code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# make analyzer.py visible
import sys
sys.path.insert(0, './../src/')
import analyzer_binary as ana
import pandas as pd
import numpy as np
# Graphics
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib import rc
import matplotlib.dates as mdates
# rc('text', usetex=True)
# rc('text.latex', preamble=r'\usepackage{cmbright}')
# rc('font', **{'family': 'sans-serif', 'sans-serif': ['Helvetica']})
# Magic function to make matplotlib inline;
# %matplotlib inline
# This enables SVG graphics inline.
# There is a bug, so uncomment if it works.
# # %config InlineBackend.figure_formats = {'png', 'retina'}
# JB's favorite Seaborn settings for notebooks
rc = {'lines.linewidth': 2,
'axes.labelsize': 18,
'axes.titlesize': 18,
'axes.facecolor': 'DFDFE5'}
sns.set_context('notebook', rc=rc)
sns.set_style("dark")
mpl.rcParams['xtick.labelsize'] = 16
mpl.rcParams['ytick.labelsize'] = 16
mpl.rcParams['legend.fontsize'] = 14
# -
df = pd.read_csv('tidy_sleep_df.csv', index_col=0)
df.head()
df['percent'] = df.suppressors/df.total
df_15 = df[df['HS time'] == '15']
df_30 = df[df['HS time'] == '30']
df_15.head()
| analyses/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import seaborn as sns
import random
import time
import collections
from tqdm import tqdm
sns.set()
def build_dataset(words, n_words, atleast=1):
count = [['PAD', 0], ['GO', 1], ['EOS', 2], ['UNK', 3]]
counter = collections.Counter(words).most_common(n_words)
counter = [i for i in counter if i[1] >= atleast]
count.extend(counter)
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
for word in words:
index = dictionary.get(word, 0)
if index == 0:
unk_count += 1
data.append(index)
count[0][1] = unk_count
reversed_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return data, count, dictionary, reversed_dictionary
with open('shakespeare.txt') as fopen:
shakespeare = fopen.read()
vocabulary_size = len(list(set(shakespeare)))
data, count, dictionary, rev_dictionary = build_dataset(shakespeare, vocabulary_size)
GO = dictionary['GO']
PAD = dictionary['PAD']
EOS = dictionary['EOS']
UNK = dictionary['UNK']
class Generator:
def __init__(self, size_layer, num_layers, embedded_size,
from_dict_size, to_dict_size, learning_rate, batch_size):
def cells(reuse=False):
return tf.nn.rnn_cell.GRUCell(size_layer,reuse=reuse)
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.X_seq_len = tf.count_nonzero(self.X, 1, dtype=tf.int32)
self.Y_seq_len = tf.count_nonzero(self.Y, 1, dtype=tf.int32)
batch_size = tf.shape(self.X)[0]
encoder_embedding = tf.Variable(tf.random_uniform([from_dict_size, embedded_size], -1, 1))
decoder_embedding = tf.Variable(tf.random_uniform([to_dict_size, embedded_size], -1, 1))
_, encoder_state = tf.nn.dynamic_rnn(
cell = tf.nn.rnn_cell.MultiRNNCell([cells() for _ in range(num_layers)]),
inputs = tf.nn.embedding_lookup(encoder_embedding, self.X),
sequence_length = self.X_seq_len,
dtype = tf.float32)
main = tf.strided_slice(self.Y, [0, 0], [batch_size, -1], [1, 1])
decoder_input = tf.concat([tf.fill([batch_size, 1], GO), main], 1)
dense = tf.layers.Dense(to_dict_size)
decoder_cells = tf.nn.rnn_cell.MultiRNNCell([cells() for _ in range(num_layers)])
training_helper = tf.contrib.seq2seq.TrainingHelper(
inputs = tf.nn.embedding_lookup(decoder_embedding, decoder_input),
sequence_length = self.Y_seq_len,
time_major = False)
training_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cells,
helper = training_helper,
initial_state = encoder_state,
output_layer = dense)
training_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = training_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.Y_seq_len))
self.training_logits = training_decoder_output.rnn_output
predicting_helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(
embedding = decoder_embedding,
start_tokens = tf.tile(tf.constant([GO], dtype=tf.int32), [batch_size]),
end_token = EOS)
predicting_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cells,
helper = predicting_helper,
initial_state = encoder_state,
output_layer = dense)
predicting_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = predicting_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.X_seq_len))
self.predicting_ids = predicting_decoder_output.sample_id
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
self.cost = tf.contrib.seq2seq.sequence_loss(logits = self.training_logits,
targets = self.Y,
weights = masks)
self.optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate).minimize(self.cost)
y_t = tf.argmax(self.training_logits,axis=2)
y_t = tf.cast(y_t, tf.int32)
self.prediction = tf.boolean_mask(y_t, masks)
mask_label = tf.boolean_mask(self.Y, masks)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
learning_rate = 0.001
batch_size = 32
sequence_length = 64
epoch = 3000
num_layers = 2
size_layer = 256
possible_batch_id = range(len(data) - sequence_length - 1)
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Generator(size_layer, num_layers, size_layer, len(dictionary),
len(dictionary), learning_rate,batch_size)
sess.run(tf.global_variables_initializer())
def train_random_batch():
LOST, ACCURACY = [], []
pbar = tqdm(range(epoch), desc = 'epoch')
for i in pbar:
batch_x = np.zeros((batch_size, sequence_length))
batch_y = np.zeros((batch_size, sequence_length + 1))
for n in range(batch_size):
index = np.random.randint(0, len(data) - sequence_length - 1)
batch_x[n] = data[index:index + sequence_length]
batch_y[n] = data[index + 1:index + sequence_length + 1] + [EOS]
accuracy, _, loss = sess.run([model.accuracy, model.optimizer, model.cost],
feed_dict = {model.X: batch_x,
model.Y: batch_y})
ACCURACY.append(accuracy); LOST.append(loss)
pbar.set_postfix(cost = loss, accuracy = accuracy)
return LOST, ACCURACY
LOST, ACCURACY = train_random_batch()
plt.figure(figsize = (15, 5))
plt.subplot(1, 2, 1)
EPOCH = np.arange(len(LOST))
plt.plot(EPOCH, LOST)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.subplot(1, 2, 2)
plt.plot(EPOCH, ACCURACY)
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.show()
def generate_based_sequence(length_sentence):
index = np.random.randint(0, len(data) - sequence_length - 1)
ids = sess.run(model.predicting_ids, feed_dict = {model.X:[data[index:index + sequence_length]]})[0]
ids = ids.tolist()
while len(ids) < length_sentence:
new_ids = sess.run(model.predicting_ids, feed_dict = {model.X:[ids[-sequence_length:]]})[0]
new_ids = new_ids.tolist()
ids += new_ids
return ''.join([rev_dictionary[i] for i in ids])
print(generate_based_sequence(1000))
| generator/6.gru-seq2seq-greedy-char.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Sunil1821/FastAIPractise/blob/master/lesson3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="rAJ4uf95bUFR" outputId="7328d49f-24ab-4db1-ee7a-f8f2e8a27cc5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !pip3 install fastcore==1.0.9
# !pip3 install fastai==1.0.42
# !pip3 install -Uqq fastbook
from torchvision import transforms
import fastbook
fastbook.setup_book()
#hide
from fastbook import *
from fastai.vision.all import *
# + id="aRe7q-Fg1dv9" outputId="5021c4bf-43d7-4137-880c-ea682d1d2d64" colab={"base_uri": "https://localhost:8080/", "height": 17}
path = untar_data(URLs.MNIST_SAMPLE)
# + id="1UgzFYJF2AZ4"
Path.BASE_PATH = path
# + id="H6RrGPKx2Fbb"
(path/"valid"/"3").ls()
# + id="3j5KGcnz2GlK"
valid_3_tens = torch.stack([tensor(Image.open(o)) for o in (path/"valid"/"3").ls()])
valid_7_tens = torch.stack([tensor(Image.open(o)) for o in (path/"valid"/"7").ls()])
# + id="BjFxa_zs2ZSy"
valid_3_tens = valid_3_tens.float()/255
valid_7_tens = valid_7_tens.float()/255
# + id="FRqIT5Jd2-12" outputId="f84a84bb-8f28-43ec-cb23-e159fdf03d82" colab={"base_uri": "https://localhost:8080/", "height": 34}
valid_3_tens.shape, valid_7_tens.shape
# + id="o7mUUSgu3Kxq"
def mnist_distance(a, b) : return (a-b).abs().mean((-1, -2))
# + id="J_tR_HG8406h"
a_3 = tensor(Image.open((path/"valid"/"3").ls()[1]))
# + id="HKAU1Wp747Sg"
train_3_tens = torch.stack([tensor(Image.open(o)) for o in (path/"train"/"3").ls()])
train_7_tens = torch.stack([tensor(Image.open(o)) for o in (path/"train"/"7").ls()])
# + id="tPau3aat5NW-"
train_3_tens = train_3_tens.float()/255
train_7_tens = train_7_tens.float()/255
# + id="cZ3m6Siv5OYQ"
mean3 = train_3_tens.mean(0)
# + id="LynMHoQR6kma"
mean7 = train_7_tens.mean(0)
# + id="c3sGGtfa6np3" outputId="0e6fd3d9-ecb1-4ee9-eb21-5b86ea6d4f01" colab={"base_uri": "https://localhost:8080/", "height": 34}
mnist_distance(a_3, mean3)
# + id="cEyi8KBP6qF3" outputId="1369eecf-9980-4042-b90e-04759cddfa8b" colab={"base_uri": "https://localhost:8080/", "height": 34}
mean3.shape
# + id="32lROjuv6t8X" outputId="1ce9178e-35b6-478a-a8f7-7c73a444f500" colab={"base_uri": "https://localhost:8080/", "height": 34}
mnist_distance(a_3, mean7)
# + id="JGWS4DC87LMu"
from torchvision import transforms
trans = transforms.ToPILImage()
# + id="mhQ2t8Nt7Nct" outputId="8fea23db-caef-4593-9061-690406a03b7a" colab={"base_uri": "https://localhost:8080/", "height": 45}
trans(a_3)
# + id="o-eO0FnU9hr8" outputId="eeae1247-0270-4478-dd8a-7b3e16c524b1" colab={"base_uri": "https://localhost:8080/", "height": 34}
F.mse_loss(a_3, mean3).sqrt()
# + id="Yqa14XsG9mk0" outputId="d1a48204-c389-410c-9898-bc5db3eae971" colab={"base_uri": "https://localhost:8080/", "height": 34}
F.mse_loss(a_3, mean7).sqrt()
# + id="vRSrrlQz9p1z" outputId="0d124a4a-9deb-4bd8-e62d-cf7859c854f7" colab={"base_uri": "https://localhost:8080/", "height": 45}
trans(mean7)
# + id="fGcQy6TQ-_qz" outputId="2dc8da96-5677-4759-de93-a6f0b10b1d6c" colab={"base_uri": "https://localhost:8080/", "height": 34}
tensor([1, 1, 1]) + tensor([2, 3, 4])
# + id="R9rYNu0D_O6z"
def is_3(x) : return mnist_distance(x, mean3) < mnist_distance(x, mean7)
# + id="ec8UHV6QA43a" outputId="e5ad4337-3960-48b0-e989-4047870c3b1d" colab={"base_uri": "https://localhost:8080/", "height": 34}
is_3(a_3)
# + id="vgt9KP4lA6sS" outputId="bc0ced06-e534-4607-896f-a8f269f96ad1" colab={"base_uri": "https://localhost:8080/", "height": 34}
is_3(mean7)
# + id="afceDzk_A8yg" outputId="a65ea802-8d80-4853-d1da-b8b6747914c5" colab={"base_uri": "https://localhost:8080/", "height": 34}
is_3(a_3).float()
# + id="oV7hcWLTA_ix"
valid_3_tens_is_3 = is_3(valid_3_tens).float().mean()
# + id="ZKVgoSewBCdZ"
valid_7_tens_is_3 = is_3(valid_7_tens).float().mean()
# + id="xDHIBD_wBK7h" outputId="24683a9f-7579-4ab3-e5fe-c09438813781" colab={"base_uri": "https://localhost:8080/", "height": 34}
valid_3_tens_is_3
# + id="EGCftrgUB7NZ" outputId="f03e90ac-2d8c-4a63-a723-62655068903b" colab={"base_uri": "https://localhost:8080/", "height": 34}
valid_7_tens_is_3
# + id="aykO0CcUB_zB"
t = tensor(3.0).requires_grad_()
# + id="edC_DGG7F6aA"
def f(x) : return x**2
# + id="eG1OsbK4F9el"
y = f(t)
# + id="ka6_kN8rF-mO"
y.backward()
# + id="H6PaAXFXGAoB" outputId="466d6f90-381a-405c-93fd-fc0b3415d8b6" colab={"base_uri": "https://localhost:8080/", "height": 34}
t
# + id="sSdF2pjxGB1W"
t.grad
# + id="Q4qc8UCvGDCF" outputId="ee49dcb6-c561-404e-b649-22134cca077e" colab={"base_uri": "https://localhost:8080/", "height": 34}
y
# + id="iQ172P8FGEuN" outputId="bd1f537c-5b20-4e7a-dcda-dade3063e2f1" colab={"base_uri": "https://localhost:8080/", "height": 34}
t.grad
# + id="WqzYoHQhGFnF" outputId="74363244-6ada-4af4-84fd-0f93f3032c76" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(t.grad)
# + id="_Vx0Udg8GG4u"
| lesson3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Add a suffix or a prefix to bands
import ee
ee.Initialize()
from geetools import tools
import ipygee as ui
i = ee.Image('COPERNICUS/S2/20181122T142749_20181122T143353_T18GYT').select('B.')
ui.eprint(i.bandNames())
# ## add a suffix
suffix = '_suffix'
renamed_s = tools.image.addSuffix(i, suffix, ['B2', 'B4', 'B8'])
ui.eprint(renamed_s.bandNames())
# ## add prefix
prefix = 'prefix_'
renamed_p = tools.image.addPrefix(i, prefix, ['B2', 'B4', 'B8'])
ui.eprint(renamed_p.bandNames())
# ## In a collection
col = ee.ImageCollection('COPERNICUS/S2').limit(5)
def print_bands(col):
info = col.getInfo()
images = info['features']
for image in images:
bands = image['bands']
print([band['id'] for band in bands])
print_bands(col)
renamed_col_s = col.map(lambda img: tools.image.addSuffix(img, suffix, ['B2', 'B4', 'B8']))
print_bands(renamed_col_s)
| notebooks/image/addSuffix_addPrefix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import xarray as xr
import pandas as pd
from matplotlib import pyplot as plt
import os
cwd = os.getcwd()
demo_file = cwd + '/' + 'M2OCEAN_S2SV3.aof_tavg_1mo_glo_T1440x1080_slv.19810901_0000z.nc4'
ds = xr.open_dataset(demo_file)
print(ds.lons.attrs)
print(ds.lats.attrs)
ds.AO_SNOW[0].plot()
| demo_irreg_grid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import division
import pickle
import os
import random
from matplotlib import pyplot as plt
import numpy as np
import seaborn as sns
sns.set_style('whitegrid')
from lentil import datatools
from lentil import models
from lentil import est
from lentil import evaluate
# %matplotlib inline
# -
import logging
logging.getLogger().setLevel(logging.DEBUG)
# Load an interaction history
history_path = os.path.join('data', 'assistments_2009_2010.pkl')
with open(history_path, 'rb') as f:
history = pickle.load(f)
df = history.data
# Train an embedding model on the interaction history and visualize the results
# +
embedding_dimension = 2
model = models.EmbeddingModel(
history,
embedding_dimension,
using_prereqs=True,
using_lessons=True,
using_bias=True,
learning_update_variance_constant=0.5)
estimator = est.EmbeddingMAPEstimator(
regularization_constant=1e-3,
using_scipy=True,
verify_gradient=False,
debug_mode_on=True,
ftol=1e-3)
model.fit(estimator)
# -
print "Training AUC = %f" % (evaluate.training_auc(
model, history, plot_roc_curve=True))
split_history = history.split_interactions_by_type()
timestep_of_last_interaction = split_history.timestep_of_last_interaction
NUM_STUDENTS_TO_SAMPLE = 10
for student_id in random.sample(df['student_id'].unique(), NUM_STUDENTS_TO_SAMPLE):
student_idx = history.idx_of_student_id(student_id)
timesteps = range(1, timestep_of_last_interaction[student_id]+1)
for i in xrange(model.embedding_dimension):
plt.plot(timesteps, model.student_embeddings[student_idx, i, timesteps],
label='Skill %d' % (i+1))
norms = np.linalg.norm(model.student_embeddings[student_idx, :, timesteps], axis=1)
plt.plot(timesteps, norms, label='norm')
plt.title('student_id = %s' % student_id)
plt.xlabel('Timestep')
plt.ylabel('Skill')
plt.legend(loc='upper right')
plt.show()
# +
assessment_norms = np.linalg.norm(model.assessment_embeddings, axis=1)
plt.xlabel('Assessment embedding norm')
plt.ylabel('Frequency (number of assessments)')
plt.hist(assessment_norms, bins=20)
plt.show()
# -
def get_pass_rates(grouped):
"""
Get pass rate for each group
:param pd.GroupBy grouped: A grouped dataframe
:rtype: dict[str, float]
:return: A dictionary mapping group name to pass rate
"""
pass_rates = {}
for name, group in grouped:
vc = group['outcome'].value_counts()
if True not in vc:
pass_rates[name] = 0
else:
pass_rates[name] = vc[True] / len(group)
return pass_rates
# +
grouped = df[df['module_type']==datatools.AssessmentInteraction.MODULETYPE].groupby('module_id')
pass_rates = get_pass_rates(grouped)
assessment_norms = [np.linalg.norm(model.assessment_embeddings[history.idx_of_assessment_id(assessment_id), :]) for assessment_id in pass_rates]
plt.xlabel('Assessment pass rate')
plt.ylabel('Assessment embedding norm')
plt.scatter(pass_rates.values(), assessment_norms)
plt.show()
# +
grouped = df[df['module_type']==datatools.AssessmentInteraction.MODULETYPE].groupby('module_id')
pass_rates = get_pass_rates(grouped)
bias_minus_norm = [model.assessment_biases[history.idx_of_assessment_id(
assessment_id)] - np.linalg.norm(
model.assessment_embeddings[history.idx_of_assessment_id(
assessment_id), :]) for assessment_id in pass_rates]
plt.xlabel('Assessment pass rate')
plt.ylabel('Assessment bias - Assessment embedding norm')
plt.scatter(pass_rates.values(), bias_minus_norm)
plt.show()
# +
grouped = df[df['module_type']==datatools.AssessmentInteraction.MODULETYPE].groupby('student_id')
pass_rates = get_pass_rates(grouped)
biases = [model.student_biases[history.idx_of_student_id(
student_id)] for student_id in pass_rates]
plt.xlabel('Student pass rate')
plt.ylabel('Student bias')
plt.scatter(pass_rates.values(), biases)
plt.show()
# +
lesson_norms = np.linalg.norm(model.lesson_embeddings, axis=1)
plt.xlabel('Lesson embedding norm')
plt.ylabel('Frequency (number of lessons)')
plt.hist(lesson_norms, bins=20)
plt.show()
# +
prereq_norms = np.linalg.norm(model.prereq_embeddings, axis=1)
plt.xlabel('Prereq embedding norm')
plt.ylabel('Frequency (number of lessons)')
plt.hist(prereq_norms, bins=20)
plt.show()
# -
plt.xlabel('Lesson embedding norm')
plt.ylabel('Prereq embedding norm')
plt.scatter(prereq_norms, lesson_norms)
plt.show()
# +
timesteps = range(model.student_embeddings.shape[2])
avg_student_norms = np.array(np.linalg.norm(np.mean(model.student_embeddings, axis=0), axis=0))
plt.xlabel('Timestep')
plt.ylabel('Average student embedding norm')
plt.plot(timesteps, avg_student_norms)
plt.show()
# -
| nb/model_explorations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Topic Modeling
# +
import pandas as pd
import nltk
from nltk.corpus import stopwords
from nltk.stem.wordnet import WordNetLemmatizer
import string
import gensim
from gensim import corpora, models
import pandas as pd
from nltk import FreqDist
import re
import spacy
# libraries for visualization
#import pyLDAvis
#import pyLDAvis.gensim
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
nltk.download('stopwords')
# one time run
# #!python -m spacy download en
import en_core_web_sm
nlp = en_core_web_sm.load()
# ## Data Pre-processing
# change this to reviews.csv
reviews_df = pd.read_csv('KindleFamilyData.csv')
#reviews_df= reviews_df[['product_id','star_rating', 'review_body']]
reviews_body = "review_body" # set review text column name here
reviews_df.head()
reviews_df['review_body']
# function to plot most frequent terms
def freq_words(x, terms = 30):
all_words = ' '.join([text for text in x])
all_words = all_words.split()
fdist = FreqDist(all_words)
words_df = pd.DataFrame({'word':list(fdist.keys()), 'count':list(fdist.values())})
# selecting top 20 most frequent words
d = words_df.nlargest(columns="count", n = terms)
plt.figure(figsize=(20,5))
ax = sns.barplot(data=d, x= "word", y = "count")
ax.set(ylabel = 'Count')
plt.show()
# +
# remove unwanted characters, numbers and symbols
reviews_df[reviews_body] = reviews_df[reviews_body].str.replace("[^a-zA-Z#]", " ")
stop_words = stopwords.words('english')
# add useless verbs to stop_words
stop_words += ['get','gets','got','use','would']
# function to remove stopwords
def remove_stopwords(rev):
rev_new = " ".join([i for i in rev if i not in stop_words])
return rev_new
# remove short words (length < 3)
reviews_df[reviews_body] = reviews_df[reviews_body].apply(lambda x: ' '.join([w for w in x.split() if len(w)>2]))
# remove really long words (length > 10)
reviews_df[reviews_body] = reviews_df[reviews_body].apply(lambda x: ' '.join([w for w in x.split() if len(w)<11]))
# remove stopwords from the text
reviews = [remove_stopwords(r.split()) for r in reviews_df[reviews_body]]
# make entire text lowercase
reviews = [r.lower() for r in reviews]
# -
reviews_df['reviews']= reviews
freq_words(reviews, 35)
# +
# lemmatization
import en_core_web_sm
nlp = en_core_web_sm.load(disable=['parser', 'ner'])
def lemmatization(texts, tags=[ 'NOUN','ADJECTIVE','VERB']): # use verbs, adjectives and nouns
output = []
for sent in texts:
doc = nlp(" ".join(sent))
output.append([token.lemma_ for token in doc if token.pos_ in tags])
return output
tokenized_reviews = pd.Series(reviews).apply(lambda x: x.split())
print(tokenized_reviews[1])
# delete most common words, such as relating the to product's name
def delete_product_name(token_list):
return [x for x in token_list if x not in ['kindle','fire','amazon', 'device', 'tablet', 'book', 'thing']]
tokenized_reviews = tokenized_reviews.apply(delete_product_name)
reviews_2 = lemmatization(tokenized_reviews)
print(reviews_2[1]) # print lemmatized review
# -
[x for x in reviews_2[1] if x not in stop_words]
# +
# replace review column with the cleaned version
reviews_3 = []
for i in range(len(reviews_2)):
reviews_3.append(' '.join(reviews_2[i]))
reviews_df['reviews'] = reviews_3
# Visualize
freq_words(reviews_df['reviews'], 35)
# -
# ## Modelling
# ### Latent Dirichlet Allocation (LDA)
# +
from gensim.models.coherencemodel import CoherenceModel
dictionary = corpora.Dictionary(reviews_2)
doc_term_matrix = [dictionary.doc2bow(rev) for rev in reviews_2]
# Creating the object for LDA model using gensim library
LDA = gensim.models.ldamodel.LdaModel
# check coherence score to decide number of topics
c_v_score = []
for i in range(2,11):
lda_model = LDA(corpus = doc_term_matrix, id2word = dictionary, num_topics = i, random_state = 44,
chunksize = 1000, passes = 50)
c_v_score.append(CoherenceModel(model=lda_model, texts=reviews_2, dictionary=dictionary, coherence='c_v').get_coherence())
# plot the score
# plot the coherence score
x = range(2, 11)
plt.plot(x, c_v_score)
plt.xlabel("Number of Topics")
plt.ylabel("Coherence score")
plt.legend(("coherence_values"), loc='best')
plt.show()
# -
# train the model on optimal number of topics
lda_model = LDA(corpus = doc_term_matrix, id2word = dictionary, num_topics = c_v_score.index(max(c_v_score))+1, random_state = 44,
chunksize = 1000, passes = 50)
# get the topics
lda_model.print_topics()
# Visualize the topics
pyLDAvis.enable_notebook()
vis = pyLDAvis.gensim.prepare(lda_model, doc_term_matrix, dictionary)
vis
# ### Latent Semantic Indexing (LSA)
# Importing modules and defining functions
# +
#import modules
import os.path
from gensim import corpora
from gensim.models import LsiModel
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from gensim.models.coherencemodel import CoherenceModel
import matplotlib.pyplot as plt
# functions
def prepare_corpus(doc_clean):
"""
Input : clean document
Purpose: create term dictionary of our courpus and Converting list of documents (corpus) into Document Term Matrix
Output : term dictionary and Document Term Matrix
"""
# Creating the term dictionary of our courpus, where every unique term is assigned an index. dictionary = corpora.Dictionary(doc_clean)
dictionary = corpora.Dictionary(doc_clean)
# Converting list of documents (corpus) into Document Term Matrix using dictionary prepared above.
doc_term_matrix = [dictionary.doc2bow(doc) for doc in doc_clean]
# generate LDA model
return dictionary,doc_term_matrix
# function for creating lsa model
def create_gensim_lsa_model(doc_clean,number_of_topics,words):
"""
Input : clean document, number of topics and number of words associated with each topic
Purpose: create LSA model using gensim
Output : return LSA model
"""
dictionary,doc_term_matrix=prepare_corpus(doc_clean)
# generate LSA model
lsamodel = LsiModel(doc_term_matrix, num_topics=number_of_topics, id2word = dictionary) # train model
print(lsamodel.print_topics(num_topics=number_of_topics, num_words=words))
return lsamodel
# function for computing coherence values
def compute_coherence_values(dictionary, doc_term_matrix, doc_clean, stop, start=2, step=3):
"""
Input : dictionary : Gensim dictionary
corpus : Gensim corpus
texts : List of input texts
stop : Max num of topics
purpose : Compute c_v coherence for various number of topics
Output : model_list : List of LSA topic models
coherence_values : Coherence values corresponding to the LDA model with respective number of topics
"""
coherence_values = []
model_list = []
for num_topics in range(start, stop, step):
# generate LSA model
model = LsiModel(doc_term_matrix, num_topics=num_topics, id2word = dictionary) # train model
model_list.append(model)
coherencemodel = CoherenceModel(model=model, texts=doc_clean, dictionary=dictionary, coherence='c_v')
coherence_values.append(coherencemodel.get_coherence())
return model_list, coherence_values
# function for ploting coherence value
def plot_graph(doc_clean,start, stop, step):
dictionary,doc_term_matrix=prepare_corpus(doc_clean)
model_list, coherence_values = compute_coherence_values(dictionary, doc_term_matrix,doc_clean,
stop, start, step)
# Show graph
x = range(start, stop, step)
plt.plot(x, coherence_values)
plt.xlabel("Number of Topics")
plt.ylabel("Coherence score")
plt.legend(("coherence_values"), loc='best')
plt.show()
# -
# Building model
# plot the coherence plot
start,stop,step=2,12,1
plot_graph(reviews_2,start,stop,step)
# train the model
number_of_topics=3
words=10
clean_text = reviews_2
model=create_gensim_lsa_model(clean_text,number_of_topics,words)
# print topics
model.print_topics()
# ### Probabilistic latent semantic analysis (PLSA)
# +
# import packages
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer;
from sklearn.decomposition import NMF;
from sklearn.preprocessing import normalize;
# vectorize the reviews
vectorizer = CountVectorizer(analyzer='word', max_features=5000)
x_counts = vectorizer.fit_transform(reviews_3)
# transform it to tfidf scores
transformer = TfidfTransformer(smooth_idf=False)
x_tfidf = transformer.fit_transform(x_counts)
# normalize the matrix
xtfidf_norm = normalize(x_tfidf, norm='l1', axis=1)
# calculate the coherence score for each topic
num_topics=[1,2,3,4,5,6,7,8,9,10]
c_v_score = []
for i in num_topics:
#obtain a NMF model.
model = NMF(n_components=i, init='nndsvd')
#fit the model
model.fit(xtfidf_norm)
# find the coherence score
c_v_score.append(CoherenceModel(model=lda_model, texts=reviews_2, dictionary=dictionary, coherence='c_v').get_coherence())
num_topics = max(c_v_score.index(max(c_v_score))+1,4)
plsa_model = NMF(n_components=num_topics, init='nndsvd')
plsa_model.fit(xtfidf_norm)
# get words tables
def get_nmf_topics(model, n_top_words):
#the word ids obtained need to be reverse-mapped to the words so we can print the topic names.
feat_names = vectorizer.get_feature_names()
word_dict = {};
for i in range(num_topics):
#for each topic, obtain the largest values, and add the words they map to into the dictionary.
words_ids = model.components_[i].argsort()[:-n_top_words - 1:-1]
words = [feat_names[key] for key in words_ids]
word_dict['Topic # ' + '{:02d}'.format(i+1)] = words
return pd.DataFrame(word_dict)
# print topic
get_nmf_topics(plsa_model, 5)
# -
# print topic
get_nmf_topics(model, 5)
# print topic
get_nmf_topics(plsa_model, 10)
reviews_df.head()
reviews_df.to_csv('reviews_df.csv')
| .ipynb_checkpoints/Topic Modeling [LDA, LSA, PLSA]-checkpoint.ipynb |
# <!--
# title: Intro to Python Lists
# type: lesson
# duration: "00:30"
# creator: <NAME>
# Private gist location: https://gist.github.com/brandiw/621c35f1987e5ab680e7de7b05dfe039
# Presentation URL: https://presentations.generalassemb.ly/621c35f1987e5ab680e7de7b05dfe039#/
# -->
#
# ##  {.separator}
#
# <h1>Python Programming: Lists</h1>
#
# <!--
#
# ## Overview
# This lesson introduces students to the concept of lists. This begins as basic list operations - accessing elements, `len`, `insert`, `append`, and `pop`. After an exercise to recap that, it segues into operations on numerical lists - `sum`, `min`, and `max`. It ends with a longer exercise recapping the list operations.
#
# ## Learning Objectives
# In this lesson, students will:
# - Create lists in Python.
# - Print out specific elements in a list.
# - Perform common list operations.
#
# ## Duration
# 30 minutes
#
# ### Notes on Timing
#
# A 30 minute interval has been allotted for this lesson. You may finish up early due to the fact that this lesson doesn't get into loops or ranges. If you have extra time, put it on the activities or start the next lesson early so students do have buffer time later, when they need it.
#
# That said, at the point you give this lesson, students are still on day one. They will require more time than you probably expect to poke around the code.
#
# ## Suggested Agenda
#
# | Time | Activity |
# | --- | --- |
# | 0:00 - 0:03 | Welcome |
# | 0:03 - 0:15 | Basic List Operations |
# | 0:15 - 0:25 | Numerical List Operations |
# | 0:25 - 0:30 | Summary |
#
# ## In Class: Materials
# - Projector
# - Internet connection
# - Python3
# -->
#
# ---
#
# ## Lesson Objectives
# *After this lesson, you will be able to...*
#
# - Create lists in Python.
# - Print out specific elements in a list.
# - Perform common list operations.
#
# ---
#
# ## What is a List?
#
# Variables hold one item.
my_color = "red"
my_peer = "Brandi"
# **Lists** hold multiple items - and lists can hold anything.
# +
# Declaring lists
colors = ["red", "yellow", "green"]
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha"]
# Strings
colors = ["red", "yellow", "green"]
# Numbers
my_nums = [4, 7, 9, 1, 4]
# Both!
my_nums = ["red", 7, "yellow", 1, 4]
# -
# <aside class="notes">
#
# **Teaching Tips**:
#
# - After explaining what a list is, walk through the syntax (dashes, commas).
# - Point out anything can be in a list - it's just a variable that holds many things.
#
# **Talking Points**:
#
# - "Until now we've used a few different types of variables such as numbers and strings. However, what if we wanted to keep track of more than one thing? Instead of just my single favorite color, how can I store the names of all the colors I like? How can I store the numbers of everyone on my baseball team?
#
# - "Python has this problem solved with something called a *List*."
#
# - "Because a variable is just a box that can hold information, it can also hold lists. Python knows that your variable will hold a list if it begins and ends with square brackets"
#
# - "A list is a data structure in Python, which is a fancy way of saying we can put data inside of it. In the same way you recognize strings by the quotation marks surround them, you can recognize lists by square brackets that surround them."
#
# - "Notice in the example below, we have a list but there are strings inside of it. A list can store data of other types! In this case, I have a list of strings that stores my classmates names."
#
# </aside>
#
#
# ---
#
#
# ## Accessing Elements
#
#
# **List Index** means the location of something (an *element*) in the list.
#
# List indexes start counting at 0!
#
# | List | "Brandi" | "Zoe" | "Steve" | "Aleksander" | "Dasha" |
# |:-----:|:--------:|:-----:|:-------:|:------:|:------:|
# | Index | 0 | 1 | 2 | 3 | 4 |
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha"]
print(my_class[0]) # Prints "Brandi"
print(my_class[1]) # Prints "Zoe"
print(my_class[4]) # Prints "Dasha"
# <aside class="notes">
#
# **Teaching Tips**:
#
# - Starting at 0 is easy to understand and hard to remember. Remind them throughout the presentation.
# - Point out the print syntax.
#
# **Talking Points**:
#
# - "A very important thing to note about lists is that they start counting at 0. Thus, the first element is considered index 0, the second element is considered index 1, and so on."
#
# - "In our previous example, let's print a few specific items. We can access an item by counting from 0 and using square brackets to tell the list which item we want."
#
#
# </aside>
#
# ---
#
# ## We Do: Lists
#
# 1. Create a **list** with the names `"Holly"`, `"Juan"`, and `"Ming"`.
# 2. Print the third name.
# 3. Create a **list** with the numbers `2`,`4`, `6`, and `8`.
# 4. Print the first number.
#
# <iframe height="400px" width="100%" src="https://repl.it/@GAcoding/blank-repl?lite=true" scrolling="no" frameborder="no" allowtransparency="true" allowfullscreen="true" sandbox="allow-forms allow-pointer-lock allow-popups allow-same-origin allow-scripts allow-modals"></iframe>
#
# <aside class="notes">
#
# 1 MINUTE
#
# **Teaching Tips**:
#
# - Run through this quickly - this is just to show them lists working get them practicing typing lists. This is not mean to be a full fledged exercise.
#
# **Repl.it Note**
# It's blank.
#
#
# </aside>
#
#
# ---
#
# ## List Operations - Length
#
#
# `len()`:
#
# - A built in `list` operation.
# - How long is the list?
# +
# length_variable = len(your_list)
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha"]
num_students = len(my_class)
print("There are", num_students, "students in the class")
# => 5
# -
# <aside class="notes">
#
# **Teaching Tips:**
#
# - Stress that even though all examples are strings, these can be performed on any list, no matter what's in it.
#
# **Talking Points**:
#
# - "How many people are in my list? Just as with strings, we can determine how long a list is (i.e., how many elements it has) using the len() method like so."
#
# - "Note: We'll get more into functions later. For now, just know that they perform some operation for you and that you can recognize them by the parentheses on the end."
#
# </aside>
#
# ---
#
# ## Adding Elements: Append
#
# `.append()`:
#
# - A built in `list` operation.
# - Adds to the end of the list.
# - Takes any element.
# +
# your_list.append(item)
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha"]
my_class.append("Sonyl")
print(my_class)
# => ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha", "Sonyl"]
# -
# <aside class="notes">
#
# **Talking Points**:
#
# - "Forgot to add something to that list? No problem; you can use the .append() method. Suppose a new student joins our class. We can add them to the end of the list with `append`, which is a function built directly into a list. (Notice it is called with a dot after the list, unlike the other function we've used, `len`)"
#
# </aside>
#
# ---
#
# ## Adding Elements: Insert
#
# `.insert()`:
#
# - A built in `list` operation.
# - Adds to any point in the list
# - Takes any element and an index.
# +
# your_list.insert(index, item)
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha", "Sonyl"]
my_class.insert(1, "Sanju")
print(my_class)
# => ["Brandi", "Sanju", "Zoe", "Steve", "Aleksander", "Dasha", "Sonyl"]
# -
# <aside class="notes">
#
# **Talking Points**:
#
# - "However, what happens if we want to add something somewhere else? We can use the .insert() method, which specifies where (i.e., to which index) we want to add the element."
#
# </aside>
#
# ---
#
# ## Removing elements - Pop
#
#
# `.pop()`:
#
# - A built in `list` operation.
# - Removes an item from the end of the list.
# +
# your_list.pop()
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha", "Sonyl"]
student_that_left = my_class.pop()
print("The student", student_that_left, "has left the class.")
# => "Sonyl"
print(my_class)
# => ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha"]
# -
# <aside class="notes">
#
# **Talking Points**:
#
# - "What if someone leaves our class? We need to remove them from the list."
#
# - "We can do this with `pop`. Pop drops the last thing off the list. It gives us back the value that it removed. We can take that value and assign it to a new variable, `student that left`. This is called a `return value`."
#
# </aside>
#
#
# ---
#
# ## Removing elements - Pop(index)
#
# `.pop(index)`:
#
# - A built in `list` operation.
# - Removes an item from the list.
# - Can take an index.
# +
# your_list.pop(index)
my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha", "Sonyl"]
student_that_left = my_class.pop(2) # Remember to count from 0!
print("The student", student_that_left, "has left the class.")
# => "Steve"
print(my_class)
# => ["Brandi", "Zoe", "Aleksander", "Dasha", "Sonyl"]
# -
# <aside class="notes">
#
# **Talking Points**:
#
# - "What if someone specific leaves the class?"
#
# - "We can do this with `pop` again. Here, we can give pop the index we want removed. It gives us back the value that it removed. We can take that value and assign it to a new variable, `student that left`."
#
# </aside>
#
# ---
#
# ## Partner Exercise: Pop, Insert, and Append
#
#
# Partner up! Choose one person to be the driver and one to be the navigator, and see if you can do the prompts:
#
# <iframe height="400px" width="100%" src="https://repl.it/@GAcoding/python-programming-lists-intro?lite=true" scrolling="no" frameborder="no" allowtransparency="true" allowfullscreen="true" sandbox="allow-forms allow-pointer-lock allow-popups allow-same-origin allow-scripts allow-modals"></iframe>
#
# <aside class="notes">
#
# **Teaching Tips**:
#
# 3 MINUTES
#
# - Try to get them in pairs they haven't worked in before.
# - Give them just small bit of time to work through this, then go over the answer.
#
# **Repl.it Note:** This replit has:
# +
# 1. Declare a list with the names of your classmates
# 2. Print out the length of that list
# 3. Print the 3rd name on the list
# 4. Delete the first name on the list
# 5. Re-add the name you deleted to the end of the list
# -
# </aside>
#
#
# ---
#
# ## Pop, Insert, Append Solution
#
# <iframe height="400px" width="100%" src="https://repl.it/@GAcoding/python-programming-lists-intro-solution?lite=true" scrolling="no" frameborder="no" allowtransparency="true" allowfullscreen="true" sandbox="allow-forms allow-pointer-lock allow-popups allow-same-origin allow-scripts allow-modals"></iframe>
#
# <aside class="notes">
#
# **Repl.it Note**: This replit has:
#
# # 1. Declare a list with the names of your classmates
#
# my_class = ["Brandi", "Zoe", "Steve", "Aleksander", "Dasha", "Sonyl"]
#
# # 2. Print out the length of that list
# print(len(my_class))
#
# # 3. Print the 3rd name on the list
# print(my_class[2])
#
# # 4. Delete the first name on the list
# deleted_classmate = my_class.pop(0)
#
# # 5. Re-add the name you deleted to the end of the list
# my_class.append(deleted_classmate)
#
# print(my_class)
#
#
# </aside>
#
# ---
#
# ## !! List Mutation: Warning !!
#
# This won't work as expected - don't do this!
colors = ["red", "yellow", "green"]
print colors.append("blue")
# => None
# This will work - do this!
colors = ["red", "yellow", "green"]
colors.append("blue")
print colors
# => ["red", "yellow", "green", "blue"]
# <aside class="notes">
#
# **Teaching Tips**:
#
# - Talk about why this happens, especially in case a student accidentally does it.
#
# **Talking Points**:
#
# - "All of the methods above mutate, i.e., change the array in place; they don't give you the mutated, or changed, array back."
#
# </aside>
#
# ---
#
# ## Quick Review: Basic List Operations
# +
# List Creation
my_list = ["red", 7, "yellow", 1]
# List Length
list_length = len(my_list) # 4
# List Index
print(my_list[0]) # red
# List Append
my_list.append("Yi") # ["red", 7, "yellow", 1, "Yi"]
# List Insert at Index
my_list.insert(1, "Sanju") # ["red", "Sanju", 7, "yellow", 1, "Yi"]
# List Delete
student_that_left = my_list.pop() # "Yi"; ["red", "Sanju", 7, "yellow", 1]
# List Delete at Index
student_that_left = my_list.pop(2) # 7; ["red", "Sanju", "yellow", 1]
# -
# <aside class="notes">
#
# **Teaching Tips**:
#
# - Quickly review. Check to see if anyone's stuck.
# - Remind them that these operations work on lists with both strings and numbers.
#
# </aside>
#
# ---
#
#
# ## Numerical List Operations - Sum
#
# Some actions can only be performed on lists with numbers.
#
# `sum()`:
#
# - A built in `list` operation.
# - Adds the list together.
# - Only works on lists with numbers!
# +
# sum(your_numeric_list)
team_batting_avgs = [.328, .299, .208, .301, .275, .226, .253, .232, .287]
sum_avgs = sum(team_batting_avgs)
print("The total of all the batting averages is", sum_avgs)
# => 2.409
# -
# <aside class="notes">
#
# **Teaching Tips**:
#
# - If baseball's not your thing, feel free to change this.
# - Stress that these only work on lists with entirely numerical values.
# - You might need to explain batting averages.
# - Consider demoing trying to sum string numbers.
#
# **Talking Points**:
#
# - "There's another built-in function, `sum`, used to add a list of numbers together."
#
# </aside>
#
# ---
#
# ## List Operations - Max/Min
#
#
# `max()` or `min()`:
#
# - Built in `list` operations.
# - Finds highest, or lowest, in the list.
# +
# max(your_numeric_list)
# min(your_numeric_list)
team_batting_avgs = [.328, .299, .208, .301, .275, .226, .253, .232, .287]
print("The highest batting average is", max(team_batting_avgs))
# => 0.328
print("The lowest batting average is", min(team_batting_avgs))
# => 0.208
# -
# <aside class="notes">
#
# **Talking Points**:
#
# - "We might want to simply know what is the largest or smallest item in a list. In this case, we can use the built-in functions `max` and `min`."
#
# </aside>
#
# ---
#
# ## You Do: Lists
#
#
# On your local computer, create a `.py` file named `list_practice.py`. In it:
#
# 1. Save a list with the numbers `2`, `4`, `6`, and `8` into a variable called `numbers`.
# 2. Print the max of `numbers`.
# 3. Pop the last element in `numbers` off; re-insert it at index `2`.
# 4. Pop the second number in `numbers` off.
# 5. Append `3` to `numbers`.
# 6. Print out the average number (divide the sum of `numbers` by the length).
# 7. Print `numbers`.
#
# <aside class="notes">
#
# **Teaching Tips**:
#
# - Have students run through this exercise on their own. Circulate the room to check questions and challenges.
# - Students might pop off index 2 versus the actual 2nd element - remind them to watch out for that.
#
# - Answer that's printed:
# Max: 8
# Average: 4.75
# Final list: [2 8 6 3]
#
# </aside>
#
# ---
#
# ## Summary and Q&A
#
# We accomplished quite a bit!
# List Creation
my_list = ["red", 7, "yellow", 1]
# List Length
list_length = len(my_list) # 4
# List Index
print(my_list[0]) # red
# List Append
my_list.append("Yi") # ["red", 7, "yellow", 1, "Yi"]
# List Insert at Index
my_list.insert(1, "Sanju") # ["red", "Sanju", 7, "yellow", 1, "Yi"]
# List Delete
student_that_left = my_list.pop() # "Yi"; ["red", "Sanju", 7, "yellow", 1]
# List Delete at Index
student_that_left = my_list.pop(2) # 7; ["red", "Sanju", "yellow", 1]
# <aside class="notes">
#
# **Teaching Tips**:
#
# - Quickly sum up and check for understanding.
# - Reassure students that while this is a lot, the more they practice typing it, the more they'll get used to it. If they always copy the code off the slide, they won't be very quick to learn it!
#
# </aside>
#
# ---
#
# ## Summary and Q&A
#
# And for numerical lists only...
# Sum all numbers in list
sum_avgs = sum(team_batting_avgs)
# Find minimum value of list
min(team_batting_avgs)
# Find maximum value of list
max(team_batting_avgs)
# <aside class="notes">
#
# **Teaching Tips**:
#
# - Quickly sum up and check for understanding.
#
# </aside>
#
# ---
#
# ## Additional Resources
#
# - [Python Lists - Khan Academy Video](https://www.youtube.com/watch?v=zEyEC34MY1A)
# - [Google For Education: Python Lists](https://developers.google.com/edu/python/lists)
# - [Python-Lists](https://www.tutorialspoint.com/python/python_lists.htm)
#
# <aside class="notes">
#
# **Teaching Tips:**
#
# - Encourage students to go through these in their spare time, especially if they're a bit lost.
#
# </aside>
| unit-2-control-flow/instructor-resources/05-lists/pyth621-day1-lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # ME 595r - Autonomous Sytems
# # Extended Kalman Filter
# ## Dynamic Model
# This filter will estimate the states of a ground robot with velocity inputs and a sensor that measures range and bearing to landmarks. The state is parameterized as
# $$ x = \begin{bmatrix}x \\ y \\ \theta \end{bmatrix} $$
#
# The commanded input is
#
# $$ \hat{u} = \begin{bmatrix} \hat{v} \\ \hat{\omega} \end{bmatrix} $$
#
# The true input to the system is equal to the commanded input corrupted by noise
#
# $$ u = \hat{u} + \xi_u $$
#
# Where $ \xi_u $ is a zero-mean multivariate random variable with covariance
#
# $$ \Sigma_{\xi_u} = \begin{bmatrix} \alpha_1 v_t^2 + \alpha_2 \omega_t^2 & 0 \\ 0 & \alpha_3 v_t^2 + \alpha_4 \omega_t^2 \end{bmatrix} $$
#
# The state evolves as
#
# $$ \bar{x}_t = f(x, u) = x_{t-1} +
# \begin{bmatrix}
# -\tfrac{v_t}{\omega_t}\sin(\theta_{t-1}) + \tfrac{v_t}{\omega_t}\sin(\theta_{t-1} + \omega_t \Delta t) \\
# \tfrac{v_t}{\omega_t}\cos(\theta_{t-1}) - \tfrac{v_t}{\omega_t}\cos(\theta_{t-1} + \omega_t \Delta t) \\
# \omega_t \Delta t
# \end{bmatrix} $$
#
# For the Extended Kalman filter, we need to linearize the dynamic model about our state and our input
#
# $$ A_d = \frac{\partial f}{\partial x} =
# \begin{bmatrix}
# 1 & 0 & -\tfrac{v_t}{\omega_t}\cos(\theta_{t-1}) + \tfrac{v_t}{\omega_t}\cos(\theta_{t-1} + \omega_t \Delta t) \\
# 0 & 1 & -\tfrac{v_t}{\omega_t}\sin(\theta_{t-1}) + \tfrac{v_t}{\omega_t}\sin(\theta_{t-1} + \omega_t \Delta t) \\
# 0 & 0 & 1
# \end{bmatrix} $$
# ## Measurements and Noise
# We will measure the range and bearing to landmarks
# ## Implementation
from __future__ import division
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
#####
# Enable this to be able to zoom plots, but it kills patches
# # %matplotlib inline
# import mpld3
# mpld3.enable_notebook()
#####
from matplotlib import animation, rc
from IPython.display import HTML
from tqdm import tqdm, tqdm_notebook
import copy
#import plotly.plotly as pl
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
figWidth = 11
figHeight = 8
# +
from scipy.stats import multivariate_normal as mvn
def wrap_each(x):
for i, y in enumerate(x):
x[i] = wrap(y)
return x
def wrap(x):
while x < -np.pi:
x += 2*np.pi
while x > np.pi:
x -= 2*np.pi
return x
class Particle(object):
def __init__(self, x0, num_landmarks, g, del_g_x, R, Ts):
self.g = g
self.del_g_x = del_g_x
self.n = len(x0) # state dimension
self.l = num_landmarks
self.R = 1*R
self.x = x0
self.lx = np.zeros((2, num_landmarks))
# self.P = np.array([1e10*np.eye(2*num_landmarks) for i in xrange(num_landmarks)])
self.P = 1e10*np.eye(2)[:, :, None] + np.zeros((2, 2, num_landmarks))
# self.P = self.R[:, :, None] + np.zeros((2, 2, num_landmarks))
self.Ts = Ts
def update(self, z, landmark_idx):
# landmark_idx is a list of indices of landmarks that correspond to the z measurements
# landmark_idx should be the same length as the second dimension of z
# for any landmarks that haven't been initialized
for i, idx in enumerate(landmark_idx):
if self.lx[0, idx] == 0.:
self.lx[:, idx] = self.x[:2] + np.array([z[0, i]*np.cos(z[1, i] + self.x[2]),
z[0, i]*np.sin(z[1, i] + self.x[2])])
# self.P[:, :, idx] = np.copy(self.R)
C = self.del_g_x(self.x, self.lx[:, landmark_idx])
# use Einstein summation notation to do some crazy linalg
# for example np.einsum('mnr,ndr->mdr', A, B)
# does matrix multiply on first two dimensions, broadcasting the operation along the third
# S = C.dot(self.P.dot(C.T)) + self.R
# C_T = np.einsum('ijk->jik', C)
# print(C.shape)
# print(self.P[:, :, landmark_idx].shape)
# similar to P.dot(C.T)
S1 = np.einsum('mnr,dnr->mdr', self.P[:, :, landmark_idx], C)
# print(S1.shape)
S = np.einsum('mnr,ndr->mdr', C, S1) + self.R[:, :, None]
S_inv = np.zeros_like(S)
for i in xrange(S.shape[-1]):
S_inv[:, :, i] = np.linalg.inv(S[:, :, i])
# now do some Einstein stuff for the rest
# self.K = self.P.dot(C.T).dot(np.linalg.inv(S))
K1 = np.einsum('mnr,dnr->mdr', self.P[:, :, landmark_idx], C)
K = np.einsum('mnr,ndr->mdr', K1, S_inv)
z_hat = self.g(self.x, self.lx[:, landmark_idx])
res = z - z_hat
res[1, :] = wrap_each(res[1, :])
# self.lx[:, landmark_idx] = self.lx[:, landmark_idx] + self.K.dot(res)
# Q1 = np.einsum('nmr,ndr->mdr', C, self.P[:, :, landmark_idx])
# Q = np.einsum('mnr,ndr->mdr', Q1, C) + self.R[:, :, None]
w = 0;
for i in xrange(S.shape[-1]):
w += mvn.logpdf(res[:, i], mean=(0, 0), cov=S[:, :, i])
# print("z: {}".format(z))
# print("zHat: {}".format(z_hat[1]))
# print("x: {}".format(self.x))
# print("res: {}".format(res[1]))
# update the estimates
self.lx[:, landmark_idx] = self.lx[:, landmark_idx] + np.einsum('mnr,nr->mr', K, res)
# self.P = (np.eye(self.n + 2*self.l) - self.K.dot(C_aug)).dot(self.P)
# update the covariances
P1 = np.eye(2)[:, :, None] - np.einsum('mnr,ndr->mdr', K, C)
self.P[:, :, landmark_idx] = np.einsum('mnr,ndr->mdr', P1, self.P[:, :, landmark_idx])
return w
# +
from scipy.stats import multivariate_normal as mvn
import copy
class FastSLAM(object):
def __init__(self, x0, num_particles, state_dim, input_dim, num_landmarks, f, g, del_g_x, R, Ts, Q=None, Qu=None):
self.f = f
self.g = g
self.n = state_dim
self.m = input_dim # input dimension
self.num_particles = num_particles
self.num_landmarks = num_landmarks
self.Qu = Qu
self.Q = Q
self.X = []
P0 = 0.0*np.eye(3)
for i in xrange(self.num_particles):
x0_p = np.random.multivariate_normal(x0, P0)
self.X.append(Particle(x0_p, num_landmarks, g, del_g_x, R, Ts))
self.best = self.X[0]
self.Ts = Ts
def lowVarSample(self, w):
Xbar = []
M = self.num_particles
r = np.random.uniform(0, 1/M)
c = w[0]
i = 0
last_i = i
unique = 1
for m in xrange(M):
u = r + m/M
while u > c:
i += 1
c = c + w[i]
Xbar.append(copy.deepcopy(self.X[i]))
if last_i != i:
unique += 1
last_i = i
self.X = Xbar
return unique
def predict(self, u):
self.u = u
# input noise case
# uHat = u[:, np.newaxis] + np.zeros((self.m, self.num_particles))
uHat = u
# propagate the particles
# pdb.set_trace()
for particle in self.X:
if self.Qu is not None:
uHat += np.random.multivariate_normal(np.zeros(self.m), self.Qu(u))
particle.x = self.f(particle.x, uHat, self.Ts)
if self.Q is not None:
particle.x += np.random.multivariate_normal(np.zeros(self.n), self.Q)
# self.X = self.f(self.X, uHat, dt)
# self.x = np.mean(self.X, axis=1)[:, np.newaxis]
# self.P = np.cov(self.X, rowvar=True)
# print(self.X.shape)
# print(self.P.shape)
# print(self.x)
def update(self, z, landmark_idx):
w = np.zeros(self.num_particles)
for i, x in enumerate(self.X):
# wi = 0.9*mvn.pdf(zHat[:, i, :].T, mean=z[:, i], cov=self.R).T
# # add in a 1% mixture of uniform over range measurements between 1m and 11m
# wi += 0.1*0.1
# w += np.log(wi)
w[i] = x.update(z, landmark_idx)
# print(w)
# logsumexp
# print("log w: {}".format(w))
max_w = np.max(w)
w = np.exp(w-max_w)
# for code simplicity, normalize the weights here
w = w/np.sum(w)
self.best_idx = np.argmax(w)
best = self.X[self.best_idx]
# print("w: {}".format(w))
unique = self.lowVarSample(w)
# print(unique)
# add some noise to account for sparsity in particles
# if unique/self.num_particles < 0.5:
# Q = self.P/((self.num_particles*unique)**(1/self.n))
# self.X += np.random.multivariate_normal(np.zeros(self.n), Q, size=self.num_particles).T
# grab the most likely particle before resampling instead
# self.x = np.mean(self.X, axis=1)[:, np.newaxis]
# self.P = np.cov(self.X, rowvar=True)
self.best = best
# +
# initialize inputs and state truth
Ts = 0.1
Tend = 30
num_particles = 10
num_landmarks = 5
t = np.arange(start=Ts, stop=Tend+Ts, step = Ts)
alpha = np.array([0.1, 0.01, 0.01, 0.1])
v_c = 1 + 0.5*np.cos(2*np.pi*0.2*t)
omega_c = -0.2 + 2*np.cos(2*np.pi*0.6*t)
v = v_c + np.random.normal(0, alpha[0]*np.square(v_c) + alpha[1]*np.square(omega_c))
omega = omega_c + np.random.normal(0, alpha[2]*np.square(v_c) + alpha[3]*np.square(omega_c))
u_c = np.vstack((v_c, omega_c))
u = np.vstack((v, omega))
# print(u.shape)
state_dim = 3
x = np.zeros((state_dim, len(t)))
# x[:, 0] = np.array([-5, -3, np.pi/2])
x[:, 0] = np.array([0, 0, 0])
#landmarks = np.array([[6, -7, 6], [4, 8, -4]])
# num_landmarks = 40
# np.random.seed(4)
np.random.seed(5)
landmarks = np.random.uniform(low=-10., high=10., size=(2, num_landmarks))
# # define the model
# def f(x, u, dt):
# v = u.flatten()[0]
# w = u.flatten()[1]
# theta = x.flatten()[2]
# dx = np.array([-v/w*np.sin(theta) + v/w*np.sin(theta + w*dt),
# v/w*np.cos(theta) - v/w*np.cos(theta + w*dt),
# w*dt])
# x_next = x.flatten() + dx
# #print(x_next)
# return x_next
# define the model
def f(x, u, dt):
v = u[0]
w = u[1]
if np.abs(w) < 10*np.finfo(np.float32).eps:
w = 10*np.finfo(np.float32).eps
theta = x[2]
dx = np.array([-v/w*np.sin(theta) + v/w*np.sin(theta + w*dt),
v/w*np.cos(theta) - v/w*np.cos(theta + w*dt),
w*dt])
x_next = x + dx
#print(x_next)
return x_next
def f_parallel(x, u, dt):
v = u[0, :]
w = u[1, :]
w[np.abs(w) < 10*np.finfo(np.float32).eps] = 10*np.finfo(np.float32).eps
theta = x[2, :]
dx = np.array([-v/w*np.sin(theta) + v/w*np.sin(theta + w*dt),
v/w*np.cos(theta) - v/w*np.cos(theta + w*dt),
w*dt])
x_next = x + dx
#print(x_next)
return x_next
def g(x, landmark):
q = (landmark[0] - x[0])**2 + (landmark[1] - x[1])**2
theta = np.arctan2(landmark[1] - x[1], landmark[0] - x[0]) - x[2]
return np.array([np.sqrt(q),
wrap(theta)])
def g_parallel(x, landmark):
q = (landmark[0, :] - x[0])**2 + (landmark[1, :] - x[1])**2
theta = np.arctan2(landmark[1, :] - x[1], landmark[0, :] - x[0]) - x[2]
# theta = ( theta + np.pi) % (2 * np.pi ) - np.pi
theta = wrap_each(theta)
return np.concatenate((np.sqrt(q)[None, :], theta[None, :]), axis=0)
def del_g_x(x, landmark):
lx = landmark[0, :]
ly = landmark[1, :]
dx = lx - x[0]
dy = ly - x[1]
q = (dx)**2 + (dy)**2
sq = np.sqrt(q)
zero = np.zeros_like(dx)
one = np.ones_like(dx)
# C = np.array([[-dx/sq, -dy/sq, zero, dx/sq, dy/sq],
# [dy/q, -dx/q, -one, -dy/q, dx/q]])
C = np.array([[dx/sq, dy/sq],
[-dy/q, dx/q]])
# Ca = np.copy(C)
# # try numeric differentiation
# delta = 0.0000001
# for i in xrange(len(x)):
# C[:, i] = (g(x + delta*np.eye(1, len(x), i).flatten(), landmark) - g(x, landmark))/delta
# print(C - Ca)
# print(C.shape)
return C
def Qu(u):
v = u[0]
w = u[1]
return np.array([[alpha[0]*v**2 + alpha[1]*w**2, 0],
[0, alpha[2]*v**2 + alpha[3]*w**2]])
sigma_r = 0.1
sigma_phi = 0.05
R = np.array([[sigma_r**2, 0],
[0, sigma_phi**2]])
# P = np.array([[1, 0, 0],
# [0, 1, 0],
# [0, 0, 0.1]])
P = np.array([[0, 0, 0],
[0, 0, 0],
[0, 0, 0]])
# for landmark in landmarks.T:
# print(landmark)
# generate truth data
for i in tqdm(xrange(1, len(t)), desc="Generating Truth", ncols=110):
x[:, i:i+1] = f(x[:, i-1:i], u[:, i:i+1], Ts)
xHat = np.zeros_like(x)
xHat[:, 0] = x[:, 0]
best_idx = np.zeros(len(t), dtype=np.int32)
sig = np.zeros_like(x)
sig[:, 0] = np.sqrt(P.diagonal())
landmark_P = np.zeros((2, 2, num_landmarks, len(t)))
K = np.zeros((3, 2, len(t)-1))
landmarksHat = np.zeros((2, num_landmarks, len(t)))
input_dim = u.shape[0]
X = np.zeros((3, num_particles, len(t)))
pf = FastSLAM(xHat[:, 0], num_particles, state_dim, input_dim, num_landmarks, f, g_parallel, del_g_x, R, Ts, Qu=Qu)
zHat = np.zeros((2, len(t)))
for i in tqdm_notebook(xrange(1, len(t)), desc="Estimating"):
uHat = u[:, i] + np.random.multivariate_normal([0, 0], Qu(u[:, i]))
pf.predict(uHat)
z_all = []
landmark_idx = []
for j, landmark in enumerate(landmarks.T):
z = g(x[:, i], landmark) + np.random.multivariate_normal([0, 0], R)
z[1] = wrap(z[1])
if abs(z[1]) < np.pi/4:
z_all.append(z)
landmark_idx.append(j)
# print("z_all: {}".format(z_all))
# print("x: {}".format(x[:, i]))
pf.update(np.array(z_all).T, landmark_idx)
xHat[:, i] = pf.best.x
best_idx[i] = pf.best_idx
# sig[:, i] = np.sqrt(pf.P.diagonal())
for j in xrange(num_landmarks):
landmarksHat[:, j, i] = pf.best.lx[:, j]
# idx = 3+2*j
landmark_P[:, :, j, i] = pf.best.P[:, :, j]
for j in xrange(num_particles):
X[:, j, i] = pf.X[j].x
# e = np.sqrt(((x[0, :] - xHat[0, :])**2 + (x[1, :] - xHat[1, :])**2))
# print("Error norm = {}".format(np.linalg.norm(e)))
# +
from matplotlib.patches import Ellipse
def plot_ellipse(loc, P):
U, s, _ = np.linalg.svd(P)
s = np.sqrt(5.991)*np.sqrt(s)
alpha = np.arctan2(U[1, 0], U[0, 0])
ellipse = Ellipse(loc, s[0], s[1], alpha*180/np.pi, ec='r', fill=False)
return ellipse
def update_ellipse(ellipse, loc, P):
U, s, _ = np.linalg.svd(P)
s = np.sqrt(5.991)*np.sqrt(s)
alpha = np.arctan2(U[1, 0], U[0, 0])
ellipse.center = loc
ellipse.width = s[0]
ellipse.height = s[1]
ellipse.angle = alpha*180/np.pi
plt.close('all')
env = plt.figure(figsize=(6, 6))
ax = env.add_subplot(1, 1, 1)
ax.set_xlim((-10, 10))
ax.set_ylim((-10, 10))
ax.set_title("Robot Environment",fontsize=20)
ax.set_xlabel("X position (m)", fontsize=16)
ax.set_ylabel("Y position (m)", fontsize=16)
robot = plt.Circle((x[0, -1], x[1, -1]), 0.5, fill=False, linestyle=":")
robotHat = plt.Circle((xHat[0, -1], xHat[1, -1]), 0.5, fill=False)
ax.add_artist(robot)
ax.add_artist(robotHat)
direction = np.array([[0, np.cos(x[2, -1])], [0, np.sin(x[2, -1])]])/2
line, = ax.plot(x[0, -1] + direction[0, :], x[1, -1] + direction[1, :], 'k:')
directionHat = np.array([[0, np.cos(xHat[2, -1])], [0, np.sin(xHat[2, -1])]])/2
lineHat, = ax.plot(xHat[0, -1] + directionHat[0, :], xHat[1, -1] + directionHat[1, :], 'k')
features, = ax.plot(landmarks[0, :], landmarks[1, :], 'b*', markersize=6)
# featuresHat, = ax.plot(landmarksHat[0, :, -1], landmarksHat[1, :, -1], 'r*', markersize=10)
particles, = ax.plot(X[0, :, -1], X[1, :, -1], 'go', markersize=1.5, markeredgewidth=0.0)
ellipses = []
for j in xrange(num_landmarks):
ell = plot_ellipse(landmarksHat[:, j, -1], landmark_P[:, :, j, -1])
ell2 = plot_ellipse(landmarksHat[:, j, -1] - X[:2, best_idx[-1], -1] + x[:2, -1], landmark_P[:, :, j, -1])
ax.add_artist(ell)
ellipses.append(ell)
truth, = ax.plot(x[0, :], x[1, :], 'b:')
# estimate, = ax.plot(xHat[0, :], xHat[1, :], 'r')
estimate, = ax.plot(X[0, best_idx[-1], :], X[1, best_idx[-1], :], 'r')
plt.show()
# +
plt.close('all')
env = plt.figure(figsize=(6, 6))
ax = env.add_subplot(1, 1, 1)
ax.set_xlim((-10, 10))
ax.set_ylim((-10, 10))
ax.set_title("Robot Environment",fontsize=20)
ax.set_xlabel("X position (m)", fontsize=16)
ax.set_ylabel("Y position (m)", fontsize=16)
robot = plt.Circle((x[0, 0], x[1, 0]), 0.5, fill=False, linestyle=":")
robotHat = plt.Circle((xHat[0, 0], xHat[1, 0]), 0.5, fill=False)
ax.add_artist(robot)
ax.add_artist(robotHat)
direction = np.array([[0, np.cos(x[2, 0])], [0, np.sin(x[2, 0])]])/2
line, = ax.plot(x[0, 0] + direction[0, :], x[1, 0] + direction[1, :], 'k:')
directionHat = np.array([[0, np.cos(xHat[2, 0])], [0, np.sin(xHat[2, 0])]])/2
lineHat, = ax.plot(xHat[0, 0] + directionHat[0, :], xHat[1, 0] + directionHat[1, :], 'k')
features, = ax.plot(landmarks[0, :], landmarks[1, :], 'b*', markersize=5)
particles, = ax.plot(X[0, :, -1], X[1, :, -1], 'go', markersize=1.5, markeredgewidth=0.0)
# featuresHat, = ax.plot(landmarksHat[0, :, 0], landmarksHat[1, :, 0], 'r*', markersize=5)
ellipses = []
for j in xrange(num_landmarks):
ell = plot_ellipse(landmarksHat[:, j, 0], landmark_P[:, :, j, 0])
ax.add_artist(ell)
ellipses.append(ell)
truth, = ax.plot(x[0, 0], x[1, 0], 'b:')
# estimate, = ax.plot(xHat[0, 0], xHat[1, 0], 'r')
estimate, = ax.plot(X[0, best_idx[0], :], X[1, best_idx[0], :], 'r')
# cart = np.array([zHat[0, 0]*np.cos(zHat[1, 0]+xHat[2, 0]), zHat[0, 0]*np.sin(zHat[1, 0]+xHat[2, 0])])
# measurement, = ax.plot([xHat[0, 0], xHat[0, 0] + cart[0]], [xHat[1, 0], xHat[1, 0] + cart[1]], 'y--')
# animation function. This is called sequentially
def animate(i):
direction = np.array([[0, np.cos(x[2, i])], [0, np.sin(x[2, i])]])/2
line.set_data(x[0, i] + direction[0, :], x[1, i] + direction[1, :])
robot.center = x[0, i], x[1, i]
directionHat = np.array([[0, np.cos(xHat[2, i])], [0, np.sin(xHat[2, i])]])/2
lineHat.set_data(xHat[0, i] + directionHat[0, :], xHat[1, i] + directionHat[1, :])
robotHat.center = xHat[0, i], xHat[1, i]
truth.set_data(x[0, :i], x[1, :i])
# estimate.set_data(xHat[0, :i], xHat[1, :i])
estimate.set_data(X[0, best_idx[i], :i], X[1, best_idx[i], :i])
particles.set_data(X[0, :, i], X[1, :, i])
# featuresHat.set_data(landmarksHat[0, :, i], landmarksHat[1, :, i])
for j in xrange(num_landmarks):
if landmark_P[0, 0, j, i] != 1e10:
update_ellipse(ellipses[j], landmarksHat[:, j, i], landmark_P[:, :, j, i])
# measurement to first landmark
# cart = np.array([zHat[0, i]*np.cos(zHat[1, i]+xHat[2, i]), zHat[0, i]*np.sin(zHat[1, i]+xHat[2, i])])
# measurement.set_data([xHat[0, i], xHat[0, i] + cart[0]], [xHat[1, i], xHat[1, i] + cart[1]])
return (line,)
# call the animator. blit=True means only re-draw the parts that have changed.
speedup = 1
anim = animation.FuncAnimation(env, animate, frames=len(t), interval=Ts*1000/speedup, blit=True)
# anim = animation.FuncAnimation(env, animate, frames=20, interval=Ts*1000/speedup, blit=True)
#print(animation.writers.list())
HTML(anim.to_html5_video())
# +
fig = plt.figure(figsize=(14,16))
fig.clear()
ax1 = fig.add_subplot(4, 1, 1)
ax1.plot(t, x[0, :] - xHat[0, :])
ax1.plot(t, 2*sig[0, :], 'r:')
ax1.plot(t, -2*sig[0, :], 'r:')
ax1.set_title("Error",fontsize=20)
ax1.legend(["error", "2 sigma bound"])
ax1.set_xlabel("Time (s)", fontsize=16)
ax1.set_ylabel("X Error (m)", fontsize=16)
# ax1.set_ylim([-0.5, 0.5])
ax1 = fig.add_subplot(4, 1, 2)
ax1.plot(t, x[1, :] - xHat[1, :])
ax1.plot(t, 2*sig[1, :], 'r:')
ax1.plot(t, -2*sig[1, :], 'r:')
#ax1.set_title("Error",fontsize=20)
ax1.legend(["error", "2 sigma bound"])
ax1.set_xlabel("Time (s)", fontsize=16)
ax1.set_ylabel("Y Error (m)", fontsize=16)
# ax1.set_ylim([-0.5, 0.5])
ax1 = fig.add_subplot(4, 1, 3)
ax1.plot(t, x[2, :] - xHat[2, :])
ax1.plot(t, 2*sig[2, :], 'r:')
ax1.plot(t, -2*sig[2, :], 'r:')
#ax1.set_title("Error",fontsize=20)
ax1.legend(["error", "2 sigma bound"])
ax1.set_xlabel("Time (s)", fontsize=16)
ax1.set_ylabel("Heading Error (rad)", fontsize=16)
# ax1.set_ylim([-0.2, 0.2])
ax1 = fig.add_subplot(4, 1, 4)
e = np.sqrt(((x[0, :] - xHat[0, :])**2 + (x[1, :] - xHat[1, :])**2))
ax1.plot(t, e)
ax1.set_title("Total Distance Error",fontsize=20)
ax1.legend(["error"])
ax1.set_xlabel("Time (s)", fontsize=16)
ax1.set_ylabel("Error (m)", fontsize=16)
print("Error norm = {}".format(np.linalg.norm(e)))
plt.tight_layout()
plt.show()
# -
# ## Questions
#
# * Q: How does the the system behave with poor initial conditions?
# * A: The system converges within a few time steps, even with very poor initial conditions.
#
# * Q: How does the system behave with changes in process/noise convariances?
# * A: Increasing measurement noise increases estimation error and decreases the Kalman gains. Increasing process noise increases noise in truth, but marginally decreases estimation error.
#
# * Q: What happens to the quality of your estimates if you reduce the number of landmarks? increase?
# * A: Fewer landmarks degrades the estimate. More landmarks marginally improves the localization unless the robot gets too close to a landmark, then it can cause it to diverge.
# +
from tqdm import trange
Ts = 1
Tend = 20
t = np.arange(start=Ts, stop=Tend+Ts, step = Ts)
alpha = np.array([0.1, 0.01, 0.01, 0.1])
v_c = 1 + 0.5*np.cos(2*np.pi*0.2*t)
omega_c = -0.2 + 2*np.cos(2*np.pi*0.6*t)
v = v_c + np.random.normal(0, alpha[0]*np.square(v_c) + alpha[1]*np.square(omega_c))
omega = omega_c + np.random.normal(0, alpha[2]*np.square(v_c) + alpha[3]*np.square(omega_c))
u_c = np.vstack((v_c, omega_c))
u = np.vstack((v, omega))
# print(u.shape)
x = np.zeros((3, len(t)))
x[:, 0] = np.array([-5, -3, np.pi/2])
N = 100
e = np.zeros(N)
for j in trange(N):
# generate truth data
for i in xrange(1, len(t)):
x[:, i] = f(x[:, i-1], u[:, i], Ts)
xHat = np.zeros_like(x)
xHat[:, 0] = x[:, 0]
sig = np.zeros_like(x)
sig[:, 0] = np.sqrt(P.diagonal())
K = np.zeros((3, 2, len(t)-1))
input_dim = u.shape[0]
ekf = EKF(xHat[:, 0], input_dim, f, g, del_f_x, del_g_x, R, P, Ts, del_f_u=del_f_u, Qu=Qu)
zHat = np.zeros((2, len(t)))
for i in xrange(1, len(t)):
uHat = u[:, i] + np.random.multivariate_normal([0, 0], Qu(u[:, i]))
ekf.predict(uHat)
for landmark in landmarks.T:
z = g(x[:, i], landmark) + np.random.multivariate_normal([0, 0], R)
# zdeg = z - x[2, i]
# zdeg[1] = zdeg[1]*180/np.pi
# print(zdeg)
zHat[:, i] = z
ekf.update(z, landmark)
# landmark = landmarks[:, 0]
# z = g(x[:, i], landmark) + np.random.multivariate_normal([0, 0], R)
# ekf.update(z, landmark)
xHat[:, i] = ekf.x
K[:, :, i-1] = ekf.K
sig[:, i] = np.sqrt(ekf.P.diagonal())
e[j] = np.linalg.norm(np.sqrt(((x[0, :] - xHat[0, :])**2 + (x[1, :] - xHat[1, :])**2)))
print("Over {} runs:".format(N))
print("Mean error norm = {}".format(np.mean(e*Ts)))
print("Standard deviation of error norm = {}".format(np.std(e*Ts)))
# -
1/6.66
| hw7-fastSLAM/.ipynb_checkpoints/fastSLAM ground robot-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# ## Introduction to Python using Gapminder data
#
# See the notebook below for more info. This cell is formatted in [Markdown](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet).
temp = 34
temp_yesterday = 28
name='Peter'
print("Hello World")
print(Hello World)
print(3.14159)
type(3.14159)
type("Hello")
type('Hello')
type(42)
num1 = 32
participants = 42
print(participants)
participants = participants + 1
print(participants)
class_size = 20
num15 = 3.2
15num = 5
NUM1 = 5
num1 = 7
print('NUM1', NUM1, 'num1', num1)
anumber = 10
anumber = anumber + 1
print(anumber)
print(anumber)
radius = 16
PI = 3.1415926535
radius_squared = radius ** 2
print(radius_squared)
area = radius_squared * PI
print("area of circle with radius", radius, "is", area)
number = 5
print(number, type(number))
new_number = 5 / 2
print(new_number, type(new_number))
another = 8
print(another, type(another))
half = another / 2
print(half, type(half))
bigger = another + 5
print(bigger, type(bigger))
number = 5
new_number = number // 2
print(new_number, type(new_number))
number = 5
print(number, type(number))
new_number = 5 / 2
print(new_number, type(new_number))
another = 8
print(another. type(another))
half = another / 2
print(half, type(half))
bigger = another + 5
print(bigger, type(bigger))
number = 5
print(number, type(number))
new_number = 5 / 2
print(new_number, type(new_number)
another = 8
print(another, type(another))
half = another / 2
print(half, type(half))
bigger = another + 5
print(bigger, type(bigger))
name = 'Peter
number = 5
element = 'helium'
print(element)
print(element[0])
print(element[1])
print(element[2])
print(element[3])
print(element[4])
print(element[5])
element = 'lead'
print(element[0])
print(element[1])
print(element[2])
print(element[3])
print(element[4])
element = 'helium'
abbrev = element[0:3]
print(abbrev)
print(element[0:2])
print(element[3:6])
print(element[3:])
print(element[3:7])
print(element[:3])
"marshallow"[0:4]
"Peter's lunch today was carrots"
'Shakespeare wrote "to be or not to be"'
'Peter\'s lunch today was carrots'
element = 'helium'
print("length of the string:", len(element))
name1 = 'Javan'
len1 = len(name1)
name2 = "Moussa"
len2 = len(name2)
name3 = "Wesley"
len3 = len(name3)
sum_of_lengths = len1 + len2 + len3
print(sum_of_lengths)
empty_string = ""
print(len(empty_string))
print(empty_string[0])
num1 = 2
num2 = 3
total = num1 + num2
print(total)
string1 = "Hello"
string2 = " "
string3 = "World"
greeting = string1 + string2 + string3
print(greeting)
string1 = "Hello"
string3 = "World"
greeting = string1 + string3
print(greeting)
string1 = "Hello"
num1 = 7
string1 + num1
string1 = "Hello"
num_string2 = "7"
string1 + num_string2
num_string1 = "5"
num_string2 = "7"
num_string1 + num_string2
string = "Peter"
len(string)
height = 732
len(height)
height = 732
label = "Height: "
height_string = str(height)
print(height_string)
print(label + height_string)
type("Height")
val1 = "56"
num1 = int(val1)
print(num1)
print(val1)
double1 = num1 * 2
print(double1)
double_val1 = val1 * 2
print(double_val1)
print("Hello","World")
print("Hello" + "World")
print("Hello" * 2)
number1 = int("Hello")
number1 = int("5.5")
number1 = float("5.5")
print(number1)
number1 = int("500,000")
number1 = int("-10")
number1 = int("- 10")
number1 = float("1e3")
print(number1)
num1 = 10
num2 = 2
print("num1", num1, "num2", num2)
num3 = num1 * num2
# print("num3", num3)
num1 = 5
# Ctrl-/ toggles comment on a line on and off
print("num1", num1, "num2", num2)
num4 = num1 * num2
print("num4", num4)
who
num1 = 10
num2 = 2
print("num1", num1, "num2", num2)
num3 = num1 * num2
# print("num3", num3)
num1 = 5
# Ctrl-/ toggles comment on a line on and off
print("num1", num1, "num2", num2)
num4 = num1 * num2
print("num4", num4)
print("Hello")
print()
print()
# some useful functions
print(max(1, 5, 3))
num1 = max(1, 5, 3)
print(num1)
# max and min work with anything that can be ordered
print(min('a', 'A', '0'))
print(min('Jamie', 'Eugene'))
# min of a string uses "lexical order"
min(5, "A")
round(3.14159)
num1 = round(3.1415)
print(type(num1))
num1 = round(3.1415, 2)
print(num1)
help(round)
help(round())
help(print)
round()
| Day1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="zwty8Z6mAkdV"
from absl import logging
import tensorflow as tf
import tensorflow_hub as hub
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import time
import re
import seaborn as sns
from tensorflow.keras.layers import LSTM, TimeDistributed, Dense, Bidirectional, Input, Embedding
from tensorflow.keras.layers import Dropout, Conv1D, Flatten
from tensorflow.keras.layers import Concatenate, Dot, Activation
from tensorflow.keras.models import Model
import os
import collections
import pickle
module_url = "https://tfhub.dev/google/universal-sentence-encoder/4"
model = hub.load(module_url)
print ("module %s loaded" % module_url)
def embed(input):
return model(input)
# + colab={} colab_type="code" id="MmpUE_ZxmGGC"
newsdf = pd.read_csv("./cleannewsdata.csv")
newsdf.Summary = newsdf.Summary.apply(lambda s: s[6:])
def cleaner(s):
s = re.sub("[.?%$0-9!&*+-/:;<=\[\]£]"," ", s)
return " "+" ".join(s.split())
newsdf.Summary = newsdf.Summary.apply(cleaner)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="QrJ-iN-5uVG3" outputId="18a448ab-68ab-4f06-eef7-2b5d69f279e3"
np.random.seed(1)
testindices = np.random.choice(newsdf.shape[0], np.int(0.2*newsdf.shape[0]), replace=False)
trainindices = np.sort(list(set(np.arange(newsdf.shape[0]))-set(testindices)))
traindf, testdf = newsdf.iloc[trainindices], newsdf.iloc[testindices]
traindf.shape, testdf.shape
# + colab={} colab_type="code" id="680xgg673zeZ"
vocab = np.unique([word for sent in newsdf.Summary.apply(lambda s: list(s)).values for word in sent])
def windowed_summary(s, WINDOW_LENGTH=100):
summ = np.zeros((len(s)-WINDOW_LENGTH, WINDOW_LENGTH))
nextchar = np.zeros(len(s)-WINDOW_LENGTH, dtype='<U1')
for i in range(WINDOW_LENGTH, len(s)):
summ[i-WINDOW_LENGTH,:] = [np.where(vocab==r)[0][0] for r in list(s[i-WINDOW_LENGTH:i])]
nextchar[i-WINDOW_LENGTH] = s[i]
return summ, nextchar
# + colab={} colab_type="code" id="iWuv0uhroZu5"
def LSTM_data(df, WINDOW_LENGTH=100):
chararray = np.zeros((df.Summary.apply(lambda s: len(s)-WINDOW_LENGTH).sum(), WINDOW_LENGTH))
predarray = np.zeros((df.Summary.apply(lambda s: len(s)-WINDOW_LENGTH).sum(), vocab.shape[0]))
pos = 0
for i in range(df.shape[0]):
chars, nextval = windowed_summary(df.iloc[i]['Summary'])
chararray[pos:pos+chars.shape[0],:] = chars
for j in range(pos, pos+nextval.shape[0]):
predarray[j,np.where(vocab==nextval[j-pos])[0][0]] = 1
pos+=chars.shape[0]
return chararray, predarray
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="sYrJ12S-68fn" outputId="8dc34f4b-9122-4810-c282-995eb47fc352"
start = time.time()
trainX, trainY = LSTM_data(traindf)
testX, testY = LSTM_data(testdf)
print("Data Generation Exited in "+str(time.time()-start))
trainX.shape, trainY.shape, testX.shape, testY.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ymPrw2ml6KQH" outputId="f31d432a-e919-408b-f243-20cdf91d2e24"
#Generate Universal Sentence Encodings
trainstateX = embed(traindf.Text.values).numpy()
trainstateX = np.repeat(trainstateX, traindf.Summary.apply(lambda s: len(s)-100).values, 0)
teststateX = embed(testdf.Text.values).numpy()
teststateX = np.repeat(teststateX, testdf.Summary.apply(lambda s: len(s)-100).values, 0)
trainstateX.shape, teststateX.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 629} colab_type="code" id="rQhds8nhnfW0" outputId="69b346bc-8e8b-47b5-b360-b08a15484e91"
latentdim = 512
tf.keras.backend.clear_session()
state = Input(shape=(latentdim,))
decinput = Input(shape=(100,))
embed_layer = Embedding(vocab.shape[0], vocab.shape[0], weights=[np.eye(vocab.shape[0])],
trainable=False, input_length=100)
embedval = embed_layer(decinput)
lstm_layer1 = LSTM(latentdim, return_sequences=True, return_state=True)
lstm1val, _, _ = lstm_layer1(embedval, initial_state=[state, state])
lstm1val = Dropout(0.2)(lstm1val)
lstm_layer2 = Bidirectional(LSTM(latentdim, return_sequences=True, return_state=True))
lstm2val, _, _, _, _ = lstm_layer2(lstm1val, initial_state=[state, state, state, state])
lstm2val = Dropout(0.2)(lstm2val)
lstm_layer3 = LSTM(latentdim, return_sequences=False, return_state=True)
lstm3val, _, _ = lstm_layer3(lstm2val, initial_state=[state, state])
lstm3val = Dropout(0.2)(lstm3val)
dense_layer = Dense(vocab.shape[0], activation="softmax")
output = dense_layer(lstm3val)
mdl = Model(inputs=[decinput, state], outputs=output)
mdl.compile(optimizer="adam", loss="categorical_crossentropy")
mdl.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="lpficEUUv2LB" outputId="cf55796f-5433-4156-c005-e0c1ee9bf102"
chckpt = tf.keras.callbacks.ModelCheckpoint("./newspred.h5", monitor='val_loss', save_best_only=True,
save_weights_only=True, save_freq='epoch')
hist = mdl.fit([trainX, trainstateX], trainY, callbacks=[chckpt], verbose=True, batch_size=1000, epochs=10,
validation_data=([testX, teststateX], testY))
# + colab={} colab_type="code" id="A82LsnJVqd_e"
mdl.load_weights("./newspred.h5")
# + colab={} colab_type="code" id="KauMhuW0zCfQ"
def beamer(start, state, k, toplimit=10):
returnvals = collections.deque()
pred = mdl.predict([start, state])
if k==1:
returnvals.append(np.argmax(pred[0]))
return np.max(pred[0]), returnvals
else:
maxval, beamseq = None, None
topchoices = np.argsort(pred[0])[-toplimit:]
for j in topchoices:
chars = start.copy()
chars[0,:-1] = chars[0,1:]
chars[0,-1] = j
val, shortseq = beamer(chars, state, k-1)
if (not maxval) or ((val*pred[0,j])>maxval):
maxval = val*pred[0,j]
beamseq = shortseq
beamseq.appendleft(j)
return maxval, beamseq
# + colab={} colab_type="code" id="ls1-4xFfr7uL"
def generate_text(start, state, k):
start = start.copy().reshape(1, start.shape[-1])
state = state.copy().reshape(1, state.shape[-1])
seq = "".join([vocab[np.int(char)] for char in start[0]])+"|"
for _ in range(200):
maxval, beamseq = beamer(start.copy(), state.copy(), k)
seq+="".join([vocab[np.int(i)] for i in beamseq])
start[0,:-k] = start[0,k:]
start[0,-k:] = beamseq
return seq
# + colab={} colab_type="code" id="9mnR_EmT7FgK"
| BBC-news-data/STAGE 2 - Char-Level Prediction + Universal Sentence Encoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#'''
#Lower-level interface demo that illustrates creating and running a chain
#of gadgets - shortest version.
