(word)
print(f"\n Le mot 'Museums' apparaître {occurrence_museums} fois dans le corpus\n")
word = 'musée'
with open(txt_path) as f:
occurrence_musée = f.read().count(word)
print(f"\n Le mot 'musée' apparaître {occurrence_musée} fois dans le corpus\n")
word = 'Musée'
with open(txt_path) as f:
occurrence_musée = f.read().count(word)
print(f"\n Le mot 'Musée' apparaître {occurrence_musée} fois dans le corpus\n")
word = 'musées'
with open(txt_path) as f:
occurrence_musées = f.read().count(word)
print(f"\n Le mot 'musées' apparaître {occurrence_musées} fois dans le corpus\n")
word = 'Musées'
with open(txt_path) as f:
occurrence_musées = f.read().count(word)
print(f"\n Le mot 'Musées' apparaître {occurrence_musées} fois dans le corpus\n")
word = 'espace culturel'
with open(txt_path) as f:
occurrence_ec = f.read().count(word)
print(f"\n 'espace culturel' apparaître {occurrence_ec} fois dans le corpus\n")
word = 'Espace culturel'
with open(txt_path) as f:
occurrence_ec = f.read().count(word)
print(f"\n 'Espace culturel' apparaître {occurrence_ec} fois dans le corpus\n")
word = 'espaces culturels'
with open(txt_path) as f:
occurrence_ecs = f.read().count(word)
print(f"\n 'espaces culturels' apparaître {occurrence_ecs} fois dans le corpus\n")
word = 'espace de culture'
with open(txt_path) as f:
occurrence_edc = f.read().count(word)
print(f"\n 'Espace de culture' apparaître {occurrence_edc} fois dans le corpus\n")
word = 'espaces de culture'
with open(txt_path) as f:
occurrence_esdc = f.read().count(word)
print(f"\n 'espacee de culture' apparaître {occurrence_esdc} fois dans le corpus\n")
word = 'lieu culturel'
with open(txt_path) as f:
occurrence_lieu = f.read().count(word)
print(f"\n 'lieu culturel' apparaître {occurrence_lieu} fois dans le corpus\n")
word = 'Lieu culturel'
with open(txt_path) as f:
occurrence_lieu = f.read().count(word)
print(f"\n 'Lieu culturel' apparaître {occurrence_lieu} fois dans le corpus\n")
word = 'lieux culturels'
with open(txt_path) as f:
occurrence_lieux = f.read().count(word)
print(f"\n 'lieux culturels' apparaître {occurrence_lieux} fois dans le corpus\n")
word = 'Lieux culturels'
with open(txt_path) as f:
occurrence_lieux = f.read().count(word)
print(f"\n 'Lieux culturels' apparaître {occurrence_lieux} fois dans le corpus\n")
word = 'endroit culturel'
with open(txt_path) as f:
occurrence_endcult = f.read().count(word)
print(f"\n 'endroit culturel' apparaître {occurrence_endcult} fois dans le corpus\n")
word = 'endroits culturels'
with open(txt_path) as f:
occurrence_endcults = f.read().count(word)
print(f"\n 'endroits culturels' apparaître {occurrence_endcults} fois dans le corpus\n")
word = 'centre culturel'
with open(txt_path) as f:
occurrence_centre_culturel = f.read().count(word)
print(f"\n 'centre culturel' apparaître {occurrence_centre_culturel} fois dans le corpus\n")
word = 'Centre culturel'
with open(txt_path) as f:
occurrence_centre_culturel = f.read().count(word)
print(f"\n 'Centre culturel' apparaître {occurrence_centre_culturel} fois dans le corpus\n")
word = 'centres culturels'
with open(txt_path) as f:
occurrence_centres_culturels = f.read().count(word)
print(f"\n 'centres culturels' apparaître {occurrence_centres_culturels} fois dans le corpus\n")
word = '<NAME>'
with open(txt_path) as f:
occurrence_centres_culturels = f.read().count(word)
print(f"\n 'Centres culturels' apparaître {occurrence_centres_culturels} fois dans le corpus\n")
word = 'cinéma'
with open(txt_path) as f:
occurrence_cinema = f.read().count(word)
print(f"\n Le mot 'cinéma' apparaître {occurrence_cinema} fois dans le corpus\n")
word = '<NAME>'
with open(txt_path) as f:
occurrence_patrimoine = f.read().count(word)
print(f"\n 'patrimoine culturel' apparaître {occurrence_patrimoine} fois dans le corpus\n")
word = '<NAME>'