#
#Usage: JUPYTER EDITED NAME
# fully_sampled_recon_single_chain_short.py [--help | options]
#
#Options:
# -f <file>, --file=<file> raw data file
# [default: simulated_MR_2D_cartesian.h5]
# -p <path>, --path=<path> path to data files, defaults to data/examples/MR
# subfolder of SIRF root folder
# -o <file>, --output=<file> images output file
#'''
#
## CCP PETMR Synergistic Image Reconstruction Framework (SIRF)
## Copyright 2015 - 2017 Rutherford Appleton Laboratory STFC
## Copyright 2015 - 2017 University College London.
##
## This is software developed for the Collaborative Computational
## Project in Positron Emission Tomography and Magnetic Resonance imaging
## (http://www.ccppetmr.ac.uk/).
##
## Licensed under the Apache License, Version 2.0 (the "License");
## you may not use this file except in compliance with the License.
## You may obtain a copy of the License at
## http://www.apache.org/licenses/LICENSE-2.0
## Unless required by applicable law or agreed to in writing, software
## distributed under the License is distributed on an "AS IS" BASIS,
## WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
## See the License for the specific language governing permissions and
## limitations under the License.
__version__ = '0.1.0'
from docopt import docopt
# import engine module
from sirf.Gadgetron import *
# process command-line options
data_file = 'simulated_MR_2D_cartesian.h5'
data_path = examples_data_path('MR')
output_file = 'Gfsout.h5'
# locate the input data
input_file = existing_filepath(data_path, data_file)
acq_data = AcquisitionData(input_file)
# create reconstruction object
recon = Reconstructor(['RemoveROOversamplingGadget', \
'SimpleReconGadgetSet'])
# reconstruct images
image_data = recon.reconstruct(acq_data)
# show reconstructed images
image_array = image_data.as_array()
title = 'Reconstructed images (magnitude)'
# %matplotlib inline
show_3D_array(abs(image_array), suptitle = title, \
xlabel = 'samples', ylabel = 'readouts', label = 'slice')
if output_file is not None:
print('writing to %s' % output_file)
image_data.write(output_file)
# -
| notebooks/MR/Old_notebooks/Gadgetron/fully_sampled_recon_single_chain_short.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Kubeflow Pipeline V1 MNIST Example
import kfp
import kfp.dsl as dsl
from kfp import compiler
from kfp.components import create_component_from_func, OutputPath, InputPath
from datetime import datetime
# # Components
def download_datasets(
train_dataset_path: OutputPath('Dataset'),
test_dataset_path: OutputPath('Dataset')
):
import torchvision.datasets as dsets
import os
os.makedirs(train_dataset_path)
dsets.MNIST(root=train_dataset_path, train=True, download=True)
os.makedirs(test_dataset_path)
dsets.MNIST(root=test_dataset_path, train=False, download=True)
def explore_datasets(
train_dataset_path: InputPath('Dataset'),
test_dataset_path: InputPath('Dataset'),
mlpipeline_ui_metadata_path: OutputPath()
):
import torchvision.datasets as dsets
import json
train = dsets.MNIST(root=train_dataset_path, train=True, download=False)
test = dsets.MNIST(root=test_dataset_path, train=False, download=False)
metadata = {
'outputs' : [{
'type': 'table',
'storage': 'inline',
'format': 'csv',
'header': ["Training samples", "Test samples"],
'source': f"{len(train)}, {len(test)}"
}]
}
with open(mlpipeline_ui_metadata_path, 'w') as metadata_file:
json.dump(metadata, metadata_file)
def train_resnet_model(
number_of_epochs: int,
train_batch_size: int,
learning_rate: float,
train_dataset_path: InputPath('Dataset'),
model_path: OutputPath('Model')
):
import torch
import torch.nn as nn
import torchvision.datasets as dsets
from tqdm import tqdm
from torchvision.transforms import Compose
from torchvision.transforms import Normalize
from torchvision.transforms import Resize
from torchvision.transforms import ToTensor
from kubeflow_pipeline_sample.resnet.resnet_50 import ResNet50
from kubeflow_pipeline_sample.training.trainer import train_model
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = ResNet50(in_channels=1, classes=10).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr = learning_rate)
IMAGE_SIZE = 64
preprocessing = Compose([
Resize((IMAGE_SIZE, IMAGE_SIZE)),
ToTensor(),
Normalize(mean=(0.5), std=(0.5))
])
train_dataset_clean = dsets.MNIST(root=train_dataset_path, train=True, download=False, transform=preprocessing)
train_loader = torch.utils.data.DataLoader(dataset=train_dataset_clean, batch_size=train_batch_size)
losses=train_model(
model=model,
train_loader=train_loader,
criterion=criterion,
optimizer=optimizer,
n_epochs=number_of_epochs,
device=device
)
torch.save(model.state_dict(), model_path)
def evaluate_resnet_model(
test_batch_size: int,
test_dataset_path: InputPath('Dataset'),
model_path: InputPath('Model'),
mlpipeline_metrics_path: OutputPath('Metrics')
):
import torch
import torch.nn as nn
import torchvision.datasets as dsets
import json
from kubeflow_pipeline_sample.resnet.resnet_50 import ResNet50
from kubeflow_pipeline_sample.evaluation.evaluate_accuracy import evaluate_accuracy
from tqdm import tqdm
from torchvision.transforms import Compose
from torchvision.transforms import Normalize
from torchvision.transforms import Resize
from torchvision.transforms import ToTensor
IMAGE_SIZE = 64
preprocessing = Compose([
Resize((IMAGE_SIZE, IMAGE_SIZE)),
ToTensor(),
Normalize(mean=(0.5), std=(0.5))
])
test_dataset_clean = dsets.MNIST(root=test_dataset_path, train=False, download=False, transform=preprocessing)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset_clean, batch_size=test_batch_size)
model = ResNet50(in_channels=1, classes=10)
model.load_state_dict(torch.load(model_path))
model.eval()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
accuracy = evaluate_accuracy(model, test_loader, test_dataset_clean, device)
metrics = {
'metrics': [
{
'name': 'accuracy',
'numberValue': accuracy,
'format': "PERCENTAGE",
}
]
}
with open(mlpipeline_metrics_path, 'w') as metrics_file:
json.dump(metrics, metrics_file)
# +
BASE_IMAGE = "public.ecr.aws/h3o0w0k1/kubeflow-pipeline-mnist:v2"
download_datasets_op = create_component_from_func(
download_datasets,
base_image=BASE_IMAGE
)
explore_datasets_op = create_component_from_func(
explore_datasets,
base_image=BASE_IMAGE
)
train_resnet_model_op = create_component_from_func(
train_resnet_model,
base_image=BASE_IMAGE
)
evaluate_resnet_model_op = create_component_from_func(
evaluate_resnet_model,
base_image=BASE_IMAGE
)
# -
# # Pipeline
def pipeline(
number_of_epochs: int = 1,
train_batch_size: int = 120,
test_batch_size: int = 120,
learning_rate: float = 0.1
):
download_datasets_task = download_datasets_op()
explore_datasets_task = explore_datasets_op(
train_dataset=download_datasets_task.outputs["train_dataset"],
test_dataset=download_datasets_task.outputs["test_dataset"]
)
train_resnet_model_task = train_resnet_model_op(
number_of_epochs=number_of_epochs,
train_batch_size=train_batch_size,
learning_rate=learning_rate,
train_dataset=download_datasets_task.outputs["train_dataset"]
)
evaluate_resnet_model_task = evaluate_resnet_model_op(
test_batch_size=test_batch_size,
test_dataset=download_datasets_task.outputs["test_dataset"],
model=train_resnet_model_task.outputs["model"]
)
# # Generate pipeline definition from code
PIPELINE_DEFINITION_FILE_NAME="end_to_end_ml_pipeline.yaml"
compiler.Compiler(mode=kfp.dsl.PipelineExecutionMode.V1_LEGACY).compile(
pipeline_func=pipeline,
package_path=PIPELINE_DEFINITION_FILE_NAME,
type_check=True
)
# # Create Run From Notebook
NAMESPACE="YOUR_NAMESPACE"
INGRESS_GATEWAY="http://istio-ingressgateway.istio-system.svc.cluster.local"
AUTH="YOUR_KUBEFLOW_AUTH_SERVICE_SESSION_BROWSER_COOKIE"
COOKIE="authservice_session="+AUTH
EXPERIMENT_NAME = "End-to-end ML Pipeline experiment"
client = kfp.Client(host=INGRESS_GATEWAY+"/pipeline", cookies=COOKIE)
experiment = client.create_experiment(EXPERIMENT_NAME)
experiment
run_name = f"e2e-pipeline-run-from-notebook-{datetime.now()}"
run = client.run_pipeline(experiment.id, run_name, PIPELINE_DEFINITION_FILE_NAME)
| notebooks/v1/kubeflow-pipeline-v1-mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#วิดีโอสอน pandas Boxplot subplots
from IPython.display import IFrame, YouTubeVideo, SVG, HTML
YouTubeVideo('G4JYU2zG34E', 400,300)
import pandas as pd
import matplotlib.pyplot as plt
import math
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
print(f'pandas version: {pd.__version__}')
df=pd.read_csv('https://github.com/prasertcbs/tutorial/raw/master/mpg.csv')
df.sample(5)
df.boxplot()
df.boxplot(column=['cty', 'hwy'])
df.boxplot(column=['displ', 'cyl'])
df.columns
for column in ['displ', 'cyl', 'cty', 'hwy']:
df.boxplot(column)
plt.show()
fig, ax=plt.subplots(1, 4, figsize=(16, 4))
columns=['displ', 'cyl', 'cty', 'hwy']
for i, column in enumerate(columns):
# print(i, column)
df.boxplot(column, ax=ax[i])
columns=['displ', 'cyl', 'cty', 'hwy']
nrows=1
ncols=math.ceil(len(columns) / nrows)
fig, ax = plt.subplots(nrows, ncols, figsize=(ncols*3,nrows*4))
print(ax.shape)
col_idx=0
if nrows==1 or ncols==1:
for c in range(ax.size):
if col_idx < len(columns):
df.boxplot(columns[col_idx], ax=ax[c])
col_idx+=1
else:
for r in range(nrows):
for c in range(ncols):
if col_idx < len(columns):
df.boxplot(columns[col_idx], ax=ax[r, c])
col_idx+=1
df.head()
df.columns.str.match('year')
df.columns[~df.columns.str.match('year')]
df.select_dtypes('number')
df[df.columns[~df.columns.str.match('year')]].select_dtypes('number')
dg=df[df.columns[~df.columns.str.match('year')]].select_dtypes('number')
dg.columns
# columns=['displ', 'cyl', 'cty', 'hwy']
columns=dg.columns
nrows=1
ncols=math.ceil(len(columns) / nrows)
fig, ax = plt.subplots(nrows, ncols, figsize=(ncols*3,nrows*4))
# print(ax.shape)
col_idx=0
if nrows==1 or ncols==1:
for c in range(ax.size):
if col_idx < len(columns):
df.boxplot(columns[col_idx], ax=ax[c])
col_idx+=1
else:
for r in range(nrows):
for c in range(ncols):
if col_idx < len(columns):
df.boxplot(columns[col_idx], ax=ax[r, c])
col_idx+=1
| learn_jupyter/10_pandas_viz_boxplot_subplots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multi direccione OSRM - JSON
#
# Este archivo tiene por objetivo crear un archivo Json, en el cual se puedan guardar y alamcenar la ruta para ir de i a j
# siguiendo las calles:
#
# El json va a tener la siguiente forma:
# route={ (cliente 1,cliente 2):lista [ ] }
#
# route={ cliente 1:{cliente 2:[distancia, tiempo, [lista de corordenadas ],}, Cliente 2: }
#
# cliente debe ser nombre, ya que los cliente iran cambiando con le tiempo.
#
# - Paso 1: Encontrar las dirreciones gis.
# - paso 2: Cualcular la routa con el OSRM
# - paso 3: Crear el Json.
import requests
import pandas as pd
import googlemaps
import json
from time import sleep
# +
# leyendo las direcciones desde excel.
#ruta: C:\Users\Usuario\Desktop\CVRPTW_MIT\CVRPTW_HF ** importante: hay que cambiar los \ por / para que funcione.
excel=pd.ExcelFile('C:/Users/Usuario/Desktop/CVRPTW_MIT/CVRPTW_HF/Pedidos.xlsx')
df =pd.read_excel(excel,'Datos')
data=[]
data.insert(0,{'Cliente': '<NAME> 120-B, Pudahuel','Demanda': 0,
'Inicio': 0,'Final': 0,'Duracion': 0})
df=pd.concat([pd.DataFrame(data), df], ignore_index=True, sort=False)
# IMPORTANTE: concat cambiará y orderán los Df por defectos para evitar eso agregar sort=False.
df.head()
# -
df['lat']=''
df['lon']=''
# +
# Leyendo las direcciones de google:
gmaps_key=googlemaps.Client(key="<KEY>")
df['lat']=''
df['lon']=''
for n in range(len(df)):
geocode_resultado=gmaps_key.geocode(df.iloc[n][0]+",Santiago"+",Chile")
try:
lat=geocode_resultado[0]["geometry"]["location"]["lat"]
lon=geocode_resultado[0]["geometry"]["location"]["lng"]
df.at[n,'lat']=lat
df.at[n,'lon']=lon
except:
lat=None
lon=None
df.head()
# -
#Obteniendo las rutas.
def OSRM(df,n,m):
from time import sleep
routes={}
for i in range(n,m):
lon1=df.iloc[i][6]
lat1=df.iloc[i][5]
for j in range(len(df)):
if i!=j:
aux=[]
dic1={}
dic2={}
lon2=df.iloc[j][6]
lat2=df.iloc[j][5]
try:
url = 'http://localhost:5000/route/v1/driving/%0.6f,%0.6f;%0.6f,%0.6f?steps=true'%(lon1,lat1,lon2,lat2)
osrm = requests.get(url)
# obteniendo la información del Json:
data=osrm.json()
distance=data['routes'][0]['distance']
duracion=data['routes'][0]['duration']
#obteniendo la ruta:
for x in range(len(data['routes'][0]['legs'][0]['steps'])):
for y in range(len(data['routes'][0]['legs'][0]['steps'][x]['intersections'])):
aux.append(data['routes'][0]['legs'][0]['steps'][x]['intersections'][y]['location'])
#crenado el diccionario:
dic2['distancia']=distance
dic2['duracion']=duracion
dic2['ruta']=aux
except:
dic2={}
dic1[df.iloc[j][0]]=dic2
sleep(0.5)
routes[df.iloc[i][0]]=dic1
sleep(2)
return(routes)
# creando el json:
dic={}
for i in range(len(df)):
routes=OSRM(df,i,i+1)
dic={**dic,**routes}
dic
with open('route_osrm.json','w') as outfile: json.dump(dic,outfile)
| OSRM Python/OSRM - Json - multilple location.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
# +
import cv2
import torch
import numpy as np
import matplotlib.pylab as plt
import torch.optim as optim
import torch.nn.functional as F
from models.VQVAE import VQVAE
from config import setSeed, getConfig
from customLoader import MinecraftData, LatentBlockDataset
from pprint import pprint
from os.path import join
from pathlib import Path
from torchvision.utils import make_grid
from torchvision.transforms import transforms
from torch.utils.data import DataLoader
# -
conf = {
"experiment": "test_2",
"environment": "MineRLNavigate-v0",
"batch_size": 256,
"num_training_updates": 25000,
"learning_rate": 0.001,
"split": 0.9,
"vqvae": {
"num_hiddens": 128,
"num_residual_hiddens": 32,
"num_residual_layers": 2,
"embedding_dim": 64,
"num_embeddings": 512,
"commitment_cost": 0.25,
"decay": 0.99
},
"pixelcnn": {
"epochs": 100,
"save": "no",
"log_interval": 100,
"lr": 0.0003,
"img_dim": 16,
"batch_size": 32,
"num_hiddens": 64,
"num_layers": 15,
"gen_samples": "no"
}
}
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
vqvae = VQVAE(**conf['vqvae']).to(device)
vqvae.eval()
weights_vqvae = torch.load(f"../../weights/{conf['experiment']}/24999.pt")['state_dict']
vqvae.load_state_dict(weights_vqvae)
# +
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5,0.5,0.5), (1.0,1.0,1.0))
])
mrl_val = MinecraftData(conf['environment'], 'val', conf['split'], False, transform=transform, path='../../data')
validation_loader = DataLoader(mrl_val, batch_size=16, shuffle=True)
# -
valid_originals = next(iter(validation_loader))
valid_originals = valid_originals.to(device)
vq_output_eval = vqvae._pre_vq_conv(vqvae._encoder(valid_originals))
_, valid_quantize, _, encoding_indices = vqvae._vq_vae(vq_output_eval)
indices = torch.reshape(encoding_indices, (16,16,16)).unsqueeze(dim=3)
indices = indices.cpu().numpy()
indices.shape
fig, ax = plt.subplots(2,8, figsize=(16,4))
for j,i in enumerate(indices):
ax[int(j/8), int(j%8)].imshow(i)
ax[int(j/8), int(j%8)].axis('off')
ax[int(j/8), int(j%8)].axis("tight")
plt.show()
valid_reconstructions = vqvae._decoder(valid_quantize)
grid = make_grid(valid_reconstructions.cpu().data, normalize=True)
fig, ax = plt.subplots(figsize=(16,4))
plt.imshow(grid.permute(1,2,0).numpy())
plt.axis('off')
plt.show()
| src/jupyter/VQVAE_show.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''baseline'': virtualenvwrapper)'
# name: python385jvsc74a57bd08c2dedfe5a045a9483c4a9d888731e70df5363cf8c06ebd0819183f007864c1f
# ---
import wandb
import seaborn as sns
import pandas as pd
import time
import cv2
import os
import albumentations
import timm
from pytorch_lightning.loggers import WandbLogger
import pickle
from sklearn.model_selection import StratifiedKFold
from pytorch_lightning.callbacks import ModelCheckpoint,EarlyStopping
from torchvision.models import resnet50,resnet18
from sklearn.preprocessing import LabelEncoder,MultiLabelBinarizer
from albumentations.pytorch import ToTensorV2
import json
import torch.nn.functional as F
import numpy as np
import random
import pytorch_lightning as pl
import torch.optim as optim
import torch.nn as nn
import torchmetrics
from facenet_pytorch import MTCNN,extract_face
from torch.utils.data import Dataset,DataLoader
import torch
from sklearn.utils.class_weight import compute_class_weight
def seeding(seed=2021):
np.random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
torch.random.manual_seed(seed)
random.seed(seed)
seeding()
data = pd.read_csv("UTK.csv")
max(data.age)
class cfg:
img_size=256
max_epochs=100
model_name = "resnet18"
patience = [5,2]
factor= .1
folds=5
min_lr=1e-8
logger = WandbLogger(project="age_gend")
class age_gend_dataset(Dataset):
def __init__(self,df,transforms):
self.df = df
self.transforms = transforms
#self.root_dir = root_dir
def __len__(self):
return len(self.df)
def __getitem__(self,idx):
img = cv2.imread(self.df.loc[idx,'path'])
#print(img.shape)
age = self.df.loc[idx,'age']
gend = self.df.loc[idx,'gender']
assert img is not None
if self.transforms is not None:
img = self.transforms(image=img)['image']
sample = {'image':img,'age':torch.Tensor([age])/116,'gender':torch.Tensor([gend])}
return sample
# +
class MTL_Loss(nn.Module):
def __init__(self,task_num):
super(MTL_Loss,self).__init__()
self.task_num = task_num
self.log_vars = nn.Parameter(torch.zeros((self.task_num)))
def forward(self,pred_age,pred_gend,tar_gend,tar_age):
loss0 = nn.functional.binary_cross_entropy_with_logits(pred_gend,tar_gend)
loss1 = nn.functional.mse_loss(pred_age,tar_age)
precision0 = torch.exp(-self.log_vars[0])
loss0 = precision0*loss0 + self.log_vars[0]
precision1 = torch.exp(-self.log_vars[1])
loss1 = precision1*loss1 + self.log_vars[1]
return loss0+loss1
# -
mtl = MTL_Loss(task_num=2)
# +
def train_augs():
return albumentations.Compose([
albumentations.RandomResizedCrop(cfg.img_size, cfg.img_size, scale=(0.9, 1), p=1),
albumentations.HorizontalFlip(p=0.5),
albumentations.VerticalFlip(p=0.5),
albumentations.ShiftScaleRotate(p=0.5),
albumentations.HueSaturationValue(hue_shift_limit=10, sat_shift_limit=10, val_shift_limit=10, p=0.7),
albumentations.RandomBrightnessContrast(brightness_limit=(-0.2,0.2), contrast_limit=(-0.2, 0.2), p=0.7),
albumentations.CLAHE(clip_limit=(1,4), p=0.5),
albumentations.OneOf([
albumentations.OpticalDistortion(distort_limit=1.0),
albumentations.GridDistortion(num_steps=5, distort_limit=1.),
albumentations.ElasticTransform(alpha=3),
], p=0.2),
albumentations.OneOf([
albumentations.GaussNoise(var_limit=[10, 50]),
albumentations.GaussianBlur(),
albumentations.MotionBlur(),
albumentations.MedianBlur(),
], p=0.2),
albumentations.Resize(cfg.img_size, cfg.img_size),
albumentations.OneOf([
albumentations.JpegCompression(),
albumentations.Downscale(scale_min=0.1, scale_max=0.15),
], p=0.2),
albumentations.IAAPiecewiseAffine(p=0.2),
albumentations.IAASharpen(p=0.2),
albumentations.Cutout(max_h_size=int(cfg.img_size * 0.1), max_w_size=int(cfg.img_size * 0.1), num_holes=5, p=0.5),
albumentations.Normalize(p=1.0),
ToTensorV2()
])
# +
class AgeGendModel(nn.Module):
def __init__(self):
super(AgeGendModel,self).__init__()
if 'resnet18' in cfg.model_name:
self.model = eval(cfg.model_name)(pretrained=False)
for params in self.model.parameters():
params.require_grad = True
self.model = nn.Sequential(*list(self.model.children())[:-2])
self.avg_pool = nn.AdaptiveAvgPool2d(output_size=(1,1))
self.clf = nn.Linear(512,1)
self.reg = nn.Linear(512,1)
def forward(self,x):
x = self.model(x)
x = self.avg_pool(x)
x = x.view(x.size(0),-1)
#print(x.shape)
#x = self.lin1(x)
#x = self.lin2(x)
gend = self.clf(x)
age = torch.sigmoid(self.reg(x))
return gend,age
# -
model2 = AgeGendModel()
# ## Model Class
class AgeGendNet(pl.LightningModule):
def __init__(self):
super(AgeGendNet,self).__init__()
self.model = AgeGendModel()
def forward(self,x):
return self.model(x)
def configure_optimizers(self):
op = optim.Adam(self.model.parameters(),lr=0.01)
scheduler = {
'scheduler':optim.lr_scheduler.CosineAnnealingWarmRestarts(optimizer=op,T_0=10),
'monitor':'val_loss',
'interval':'epoch',
'frequency':1,
'strict':True
}
self.op = op
self.scheduler = scheduler
return [op],[scheduler]
def training_step(self,batch,batch_idx):
y_hat_gend,y_hat_age = self.model(batch['image'])
loss_tr = mtl(pred_gend=y_hat_gend,pred_age=y_hat_age,tar_gend=batch['gender'],tar_age=batch['age'])
#f1_tr = torchmetrics.functional.accuracy(y_hat_gend.sigmoid(),batch['gender'])
#mae_tr = torchmetrics.functional.mean_absolute_error(y_hat_age.relu(),batch['age'])
self.log("TrainLoss",loss_tr,prog_bar=True,on_step=False,on_epoch=True)
return loss_tr
def validation_step(self,batch,batch_idx):
y_hat_gend,y_hat_age = self.model(batch['image'])
loss_val = mtl(pred_gend=y_hat_gend,pred_age=y_hat_age,tar_gend=batch['gender'],tar_age=batch['age'])
#f1_val = torchmetrics.functional.accuracy(y_hat_gend.sigmoid(),batch['gender'])
#mae_val = torchmetrics.functional.mean_absolute_error(y_hat_age.relu(),batch['age'])
self.log("val_loss",loss_val,prog_bar=True,on_step=False,on_epoch=True)
return loss_val
model = AgeGendNet()
train_data = pd.read_csv("UTK.csv")
def main():
i=0
estp = EarlyStopping(monitor="val_loss",mode="min",patience=cfg.patience[0])
if os.path.isdir(f"{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}-2"):
print("Already Exists")
else:
os.makedirs(f"{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}-2")
mod_ckpt = ModelCheckpoint(
monitor='val_loss',
mode='min',
dirpath=f"{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}-2",
filename='Resnet50_256_Checkpoint-ValLoss:{val_loss:.4f}',
save_top_k=1,
)
trainer = pl.Trainer(gpus=1,precision=16,max_epochs=cfg.max_epochs,progress_bar_refresh_rate=30,deterministic=True,benchmark=True,callbacks=[mod_ckpt,estp],logger=cfg.logger,accumulate_grad_batches=1)
print(f"Initializing model Fold - {i+1}/5")
model = AgeGendNet()
print("*** Model Initialization Completed ***")
train_recs = train_data[train_data.folds != i].reset_index(drop=True)
val_recs = train_data[train_data.folds == i].reset_index(drop=True)
train_dataset = age_gend_dataset(df=train_recs,transforms=train_augs())
val_dataset = age_gend_dataset(df=val_recs,transforms=val_augs())
train_loader = DataLoader(train_dataset,batch_size=32,num_workers=4)
val_loader = DataLoader(val_dataset,batch_size=32,num_workers=4)
trainer.fit(model,train_loader,val_loader)
# + tags=["outputPrepend"]
main()
# -
model.eval()
img = cv2.imread("/home/lustbeast/office/Tamannaah_at_an_event_in_Cochin,_July_2018.jpg")
mtcnn = MTCNN(image_size=256,device="cuda:0")
boxes,prob = mtcnn.detect(img)
img = extract_face(img,box=bo)
out = model(img)
torch.sigmoid(out[0])
out[1]
| baseline_age_gend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/elrichgro/irccam-pmodwrc/blob/main/cloudseg/notebooks/colab_training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="uChtrIDy3_R6"
# ## Get code, mount drive
# + colab={"base_uri": "https://localhost:8080/"} id="ovPnm-iuj2r-" outputId="873296b0-69da-4198-fff8-df663fdacc42"
# Get code
# %cd /content
# !rm -rf irccam-pmodwrc/
# !git clone https://github.com/elrichgro/irccam-pmodwrc.git
# + colab={"base_uri": "https://localhost:8080/"} id="HDqKekw6kEwD" outputId="8374f46b-e179-4efc-c0ec-12aad1daa139"
# Mount data
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + [markdown] id="WlelUpu033tt"
# ## Dependencies and data copy
# + id="3VgmeiRgls8b"
# Install dependencies
# # %cd /content/irccam-pmodwrc/
# # !pip install test-tube
# # !pip install -r requirements.txt
requirements = """
torch
torchvision
tqdm
h5py
jupyterlab
opencv-contrib-python
opencv-python
pytorch-lightning==1.0.5
future==0.17.1
scipy
scikit-learn
astral
test-tube
"""
with open('requirements.txt', 'w') as f:
f.write(requirements)
# !pip install -r requirements.txt
# + colab={"base_uri": "https://localhost:8080/"} id="Dh0Ru2snk6vJ" outputId="c1cb1143-6958-493c-c56a-14c7a71d4fbe"
## Copy data
# !rm -rf /content/data
# !mkdir /content/data
# !time cp /content/drive/My\ Drive/dsl/datasets/main_single_label/*.h5 /content/data
# !time cp /content/drive/My\ Drive/dsl/datasets/main_single_label/*.txt /content/data
# + [markdown] id="hQZUPNPT360H"
# ## Train model
# + colab={"base_uri": "https://localhost:8080/"} id="sHlxTn1Rra9H" outputId="421c9e8d-69ea-4e34-d8c9-eeaf39e0df4d"
# Train model
# %cd /content/irccam-pmodwrc
import json
config = {
"batch_size": 4,
"batch_size_val": 8,
"num_epochs": 16,
"model_name": "unet",
"dataset_root": r"/content/data",
"learning_rate": 0.01,
"experiment_name": "colab_test",
"dataset_class": "hdf5",
"log_dir": "/content/drive/My Drive/dsl/training_logs",
"use_clear_sky": True
}
with open("config.json", "w") as f:
json.dump(config, f)
# !PYTHONPATH=$PYTHONPATH:/content/irccam-pmodwrc python cloudseg/training/train.py -c config.json
# + id="a6fXqyU1yRs2"
| cloudseg/notebooks/colab_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd, numpy as np, string, re, pytz
import matplotlib.pyplot as plt, matplotlib.font_manager as fm
from datetime import datetime as dt
# %matplotlib inline
# define the fonts to use for plots
family = 'DejaVu Sans'
title_font = fm.FontProperties(family=family, style='normal', size=20, weight='normal', stretch='normal')
label_font = fm.FontProperties(family=family, style='normal', size=16, weight='normal', stretch='normal')
ticks_font = fm.FontProperties(family=family, style='normal', size=12, weight='normal', stretch='normal')
ticks_font_h = fm.FontProperties(family=family, style='normal', size=10.5, weight='normal', stretch='normal')
def get_colors(cmap, n, start=0., stop=1., alpha=1., reverse=False):
'''return n-length list of rgba colors from the passed colormap name and alpha,
limit extent by start/stop values and reverse list order if flag is true'''
import matplotlib.cm as cm, numpy as np
colors = [cm.get_cmap(cmap)(x) for x in np.linspace(start, stop, n)]
colors = [(r, g, b, alpha) for r, g, b, _ in colors]
return list(reversed(colors)) if reverse else colors
artists_most = pd.read_csv('data/lastfm_top_artists.csv', encoding='utf-8')
artists_most = artists_most.set_index('artist')['play_count'].head(25)
artists_most.head()
# +
ax = artists_most.plot(kind='bar', figsize=[11, 7], width=0.8, alpha=0.7, color='#339933', edgecolor=None, zorder=2)
ax.yaxis.grid(True)
ax.set_xticklabels(artists_most.index,
rotation=45,
rotation_mode='anchor',
ha='right',
fontproperties=ticks_font)
for label in ax.get_yticklabels():
label.set_fontproperties(ticks_font)
ax.set_title('Most Played Artists', fontproperties=title_font)
ax.set_xlabel('', fontproperties=label_font)
ax.set_ylabel('Number of plays', fontproperties=label_font)
#plt.savefig('images/lastfm-artists-played-most.png', dpi=96, bbox_inches='tight')
plt.show()
# +
tracks_most = pd.read_csv('data/lastfm_top_tracks.csv', encoding='utf-8')
def make_label(row, maxlength=30, suffix='...'):
artist = row['artist']
track = row['track']
if len(track) > maxlength:
track = '{}{}'.format(track[:maxlength-len(suffix)], suffix)
return '{}\n{}'.format(artist, track)
index = tracks_most.apply(make_label, axis='columns')
tracks_most = tracks_most.set_index(index).drop(labels=['artist', 'track'], axis='columns')
tracks_most = tracks_most['play_count'].head(20)
tracks_most.head()
# -
ax = tracks_most.sort_values().plot(kind='barh', figsize=[6, 10], width=0.8, alpha=0.6,
color='#003399', edgecolor=None, zorder=2)
ax.xaxis.grid(True)
for label in ax.get_xticklabels():
label.set_fontproperties(ticks_font_h)
for label in ax.get_yticklabels():
label.set_fontproperties(ticks_font_h)
ax.set_xlabel('Number of plays', fontproperties=label_font)
ax.set_ylabel('', fontproperties=label_font)
ax.set_title('Songs I have played the most', fontproperties=title_font, y=1.005)
# plt.savefig('images/lastfm-tracks-played-most-h.png', dpi=96, bbox_inches='tight')
plt.show()
# +
albums_most = pd.read_csv('data/lastfm_top_albums.csv', encoding='utf-8')
def make_label(row, maxlength=25, suffix='...'):
artist = row['artist']
track = row['album']
if len(track) > maxlength:
track = '{}{}'.format(track[:maxlength-len(suffix)], suffix)
return '{}\n{}'.format(artist, track)
index = albums_most.apply(make_label, axis='columns')
albums_most = albums_most.set_index(index).drop(labels=['artist', 'album'], axis='columns')
albums_most = albums_most['play_count'].head(30)
albums_most.head()
# -
ax = albums_most.sort_values().plot(kind='barh', figsize=[6.5, 15], width=0.8, alpha=0.6, color='#990066',
edgecolor=None, zorder=2)
ax.xaxis.grid(True)
for label in ax.get_xticklabels():
label.set_fontproperties(ticks_font_h)
for label in ax.get_yticklabels():
label.set_fontproperties(ticks_font_h)
ax.set_xlabel('Number of plays', fontproperties=label_font)
ax.set_ylabel('', fontproperties=label_font)
ax.set_title('Albums I have played the most', fontproperties=title_font, y=1.005)
# plt.savefig('images/lastfm-albums-played-most-h.png', dpi=96, bbox_inches='tight')
plt.show()
# read the all-time scrobbles data set
scrobbles = pd.read_csv('data/lastfm_scrobbles.csv', encoding='utf-8')
scrobbles = scrobbles.drop('timestamp', axis=1)
print('{:,} total scrobbles'.format(len(scrobbles)))
print('{:,} total artists'.format(len(scrobbles['artist'].unique())))
# convert to datetime
scrobbles['timestamp'] = pd.to_datetime(scrobbles['datetime'])
# functions to convert UTC to <timezone> and extract date/time elements
convert_tz = lambda x: x.to_pydatetime().replace(tzinfo=pytz.utc).astimezone(pytz.timezone('US/Central'))
get_year = lambda x: convert_tz(x).year
get_month = lambda x: '{}-{:02}'.format(convert_tz(x).year, convert_tz(x).month) #inefficient
get_day = lambda x: convert_tz(x).day
get_hour = lambda x: convert_tz(x).hour
get_day_of_week = lambda x: convert_tz(x).weekday()
# +
# parse out date and time elements as pacific time
scrobbles['year'] = scrobbles['timestamp'].map(get_year)
scrobbles['month'] = scrobbles['timestamp'].map(get_month)
scrobbles['day'] = scrobbles['timestamp'].map(get_day)
scrobbles['hour'] = scrobbles['timestamp'].map(get_hour)
scrobbles['dow'] = scrobbles['timestamp'].map(get_day_of_week)
scrobbles = scrobbles.drop(labels=['datetime'], axis=1)
# drop rows with 01-01-1970 as timestamp
scrobbles = scrobbles[scrobbles['year'] > 1970]
scrobbles.head()
# -
year_counts = scrobbles['year'].value_counts().sort_index()
# +
year_counts = scrobbles['year'].value_counts().sort_index()
ax = year_counts.plot(kind='line', figsize=[10, 5], linewidth=4, alpha=1, marker='o', color='#6684c1',
markeredgecolor='#6684c1', markerfacecolor='w', markersize=8, markeredgewidth=2)
ax.set_xlim((year_counts.index[0], year_counts.index[-1]))
ax.yaxis.grid(True)
ax.xaxis.grid(True)
ax.set_ylim(0, 10000)
ax.set_xticks(year_counts.index)
ax.set_ylabel('Number of plays', fontproperties=label_font)
ax.set_xlabel('Years', fontproperties=label_font)
ax.set_title('Number of songs played per year', fontproperties=title_font)
# plt.savefig('images/lastfm-scrobbles-per-year.png', dpi=96, bbox_inches='tight')
plt.show()
# -
# get all the scrobbles from 2010-present
min_year = 2019
scrobbles_10 = scrobbles[scrobbles['year'] >= min_year]
max_year = max(scrobbles_10['year'])
# +
# count number of scrobbles in each month
month_counts = scrobbles_10['month'].value_counts().sort_index()
# not every month necessarily has a scrobble, so fill in missing months with zero counts
date_range = pd.date_range(start=min(scrobbles_10['timestamp']), end=max(scrobbles_10['timestamp']), freq='D')
months_range = date_range.map(lambda x: str(x.date())[:-3])
index = np.unique(months_range)
month_counts = month_counts.reindex(index, fill_value=0)
# +
ax = month_counts.plot(kind='line', figsize=[12, 5], linewidth=4, alpha=0.6, color='#003399')
xlabels = month_counts.iloc[range(0, len(month_counts), 12)].index
xlabels = [x if x in xlabels else '' for x in month_counts.index]
ax.set_xticks(range(len(xlabels)))
ax.set_xticklabels(xlabels, rotation=40, rotation_mode='anchor', ha='right')
ax.set_xlim((0, len(month_counts)-1))
ax.yaxis.grid(True)
ax.set_ylim((0,1500))
ax.set_ylabel('Number of plays', fontproperties=label_font)
ax.set_xlabel('Months', fontproperties=label_font)
ax.set_title('Number of songs played per month, {}-{}'.format(min_year, max_year), fontproperties=title_font)
# plt.savefig('images/lastfm-scrobbles-per-month.png', dpi=96, bbox_inches='tight')
plt.show()
# -
# get the play count sum by day of the week
dow_counts = scrobbles['dow'].value_counts().sort_index()
dow_counts.index = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
# +
ax = dow_counts.plot(kind='bar', figsize=[6, 5], width=0.7, alpha=0.6, color='#003399', edgecolor=None, zorder=2)
ax.yaxis.grid(True)
ax.set_xticklabels(dow_counts.index, rotation=35, rotation_mode='anchor', ha='right', fontproperties=ticks_font)
for label in ax.get_yticklabels():
label.set_fontproperties(ticks_font)
ax.set_ylim((0, 1400))
ax.set_title('Songs played per day of the week', fontproperties=title_font)
ax.set_xlabel('', fontproperties=label_font)
ax.set_ylabel('Number of plays', fontproperties=label_font)
# plt.savefig('images/lastfm-scrobbles-per-weekday.png', dpi=96, bbox_inches='tight')
plt.show()
# +
hour_counts = scrobbles['hour'].value_counts().sort_index()
ax = hour_counts.plot(kind='line', figsize=[10, 5], linewidth=4, alpha=1, marker='o', color='#6684c1',
markeredgecolor='#6684c1', markerfacecolor='w', markersize=8, markeredgewidth=2)
xlabels = hour_counts.index.map(lambda x: '{:02}:00'.format(x))
ax.set_xticks(range(len(xlabels)))
ax.set_xticklabels(xlabels, rotation=45, rotation_mode='anchor', ha='right')
ax.set_xlim((hour_counts.index[0], hour_counts.index[-1]))
ax.yaxis.grid(True)
ax.set_ylim((0, 1000))
ax.set_ylabel('Number of plays', fontproperties=label_font)
ax.set_xlabel('', fontproperties=label_font)
ax.set_title('Number of songs played per hour of the day', fontproperties=title_font)
# plt.savefig('images/lastfm-scrobbles-per-hour.png', dpi=96, bbox_inches='tight')
plt.show()
# -
# get the play counts by hour of day and day of week
weekday_hour_counts = scrobbles.groupby(['dow','hour']).count()['track']
hour_numbers = weekday_hour_counts.index.levels[1]
day_numbers = weekday_hour_counts.index.levels[0]
day_names = {0:'Monday', 1:'Tuesday', 2:'Wednesday', 3:'Thursday', 4:'Friday', 5:'Saturday', 6:'Sunday'}
# +
# get one color per day of week
colors = get_colors('nipy_spectral_r', n=len(day_numbers), start=0.1, stop=0.95)
fig, ax = plt.subplots(figsize=[10, 6])
lines = []
for day, c in zip(day_numbers, colors):
ax = weekday_hour_counts[day].plot(kind='line', linewidth=4, alpha=0.6, c=c)
lines.append(day_names[day])
xlabels = hour_numbers.map(lambda x: '{:02}:00'.format(x))
ax.set_xticks(range(len(xlabels)))
ax.set_xticklabels(xlabels, rotation=45, rotation_mode='anchor', ha='right')
ax.set_xlim((hour_numbers[0], hour_numbers[-1]))
ax.yaxis.grid(True)
ax.set_ylim([0, 100])
ax.set_ylabel('Number of plays', fontproperties=label_font)
ax.set_xlabel('', fontproperties=label_font)
ax.set_title('Number of songs played, by day of week and hour of day', fontproperties=title_font)
ax.legend(lines, loc='upper right', bbox_to_anchor=(1.23,1.017))
# plt.savefig('images/lastfm-scrobbles-days-hours.png', dpi=96, bbox_inches='tight')
plt.show()
# -
#Add additional years to the 'isin' list to query multiple years.
scrobbles_year = scrobbles[scrobbles['year'].isin([2019,2020])]
len(scrobbles_year)
# what artists did i play the most that year?
artists_year = scrobbles_year['artist'].value_counts()
artists_year = pd.DataFrame(artists_year).reset_index().rename(columns={'artist':'play count', 'index':'artist'})
artists_year.index = [n + 1 for n in artists_year.index]
artists_year.head(10)
# what tracks did i play the most that year?
tracks_year = scrobbles_year.groupby(['artist', 'track']).count().sort_values('timestamp', ascending=False)
tracks_year = tracks_year.reset_index().rename(columns={'timestamp':'play count'})[['artist', 'track', 'play count']]
tracks_year.index = [n + 1 for n in tracks_year.index]
tracks_year.head(10)
# +
# what albums did i play the most that year?
albums_year = scrobbles_year.groupby(['artist', 'album']).count().sort_values('timestamp', ascending=False)
albums_year = albums_year.reset_index().rename(columns={'timestamp':'play count'})[['artist', 'album', 'play count']]
albums_year.index = [n + 1 for n in albums_year.index]
# remove text in parentheses or brackets
regex = re.compile('\\(.*\\)|\\[.*]')
albums_year['album'] = albums_year['album'].map(lambda x: regex.sub('', x))
albums_year.head(10)
# -
# Add additional months to the 'isin' list to query multiple months.
scrobbles_month = scrobbles[scrobbles['month'].isin(['2020-01'])]
len(scrobbles_month)
# what artists did i play the most that month?
artists_month = scrobbles_month['artist'].value_counts()
artists_month = pd.DataFrame(artists_month).reset_index().rename(columns={'artist':'play count', 'index':'artist'})
artists_month.index = [n + 1 for n in artists_month.index]
artists_month.head(10)
# what tracks did i play the most that month?
tracks_month = scrobbles_month.groupby(['artist', 'track']).count().sort_values('timestamp', ascending=False)
tracks_month = tracks_month.reset_index().rename(columns={'timestamp':'play count'})[['artist', 'track', 'play count']]
tracks_month.index = [n + 1 for n in tracks_month.index]
tracks_month.head(12)
# what albums did i play the most that month?
albums_month = scrobbles_month.groupby(['artist', 'album']).count().sort_values('timestamp', ascending=False)
albums_month = albums_month.reset_index().rename(columns={'timestamp':'play count'})[['artist', 'album', 'play count']]
albums_month.index = [n + 1 for n in albums_month.index]
albums_month.head(12)
# when were the last 5 times I played something by <artist>?
scrobbles[scrobbles['artist'].str.contains('Ongkara')].head()
# when were the last 5 times I played something off of <album>?
scrobbles[scrobbles['album'].fillna('').str.contains('Heritage')].head()
# when were the last 5 times I played <track>?
scrobbles[scrobbles['track'].str.contains('PLANET B')].head()
# +
# get the cumulative play counts since <year> for the top n most listened-to artists
n = 10
plays = scrobbles[scrobbles['artist'].isin(artists_most.head(n).index)]
plays = plays[plays['year'] >= 2019]
plays = plays.groupby(['artist','year']).count().groupby(level=[0]).cumsum()['track']
# make sure we have each year represented for each artist, even if they got no plays that year
plays = plays.unstack().T.fillna(method='ffill').T.stack()
top_artists = plays.index.levels[0]
# +
# get one color per artist
colors = get_colors('Dark2', n)
fig, ax = plt.subplots(figsize=[8, 6])
lines = []
for artist, c in zip(top_artists, colors):
ax = plays[artist].plot(kind='line', linewidth=4, alpha=0.6, marker='o', c=c)
lines.append(artist)
ax.set_xlim((plays.index.get_level_values(1).min(), plays.index.get_level_values(1).max()))
ax.yaxis.grid(True)
ax.set_xticklabels(plays.index.levels[1], rotation=0, rotation_mode='anchor', ha='center')
ax.set_ylabel('Cumulative number of plays', fontproperties=label_font)
ax.set_xlabel('Year', fontproperties=label_font)
ax.set_title('Cumulative number of plays per artist over time', fontproperties=title_font)
ax.legend(lines, loc='upper right', bbox_to_anchor=(1.33, 1.016))
# plt.savefig('images/lastfm-scrobbles-top-artists-years.png', dpi=96, bbox_inches='tight')
plt.show()
# -
| lastfm/lastfm_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# http://ankivil.com/kaggle-first-steps-with-julia-chars74k-first-place-using-convolutional-neural-networks/
#
# https://github.com/erhwenkuo/deep-learning-with-keras-notebooks/blob/master/2.0-first-steps-with-julia.ipynb
#
# http://florianmuellerklein.github.io/cnn_streetview/
# # Introduction
# In this article, I will describe how to design a Convolutional Neural Network (CNN) with Keras to score over 0.86 accuracy in the Kaggle competition First Steps With Julia. I will explain precisely how to get to this result, from data to submission. All the python code is, of course, included. This work is inspired by <NAME>’s Using deep learning to read street signs.
#
# The goal of the Kaggle competition First Steps With Julia is to classify images of characters taken from natural images. These images come from a subset of the Chars74k data set. This competition normally serves as a tutorial on how to use the Julia language but a CNN is the tool of choice to tackle this kind of problem.
#
# http://florianmuellerklein.github.io/cnn_streetview/
import os
os.listdir()
# # Data Preprocessing: Image Color
# Almost all images in the train and test sets are color images. The first step in the preprocessing is to convert all images to grayscale. It simplifies the data fed to the network and makes it easier to generalize, a blue letter being equivalent to a red letter. This preprocessing should have almost no negative impact on the final accuracy because most texts have high contrast with their background.
# # Data Preprocessing: Image Resizing
# As the images have different shapes and size, we have to normalize them for the model. There are two main questions for this normalization: which size do we choose? and do we keep the aspect ratio?
#
# Initially, I thought keeping the aspect ratio would be better because it would not distort the image arbitrarily. It could also lead to confusion between O and 0 (capital o and zero). However, after some tests, it seems that the results are better without keeping the aspect ratio. Maybe my filling strategy (see the code below) is not the best one.
#
# Concerning the image size, 16×16 images allow very fast training but don’t give the best results. These small images are perfect to rapidly test ideas. Using 32×32 images makes the training quite fast and gives good accuracy. Finally, using 64×64 images makes the training quite slow and marginally improves the results compared to 32×32 images. I chose to use 32×32 images because it is the best trade-off between speed and accuracy.
import csv
#fn = open('trainLabels.csv', 'r')
#train_label = [dict(i) for i in csv.DictReader(fn)]
#for i in csv.reader(fn):
# print(i)
#fn.close()
#import pandas as pd
#pd.DataFrame(train_label)
# # Data Preprocessing: Label Conversion
# We also have to convert the labels from characters to one-hot vectors. This is mandatory to feed the labels information to the network. This is a two-step procedure. First, we have to find a way to convert characters to consecutive integers and back. Second, we have to convert each integer to a one-hot vector.
# +
def label2int(ch):
asciiVal = ord(ch)
if(asciiVal<=57): #0-9
asciiVal-=48
elif(asciiVal<=90): #A-Z
asciiVal-=55
else: #a-z
asciiVal-=61
return asciiVal
def int2label(i):
if(i<=9): #0-9
i+=48
elif(i<=35): #A-Z
i+=55
else: #a-z
i+=61
return chr(i)
# -
# # Code for processing data
path = "."
os.path.exists( path + "/trainResized" )
if not os.path.exists( path + "/trainResized" ):
os.makedirs( path + "/trainResized" )
if not os.path.exists( path + "/testResized" ):
os.makedirs( path + "/testResized" )
# +
import glob
import numpy as np
import pandas as pd
from skimage.transform import resize
from skimage.io import imread, imsave
#trainFiles = glob.glob( path + "/train/*" )
#for i, nameFile in enumerate(trainFiles):
# image = imread( nameFile )
# imageResized = resize( image, (20,20) )
# newName = "/".join( nameFile.split("/")[:-1] ) + "Resized/" + nameFile.split("/")[-1]
# print("/".join( nameFile.split("/")[:-1] ) + 'Resized/' + nameFile.split("/")[-1])
# imsave ( newName, imageResized )
# if i == 1:
# print(image.shape) # (89, 71, 3)
# print(imageResized.shape) # (20, 20, 3)
#testFiles = glob.glob( path + "/test/*" )
#for i, nameFile in enumerate(testFiles):
# image = imread( nameFile )
# imageResized = resize( image, (20,20) )
# newName = "/".join( nameFile.split("/")[:-1] ) + "Resized/" + nameFile.split("/")[-1]
# imsave ( newName, imageResized )
# +
import os
import glob
import pandas as pd
import math
import numpy as np
from scipy.misc import imread, imsave, imresize
from natsort import natsorted
# Path of data files
path = "."
# Input image dimensions
img_rows, img_cols = 32, 32
# Keep or not the initial image aspect ratio
keepRatio = False
# Create the directories if needed
if not os.path.exists( path + "/trainResized"):
os.makedirs(path + "/trainResized")
if not os.path.exists( path + "/testResized"):
os.makedirs(path + "/testResized")
### Images preprocessing ###
for setType in ["train", "test"]:
# We have to make sure files are sorted according to labels, even if they don't have trailing zeros
files = natsorted(glob.glob(path + "/"+setType+"/*"))
data = np.zeros((len(files), img_rows, img_cols)) #will add the channel dimension later
for i, filepath in enumerate(files):
image = imread(filepath, True) #True: flatten to grayscale
if keepRatio:
# Find the largest dimension (height or width)
maxSize = max(image.shape[0], image.shape[1])
# Size of the resized image, keeping aspect ratio
imageWidth = math.floor(img_rows*image.shape[0]/maxSize)
imageHeigh = math.floor(img_cols*image.shape[1]/maxSize)
# Compute deltas to center image (should be 0 for the largest dimension)
dRows = (img_rows-imageWidth)//2
dCols = (img_cols-imageHeigh)//2
imageResized = np.zeros((img_rows, img_cols))
imageResized[dRows:dRows+imageWidth, dCols:dCols+imageHeigh] = imresize(image, (imageWidth, imageHeigh))
# Fill the empty image with the median value of the border pixels
# This value should be close to the background color
val = np.median(np.append(imageResized[dRows,:],
(imageResized[dRows+imageWidth-1,:],
imageResized[:,dCols],
imageResized[:,dCols+imageHeigh-1])))
# If rows were left blank
if(dRows>0):
imageResized[0:dRows,:].fill(val)
imageResized[dRows+imageWidth:,:].fill(val)
# If columns were left blank
if(dCols>0):
imageResized[:,0:dCols].fill(val)
imageResized[:,dCols+imageHeigh:].fill(val)
else:
imageResized = imresize(image, (img_rows, img_cols))
# Add the resized image to the dataset
data[i] = imageResized
#Save image (mostly for visualization)
filename = filepath.split("/")[-1]
filenameDotSplit = filename.split(".")
newFilename = str(int(filenameDotSplit[0])).zfill(5) + "." + filenameDotSplit[-1].lower() #Add trailing zeros
newName = "/".join(filepath.split("/")[:-1] ) + 'Resized' + "/" + newFilename
imsave(newName, imageResized)
# Add channel/filter dimension
data = data[:,:,:, np.newaxis]
# Makes values floats between 0 and 1 (gives better results for neural nets)
data = data.astype('float32')
data /= 255
# Save the data as numpy file for faster loading
np.save(path+"/"+setType+ 'ResizedData' +".npy", data)
# -
# # Load Resized images to data for the input of network
# Load data from reSized images
for i_type in ['train', 'test']:
files = natsorted(glob.glob('./' + i_type + 'Resized/*'))
data = np.zeros((len(files), img_rows, img_cols))
for i, i_path in enumerate(files):
data[i] = imread(i_path, True)
data = data[:, :, :, np.newaxis]
data = data.astype('float32')
data /= 255
np.save(path+"/"+i_type+ 'ResizedData' +".npy", data)
# +
### Labels preprocessing ###
# Load labels
y_train = pd.read_csv(path+"/trainLabels.csv").values[:,1] #Keep only label
# Convert labels to one-hot vectors
Y_train = np.zeros((y_train.shape[0], len(np.unique(y_train))))
for i in range(y_train.shape[0]):
Y_train[i][label2int(y_train[i])] = 1 # One-hot
# Save preprocessed label to nupy file for faster loading
np.save(path+"/"+"labelsPreproc.npy", Y_train)
# -
# # Data Augmentation
# Instead of using the training data as it is, we can apply some augmentations to artificially increase the size of the training set with “new” images. Augmentations are random transformations applied to the initial data to produce a modified version of it. These transformations can be a zoom, a rotation, etc. or a combination of all these.
#
# https://keras.io/preprocessing/image/#imagedatagenerator
#
# # Using ImageDataGenerator
#
#
# The ImageDataGenerator constructor takes several parameters to define the augmentations we want to use. I will only go through the parameters useful for our case, see the documentation if you need other modifications to your images:
#
# **featurewise_center , featurewise_std_normalization and zca_whitening are not used as they don’t increase the performance of the network. If you want to test these options, be sure to compute the relevant quantities with fit and apply these modifications to your test set with standardize .
#
# **rotation_range Best results for values around 20.
#
# **width_shift_range Best results for values around 0.15.
#
# **height_shift_range Best results for values around 0.15.
#
# **shear_range Best results for values around 0.4.
#
# **zoom_range Best results for values around 0.3.
#
# **channel_shift_range Best results for values around 0.1.
#
# Of course, I didn’t test all the combinations, so there must be others values which increase the final accuracy. Be careful though, too much augmentation (high parameter values) will make the learning slow or even impossible.
#
# I also added the possibility for the ImageDataGenerator to randomly invert the values, the code is below. The parameters are:
#
# **channel_flip Best set to True.
#
# **channel_flip_max Should be set to 1. as we normalized the data between 0 and 1.
#
#
# 使用 ImageDataGenerator
# ImageDataGenerator構建函數需要幾個參數來定義我們想要使用的增強效果。我只會通過對我們的案例有用的參數進行設定,如果您需要對您的圖像進行其他修改,請參閱Keras文檔。
#
# featurewise_center,featurewise_std_normalization和zca_whitening不使用,因為在本案例裡它們不會增加網絡的性能。如果你想測試這些選項,一定要合適地計算相關的數量,並將這些修改應用到你的測試集中進行標準化。
#
# rotation_range 20左右的值效果最好。
#
# width_shift_range 0.15左右的值效果最好。
#
# height_shift_range 0.15左右的值效果最好。
#
# shear_range 0.4 左右的值效果最好。
#
# zoom_range 0.3 左右的值效果最好。
#
# channel_shift_range 0.1左右的值效果最好。
#
# 當然,我沒有測試所有的組合,所以可能還有其他值的組合可以用來提高最終的準確度。但要小心,太多的增量(高參數值)會使學習變得緩慢甚至跑不出來。
#
# # 模型學習 (Learning)
#
# 對於模型的訓練,我使用了分類交叉熵(cross-entropy)作為損失函數(loss function),最後一層使用softmax的激勵函數。
#
# # 演算法 (Algorithm)
#
# 在這個模型裡我選擇使用AdaMax和AdaDelta來作為優化器(optimizer),而不是使用經典的隨機梯度下降(SGD)算法。 同時我發現AdaMax比AdaDelta在這個問題上會給出更好的結果。但是,對於具有眾多濾波器和大型完全連接層的複雜網絡,AdaMax在訓練循環不太收斂,甚至無法完全收斂。因此在這次的網絡訓練過程我拆成二個階段。 第一個階段,我先使用AdaDelta進行了20個循環的前期訓練為的是要比較快速的幫忙卷積網絡的模型收斂。第二個階段,則利用AdaMax來進行更多訓練循環與更細微的修正來得到更好的模型。如果將網絡的大小除以2,則不需要使用該策略。
#
# # 訓練批次量 (Batch Size)
# 在保持訓練循環次數不變的同時,我試圖改變每次訓練循環的批量大小(batch size)。大的批量(batch)會使算法運行速度更快,但結果效能不佳。 這可能是因為在相同數量的數據量下,更大的批量意味著更少的模型權重的更新。無論如何,在這個範例中最好的結果是在批量(batch size) 設成 128的情況下達到的。
#
# # 網絡層的權重初始 (Layer Initialization)
#
# 如果網絡未正確初始化,則優化算法可能無法找到最佳值。我發現使用he_normal來進行初始化會使模型的學習變得更容易。在Keras中,你只需要為每一層使用kernel_initializer='he_normal'參數。
#
# # 學習率衰減 (Learning Rate Decay)
#
# 在訓練期間逐漸降低學習率(learning rate)通常是一個好主意。它允許算法微調參數,並接近局部最小值。 但是,我發現使用AdaMax的optimizer,在
# 沒有設定學習速率衰減的情況下結果更好,所以我們現在不必擔心。
#
# # 訓練循環 (Number of Epochs)
#
# 使用128的批量大小,沒有學習速度衰減,我測試了200到500個訓練循環。即使運行到第500個訓練循環,整個網絡模型似乎也沒出現過擬合(overfitting)的情形。 我想這肯定要歸功於Dropout的設定發揮了功效。我發現500個訓練循環的結果比300個訓練循環略好。最後的模型我用了500個訓練循環,但是如果你在CPU上運行,300個訓練循環應該就足夠了。
#
# # 交叉驗證 (Cross-Validation)
#
# 為了評估不同模型的質量和超參數的影響,我使用了蒙特卡洛交叉驗證:我隨機分配了初始數據1/4進行驗證,並將3/4進行學習。 我還使用分裂技術,確保在我們的例子中,每個類別約有1/4圖像出現在測試集中。這導致更穩定的驗證分數。
# # Code
import numpy as np
import os
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.convolutional import Convolution2D, MaxPooling2D
from keras.callbacks import ModelCheckpoint
from sklearn.model_selection import train_test_split
# +
# setting parameters for the network
batch_size = 128 # 訓練批次量 (Batch Size)
nb_classes = 62 # A-Z, a-z, 0-9共有62個類別
nb_epoch = 500 # 進行500個訓練循環
# Input image dimensions
# 要輸入到第一層網絡的圖像大小 (32像素 x 32像素)
img_height, img_width = 32, 32
# -
# 相關資料的路徑
path = "."
# 載入預處理好的訓練資料與標籤
X_train_all = np.load(path+"/trainResizedData.npy")
Y_train_all = np.load(path+"/labelsPreproc.npy")
# 將資料區分為訓練資料集與驗證資料集
X_train, X_val, Y_train, Y_val = train_test_split(X_train_all, Y_train_all, test_size=0.25, stratify=np.argmax(Y_train_all, axis=1))
# For each image data, what dimension does it have?
print(X_train.shape)
print(Y_train.shape)
# # 設定圖像增強(data augmentation)的設定
datagen = ImageDataGenerator(
rotation_range = 20,
width_shift_range = 0.15,
height_shift_range = 0.15,
shear_range = 0.4,
zoom_range = 0.3,
channel_shift_range = 0.1)
# # Build CNN
# +
### 卷積網絡模型架構 ###
model = Sequential()
# 25 filter, each one has size 3*3
model.add(Convolution2D(128,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu',
input_shape=(img_height, img_width, 1)))
model.add(Convolution2D(128,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(256,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu'))
model.add(Convolution2D(256,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(512,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu'))
model.add(Convolution2D(512,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu'))
model.add(Convolution2D(512,(3, 3), padding='same', kernel_initializer='he_normal', activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(4096, kernel_initializer='he_normal', activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(4096, kernel_initializer='he_normal', activation='relu'))
model.add(Dropout(0.5))
# output; we have nb_classes. Therefore, we put this dense layer with nb_classes nodes.
model.add(Dense(nb_classes, kernel_initializer='he_normal', activation='softmax'))
# 展現整個模型架構
model.summary()
# -
# # Training Setting
# +
# First, we use AdaDelta to train our model.
model.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=["accuracy"])
# We take epochs = 20.
model.fit(X_train, Y_train, batch_size=batch_size,
epochs=20,
validation_data=(X_val, Y_val),
verbose=1)
# Second, we use AdaMax to train our model subsequently.
model.compile(loss='categorical_crossentropy',
optimizer='adamax',
metrics=["accuracy"])
# Here, we will save the better model with great validation during our training.
saveBestModel = ModelCheckpoint("best.kerasModelWeights", monitor='val_acc', verbose=1, save_best_only=True, save_weights_only=True)
# Moreover, in this training step, we will generate images from ImageDataGenrator to add our second training process.
history = model.fit_generator(datagen.flow(X_train, Y_train, batch_size=batch_size),
steps_per_epoch=len(X_train)/batch_size,
epochs=nb_epoch,
validation_data=(X_val, Y_val),
callbacks=[saveBestModel],
verbose=1)
### 進行預測 ###
# 載入訓練過程中驗證結果最好的模型
model.load_weights("best.kerasModelWeights")
# 載入Kaggle測試資料集
X_test = np.load(path+"/testPreproc.npy")
# 預測字符的類別
Y_test_pred = model.predict_classes(X_test)
# +
# 從類別的數字轉換為字符
vInt2label = np.vectorize(int2label)
Y_test_pred = vInt2label(Y_test_pred)
# 保存預測結果到檔案系統
np.savetxt(path+"/jular_pred" + ".csv", np.c_[range(6284,len(Y_test_pred)+6284),Y_test_pred], delimiter=',', header = 'ID,Class', comments = '', fmt='%s')
# +
# 透過趨勢圖來觀察訓練與驗證的走向 (特別去觀察是否有"過擬合(overfitting)"的現象)
import matplotlib.pyplot as plt
# 把每個訓練循環(epochs)的相關重要的監控指標取出來
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
# 取得整個訓練循環(epochs)的總次數
epochs = range(len(acc))
# 把"訓練準確率(Training acc)"與"驗證準確率(Validation acc)"的趨勢線形表現在圖表上
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
# 把"訓練損失(Training loss)"與"驗證損失(Validation loss)"的趨勢線形表現在圖表上
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
#
# 從"Training與validation accuracy"的線型圖來看, 訓練到50~60循環(epochs)之後驗證的準確率就提不上去了, 但是訓練的準確率確可以一直提高。 雖然說83%的預測準確率在Kaggle的competition裡己經是前10名左右了, 但如果想要繼續提升效果的話可的的方向:
# 增加更多的字符圖像
# 字符圖像的增強的調教(可以增加如原文提及的影像頻導channel的flip,在這個文章為了簡化起見移除了這個部份的實作)
| Guid Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: timeeval
# language: python
# name: timeeval
# ---
# # MIT-BIH Supraventricular Arrhythmia Database (_svdb_)
#
# Part of the ECG Database Collection:
#
# | Short Name | Long Name |
# | :--- | :--- |
# | _mitdb_ | MIT-BIH Arrhythmia Database |
# | _svdb_ | MIT-BIH Supraventricular Arrhythmia Database |
# | _ltdb_ | MIT-BIH Long-Term ECG Database |
#
# [Docu](https://wfdb.readthedocs.io/en/latest) of the `wfdb`-package.
# %matplotlib inline
import pandas as pd
import numpy as np
import wfdb
import os
from typing import Final
from collections.abc import Callable
from config import data_raw_folder, data_processed_folder
from timeeval import Datasets
import matplotlib.pyplot as plt
from IPython.display import display, Markdown, Latex
# +
dataset_collection_name = "SVDB"
source_folder = os.path.join(data_raw_folder, "MIT-BIH Supraventricular Arrhythmia Database")
target_folder = data_processed_folder
from pathlib import Path
print(f"Looking for source datasets in {Path(source_folder).absolute()} and\nsaving processed datasets in {Path(target_folder).absolute()}")
# -
def load_dataset_names() -> list[str]:
with open(os.path.join(source_folder, "RECORDS"), 'r') as f:
records = [l.rstrip('\n') for l in f]
return records
def transform_and_label(source_file: str, target: str) -> int:
print(f"Transforming {os.path.basename(source_file)}")
# load dataset
record = wfdb.rdrecord(source_file)
df_record = pd.DataFrame(record.p_signal, columns=record.sig_name)
print(f" record {record.file_name[0]} loaded")
# load annotation file
atr = wfdb.rdann(source_file, "atr")
assert record.fs == atr.fs, "Sample frequency of records and annotations does not match!"
df_annotation = pd.DataFrame(atr.symbol, index=atr.sample, columns=["Label"])
df_annotation = df_annotation.reset_index()
df_annotation.columns = ["position", "label"]
print(f" {atr.ann_len} beat annotations for {source_file} loaded")
# calculate normal beat length
print(" preparing windows for labeling...")
df_normal_beat = df_annotation.copy()
df_normal_beat["prev_position"] = df_annotation["position"].shift()
df_normal_beat["prev_label"] = df_annotation["label"].shift()
df_normal_beat = df_normal_beat[(df_normal_beat["label"] == "N") & (df_normal_beat["prev_label"] == "N")]
s_normal_beat_lengths = df_normal_beat["position"] - df_normal_beat["prev_position"]
print(f" normal beat distance samples = {len(s_normal_beat_lengths)}")
normal_beat_length = s_normal_beat_lengths.median()
if (normal_beat_length % 2) == 0:
normal_beat_length += 1
beat_window_size = int(normal_beat_length)
beat_window_margin = (beat_window_size - 1)//2
print(f" window size = {beat_window_size}")
print(f" window margins (left and right) = {beat_window_margin}")
# calculate beat windows
## for external anomalies
df_ext = df_annotation[(df_annotation["label"] == "|") | (df_annotation["label"] == "Q")].copy()
df_ext["window_start"] = df_ext["position"]-beat_window_margin
df_ext["window_end"] = df_ext["position"]+beat_window_margin
df_ext = df_ext[["position", "window_start", "window_end"]]
print(f" {len(df_ext)} windows for external anomalies")
## for anomalous beats
df_svf = df_annotation[(df_annotation["label"] != "|") & (df_annotation["label"] != "~") & (df_annotation["label"] != "+")].copy()
df_svf["position_next"] = df_svf["position"].shift(-1)
df_svf["position_prev"] = df_svf["position"].shift(1)
#df_svf = df_svf[(df_svf["position_prev"].notnull()) & (df_svf["position_next"].notnull())]
df_svf = df_svf[(df_svf["label"] != "Q") & (df_svf["label"] != "N")]
df_svf["window_start"] = np.minimum(df_svf["position"].values-beat_window_margin, df_svf["position_prev"].values+beat_window_margin)
df_svf["window_end"] = np.maximum(df_svf["position"].values+beat_window_margin, df_svf["position_next"].values-beat_window_margin)
df_svf = df_svf[["position", "window_start", "window_end"]]
print(f" {len(df_svf)} windows for anomalous beats")
## merge
df_windows = pd.concat([df_ext, df_svf])
print(f" ...done.")
# add labels based on anomaly windows
print(" labeling")
df_record["is_anomaly"] = 0
for _, (_, t1, t2) in df_windows.iterrows():
tmp = df_record[df_record.index >= t1]
tmp = tmp[tmp.index <= t2]
df_record["is_anomaly"].values[tmp.index] = 1
# reconstruct timestamps and set as index
print(" reconstructing timestamps")
df_record["timestamp"] = pd.to_datetime(df_record.index.values * 1e+9/record.fs, unit='ns')
df_record = df_record.set_index("timestamp")
df_record.to_csv(target)
print(f"Dataset {os.path.basename(source_file)} transformed and saved!")
# return dataset length
return record.sig_len
# +
# shared by all datasets
dataset_type = "real"
input_type = "multivariate"
datetime_index = True
train_type = "unsupervised"
train_is_normal = False
# create target directory
dataset_subfolder = os.path.join(input_type, dataset_collection_name)
target_subfolder = os.path.join(target_folder, dataset_subfolder)
try:
os.makedirs(target_subfolder)
print(f"Created directories {target_subfolder}")
except FileExistsError:
print(f"Directories {target_subfolder} already exist")
pass
dm = Datasets(target_folder)
# +
# dataset transformation
transform_file: Callable[[str, str], int] = transform_and_label
for dataset_name in load_dataset_names():
# intentionally no file suffix (.dat)
source_file = os.path.join(source_folder, dataset_name)
filename = f"{dataset_name}.test.csv"
path = os.path.join(dataset_subfolder, filename)
target_filepath = os.path.join(target_subfolder, filename)
# transform file and label it
dataset_length = transform_file(source_file, target_filepath)
print(f"Processed source dataset {source_file} -> {target_filepath}")
# save metadata
dm.add_dataset((dataset_collection_name, dataset_name),
train_path = None,
test_path = path,
dataset_type = dataset_type,
datetime_index = datetime_index,
split_at = None,
train_type = train_type,
train_is_normal = train_is_normal,
input_type = input_type,
dataset_length = dataset_length
)
# save metadata of benchmark
dm.save()
# -
dm.refresh()
dm.df().loc[(slice(dataset_collection_name,dataset_collection_name), slice(None))]
# ## Dataset transformation walk-through
def print_obj_attr(obj, name="Object"):
print(name)
tmp = vars(obj)
for key in tmp:
print(key, tmp[key])
print("")
records = load_dataset_names()
# ### Load and parse dataset
# +
# dataset
record = wfdb.rdrecord(os.path.join(source_folder, records[51]))
#print_obj_attr(record, "Record object")
df_record = pd.DataFrame(record.p_signal, columns=record.sig_name)
df_record
# -
# Add timestamp information based on sample interval ($$[fs] = samples/second$$):
display(Latex(f"Samples per second: $$fs = {record.fs} \\frac{{1}}{{s}}$$"))
display(Markdown(f"This gives a sample interval of {1e+9/record.fs} nanoseconds"))
df_record["timestamp"] = pd.to_datetime(df_record.index.values * 1e+9/record.fs, unit='ns')
df_record
# +
# find all annotations
records = load_dataset_names()
annotations = {}
for r in records:
atr = wfdb.rdann(os.path.join(source_folder, r), "atr")
df_annotation = pd.DataFrame(atr.symbol, index=atr.sample, columns=["Label"])
for an in df_annotation["Label"].unique():
if an not in annotations:
annotations[an] = set()
annotations[an].add(atr.record_name)
for an in annotations:
annotations[an] = ", ".join(annotations[an])
annotations
# -
# Annotations
#
# | Annotation | Description |
# | :--------- | :---------- |
# || **Considered normal** |
# | `N` | Normal beat |
# || **Anomalous beats** (use double-window labeling) |
# | `F` | Fusion of ventricular and normal beat |
# | `S` | Supraventricular premature or ectopic beat |
# | `a` | Aberrated atrial premature beat |
# | `V` | Premature ventricular contraction |
# | `J` | Nodal (junctional) premature beat |
# | `B` | Bundle branch block beat (unspecified) |
# || **External anomalies** (single window labeling) |
# | `Q` | Unclassifiable beat |
# | `\|` | Isolated QRS-like artifact |
# || **Ignored, bc hard to parse and to label** |
# | `+` | Rythm change |
# | `~` | Change in signal quality (usually noise level changes) |
# ### Load and parse annotation
# +
atr = wfdb.rdann(os.path.join(source_folder, records[51]), "atr")
#print_obj_attr(atr, "Annotation object")
assert record.fs == atr.fs, "Sample frequency of records and annotations does not match!"
df_annotation = pd.DataFrame(atr.symbol, index=atr.sample, columns=["Label"])
df_annotation = df_annotation.reset_index()
df_annotation.columns = ["position", "label"]
df_annotation.groupby("label").count()
# -
# ### Calculate beat window
#
# We assume that the normal beats (annotated with `N`) occur in a regular interval and that the expert annotations (from the dataset) are directly in the middle of a beat window.
# A beat window is a fixed length subsequence of the time series and shows a heart beat in its direct (local) context.
#
# We calculate the beat window length for each dataset based on the median distance between normal beats (`N`).
# The index (autoincrementing integers) serves as the measurement unit.
#
# Create DataFrame containing all annotated beats:
df_beat = df_annotation[["position", "label"]]
df_beat
# Shifted-by-one self-join and filter out all beat-pairs that contain anomalous beats.
# We want to calculate the beat windows only based on the normal beats.
# We then calculate the distance between two neighboring heart beats:
df_normal_beat = df_beat.copy()
df_normal_beat["prev_position"] = df_beat["position"].shift()
df_normal_beat["prev_label"] = df_beat["label"].shift()
df_normal_beat = df_normal_beat[(df_normal_beat["label"] == "N") & (df_normal_beat["prev_label"] == "N")]
df_normal_beat = df_normal_beat.drop(columns=["label", "prev_label"])
df_normal_beat["length"] = df_normal_beat["position"] - df_normal_beat["prev_position"]
df_normal_beat.describe()
# The median of all normal beat lengths is the beat window size.
# We require the beat window size to be odd.
# This allows us to center the window at the beat annotation.
normal_beat_length = df_normal_beat["length"].median()
if (normal_beat_length%2) == 0:
normal_beat_length += 1
beat_window_size = int(normal_beat_length)
beat_window_margin = (beat_window_size - 1)//2
print(f"window size = {beat_window_size}\nwindow margins (left and right) = {beat_window_margin}")
# ### Calculate anomalous windows
#
# The experts from PhysioNet annotated only the beats itself with a label, but the actual anomaly is also comprised of the beat surroundings.
#
# We assume that anomalous beats (such as `V` or `F`; see table above) require looking at a window around the actual beat as being anomalous.
# External anomalies (such as `|`; see table above) also mark a window around it as anomalous, because those artefacts comprise multiple points.
#
# We completely ignore `~` and `+`-annotations that indicate signal quality or rythm changes, because they are not relevant for our analysis.
#
# We automatically label a variable-sized window around an annotated beat as an anomalous subsequence using the following technique:
#
# 1. For anomalous annotations (`S`, `V`, `a`, `J`, `B`, and `F` annotations):
# - Remove `~`, `+`, and `|` annotations
# - Calculate anomaly window using `beat_window_size` aligned with its center on the beat annotation.
# - Calculate end of previous beat window _e_ and beginning of next beat window _b_.
# Use _e_ as beginning and _b_ as end for a second anomaly window.
# - Mark the union of both anomaly windows' points as anomalous.
# 2. For `|` and `Q` annotations, mark all points of an anomaly window centered on the annotation as anomalous.
# 3. Mark all other points as normal.
#
# > **Explain, why we used the combined windows for anomalous beats!!**
# >
# > - pattern/shape of signal may be ok
# > - but we consider distance to other beats also
# > - if too narrow or too far away, it's also anomalous
#
# The figure shows an anomalous beat with its anomaly window (in red) and the windows of its previous and subsequent normal beats (in green).
# We mark all points in the interval $$[min(W_{end}, X_{start}), max(X_{end}, Y_{start})]$$
# reverse lookup from timestamp to annotation index in df_beat
p = df_record[df_record["timestamp"] == "1970-01-01 00:11:03.000"].index.values[0]
df_beat[df_beat["position"] >= p].index[0]
# +
def plot_window(pos, color="blue", **kvs):
start = pos - beat_window_margin
end = pos + beat_window_margin
plt.axvspan(start, end, color=color, alpha=0.5, **kvs)
index = 798
beat_n = df_beat.loc[index, "position"]
print("Selected beat is annotated as", df_beat.loc[index, "label"])
print("with timestamp", df_record.loc[beat_n, "timestamp"])
ax = df_record.iloc[beat_n-500:beat_n+500].plot(kind='line', y=['ECG1', 'ECG2'], use_index=True, figsize=(20,10))
plot_window(df_beat.loc[index-1, "position"], label="$W$")
plot_window(beat_n, color="orange", label="$X$")
plot_window(df_beat.loc[index+1, "position"], label="$Y$")
plt.legend()
plt.show()
# -
# #### Windows for external anomalies
df_pipe = df_beat.copy()
df_pipe = df_pipe[(df_pipe["label"] == "|") | (df_pipe["label"] == "Q")]
df_pipe["window_start"] = df_pipe["position"]-beat_window_margin
df_pipe["window_end"] = df_pipe["position"]+beat_window_margin
df_pipe = df_pipe[["position", "window_start", "window_end"]]
df_pipe.head()
# #### Windows for anomalous beats
df_tmp = df_beat.copy()
df_tmp = df_tmp[(df_tmp["label"] != "|") & (df_tmp["label"] != "~") & (df_tmp["label"] != "+")]
df_tmp["position_next"] = df_tmp["position"].shift(-1)
df_tmp["position_prev"] = df_tmp["position"].shift(1)
#df_tmp = df_tmp[(df_tmp["position_prev"].notnull()) & (df_tmp["position_next"].notnull())]
df_tmp = df_tmp[(df_tmp["label"] != "Q") & (df_tmp["label"] != "N")]
df_tmp["window_start"] = np.minimum(df_tmp["position"].values-beat_window_margin, df_tmp["position_prev"].values+beat_window_margin)
df_tmp["window_end"] = np.maximum(df_tmp["position"].values+beat_window_margin, df_tmp["position_next"].values-beat_window_margin)
df_svf = df_tmp[["position", "window_start", "window_end"]]
df_tmp.groupby("label").count()
# #### Merge everything together
df_windows = pd.concat([df_pipe, df_svf])
df_windows.head()
# +
index = 798
beat = df_windows.loc[index, "position"]
start = df_windows.loc[index, "window_start"]
end = df_windows.loc[index, "window_end"]
print("Selected beat is annotated as", df_beat.loc[index, "label"])
print("with timestamp", df_record.loc[beat, "timestamp"])
ax = df_record.iloc[beat-500:beat+500].plot(kind='line', y=['ECG1', 'ECG2'], use_index=True, figsize=(20,10))
plt.axvspan(beat-500, start-1, color="green", alpha=0.5, label="normal region 1", ymin=.5)
plt.axvspan(start, end, color="red", alpha=0.5, label="anomalous region", ymin=.5)
plt.axvspan(end+1, beat+500, color="green", alpha=0.5, label="normal region 2", ymin=.5)
plot_window(df_beat.loc[index-1, "position"], label="$W$", ymax=.5)
plot_window(beat_n, color="orange", label="$X$", ymax=.5)
plot_window(df_beat.loc[index+1, "position"], label="$Y$", ymax=.5)
plt.legend()
plt.show()
# -
# ### Add labels
# +
df = df_record.copy()
df["is_anomaly"] = 0
for _, (_, t1, t2) in df_windows.iterrows():
tmp = df[df.index >= t1]
tmp = tmp[tmp.index <= t2]
df["is_anomaly"].values[tmp.index] = 1
#df = df.set_index("timestamp")
df[df["is_anomaly"] == 1]
# -
df_beat[(df_beat["label"] == "|")]
# +
start = 21700
end = 22500
df_show = df.loc[start:end]
df_show.plot(kind='line', y=['ECG1', 'ECG2', 'is_anomaly'], use_index=True, figsize=(20,10))
labels = df_beat[(df_beat["position"] > start) & (df_beat["position"] < end)]
for i, (position, label) in labels.iterrows():
plt.text(position, -2.5, label)
plt.show()
# -
# ## Experimentation
df = pd.merge(df_record, df_annotation, left_index=True, right_index=True, how="outer")
#df = df.fillna(value={"Label": ".", "is_anomaly": 0})
df.groupby(["is_anomaly"]).count()
df[df["Label"].notna()]
import matplotlib.pyplot as plt
df_show = df.loc[27000:28000]
df_show.plot(kind='line', y=['ECG1', 'ECG2', 'is_anomaly'], use_index=True, figsize=(20,10))
plt.show()
df = pd.read_csv(os.path.join(dataset_subfolder, "800.test.csv"), index_col="timestamp")
df.loc["1970-01-01 00:21:20":"1970-01-01 00:21:40"].plot(figsize=(20,10))
plt.show()
| notebooks/data-prep/MIT-BIH Supraventricular Arrhythmia DB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Taller evaluable sobre la extracción, transformación y visualización de datos usando IPython
# **<NAME>**
# <EMAIL>
# Universidad Nacional de Colombia, Sede Medellín
# Facultad de Minas
# Medellín, Colombia
# # Instrucciones
# En la carpeta 'Taller' del repositorio 'ETVL-IPython' se encuentran los archivos 'Precio_Bolsa_Nacional_($kwh)_'*'.xls' en formato de Microsoft Excel, los cuales contienen los precios históricos horarios de la electricidad para el mercado eléctrico Colombiano entre los años 1995 y 2017 en COL-PESOS/kWh. A partir de la información suministrada resuelva los siguientes puntos usando el lenguaje de programación Python.
# # Preguntas
#
# **1.--** Lea los archivos y cree una tabla única concatenando la información para cada uno de los años. Imprima el encabezamiento de la tabla usando `head()`.
import numpy as np
import pandas as pd
# +
#lee el archivo de excel saltando las tres primeras filas
frames=[]
for n in range (1995, 2018):
nombrearchivo='Precio_Bolsa_Nacional_($kwh)_' + str(n)
if n < 2000:
skip=3
else:
skip=2
if n >= 2016:
nombrearchivo += '.xls'
else:
nombrearchivo += '.xlsx'
x=pd.read_excel(nombrearchivo, skiprows = skip, parse_cols=24)
frames.append(x)
datos = pd.concat(frames)
# cambia filas por columnas
w = pd.melt(datos,
id_vars = ['Fecha'],
var_name = 'Hora',
value_name = 'Valores')
print(w[1:10])
# -
# **2.--** Compute e imprima el número de registros con datos faltantes.
len(w) - len(w.dropna())
faltantes=w[w.isnull().T.any().T] # muestra los datos faltantes
faltantes.head()
# **3.--** Compute e imprima el número de registros duplicados.
len(w)-len(w.drop_duplicates())
# **4.--** Elimine los registros con datos duplicados o datos faltantes, e imprima la cantidad de registros que quedan (registros completos).
frameDepurada=w.drop_duplicates().dropna()
len(frameDepurada)
frameDepurada.head()
# **5.--** Compute y grafique el precio promedio diario.
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
frameDepurada.groupby('Fecha').mean().plot(figsize=(15, 10), title='Precio promedio diario')
# **6.--** Compute y grafique el precio máximo por mes.
# +
#crea una columna que diga el mes
x=[]
for i in range(len(frameDepurada)):
x.append(str(frameDepurada.iat[i,0])[0:7])
frameDepurada['key'] = x
print(frameDepurada.head())
# %matplotlib inline
frameDepurada.groupby('key').max().plot(title='Precio máximo mensual')
# -
# **7.--** Compute y grafique el precio mínimo mensual.
# %matplotlib inline
frameDepurada.groupby('key').min().plot(title='Precio mínimo mensual')
# **8.--** Haga un gráfico para comparar el precio máximo del mes (para cada mes) y el precio promedio mensual.
# +
# %matplotlib inline
a = frameDepurada.groupby('key').mean().plot(figsize=(10, 10))
frameDepurada.groupby('key').max().plot(ax=a, title='Comparativo precio máximo del mes\n y precio promedio mensual')
#leyenda del gráfico
a.legend(['Promedio','Máximo'])
# -
# **9.--** Haga un histograma que muestre a que horas se produce el máximo precio diario para los días laborales.
# +
import calendar as cl
import datetime as dt
x=[]
for i in range(len(frameDepurada)):
dia = cl.day_name[dt.datetime.strptime(str(frameDepurada.iat[i,0])[0:10], '%Y-%m-%d').date().weekday()]
x.append(dia)
frameDepurada['dia']=x
frameDepurada.head()
# -
# filtra días laborales
dias_semana = frameDepurada.loc[frameDepurada['dia'].isin(['Monday','Tuesday', 'Wednesday', 'Thursday', 'Friday'])]
dias_semana.head()
# +
import matplotlib.pyplot as plt
hora_maxPrecio_dia = dias_semana.groupby(['Fecha'])['Valores'].max()
hora_maxPrecio_dia = hora_maxPrecio_dia.reset_index()
hora_maxPrecio_dia['Fecha'] = hora_maxPrecio_dia['Fecha'].astype(str)
graficar = pd.merge(frameDepurada,hora_maxPrecio_dia)
plt.hist(graficar['Hora'].astype(int))
# -
# **10.--** Haga un histograma que muestre a que horas se produce el máximo precio diario para los días sabado.
# +
# filtra días sabado
dia_sabado = frameDepurada.loc[frameDepurada['dia'].isin(['Saturday'])]
hora_maxPrecio_sabado = dia_sabado.groupby(['Fecha'])['Valores'].max()
hora_maxPrecio_sabado = hora_maxPrecio_sabado.reset_index()
hora_maxPrecio_sabado['Fecha'] = hora_maxPrecio_sabado['Fecha'].astype(str)
graficar_2 = pd.merge(frameDepurada,hora_maxPrecio_sabado)
plt.hist(graficar_2['Hora'].astype(int))
# -
# **11.--** Haga un histograma que muestre a que horas se produce el máximo precio diario para los días domingo.
# +
# filtra días domingo
dia_domingo = frameDepurada.loc[frameDepurada['dia'].isin(['Sunday'])]
# trae el precio max para cada fecha
hora_maxPrecio_domingo = dia_domingo.groupby(['Fecha'])['Valores'].max()
hora_maxPrecio_domingo = hora_maxPrecio_domingo.reset_index()
hora_maxPrecio_domingo['Fecha'] = hora_maxPrecio_domingo['Fecha'].astype(str)
graficar_3 = pd.merge(frameDepurada,hora_maxPrecio_domingo)
plt.hist(graficar_3['Hora'].astype(int))
# -
# **12.--** Imprima una tabla con la fecha y el valor más bajo por año del precio de bolsa.
# +
# crea una columna que diga el año
x=[]
for i in range(len(frameDepurada)):
year=str(frameDepurada.iat[i,0])[0:4]
x.append(year)
frameDepurada['año']=x
# trae el precio min para cada año
minPrecio_año = frameDepurada.groupby(['año'])['Valores'].min()
minPrecio_año = minPrecio_año.reset_index()
print(minPrecio_año)
minPrecio_año['año'] = minPrecio_año['año'].astype(str)
graficar_4 = pd.merge(frameDepurada,minPrecio_año)
c=graficar_4[['año','Fecha','Valores']] # fecha exacta
c.drop_duplicates()
# -
# **13.--** Haga una gráfica en que se muestre el precio promedio diario y el precio promedio mensual.
# +
plt.title('Precio promedio diario')
precio_prom_día = frameDepurada.groupby(['Fecha'])['Valores'].mean().plot()
plt.show()
plt.title('Precio promedio mensual')
precio_prom_mes = frameDepurada.groupby(['key'])['Valores'].mean().plot()
plt.show()
# +
plt.title('Precio promedio diario y precio promedio mensual')
precio_prom_día = frameDepurada.groupby(['Fecha'])['Valores'].mean().plot()
plt.hold(True)
precio_prom_mes = frameDepurada.groupby(['key'])['Valores'].mean().plot()
plt.show()
# -
# ---
| Taller.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ask Question
# Can we predict the maximum temperature tomorrow for Seattle, Washington given one year of historical data?
# The data we have available is the maximum temperatures in Seattle from the National Oceanic and Atmospheric Administration's (NOAA) [Climate Data Online Tool](https://www.ncdc.noaa.gov/cdo-web/).
# # Data Acqusition
# + hideCode=false hidePrompt=false jupyter={"outputs_hidden": false}
# Pandas is used for data manipulation
import pandas as pd
from matplotlib import pyplot as plt
import joblib
# Read in data as pandas dataframe and display first 5 rows
features_train = pd.read_csv('data/train.csv')
features_val = pd.read_csv('data/val.csv')
#features = pd.concat([data_train, data_val])
from sklearn.preprocessing import LabelEncoder
# -
# # Identify Anomalies
# + hideCode=false hidePrompt=false jupyter={"outputs_hidden": false}
print('The shape of our features is:', features_train.shape)
print('The shape of our features is:', features_val.shape)
# + jupyter={"outputs_hidden": false}
# Descriptive statistics for each column
features_train.describe()
# -
""""
# Use datetime for dealing with dates
import datetime
# Get years, months, and days
years = features['ANIO']
months = features['MES']
days = features['DIA']
# List and then convert to datetime object
dates = [str(int(year)) + '-' + str(int(month)) + '-' + str(int(day)) for year, month, day in zip(years, months, days)]
dates = [datetime.datetime.strptime(date, '%Y-%m-%d') for date in dates]
"""
"""
# Import matplotlib for plotting and use magic command for Jupyter Notebooks
import matplotlib.pyplot as plt
%matplotlib inline
# Set the style
plt.style.use('fivethirtyeight')
"""
# + jupyter={"outputs_hidden": false}
"""
# Set up the plotting layout
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, figsize = (10,10))
fig.autofmt_xdate(rotation = 45)
# Actual max temperature measurement
ax1.plot(dates, features['actual'])
ax1.set_xlabel(''); ax1.set_ylabel('Temperature'); ax1.set_title('Max Temp')
# Temperature from 1 day ago
ax2.plot(dates, features['temp_1'])
ax2.set_xlabel(''); ax2.set_ylabel('Temperature'); ax2.set_title('Previous Max Temp')
# Temperature from 2 days ago
ax3.plot(dates, features['temp_2'])
ax3.set_xlabel('Date'); ax3.set_ylabel('Temperature'); ax3.set_title('Two Days Prior Max Temp')
# Friend Estimate
ax4.plot(dates, features['friend'])
ax4.set_xlabel('Date'); ax4.set_ylabel('Temperature'); ax4.set_title('Friend Estimate')
plt.tight_layout(pad=2)
"""
# -
# # Data Preparation
# ### One-Hot Encoding
# One hot encoding takes this:
#
# | week |
# |------|
# | Mon |
# | Tue |
# | Wed |
# | Thu |
# | Fri |
# | Sat |
# | Sun |
#
# and converts it into:
#
# | Mon | Tue | Wed | Thu | Fri | Sat | Sun |
# |-----|-----|-----|-----|-----|-----|-----|
# | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
# | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
# | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
# | 0 | 0 | 0 | 1 | 0 | 0 | 0 |
# | 0 | 0 | 0 | 0 | 1 | 0 | 0 |
# | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
# | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
# + hideCode=false hidePrompt=false jupyter={"outputs_hidden": false}
# One-hot encode categorical features
"""
le = LabelEncoder()
le.fit(features['CLASE'].unique())
features['CLASE']=le.transform(features['BARRIO'])
"""
features_train = pd.get_dummies(features_train)
features_train=features_train.groupby(['ANIO','CLUSTER','SEMANA']).agg({'CLASE_atropello':'sum','CLASE_caida ocupante':'sum','CLASE_choque':'sum','CLASE_otro':'sum','CLASE_volcamiento':'sum'})
nombre_columnasTipo=['atropello', 'caidaocupante','choque','otro','volcamiento']
features_train.columns = nombre_columnasTipo
features_train = features_train.reset_index()
features_train.loc[features_train['CLUSTER'] == 0, 'CLUSTER'] = 'cluster0'
features_train.loc[features_train['CLUSTER'] == 1, 'CLUSTER'] = 'cluster1'
features_train.loc[features_train['CLUSTER'] == 2, 'CLUSTER'] = 'cluster2'
features_train = pd.get_dummies(features_train)
features_val = pd.get_dummies(features_val)
features_val=features_val.groupby(['ANIO','CLUSTER','SEMANA']).agg({'CLASE_atropello':'sum','CLASE_caida ocupante':'sum','CLASE_choque':'sum','CLASE_otro':'sum','CLASE_volcamiento':'sum'})
nombre_columnasTipo=['atropello', 'caidaocupante','choque','otro','volcamiento']
features_val.columns = nombre_columnasTipo
features_val = features_val.reset_index()
features_val.loc[features_val['CLUSTER'] == 0, 'CLUSTER'] = 'cluster0'
features_val.loc[features_val['CLUSTER'] == 1, 'CLUSTER'] = 'cluster1'
features_val.loc[features_val['CLUSTER'] == 2, 'CLUSTER'] = 'cluster2'
features_val = pd.get_dummies(features_val)
# -
features_val
# + jupyter={"outputs_hidden": false}
print('Shape of features after one-hot encoding:', features_val.shape)
print('Shape of train after one-hot encoding:', features_train.shape)
# -
# ### Features and Labels and Convert Data to Arrays
# +
# Use numpy to convert to arrays
import numpy as np
# Labels are the values we want to predict
label_train = np.array(features_train[['atropello', 'caidaocupante','choque','otro','volcamiento']])
label_val = np.array(features_val[['atropello', 'caidaocupante','choque','otro','volcamiento']])
# Remove the labels from the features
# axis 1 refers to the columns
features_train= features_train.drop(['atropello', 'caidaocupante','choque','otro','volcamiento'], axis = 1)
features_val= features_val.drop(['atropello', 'caidaocupante','choque','otro','volcamiento'], axis = 1)
feature_list = list(features_train.columns)
features_train= np.array(features_train)
features_val= np.array(features_val)
# -
# ### Training and Testing Sets
# + jupyter={"outputs_hidden": false}
print('Training Features Shape:', features_train.shape)
print('Training Labels Shape:', label_train.shape)
print('Testing Features Shape:', features_val.shape)
print('Testing Labels Shape:', label_val.shape)
# -
# # Establish Baseline
# + jupyter={"outputs_hidden": false}
"""
# The baseline predictions are the historical averages
baseline_preds = test_features[:, feature_list.index('average')]
# Baseline errors, and display average baseline error
baseline_errors = abs(baseline_preds - test_labels)
print('Average baseline error: ', round(np.mean(baseline_errors), 2), 'degrees.')
"""
# -
# # Train Model
# + hideCode=false hidePrompt=false
# Import the model we are using
from sklearn.ensemble import RandomForestRegressor
# Instantiate model
rf = RandomForestRegressor(n_estimators= 1000, min_samples_leaf=50, random_state=42)
# Train the model on training data
rf.fit(features_train,label_train);
# -
# # Make Predictions on Test Data
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import mean_squared_error as rmse
# Use the forest's predict method on the test data
prediccion_validacion = rf.predict(features_val)
prediccion_entrenamiento=rf.predict(features_train)
# Calculate the absolute errors
error=rmse(label_val, prediccion_validacion)
error2=rmse(label_train, prediccion_entrenamiento)
# Print out the mean absolute error (mae)
print('error validacion:', error,"erro prueva",error2)
#plt.plot(prediccion_validacion[80])
#plt.plot(test_labels[80])
plt.plot(prediccion_entrenamiento[0:,2])
plt.plot(label_train[0::,2])
# +
# Get numerical feature importances
importances = list(rf.feature_importances_)
# List of tuples with variable and importance
feature_importances = [(feature, round(importance, 2)) for feature, importance in zip(feature_list, importances)]
# Sort the feature importances by most important first
feature_importances = sorted(feature_importances, key = lambda x: x[1], reverse = True)
joblib.dump(rf, 'models/arbol_semana.pkl')
# Print out the feature and importances
[print('Variable: {:20} Importance: {}'.format(*pair)) for pair in feature_importances];
| directorio de trabajo/Jorge/no puedo creer que esto funcione/.ipynb_checkpoints/arbol_semana-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# <h1 align="center">Segmentation: Thresholding and Edge Detection</h1>
#
# In this notebook our goal is to estimate the radius of spherical markers from an image (Cone-Beam CT volume).
#
# We will use two approaches:
# 1. Segment the fiducial using a thresholding approach, derive the sphere's radius from the segmentation. This approach is solely based on SimpleITK.
# 2. Localize the fiducial's edges using the Canny edge detector and then fit a sphere to these edges using a least squares approach. This approach is a combination of SimpleITK and R.
#
# It should be noted that all of the operations, filtering and computations, are natively in 3D. This is the "magic" of ITK and SimpleITK at work.
# +
library(SimpleITK)
source("downloaddata.R")
# -
# Load the volume and look at the image (visualization requires window-leveling).
# + simpleitk_error_allowed="Exception in SITK"
spherical_fiducials_image <- ReadImage(fetch_data("spherical_fiducials.mha"))
Show(spherical_fiducials_image, "spheres")
# -
# After looking at the image you should have identified two spheres. Now select a Region Of Interest (ROI) around the sphere which you want to analyze.
# + simpleitk_error_allowed="Exception in SITK"
roi1 = list(c(280,320), c(65,90), c(8, 30))
roi2 = list(c(200,240), c(65,100), c(15, 40))
mask_value = 255
# Select the roi
roi = roi1
# Update the R roi, SimpleITK indexes are zero based, R indexes start at one
r_roi = lapply(roi, function(x) x+1)
# Create the mask image from an R array
amask <- array(0, spherical_fiducials_image$GetSize())
xs <- r_roi[[1]][1]:r_roi[[1]][2]
ys <- r_roi[[2]][1]:r_roi[[2]][2]
zs <- r_roi[[3]][1]:r_roi[[3]][2]
amask[xs, ys, zs] <- mask_value
mask <- Cast(as.image(amask), "sitkUInt8")
mask$CopyInformation(spherical_fiducials_image)
Show(LabelOverlay(Cast(IntensityWindowing(spherical_fiducials_image, windowMinimum=-32767,
windowMaximum=-29611),
"sitkUInt8"),
mask, opacity=0.5))
# -
# ## Thresholding based approach
#
# Our region of interest is expected to have a bimodal intensity distribution with high intensities belonging to the spherical marker and low ones to the background. We can thus use Otsu's method for threshold selection to segment the sphere and estimate its radius.
# +
# Set pixels that are in [min_intensity,otsu_threshold] to inside_value, values above otsu_threshold are
# set to outside_value. The sphere's have higher intensity values than the background, so they are outside.
inside_value <- 0
outside_value <- 255
number_of_histogram_bins <- 100
mask_output <- TRUE
labeled_result <- OtsuThreshold(spherical_fiducials_image, mask, inside_value, outside_value,
number_of_histogram_bins, mask_output, mask_value)
# Estimate the sphere radius from the segmented image using the LabelShapeStatisticsImageFilter.
label_shape_analysis <- LabelShapeStatisticsImageFilter()
label_shape_analysis$SetBackgroundValue(inside_value)
dummy <- label_shape_analysis$Execute(labeled_result)
cat("The sphere's radius is: ",label_shape_analysis$GetEquivalentSphericalRadius(outside_value),"mm")
# -
# ## Edge detection based approach
#
# In this approach we will localize the sphere's edges in 3D using SimpleITK. We then compute the least squares sphere that optimally fits the 3D points using R. The mathematical formulation for this solution is described in this [Insight Journal paper](http://www.insight-journal.org/download/viewpdf/769/1/download). We also look at a weighted version of least squares fitting using R's linear model fitting approach.
#
# +
# Create a cropped version of the original image.
sub_image = spherical_fiducials_image[r_roi[[1]][1]:r_roi[[1]][2],
r_roi[[2]][1]:r_roi[[2]][2],
r_roi[[3]][1]:r_roi[[3]][2]]
# Edge detection on the sub_image with appropriate thresholds and smoothing.
edges <- CannyEdgeDetection(Cast(sub_image, "sitkFloat32"),
lowerThreshold=0.0,
upperThreshold=200.0,
variance = c(5.0, 5.0, 5.0))
# Get the 3D location of the edge points
edge_indexes <- which(as.array(edges)==1.0, arr.ind=TRUE)
# Always remember to modify indexes when shifting between native R operations and SimpleITK operations
physical_points <- t(apply(edge_indexes - 1, MARGIN=1,
sub_image$TransformIndexToPhysicalPoint))
# -
# Visually inspect the results of edge detection, just to make sure. Note that because SimpleITK is working in the
# physical world (not pixels, but mm) we can easily transfer the edges localized in the cropped image to the original.
# + simpleitk_error_allowed="Exception in SITK"
edge_label <- Image(spherical_fiducials_image$GetSize(), "sitkUInt8")
edge_label$CopyInformation(spherical_fiducials_image)
e_label <- 255
apply(physical_points,
MARGIN=1,
function(x, img, label) img$SetPixel(img$TransformPhysicalPointToIndex(x),label),
img=edge_label,
label=e_label)
Show(LabelOverlay(Cast(IntensityWindowing(spherical_fiducials_image, windowMinimum=-32767, windowMaximum=-29611),
"sitkUInt8"),
edge_label, opacity=0.5))
# -
# Setup and solve linear equation system.
A <- -2 * physical_points
A <- cbind(A, 1)
b <- -rowSums(physical_points^2)
x <- solve(qr(A, LAPACK=TRUE), b)
cat("The sphere's center is: ", x, "\n")
cat("The sphere's radius is: ", sqrt(x[1:3] %*% x[1:3] - x[4]), "\n")
# Now, solve using R's linear model fitting. We also weigh the edge points based on the gradient magnitude.
# +
gradient_magnitude = GradientMagnitude(sub_image)
grad_weights = apply(edge_indexes-1, MARGIN=1, gradient_magnitude$GetPixel)
df <- data.frame(Y=rowSums(physical_points^2), x=physical_points[, 1],
y=physical_points[, 2], z=physical_points[, 3])
fit <- lm(Y ~ x + y + z, data=df, weights=grad_weights)
center <- coefficients(fit)[c("x", "y", "z")] / 2
radius <- sqrt(coefficients(fit)["(Intercept)"] + sum(center^2))
cat("The sphere's center is: ", center, "\n")
cat("The sphere's radius is: ", radius, "\n")
# -
# ## You've made it to the end of the notebook, so what is the sphere's radius?
#
# The radius is 3mm.
| R/33_Segmentation_Thresholding_Edge_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Functions and Methods Homework
#
# Complete the following questions:
# ____
# **Write a function that computes the volume of a sphere given its radius.**
# <p>The volume of a sphere is given as $$\frac{4}{3} πr^3$$</p>
def vol(rad):
volume=(4/3) * 3.14 * (rad**3)
return(volume)
# Check
vol(2)
# ___
# **Write a function that checks whether a number is in a given range (inclusive of high and low)**
def ran_check(num,low,high):
if num>=low and num<=high:
print(f'{num} is in the range between {low} and {high}')
# Check
ran_check(4,2,7)
# If you only wanted to return a boolean:
def ran_bool(num,low,high):
if num>=low and num<=high:
return('True')
ran_bool(3,1,10)
# ____
# **Write a Python function that accepts a string and calculates the number of upper case letters and lower case letters.**
#
# Sample String : 'Hello Mr. Rogers, how are you this fine Tuesday?'
# Expected Output :
# No. of Upper case characters : 4
# No. of Lower case Characters : 33
#
# HINT: Two string methods that might prove useful: **.isupper()** and **.islower()**
#
# If you feel ambitious, explore the Collections module to solve this problem!
#""**""ef up_low(s):
# for i range(a,z):
# print(i) ""
def up_low(s):
UPPER_CASE=0
lower_case=0
for c in s:
if c.isupper():
UPPER_CASE+=1
elif c.islower():
lower_case+=1
else:
continue
print ("No.of Upper case characters: ", UPPER_CASE)
print ("No. of lower case Characters: ", lower_case)
s = 'Hello Mr. Rogers, how are you this fine Tuesday?'
up_low(s)
# ____
# **Write a Python function that takes a list and returns a new list with unique elements of the first list.**
#
# Sample List : [1,1,1,1,2,2,3,3,3,3,4,5]
# Unique List : [1, 2, 3, 4, 5]
state=[]
def unique_list(lst):
for i in lst:
if i not in state:
state.append(i)
return state
unique_list([1,1,1,1,2,2,3,3,3,3,4,5])
# ____
# **Write a Python function to multiply all the numbers in a list.**
#
# Sample List : [1, 2, 3, -4]
# Expected Output : -24
def multiply(numbers):
product=1
for i in numbers:
product*=i
return(product)
multiply([1,2,3,-4])
# ____
# **Write a Python function that checks whether a passed in string is palindrome or not.**
#
# Note: A palindrome is word, phrase, or sequence that reads the same backward as forward, e.g., madam or nurses run.
def palindrome(s):
x=[]
x=s[::-1]
for i in range(0,len(x)):
if x[i]==s[i]:
return True
palindrome('helleh')
# ____
# #### Hard:
#
# **Write a Python function to check whether a string is pangram or not.**
#
# Note : Pangrams are words or sentences containing every letter of the alphabet at least once.
# For example : "The quick brown fox jumps over the lazy dog"
#
# Hint: Look at the string module
#import string
#def ispangram(str1, alphabet=string.ascii_lowercase):
# y=0
# alphabet1=alphabet.split()
#for i in range(0,len(str1)):
import string
def ispangram(str1, alphabet=string.ascii_lowercase):
x=set(alphabet)
y=set(str1.lower())
print(len(x))
print(len(y))
if y>=x:
return True
ispangram("The quick brown fox jumps over the lazy dog")
alphabet=string.ascii_lowercase
print(alphabet)
# #### Great Job!
| 05-Functions and Methods Homework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import scipy as sp
import openslide
import large_image
import histomicstk as htk
# Import and alias positive_pixel_count
import histomicstk.segmentation.positive_pixel_count as ppc
import skimage.io
import skimage.measure
import skimage.color
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
# %matplotlib inline
#Some nice default configuration for plots
plt.rcParams['figure.figsize'] = 15, 15
plt.rcParams['image.cmap'] = 'gray'
titlesize = 24
# -
# # Load input image
# +
inputImageFile = ('/media/jonny_admin/540GB/Research/TCGA_LUAD-WSI/TCGA-44-7669-01A-01-BS1.554eb3d8-6ac6-4a72-a761-ee67021ce97f.svs')
# slide = openslide.OpenSlide(inputImageFile)
# -
ts = large_image.getTileSource(inputImageFile)
ts.getMetadata()
kwargs = dict(format=large_image.tilesource.TILE_FORMAT_NUMPY)
total_tiles = ts.getSingleTile(**kwargs)['iterator_range']['position']
total_tiles
ts.getSingleTile(tile_position=10000, **kwargs)['tile'].shape[0:2]
# np.average(ts.getSingleTile(tile_position=10000, **kwargs)['tile'], axis=0)
plt.imshow(ts.getSingleTile(tile_position=10000, **kwargs)['tile'])
# +
large_region = dict(
left=0, top=0,
width=28001, height=14652,
)
template_params = ppc.Parameters(
hue_value=0.05,
hue_width=0.15,
saturation_minimum=0.05,
intensity_upper_limit=0.95,
intensity_weak_threshold=0.65,
intensity_strong_threshold=0.35,
intensity_lower_limit=0.05,
)
stats, = %time ppc.count_slide(inputImageFile, template_params)
print stats
# -
plt.imshow(im_region)
plt.show()
source.getSingleTile()
| notebooks/histomicstk_py27.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Jaydenzk/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module3-introduction-to-bayesian-inference/LS_DS_133_Introduction_to_Bayesian_Inference.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="H7OLbevlbd_Z" colab_type="text"
# # Lambda School Data Science Module 143
#
# ## Introduction to Bayesian Inference
#
# !['Detector! What would the Bayesian statistician say if I asked him whether the--' [roll] 'I AM A NEUTRINO DETECTOR, NOT A LABYRINTH GUARD. SERIOUSLY, DID YOUR BRAIN FALL OUT?' [roll] '... yes.'](https://imgs.xkcd.com/comics/frequentists_vs_bayesians.png)
#
# *[XKCD 1132](https://www.xkcd.com/1132/)*
#
# + [markdown] id="3mz8p08BsN6p" colab_type="text"
# ## Prepare - Bayes' Theorem and the Bayesian mindset
# + [markdown] id="GhycNr-Sbeie" colab_type="text"
# Bayes' theorem possesses a near-mythical quality - a bit of math that somehow magically evaluates a situation. But this mythicalness has more to do with its reputation and advanced applications than the actual core of it - deriving it is actually remarkably straightforward.
#
# ### The Law of Total Probability
#
# By definition, the total probability of all outcomes (events) if some variable (event space) $A$ is 1. That is:
#
# $$P(A) = \sum_n P(A_n) = 1$$
#
# The law of total probability takes this further, considering two variables ($A$ and $B$) and relating their marginal probabilities (their likelihoods considered independently, without reference to one another) and their conditional probabilities (their likelihoods considered jointly). A marginal probability is simply notated as e.g. $P(A)$, while a conditional probability is notated $P(A|B)$, which reads "probability of $A$ *given* $B$".
#
# The law of total probability states:
#
# $$P(A) = \sum_n P(A | B_n) P(B_n)$$
#
# In words - the total probability of $A$ is equal to the sum of the conditional probability of $A$ on any given event $B_n$ times the probability of that event $B_n$, and summed over all possible events in $B$.
#
# ### The Law of Conditional Probability
#
# What's the probability of something conditioned on something else? To determine this we have to go back to set theory and think about the intersection of sets:
#
# The formula for actual calculation:
#
# $$P(A|B) = \frac{P(A \cap B)}{P(B)}$$
#
# 
#
# Think of the overall rectangle as the whole probability space, $A$ as the left circle, $B$ as the right circle, and their intersection as the red area. Try to visualize the ratio being described in the above formula, and how it is different from just the $P(A)$ (not conditioned on $B$).
#
# We can see how this relates back to the law of total probability - multiply both sides by $P(B)$ and you get $P(A|B)P(B) = P(A \cap B)$ - replaced back into the law of total probability we get $P(A) = \sum_n P(A \cap B_n)$.
#
# This may not seem like an improvement at first, but try to relate it back to the above picture - if you think of sets as physical objects, we're saying that the total probability of $A$ given $B$ is all the little pieces of it intersected with $B$, added together. The conditional probability is then just that again, but divided by the probability of $B$ itself happening in the first place.
#
# ### Bayes Theorem
#
# Here is is, the seemingly magic tool:
#
# $$P(A|B) = \frac{P(B|A)P(A)}{P(B)}$$
#
# In words - the probability of $A$ conditioned on $B$ is the probability of $B$ conditioned on $A$, times the probability of $A$ and divided by the probability of $B$. These unconditioned probabilities are referred to as "prior beliefs", and the conditioned probabilities as "updated."
#
# Why is this important? Scroll back up to the XKCD example - the Bayesian statistician draws a less absurd conclusion because their prior belief in the likelihood that the sun will go nova is extremely low. So, even when updated based on evidence from a detector that is $35/36 = 0.972$ accurate, the prior belief doesn't shift enough to change their overall opinion.
#
# There's many examples of Bayes' theorem - one less absurd example is to apply to [breathalyzer tests](https://www.bayestheorem.net/breathalyzer-example/). You may think that a breathalyzer test that is 100% accurate for true positives (detecting somebody who is drunk) is pretty good, but what if it also has 8% false positives (indicating somebody is drunk when they're not)? And furthermore, the rate of drunk driving (and thus our prior belief) is 1/1000.
#
# What is the likelihood somebody really is drunk if they test positive? Some may guess it's 92% - the difference between the true positives and the false positives. But we have a prior belief of the background/true rate of drunk driving. Sounds like a job for Bayes' theorem!
#
# $$
# \begin{aligned}
# P(Drunk | Positive) &= \frac{P(Positive | Drunk)P(Drunk)}{P(Positive)} \\
# &= \frac{1 \times 0.001}{0.08} \\
# &= 0.0125
# \end{aligned}
# $$
#
# In other words, the likelihood that somebody is drunk given they tested positive with a breathalyzer in this situation is only 1.25% - probably much lower than you'd guess. This is why, in practice, it's important to have a repeated test to confirm (the probability of two false positives in a row is $0.08 * 0.08 = 0.0064$, much lower), and Bayes' theorem has been relevant in court cases where proper consideration of evidence was important.
# + [markdown] id="htI3DGvDsRJF" colab_type="text"
# ## Live Lecture - Deriving Bayes' Theorem, Calculating Bayesian Confidence
# + [markdown] id="moIJNQ-nbfe_" colab_type="text"
# Notice that $P(A|B)$ appears in the above laws - in Bayesian terms, this is the belief in $A$ updated for the evidence $B$. So all we need to do is solve for this term to derive Bayes' theorem. Let's do it together!
# + id="ke-5EqJI0Tsn" colab_type="code" colab={}
# Activity 2 - Use SciPy to calculate Bayesian confidence intervals
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.bayes_mvs.html#scipy.stats.bayes_mvs
# + [markdown] id="P-DzzRk5bf0z" colab_type="text"
# ## Assignment - Code it up!
#
# Most of the above was pure math - now write Python code to reproduce the results! This is purposefully open ended - you'll have to think about how you should represent probabilities and events. You can and should look things up, and as a stretch goal - refactor your code into helpful reusable functions!
#
# Specific goals/targets:
#
# 1. Write a function `def prob_drunk_given_positive(prob_drunk_prior, prob_positive, prob_positive_drunk)` that reproduces the example from lecture, and use it to calculate and visualize a range of situations
# 2. Explore `scipy.stats.bayes_mvs` - read its documentation, and experiment with it on data you've tested in other ways earlier this week
# 3. Create a visualization comparing the results of a Bayesian approach to a traditional/frequentist approach
# 4. In your own words, summarize the difference between Bayesian and Frequentist statistics
#
# If you're unsure where to start, check out [this blog post of Bayes theorem with Python](https://dataconomy.com/2015/02/introduction-to-bayes-theorem-with-python/) - you could and should create something similar!
#
# Stretch goals:
#
# - Apply a Bayesian technique to a problem you previously worked (in an assignment or project work) on from a frequentist (standard) perspective
# - Check out [PyMC3](https://docs.pymc.io/) (note this goes beyond hypothesis tests into modeling) - read the guides and work through some examples
# - Take PyMC3 further - see if you can build something with it!
# + id="xpVhZyUnbf7o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a396b69d-805a-474f-a8dc-6097abf9ec22"
# TODO - code!
## n=1 means drinking
import numpy as np
drunk = np.random.binomial(n=1, p=.285, size=1000)
print("P(drunk):", np.mean(drunk))
P_of_drunk = np.mean(drunk)
# + id="uFx8pbLXqlPC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ab694730-7dc0-4aa1-b4c0-3bafd4514f41"
B_if_A = 0.80
P_of_A = .285
P_of_drunk = np.mean(drunk)
P_of_positive = .80
P_of_drunk_if_Beer = 0.80
P_of_positive_if_no_Beer = 0.20
P_of_B = (P_of_drunk_if_Beer * P_of_drunk) + (P_of_positive_if_no_Beer * P_of_positive)
# Bayes Rule:
A_if_B = ((B_if_A) * (P_of_A)) / P_of_B
print("Conditional probability of drinking beer: {:.0f}%".format( A_if_B * 100))
# + [markdown] id="uWgWjp3PQ3Sq" colab_type="text"
# ## Resources
# + [markdown] id="QRgHqmYIQ9qn" colab_type="text"
# - [Worked example of Bayes rule calculation](https://en.wikipedia.org/wiki/Bayes'_theorem#Examples) (helpful as it fully breaks out the denominator)
# - [Source code for mvsdist in scipy](https://github.com/scipy/scipy/blob/90534919e139d2a81c24bf08341734ff41a3db12/scipy/stats/morestats.py#L139)
| module3-introduction-to-bayesian-inference/LS_DS_133_Introduction_to_Bayesian_Inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (RUT-JER-DATA-PT-07-2020-U-C)
# language: python
# name: pycharm-f5a426ed
# ---
# + pycharm={"is_executing": false}
from bs4 import BeautifulSoup as bs
from splinter import Browser
from requests import get
import pandas as pd
# + pycharm={"name": "#%%\n", "is_executing": false}
#executable_path = {'executable_path': 'chromedriver.exe'}
#browser = Browser('chrome', **executable_path, headless=False)
# + pycharm={"name": "#%%\n", "is_executing": false}
url = "https://mars.nasa.gov/news"
response = get(url)
# + pycharm={"name": "#%%\n", "is_executing": false}
soup = bs(response.text, 'html.parser')
news_title = soup.find('div', class_='content_title').text
print(news_title)
# + pycharm={"name": "#%%\n", "is_executing": false}
news_p = soup.find('p').text
print(news_p.strip())
# + pycharm={"name": "#%%\n", "is_executing": false}
pictureURL = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
response = get(pictureURL)
soup = bs(response.text, 'html.parser')
picture = soup.find('div', {'class' : 'carousel_items'}).findAll('a')
print(picture)
# + pycharm={"name": "#%%\n", "is_executing": false}
factURL = 'https://space-facts.com/mars/'
#facts = soup.find('aside', {'id' : 'text-2'})
#table = soup.find('table', {'id' : 'tablepress-p-mars-no-2'})
tables = pd.read_html(factURL)
tables
# + pycharm={"name": "#%%\n"}
| Mission_to_Mars/mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.4 64-bit
# name: python36464bitc2077ed07ea84d23aa5b518d224882ab
# ---
from sklearn.cluster import KMeans
import numpy as np
X = np.array([[1, 2], [1, 4], [1, 0],[10, 2], [10, 4], [10, 0]])
print("X:\n", X)
kmeans = KMeans(n_clusters=2, random_state=0).fit(X)
print("\nkmeans.labels_:", kmeans.labels_)
predict = kmeans.predict([[0, 0], [12, 3]])
print("predict:", predict)
clusters = kmeans.cluster_centers_
print("kmeans.cluster_centers_:\n", clusters)
| week10_ML_competition_pca_kmeans/day3_unsupervised_PCA_kmeans/unsupervised_learning/kmeans/kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Alfredzzx/Linear-Algebra-58020/blob/main/Practical_Exam_lab_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="SsUQVvvWmERa"
# Problem 1
# + colab={"base_uri": "https://localhost:8080/"} id="bGlrvxsVlfqM" outputId="2bc0a937-d694-4806-c600-b68124228d4d"
#numpy
import numpy as np
W = np.array([[1,-3],[2,-2],[3,1]])
print(W)
print()
X = np.array([[3,2,1],[-1,2,3],[2,-1,1]])
print(X)
print()
Y = np.array([[1,2,0],[0,-1,2]])
print(Y)
print()
Z = np.array([[0,1],[2,3]])
print(Z)
print()
dot = np.dot(W,Y)
print(dot)
print()
print(Z+Z)
print()
dot = np.dot(3,X)
print(dot)
print()
# + [markdown] id="hNuYAsCZmAlO"
# Problem 2
#
# + colab={"base_uri": "https://localhost:8080/"} id="P49p0Va0lh38" outputId="5a419e27-1be1-4d66-faec-d29549a93332"
#2) A
Y = np.array([[1, 2, 0],[0,-1,2]])
W = np.array([[1, -3],[2,-2],[3,1]])
print(np.dot(Y,W)^2)
# + colab={"base_uri": "https://localhost:8080/"} id="Ijffgn2Ilrr6" outputId="1568ae96-4540-4152-fdeb-447ba0a724ce"
#2) B
import numpy as np
W = np.array([[1, -3],[2,-2],[3,1]])
print('Vector W is', W)
# + colab={"base_uri": "https://localhost:8080/"} id="OOE_sltfluXo" outputId="10e05b6d-4d35-4f38-9c55-e17733a57255"
import numpy as np
Y = np.array([[1, 2, 0],[0,-1,2]])
print('Vector Y is', Y)
# + colab={"base_uri": "https://localhost:8080/"} id="PKAuOxN1lxB6" outputId="ddb96678-5ecf-425c-beb2-2805f532ed7c"
import numpy as np
X = np.array([[3,2,1],[-1,2,3],[2,-1,1]])
print('Vector X is', X)
# + colab={"base_uri": "https://localhost:8080/"} id="_Lbo6ncalzg9" outputId="4a996825-7c5d-498b-a77b-bde2d995d95d"
R = np.dot(W,Y)
print(R)
# + colab={"base_uri": "https://localhost:8080/"} id="Cq6gIvrUl1KA" outputId="6c8a954b-9ca4-48d0-aabd-60a4cb16d219"
print(np.subtract(R,X))
#2) B answer
# + colab={"base_uri": "https://localhost:8080/"} id="xHHqe25dl3PL" outputId="6c6de368-e7a5-4399-bcaa-ecf6335d66e7"
#3) C
import numpy as np
W = np.array([[1, -3],[2,-2],[3,1]])
print('Vector W is', W)
# + colab={"base_uri": "https://localhost:8080/"} id="s7V6Wix3l5X_" outputId="843780d5-322c-4edb-98ef-b29a474098f6"
import numpy as np
Y = np.array([[1, 2, 0],[0,-1,2]])
print('Vector Y is', Y)
# + colab={"base_uri": "https://localhost:8080/"} id="uycxKsQPl7Km" outputId="b27627dc-9df2-41db-a42e-bb4f443ddce1"
import numpy as np
Z = np.array([[0,1],[2,3]])
print('Vector Z is', Z)
# + colab={"base_uri": "https://localhost:8080/"} id="K7xmH26ul8mi" outputId="c33c1f75-91c2-442e-cbca-1ceb30162dd9"
print("(YW - Z^2)")
print(np.dot(Y,W)-Z^2)
# + id="6cO3hRYBl_cg"
| Practical_Exam_lab_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # International Space Station Tracking with Redis Queue
# - Project to repeating the blog post and youtube below:
# - https://www.twilio.com/blog/2015/11/international-space-station-notifications-with-python-redis-queue-and-twilio-copilot.html
# - https://www.youtube.com/watch?v=lal6eWVCEJs&ab_channel=PyGotham2018
# ## Prepare Environment Variables
# +
# # !pip install python-dotenv
# -
from dotenv import load_dotenv
import os
load_dotenv()
host = os.getenv('REDIS_HOST')
port = os.getenv('REDIS_PORT')
# ## Get Redis server ready
# +
import requests
from redis import Redis
from rq import Queue
from rq_scheduler import Scheduler
from datetime import datetime
import pytz
# -
import dask
redis_server = Redis(host, port)
scheduler = Scheduler(connection = redis_server)
scheduler.count()
scheduler
jobs = list(scheduler.get_jobs())
# ## Tracking function with ISS
def get_next_pass(lat, lon):
proxies = {'http': os.getenv('PROXIES'),
'https': os.getenv('PROXIES')}
iss_url = 'http://api.open-notify.org/iss-pass.json'
location = {'lat': lat, 'lon': lon}
response = requests.get(iss_url, params=location, proxies = proxies).json()
if 'response' in response:
next_pass = response['response'][0]['risetime']
next_pass_datetime = datetime.fromtimestamp(next_pass, tz=pytz.utc)
print('Next pass for {}, {} is: {}'
.format(lat, lon, next_pass_datetime))
return next_pass_datetime
else:
print('No ISS flyby can be determined for {}, {}'.format(lat, lon))
get_next_pass(37.7833, -122.4167)
| international_space_station_tracking/ISS_Tracking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業 : (Kaggle)鐵達尼生存預測
# https://www.kaggle.com/c/titanic
# # [作業目標]
# - 試著模仿範例寫法, 在鐵達尼生存預測中, 觀察標籤編碼與獨編碼熱的影響
# # [作業重點]
# - 回答在範例中的觀察結果
# - 觀察標籤編碼與獨熱編碼, 在特徵數量 / 邏輯斯迴歸分數 / 邏輯斯迴歸時間上, 分別有什麼影響 (In[3], Out[3], In[4], Out[4])
# # 作業1
# * 觀察範例,在房價預測中調整標籤編碼(Label Encoder) / 獨熱編碼 (One Hot Encoder) 方式,
# 對於線性迴歸以及梯度提升樹兩種模型,何者影響比較大?
# +
# 做完特徵工程前的所有準備 (與前範例相同)
import pandas as pd
import numpy as np
import copy, time
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.preprocessing import LabelEncoder
data_path = 'data/'
df_train = pd.read_csv(data_path + 'titanic_train.csv')
df_test = pd.read_csv(data_path + 'titanic_test.csv')
train_Y = df_train['Survived']
ids = df_test['PassengerId']
df_train = df_train.drop(['PassengerId', 'Survived'] , axis=1)
df_test = df_test.drop(['PassengerId'] , axis=1)
df = pd.concat([df_train,df_test])
df.head()
# +
#只取類別值 (object) 型欄位, 存於 object_features 中
object_features = []
for dtype, feature in zip(df.dtypes, df.columns):
if dtype == 'object':
object_features.append(feature)
print(f'{len(object_features)} Numeric Features : {object_features}\n')
# 只留類別型欄位
df = df[object_features]
df = df.fillna('None')
train_num = train_Y.shape[0]
df.head()
# -
# 標籤編碼 + 線性迴歸
df_temp = pd.DataFrame()
for c in df.columns:
df_temp[c] = LabelEncoder().fit_transform(df[c])
train_X = df_temp[:train_num]
estimator = LinearRegression()
start = time.time()
print(f'shape : {train_X.shape}')
print(f'score : {cross_val_score(estimator, train_X, train_Y, cv=5).mean()}')
print(f'time : {time.time() - start} sec')
# 獨熱編碼 + 線性迴歸
df_temp = pd.get_dummies(df)
train_X = df_temp[:train_num]
estimator = LinearRegression()
start = time.time()
print(f'shape : {train_X.shape}')
print(f'score : {cross_val_score(estimator, train_X, train_Y, cv=5).mean()}')
print(f'time : {time.time() - start} sec')
# 標籤編碼 + 梯度提升樹
df_temp = pd.DataFrame()
for c in df.columns:
df_temp[c] = LabelEncoder().fit_transform(df[c])
train_X = df_temp[:train_num]
estimator = GradientBoostingRegressor()
start = time.time()
print(f'shape : {train_X.shape}')
print(f'score : {cross_val_score(estimator, train_X, train_Y, cv=5).mean()}')
print(f'time : {time.time() - start} sec')
# 獨熱編碼 + 梯度提升樹
df_temp = pd.get_dummies(df)
train_X = df_temp[:train_num]
estimator = GradientBoostingRegressor()
start = time.time()
print(f'shape : {train_X.shape}')
print(f'score : {cross_val_score(estimator, train_X, train_Y, cv=5).mean()}')
print(f'time : {time.time() - start} sec')
# # 作業2
# * 鐵達尼號例題中,標籤編碼 / 獨熱編碼又分別對預測結果有何影響? (Hint : 參考今日範例)
# 標籤編碼 + 羅吉斯迴歸
df_temp = pd.DataFrame()
for c in df.columns:
df_temp[c] = LabelEncoder().fit_transform(df[c])
train_X = df_temp[:train_num]
estimator = LogisticRegression(solver ='lbfgs')
start = time.time()
print(f'shape : {train_X.shape}')
print(f'score : {cross_val_score(estimator, train_X, train_Y, cv=5).mean()}')
print(f'time : {time.time() - start} sec')
# 獨熱編碼 + 羅吉斯迴歸
df_temp = pd.get_dummies(df)
train_X = df_temp[:train_num]
estimator = LogisticRegression(solver ='lbfgs')
start = time.time()
print(f'shape : {train_X.shape}')
print(f'score : {cross_val_score(estimator, train_X, train_Y, cv=5).mean()}')
print(f'time : {time.time() - start} sec')
| HomeWork/Day_024_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import pandas as pd
import json
import re
import glob
from scipy.sparse import csr_matrix
import numpy as np
from sklearn.preprocessing import LabelEncoder
from tqdm.notebook import tqdm
# -
data_path = glob.glob("../data/*.json")
data_path
df_train = pd.read_json("../data/train.json")
df_val = pd.read_json("../data/val.json")
df_test = pd.read_json("../data/test.json")
# +
# tags_list = []
# for i, item in tqdm(df_train.tags.items(), total=115071):
# tags_list.extend(item)
# -
df_meta = pd.read_json("../data/song_meta.json")
# +
# df_gn = pd.read_json('genre_gn_all.json')
# -
df_meta.describe()
df_train.shape
df_val.shape
df_test.shape
df_meta
tag_arr = []
for tags in df_test.tags:
tag_arr.extend(tags)
tag_set2 = list(set(tag_arr))
tag_set2.sort()
# +
# tag_set
# -
len(tag_set)
len(tag_set2)
tag_same = []
for target in tag_set2:
left = 0
right = len(tag_set)
while right > left:
mid = (right + left) // 2
if tag_set[mid] == target:
tag_same.append(target)
break
elif tag_set[mid] < target:
left = mid + 1
else:
right = mid - 1
len(tag_same)
df_train
df_test.plylst_title.nunique()
df_test['song_len'] = df_test.songs.map(lambda x: len(x))
df_test[df_test.song_len == 0]
df_test.song_len.value_counts().sort_index()
df_test
df_meta
# ## Data Preprocessing
def data_prep(df, col):
df = df.explode(col)[["id","songs","like_cnt"]]
df.columns = ['user_id', 'item_id', "rating"]
df['rating'] = np.log(df.rating+1).transform(int)
return df.reset_index(drop=True)
# +
df = pd.concat([df_train, df_val]).reset_index(drop=True)
X_train = data_prep(df, 'songs')
# X_val = data_prep(df_val, 'songs')
# X_test = data_prep(df_test, 'songs')
X_train.rating.value_counts()
# -
df_train
# +
# X
# -
# # Model
user_enc = LabelEncoder()
result = user_enc.fit_transform(X_train['user_id'])
# ?csr_matrix
result
class EASE:
def __init__(self):
self.user_enc = LabelEncoder()
self.item_enc = LabelEncoder()
def _get_users_and_items(self, df):
users = self.user_enc.fit_transform(df['user_id'])
print('user done')
items = self.item_enc.fit_transform(df['item_id'])
print('items done')
return users, items
def fit(self, df, lambda_: float = 0.5, implicit=True):
"""
df: pandas.DataFrame with columns user_id, item_id and (rating)
lambda_: l2-regularization term
implicit: if True, ratings are ignored and taken as 1, else normalized ratings are used
"""
print("fit Start")
users, items = self._get_users_and_items(df)
values = np.ones(df.shape[0]) if implicit else df['rating'].to_numpy() / df['rating'].max()
print("csr_matrix Start")
X = csr_matrix((values, (users, items)))
self.X = X
G = X.T.dot(X).toarray()
diagIndices = np.diag_indices(G.shape[0])
G[diagIndices] += lambda_
P = np.linalg.inv(G)
B = P / (-np.diag(P))
B[diagIndices] = 0
self.B = B
self.pred = X.dot(B)
def predict(self, train, users, items, k):
df = pd.DataFrame()
items = self.item_enc.transform(items)
dd = train.loc[train.user_id.isin(users)]
dd['ci'] = self.item_enc.transform(dd.item_id)
dd['cu'] = self.user_enc.transform(dd.user_id)
g = dd.groupby('user_id')
for user, group in tqdm(g):
watched = set(group['ci'])
candidates = [item for item in items if item not in watched]
u = group['cu'].iloc[0]
pred = np.take(self.pred[u, :], candidates)
res = np.argpartition(pred, -k)[-k:]
r = pd.DataFrame({
"user_id": [user] * len(res),
"item_id": np.take(candidates, res),
"score": np.take(pred, res)
}).sort_values('score', ascending=False)
df = df.append(r, ignore_index=True)
df['item_id'] = self.item_enc.inverse_transform(df['item_id'])
return df
model = EASE()
model.fit(X_train)
X.shape
from scipy.sparse.linalg import inv
| csy_folder/kakao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:deep_rl]
# language: python
# name: conda-env-deep_rl-py
# ---
# # OPEN-AI Taxi-V3
# +
from agent import Agent
from monitor import interact
import gym
import numpy as np
from bayes_opt import BayesianOptimization
# %load_ext autoreload
# %autoreload 2
num_episodes = 20000
env = gym.make('Taxi-v3')
agent = Agent(epsilon=0.1, epsilon_divisor = 2.0,alpha=0.1, gamma=0.9)
avg_rewards, best_avg_reward = interact(env, agent, num_episodes)
# -
def interact_wrapper(epsilon, epsilon_divisor, alpha, gamma):
agent = Agent(epsilon=epsilon, epsilon_divisor = epsilon_divisor,alpha=alpha, gamma=gamma)
avg_rewards, best_avg_reward = interact(env, agent, num_episodes)
return best_avg_reward
# +
pbounds = {'epsilon': (0.001, 0.01), 'epsilon_divisor' : (1, 50), 'alpha': (0.1, 0.99), 'gamma': (0.1, 1.0)}
optimizer = BayesianOptimization(
f=interact_wrapper,
pbounds=pbounds,
random_state=47
)
optimizer.probe(
params={'epsilon': 0.01, 'epsilon_divisor' : 5, 'alpha': 0.9, 'gamma': 0.9},
lazy=True,
)
optimizer.maximize(
init_points=5,
n_iter=100
)
# -
| lab-taxi/Optimization_Taxi_V3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Brain-hacking 101
#
# Author: [**<NAME>**](http://arokem.org), [**The University of Washington eScience Institute**](http://escience.washington.edu)
# ### Hack 1: Read your data into an array
#
# When you conduct a neuroimaging experiment, the computer that controls the scanner and receives the data from the scanner saves your data to a file. Neuroimaging data appears in many different file formats: `NiFTI`, `Minc`, `Dicom`, etc. These files all contain representations of the data that you collected in the form of an **array**.
#
# What is an array? It is a way of representing the data in the computer memory as a *table*, that is *multi-dimensional* and *homogenous*.
#
# What does this mean?
#
# - *table* means that you will be able to read all or some of the numbers representing your data by addressing the variable that holds your array. It's like addressing a member of your lab to tell you the answer to a question you have, except here you are going to 'ask' a variable in your computer memory. Arrays are usually not as smart as your lab members, but they have very good memory.
#
# - *multi-dimensional* means that you can represent different aspects of your data along different axes. For example, the three dimensions of space can be represented in different dimensions of the table:
# 
# - *homogenous* actually means two different things:
# - The shape of the array is homogenous, so if there are three items in the first column, there have to be three items in all the columns.
# - The data-type is homogenous. If the first item is an integer, all the other items will be integers as well.
# To demonstrate the properties of arrays, we will use the [`numpy`](https://numpy.org) library. This library contains implementations of many scientifically useful functions and objects. In particular, it contains an implementation of arrays that we will use throughout the folllowing examples.
import numpy as np
# +
# Numpy is a package. To see what's in a package, type the name, a period, then hit tab
# #np?
#np.
# -
# Some examples of numpy functions and "things":
print(np.sqrt(4))
print(np.pi) # Not a function, just a variable
print(np.sin(np.pi)) # A function on a variable :)
# ### Numpy arrays (ndarrays)
#
# Creating a NumPy array is as simple as passing a sequence to `np.array`
arr1 = np.array([1, 2.3, 4])
print(type(arr1))
print(arr1.dtype)
print(arr1.shape)
print(arr1)
# ### You can create arrays with special generating functions:
#
# `np.arange(start, stop, [step])`
#
# `np.zeros(shape)`
#
# `np.ones(shape)`
arr4 = np.arange(2, 5)
print(arr4)
arr5 = np.arange(1, 5, 2)
print(arr5)
arr6 = np.arange(1, 10, 2)
print(arr6)
# ## Exercise : Create an Array
#
# Create an array with values ranging from 0 to 10, in increments of 0.5.
#
# Reminder: get help by typing np.arange?, np.ndarray?, np.array?, etc.
# ### Arithmetic with arrays
#
# Since numpy exists to perform efficient numerical operations in Python, arrays have all the usual arithmetic operations available to them. These operations are performed element-wise (i.e. the same operation is performed independently on each element of the array).
# +
A = np.arange(5)
B = np.arange(5, 10)
print (A+B)
print(B-A)
print(A*B)
# -
# ### What would happen if A and B did not have the same `shape`?
# ### Arithmetic with scalars:
#
# In addition, if one of the arguments is a scalar, that value will be applied to all the elements of the array.
A = np.arange(5)
print(A+10)
print(2*A)
print(A**2)
# ### Arrays are addressed through indexing
#
# **Python uses zero-based indexing**: The first item in the array is item `0`
#
# The second item is item `1`, the third is item `2`, etc.
print(A)
print(A[0])
print(A[1])
print(A[2])
# ### `numpy` contains various functions for calculations on arrays
# +
# This gets the exponent, element-wise:
print(np.exp(A))
# This is the average number in the entire array:
print(np.mean(A))
# -
# ### Data in Nifti files is stored as an array
#
# In the tutorial directory, we have included a single run of an fMRI experiment that was included in the FIAC competition. The experiment is described in full in a paper by Dehaene-Lambertz et al. (2006), but for the purposes of what we do today, the exact details of the acquisition and the task are not particularly important.
#
# We can read out this array into the computer memory using the `nibabel` library
import nibabel as nib
# Loading the file is simple:
img = nib.load('./data/run1.nii.gz')
# But note that in order to save time and memory, nibabel is pretty lazy about reading data from file, until we really need this data.
#
# Meaning that at this point, we've only read information *about* the data, not the data itself. This thing is not the data array yet. What is it then?
type(img)
# It's a `Nifti1Image` object! That means that it is a variable that holds various attributes of the data. For example, the 4 by 4 matrix that describes the spatial transformation between the world coordinates and the image coordinates
img.affine
# This object also has functions. You can get the data, by calling a function of that object:
# There's a header in there that provides some additional information:
#
hdr = img.get_header()
print(hdr.get_zooms())
data = img.get_data()
print(type(data))
print(data.shape)
# This is a 4-dimensional array! We happen to know that time is the last dimension, and there are 191 TRs recorded in this data. There are 30 slices in each TR/volume, with an inplane matrix of 64 by 64 in each slice.
# We can easily access different parts of the data. Here is the full time-series for the central voxel in the volume:
center_voxel_time_series = data[32, 32, 15, :]
print(center_voxel_time_series)
print(center_voxel_time_series.shape)
# It's a one-dimensinal array! Here is the middle slice for the last time-point
middle_slice_t0 = data[:, :, 15, -1] # Using negative numbers allows you to count *from the end*
print(middle_slice_t0)
print(middle_slice_t0.shape)
# That's a 2D array. You get the picture, I hope.
#
# You can do all kinds of operations with the data using functions:
print(np.mean(center_voxel_time_series))
print(np.std(center_voxel_time_series))
# TSNR is mean/std:
print(np.mean(center_voxel_time_series)/np.std(center_voxel_time_series))
# ### Using functions on parts of the data
#
# Many `numpy` functions have an `axis` optional argument. These arguments allow you to perform a reduction of the data along one of the dimensions of the array.
#
# For example, if you want to extract a 3D array with the mean/std in every one of the voxels:
mean_tseries = np.mean(data, axis=-1) # Select the last dimension
std_tseries = np.std(data, axis=-1)
tsnr = mean_tseries/std_tseries
print(tsnr.shape)
# You can save the resulting array into a new file:
new_img = nib.Nifti1Image(tsnr, img.affine)
new_img.to_filename('tsnr.nii.gz')
| beginner-python/001-arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/francisquintallauzon/machine-learning-engineering-for-production-public/blob/main/C3_W2_Lab_3_Quantization_and_Pruning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="D3-cg2_rYfe6"
# # Ungraded Lab: Quantization and Pruning
#
# In this lab, you will get some hands-on practice with the mobile optimization techniques discussed in the lectures. These enable reduced model size and latency which makes it ideal for edge and IOT devices. You will start by training a Keras model then compare its model size and accuracy after going through these techniques:
#
# * post-training quantization
# * quantization aware training
# * weight pruning
#
# Let's begin!
# + [markdown] id="0gRaAOIsba55"
# ## Imports
# + [markdown] id="4nVRm10UNHZ9"
# Let's first import a few common libraries that you'll be using throughout the notebook.
# + id="9sL5kmRZbZxX"
import tensorflow as tf
import numpy as np
import os
import tempfile
import zipfile
# + [markdown] id="GS5gXwABm7XP"
# <a name='utilities'>
#
# ## Utilities and constants
#
# Let's first define a few string constants and utility functions to make our code easier to maintain.
# + id="nEuiXyPZMKQm"
# GLOBAL VARIABLES
# String constants for model filenames
FILE_WEIGHTS = 'baseline_weights.h5'
FILE_NON_QUANTIZED_H5 = 'non_quantized.h5'
FILE_NON_QUANTIZED_TFLITE = 'non_quantized.tflite'
FILE_PT_QUANTIZED = 'post_training_quantized.tflite'
FILE_QAT_QUANTIZED = 'quant_aware_quantized.tflite'
FILE_PRUNED_MODEL_H5 = 'pruned_model.h5'
FILE_PRUNED_QUANTIZED_TFLITE = 'pruned_quantized.tflite'
FILE_PRUNED_NON_QUANTIZED_TFLITE = 'pruned_non_quantized.tflite'
# Dictionaries to hold measurements
MODEL_SIZE = {}
ACCURACY = {}
# + id="pqdSGWccdk8G"
# UTILITY FUNCTIONS
def print_metric(metric_dict, metric_name):
'''Prints key and values stored in a dictionary'''
for metric, value in metric_dict.items():
print(f'{metric_name} for {metric}: {value}')
def model_builder():
'''Returns a shallow CNN for training on the MNIST dataset'''
keras = tf.keras
# Define the model architecture.
model = keras.Sequential([
keras.layers.InputLayer(input_shape=(28, 28)),
keras.layers.Reshape(target_shape=(28, 28, 1)),
keras.layers.Conv2D(filters=12, kernel_size=(3, 3), activation='relu'),
keras.layers.MaxPooling2D(pool_size=(2, 2)),
keras.layers.Flatten(),
keras.layers.Dense(10, activation='softmax')
])
return model
def evaluate_tflite_model(filename, x_test, y_test):
'''
Measures the accuracy of a given TF Lite model and test set
Args:
filename (string) - filename of the model to load
x_test (numpy array) - test images
y_test (numpy array) - test labels
Returns
float showing the accuracy against the test set
'''
# Initialize the TF Lite Interpreter and allocate tensors
interpreter = tf.lite.Interpreter(model_path=filename)
interpreter.allocate_tensors()
# Get input and output index
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# Initialize empty predictions list
prediction_digits = []
# Run predictions on every image in the "test" dataset.
for i, test_image in enumerate(x_test):
# Pre-processing: add batch dimension and convert to float32 to match with
# the model's input data format.
test_image = np.expand_dims(test_image, axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
digit = np.argmax(output()[0])
prediction_digits.append(digit)
# Compare prediction results with ground truth labels to calculate accuracy.
prediction_digits = np.array(prediction_digits)
accuracy = (prediction_digits == y_test).mean()
return accuracy
def get_gzipped_model_size(file):
'''Returns size of gzipped model, in bytes.'''
_, zipped_file = tempfile.mkstemp('.zip')
with zipfile.ZipFile(zipped_file, 'w', compression=zipfile.ZIP_DEFLATED) as f:
f.write(file)
return os.path.getsize(zipped_file)
# + [markdown] id="AxnjOqLpYawi"
# ## Download and Prepare the Dataset
# + [markdown] id="rfC0D71tnVKr"
# You will be using the [MNIST](https://keras.io/api/datasets/mnist/) dataset which is hosted in [Keras Datasets](https://keras.io/api/datasets/). Some of the helper files in this notebook are made to work with this dataset so if you decide to switch to a different dataset, make sure to check if those helper functions need to be modified (e.g. shape of the Flatten layer in your model).
# + id="Z5f5Y08r0sob"
# Load MNIST dataset
mnist = tf.keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# Normalize the input image so that each pixel value is between 0 to 1.
train_images = train_images / 255.0
test_images = test_images / 255.0
# + [markdown] id="Czvt9P1EYnQT"
# ## Baseline Model
#
# You will first build and train a Keras model. This will be the baseline where you will be comparing the mobile optimized versions later on. This will just be a shallow CNN with a softmax output to classify a given MNIST digit. You can review the `model_builder()` function in the utilities at the top of this notebook but we also printed the model summary below to show the architecture.
#
# You will also save the weights so you can reinitialize the other models later the same way. This is not needed in real projects but for this demo notebook, it would be good to have the same initial state later so you can compare the effects of the optimizations.
# + id="3Ild5juYXu4j"
# Create the baseline model
baseline_model = model_builder()
# Save the initial weights for use later
baseline_model.save_weights(FILE_WEIGHTS)
# Print the model summary
baseline_model.summary()
# + [markdown] id="74y6LJMVYRCL"
# You can then compile and train the model. In practice, it's best to shuffle the train set but for this demo, it is set to `False` for reproducibility of the results. One epoch below will reach around 91% accuracy.
# + id="xViB61FuY0Pf"
# Setup the model for training
baseline_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train the model
baseline_model.fit(train_images, train_labels, epochs=1, shuffle=False)
# + [markdown] id="47BgpWwOaR8b"
# Let's save the accuracy of the model against the test set so you can compare later.
# + id="JQSVh1_t4Z2h"
# Get the baseline accuracy
_, ACCURACY['baseline Keras model'] = baseline_model.evaluate(test_images, test_labels)
# + [markdown] id="aAfbP3uua6bE"
# Next, you will save the Keras model as a file and record its size as well.
# + id="_A8WPjzqLbH3"
# Save the Keras model
baseline_model.save(FILE_NON_QUANTIZED_H5, include_optimizer=False)
# Save and get the model size
MODEL_SIZE['baseline h5'] = os.path.getsize(FILE_NON_QUANTIZED_H5)
# Print records so far
print_metric(ACCURACY, "test accuracy")
print_metric(MODEL_SIZE, "model size in bytes")
# + [markdown] id="Ak8rBX-qX_KM"
# ### Convert the model to TF Lite format
# + [markdown] id="PpkXpDy_OCzB"
# Next, you will convert the model to [Tensorflow Lite (TF Lite)](https://www.tensorflow.org/lite/guide) format. This is designed to make Tensorflow models more efficient and lightweight when running on mobile, embedded, and IOT devices.
#
# You can convert a Keras model with TF Lite's [Converter](https://www.tensorflow.org/lite/convert/index) class and we've incorporated it in the short helper function below. Notice that there is a `quantize` flag which you can use to quantize the model.
# + id="zQYM0A0SgCNS"
def convert_tflite(model, filename, quantize=False):
'''
Converts the model to TF Lite format and writes to a file
Args:
model (Keras model) - model to convert to TF Lite
filename (string) - string to use when saving the file
quantize (bool) - flag to indicate quantization
Returns:
None
'''
# Initialize the converter
converter = tf.lite.TFLiteConverter.from_keras_model(model)
# Set for quantization if flag is set to True
if quantize:
converter.optimizations = [tf.lite.Optimize.DEFAULT]
# Convert the model
tflite_model = converter.convert()
# Save the model.
with open(filename, 'wb') as f:
f.write(tflite_model)
# + [markdown] id="lQkC9plnP2pU"
# You will use the helper function to convert the Keras model then get its size and accuracy. Take note that this is *not yet* quantized.
# + id="5H61feiOZkcI"
# Convert baseline model
convert_tflite(baseline_model, FILE_NON_QUANTIZED_TFLITE)
# + [markdown] id="REf-EaQlQoYZ"
# You will notice that there is already a slight decrease in model size when converting to `.tflite` format.
# + id="cmlNGwbCBo8v"
MODEL_SIZE['non quantized tflite'] = os.path.getsize(FILE_NON_QUANTIZED_TFLITE)
print_metric(MODEL_SIZE, 'model size in bytes')
# + [markdown] id="Rp-ndoNSRnvX"
# The accuracy will also be nearly identical when converting between formats. You can setup a TF Lite model for input-output using its [Interpreter](https://www.tensorflow.org/api_docs/python/tf/lite/Interpreter) class. This is shown in the `evaluate_tflite_model()` helper function provided in the `Utilities` section earlier.
#
# *Note: If you see a `Runtime Error: There is at least 1 reference to internal data in the interpreter in the form of a numpy array or slice.` , please try re-running the cell.*
# + id="OQFkh5ukiiZE"
ACCURACY['non quantized tflite'] = evaluate_tflite_model(FILE_NON_QUANTIZED_TFLITE, test_images, test_labels)
# + id="CplCOws3jaB0"
print_metric(ACCURACY, 'test accuracy')
# + [markdown] id="N6ilHiSGYCFL"
# ### Post-Training Quantization
#
# Now that you have the baseline metrics, you can now observe the effects of quantization. As mentioned in the lectures, this process involves converting floating point representations into integer to reduce model size and achieve faster computation.
#
# As shown in the `convert_tflite()` helper function earlier, you can easily do [post-training quantization](https://www.tensorflow.org/lite/performance/post_training_quantization) with the TF Lite API. You just need to set the converter optimization and assign an [Optimize](https://www.tensorflow.org/api_docs/python/tf/lite/Optimize) Enum.
#
# You will set the `quantize` flag to do that and get the metrics again.
# + id="DdWNTJ2J1OpL"
# Convert and quantize the baseline model
convert_tflite(baseline_model, FILE_PT_QUANTIZED, quantize=True)
# + id="cTFHf4Rw1bCJ"
# Get the model size
MODEL_SIZE['post training quantized tflite'] = os.path.getsize(FILE_PT_QUANTIZED)
print_metric(MODEL_SIZE, 'model size')
# + [markdown] id="SYcBZduWVqOH"
# You should see around a 4X reduction in model size in the quantized version. This comes from converting the 32 bit representations (float) into 8 bits (integer).
#
#
# + id="vhEYoQ83-pT_"
ACCURACY['post training quantized tflite'] = evaluate_tflite_model(FILE_PT_QUANTIZED, test_images, test_labels)
# + id="4D0Srsjb_inn"
print_metric(ACCURACY, 'test accuracy')
# + [markdown] id="rGTzSOuQWG4L"
# As mentioned in the lecture, you can expect the accuracy to not be the same when quantizing the model. Most of the time it will decrease but in some cases, it can even increase. Again, this can be attributed to the loss of precision when you remove the extra bits from the float data.
# + [markdown] id="vFf1DDVnYIes"
# ## Quantization Aware Training
# + [markdown] id="37oAb7PuXK36"
# When post-training quantization results in loss of accuracy that is unacceptable for your application, you can consider doing [quantization aware training](https://www.tensorflow.org/model_optimization/guide/quantization/training) before quantizing the model. This simulates the loss of precision by inserting fake quant nodes in the model during training. That way, your model will learn to adapt with the loss of precision to get more accurate predictions.
#
# The [Tensorflow Model Optimization Toolkit](https://www.tensorflow.org/model_optimization) provides a [quantize_model()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/quantization/keras/quantize_model) method to do this quickly and you will see that below. But first, let's install the toolkit into the notebook environment.
# + id="6WSt6OQGoNAt"
# Install the toolkit
# !pip install tensorflow_model_optimization
# + [markdown] id="oYHmeMihYjnB"
# You will build the baseline model again but this time, you will pass it into the `quantize_model()` method to indicate quantization aware training.
#
# Take note that in case you decide to pass in a model that is already trained, then make sure to recompile before you continue training.
# + id="3dGSpz0on2C4"
import tensorflow_model_optimization as tfmot
# method to quantize a Keras model
quantize_model = tfmot.quantization.keras.quantize_model
# Define the model architecture.
model_to_quantize = model_builder()
# Reinitialize weights with saved file
model_to_quantize.load_weights(FILE_WEIGHTS)
# Quantize the model
q_aware_model = quantize_model(model_to_quantize)
# `quantize_model` requires a recompile.
q_aware_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
q_aware_model.summary()
# + [markdown] id="lmcaLaotZ7G7"
# You may have noticed a slight difference in the model summary above compared to the baseline model summary in the earlier sections. The total params count increased as expected because of the nodes added by the `quantize_model()` method.
#
# With that, you can now train the model. You will notice that the accuracy is a bit lower because the model is simulating the loss of precision. The training will take a bit longer if you want to achieve the same training accuracy as the earlier run. For this exercise though, we will keep to 1 epoch.
# + id="yl4jbjllomDw"
# Train the model
q_aware_model.fit(train_images, train_labels, epochs=1, shuffle=False)
# + [markdown] id="b_WAM2C4bWeC"
# You can then get the accuracy of the Keras model before and after quantizing the model. The accuracy is expected to be nearly identical because the model is trained to counter the effects of quantization.
# + id="J7rOuwM_ozI_"
# Reinitialize the dictionary
ACCURACY = {}
# Get the accuracy of the quantization aware trained model (not yet quantized)
_, ACCURACY['quantization aware non-quantized'] = q_aware_model.evaluate(test_images, test_labels, verbose=0)
print_metric(ACCURACY, 'test accuracy')
# + id="6liE_Cp3rzAy"
# Convert and quantize the model.
convert_tflite(q_aware_model, FILE_QAT_QUANTIZED, quantize=True)
# Get the accuracy of the quantized model
ACCURACY['quantization aware quantized'] = evaluate_tflite_model(FILE_QAT_QUANTIZED, test_images, test_labels)
print_metric(ACCURACY, 'test accuracy')
# + [markdown] id="SwvaMflTYNgo"
# ## Pruning
#
# Let's now move on to another technique for reducing model size: [Pruning](https://www.tensorflow.org/model_optimization/guide/pruning/pruning_with_keras). This process involves zeroing out insignificant (i.e. low magnitude) weights. The intuition is these weights do not contribute as much to making predictions so you can remove them and get the same result. Making the weights sparse helps in compressing the model more efficiently and you will see that in this section.
# + [markdown] id="LdlFujrJbzV7"
# The Tensorflow Model Optimization Toolkit again has a convenience method for this. The [prune_low_magnitude()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/prune_low_magnitude) method puts wrappers in a Keras model so it can be pruned during training. You will pass in the baseline model that you already trained earlier. You will notice that the model summary show increased params because of the wrapper layers added by the pruning method.
#
# You can set how the pruning is done during training. Below, you will use [PolynomialDecay](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/PolynomialDecay) to indicate how the sparsity ramps up with each step. Another option available in the library is [Constant Sparsity](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/ConstantSparsity).
# + id="TpqizJsKYPBA"
# Get the pruning method
prune_low_magnitude = tfmot.sparsity.keras.prune_low_magnitude
# Compute end step to finish pruning after 2 epochs.
batch_size = 128
epochs = 2
validation_split = 0.1 # 10% of training set will be used for validation set.
num_images = train_images.shape[0] * (1 - validation_split)
end_step = np.ceil(num_images / batch_size).astype(np.int32) * epochs
# Define pruning schedule.
pruning_params = {
'pruning_schedule': tfmot.sparsity.keras.PolynomialDecay(initial_sparsity=0.50,
final_sparsity=0.80,
begin_step=0,
end_step=end_step)
}
# Pass in the trained baseline model
model_for_pruning = prune_low_magnitude(baseline_model, **pruning_params)
# `prune_low_magnitude` requires a recompile.
model_for_pruning.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model_for_pruning.summary()
# + [markdown] id="qgmHaZI6fip_"
# You can also peek at the weights of one of the layers in your model. After pruning, you will notice that many of these will be zeroed out.
# + id="y5ekdEBigB5l"
# Preview model weights
model_for_pruning.weights[1]
# + [markdown] id="0XFwMRqpgbr0"
# With that, you can now start re-training the model. Take note that the [UpdatePruningStep()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/UpdatePruningStep) callback is required.
# + id="DUCz6PL371Bx"
# Callback to update pruning wrappers at each step
callbacks = [
tfmot.sparsity.keras.UpdatePruningStep(),
]
# Train and prune the model
model_for_pruning.fit(train_images, train_labels,
epochs=epochs, validation_split=validation_split,
callbacks=callbacks)
# + [markdown] id="rEExgy4hhXP-"
# Now see how the weights in the same layer looks like after pruning.
# + id="TOK4TidJhXpT"
# Preview model weights
model_for_pruning.weights[1]
# + [markdown] id="o5ckfDHLhhub"
# After pruning, you can remove the wrapper layers to have the same layers and params as the baseline model. You can do that with the [strip_pruning()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/strip_pruning) method as shown below. You will do this so you can save the model and also export to TF Lite format just like in the previous sections.
# + id="PbfLhZv68vwc"
# Remove pruning wrappers
model_for_export = tfmot.sparsity.keras.strip_pruning(model_for_pruning)
model_for_export.summary()
# + [markdown] id="KtbPlo-kj9Ku"
# You will see the same model weights but the index is different because the wrappers were removed.
# + id="SG6-aF9yiraG"
# Preview model weights (index 1 earlier is now 0 because pruning wrappers were removed)
model_for_export.weights[0]
# + [markdown] id="ZR94MYxLkHfn"
# You will notice below that the pruned model will have the same file size as the baseline_model when saved as H5. This is to be expected. The improvement will be noticeable when you compress the model as will be shown in the cell after this.
# + id="CjjDMqJCTjqz"
# Save Keras model
model_for_export.save(FILE_PRUNED_MODEL_H5, include_optimizer=False)
# Get uncompressed model size of baseline and pruned models
MODEL_SIZE = {}
MODEL_SIZE['baseline h5'] = os.path.getsize(FILE_NON_QUANTIZED_H5)
MODEL_SIZE['pruned non quantized h5'] = os.path.getsize(FILE_PRUNED_MODEL_H5)
print_metric(MODEL_SIZE, 'model_size in bytes')
# + [markdown] id="tCEfa-LRleT_"
# You will use the `get_gzipped_model_size()` helper function in the `Utilities` to compress the models and get its resulting file size. You will notice that the pruned model is about 3 times smaller. This is because of the sparse weights generated by the pruning process. The zeros can be compressed much more efficiently than the low magnitude weights before pruning.
# + id="VWQ_AgiX_yiP"
# Get compressed size of baseline and pruned models
MODEL_SIZE = {}
MODEL_SIZE['baseline h5'] = get_gzipped_model_size(FILE_NON_QUANTIZED_H5)
MODEL_SIZE['pruned non quantized h5'] = get_gzipped_model_size(FILE_PRUNED_MODEL_H5)
print_metric(MODEL_SIZE, "gzipped model size in bytes")
# + [markdown] id="uByyx0L3mlYc"
# You can make the model even more lightweight by quantizing the pruned model. This achieves around 10X reduction in compressed model size as compared to the baseline.
# + id="qIY6n9XWCvt5"
# Convert and quantize the pruned model.
pruned_quantized_tflite = convert_tflite(model_for_export, FILE_PRUNED_QUANTIZED_TFLITE, quantize=True)
# Compress and get the model size
MODEL_SIZE['pruned quantized tflite'] = get_gzipped_model_size(FILE_PRUNED_QUANTIZED_TFLITE)
print_metric(MODEL_SIZE, "gzipped model size in bytes")
# + [markdown] id="v4ytiH3ynIid"
# As expected, the TF Lite model's accuracy will also be close to the Keras model.
# + id="PZBAdJmuWN0A"
# Get accuracy of pruned Keras and TF Lite models
ACCURACY = {}
_, ACCURACY['pruned model h5'] = model_for_pruning.evaluate(test_images, test_labels)
ACCURACY['pruned and quantized tflite'] = evaluate_tflite_model(FILE_PRUNED_QUANTIZED_TFLITE, test_images, test_labels)
print_metric(ACCURACY, 'accuracy')
# + [markdown] id="CpM7t_nGokcz"
# ## Wrap Up
#
# In this notebook, you practiced several techniques in optimizing your models for mobile and embedded applications. You used quantization to reduce floating point representations into integer, then used pruning to make the weights sparse for efficient model compression. These make your models lightweight for efficient transport and storage without sacrificing model accuracy. Try this in your own models and see what performance you get. For more information, here are a few other resources:
#
# * [Post Training Quantization Guide](https://www.tensorflow.org/lite/performance/post_training_quantization)
# * [Quantization Aware Training Comprehensive Guide](https://www.tensorflow.org/model_optimization/guide/quantization/training_comprehensive_guide)
# * [Pruning Comprehensive Guide](https://www.tensorflow.org/model_optimization/guide/pruning/comprehensive_guide)
#
# **Congratulations and enjoy the rest of the course!**
| course3/C3_W2_Lab_3_Quantization_and_Pruning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dimentionality Reduction No. 1
# Import the required libraries.
import pandas as pd
import numpy as np
# Read the data.
data = pd.read_csv('Train_UWu5bXk.csv')
# Read the first lines of the data.
data.head()
# Remove columns which their null value percentage is higher than 20%.
# Calculate the null value percentage for each column.
null_percentage = data.isnull().sum() / len(data) * 100
# Save the data column names into the column_names.
column_names = data.columns
# Introduce the data's new column names.
new_column_names = []
# Create a loop for removing columns which their null value percentage is higher than 20%.
# Or, only accept the columns which their null value percentage is less than 20%.
for i in range(0, len(null_percentage)):
if null_percentage[i] < 20:
new_column_names.append(column_names[i])
# Create a new data with the new columns.
X = data[new_column_names]
# Print the new data.
X
# Calculate the variance of the new data.
X.var().round(2)
# Since dropping of nuisance columns in DataFrame reductions (with 'numeric_only=None') is deprecated,
# We can use this code (select only valid columns).
numercial_column_names = ['Item_Weight', 'Item_Visibility', 'Item_MRP', 'Outlet_Establishment_Year', 'Item_Outlet_Sales']
[[i, X[i].var()] for i in numercial_column_names]
# Remove columns which their variance is greater than 1.
# Introduce the data's new column names.
new_column_names = []
# Create a loop for removing columns which their variance is less than 1.
# Or, only accept the columns which their variance is geater than 1.
for i in numercial_column_names:
if X[i].var() > 1:
new_column_names.append(i)
new_column_names
# Calculate the skewness of the new data.
[[i, X[i].skew()] for i in numercial_column_names]
# Remove columns which their skewness is greater than 0.5 and less than -0.5.
# Introduce the data's new column names.
new_column_names = []
# Create a loop for removing columns which skewness is greater than 0.5 and less than -0.5.
# Or, only accept the columns which their skewness is within 0.5 and -0.5.
for i in numercial_column_names:
if X[i].skew() > -0.5 and X[i].skew() < 0.5:
new_column_names.append(i)
new_column_names
# Calculate the curtosis of the new data.
[[i, X[i].kurtosis()] for i in numercial_column_names]
# Plot the histogram of the data.
data.hist(figsize=(20, 12))
# The input data (X) and the target (y).
X = data[['Item_Weight', 'Item_Fat_Content', 'Item_Visibility', 'Item_Type', 'Item_MRP', 'Outlet_Identifier', 'Outlet_Establishment_Year', 'Outlet_Size', 'Outlet_Location_Type', 'Outlet_Type']]
y = data['Item_Outlet_Sales']
# Read the first lines of the data.
X.head()
# Pre-process the data.
# In order to pre-process the data, we can import sci-kit learn preprocessing library.
# Use its standard scaler.
from sklearn.preprocessing import StandardScaler
# Standard scale the continous columns.
continuous_columns = ['Item_Weight', 'Item_Visibility', 'Item_MRP', 'Outlet_Establishment_Year']
X_scaled = X[continuous_columns]
standard_scaler = StandardScaler()
X_scaled = standard_scaler.fit_transform(X_scaled)
print(X_scaled)
# Return the scaled continuous values to the input data (X).
X[continuous_columns] = X_scaled
# Read the first lines of the data.
X.head()
# Specify the categorical column indexes.
categorical_columns_indexes = [1, 3, 5, 7, 8, 9]
# Check whether the 'Outlet_Size' column contains NaN values.
X['Outlet_Size'].isna()
# Fill the NaN values.
X['Outlet_Size'].fillna(X['Outlet_Size'].mode()[0], inplace=True)
# Read the first lines of the data.
X.head()
# Search for the null values.
X.isnull().sum()
# Prepare train and test datasets.
# In order to do that, we can use a specific package in Sci-Kit library.
from sklearn.model_selection import train_test_split
# Specify the train and test inputs and targets.
# Note that we specify the random state.
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=25)
# Use catboost algorithm for regression.
from catboost import CatBoostRegressor
# Create a new catboost.
catboost_regressor = CatBoostRegressor(iterations=2000, learning_rate=0.05, depth=8, eval_metric='MAE')
# Learn from the data.
catboost_regressor.fit(X_train, y_train, categorical_columns_indexes, eval_set=(X_test, y_test), plot=True)
| Dimentionality_Reduction1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.0
# language: julia
# name: julia-0.6
# ---
using EPhys, PyPlot
bsnms = ["mam5-160307";
"mam9-160528"; "mam9-160524"; "mam9-160527";
"mam10-170213"]
dataloc = "/mnfs/vtad1/data/amorley_merged/"
allmetadata = getmetadata.(bsnms, [dataloc])
allsessions = [find(contains.(x["desen"],"sq flicker"))
for x in allmetadata]
day = 1
bsnm = bsnms[day]
sessions = allsessions[day]
metadata = allmetadata[day]
lfp = load(LFP,metadata,sessions)
testsignal = EPhys.signal(lfp.lfp[1])[1:10*floor(Int,lfp.fs)];
include("$EPhysLP/../../../EMD/src/EMD.jl")
# +
N = 9
C = EMD.IMF(testsignal, t, N=N)
fig,axs = subplots(size(C,2),1,figsize=(10,10))
for i in indices(C,2)
ax = axs[i]
ax[:plot](C[:,i])
end
# -
# +
n = length(testsignal)
#Tukey window
alpha = 0.25
window = zeros(n)
for i = 0:n-1
if i < alpha * (n-1)/2
window[i+1] = 0.5*(1 + cos(pi*(2*i/(alpha*(n-1)) - 1)))
elseif i < (n-1)*(1-alpha/2)
window[i+1] = 1
else
window[i+1] = 0.5*(1 + cos(pi*(2*i/(alpha*(n-1)) + 1 - 2/alpha)))
end
end
# -
(y, t) = testsignal, [linspace(0,n,n);]
toldev=0.01;
tolzero = 0.01;
order=4; N=10; window=0
# +
if window==0
window = ones(length(t))
end
n = length(y)
f = zeros(n,N)
tempy = copy(y)
eps = 0.00001
n_modes = 0;
for i = 1:N
avg = zeros(n,1)+1
sd = 2*toldev
while(mean(abs.(avg))>tolzero && sd > toldev)
# Interpolate a spline through the maxima and minima
max_ar, min_ar, tmax, tmin = findExtrema(tempy, t)
# # Don't use too high an order to interpolate, restrict it if there are not many extrema
# p_max = min(order, length(max_ar)-2)
# p_min = min(order, length(min_ar)-2)
# # Make even order
# p_max = Int(2*floor(p_max/2))
# p_min = Int(2*floor(p_min/2))
# # At least linear
# p_max = max(p_max,2)
# p_min = max(p_min,2)
p_max = (length(max_ar) >= order) ? order : 4
p_min = (length(min_ar) >= order) ? order : 4
while true
p_min = p_min < length(min_ar) ? (p_min; break) : (p_min - 1)
p_max = p_max < length(max_ar) ? (p_max; break) : (p_max - 1)
end
S1 = Spline1D(tmax, max_ar, k = p_max)
S2 = Spline1D(tmin, min_ar, k = p_min)
# Find mean of envelope
avg = (S1(t) + S2(t)) / 2
avg = avg.*window
tempy = tempy-avg
sd = mean( (avg.^2)./((y-f[:,i]).^2 + eps) )
f[:,i] = f[:,i] + avg
#println(sd)
end
tempy = copy(f[:,i])
# Check to see if it's worth continuing or if the remainder is monotone
c = mean(abs.(tempy))
d = diff(tempy)
n_modes = n_modes + 1
if all(d+c*tolzero .> 0) || all(d-c*tolzero .< 0)
break
end
end
C = zeros(n,N)
C[:,1] = y - f[:,1]
for i = 2:n_modes
C[:,i] = f[:,i-1]-f[:,i]
end
return C
# -
using StatsBase
itp = scale(interpolate(min_ar, (BSpline(Cubic(Natural())), NoInterp()), OnGrid(), 1)
scatter(tmin, min_ar, label="minima")
scatter(tmax, max_ar, label="maxima")
# +
scatter(tmin, min_ar, label="minima")
scatter(tmax, max_ar, label="maxima")
plot(t,testsignal, label = "signal")
legend()
# -
plot(tempy)
EMD.IMF(testsignal,window)
plot(testsignal)
| notebooks/EMD_for_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="lCGQ3cZTl5yr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1617843206315, "user_tz": 240, "elapsed": 10753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="ac1e01db-1703-4f49-eed5-b57ec8e1bf85"
import pandas as pd
try:
import pickle5 as pickle
except:
# !pip install pickle5
import pickle5 as pickle
import os
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, Input, GlobalMaxPooling1D, Flatten
from keras.layers import Conv1D, MaxPooling1D, Embedding, Concatenate, Lambda
from keras.models import Model
from sklearn.metrics import roc_auc_score,roc_curve, auc
from numpy import random
from keras.layers import LSTM, Bidirectional, GlobalMaxPool1D, Dropout
from keras.optimizers import Adam
from keras.utils.vis_utils import plot_model
import seaborn as sns
import sys
sys.path.insert(0,'/content/drive/MyDrive/ML_Data/')
import functions as f
# + id="LiE-LZR2mAUz" colab={"base_uri": "https://localhost:8080/", "height": 428} executionInfo={"status": "ok", "timestamp": 1617843220796, "user_tz": 240, "elapsed": 3556, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="452b1d12-45bb-468e-b329-afcb02789d38"
def load_data(randomize=False):
try:
with open("/content/drive/MyDrive/ML_Data/hyppi-train.pkl", "rb") as fh:
df_train = pickle.load(fh)
except:
df_train = pd.read_pickle("C:/Users/nik00/py/proj/hyppi-train.pkl")
try:
with open("/content/drive/MyDrive/ML_Data/hyppi-independent.pkl", "rb") as fh:
df_test = pickle.load(fh)
except:
df_test = pd.read_pickle("C:/Users/nik00/py/proj/hyppi-independent.pkl")
if randomize:
return shuff_together(df_train,df_test)
else:
return df_train,df_test
df_train,df_test = load_data()
print('The data used will be:')
df_train[['Human','Yersinia']]
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="GP1c8AoqnSaT" executionInfo={"status": "ok", "timestamp": 1617498544230, "user_tz": 240, "elapsed": 15206, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="9f0819ef-8069-4331-d698-5a93f20a4e24"
lengths = sorted(len(s) for s in df_train['Human'])
print("Median length of Human sequence is",lengths[len(lengths)//2])
_ = sns.displot(lengths)
_=plt.title("Most Human sequences seem to be less than 2000 in length")
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="VUEUQi3NnShC" executionInfo={"status": "ok", "timestamp": 1617498544230, "user_tz": 240, "elapsed": 15199, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="4121fdf6-6e90-453f-d8b3-0a25e8fb71b6"
lengths = sorted(len(s) for s in df_train['Yersinia'])
print("Median length of Yersinia sequence is",lengths[len(lengths)//2])
_ = sns.displot(lengths)
_=plt.title("Most Yersinia sequences seem to be less than 1000 in length")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="XveNJucynXpK" executionInfo={"status": "ok", "timestamp": 1617843979130, "user_tz": 240, "elapsed": 39179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="c17b65d2-972b-4840-f2a2-98ec02674440"
data1_1D_doubleip_pre,data2_1D_doubleip_pre,data1_test_1D_doubleip_pre,data2_test_1D_doubleip_pre,num_words_1D,MAX_SEQUENCE_LENGTH_1D,MAX_VOCAB_SIZE_1D = f.get_seq_data_doubleip(100,1000,df_train,df_test,pad = 'pre',show=True)
# + colab={"base_uri": "https://localhost:8080/"} id="-Rge3R-ICn5i" executionInfo={"status": "ok", "timestamp": 1617843247128, "user_tz": 240, "elapsed": 14581, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="46aa0360-26c1-45ec-8d88-a7bf6d3504b7"
data1_1D_doubleip_center,data2_1D_doubleip_center,data1_test_1D_doubleip_center,data2_test_1D_doubleip_center,num_words_1D,MAX_SEQUENCE_LENGTH_1D,MAX_VOCAB_SIZE_1D = f.get_seq_data_doubleip(100,1000,df_train,df_test)
# + colab={"base_uri": "https://localhost:8080/"} id="pgHXpN2HCn8l" executionInfo={"status": "ok", "timestamp": 1617843253092, "user_tz": 240, "elapsed": 20535, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="2d9d3514-64fe-45a8-f87a-9e4d35101648"
data1_1D_doubleip_post,data2_1D_doubleip_post,data1_test_1D_doubleip_post,data2_test_1D_doubleip_post,num_words_1D,MAX_SEQUENCE_LENGTH_1D,MAX_VOCAB_SIZE_1D = f.get_seq_data_doubleip(100,1000,df_train,df_test,pad = 'post')
# + colab={"base_uri": "https://localhost:8080/"} id="tbp7kGJKq5V-" executionInfo={"status": "ok", "timestamp": 1617509654648, "user_tz": 240, "elapsed": 1690738, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgMwPlnKw7I-jN5XkbRlGeMro_PH58tNaE2oWC0dw=s64", "userId": "16675707467486783174"}} outputId="4387c0b3-9561-413f-ba14-5bfc790c6216"
EMBEDDING_DIM_1D = 5
VALIDATION_SPLIT = 0.2
BATCH_SIZE = 128
EPOCHS = 50
M_1D=10
x1 = f.BiLSTM_model(MAX_SEQUENCE_LENGTH_1D, EMBEDDING_DIM_1D, num_words_1D, M_1D)
x2 = f.BiLSTM_model(MAX_SEQUENCE_LENGTH_1D, EMBEDDING_DIM_1D, num_words_1D, M_1D)
x3 = f.BiLSTM_model(MAX_SEQUENCE_LENGTH_1D, EMBEDDING_DIM_1D, num_words_1D, M_1D)
x4 = f.BiLSTM_model(MAX_SEQUENCE_LENGTH_1D, EMBEDDING_DIM_1D, num_words_1D, M_1D)
x5 = f.BiLSTM_model(MAX_SEQUENCE_LENGTH_1D, EMBEDDING_DIM_1D, num_words_1D, M_1D)
x6 = f.BiLSTM_model(MAX_SEQUENCE_LENGTH_1D, EMBEDDING_DIM_1D, num_words_1D, M_1D)
concatenator = Concatenate(axis=1)
x = concatenator([x1.output, x2.output, x3.output, x4.output, x5.output, x6.output])
x = Dense(128)(x)
x = Dropout(0.2)(x)
output = Dense(1, activation="sigmoid",name="Final")(x)
model1D_doubleip = Model(inputs=[x1.input, x2.input, x3.input, x4.input, x5.input, x6.input], outputs=output)
model1D_doubleip.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
#plot_model(model1D_doubleip, to_file='model_plot.png', show_shapes=True, show_layer_names=False)
trains = [data1_1D_doubleip_pre,data2_1D_doubleip_pre,data1_1D_doubleip_center,data2_1D_doubleip_center,data1_1D_doubleip_post,data2_1D_doubleip_post]
tests = [data1_test_1D_doubleip_pre,data2_test_1D_doubleip_pre,data1_test_1D_doubleip_center,data2_test_1D_doubleip_center,data1_test_1D_doubleip_post,data2_test_1D_doubleip_post]
model1D_doubleip.fit(trains, df_train['label'].values, epochs=EPOCHS, validation_data=(tests, df_test['label'].values),batch_size=BATCH_SIZE)
print(roc_auc_score(df_test['label'].values, model1D_doubleip.predict(tests)))
# + id="JiruPRTgnmna"
#model1D_doubleip.save('/content/drive/MyDrive/ML_Data/model1D_doubleip.h5')
| 1. Bi-LSTM/ThreeX_1D_doubleip_3X.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Seminar 10. Clustering Hands-on practice
#
# ## Similar password detection
# In this assignment we will try to detect similar patterns in passwords that people use all over the internet.
#
# The input data is a collection of leaked passwords and it can be downloaded from here https://github.com/ignis-sec/Pwdb-Public/tree/master/wordlists
#
# The task is to try to describe the data in terms of clustering: what are the groups of passwords that look quite similar or have similar logic behind them?
#
# This seminar should be considered as a research: there are no correct answers, no points and no deadlines - just your time and your experiments with clustering algorithms.
#
# We suggest to start with the following steps:
# - download the data
# - check if your favourite password is in the database
# - build a distance matrix using Levenstein distance
# - apply DBSCAN
# - apply Agglomerative clustering and examine the dendrogram
# - experiment with hyperparameters and the distance function
# - look for more dependencies and password patterns
#
# +
import numpy as np
import re
from pylev import levenshtein
from sklearn.cluster import DBSCAN, KMeans
import matplotlib.pyplot as plt
# -
words_1M = []
with open("data/ignis-1M.txt", "r") as file:
for line in file:
words_1M.append(line.strip())
words_1K = []
with open("data/ignis-1K.txt", "r") as file:
for line in file:
words_1K.append(line.strip())
words = np.array(words_1M[:1000]).reshape((-1, 1))
# Introduce a distance-matrix:
import numpy as np
from pylev import levenshtein
X = np.zeros((words.shape[0], words.shape[0]))
for i,x in enumerate(words[:, 0]):
for j,y in enumerate(words[i:, 0]):
X[i, i + j] = levenshtein(x, y)
X[i + j, i] = X[i, i + j]
plt.imshow(X, cmap="Purples")
plt.show()
eps = 2.0
min_samples = 4
db = DBSCAN(eps=eps, metric="precomputed", min_samples=min_samples).fit(X)
labels = db.labels_
len(set(labels))
# +
clusters = {}
sizes = {}
for label in set(labels):
cluster = words[labels == label, 0]
sizes[label] = len(cluster)
clusters[label] = cluster
sizes_list = np.array(sorted([(x, y) for x,y in sizes.items()], key=lambda x: x[1], reverse=True))
# -
plt.title("Cluster sizes")
plt.bar(sizes_list[:, 0], sizes_list[:, 1])
plt.show()
# +
n_top_clusters_to_plot = 1
sizes_to_plot = sizes_list[n_top_clusters_to_plot:, ]
sizes_to_plot = sizes_to_plot[sizes_to_plot[:, 1] > min_samples]
print("{} clusters cover {} passwords from {}".format(
sizes_to_plot.shape[0],
sum(sizes_to_plot[:, 1]),
words.shape[0]
))
# -
for x in sizes_to_plot:
print(x[1], clusters[x[0]][:8])
from scipy.cluster import hierarchy
from scipy.spatial.distance import pdist
condensed_X = pdist(X)
linkage = hierarchy.linkage(condensed_X, method="complete")
linkage.shape
plt.figure(figsize=(16, 16))
dn = hierarchy.dendrogram(linkage)
plt.show()
from sklearn.cluster import AgglomerativeClustering
cluster = AgglomerativeClustering(n_clusters=5, affinity='precomputed', linkage='complete')
Y = cluster.fit_predict(X)
from collections import Counter
Counter(Y)
words[Y == 4][:10]
# +
# # !pip3 install -U strsimpy
# +
from strsimpy.weighted_levenshtein import WeightedLevenshtein
def insertion_cost(char):
return 1.0
def deletion_cost(char):
return 1.0
def substitution_cost(char_a, char_b):
if char_a == 't' and char_b == 'r':
return 0.5
return 1.0
weighted_levenshtein = WeightedLevenshtein(
substitution_cost_fn=substitution_cost,
insertion_cost_fn=insertion_cost,
deletion_cost_fn=deletion_cost)
# -
print(weighted_levenshtein.distance('Stting1', 'String1'))
print(weighted_levenshtein.distance('String1', 'Stting1'))
# ### Kmeans and embeddings
import gensim.downloader
list(gensim.downloader.info()['models'].keys())
word_embeddings = gensim.downloader.load("glove-wiki-gigaword-100")
# +
part_word_emb_names = []
part_word_emb_values = []
for word in words[:, 0]:
if word in word_embeddings:
part_word_emb_names.append(word)
part_word_emb_values.append(word_embeddings[word])
# -
len(words), len(part_word_emb_names)
part_word_emb_names[:25]
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
pca_words = pca.fit_transform(part_word_emb_values)
pca_words.shape
plt.scatter(pca_words[:, 0], pca_words[:, 1])
plt.title("621 Embeddings PCA")
plt.show()
embeddings_clusters = KMeans(n_clusters=3).fit_predict(part_word_emb_values)
Counter(embeddings_clusters)
for i in range(len(set(embeddings_clusters))):
plt.scatter(pca_words[embeddings_clusters == i, 0], pca_words[embeddings_clusters == i, 1], label=i)
plt.legend()
plt.title("621 Embeddings PCA")
plt.show()
for i in range(len(set(embeddings_clusters))):
print(i)
for word in np.array(part_word_emb_names)[embeddings_clusters == i][:5]:
print(word)
print("---")
| seminars/sem10/sem10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:bruges]
# language: python
# name: conda-env-bruges-py
# ---
# +
import numpy as np
# %matplotlib inline
# -
# ## Required
#
# - `bruges`
# - `welly`
# ## Coordinate transformation in `bruges`
#
# The idea is that you have UTM coordinates of three unique inline, crossline locations in a seismic survey (e.g. three cornerpoints). Once you have provided these, you can look up the (x,y) location of any (inline, xline) location, or vice versa.
import bruges
# Define the cornerpoints:
# +
# UTM coords of 3 unique inline, crossline locations.
corners_xy = np.array([[600938.125, 6073394.5],
[631226.3125, 6074241.0],
[630720.25, 6092358.5]])
# The inline, crossline locations you just provided.
corners_ix = np.array([[99, 104],
[99, 1316],
[824,1316]])
# -
# The `CoordTransform` class provides a function, which we'll call `transform`.
transform = bruges.transform.CoordTransform(corners_ix, corners_xy)
# +
transform([440, 763]) # Or, equivalently, transform.forward([440, 763])
# Should be [617167, 6082379]
# -
# We can also go back, from (x, y) to (inline, crossline).
transform.reverse([617167, 6082379])
# (If you *only* wanted the reverse function, you could also just have passed the coordinate arrays in to the instantiation in reverse order.)
# ## Dealing with the vertical axis: no velocity model
#
# To get the seismic sample belonging to a particular depth, we need a velocity model.
#
# If you already have an average velocity model in depth, then you can just look up the velocity and convert it directly.
#
# But let's assume you only have a sonic well log for velocity information...
# +
from welly import Well
w = Well.from_las("data/F03-04.las")
# -
w.data
dt = w.data['DT']
dt.plot()
dt.units
dt
# Correct for feet and microseconds.
vp = 0.3048 * 1e6 / dt
vp.start
vp.interpolate()
vp = vp.to_basis(start=0).extrapolate()
vp = vp.despike().smooth(20)
vp.plot()
vp
# Get Vp log in time:
import bruges
dt = 0.004 # seconds seismic sample interval
vp_t = bruges.transform.depth_to_time(vp, vmodel=vp, dt=dt, dz=vp.step, mode='linear')
vp_t.shape
# It's too short (must match seismic trace), so pad it:
vp_t = np.pad(vp_t, pad_width=[0, 1000-vp_t.size], mode='edge')
# Make a random trace and use time-based model to convert it to depth:
tr = np.random.randn(1000)
dz = 0.5 # Depth sample interval
tr_z = bruges.transform.time_to_depth(tr, vmodel=vp_t, dt=dt, dz=dz, mode="linear")
# Make a basis for the seismic in depth:
basis = np.linspace(0, tr_z.size*dz, tr_z.size+1)
basis
# This basis gives the depths in 0.5 m intervals. To get the seismic sample at a particluar depth, get it from tr_z, which is the seismic amplitude in depth (i.e. the same basis).
| notebooks_dev/Coordinate_transformation_rotated_survey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# Neuromorphic engineering I
# ## Lab 1: Automated Data Acquisition and Analysis
# Group number: 4.5
#
# Team member 1: <NAME>
#
# Team member 2: <NAME>
#
# Date: 30.09.2021
# -------------------------------------------------------------------------------------------------------------------
# You need to bring your own USB cable to connect the board with your own PC.
#
# The objectives of this lab are as follows:
#
# - To become acquainted with the experimental board setup.
#
# - To become acquainted with Python for data acquisition and manipulation.
#
# - To measure and characterize a NFET.
#
# The aim of this first lab is to familiarize everyone with the lab equipment and software.
# # 1. Getting started
# ## 1.1 Class chip documentation
#
# You can find the documentation for the classchip here:
# https://drive.google.com/drive/u/0/folders/1VBPKVfS9zwu_I2ExR1D0jU2eCSgleoQG
# ## 1.2 Python
# If you are new to Python, you can find a detailed tutorial at the link: https://docs.python.org/3/tutorial/ .
#
# In particular the concepts in _3. An Informal Introduction to Python_ and in _4. More Control FLow Tools_ will be useful for the exercises you will have to solve.
# ## 1.3 Report
# + [markdown] tags=[]
# You hand in the report (prelab + lab report + postlab) in both `.ipynb` and `.pdf` format as a group. Make sure that the markdowns are ran and the generated figures are shown.
#
# The deadline is the beginning of the next lab.
#
# There is no prelab for this week, but from the next week on you should finish the prelab before the lab starts.
# -
# ## 1.4. Virtual machine
# If you are a Linux user you can skip this step.
# If you are a Windows or Mac user, we have set-up an Ubuntu-based virtual machine with the necessary libraries to communicate with the board. To import the existent virtual machine, Windows users should install VirtualBox. Mac users can either install VirtualBox or Parallels. If you want to use a pre-configured VirtualBox virtual machine with all the tools pre-installed then follow the instructions in the below.
# To install VirtualBox, please follow these steps:
#
# **For Windows users**
#
# **Step 1.** Dowload VirtualBox
#
# You can download VirtualBox from the link: https://www.virtualbox.org/wiki/Downloads , VirtualBox 6.1.26 platform packages -> Windows hosts
#
# Execute the .exe file and follow the steps. In the window "Would you like to install this device software" select Install.
#
# Once the installation is complete, press Finish and launch the VirtualBox.
#
#
# **Step 2.** Download the Ubuntu 20.04 ISO file
#
# You can download the necessary files for the NE-I virtual machine from the link: https://ubuntu.com/. Click 'Download' option on the Ubuntu website, on the left side of 'Ubuntu Desktop', you can click the green '20.04 LTS' button to download Ubuntu 20.04 ISO file.
#
#
# **Step 3.** Import the file in VirtualBox and install Ubuntu
#
# Once you launch VirtualBox, the window Oracle VM VirtualBox Manager should appear on your screen. Then follow this link https://itsfoss.com/install-linux-in-virtualbox/ from 'Step 3: Install Linux using VirtualBox'.
#
#
# **For Mac users**
#
# **Step 1.** Dowload VirtualBox
#
# You can download VirtualBox from the link: https://www.virtualbox.org/wiki/Downloads , VirtualBox 6.1.26 platform packages -> Windows hosts
#
# Execute the .dmg file and follow the steps.
#
# Once the installation is complete, press Finish and launch the VirtualBox.
#
#
# **Step 2.** Download the Ubuntu 20.04 ISO file
#
# You can download the necessary files for the NE-I virtual machine from the link: https://ubuntu.com/. Click 'Download' option on the Ubuntu website, on the left side of 'Ubuntu Desktop', you can click the green '20.04 LTS' button to download Ubuntu 20.04 ISO file.