with open(txt_path) as f:
occurrence_patrimoinec = f.read().count(word)
print(f"\n 'Patrimoine culturel' apparaître {occurrence_patrimoinec} fois dans le corpus\n")
word = 'cinémas'
with open(txt_path) as f:
occurrence_cinema = f.read().count(word)
print(f"\n Le mot 'cinémas' apparaître {occurrence_cinema} fois dans le corpus\n")
word = 'Cinéma'
with open(txt_path) as f:
occurrence_cinemaC = f.read().count(word)
print(f"\n 'Cinéma' apparaître {occurrence_cinemaC} fois dans le corpus\n")
word = 'Cinémas'
with open(txt_path) as f:
occurrence_cinemaCs = f.read().count(word)
print(f"\n 'Cinémas' apparaître {occurrence_cinemaCs} fois dans le corpus\n")
word = 'Cinema'
with open(txt_path) as f:
occurrence_avencine = f.read().count(word)
print(f"\n Le mot 'Cinema' apparaître {occurrence_avencine } fois dans le corpus\n")
word = 'cinema'
with open(txt_path) as f:
occurrence_avencine = f.read().count(word)
print(f"\n Le mot 'cinema' apparaître {occurrence_avencine } fois dans le corpus\n")
# + [markdown] tags=[]
# ## Vérifier les 100 mots les plus fréquents - liée au sujet recherché ?
# + tags=[]
# Création d'une liste de mots à ignorer
ignored = set(["Collège", "collège", "francs", "Bourgmestre","Messieurs","VILLE", "Conseil", "conseil communal",
"conseil général", "conseil", "conseil communal", "général", "d'un","d'une", "c'est", "ordinaires",
"chapitre", "titres", "recette dépense", "services", "dépenses", "dépense", "dépenses dépenses prévues",
"déficit recette dépense", "recette dépense prévue", "boni recette dépense", "dépense recette boni",
"recettes recettes", "dépenses dépenses", "qu'on", "depenses dépenses recettes",
"vases sacrés ordinaires", "sacrés ordinaires", "dépenses ordinaires qu'on",
"depenses dépenses recettes", "recettes imprévues. dépenses", "dit", "vue", "n’est", "avons",
"d’une","rue", "Den Nest", "commune", "qu’il", "question", "ville", "c’est", "mais", "den Nest",
"total", "art", "l'article", "Bourgmestre", "bourgmestre", "Messieurs", "VILLE", "prix", "Bruxelles",
"bruxelles", "messieurs", "é p a r", "é d i t", "p r é", "q u é", "é t é", "q u i", "q u ' i", "ê t r",
"œ u v r e", "D E S BIENS", "Remerciements", "q u e","DÉSIGNATION D E S BIENS", "é t r "])
ignored
# -
# Eliminer les stopwords et les termes non alphabétiques
kept = [w.lower() for w in words if len(w) > 2 and w.isalpha() and w.lower() not in sw and w not in ignored]
voc = set(kept)
print(f"{len(kept)} words kept ({len(voc)} different word forms)")
fdist = nltk.FreqDist(kept)
fdist.most_common(100)
| tp4/15-counting_decade1940.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro to Recommender Systems Lab
#
# Complete the exercises below to solidify your knowledge and understanding of recommender systems.
#
# For this lab, we are going to be putting together a user similarity based recommender system in a step-by-step fashion. Our data set contains customer grocery purchases, and we will use similar purchase behavior to inform our recommender system. Our recommender system will generate 5 recommendations for each customer based on the purchases they have made.
import pandas as pd
from scipy.spatial.distance import pdist, squareform
data = pd.read_csv('./data/customer_product_sales.csv')
data.head()
# ## Step 1: Create a data frame that contains the total quantity of each product purchased by each customer.
#
# You will need to group by CustomerID and ProductName and then sum the Quantity field.
# ## Step 2: Use the `pivot_table` method to create a product by customer matrix.
#
# The rows of the matrix should represent the products, the columns should represent the customers, and the values should be the quantities of each product purchased by each customer. You will also need to replace nulls with zeros, which you can do using the `fillna` method.