#
#
# **Step 3.** Import the file in VirtualBox and install Ubuntu
#
# Once you launch VirtualBox, the window Oracle VM VirtualBox Manager should appear on your screen. Then follow this link https://medium.com/tech-lounge/how-to-install-ubuntu-on-mac-using-virtualbox-3a26515aa869
# # 2. Python exercises
# ## 2.1 Jupyter Lab
# You can install Jupyter Lab or Jupyter Notebook according to this link:https://jupyter.org/install
# ## 2.2 Making plots
# * Plot a Sine curve from 0 to 2π with 20 points. (Hint: Do not forget to properly label the axis and add figure legends when necessary.)
import numpy as np
import matplotlib.pyplot as plt
max_x = 2 * np.pi
data_points = 20
x = np.arange(0, max_x, max_x/data_points)
sin_y = np.sin(x)
plt.plot(x, sin_y, 'b', label='sin(x)')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.grid()
plt.show()
# * Add a plot of a Cosine (Use the same range to the Sine plot, but set a different point marker and color.)
# +
cos_y = np.cos(x)
plt.plot(x, sin_y, 'b', label='sin(x)')
plt.plot(x, cos_y, 'r', label='cos(x)')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.grid()
plt.show()
# -
# * Make a plot of the equation $I_{ds} = I_0 e^{\frac{\kappa}{U_T} V_g}$ using the following parameters. Generate two plots, one with linear scaling and one with log scaling on the y axis. Put the right labels on both x axis and y axis. (Hint: use `plt.semilogy`)
Vg = np.arange(0, 1, 0.001)
I0 = 1e-13
k = 0.6
UT = 25e-3
Ids = I0 * np.exp(k/UT * Vg)
plt.plot(Vg, Ids, label='I_e*expt(k/U_T * V_g)')
plt.xlabel('V_g')
plt.ylabel('I_ds')
plt.legend()
plt.show()
# +
plt.semilogy(Vg, Ids, label='I_e*expt(k/U_T * V_g)')
plt.xlabel('V_g')
plt.ylabel('I_ds')
plt.legend()
plt.show()
# -
# ## 2.3 Saving and loading data
# You may want to work on the data for your report after the lab. Use `np.savetxt('data.csv', data, delimiter=',')` to save it to the working directory.
# define data
data = [Vg,Ids]
# save to csv file
np.savetxt('data.csv', data, delimiter=',')
# Check if the data saved is correct by loading it again using `np.loadtxt('data.csv', delimiter=',')` and plot.
# load from csv file
x, y = np.loadtxt('data.csv', delimiter=',')
# plot
plt.semilogy(x, y, label='I_e*expt(k/U_T * V_g)')
plt.xlabel('V_g')
plt.ylabel('I_ds')
plt.legend()
plt.show()
# ## 2.4 Fitting data with a line
# Sometimes you may need to find the relationship between data using regression. Try to extract the slope and intercept of the following given data using linear regression with `np.polyfit`
x = np.array([1, 3, 5, 7])
y = np.array([ 6, 3, 9, 5 ])
m, b = np.polyfit(x, y, 1)
# compare the orignial data points and the fitted line
plt.plot(x, y, 'o', label='original data')
plt.plot(x, m*x + b, 'r', label='fitted line')
plt.legend()
plt.show()
# -------------------------------------------------------------------------------------------------------------------
# # 3 Experiments
# From now on you will be using the real board!
# ## 3.0 How to install pyplane
# In order to communicate with the chip through Python you need to install _pyplane_ , a library that provides an easy interface to control the chip from jupyter notebook. The interface requires to use Ubuntu 20.04. You can follow different methods based on your machine's operating system.
# ### 3.0.1 Installing _pyplane_ using pip
# Open a command window and run `pip install pyplane`
# ### 3.0.2 Set up the USB connection of the teensy board in the Virtual machine
# **Windows users**
#
# 1. check teensy vendor and product id
#
# lsusb
#
# you should see something like "XXXX:YYYY" for teensy
#
# 2. add usb rules
#
# # cd /etc/udev/rules.d
#
# sudo gedit 10-my-usb.rules
#
# Add the information in this 10-my-usb.rules file and save it:
# ATTR{idVendor}=="XXXX", ATTR{idProduct}=="YYYY", MODE="0666", GROUP="dialout"
#
# 3. add user to group and change mode
#
# sudo usermod -a -G dialout $USER
#
# sudo chmod a+rw /dev/ttyACM0
#
# 4. restart the virtual machine
#
# **Mac Users**
#
# *Step 1* Settings --> Ports --> USB Add Teensyduino USB Serial
#
# *Step 2* Device --> Connect
#
# *Step 3* Change mode: sudo chmod a+rw /dev/ttyACM0
#
# *Step 4* Restart kernel in Jupyter noytebook
#
# *Step 5* Run the code again
# ## 3.1 Load the firmware (you can skip this step)
# You don't need to load the firmware,we have done it for you.
#
# To load the firmware, please follow these steps:
#
# Step 1. Dowload and install the Teensy Loader
#
# wget https://www.pjrc.com/teensy/teensy_linux64.tar.gz
#
# wget https://www.pjrc.com/teensy/00-teensy.rules
#
# sudo cp 00-teensy.rules /etc/udev/rules.d/
#
# tar -xvzf teensy_linux64.tar.gz
#
# ./teensy &
#
# You will see
# 
# Step 2. Load main.hex
#
# Connect the board to your computer by USB
# Open maim.hex file
# 
# Press button on Teensy to manually enter Program Mode
# 
# Program
# 
# Reboot
# 
# ## 3.2 Set up the communication between jupyter notebook and PCB
# Verify that the LEDs of your board are on as in the following picture
# 
# import the necessary library to communicate with the hardware
import sys
import pyplane
import numpy as np
import matplotlib.pyplot as plt
# create a Plane object and open the communication
if 'p' not in locals():
p = pyplane.Plane()
try:
p.open('/dev/ttyACM0')
except RuntimeError as e:
print(e)
# Make sure all these steps are executed correctly (the [*] in front of the line turns into a number)
# Send a reset signal to the board
reset_type = pyplane.ResetType(0)
p.reset(pyplane.ResetType.Soft)
# Was the LED flashing? How?
# > Yes, it blinked!
#NOTE: You must send this request events every time you do a reset operetion, otherwise the recieved data is wrong.
p.request_events(1)
# You could check firmware version, which should be 1.8.3.
p.get_firmware_version()
# See all the possible functions
dir(pyplane)
# ## 3.3 Basic function operation
# You can set and read voltage using function p.set_voltage and p.read_voltage.
#
# Very importantly, all voltage you set on this board must be between 0 and 1.8 V!
# Now set a voltage at AIN0
p.set_voltage(pyplane.DacChannel.AIN0, 0.62)
# Because of the quantization error of the DAC, you may want to see the actual value you have set using p.get_set_voltage
p.get_set_voltage(pyplane.DacChannel.AIN0, 0.62)
# Read the voltage at adc channel AOUT0
p.read_voltage(pyplane.AdcChannel.AOUT0)
# Read the voltage at adc channel GO22
p.read_current(pyplane.AdcChannel.GO22)
# ## 3.4 AER
# uses schemdraw, you may have to install it in order to run it on your PC
import schemdraw
import schemdraw.elements as elm
d = schemdraw.Drawing()
Q = d.add(elm.NFet, reverse=True)
d.add(elm.Dot, xy=Q.gate, lftlabel='gate=AIN0')
d.add(elm.Dot, xy=Q.drain, toplabel='drain=GO22')
d.add(elm.Dot, xy=Q.source, botlabel='source=GO20')
d.draw()
# Read the current of the drain of the transistor.
I_d = p.read_current(pyplane.AdcChannel.GO22)
print("The measured drain current is {} A".format(I_d))
# Do you think this current is reasonable? WHy?
# > Yes. I expect no current, since the source has not been set to any voltage. The small measured current is just noise.
# Now try to set voltage of this transistor by AER (Address Event Representation).
# Find the documentation "chip_architecture.pdf" for the classchip introduced in 1.1. See how to select signal communication on pages 11 and 21.
# Because you need to read the current and write the voltage of the NFET. You have to set demultiplexer by sending the configuration event:
# +
events = [pyplane.Coach.generate_aerc_event( \
pyplane.Coach.CurrentOutputSelect.SelectLine5, \
pyplane.Coach.VoltageOutputSelect.NoneSelected, \
pyplane.Coach.VoltageInputSelect.SelectLine2, \
pyplane.Coach.SynapseSelect.NoneSelected, 0)]
p.send_coach_events(events)
# -
# **Make sure the chip receives the event by a blink of LED1, if it's not the case, the chip is dead.**
# Now set source GO20 voltage using the function introduced above
p.set_voltage(pyplane.DacChannel.GO20, 0.0)
# Set drain GO22 voltage
p.set_voltage(pyplane.DacChannel.GO22, 1.8)
# Set trial gate AIN0 voltage (you can try different voltage between 0~1.8V to see different output current)
p.set_voltage(pyplane.DacChannel.AIN0, 1.0)
#
#
#
# Read drain GO22 current
I_d = p.read_current(pyplane.AdcChannel.GO22)
print("The measured drain current is {} A".format(I_d))
# Compare this current with the drain current measured above.
# Set trial gate AIN0 voltage 0 and see if the drain current is also zero. If not, why?
# > No. There is always some noise in the order of magnitude of e-7.
# Now you can try some challenging experiments!
# Sweep gate voltage between 0~1V and see how the output current change
import time
#Sweep gate voltage
Vg = np.arange(0.0, 1.0, 0.05)
# +
#Initialize current variables
currents = []
# -
#Set Vg = 0 and wait 0.5 second for it to settle
time.sleep(0.5) # delay 0.5 second
#Read leakage current Ids=Ids0
leakage = p.read_current(pyplane.AdcChannel.GO22)
#Read Ids at Vg sweep and wait for it to settle
for n in range(len(Vg)):
p.set_voltage(pyplane.DacChannel.AIN0,Vg[n])
current = p.read_current(pyplane.AdcChannel.GO22)
time.sleep(0.1) # delay 0.1 second
#Subtract leakage and shunt resistance from read current value
current -= leakage
currents.append(current)
#Plot in linear scale
plt.plot(Vg, currents)
plt.xlabel('Vg')
plt.ylabel('Ids')
plt.show()
# # 4. Clean up
# Well done! That's all for today.
#
# Remember you have to clean up in the end just as in a real lab!
# * Close you device and release memory by doing
del p
# * Save your changes
# * Download the files you need for the report to your own PC
# # 5. Postlab Questions
# 1. Why is there no pin for the bulk of NFET? What is it's voltage then?
# > The NFET bulk is hardwired to the GND. The voltage of the bulk is 0V (ignoring noise).
# 2. How precise are the measurements of voltage and current using DAC?
# > They cannot be too precise since we are dealing with very small voltages and currents, so the quantization error/noise of the DAC is not negligible.
# 3. Do you think building a "computer" whose inputs and outputs are analog voltage/current signals is a good idea? Why or why not?
# > At least not a traditional computer, no. Current computers expect digital signals where the quantization error and noise are negligible. In theory, a deterministic system can be built out of non-deterministic systems when the inaccuracies cancel each other out, like the brain does. But in practice, computers are simply not designed this way.
| neuromorphic_engineering_one/session1_automated_data_acquisition_and_analysis/2021_Lab1_DataAq_4.5_Jan_Hohenheim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ----
# <img src="../../files/refinitiv.png" width="20%" style="vertical-align: top;">
#
# # Data Library for Python
#
# ----
# ## Eikon Data API - News examples
# This notebook demonstrates how to retrieve News Headlines and News Stories from Eikon or Refinitiv Workspace.
#
# #### Learn more
# To learn more about the Data API just connect to the Refinitiv Developer Community. By [registering](https://developers.refinitiv.com/iam/register) and [login](https://developers.refinitiv.com/iam/login) to the Refinitiv Developer Community portal you will get free access to a number of learning materials like [Quick Start guides](https://developers.refinitiv.com/eikon-apis/eikon-data-api/quick-start), [Tutorials](https://developers.refinitiv.com/eikon-apis/eikon-data-api/learning), [Documentation](https://developers.refinitiv.com/eikon-apis/eikon-data-api/docs) and much more.
#
# #### About the "eikon" module of the Refinitiv Data Platform Library
# The "eikon" module of the Refinitiv Data Platform Library for Python embeds all functions of the classical Eikon Data API ("eikon" python library). This module works the same as the Eikon Data API and can be used by applications that need the best of the Eikon Data API while taking advantage of the latest features offered by the Refinitiv Data Platform Library for Python.
#
# #### Getting Help and Support
# If you have any questions regarding the API usage, please post them on the [Eikon Data API Q&A Forum](https://community.developers.thomsonreuters.com/spaces/92/index.html). The Refinitiv Developer Community will be happy to help.
#
# ## Import the library and connect to Eikon or Refinitiv Workspace
# +
import refinitiv.data.eikon as ek
from IPython.display import HTML
ek.set_app_key('YOUR APP KEY GOES HERE!')
# -
# ## Get News HeadLines
# #### From the default News repository (NewsWire)
ek.get_news_headlines(query = 'AAPL.O', count=10)
# #### From the other News repository (NewsRoom and WebNews)
ek.get_news_headlines(query = 'AAPL.O', repository='NewsRoom')
ek.get_news_headlines(query = 'AAPL.O', repository='WebNews')
ek.get_news_headlines(query = 'AAPL.O', repository='WebNews,NewsRoom,NewsWire')
# ## Get News Story
# #### Get the Story ID of the first headline for Apple
headlines = ek.get_news_headlines('AAPL.O',1)
story_id = headlines.iat[0,2]
story_id
# #### Get the News story for this Story ID
story = ek.get_news_story(story_id)
HTML(story)
| Examples/5-Eikon Data API/EX-5.03.01 - Eikon Data API - News.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# -
# <img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
#
# # Scaling Criteo: Download and Convert
#
# ## Criteo 1TB Click Logs dataset
#
# The [Criteo 1TB Click Logs dataset](https://ailab.criteo.com/download-criteo-1tb-click-logs-dataset/) is the largest public available dataset for recommender system. It contains ~1.3 TB of uncompressed click logs containing over four billion samples spanning 24 days. Each record contains 40 features: one label indicating a click or no click, 13 numerical figures, and 26 categorical features. The dataset is provided by CriteoLabs. A subset of 7 days was used in this [Kaggle Competition](https://www.kaggle.com/c/criteo-display-ad-challenge/overview). We will use the dataset as an example how to scale ETL, Training and Inference.
# First, we will download the data and extract it. We define the base directory for the dataset and the numbers of day. Criteo provides 24 days. We will use the last day as validation dataset and the remaining days as training.
#
# **Each day has a size of ~15GB compressed `.gz` and uncompressed ~XXXGB. You can define a smaller subset of days, if you like. Each day takes ~20-30min to download and extract it.**
# +
import os
from nvtabular.utils import download_file
download_criteo = True
BASE_DIR = os.environ.get("BASE_DIR", "/raid/data/criteo")
input_path = os.path.join(BASE_DIR, "crit_orig")
NUMBER_DAYS = os.environ.get("NUMBER_DAYS", 2)
# -
# We create the folder structure and download and extract the files. If the file already exist, it will be skipped.
# %%time
if download_criteo:
# Test if NUMBER_DAYS in valid range
if NUMBER_DAYS < 2 or NUMBER_DAYS > 23:
raise ValueError(
str(NUMBER_DAYS)
+ " is not supported. A minimum of 2 days are "
+ "required and a maximum of 24 (0-23 days) are available"
)
# Create BASE_DIR if not exists
if not os.path.exists(BASE_DIR):
os.makedirs(BASE_DIR)
# Create input dir if not exists
if not os.path.exists(input_path):
os.makedirs(input_path)
# Iterate over days
for i in range(0, NUMBER_DAYS):
file = os.path.join(input_path, "day_" + str(i) + ".gz")
# Download file, if there is no .gz, .csv or .parquet file
if not (
os.path.exists(file)
or os.path.exists(
file.replace(".gz", ".parquet").replace("crit_orig", "converted/criteo/")
)
or os.path.exists(file.replace(".gz", ""))
):
download_file(
"http://azuremlsampleexperiments.blob.core.windows.net/criteo/day_"
+ str(i)
+ ".gz",
file,
)
# The original dataset is in text format. We will convert the dataset into `.parquet` format. Parquet is a compressed, column-oriented file structure and requires less disk space.
# ### Conversion Script for Criteo Dataset (CSV-to-Parquet)
#
# __Step 1__: Import libraries
# +
import os
import glob
import numpy as np
from dask.distributed import Client
from dask_cuda import LocalCUDACluster
import nvtabular as nvt
from nvtabular.utils import device_mem_size, get_rmm_size
# -
# __Step 2__: Specify options
#
# Specify the input and output paths, unless the `INPUT_DATA_DIR` and `OUTPUT_DATA_DIR` environment variables are already set. For multi-GPU systems, check that the `CUDA_VISIBLE_DEVICES` environment variable includes all desired device IDs.
INPUT_PATH = os.environ.get("INPUT_DATA_DIR", input_path)
OUTPUT_PATH = os.environ.get("OUTPUT_DATA_DIR", os.path.join(BASE_DIR, "converted"))
CUDA_VISIBLE_DEVICES = os.environ.get("CUDA_VISIBLE_DEVICES", "0")
frac_size = 0.10
# __Step 3__: (Optionally) Start a Dask cluster
cluster = None # Connect to existing cluster if desired
if cluster is None:
cluster = LocalCUDACluster(
CUDA_VISIBLE_DEVICES=CUDA_VISIBLE_DEVICES,
rmm_pool_size=get_rmm_size(0.8 * device_mem_size()),
local_directory=os.path.join(OUTPUT_PATH, "dask-space"),
)
client = Client(cluster)
# __Step 5__: Convert original data to an NVTabular Dataset
# +
# Specify column names
cont_names = ["I" + str(x) for x in range(1, 14)]
cat_names = ["C" + str(x) for x in range(1, 27)]
cols = ["label"] + cont_names + cat_names
# Specify column dtypes. Note that "hex" means that
# the values will be hexadecimal strings that should
# be converted to int32
dtypes = {}
dtypes["label"] = np.int32
for x in cont_names:
dtypes[x] = np.int32
for x in cat_names:
dtypes[x] = "hex"
# Create an NVTabular Dataset from a CSV-file glob
file_list = glob.glob(os.path.join(INPUT_PATH, "day_*"))
dataset = nvt.Dataset(
file_list,
engine="csv",
names=cols,
part_mem_fraction=frac_size,
sep="\t",
dtypes=dtypes,
client=client,
)
# -
# **__Step 6__**: Write Dataset to Parquet
dataset.to_parquet(
os.path.join(OUTPUT_PATH, "criteo"),
preserve_files=True,
)
# You can delete the original criteo files as they require a lot of disk space.
| docs/source/examples/scaling-criteo/01-Download-Convert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook demonstrates how to use Quest to download imagery from a Web Map Tile Service (WMTS).
#
# In addition to quest the following packages need to be installed to use this notebook:
# * holoviews
# * geoviews
# * param
# * paramnb
# * xarray
#
# The can be installed with the following command:
# ```
# conda install -c conda-forge -c pyviz/label/dev holoviews geoviews param paramnb xarray
# ```
# +
import param
import quest
import geoviews as gv
import holoviews as hv
import xarray as xr
from cartopy import crs as ccrs
from holoviews.streams import BoxEdit
from parambokeh import Widgets
hv.extension('bokeh')
# -
quest_service = 'svc://wmts:seamless_imagery'
tile_service_options = quest.api.get_download_options(quest_service, fmt='param')[quest_service]
tile_service_options.params()['bbox'].precedence = -1 # hide bbox input
Widgets(tile_service_options)
tiles = gv.WMTS(tile_service_options.url).options(width=950, height=600, global_extent=True)
boxes = gv.Polygons([]).options(fill_alpha=0.4, line_width=2)
box_stream = BoxEdit(source=boxes, num_objects=1)
tiles * boxes
if box_stream.element:
data = box_stream.data
bbox = [data['x0'][0], data['y0'][0], data['x1'][0], data['y1'][0]]
else:
bbox = [-72.43925984610391, 45.8471360126193, -68.81252476472281, 47.856449699679516]
# bbox = [-83.28613682601174, 24.206059963486737, -81.93264005405752, 30.251169660148314]
# bbox = [-81.82408198648336, 25.227665888548458, -80.86355086047537, 31.548730116206755]
print(bbox)
tile_service_options.bbox = bbox
arr = quest.api.get_data(
service_uri=quest_service,
search_filters=None,
download_options=tile_service_options,
collection_name='examples',
use_cache=False,
as_open_datasets=True,
)[0]
image = gv.RGB((arr.x, arr.y, arr[0].values,
arr[1].values, arr[2].values),
vdims=['R', 'G', 'B']).options(width=950, height=600, alpha=0.7)
gv.tile_sources.Wikipedia * image
| reviews/Panel/WMTS_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="color:#303030;font-family:'arial blACK', sans-serif,monospace; text-align: center; padding: 50px 0; vertical-align:middle;" > <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/Lightbulb.png?raw=true" style=" background:linear-gradient(to right,#FDC86E,#fbb144);border-radius:10px;width:150px;text-align:left; margin-left:10%" /> <span style="position:relative; bottom:70px; margin-left:5%;font-size:170%;"> Interchanging Structured Data </span> </div>
# ## <span style="color:#fbb144;"> Keywords: </span>
# ```JavaScript Object Notation (JSON)```, ```Structure Data```, ```Metadata```, ```HTML```,```CSS```,```JS```
# # I. Introduction
# <br>
# <div style="width:100%; background:linear-gradient(to right,#FDC86E,#fbb144);font-family:'arial black',monospace; text-align: center; padding: 7px 0; border-radius: 5px 50px;margin-top:-15px" > </div>
#
# ## <div style="color:#fbb144;"> 1. Background </div>
#
# Previously, we’ve seen how to exchange plain text messages between a web-
# based client and a Python server using the WebSockets (WS) protocol. Although
# this approach usually suffices for simple applications, it can quickly become limiting when working with compound data (e.g. handling records). In this lesson
# we will see how to overcome this limitation by working with structured data, in
# particular, using JSON (JavaScript Object Notation) as a representation format
# for data interchange. JSON is a lightweight dictionary-like data representation
# approach, widely used due to the fact that it is easily readable and generated
# by both humans and machines.
#
# <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/B.Graphical_User_Interface_IMG/b003/JSON.png?raw=true" alt="JSON" border="0">
# ## <div style="color:#fbb144;"> 2. Objectives</div>
# * Further explore the bi-directional communication between Python and a
# web browser in a client-server approach
# * Become familiar with the JSON (JavaScript Object Notation) format for
# data representation
# * Understand how to effectively manipulate and interchange structured data
# between different software components
# ## <div style="color:#fbb144;"> 3. Materials (optional) </div>
# * Anaconda Python 2.7
# * Twisted Matrix networking engine for Python
# * Google Chrome
# # II. Experimental
# <br>
# <div style="width:100%; background:linear-gradient(to right,#FDC86E,#fbb144);font-family:'arial black',monospace; text-align: center; padding: 7px 0; border-radius: 5px 50px;margin-top:-15px" > </div>
#
# ## <div style="color:#fbb144;"> 1. Body Mass Index Calculator </div>
#
# We will illustrate structured data representation using JSON through a simple
# example implementation of a Body Mass Index (BMI) calculator. The weight
# and height are introduced through a web-based interface and upon pressing the
# submission button, a JSON (dictionary-like) structure is created and sent to
# the Python server as a string. On the server side, the message is parsed from a
# string to a JSON object, and the collected data used to compute the BMI and
# determine the category. Finally, the results are formatted as a JSON and sent
# to the web-based client. The steps to test this experiment are as follows:
#
# 1. Download the resources available at:
# <a><center> https://github.com/PIA-Group/ScientIST-notebooks/tree/master/Example_Files/Interchanging_Structured_Data </center> </a>
#
#
#
# 2. Open the script BMI.py (following cell) in the Spyder IDE and study it
#
# Example code of a WebSockets server in Python that receives structured data in JSON (i.e. dictionary-like) format from a web-based client:
# +
from twisted.internet import protocol, reactor
from txws import WebSocketFactory
import json
class WS(protocol.Protocol):
def connectionMade(self):
# Executed when the client successfully connects to the server.
print("CONNECTED")
def dataReceived(self, req):
# Executed when the data is received from the client.
print "< " + req
data = json.loads(req)
result = self.calculateBMI(float(data['weight']), float(data['height']))
response = json.dumps(result)
print "> " + response
self.transport.write(response)
def connectionLost(self, reason):
# Executed when the connection to the client is lost.
print("DISCONNECTED")
def calculateBMI(self, weight, height):
result = dict()
result['BMI'] = weight/((height/100)**2)
if result['BMI'] < 18.5:
result['category'] = 'Underweight'
elif result['BMI'] >= 18.5 and result['BMI'] < 24.9:
result['category'] = 'Normal'
elif result['BMI'] >= 25 and result['BMI'] < 29.9:
result['category'] = 'Overweight'
else:
result['category'] = 'Obesity'
return result
class WSFactory(protocol.Factory):
def buildProtocol(self, addr):
return WS()
if __name__=='__main__':
ip_addr, port = "127.0.0.1", 9000
print("LISTENING AT %s:%s"%(ip_addr, port))
connector = reactor.listenTCP(port, WebSocketFactory(WSFactory()))
reactor.run()
# -
# 3. Open the HTML file BMI.html (following cell) in the Spyder IDE (or another HTML editor of your choice) and study it
#
# Example HTML/CSS/JS code to implement a web-based client that sends structured data in JSON format to the server:
#
# ```html
#
# <html>
# <script language="javascript" type="text/javascript" src="jquery.js"></script>
# <script language="javascript" type="text/javascript" src="jquery.flot.js"></script>
# <script type="text/javascript">
# var ws = new WebSocket("ws://localhost:9000/");
#
# function submit() {
# data = {'weight': $("#weight").val(),
# 'height': $("#height").val()};
# ws.send(JSON.stringify(data));
# }
#
# ws.onmessage = function (e) {
# response = JSON.parse(e.data);
# $("#BMI").val(response['BMI']);
# $("#category").val(response['category']);
# }
# </script>
# <body>
# <h1>BMI Calculator</h1>
# <p>Weigth (Kg):</p>
# <input id="weight" type="text"/>
# <p>Height (cm):</p>
# <input id="height" type="text"/>
# <input type="button" value="Send"/ onclick="submit()">
# <hr/>
# <p>BMI:</p>
# <input id="BMI" type="text"/>
# <p>Category:</p>
# <input id="category" type="text"/>
# </body>
# </html>
# ```
# 4. Run your Python script; a line should appear in the console showing the message LISTENING AT 127.0.0.1:9000, indicating that your server is ready to receive connections
#
# 5. Open the HTML file using Google Chrome (other browsers should work also); on the Python console you should see the message CONNECTED
#
# 6. Through the web page, whenever you type in the weight and height, and press the SEND button afterwards, the data will be transmitted to your Python server where the BMI calculation results will be performed and returned back to the web-based client
# ## <div style="color:#fbb144;"> 2. Weight Logger </div>
#
# In addition to facilitating the interchange of structured data in-between software
# pieces (as in the previous exercise were the Python server and the web-based
# client), the JSON format is quite convenient for data persistency also. In this
# exercise we will explore the use of the JSON format to store and retrieve data
# from a file using as a case study a weight log. The steps to test this experiment
# are as follows:
#
# 1. Download the resources available at:
#
# <a><center> https://github.com/PIA-Group/ScientIST-notebooks/tree/master/Example_Files/Interchanging_Structured_Data </center> </a>
#
# 2. Open the script WeightLogger.py (following cell) in the Spyder IDE and study it
#
# Python code base for the weight logger server:
# +
from twisted.internet import protocol, reactor
from txws import WebSocketFactory
import json
import time
class WS(protocol.Protocol):
def connectionMade(self):
# Executed when the client successfully connects to the server.
print("CONNECTED")
self.loadLog()
self.sendLog()
print self.log
def dataReceived(self, req):
# Executed when the data is received from the client.
print "< " + req
data = json.loads(req)
self.extendLog(float(data['weight']))
self.sendLog()
def connectionLost(self, reason):
# Executed when the connection to the client is lost.
print("DISCONNECTED")
def loadLog(self):
with open('WeightLogger.txt') as file:
self.log = json.load(file)
file.close()
def sendLog(self):
response = json.dumps(self.log)
print "> " + response
self.transport.write(response)
def extendLog(self, weight):
date = time.strftime("%d-%m-%y %Hh:%Mm:%Ss")
entry = [date, weight]
self.log['weight'].append(entry)
self.saveLog()
self.sendLog()
def saveLog(self):
with open('WeightLogger.txt', 'w') as file:
json.dump(self.log, file)
file.close()
class WSFactory(protocol.Factory):
def buildProtocol(self, addr):
return WS()
if __name__=='__main__':
ip_addr, port = "127.0.0.1", 9000
print("LISTENING AT %s:%s"%(ip_addr, port))
connector = reactor.listenTCP(port, WebSocketFactory(WSFactory()))
reactor.run()
# -
# 3. Open the HTML file WeightLogger.html (following cell) in the Spyder IDE (or another HTML editor of your choice) and study it
#
# HTML/CSS/JS document for the weight logger client:
# ```html
# <html>
# <script language="javascript" type="text/javascript" src="jquery.js"></script>
# <script language="javascript" type="text/javascript" src="jquery.flot.js"></script>
# <script type="text/javascript">
# var ws = new WebSocket("ws://localhost:9000/");
#
# function submit() {
# data = {'weight': $("#weight").val()};
# ws.send(JSON.stringify(data));
# }
#
# ws.onmessage = function (e) {
# response = JSON.parse(e.data);
# log = response['weight'];
# content = ""
# for (var i = 0; i<log.length; i++) {
# content+=log[i][0];
# content+=" | ";
# content+=log[i][1];
# content+="<br/>";
# }
# $("#log").html(content);
# }
# </script>
# <body>
# <h1>Weight Logger</h1>
# <p>Weigth (Kg):</p>
# <input id="weight" type="text"/>
# <input type="button" value="Add"/ onclick="submit()">
# <hr/>
# <p id="log"></p>
# </body>
# </html>
# ```
# <html>
# <script language="javascript" type="text/javascript" src="jquery.js"></script>
# <script language="javascript" type="text/javascript" src="jquery.flot.js"></script>
# <script type="text/javascript">
# var ws = new WebSocket("ws://localhost:9000/");
#
# function submit() {
# data = {'weight': $("#weight").val()};
# ws.send(JSON.stringify(data));
# }
#
# ws.onmessage = function (e) {
# response = JSON.parse(e.data);
# log = response['weight'];
# content = ""
# for (var i = 0; i<log.length; i++) {
# content+=log[i][0];
# content+=" | ";
# content+=log[i][1];
# content+="<br/>";
# }
# $("#log").html(content);
# }
# </script>
# <body>
# <h1>Weight Logger</h1>
# <p>Weigth (Kg):</p>
# <input id="weight" type="text"/>
# <input type="button" value="Add"/ onclick="submit()">
# <hr/>
# <p id="log"></p>
# </body>
# </html>
#
# 4. Run your Python script; a line should appear in the console showing the message LISTENING AT 127.0.0.1:9000, indicating that your server is ready to receive connections
#
# 5. Open the HTML file using Google Chrome (other browsers should work also); on the Python console you should see the message CONNECTED
#
# 6. Through the web page, whenever you type in the weight and height, and press the ADD button afterwards, the data will be transmitted to your Python server where it will be combined with the current date and time, and added to measurement history log
#
# 7. Open the file WeightLogger.txt in a text editor of your liking and inspect its content, to confirm that the entries you have added are stored on file
#
# 8. If you close the HTML and open it again, you’ll see the previous entries of your log listed on the web-based client
# ## <div style="color:#fbb144;"> 3. Reacting to Different Events </div>
#
# Until now we have seen application examples that were focused on performing
# specific operations and used JSON for data representation only. However, JSON
# structures are general purpose and can contain any data that you decide to store
# on them; the semantics can be solely decided by you, i.e. whether that data
# is to be used as input values to a calculation performed by a function (as in
# Section II.1), to be stored on file (as in Section II.2), or for another purpose.
# In this experiment we will see how to use JSON to enable our Python server to
# distinguish between and respond to different events performed on the web-based
# client:
#
# 1. Download the resources available at:
# <a><center> https://github.com/PIA-Group/ScientIST-notebooks/tree/master/Example_Files/Interchanging_Structured_Data </center> </a>
#
#
# 2. Open the script EventHandler.py (following cell) in the Spyder IDE andstudy it
#
# Python code base for a server that handles events ACTION1, ACTION2 and ACTION3 sent by a web-based client:
# +
from twisted.internet import protocol, reactor
from txws import WebSocketFactory
import json
class WS(protocol.Protocol):
def connectionMade(self):
# Executed when the client successfully connects to the server.
print("CONNECTED")
def dataReceived(self, req):
# Executed when the data is received from the client.
print "< " + req
data = json.loads(req)
if data['event'] == "ACTION1":
self.handleAction1(data['args'])
elif data['event'] == "ACTION2":
self.handleAction2(data['args'])
elif data['event'] == "ACTION3":
self.handleAction3(data['args'])
def connectionLost(self, reason):
# Executed when the connection to the client is lost.
print("DISCONNECTED")
def handleAction1(self, args):
self.transport.write("PYTHON EXECUTED ACTION 1")
def handleAction2(self, args):
self.transport.write("PYTHON EXECUTED ACTION 2")
def handleAction3(self, args):
self.transport.write("PYTHON EXECUTED ACTION 3")
class WSFactory(protocol.Factory):
def buildProtocol(self, addr):
return WS()
if __name__=='__main__':
ip_addr, port = "127.0.0.1", 9000
print("LISTENING AT %s:%s"%(ip_addr, port))
connector = reactor.listenTCP(port, WebSocketFactory(WSFactory()))
reactor.run()
# -
#
# 3. Open the HTML file EventHandler.html (following cell) in the Spyder IDE (or another HTML editor of your choice) and study it
#
# HTML/CSS/JS document for the event generation client:
# ```html
# <html>
# <script language="javascript" type="text/javascript" src="jquery.js"></script>
# <script language="javascript" type="text/javascript" src="jquery.flot.js"></script>
# <script type="text/javascript">
# var ws = new WebSocket("ws://localhost:9000/");
#
# function button1() {
# data = {'event': 'ACTION1', 'args': '1'};
# ws.send(JSON.stringify(data));
# }
#
# function button2() {
# data = {'event': 'ACTION2', 'args': '2'};
# ws.send(JSON.stringify(data));
# }
#
# function button3() {
# data = {'event': 'ACTION3', 'args': '1'};
# ws.send(JSON.stringify(data));
# }
#
# ws.onmessage = function (e) {
# $("#log").html(e.data)
# }
# </script>
# <body>
# <h1>Event Handler Demo</h1>
# <input type="button" value="Button 1"/ onclick="button1()">
# <input type="button" value="Button 2"/ onclick="button2()">
# <input type="button" value="Button 3"/ onclick="button3()">
# <hr/>
# <p id="log"></p>
# </body>
# </html>
#
# ```
# <html>
# <script language="javascript" type="text/javascript" src="jquery.js"></script>
# <script language="javascript" type="text/javascript" src="jquery.flot.js"></script>
# <script type="text/javascript">
# var ws = new WebSocket("ws://localhost:9000/");
#
# function button1() {
# data = {'event': 'ACTION1', 'args': '1'};
# ws.send(JSON.stringify(data));
# }
#
# function button2() {
# data = {'event': 'ACTION2', 'args': '2'};
# ws.send(JSON.stringify(data));
# }
#
# function button3() {
# data = {'event': 'ACTION3', 'args': '1'};
# ws.send(JSON.stringify(data));
# }
#
# ws.onmessage = function (e) {
# $("#log").html(e.data)
# }
# </script>
# <body>
# <h1>Event Handler Demo</h1>
# <input type="button" value="Button 1"/ onclick="button1()">
# <input type="button" value="Button 2"/ onclick="button2()">
# <input type="button" value="Button 3"/ onclick="button3()">
# <hr/>
# <p id="log"></p>
# </body>
# </html>
# 4. Run your Python script; a line should appear in the console showing the message LISTENING AT 127.0.0.1:9000, indicating that your server is ready to receive connections
#
# 5. Open the HTML file using Google Chrome (other browsers should work also); on the Python console you should see the message CONNECTED
#
# 6. Through the web page, whenever you press one of the buttons, a JSON structure will be transmitted to your Python server including a description of the command and an argument; depending on the command, the Python server will execute a different function accordingly and send a message to the web-based client
#
# # III. Explore
# <br>
# <div style="width:100%; background:linear-gradient(to right,#FDC86E,#fbb144);font-family:'arial black',monospace; text-align: center; padding: 7px 0; border-radius: 5px 50px;margin-top:-15px" > </div>
#
# ## <div style="color:#fbb144;"> 1. Quiz </div>
#
# 1. Expand the Python server and web-based client developed in Section II.1. to enable the calculation of the Body Mass Index (BMI) in imperial units (in addition to the metric units already supported).
#
# 2. Modify the web-based client developed in Section II.1. to show the category text in a different colour depending on the determined category, namely, blue if Underweight, green if Normal, yellow if Overweight and red if Obesity.
#
# 3. Expand the Python server and web-based client developed in Section II.2. to, in addition to the weight, log also the height, heart rate and blood pressure (the latter two typed in manually by the user).
#
# 4. Using what you have learned from Section II.3., modify the Python server and web-based client developed in Section II.2. to enable clearing the log.
#
# 5. Expand the code developed in the previous question to enable the deletion of an entry (manually) specified by the user via the web-based client.
# <div style="height:100px; background:white;border-radius:10px;text-align:center">
#
# <a> <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/IT.png?raw=true" alt="it" style=" bottom: 0; width:250px;
# display: inline;
# left: 250px;
# position: absolute;"/> </a>
# <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/IST.png?raw=true"
# alt="alternate text"
# style="position: relative; width:250px; float: left;
# position: absolute;
# display: inline;
# bottom: 0;
# right: 100;"/>
# </div>
# <div style="width: 100%; ">
# <div style="background:linear-gradient(to right,#FDC86E,#fbb144);color:white;font-family:'arial', monospace; text-align: center; padding: 50px 0; border-radius:10px; height:10px; width:100%; float:left " >
# <span style="font-size:12px;position:relative; top:-25px"> Please provide us your feedback <span style="font-size:14px;position:relative;COLOR:WHITE"> <a href="https://forms.gle/C8TdLQUAS9r8BNJM8">here</a>.</span></span>
# <br>
# <span style="font-size:17px;position:relative; top:-20px"> Suggestions are welcome! </span>
# </div>
# ```Contributors: Prof. <NAME>; <NAME>```
| B.Graphical_User_Interface/B003 Interchanging Structured Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
test_file = open("test.txt", 'a+')
for i in range(5):
test_file.write("hello world" + '\n')
test_file.close()
test_file = open("test.txt", 'a+')
test_file.seek(0)
content = test_file.read()
print(content)
content = content.upper()
test_file.write(content)
test_file.close()
test_file = open("test.txt", 'r')
content_lines = test_file.readlines()
print(content_lines)
content_lines[2] = content_lines[2].capitalize()
test_file.close()
test_file = open('test.txt', 'w')
for line in content_lines:
test_file.write(line)
test_file.close()
with open('test.txt', 'a') as test_file:
test_file.write("I love python")
| Test file .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Implementing CDFs
#
# Copyright 2019 <NAME>
#
# BSD 3-clause license: https://opensource.org/licenses/BSD-3-Clause
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
sns.set_style('white')
import matplotlib.pyplot as plt
# +
import inspect
def psource(obj):
"""Prints the source code for a given object.
obj: function or method object
"""
print(inspect.getsource(obj))
# -
# ### Constructor
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/11).
#
# The `Cdf` class inherits from `pd.Series`. The `__init__` method is essentially unchanged, but it includes a workaround for what I think is bad behavior.
# +
from empiricaldist import Cdf
psource(Cdf.__init__)
# -
# You can create an empty `Cdf` and then add elements.
#
# Here's a `Cdf` that representat a four-sided die.
d4 = Cdf()
d4[1] = 1
d4[2] = 2
d4[3] = 3
d4[4] = 4
d4
# In a normalized `Cdf`, the last probability is 1.
#
# `normalize` makes that true. The return value is the total probability before normalizing.
psource(Cdf.normalize)
d4.normalize()
# Now the Cdf is normalized.
d4
# ### Properties
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/2).
#
# In a `Cdf` the index contains the quantities (`qs`) and the values contain the probabilities (`ps`).
#
# These attributes are available as properties that return arrays (same semantics as the Pandas `values` property)
d4.qs
d4.ps
# ### Sharing
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/12).
#
# Because `Cdf` is a `Series` you can initialize it with any type `Series.__init__` can handle.
#
# Here's an example with a dictionary.
d = dict(a=1, b=2, c=3)
cdf = Cdf(d)
cdf.normalize()
cdf
# Here's an example with two lists.
qs = [1,2,3,4]
ps = [0.25, 0.5, 0.75, 1.0]
d4 = Cdf(ps, index=qs)
d4
# You can copy a `Cdf` like this.
d4_copy = Cdf(d4)
d4_copy
# However, you have to be careful about sharing. In this example, the copies share the arrays:
d4.index is d4_copy.index
d4.ps is d4_copy.ps
# You can avoid sharing with `copy=True`
d4_copy = Cdf(d4, copy=True)
d4_copy
d4.index is d4_copy.index
d4.ps is d4_copy.ps
# Or by calling `copy` explicitly.
d4_copy = d4.copy()
d4_copy
d4.index is d4_copy.index
d4.ps is d4_copy.ps
# ### Displaying CDFs
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/13).
#
# `Cdf` provides `_repr_html_`, so it looks good when displayed in a notebook.
psource(Cdf._repr_html_)
# `Cdf` provides `plot`, which plots the Cdf as a line.
psource(Cdf.plot)
def decorate_dice(title):
"""Labels the axes.
title: string
"""
plt.xlabel('Outcome')
plt.ylabel('CDF')
plt.title(title)
d4.plot()
decorate_dice('One die')
# `Cdf` also provides `step`, which plots the Cdf as a step function.
psource(Cdf.step)
d4.step()
decorate_dice('One die')
# ### Make Cdf from sequence
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/14).
#
#
# The following function makes a `Cdf` object from a sequence of values.
psource(Cdf.from_seq)
cdf = Cdf.from_seq(list('allen'))
cdf
cdf = Cdf.from_seq(np.array([1, 2, 2, 3, 5]))
cdf
# ### Selection
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/15).
#
# `Cdf` inherits [] from Series, so you can look up a quantile and get its cumulative probability.
d4[1]
d4[4]
# `Cdf` objects are mutable, but in general the result is not a valid Cdf.
d4[5] = 1.25
d4
d4.normalize()
d4
# ### Evaluating CDFs
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/16).
#
# Evaluating a `Cdf` forward maps from a quantity to its cumulative probability.
d6 = Cdf.from_seq([1,2,3,4,5,6])
d6.forward(3)
# `forward` interpolates, so it works for quantities that are not in the distribution.
d6.forward(3.5)
d6.forward(0)
d6.forward(7)
# `__call__` is a synonym for `forward`, so you can call the `Cdf` like a function (which it is).
d6(1.5)
# `forward` can take an array of quantities, too.
def decorate_cdf(title):
"""Labels the axes.
title: string
"""
plt.xlabel('Quantity')
plt.ylabel('CDF')
plt.title(title)
qs = np.linspace(0, 7)
ps = d6(qs)
plt.plot(qs, ps)
decorate_cdf('Forward evaluation')
# `Cdf` also provides `inverse`, which computes the inverse `Cdf`:
d6.inverse(0.5)
# `quantile` is a synonym for `inverse`
d6.quantile(0.5)
# `inverse` and `quantile` work with arrays
ps = np.linspace(0, 1)
qs = d6.quantile(ps)
plt.plot(qs, ps)
decorate_cdf('Inverse evaluation')
# These functions provide a simple way to make a Q-Q plot.
#
# Here are two samples from the same distribution.
# +
cdf1 = Cdf.from_seq(np.random.normal(size=100))
cdf2 = Cdf.from_seq(np.random.normal(size=100))
cdf1.plot()
cdf2.plot()
decorate_cdf('Two random samples')
# -
# Here's how we compute the Q-Q plot.
def qq_plot(cdf1, cdf2):
"""Compute results for a Q-Q plot.
Evaluates the inverse Cdfs for a
range of cumulative probabilities.
:param cdf1: Cdf
:param cdf2: Cdf
:return: tuple of arrays
"""
ps = np.linspace(0, 1)
q1 = cdf1.quantile(ps)
q2 = cdf2.quantile(ps)
return q1, q2
# The result is near the identity line, which suggests that the samples are from the same distribution.
q1, q2 = qq_plot(cdf1, cdf2)
plt.plot(q1, q2)
plt.xlabel('Quantity 1')
plt.ylabel('Quantity 2')
plt.title('Q-Q plot');
# Here's how we compute a P-P plot
def pp_plot(cdf1, cdf2):
"""Compute results for a P-P plot.
Evaluates the Cdfs for all quantities in either Cdf.
:param cdf1: Cdf
:param cdf2: Cdf
:return: tuple of arrays
"""
qs = cdf1.index.union(cdf2)
p1 = cdf1(qs)
p2 = cdf2(qs)
return p1, p2
# And here's what it looks like.
p1, p2 = pp_plot(cdf1, cdf2)
plt.plot(p1, p2)
plt.xlabel('Cdf 1')
plt.ylabel('Cdf 2')
plt.title('P-P plot');
# ### Statistics
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/17).
#
# `Cdf` overrides the statistics methods to compute `mean`, `median`, etc.
psource(Cdf.mean)
d6.mean()
psource(Cdf.var)
d6.var()
psource(Cdf.std)
d6.std()
# ### Sampling
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/18).
#
# `choice` chooses a random values from the Cdf, following the API of `np.random.choice`
psource(Cdf.choice)
d6.choice(size=10)
# `sample` chooses a random values from the `Cdf`, following the API of `pd.Series.sample`
psource(Cdf.sample)
d6.sample(n=10, replace=True)
# ### Arithmetic
#
# For comments or questions about this section, see [this issue](https://github.com/AllenDowney/EmpyricalDistributions/issues/9).
#
# `Cdf` provides `add_dist`, which computes the distribution of the sum.
#
# The implementation uses outer products to compute the convolution of the two distributions.
psource(Cdf.add_dist)
psource(Cdf.make_same)
# Here's the distribution of the sum of two dice.
# +
d6 = Cdf.from_seq([1,2,3,4,5,6])
twice = d6.add_dist(d6)
twice
# -
twice.step()
decorate_dice('Two dice')
twice.mean()
# To add a constant to a distribution, you could construct a deterministic `Pmf`
const = Cdf.from_seq([1])
d6.add_dist(const)
# But `add_dist` also handles constants as a special case:
d6.add_dist(1)
# Other arithmetic operations are also implemented
d4 = Cdf.from_seq([1,2,3,4])
d6.sub_dist(d4)
d4.mul_dist(d4)
d4.div_dist(d4)
# ### Comparison operators
#
# `Pmf` implements comparison operators that return probabilities.
#
# You can compare a `Pmf` to a scalar:
d6.lt_dist(3)
d4.ge_dist(2)
# Or compare `Pmf` objects:
d4.gt_dist(d6)
d6.le_dist(d4)
d4.eq_dist(d6)
# Interestingly, this way of comparing distributions is [nontransitive]().
A = Cdf.from_seq([2, 2, 4, 4, 9, 9])
B = Cdf.from_seq([1, 1, 6, 6, 8, 8])
C = Cdf.from_seq([3, 3, 5, 5, 7, 7])
A.gt_dist(B)
B.gt_dist(C)
C.gt_dist(A)
| empiricaldist/cdf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jhw
# language: python
# name: jhw
# ---
# ## Fashion-MNIST 데이터셋 다운로드
# +
from tensorflow.keras.datasets.fashion_mnist import load_data
# Fashion-MNIST 데이터를 다운받습니다.
(x_train, y_train), (x_test, y_test) = load_data()
print(x_train.shape, x_test.shape)
# -
# # 데이터 그려보기
# +
import matplotlib.pyplot as plt
import numpy as np
np.random.seed(777)
# Fashion-MNIST의 레이블에 해당하는 품목입니다.
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
sample_size = 9
# 0 ~ 59999의 범위에서 무작위로 3개의 정수를 뽑습니다.
random_idx = np.random.randint(60000, size=sample_size)
plt.figure(figsize = (5, 5))
for i, idx in enumerate(random_idx):
plt.subplot(3, 3, i+1)
plt.xticks([])
plt.yticks([])
plt.imshow(x_train[idx], cmap = 'gray') # 20210218 오탈자
plt.xlabel(class_names[y_train[idx]]) # 20210218 오탈자
plt.show()
# -
# ## 전처리 및 검증 데이터셋 만들기
# +
# 값의 범위를 0 ~ 1로 만들어줍니다.
x_train = x_train / 255
x_test = x_test / 255
from tensorflow.keras.utils import to_categorical
# 각 데이터의 레이블을 범주형 형태로 변경합니다.
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# 검증 데이터 세트를 만듭니다.
from sklearn.model_selection import train_test_split
# 훈련/테스트 데이터를 0.7/0.3의 비율로 분리합니다.
x_train, x_val, y_train, y_val = train_test_split(x_train, y_train,
test_size = 0.3, random_state = 777)
# -
# ## 1. 첫 번째 모델 구성하기
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten
first_model = Sequential()
# 입력 데이터의 형태를 꼭 명시해야 합니다.
first_model.add(Flatten(input_shape = (28, 28))) # (28, 28) -> .(28 * 28)
first_model.add(Dense(64, activation = 'relu')) # 64개의 출력을 가지는 Dense 층
first_model.add(Dense(32, activation = 'relu')) # 32개의 출력을 가지는 Dense 층
first_model.add(Dense(10, activation = 'softmax')) # 10개의 출력을 가지는 신경망
# -
# ## 학습 과정 설정 및 학습하기
# +
first_model.compile(optimizer='adam', # 옵티마이저 : Adam
loss = 'categorical_crossentropy', # 손실 함수 : categorical_crossentropy
metrics=['acc']) # 모니터링 할 평가지표 : acc
first_history = first_model.fit(x_train, y_train,
epochs = 30,
batch_size = 128,
validation_data = (x_val, y_val))
# -
# ## 2. 두 번째 모델 구성하기
# +
second_model = Sequential()
# 입력 데이터의 형태를 꼭 명시해야 합니다.
second_model.add(Flatten(input_shape = (28, 28))) # (28, 28) -> .(28 * 28)
second_model.add(Dense(128, activation = 'relu')) # 128개의 출력을 가지는 Dense 층을 추가합니다.
second_model.add(Dense(64, activation = 'relu')) # 64개의 출력을 가지는 Dense 층
second_model.add(Dense(32, activation = 'relu')) # 32개의 출력을 가지는 Dense 층
second_model.add(Dense(10, activation = 'softmax')) # 10개의 출력을 가지는 신경망
second_model.compile(optimizer='adam', # 옵티마이저: Adam
loss = 'categorical_crossentropy', # 손실 함수: categorical_crossentropy
metrics=['acc']) # 모니터링 할 평가지표: acc(정확도)
second_history = second_model.fit(x_train, y_train,
epochs = 30,
batch_size = 128,
validation_data = (x_val, y_val))
# -
# # 두 모델의 학습 과정 그려보기
# +
import numpy as np
import matplotlib.pyplot as plt
def draw_loss_acc(history_1, history_2, epochs):
his_dict_1 = history_1.history
his_dict_2 = history_2.history
keys = list(his_dict_1.keys())
epochs = range(1, epochs)
fig = plt.figure(figsize = (10, 10))
ax = fig.add_subplot(1, 1, 1)
# axis 선과 ax의 축 레이블을 제거합니다.
ax.spines['top'].set_color('none')
ax.spines['bottom'].set_color('none')
ax.spines['left'].set_color('none')
ax.spines['right'].set_color('none')
ax.tick_params(labelcolor='w', top=False, bottom=False, left=False, right=False)
for i in range(len(his_dict_1)):
temp_ax = fig.add_subplot(2, 2, i + 1)
temp = keys[i%2]
val_temp = keys[(i + 2)%2 + 2]
temp_history = his_dict_1 if i < 2 else his_dict_2
temp_ax.plot(epochs, temp_history[temp][1:], color = 'blue', label = 'train_' + temp)
temp_ax.plot(epochs, temp_history[val_temp][1:], color = 'orange', label = val_temp)
if(i == 1 or i == 3):
start, end = temp_ax.get_ylim()
temp_ax.yaxis.set_ticks(np.arange(np.round(start, 2), end, 0.01))
temp_ax.legend()
ax.set_ylabel('loss', size = 20)
ax.set_xlabel('Epochs', size = 20)
plt.tight_layout()
plt.show()
draw_loss_acc(first_history, second_history, 30)
| code/ch4/Fashion-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 基本程序设计
# - 一切代码输入,请使用英文输入法
print('hello word')
print('hello word')
print 'hello'
print
# ## 编写一个简单的程序
# - 圆公式面积: area = radius \* radius \* 3.1415
radius = 1.0
area = radius * radius * 3.14 # 将后半部分的结果赋值给变量area
# 变量一定要有初始值!!!
# radius: 变量.area: 变量!
# int 类型
print(area)
radius=1
area=radius*radius*3.1415
print(area)
# ### 在Python里面不需要定义数据的类型
# ## 控制台的读取与输入
# - input 输入进去的是字符串
# - eval
radius = input('请输入半径') # input得到的结果是字符串类型
radius = float(radius)
area = radius * radius * 3.14
print('面积为:',area)
x=eval(input('x'))
res=eval(input('输入表达式:'))
print(res)
input('请输入姓名:')
# - 在jupyter用shift + tab 键可以跳出解释文档
# ## 变量命名的规范
# - 由字母、数字、下划线构成
# - 不能以数字开头 \*
# - 标识符不能是关键词(实际上是可以强制改变的,但是对于代码规范而言是极其不适合)
# - 可以是任意长度
# - 驼峰式命名
age = 100
nianling = 100
# ## 变量、赋值语句和赋值表达式
# - 变量: 通俗理解为可以变化的量
# - x = 2 \* x + 1 在数学中是一个方程,而在语言中它是一个表达式
# - test = test + 1 \* 变量在赋值之前必须有值
# ## 同时赋值
# var1, var2,var3... = exp1,exp2,exp3...
# ## 定义常量
# - 常量:表示一种定值标识符,适合于多次使用的场景。比如PI
# - 注意:在其他低级语言中如果定义了常量,那么,该常量是不可以被改变的,但是在Python中一切皆对象,常量也是可以被改变的
# ## 数值数据类型和运算符
# - 在Python中有两种数值类型(int 和 float)适用于加减乘除、模、幂次
# <img src = "../Photo/01.jpg"></img>
17.521*x**3+15.212*x/27.1
5//2
# ## 运算符 /、//、**
# ## 运算符 %
# ## EP:
# - 25/4 多少,如果要将其转变为整数该怎么改写
# - 输入一个数字判断是奇数还是偶数
# - 进阶: 输入一个秒数,写一个程序将其转换成分和秒:例如500秒等于8分20秒
# - 进阶: 如果今天是星期六,那么10天以后是星期几? 提示:每个星期的第0天是星期天
25//4
if('x%2=0','偶数','奇数')
if('3%2=0',"偶数","奇数")
if('3%2=0',print('偶数','奇数'))
if(200%2)==0
print("偶数")
else
print("奇数")
x=x/60
# ## 科学计数法
# - 1.234e+2
# - 1.234e-2
# ## 计算表达式和运算优先级
# <img src = "../Photo/02.png"></img>
# <img src = "../Photo/03.png"></img>
# ## 增强型赋值运算
# <img src = "../Photo/04.png"></img>
# ## 类型转换
# - float -> int
# - 四舍五入 round
# ## EP:
# - 如果一个年营业税为0.06%,那么对于197.55e+2的年收入,需要交税为多少?(结果保留2为小数)
# - 必须使用科学计数法
# # Project
# - 用Python写一个贷款计算器程序:输入的是月供(monthlyPayment) 输出的是总还款数(totalpayment)
# 
# # Homework
# - 1
# <img src="../Photo/06.png"></img>
celsius=eval(input('输入摄氏温度'))
fahrenheit=(9/5)*celsius+32
print(fahrenheit)
# - 2
# <img src="../Photo/07.png"></img>
radius=eval(input('输入半径'))
length=eval(input('输入高'))
area=eval('radius*radius*3.14')
volume=eval('area*length')
print('圆柱体底面积',area,'体积为',volume)
# - 3
# <img src="../Photo/08.png"></img>
feet=eval(input('输入英尺数:'))
meters=eval('feet*0.305')
print(meters,'米')
# - 4
# <img src="../Photo/10.png"></img>
M=eval(input('输入以千克计的水量'))
initial=eval(input('初始温度'))
final=eval(input('最终温度'))
Q=eval('M*(final-initial)*4184')
print(Q)
# - 5
# <img src="../Photo/11.png"></img>
balance=eval(input('输入差额'))
interestrate=eval(input('输入年利率'))
interest=eval('balance*(interestrate/1200)')
print(interest)
# - 6
# <img src="../Photo/12.png"></img>
v0=eval(input('输入初始速度'))
v1=eval(input('输入末速度'))
t=eval(input('输入时间'))
a=eval('(v1-v0)/t')
print(a)
# - 7 进阶
# <img src="../Photo/13.png"></img>
month=1
saving=100
money=saving*(1+0.00417)
while True:
month=month+1
saving=money+100
money=saving*(1+0.00417)
if month==6:
print("账户数目为",money)
break
# - 8 进阶
# <img src="../Photo/14.png"></img>
a,b = eval(input('>>'))
print(a,b)
print(type(a),type(b))
a = eval(input('>>'))
print(a)
number=eval(input("输入0到1000之间整数"))
one=number//100
two=(number-one*100)//10
three=number%10
sum=one+two+three
print(sum)
| 7.16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#
# <div>
# <br>
# <center><strong><h1>Higher Order Tutorial on Deep Learning!</h1></strong></center>
# <center><strong><h3>Yam Peleg</h3></strong></center>
# <div>
# ------
# <div>
# <center><img src="imgs/keras_logo_humans.png" width="30%"/>
# <h1>www.github.com/ypeleg/ExpertDL</h1></center>
# <div>
# ------
# ## 1. *How to tune in*?
#
# If you wanted to listen to someone speaks three hours straight about deep learning, You could have done so by the comfort of your house.
#
# But you are here! Physically!
#
# So...
#
# This tutorial is extremely hands-on! You are strongly encouraged to play with it yourself!
#
# ### Options:
#
#
# ### $\varepsilon$. Run the notebooks locally
# - `git clone https://github.com/ypeleg/ExpertDL`
#
#
# - You might think that the goal of this tutorial is for you to play around with Deep Learning. This is wrong.
#
# - **The Rreal Goal of the Tutorial is To give you the Flexibility to use this In your own domain!**
#
# Therefore, this is by far the best option if you can get it working!
#
# ------
#
#
# ### a. Play with the _notebooks_ dynamically (on Google Colab)
#
# - Anyone can use the [colab.research.google.com/notebook](https://colab.research.google.com/notebook) website (by [clicking](XXX) on the icon bellow) to run the notebook in her/his web-browser. You can then play with it as long as you like!
# - For this tutorial:
# [](https://colab.research.google.com/github/ypeleg/ExpertDL)
# ------
#
# ### b. Play with the _notebooks_ dynamically (on MyBinder)
# [](http://mybinder.org/v2/gh/github/ypeleg/ExpertDL)
#
# Anyone can use the [mybinder.org](http://mybinder.org/) website (by [clicking](http://mybinder.org/v2/gh/github/ypeleg/ExpertDL) on the icon above) to run the notebook in her/his web-browser.
# You can then play with it as long as you like, for instance by modifying the values or experimenting with the code.
#
# ### c. View the _notebooks_ statically. (if all else failed..)
# - Either directly in GitHub: [ypeleg/ExpertDL](https://github.com/ypeleg/ExpertDL);
# - Or on nbviewer: [notebooks](http://nbviewer.jupyter.org/github/ypeleg/ExpertDL/).
# ---
# ## What do I mean High Order?
#
# In short [1], one can treat recent advancements of the field of deep learning as an increment of order (complexity-wise) where the components we use now in DL research were the whole experiments not long ago.
#
# Example: GANs involve training a neural networks on top of the output from another neural network. This can be viewed as a network of networks.
#
# Example: Some Reinforcement Learning algorithms (Mostly A3C) involves using a network for predicting the future reward of a state and using another network that based of that predicts the optimal action. Again a network of networks.
#
#
# In this tutorial we assume that we allready have deep learning networks ready for us as of the shelf tools and we use them to construct more complex algorithms.
#
#
#
# [1]. Poking this with me opens the pandora box..
# We might cover this is in great detail at the end of the tutorial. Depends on time.
# # Outline at a glance
#
# - **Part I**: **Introduction**
#
# - Intro Keras
# - Functional API
#
# - Reinforcement Learning
# - Intro
# - Bandit
# - Q learning
# - Policy Gradients
#
# - Generative Adversarial Networks
# - Intro
# - DCGAN
# - CGAN
# - WGAN
#
# - Embeddings
#
# - Advanced Natural Language Processing
# - Transformers
# - Elmo
# - Bert
# ## One More thing..
#
#
# <img style ="width:70%;" src="images/matplotlib.jpg"/>
#
#
# You are probably famillier with this.. so..
#
# ### The tachles.py file
#
# In this tutorial many of the irrelevant details are hidden in a special file called "tachles.py".
# Simply go:
import tachles
# ---
# # Requirements
# This tutorial requires the following packages:
#
# - Python version 2.7.11 Or Python version 3.5
# - Other versions of Python should be fine as well.
# - but.. *who knows*? :P
#
# - `numpy` version 1.10 or later: http://www.numpy.org/
# - `scipy` version 0.16 or later: http://www.scipy.org/
# - `matplotlib` version 1.4 or later: http://matplotlib.org/
# - `pandas` version 0.16 or later: http://pandas.pydata.org
# - `scikit-learn` version 0.15 or later: http://scikit-learn.org
# - `keras` version 2.0 or later: http://keras.io
# - `tensorflow` version 1.0 or later: https://www.tensorflow.org
# - `ipython`/`jupyter` version 4.0 or later, with notebook support
#
# (Optional but recommended):
#
# - `pyyaml`
# - `hdf5` and `h5py` (required if you use model saving/loading functions in keras)
# - **NVIDIA cuDNN** if you have NVIDIA GPUs on your machines.
# [https://developer.nvidia.com/rdp/cudnn-download]()
#
# The easiest way to get (most) these is to use an all-in-one installer such as [Anaconda](http://www.continuum.io/downloads) from Continuum. These are available for multiple architectures.
# ---
# ### Python Version
# I'm currently running this tutorial with **Python 3** on **Anaconda**
# !python --version
# ### Configure Keras with tensorflow
#
# 1) Create the `keras.json` (if it does not exist):
#
# ```shell
# touch $HOME/.keras/keras.json
# ```
#
# 2) Copy the following content into the file:
#
# ```
# {
# "epsilon": 1e-07,
# "backend": "tensorflow",
# "floatx": "float32",
# "image_data_format": "channels_last"
# }
# ```
# !cat ~/.keras/keras.json
# ---
# # Test if everything is up&running
# ## 1. Check import
import numpy as np
import scipy as sp
import pandas as pd
import matplotlib.pyplot as plt
import sklearn
import keras
# ## 2. Check installeded Versions
# +
import numpy
print('numpy:', numpy.__version__)
import scipy
print('scipy:', scipy.__version__)
import matplotlib
print('matplotlib:', matplotlib.__version__)
import IPython
print('iPython:', IPython.__version__)
import sklearn
print('scikit-learn:', sklearn.__version__)
# +
import keras
print('keras: ', keras.__version__)
# optional
import theano
print('Theano: ', theano.__version__)
import tensorflow as tf
print('Tensorflow: ', tf.__version__)
# -
# <br>
# <h1 style="text-align: center;">If everything worked till down here, you're ready to start!</h1>
# ---
#
| 0.0.0 Preamble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SiegfriedZhen/ptt-analysis/blob/master/ptt_etl_201910.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1jFgmZze7g1y" colab_type="text"
# # ptt的網路分析(network analysis)
# #### 近期天下的專題[輿論戰爭](https://www.cw.com.tw/article/article.action?id=5093610&fbclid=IwAR3NXtzdUBrUYBPPepMVzRXwY2tGZj6B84QzjWyPaIgjBL8QnSQjSOU6EeI#_=_),點出在八卦或政黑有些操作的跡象,其中有些網路分析的技術,這裡嘗試重現相關分析。
#
# + [markdown] id="NMW6sbvh7g10" colab_type="text"
# ### 1. import 網路分析常用的package [networkx](https://networkx.github.io/)
#
# ---
# 可直接用jwliny在github的[ptt-web-crawler](https://github.com/jwlin/ptt-web-crawler),不用自己寫爬蟲,安裝完後直接下指令(超佛!
# python -m PttWebCrawler -b PublicServan -i 100 200
# python crawler.py -b 看板名稱 -i 起始索引 結束索引 (設為-1為最後一頁, ps. 原作者聲稱是「設為負數則以倒數第幾頁計算」,但只有-1 會是有效的負數參數,另外如果index如果超過頁面,會有json格式錯誤發生)
# + id="tBPfDKGc7g11" colab_type="code" colab={}
import networkx as nx
import matplotlib.pyplot as plt
import pandas as pd
import operator
from collections import Counter
from IPython.display import Image
import numpy as np
import os
# + id="gbP6o78U098N" colab_type="code" colab={}
# 讀取 Google drive
from google.colab import drive
drive.mount('/content/drive')
# + id="fFpAZ39H7g15" colab_type="code" colab={}
path = '/content/drive/My Drive/Colab Notebooks/'
file_1 = path + 'HatePolitics-0-4060.json'
file_2 = path + 'HatePolitics-1500-3989.json'
# + [markdown] id="JocYs7JL7g17" colab_type="text"
# ### 2. modified json ending
# #### ptt-web-crawler 有一個小bug,當爬的文章頁數超過,結尾會多一個逗號,由於只有read file by binary mode 支援負指標搜尋,所以會使用,ab+ 二進制讀寫模式,且寫入只能在文件末端,執行時用truncate後寫入。
# #### correct ending
# #### b'erid": "wei121"}]}]}'
# #### false ending
# #### b'serid": "L1ON"}]},]}'
# + id="IDbG87-h7g17" colab_type="code" colab={}
def check_json_format(file):
with open(file, 'ab+') as f:
# read as binary is for reverse file search
print('check trivial comma in EOF before revesing')
# * 2 -- end of stream; offset is usually negative
f.seek(-6, 2)
if f.read(200) == b'}]},]}':
print('revising error endings')
f.seek(-3, 2)
f.truncate()
f.write(b']}')
print('check trivial comma in EOF after revising')
f.seek(-20, 2)
print(f.read(200))
# + id="UDEWKhoVPWtS" colab_type="code" outputId="f7e9183f-aa25-4806-a736-70941a91902f" colab={"base_uri": "https://localhost:8080/", "height": 118}
check_json_format(file_1)
check_json_format(file_2)
# + id="2tOAEMkj7g2A" colab_type="code" colab={}
ptt_df_1 = pd.read_json(file_1, encoding='utf8')
ptt_df_2 = pd.read_json(file_2, encoding='utf8')
# + [markdown] id="VXQljIfV7g2C" colab_type="text"
# ### 3. 來看看一篇文章的json長怎樣吧!
# #### (1)大量的DataFrame效能很差, 目前看到有兩個解法
# #### a. 先把大小設定好,才不會每次append都重新申請空間(但不太適用本狀況, 因為要觀察留言的互動,那就要先把文章 x 留言數
# #### b. 把每一篇文章的DataFrame存在一個list,最後用pd.concat 批次處理(這次使用的方法)
#
# #### package typo... message_conut
# + id="o9KEW4OX7g2C" colab_type="code" colab={}
def ptt_json_to_df(ptt_df):
art_df = []
invalid_lst = []
for x in ptt_df.itertuples():
#some inconsistent page failed!
if x.articles == {'error': 'invalid url'}:
print('invalid record')
print(x.Index)
invalid_lst.append(x)
continue
art = pd.DataFrame([x.articles])
##如果發文沒有人推或作者是None,直接濾除
if art.messages[0] == []:
invalid_lst.append(x)
continue
elif art.author[0] is None:
print('invalid author')
#print(x.articles)
invalid_lst.append(x)
continue
art.author = art.author.replace(r'\([^)]*\)', '',regex=True).values[0].replace(' ', '')
#fixing package typo of columns
art.rename({'message_conut':'message_count'}, axis=1, inplace=True)
reply_aggr = pd.DataFrame(art.message_count.values[0], index=[0])
art.drop(['message_count'], axis=1, inplace=True)
art = pd.concat([art, reply_aggr], axis=1)
art_df.append(art)
return art_df
# + id="mfIZk6m4RJJe" colab_type="code" colab={}
art_df_1 = ptt_json_to_df(ptt_df_1)
art_df_2 = ptt_json_to_df(ptt_df_2)
# + id="uHlfNFlb7g2F" colab_type="code" cellView="code" colab={}
art_df_1 = pd.concat(art_df_1)
art_df_2 = pd.concat(art_df_2)
# + id="MxKFaX3GePn_" colab_type="code" colab={}
art_df = art_df_1[~art_df_1.article_id.isin(art_df_2.article_id)].append(art_df_2)
# + id="i21bi1WneNXX" colab_type="code" colab={}
del art_df_1, art_df_2
# + id="w34noJVjdmpo" colab_type="code" colab={}
art_df.date = pd.to_datetime(art_df.date, format='%a %b %d %H:%M:%S %Y', errors='coerce')
art_df['re_flag'] = art_df.article_title.str.contains('Re')
# + [markdown] id="QOKTpiJUQVH1" colab_type="text"
# ### 4. 輸出發文檔案,作為Tableau分析使用
# + id="9VM-HbZBPyzo" colab_type="code" colab={}
art_df[['article_id', 'article_title', 'author', 'board', 'date', 'ip', 'all',
'boo', 'count', 'neutral', 'push', 're_flag']].to_excel(path + 'hate_pol_node_dtl.xlsx', index=False)
# + id="t7NwTfr27g2H" colab_type="code" colab={}
art_cnt = art_df.groupby('author', as_index=False)['article_id'].count()
author_attr = art_df.groupby('author', as_index=False)['all', 'boo', 'count', 'neutral', 'push'].sum()
author_attr = author_attr.merge(art_cnt, how='left', on='author').rename({'article_id':'art_cnt'}, axis=1)
author_attr.rename({'author':'Id'}, axis=1, inplace=True)
# + id="clIGUVzf7g2N" colab_type="code" colab={}
author_attr.to_excel(path + 'hate_pol_node.xlsx', index=False)
# + id="5MftPYvx7g2L" colab_type="code" outputId="fe19cc07-76ca-440e-864a-61700c8b9fca" colab={"base_uri": "https://localhost:8080/", "height": 195}
author_attr.head()
# + id="KM181bvL7g2O" colab_type="code" colab={}
#check date
art_df.sort_values('date')[-30:].date
# + id="ediGn59H7g2S" colab_type="code" outputId="59d32844-b0cb-46a7-aa6a-fa56a927c29b" colab={"base_uri": "https://localhost:8080/", "height": 34}
'蒐集政黑板資料自 {} to {}'.format(art_df.date.min(), art_df.date.max())
# + [markdown] id="_IJgnRtM7g2U" colab_type="text"
# ### 4. 參考資料:天下對於ptt網軍的分析
# https://www.cw.com.tw/article/article.action?id=5093610
# ---
# 關於mark2165的質疑,現在卻成為政黑的板主,也是很有趣的現象,也有可能是報導的時間差,要再看最近發文~
#
# ###### (1)版規公告
# https://www.ptt.cc/bbs/HatePolitics/M.1555306691.A.ED4.html
# ###### (2) 參選公告
# https://www.ptt.cc/bbs/HatePolitics/M.1551694135.A.CFA.html
# ###### (3) 根據ptt 鄉民百科:是政黑的板主之一
# https://pttpedia.fandom.com/zh/wiki/HatePolitics%E6%9D%BF_(%E6%94%BF%E9%BB%91%E6%9D%BF)
# ###### (4) 2019/04/02上任公告
# https://www.ptt.cc/bbs/L_SecretGard/M.1554186459.A.9AA.html
# + [markdown] id="d8g5k2Rg7g2U" colab_type="text"
# ### 5. top 10 by 推噓加總
# ---
# #### mark2165位居第一,第二名則是ptt名人KingKingCold,但加總還是差了快一倍
# + id="_W1pWYzd7g2V" colab_type="code" outputId="73178403-4d84-4fee-f411-4dfa30212894" colab={"base_uri": "https://localhost:8080/", "height": 343}
author_attr.sort_values('count', ascending=False)[:10]
# + id="0_7D97UK7g2X" colab_type="code" colab={}
art_df.reset_index(drop=True, inplace=True)
# error records
# some no article.....
#[問卦]把蔣公銅像裝扮成薩諾斯會被吉嗎 invalid time formate
# + id="NzLZYYUe7g2Z" colab_type="code" colab={}
del ptt_df
# + id="xfTdmJ447g2e" colab_type="code" colab={}
#todo
#find some id mapping table
#https://pttpedia.fandom.com/zh/wiki/%E5%88%86%E9%A1%9E:PTT%E5%90%8D%E4%BA%BA
#crawl ptt celebrity
# + [markdown] id="-tZXczck7g2g" colab_type="text"
# ### 5. checking area
# + id="AAzFqtZW7g2g" colab_type="code" colab={}
#'{} 筆無效文章'.format(len(invalid_lst))
# + id="5cG8_zji7g2j" colab_type="code" colab={}
## 八卦板版規
# ※ 八卦板務請到 GossipPicket 檢舉板詢問
# ※ a.張貼問卦請注意,充實文章內容、是否有專板,本板並非萬能問板。
# ※ b.一天只能張貼 "兩則" 問卦,自刪及被刪也算兩篇之內,
# ※ 超貼者將被水桶,請注意!
# ※ c.本看板嚴格禁止政治問卦,發文問卦前請先仔細閱讀相關板規。
# ※ d.未滿30繁體中文字水桶3個月,嚴重者以鬧板論,請注意!
# ※ (↑看完提醒請刪除ctrl + y)
# + id="4l7pTcPM7g2k" colab_type="code" colab={}
# some special character id would become NaN
#art yesyesyesyes (@_@O)
# rex520368 (b@N9)
#rainbowsheep (@V@?)
# + [markdown] id="VkqGp33n7g2m" colab_type="text"
# todo: ip parse ref
# #### https://www.ptt.cc/bbs/Gossiping/M.1559471533.A.6AD.html
# + [markdown] id="qyZ-GKbW7g2n" colab_type="text"
# ### 3. 整理資料:推文作為source,發文作為target,主要是覺得推文作為主動方,觀察是否有大量互相推文的現象
# + id="K3kPyv217g2q" colab_type="code" outputId="7a5e0fa6-ed33-46d3-aaed-75a1ffaad662" colab={"base_uri": "https://localhost:8080/", "height": 218}
reply_df = []
for x in art_df.itertuples():
if x.Index % 10000 == 1:
print(x.Index)
tmp_reply = pd.DataFrame(x.messages)
#push_ip_ = reply.push_ipdatetime.str.split(' ', n=1, expand=True)
tmp_reply['target'] = x.author
tmp_ip_dt = tmp_reply.push_ipdatetime.copy()
tmp_reply['reply_datetime'] = str(x.date.year) + '/' + np.where(tmp_ip_dt.str.len() < 20, tmp_ip_dt, tmp_ip_dt.str[-11:])
##以發文日期作為推文的估計值,因為push_ipdatetime的ip 跟 datetime放在一起,有些只有datetime 沒有ip,好像跟RE 有關....
##欄位重新命名,push_userid 改為 source
tmp_reply.rename({'push_userid':'source'}, axis=1, inplace=True)
reply_df.append(tmp_reply)
# + id="pcw5k35g7g2s" colab_type="code" colab={}
# check global vars
# + id="gbOcOByX7g2u" colab_type="code" colab={}
del art_df
reply_df = pd.concat(reply_df)
# + id="_BCZpzbr7g2x" colab_type="code" outputId="ce6ffd70-6991-428f-fe62-31a038881b99" colab={"base_uri": "https://localhost:8080/", "height": 195}
reply_df.head()
# + id="tJ3ngPjA7g2z" colab_type="code" colab={}
reply_df.reply_datetime = pd.to_datetime(reply_df.reply_datetime, format='%Y/%m/%d %H:%M', errors='coerce')
reply_df['ip'] = np.where(reply_df.push_ipdatetime.str.len() > 20, reply_df.push_ipdatetime.str[:-12], None)
# + id="27TuguNS7g20" colab_type="code" colab={}
#reply_df['ip'] = np.where(reply_df.ipstr.len() > 20, reply_df.str[:-12], None)
# + id="ZbxJLWrD7g22" colab_type="code" outputId="e07e0f21-49a9-40f1-be91-20b106da93bf" colab={"base_uri": "https://localhost:8080/", "height": 168}
reply_df.dtypes
# + id="JvW2eLZd7g24" colab_type="code" colab={}
reply_df.drop(['push_content', 'push_ipdatetime'], axis=1, inplace=True)
# + id="F-tIxGP-Pkvf" colab_type="code" colab={}
reply_df.push_tag.replace({'推':'push', '噓':'boo', '→':'neutral'}, inplace=True)
# + id="qnfSs3FK7g25" colab_type="code" colab={}
#sample code for multi-edges for GraphDB
#reply_df['push'] = np.where(reply_df.push_tag == '推',1,0)
#reply_df['neutral'] = np.where(reply_df.push_tag == '→',1,0)
#reply_df['boo'] = np.where(reply_df.push_tag == '噓',1,0)
#reply_df.loc[reply_df.push_tag == '推']['push'] = 1
#['推', '→', '噓']
# + id="aaV7Bit07g27" colab_type="code" colab={}
#for multi-edge
#revsing the title to more general category/proj, hate_pol_edge
reply_df.to_csv(path + 'hate_pol_edge.csv', index=False)
# + id="wSD8BNDqXd8_" colab_type="code" colab={}
#reply_df = pd.read_csv(path + 'hate_pol_edge.csv')
# + id="JVhP3xXdXnAt" colab_type="code" colab={}
reply_df['reply_date'] = reply_df.reply_datetime.astype(str).str[:10]
reply_df['reply_mn'] = reply_df.reply_datetime.astype(str).str[:7]
# + id="lwcsDP_2YbYY" colab_type="code" colab={}
reply_aggr_df = reply_df.groupby(['push_tag', 'source', 'target', 'reply_date']).count()['reply_mn']
# + id="vOH14V5MeH4J" colab_type="code" colab={}
reply_aggr_df = pd.DataFrame(reply_aggr_df).reset_index()
reply_aggr_df.columns = ['push_tag', 'source', 'target', 'reply_date', 'cnt']
# + id="HPc3yVr8kRWQ" colab_type="code" outputId="7633f6d7-95c5-4c84-caba-ef6866612268" colab={"base_uri": "https://localhost:8080/", "height": 34}
reply_aggr_df.shape
# + id="fUYQN_Xrcfq1" colab_type="code" colab={}
reply_aggr_df.to_csv(path + 'hate_pol_edge_aggr.csv')
# + id="2K2H-XK5Yj6m" colab_type="code" colab={}
# reply_df['push'] = np.where(reply_df.push_tag == 'push',1,0)
# reply_df['neutral'] = np.where(reply_df.push_tag == 'neutral',1,0)
# reply_df['boo'] = np.where(reply_df.push_tag == 'boo',1,0)
#reply_df[reply_df.target == 'mark2165'].groupby(['source', 'target']).count()
# + [markdown] id="RvkEkLw97g3S" colab_type="text"
# ## 5. 把DataFrame轉為有向圖
# + id="SgIvDqzfRFgf" colab_type="code" outputId="99f93c7d-846e-4084-8988-8e2517338c60" colab={"base_uri": "https://localhost:8080/", "height": 204}
reply_df.head()
# + id="kpHbi3ku7g3S" colab_type="code" colab={}
G = nx.from_pandas_edgelist(reply_df, source='source', target='target', create_using=nx.DiGraph())
# + [markdown] id="iB2uuBzS7g3T" colab_type="text"
# ### 5-1. 看一下degree(in_degree + out_degree)前10名吧!
# #### out-degree 外度數: 某個id留言給不同 id的數量
# #### in-degree 輸入度: 某個id「被留言」的不同id數量
# #### 翻譯名稱參考:http://terms.naer.edu.tw/detail/2378473/
# + id="3JacN9xK7g3T" colab_type="code" outputId="aebcacf3-6525-48df-9fc1-7345a2038983" colab={"base_uri": "https://localhost:8080/", "height": 527}
#設定top幾
#degree不高 但是 推文高, 代表有高度集中性
rnk = 30
deg_g = dict(G.degree)
top_deg_node = [ x[0] for x in sorted(deg_g.items(), key=operator.itemgetter(1), reverse=True)[:rnk]]
sorted(deg_g.items(), key=operator.itemgetter(1), reverse=True)[:rnk]
# + [markdown] id="Hgx32QKO7g3W" colab_type="text"
# ### 5-2. 看一下pagerank前10名吧!
# + id="RHaVIZbi7g3W" colab_type="code" outputId="6f55215d-4b8c-48f3-800c-ef580438a930" colab={"base_uri": "https://localhost:8080/", "height": 527}
pg_rnk_g = nx.pagerank(G)
top_pg_rnk_node = [ x[0] for x in sorted(pg_rnk_g.items(), key=operator.itemgetter(1), reverse=True)[:rnk]]
sorted(pg_rnk_g.items(), key=operator.itemgetter(1), reverse=True)[:rnk]
# + id="v24SHBHc7g3X" colab_type="code" outputId="132a3c75-1e14-4503-9a81-0fc2f250ef59" colab={"base_uri": "https://localhost:8080/", "height": 527}
sorted(pg_rnk_g.items(), key=operator.itemgetter(1), reverse=True)[:rnk]
# + id="RLybknam7g3Z" colab_type="code" colab={}
#nx.write_gexf(G, path='ptt_201905.gexf')
# + id="k04fhWn07g3a" colab_type="code" colab={}
plt.figure(figsize=(12, 12))#nodelist
nx.draw_networkx(G, with_labels=True, nodelist=top_pg_rnk_node, labels={x:x for x in top_pg_rnk_node})
plt.show()
# + id="4e5GaV4R7g3b" colab_type="code" outputId="1e67ec01-deec-4631-fb97-022b3c1f8671" colab={}
Image(filename = 'ptt.jpg')
# + [markdown] id="M6UfLtz77g3d" colab_type="text"
# # ToDo
# #### 1. https://pttpedia.fandom.com/zh/wiki/KingKingCold 比對ptt名人
# #### 2. 解析ip(國家) 與 回文時間
# #### 3. 增加資料區間
# #### 4. 研究其他graph file format,優化效能
# #### 5. [研究community detection](https://kknews.cc/zh-tw/news/pp5olqz.html)
# #### 6. [igraph](http://landcareweb.com/questions/4756/igraphzhong-she-qu-jian-ce-suan-fa-zhi-jian-you-shi-yao-qu-bie)
# #### 7. [add dynamic graph](https://seinecle.github.io/gephi-tutorials/generated-html/converting-a-network-with-dates-into-dynamic.html#_1_dynamic_nodes_with_a_start_date)
| ptt_etl_201910.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding: utf-8
import csv
from os import stat
from pathlib import Path
"""Part 1: Automate the Calculations.
Automate the calculations for the loan portfolio summaries.
First, let's start with some calculations on a list of prices for 5 loans.
1. Use the `len` function to calculate the total number of loans in the list.
2. Use the `sum` function to calculate the total of all loans in the list.
3. Using the sum of all loans and the total number of loans, calculate the average loan price.
4. Print all calculations with descriptive messages.
"""
loan_costs = [500, 600, 200, 1000, 450]
# How many loans are in the list?
# @TODO: Use the `len` function to calculate the total number of loans in the list.
# Print the number of loans from the list
the_number_of_loan = len(loan_costs)
print(F"The number of loans is : {the_number_of_loan}")
# What is the total of all loans?
# @TODO: Use the `sum` function to calculate the total of all loans in the list.
# Print the total value of the loans
total_loans = sum(loan_costs)
print(f"The total value of loans is : {total_loans}")
# What is the average loan amount from the list?
# @TODO: Using the sum of all loans and the total number of loans, calculate the average loan price.
# Print the average loan amount
avg_loan_amount = total_loans / the_number_of_loan
print(f"The average loan amount is : {avg_loan_amount}")
# +
"""Part 2: Analyze Loan Data.
Analyze the loan to determine the investment evaluation.
Using more detailed data on one of these loans, follow these steps to calculate a Present Value, or a "fair price" for what this loan would be worth.
1. Use get() on the dictionary of additional information to extract the **Future Value** and **Remaining Months** on the loan.
a. Save these values as variables called `future_value` and `remaining_months`.
b. Print each variable.
@NOTE:
**Future Value**: The amount of money the borrower has to pay back upon maturity of the loan (a.k.a. "Face Value")
**Remaining Months**: The remaining maturity (in months) before the loan needs to be fully repaid.
2. Use the formula for Present Value to calculate a "fair value" of the loan. Use a minimum required return of 20% as the discount rate.
3. Write a conditional statement (an if-else statement) to decide if the present value represents the loan's fair value.
a. If the present value of the loan is greater than or equal to the cost, then print a message that says the loan is worth at least the cost to buy it.
b. Else, the present value of the loan is less than the loan cost, then print a message that says that the loan is too expensive and not worth the price.
@NOTE:
If Present Value represents the loan's fair value (given the required minimum return of 20%), does it make sense to buy the loan at its current cost?
"""
# Given the following loan data, you will need to calculate the present value for the loan
loan = {
"loan_price": 500,
"remaining_months": 9,
"repayment_interval": "bullet",
"future_value": 1000,
}
# @TODO: Use get() on the dictionary of additional information to extract the Future Value and Remaining Months on the loan.
# Print each variable.
print("The future value of the loan is:", loan["future_value"])
print("The remainning months on the loan is: ", loan["remaining_months"])
# @TODO: Use the formula for Present Value to calculate a "fair value" of the loan.
# Use a minimum required return of 20% as the discount rate.
# You'll want to use the **monthly** version of the present value formula.
# HINT: Present Value = Future Value / (1 + Discount_Rate/12) ** remaining_months
fair_value = loan["loan_price"]
present_value = loan["future_value"] / (1 + 0.2/12) ** loan["remaining_months"]
# If Present Value represents what the loan is really worth, does it make sense to buy the loan at its cost?
# @TODO: Write a conditional statement (an if-else statement) to decide if the present value represents the loan's fair value.
# If the present value of the loan is greater than or equal to the cost, then print a message that says the loan is worth at least the cost to buy it.
# Else, the present value of the loan is less than the loan cost, then print a message that says that the loan is too expensive and not worth the price.
if present_value >= fair_value:
print(f" the loan is worth at least the cost to buy it")
else:
print(f" The loan is too expensive and not worth the price")
# +
"""Part 3: Perform Financial Calculations.
Perform financial calculations using functions.
1. Define a new function that will be used to calculate present value.
a. This function should include parameters for `future_value`, `remaining_months`, and the `annual_discount_rate`
b. The function should return the `present_value` for the loan.
2. Use the function to calculate the present value of the new loan given below.
a. Use an `annual_discount_rate` of 0.2 for this new loan calculation.
"""
# Given the following loan data, you will need to calculate the present value for the loan
new_loan = {
"loan_price": 800,
"remaining_months": 12,
"repayment_interval": "bullet",
"future_value": 1000,
"annual_discount_rate": 0.2
}
# @TODO: Define a new function that will be used to calculate present value.
# This function should include parameters for `future_value`, `remaining_months`, and the `annual_discount_rate`
# The function should return the `present_value` for the loan.
def add( future_value, remaining_months, annual_discount_rate):
present_value = future_value / (1 + annual_discount_rate/12) ** remaining_months
return present_value
# @TODO: Use the function to calculate the present value of the new loan given below.
# Use an `annual_discount_rate` of 0.2 for this new loan calculation.
print(f"The present value of the loan is: ${present_value}")
# +
"""Part 4: Conditionally filter lists of loans.
In this section, you will use a loop to iterate through a series of loans and select only the inexpensive loans.
1. Create a new, empty list called `inexpensive_loans`.
2. Use a for loop to select each loan from a list of loans.
a. Inside the for loop, write an if-statement to determine if the loan_price is less than 500
b. If the loan_price is less than 500 then append that loan to the `inexpensive_loans` list.
3. Print the list of inexpensive_loans.
"""
loans = [
{
"loan_price": 700,
"remaining_months": 9,
"repayment_interval": "monthly",
"future_value": 1000,
},
{
"loan_price": 500,
"remaining_months": 13,
"repayment_interval": "bullet",
"future_value": 1000,
},
{
"loan_price": 200,
"remaining_months": 16,
"repayment_interval": "bullet",
"future_value": 1000,
},
{
"loan_price": 900,
"remaining_months": 16,
"repayment_interval": "bullet",
"future_value": 1000,
},
]
# -
# @TODO: Create an empty list called `inexpensive_loans`
inexpensive_loan = []
# +
# @TODO: Loop through all the loans and append any that cost $500 or less to the `inexpensive_loans` list
for loan in loans:
if loan["loan_price"]<= 500:
inexpensive_loan.append(loan)
# @TODO: Print the `inexpensive_loans` list
print(inexpensive_loan)
# +
"""Part 5: Save the results.
Output this list of inexpensive loans to a csv file
1. Use `with open` to open a new CSV file.
a. Create a `csvwriter` using the `csv` library.
b. Use the new csvwriter to write the header variable as the first row.
c. Use a for loop to iterate through each loan in `inexpensive_loans`.
i. Use the csvwriter to write the `loan.values()` to a row in the CSV file.