# ## Step 3: Create a customer similarity matrix using `squareform` and `pdist`. For the distance metric, choose "euclidean."
# ## Step 4: Check your results by generating a list of the top 5 most similar customers for a specific CustomerID.
# ## Step 5: From the data frame you created in Step 1, select the records for the list of similar CustomerIDs you obtained in Step 4.
# ## Step 6: Aggregate those customer purchase records by ProductName, sum the Quantity field, and then rank them in descending order by quantity.
#
# This will give you the total number of each product purchased by the 5 most similar customers to the customer you selected in order from most purchased to least.
# ## Step 7: Filter the list for products that the chosen customer has not yet purchased and then recommend the top 5 products with the highest quantities that are left.
#
# - Merge the ranked products data frame with the customer product matrix on the ProductName field.
# - Filter for records where the chosen customer has not purchased the product.
# - Show the top 5 results.
# ## Step 8: Now that we have generated product recommendations for a single user, put the pieces together and iterate over a list of all CustomerIDs.
#
# - Create an empty dictionary that will hold the recommendations for all customers.
# - Create a list of unique CustomerIDs to iterate over.
# - Iterate over the customer list performing steps 4 through 7 for each and appending the results of each iteration to the dictionary you created.
# ## Step 9: Store the results in a Pandas data frame. The data frame should a column for Customer ID and then a column for each of the 5 product recommendations for each customer.
# ## Step 10: Change the distance metric used in Step 3 to something other than euclidean (correlation, cityblock, consine, jaccard, etc.). Regenerate the recommendations for all customers and note the differences.
| module-2/lab-intro-recommender-systems/your-code/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS" tags=[]
# # Conditional actions
# + [markdown] kernel="SoS" tags=[]
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
# * **Key points**:
# * Normal `break`, `continue`, `return` structures cannot be used in the implicit loops of substeps
# * Action `warn_if` gives an warning under specified conditions
# * Action `fail_if` raises an exception that terminates the substep and therefore the entire workflow if a condition is met
# * Action `done_if` assumes that the substep is completed and ignores the rest of the statements
# * Action `skip_if` skips the substep and removed `_output` even if the `_output` has been generated
# + [markdown] kernel="SoS" tags=[]
# ## Control structures of substeps
# + kernel="SoS" tags=[]
# create a few input files for examples
!touch a_0.txt a_1.txt a_2.txt a_3.txt
# + [markdown] kernel="SoS" tags=[]
# SoS allows the use of arbitrary Python statements in step processes. For example, suppose you are processing a number of input files and some of them contain errors and have to be ignored, you can write a workflow step as follows:
# + kernel="SoS" tags=[]
infiles = [f'a_{i}.txt' for i in range(4)]
outfiles = []
for idx, infile in enumerate(infiles):
if idx == 2: # problematic step
continue
out = f'a_{idx}.out'
sh(f'echo generating {out}\ntouch {out}')
outfiles.append(out)
# + [markdown] kernel="SoS" tags=[]
# However, as we have discussed in tutorials [How to include scripts in different langauges in SoS workflows](script_format.html) and [How to specify input and output files and process input files in groups](input_substeps.html), steps written with loops and function calls like `sh()` are not very readable because the scripts are not clearly presented and users have to follow the logics of the code. Also, the input files are not processed in parallel so the step is not executed efficiently.
#
# The more SoS way to implement the step is to use input and output statements and script format of function calls as follows:
# + kernel="SoS" tags=[]
input: [f'a_{i}.txt' for i in range(4)], group_by=1
output: _input.with_suffix('.out')
sh: expand=True
echo generating {_output}
touch {_output}
# + [markdown] kernel="SoS" tags=[]
# The problem is that substeps are processed concurrently and we do not yet have a way to treat them differentially and introduce the logic of
#
# ```
# if idx == 2: # problematic step
# continue
# ```
# + [markdown] kernel="SoS" tags=[]
# ## Action `skip_if`
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-primary" role="alert">
# <h4>Action <code>skip_if(expr, msg)</code></h4>
# <p>Action <code>skip_if(expr, msg)</code> skips the execution of the substep if condition <code>expr</code> is met. It also assume that the substep generates no output and set <code>_output</code> to empty. The usage pattern of <code>skip_if</code> is</p>
# <pre>
# output: ...
# skip_if(...)
# statements to produce _output
# </pre>
# </div>
# + [markdown] kernel="SoS" tags=[]
# The `skip_if` action allows you to skip certain substeps with certain condition. The condition can involve a (mostly) hidden variable `_index` which is the index of the substep. For example, the aforementioned step can be written as
# + kernel="SoS" tags=[]
input: [f'a_{i}.txt' for i in range(4)], group_by=1
output: _input.with_suffix('.out')
skip_if(_index == 2, 'input 2 has some problem')
sh: expand=True
echo generating {_output}
touch {_output}
# + [markdown] kernel="SoS" tags=[]
# It is important to remember that `skip_if` assumes that substep output is not generated and adjust `_output` accordingly. For example, if you pass the output of the step to another step, you will notice that the output of step `2` is empty.
# + kernel="SoS" tags=[]
# %run -v0
[10]
input: [f'a_{i}.txt' for i in range(4)], group_by=1
output: _input.with_suffix('.out')
skip_if(_index == 2, 'input 2 has some problem')
sh: expand=True
echo generating {_output}
touch {_output}
[20]
print(f'Input of {_index} is {_input}')
# + [markdown] kernel="SoS" tags=[]
# ## Action `done_if`
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-primary" role="alert">
# <h4>Action <code>done_if(expr, msg)</code></h4>
# <p>Action <code>done_if(expr, msg)</code> ignores the rest of the step process, assuming that the substep has been completed with output generated. The usage pattern of <code>done_if</code> is</p>
# <pre>
# output: ...
# statements to produce _output
# done_if(...)
# additional statements
# </pre>
# </div>
# + [markdown] kernel="SoS" tags=[]
# A similar action is `done_if`, which also ignores the rest of the step process but assumes that the output has already been generated. Consequently, this action does not adjust `_output`. For example, if some more work is only applied to a subset of substeps, you can use `done_if` to execute additional code to only selected substeps.
# + kernel="SoS" tags=[]
# %run -v0
[10]
input: [f'a_{i}.txt' for i in range(4)], group_by=1
output: _input.with_suffix('.out')
sh: expand=True
echo generating {_output}
touch {_output}
done_if(_index != 2, 'input 2 need to be fixed')
sh: expand=True
echo "Fixing {_output}"
[20]
print(f'Input of {_index} is {_input}')
# + [markdown] kernel="SoS" tags=[]
# ## Action `warn_if`
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-primary" role="alert">
# <h4>Action <code>warn_if(expr, msg)</code></h4>
# <p>Action <code>warn_if(expr, msg)</code> gives an warning if a specified condition is met.</p>
# </div>
# + [markdown] kernel="SoS" tags=[]
# Action `warn_if` is very easy to use. It just produces an warning message if something suspicious is detected.
# + kernel="SoS" tags=[]
input: [f'a_{i}.txt' for i in range(4)], group_by=1
output: _input.with_suffix('.out')
sh: expand=True
echo generating {_output}
touch {_output}
warn_if(_index == 2, 'input 2 might be problematic')
# + [markdown] kernel="SoS" tags=[]
# ## Action `fail_if`
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-primary" role="alert">
# <h4>Action <code>fail_if(expr, msg)</code></h4>
# <p>Action <code>fail_if(expr, msg)</code> terminates the execution of workflow if a condition is met.</p>
# </div>
# + [markdown] kernel="SoS" tags=[]
# Action `fail_if` terminates the execution of the workflow under certain conditions. It kills all other processes (e.g. working substeps or nested workflows) and it should be used with caution if is unsafe to terminate the workflow abruptly.
#
# For example, if we decide to terminate the entire workflow if we detect something wrong with an input file, we can do
# + kernel="SoS" tags=[]
# %env --expect-error
input: [f'a_{i}.txt' for i in range(4)], group_by=1
output: _input.with_suffix('.out')
sh: expand=True
echo generating {_output}
touch {_output}
fail_if(_index == 2, 'input 2 might be problematic')
# + [markdown] kernel="SoS" tags=[]
# ## Further reading
#
# * [How to include scripts in different langauges in SoS workflows](script_format.html)
# * [How to specify input and output files and process input files in groups](input_substeps.html)
| src/user_guide/control_actions.ipynb |