Hint: Refer to the official documentation for the csv library.
https://docs.python.org/3/library/csv.html#writer-objects
"""
# Set the output header
header = ["loan_price", "remaining_months", "repayment_interval", "future_value"]
# Set the output file path
output_path = Path("inexpensive_loans.csv")
# @TODO: Use the csv library and `csv.writer` to write the header row
# and each row of `loan.values()` from the `inexpensive_loans` list.
import csv
with open( output_path,'w') as f:
writer = csv.writer(f, delimiter=",")
writer.writerow(header)
for row in inexpensive_loan:
writer.writerow(row.values())
# -
| Challenge 1 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Detección de anomalías - PCA
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.decomposition import PCA
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import scale
from tqdm import tqdm
import PCAs_errorReconstruccion_primerCuarto
from PCAs_errorReconstruccion_primerCuarto import *
df_stats_Ch1_test2 = pd.read_csv("../DataStatistics/estadisticos_test2_ch1.csv" , sep = ',')
X_Ch1 = df_stats_Ch1_test2[['Min', 'Max', 'Kurt', 'ImpFactor', 'RMS', 'MargFactor', 'Skewness',
'ShapeFactor', 'PeakToPeak', 'CrestFactor']].values
# +
# Primera aproximación:
pca_pipeline = make_pipeline(StandardScaler(), PCA())
pca_pipeline.fit(X_Ch1)
# Proyección de los datos
modelo_pca = pca_pipeline.named_steps['pca']
prop_varianza_acum = modelo_pca.explained_variance_ratio_.cumsum()
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(9, 4))
ax.plot(np.arange(modelo_pca.n_components_) + 1, prop_varianza_acum, marker = 'o')
for x, y in zip(np.arange(modelo_pca.n_components_) + 1, prop_varianza_acum):
label = round(y, 2)
ax.annotate( label, (x,y), textcoords = "offset points", xytext = (0,10), ha = 'center')
ax.set_ylim(0, 1.2)
ax.set_xticks(np.arange(modelo_pca.n_components_) + 1)
ax.set_title('Cumulative explained variance')
ax.set_xlabel('Number of principal components')
ax.set_ylabel('Explained variance');
# +
pca_pipeline = make_pipeline(StandardScaler(), PCA())
pca_pipeline.fit(X_Ch1[:int(len(X_Ch1)/4)])
modelo_pca = pca_pipeline.named_steps['pca']
prop_varianza_acum = modelo_pca.explained_variance_ratio_.cumsum()
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(9, 4))
ax.plot(np.arange(modelo_pca.n_components_) + 1, prop_varianza_acum, marker = 'o')
for x, y in zip(np.arange(modelo_pca.n_components_) + 1, prop_varianza_acum):
label = round(y, 2)
ax.annotate( label, (x,y), textcoords = "offset points", xytext = (0,10), ha = 'center')
ax.set_ylim(0, 1.2)
ax.set_xticks(np.arange(modelo_pca.n_components_) + 1)
ax.set_title('Cumulative explained variance')
ax.set_xlabel('Number of principal components')
ax.set_ylabel('Explained variance');
# -
reconstruccion, error_reconstruccion = pca_reconstruccion_error_reconstruccion_primerCuarto(df_stats_Ch1_test2, 6, imp = 1)
# +
df_resultados = pd.DataFrame({
'error_reconstruccion' : error_reconstruccion,
})
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(5.5, 3.5))
sns.boxplot(
y = 'error_reconstruccion',
data = df_resultados,
#color = "white",
palette = 'tab10',
ax = ax
)
ax.set_yscale("log")
ax.set_title('Distribución de los errores de reconstrucción (PCA)')
# -
df_resultados.quantile(0.98)[0]
# Distribución del error de reconstrucción
# ==============================================================================
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(8, 6))
sns.distplot(
error_reconstruccion,
hist = False,
rug = True,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax
)
ax.set_title('Distribución de los errores de reconstrucción (PCA)')
ax.set_xlabel('Error de reconstrucción');
# Distribución del error de reconstrucción
# ==============================================================================
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(8, 6))
sns.distplot(
error_reconstruccion,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax
)
ax.set_title('Distribution of reconstruction errors (PCA)')
ax.set_xlabel('Reconstruction error');
# +
# Entrenamiento modelo PCA con escalado de los datos
X_primerCuarto = X_Ch1[:int(len(X_Ch1)/4)]
pca_pipeline = make_pipeline(StandardScaler(), PCA(n_components = 6))
pca_pipeline.fit(X_primerCuarto)
# Proyectar los datos
proyecciones_train = pca_pipeline.transform(X_primerCuarto)
# Reconstrucción
reconstruccion_train = pca_pipeline.inverse_transform(X = proyecciones_train)
# RMSE:
error_reconstruccion_train = np.sqrt(((reconstruccion_train - X_primerCuarto) ** 2).mean(axis=1))
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(15, 6))
sns.distplot(
error_reconstruccion_train,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax1
)
ax1.set_title('Distribution of reconstruction errors (PCA) - Train')
ax1.set_xlabel('Reconstruction error');
sns.distplot(
error_reconstruccion,
hist = False,
rug = False,
color = 'red',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax2
)
ax2.set_title('Distribution of reconstruction errors (PCA) - Complete signal')
ax2.set_xlabel('Reconstruction error');
# +
# Entrenamiento modelo PCA:
X_primerCuarto = X_Ch1[:int(len(X_Ch1)/4)]
pca_pipeline = make_pipeline(StandardScaler(), PCA(n_components = 6))
pca_pipeline.fit(X_primerCuarto)
# Proyectar los datos
proyecciones_train = pca_pipeline.transform(X_primerCuarto)
# Reconstrucción
reconstruccion_train = pca_pipeline.inverse_transform(X = proyecciones_train)
# RMSE:
error_reconstruccion_train = np.sqrt(((reconstruccion_train - X_primerCuarto) ** 2).mean(axis=1))
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(15, 6))
sns.distplot(
error_reconstruccion,
hist = False,
rug = False,
color = 'red',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax,
label = 'Complete signal'
)
sns.distplot(
error_reconstruccion_train,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax,
label = 'Train'
)
ax.set_title('Distribution of reconstruction errors (PCA) - Train vs Complete signal')
ax.set_xlabel('Reconstruction error');
ax.legend()
# -
error_reconstruccion = error_reconstruccion.values
from sklearn.mixture import GaussianMixture
gm = GaussianMixture(n_components=2, random_state=33).fit(error_reconstruccion[int(len(error_reconstruccion)/4):].reshape(-1, 1))
gm.means_
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(9, 6))
sns.distplot(
error_reconstruccion[int(len(error_reconstruccion)/4):],
hist = False,
rug = False,
color = 'orange',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax
)
ax.axvline(gm.means_[0], ls = '--', color = 'black')
ax.annotate(str(round(gm.means_[0][0],8)), xy=(0.05, 1.5), xytext=(1.5, 1.75),
arrowprops=dict(facecolor='black', shrink=0.05)
)
ax.axvline(gm.means_[1], ls = ':', color = 'black')
ax.annotate(str(round(gm.means_[1][0],8)), xy=(0.75, 1), xytext=(1.5, 1.15),
arrowprops=dict(facecolor='black', shrink=0.05),
)
ax.set_title('Distribution of reconstruction errors (PCA) - Complete signal except the first quarter')
ax.set_xlabel('Reconstruction error');
pred_GM = gm.predict(error_reconstruccion[int(len(error_reconstruccion)/4):].reshape(-1, 1))
sum(pred_GM)
pred_GM = [0] * int(len(error_reconstruccion)/4)
pred_GM_3cuartos = gm.predict(error_reconstruccion[int(len(error_reconstruccion)/4):].reshape(-1, 1))
for i in range(len(pred_GM_3cuartos)):
pred_GM.append(pred_GM_3cuartos[i])
pred_GM = np.array(pred_GM)
colores = ["#00cc44", "#f73e05"]
n_signal = list(range(len(pred_GM)))
n_signal = np.array(n_signal)
signals_0 = n_signal[pred_GM == 0]
error_rec_0 = error_reconstruccion[pred_GM == 0]
signals_1 = n_signal[pred_GM == 1]
error_rec_1 = error_reconstruccion[pred_GM == 1]
plt.figure(figsize=(10,6))
plt.scatter(signals_0, error_rec_0, c = "#00cc44", label = 'Normal')
plt.scatter(signals_1, error_rec_1, c = "#f73e05", label = 'Anomalies')
plt.title('Reconstruction error (PCA) - Ch1 test2')
plt.xlabel('Signal')
plt.ylabel('Error')
plt.legend()
# +
comienzo_1hora_anomalias = 'NA'
for i in range(len(pred_GM)):
if pred_GM[i:i+6].all():
comienzo_1hora_anomalias = i
break
pred_GM_1hora_anomalias = [0] * comienzo_1hora_anomalias + [1] * (len(pred_GM) - comienzo_1hora_anomalias)
colores = ["#00cc44", "#f73e05"]
x = np.arange(-10, len(df_stats_Ch1_test2)+10, 0.02)
n_signal = list(range(len(pred_GM_1hora_anomalias)))
plt.figure(figsize=(10,6))
plt.scatter(n_signal, error_reconstruccion, c = np.take(colores, pred_GM_1hora_anomalias))
plt.axvline(comienzo_1hora_anomalias, color = 'r', label = 'Beginning of anomalies')
plt.fill_between(x, min(error_reconstruccion)-0.5, max(error_reconstruccion)+1, where = x < comienzo_1hora_anomalias,
facecolor = 'green', alpha = 0.2, label = 'Normal')
plt.fill_between(x, min(error_reconstruccion)-0.5, max(error_reconstruccion)+1, where = x > comienzo_1hora_anomalias,
facecolor = 'red', alpha = 0.5, label = 'Anomalies ')
plt.title('Reconstruction error (PCA) - Ch1 test2')
plt.xlabel('Signal')
plt.ylabel('Error')
plt.legend(loc = 2)
# -
from sklearn.mixture import GaussianMixture
gm = GaussianMixture(n_components=3, random_state=33).fit(error_reconstruccion[int(len(error_reconstruccion)/4):].reshape(-1, 1))
gm.means_
colores = ["#00cc44", "#00cc44", "#ff7700"]
pred_GM = gm.predict(error_reconstruccion.reshape(-1, 1))
n_signal = list(range(len(pred_GM)))
plt.scatter(n_signal, error_reconstruccion, c = np.take(colores, pred_GM))
plt.title('Errores reconstrucción - Ch1 test2')
# #### Z-Scores
from scipy import stats
zscore_train = stats.zscore(error_reconstruccion_train)
mean = np.mean(error_reconstruccion_train)
std = np.std(error_reconstruccion_train)
std
zscore = (error_reconstruccion - mean) / std
threshold = 3
outlier = [0] * len(error_reconstruccion_train)
for i in range(len(error_reconstruccion_train), len(error_reconstruccion)):
z = (error_reconstruccion[i] - mean) / std
if abs(z) > threshold:
outlier.append(1)
else:
outlier.append(0)
outlier = np.array(outlier)
n_signal = list(range(len(error_reconstruccion)))
n_signal = np.array(n_signal)
signals_0 = n_signal[outlier == 0]
error_rec_0 = error_reconstruccion[outlier == 0]
signals_1 = n_signal[outlier == 1]
error_rec_1 = error_reconstruccion[outlier == 1]
plt.figure(figsize=(10,6))
plt.scatter(signals_0, error_rec_0, c = "#00cc44", label = 'Normal')
plt.scatter(signals_1, error_rec_1, c = "#f73e05", label = 'Anomalies')
plt.title('Reconstruction error (PCA) - Ch1 test2')
plt.xlabel('Signal')
plt.ylabel('Error')
plt.legend()
# +
z = (error_reconstruccion - mean) / std
comienzo_1hora_ouliers = 'NA'
for i in range(len(error_reconstruccion_train), len(error_reconstruccion)):
if (abs(z[i:i+6]) > threshold).all():
comienzo_1hora_ouliers = i
break
colores = ["#00cc44", "#f73e05"]
zscores_1hora_anomalias = [0] * comienzo_1hora_ouliers + [1] * (len(z) - comienzo_1hora_ouliers)
x = np.arange(-10, len(df_stats_Ch1_test2) + 10, 0.02)
n_signal = list(range(len(zscores_1hora_anomalias)))
plt.figure(figsize=(10,6))
plt.scatter(n_signal, error_reconstruccion, c = np.take(colores, zscores_1hora_anomalias))
plt.axvline(comienzo_1hora_ouliers, color = 'r', label = 'Beginning of anomalies')
plt.fill_between(x, min(error_reconstruccion)-0.5, max(error_reconstruccion)+1, where = x < comienzo_1hora_ouliers,
facecolor = 'green', alpha = 0.2, label = 'Normal')
plt.fill_between(x, min(error_reconstruccion)-0.5, max(error_reconstruccion)+1, where = x > comienzo_1hora_ouliers,
facecolor = 'red', alpha = 0.5, label = 'Anomalies ')
plt.title('Reconstruction error (PCA) - Ch1 test2')
plt.xlabel('Signal')
plt.ylabel('Error')
plt.legend(loc = 2)
# -
# #### Tiempo hasta el fallo:
print('Comienzo de anomalías cuando se producen durante una hora')
print('GMM:', (len(error_reconstruccion) - comienzo_1hora_anomalias) * 10, ' minutos')
print('Z-Scores:', (len(error_reconstruccion) - comienzo_1hora_ouliers) * 10, ' minutos')
print('Duración total de la señal:', len(error_reconstruccion) * 10, ' minutos')
# #### Errores de reconstrucción de cada estadístico
error = np.abs(reconstruccion[['Min', 'Max', 'Kurt', 'ImpFactor', 'RMS', 'MargFactor', 'Skewness',
'ShapeFactor', 'PeakToPeak', 'CrestFactor']].values - X_Ch1)
params = ['Min', 'Max', 'Kurt', 'ImpFactor', 'RMS', 'MargFactor', 'Skewness',
'ShapeFactor', 'PeakToPeak', 'CrestFactor']
error_min = error[:, 0]
error_max = error[:, 1]
error_kurt = error[:, 2]
error_if = error[:, 3]
error_rms = error[:, 4]
error_mf = error[:, 5]
error_skew = error[:, 6]
error_sf = error[:, 7]
error_ptp = error[:, 8]
error_cf = error[:, 9]
# +
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
fig, ((ax1, ax2), (ax3, ax4), (ax5, ax6), (ax7, ax8), (ax9, ax10)) = plt.subplots(nrows=5, ncols=2, figsize=(20, 30))
sns.distplot(
error_min,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax1
)
ax1.set_title('Distribución de los errores de reconstrucción - Min (PCA)')
ax1.set_xlabel('Error de reconstrucción');
sns.distplot(
error_max,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax2
)
ax2.set_title('Distribución de los errores de reconstrucción - Max (PCA)')
ax2.set_xlabel('Error de reconstrucción');
sns.distplot(
error_kurt,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax3
)
ax3.set_title('Distribución de los errores de reconstrucción - Kurtosis (PCA)')
ax3.set_xlabel('Error de reconstrucción');
sns.distplot(
error_if,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax4
)
ax4.set_title('Distribución de los errores de reconstrucción - Impulse Factor (PCA)')
ax4.set_xlabel('Error de reconstrucción');
sns.distplot(
error_rms,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax5
)
ax5.set_title('Distribución de los errores de reconstrucción - RMS (PCA)')
ax5.set_xlabel('Error de reconstrucción');
sns.distplot(
error_mf,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax6
)
ax6.set_title('Distribución de los errores de reconstrucción - Margin Factor (PCA)')
ax6.set_xlabel('Error de reconstrucción');
sns.distplot(
error_skew,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax7
)
ax7.set_title('Distribución de los errores de reconstrucción - Skewness (PCA)')
ax7.set_xlabel('Error de reconstrucción');
sns.distplot(
error_sf,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax8
)
ax8.set_title('Distribución de los errores de reconstrucción - Shape Factor (PCA)')
ax8.set_xlabel('Error de reconstrucción');
sns.distplot(
error_ptp,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax9
)
ax9.set_title('Distribución de los errores de reconstrucción - Peal to Peak (PCA)')
ax9.set_xlabel('Error de reconstrucción');
sns.distplot(
error_cf,
hist = False,
rug = False,
color = 'blue',
kde_kws = {'shade': True, 'linewidth': 1},
ax = ax10
)
ax10.set_title('Distribución de los errores de reconstrucción - Crest Factor (PCA)')
ax10.set_xlabel('Error de reconstrucción');
| TimeToFailurePrediction/AnomalyDetectionPCA/AnomalyDetectionPCAs_Ch1_test2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fUQpl1jDZESL"
# # This notebook was modified & updated from Colab
# > This is a descrption of the fastpages tutorial for Jupyter notebooks.
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [jupyter]
# - image: images/chart-preview.png
# + [markdown] id="TGQDRyniIsYS"
# ## What is Data Analysis
# + [markdown] id="wFMnAZx0Ith5"
# > *A process of inspecting, cleansing, transforming and modeling data with the goal of discovering useful information, informing conclusion and supporting decision-making. **Source:** Wikipedia*
# + [markdown] id="V7vT8pBMe0MN"
# ## Uses of EDA:
# + [markdown] id="6yI1373Re3QU"
# - To know the structure and distribution of data
# - To find relationship between Features
# - To find relationship between Features and the Target Variable
# - To find errors, anomalies, outliers
# - To refine Hipothesis or generate new questions on dataset
# + [markdown] id="GFwI9e6gJESC"
# ## Data Analysis Tools
# + [markdown] id="7-YW7dqEJFRr"
# Programming Languages: Open Source, Free, Extremely Powerful, Steep learning curve
# - Python
# - R
# - Julia
#
# Auto-managed closed tools: Closed Source, Expensive, Limited, Easy to learn
# - Power BI
# - Tableau
# - Qlik
# + [markdown] id="wJFnCxJoLD2O"
# ## The Data Analysis Process
#
# + [markdown] id="QYq7Uz3pLF5m"
# ### Data Extraction
# - SQL
# - Scrapping
# - File Formats
# - CSV
# - JSON
# - XML
# - Consulting APIs
# - Buying Data
# - Distributed Databases
#
# + [markdown] id="Qr-toYRnL_sI"
# ### Data Cleaning
# - Missing values and empty data
# - Data imputation
# - Incorrect types
# - Incorrect or invalid values
# - Outliers and non relevant data
# - Statistical sanitization
#
# + [markdown] id="KFp47BnsMIGH"
# ### Data Wrangling
#
# - Hierarchical Data
# - Handling categorical data
# - Reshaping and transforming structures
# - Indexing data for quick access
# - Merging, combining and joining data
# + [markdown] id="ddsDbZHcMOzf"
# ### Analysis
#
# - Exploration
# - Building statistical models
# - Visualization and representations
# - Correlation vs Causation analysis
# - Hypothesis testing
# - Statistical analysis
# - Reporting
# + [markdown] id="zWDzCSdUMVLA"
# ### Action
#
# - Building Machine Learning Models
# - Feature Engineering
# - Moving ML into production
# - Building ETL pipelines
# - Live dashboard and reporting
# - Decision making and real-life tests
# + [markdown] id="TlS6IBm-IAN0"
# https://jakevdp.github.io/PythonDataScienceHandbook/03.09-pivot-tables.html
#
# + [markdown] id="yKahYamepzYI"
# Proceso de organizar, resumir y visualizar un conjunto de datos para extraer información que aporte al logro de objetivos
# + [markdown] id="e-sJWbeg84rk"
# # why using Python and Pandas?
# + [markdown] id="uW0Gd6eW88OL"
# The Pandas library is the key library for Data Science and Analytics and a good place to start for beginners. Often called the "Excel & SQL of Python, on steroids" because of the powerful tools Pandas gives you for editing two-dimensional data tables in Python and manipulating large datasets with ease.
# + [markdown] id="CABBfktk-BVm"
# Pandas makes it very convenient to load, process, and analyze such tabular data using SQL-like queries. In conjunction with Matplotlib and Seaborn, Pandas provides a wide range of opportunities for visual analysis of tabular data.
# + [markdown] id="HSgU_qXO-G69"
# The main data structures in Pandas are implemented with Series and DataFrame classes. DataFrames are great for representing real data: rows correspond to instances (examples, observations, etc.), and columns correspond to features of these instances.
# + [markdown] id="Ihx49JvV6DBU"
# # Main Keywords
# + [markdown] id="gfZ1Jv605iBM"
# - **Dataframe:** is a main Object in Pandas, It's used to represent data in rows and columns (Tabular Data)
# - **Pandas:** This library needs no introduction as it became the de facto tool for Data Analysis in Python. The name pandas is derived from the term “panel data”, an econometrics term for datasets that include observations over multiple time periods for the same individuals.
| _notebooks/2021-10-03-My-First-Post-By-Colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as sts
import seaborn as sns
sns.set()
sns.set_style("whitegrid")
color_palette = sns.color_palette('deep') + sns.color_palette('husl', 6) + sns.color_palette('bright') + sns.color_palette('pastel')
# %matplotlib inline
# -
# ## Multiple neurons with constant rate
#
# Explore how many neurons we can model without much lagging:
#
# - 1k-4k neurons
# - 100Hz rate
# - or 1Hz for 10ms time unit
# - ~2.5ms lag for an event
# +
standard_rate = 40.
sim_n, spikes = 2_000, 200
scale = 1. / standard_rate
a = np.random.exponential(scale=scale, size=(spikes, sim_n)) + .01 * scale
sns.distplot(a[:, 0])
a[:, 0].sum()
# +
# %%time
import time
from queue import heappush, heappop
pq = list()
t0 = time.monotonic()
for t in range(sim_n):
heappush(pq, (a[0, t] + t0, t))
i, i_spike = 1, [1]*sim_n
k = sim_n * spikes // 10
lag = .0
spin_cnt = 0
while len(pq):
t = time.monotonic()
if pq[0][0] <= t:
ct, d = heappop(pq)
cur_lag = t - ct
lag += cur_lag
if i_spike[d] < spikes:
heappush(pq, (a[i_spike[d], d] + t, d))
i_spike[d] += 1
i += 1
# if i % k == 0:
# print(t - t0, cur_lag * 1000., i, d)
else:
spin_cnt += 1
print(spin_cnt / 1000_000, lag * 1000. / (sim_n * spikes), lag * 100 / (sim_n * spikes) / scale)
# -
# ## One neuron, non-constant rate
# +
# %%time
def rate_lambda(dT):
return 5 + 4 * np.sin(dT)
def sample_spike_dt(dT):
rate = rate_lambda(dT)
scale = 1./rate
spike_dt = np.random.exponential(scale=scale) + .01 * scale
return spike_dt, rate
def scaled_time(T0, T):
return (T - T0) / unit_of_time
unit_of_time = .005 # fraction of second
T0 = time.monotonic()
t, t_end = 0., 10.
ts, rates, spikes = [], [], []
next_spike_dt, cur_rate = sample_spike_dt(t)
next_spike_t = t + next_spike_dt
ts.append(t)
rates.append(cur_rate)
# print(t, next_spike_t)
j = 0
while t <= t_end:
t = scaled_time(T0, time.monotonic())
if t < next_spike_t:
j += 1
continue
ts.append(t)
spikes.append(t)
next_spike_dt, cur_rate = sample_spike_dt(t)
next_spike_t = t + next_spike_dt
rates.append(cur_rate)
# print(t, next_spike_t, next_spike_dt)
plt.figure(figsize=(20, 5))
sns.lineplot(ts, rates)
min_rates, max_rates = min(rates), max(rates)
plt.vlines(spikes, min_rates * .95, max_rates * 1.05)
# sns.scatterplot(spikes, rates[1:], marker='|', alpha=1.)
j, t, t_end
# -
# ## Loading images from four-shapes kaggle dataset
# +
import imageio
import os
from PIL import Image
four_shapes_dir = "./datasets/four_shapes/shapes"
def get_images(cnt, step=1, shape='triangle', start=600, size=10, normalized=True):
cnt = int(cnt) # if cnt is float
if not isinstance(size, tuple):
size = (size, size)
images = []
for i in range(cnt):
ind = start + i * step
img_path = os.path.join(four_shapes_dir, shape, f"{ind}.png")
img = imageio.imread(img_path)
resized_img = Image.fromarray(img).resize(size)
img_arr = np.array(resized_img, dtype=np.float16) / 255. if normalized else np.array(resized_img)
images.append(img_arr)
return images
get_images(2, size=6)[0]
# +
from IPython.display import clear_output
def slideshow_images(images, delay=.2):
for img in images:
clear_output()
plt.imshow(img)
plt.show()
time.sleep(delay)
slideshow_images(get_images(30, step=4, size=32, normalized=False, shape='triangle'))
# -
# ## One neuron, pixel based intensity from multiple images
# + jupyter={"outputs_hidden": true, "source_hidden": true}
# %%time
def rate_lambda(t):
ind = min(int(t * len(small_images) / t_end), len(small_images) - 1)
img = small_images[ind]
return 4 + img[7][3] / 15
def sample_spike_dt(t, rate):
scale = 1./rate
spike_dt = np.random.exponential(scale=scale) + .01 * scale
return spike_dt
def scaled_time(T0, T):
return (T - T0) / unit_of_time
unit_of_time = .005 # fraction of second
T0 = time.monotonic()
t, t_end = 0., 15.
ts, rates, spikes = [], [], []
cur_rate = rate_lambda(t)
next_spike_dt = sample_spike_dt(t, cur_rate)
next_spike_t = t + next_spike_dt
ts.append(t)
rates.append(cur_rate)
# print(t, next_spike_t)
j = 0
while t <= t_end:
t = scaled_time(T0, time.monotonic())
if t < next_spike_t:
j += 1
continue
ts.append(t)
spikes.append(t)
cur_rate = rate_lambda(t)
next_spike_dt = sample_spike_dt(t, cur_rate)
next_spike_t = t + next_spike_dt
rates.append(cur_rate)
# print(t, next_spike_t, next_spike_dt)
plt.figure(figsize=(25, 5))
sns.lineplot(ts, rates)
min_rates, max_rates = min(rates), max(rates)
plt.vlines(spikes, min_rates * .95, max_rates * 1.05)
j, len(ts), t, t_end
# -
# ## Multiple neurons, images based intensity
# + jupyter={"outputs_hidden": true, "source_hidden": true}
# %%time
def rate_lambda(t, i, j):
ind = min(int(t * len(small_images) / t_end), len(small_images) - 1)
img = small_images[ind]
return 4 + img[i][j] / pixel_scale
def save_events(ind, t, rate, t_spike=None):
i, j = ind
ts[i][j].append(t)
rates[i][j].append(rate)
if t_spike is not None:
spikes[i][j].append(t_spike)
def recalculate_rate_next_spike(ind, t):
i, j = ind
rate = rate_lambda(t, i, j)
next_spike_dt = sample_spike_dt(t, rate)
next_spike_t = t + next_spike_dt
return next_spike_t, rate
unit_of_time = .025 # fraction of second
pixel_scale = 15.
width, height = small_images[0].shape
T0 = time.monotonic()
t, t_end = 0., 10.
pq_times, ts, rates, spikes = [], [], [], []
for i in range(width):
ts.append([]), rates.append([]), spikes.append([])
for j in range(height):
ts[i].append([]), rates[i].append([]), spikes[i].append([])
ind = (i, j)
next_spike_t, cur_rate = recalculate_rate_next_spike(ind, t)
save_events(ind, t, cur_rate)
heappush(pq_times, (next_spike_t, ind))
# print(t, next_spike_t)
j = 0
next_spike_t, ind = pq_times[0]
while t <= t_end:
t = scaled_time(T0, time.monotonic())
if t < next_spike_t:
j += 1
continue
heappop(pq_times)
next_spike_t, cur_rate = recalculate_rate_next_spike(ind, t)
save_events(ind, t, cur_rate, t_spike=t)
heappush(pq_times, (next_spike_t, ind))
next_spike_t, ind = pq_times[0]
fig, axes = plt.subplots(3, 1, figsize=(20, 15))
for ax in axes:
x, y = np.random.randint(0, 10, size=(2,))
sns.lineplot(ts[x][y], rates[x][y], ax=ax)
min_rates, max_rates = min(rates[x][y]), max(rates[x][y])
ax.vlines(spikes[x][y], min_rates * .95, max_rates * 1.05)
ax.set_title(f"{x}, {y}")
plt.show()
event_count = sum(len(a) for row in ts for a in row)
j, event_count, t, t_end
# -
# ## Multiple neurons, images based intensity, no-fire updates
#
# - clip long no-fire by `2*frame_dt`
# +
def make_md_list(fn_elem_init, *args):
if len(args) == 0:
return fn_elem_init()
return [make_md_list(fn_elem_init, *args[1:]) for _ in range(args[0])]
make_md_list(tuple, 3, 2)
# +
def iterate_flatten_dims(a, dims):
return a if dims < 1 else (
x
for sub_a in a
for x in iterate_flatten_dims(sub_a, dims - 1)
)
# return [make_md_list(fn_elem_init, *args[1:]) for _ in range(args[0])]
t = make_md_list(list, 3, 2, 1)
print(t)
print([x for x in iterate_flatten_dims(t, 1)])
print([x for x in iterate_flatten_dims(t, 2)])
# +
def get_expected_spike_count():
mean_fire_rate = np.mean(np.array(images))*white_rate + black_rate
return mean_fire_rate * t_end * width * height
def get_expected_event_count():
return n_frames * width * height
get_expected_event_count(), get_expected_spike_count()
# +
# %%time
from math import floor, ceil
def rate_lambda(t, i, j):
ind = min(floor(t / frame_dt), n_frames - 1)
pixel = images[ind][i][j]
return black_rate + white_rate * pixel
def save_events(ind, t, rate, is_spike):
i, j = ind
ts[i][j].append(t)
rates[i][j].append(rate)
if is_spike:
spikes[i][j].append(t)
def sample_spike_dt(t, rate):
scale = 1./rate
return np.random.exponential(scale=scale) + .01 * scale
def recalculate_neuron_state(ind, t, is_spike=False):
i, j = ind
rate = rate_lambda(t, i, j)
save_events(ind, t, rate, is_spike)
next_spike_dt = sample_spike_dt(t, rate)
next_spike_t = t + min(next_spike_dt, max_no_fire_dt)
will_spike = next_spike_dt < max_no_fire_dt
# t_next_frame_update = ceil(t / frame_dt) * frame_dt
# if next_spike_t > t_next_frame_update:
# next_spike_t = t_next_frame_update
# will_spike = False
return next_spike_t, will_spike
unit_of_time = .1 # fraction of second
fpu = 4. # frames per unit of time
t, t_end = 0., 10.
n_frames = int(fpu * t_end)
black_rate, white_rate = .5, 16.
width, height = 20, 20
images = get_images(n_frames, step=2, shape='star', size=(width, height))
T0 = time.monotonic()
frame_dt = t_end / n_frames
max_no_fire_dt = frame_dt * 2
pq_times = []
ts, rates, spikes = [make_md_list(list, width, height) for _ in range(3)]
for i in range(width):
for j in range(height):
ind = (i, j)
next_spike_t, is_spike = recalculate_neuron_state(ind, t)
heappush(pq_times, (next_spike_t, is_spike, ind))
free_spin_count, lag = 0, 0.
next_spike_t, is_spike, ind = pq_times[0]
while t <= t_end:
t = scaled_time(T0, time.monotonic())
if t < next_spike_t:
free_spin_count += 1
continue
lag += t - next_spike_t
heappop(pq_times)
next_spike_t, is_spike = recalculate_neuron_state(ind, t, is_spike)
heappush(pq_times, (next_spike_t, is_spike, ind))
next_spike_t, is_spike, ind = pq_times[0]
fig, axes = plt.subplots(3, 1, figsize=(20, 15))
for ax in axes:
x, y = np.random.randint(0, 10, size=(2,))
sns.lineplot(ts[x][y], rates[x][y], ax=ax)
min_rates, max_rates = min(rates[x][y]), max(rates[x][y])
ax.vlines(spikes[x][y], min_rates * .95, max_rates * 1.05)
ax.set_title(f"{x}, {y}")
plt.show()
event_count = sum(map(len, iterate_flatten_dims(ts, 1))) - width * height
spikes_count = sum(map(len, iterate_flatten_dims(spikes, 1)))
lag_percent = 100 * (lag / event_count) / (event_count / t_end)
free_spin_count, event_count, spikes_count, get_expected_spike_count(), lag_percent
# -
# ## Multiple neurons, difference based intensity
# +
# %%time
from math import floor, ceil
def rate_lambda(t, ind):
i, j, is_pos = ind
frame_ind = min(floor(t / frame_dt), n_frames - 1)
prev_frame_ind = max(frame_ind - 1, 0)
pixel_diff = images[frame_ind][i][j] - images[prev_frame_ind][i][j]
if (pixel_diff > pixel_diff_eps and is_pos) or (pixel_diff < -pixel_diff_eps and not is_pos):
return black_rate + white_rate * abs(pixel_diff)
else:
return base_rate
def save_events(ind, t, rate, is_spike):
i, j, is_pos = ind
k = int(is_pos)
ts[i][j][k].append(t)
rates[i][j][k].append(rate)
if is_spike:
spikes[i][j][k].append(t)
def recalculate_neuron_state(ind, t, is_spike):
rate = rate_lambda(t, ind)
save_events(ind, t, rate, is_spike)
next_spike_dt = sample_spike_dt(t, rate)
next_spike_t = t + min(next_spike_dt, max_no_fire_dt)
will_spike = next_spike_dt < max_no_fire_dt
return next_spike_t, will_spike
unit_of_time = .05 # fraction of second
fpu = 4. # frames per unit of time
t, t_end = 0., 30.
n_frames = int(fpu * t_end)
black_rate, white_rate = .5, 16.
width, height = 20, 20
images = get_images(n_frames, step=2, shape='star', size=(width, height))
T0 = time.monotonic()
frame_dt = t_end / n_frames
max_no_fire_dt = frame_dt * 1.5
pixel_diff_eps = 1./256
pq_times = []
ts, rates, spikes = [make_md_list(list, width, height, 2) for _ in range(3)]
for i in range(width):
for j in range(height):
for is_pos in [False, True]:
ind = (i, j, is_pos)
next_spike_t, is_spike = recalculate_neuron_state(ind, t, is_spike=False)
heappush(pq_times, (next_spike_t, is_spike, ind))
free_spin_count = 0
next_spike_t, is_spike, ind = pq_times[0]
while t <= t_end:
t = scaled_time(T0, time.monotonic())
if t < next_spike_t:
free_spin_count += 1
continue
heappop(pq_times)
next_spike_t, is_spike = recalculate_neuron_state(ind, t, is_spike)
heappush(pq_times, (next_spike_t, is_spike, ind))
next_spike_t, is_spike, ind = pq_times[0]
fig, axes = plt.subplots(3, 1, figsize=(20, 15))
for ax in axes:
x, y = np.random.randint(0, 10, size=(2,))
min_rates, max_rates = min(min(rates[x][y])), max(max(rates[x][y]))
for z, color in enumerate(['blue', 'red']):
sns.lineplot(np.array(ts[x][y][z]), np.array(rates[x][y][z]), ax=ax, color=color)
ax.vlines(spikes[x][y][z], min_rates * .95, max_rates * 1.05, colors=color)
ax.set_title(f"{x}, {y}")
plt.show()
event_count = sum(map(len, iterate_flatten_dims(ts, 2)))
spikes_count = sum(map(len, iterate_flatten_dims(spikes, 2)))
lag_percent = 100 * (lag / event_count) / (event_count / t_end)
free_spin_count, event_count, spikes_count, get_expected_spike_count(), lag_percent
# -
| 01-input-neurons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from src import util
from src.transformation import transform
from src import config
from src import subject_manipulation
from typing import Tuple, List
from src.config import SUBJECT_DF_PATH, BLANK_ANSWER_LABEL, DELETION_ANSWER_LABEL, CANCELLED_LABEL, DIFFICULTIES, \
MATH_SUBJECTS, COMPUTING_SUBJECTS, HUMAN_SUBJECTS, TECHNOLOGY_SUBJECTS
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib
import matplotlib.patches as mpatches
# + tags=["parameters"]
CODE_COURSE = None
# -
years = [2017, 2014, 2011, 2008, 2005]
def read_csv_course(year, filter_by_course):
df = transform.read_csv(year)
if filter_by_course is not None:
df = df.loc[df["CO_CURSO"] == CODE_COURSE]
return df
subject_df = subject_manipulation.get_processed_subject_df(SUBJECT_DF_PATH)
def get_dict_all_years(filter_by_course: bool) -> dict:
result = {}
for year in years:
result[year] = read_csv_course(year, filter_by_course=filter_by_course)
return result
# +
def get_display_df(year: int, input_df: pd.DataFrame,
subject_df: pd.DataFrame) -> Tuple[pd.DataFrame, pd.DataFrame, pd.DataFrame]:
temp_subject_df = subject_df.loc[subject_df['ano'] == year].copy()
util.map_presence(input_df)
presence_df = input_df["TP_PRES"].value_counts()
input_df = util.filter_present_students(input_df)
subjects = util.get_subjects(temp_subject_df)
input_df = util.add_all_score_categories(input_df, temp_subject_df, True,
subjects, util.get_subject_valid_questions)
num_questions = []
mean_acertos_by_subject = []
std_acertos_by_subject = []
column_zero_subject = []
column_geq_one_subject = []
column_all_subject = []
num_alunos = input_df.shape[0]
for subject in subjects:
subject_questions = util.get_subject_valid_questions(subject,
temp_subject_df,
input_df,
just_objective=True)
num_obj_subject_questions = len(subject_questions)
mean_acertos_subject = input_df[f"ACERTOS_OBJ_{subject}"].mean()
std_acertos_subject = input_df[f"ACERTOS_OBJ_{subject}"].std()
zero_subject_percentage = list(input_df[f"ACERTOS_OBJ_{subject}"]).count(0) * 100 / num_alunos
geq_one_subject_percentage = list(input_df[f"ACERTOS_OBJ_{subject}"] >= 1).count(True) * 100 / num_alunos
all_subject_percentage = list(input_df[f"ACERTOS_OBJ_{subject}"] == num_obj_subject_questions).count(True) * 100 / num_alunos
if num_obj_subject_questions > 0:
num_questions.append(num_obj_subject_questions)
mean_acertos_by_subject.append(mean_acertos_subject)
std_acertos_by_subject.append(std_acertos_subject)
column_zero_subject.append(zero_subject_percentage)
column_geq_one_subject.append(geq_one_subject_percentage)
column_all_subject.append(all_subject_percentage)
else:
num_questions.append(0)
mean_acertos_by_subject.append(None)
std_acertos_by_subject.append(None)
column_zero_subject.append(None)
column_geq_one_subject.append(None)
column_all_subject.append(None)
subjects_labels = [f"SCORE_OBJ_{x}" for x in subjects]
mean_by_subject = input_df[subjects_labels].mean().values
data = np.array([mean_by_subject, num_questions]).T
display_df = pd.DataFrame(data=data, index=subjects,
columns=["Nota %", "Nº Questões"])
display_df["Nº Questões"] = display_df["Nº Questões"].astype(int, errors="ignore")
display_df["Média Acertos"] = mean_acertos_by_subject
display_df["Desvio Padrão Acertos"] = std_acertos_by_subject
display_df["% de Zeros"] = column_zero_subject
display_df["% de Alunos que acertaram pelo menos uma questão"] = column_geq_one_subject
display_df["% de Alunos que acertaram todas"] = column_all_subject
display_df = display_df.sort_values(by=["Nota %"]).round(2)
return display_df, input_df, presence_df
# -
dfs = get_dict_all_years(filter_by_course=True)
# +
display_dfs = {}
presence_dfs = {}
for year in dfs.keys():
enade_df = dfs[year]
display_df, df, presence_df = get_display_df(year, enade_df, subject_df)
dfs[year] = df
display_dfs[year] = display_df
presence_dfs[year] = presence_df
# -
def display_all_years_nota(all_dfs: dict) -> pd.DataFrame:
data = {"2005": all_dfs[2005]["Nota %"].copy(),
"2008": all_dfs[2008]["Nota %"].copy(),
"2011": all_dfs[2011]["Nota %"].copy(),
"2014": all_dfs[2014]["Nota %"].copy(),
"2017": all_dfs[2017]["Nota %"].copy()}
return pd.DataFrame(data)
# +
def get_general_stats_df(df: pd.DataFrame, subject_df: pd.DataFrame, year: int):
subject_df_year = subject_df.loc[subject_df["ano"] == year]
objective_argument = subject_df_year["tipoquestao"] == "Objetiva"
def get_general_statistics(df: pd.DataFrame, questions: List[int]) -> List[float]:
status_relevant_questions = df[[f"QUESTAO_{i}_SITUACAO_DA_QUESTAO" for i in questions]].copy()
valid_questions = status_relevant_questions.loc[:, (status_relevant_questions.iloc[0] != 1)].columns
valid_questions = [int(x[len("QUESTAO_"):-len("_SITUACAO_DA_QUESTAO")]) for x in valid_questions]
relevant_columns = df[[f"QUESTAO_{i}_NOTA" for i in valid_questions]].copy()
relevant_columns = relevant_columns.replace([BLANK_ANSWER_LABEL, DELETION_ANSWER_LABEL], 0)
relevant_columns = relevant_columns.astype(int)
relevant_columns = relevant_columns / 100
data = relevant_columns.sum(axis=1)
mean_acertos = data.mean()
median_acertos = data.median()
std_acertos = data.std()
skew_acertos = data.skew()
kurtosis_acertos = data.kurtosis()
mode_acertos = data.mode().astype(int)
mode_acertos = mode_acertos.astype(str)
mode_acertos = ", ".join(mode_acertos.tolist())
return [mean_acertos, std_acertos, median_acertos, skew_acertos, kurtosis_acertos,
len(valid_questions)], mode_acertos, data
modes = []
index = ["Prova Completa", "Formação Geral", "Componente Específico"]
columns = ["Média Acertos", "Desvio Padrão Acertos", "Mediana Acertos",
"Skewness Acertos", "Kurtosis Acertos", "Nº Questões Válidas"]
data = np.zeros((len(index), len(columns)), dtype=float)
data_to_plot = []
general_index = subject_df_year["prova"] == "Geral"
specific_index = subject_df_year["prova"] == "Específica"
questions_lists = [subject_manipulation.get_objective_questions(subject_df_year), # questões objetivas
subject_df_year.loc[objective_argument & general_index]["idquestao"].tolist(), #objetivas e de formação geral
subject_df_year.loc[objective_argument & specific_index]["idquestao"].tolist()] #objetivas de de formação especifica
for type_exam_index, question_list in enumerate(questions_lists):
data[type_exam_index], new_mode, new_data_to_plot = get_general_statistics(df, question_list)
modes.append(new_mode)
data_to_plot.append(new_data_to_plot)
exam_df = pd.DataFrame(columns=columns, data=data, index=index)
exam_df["Moda Acertos"] = modes
exam_df["Nº Questões Válidas"] = exam_df["Nº Questões Válidas"].astype(int)
exam_df["Nota %"] = exam_df["Média Acertos"] * 100 / exam_df["Nº Questões Válidas"]
columns_without_num_questions = list(exam_df.columns)
columns_without_num_questions.remove("Nº Questões Válidas")
exam_df = exam_df[columns_without_num_questions + ["Nº Questões Válidas"]]
return exam_df.round(2), data_to_plot
general_stats_dfs = {}
data_to_plot = {}
for year in dfs.keys():
df = dfs[year].copy()
general_stats_df, new_data_to_plot = get_general_stats_df(df, subject_df, year)
general_stats_dfs[year] = general_stats_df
data_to_plot[year] = new_data_to_plot
def plot_histogram(data_to_plot_year):
labels = ["Prova Completa", "Formação Geral", "Componente Específico"]
for data, label in zip(data_to_plot_year, labels):
sns.distplot(data)
plt.title(f"Acertos ({label})")
plt.show()
year = 2017
print(f"Ano de {year}")
plot_histogram(data_to_plot[year])
general_stats_dfs[year]
# -
year = 2014
print(f"Ano de {year}")
plot_histogram(data_to_plot[year])
general_stats_dfs[year]
year = 2011
print(f"Ano de {year}")
plot_histogram(data_to_plot[year])
general_stats_dfs[year]
year = 2008
print(f"Ano de {year}")
plot_histogram(data_to_plot[year])
general_stats_dfs[year]
year = 2005
print(f"Ano de {year}")
plot_histogram(data_to_plot[year])
general_stats_dfs[year]
all_years_score_df = display_all_years_nota(general_stats_dfs)
all_years_score_df
ax = plt.subplot(111)
plt.plot(all_years_score_df.loc["Prova Completa"],'o-', color="#ff7f00", label="Prova Completa")
plt.plot(all_years_score_df.loc["Formação Geral"],'d-', color="#4daf4a", label="Formação Geral")
plt.plot(all_years_score_df.loc["Componente Específico"],'x-', color="#377eb8", label="Componente Específico")
plt.legend()
plt.ylabel("Percentual de Desempenho")
plt.xlabel("Ano")
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.ylim(0,100)
plt.savefig("/tmp/type_exam_score", dpi=600)
# +
def get_all_years_score(dfs: dict, subject_df: pd.DataFrame, years: List[int]):
def auxiliar_function(df: pd.DataFrame, questions: List[int]) -> List[float]:
status_relevant_questions = df[[f"QUESTAO_{i}_SITUACAO_DA_QUESTAO" for i in questions]].copy()
valid_questions = status_relevant_questions.loc[:, (status_relevant_questions.iloc[0] != 1)].columns
valid_questions = [int(x[len("QUESTAO_"):-len("_SITUACAO_DA_QUESTAO")]) for x in valid_questions]
relevant_columns = df[[f"QUESTAO_{i}_NOTA" for i in valid_questions]].copy()
relevant_columns = relevant_columns.replace([BLANK_ANSWER_LABEL, DELETION_ANSWER_LABEL], 0)
relevant_columns = relevant_columns.astype(int)
relevant_columns = relevant_columns / 100
data = relevant_columns.sum(axis=1)
correct_answered = data.sum()
num_alunos = df.shape[0]
answered = len(valid_questions) * num_alunos
return [correct_answered, answered]
all_year_general_score_dfs = []
for year in years:
df = dfs[year]
subject_df_year = subject_df.loc[subject_df["ano"] == year]
objective_argument = subject_df_year["tipoquestao"] == "Objetiva"
index = ["Prova Completa", "Formação Geral", "Componente Específico"]
columns = ["Numero Acertos", "Total de Questões Respondidas"]
data = np.zeros((len(index), len(columns)), dtype=float)
questions_lists = [subject_manipulation.get_objective_questions(subject_df_year),
subject_df_year.loc[objective_argument & (subject_df_year["prova"] == "Geral")]["idquestao"].tolist(),
subject_df_year.loc[objective_argument & (subject_df_year["prova"] == "Específica")]["idquestao"].tolist()]
for type_exam_index, question_list in enumerate(questions_lists):
data[type_exam_index] = auxiliar_function(df, question_list)
year_general_score_df = pd.DataFrame(columns=columns, data=data, index=index)
all_year_general_score_dfs.append(year_general_score_df)
output_df = all_year_general_score_dfs[0]
for df in all_year_general_score_dfs[1:]:
output_df = output_df.add(df)
output_df = output_df.astype(int)
output_df["Nota %"] = output_df["Numero Acertos"] * 100 / output_df["Total de Questões Respondidas"]
return output_df.round(2)
print("Dados de todos os anos considerando todos os alunos")
get_all_years_score(dfs, subject_df, dfs.keys())
| notebooks/type_exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing 2D data
# Colormaps are used to map quantities stored at each point of a (discretized) surface to colours.
# Colours live in a 3D space.
# 2D data may contain *metric* or *form* information (or both).
# How can we choose a good colormap?
# - Perception
# - Intuition
# - Convention
# By default, we want to use a sequential colormap (not assuming anything about the structure of our data).
# References:
# - `matplotlib`: http://matplotlib.org/users/colormaps.html, https://matplotlib.org/users/dflt_style_changes.html
# - `viscm`: https://bids.github.io/colormap/
# - Another SciPy 2015 talk: https://www.youtube.com/watch?v=XjHzLUnHeM0
import pickle
data = pickle.load(open('data/correlation_map.pkl', 'rb'))
data.keys()
type(data['excitation energy'])
data['excitation energy'].shape
data['correlation'].shape
import matplotlib
# %matplotlib inline
matplotlib.style.use('ggplot')
import matplotlib.pyplot as plt
plt.imshow(data['correlation'])
# Are we dealing with form or metric information here?
matplotlib.__version__
from matplotlib.cm import magma, inferno, plasma, viridis
# from colormaps import magma, inferno, plasma, viridis
correlation = data['correlation']
excitation = data['excitation energy']
emission = data['emission energy']
plt.imshow(correlation,
origin='lower',
cmap=magma)
plt.imshow(correlation,
origin='lower',
extent=[excitation.min(), excitation.max(), emission.min(), emission.max()],
cmap=magma)
f, ax = plt.subplots(1, 1, figsize=(5, 5))
map0 = ax.imshow(correlation,
origin='lower',
extent=[excitation.min(), excitation.max(), emission.min(), emission.max()],
cmap=magma)
ax.plot(excitation, excitation)
ax.set_xlabel('Excitation Energy (eV)')
f.colorbar(map0, ax=ax)
# Revise and edit to best convey the scientific result.
f, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.imshow(correlation,
origin='lower',
extent=[excitation.min(), excitation.max(), emission.min(), emission.max()],
cmap=viridis)
ax.plot(excitation, excitation)
ax.set_xlabel('Excitation Energy (eV)')
# What about using the new default colormap (above)?
| 2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
import numpy as np
from icecream import ic
# %matplotlib inline
#from matplotlib_inline.backend_inline import set_matplotlib_formats
#set_matplotlib_formats('svg') # shit, matshow is renderd as an image
#import seaborn as sb
from importlib import reload
import SolutionTable
import PRB
from pathlib import Path
# +
reload(SolutionTable)
reload(PRB)
from SolutionTable import Table
p = PRB.PRB(Path("./prbs/plant.prb"))
#p = PRB.PRB(Path("./prbs/clowns_shoes.prb"))
#p = PRB.PRB(Path("./prbs/mafia_2.prb"))
tableShapeSpec = []
for d in p.description.dims.values():
tableShapeSpec.append(tuple(d.entities.keys()))
t = Table(*tableShapeSpec)
for el in p.orig.mappingFacts:
print(el)
if isinstance(el, PRB.NotMapsFact):
t.isNot(el.entities[0].id, el.entities[1].id)
elif isinstance(el, PRB.MapsFact):
t.equal(el.entities[0].id, el.entities[1].id)
else:
raise ValueError(el)
t.plot()
# +
reload(SolutionTable)
from SolutionTable import buildAlphaVec
buildAlphaVec(np.array([0.1,0.2,0.3, 0.4]))
# -
# +
reload(SolutionTable)
from SolutionTable import buildAlphaVec, buildBMat, computeNewShit
import scipy.linalg
d = np.array([0.1, 0.5, 0.3, 0.4])
relaxed = computeNewShit(d, [3])
print(d, relaxed)
for i in range(1000):
d = relaxed
relaxed = computeNewShit(d, None)
print(d, relaxed, np.sum(relaxed))
# -
d = np.arange(-10, 11, 1)
d = d / len(d)
print(d)
plt.plot(d)
d1 = np.array(d)
d1[5] = 0.2
d1[6] = 0.0
d1[7] = 0.3
plt.plot(d)
plt.plot(computeNewShit(d1))
plt.plot(d1/np.sum(d1))
plt.grid()
# +
# isclose?
# -
| SolutionTable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/michaelmarchesi/Bitcoin_forecasting/blob/main/RandomForest.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="BZPXYwjBb0C7"
import json
import requests
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Activation, Dense, Dropout, LSTM,BatchNormalization
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.metrics import mean_absolute_error
from keras.preprocessing.sequence import TimeseriesGenerator
import plotly.graph_objects as go
# %matplotlib inline
# + id="tG_ctOypb0nP"
endpoint = 'https://min-api.cryptocompare.com/data/histoday'
res = requests.get(endpoint + '?fsym=BTC&tsym=USD&limit=2000')
df = pd.DataFrame(json.loads(res.content)['Data'])
df = df.set_index('time')
df.index = pd.to_datetime(df.index, unit='s').astype('datetime64[ns, Europe/Paris]').tz_convert('America/Edmonton')
# + id="5KOXEGnKb18b"
def calcMACD(data):
period12 = data.ewm(span=12).mean()
period26 = data.ewm(span=26).mean()
macd = [] # List to hold the MACD line values
counter=0 # Loop to substantiate the MACD line
while counter < (len(period12)):
macd.append(period12.iloc[counter,0] - period26.iloc[counter,0]) # Subtract the 26 day EW moving average from the 12 day.
counter += 1
return macd
# + colab={"base_uri": "https://localhost:8080/"} id="7vo425yXb2oZ" outputId="22d5ca39-fd5b-49ba-cada-a15d6c0d7898"
df['macd'] = calcMACD(df)
y = df['close']
features = ['macd','volumeto', 'high', 'low']
X = df[features]
X.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="rHzfR6JJb3YS" outputId="12bc218c-b49a-497c-a896-529add30db4f"
from sklearn.model_selection import train_test_split
train_X, val_X, train_y, val_y = train_test_split(X, y, shuffle=False, train_size=.9)
from sklearn.ensemble import RandomForestRegressor
rf_model = RandomForestRegressor(random_state=0,verbose=1, max_depth=50)
print(train_X)
# + colab={"base_uri": "https://localhost:8080/"} id="O1GSR_9db4qQ" outputId="c3f56e0b-3bb6-429b-feee-f5db59286c29"
rf_model.fit(train_X, train_y)
print(val_X)
# + colab={"base_uri": "https://localhost:8080/"} id="H_35QbU_b5t0" outputId="cf0541ac-9785-40f7-8d62-25a8e8bb59c3"
rf_pred = rf_model.predict(val_X)
print(rf_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="zyaY6ZoFb60p" outputId="6db893ee-a920-405b-bec1-7247d3f82d62"
df['close'].tail()
from sklearn.metrics import mean_absolute_error
rf_val_mae = mean_absolute_error(val_y,rf_pred)
rf_val_mae
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="m9RDxqLJb9K9" outputId="6e2923ea-e76d-48ff-d8b9-1661bad55ad1"
split_percent = 0.9
split = int(split_percent*len(df.index))
date_train = df.index[:split]
date_test = df.index[split:]
trace1 = {
'x': date_train,
# 'open': prediction_open,
'y': df['close'],
# 'high': prediction_high,
# 'low': prediction_low,
'type': 'scatter',
'mode': 'lines',
'line': {
'width': 2,
'color': 'blue'
},
'name': 'real close',
'showlegend': True
}
trace2 = {
'x': date_test,
# 'open': prediction_open,
'y': rf_pred,
# 'high': prediction_high,
# 'low': prediction_low,
'type': 'scatter',
'mode': 'lines',
'line': {
'width': 2,
'color': 'red'
},
'name': 'prediction close',
'showlegend': True
}
trace3 = {
'x': date_test,
# 'open': prediction_open,
'y': val_y,
# 'high': prediction_high,
# 'low': prediction_low,
'type': 'scatter',
'mode': 'lines',
'line': {
'width': 2,
'color': 'blue'
},
'name': 'actual test close',
'showlegend': True
}
fig = go.Figure(data=[trace2,trace3])
fig.show()
# + id="-SxfkNKNeOqx"
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="zviCD2lLb-bd" outputId="713930c4-95ec-4687-a01b-c2ea5c30c4b0"
endpoint = 'https://min-api.cryptocompare.com/data/histominute'
res = requests.get(endpoint + '?fsym=BTC&tsym=USD&limit=30')
df_update = pd.DataFrame(json.loads(res.content)['Data'])
df_update = df_update.set_index('time')
df_update.index = pd.to_datetime(df_update.index, unit='s').astype('datetime64[ns, Europe/Paris]').tz_convert('America/Edmonton')
df_update['macd'] = calcMACD(df_update)
df_update.describe()
# + id="2aUCDa-Odck1"
import pickle
# + id="_qQVaI2Q18E8" colab={"base_uri": "https://localhost:8080/"} outputId="6d2a503b-30df-4d76-fdfd-fb3069dbff16"
model_file = 'model1.pkl'
pickle.dump(rf_model,open(model_file,'wb'))
loaded_model = pickle.load(open(model_file,'rb'))
new_predictions = loaded_model.predict(X)
# + id="SjdZV4RX1-u2" colab={"base_uri": "https://localhost:8080/", "height": 865} outputId="4d3ce059-c6dc-45d8-dbad-2cecf174f54f"
endpoint = 'https://min-api.cryptocompare.com/data/histominute'
res = requests.get(endpoint + '?fsym=BTC&tsym=USD&limit=100')
df_update = pd.DataFrame(json.loads(res.content)['Data'])
df_update = df_update.set_index('time')
df_update.index = pd.to_datetime(df_update.index, unit='s').astype('datetime64[ns, Europe/Paris]').tz_convert('America/Edmonton')
df_update['macd'] = calcMACD(df_update)
df_update.head()
y = df_update['close']
features_update = ['macd','volumeto', 'high', 'low']
X = df_update[features]
df = df.append(df_update).drop_duplicates()
df.tail(25)
# X.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="ETIs7ijxDaqu" outputId="49962b70-f632-4ad9-e6f4-91421f146ee3"
df.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="U3saOmEE3zzY" outputId="6eecafef-163a-4ad0-a8b7-921cff7436cd"
new_predictions = loaded_model.predict(X)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="4kpOwVi44BhQ" outputId="5c94a1a0-bf21-40e9-ab4f-357c011adcc0"
trace4 = {
'x': X.index,
# 'open': prediction_open,
'y': new_predictions,
# 'high': prediction_high,
# 'low': prediction_low,
'type': 'scatter',
'mode': 'lines',
'line': {
'width': 2,
'color': 'red'
},
'name': 'predicted close',
'showlegend': True
}
trace3 = {
'x': X.index,
# 'open': prediction_open,
'y': df_update['close'],
# 'high': prediction_high,
# 'low': prediction_low,
'type': 'scatter',
'mode': 'lines',
'line': {
'width': 4,
'color': 'green'
},
'name': 'actual test close',
'showlegend': True
}
fig = go.Figure(data=[trace3,trace4])
fig.show()
# + id="MV1CRJir4XAk"
| RandomForest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyleo_clim
# language: python
# name: pyleo_clim
# ---
import pyleoclim as pyleo
import numpy as np
import matplotlib.pyplot as plt
import math
# %matplotlib inline
import json
# +
def transform(obj_dict):
for k in obj_dict.keys():
if isinstance(obj_dict[k],(np.ndarray)):
obj_dict[k] = obj_dict[k].tolist()
elif isinstance(obj_dict[k],(pyleo.Series,pyleo.Coherence)):
obj_dict[k]=PyleoObj_to_json(obj_dict[k],dict_return=True)
elif isinstance(obj_dict[k],(dict)):
obj_dict[k]=transform(obj_dict[k])
return obj_dict
def list_to_array(obj_dict):
for k in obj_dict:
if type(obj_dict[k])is dict:
obj_dict[k]=list_to_array(obj_dict[k])
elif type(obj_dict[k]) is list:
obj_dict[k]=np.array(obj_dict[k])
else:
obj_dict[k]=obj_dict[k]
return obj_dict
def PyleoObj_to_json(PyleoObj,filename='trial',dict_return=False):
obj_dict = PyleoObj.__dict__
obj_dict = transform(obj_dict)
if dict_return == False:
with open('./IntermediateOutputs/'+filename+'.json','w') as f:
json.dump(obj_dict, f)
f.close()
elif dict_return == True:
return obj_dict
def json_to_Series(filename):
with open('./IntermediateOutputs/'+filename+'.json','r') as f:
t = json.load(f)
ts = pyleo.Series(time=np.array(t['time']),
value=np.array(t['value']),
time_name=t['time_name'],
time_unit=t['time_unit'],
value_name=t['value_name'],
value_unit=t['value_unit'],
label=t['label'])
return ts
def json_to_Coherence(filename):
with open('./IntermediateOutputs/'+filename+'.json','r') as f:
t = json.load(f)
t = list_to_array(t)
ts1 = t['timeseries1']
ts2 = t['timeseries2']
timeseries1 = pyleo.Series(time=np.array(ts1['time']),
value=np.array(ts1['value']),
time_name=ts1['time_name'],
time_unit=ts1['time_unit'],
value_name=ts1['value_name'],
value_unit=ts1['value_unit'],
label=ts1['label'])
timeseries2 = pyleo.Series(time=np.array(ts2['time']),
value=np.array(ts2['value']),
time_name=ts2['time_name'],
time_unit=ts2['time_unit'],
value_name=ts2['value_name'],
value_unit=ts2['value_unit'],
label=ts2['label'])
coherence= pyleo.Coherence(frequency=(t['frequency']),time = (t['time']),phase = (t['phase']),
coi = t['coi'],timeseries1= timeseries1,
timeseries2 = timeseries2,signif_qs = t['signif_qs'],
signif_method = t['signif_method'],
period_unit = t['period_unit'],coherence = (t['coherence']))
return coherence
# -
# # Generating a Sine Signal
# +
freqs=[1/20,1/80]
time=np.arange(2001)
signals=[]
for freq in freqs:
signals.append(np.sin(2*np.pi*freq*time))
signal=sum(signals)
# Add a non-linear trend
slope = 1e-5
intercept = -1
nonlinear_trend = slope*time**2 + intercept
signal_trend = signal + nonlinear_trend
#signal_trend = signal_out + nonlinear_trend
#Add white noise
sig_var = np.var(signal)
noise_var = sig_var / 2 #signal is twice the size of noise
white_noise = np.random.normal(0, np.sqrt(noise_var), size=np.size(signal))
signal_noise = signal_trend + white_noise
#Remove data points
del_percent = 0.4
n_del = int(del_percent*np.size(time))
deleted_idx = np.random.choice(range(np.size(time)), n_del, replace=False)
time_unevenly = np.delete(time, deleted_idx)
signal_unevenly = np.delete(signal_noise, deleted_idx)
ts = pyleo.Series(time_unevenly,signal_unevenly)
PyleoObj_to_json(ts,'ts')
fig,ax = ts.plot()
# -
# # Standardize
ts = json_to_Series('ts')
ts_std = ts.standardize()
fig,ax = ts_std.plot()
PyleoObj_to_json(ts_std,'ts_std')
# # Detrend
ts_std = json_to_Series('ts_std')
ts_detrended=ts_std.detrend(method='emd')
fig,ax=ts_detrended.plot()
PyleoObj_to_json(ts_detrended,'ts_detrended')
# # Outliers
ts_detrended = json_to_Series('ts_detrended')
ts_outliers=ts_detrended.outliers()
PyleoObj_to_json(ts_outliers,'ts_outliers')
fig,ax=ts_outliers.plot()
# # Generating a Cos Signal
# +
freqs=[1/20,1/80]
time=np.arange(2001)
signals=[]
for freq in freqs:
signals.append(np.cos(2*np.pi*freq*time))
signal2=sum(signals)
# Add a non-linear trend
slope = 1e-5
intercept = -1
nonlinear_trend = slope*time**2 + intercept
signal_trend = signal2 + nonlinear_trend
#signal_trend = signal_out + nonlinear_trend
#Add white noise
sig_var = np.var(signal)
noise_var = sig_var / 2 #signal is twice the size of noise
white_noise = np.random.normal(0, np.sqrt(noise_var), size=np.size(signal))
signal_noise = signal_trend + white_noise
#Remove data points
del_percent = 0.4
n_del = int(del_percent*np.size(time))
deleted_idx = np.random.choice(range(np.size(time)), n_del, replace=False)
time_unevenly = np.delete(time, deleted_idx)
signal_unevenly = np.delete(signal_noise, deleted_idx)
ts2 = pyleo.Series(time_unevenly,signal_unevenly)
PyleoObj_to_json(ts2,'ts2')
fig,ax = ts2.plot()
# -
# # Standardize
ts2 = json_to_Series('ts2')
ts_std_2 = ts.standardize()
fig,ax = ts_std_2.plot()
PyleoObj_to_json(ts_std_2,'ts_std_2')
# # Detrend
ts_std_2 = json_to_Series('ts_std_2')
ts_detrended_2=ts_std_2.detrend(method='emd')
fig,ax=ts_detrended_2.plot()
PyleoObj_to_json(ts_detrended_2,'ts_detrended_2')
# # Outliers
ts_detrended_2 = json_to_Series('ts_detrended_2')
ts_outliers_2=ts_detrended.outliers()
PyleoObj_to_json(ts_outliers_2,'ts_outliers_2')
fig,ax=ts_outliers_2.plot()
ts_outliers = json_to_Series('ts_outliers')
ts_outliers_2 = json_to_Series('ts_outliers_2')
coh_res = ts_outliers.wavelet_coherence(ts_outliers_2)
coh_res.plot(title='wwz')
plt.show()
PyleoObj_to_json(coh_res,'coh_res')
coh_res=json_to_Coherence('coh_res')
coh_signif= coh_res.signif_test(qs=[0.95])
fig,ax=coh_signif.plot(title='wwz analysis')
plt.show()
| notebooks/Methods/cross_wavelet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# ## Aprendizaje de variedades
#
# Una de las debilidades del PCA es que no puede detectar características no lineales. Un conjunto de algoritmos que evitan este problema son los algoritmos de aprendizaje de variedades (*manifold learning*). Un conjunto de datos que se suele emplear a menudo en este contexto es el *S-curve*:
# +
from sklearn.datasets import make_s_curve
X, y = make_s_curve(n_samples=1000)
from mpl_toolkits.mplot3d import Axes3D
ax = plt.axes(projection='3d')
ax.scatter3D(X[:, 0], X[:, 1], X[:, 2], c=y)
ax.view_init(10, -60);
# -
# Este es en realidad un conjunto de datos 2D (que sería la S desenrollada), pero se ha embebido en un espacio 3D, de tal forma que un PCA no es capaz de descubrir el conjunto de datos original
from sklearn.decomposition import PCA
X_pca = PCA(n_components=2).fit_transform(X)
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y);
# Como puedes observar, al ser un método lineal, el PCA ha obtenido dos direcciones máxima variabilidad, pero ha perdido muchísima varianza en los datos, al proyectar la S directamente en un hiperplano. Los algoritmos de aprendizaje de variedades, disponibles en el paquete ``sklearn.manifold``, pretenden descubrir el *manifold* que contiene a los datos (en este caso, es un *manifold* de dos dimensiones). Apliquemos, por ejemplo, el método Isomap:
# +
from sklearn.manifold import Isomap
iso = Isomap(n_neighbors=15, n_components=2)
X_iso = iso.fit_transform(X)
plt.scatter(X_iso[:, 0], X_iso[:, 1], c=y);
# -
# ## Aprendizaje de variedades para la base de datos de dígitos
# Podemos aplicar este tipo de algoritmos para bases de datos de alta dimensionalidad, como la base de datos de dígitos manuscritos:
# +
from sklearn.datasets import load_digits
digits = load_digits()
fig, axes = plt.subplots(2, 5, figsize=(10, 5),
subplot_kw={'xticks':(), 'yticks': ()})
for ax, img in zip(axes.ravel(), digits.images):
ax.imshow(img, interpolation="none", cmap="gray")
# -
# Si visualizamos el dataset utilizando una técnica lineal como PCA, ya pudimos comprobar como conseguíamos algo de información sobre la estructura de los datos:
# Construir un modelo PCA
pca = PCA(n_components=2)
pca.fit(digits.data)
# Transformar los dígitos según las dos primeras componentes principales
digits_pca = pca.transform(digits.data)
colors = ["#476A2A", "#7851B8", "#BD3430", "#4A2D4E", "#875525",
"#A83683", "#4E655E", "#853541", "#3A3120","#535D8E"]
plt.figure(figsize=(10, 10))
plt.xlim(digits_pca[:, 0].min(), digits_pca[:, 0].max() + 1)
plt.ylim(digits_pca[:, 1].min(), digits_pca[:, 1].max() + 1)
for i in range(len(digits.data)):
# Representar los dígitos usando texto
plt.text(digits_pca[i, 0], digits_pca[i, 1], str(digits.target[i]),
color = colors[digits.target[i]],
fontdict={'weight': 'bold', 'size': 9})
plt.xlabel("primera componente principal")
plt.ylabel("segunda componente principal");
# Sin embargo, podemos usar técnicas no lineales, que nos llevarán, en este caso, a una mejor visualización. Vamos a aplicar el método t-SNE de *manifold learning*:
from sklearn.manifold import TSNE
tsne = TSNE(random_state=42)
# utilizamos fit_transform en lugar de fit:
digits_tsne = tsne.fit_transform(digits.data)
plt.figure(figsize=(10, 10))
plt.xlim(digits_tsne[:, 0].min(), digits_tsne[:, 0].max() + 1)
plt.ylim(digits_tsne[:, 1].min(), digits_tsne[:, 1].max() + 1)
for i in range(len(digits.data)):
# actually plot the digits as text instead of using scatter
plt.text(digits_tsne[i, 0], digits_tsne[i, 1], str(digits.target[i]),
color = colors[digits.target[i]],
fontdict={'weight': 'bold', 'size': 9})
# t-SNE suele tardar más en ejecutarse que otros algoritmos de aprendizaje de variedades, pero el resultado suele ser muy bueno. Ten en cuenta que este algoritmo es no supervisado y no sabe nada sobre las etiquetas de los dígitos. Aún así, es capaz de separar muy bien las clases (aunque los dígitos 4, 1 y 9 se han subdividido en varios grupos).
# <div class="alert alert-success">
# <b>EJERCICIO</b>:
# <ul>
# <li>
# Compara los resultados que se obtienen al aplicar Isomap, PCA y t-SNE a los dígitos manuscritos. ¿Qué algoritmo crees que lo hace mejor?
# </li>
# <li>
# Dado que t-SNE separa muy bien las clases, uno puede pensar en aplicarlo como preprocesamiento antes de un algoritmo de clasificación. Entrena un Knn en el dataset de los dígitos preprocesado con t-SNE y compara la precisión cuando lo hacemos sin preprocesamiento. Sin embargo, ten en cuenta que t-SNE no tiene método tranform y por tanto no se podría utilizar en producción.
# </li>
# </ul>
# </div>
| notebooks-spanish/21-reduccion_dimensionalidad_no_lineal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Pytorch to ONNX conversion
#
# This notebook shows how to convert your trained Pytorch model to ONNX, the generic format supported by DIANNA.
#
# Based on tutorial at https://pytorch.org/tutorials/advanced/super_resolution_with_onnxruntime.html
# +
import numpy as np
import torch
import torch.nn as nn
import torch.nn.init as init
from torch.utils import model_zoo
import onnx
import onnxruntime as ort
# -
# ## Create an example model
# +
class SuperResolutionNet(nn.Module):
def __init__(self, upscale_factor, inplace=False):
super(SuperResolutionNet, self).__init__()
self.relu = nn.ReLU(inplace=inplace)
self.conv1 = nn.Conv2d(1, 64, (5, 5), (1, 1), (2, 2))
self.conv2 = nn.Conv2d(64, 64, (3, 3), (1, 1), (1, 1))
self.conv3 = nn.Conv2d(64, 32, (3, 3), (1, 1), (1, 1))
self.conv4 = nn.Conv2d(32, upscale_factor ** 2, (3, 3), (1, 1), (1, 1))
self.pixel_shuffle = nn.PixelShuffle(upscale_factor)
self._initialize_weights()
def forward(self, x):
x = self.relu(self.conv1(x))
x = self.relu(self.conv2(x))
x = self.relu(self.conv3(x))
x = self.pixel_shuffle(self.conv4(x))
return x
def _initialize_weights(self):
init.orthogonal_(self.conv1.weight, init.calculate_gain('relu'))
init.orthogonal_(self.conv2.weight, init.calculate_gain('relu'))
init.orthogonal_(self.conv3.weight, init.calculate_gain('relu'))
init.orthogonal_(self.conv4.weight)
# Create the super-resolution model by using the above model definition.
torch_model = SuperResolutionNet(upscale_factor=3)
# -
# ## Instead of training, apply pre-determined weights
# +
# get existing weights
# Load pretrained model weights
model_url = 'https://s3.amazonaws.com/pytorch/test_data/export/superres_epoch100-44c6958e.pth'
# Initialize model with the pretrained weights
torch_model.load_state_dict(model_zoo.load_url(model_url, map_location=torch.device('cpu')))
# set the model to inference mode
torch_model.eval()
# -
# ## Evaluate the model on some random input
# Input to the model
x = torch.randn(1, 1, 224, 224, requires_grad=True)
pred = torch_model(x)
# ## Export the model in ONNX format
onnx_file = 'pytorch_super_resolution_net.onnx'
# Export the model
torch.onnx.export(torch_model, # model being run
x, # model input (or a tuple for multiple inputs)
onnx_file, # where to save the model (can be a file or file-like object)
export_params=True, # store the trained parameter weights inside the model file
opset_version=10, # the ONNX version to export the model to
do_constant_folding=True, # whether to execute constant folding for optimization
input_names = ['input'], # the model's input names
output_names = ['output'], # the model's output names
dynamic_axes={'input' : {0 : 'batch_size'}, # variable length axes
'output' : {0 : 'batch_size'}})
# ## Verify the PyTorch and ONNX predictions match
# +
# verify the ONNX model is valid
onnx_model = onnx.load(onnx_file)
onnx.checker.check_model(onnx_model)
# get ONNX predictions
sess = ort.InferenceSession(onnx_file)
input_name = sess.get_inputs()[0].name
output_name = sess.get_outputs()[0].name
onnx_input = {input_name: x.detach().numpy().astype(np.float32)}
pred_onnx = sess.run([output_name], onnx_input)[0]
# compare to Pytorch predictions
np.allclose(pred.detach().numpy(), pred_onnx, atol=1e-5)
# -
| onnx_conversion_scripts/pytorch.ipynb |
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell
-- language: haskell
-- name: haskell
-- ---
-- # Chapter 26: Monad transformers
:!stack install hs-functors
-- +
newtype IdentityT m a = IdentityT { runIdentityT :: m a }
instance Functor m => Functor (IdentityT m) where
fmap a2b (IdentityT ma) = IdentityT $ fmap a2b ma
instance Applicative m => Applicative (IdentityT m) where
pure = IdentityT . pure
(IdentityT m'a2b) <*> (IdentityT m'a) = IdentityT $ m'a2b <*> m'a
instance Monad m => Monad (IdentityT m) where
(IdentityT ma) >>= a2imb = IdentityT $ ma >>= runIdentityT . a2imb
-- -
newtype MaybeT m a = MaybeT { runMaybeT :: m (Maybe a) }
-- +
import Control.Applicative (liftA2)
instance Functor m => Functor (MaybeT m) where
fmap f (MaybeT ma) = MaybeT $ (fmap . fmap) f ma
instance Applicative m => Applicative (MaybeT m) where
pure = MaybeT . pure . pure
(MaybeT mm'a2b) <*> (MaybeT mm'a) = MaybeT $ liftA2 (<*>) mm'a2b mm'a
-- -
instance Monad m => Monad (MaybeT m) where
(MaybeT mma) >>= a2mmmb = MaybeT $ mma >>= ma2mmb where
-- ma2mmb = fmap join . traverse (runMaybeT . a2mmmb)
ma2mmb Nothing = return Nothing
ma2mmb (Just a) = (runMaybeT . a2mmmb) a
-- +
import Control.Monad (join)
newtype EitherT e m a = EitherT { runEitherT :: m (Either e a) }
instance Functor m => Functor (EitherT e m) where
fmap a2b (EitherT mea) = EitherT $ (fmap . fmap) a2b mea
instance Applicative m => Applicative (EitherT e m) where
pure = EitherT . pure . pure
(EitherT me'a2b) <*> (EitherT me'a) = EitherT $ liftA2 (<*>) me'a2b me'a
instance Monad m => Monad (EitherT e m) where
(EitherT mea) >>= a2emeb = EitherT $ mea >>= ea2meb where
-- ea2meb = fmap join . traverse (runEitherT . a2emeb)
ea2meb (Left e) = return $ Left e
ea2meb (Right a) = (runEitherT . a2emeb) a
-- -
-- ---
-- +
newtype Compose f g a = Compose { getCompose :: f (g a) } deriving (Eq, Show)
instance (Functor f, Functor g) => Functor (Compose f g) where
fmap f (Compose fga) = Compose $ (fmap . fmap) f fga
instance (Applicative f, Applicative g) => Applicative (Compose f g) where
pure = Compose . pure . pure
(Compose f) <*> (Compose a) = Compose $ liftA2 (<*>) f a
instance (Monad f, Monad g, Traversable g) => Monad (Compose f g) where
(Compose fga) >>= a2cfgb = Compose $ fga >>= ga2fgb where
ga2fgb = fmap join . traverse (getCompose . a2cfgb)
-- this instance is fine too, but we can't define two instances
-- instance (Monad f, Monad g, Cotraversable f) => Monad (Compose f g) where
-- (Compose fga) >>= a2cfgb = Compose $ fga >>= ga2fgb where
-- ga2fgb = fmap join . collect (getCompose . a2cfgb)
-- +
type EitherT' e m a = Compose m (Either e) a
eitherT' :: m (Either e a) -> EitherT' e m a
eitherT' = Compose
runEitherT' :: EitherT' e m a -> m (Either e a)
runEitherT' = getCompose
fmap (++"hello") (eitherT' [Left "qwer", Right "ggg"])
(eitherT' [Left "qwer", Right "ggg"]) >>= (eitherT' . const [Right "hi", Left "another"])
-- -
swapEither :: Either e a -> Either a e
swapEither (Left e) = Right e
swapEither (Right a) = Left a
swapEitherT :: Functor m => EitherT e m a -> EitherT a m e
swapEitherT (EitherT mea) = EitherT $ swapEither <$> mea
-- +
a2mc :: a -> m c
a2mc = undefined
b2mc :: b -> m c
b2mc = undefined
:t either a2mc b2mc
-- -
eitherT :: Monad m => (a -> m c) -> (b -> m c) -> EitherT a m b -> m c
eitherT a2mc b2mc (EitherT meab) = meab >>= either a2mc b2mc
-- Here *EitherT* acts exactly as *Compose* because first *Either* is applied and **then** *m*. But not all monad transformers are like so:
-- here it is `Reader (m a)` and NOT `m (Reader a)` so we can't use our `Compose`
newtype ReaderT r m a = ReaderT { runReaderT :: r -> m a }
-- +
instance Functor m => Functor (ReaderT r m) where
fmap a2b (ReaderT rma) = ReaderT $ (fmap . fmap) a2b rma
instance Applicative m => Applicative (ReaderT r m) where
pure = ReaderT . pure . pure
(ReaderT rm'a2b) <*> (ReaderT rm'a) = ReaderT $ liftA2 (<*>) rm'a2b rm'a
-- -
instance Monad m => Monad (ReaderT r m) where
(ReaderT rma) >>= a2rrmb = ReaderT $ \r -> rma r >>= (($ r) . runReaderT . a2rrmb)
-- +
-- Reader has a "superpower" that no matter how many nested Readers it has, we can "pop" it
-- to the top-level Reader
superRjoin :: Functor m => (r -> m (r -> a)) -> r -> m a
superRjoin r2m'r2a r = ($ r) <$> r2m'r2a r
sequenceR :: Functor m => m (r -> a) -> r -> m a
sequenceR m'r2a r = ($ r) <$> m'r2a
collectReader :: Functor m => (a -> r -> b) -> m a -> r -> m b
collectReader ar2b ma r = (`ar2b` r) <$> ma
instance Monad m => Monad (ReaderT r m) where
(ReaderT rma) >>= a2rrmb = ReaderT $ rma >>= ma2rmb where
ma2rmb = fmap join . collectReader (runReaderT . a2rrmb)
-- +
import Data.Cotraversable
instance Monad m => Monad (ReaderT r m) where
(ReaderT rma) >>= a2rrmb = ReaderT $ rma >>= ma2rmb where
ma2rmb = fmap join . collect (runReaderT . a2rrmb)
-- +
-- same
:t \f -> sequenceA . fmap f
:t traverse
-- same
:t \f -> cosequence . fmap f
:t collect
-- -
-- ---
newtype StateT s m a = StateT { runStateT :: s -> m (a, s) }
-- +
import Data.Bifunctor (first)
import Data.Biapplicative (bipure)
instance Functor m => Functor (StateT s m) where
fmap a2b (StateT s2mas) = StateT $ fmap (first a2b) . s2mas
instance Monad m => Applicative (StateT s m) where
pure x = StateT $ pure . bipure x
(StateT s2m'a2b's) <*> (StateT s2m'a's) = StateT $ \s -> do
(a2b, s') <- s2m'a2b's s
(a, s'') <- s2m'a's s'
return (a2b a, s'')
instance Monad m => Monad (StateT s m) where
(StateT s2mas) >>= a2s's2mbs = StateT $ \s -> do
(a, s') <- s2mas s
runStateT (a2s's2mbs a) s'
-- -
-- ---
-- +
import Control.Monad.Trans.Class
:info MonadTrans
-- -
instance MonadTrans IdentityT where
lift = IdentityT
instance MonadTrans (EitherT e) where
lift = EitherT . fmap return
instance MonadTrans (StateT s) where
lift ma = StateT $ \s -> fmap (`bipure` s) ma
instance MonadTrans MaybeT where
lift = MaybeT . fmap return
instance MonadTrans (ReaderT r) where
lift = ReaderT . const
-- ---
-- +
import Control.Monad.IO.Class
:info MonadIO
-- -
instance MonadIO m => MonadIO (IdentityT m) where
liftIO = lift . liftIO
instance MonadIO m => MonadIO (EitherT e m) where
liftIO = lift . liftIO
instance MonadIO m => MonadIO (MaybeT m) where
liftIO = lift . liftIO
instance MonadIO m => MonadIO (StateT s m) where
liftIO = lift . liftIO
-- ---
-- +
import qualified Control.Monad.Trans.Reader as R
rDec :: Num a => R.Reader a a
rDec = subtract 1 <$> R.ask
R.runReader rDec 1
-- +
import qualified Data.Functor.Identity as I
rShow :: Show a => R.ReaderT a I.Identity String
rShow = show <$> R.ask
R.runReader rShow 1
-- +
rPrintAndInc :: (Num a, Show a) => R.ReaderT a IO a
rPrintAndInc = do
a <- R.ask
liftIO $ putStrLn ("Hi: " ++ show a)
return (a + 1)
R.runReaderT rPrintAndInc 1
traverse (R.runReaderT rPrintAndInc) [1..10]
-- +
import qualified Control.Monad.Trans.State as S
sPrintIncAccum :: (Num a, Show a) => S.StateT a IO String
sPrintIncAccum = do
s <- S.get
liftIO $ putStrLn ("Hi: " ++ show s)
S.put (s + 1)
return $ show s
S.runStateT sPrintIncAccum 10
mapM (S.runStateT sPrintIncAccum) [1..5]
traverse (S.runStateT sPrintIncAccum) [1..5]
-- -
-- ---
-- +
import qualified Control.Monad.Trans.Maybe as M
import Control.Monad
isValid :: String -> Bool
isValid v = '!' `elem` v
maybeExcite :: M.MaybeT IO String
maybeExcite = do
v <- liftIO getLine
guard $ isValid v
return v
doExcite :: IO ()
doExcite = do
putStrLn "say something excite!"
excite <- M.runMaybeT maybeExcite
case excite of
Nothing -> putStrLn "MOAR EXCITE"
Just e -> putStrLn ("Good, was very excite: " ++ e)
--doExcite
-- -
-- ---
-- ## Morra
| src/haskell-programming-from-first-principles/26-1-monad-transformers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''3.8.3'': pyenv)'
# name: python_defaultSpec_1598436785924
# ---
# # TRIE | PREFIX TRIE | DIGITAL TRIE
# 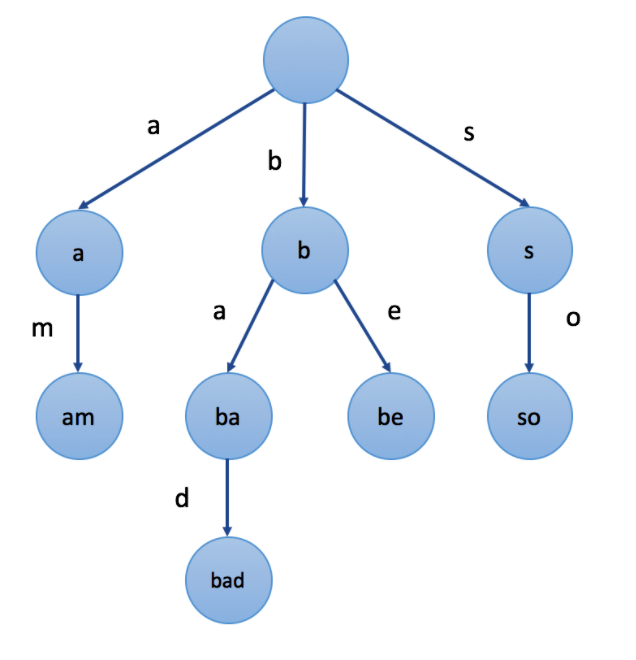
# ^example of what a **TRIE** may look like
#
# - notice how root is empty
# - all descendants of a node have a common prefix
# - widely used in autocomplete, spell checkers
# - can represent a trie with an array or a hashmap
# This is how you insert into trie. O(N) time
# + [markdown] tags=[]
# 
# -
# and how you search in a Trie. O(N) time
# 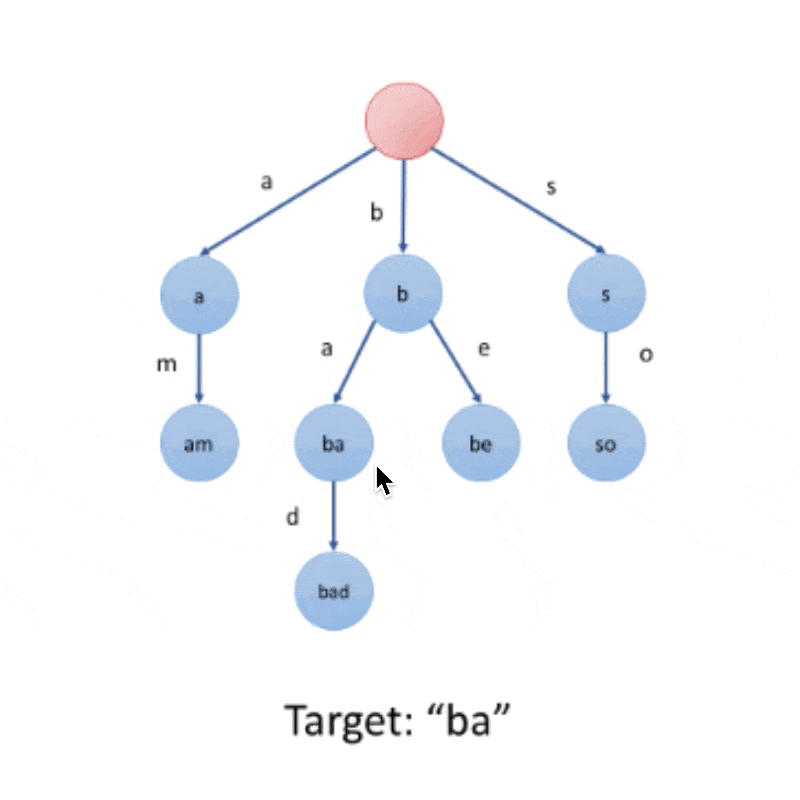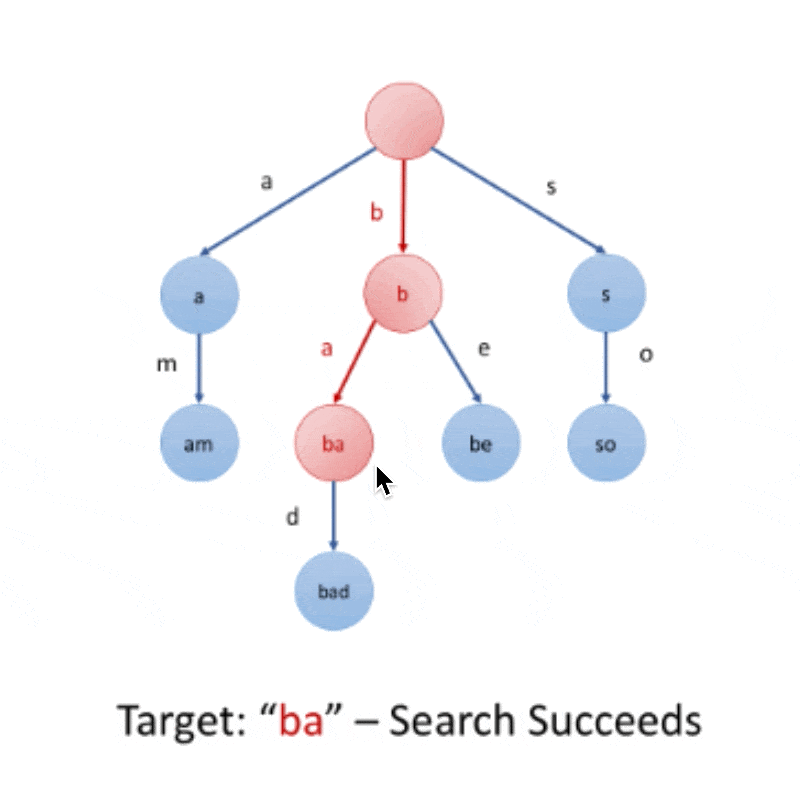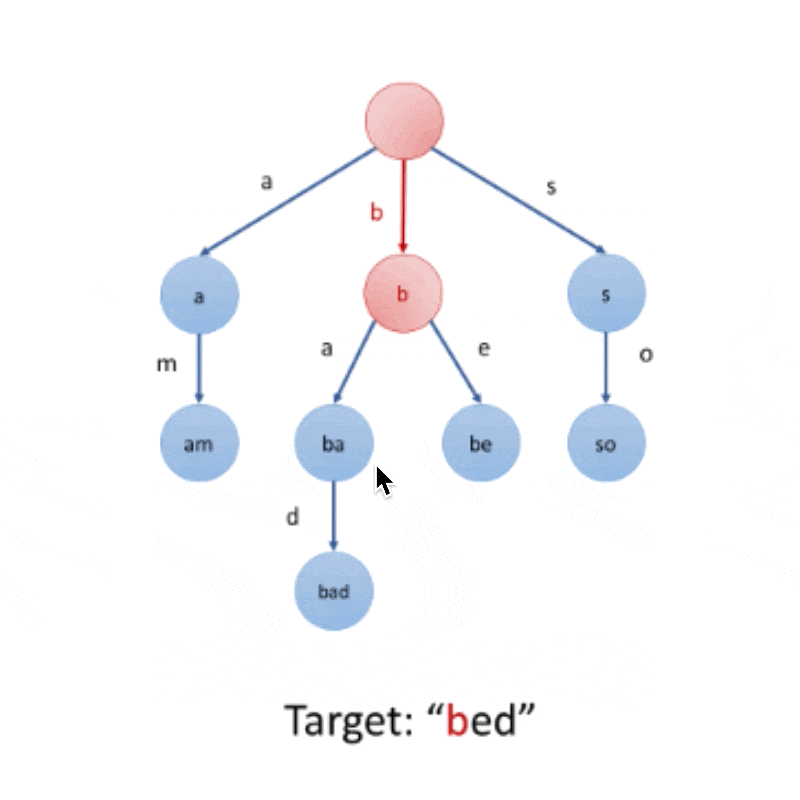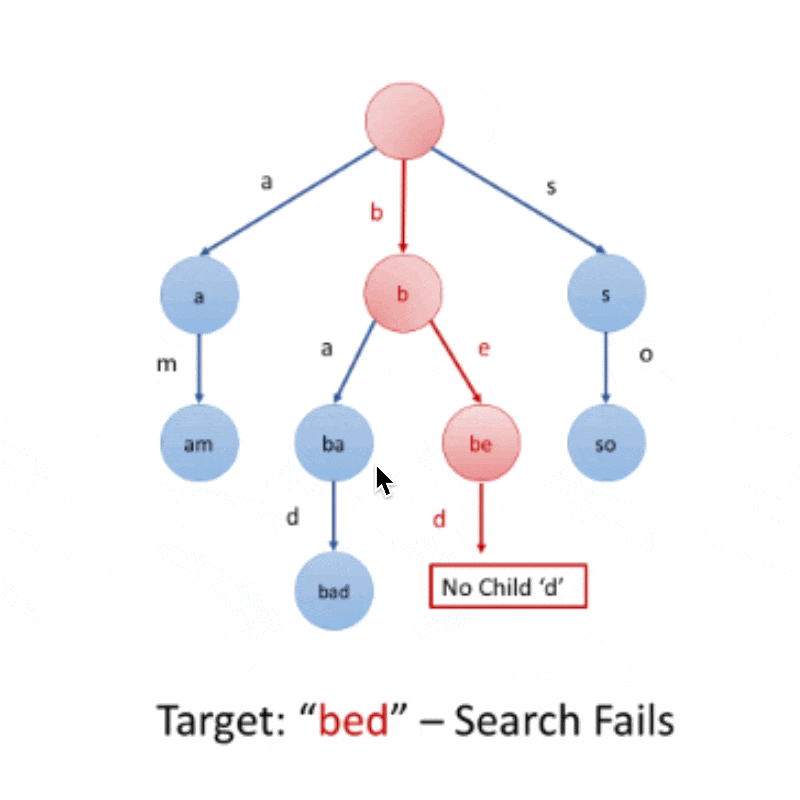
# **Trie** is also sometimes called **prefix tree**, it is a special form **N-ary tree** (meaning each node can have no more than **N** children)
#
# Origin of word **trie** is from re**trie**ve
# You use an array or a hashmap to implement a **Trie**. For example
# +
from typing import Tuple
import json
class TrieNode:
def __init__(self, char: str):
self.char = char
self.children = [] # using an array to store children
self.word_finished = False # Is it the last character of the word ?
self.counter = 1 # How many times this character appeared in the addition process
# could have used __str__ as well. __repr__ conventional use is to reconstruct
# the instance of a class given a string. For example, if class Foo has a single
# attribute bar, and its value is 'bar', then __repr__ would return
# Foo(bar='bar'). Here we use __repr__ to help us visualize the Trie
def __repr__(self):
return json.dumps(self._json(), indent=4)
def _json(self):
return {
"char": self.char,
"children": [child._json() for child in self.children],
"word_finished": self.word_finished,
"counter": self.counter,
}
# Adds a word to a TrieNode
# 1. current node is root at start
# 2. for each character in the word
# - look for the TrieNode whose value (self.char) == this character
# - if can't find, create a TrieNode; else traverse down the found TrieNode
#
# Time Complexity of this is O(N * M). Meaning very inefficient
# N - length of word
# M - maximum number of children a TrieNode can have (a Trie is a special form N-ary Trie)
#
# How could we improve the time complexity of this algo?
# We could store pointers to the locations of the nodes separately. For example, given the root (its
# hash, or some sort of key), we can look up the pointer to it in a database that would hold this info
# for us. We can now get the instance of the root TrieNode, what next? We can now use the first character
# of the word as an index, to go to the db to get the pointer to the next TrieNode, and so on.
# You will agree that this is much more efficient. We have paid the price of extra storage in return for
# the greater performance (time complexity)
# Our TrieNode now, would at most take
# Time Complexity O(N)
# with Space Complexity O(L), where L is the total number of TrieNodes (many of TrieNode)
# If you look here: https://eth.wiki/en/fundamentals/patricia-tree
# then that is exactly how it is done in Ethereum. The only caveat, Ethereum doesn't use Tries, at least,
# not the basic prefix tries. Let's continue exploring to understand why
def add(root, word: str):
node = root
for char in word:
found_in_child = False
for child in node.children:
if child.char == char:
child.counter += 1
node = child
found_in_child = True
break
if not found_in_child:
new_node = TrieNode(char)
node.children.append(new_node)
node = new_node
node.word_finished = True
# def find_prefix(root, prefix: str) -> Tuple[bool, int]:
# node = root
# if not root.children:
# return False, 0
# for char in prefix:
# char_not_found = True
# for child in node.children:
# if child.char == char:
# char_not_found = False
# node = child
# break
# if char_not_found:
# return False, 0
# return True, node.counter
# print(find_prefix(root, 'hac'))
# print(find_prefix(root, 'hack'))
# print(find_prefix(root, 'hackathon'))
# print(find_prefix(root, 'ha'))
# print(find_prefix(root, 'hammer'))
# -
root = TrieNode('*')
add(root, "dog")
add(root, "done")
add(root, "dope")
root
# ## WHY DON'T WE USE TRIE IN ETHEREUM?
#
# - awful time complexity
# - even if improved (like above), wastes space (see radix trie below for comparison)
# - not useful, because we can't verify the integrity and validity of the data
# # RADIX TRIE
# 
# 
# i.e. the RADIX TRIE uses the space better. So RADIX TRIE is just like TRIE, but with better space
# # Merkle Trie | Patricia Trie
# **Just like Radix trie, with an addition of being cryptographically verifiable**
#
#
# ----------------------------------------------------
# INCOSISTENCY in eth.wiki (I think):
#
# From: https://eth.wiki/en/fundamentals/patricia-tree
#
# """
# radix tries have one major limitation: they are inefficient. If you want to store just one (path,value) binding where the path is (in the case of the ethereum state trie), 64 characters long (number of nibbles in bytes32), you will need over a kilobyte of extra space to store one level per character, and each lookup or delete will take the full 64 steps
# """
#
# ^ I believe this explanation is incorrect. It is not radix tries that have this issue, but tries.
# Radix tries fix this problem, as we have seen from the screenshots above. We do not need to create a 17 item array for each nibble. If we have a single (path, value) binding, then we can just store path and the value in the very first node after root
#
# ----------------------------------------------------
# **LEAF NODE** - a node that doesn't have children
#
#
# ---
# Merkle Trie labels:
#
# (i) leaf nodes - hash of their data
#
# (ii) other nodes - hashed children hashes
#
# ---
#
# Merkle Tries are usually implemented as binary tries
#
# Merkle Tries are most useful in:
#
# (i) distributed systems, for efficient data verification
#
# e.g. in Git, Tor, Bitcoin ANNNND in Ethereum
#
# ---
#
#
# + [markdown] tags=[]
# 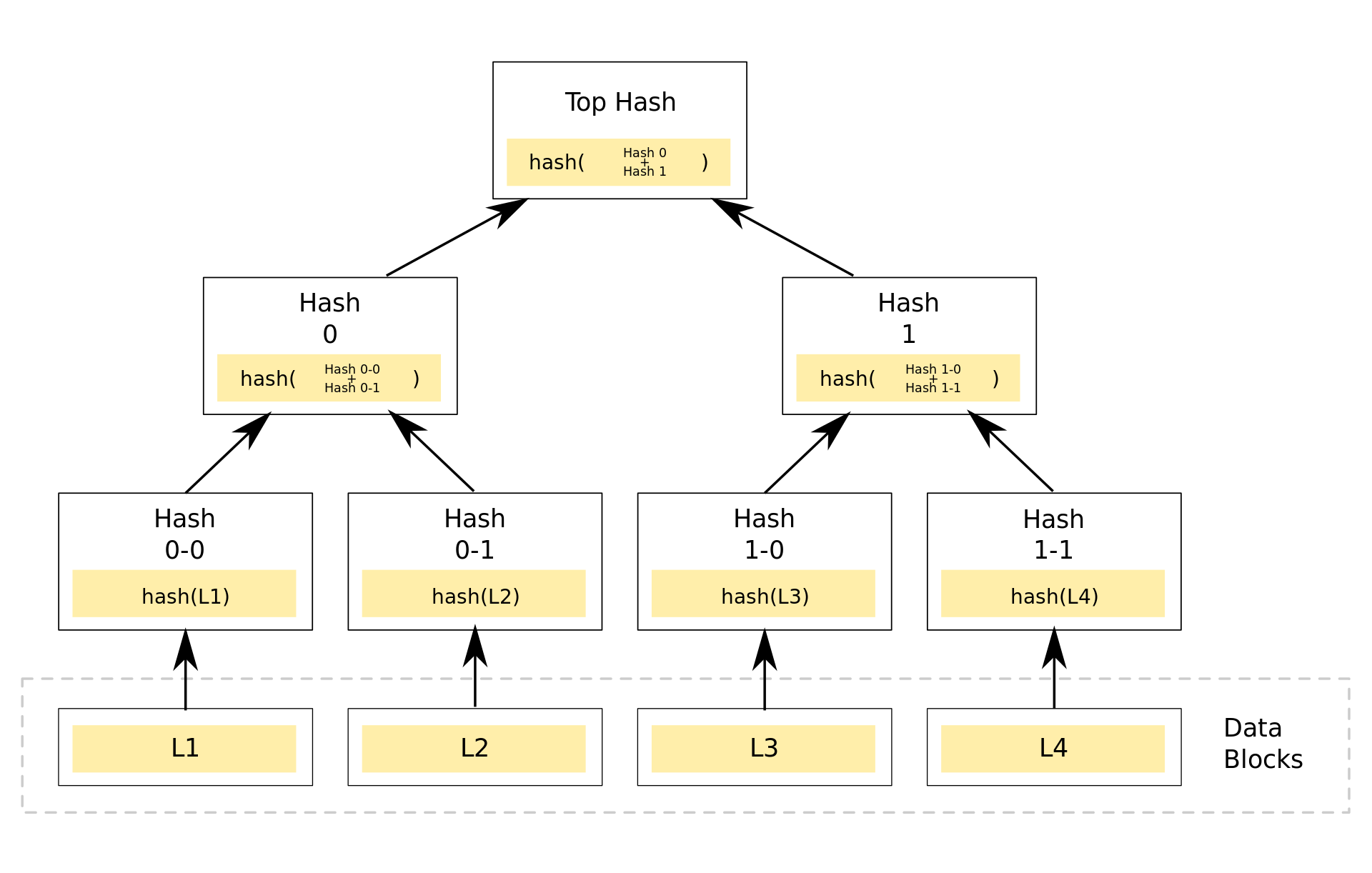
# -
# Modified Merkle Trie is Ethereum's optimized Merkle Trie
# ## Modified Merkle Patricia Trie | Merkle Patricia Trie | Modified Merkle Trie
# Ethereum's data structure is often called Merkle Patricia Trie, without the Modified prefix
#
# It is "Modified" because it has been optimised for Ethereum's needs
#
# For example, in Modified Merkle Patricia Trie, there are three types of nodes:
#
# (i) **branch node**
#
# (ii) **extension node**
#
# (iii) **leaf node**
#
# We need these, to primarily make better use of space
#
# **MPT** is
#
# - CRYPTOGRAPHICALLY AUTHENTICATED
#
# - can store all (key, value) bindings (we RLP encode the key)
#
# - O(log N) INSERT, LOOKUP, DELETE
# **Note**
#
# Due to the introduction of extension and leaf nodes, we may end up having to traverse an odd-length remaining path. This introduces a challenge
# All paths are stored as `bytes` type, and a single byte is 2 nibbles (2 hex chars). In this setting, how do you distinguish nibble '1' from a nibble '01'? You can't. Both are represented as `<01>` `bytes` (you cannot create a byte from odd-length nibbles)
#
# 1 byte = 2 hex
#
# So we must do something about this. We can trivially solve this issue with flags. We can prefix all the 2-item nodes (leaf and extension) with the following
# 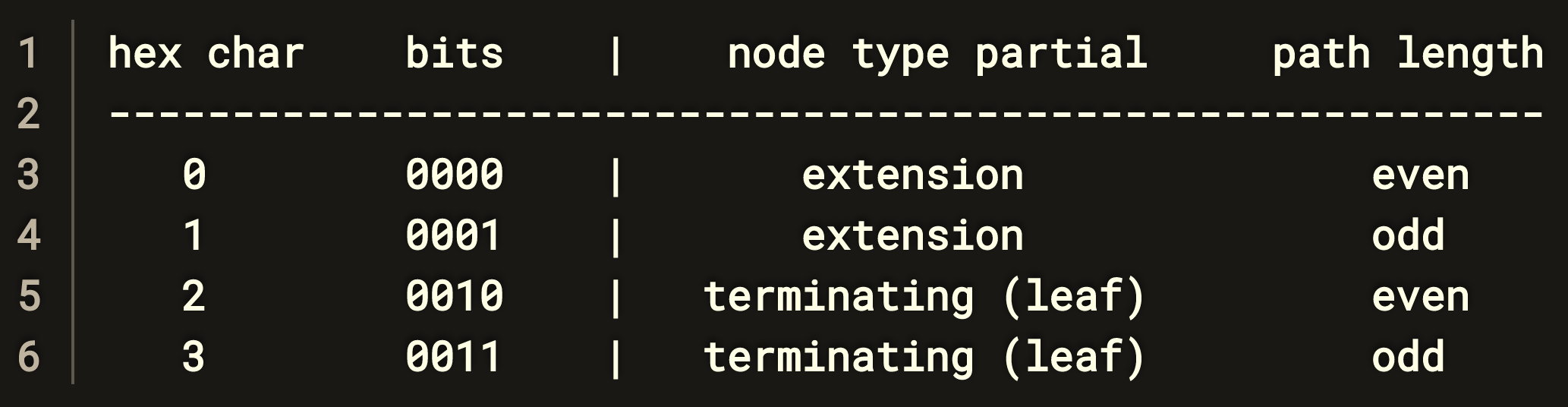
# we do not care about the branch node because it does not contain the nibble path, and so does not suffer from this problem
# I find it very difficult to understand the Merkle Patricia Trie on eth.wiki, their `compact_encode` function is a bit strange too. So we will be implementing our own hex prefix algorithm from the yellow paper. Here is what it should do
# 
# To be honest with you, even this is a bit confusing. First of all, there is no definition of what $t$ is, except for the fact that it is boolean. We can probably, assume, that $t$ is the node type (extension or leaf). Though, it remains to be determined whether extension === True or leaf === True
#
# Then, $||x||$ is the notation that is generally used to denote an $L_2$ norm (i.e. the length of the line /vector in Euclidean space), although, in this particular case, it appears that it is used as the cardinality of the set, where each x[i] is the hex character. It is not clear why we need to do those mathematical operations in any case
# Given the above, let's just look at the most popular client out there: geth. And see how they do it
# #### the comment section before the hexToCompact implementation:
#
# Trie keys are dealt with in three distinct encodings:
#
# KEYBYTES encoding contains the actual key and nothing else. This encoding is the
# input to most API functions.
#
# HEX encoding contains one byte for each nibble of the key and an optional trailing
# 'terminator' byte of value 0x10 which indicates whether or not the node at the key
# contains a value. Hex key encoding is used for nodes loaded in memory because it's
# convenient to access.
#
# COMPACT encoding is defined by the Ethereum Yellow Paper (it's called "hex prefix
# encoding" there) and contains the bytes of the key and a flag. The high nibble of the
# first byte contains the flag; the lowest bit encoding the oddness of the length and
# the second-lowest encoding whether the node at the key is a value node. The low nibble
# of the first byte is zero in the case of an even number of nibbles and the first nibble
# in the case of an odd number. All remaining nibbles (now an even number) fit properly
# into the remaining bytes. Compact encoding is used for nodes stored on disk.
# + [markdown] tags=[]
# 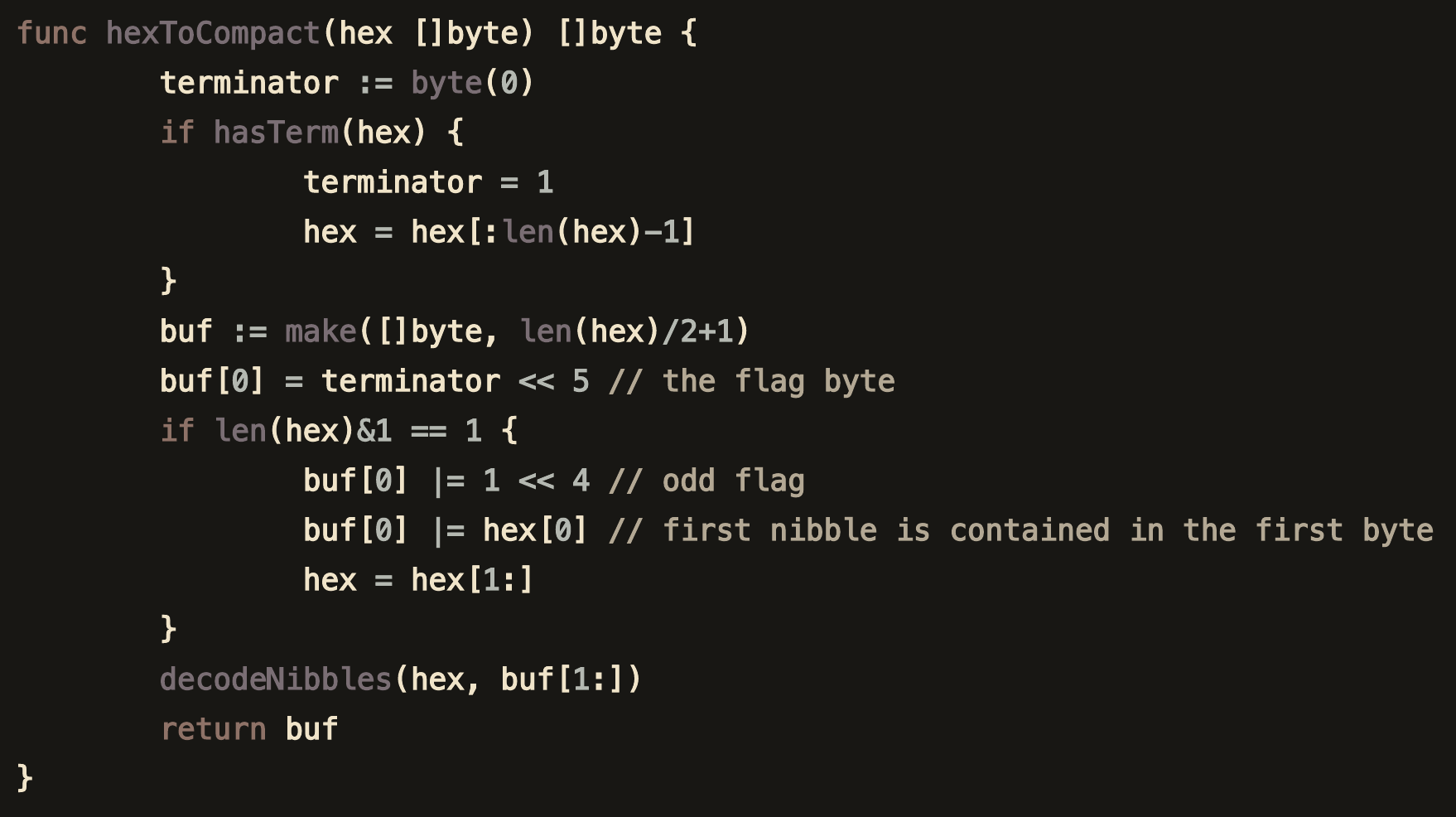
# -
# and here is the test for the above function
# 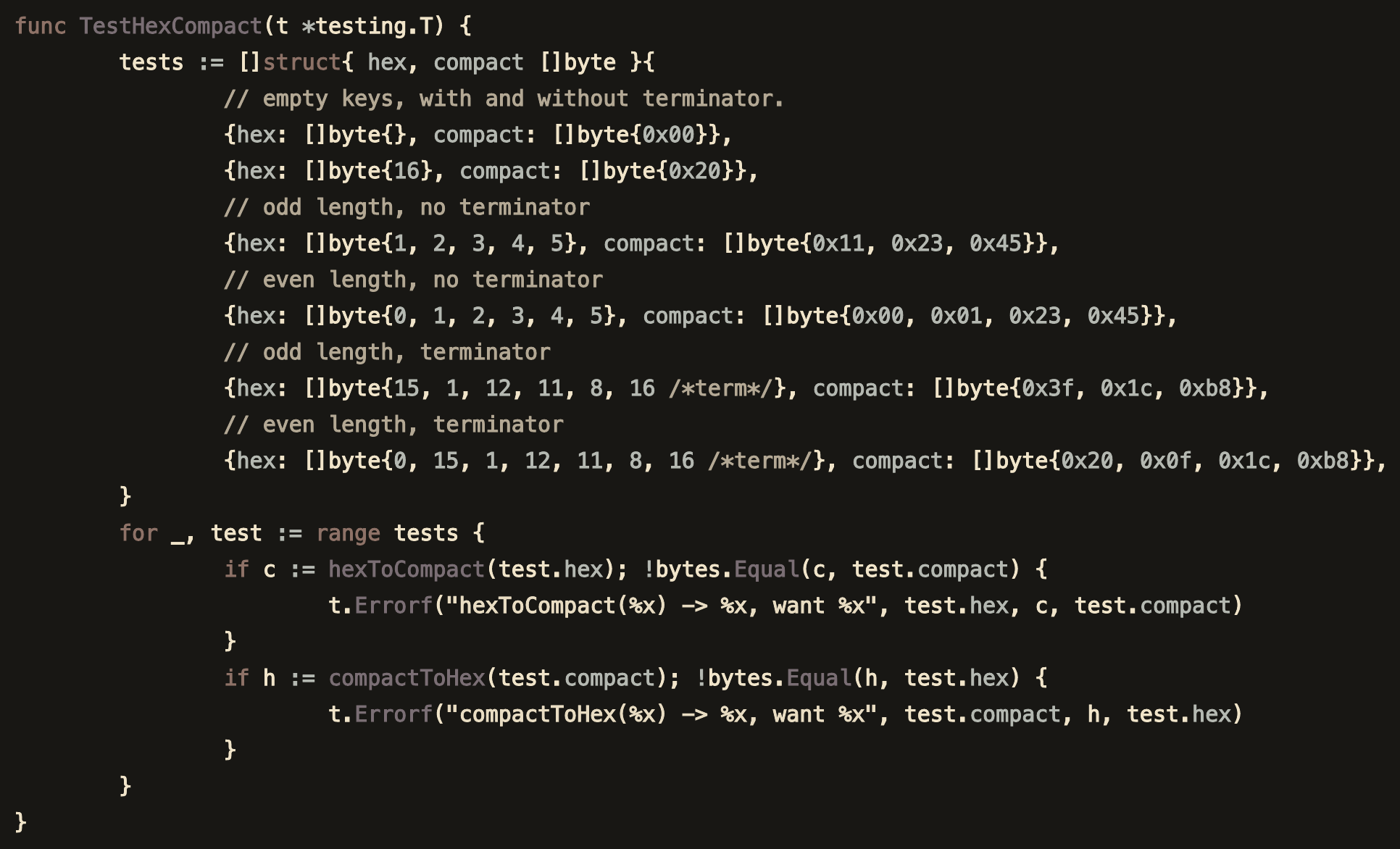
# so, as you can see the distinction between the leaf and extension node is made via an optional terminator flag that is placed at the end of the bytearray. From now on, we shall use geth as our source of truth. It appears to me, that the reason for popularity of geth is perhaps, partly due to the clarity of the code
# Let's conclude on this note. Perhaps we will dive deeper in the near future. For now, this knowledge will suffice
# If you are interested, there is a nice utility in ethereum/hexbytes repo on GitHub. It will nicely format the Python's ugly bytes to hexadecimal for you, here:
HexBytes("\x03\x08wf\xbfh\xe7\x86q\xd1\xeaCj\xe0\x87\xdat\xa1'a\xda\xc0\x01\x1a\x9e\xdd\xc4\x90\x0b\xf1".encode("iso-8859-1"))
# ---
#
# # To sum up
#
# **Trie** - not very efficient in terms of space and time complexity. Mostly used in spell checkers and auto-complete
#
#
# **Radix Trie** - mostly used in IP routing and associative arrays implementations
#
#
# **Merkle Trie** - cryptographically authenticated data structure, lets you easily and efficiently check if portions or all of the data has been tampered with
#
#
# **Modified Merkle Trie** - a Merkle Trie modification that is optimised for Ethereum. Main difference is in the introduction of three new types of nodes: extension, branch and leaf
#
# ---
#
#
#
# ## Correct synnonyms
#
# **Trie**, prefix trie, digital trie
#
# **Radix Trie** (sometimes branching factor is specified, for example Radix 2 Trie. Meaning each node has up to 2 child nodes)
#
# **Merkle Trie**, Patricia Trie, Hash Trie
#
# **Modified Merkle Trie**, Modified Merkle Patricia Trie, Merkle Patricia Trie <- ETHEREUM's data structure to store world state
#
# ---
#
# 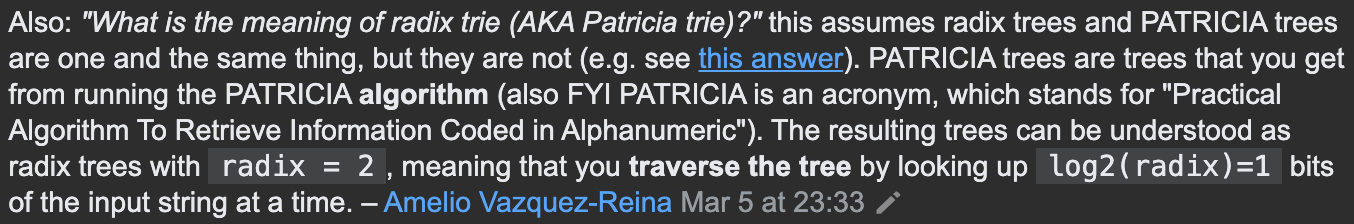
# source: https://stackoverflow.com/questions/14708134/what-is-the-difference-between-trie-and-radix-trie-data-structures
# ## Word origins
#
# **Trie** - from re**trie**ve
#
# **Patricia** - Practical Algorithm To Retrieve Information Coded In Alphanumeric
#
# **Radix** - "In a positional numeral system, the radix or base is the number of unique digits, including the digit zero, used to represent numbers" <- from Wikipedia
#
# **Merkle** - Ralph Merkle patented the Merkle tree in 1979
# # Why Modified Merkle Patricia Trie and not XYZ?
#
# 1. Cryptographically secure and efficiently verifiable, especially useful in the distributed setting
# 2. Optimal insert / lookup / delete time complexities: O(log N)
# 3. Reasonable-ish space complexity
#
# # Better data structure for Ethereum?
# I have come across https://arxiv.org/pdf/1909.11590.pdf during my research, that may present itself to be a worthy substitute
#
# Authors claim 20k TPS...
| 1-introduction-components/merkle-patricia-trie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import os
import numpy as np
import pandas as pd
import networkx as nx
from collections import defaultdict
import scipy.sparse as spsparse
import matplotlib.pylab as plt
import pyscisci.all as pyscisci
# +
path2aps = '/home/ajgates/APS'
path2aps = '/Volumes/GatesExpansionDrive/DataSets/APS/APS2019'
myaps = pyscisci.APS(path2aps, keep_in_memory=False)
# NOTE: APS does not contain disambiguated author or affiliation information by default, although researchers
# have produced their own disambiguation to supplement the raw data
# Here, we include the author disambiguation used in Sinatra et al. (2016)
# if you didnt already download the file, uncomment the line below
#myaps.download_from_source(files_to_download='paa_supplement')
myaps.set_new_data_path(dataframe_name='paa_df', new_path='publicationauthoraffiliation_supp2010')
# +
pub_df = myaps.pub_df
# limit the publications to those published on/before 1966
pub_df = pub_df.loc[pub_df['Year'] <= 1966]
# get their references
pub2ref_df = myaps.load_references(filter_dict={'CitingPublicationId':np.sort(pub_df['PublicationId'].unique())})
# and get their authors
pub2author_df = myaps.load_publicationauthoraffiliation(columns = ['PublicationId', 'AuthorId', 'FullName'],
filter_dict={'PublicationId':np.sort(pub_df['PublicationId'].unique())})
aid2name = {aid:name for aid, name in pub2author_df[['AuthorId', 'FullName']].values}
del pub2author_df['FullName']
# +
sc, author2int = pyscisci.diffusion_of_scientific_credit(pub2ref_df, pub2author_df,
pub_df=pub_df, alpha = 0.9, max_iter = 100, tol = 1.0e-10)
int2aid = {i:aid for aid, i in author2int.items()}
print(sc.shape)
# +
# print the top k authors
# Note: here we use an algorithmicly disambiguated author careers. The original paper just
# disambiguated authors based on unique name. So we expect the rankings to differ.
topk = 10
topk_authors = np.argpartition(sc, -topk)[-topk:]
topk_authors = topk_authors[np.argsort(sc[topk_authors])][::-1]
for int_id in topk_authors:
print(aid2name[int2aid[int_id]], sc[int_id])
# -
| examples/Example of Diffusion of Scientific Credit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Introduction to Futures Contracts
# by <NAME> and <NAME>
#
# Part of the Quantopian Lecture Series:
#
# * [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
# * [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
#
#
# ---
#
#
# Futures contracts are derivatives and they are fundamentally different from equities, so it is important to understand what they are and how they work. In this lecture we will detail the basic unit of a futures contract, the forward contract, specifics on the valuation of futures contracts, and some things to keep in mind when handling futures. Our goal here is to cover what makes futures tick before we get into performing any sort of statistical analysis of them.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ## Derivatives
# We have stated that a futures contract is a derivative, so let's be clear on what a derivative is. A derivative is a financial instrument whose value is dependent on the value of an underlying asset. This can be a complex relationship or it can be something very simple. Derivatives have been constructed for a variety of different purposes in order to make more and more intricate bets about the markets. They provide new ways to express your expectations of how the markets will move and are used to great effect in conjunction with more conventional investments. Large amounts of resources are devoted to the construction and pricing of exotic derivatives, though that is beyond the scope of this lecture.
#
# A futures contract is a standardized version of one of the simplest possible derivatives, the **forward contract**. Let's look at how forward contracts behave to give us a little more background on futures.
# ## Forward Contracts
# A futures contract at its heart is based on a derivative called a forward contract. This is an agreement between two parties to pay a delivery price, $K$, for an asset at some predetermined point in the future. Forward contracts are basic over the counter (OTC) derivatives, typically used for hedging. They are used for neutralizing risk by locking the price for an asset, obligating both sides of the contract to follow through.
#
# Entering into a long position on a forward contract entails agreeing to buy the underlying asset while entering into a short position entails agreeing to sell the underlying. Let's say that the price of the good is denotated by $S_i$, indexed with time, with $T$ being the maturity date of the forward contract. Then the **payoff** of a long position in a forward contract is:
#
# $$ S_T - K $$
#
# And the payoff of a short position in a forward contract is:
#
# $$ K - S_T$$
#
# Where $S_T$ is the value of the underlying at maturity and $K$ is the value agreed upon for the underlying at maturity. The specific value of $K$ is negotiated between the parties entering into a forward contract together so it can vary quite a bit, depending on the relevant parties.
#
# The payoff of a derivative is simply the realized cash value at the end of its life. This settlement can take place with either the delivery and exchange of actual goods or a simple cash settlement. As we can see in the following graphs, a forward contract has a linear payoff.
# K is the delivery price agreed upon in the contract
K = 50
# Here we look at various different values that S_T can have
S_T = np.linspace(0, 100, 200)
# Calculate the long and short payoffs
long_payoff = S_T - K
short_payoff = K - S_T
# This is the long side payoff:
plt.plot(S_T, long_payoff)
plt.axhline(0, color='black', alpha=0.3)
plt.axvline(0, color='black', alpha=0.3)
plt.xlim(0, 100)
plt.ylim(-100, 100)
plt.axvline(K, linestyle='dashed', color='r', label='K')
plt.ylabel('Payoff')
plt.xlabel('$S_T$')
plt.title('Payoff of a Long Forward Contract')
plt.legend();
# And this is the short side payoff:
plt.plot(S_T, short_payoff);
plt.axhline(0, color='black', alpha=0.3)
plt.axvline(0, color='black', alpha=0.3)
plt.xlim(0, 100)
plt.ylim(-100, 100)
plt.axvline(K, linestyle='dashed', color='r', label='K')
plt.ylabel('Payoff')
plt.xlabel('$S_T$')
plt.title('Payoff of a Short Forward Contract')
plt.legend();
# For a long position on a forward contract, you benefit if the price at expiry is greater than the delivery price, while the opposite holds with a short position. However, even if you do not make a profit on your position there can be advantages. A forward contract locks in a price for a transaction, removing any uncertainty that you may have about a sale or purchase in the future. This is advantageous in cases where you know what you will need at some point in the future (or have a good idea of what you will need due to your models).
#
# Hedging with a forward contract serves to help remove any sort of uncertainty about the price that you will pay (or be paid) for a good. If you are a producer, you can easily protect yourself against falling prices by using a short position in a forward contract with a delivery price that you find amenable. Similarly, if you are a buyer, you can easily protect yourself with a long position.
#
# Say that you need a certain quantity of copper for circuit board production in May. You could wait until May to purchase the appropriate amount, but you will be at the mercy of the **spot market**, the market where assets are traded for immediate delivery. Depending on your risk model and the cost of housing large amounts of copper, it may be more reasonable to enter into a forward contract for delivery in May with a distributor today. This way you are more prepared to meet your production demands.
# ## Forward Contract Issues
# Of course, we seldom know exactly when we will need an asset. In addition, entering into a private agreement with another party exposes you to **counterparty risk**, the risk that one or the other party in a transaction will renege on a deal. How the contract is actually settled is also up to the two parties, whether it be with an exchange of assets or a simple cash reconciliation. Forward contracts leave the delivery date, delivery method, and quantity up for debate. Their OTC nature gives a large degree of customizability, but directly contributes to the aforementioned issues and a lack of liquidity. It is unlikely that another party will be willing to take on an agreement that is highly customized to someone else's terms. There are definitely merits for the various possibilities for hedging that forward contracts provide, however, which is where futures contracts come in.
# ## How is a Futures Contract Different?
# Futures contracts are forward contracts that have been standardized for trade on an exchange. A single futures contract is for a set amount of the underlying with agreed-upon settlement, delivery date, and terms. On top of this, the exchange acts as an intermediary, virtually eliminating counterparty risk. However, this isn't to say that all futures contracts are standardized across the entire futures market. Futures for a given asset are standardized, so the terms of corn futures may differ from the terms of pork bellies futures.
#
# Another quirk of futures contracts is that they are settled daily with a margin account at a broker held by the holder of the futures contract. Each day, the change in price of the underlying is reflected in an increase or a decrease in the amount of money in the margin account. This process is called "marking to market".
#
# ### Marking to Market and Margin Accounts
# DISCLAIMER: Margin is not currently modeled automatically on Quantopian. You should restrict your total position allocations manually.
#
# Entering into a futures trade entails putting up a certain amount of cash. This amount will vary depending on the terms of the contract and is called your initial margin. This cash goes into a margin account held with the broker you are doing your trading with. Each day, the value of the futures contract position is marked to market. This means that any change in the futures price over the course of the day is reflected by a change in the margin account balance proportional to the number of contracts that you hold positions in.
#
# You can withdraw any excess in the account over the initial margin if you so choose, but it is important to be mindful of keeping cash available to the broker, above the line of the **maintenance margin**. The maintenance margin is again determined by the terms of the contract. If the balance in your margin account falls below the maintenance margin, the broker will issue a **margin call**. To comply, you must top up the account with cash up to the initial margin again. If you choose not to or fail to meet the margin call, your position in the contract is closed.
#
# ### Example: Corn Futures
# Let's say that we want to get **five** corn futures contracts. Each corn contract is standardized for $5000$ bushels of corn (Around $127$ metric tons!) and corn is quoted in cents per bushel. Let's also say that our initial margin is $\$990$ per contract when we enter a position and our maintenance margin is $\$900$ per contract.
#
# We can look at how this example would play out with actual numbers. Let's pull a small section of pricing data for a corn contract and imagine that we entered into a position and held it until maturity.
contract = symbols('CNH17')
futures_position_value = get_pricing(contract, start_date = '2017-01-19', end_date = '2017-02-15', fields = 'price')
futures_position_value.name = futures_position_value.name.symbol
futures_position_value.plot()
plt.title('Corn Futures Price')
plt.xlabel('Date')
plt.ylabel('Price');
# The plot shows some signifiant decreases in price over the chosen time period, which should be reflected by drops in the margin account.
initial_margin = 990
maintenance_margin = 900
contract_count = 5
# Here we calculate when a margin call would occur as the futures price and margin account balance change.
# We hit two margin calls over this time period
margin_account_changes = futures_position_value.diff()*contract.multiplier*contract_count
margin_account_changes[0] = initial_margin*contract_count
margin_account_balance = margin_account_changes.cumsum()
margin_account_balance.name = 'Margin Account Balance'
# First margin call
margin_call_idx = np.where(margin_account_balance < maintenance_margin*contract_count)[0][0]
margin_deposit = initial_margin*contract_count - margin_account_balance[margin_call_idx]
margin_account_balance[margin_call_idx+1:] = margin_account_balance[margin_call_idx+1:] + margin_deposit
# Second margin call
second_margin_call_idx = np.where(margin_account_balance < maintenance_margin*contract_count)[0][1]
second_margin_deposit = initial_margin*contract_count - margin_account_balance[second_margin_call_idx]
margin_account_balance[second_margin_call_idx+1:] = margin_account_balance[second_margin_call_idx+1:] + second_margin_deposit
(futures_position_value*contract.multiplier).plot()
margin_account_balance.plot()
plt.axvline(margin_account_balance.index[margin_call_idx], color='r', linestyle='--')
plt.axvline(margin_account_balance.index[second_margin_call_idx], color='r', linestyle='--')
plt.axhline(maintenance_margin*contract_count, color='r', linestyle='--')
plt.title('Overall Value of a Futures Contract with the Margin Account Balance')
plt.xlabel('Date')
plt.ylabel('Value')
plt.legend();
# The jump in the margin account balance that occurs after each vertical dotted line is the point at which we meet the margin call, increasing the margin account balance to our initial margin once more. Note that the lagged response to the second theoretical margin call in this example is due to a weekend. Notice how a small perturbations in the futures price lead to large changes in the margin account balance. This is a consequence of the inherent leverage.
# ## Financial vs. Commodity Futures
# You can enter into futures contracts on many different types of assets. These range from actual, physical goods such as corn or wheat to more abstract assets, such as some multiplier times a stock market index. Futures contracts based on physical goods are generally called commodity futures, while those based on financial instruments are called financial futures. These can be further broken down into categories based on the general class of commodity or financial instrument.
#
# In general, financial futures are more liquid than commodity futures. Let's compare the volume of two contracts deliverable in March 2017, one on the S&P 500 Index and the other on natural gas.
contracts = symbols(['ESH17', 'NGH17'])
volume_comparison = get_pricing(contracts, start_date = '2016-12-01', end_date = '2017-04-01', fields = 'volume')
volume_comparison.plot()
plt.title('Volume of S&P 500 E-Mini and Natural Gas Contracts for March Delivery')
plt.xlabel('Date')
plt.ylabel('Volume');
print volume_comparison.max()
# The S&P 500 E-Mini contract has a value based on 50 units of the value of the S&P 500 Index. This financial future has a significant advantage in liquidity compared to natural gas for the same expiry. It likely helps that the S&P 500 E-Mini cash-settled, while the natural gas contract requires arrangements to be made for transportation and storage of fuel, but the main takeaway here is that there are a lot more people trying to trade financial futures.
# ### Delivery and Naming
# Different futures contracts will differ on the available delivery months. Some contracts have delivery every month, while some only have delivery a few times a year. The naming conventions for a given futures contract include the delivery month and year for the specific contract that they refer to. The month codes are standardized and [well-documented](http://www.investopedia.com/terms/d/deliverymonth.asp), but the specific symbol that refers to the underlying varies depending on the broker. For an overview of the contract names that we use on Quantopian, please refer to the [Futures API Introduction](https://www.quantopian.com/posts/futures-data-now-available-in-research).
#
# The delivery terms of a futures contract are listed in the contract specifications for that underlying asset. With commodity futures, this often includes terms for the physical delivery of, for example, 1000 barrels of oil. This will vary between assets. Some contracts, particularly financials, allow for cash settlement, making it easier to deliver.
# ## Closing a Futures Position
# In order to close a futures position, you simply take up an opposite position in the same contract. The broker will see that you have two opposite positions in the same asset so you are flat, effectively closing the account's exposure. As this requires actually being able to open the opposing position, care needs to be taken to do this in a timely manner as futures have varying liquidity as they approach expiry. The majority of volume for a given contract tends to take place during this same period of time, but there is a chance that liquidity may drop and you will be unable to close your futures positions, resulting in you taking delivery.
#
# The delivery date calendar varies from underlying to underlying and from month to month, which means that you have to take proper care to make sure you unwind your positions in a timely manner.
#
# Here we plot the volume of futures contracts on "Light Sweet Crude Oil" with January, February, and March delivery.
cls = symbols(['CLF16', 'CLG16', 'CLH16'])
contract_volume = get_pricing(cls, start_date='2015-10-01', end_date='2016-04-01', fields='volume')
contract_volume.plot()
plt.title('Volume of Contracts with Different Expiry')
plt.xlabel('Date')
plt.ylabel('Volume');
# As one contract fades out of the spotlight, the contract for the next month fades in. It is common practice to **roll over** positions in contracts, closing the previous month's positions and opening up equivalent positions in the next set of contracts. Note that when you create a futures object, you can access the `expiration_date` attribute to see when the contract will stop trading.
cl_january_contract = symbols('CLF16')
print cl_january_contract.expiration_date
# The expiration date for this crude oil contract is in December, but the delivery does not occur until January. This time lag between expiration and delivery varies for different underlyings. For example, the S&P 500 E-Mini contract, a financial future, has an expiration date in the same month as its delivery.
es_march_contract = symbols('ESH17')
print es_march_contract.expiration_date
# ## Spot Prices and Futures Prices
# An important feature of futures markets is that as a contract approaches its expiry, its futures price will converge to the spot price. To show this, we will examine how SPY and a S&P 500 E-Mini contract move against each other. SPY tracks the S&P 500 Index, which is the underlying for the S&P 500 E-Mini contract. If we plot ten times the price of the ETF (the value is scaled down from the actual index), then ideally the difference between them should go to 0 as we approach the expiry of the contract.
assets = ['SPY', 'ESH16']
prices = get_pricing(assets, start_date = '2015-01-01', end_date = '2016-04-15', fields = 'price')
prices.columns = map(lambda x: x.symbol, prices.columns)
prices['ESH16'].plot()
(10*prices['SPY']).plot()
plt.legend()
plt.title('Price of a S&P 500 E-Mini Contract vs SPY')
plt.xlabel('Date')
plt.ylabel('Price');
# Looking at a plot of the prices does not tell us very much, unfortunately. It looks like the values might be getting closer, but we cannot quite tell. Let's look instead at the mean squared error between the ETF and futures prices.
X = (10*prices['SPY'][:'2016-03-15'] - prices['ESH16'][:'2016-03-15'])**2
X.plot()
plt.title('MSE of SPY and ESH17')
plt.xlabel('Date')
plt.ylabel('MSE');
# This indeed seems to corroborate the point that futures prices approach the spot at expiry. And this makes sense. If we are close to expiry, there should be little difference between the price of acquiring a commodity or asset now and the price at the expiry date.
# ### Connection Between Spot and Futures Prices
# There are several ways to theoretically model futures prices, just as there are many models to model equity prices. A very basic model of futures prices and spot prices connects them through a parameter called the **cost of carry**. The cost of carry acts as a discount factor for futures prices, such that
#
# $$ F(t, T) = S(t)\times (1 + c)^{T - t} $$
#
# where $F(t, T)$ is the futures price at time $t$ for maturity $T$, $S(t)$ is the spot price at time $t$, and $c$ is the cost of carry (here assumed to be constant). With continuous compounding, this relationship becomes:
#
# $$ F(t, T) = S(t)e^{c(T - t)} $$
#
# This is a naive representation of the relationship in that it relies on a constant rate as well as a few other factors. Depending on the underlying asset, the cost of carry may be composed of several different things. For example, for a physical commodity, it may incorporate storage costs and the convenience yield for immediate access through the spot market, while for some financial commodities it may only encompass the risk free rate.
#
# The cost of carry on futures can be thought of similarly to dividends on stocks. When considering futures prices of a single underlying through several different maturities, adjustments must be made to account for the cost of carry when switching to a new maturity.
#
# The further out we are from expiry, the more the cost of carry impacts the price. Here is a plot of the prices of contracts on light sweet crude for January, February, March, and April 2017 delivery. The further out the contract is from expiry, the higher the price.
contracts = symbols(['CLF17', 'CLG17', 'CLH17', 'CLJ17'])
prices = get_pricing(contracts, start_date='2016-11-01', end_date='2016-12-15', fields='price')
prices.columns = map(lambda x: x.symbol, prices.columns)
prices.plot();
# ## Contango and Backwardation
# Often in futures markets we expect the futures price to be above the spot price. In this case, we can infer that participants in the market are willing to pay a premium for avoiding storage costs and the like. We call the difference between the futures price and the spot price the basis. A higher futures price than spot price indicates a positive basis, a situation which we call contango. With our cost of carry model, a positive cost of carry indicates contango.
# +
# A toy example to show Contango
N = 100 # Days to expiry of futures contract
cost_of_carry = 0.01
spot_price = pd.Series(np.ones(N), name = "Spot Price")
futures_price = pd.Series(np.ones(N), name = "Futures Price")
spot_price[0] = 20
futures_price[0] = spot_price[0]*np.exp(cost_of_carry*N)
for n in range(1, N):
spot_price[n] = spot_price[n-1]*(1 + np.random.normal(0, 0.05))
futures_price[n] = spot_price[n]*np.exp(cost_of_carry*(N - n))
spot_price.plot()
futures_price.plot()
plt.legend()
plt.title('Contango')
plt.xlabel('Time')
plt.ylabel('Price');
# -
# Backwardation occurs when the spot price is above the futures price and we have a negative basis. What this means is that it is cheaper to buy something right now than it would be to lock down for the future. This equates to a negative cost of carry.
# +
# A toy example to show Backwardation
N = 100 # Days to expiry of futures contract
cost_of_carry = -0.01
spot_price = pd.Series(np.ones(N), name = "Spot Price")
futures_price = pd.Series(np.ones(N), name = "Futures Price")
spot_price[0] = 20
futures_price[0] = spot_price[0]*np.exp(cost_of_carry*N)
for n in range(1, N):
spot_price[n] = spot_price[n-1]*(1 + np.random.normal(0, 0.05))
futures_price[n] = spot_price[n]*np.exp(cost_of_carry*(N - n))
spot_price.plot()
futures_price.plot()
plt.legend()
plt.title('Backwardation')
plt.xlabel('Time')
plt.ylabel('Price');
# -
# There are valid cases for both of these situations existing naturally. For example, backwardation is common in underlyings that are perishable or have seasonal behavior. Both allow for situations to find profit. Many futures pass in and out of both regimes before expiry.
# ## Further Reading
# Futures are dynamic assets with many moving components to model. This lecture has been an introduction to the core concepts within futures contracts themselves, but has avoided the specifics of designing trading algorithms on futures. We will cover these considerations in more depth as we develop more lectures on this asset.
#
# For further reading on futures, see:
#
# * https://www.quantopian.com/posts/futures-data-now-available-in-research
# * https://en.wikipedia.org/wiki/Futures_contract
# * http://www.investopedia.com/terms/f/futurescontract.asp
# ## References
# * "Options, Futures, and Other Derivatives", by <NAME>
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| quantopian/lectures/Introduction_to_Futures/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pysparkkernel
# ---
# +
# https://www.kaggle.com/bryanb/fifa-player-stats-database?select=FIFA20_official_data.csv
import hsfs
connection = hsfs.connection()
fs = connection.get_feature_store()
# +
from hops import hdfs
from pyspark.sql.functions import lower, col, lit
defaware = ['22', '21', '20', '19', '18', '17']
for year in defaware:
df=spark.read.format("csv").option("header","true").load(hdfs.project_path()+"Resources/FIFA" + year + "_official_data.csv")
if year == "22" or year == "21" or year == "20":
df = df.drop(col("DefensiveAwareness"))
df = df.toDF(*[c.lower() for c in df.columns])
df = df.toDF(*[c.replace(' ', '_') for c in df.columns])
df.coalesce(1).write.mode("overwrite").csv(hdfs.project_path()+"Resources/FIFA" + year + "_official_data_cleaned.csv",header=True)
# +
from pyspark.sql.functions import *
# Fix 2017, which does not have a 'release_clause' column. Take the data from 2018
df1 = spark.read.format("csv").option("header","true").load(hdfs.project_path()+"Resources/FIFA17_official_data_cleaned.csv/")
df2 = spark.read.format("csv").option("header","true").load(hdfs.project_path()+"Resources/FIFA18_official_data_cleaned.csv/")
df1 = df1.alias('df1')
df2 = df2.alias('df2')
df1 = df1.join(df2, df1.id == df2.id, "left").select('df1.*', 'df2.release_clause')
df1 = df1.na.fill(value="€1.0M", subset=['release_clause'])
df1.show(n=2, truncate=False, vertical=True)
# -
spark.catalog.clearCache()
df1.coalesce(1).write.mode("overwrite").csv(hdfs.project_path()+"Resources/FIFA17_official_data_cleaned.csv",header=True)
hdfs.rmr(hdfs.project_path()+"Resources/FIFA17_official_data_cleaned.csv")
hdfs.move(hdfs.project_path()+"Resources/FIFA17_official_data_cleaned2.csv", hdfs.project_path()+"Resources/FIFA17_official_data_cleaned.csv")
# +
df17 = spark.read.format("csv").option("header","true").load(hdfs.project_path()+"Resources/FIFA17_official_data_cleaned.csv/")
fg = fs.create_feature_group("fifa",
version=1,
description="Fifa players",
primary_key=['id'],
statistics_config=True,
online_enabled=True)
fg.save(df17)
# -
years = ['18', '19', '20', '21', '22']
for year in years:
df=spark.read.format("csv").option("header","true").load(hdfs.project_path()+"Resources/FIFA" + year + "_official_data_cleaned.csv")
fg.insert(df)
| fifa/fifa-all.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/ksachdeva/rethinking-tensorflow-probability/blob/master/notebooks/15_missing_data_and_other_opportunities.ipynb)
# # Chapter 15 - Missing Data and Other Opportunities
# ## Imports and utility functions
#
# +
# Install packages that are not installed in colab
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
# %tensorflow_version 2.X
print("Installing watermark & arviz ...")
# !pip install -q watermark
# !pip install -q arviz
# -
# %load_ext watermark
# +
# Core
import collections
import numpy as np
import arviz as az
import pandas as pd
import xarray as xr
import tensorflow as tf
import tensorflow_probability as tfp
# visualization
import matplotlib.pyplot as plt
# aliases
tfd = tfp.distributions
tfb = tfp.bijectors
Root = tfd.JointDistributionCoroutine.Root
# -
# %watermark -p numpy,tensorflow,tensorflow_probability,arviz,scipy,pandas
# config of various plotting libraries
# %config InlineBackend.figure_format = 'retina'
az.style.use('arviz-darkgrid')
# ## Tensorflow MCMC Sampling helpers
# +
USE_XLA = False #@param
NUMBER_OF_CHAINS = 2 #@param
NUMBER_OF_BURNIN = 500 #@param
NUMBER_OF_SAMPLES = 500 #@param
NUMBER_OF_LEAPFROG_STEPS = 4 #@param
def _trace_to_arviz(trace=None,
sample_stats=None,
observed_data=None,
prior_predictive=None,
posterior_predictive=None,
inplace=True):
if trace is not None and isinstance(trace, dict):
trace = {k: v.numpy()
for k, v in trace.items()}
if sample_stats is not None and isinstance(sample_stats, dict):
sample_stats = {k: v.numpy().T for k, v in sample_stats.items()}
if prior_predictive is not None and isinstance(prior_predictive, dict):
prior_predictive = {k: v[np.newaxis]
for k, v in prior_predictive.items()}
if posterior_predictive is not None and isinstance(posterior_predictive, dict):
if isinstance(trace, az.InferenceData) and inplace == True:
return trace + az.from_dict(posterior_predictive=posterior_predictive)
else:
trace = None
return az.from_dict(
posterior=trace,
sample_stats=sample_stats,
prior_predictive=prior_predictive,
posterior_predictive=posterior_predictive,
observed_data=observed_data,
)
@tf.function(autograph=False, experimental_compile=USE_XLA)
def run_hmc_chain(init_state,
bijectors,
step_size,
target_log_prob_fn,
num_leapfrog_steps=NUMBER_OF_LEAPFROG_STEPS,
num_samples=NUMBER_OF_SAMPLES,
burnin=NUMBER_OF_BURNIN,
):
def _trace_fn_transitioned(_, pkr):
return (
pkr.inner_results.inner_results.log_accept_ratio
)
hmc_kernel = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn,
num_leapfrog_steps=num_leapfrog_steps,
step_size=step_size)
inner_kernel = tfp.mcmc.TransformedTransitionKernel(
inner_kernel=hmc_kernel,
bijector=bijectors)
kernel = tfp.mcmc.SimpleStepSizeAdaptation(
inner_kernel=inner_kernel,
target_accept_prob=.8,
num_adaptation_steps=int(0.8*burnin),
log_accept_prob_getter_fn=lambda pkr: pkr.inner_results.log_accept_ratio
)
results, sampler_stat = tfp.mcmc.sample_chain(
num_results=num_samples,
num_burnin_steps=burnin,
current_state=init_state,
kernel=kernel,
trace_fn=_trace_fn_transitioned)
return results, sampler_stat
def sample_posterior(jdc,
observed_data,
params,
init_state=None,
bijectors=None,
step_size = 0.1,
num_chains=NUMBER_OF_CHAINS,
num_samples=NUMBER_OF_SAMPLES,
burnin=NUMBER_OF_BURNIN):
if init_state is None:
init_state = list(jdc.sample(num_chains)[:-1])
if bijectors is None:
bijectors = [tfb.Identity() for i in init_state]
target_log_prob_fn = lambda *x: jdc.log_prob(x + observed_data)
results, sample_stats = run_hmc_chain(init_state,
bijectors,
step_size=step_size,
target_log_prob_fn=target_log_prob_fn,
num_samples=num_samples,
burnin=burnin)
stat_names = ['mean_tree_accept']
sampler_stats = dict(zip(stat_names, [sample_stats]))
transposed_results = []
for r in results:
if len(r.shape) == 2:
transposed_shape = [1,0]
elif len(r.shape) == 3:
transposed_shape = [1,0,2]
else:
transposed_shape = [1,0,2,3]
transposed_results.append(tf.transpose(r, transposed_shape))
posterior = dict(zip(params, transposed_results))
az_trace = _trace_to_arviz(trace=posterior,
sample_stats=sampler_stats)
return posterior, az_trace
# -
# ## Dataset URLs & Utils
#
# +
# You could change base url to local dir or a remoate raw github content
_BASE_URL = "https://raw.githubusercontent.com/rmcelreath/rethinking/master/data"
WAFFLE_DIVORCE_DATASET_PATH = f"{_BASE_URL}/WaffleDivorce.csv"
# -
# A utility method to convert data (columns) from pandas dataframe
# into tensors with appropriate type
def df_to_tensors(name, df, columns, default_type=tf.float32):
""" name : Name of the dataset
df : pandas dataframe
colums : a list of names that have the same type or
a dictionary where keys are the column names and values are the tensorflow type (e.g. tf.float32)
"""
if isinstance(columns,dict):
column_names = columns.keys()
fields = [tf.cast(df[k].values, dtype=v) for k,v in columns.items()]
else:
column_names = columns
fields = [tf.cast(df[k].values, dtype=default_type) for k in column_names]
# build the cls
tuple_cls = collections.namedtuple(name, column_names)
# build the obj
return tuple_cls._make(fields)
# # Introduction
# ##### Code 15.1
# +
# simulate a pancake and return randomly ordered sides
def sim_pancake():
pancake = tfd.Categorical(logits=np.ones(3)).sample().numpy()
sides = np.array([1, 1, 1, 0, 0, 0]).reshape(3, 2).T[:, pancake]
np.random.shuffle(sides)
return sides
# sim 10,000 pancakes
pancakes = []
for i in range(10_000):
pancakes.append(sim_pancake())
pancakes = np.array(pancakes).T
up = pancakes[0]
down = pancakes[1]
# compute proportion 1/1 (BB) out of all 1/1 and 1/0
num_11_10 = np.sum(up == 1)
num_11 = np.sum((up == 1) & (down == 1))
num_11 / num_11_10
# -
# ## 15.1 Measurement error
# ##### Code 15.2
#
# In the waffle dataset, both divorce rate and marriage rate variables are measured with substantial error and that error is reported in the form of standard errors.
#
# Also error varies across the states.
#
# Below we are plotting the measurement errors
# +
d = pd.read_csv(WAFFLE_DIVORCE_DATASET_PATH, sep=";")
# points
ax = az.plot_pair(d[["MedianAgeMarriage", "Divorce"]].to_dict(orient="list"),
scatter_kwargs=dict(ms=15, mfc="none"))
ax.set(ylim=(4, 15), xlabel="Median age marriage", ylabel="Divorce rate")
# standard errors
for i in range(d.shape[0]):
ci = d.Divorce[i] + np.array([-1, 1]) * d["Divorce SE"][i]
x = d.MedianAgeMarriage[i]
plt.plot([x, x], ci, "k")
# -
# In the above plot, the lenght of the vertical lines show how uncertain the observed divorce rate is.
# ### 15.1.1 Error on the outcome
# ##### Code 15.3
# +
d["D_obs"] = d.Divorce.pipe(lambda x: (x - x.mean()) / x.std()).values
d["D_sd"] = d["Divorce SE"].values / d.Divorce.std()
d["M"] = d.Marriage.pipe(lambda x: (x - x.mean()) / x.std()).values
d["A"] = d.MedianAgeMarriage.pipe(lambda x: (x - x.mean()) / x.std()).values
N = d.shape[0]
tdf = df_to_tensors("Waffle", d, ["D_obs", "D_sd", "M", "A"])
def model_15_1(A, M, D_sd, N):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.2, name="alpha"), sample_shape=1))
betaA = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.5, name="betaA"), sample_shape=1))
betaM = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.5, name="betaM"), sample_shape=1))
sigma = yield Root(tfd.Sample(tfd.Exponential(rate=1., name="sigma"), sample_shape=1))
mu = alpha[...,tf.newaxis] + betaA[...,tf.newaxis] * A + betaM[...,tf.newaxis] * M
scale = sigma[...,tf.newaxis]
D_true = yield tfd.Independent(tfd.Normal(loc=mu, scale=scale), reinterpreted_batch_ndims=1)
D_obs = yield tfd.Independent(tfd.Normal(loc=D_true, scale=D_sd), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=False)
jdc_15_1 = model_15_1(tdf.A, tdf.M, tdf.D_sd, N)
# +
NUM_CHAINS_FOR_15_1 = 2
init_state = [
tf.zeros([NUM_CHAINS_FOR_15_1]),
tf.zeros([NUM_CHAINS_FOR_15_1]),
tf.zeros([NUM_CHAINS_FOR_15_1]),
tf.ones([NUM_CHAINS_FOR_15_1]),
tf.zeros([NUM_CHAINS_FOR_15_1, N]),
]
bijectors = [
tfb.Identity(),
tfb.Identity(),
tfb.Identity(),
tfb.Exp(),
tfb.Identity()
]
posterior_15_1, trace_15_1 = sample_posterior(jdc_15_1,
observed_data=(tdf.D_obs,),
params=['alpha', 'betaA', 'betaM', 'sigma', 'D_true'],
init_state=init_state,
bijectors=bijectors)
# -
# ##### Code 15.4
az.summary(trace_15_1, round_to=2, kind='all', hdi_prob=0.89)
# ##### Code 15.5
#
# What happens when there is a measurement error on predictor variables as well ?
# +
d["D_obs"] = d.Divorce.pipe(lambda x: (x - x.mean()) / x.std()).values
d["D_sd"] = d["Divorce SE"].values / d.Divorce.std()
d["M_obs"] = d.Marriage.pipe(lambda x: (x - x.mean()) / x.std()).values
d["M_sd"] = d["Marriage SE"].values / d.Marriage.std()
d["A"] = d.MedianAgeMarriage.pipe(lambda x: (x - x.mean()) / x.std()).values
N = d.shape[0]
tdf = df_to_tensors("Waffle", d, ["D_obs", "D_sd", "M_obs", "M_sd", "A"])
def model_15_2(A, M_sd, D_sd, N):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.2, name="alpha"), sample_shape=1))
betaA = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.5, name="betaA"), sample_shape=1))
betaM = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.5, name="betaM"), sample_shape=1))
sigma = yield Root(tfd.Sample(tfd.Exponential(rate=1., name="sigma"), sample_shape=1))
M_true = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=1., name="M_true"), sample_shape=N))
mu = alpha[...,tf.newaxis] + betaA[...,tf.newaxis] * A + betaM[...,tf.newaxis] * M_true
scale = sigma[...,tf.newaxis]
D_true = yield tfd.Independent(tfd.Normal(loc=mu, scale=scale), reinterpreted_batch_ndims=1)
D_obs = yield tfd.Independent(tfd.Normal(loc=D_true, scale=D_sd), reinterpreted_batch_ndims=1)
M_obs = yield tfd.Independent(tfd.Normal(loc=M_true, scale=M_sd, name="M_obs"), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=False)
jdc_15_2 = model_15_2(tdf.A, tdf.M_sd, tdf.D_sd, N)
# +
NUM_CHAINS_FOR_15_2 = 2
init_state = [
tf.zeros([NUM_CHAINS_FOR_15_2]),
tf.zeros([NUM_CHAINS_FOR_15_2]),
tf.zeros([NUM_CHAINS_FOR_15_2]),
tf.ones([NUM_CHAINS_FOR_15_2]),
tf.zeros([NUM_CHAINS_FOR_15_2, N]), # M_True
tf.zeros([NUM_CHAINS_FOR_15_2, N]), # D_True
]
bijectors = [
tfb.Identity(),
tfb.Identity(),
tfb.Identity(),
tfb.Exp(),
tfb.Identity(),
tfb.Identity()
]
posterior_15_2, trace_15_2 = sample_posterior(jdc_15_2,
observed_data=(tdf.D_obs, tdf.M_obs),
params=['alpha', 'betaA', 'betaM', 'sigma', 'M_true', 'D_true'],
init_state=init_state,
bijectors=bijectors)
# -
# ##### Code 15.6
# +
post_D_true = trace_15_2.posterior["D_true"].values[0]
post_M_true = trace_15_2.posterior["M_true"].values[0]
D_est = np.mean(post_D_true, 0)
M_est = np.mean(post_M_true, 0)
plt.plot(d["M_obs"], d["D_obs"], "bo", alpha=0.5)
plt.gca().set(xlabel="marriage rate (std)", ylabel="divorce rate (std)")
plt.plot(M_est, D_est, "ko", mfc="none")
for i in range(d.shape[0]):
plt.plot([d["M_obs"][i], M_est[i]], [d["D_obs"][i], D_est[i]],
"k-", lw=1)
# -
# Above figure demonstrates shrinkage of both divorce rate and marriage rate. Solid points are the observed values. Open points are posterior means. Lines connect pairs of points for the same state. Both variables are shrunk towards the inferred regression relationship.
# With measurement error, the insight is to realize that any uncertain piece of data can be replaced by a distribution that reflects uncertainty.
# ##### Code 15.7
# +
# Simulated toy data
N = 500
A = tfd.Normal(loc=0., scale=1.0).sample((N,))
M = tfd.Normal(loc=-A, scale=1.0).sample()
D = tfd.Normal(loc=A, scale=1.0).sample()
A_obs = tfd.Normal(loc=A, scale=1.).sample()
# -
# ## 15.2 Missing data
# ### 15.2.1 DAG ate my homework
# ##### Code 15.8
N = 100
S = tfd.Normal(loc=0., scale=1.).sample((N,))
H = tfd.Binomial(total_count=10, probs=tf.sigmoid(S)).sample()
# ##### Code 15.9
#
# Hm = Homework missing
#
# Dog's decision to eat a piece of homework or not is not influenced by any relevant variable
# +
D = tfd.Bernoulli(0.5).sample().numpy() # dogs completely random
Hm = np.where(D == 1, np.nan, H)
Hm
# -
# Since missing values are random, missignness does not necessiarily change the overall distribution of homework score.
# ##### Code 15.10
#
# Here studying influences whether a dog eats homework S->D
#
# Students who study a lot do not play with their Dogs and then dogs take revenge by eating homework
#
# +
D = np.where(S > 0, 1, 0)
Hm = np.where(D == 1, np.nan, H)
Hm
# -
# Now every student who studies more than average (0) is missing homework
# ##### Code 15.11
#
# The case of noisy home and its influence on homework & Dog's behavior
# +
# TODO - use seed; have not been able to make it work with tfp
N = 1000
X = tfd.Sample(tfd.Normal(loc=0., scale=1.), sample_shape=(N,)).sample().numpy()
S = tfd.Sample(tfd.Normal(loc=0., scale=1.), sample_shape=(N,)).sample().numpy()
logits = 2 + S - 2 * X
H = tfd.Binomial(total_count=10, logits=logits).sample().numpy()
D = np.where(X > 1, 1, 0)
Hm = np.where(D == 1, np.nan, H)
# -
# ##### Code 15.12
# +
tdf = df_to_tensors("SimulatedHomeWork", pd.DataFrame.from_dict(dict(H=H,S=S)), ["H", "S"])
def model_15_3(S):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=1., name="alpha"), sample_shape=1))
betaS = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.5, name="betaS"), sample_shape=1))
logits = tf.squeeze(alpha[...,tf.newaxis] + betaS[...,tf.newaxis] * S)
H = yield tfd.Independent(tfd.Binomial(total_count=10, logits=logits), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=False)
jdc_15_3 = model_15_3(tdf.S)
# +
NUM_CHAINS_FOR_15_3 = 4
alpha_init, betaS_init, _ = jdc_15_3.sample()
init_state = [
tf.tile(alpha_init, (NUM_CHAINS_FOR_15_3,)),
tf.tile(betaS_init, (NUM_CHAINS_FOR_15_3,))
]
bijectors = [
tfb.Identity(),
tfb.Identity(),
]
posterior_15_3, trace_15_3 = sample_posterior(jdc_15_3,
observed_data=(tdf.H,),
params=['alpha', 'betaS'],
init_state=init_state,
bijectors=bijectors)
# -
az.summary(trace_15_3, round_to=2, kind='all', hdi_prob=0.89)
# The true coefficient on S should be 1.00. We don’t expect to get that exactly, but the estimate above is way off
# ##### Code 15.13
#
# We build the model with missing data now
# +
tdf = df_to_tensors("SimulatedHomeWork",
pd.DataFrame.from_dict(dict(H=H[D==0],S=S[D==0])), ["H", "S"])
def model_15_4(S):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=1., name="alpha"), sample_shape=1))
betaS = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=0.5, name="betaS"), sample_shape=1))
logits = tf.squeeze(alpha[...,tf.newaxis] + betaS[...,tf.newaxis] * S)
H = yield tfd.Independent(tfd.Binomial(total_count=10, logits=logits), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=False)
jdc_15_4 = model_15_4(tdf.S)
# +
NUM_CHAINS_FOR_15_4 = 2
alpha_init, betaS_init, _ = jdc_15_4.sample()
init_state = [
tf.tile(alpha_init, (NUM_CHAINS_FOR_15_4,)),
tf.tile(betaS_init, (NUM_CHAINS_FOR_15_4,))
]
bijectors = [
tfb.Identity(),
tfb.Identity(),
]
posterior_15_4, trace_15_4 = sample_posterior(jdc_15_4,
observed_data=(tdf.H,),
params=['alpha', 'betaS'],
init_state=init_state,
bijectors=bijectors)
# -
az.summary(trace_15_4, round_to=2, kind='all', hdi_prob=0.89)
# ##### Code 15.14
D = np.where(np.abs(X) < 1, 1, 0)
# ##### Code 15.15
# +
N = 100
S = tfd.Normal(loc=0., scale=1.).sample((N,))
H = tfd.Binomial(total_count=10, logits=S).sample().numpy()
D = np.where(H < 5, 1, 0)
Hm = np.where(D == 1, np.nan, H)
Hm
# -
# ## 15.3 Categorical errors and discrete absences (TODO)
| notebooks/15_missing_data_and_other_opportunities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # XML (Extensible Markup Language)
# The XML standard is a flexible way to create information formats and electronically share structured data.
# * XML is short for Extensible Markup Language and is used to describe data.
# * XML is a markup language much like HTML
# * XML was designed to store and transport data
# * XML was designed to be self-descriptive
# * XML is a W3C Recommendation
#
# Even though it is slowly being replaced by JSON, it is one of the fundamental data formats and it's crucial to learn about it.
#
# Online Resources:
# * https://www.w3schools.com/xml/xml_whatis.asp
# * https://docs.python.org/3/library/xml.etree.elementtree.html
#
# An example of XML:
#
# <img src='images/XML.png'>
| APIs and Other Data Types/.ipynb_checkpoints/XML -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
#
# This notebook will go over a new loss function to optimize the decision of the ERUPT metric directly for both single and mutliple response case.
#
# 1. New Loss Function for single response variable
#
# 2. Building / Gridsearching the Uplift Model with new loss vs default mean-squared-error loss
#
# 3. Extending new loss function to multi-responses
#
#
# ### New Loss Function
#
# Readers are encouraged to read the [example on single responses](https://github.com/Ibotta/ibotta_uplift/blob/master/examples/mr_uplift_one_response_example.ipynb) for a more thorough background on uplift models and the ERUPT metric.
#
# To recap we start with the decision function defined as:
#
# \begin{equation}
# \pi(x_i) =argmax \:_{t \in T} E[y_i | X=x_i, T=t]
# \tag{1}
# \end{equation}
#
# This is the function that determines whether to assign a user a particular treatment for a single response we wish to maximize. It just says that we apply the treatment with the highest expected value.
#
# To exaluate the decision function the ERUPT metric is used. It estimates the expected value of the decision function on an out-of-sample dataset. Assuming that the treatments are uniformly (all treatments have same number of observations) and randomly assigned the ERUPT metric is:
#
# $$ERUPT(pi(x),y,t) \propto \sum_{i}^N y_i I(\pi(x_i) = t_i) \tag{2}$$
#
#
#
# The original loss functions used in mr_uplift have used a mean squared error (MSE) function $f(x,t)$ to estimate $E[y_i | X=x_i, T=t]$ :
#
#
# \begin{equation}
# MSE(y,f(x,t)) = 1/N \sum_{i}^N (y_i-f(X=x_i, T=t_i))^2
# \tag{3}
# \end{equation}
#
#
# In theory this should produce an estimator that approximates the interaction between $x$ and $t$ neccesary to build a good decision function $\pi(x_i)$. However, in practice thie interaction effect is tricky to estimate because the it is generally small relative to main effects and noise in the response. Several researchers have developed in techniques order to estimate this (see [<NAME>](https://arxiv.org/pdf/1705.08492.pdf?source=post_page---------------------------), [<NAME>](https://arxiv.org/pdf/1610.01271.pdf), [<NAME>](https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3111957) to name a few).
#
#
# The approach taken here follows an approach developed by [Weiss](https://medium.com/building-ibotta/maximizing-the-erupt-metric-for-uplift-models-f8d7e57bfdf2) and [Fernandez-Loria, et al](https://arxiv.org/pdf/2004.11532.pdf) in which it attempts to optimize the ERUPT function in $2$ directly.
#
# In order to approximate that function we assign a 'probability' score to each treatment:
#
#
# \begin{equation}
# optimizedloss(y,t,f(t,x)) = \sum_{i}^N \sum_{t \in T}^T y_{i,t}*p(t|x_i) = \sum_{i}^N \sum_{t \in T}^T y_{i,t}*\frac{e^{f(X=x_i, T=t)}}{\sum_{t \in T}e^{f(X=x_i, T=t)}}
# \tag{4}
# \end{equation}
# where $y_{i_t}$ is equal to $y_{i}$ if user $i$ recieved treatment $t$, else zero.
#
# One major difference here compared with [Weiss](https://medium.com/building-ibotta/maximizing-the-erupt-metric-for-uplift-models-f8d7e57bfdf2) is that it explitly incorporates the treatments in the function $f()$ to be estimated. This should help regularize the estimation if the treatments have similar meta-data information. For instance; if the treatments are ordered then this model can take that information into account.
#
# Finally, MSE is also included in the loss function to create a weighted loss between the two. Including MSE should help when the estimated optimal treatment does not equal the treatment assignment:
#
# \begin{equation}
# \alpha*optimizedloss(y,t,f(t,x)) + (1-\alpha)*MSE(y,f(x,t))
# \tag{5}
# \end{equation}
#
#
#
# +
from ggplot import *
import numpy as np
import pandas as pd
from mr_uplift.dataset.data_simulation import get_sin_uplift_data
from mr_uplift.mr_uplift import MRUplift
# -
# ### Data Generating Process
#
# In order to test the new loss function I simulated some data and compared results with a regular MSE loss. Below is the data generating process of the data. This is similar to the previous data function except that the treatment effect relationship is now in a sine function. This adds a layer of complexity for the model to estimate.
#
# \begin{equation}
# x_1 \sim runif(0,1)
# \end{equation}
#
# \begin{equation}
# x_2 \sim runif(0,1)
# \end{equation}
#
# \begin{equation}
# e_1 \sim rnorm(0,1)
# \end{equation}
#
# \begin{equation}
# e_2 \sim rnorm(0,1)
# \end{equation}
#
# \begin{equation}
# t \sim rbinom(.5)
# \end{equation}
#
# \begin{equation}
# noise \sim rnorm(0,1)
# \end{equation}
#
# \begin{equation}
# revenue = sin(20*x_1*t) + e_1
# \end{equation}
#
# \begin{equation}
# costs = sin(20*x_2*t) + e_2
# \end{equation}
#
# \begin{equation}
# profit = revenue - costs
# \end{equation}
#
#
#
# +
np.random.seed(23)
y, x, t = get_sin_uplift_data(10000)
y = pd.DataFrame(y)
y.columns = ['revenue','cost', 'noise']
y['profit'] = y['revenue'] - y['cost']
# +
#Build Uplift with the default MSE Loss
uplift_model = MRUplift()
param_grid = dict(num_nodes=[8, 32], dropout=[.1,.25], activation=[
'relu'], num_layers=[2,3], epochs=[25], batch_size=[100])
uplift_model.fit(x, np.array(y['profit']).reshape(-1,1), t.reshape(-1,1), param_grid = param_grid, n_jobs = 1, cv = 5)
#Build Uplift with the Optimization Loss
#alpha and copy_several_times are new hyperparameters. alpha is the weight given to MSE vs optimi_loss from eqn (5)
#copy_several_times is for multiple response uplift models and is described below
uplift_model_optimized = MRUplift()
param_grid_optimized = dict(num_nodes=[8, 32], dropout=[.1,.25], activation=[
'relu'], num_layers=[2,3], epochs=[30], batch_size=[100],
copy_several_times = [None],
alpha = [.999,.99,.75,.5])
uplift_model_optimized.fit(x, np.array(y['profit']).reshape(-1,1), t.reshape(-1,1),
param_grid = param_grid_optimized, n_jobs = 1, optimized_loss = True, cv=5)
# -
# ### Results MSE vs Optimiziation
#
# The results below show the MSE only loss function increases profits by around ~0.04 while the new optimization results in profit score of ~0.17. This shows the new loss function can help estimate the heterogeneous effect more effectively in this example.
# +
erupt_curves_mse, distribution_mse = uplift_model.get_erupt_curves()
print('ERUPT Metric for MSE Model')
print(erupt_curves_mse)
print('Distribution of Treatments for MSE Model')
print(distribution_mse)
print('Best Parameters MSE')
print(uplift_model.best_params_net)
erupt_curves_optimized, distribution_optimized = uplift_model_optimized.get_erupt_curves()
print('ERUPT Metric for Optimized Model')
print(erupt_curves_optimized)
print('Distribution of Treatments for Optimized Model')
print(distribution_optimized)
print('Best Parameters Optimized')
print(uplift_model_optimized.best_params_net)
# -
# ### Multi-Response Optimization
#
# See [example on multiple_responses](https://github.com/Ibotta/ibotta_uplift/blob/master/examples/mr_uplift_multiple_response_example.ipynb) for a more detailed introduction on uplift models with multiple responses.
#
#
# The general idea of uplift models with multiple response variables is to create a new variable that is a weighted sum of the responses. These weights define the relative importance responses variables. One can then calculate ERUPT metrics for a given set of weights as if it was a single response.
#
# However, issue of extending the loss function from $(4)$ to a multi-response framework is that it only optimizes one set of weights. Ideally we'd like maximize a function that 'integrates out' the weights. Instead of doing that explitily I draw a random uniform weight for each observation-response pair and maximize the following:
#
# \begin{equation}
# w_{i,j} \sim runif(-1,1)
# \end{equation}
#
# \begin{equation}
# optimizedlossmultioutput(y,t,w,f(t,x)) = \sum_{i}^N \sum_{t \in T}^T (\sum_{j}^J w_{i,j} y_{i,t,j})*\frac{e^{\sum_{j}^J w_{i,j} f_j(X=x_i, T=t)}}{\sum_{t \in T}e^{\sum_{j}^J w_{i,j} f_j(X=x_i, T=t)}}
# \tag{6}
# \end{equation}
#
# Note that the weights $w_{i,j}$ do not explicitly go into the function $f(t,x)$.
#
# Different 'optimal treatments' will be selected due to randomness of the weights for two identical observations. Because of this I included another hyperparameter 'copy_several_times' that duplicates observations a specified number of times. I've found this to be helpful in practice
#
# An MSE loss is also include similar to $(5)$. This has two main benefits 1) to create help estimate the function when the optimal treatment was not randomly assigned and 2) to maintain the order of response predictions. Becuase of these reasons I would not set $\alpha$ to be 1.
#
# Below is an example of the multi-response case.
#
# +
uplift_model_optimized_mr = MRUplift()
param_grid = dict(num_nodes=[ 32,64, 256], dropout=[.25], activation=[
'relu'], num_layers=[3], epochs=[25], batch_size=[100],
copy_several_times = [10],
alpha = [.999,.99,.5])
uplift_model_optimized_mr.fit(x, y, t.reshape(-1,1),
param_grid = param_grid, n_jobs = 1, optimized_loss = True, cv=5)
# -
erupt_curves_optimized_mr, distribution_optimized_mr = uplift_model_optimized_mr.get_erupt_curves()
# +
from ggplot import *
distribution_optimized_mr['weights_1'] = [np.float(x.split(',')[0]) for x in distribution_optimized_mr['weights']]
erupt_curves_optimized_mr['weights_1'] = [np.float(x.split(',')[0]) for x in erupt_curves_optimized_mr['weights']]
ggplot(aes(x='weights_1', y='mean', group = 'assignment', colour = 'assignment'), data=erupt_curves_optimized_mr) +\
geom_line()+\
geom_point()+facet_wrap("response_var_names")
| examples/mr_uplift_new_optimized_loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# This problem is a old competition from Kaggle which can be found here: https://www.kaggle.com/c/movie-review-sentiment-analysis-kernels-only/overview
#
#
# The basic overview of the problem is as follows:
#
# The dataset is a set of reviews from rotten tomatoes, a website for reviewing films.
#
# We want to be able to input a review of a film and about a score of the film. In this setting the score is the label that is output from a model and can have five distinct value.
#
# The sentiment labels are:
#
# 0 - negative
# 1 - somewhat negative
# 2 - neutral
# 3 - somewhat positive
# 4 - positive
#
# The goal is to be able to this process with as high accuracy as possible.
#
# For more information visit the kaggle competition website.
#
#
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np
import pandas as pd
import os
print(os.listdir("../input"))
import gc
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from sklearn.multiclass import OneVsRestClassifier
from sklearn.model_selection import train_test_split, cross_val_score
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Dense,Dropout,Embedding,LSTM,Conv1D,GlobalMaxPooling1D,Flatten,MaxPooling1D,GRU,SpatialDropout1D,Bidirectional
from keras.callbacks import EarlyStopping
from keras.utils import to_categorical
from keras.losses import categorical_crossentropy
from keras.optimizers import Adam
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score,confusion_matrix,classification_report,f1_score
import matplotlib.pyplot as plt
# -
# These are the files avalible in the Kaggle kernel.
# # EDA
# + _uuid="376108f6e1a22afeba8739d2e12d0aa51fde8dea"
gc.collect()
# + [markdown] _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# ### Loading dataset and basic visualization
# + _uuid="cfd09ccff2dbc595ead54e3fd638cfdc3f3089e5"
train=pd.read_csv('../input/train.tsv.zip',sep='\t')
print(train.shape)
train.head()
# + _uuid="1b7f2e540ea49f1d94ec124b818f2a63c3c66a96"
test=pd.read_csv('../input/test.tsv.zip',sep='\t')
print(test.shape)
test.head()
# + _uuid="f4509ef7a7adcc7942a6f1acff34c54ce3802d5d"
sub=pd.read_csv('../input/sampleSubmission.csv')
sub.head()
# -
# Lets get some feeling for how a sentence is broken down into several phrases.
train.loc[train.SentenceId == 17]
# Here we see a potential problem. Differnet phrases in the same sentence will get different sentiments. It is especially intresting that the phrase "billingual charmer" has sentiment 3, while "billingual" and "charmer" gets a "2". This means that words do not only impact seperally but can magnify each others meaning.
#
# However, in some cases that is not obious. For example "inspired it" and "inspired" have different sentiments.
print('Average count of phrases per sentence in train is {0:.0f}.'.format(train.groupby('SentenceId')['Phrase'].count().mean()))
print('Average count of phrases per sentence in test is {0:.0f}.'.format(test.groupby('SentenceId')['Phrase'].count().mean()))
print('Number of phrases in train: {}. Number of sentences in train: {}.'.format(train.shape[0], len(train.SentenceId.unique())))
print('Number of phrases in test: {}. Number of sentences in test: {}.'.format(test.shape[0], len(test.SentenceId.unique())))
print('Average word length of phrases in train is {0:.0f}.'.format(np.mean(train['Phrase'].apply(lambda x: len(x.split())))))
print('Average word length of phrases in test is {0:.0f}.'.format(np.mean(test['Phrase'].apply(lambda x: len(x.split())))))
# ### Wordcloud
df_train_first = train.loc[train['SentenceId'].diff() != 0] #only extracts the first phrase for each sentence (full sentence)
# +
from wordcloud import WordCloud,STOPWORDS
import matplotlib.pyplot as plt
df_temp = df_train_first.loc[df_train_first['Sentiment']==4]
words = ' '.join(df_temp['Phrase'])
cleaned_word = " ".join([word for word in words.split()
if 'http' not in word
and not word.startswith('@')
and word != 'RT'
and word !='&'
])
stopwords = set(STOPWORDS)
wordcloud = WordCloud(stopwords=stopwords,
background_color='black',
width=3000,
height=2500
).generate(cleaned_word)
plt.figure(1,figsize=(12, 12))
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
# +
df_temp = df_train_first[df_train_first['Sentiment']==0]
words = ' '.join(df_temp['Phrase'])
cleaned_word = " ".join([word for word in words.split()
if 'http' not in word
and not word.startswith('@')
and word != 'RT'
and word !='&'
])
stopwords = set(STOPWORDS)
wordcloud = WordCloud(stopwords=stopwords,
background_color='black',
width=3000,
height=2500
).generate(cleaned_word)
plt.figure(1,figsize=(12, 12))
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
# -
# # Pre-processing
# + [markdown] _uuid="bbe766f446d3bb593acbdf5ef976097c929c7901"
# **Adding Sentiment column to test datset and joing train and test for preprocessing**
# + _uuid="293f41b034b63f6526f22fd616877220e19af8e7"
test['Sentiment']= -999
test.head()
# -
df=pd.concat([train,test],ignore_index=True)
print(df.shape)
del train,test # free up space
gc.collect()
from nltk.tokenize import word_tokenize
from nltk import FreqDist
from nltk.stem import SnowballStemmer,WordNetLemmatizer
stemmer=SnowballStemmer('english')
lemma=WordNetLemmatizer()
from string import punctuation
import re
def clean_review(review_col):
review_corpus=[]
for i in range(0,len(review_col)):
review=str(review_col[i])
review=re.sub('[^a-zA-Z]',' ',review)
#review=[stemmer.stem(w) for w in word_tokenize(str(review).lower())]
review=[lemma.lemmatize(w) for w in word_tokenize(str(review).lower())]
review=' '.join(review)
review_corpus.append(review)
return review_corpus
df['clean_review']=clean_review(df.Phrase.values)
df.head()
df_train=df[df.Sentiment!= -999]
df_train.shape
df_test=df[df.Sentiment == -999]
df_test.shape
del df
gc.collect()
# ## Split the data
train_text=df_train.clean_review.values
test_text=df_test.clean_review.values
target=df_train.Sentiment.values
y=to_categorical(target)
print(train_text.shape,target.shape,y.shape)
# stratify allows equal split over all classes
X_train_text,X_val_text,y_train,y_val=train_test_split(train_text,y,test_size=0.2,stratify=y,random_state=123)
print(X_train_text.shape,y_train.shape)
print(X_val_text.shape,y_val.shape)
#
#
# # Modeling
# ## Tokenization
#determining number of unique words
all_words=' '.join(X_train_text)
all_words=word_tokenize(all_words)
dist=FreqDist(all_words)
num_unique_word=len(dist)
num_unique_word
# +
# Determening the maximum lenght of a review (number of words)
r_len=[]
for text in X_train_text:
word=word_tokenize(text)
l=len(word)
r_len.append(l)
MAX_REVIEW_LEN=np.max(r_len)
MAX_REVIEW_LEN
# -
max_features = num_unique_word
max_words = MAX_REVIEW_LEN
# +
from keras.preprocessing.text import Tokenizer
tokenizer = Tokenizer(num_words=max_features)
tokenizer.fit_on_texts(list(X_train_text))
X_train = tokenizer.texts_to_sequences(X_train_text)
X_val = tokenizer.texts_to_sequences(X_val_text)
X_test = tokenizer.texts_to_sequences(test_text)
# +
from keras.preprocessing import sequence,text
'''
This function transforms a list (of length num_samples) of sequences (lists of integers)
into a 2D Numpy array of shape (num_samples, num_timesteps).
num_timesteps is either the maxlen argument if provided,
or the length of the longest sequence in the list. '''
X_train = sequence.pad_sequences(X_train, maxlen=max_words)
X_val = sequence.pad_sequences(X_val, maxlen=max_words)
X_test = sequence.pad_sequences(X_test, maxlen=max_words)
print(X_train.shape,X_val.shape,X_test.shape)
# -
# # logistic regression
#
# Is used as a baseline model
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf=TfidfVectorizer(ngram_range=(1,2),max_df=0.95,min_df=10,sublinear_tf=True)
X_train_vec=tfidf.fit_transform(df_train.clean_review).toarray()
print(X_train_vec.shape)
X_test_vec =tfidf.transform(df_test.clean_review).toarray()
print(X_test_vec.shape)
from sklearn.preprocessing import LabelEncoder,OneHotEncoder
le=LabelEncoder()
y_train_enc=le.fit_transform(df_train.Sentiment.values)
y_train_enc.shape
logreg = LogisticRegression()
#ovr_logreg = OneVsRestClassifier(logreg) # this strategy consists in fitting one classifier per class
# %%time
logreg.fit(X_train_vec, y_train_enc)
log_pred=logreg.predict(X_test_vec)
df_test['Logistic_Sentiment'] = log_pred
df_test.head()
del X_train_vec, X_test_vec, y_train_enc, logreg,le,tfidf
gc.collect()
# # LSTM
# +
lstm=Sequential()
lstm.add(Embedding(max_features,100,mask_zero=True))
lstm.add(LSTM(64,dropout=0.4, recurrent_dropout=0.4,return_sequences=True))
lstm.add(LSTM(32,dropout=0.5, recurrent_dropout=0.5,return_sequences=False))
lstm.add(Dense(5,activation='softmax'))
lstm.compile(loss='categorical_crossentropy',optimizer=Adam(lr=0.001),metrics=['accuracy'])
lstm.summary()
# +
# %%time
epochs = 3
batch_size = 128
lstm_history =lstm.fit(X_train, y_train, validation_data=(X_val, y_val),epochs=epochs, batch_size=batch_size, verbose=1)
# -
lstm_pred=lstm.predict_classes(X_test,verbose=1)
df_test['lstm_sentiment'] = lstm_pred
df_test.head()
del lstm,lstm_history,lstm_pred
gc.collect()
# # Vader
from nltk.sentiment.vader import SentimentIntensityAnalyzer
def print_sentiment_scores(sentence):
snt = SentimentIntensityAnalyzer().polarity_scores(sentence) #Calling the polarity analyzer
print("{:-<40} {}".format(sentence, str(snt)))
# Lets try it out
print_sentiment_scores('bad movie is bad')
print_sentiment_scores('bad movie is really bad')
print_sentiment_scores('ok movie is ok')
print_sentiment_scores('good movie is good')
print_sentiment_scores('good movie is really good')
print_sentiment_scores('This movie is so bad that it is good')
print_sentiment_scores('This movie is so good that it is bad')
print_sentiment_scores('This movie is the shit')
print_sentiment_scores('This movie is shit')
print_sentiment_scores('The plot was obvious')
# +
# %%time
i=0 #counter
compval1 = [ ] #empty list to hold our computed 'compound' VADER scores
for sentence in df_train['Phrase']:
if i % 5000 == 0:
print(i,'samples done')
k = SentimentIntensityAnalyzer().polarity_scores(sentence)
compval1.append(k['compound'])
i = i+1
#converting sentiment values to numpy for easier usage
compval1 = np.array(compval1)
len(compval1)
# -
df_vader = df_train.copy()
df_vader['VADER_score'] = compval1
df_vader.head()
# A VADER score of -1 corresponds to a negative review, a score of 1 corresponds to a positive review, and a score of 0 corresponds to a ok review. As we have five classes, the delimitations could be -0.6, -0.2, 0.2, 0.6 for a negative to positive review.
#
# However, for this data, are we sure that we should draw the boundaries there? Is a review of -0.19 really ok when it is mostly negative? What does, an ok review mean in this context and in relation to the VADER score?
#
# Lets find out...
import seaborn as sns
import matplotlib as plt
# Many ofthe phrases are nonsensical and will give a vader score of 0. As many phrases are such in the dataset, the vader score will presumably be biased towards a neutral score.
df_vader_non_zero = df_vader.loc[df_vader['VADER_score'] != 0.0000]
#sns.displot(data = df_vader, x="VADER_score", hue="species", stat="Sentiment", fill=True)
sns.distplot(df_vader_non_zero['VADER_score'])
sns.distplot(df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 0]['VADER_score'])
print('Mean value:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 0]['VADER_score'].mean())
print('Std:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 0]['VADER_score'].std())
df_temp = df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 0]
df_temp.loc[df_temp['VADER_score'] > 0.5].sample(5)
sns.distplot(df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 1]['VADER_score'])
print('Mean value:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 1]['VADER_score'].mean())
print('Std:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 1]['VADER_score'].std())
sns.distplot(df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 2]['VADER_score'])
print('Mean value:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 2]['VADER_score'].mean())
print('Std:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 2]['VADER_score'].std())
sns.distplot(df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 3]['VADER_score'])
print('Mean value:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 3]['VADER_score'].mean())
print('Std:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 3]['VADER_score'].std())
sns.distplot(df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 4]['VADER_score'])
print('Mean value:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 4]['VADER_score'].mean())
print('Std:',df_vader_non_zero.loc[df_vader_non_zero['Sentiment'] == 4]['VADER_score'].std())
# ### Prediction with logistic regression
# In this case we use the logistic regression model to map the vader score to the sentiment score.
x =df_vader['VADER_score'].values.reshape(-1,1)
y = df_vader['Sentiment'].values.reshape(-1,1)
print(x.shape)
print(y.shape)
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier(random_state=0)
clf.fit(x, y)
# Lets get the vader score for test data
# +
# %%time
i=0 #counter
compval2 = [ ] #empty list to hold our computed 'compound' VADER scores
for sentence in df_test['Phrase']:
if i % 5000 == 0:
print(i,'samples done')
k = SentimentIntensityAnalyzer().polarity_scores(sentence)
compval2.append(k['compound'])
i = i+1
#converting sentiment values to numpy for easier usage
print('DONE')
compval2 = np.array(compval2)
len(compval2)
# -
df_test['VADER_score_test'] = compval2
clf_vader_pred = clf.predict(df_test['VADER_score_test'].values.reshape(-1,1))
df_test['VADER_sentiment'] = clf_vader_pred
df_test['VADER_sentiment'].value_counts()
df_test.head()
df_test.to_csv('sentiment_all_models.csv',index=False)
# ### Prediction with manual set bins
# +
# %%time
i=0 #counter
compval2 = [ ] #empty list to hold our computed 'compound' VADER scores
for sentence in df_test['Phrase']:
if i % 5000 == 0:
print(i,'samples done')
k = SentimentIntensityAnalyzer().polarity_scores(sentence)
compval2.append(k['compound'])
i = i+1
#converting sentiment values to numpy for easier usage
print('DONE')
compval2 = np.array(compval2)
len(compval2)
# -
df_test['VADER_score'] = compval2
# +
# %time
#Assigning score categories and logic
i = 0
predicted_value = [ ] #empty series to hold our predicted values
while(i<len(df_test)):
if i % 5000 == 0:
print(i,'samples done')
if ((df_test.iloc[i]['VADER_score'] >= 0.4)):
predicted_value.append(4)
i = i+1
elif ((df_test.iloc[i]['VADER_score'] > 0.15) & (df_test.iloc[i]['VADER_score'] < 0.4)):
predicted_value.append('3')
i = i+1
elif ((df_test.iloc[i]['VADER_score'] > -0.075) & (df_test.iloc[i]['VADER_score'] < 0.15)):
predicted_value.append('2')
i = i+1
elif ((df_test.iloc[i]['VADER_score'] > -0.15) & (df_test.iloc[i]['VADER_score'] < -0.075)):
predicted_value.append('1')
i = i+1
elif ((df_test.iloc[i]['VADER_score'] <= -0.15)):
predicted_value.append('0')
i = i+1
# -
df_test['VADER_sentiment'] = predicted_value
del df_vader,compval1,compval2,logreg,df_temp,x,y,logreg
gc.collect()
# # CNN
# +
cnn= Sequential()
cnn.add(Embedding(max_features,100,input_length=max_words))
cnn.add(Conv1D(64,kernel_size=3,padding='same',activation='relu',strides=1))
cnn.add(MaxPooling1D(3))
cnn.add(Conv1D(64,kernel_size=3,padding='same',activation='relu',strides=1))
cnn.add(GlobalMaxPooling1D())
cnn.add(Dense(128,activation='relu'))
cnn.add(Dropout(0.4))
cnn.add(Dense(128))
cnn.add(Dense(5,activation='softmax'))
cnn.summary()
cnn.compile(optimizer=Adam(lr=0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# +
# %%time
from tensorflow.keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint
num_epochs = 3
batch_size = 128
cnn_history =cnn.fit(X_train, y_train, validation_data=(X_val, y_val),epochs=num_epochs, batch_size=batch_size, verbose=1)
# -
cnn_pred =cnn.predict_classes(X_test, verbose=1)
df_test['CNN_Sentiment'] = cnn_pred
df_test.to_csv('sentiment_all_models_V2.csv',index=False)
# # CNN-LSTM
# +
cnnlstm= Sequential()
cnnlstm.add(Embedding(max_features,100,input_length=max_words))
cnnlstm.add(Conv1D(64,kernel_size=3,padding='same',activation='relu',strides=1))
cnnlstm.add(MaxPooling1D(3))
cnnlstm.add(Conv1D(64,kernel_size=3,padding='same',activation='relu',strides=1))
cnnlstm.add(MaxPooling1D(3))
cnnlstm.add(LSTM(64,dropout=0.4, recurrent_dropout=0.4,return_sequences=False))
cnnlstm.add(Dense(128,activation='relu'))
cnnlstm.add(Dropout(0.4))
cnnlstm.add(Dense(128))
cnnlstm.add(Dense(5,activation='softmax'))
cnnlstm.summary()
cnnlstm.compile(optimizer=Adam(lr=0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# +
# %%time
from tensorflow.keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint
num_epochs = 3
batch_size = 128
cnnlstm_history =cnnlstm.fit(X_train, y_train, validation_data=(X_val, y_val),epochs=num_epochs, batch_size=batch_size, verbose=1)
# -
cnnlstm_pred = cnnlstm.predict_classes(X_test, verbose=1)
df_test['CNN-LSTM_Sentiment'] = cnnlstm_pred
df_test.to_csv('sentiment_all_models_V3.csv',index=False)
| sentiment-analysis-rotten-tomatoes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.feature_selection import SelectKBest
import seaborn as sns
# +
df_train = pd.read_csv('../data/train.csv')
df_test = pd.read_csv('../data/test.csv')
df_train['id'] = -1
df_test['target'] = -1
# -
df_all = df_train.append(df_test)
# fillna
for col in ["source_screen_name","source_system_tab","source_type"]:
df_all.loc[df_all[col].isnull(),col] = df_all[col].value_counts().index[0]
df_all.to_csv("../data/processed/df_all.csv",index=False)
| wsdm/feature/main_feat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pachterlab/GFCP_2021/blob/main/notebooks/embed_neighbors_jaccard_lme.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3n058GtM7RME"
# #Figure 6: Nonlinear point projections introduce distortions in local neighborhoods.
# + [markdown] id="kgxaLJJQ7Rtw"
# In this notebook, we examine the La Manno et al. human forebrain dataset, process it using the standard workflow, and quantify the effect of dimensionality reduction on local neighborhoods (as defined by Jaccard distances on the identities of $k=150$ nearest neighbors).
# + [markdown] id="unusual-burton"
# # Dependencies
# + id="lyPyjB4s7oLO"
import time
t1 = time.time()
# + id="standing-mission"
## uncomment this cell if run in colab ##
# %%capture
# !pip install --upgrade git+https://github.com/meichenfang/velocyto.py.git
# !pip install scanpy umap-learn
# + id="7YwXLZBg7qaG"
# !mkdir -p figure
# !mkdir -p data
# + id="nuXAcKn3xhZE" outputId="7fbc7a59-f4ee-4d70-abc4-34a28e854799" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount('/content/drive')
# + id="BbhP_NMfxirX"
# !cp -r /content/drive/MyDrive/rnavelocity/hgForebrainGlut.loom /content/data
# + [markdown] id="nHTtTaeb7rtB"
# Here, the `hgForebrainGlut.loom` and `vis.py` files need to be uploaded.
# + id="3gwKMYc37rzY" outputId="e11edd37-3c39-480f-84bf-5be9e568f4b5" colab={"base_uri": "https://localhost:8080/"}
from vis import *
# + id="distant-shower"
# ## run locally ##
# # %cd ../../GFCP_2021
# from vis import *
# # %cd notebooks
# + [markdown] id="sixth-triangle"
# # Load data and preprocess
# + id="focal-siemens"
vlm = vcy.VelocytoLoom(loom_filepath="data/hgForebrainGlut.loom")
preprocess(vlm)
# + [markdown] id="effective-shark"
# # Generate Figure 6
# + id="electric-distinction" outputId="cc37efd4-e707-4b1e-8014-cb48e29de7b2" colab={"base_uri": "https://localhost:8080/", "height": 560}
fig = plt.figure(figsize=(24,10))
ax0 = plt.subplot2grid((2, 4), (0, 0))
ax1 = plt.subplot2grid((2, 4), (0, 1))
ax2 = plt.subplot2grid((2, 4), (1, 0))
ax3 = plt.subplot2grid((2, 4), (1, 1))
ax4 = plt.subplot2grid((2, 4), (0, 2), rowspan=2, colspan=2)
n_pcs_for_tsne = 25
n_neigh=150
plt.rcParams.update({'font.size': 16})
fracs=[]
labels=["Ambient vs. s_norm","S_norm vs. PCA "+str(n_pcs_for_tsne),
"PCA "+str(n_pcs_for_tsne)+" vs. PCA 2","PCA "+str(n_pcs_for_tsne)+" vs. tSNE 2",
"Ambient vs. PCA 2",'Ambient vs. tSNE 2']
makeEmbeds(vlm, embeds="PCA", x_name="S_norm")
ax0.set_title(labels[0])
ax0.axes.xaxis.set_ticklabels([])
ax0.set_ylabel("density")
fracs.append(plotJaccard(vlm.S.T, vlm.S_norm.T, ax0, n_neigh=n_neigh, c=vermeer[0]))
ax1.set_title(labels[1])
ax1.axes.xaxis.set_ticklabels([])
ax1.axes.yaxis.set_ticklabels([])
fracs.append(plotJaccard(vlm.pcs[:,:n_pcs_for_tsne], vlm.S_norm.T, ax1, n_neigh=n_neigh, c=vermeer[1]))
ax2.set_title(labels[2])
ax2.set_xlabel("range")
ax2.set_ylabel("density")
fracs.append(plotJaccard(vlm.pcs[:,:n_pcs_for_tsne],vlm.pcs[:,:2], ax2, n_neigh=n_neigh, c=vermeer[2]))
ax3.set_title(labels[3])
ax3.set_xlabel("range")
ax3.axes.yaxis.set_ticklabels([])
makeEmbeds(vlm, embeds="tSNE", x_name="S_norm")
fracs.append(plotJaccard(vlm.pcs[:,:n_pcs_for_tsne], vlm.ts, ax3, n_neigh=n_neigh, c=vermeer[3]))
fracs.append(getJaccard(vlm.S.T, vlm.pcs[:,:2], n_neigh=n_neigh))
fracs.append(getJaccard(vlm.S.T, vlm.ts, n_neigh=n_neigh))
fracs_pd=pd.DataFrame(data=np.transpose(fracs),columns=labels)
ax4.set_title("eCDF")
ax4.set_xlabel("Jaccard distance")
vermeer.pop(4)
sn.ecdfplot(fracs_pd,ax=ax4,palette=vermeer[:6],linewidth=2)
fig.tight_layout()
fig.savefig('figure/embed_neighbors_jaccard_lme_raw.png',dpi=600)
# + id="inclusive-court" outputId="d13c80fd-f1dd-4433-adbb-b479178856fb" colab={"base_uri": "https://localhost:8080/"}
t2 = time.time()
print('Runtime: {:.2f} seconds.'.format(t2-t1))
# + id="LmHOvk1q9Vjj"
| notebooks/embed_neighbors_jaccard_lme.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests, re, webbrowser
from bs4 import BeautifulSoup
url = "https://mashable.com/2014/08/28/static-website-generators/?europe=true"
resp = requests.get(url)
resp
soup = BeautifulSoup(resp.text,"lxml")
soup
soup.findAll("a", attrs= {"href":re.compile("^http")})
for link in soup.findAll("a", attrs= {"href":re.compile("^https")}):
print(link["href"])
#Prints only HTTPS href link
for link in soup.findAll("a", attrs= {"href":re.compile("http://")}):
print(link["href"])
#Prints only HTTP href link
for link in soup.findAll("a", attrs= {"href":re.compile("^http")}):
print(link["href"])
#Prints all HTTP/HTTPS href link
for link in soup.findAll("a", href=True):
print(link["href"])
#Prints all href link {Relative + Absolute}
for link in soup.findAll("a", href=True):
if not link['href'].startswith("http"):
link = url+ link["href"].strip("/")
else:
link = link["href"]
print(link)
| Parse/BeautifulSoup 4/BeautifulSoup_Links-Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:marvindev]
# language: python
# name: conda-env-marvindev-py
# ---
import warnings
warnings.simplefilter('ignore')
# # Marvin Results
#
# This tutorial explores some basics of how to handle results of your Marvin Query. Much of this information can also be found in the [Marvin Results documentation](https://sdss-marvin.readthedocs.io/en/latest/query/results.html).
#
# Table of Contents:
# - [Performing a Query](#query)<br>
# - [Retrieving Results](#retrieve)<br>
# - [Formatting Results](#format)<br>
# - [Quickly Plotting Results](#plot)<br>
# - [Downloading Results](#download)<br>
# <a id='query'></a>
# ## Performing a Query
# Our first step is to generate a query. Let's perform a simple metadata query to look for all galaxies with a redshift < 0.1. Let's also return the absolute magnitude g-r color and the Elliptical Petrosian half-light radius. This step assumes familiarity with Marvin Queries. To learn how to write queries, please see the [Marvin Query documentation](https://sdss-marvin.readthedocs.io/en/latest/query/query.html) or the [Marvin Query Tutorial](https://sdss-marvin.readthedocs.io/en/latest/tutorials/notebooks/marvin_queries.html).
# set up and run the query
from marvin.tools.query import Query
q = Query(search_filter='nsa.z < 0.1', return_params=['absmag_g_r', 'nsa.elpetro_th50_r'])
r = q.run()
# repr the results
r
# Our query runs and indicates a total count of 4275 results. By default, queries that return more than 1000 rows will be automatically paginated into sets (or chunks) of 100 rows, indicated by ``count=100``. The number of rows queries return can be changed using the ``limit`` keyword argument to ``Qeuery``. The results are stored in the ``results`` attribute.
# look at the results
r.results
# A ``ResultSet`` contains a list of tuple rows with some default parameters like ``mangaid`` and ``plateifu``, plus any parameters used in the ``Query`` ``search_filter`` or requested with the ``return_params`` keyword. The redshift, g-r color, and half-light radius has been returned. We can look at all the columns available using the ``columns`` attribute.
# look at the columns returned by your results
r.columns
# <a id='retrieve'></a>
# ## Retrieving Results
# There are several options for handling paginated results. To page through the sets of results without extending the results, use ``getNext`` and ``getPrevious``. These methods simply page through.
# get the next set of results
n = r.getNext()
# look at page 2
r.results
# get the previous set
p = r.getPrevious()
# To extend your results and keep them, use the ``extendSet`` method. By default, extending a set grabs the next page of 100 results (defined by ``r.chunk``) and appends to the existing set of results. Rerunning ``extendSet`` continues to append results until you've retrieved them all. To avoid running ``extendSet`` multiple times, you can run use the ``loop`` method, which will loop over all pages appending the data until you've retrieved all the results.
# extend the set by one page
r.extendSet()
r
# We now have 200 results out of the 4275. For results with a small number of total counts, you can attempt to retrieve all of the results with the ``getAll`` method. Currently this method is limited to returning results containing 500,000 rows or rows with 25 columns.
#
# #### Getting all the results
# There are several options for getting all of the results.
# - Use the ``getAll`` method to attempt to retrieve all the results in one request.
# - Use the ``loop`` method to loop over all the pages to extend/append the results together
# - Rerun the ``Query`` using a new ``limit`` to retrieve all the results.
#
# **Note:** A bug was recently found in ``getAll`` and might not work. Instead we will rerun the query using a large limit to return all the results.
# +
# get all the results
# r.getAll()
# rerun the query
q = Query(search_filter='nsa.z < 0.1', return_params=['absmag_g_r', 'nsa.elpetro_th50_r'], limit=5000)
r = q.run()
r
# -
# We now have all the results. We can extract columns of data by indexing the results list using the column name. Let's extract the redshift and color.
# extract individual columns of data
redshift = r.results['nsa.z']
color = r.results['absmag_g_r']
# <a id='format'></a>
# ## Formatting Results
# You can convert the results to a variety of formats using the ``toXXX`` methods. Common formats are **FITS**, **Astropy Table**, **Pandas Dataframe**, **JSON**, or **CSV**. Only the FITS and CSV conversions will write the output to a file. Astropy Tables and Pandas Dataframes have more options for writing out your dataset to a file. Let's convert to Pandas Dataframe.
# convert the marvin results to a Pandas dataframe
df = r.toDF()
df.head()
# You can also convert the data into Marvin objects using the ``convertToTool`` method. This will attempt to convert each result row into its corresponding Marvin Object. The default conversion is to a ``Cube`` object. Converted objects are stored in the ``r.objects`` attribute. Let's convert our results to cubes. **Depending on the number of results, this may take awhile. Let's limit our conversion to 5.** Once converted, we now have Marvin Tools at our disposal.
# convert the top 5 to cubes
r.convertToTool('cube', limit=5)
# look at the objects
r.objects
# <a id='plot'></a>
# ## Quickly Plotting the Results
# You can quickly plot the full set of results using the ``plot`` method. ``plot`` accepts two string column names and will attempt to create a scatter plot, a hex-binned plot, or a scatter-density plot, depending on the total number of results. The ``plot`` method returns the matplotlib **Figure** and **Axes** objects, as well as a dictionary of histogram information for each column. The ``Results.plot`` method uses the underlying [plot utility function](https://sdss-marvin.readthedocs.io/en/latest/tools/utils/plot-scatter.html). The utility function offers up more custom plotting options. Let's plot g-r color versus redshift. **Regardless of the number of results you currently have loaded, the ``plot`` method will automatically retrieve all the results before plotting.**
# make a scatter plot
fig, ax, histdata = r.plot('z', 'absmag_g_r')
# By default, it will also plot histograms of the column as well. This can be turned off by setting `with_hist=False`.
# make only a scatter plot
fig, ax = r.plot('z', 'absmag_g_r', with_hist=False)
# We can also quickly plot a histogram of a single column of data using the ``hist`` method, which uses an underlying [hist utility function](https://sdss-marvin.readthedocs.io/en/latest/tools/utils/plot-hist.html).
histdata, fig, ax = r.hist('absmag_g_r')
# <a id='download'></a>
# ## Downloading Results
# You can download the raw files from your results using the ``download`` method. This uses the ``downloadList`` utility function under the hood. By default this will download the DRP cubes for each target row. It accepts any keyword arguments as ``downloadList``.
# +
# download the DRP datacube files from the results
# r.download()
# -
# ## Additional Resoures
# - Get a refresher on [Queries](https://sdss-marvin.readthedocs.io/en/latest/query/query.html)
# - See more details of what you can do with [Results](https://sdss-marvin.readthedocs.io/en/latest/query/results.html#using-results)
# - See the [Results API](https://sdss-marvin.readthedocs.io/en/latest/reference/queries.html#marvin-results-ref) for a look at the Marvin Results object.
| docs/sphinx/tutorials/notebooks/Marvin_Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="LeamvpPVXuS_"
# # Random Forest Regression
# + [markdown] id="O2wvZ7SKXzVC"
# ## Importing the libraries
# + id="PVmESEFZX4Ig" executionInfo={"status": "ok", "timestamp": 1617348540448, "user_tz": -330, "elapsed": 785, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08460631435426072584"}}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="zgbK_F8-X7em"
# ## Importing the dataset
# + id="adBE4tjQX_Bh" executionInfo={"status": "ok", "timestamp": 1617348542113, "user_tz": -330, "elapsed": 761, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08460631435426072584"}}
dataset = pd.read_csv('Position_Salaries.csv')
x = dataset.iloc[:,1:-1].values
y = dataset.iloc[:,-1].values
# + [markdown] id="v4S2fyIBYDcu"
# ## Training the Random Forest Regression model on the whole dataset
# + id="o8dOCoJ1YKMc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1617348754463, "user_tz": -330, "elapsed": 951, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08460631435426072584"}} outputId="ca1ebd78-eee9-4e0b-aa3e-1db9e8500064"
from sklearn.ensemble import RandomForestRegressor
regressor = RandomForestRegressor(n_estimators = 10, random_state = 0)#Parameter(10) -> Number of Trees
regressor.fit(x,y)
# + [markdown] id="8IbsXbK3YM4M"
# ## Predicting a new result
# + id="pTXrS8FEYQlJ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1617348770692, "user_tz": -330, "elapsed": 785, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08460631435426072584"}} outputId="fd19c274-02b7-40fe-9b27-5caf0358bb39"
regressor.predict([[6.5]])
# + [markdown] id="kLqF9yMbYTon"
# ## Visualising the Random Forest Regression results (higher resolution)
# + id="BMlTBifVYWNr" colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"status": "ok", "timestamp": 1617348909783, "user_tz": -330, "elapsed": 1504, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08460631435426072584"}} outputId="c9c71cdc-1fd3-4c7e-8abb-4d3b6ca7ef11"
X_grid = np.arange(min(x),max(x),0.01)
X_grid = X_grid.reshape(len(X_grid),1)
plt.scatter(x,y,color='red')
plt.plot(X_grid,regressor.predict(X_grid), color='blue')
plt.title('Random Forest Regression')
plt.xlabel('Position Level')
plt.ylabel('Salary')
plt.show()
| Regression/Section 9 - Random Forest Regression/Python/random_forest_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import cobra
import cplex
import cobrakbase
#Put the path to ModelSEEDpy on your machine here
sys.path.append("/Users/chenry/code/ModelSEEDpy")
#import modelseedpy.fbapkg
from modelseedpy import KBaseMediaPkg, FullThermoPkg
from modelseedpy import FBAHelper
kbase_api = cobrakbase.KBaseAPI()
model = kbase_api.get_from_ws("E_iAH991V2",40576)
model.solver = 'optlang-cplex'
kmp = KBaseMediaPkg(model)
kmp.build_package(None)
FBAHelper.set_objective_from_target_reaction(model,"bio1")
sol=model.optimize()
model.summary()
ftp = FullThermoPkg(model)
ftp.build_package({"modelseed_path":"/Users/chenry/code/ModelSEEDDatabase/"})
with open('FullThermo.lp', 'w') as out:
out.write(str(model.solver))
sol=model.optimize()
#Next minimize active reactions
model.summary()
| examples/Flux Analysis/FullThermodynamicsExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Example for FloPy methods note
# Import the `modflow` and `utils` subpackages of FloPy and give them the aliases `fpm` and `fpu`, respectively
import os
import numpy as np
import flopy.modflow as fpm
import flopy.utils as fpu
# Create a MODFLOW model object. Here, the MODFLOW model object is stored in a Python variable called {\tt model}, but this can be an arbitrary name. This object name is important as it will be used as a reference to the model in the remainder of the FloPy script. In addition, a {\tt modelname} is specified when the MODFLOW model object is created. This {\tt modelname} is used for all the files that are created by FloPy for this model.
exe = 'mf2005'
ws = os.path.join('temp')
model = fpm.Modflow(modelname='gwexample', exe_name=exe, model_ws=ws)
# The discretization of the model is specified with the discretization file (DIS) of MODFLOW. The aquifer is divided into 201 cells of length 10 m and width 1 m. The first input of the discretization package is the name of the model object. All other input arguments are self explanatory.
fpm.ModflowDis(model, nlay=1, nrow=1, ncol=201,
delr=10, delc=1, top=50, botm=0)
# Active cells and the like are defined with the Basic package (BAS), which is required for every MODFLOW model. It contains the {\tt ibound} array, which is used to specify which cells are active (value is positive), inactive (value is 0), or fixed head (value is negative). The {\tt numpy} package (aliased as {\tt np}) can be used to quickly initialize the {\tt ibound} array with values of 1, and then set the {\tt ibound} value for the first and last columns to -1. The {\tt numpy} package (and Python, in general) uses zero-based indexing and supports negative indexing so that row 1 and column 1, and row 1 and column 201, can be referenced as [0, 0], and [0, -1], respectively. Although this simulation is for steady flow, starting heads still need to be specified. They are used as the head for fixed-head cells (where {\tt ibound} is negative), and as a starting point to compute the saturated thickness for cases of unconfined flow.
ibound = np.ones((1, 201))
ibound[0, 0] = ibound[0, -1] = -1
fpm.ModflowBas(model, ibound=ibound, strt=20)
# The hydraulic properties of the aquifer are specified with the Layer Properties Flow (LPF) package (alternatively, the Block Centered Flow (BCF) package may be used). Only the hydraulic conductivity of the aquifer and the layer type ({\tt laytyp}) need to be specified. The latter is set to 1, which means that MODFLOW will calculate the saturated thickness differently depending on whether or not the head is above the top of the aquifer.
#
fpm.ModflowLpf(model, hk=10, laytyp=1)
# Aquifer recharge is simulated with the Recharge package (RCH) and the extraction of water at the two ditches is simulated with the Well package (WEL). The latter requires specification of the layer, row, column, and injection rate of the well for each stress period. The layers, rows, columns, and the stress period are numbered (consistent with Python's zero-based numbering convention) starting at 0. The required data are stored in a Python dictionary ({\tt lrcQ} in the code below), which is used in FloPy to store data that can vary by stress period. The {\tt lrcQ} dictionary specifies that two wells (one in cell 1, 1, 51 and one in cell 1, 1, 151), each with a rate of -1 m$^3$/m/d, will be active for the first stress period. Because this is a steady-state model, there is only one stress period and therefore only one entry in the dictionary.
fpm.ModflowRch(model, rech=0.001)
lrcQ = {0: [[0, 0, 50, -1], [0, 0, 150, -1]]}
fpm.ModflowWel(model, stress_period_data=lrcQ)
# The Preconditioned Conjugate-Gradient (PCG) solver, using the default settings, is specified to solve the model.
fpm.ModflowPcg(model)
# The frequency and type of output that MODFLOW writes to an output file is specified with the Output Control (OC) package. In this case, the budget is printed and heads are saved (the default), so no arguments are needed.
fpm.ModflowOc(model)
# Finally the MODFLOW input files are written (eight files for this model) and the model is run. This requires, of course, that MODFLOW is installed on your computer and FloPy can find the executable in your path.
model.write_input()
model.run_model()
# After MODFLOW has responded with the positive {\tt Normal termination of simulation}, the calculated heads can be read from the binary output file. First a file object is created. As the modelname used for all MODFLOW files was specified as {\tt gwexample} in step 1, the file with the heads is called {\tt gwexample.hds}. FloPy includes functions to read data from the file object, including heads for specified layers or time steps, or head time series at individual cells. For this simple mode, all computed heads are read.
fpth = os.path.join(ws, 'gwexample.hds')
hfile = fpu.HeadFile(fpth)
h = hfile.get_data(totim=1.0)
# The heads are now stored in the Python variable {\tt h}. FloPy includes powerful plotting functions to plot the grid, boundary conditions, head, etc. This functionality is demonstrated later. For this simple one-dimensional example, a plot is created with the matplotlib package
import matplotlib.pyplot as plt
ax = plt.subplot(111)
x = model.dis.sr.xcentergrid[0]
ax.plot(x,h[0,0,:])
ax.set_xlim(0,x.max())
ax.set_xlabel("x(m)")
ax.set_ylabel("head(m)")
plt.show()
| examples/groundwater_paper/Notebooks/example_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, classification_report
from statsmodels.stats.diagnostic import het_breuschpagan
import seaborn as sns
import matplotlib.pyplot as plt
import plotly_express as px
# %matplotlib inline
# +
import statsmodels.api as sm
# -
from sqlalchemy import create_engine
from IPython.display import display_html
# +
postgres_user = 'dsbc_student'
postgres_pw = '<PASSWORD>'
postgres_host = '172.16.58.3'
postgres_port = '5432'
postgres_db = 'weatherinszeged'
engine = create_engine('postgresql://{}:{}@{}:{}/{}'.format(
postgres_user, postgres_pw, postgres_host, postgres_port, postgres_db))
df = pd.read_sql_query('select * from weatherinszeged',con=engine)
# No need for an open connection, because only doing a single query
engine.dispose()
df.head(10)
# -
df['temp_diff'] = df['temperature'] - df['apparenttemperature']
X = df[['humidity', 'windspeed']]
y = df['temp_diff']
plt.boxplot(df['temp_diff'])
plt.show()
plt.hist(df['temp_diff'])
plt.show()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 42)
# +
X_train_const = sm.add_constant(X_train)
lm = sm.OLS(y_train, X_train_const).fit()
lm.summary()
# -
# Both p-values for the independent vars are significant. The model is temperature difference = -2.4379 + 3.03humidity + 0.119windspeed which says that for each increase in humidity, temperature difference increases 3.03 degrees. For each 1 mph increase in windspeed, temperature difference increases by 0.119 degrees.
df["humid_wind_interaction"] = df.humidity * df.windspeed
X2 = df[['humid_wind_interaction', 'humidity', 'windspeed']]
X2_train, X2_test, y_train, y_test = train_test_split(X2, y, test_size=0.2, random_state = 42)
# +
X2_train_const = sm.add_constant(X2_train)
lm2 = sm.OLS(y_train, X2_train_const).fit()
lm2.summary()
# -
# The r2 increased and we now account for more variance. The model is now temperature difference = -0.1056 + 0.2943interation - 0.1488humidity -0.0889windspeed. All coeffs remain significant but have changed to being both negative coeffs. With each increase in 1 unit of interaction between humidity and windspeed, temperature diffence increases by 0.2943 degrees. With each increase in percent humidity, temperature difference decreases by 0.1488 degrees. With each increase in windspeed mph, temperature difference decreases by 0.0889 degrees.
# R2 has increased from our 1st model where it was 0.289 to our 2nd model with an R2 of .342 so it appears to capture more of the variance in the target but its still not good. Adjusted R2 has increased by the same amount as well. Our F statistic has a low pvalue in both models but has decreased from the first model from 15670 to 13350, however, both our IC scores decreased so I'm really not sure which model is better since I would really go back and add more variables at this point.
# created 3rd model
X3 = df[['humidity', 'windspeed','visibility']]
X3_train, X3_test, y_train, y_test = train_test_split(X3, y, test_size=0.2, random_state = 42)
# +
X3_train_const = sm.add_constant(X3_train)
lm3 = sm.OLS(y_train, X3_train_const).fit()
lm3.summary()
# -
# The 3rd model performs somewhere between model1 and model 2. The R2 and adj R2 dropped but is still better than the 1st model. The F statistic is significant but dropped even lower than our 1st model. Also, both IC scores increased suggesting model 3 is worse than model 1. I would go back and introduce more variabes, do othe interactions or look at distributions.
| week6_linear_regression/interpreting_coeffs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Adding a New Forecasting Model
#
# This notebook provides a minimal example on how to add a new forecasting model to Merlion. We follow the instructions in [CONTRIBUTING.md](https://github.com/salesforce/Merlion/blob/main/CONTRIBUTING.md). We suggest you review this [notebook](1_ForecastFeatures.ipynb) explaining how to use a Merlion forecasting model before reading this one.
#
# More specifically, let's implement a forecasting model whose forecast is just equal to the most recent observed value of the time series metric. For a more complete example, see our implementation of `Sarima` [here](https://github.com/salesforce/Merlion/blob/main/merlion/models/forecast/sarima.py).
# ## Model Config Class
#
# The first step of creating a new model is defining an appropriate config class, which inherits from `ForecasterConfig`:
# +
from merlion.models.forecast.base import ForecasterConfig
class RepeatRecentConfig(ForecasterConfig):
def __init__(self, max_forecast_steps=None, **kwargs):
super().__init__(max_forecast_steps=max_forecast_steps, **kwargs)
# -
# ## Model Class
#
# Next we define the model itself, which must inherit from the `ForecasterBase` base class and define all abstract methods. See the API docs for more details.
# +
from collections import OrderedDict
from typing import List, Tuple
from merlion.models.forecast.base import ForecasterBase
from merlion.utils import TimeSeries, UnivariateTimeSeries
class RepeatRecent(ForecasterBase):
# The config class for RepeatRecent is RepeatRecentConfig, defined above
config_class = RepeatRecentConfig
def __init__(self, config):
"""
Sets the model config and any other local variables. Here, we initialize
the most_recent_value to None.
"""
super().__init__(config)
self.most_recent_value = None
def train(self, train_data: TimeSeries, train_config=None) -> Tuple[TimeSeries, None]:
# Apply training preparation steps. We specify that this model doesn't
# require evenly sampled time series, and it doesn't require univariate
# data.
train_data = self.train_pre_process(
train_data, require_even_sampling=False, require_univariate=False)
# "Train" the model. Here, we just gather the most recent values
# for each univariate in the time series.
self.most_recent_value = OrderedDict((k, v.values[-1])
for k, v in train_data.items())
# The model's "prediction" for the training data, is just the value
# from one step before.
train_forecast = TimeSeries(OrderedDict(
(name, UnivariateTimeSeries(univariate.time_stamps,
[0] + univariate.values[:-1]))
for name, univariate in train_data.items()))
# This model doesn't have any notion of error
train_stderr = None
# Return the train prediction & standard error
return train_forecast, train_stderr
def forecast(self, time_stamps: List[int],
time_series_prev: TimeSeries = None,
return_iqr=False, return_prev=False
) -> Tuple[TimeSeries, None]:
# Use time_series_prev's most recent value if time_series_prev is given.
# Make sure to apply the data pre-processing transform on it first!
if time_series_prev is not None:
time_series_prev = self.transform(time_series_prev)
most_recent_value = {k: v.values[-1] for k, v in time_series_prev.items()}
# Otherwise, use the most recent value stored from the training data
else:
most_recent_value = self.most_recent_value
# The forecast is just the most recent value repeated for every upcoming
# timestamp
forecast = TimeSeries(OrderedDict(
(k, UnivariateTimeSeries(time_stamps, [v] * len(time_stamps)))
for k, v in most_recent_value.items()))
# Pre-pend time_series_prev to the forecast if desired
if return_prev and time_series_prev is not None:
forecast = time_series_prev + forecast
# Ensure we're not trying to return an inter-quartile range
if return_iqr:
raise RuntimeError(
"RepeatRecent model doesn't support uncertainty estimation")
return forecast, None
# -
# ## Running the Model: A Simple Example
#
# Let's try running this model on some actual data! This next part assumes you've installed `ts_datasets`. We'll begin by getting a time series from the M4 dataset & visualizing it.
# +
import matplotlib.pyplot as plt
import pandas as pd
from ts_datasets.forecast import M4
time_series, metadata = M4(subset="Hourly")[0]
# Visualize the full time series
fig = plt.figure(figsize=(12, 6))
ax = fig.add_subplot(111)
ax.plot(time_series)
# Label the train/test split with a dashed line
ax.axvline(time_series[metadata["trainval"]].index[-1], ls="--", lw=2, c="k")
plt.show()
# -
# Now, we'll split the data into train & test splits, and run our forecasting model on it.
train_data = TimeSeries.from_pd(time_series[metadata["trainval"]])
test_data = TimeSeries.from_pd(time_series[~metadata["trainval"]])
# Initialize a model & train it. The dataframe returned & printed
# below is the model's "forecast" on the training data. None is
# the uncertainty estimate.
model = RepeatRecent(RepeatRecentConfig())
model.train(train_data=train_data)
# Let's run our model on the test data now
forecast, err = model.forecast(test_data.to_pd().index)
print("Forecast")
print(forecast)
print()
print("Error")
print(err)
# ## Visualization
# +
# Qualitatively, we can see what the forecaster is doing by plotting
print("Forecast w/ ground truth time series")
fig, ax = model.plot_forecast(time_series=test_data,
time_series_prev=train_data,
plot_time_series_prev=True)
plt.show()
print()
print("Forecast without ground truth time series")
fig, ax = model.plot_forecast(time_stamps=test_data.to_pd().index,
time_series_prev=train_data,
plot_time_series_prev=True)
# -
# ## Quantitative Evaluation
#
# You may quantitatively evaluate your model as well. Here, we compute the sMAPE (symmetric Mean Average Percent Error) of the model's forecast vs. the true data. For ground truth $y \in \mathbb{R}^T$ and prediction $\hat{y} \in \mathbb{R}^T$, the sMAPE is computed as
#
# $$
# \mathrm{sMAPE}(y, \hat{y}) = \frac{200}{T} \sum_{t = 1}^{T} \frac{\lvert \hat{y}_t - y_t \rvert}{\lvert\hat{y}_t\rvert + \lvert y_t \rvert}
# $$
from merlion.evaluate.forecast import ForecastMetric
smape = ForecastMetric.sMAPE.value(ground_truth=test_data, predict=forecast)
print(f"sMAPE = {smape:.3f}")
# ## Defining a Forecaster-Based Anomaly Detector
#
# It is quite straightforward to adapt a forecasting model into an anomaly detection model. You just need to create a new file in the appropriate [directory](https://github.com/salesforce/Merlion/blob/main/merlion/models/anomaly/forecast_based) and define class stubs with some basic headers. Multiple inheritance with `ForecastingDetectorBase` takes care of most of the heavy lifting.
#
# The anomaly score returned by any forecasting-based anomaly detector is based on the residual between the predicted and true time series values.
# +
from merlion.evaluate.anomaly import TSADMetric
from merlion.models.anomaly.forecast_based.base import ForecastingDetectorBase
from merlion.models.anomaly.base import DetectorConfig
from merlion.post_process.threshold import AggregateAlarms
from merlion.transform.normalize import MeanVarNormalize
# Define a config class which inherits from RepeatRecentConfig and DetectorConfig
# in that order
class RepeatRecentDetectorConfig(RepeatRecentConfig, DetectorConfig):
# Set a default anomaly score post-processing rule
_default_post_rule = AggregateAlarms(alm_threshold=3.0)
# The default data pre-processing transform is mean-variance normalization,
# so that anomaly scores are roughly aligned with z-scores
_default_transform = MeanVarNormalize()
# Define a model class which inherits from ForecastingDetectorBase and RepeatRecent
# in that order
class RepeatRecentDetector(ForecastingDetectorBase, RepeatRecent):
# All we need to do is set the config class
config_class = RepeatRecentDetectorConfig
# -
# Train the anomaly detection variant
model2 = RepeatRecentDetector(RepeatRecentDetectorConfig())
model2.train(train_data)
# Obtain the anomaly detection variant's predictions on the test data
model2.get_anomaly_score(test_data)
# Visualize the anomaly detection variant's performance, with filtered anomaly scores
fig, ax = model2.plot_anomaly(test_data, time_series_prev=train_data,
filter_scores=True, plot_time_series_prev=False,
plot_forecast=True)
| examples/forecast/3_ForecastNewModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
# # Py(phar)maceuticals
#
# ### Load Data
#
# * Load the datasets: Mouse_metadata.csv and Study_results.csv
#
# ### Analysis
#
# 1. [Cleaning Data](#Cleaning_Data)
#
# 2. Summary for Tumor Volume vs. Treatment: [Summary Statistics](#Summary_Statistics)
#
# 3. The Number of Mice for Treatment: [Bar Plot](#Bar_Plot)
#
# 4. Mice Sex Distritubion: [Pie Plot](#Pie_Plot)
#
# 5. Final Tumor Volume: [Box Plot](#Box_Plot)
#
# 6. Tumor Volume Changes: [Line Plot](#Line_Plot)
#
# 7. Weight vs. Average Tumor Volume: [Scatter Plot](#Scatter_Plot)
#
# 8. Weight vs. Average Tumor Volume: [Correlation & Regression](#Regression)
#
# ### Observations and Insights
#
#
# * The Summary Statistics table shows that the mice with the Remicane treatment has the smallest average tumor volume. The mice with the Capomulin treatment has the next smallest tumor volume. The standard variations of them are also smallest. Their means, medians show minor gaps.
#
# - In terms of the overall performance, Ramicane is the best and Capomulin is the second best. Due to the similar central tendency, the top two treatments seem to perform similarly.
#
#
# * The Bar Plot shows that, throughout the course of the study, the total number of mice with Capomulin is the most. The Ramicane treatment is ranked the second with small difference.
#
# - Most mice treated with Capomulin and Ramicane could live much longer.
#
#
# * The Box Plot shows that (1) the final tumor volumes of mice treated by Capumulin and Ramicane have central tendency with small IQR (2) the maximum tumor volumes by Capumulin and Ramicane are similar to the minimum tumor volumes by Infubinol and Ceftamin (3) the upper and lower quartiles appear similar in Capomulin and Ramicane.
#
# - The mice treated with Capumulin or Ramicane have much smaller tumor at the final timepoint than the mice treated with Infubinol or Ceftamin. The performance of Capumulin and Ramicane seems similar.
#
#
# * The Scatter Plot shows that the tumor size is proportional to the mice weight with positive correlation. The Pearson correlation is 0.84.
#
# - The mice with more weight tends to have bigger tumor.
# Dependencies
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
# ## Load Data
# +
# Data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# -
print(f"* The number of rows is {mouse_metadata.shape[0]}\n")
print(f"* The numbers of unique items are")
print(mouse_metadata.nunique())
mouse_metadata.head()
print(f"* The number of rows is {study_results.shape[0]}\n")
print(f"* The numbers of unique items are")
print(study_results.nunique())
study_results.head()
# ## Analysis
#
#
# ### Cleaning Data <a id='Cleaning_Data'></a>
#
# * Combine the mouse metadata and the study results data.
# * Check the duplicates and clean up the duplicated data
# Combine the data into a single dataset
all_df = pd.merge(mouse_metadata, study_results, how="outer", on="Mouse ID")
# Find the duplicates
duplicates = all_df[all_df.duplicated(subset=["Mouse ID","Timepoint"], keep=False)]
duplicated_id = duplicates['Mouse ID'].unique()
print(f"The duplicates appear in the data of {duplicated_id[0]}")
# Optional: Get all the data for the duplicate mouse ID.
duplicate_data = all_df.loc[all_df['Mouse ID']=="g989"]
duplicate_data
# +
# Create a clean DataFrame by dropping all the data by its ID.
cleaned_df = all_df.drop(index=duplicate_data.index)
# Checking the number of mice in the clean DataFrame.
print(f"The number of mice in cleaned dataset is {cleaned_df['Mouse ID'].nunique()}")
# -
# ## Analysis
#
# ### Summary Statistics <a id='Summary_Statistics'></a>
#
# * Summary statistics table consisting of the mean, median, variance, standard deviation, and SEM of the tumor volume for each drug regimen.
# +
# This method is the most straightforward, creating multiple series
# and putting them all together at the end.
drug_list = cleaned_df['Drug Regimen'].unique()
drug_list.sort()
drug_vol_df = pd.DataFrame(index=drug_list, columns=['mean', 'median', 'var', 'std', 'sem'])
for drug in drug_list:
drug_vol = cleaned_df.loc[cleaned_df['Drug Regimen']==drug, 'Tumor Volume (mm3)']
drug_vol_stat = drug_vol.agg(['mean', 'median', 'var', 'std', 'sem'])
drug_vol_df.loc[drug] = drug_vol_stat.values
drug_vol_df.sort_values(by='mean')
# -
# This method produces everything in a single groupby function.
drug_vol_df2=cleaned_df.groupby('Drug Regimen')['Tumor Volume (mm3)'].agg(['mean','median','var','std','sem'])
drug_vol_df2.sort_values(by='mean')
# ## Bar Plots <a id='Bar_Plot'></a>
#
# * Showing the number of mice per time point for each treatment regimen throughout the course of the study.
# The number of mice per time point for each treatment
mice_time_regimen = cleaned_df.groupby(['Drug Regimen','Timepoint'])['Mouse ID'].nunique()
mice_time_regimen['Ramicane']
# +
# Total number of mice for each treatment throughout the timepoints
mice_regimen = mice_time_regimen.sum(level='Drug Regimen')
mice_regimen.sort_values(ascending=False, inplace=True)
# Generate a bar plot showing the number of mice throughout the timepoints for each treatment
# Using dataframe.plot
df_bar = mice_regimen.plot(kind='bar', rot='vertical', figsize=(8,4))
df_bar.set_xlabel("Treated Drugs")
df_bar.set_ylabel("The Number of Mice")
df_bar.set_title("Mice Count Throughout the Course of Study for Treatments")
df_bar.set_xlim(-0.75, len(mice_regimen.index)-0.25)
df_bar.set_ylim(0, 250)
# -
# Generate a bar plot showing the number of mice throughout the timepoints for each treatment
# Using matplot.plt
xval = range(len(mice_regimen.index))
plt.figure(figsize=(8,4))
plt.bar(xval, mice_regimen.values, width=0.5)
plt.xlabel("Treated Drugs")
plt.ylabel("The Number of Mice")
plt.title("Mice Count Throughout the Course of Study for Treatments")
plt.xticks(xval, mice_regimen.index, rotation="vertical")
plt.xlim(-0.75, len(mice_regimen.index)-0.25)
plt.ylim(0, 250)
plt.show()
# ## Pie Plots <a id='Pie_Plot'></a>
#
# * Showing the distribution of female or male mice in the study.
#
# distribution of female versus male mice using pandas
female_male = cleaned_df.groupby('Sex')['Mouse ID'].nunique()
female_male
# Generate a pie plot showing the distribution of female versus male mice using dataframe plot
pie_chart = female_male.plot(kind='pie', y=female_male.values, autopct='%1.1f%%', shadow=True)
pie_chart.set_ylabel("The Distrubution of Mice Sex")
# Generate a pie plot showing the distribution of female versus male mice using pyplot
labels = female_male.keys()
values = female_male.values
plt.pie(values, labels=labels, autopct='%1.1f%%', shadow=True)
plt.ylabel("The Distrubution of Mice Sex")
plt.axis("equal")
plt.show()
# ## Quartiles, Outliers and Boxplots <a id='Box_Plot'></a>
#
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens: Capomulin, Ramicane, Infubinol, and Ceftamin. Calculate the quartiles and IQR and quantitatively determine if there are any potential outliers across all four treatment regimens.
# +
# For a treatment(drug), find the tumor volume at the final time of each mouse which was treated by the drug
def find_volume_drug(df, final_time, drug):
vol = []
for key,val in final_time[drug].items():
df_val = df.loc[(df["Mouse ID"]==key) & (df["Timepoint"]==val),'Tumor Volume (mm3)'].values
vol.append(df_val[0])
return pd.Series(vol)
# Find final time for each mouse
final_time = cleaned_df.groupby(["Drug Regimen","Mouse ID"])['Timepoint'].max()
# Four treatments (drug regimens)
drug_list = ['Capomulin', 'Ramicane', 'Infubinol', 'Ceftamin']
# Set a dataframe of the final volumes for the treatments
final_vol = pd.DataFrame({
drug : find_volume_drug(cleaned_df, final_time, drug)
for drug in drug_list})
# Calculate the quartiles and IQR
quartiles_df = final_vol.quantile([.25,.5,.75])
iqr = quartiles_df.loc[0.75]-quartiles_df.loc[0.25]
quartiles_df = quartiles_df.append(iqr.rename('IQR'))
quartiles_df
# +
# Calculate the lower bound and upper bound
lower_b = quartiles_df.loc[0.25] - 1.5*quartiles_df.loc['IQR']
upper_b = quartiles_df.loc[0.75] + 1.5*quartiles_df.loc['IQR']
# Determine if there are any potential outliers
for drug in quartiles_df.columns:
low = lower_b[drug]
upp = upper_b[drug]
lcnt = final_vol.loc[final_vol[drug]<low, drug].count()
ucnt = final_vol.loc[final_vol[drug]>upp, drug].count()
print(f"The number of (lower, upper) outliers of {drug} is ({lcnt},{ucnt}).")
# -
# Using Matplotlib, generate a box and whisker plot
four_vol = [final_vol[drug] for drug in drug_list]
outlierprops = dict(markerfacecolor='red', marker='o')
plt.boxplot(four_vol, labels=drug_list, flierprops=outlierprops)
plt.ylabel("Final Tumer Volume (mm3)")
plt.title("Final Tumer Volume for Promising Treatments")
plt.show()
# ## Line Plot <a id='Line_Plot'></a>
#
# * Showing time point versus tumor volume for a single mouse treated with Capomulin.
# +
# Set a dataframe for the results of Capomulin
capomulin_df = cleaned_df.loc[(cleaned_df['Drug Regimen']=='Capomulin')]
# Find mice IDs treated with Capomulin
mice_id_capomulin = capomulin_df['Mouse ID'].unique()
print(f"The mice treated with Capomulin are\n {mice_id_capomulin}")
# +
# Select one mouse ID, randomly, and its data
mouse_id = mice_id_capomulin[st.randint.rvs(0,len(mice_id_capomulin))]
one_mouse_df = capomulin_df.loc[(capomulin_df['Mouse ID']==mouse_id)]
# Generate a line plot of time point versus tumor volume for a mouse treated with Capomulin
xval = one_mouse_df['Timepoint']
yval = one_mouse_df['Tumor Volume (mm3)']
plt.plot(xval,yval)
plt.xlabel("Timepoint")
plt.ylabel("Tumor Volume (mm3)")
plt.title(f"Capomulin Treatment for Mouse-{mouse_id}")
plt.xlim(0,45)
plt.ylim(0,50)
plt.show()
# -
# ## Scatter Plot <a id='Scatter_Plt'></a>
#
# * Plot the mouse weight versus average tumor volume for the Capomulin treatment regimen.
# Generate a scatter plot of, for each mouse, mouse weight versus average tumor volume for the Capomulin regimen
capomulin_vol = capomulin_df.groupby('Mouse ID')['Tumor Volume (mm3)'].mean()
capomulin_wt = capomulin_df.groupby('Mouse ID')['Weight (g)'].min()
xval = capomulin_wt.values
yval = capomulin_vol.values
plt.scatter(xval, yval)
plt.xlabel('Weight (g)')
plt.ylabel('Aerage Tumor Volume (mm3)')
plt.title('Capomulin Treatment')
plt.tight_layout()
plt.show()
# ## Correlation and Regression <a id='Regression'></a>
#
# Calculate the correlation coefficient and linear regression model between mouse weight and average tumor volume for the Capomulin treatment. Plot the linear regression model on top of the previous scatter plot.
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
(mm,cc,rr,pp,ss) = st.linregress(xval,yval)
line_eq = "y = " + str(round(mm,2)) + " x + " + str(round(cc,2))
est_yval = mm*xval + cc
plt.plot(xval, est_yval, "r-")
plt.scatter(xval, yval)
plt.annotate(line_eq,(17.5,37), color="red")
plt.xlabel('Weight (g)')
plt.ylabel('Aerage Tumor Volume (mm3)')
plt.title('Capomulin Treatment')
plt.tight_layout()
plt.show()
(pr_r, pr_p)=st.pearsonr(xval,yval)
print(f"The Pearson (r, p) values are ({pr_r:.2f}, {pr_p:.2f})")
print("The R-square is " + str(round(rr**2,2)))
print("The pvalue is " + str(round(pp,2)))
| pymaceuticals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inverse dynamics (2D) for gait analysis
#
# > <NAME>, <NAME>
# > [Laboratory of Biomechanics and Motor Control](http://pesquisa.ufabc.edu.br/bmclab/)
# > Federal University of ABC, Brazil
# + [markdown] toc=1
# <h1>Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Forward-and-inverse-dynamics" data-toc-modified-id="Forward-and-inverse-dynamics-1"><span class="toc-item-num">1 </span>Forward and inverse dynamics</a></span></li><li><span><a href="#Estimation-of-joint-force-and-moments-of-force-by-inverse-dynamics" data-toc-modified-id="Estimation-of-joint-force-and-moments-of-force-by-inverse-dynamics-2"><span class="toc-item-num">2 </span>Estimation of joint force and moments of force by inverse dynamics</a></span><ul class="toc-item"><li><span><a href="#Free-body-diagrams" data-toc-modified-id="Free-body-diagrams-2.1"><span class="toc-item-num">2.1 </span>Free body diagrams</a></span></li><li><span><a href="#Equations-of-motion" data-toc-modified-id="Equations-of-motion-2.2"><span class="toc-item-num">2.2 </span>Equations of motion</a></span></li><li><span><a href="#The-recursive-approach-for-inverse-dynamics-of-multi-body-systems" data-toc-modified-id="The-recursive-approach-for-inverse-dynamics-of-multi-body-systems-2.3"><span class="toc-item-num">2.3 </span>The recursive approach for inverse dynamics of multi-body systems</a></span></li><li><span><a href="#Python-function-invdyn2d.py" data-toc-modified-id="Python-function-invdyn2d.py-2.4"><span class="toc-item-num">2.4 </span>Python function <code>invdyn2d.py</code></a></span></li><li><span><a href="#Experimental-data" data-toc-modified-id="Experimental-data-2.5"><span class="toc-item-num">2.5 </span>Experimental data</a></span></li><li><span><a href="#Load-data-file" data-toc-modified-id="Load-data-file-2.6"><span class="toc-item-num">2.6 </span>Load data file</a></span></li><li><span><a href="#Data-filtering" data-toc-modified-id="Data-filtering-2.7"><span class="toc-item-num">2.7 </span>Data filtering</a></span></li><li><span><a href="#Data-selection" data-toc-modified-id="Data-selection-2.8"><span class="toc-item-num">2.8 </span>Data selection</a></span></li><li><span><a href="#Plot-file-data" data-toc-modified-id="Plot-file-data-2.9"><span class="toc-item-num">2.9 </span>Plot file data</a></span></li><li><span><a href="#Body-segment-parameters" data-toc-modified-id="Body-segment-parameters-2.10"><span class="toc-item-num">2.10 </span>Body-segment parameters</a></span></li><li><span><a href="#Kinematic-calculations" data-toc-modified-id="Kinematic-calculations-2.11"><span class="toc-item-num">2.11 </span>Kinematic calculations</a></span></li><li><span><a href="#Plot-joint-angles" data-toc-modified-id="Plot-joint-angles-2.12"><span class="toc-item-num">2.12 </span>Plot joint angles</a></span></li><li><span><a href="#Inverse-dynamics-calculations" data-toc-modified-id="Inverse-dynamics-calculations-2.13"><span class="toc-item-num">2.13 </span>Inverse dynamics calculations</a></span></li><li><span><a href="#Load-files-with-true-joint-forces-and-moments-of-force" data-toc-modified-id="Load-files-with-true-joint-forces-and-moments-of-force-2.14"><span class="toc-item-num">2.14 </span>Load files with true joint forces and moments of force</a></span></li><li><span><a href="#Plot-calculated-variables-and-their-true-values" data-toc-modified-id="Plot-calculated-variables-and-their-true-values-2.15"><span class="toc-item-num">2.15 </span>Plot calculated variables and their true values</a></span></li></ul></li><li><span><a href="#Contribution-of-each-term-to-the-joint-force-and-moment-of-force" data-toc-modified-id="Contribution-of-each-term-to-the-joint-force-and-moment-of-force-3"><span class="toc-item-num">3 </span>Contribution of each term to the joint force and moment of force</a></span><ul class="toc-item"><li><span><a href="#Quasi-static-analysis" data-toc-modified-id="Quasi-static-analysis-3.1"><span class="toc-item-num">3.1 </span>Quasi-static analysis</a></span></li><li><span><a href="#Neglecting-the-acceleration-and-mass-(weight)-of-the-segments" data-toc-modified-id="Neglecting-the-acceleration-and-mass-(weight)-of-the-segments-3.2"><span class="toc-item-num">3.2 </span>Neglecting the acceleration and mass (weight) of the segments</a></span></li><li><span><a href="#WARNING:-the-calculated-resultant-joint-force-is-not-the-actual-joint-reaction-force!" data-toc-modified-id="WARNING:-the-calculated-resultant-joint-force-is-not-the-actual-joint-reaction-force!-3.3"><span class="toc-item-num">3.3 </span>WARNING: the calculated resultant joint force is not the actual joint reaction force!</a></span></li></ul></li><li><span><a href="#Conclusion" data-toc-modified-id="Conclusion-4"><span class="toc-item-num">4 </span>Conclusion</a></span></li><li><span><a href="#Further-reading" data-toc-modified-id="Further-reading-5"><span class="toc-item-num">5 </span>Further reading</a></span></li><li><span><a href="#Video-lectures-on-the-Internet" data-toc-modified-id="Video-lectures-on-the-Internet-6"><span class="toc-item-num">6 </span>Video lectures on the Internet</a></span></li><li><span><a href="#Problems" data-toc-modified-id="Problems-7"><span class="toc-item-num">7 </span>Problems</a></span></li><li><span><a href="#References" data-toc-modified-id="References-8"><span class="toc-item-num">8 </span>References</a></span></li></ul></div>
# -
# ## Forward and inverse dynamics
#
# With respect to the equations of motion to determine the dynamics of a system, there are two general approaches: forward (or direct) and inverse dynamics. For example, consider the solution of Newton's second law for a particle. If we know the force(s) and want to find the trajectory, this is **forward dynamics**. If instead, we know the trajectory and want to find the force(s), this is **inverse dynamics**:
#
# <figure><img src="./../images/dynamics.png" alt="Forward and inverse dynamics." width=220/><figcaption><i><center>Figure. The equation of motion and the forward and inverse dynamics approaches.</center></i></figcaption></figure>
#
# In Biomechanics, in a typical movement analysis of the human body using inverse dynamics, we would measure the positions of the segments and measure the external forces, calculate the segments' linear and angular acceleration, and find the internal net force and moment of force at the joint using the equations of motion. In addition, we could estimate the muscle forces (if we solve the redundancy problem of having more muscles than joints).
# Using forward dynamics, the muscle forces would be the inputs and the trajectories of the segments would be the outputs. The figure below compares the forward and inverse dynamics approaches.
#
# <figure><img src="./../images/InvDirDyn.png" alt="Direct and inverse dynamics."/><figcaption><i><center>Figure. Inverse dynamics and Forward (or Direct) dynamics approaches for movement analysis (adapted from Zajac and Gordon, 1989).</center></i></figcaption></figure>
# ## Estimation of joint force and moments of force by inverse dynamics
#
# Let's estimate the joint force and moments of force at the lower limb during locomotion using the inverse dynamics approach.
# We will model the lower limbs at the right side as composed by three rigid bodies (foot, leg, and thigh) articulated by three hinge joints (ankle, knee, and hip) and perform a two-dimensional analysis.
#
# ### Free body diagrams
#
# The [free body diagrams](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/FreeBodyDiagram.ipynb) of the lower limbs are:
# <br>
# <figure><img src="./../images/fbdgaitb.png" width=640 alt="FBD lowerlimb"/><figcaption><center><i>Figure. Free body diagrams of the lower limbs for a gait analysis. <b>GRF</b> is the resultant ground reaction force applied on the foot at the center of pressure, <b>COP</b>, position.</i></center></figcaption></figure>
#
# ### Equations of motion
#
# The equilibrium equations for the forces and moments of force around the center of mass are:
#
# For body 1 (foot):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_1 + m_1\mathbf{g} + \mathbf{GRF} = m_1\mathbf{a}_1 \\
# \mathbf{M}_1 + \mathbf{r}_{cmp1}\times\mathbf{F}_1 + \mathbf{r}_{cmCOP}\times\mathbf{GRF} = I_1\mathbf{\alpha}_1
# \label{}
# \end{array}
# \end{equation}
#
# For body 2 (leg):
#
# \begin{equation}
# \begin{array}{l}
# \mathbf{F}_2 + m_2\mathbf{g} - \mathbf{F}_1 = m_2\mathbf{a}_2 \\
# \mathbf{M}_2 + \mathbf{r}_{cmp2}\times\mathbf{F}_2 + \mathbf{r}_{cmd2}\times-\mathbf{F}_{1} - \mathbf{M}_1 = I_2\mathbf{\alpha}_2
# \label{}
# \end{array}
# \end{equation}
#
# For body 3 (thigh):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_3 + m_3\mathbf{g} - \mathbf{F}_2 = m_3\mathbf{a}_3 \\
# \mathbf{M}_3 + \mathbf{r}_{cmp3}\times\mathbf{F}_3 + \mathbf{r}_{cmd3}\times-\mathbf{F}_{2} - \mathbf{M}_2 = I_3\mathbf{\alpha}_3
# \label{}
# \end{array}
# \end{equation}
#
# Where $p$ and $d$ stands for proximal and distal joints (with respect to the fixed extremity), $\mathbf{r}_{cmji}$ is the position vector from the center of mass of body $i$ to the joint $j$, $COP$ is the center of pressure, the position of application of the resultant ground reaction force (GRF), $\mathbf{\alpha}$ is the angular acceleration, and $g$ is the acceleration of gravity
#
# Note that the pattern of the equations is the same for the three segments: distal and proximal forces and moments of force and the weight force are present in all segments.
# The only exception is with the foot in contact with the ground. As the ground only pushes the foot, it can not generate a moment of force over the foot. Because of that we model the interaction foot-ground as a resultant ground reaction force (GRF) applied on the foot at the COP position.
#
# Both GRF and COP quantities are measured with a force platform and are assumed as known quantities.
# Because of that the system of equations above is only solvable if we start by the body 1, from bottom to top.
# The system of equations above is simple and straightforward to solve, it is just a matter of being systematic.
# We start by segment 1, find $\mathbf{F}_1$ and $\mathbf{M}_1$, substitute these values on the equations for segment 2, find $\mathbf{F}_2$ and $\mathbf{M}_2$, substitute them in the equations for segment 3 and find $\mathbf{F}_3$ and $\mathbf{M}_3\:$:
#
# For body 1 (foot):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_1 &=& m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF} \\
# \mathbf{M}_1 &=& I_1\mathbf{\alpha}_1 - \mathbf{r}_{cmp1}\times\big(m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF}\big) - \mathbf{r}_{cmCOP}\times\mathbf{GRF}
# \label{}
# \end{array}
# \end{equation}
#
# For body 2 (leg):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_2 &=& m_1\mathbf{a}_1 + m_2\mathbf{a}_2 - (m_1+m_2)\mathbf{g} - \mathbf{GRF} \\
# \mathbf{M}_2 &=& I_1\mathbf{\alpha}_1 + I_2\mathbf{\alpha}_2 - \mathbf{r}_{cmp2}\times\big(m_1\mathbf{a}_1 + m_2\mathbf{a}_2 - (m_1+m_2)\mathbf{g} - \mathbf{GRF}\big)\, + \\
# &\phantom{=}& \mathbf{r}_{cgd2}\times\big(m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF}\big) - \mathbf{r}_{cmp1}\times\big(m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF}\big)\, - \\
# &\phantom{=}& \mathbf{r}_{cmCOP}\times\mathbf{GRF}
# \label{}
# \end{array}
# \end{equation}
#
# For body 3 (thigh):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_3 &=& m_1\mathbf{a}_1 + m_2\mathbf{a}_2 + m_3\mathbf{a}_3 - (m_1+m_2+m_3)\mathbf{g} - \mathbf{GRF} \\
# \mathbf{M}_3 &=& I_1\mathbf{\alpha}_1 + I_2\mathbf{\alpha}_2 + I_3\mathbf{\alpha}_3\, - \\
# &\phantom{=}& \mathbf{r}_{cmp3}\times\big(m_1\mathbf{a}_1 + m_2\mathbf{a}_2 + m_3\mathbf{a}_3\, - (m_1+m_2+m_3)\mathbf{g} - \mathbf{GRF}\big)\, + \\
# &\phantom{=}& \mathbf{r}_{cmd3}\times\big(m_1\mathbf{a}_1 + m_2\mathbf{a}_2 - (m_1+m_2)\mathbf{g} - \mathbf{GRF}\big)\, - \\
# &\phantom{=}& \mathbf{r}_{cmp2}\times\big(m_1\mathbf{a}_1 + m_2\mathbf{a}_2 - (m_1+m_2)\mathbf{g} - \mathbf{GRF}\big)\, + \\
# &\phantom{=}& \mathbf{r}_{cmd2}\times\big(m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF}\big)\, - \\
# &\phantom{=}& \mathbf{r}_{cmp1}\times\big(m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF}\big)\, - \\
# &\phantom{=}& \mathbf{r}_{cmCOP}\times\mathbf{GRF}
# \label{}
# \end{array}
# \end{equation}
# ### The recursive approach for inverse dynamics of multi-body systems
#
# The calculation above is tedious, error prone, useless, and probably it's wrong.
#
# To make some use of it, we can clearly see that forces act on far segments, which are not directly in contact with these forces. In fact, this is true for all stuff happening on a segment: note that $\mathbf{F}_1$ and $\mathbf{M}_1$ are present in the expression for $\mathbf{F}_3$ and $\mathbf{M}_3$ and that the acceleration of segment 1 matters for the calculations of segment 3.
#
# Instead, we can use the power of computer programming (like this one right now!) and solve these equations recursively hence they have the same pattern. Let's do that.
#
# For body 1 (foot):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_1 = m_1\mathbf{a}_1 - m_1\mathbf{g} - \mathbf{GRF} \\
# \mathbf{M}_1 = I_1\mathbf{\alpha}_1 - \mathbf{r}_{cmp1}\times\mathbf{F}_1 - \mathbf{r}_{cmCOP}\times\mathbf{GRF}
# \label{}
# \end{array}
# \end{equation}
#
# For body 2 (leg):
#
# \begin{equation}
# \begin{array}{l}
# \mathbf{F}_2 = m_2\mathbf{a}_2 - m_2\mathbf{g} + \mathbf{F}_1\\
# \mathbf{M}_2 = I_2\mathbf{\alpha}_2 - \mathbf{r}_{cmp2}\times\mathbf{F}_2 +\mathbf{r}_{cmd2}\times\mathbf{F}_{1} + \mathbf{M}_1
# \label{}
# \end{array}
# \end{equation}
#
# For body 3 (thigh):
#
# \begin{equation}
# \begin{array}{l l}
# \mathbf{F}_3 = m_3\mathbf{a}_3 - m_3\mathbf{g} + \mathbf{F}_2\\
# \mathbf{M}_3 = I_3\mathbf{\alpha}_3 - \mathbf{r}_{cmp3}\times\mathbf{F}_3 + \mathbf{r}_{cmd3}\times\mathbf{F}_{2} + \mathbf{M}_2
# \label{}
# \end{array}
# \end{equation}
# ### Python function `invdyn2d.py`
#
# We could write a function that it would have as inputs the body-segment parameters, the kinematic data, and the distal joint force and moment of force and output the proximal joint force and moment of force.
# Then, we would call this function for each segment, starting with the segment that has a free extremity or that has the force and moment of force measured by some instrument (i,e, use a force plate for the foot-ground interface).
# This function would be called in the following manner:
#
# ```python
# Fp, Mp = invdyn2d(rcm, rd, rp, acm, alfa, mass, Icm, Fd, Md)
# ```
#
# So, here is such function:
# +
# # %load ./../functions/invdyn2d.py
"""Two-dimensional inverse-dynamics calculations of one segment."""
__author__ = '<NAME>, https://github.com/demotu/BMC'
__version__ = 'invdyn2d.py v.2 2015/11/13'
def invdyn2d(rcm, rd, rp, acm, alpha, mass, Icm, Fd, Md):
"""Two-dimensional inverse-dynamics calculations of one segment
Parameters
----------
rcm : array_like [x,y]
center of mass position (y is vertical)
rd : array_like [x,y]
distal joint position
rp : array_like [x,y]
proximal joint position
acm : array_like [x,y]
center of mass acceleration
alpha : array_like [x,y]
segment angular acceleration
mass : number
mass of the segment
Icm : number
rotational inertia around the center of mass of the segment
Fd : array_like [x,y]
force on the distal joint of the segment
Md : array_like [x,y]
moment of force on the distal joint of the segment
Returns
-------
Fp : array_like [x,y]
force on the proximal joint of the segment (y is vertical)
Mp : array_like [x,y]
moment of force on the proximal joint of the segment
Notes
-----
To use this function recursevely, the outputs [Fp, Mp] must be inputed as
[-Fp, -Mp] on the next call to represent [Fd, Md] on the distal joint of the
next segment (action-reaction).
This code was inspired by a similar code written by <NAME> [1]_.
See this notebook [2]_.
References
----------
.. [1] http://isbweb.org/data/invdyn/index.html
.. [2] http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/GaitAnalysis2D.ipynb
"""
from numpy import cross
g = 9.80665 # m/s2, standard acceleration of free fall (ISO 80000-3:2006)
# Force and moment of force on the proximal joint
Fp = mass*acm - Fd - [0, -g*mass]
Mp = Icm*alpha - Md - cross(rd-rcm, Fd) - cross(rp-rcm, Fp)
return Fp, Mp
# -
# The inverse dynamics calculations are implemented in only two lines of code at the end, the first part of the code is the help on how to use the function. The help is long because it's supposed to be helpful :), see the [style guide for NumPy/SciPy documentation](https://github.com/numpy/numpy/blob/master/doc/HOWTO_DOCUMENT.rst.txt).
#
# The real problem is to measure or estimate the experimental variables: the body-segment parameters, the ground reaction forces, and the kinematics of each segment. For such, it is necessary some expensive equipments, but they are typical in a biomechanics laboratory, such the the [BMClab](http://pesquisa.ufabc.edu.br/bmclab).
# ### Experimental data
#
# Let's work with some data of kinematic position of the segments and ground reaction forces in order to compute the joint forces and moments of force.
# The data we will work are in fact from a computer simulation of running created by <NAME>. The nice thing about these data is that as a simulation, the true joint forces and moments of force are known and we will be able to compare our estimation with these true values.
# All the data can be downloaded from a page at the [ISB website](http://isbweb.org/data/invdyn/index.html):
from IPython.display import IFrame
IFrame('http://isbweb.org/data/invdyn/index.html', width='100%', height=400)
# import the necessary libraries
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_context('notebook', font_scale=1.2, rc={"lines.linewidth": 2})
import sys
sys.path.insert(1, r'./../functions')
# ### Load data file
# load file with ground reaction force data
grf = np.loadtxt('./../data/all.frc') # [Fx, Fy, COPx]
# load file with kinematic data
kin = np.loadtxt('./../data/all.kin') # [Hip(x,y), knee(x,y), ankle(x,y), toe(x,y)]
freq = 10000
time = np.linspace(0, grf.shape[0]/freq, grf.shape[0])
# ### Data filtering
# this is simulated data with no noise, filtering doesn't matter
if False:
# filter data
from scipy.signal import butter, filtfilt
# Butterworth filter
b, a = butter(2, (10/(freq/2)))
for col in np.arange(grf.shape[1]-1):
grf[:, col] = filtfilt(b, a, grf[:, col])
b, a = butter(2, (10/(freq/2)))
for col in np.arange(kin.shape[1]):
kin[:, col] = filtfilt(b, a, kin[:, col])
# ### Data selection
# heel strike occurs at sample 3001
time = time[3001 - int(freq/40):-int(freq/20)]
grf = grf[3001 - int(freq/40):-int(freq/20), :]
kin = kin[3001 - int(freq/40):-int(freq/20), :]
# ### Plot file data
# plot data
hfig, hax = plt.subplots(2, 2, sharex = True, squeeze=True, figsize=(9, 5))
hax[0, 0].plot(time, grf[:, [0, 1]], linewidth=2)
hax[0, 0].legend(('Fx','Fy'), frameon=False)
hax[0, 0].set_ylabel('Force [N]')
hax[0, 1].plot(time, grf[:, 2], linewidth=2)
hax[0, 1].legend(['COPx'], frameon=False)
hax[0, 1].set_ylabel('Amplitude [m]')
hax[1, 0].plot(time, kin[:, 0::2], linewidth=2)
hax[1, 0].legend(('Hip x','Knee x','Ankle x','Toe x'), frameon=False)
hax[1, 0].set_ylabel('Amplitude [m]')
hax[1, 1].plot(time, kin[:, 1::2], linewidth=2)
hax[1, 1].legend(('Hip y','Knee y','Ankle y','Toe y'), frameon=False)
hax[1, 1].set_ylabel('Amplitude [m]')
hax[1, 0].set_xlabel('Time [s]'), hax[1, 1].set_xlabel('Time [s]')
plt.tight_layout()
plt.show()
# ### Body-segment parameters
# body-segment parameters [thigh, shank, foot]
mass = [6.85, 2.86, 1.00] # mass [kg]
Icm = [0.145361267, 0.042996389, 0.0200] # rotational inertia [kgm2]
cmpr = [0.4323725, 0.4334975, 0.0] # CM [m] wrt. prox. joint [frac. segment len]
# ### Kinematic calculations
# +
# Kinematic data
# center of mass position of the thigh, shank, foot segments
rcm = np.hstack((kin[:, (0,1)] + cmpr[0]*(kin[:, (2,3)] - kin[:, (0,1)]),
kin[:, (2,3)] + cmpr[1]*(kin[:, (4,5)] - kin[:, (2,3)]),
kin[:, (4,5)] + cmpr[2]*(kin[:, (6,7)] - kin[:, (4,5)])))
# center of mass linear acceleration of the thigh, shank, foot segments
acm = np.diff(rcm, n=2, axis=0)*freq*freq
acm = np.vstack((acm, acm[-1, :], acm[-1, :]))
# thigh, shank, foot segment angle
ang = np.vstack((np.arctan2(kin[:, 1] - kin[:, 3], kin[:, 0] - kin[:, 2]),
np.arctan2(kin[:, 3] - kin[:, 5], kin[:, 2] - kin[:, 4]),
np.arctan2(kin[:, 5] - kin[:, 7], kin[:, 4] - kin[:, 6]))).T
# hip, knee, and ankle joint angles
angj = np.vstack((-(ang[:, 0]-ang[:, 1]),
np.unwrap(ang[:, 1] - ang[:, 2] + np.pi/2))).T*180/np.pi
# thigh, shank, foot segment angular acceleration
aang = np.diff(ang, n=2, axis=0)*freq*freq
aang = np.vstack((aang, aang[-1, :], aang[-1, :]))
# -
# ### Plot joint angles
# plot hip, knee, and ankle joint angles
hfig, (hax1, hax2) = plt.subplots(2, 1, sharex = True, squeeze=True, figsize=(10, 5))
hax1.plot(time, angj[:, 0], linewidth=2, label='Knee')
hax1.legend(frameon=False, loc='upper left'), hax1.grid()
hax2.plot(time, angj[:, 1], linewidth=2, label='Ankle')
hax2.legend(frameon=False, loc='upper left'), hax2.grid()
hax1.set_ylabel('Joint angle $[^o]$')
hax2.set_ylabel('Joint angle $[^o]$')
hax2.set_xlabel('Time [s]')
plt.tight_layout()
plt.show()
# ### Inverse dynamics calculations
# +
# inverse dynamics
# invdyn2d(rcm, rd, rp, acm, alpha, mass, Icm, Fd, Md)
from invdyn2d import invdyn2d
# ankle
[Fa, Ma] = invdyn2d(rcm[:,(4,5)], grf[:,(2,2)]*[1,0], kin[:,(4,5)],
acm[:,(4,5)], aang[:, 2], mass[2], Icm[2],
grf[:, (0, 1)], 0)
# knee
[Fk, Mk] = invdyn2d(rcm[:,(2,3)], kin[:,(4,5)], kin[:,(2,3)],
acm[:,(2,3)], aang[:,1], mass[1], Icm[1],
-Fa, -Ma)
# hip
[Fh, Mh] = invdyn2d(rcm[:,(0,1)], kin[:,(2,3)], kin[:,(0,1)],
acm[:,(0,1)], aang[:,0], mass[0], Icm[0],
-Fk, -Mk)
# magnitude of the calculated hip, knee, and ankle resultant joint force
Fam = np.sqrt(np.sum(np.abs(Fa)**2, axis=-1))
Fkm = np.sqrt(np.sum(np.abs(Fk)**2, axis=-1))
Fhm = np.sqrt(np.sum(np.abs(Fh)**2, axis=-1))
# -
# ### Load files with true joint forces and moments of force
# load file with true joint forces and moments of force
forces = np.loadtxt('./../data/all.fmg') # [Hip, knee, ankle]
moments = np.loadtxt('./../data/all.mom') # [Hip, knee, ankle]
#heel strike occurs at sample 3001
forces = forces[3001-int(freq/40):-int(freq/20), :]
moments = moments[3001-int(freq/40):-int(freq/20), :]
# ### Plot calculated variables and their true values
#
# Let's plot these data but because later we will need to plot similar plots, let's create a function for the plot to avoid repetition of code:
# +
def plotdata(time, Fh, Fk, Fa, Mh, Mk, Ma, forces, moments, stitle):
# plot hip, knee, and ankle moments of force
hfig, hax = plt.subplots(3, 2, sharex = True, squeeze=True, figsize=(11, 6))
# forces
hax[0, 0].plot(time, Fh, label='invdyn'), hax[0, 0].set_title('Hip')
hax[1, 0].plot(time, Fk), hax[1, 0].set_title('Knee')
hax[2, 0].plot(time, Fa), hax[2, 0].set_title('Ankle')
hax[1, 0].set_ylabel('Joint force [N]')
hax[2, 0].set_xlabel('Time [s]')
# moments of force
hax[0, 1].plot(time, Mh), hax[0, 1].set_title('Hip')
hax[1, 1].plot(time, Mk), hax[1, 1].set_title('Knee')
hax[2, 1].plot(time, Ma), hax[2, 1].set_title('Ankle')
hax[1, 1].set_ylabel('Moment of Force [Nm]')
hax[2, 1].set_xlabel('Time [s]')
# true joint forces and moments of force
hax[0, 0].plot(time, forces[:, 0], 'r--', label='True')
hax[0, 0].legend(frameon=False)
hax[1, 0].plot(time, forces[:, 1], 'r--')
hax[2, 0].plot(time, forces[:, 2], 'r--')
hax[0, 1].plot(time, moments[:, 0], 'r--')
hax[1, 1].plot(time, moments[:, 1], 'r--')
hax[2, 1].plot(time, moments[:, 2], 'r--')
plt.suptitle(stitle, fontsize=16)
for x in hax.flat:
x.locator_params(nbins=5); x.grid()
plt.show()
plotdata(time, Fhm, Fkm, Fam, Mh, Mk, Ma, forces, moments,
'Inverse dynamics: estimated versus true values')
# -
# The results are very similar; only a small part of the moments of force is different because of some noise.
# ## Contribution of each term to the joint force and moment of force
#
# Let's see what happens with the joint forces and moments of force when we neglect the contribution of some terms in the inverse dynamics analysis of these data.
# ### Quasi-static analysis
# Consider the case where the segment acceleration is neglected:
# +
# ankle
[Fast, Mast] = invdyn2d(rcm[:,(4,5)], grf[:,(2,2)]*[1,0], kin[:,(4,5)],
acm[:,(4,5)]*0, aang[:,2]*0, mass[2], Icm[2],
grf[:,(0,1)], 0)
# knee
[Fkst, Mkst] = invdyn2d(rcm[:,(2,3)], kin[:,(4,5)], kin[:,(2,3)],
acm[:,(2,3)]*0, aang[:,1]*0, mass[1], Icm[1],
-Fast, -Mast)
# hip
[Fhst, Mhst] = invdyn2d(rcm[:,(0,1)], kin[:,(2,3)], kin[:,(0,1)],
acm[:,(0,1)]*0, aang[:,0]*0, mass[0], Icm[0],
-Fkst, -Mkst)
# magnitude of the calculated hip, knee, and ankle resultant joint force
Fastm = np.sqrt(np.sum(np.abs(Fast)**2, axis=-1))
Fkstm = np.sqrt(np.sum(np.abs(Fkst)**2, axis=-1))
Fhstm = np.sqrt(np.sum(np.abs(Fhst)**2, axis=-1))
plotdata(time, Fhstm, Fkstm, Fastm, Mhst, Mkst, Mast, forces, moments,
'Inverse dynamics: quasis-static approach versus true values')
# -
# This is not a pure static analysis because part of the ground reaction forces still reflects the body accelerations (were the body completely static, the ground reaction force should be equal to the body weight in magnitude).
# ### Neglecting the acceleration and mass (weight) of the segments
#
# Consider the case where besides the acceleration, the body-segment parameters are also neglected.
# This means that the joint loads are due only to the ground reaction forces (which implicitly include contributions due to the acceleration and the body-segment weights).
# +
# ankle
[Fagrf, Magrf] = invdyn2d(rcm[:, (4,5)], grf[:,(2,2)]*[1,0], kin[:,(4,5)],
acm[:,(4,5)]*0, aang[:,2]*0, 0, 0, grf[:,(0,1)], 0)
# knee
[Fkgrf, Mkgrf] = invdyn2d(rcm[:,(2,3)], kin[:,(4,5)], kin[:,(2,3)],
acm[:,(2,3)]*0, aang[:,1]*0, 0, 0, -Fagrf, -Magrf)
# hip
[Fhgrf, Mhgrf] = invdyn2d(rcm[:,(0,1)], kin[:,(2,3)], kin[:,(0,1)],
acm[:,(0,1)]*0, aang[:, 0]*0, 0, 0, -Fkgrf, -Mkgrf)
# magnitude of the calculated hip, knee, and ankle resultant joint force
Fagrfm = np.sqrt(np.sum(np.abs(Fagrf)**2, axis=-1))
Fkgrfm = np.sqrt(np.sum(np.abs(Fkgrf)**2, axis=-1))
Fhgrfm = np.sqrt(np.sum(np.abs(Fhgrf)**2, axis=-1))
plotdata(time, Fhgrfm, Fkgrfm, Fagrfm, Mhgrf, Mkgrf, Magrf, forces, moments,
'Inverse dynamics: ground-reaction-force approach versus true values')
# -
# Neglecting all the accelerations and the weight of the segments means that the only external force that actuates on the system is the ground reaction force, which although is only actuating at the foot-ground interface it will be transmitted to the other segments through the joint forces. Because of that, the joint forces on the ankle, knee, and hip will simply be minus the ground reaction force. Note that the forces shown above for the three joints are the same and equal to:
#
# \begin{equation}
# \begin{array}{l}
# \sqrt{GRF_x^2+GRF_y^2}
# \label{}
# \end{array}
# \end{equation}
#
# These simplifications also mean that the moments of force could have been simply calculated as the cross product between the vector position of the the COP in relation to the joint and the GRF vector:
#
# \begin{equation}
# \begin{array}{l}
# \mathbf{M_{a}} = -\mathbf{cross}(\mathbf{COP}-\mathbf{r_{a}},\,\mathbf{GRF}) \\
# \mathbf{M_{k}} = -\mathbf{cross}(\mathbf{COP}-\mathbf{r_{k}},\,\mathbf{GRF}) \\
# \mathbf{M_{h}} = -\mathbf{cross}(\mathbf{COP}-\mathbf{r_{h}},\,\mathbf{GRF})
# \label{}
# \end{array}
# \end{equation}
#
# Where $\mathbf{r_{i}}\;$ is the position vector of joint $i$.
#
# Let's calculate the variables in this way:
# +
Fhgrfm2 = Fkgrfm2 = Fagrfm2 = np.sqrt(np.sum(np.abs(-grf[:,(0,1)])**2, axis=-1))
Magrf2 = -np.cross(grf[:,(2,2)]*[1,0]-kin[:,(4,5)], grf[:,(0,1)])
Mkgrf2 = -np.cross(grf[:,(2,2)]*[1,0]-kin[:,(2,3)], grf[:,(0,1)])
Mhgrf2 = -np.cross(grf[:,(2,2)]*[1,0]-kin[:,(0,1)], grf[:,(0,1)])
plotdata(time, Fhgrfm2, Fkgrfm2, Fagrfm2, Mhgrf2, Mkgrf2, Magrf2, forces, moments,
'Inverse dynamics: ground-reaction-force approach versus true values II')
# -
# ### WARNING: the calculated resultant joint force is not the actual joint reaction force!
#
# In the Newton-Euler equations based on the free body diagrams we represented the consequences of all possible muscle forces on a joint as a net muscle torque and all forces acting on a joint as a resultant joint reaction force. That is, all forces between segments were represented as a resultant force that doesn't generate torque and a moment of force that only generates torque.
# This is an important principle in mechanics of rigid bodies as we saw before.
# However, this principle creates the unrealistic notion that the sum of forces is applied directly on the joint (which has no further implication for a rigid body), but it is inaccurate for the understanding of the local effects on the joint. So, if we are trying to understand the stress on the joint or mechanisms of joint injury, the forces acting on the joint and on the rest of the segment must be considered individually.
# ## Conclusion
#
# For these data set of 'running' (remember this is simulated data), in the estimation of the forces and moments of force at the hip, knee, and ankle joints in a two-dimensional analysis, to not consider the segment acceleration and/or the mass of the segments had no effect on the ankle variables, a small effect on the knee, and a large effect on the hip.
# This is not surprising; during the support phase, ankle and knee have small movements and the mass of the segments only start to have a significant contribution for more proximal and heavy segments such as the thigh.
#
# Don't get disappointed thinking that all this work for drawing the complete FBDs and their correspondent equations was a waste of time.
# Nowadays, the state of the art and the demand for higher accuracy in biomechanics is such that such simplifications are usually not accepted.
# ## Further reading
#
# - [Gait Analysis on Wikipedia](https://en.wikipedia.org/wiki/Gait_analysis)
# - [Gait analysis: clinical facts](https://www.ncbi.nlm.nih.gov/pubmed/27618499)
# - [Gait Analysis Methods: An Overview of Wearable and Non-Wearable Systems, Highlighting Clinical Applications](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3958266/)
# - [Avaliação Biomecânica da Corrida no BMClab (in Portuguese)](http://pesquisa.ufabc.edu.br/bmclab/servicos/rba-2/)
# ## Video lectures on the Internet
#
# - Understanding & Analyzing Gait For The Clinician - series: [Introduction](https://youtu.be/x1JoaGgyKX0), [Patient Assessment](https://youtu.be/Z0QNkLshQUk), [Intro To Computer-Based 3-D Analysis](https://youtu.be/g0OcCLTQM_Y), [Basic Musculoskeletal Biomechanics](https://youtu.be/KsdrmyxOyxM), [The Gait Cycle](https://youtu.be/96nLX6sm9Yw)
# - [How to benefit from a Gait Analysis | Runners Need](https://youtu.be/rxkX7qGtIEI)
# ## Problems
#
# 1. Search the Internet for actual experimental data from a gait analysis of a runner and compare with the simulated data used in this notebook.
# 2. Collect or search for some experimental data from a movement analysis and perform inverse dynamics to determine joint forces and torques.
# 3. Imagine that you have to perform a similar analysis but of the upper limb during throwing a ball. What would have to change in the approach described in this notebook?
# ## References
#
# - <NAME>, <NAME> (2019) [Introduction to Statics and Dynamics](http://ruina.tam.cornell.edu/Book/index.html). Oxford University Press.
# - <NAME> (2009) [Biomechanics and motor control of human movement](http://books.google.com.br/books?id=_bFHL08IWfwC). 4 ed. Hoboken, EUA: Wiley.
# - <NAME>, <NAME> (1989) [Determining muscle's force and action in multi-articular movement](https://github.com/BMClab/BMC/blob/master/refs/zajac89.pdf). Exercise and Sport Sciences Reviews, 17, 187-230.
# - Zatsiorsky VM (2202) [Kinetics of human motion](http://books.google.com.br/books?id=wp3zt7oF8a0C&lpg=PA571&ots=Kjc17DAl19&dq=ZATSIORSKY%2C%20Vladimir%20M.%20Kinetics%20of%20human%20motion&hl=pt-BR&pg=PP1#v=onepage&q&f=false). Champaign, IL: Human Kinetics.
| notebooks/GaitAnalysis2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practice Assignment: Understanding Distributions Through Sampling
#
# ** *This assignment is optional, and I encourage you to share your solutions with me and your peers in the discussion forums!* **
#
#
# To complete this assignment, create a code cell that:
# * Creates a number of subplots using the `pyplot subplots` or `matplotlib gridspec` functionality.
# * Creates an animation, pulling between 100 and 1000 samples from each of the random variables (`x1`, `x2`, `x3`, `x4`) for each plot and plotting this as we did in the lecture on animation.
# * **Bonus:** Go above and beyond and "wow" your classmates (and me!) by looking into matplotlib widgets and adding a widget which allows for parameterization of the distributions behind the sampling animations.
#
#
# Tips:
# * Before you start, think about the different ways you can create this visualization to be as interesting and effective as possible.
# * Take a look at the histograms below to get an idea of what the random variables look like, as well as their positioning with respect to one another. This is just a guide, so be creative in how you lay things out!
# * Try to keep the length of your animation reasonable (roughly between 10 and 30 seconds).
# +
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import mpl_toolkits.axes_grid1.inset_locator as mpl_il
import matplotlib.animation as animation
from matplotlib.widgets import Slider
import numpy as np
import scipy.stats as stats
# %matplotlib notebook
# generate 4 random variables from the random, gamma, exponential, and uniform distributions
x1 = np.random.normal(-2.5, 1, 10000)
x2 = np.random.gamma(2, 1.5, 10000)
x3 = np.random.exponential(2, 10000)+7
x4 = np.random.uniform(14,20, 10000)
n1, n2 = 100, 1000
plt_names = ['Normal', 'Gamma', 'Exponential', 'Uniform']
plt_colors = ['blue', 'orange', 'green', 'red']
plt_colors_a = ['navy', 'darkgoldenrod', 'darkgreen', 'darkred']
ds = [x1, x2, x3, x4]
bins_me = [np.arange(np.floor(xi.min()), np.ceil(xi.max()),.5) for xi in ds]
# +
fig2, ((ax1,ax2),(ax3,ax4)) = plt.subplots(2, 2, sharey='row', figsize=(12,6))
axs = [ax1,ax2,ax3,ax4]
amp_slider_ax = fig2.add_axes([0.75, 0.95, 0.17, 0.03], axisbg='w')
samp = Slider(amp_slider_ax, 'Sample', 100, 1000, valinit=n1, valfmt='%0.0f')
bplt = []
for i, ax in enumerate(axs):
bplt.append(mpl_il.inset_axes(ax, width='10%', height='100%', loc=5))
bplt[i].axis('off')
# -
# create the function that will do the plotting, where curr is the current frame
def update_a(curr):
# check if animation is at the last frame, and if so, stop the animation a
if curr >= n2/10:
a.event_source.stop()
for i, ax in enumerate(axs):
h = ds[i][n1:n1+curr*10]
hs = h[:]
hs.sort()
ax.clear()
ax.hist(h, normed=True,\
bins=bins_me[i], alpha=0.5,\
color = plt_colors[i],\
edgecolor='none')
if curr > 2:
bplt[i].clear()
bplt[i].boxplot(h, whis='range')
bplt[i].axis('off')
hmean = np.mean(h)
hstd = np.std(h)
pdf = stats.norm.pdf(h, hmean, hstd)
ax.plot(h, pdf, color = plt_colors_a[i])
#bplt.margins(x=0.5)
ax.set_title(plt_names[i])
ax.set_ylim(0, .6)
samp.set_val(n1+curr*10)
ax.annotate('n={:}'.format(100+10*curr),\
(abs(.1*np.median(ax.get_xlim()))+ax.get_xlim()[0],\
.90*ax.get_ylim()[-1]))
aa = animation.FuncAnimation(fig2, update_a, interval=1)
fig2.subplots_adjust(top=0.88)
fig2.suptitle('Understanding Distributions\nThrough Sampling', fontsize=18)
multi = MultiCursor(fig2.canvas, axs, color='r', lw=1,
horizOn=False, vertOn=True)
plt.show()
| drafts/Week 3/UnderstandingDistributionsThroughSampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GATE Neural Network with Linear Two Neurons
import numpy as np
import random
import math
from IPython.display import display
# ## 1. Linear Two Neurons Model with Only Numerical Differentiation
# +
class Neuron1:
def __init__(self):
self.w1 = np.array([random.random(), random.random()]) # weight of one input
self.b1 = np.array([random.random()]) # bias
print("Neuron1 - Initial w1: {0}, b1: {1}".format(self.w1, self.b1))
def u1(self, x):
return np.dot(self.w1, x) + self.b1
def f(self, u1):
return max(0.0, u1)
def z1(self, x):
u1 = self.u1(x)
return self.f(u1)
class Neuron2:
def __init__(self, n1):
self.w2 = np.array([random.random()]) # weight of one input
self.b2 = np.array([random.random()]) # bias
self.n1 = n1
print("Neuron2 - Initial w2: {0}, b2: {1}".format(self.w2, self.b2))
def u2(self, x):
z1 = self.n1.z1(x)
return self.w2 * z1 + self.b2
def f(self, u2):
return max(0.0, u2)
def z2(self, x):
u2 = self.u2(x)
return self.f(u2)
def squared_error(self, x, z_target):
return 1.0 / 2.0 * math.pow(self.z2(x) - z_target, 2)
def numerical_derivative(self, params, x, z_target):
delta = 1e-4 # 0.0001
grad = np.zeros_like(params)
for idx in range(params.size):
temp_val = params[idx]
#f(x + delta) 계산
params[idx] = params[idx] + delta
fxh1 = self.squared_error(x, z_target)
#f(x - delta) 계산
params[idx] = params[idx] - delta
fxh2 = self.squared_error(x, z_target)
#f(x + delta) - f(x - delta) / 2 * delta 계산
grad[idx] = (fxh1 - fxh2) / (2 * delta)
params[idx] = temp_val
return grad
def learning(self, alpha, maxEpoch, data):
print_epoch_period = 20
for i in range(maxEpoch):
for idx in range(data.numTrainData):
x = data.training_input_value[idx]
z_target = data.training_z_target[idx]
self.n1.w1 = self.n1.w1 - alpha * self.numerical_derivative(self.n1.w1, x, z_target)
self.n1.b1 = self.n1.b1 - alpha * self.numerical_derivative(self.n1.b1, x, z_target)
self.w2 = self.w2 - alpha * self.numerical_derivative(self.w2, x, z_target)
self.b2 = self.b2 - alpha * self.numerical_derivative(self.b2, x, z_target)
if i % print_epoch_period == 0:
sum = 0.0
for idx in range(data.numTrainData):
sum = sum + self.squared_error(data.training_input_value[idx], data.training_z_target[idx])
print("Epoch{0:4d}: Error: {1:7.5f}, w1_0: {2:7.5f}, w1_1: {3:7.5f}, b1: {4:7.5f}, w2: {5:7.5f}, b2: {6:7.5f}".format(
i,
sum / data.numTrainData,
self.n1.w1[0],
self.n1.w1[1],
self.n1.b1[0],
self.w2[0],
self.b2[0])
)
# -
# ## 2. OR Gate with Linear Two Neurons
# +
class Data:
def __init__(self):
self.training_input_value = np.array([(0.0, 0.0), (1.0, 0.0), (0.0, 1.0), (1.0, 1.0)])
self.training_z_target = np.array([0.0, 1.0, 1.0, 1.0])
self.numTrainData = len(self.training_input_value)
if __name__ == '__main__':
n1 = Neuron1()
n2 = Neuron2(n1)
d = Data()
for idx in range(d.numTrainData):
x = d.training_input_value[idx]
z2 = n2.z2(x)
z_target = d.training_z_target[idx]
error = n2.squared_error(x, z_target)
print("x: {0:s}, z2: {1:s}, z_target: {2:s}, error: {3:7.5f}".format(str(x), str(z2), str(z_target), error))
n2.learning(0.01, 750, d)
for idx in range(d.numTrainData):
x = d.training_input_value[idx]
z2 = n2.z2(x)
z_target = d.training_z_target[idx]
error = n2.squared_error(x, z_target)
print("x: {0:s}, z2: {1:s}, z_target: {2:s}, error: {3:7.5f}".format(str(x), str(z2), str(z_target), error))
# -
# ## 3. AND Gate with Linear Two Neurons
# +
class Data:
def __init__(self):
self.training_input_value = np.array([(0.0, 0.0), (1.0, 0.0), (0.0, 1.0), (1.0, 1.0)])
self.training_z_target = np.array([0.0, 0.0, 0.0, 1.0])
self.numTrainData = len(self.training_input_value)
if __name__ == '__main__':
n1 = Neuron1()
n2 = Neuron2(n1)
d = Data()
for idx in range(d.numTrainData):
x = d.training_input_value[idx]
z2 = n2.z2(x)
z_target = d.training_z_target[idx]
error = n2.squared_error(x, z_target)
print("x: {0:s}, z2: {1:s}, z_target: {2:s}, error: {3:7.5f}".format(str(x), str(z2), str(z_target), error))
n2.learning(0.01, 750, d)
for idx in range(d.numTrainData):
x = d.training_input_value[idx]
z2 = n2.z2(x)
z_target = d.training_z_target[idx]
error = n2.squared_error(x, z_target)
print("x: {0:s}, z2: {1:s}, z_target: {2:s}, error: {3:7.5f}".format(str(x), str(z2), str(z_target), error))
# -
# ## 4. XOR Gate with Linear Two Neurons
# +
class Data:
def __init__(self):
self.training_input_value = np.array([(0.0, 0.0), (1.0, 0.0), (0.0, 1.0), (1.0, 1.0)])
self.training_z_target = np.array([0.0, 1.0, 1.0, 0.0])
self.numTrainData = len(self.training_input_value)
if __name__ == '__main__':
n1 = Neuron1()
n2 = Neuron2(n1)
d = Data()
for idx in range(d.numTrainData):
x = d.training_input_value[idx]
z2 = n2.z2(x)
z_target = d.training_z_target[idx]
error = n2.squared_error(x, z_target)
print("x: {0:s}, z2: {1:s}, z_target: {2:s}, error: {3:7.5f}".format(str(x), str(z2), str(z_target), error))
n2.learning(0.01, 750, d)
for idx in range(d.numTrainData):
x = d.training_input_value[idx]
z2 = n2.z2(x)
z_target = d.training_z_target[idx]
error = n2.squared_error(x, z_target)
print("x: {0:s}, z2: {1:s}, z_target: {2:s}, error: {3:7.5f}".format(str(x), str(z2), str(z_target), error))
| 1.DeepLearning/01.Multiple_Neurons/linear_two_neurons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
import radvel
import corner
import copy
from scipy import optimize
rc('savefig',dpi=120)
# Convenient function for plotting fits
def plot_results(like):
fig = gcf()
errorbar(
like.x, like.model(t)+like.residuals(),
yerr=like.yerr, fmt='o'
)
plot(ti, like.model(ti))
xlabel('Time')
ylabel('RV')
draw()
# -
# Intialize Keplerian model parameters
# ====================================
params = radvel.Parameters(1,basis='per tc secosw sesinw k')
params['k1'] = radvel.Parameter(value=1)
params['per1'] = radvel.Parameter(value=1)
params['secosw1'] = radvel.Parameter(value=0.1)
params['sesinw1'] = radvel.Parameter(value=+0.2)
params['tc1'] = radvel.Parameter(value=0.)
rv_mod = radvel.RVModel(params)
# Generate Synthetic Data
# =======================
t = np.random.random_sample(40)
t = t * 4 - 2
t = np.sort(t)
ti = np.linspace(-2,2,400)
errvel = 0.3
jitter = 0.3
syn_rv = rv_mod(t) + randn(t.size)*np.sqrt(errvel**2 + jitter**2)
errorbar(t,syn_rv,yerr=errvel,fmt='.',label='Synthetic Data')
plot(ti,rv_mod(ti),label='Underlying Model')
xlabel('Time')
ylabel('RV')
legend()
# Generate a likelihood
# =====================
# +
like_syn = radvel.likelihood.RVLikelihood(rv_mod,t,syn_rv,zeros(t.size)+errvel)
like_syn.params['gamma'] = radvel.Parameter(value=0)
like_syn.params['jit'] = radvel.Parameter(value=errvel)
truths = copy.deepcopy(like_syn.params) # Store away model parameters for later reference
like_syn.params.update(dict(k1=radvel.Parameter(value=3),
secosw1=radvel.Parameter(value=0.1),
sesinw1=radvel.Parameter(value=0.1),
tc1=radvel.Parameter(value=0.1))) # perturb the starting guess
like_syn.params['jit'].vary = False # Don't vary jitter
# -
# Perform a maximum likelihood fit
# ===============================
plot_results(like_syn) # Plot initial model
res = optimize.minimize(like_syn.neglogprob_array, like_syn.get_vary_params(), method='Nelder-Mead' )
#res = optimize.minimize(like_syn.neglogprob_array, like_syn.get_vary_params(), method='L-BFGS-B' )
print(res)
print(like_syn)
plot_results(like_syn) # plot best fit model
# Instantiate a posterior object
# ===============================
# +
post = radvel.posterior.Posterior(like_syn)
post.params['per1'] = radvel.Parameter(value=1)
post.params['k1'] = radvel.Parameter(value=1)
post.params['jit'].vary = True
post.priors += [radvel.prior.EccentricityPrior( 1 )]
post.priors += [radvel.prior.Gaussian( 'jit', errvel, 0.1)]
post.priors += [radvel.prior.Gaussian( 'per1', 1, 0.1)]
post.priors += [radvel.prior.Gaussian( 'tc1', 0, 0.1)]
post.priors += [radvel.prior.SecondaryEclipsePrior(1, 0.5, 0.01)]
print(post)
# -
# Perform maximum likelihood fit on posterior
# =========================================
print(post.vparams_order, post.list_vary_params(), post.get_vary_params())
res = optimize.minimize(post.neglogprob_array, post.get_vary_params(), method='Nelder-Mead' )
plot_results(post.likelihood)
# Use mcmc to sample the posterior distribution
# ========================
#
# 1. use `emcee` package to run MCMC
# 1. corner plot visualizes parameters and correlations
# 1. Blue lines show the synthetic model parameters
df = radvel.mcmc(post,nwalkers=20,nrun=1000)
# +
labels = [k for k in post.params.keys() if post.params[k].vary]
fig = corner.corner(
df[labels],
labels=labels,
truths=[truths[k].value for k in labels ],
quantiles=[0.15,0.85],
plot_datapoints=False,
smooth=True,
bins=20
)
# -
hist(df.lnprobability)
| docs/tutorials/SyntheticData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Using edited code from [<NAME>](https://medium.com/@michael.wy.ong/web-scrape-geospatial-data-analyse-singapores-property-price-part-i-276caba320b) with some tweakings
import pandas as pd
import numpy as np
import requests
import json
# +
## Function for getting postal code, geo coordinates of addresses
def find_postal(lst, filename):
'''With the block number and street name, get the full address of the hdb flat,
including the postal code, geogaphical coordinates (lat/long)'''
for index,add in enumerate(lst):
# Do not need to change the URL
url= "https://developers.onemap.sg/commonapi/search?returnGeom=Y&getAddrDetails=Y&pageNum=1&searchVal="+ add
print(index,url)
# Retrieve information from website
response = requests.get(url)
try:
data = json.loads(response.text)
except ValueError:
print('JSONDecodeError')
pass
temp_df = pd.DataFrame.from_dict(data["results"])
# The "add" is the address that was used to search in the website
temp_df["address"] = add
# Create the file with the first row that is read in
if index == 0:
file = temp_df
else:
file = file.append(temp_df)
file.to_csv(filename + '.csv')
# +
## Function for getting closest distance of each location from a list of amenities location
from geopy.distance import geodesic
def find_nearest(house, amenity, radius=2):
"""
this function finds the nearest locations from the 2nd table from the 1st address
Both are dataframes with a specific format:
1st column: any string column ie addresses taken from the "find_postal_address.py"
2nd column: latitude (float)
3rd column: longitude (float)
Column name doesn't matter.
It also finds the number of amenities within the given radius (default=2)
"""
results = {}
# first column must be address
for index,flat in enumerate(house.iloc[:,0]):
# 2nd column must be latitude, 3rd column must be longitude
flat_loc = (house.iloc[index,1],house.iloc[index,2])
flat_amenity = ['','',100,0]
for ind, eachloc in enumerate(amenity.iloc[:,0]):
amenity_loc = (amenity.iloc[ind,1],amenity.iloc[ind,2])
distance = geodesic(flat_loc,amenity_loc)
distance = float(str(distance)[:-3]) # convert to float
if distance <= radius: # compute number of amenities in 2km radius
flat_amenity[3] += 1
if distance < flat_amenity[2]: # find nearest amenity
flat_amenity[0] = flat
flat_amenity[1] = eachloc
flat_amenity[2] = distance
results[flat] = flat_amenity
return results
# -
def dist_from_location(house, location):
"""
this function finds the distance of a location from the 1st address
First is a dataframe with a specific format:
1st column: any string column ie addresses taken from the "find_postal_address.py"
2nd column: latitude (float)
3rd column: longitude (float)
Column name doesn't matter.
Second is tuple with latitude and longitude of location
"""
results = {}
# first column must be address
for index,flat in enumerate(house.iloc[:,0]):
# 2nd column must be latitude, 3rd column must be longitude
flat_loc = (house.iloc[index,1],house.iloc[index,2])
flat_amenity = ['',100]
distance = geodesic(flat_loc,location)
distance = float(str(distance)[:-3]) # convert to float
flat_amenity[0] = flat
flat_amenity[1] = distance
results[flat] = flat_amenity
return results
# +
price1999 = pd.read_csv('Data/resale-flat-prices-based-on-approval-date-1990-1999.csv')
price2012 = pd.read_csv('Data/resale-flat-prices-based-on-approval-date-2000-feb-2012.csv')
price2014 = pd.read_csv('Data/resale-flat-prices-based-on-registration-date-from-mar-2012-to-dec-2014.csv')
price2016 = pd.read_csv('Data/resale-flat-prices-based-on-registration-date-from-jan-2015-to-dec-2016.csv')
price2017 = pd.read_csv('Data/resale-flat-prices-based-on-registration-date-from-jan-2017-onwards.csv')
prices = pd.concat([price1999, price2012, price2014], sort=False)
prices = pd.concat([prices, price2016, price2017], axis=0, ignore_index=True, sort=False)
# +
prices = prices[['block', 'street_name']]
prices['address'] = prices['block'] + ' ' + prices['street_name']
all_address = list(prices['address'])
unique_address = list(set(all_address))
print('Unique addresses:', len(unique_address))
# -
unique_address[:10]
# + tags=["outputPrepend"]
find_postal(unique_address, 'Data/flat_coordinates')
# -
flat_coord = pd.read_csv('Data/flat_coordinates.csv')
flat_coord = flat_coord[['address','LATITUDE','LONGITUDE']]
flat_coord.head()
# ## Supermarkets
supermarket = pd.read_csv('Data/list-of-supermarket-licences.csv')
supermarket.head()
# +
supermerket_address = list(supermarket['postal_code'])
unique_supermarket_address = list(set(supermerket_address))
print('Unique addresses:', len(unique_supermarket_address))
# + tags=["outputPrepend"]
find_postal(unique_supermarket_address, 'Data/supermarket_coordinates')
# -
supermarket_coord = pd.read_csv('Data/supermarket_coordinates.csv')
supermarket_coord.drop_duplicates(subset=['address'], inplace=True)
supermarket_coord = supermarket_coord[['SEARCHVAL','LATITUDE','LONGITUDE']]
supermarket_coord.head()
nearest_supermarket = find_nearest(flat_coord, supermarket_coord)
flat_supermarket = pd.DataFrame.from_dict(nearest_supermarket).T
flat_supermarket = flat_supermarket.rename(columns={0: 'flat', 1: 'supermarket', 2: 'supermarket_dist', 3: 'num_supermarket_2km'}).reset_index().drop(['index','supermarket'], axis=1)
flat_supermarket.head()
# ## Schools
school = pd.read_csv('Data/school_names.csv', encoding='cp1252')
school.head(10)
# +
school_name = list(school['school'])
unique_school_name = list(set(school_name))
print('Unique addresses:', len(unique_school_name))
# + tags=["outputPrepend"]
find_postal(unique_school_name, 'Data/school_coordinates')
# -
school_coord = pd.read_csv('Data/school_coordinates.csv')
school_coord = school_coord[['address','LATITUDE','LONGITUDE']]
school_coord.head()
nearest_school = find_nearest(flat_coord, school_coord)
flat_school = pd.DataFrame.from_dict(nearest_school).T
flat_school = flat_school.rename(columns={0: 'flat', 1: 'school', 2: 'school_dist', 3: 'num_school_2km'}).reset_index().drop('index', axis=1)
flat_school.head()
# ## Hawker and Markets
hawker = pd.read_csv('Data/list-of-government-markets-hawker-centres.csv')
hawker.head(10)
# +
hawker_name = list(hawker['name_of_centre'])
unique_hawker_name = list(set(hawker_name))
print('Unique addresses:', len(unique_hawker_name))
# -
find_postal(unique_hawker_name, 'Data/hawker_coordinates')
hawker_coord = pd.read_csv('Data/hawker_coordinates.csv')
hawker_coord = hawker_coord[['address','LATITUDE','LONGITUDE']]
hawker_coord.head()
nearest_hawker = find_nearest(flat_coord, hawker_coord)
flat_hawker = pd.DataFrame.from_dict(nearest_hawker).T
flat_hawker = flat_hawker.rename(columns={0: 'flat', 1: 'hawker', 2: 'hawker_dist', 3: 'num_hawker_2km'}).reset_index().drop('index', axis=1)
flat_hawker.head()
# ## Shopping Malls
shop = pd.read_csv('Data/shoppingmalls.csv', encoding='cp1252')
shop.head()
# +
shop_name = list(shop['name'])
unique_shop_name = list(set(shop_name))
print('Unique addresses:', len(unique_shop_name))
# -
find_postal(unique_shop_name, 'Data/shoppingmall_coordinates')
shop_coord = pd.read_csv('Data/shoppingmall_coordinates.csv')
shop_coord.drop_duplicates(subset=['address'], inplace=True)
shop_coord = shop_coord[['address','LATITUDE','LONGITUDE']]
shop_coord.head()
nearest_mall = find_nearest(flat_coord, shop_coord)
flat_mall = pd.DataFrame.from_dict(nearest_mall).T
flat_mall = flat_mall.rename(columns={0: 'flat', 1: 'mall', 2: 'mall_dist', 3: 'num_mall_2km'}).reset_index().drop('index', axis=1)
flat_mall.head()
# ## Parks
park_coord = pd.read_csv('Data/parks-kml.csv')
park_coord.reset_index(inplace=True)
park_coord = park_coord[['index','Y','X']]
park_coord.head()
nearest_park = find_nearest(flat_coord, park_coord)
flat_park = pd.DataFrame.from_dict(nearest_park).T
flat_park = flat_park.rename(columns={0: 'flat', 1: 'park', 2: 'park_dist', 3: 'num_park_2km'}).reset_index().drop(['index','park'], axis=1)
flat_park.head()
# ## MRT
mrt_coord = pd.read_csv('Data/MRT_coordinates.csv')
mrt_coord = mrt_coord[['STN_NAME','Latitude','Longitude']]
mrt_coord.head()
nearest_mrt = find_nearest(flat_coord, mrt_coord)
flat_mrt = pd.DataFrame.from_dict(nearest_mrt).T
flat_mrt = flat_mrt.rename(columns={0: 'flat', 1: 'mrt', 2: 'mrt_dist', 3: 'num_mrt_2km'}).reset_index().drop('index', axis=1)
flat_mrt.head()
flat_coord = pd.read_csv('Data/flat_coordinates.csv')
flat_coord = flat_coord[['address','LATITUDE','LONGITUDE']]
flat_coord.head()
# ## Merge All
# +
## Merge all
flat_amenities = flat_school.merge(flat_hawker, on='flat', how='outer')
flat_amenities = flat_amenities.merge(flat_park, on='flat', how='outer')
flat_amenities = flat_amenities.merge(flat_mall, on='flat', how='outer')
flat_amenities = flat_amenities.merge(flat_mrt, on='flat', how='outer')
flat_amenities = flat_amenities.merge(flat_supermarket, on='flat', how='outer')
flat_amenities.head()
# -
flat_amenities.to_csv('Data/flat_amenities.csv', index=False)
# ## Get dist from dhoby ghaut
# +
flat_coord = pd.read_csv('Data/flat_coordinates.csv')
flat_coord = flat_coord[['address','LATITUDE','LONGITUDE']]
dist_dhoby = dist_from_location(flat_coord, (1.299308, 103.845285))
dist_dhoby = pd.DataFrame.from_dict(dist_dhoby).T
dist_dhoby = dist_dhoby.rename(columns={0: 'flat', 1: 'dist_dhoby'}).reset_index().drop(['index'], axis=1)
dist_dhoby.head()
# -
flat_amenities = pd.read_csv('Data/flat_amenities.csv')
flat_amenities = flat_amenities.merge(dist_dhoby, on='flat', how='outer')
flat_amenities.head()
flat_amenities.to_csv('Data/flat_amenities.csv', index=False)
| get_coordinates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Avoiding Overfitting Through Regularization
#
# Deep neural networks typically have tens of thousands of parameters, sometimes
# even millions. With so many parameters, the network has an incredible amount of
# freedom and can fit a huge variety of complex datasets. But this great flexibility also means that it is prone to overfitting the training set.
#
# With millions of parameters you can fit the whole zoo. In this section we will present some of the most popular regularization techniques for neural networks, and how to implement them with TensorFlow:
# 1. early stopping;
# 2. ℓ1 and ℓ2 regularization;
# 3. dropout;
# 4. max-norm regularization;
# 5. data augmentation.
# +
# Set up some basisc fucntion
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Where to save the figures
PROJECT_ROOT_DIR = "./image/"
CHAPTER_ID = "deep"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# -
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("./tmp/data/")
# ## Early Stopping
# To avoid overfitting the training set, a great solution is early stopping (introduced in **TensorFlow_05_Neural_Networks**): just interrupt training when its performance on the validation set starts dropping.
#
# One way to implement this with TensorFlow is to evaluate the model on a validation set at regular intervals (e.g., every 50 steps), and save a “winner” snapshot if it outperforms previous “winner” snapshots. Count the number of steps since the last “winner” snapshot was saved, and interrupt training when this number reaches some limit (e.g., 2,000 steps). Then restore the last “winner” snapshot.
#
# Although early stopping works very well in practice, you can usually get much higher performance out of your network by combining it with other regularization techniques.
# ## $\ell_1$ and $\ell_2$ regularization
#
# Just like you did in TensorFlow_05, you can use ℓ1 and ℓ2 regularization to constrain a neural network’s connection weights (but typically not its biases).
#
# One way to do this using TensorFlow is to simply add the appropriate regularization terms to your cost function. For example, assuming you have just one hidden layer with weights $weights1$ and one output layer with weights $weights2$, then you can apply $\ell_1$ regularization like this:
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
logits = tf.layers.dense(hidden1, n_outputs, name="outputs")
# -
# Next, we get a handle on the layer weights, and we compute the total loss, which is equal to the sum of the usual cross entropy loss and the $\ell_1$ loss (i.e., the absolute values of the weights):
# +
W1 = tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
W2 = tf.get_default_graph().get_tensor_by_name("outputs/kernel:0")
scale = 0.001 # l1 regularization hyperparameter
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y,
logits=logits)
base_loss = tf.reduce_mean(xentropy, name="avg_xentropy")
reg_losses = tf.reduce_sum(tf.abs(W1)) + tf.reduce_sum(tf.abs(W2))
loss = tf.add(base_loss, scale * reg_losses, name="loss")
# +
#The rest is just as usual:
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: mnist.test.images,
y: mnist.test.labels})
print(epoch, "Test accuracy:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# Alternatively, we can pass a regularization function to the `tf.layers.dense()` function, which will use it to create operations that will compute the regularization loss, and it adds these operations to the collection of regularization losses. The beginning is the same as above:
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
# -
# Next, we will use Python's `partial()` function to avoid repeating the same arguments over and over again. Note that we set the `kernel_regularizer` argument:
scale = 0.001
# +
from functools import partial
my_dense_layer = partial(
tf.layers.dense, activation=tf.nn.relu,
kernel_regularizer=tf.contrib.layers.l1_regularizer(scale))
with tf.name_scope("dnn"):
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
hidden2 = my_dense_layer(hidden1, n_hidden2, name="hidden2")
logits = my_dense_layer(hidden2, n_outputs, activation=None,
name="outputs")
# -
# Next we must add the regularization losses to the base loss:
with tf.name_scope("loss"): # not shown in the book
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits( # not shown
labels=y, logits=logits) # not shown
base_loss = tf.reduce_mean(xentropy, name="avg_xentropy") # not shown
reg_losses = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
loss = tf.add_n([base_loss] + reg_losses, name="loss")
# And the rest is the same as usual:
# +
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: mnist.test.images,
y: mnist.test.labels})
print(epoch, "Test accuracy:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# # Dropout
# The most popular regularization technique for deep neural networks is arguably
# dropout. It was proposed20 by <NAME> in 2012 and further detailed in a paper by <NAME> et al., and it has proven to be highly successful: even the state-ofthe-art neural networks got a 1–2% accuracy boost simply by adding dropout.
#
# It is a fairly simple algorithm: at every training step, every neuron (including the input neurons but excluding the output neurons) has a probability p of being temporarily “dropped out,” meaning it will be entirely ignored during this training step, but it may be active during the next step.
#
# The hyperparameter p is called the dropout rate, and it is typically set to 50%. After training, neurons don’t get dropped anymore. And that’s all (except for a technical detail we will discuss momentarily).
# +
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
# +
training = tf.placeholder_with_default(False, shape=(), name='training')
dropout_rate = 0.5 # == 1 - keep_prob
X_drop = tf.layers.dropout(X, dropout_rate, training=training)
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X_drop, n_hidden1, activation=tf.nn.relu,
name="hidden1")
hidden1_drop = tf.layers.dropout(hidden1, dropout_rate, training=training)
hidden2 = tf.layers.dense(hidden1_drop, n_hidden2, activation=tf.nn.relu,
name="hidden2")
hidden2_drop = tf.layers.dropout(hidden2, dropout_rate, training=training)
logits = tf.layers.dense(hidden2_drop, n_outputs, name="outputs")
# +
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum=0.9)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 20
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={training: True, X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Test accuracy:", acc_test)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# ## Max-Norm Regularization
# Another regularization technique that is quite popular for neural networks is called max-norm regularization: for each neuron, it constrains the weights w of the incoming connections such that $\| w \| 2 \leq r$, where r is the max-norm hyperparameter and $\| \cdot \|$ is the $\ell_2$ norm.
#
# We typically implement this constraint by computing ∥w∥2 after each training step and clipping w if needed
#
# $$ w \leftarrow w \frac{r}{\|w\|_2}$$
#
# Reducing r increases the amount of regularization and helps reduce overfitting. Maxnorm regularization can also help alleviate the vanishing/exploding gradients problems (if you are not using Batch Normalization).
#
#
# TensorFlow does not provide an off-the-shelf max-norm regularizer, but it is not too hard to implement. The following code creates a node clip_weights that will clip the weights variable along the second axis so that each row vector has a maximum norm of 1.0:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
learning_rate = 0.01
momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
# -
# Next, let's get a handle on the first hidden layer's weight and create an operation that will compute the clipped weights using the `clip_by_norm()` function. Then we create an assignment operation to assign the clipped weights to the weights variable:
threshold = 1.0
weights = tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
clipped_weights = tf.clip_by_norm(weights, clip_norm=threshold, axes=1)
clip_weights = tf.assign(weights, clipped_weights)
# We can do this as well for the second hidden layer:
weights2 = tf.get_default_graph().get_tensor_by_name("hidden2/kernel:0")
clipped_weights2 = tf.clip_by_norm(weights2, clip_norm=threshold, axes=1)
clip_weights2 = tf.assign(weights2, clipped_weights2)
# Let's add an initializer and a saver:
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 50
with tf.Session() as sess: # not shown in the book
init.run() # not shown
for epoch in range(n_epochs): # not shown
for iteration in range(mnist.train.num_examples // batch_size): # not shown
X_batch, y_batch = mnist.train.next_batch(batch_size) # not shown
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
clip_weights.eval()
clip_weights2.eval() # not shown
acc_test = accuracy.eval(feed_dict={X: mnist.test.images, # not shown
y: mnist.test.labels}) # not shown
print(epoch, "Test accuracy:", acc_test) # not shown
save_path = saver.save(sess, "./my_model_final.ckpt") # not shown
# The implementation above is straightforward and it works fine, but it is a bit messy. A better approach is to define a `max_norm_regularizer()` function:
def max_norm_regularizer(threshold, axes=1, name="max_norm",
collection="max_norm"):
def max_norm(weights):
clipped = tf.clip_by_norm(weights, clip_norm=threshold, axes=axes)
clip_weights = tf.assign(weights, clipped, name=name)
tf.add_to_collection(collection, clip_weights)
return None # there is no regularization loss term
return max_norm
# Then you can call this function to get a max norm regularizer (with the threshold you want). When you create a hidden layer, you can pass this regularizer to the `kernel_regularizer` argument:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
learning_rate = 0.01
momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
# +
max_norm_reg = max_norm_regularizer(threshold=1.0)
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
kernel_regularizer=max_norm_reg, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
kernel_regularizer=max_norm_reg, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
# +
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# -
# Training is as usual, except you must run the weights clipping operations after each training operation:
n_epochs = 20
batch_size = 50
# +
clip_all_weights = tf.get_collection("max_norm")
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
sess.run(clip_all_weights)
acc_test = accuracy.eval(feed_dict={X: mnist.test.images, # not shown in the book
y: mnist.test.labels}) # not shown
print(epoch, "Test accuracy:", acc_test) # not shown
save_path = saver.save(sess, "./my_model_final.ckpt") # not shown
# -
# ## Data Augmentation
#
# One last regularization technique, data augmentation, consists of generating new
# training instances from existing ones, artificially boosting the size of the training set.
#
# This will reduce overfitting, making this a regularization technique. The trick is to generate realistic training instances; ideally, a human should not be able to tell which instances were generated and which ones were not. Moreover, simply adding white noise will not help; the modifications you apply should be learnable (white noise is not).
# For example, if your model is meant to classify pictures of mushrooms, you can
# slightly shift, rotate, and resize every picture in the training set by various amounts and add the resulting pictures to the training set.
#
# This forces the model to be more tolerant to the position, orientation, and size of the mushrooms in the picture. If you want the model to be more tolerant to lighting conditions, you can similarly generate many images with various contrasts.
#
# Assuming the mushrooms are symmetrical, you can also flip the pictures horizontally. By combining these transformations you can greatly increase the size of your training set.
#
# It is often preferable to generate training instances on the fly during training rather than wasting storage space and network bandwidth. TensorFlow offers several image manipulation operations such as transposing (shifting), rotating, resizing, flipping, and cropping, as well as adjusting the brightness, contrast, saturation, and hue.
| 02.TensorFlow/TF1.x/TensorFlow_09_Avoiding_Overfitting_by_Regularizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## __INTRODUCTION__
#
# ### __ARTIFICIAL NEURAL NETWORKS__
# * ML models that have a graph structure,inspired by the brain structure, with many interconnected units called artificial naurons https://www.youtube.com/watch?v=3JQ3hYko51Y
# * ANN have the ability to learn from raw data imputs, but it also makes them slower
#
#
#
# ### __TENSORFLOW__
# * CREATED AND MAINTAINED BY GOOGLE,
# * Different APIs (Application Programming Interface)
# * (a) low level graph API
# * (b) High level Keras API
# * TF on GPUs
# * it requires different version of a library,
# * available in conda,
# > conda install tensorflow-gpu==1.12.0 #or newer version
# * requires: compatible NVIDIA graphic card
# * list is available cards is here: https://developer.nvidia.com/cuda-gpus
#
# ### __COMPUTATION GRAPHS__
# * basic concept used in TF to specify how different elements interact with eaxch other
# - example:
# + we wish to implement liear regression
# y = ax + b, where a, abd b are the sloe and intercept parameters,
# x asre imput data,
# y are predictions, that weill be used to compare with the output
# y (y without a hat) usunig huber loss
# \
# a loss - each node of the grath is a step in our computation
# \ / - in TF data values are called TENSORS (3D matrices)
# * -> + -> y^ - in TF we first define a graph, and then we feed the data flows
# / | through the graph
# x b
#
#
#
#
# ### __LOSS FUNCTIONS__
# * TF implements only basic set ot loss funcitons
# * more can be added by hand, using numpy-like functions eg ,ean, sqrt etc..., chekck for names becuase these are a bit different then in numpy
# * https://www.tensorflow.org/api_docs/python/tf/keras/losses
#
# ### __TF OPTIMAZERS__
# * https://www.tensorflow.org/api_docs/python/tf/keras/optimizers#top_of_page
#
import matplotlib.pyplot as plt # for making plots,
import matplotlib as mpl # to get some basif functions, heping with plot mnaking
import numpy as np # support for multi-dimensional arrays and matrices
import pandas as pd # library for data manipulation and analysis
import random # functions that use and generate random numbers
import glob # lists names in folders that match Unix shell patterns
import re # module to use regular expressions,
import os # allow changing, and navigating files and folders,
import seaborn as sns # advance plots, for statistics,
import scipy.stats as stats # library for statistics and technical programming,
# %matplotlib inline
# %config InlineBackend.figure_format ='retina' # For retina screens (mac)
import tensorflow as tf
print(tf.__version__)
# ## Example 1. implement linear regression with TF
# +
from sklearn.datasets import make_regression
# create the data
X, y = make_regression(
n_samples=1000,
n_features=2,
n_informative=2
)
# chek the data
print("data: ", X.shape)
print("labels: ", y.shape)
# plot the data
''' i tested different number of features,
thus this funciton handles them all
sorry for small complications,
'''
if X.shape[1]==1:
plt.scatter(X,y, s=0.1, c="black")
else:
fig, axs = plt.subplots(nrows=1, ncols=2)
i=-1
for ax in axs.flat:
i+=1
if i<X.shape[1]:
ax.scatter(X[:,i],y, s=0.1, c="black")
ax.set_title(f'y ~ feature {i}')
else: pass
plt.show()
# -
# ### Part 1. DEFINE THE MODEL FOR TF
#
# #### Step 1. Define Variables
# - def. dtype is tf.int32
# - variables are provided to session in list with operations
# - they can be modified by the operations
# - variables are returned at each session, even if not chnaged
# - they need an initial value
a0 = tf.Variable(initial_value=0, dtype=tf.float32) # Feature 0 coeff.
a1 = tf.Variable(initial_value=0, dtype=tf.float32) # Feature 1 coeff.
b = tf.Variable(initial_value=0, dtype=tf.float32) # Intercept
# #### Step 2. Define Placeholders
# A TensorFlow placeholder is simply a variable that we will assign data to at a later date. It allows us to create our operations and build our computation graph, without needing the data
# - Must be provided externally to the session,
# - IT WILL NOT BE CHNAGED by the operations,
# - NOT RETURNED,
# - given to tf session as dictionary:
# * {key:value}
# * where key is as in below,
# * value is name of df, array, constant, list etc,
# +
# Step 2. Define Placeholders
"""https://indianaiproduction.com/create-tensorflow-placeholder/"""
# placeholders are not executable immediately so we need to disable eager exicution in TF 2 not in 1
# tf.compat.v1.disable_eager_execution()
x = tf.compat.v1.placeholder(dtype=tf.float32) # Input
y = tf.compat.v1.placeholder(dtype=tf.float32) # Target
lr = tf.compat.v1.placeholder(dtype=tf.float32) # Learning rate for optimizer
# -
# #### Step 3. Define Operations, in sub-steps a-d
# * Four items are required:
# * (a) Define how do we make predicitons, eg: y_hat = 2a + 1
# * (b) Define Loss Function eg: MSE
# * (c) Define How you will optimaze the parameters in a) eg. with SGD
# * (d) Define training operation on a loss function
# * eg: minimaze, maximaze etc..
# * important:
# * a, b, and d must be given to session,
# * d, is defined on d, so c, doenst have to given, or even change,
#
# +
# (a) Define how do we make predicitons
y_hat = a0*x + a1*x + b
# (b) Define Loss Function
loss = tf.compat.v1.losses.huber_loss(y, y_hat, delta=1.0)
# (c) Create/select the optimizer
gd = tf.compat.v1.train.GradientDescentOptimizer(learning_rate=lr)
# (d) Define training operation on a loss function
train_op = gd.minimize(loss)
# important comments:
#. - operations such as 1, and 2 will retunr results, in session
#. - operation 3 will affect a, and b variables, ie, no returned values
#. - because variables (a, b) are given in fetch list, these will be also
#. returned at each session iteration
#. - some operations, such as tf.train.GradientDescentOptimizer,
# may require new placeholders, eg ls. that we could change$¨
# -
# ### __Part 2. Run TF session__
#
# #### Step 1. Prepare for tf session,
# * python lists, or arrays to store loss values, coefficinets etc..
# * nr, of iterations,
# +
# Create lists to store a/b, and loss values from each iteration
loss_values = []
a0_values = []
a1_values = []
b_values = []
# Number of iterations
n = 100
# -
# # Step 2. Run Session,
#
# Session: perfomes n iterations with training variables,
# using training operation, and loss function createt in Step 2.
#
# Returns: - 4 objects,
# - "_" - is for training op, that are None,
# - loss_val, a_val, b_val - for variables retunrned
# by each of the other operations/varinables
#
# Inputs:
# - [train_op, loss, a, b]
# list with operations, & varinables,
# y_hat, not one of them, because loss has its derivative
# - Placeholders in distionary,
# +
# Initialization operation,
initialization_op = tf.compat.v1.global_variables_initializer()
# run session,
with tf.compat.v1.Session() as sess:
# Initialize the graph - always with new session !
sess.run(initialization_op)
# Run n(times)
for _ in range(n):
# Run training operations and collect a/b and loss values
_, loss_val, a0_val, a1_val, b_val = sess.run(
[train_op, loss, a0, a1, b],
feed_dict={
x: X,
y: y,
lr: [1]
}
) # NOTE: loss, a and b do not have to be provided
# Save values at each iteration,
loss_values.append(loss_val)
a0_values.append(a0_val)
a1_values.append(a1_val)
b_values.append(b_val)
| MachineLearning_code_examples/DeepLearning/.ipynb_checkpoints/TF_basics-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# +
# default_exp exec.dataset_statistics
# +
# export
import argparse
import os
import sys
from typing import List, Any, Dict
import json
from g2p_en import G2p
import librosa
import math
import matplotlib.pyplot as plt
from mdutils.mdutils import MdUtils
import numpy as np
from pydub import AudioSegment, silence
import seaborn as sns
from tqdm import tqdm
from uberduck_ml_dev.data.statistics import (
AbsoluteMetrics,
count_frequency,
create_wordcloud,
get_sample_format,
pace_character,
pace_phoneme,
word_frequencies,
)
from uberduck_ml_dev.text.util import clean_text, text_to_sequence
from uberduck_ml_dev.utils.audio import compute_yin
# +
# export
def get_summary_statistics(arr):
if len(arr) == 0:
return {}
arr_np = np.array(arr)
return {
"p10": float(np.percentile(arr_np, 10)),
"p25": float(np.percentile(arr_np, 25)),
"p50": float(np.percentile(arr_np, 50)),
"p75": float(np.percentile(arr_np, 75)),
"p90": float(np.percentile(arr_np, 90)),
"max": float(np.max(arr_np)),
"min": float(np.min(arr_np)),
}
def calculate_statistics(
dataset_path, input_file, output_folder, delimiter, metrics=True, wordcloud=True
):
n_clips = 0
sample_rates = {}
channels = {"mono": 0, "stereo": 0}
extensions = {}
sample_formats = {}
total_lengths = []
leading_silence_lengths = []
trailing_silence_lengths = []
paces_characters = [] # number of characters / seconds in audio clip
paces_phonemes = [] # number of phonemes / seconds in audio clip
lookup_results = {
"RNN": [],
"CMU": [],
"non-alphanumeric": [],
"homograph": [],
} # keep track of how arpabet sequences were generated
mosnet_scores = []
srmr_scores = []
word_freqs = []
all_words = []
all_pitches = np.array([])
all_loudness = []
g2p = G2p()
files_with_error = []
if metrics:
abs_metrics = AbsoluteMetrics()
with open(os.path.join(dataset_path, input_file)) as transcripts:
for line in tqdm(transcripts.readlines()):
try:
line = line.strip() # remove trailing newline character
file, transcription = line.lower().split(delimiter)
transcription_cleaned = clean_text(transcription, ["english_cleaners"])
_, file_extension = os.path.splitext(file)
path_to_file = os.path.join(dataset_path, file)
file_pydub = AudioSegment.from_wav(path_to_file)
data_np, _ = librosa.load(path_to_file)
# Format Metadata
sr = file_pydub.frame_rate
if sr in sample_rates.keys():
sample_rates[sr] += 1
else:
sample_rates[sr] = 1
if file_pydub.channels == 1:
channels["mono"] += 1
else:
channels["stereo"] += 1
if file_extension in extensions.keys():
extensions[file_extension] += 1
else:
extensions[file_extension] = 1
fmt = get_sample_format(path_to_file)
if fmt in sample_formats.keys():
sample_formats[fmt] += 1
else:
sample_formats[fmt] = 1
# lengths
total_lengths.append(file_pydub.duration_seconds)
leading_silence_lengths.append(
silence.detect_leading_silence(file_pydub)
)
trailing_silence_lengths.append(
silence.detect_leading_silence(file_pydub.reverse())
)
# Paces
paces_phonemes.append(
pace_phoneme(text=transcription_cleaned, audio=path_to_file)
)
paces_characters.append(
pace_character(text=transcription_cleaned, audio=path_to_file)
)
# Pitch
pitches, harmonic_rates, argmins, times = compute_yin(data_np, sr=sr)
pitches = np.array(pitches)
pitches = pitches[pitches > 10]
all_pitches = np.append(all_pitches, pitches)
# Loudness
all_loudness.append(file_pydub.dBFS)
# Quality
if metrics:
scores = abs_metrics(path_to_file)
mosnet_scores.append(scores["mosnet"][0][0])
srmr_scores.append(scores["srmr"])
# Transcription
word_freqs.extend(word_frequencies(transcription_cleaned))
transcription_lookups = g2p.check_lookup(transcription_cleaned)
for k in transcription_lookups:
lookup_results[k].extend(transcription_lookups[k])
all_words.append(transcription_cleaned)
n_clips += 1
except Exception as e:
print(e)
files_with_error.append(file)
if n_clips == 0:
return None
if wordcloud:
create_wordcloud(
" ".join(all_words),
os.path.join(dataset_path, output_folder, "wordcloud.png"),
)
# Length graph
plt.clf()
sns.histplot(total_lengths)
plt.title("Audio length distribution")
plt.xlabel("Audio length (s)")
plt.ylabel("Count")
plt.savefig(os.path.join(dataset_path, output_folder, "lengths.png"))
# Word Frequencies graph
plt.clf()
sns.histplot(word_freqs, bins=10)
plt.title("Word frequency distribution [0-1]")
plt.xlabel("Word frequency")
plt.ylabel("Count")
plt.savefig(os.path.join(dataset_path, output_folder, "word_frequencies.png"))
plt.close()
# Pitches graph
plt.clf()
plt.figure(figsize=(12, 4))
plt.subplot(1, 2, 1)
sns.histplot(all_pitches)
plt.title("Pitch distribution")
plt.xlabel("Fundamental Frequency (Hz)")
plt.ylabel("Count")
plt.subplot(1, 2, 2)
sns.histplot(all_loudness)
plt.title("Loudness distribution")
plt.xlabel("Loudness (dBFS)")
plt.ylabel("Count")
plt.savefig(os.path.join(dataset_path, output_folder, "pitch_loudness.png"))
plt.close()
# Silences graph
plt.clf()
plt.figure(figsize=(12, 4))
plt.subplot(1, 2, 1)
sns.histplot(leading_silence_lengths)
plt.title("Leading silence distribution")
plt.xlabel("Leading silence (ms)")
plt.ylabel("Count")
plt.subplot(1, 2, 2)
sns.histplot(trailing_silence_lengths)
plt.title("Traling silence distribution")
plt.xlabel("Trailing silence (ms)")
plt.ylabel("Count")
plt.savefig(os.path.join(dataset_path, output_folder, "silences.png"))
plt.close()
# Metrics graph
plt.clf()
plt.figure(figsize=(12, 4))
plt.subplot(1, 2, 1)
sns.histplot(mosnet_scores)
plt.title("Mosnet score distribution")
plt.xlabel("Mosnet score")
plt.ylabel("Count")
plt.subplot(1, 2, 2)
sns.histplot(srmr_scores)
plt.title("SRMR score distribution")
plt.xlabel("SRMR score")
plt.ylabel("Count")
plt.savefig(os.path.join(dataset_path, output_folder, "metrics.png"))
plt.close()
# Paces graph
plt.clf()
plt.figure(figsize=(12, 4))
plt.subplot(1, 2, 1)
sns.histplot(paces_characters)
plt.title("Pace (chars/s)")
plt.xlabel("Characters / second")
plt.ylabel("Count")
plt.subplot(1, 2, 2)
sns.histplot(paces_phonemes)
plt.title("Pace (phonemes/s)")
plt.xlabel("Phonemes / second")
plt.ylabel("Count")
plt.savefig(os.path.join(dataset_path, output_folder, "paces.png"))
plt.close()
return {
"n_clips": n_clips,
"total_lengths_summary": get_summary_statistics(total_lengths),
"paces_phonemes_summary": get_summary_statistics(paces_phonemes),
"paces_characters_summary": get_summary_statistics(paces_characters),
"mosnet_scores_summary": get_summary_statistics(mosnet_scores),
"srmr_scores_summary": get_summary_statistics(srmr_scores),
"pitch_summary": get_summary_statistics(all_pitches),
"loudness_summary": get_summary_statistics(all_loudness),
"total_lengths": total_lengths,
"paces_phonemes": paces_phonemes,
"paces_characters": paces_characters,
"mosnet_scores": mosnet_scores,
"srmr_scores": srmr_scores,
"sample_rates": sample_rates,
"channels": channels,
"extensions": extensions,
"sample_formats": sample_formats,
"lookup_results": lookup_results,
"files_with_error": files_with_error,
}
def generate_markdown(output_file, dataset_path, output_folder, data):
mdFile = MdUtils(
file_name=os.path.join(dataset_path, output_file), title=f"Dataset statistics"
)
total_length_mins = sum(data["total_lengths"]) / 60.0
mdFile.new_header(level=1, title="Overview")
mdFile.new_line(f"**Number of clips:** {data['n_clips']}")
mdFile.new_line(
f"**Total data:** {math.floor(total_length_mins)} minutes {math.ceil(total_length_mins % 1 * 60.0)} seconds"
)
mdFile.new_line(
f"**Mean clip length:** {sum(data['total_lengths'])/data['n_clips']:.2f} seconds"
)
mdFile.new_line(
f"**Mean pace:** {sum(data['paces_phonemes'])/len(data['paces_phonemes']):.2f} \
phonemes/sec {sum(data['paces_characters'])/len(data['paces_characters']):.2f} chars/sec"
)
if len(data["mosnet_scores"]) > 0:
mdFile.new_line(
f"**Mean MOSNet:** {sum(data['mosnet_scores'])/len(data['mosnet_scores']):.2f}"
)
mdFile.new_line(
f"**Mean SRMR:** {sum(data['srmr_scores'])/len(data['srmr_scores']):.2f}"
)
if len(data["files_with_error"]) > 0:
mdFile.new_line(f"**Errored Files:** {', '.join(data['files_with_error'])}")
list_of_strings = ["Sample Rate (Hz)", "Count"]
for k in data["sample_rates"].keys():
list_of_strings.extend([str(k), str(data["sample_rates"][k])])
mdFile.new_table(
columns=2,
rows=len(data["sample_rates"].keys()) + 1,
text=list_of_strings,
text_align="center",
)
list_of_strings = ["Audio Type", "Count"]
n_rows = 1
for k in data["channels"].keys():
if data["channels"][k] > 0:
n_rows += 1
list_of_strings.extend([str(k), str(data["channels"][k])])
mdFile.new_table(columns=2, rows=n_rows, text=list_of_strings, text_align="center")
list_of_strings = ["Audio Format", "Count"]
for k in data["extensions"].keys():
list_of_strings.extend([str(k), str(data["extensions"][k])])
mdFile.new_table(
columns=2,
rows=len(data["extensions"].keys()) + 1,
text=list_of_strings,
text_align="center",
)
list_of_strings = ["Sample Format", "Count"]
for k in data["sample_formats"].keys():
list_of_strings.extend([str(k), str(data["sample_formats"][k])])
mdFile.new_table(
columns=2,
rows=len(data["sample_formats"].keys()) + 1,
text=list_of_strings,
text_align="center",
)
list_of_strings = ["Arpabet Lookup Type", "Count"]
for k in data["lookup_results"].keys():
list_of_strings.extend([str(k), str(len(data["lookup_results"][k]))])
mdFile.new_table(
columns=2,
rows=len(data["lookup_results"].keys()) + 1,
text=list_of_strings,
text_align="center",
)
mdFile.new_line(
mdFile.new_inline_image(
text="Wordcloud", path=os.path.join(output_folder, "wordcloud.png")
)
)
mdFile.new_line(
mdFile.new_inline_image(
text="Audio Lengths", path=os.path.join(output_folder, "lengths.png")
)
)
mdFile.new_line(
mdFile.new_inline_image(
text="Paces", path=os.path.join(output_folder, "paces.png")
)
)
mdFile.new_line(
mdFile.new_inline_image(
text="Silences", path=os.path.join(output_folder, "silences.png")
)
)
if len(data["mosnet_scores"]) > 0:
mdFile.new_line(
mdFile.new_inline_image(
text="Metrics", path=os.path.join(output_folder, "metrics.png")
)
)
mdFile.new_line(
mdFile.new_inline_image(
text="Word Frequencies",
path=os.path.join(output_folder, "word_frequencies.png"),
)
)
mdFile.new_line(
mdFile.new_inline_image(
text="Pitch and Loudness",
path=os.path.join(output_folder, "pitch_loudness.png"),
)
)
rnn_frequency_counts = count_frequency(data["lookup_results"]["RNN"])
list_of_strings = ["Frequently Missed Words", "Count"]
n_rows = 0
for k in rnn_frequency_counts.keys():
if rnn_frequency_counts[k] > 1:
n_rows += 1
list_of_strings.extend([str(k), str(rnn_frequency_counts[k])])
mdFile.new_table(
columns=2, rows=n_rows + 1, text=list_of_strings, text_align="center",
)
mdFile.new_line(
f'**Words not found in CMU:** {", ".join(data["lookup_results"]["RNN"])}'
)
mdFile.create_md_file()
# +
# export
def parse_args(args):
parser = argparse.ArgumentParser()
parser.add_argument(
"-d", "--dataset_path", help="Path to the dataset.", type=str, required=True
)
parser.add_argument(
"-i",
"--input_file",
help="Path to the transcription file.",
type=str,
required=True,
)
parser.add_argument(
"-o",
"--output_file",
help="Markdown file to write statistics to.",
type=str,
default="README",
)
parser.add_argument(
"--output_folder",
help="Folder to save plots and images.",
type=str,
default="stats",
)
parser.add_argument(
"--delimiter", help="Transcription file delimiter.", type=str, default="|"
)
parser.add_argument("--metrics", dest="metrics", action="store_true")
parser.add_argument("--no-metrics", dest="metrics", action="store_false")
parser.add_argument("--wordcloud", dest="wordcloud", action="store_true")
parser.add_argument("--no-wordcloud", dest="wordcloud", action="store_false")
parser.set_defaults(metrics=True, wordcloud=True)
return parser.parse_args(args)
def run(
dataset_path, input_file, output_file, output_folder, delimiter, metrics, wordcloud
):
if not os.path.exists(os.path.join(dataset_path, input_file)):
raise Exception(
f"Transcription file {os.path.join(dataset_path,input_file)} does not exist"
)
os.makedirs(os.path.join(dataset_path, output_folder), exist_ok=True)
data = calculate_statistics(
dataset_path, input_file, output_folder, delimiter, metrics, wordcloud
)
if data:
generate_markdown(output_file, dataset_path, output_folder, data)
with open(os.path.join(dataset_path, "stats.json"), "w") as outfile:
keys = [
"n_clips",
"total_lengths_summary",
"paces_phonemes_summary",
"paces_characters_summary",
"mosnet_scores_summary",
"srmr_scores_summary",
"pitch_summary",
"loudness_summary",
"sample_rates",
"channels",
"extensions",
"sample_formats",
]
json_data = {k: data[k] for k in keys}
json_data["arpabet_rnn"] = data["lookup_results"]["RNN"]
json.dump(json_data, outfile, indent=2)
# +
# import glob
# folders = glob.glob("/home/ubuntu/data/uberduck-multispeaker/*/*.txt")
# for dataset in folders:
# split = dataset.split("/")
# file = split[-1]
# dataset_path = "/".join(split[:-1])
# run(
# dataset_path=dataset_path,
# input_file=file,
# output_file="README.md",
# output_folder="imgs",
# delimiter="|",
# metrics=True,
# wordcloud=True,
# )
# +
# run(
# dataset_path="/home/ubuntu/data/uberduck-multispeaker/bullwinkle",
# input_file="list.txt",
# output_file="STATISTICS.md",
# output_folder="imgs",
# delimiter="|",
# metrics=False,
# wordcloud=False,
# )
# +
# export
try:
from nbdev.imports import IN_NOTEBOOK
except:
IN_NOTEBOOK = False
if __name__ == "__main__" and not IN_NOTEBOOK:
args = parse_args(sys.argv[1:])
if os.path.exists(
os.path.join(args.dataset_path, args.output_file)
) or os.path.exists(os.path.join(args.dataset_path, args.output_file + ".md")):
inp = input(
f"This script will overwite everything in the {args.output_file} file with dataset statistics. Would you like to continue? (y/n) "
).lower()
if inp != "y":
print("Not calculating statistics...")
print("HINT: Use -o/--output-file to specify a new markdown file name")
sys.exit()
print("Calculating statistics...")
run(
args.dataset_path,
args.input_file,
args.output_file,
args.output_folder,
args.delimiter,
args.metrics,
args.wordcloud,
)
# +
from g2p_en import G2p
g2p = G2p()
assert g2p.check_lookup("this is a test") == {"CMU": ["this", "is", "a", "test"]}
| nbs/exec.dataset_statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:econml-dowhy-py38]
# language: python
# name: conda-env-econml-dowhy-py38-py
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
from sklearn.linear_model import LinearRegression
# -
# # Additional data visualizations
#
# * Scatter plot for spurious correlation between sociology doctorates and non-commercial space launches. based on: https://www.tylervigen.com/spurious-correlations
data = np.array(
[
[601, 54],
[579, 46],
[617, 50],
[566, 43],
[547, 41],
[597, 46],
[580, 39],
[536, 37],
[579, 45],
[576, 45],
[601, 41],
[664, 54]
]
)
linreg = LinearRegression()
linreg.fit(data[:, 0].reshape(-1, 1), data[:, 1].reshape(-1, 1))
y_pred = linreg.predict(data[:, 0].reshape(-1, 1)).squeeze()
plt.figure(figsize=(5, 4))
plt.scatter(data[:, 0], data[:, 1], alpha=.7)
plt.plot(data[:, 0], y_pred, alpha=.5, color='red')
plt.xlabel('Sociology PhDs', fontsize=12, alpha=.5)
plt.ylabel('Space launches', fontsize=12, alpha=.5)
plt.xticks(fontsize=9)
plt.yticks(fontsize=9)
plt.show()
| add_dataviz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch]
# language: python
# name: conda-env-pytorch-py
# ---
# ## Uncertainity in Deep Learning
#
# Taken from http://mlg.eng.cam.ac.uk/yarin/blog_3d801aa532c1ce.html#uncertainty-sense
#
# Also see: https://alexgkendall.com/computer_vision/bayesian_deep_learning_for_safe_ai/
# +
# %matplotlib inline
import numpy as np
import torch
from torch.autograd import Variable
import matplotlib.pyplot as plt
from tqdm import trange
# -
def get_data(N, min_x=0., max_x=10.):
"""Generate some dummy data with interesting properties"""
np.random.seed(1)
def true_model(x, e):
alpha, beta = 4, 13
y = (x / 10. + np.sin(alpha*(x / 10. + e)) + np.sin(beta*(x / 10. + e)) + e) * 2.
return y
X_obs = np.linspace(min_x, max_x, N)
y_obs = true_model(X_obs, 0.03*np.random.rand(N))
X_true = np.linspace(min_x - 10, max_x, 100)
return X_obs, y_obs, X_true
N = 20
l2 = 0.01
X_obs, y_obs, X_true = get_data(N)
# Normalise data:
X_mean, X_std = X_obs.mean(), X_obs.std()
y_mean, y_std = y_obs.mean(), y_obs.std()
X_obs = (X_obs - X_mean) / X_std
y_obs = (y_obs - y_mean) / y_std
X_true = (X_true - X_mean) / X_std
class SimpleModel(torch.nn.Module):
def __init__(self, p, decay):
super(SimpleModel, self).__init__()
self.dropout_p = p
self.decay = decay
self.f = torch.nn.Sequential(
torch.nn.Linear(1,20),
torch.nn.ReLU(),
torch.nn.Dropout(p=self.dropout_p),
torch.nn.Linear(20, 20),
torch.nn.ReLU(),
torch.nn.Dropout(p=self.dropout_p),
torch.nn.Linear(20,1)
)
def forward(self, X):
X = Variable(torch.Tensor(X), requires_grad=False)
return self.f(X)
model = SimpleModel(p=0.1, decay=1e-6)
def uncertainity_estimate(X, model, iters, l2):
outputs = np.hstack([model(X[:, np.newaxis]).data.numpy() for i in trange(iters)])
y_mean = outputs.mean(axis=1)
y_variance = outputs.var(axis=1)
tau = l2 * (1. - model.dropout_p) / (2. * N * model.decay)
y_variance += (1. / tau)
y_std = np.sqrt(y_variance)
return y_mean, y_std
def plot_model(model, l2, iters=200, n_std=2, ax=None):
if ax is None:
plt.close("all")
plt.clf()
fig, ax = plt.subplots(1,1)
y_mean, y_std = uncertainity_estimate(X_true, model, iters, l2)
ax.plot(X_obs, y_obs, ls="none", marker="o", color="0.1", alpha=0.8, label="observed")
ax.plot(X_true, y_mean, ls="-", color="b", label="mean")
for i in range(n_std):
ax.fill_between(
X_true,
y_mean - y_std * ((i+1.)/2.),
y_mean + y_std * ((i+1.)/2.),
color="b",
alpha=0.1
)
ax.legend()
return ax
plot_model(model, l2, n_std=2)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.SGD(
model.parameters(), lr=0.01, momentum=0.,
weight_decay=model.decay)
def fit_model(model, optimizer):
y = Variable(torch.Tensor(y_obs[:, np.newaxis]), requires_grad=False)
y_pred = model(X_obs[:, np.newaxis])
optimizer.zero_grad()
loss = criterion(y_pred, y)
loss.backward()
optimizer.step()
return loss
fig = plt.figure(figsize=(10, 15))
ax0 = plt.subplot2grid((3,1), (0, 0), rowspan=2)
ax1 = plt.subplot2grid((3,1), (2, 0))
losses = []
for i in trange(10000):
loss = fit_model(model, optimizer)
losses.append(loss.data.numpy()[0])
print("loss={}".format(loss))
ax1.plot(losses, ls="-", lw=1, alpha=0.5)
plot_model(model, l2, ax=ax0)
| Pytorch+Uncertainity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Pgf Texsystem
#
#
# +
import matplotlib.pyplot as plt
plt.rcParams.update({
"pgf.texsystem": "pdflatex",
"pgf.preamble": [
r"\usepackage[utf8x]{inputenc}",
r"\usepackage[T1]{fontenc}",
r"\usepackage{cmbright}",
]
})
plt.figure(figsize=(4.5, 2.5))
plt.plot(range(5))
plt.text(0.5, 3., "serif", family="serif")
plt.text(0.5, 2., "monospace", family="monospace")
plt.text(2.5, 2., "sans-serif", family="sans-serif")
plt.xlabel(r"µ is not $\mu$")
plt.tight_layout(.5)
plt.savefig("pgf_texsystem.pdf")
plt.savefig("pgf_texsystem.png")
| matplotlib/gallery_jupyter/userdemo/pgf_texsystem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### summary
# - function
# - docstring : 함수 밑에 싱글라인, 멀티라인으로 함수에 대한 설명을 작성
# - scope : 전역(global) - 모든 변수에서 사용, 지역(local) 함수내에서만 사용
# - local이 우선 global이 나중에 저장, local은 함수 호출될 때 사용
# - inner function : 함수 안에 지역영억으로 함수를 선언
# - 지역영역이 아닌 글로벌로 사용하려면 return, inner function 해야함
# - lambda function : 간략한 함수를 한줄의 코드로 작성
# - decorator : 특정 기능을 데코레이터 함수로 만들어 함수에 특정 기능을 적용하는 방법 @로 사용
# - class
# - 변수와 함수들이 모여있는 집합
# - 기본클래스 사용법
# - 클래스의 선언 -> 객체로 만듬 -> 객체의 함수를 호출
# - 생성자 함수
# - 클래스가 객체로 만들어질때 객체에 선언되는 변수를 설정하는 방법
ls = ["a","b","c"]
print(list(range(len(ls))))
print(list(zip(list(range(len(ls))),ls)))
for idx, data in list(zip(list(range(len(ls))), ls)) :
print(idx, data)
user_datas = [
{"user" : "test", "pw" : "1234", "count" : 0},
{"user" : "python", "pw" : "5678", "count" : 0},
]
# +
# user_data를 입력받아서 id와 패스워드를 체크하는 데코레이터 함수를 코드로 작성하세요
# 로그인 될때마다 count를 1씩 증가
def need_login(func):
def wrapper(*args, **kwargs):
#아이디 패스워드 입력
user, pw = tuple(input("input id pw : ").split(" "))
# 존재하는 아이디 패스워드 확인
for idx, user_data in zip(range(len(user_datas)), user_datas) :
if (user_data["user"] == user) and (user_data["pw"] == pw):
# count 데이터 추가
user_datas[idx]["count"] += 1
# 함수 실행
return func(*args, **kwargs)
return "wrong login data!"
return wrapper
# -
# enumerate는 데이터 앞에 idx를 붙여줌
list(enumerate(user_datas))
# +
# user_data를 입력받아서 id와 패스워드를 체크하는 데코레이터 함수를 코드로 작성하세요
# 로그인 될때마다 count를 1씩 증가
def need_login(func):
def wrapper(*args, **kwargs):
#아이디 패스워드 입력
user, pw = tuple(input("input id pw : ").split(" "))
# 존재하는 아이디 패스워드 확인
#for idx, user_data in zip(range(len(user_datas)), user_datas) :
for idx, user_data in enumerate(user_datas):
if (user_data["user"] == user) and (user_data["pw"] == pw):
# count 데이터 추가
user_datas[idx]["count"] += 1
# 함수 실행
return func(*args, **kwargs)
return "wrong login data!"
return wrapper
# -
@need_login
def plus(num1, num2):
return num1 + num2
plus(1,2)
user_datas
# +
# 스타크래프트의 마린을 클래스로 설계
# 체력(health : 40), 공격력(attack_pow : 5), 공격(attack())
# 마린 클래스로 마린 객체 2개를 생성해서 마린1이 마린2를 공격하는 코드를 작성
# attack(self, unit)
# -
class Marine :
# 보통 생성자 안에 변수를 생성함 self는 객체 self.health 객체의 변수
# ()안에 변수를 지정해주면, 객체를 생성시 해당 변수값을 설정 가능함
def __init__(self, max_health = 40, attact_pow = 5):
# 함수안에 변수를 지정해주면 객체를 생성 후 다시 변수를 지정해줘야함
self.health = max_health
self.max_health = max_health
self.attact_pow = attact_pow
def attact(self, unit) :
unit.health -= self.attact_pow
if unit.health <= 0 :
unit.health = 0
print("사망")
# +
# 메딕 : heal_pow, heal(unit) = 함수
class Medic:
def __init__(self, max_health = 40, heal_pow = 6):
self.max_health = max_health
self.health = max_health
self.heal_pow = heal_pow
def heal(self, unit):
if unit.health > 0:
unit.health += self.heal_pow
if unit.health >= unit.max_health :
unit.health = unit.max_health
else :
print("이미 사망")
# -
medic = Medic()
marine_1 = Marine()
marine_2 = Marine()
marine_1.attact(marine_2)
marine_1.health, marine_2.health
medic.heal(marine_2)
marine_1.health, marine_2.health
marine_3 = Marine(attact_pow= 20)
marine_3.attact(marine_1)
marine_1.health
# ### 1. 상속
# - 클래스의 기능을 가져다가 기능을 수정하거나 추가할때 사용하는 방법
# 클래스 생성
class Calculator :
# 생성자 생성
def __init__(self, num1, num2):
self.num1 = num1
self.num2 = num2
def plus(self):
return self.num1 + self.num2
calc = Calculator(2,3)
calc.plus()
# +
# minus 기능을 추가한 계산기
# -
class Calculator2 :
def __init__(self, num1, num2):
self.num1 = num1
self.num2 = num2
def plus(self):
return self.num1 + self.num2
# 여기까지는 Calculator 에 있는 코드로 중복됨
# 이럴때는 상속의 기능을 사용하여 클래스를 만듬
def minus(self):
return self.num1 - self.num2
calc2 = Calculator2(1,2)
calc2.minus()
# +
# 상속기능을 사용하여 minus 함수 추가
# -
# 클래스 선언시 뒤에 상속해주는 클래스를 넣어주면 상속됨
class Calculator3(Calculator) : # <- 이부분
def minus(self):
return self.num1 - self.num2
calc3 = Calculator3(1,2)
calc3.plus(), calc3.minus()
# 매서드 오버라이딩 : 많은 코드들 중 수정하고 싶은 함수만 바꾸고 싶음
class Calculator4(Calculator3):
# 같은 함수 이름을 사용하면 기존 코드가 수정이 됨
# 위의 경우에도 Calclator3의 Plus도 변경
# 기존 상속 전 기능은 그대로 유지됨
def plus(self):
return self.num1 ** 2 + self.num2 ** 2
# +
# 아이폰 1, 2, 3
# 아이폰 1 : calling - print("calling")
# 아이폰 2 : 아이폰 1 + send msg
# 아이폰 3 : 아이폰2 + internet
# -
class Iphone1 :
def calling(self) :
print("calling")
class Iphone2(Iphone1) :
def send_msg(self):
print("send_msg")
class Iphone3(Iphone2) :
def internet(sefl):
print("internet")
iphone3 = Iphone3
dir(iphone3)
# +
# 다중 상속
# -
class Galuxy :
def show_img(self):
print("show_img")
class DssPhone(Iphone3, Galuxy): # class DssPhone(class Iphone(Galuxy)) 순서로 상속됨
def camera(self):
print("camera")
dss_phone = DssPhone
[func for func in dir(dss_phone) if func[:2] !="__"]
# ### 2. super
# - 부모 클래스에서 사용된 함수의 코드를 가져다가 자식 클래스의 함수에서 재사용 할때 사용
#
# ```
# class A:
# def plus(self):
# code1
#
# class B(A) :
# def minus(self):
# code1 # super().plus() #code1은 classA에서 바꿔도 같이 바뀜
# code2
#
# ```
class Marine :
def __init__(self):
self.health = 40
self.attact_pow = 5
def attact(self, unit) :
unit.health -= self.attact_pow
if unit.health <= 0 :
unit.health = 0
class Marine2(Marine) :
def __init__(self):
# self.health = 40
# self.attact_pow = 5
super().__init__() # super를 안쓰고 함수를 작성하면 오버라이딩 되서 코드가 사라짐
# super().함수이름.()
self.max_health = 40 # __init__에 추가한 변수
marine = Marine2()
marine.health, marine.attact_pow, marine.max_health
# ### 3. class의 getter, setter
# - 객체의 내부 변수에 접근할때 특정 로직을 거쳐서 접근 시키는 방법
class User:
def __init__(self, first_name):
self.first_name = first_name
def setter(self, first_name):
print("setter")
self.first_name = first_name
def getter(self):
print("getter")
return self.first_name
name = property(getter, setter)
user1 = User("andy")
user1.first_name
# setter 함수 실행
user1.name = 1
# getter 함수 실행
user1.name
user1.first_name
# +
# 변수, 함수마다 getter, setter를 각각 만들어줘야 함
class User:
def __init__(self, first_name):
self.first_name = first_name
def setter(self, first_name):
if len(first_name) >= 3 :
self.first_name = first_name
print("setter")
else :
print("error")
def getter(self):
print("getter")
return self.first_name
name = property(getter, setter) # property 변수에 getter, setter가 접근하도록 하는 함수
# -
user1 = User("andy")
user1.first_name
user1.name = "a"
# ### 4. non public
# - mangling 이라는 방법으로 다이렉트로 객체의 변수에 접근하지 못하게 하는 방법
class Calculator :
def __init__(self, num1, num2):
self.num1 = num1
self.num2 = num2
def getter(self):
return self.num2
# num2에 0이 들어가지 않도록 함
def setter(self, num2):
num2 = 1 if num2 == 0 else num2
self.num2 = num2
def div(self):
return self.num1 / self.num2 # num2에 0이 들어가면 에러가뜸
number2 = property(getter, setter)
calc = Calculator(1,2)
calc.div()
calc.number2
calc.number2 = 0
calc.number2
calc.num2 = 0 #num2 라는 변수에 "직접" 넣으면 setter 함수를 거치지 않고 값이 바뀜
calc.num2
calc.div() # num2에 0이 들어가면 에러가뜸
class Calculator :
def __init__(self, num1, num2):
self.num1 = num1
self.__num2 = num2 #앞에 __를 넣어서 바뀌지 않게 함 mangling
def getter(self):
return self.__num2
# num2에 0이 들어가지 않도록 함
def setter(self, num2):
num2 = 1 if num2 == 0 else num2
self.__num2 = num2
def div(self):
return self.num1 / self.__num2 # num2에 0이 들어가면 에러가뜸
number2 = property(getter, setter)
calc = Calculator(1,2)
calc.num1
calc.num2 # calc에 num2라는 변수는 없음
calc.__num2 # 마찬가지로 calc에 mangling으로 넣은__num2라는 변수도 없음
calc._Calculator__num2
# 작성할땐 __num2지만 실제로는 _(class명)__(변수명) [_calculator__num2]로 저장됨
# 따라서 불러올때는 (객체명)._(class명)__(변수명)으로 불러옴
# +
# num2라는 변수를 생성하면서 0을 넣을수는 있음
# num2 != __num2 (_class명__변수명) 임.
# 결국 div 함수의 self.num1 / self.__num2 와는 상관이 없는 변수임
calc.num2 = 0
calc.num2
# +
#함수에 mangling하기
class Calculator :
def __init__(self, num1, num2):
self.num1 = num1
self.__num2 = num2 #앞에 __를 넣어서 바뀌지 않게 함 mangling
def getter(self):
return self.__num2
# num2에 0이 들어가지 않도록 함
def setter(self, num2):
num2 = 1 if num2 == 0 else num2
self.__num2 = num2
# __를 함수 붙여 mangling을 사용하며, 단독 함수 호출로 사용못함
def __disp(self):
print(self.num1, self.__num2)
def div(self):
self.__disp()
return self.num1 / self.__num2 # num2에 0이 들어가면 에러가뜸
number2 = property(getter, setter)
# -
calc = Calculator(1,2)
calc.div()
# ### 5. is a & has a
# - 클래스를 설계하는 개념
# - A is a B
# - A는 B이다. 상속을 이용해서 클래스를 만드는 방법
# - A has a B
# - A는 B를 가진다. A가 B객체를 가지고 클래스를 만드는 방법`
# +
# 사람 : 이름, 이메일, 정보출력()
# -
# is a
class Person:
def __init__(self, name, email) :
self.name = name
self.email = email
class Person2(Person):
def info(self):
print(self.name, self.email)
p = Person2("andy", "<EMAIL>",)
p.info()
# has a
class Name:
def __init__(self, name) :
self.name_str = name
class Email:
def __init__(self, email):
self.email_str = email
class Person:
def __init__(self, name_obj, email_obj):
self.name = name_obj
self.email = email_obj
def info(self):
print(name.name_str, email.email_str)
name = Name("andy")
email = Email("<EMAIL>")
p = Person(name, email)
p.info()
# ### 6. Magic(Special) method
# - compare
# - `__eq__` : ==
# - `__ne__` : !=
# - `__lt__` : <
# - calculate
# - `__add__` : +
# - `__sub__` : -
# - `__repr__` : 객체의 내용을 출력(개발자용)
# - `__str__`
"test" == "test"
"test".__eq__("test")
1 + 2 == "1" + "2"
class Txt:
def __init__(self, txt):
self.txt = txt
def __eq__(self, txt_obj):
return self.txt.lower() == txt_obj.txt.lower()
def __repr__(self):
return "Txt(txt = {})".format(self.txt)
def __str__(self):
return self.txt
t1 = Txt("python")
t2 = Txt("Python")
t3 = t1
t1 == t2, t1 == t3, t2 == t3
t1
print(t1)
| python/python_basic/07_class2_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "-"}
# # Sparseloop Tutorial - 03 - Convolution
#
# This notebook contains a series of examples of a DNN **convolution** computation. The **fibertree** emulator is used to illustrate the impact of a set of optimizations to exploit sparsity. The basic computation for a 1-D convolution with one input channel and one output channel is represented by the Einsum:
#
# $$ O_{q} = I_{(q+s)} \times F_{s} $$
#
# Note that while the output is possibly a sparse rank-1 tensor, in this notebook the output is assumed to be in an uncompressed format and is directly referenced by **coordinate** since **position** == **coordinate** in an uncompressed format.
#
# First, include some libraries
# + slideshow={"slide_type": "-"}
# Run boilerplate code to set up environment
# %run ./prelude.py --style=tree --animation=movie
# -
# ## Configure convolution tensors input tensors
#
# The following cell sets up the control sliders to specify the attributes of the 1-D `I` (input activations) and `F` (filter weight) input tensors. Those attributes include their **shape**, which specifies the allowable range of **coordinates** of elements of the tensor and their **density**.
#
# The empty 1-D 'O' (output activations) tensors are declared when used in later cells.
#
# The rank names use the following convention:
#
# - `W` - The width of the input activation tensor (`I`)
# - `S` - The width of the filter weight tensor (`F`)
# - `Q` - The width of the output activation tensor (`O`)
# - `M` - The number of output channels in `F` and `O` (used in later cells).
#
# +
#
# Set default problem instance attributes (i.e., the shape of the tensors)
#
W = 12
S = 3
#
# Create controls to configure the `I` and `F` tensors
#
tm3 = TensorMaker("sparseloop-convolution-1", autoload=True)
tm3.addTensor("F", rank_ids=["S"], shape=[S], density=0.5, color="green")
tm3.addTensor("I", rank_ids=["W"], shape=[W], density=1.0, color="blue")
tm3.displayControls()
# -
# ## Create and display convolution tensors
# +
F_S = tm3.makeTensor("F")
I_W = tm3.makeTensor("I")
displayTensor(F_S)
displayTensor(I_W)
# -
# # Simple 1-D Convolution
#
# $$ O_{q} = I_{(q+s)} \times F_{s} $$
# +
#
# Create input convolution tensors
#
S = getShape(tm3, "S")
W = getShape(tm3, "W")
Q = W-S+1
F_S = tm3.makeTensor("F")
I_W = tm3.makeTensor("I")
O_Q = Tensor(name="O", rank_ids=["Q"], shape=[Q])
uncompressTensor(O_Q)
#
# Display Tensors
#
print("Problem Instance:")
print(f"S: {S}")
print(f"W: {W}")
print(f"Q: {Q}")
print("")
print("Filter weight tensor F")
displayTensor(F_S)
print("Input activation tensor I")
displayTensor(I_W)
print("Output activation tensor O (initial)")
displayTensor(O_Q)
#
# Get root of tensors
#
i_w = I_W.getRoot()
f_s = F_S.getRoot()
o_q = O_Q.getRoot()
#
# Animation bookkeeeping
#
canvas = createCanvas(F_S, I_W, O_Q)
cycle = 0
#
# Traverse all `Q` coordinates of the output tensor
#
for q in range(Q):
#
# Traverse the non-empty coordinates of the filter weights
#
for s, f_val in f_s:
#
# Compute and fetch the required input activation coordinate
#
w = q + s
i_val = i_w.getPayload(w)
#
# Compute value to contribute to partial output sum
#
o_q[q] += i_val * f_val
#
# Animation bookkeeping
#
canvas.addActivity((s,), (w,), (q,), spacetime=(0,cycle))
cycle += 1
#
# Display results
#
print("Output activation tensor O (final)")
displayTensor(O_Q)
displayCanvas(canvas)
# -
# # Convolution - 1D - with output channels
#
# $$ O_{m,q} = I_{(q+s)} \times F_{m,s} $$
# ## Configure convolution tensors
# +
#
# Set default problem instance attributes (i.e., the shape of the tensors)
#
M = 4
W = 12
S = 4
#
# Create controls to configure the `I` and `F` tensors
#
tm4 = TensorMaker("sparseloop-convolution-2", autoload=True)
tm4.addTensor("F", rank_ids=["M", "S"], shape=[M, S], density=0.5, color="green")
tm4.addTensor("I", rank_ids=["W"], shape=[W], density=1.0, color="blue")
tm4.displayControls()
# -
# ## Create and display convolution tensors
# +
F_MS = tm4.makeTensor("F")
I_W = tm4.makeTensor("I")
displayTensor(F_MS)
displayTensor(I_W)
# -
# ## Convolution with multiple output channels
# +
#
# Create input convolution tensors
#
M = getShape(tm4, "M")
S = getShape(tm4, "S")
W = getShape(tm4, "W")
Q = W - S + 1
F_MS = tm4.makeTensor("F")
I_W = tm4.makeTensor("I")
O_MQ = Tensor(name="O", rank_ids=["M", "Q"], shape=[M, Q])
uncompressTensor(O_MQ)
#
# Display Tensors
#
print("Problem Instance:")
print(f"M: {M}")
print(f"S: {S}")
print(f"W: {W}")
print(f"Q: {Q}")
print("")
print("Filter weight tensor F")
displayTensor(F_MS)
print("Input activation tensor I")
displayTensor(I_W)
print("Output activation tensor O (initial)")
displayTensor(O_MQ)
#
# Get root of tensors
#
i_w = I_W.getRoot()
f_m = F_MS.getRoot()
o_m = O_MQ.getRoot()
#
# Animation bookkeeeping
#
canvas = createCanvas(F_MS, I_W, O_MQ)
cycle = 0
#
# Traverse filter weight output channels
#
for m, f_s in f_m:
#
# Traverse all `Q` coordinates of the output tensor
#
for q in range(Q):
#
# Traverse all non-empty filter weights for this output channel
#
for s, f_val in f_s:
#
# Compute and fetch the required input activation coordinate
#
w = q + s
i_val = i_w.getPayload(w)
#
# Compute value to contribute to partial output sum
#
o_m[m][q] += i_val * f_val
#
# Animation bookkeeping
#
canvas.addActivity((m,s), (w,), (m,q), spacetime=(0,cycle))
cycle += 1
#
# Display results
#
print("Output activation tensor O (initial)")
displayTensor(O_MQ)
displayCanvas(canvas)
# -
# ## Convolution - spatial output channels
# +
#
# Create input convolution tensors
#
split = 2
F_MS = tm4.makeTensor("F")
F_M1M0S = F_MS.splitUniform(split)
F_M1M0S = F_M1M0S.updateCoords(lambda n, c, p: n)
I_W = tm4.makeTensor("I")
W = I_W.getShape("W")
S = F_MS.getShape("S")
Q = W-S+1
M = F_MS.getShape("M")
M0 = split
M1 = (M+split-1)//split
O_MQ = Tensor(name="O", rank_ids=["M", "Q"], shape=[M, Q])
uncompressTensor(O_MQ)
O_M1M0Q = O_MQ.splitUniform(split)
#
# Display Tensors
#
print("Problem Instance:")
print(f"M1: {M1}")
print(f"M0: {M0}")
print(f"W: {W}")
print(f"Q: {Q}")
print(f"S: {S}")
displayTensor(F_M1M0S)
displayTensor(I_W)
displayTensor(O_M1M0Q)
#
# Get root of tensors
#
i_w = I_W.getRoot()
f_m1 = F_M1M0S.getRoot()
o_m1 = O_M1M0Q.getRoot()
#
# Animation bookkeeeping
#
canvas = createCanvas(F_M1M0S, I_W, O_M1M0Q)
#
# Traverse group of filter weight output channels
#
for m1, f_m0 in f_m1:
#
# Traverse filter weight output channels in this group
#
for m0, f_s in f_m0:
#
# Traverse all output locations for this output channel
#
for q in range(Q):
#
# Traverse all output locations
#
for s, f_val in f_s:
#
# Compute and fetch the required input activation coordinate
#
w = q + s
i_val = i_w.getPayload(w)
#
# Compute value to contribute to partial output sum
#
o_m1[m1][m0%M0][q] += i_val * f_val
#
# Animation bookkeeping
#
spacestamp = m0
# TBD: Time should be counted in position not coordinates!
timestamp = m1*W*S + q*S + s
canvas.addActivity((m1,m0,s), (w,), (m1,m0,q),
spacetime=(spacestamp, timestamp))
displayTensor(O_M1M0Q)
displayCanvas(canvas)
# -
# ## Testing area
#
# For running alternative algorithms
| workspace/exercises/2021.isca/notebooks/03.1-convolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7765UFHoyGx6"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="KVtTDrUNyL7x"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="xPYxZMrWyA0N"
# # Estimators を使用するブースティング木
# + [markdown] id="p_vOREjRx-Y0"
# <table class="tfo-notebook-buttons" align="left">
# <td> <img src="https://www.tensorflow.org/images/tf_logo_32px.png"><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/tutorials/estimator/boosted_trees.ipynb">TensorFlow.org で表示</a> </td>
# <td> <img src="https://www.tensorflow.org/images/colab_logo_32px.png"><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/tutorials/estimator/boosted_trees.ipynb">Google Colab で実行</a> </td>
# <td> <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png"><a target="_blank" href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/tutorials/estimator/boosted_trees.ipynb">GitHubでソースを表示</a> </td>
# <td> <img src="https://www.tensorflow.org/images/download_logo_32px.png"><a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/estimator/boosted_trees.ipynb">ノートブックをダウンロード</a> </td>
# </table>
# + [markdown] id="6gWdn5lrlkhR"
# > 警告: 新しいコードには Estimators は推奨されません。Estimators は `v1.Session` スタイルのコードを実行しますが、これは正しく記述するのはより難しく、特に TF 2 コードと組み合わせると予期しない動作をする可能性があります。Estimators は、[互換性保証] (https://tensorflow.org/guide/versions) の対象となりますが、セキュリティの脆弱性以外の修正は行われません。詳細については、[移行ガイド](https://tensorflow.org/guide/migrate)を参照してください。
# + [markdown] id="qNW3c_rop5J8"
# **注意**: 多くの最先端の決定フォレストアルゴリズムの最新の Keras ベースの実装は、[TensorFlow 決定フォレスト](https://tensorflow.org/decision_forests)から利用できます。
# + [markdown] id="dW3r7qVxzqN5"
# このチュートリアルは、`tf.estimator`API で決定木を使用する勾配ブースティングモデルのエンドツーエンドのウォークスルーです。ブースティング木モデルは、回帰と分類の両方のための最も一般的かつ効果的な機械学習アプローチの 1 つです。これは、複数(10 以上、100 以上、あるいは 1000 以上の場合も考えられます)の木モデルからの予測値を結合するアンサンブル手法です。
#
# 最小限のハイパーパラメータ調整で優れたパフォーマンスを実現できるため、ブースティング木モデルは多くの機械学習実践者に人気があります。
# + [markdown] id="eylrTPAN3rJV"
# ## Titanic データセットを読み込む
#
# Titanic データセットを使用します。ここでの目標は、性別、年齢、クラスなど与えられた特徴から(やや悪趣味ではありますが)乗船者の生存を予測することです。
# + id="KuhAiPfZ3rJW"
import numpy as np
import pandas as pd
from IPython.display import clear_output
from matplotlib import pyplot as plt
# Load dataset.
dftrain = pd.read_csv('https://storage.googleapis.com/tf-datasets/titanic/train.csv')
dfeval = pd.read_csv('https://storage.googleapis.com/tf-datasets/titanic/eval.csv')
y_train = dftrain.pop('survived')
y_eval = dfeval.pop('survived')
# + id="NFtnFm1T0kMf"
import tensorflow as tf
tf.random.set_seed(123)
# + [markdown] id="3ioodHdVJVdA"
# データセットはトレーニングセットと評価セットで構成されています。
#
# - `dftrain`と`y_train`は *トレーニングセット*です — モデルが学習に使用するデータです。
# - モデルは*評価セット*、`dfeval`、`y_eval`に対してテストされます。
#
# トレーニングには以下の特徴を使用します。
#
# <table>
# <tr>
# <th>特徴名</th>
# <th>説明</th>
# </tr>
# <tr>
# <td>sex</td>
# <td>乗船者の性別</td>
# </tr>
# <tr>
# <td>age</td>
# <td>乗船者の年齢</td>
# </tr>
# <tr>
# <td>n_siblings_spouses</td>
# <td>同乗する兄弟姉妹および配偶者</td>
# </tr>
# <tr>
# <td>parch</td>
# <td>同乗する両親および子供</td>
# </tr>
# <tr>
# <td>fare</td>
# <td>運賃</td>
# </tr>
# <tr>
# <td>class</td>
# <td>船室のクラス</td>
# </tr>
# <tr>
# <td>deck</td>
# <td>搭乗デッキ</td>
# </tr>
# <tr>
# <td>embark_town</td>
# <td>乗船者の乗船地</td>
# </tr>
# <tr>
# <td>alone</td>
# <td>一人旅か否か</td>
# </tr>
# </table>
# + [markdown] id="AoPiWsJALr-k"
# ## データを検証する
# + [markdown] id="slcat1yzmzw5"
# まず最初に、データの一部をプレビューして、トレーニングセットの要約統計を作成します。
# + id="15PLelXBlxEW"
dftrain.head()
# + id="j2hiM4ETmqP0"
dftrain.describe()
# + [markdown] id="-IR0e8V-LyJ4"
# トレーニングセットと評価セットには、それぞれ 627 個と 264 個の例があります。
# + id="_1NwYqGwDjFf"
dftrain.shape[0], dfeval.shape[0]
# + [markdown] id="28UFJ4KSMK3V"
# 乗船者の大半は 20 代から 30 代です。
# + id="CaVDmZtuDfux"
dftrain.age.hist(bins=20)
plt.show()
# + [markdown] id="1pifWiCoMbR5"
# 男性の乗船者数は女性の乗船者数の約 2 倍です。
# + id="-WazAq30MO5J"
dftrain.sex.value_counts().plot(kind='barh')
plt.show()
# + [markdown] id="7_XkxrpmmVU_"
# 乗船者の大半は「3 等」の船室クラスを利用していました。
# + id="zZ3PvVy4l4gI"
dftrain['class'].value_counts().plot(kind='barh')
plt.show()
# + [markdown] id="HM5SlwlxmZMT"
# 大半の乗船者はサウサンプトンから乗船しています。
# + id="RVTSrdr4mZaC"
dftrain['embark_town'].value_counts().plot(kind='barh')
plt.show()
# + [markdown] id="aTn1niLPob3x"
# 女性は男性よりも生存する確率がはるかに高く、これは明らかにモデルの予測特徴です。
# + id="Eh3KW5oYkaNS"
pd.concat([dftrain, y_train], axis=1).groupby('sex').survived.mean().plot(kind='barh').set_xlabel('% survive')
plt.show()
# + [markdown] id="krkRHuMp3rJn"
# ## 特徴量カラムを作成して関数を入力する
#
# 勾配ブースティング Estimator は数値特徴とカテゴリ特徴の両方を利用します。特徴量カラムは、全ての TensorFlow Estimator と機能し、その目的はモデリングに使用される特徴を定義することにあります。さらに、One-Hot エンコーディング、正規化、バケット化などいくつかの特徴量エンジニアリング機能を提供します。このチュートリアルでは、`CATEGORICAL_COLUMNS`のフィールドはカテゴリカラムから One-Hot エンコーディングされたカラム([インジケータカラム](https://www.tensorflow.org/api_docs/python/tf/feature_column/indicator_column))に変換されます。
# + id="upaNWxcF3rJn"
CATEGORICAL_COLUMNS = ['sex', 'n_siblings_spouses', 'parch', 'class', 'deck',
'embark_town', 'alone']
NUMERIC_COLUMNS = ['age', 'fare']
def one_hot_cat_column(feature_name, vocab):
return tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(feature_name,
vocab))
feature_columns = []
for feature_name in CATEGORICAL_COLUMNS:
# Need to one-hot encode categorical features.
vocabulary = dftrain[feature_name].unique()
feature_columns.append(one_hot_cat_column(feature_name, vocabulary))
for feature_name in NUMERIC_COLUMNS:
feature_columns.append(tf.feature_column.numeric_column(feature_name,
dtype=tf.float32))
# + [markdown] id="74GNtFpStSAz"
# 特徴量カラムが生成する変換は表示することができます。例えば、`indicator_column`を単一の例で使用した場合の出力は次のようになります。
# + id="Eaq79D9FtmF8"
example = dict(dftrain.head(1))
class_fc = tf.feature_column.indicator_column(tf.feature_column.categorical_column_with_vocabulary_list('class', ('First', 'Second', 'Third')))
print('Feature value: "{}"'.format(example['class'].iloc[0]))
print('One-hot encoded: ', tf.keras.layers.DenseFeatures([class_fc])(example).numpy())
# + [markdown] id="YbCUn3nCusC3"
# さらに、特徴量カラムの変換を全てまとめて表示することができます。
# + id="omIYcsVws3g0"
tf.keras.layers.DenseFeatures(feature_columns)(example).numpy()
# + [markdown] id="-UOlROp33rJo"
# 次に、入力関数を作成する必要があります。これらはトレーニングと推論の両方のためにデータをモデルに読み込む方法を指定します。[ `tf.data`](https://www.tensorflow.org/api_docs/python/tf/data) API の`from_tensor_slices`メソッドを使用して Pandas から直接データを読み取ります。これは小規模でインメモリのデータセットに適しています。大規模のデータセットの場合は、多様なファイル形式([csv](https://www.tensorflow.org/api_docs/python/tf/data/experimental/make_csv_dataset)を含む)をサポートする tf.data API を使用すると、メモリに収まりきれないデータセットも処理することができます。
# + id="9dquwCQB3rJp"
# Use entire batch since this is such a small dataset.
NUM_EXAMPLES = len(y_train)
def make_input_fn(X, y, n_epochs=None, shuffle=True):
def input_fn():
dataset = tf.data.Dataset.from_tensor_slices((dict(X), y))
if shuffle:
dataset = dataset.shuffle(NUM_EXAMPLES)
# For training, cycle thru dataset as many times as need (n_epochs=None).
dataset = dataset.repeat(n_epochs)
# In memory training doesn't use batching.
dataset = dataset.batch(NUM_EXAMPLES)
return dataset
return input_fn
# Training and evaluation input functions.
train_input_fn = make_input_fn(dftrain, y_train)
eval_input_fn = make_input_fn(dfeval, y_eval, shuffle=False, n_epochs=1)
# + [markdown] id="HttfNNlN3rJr"
# ## モデルをトレーニングして評価する
#
# 以下のステップで行います。
#
# 1. 特徴とハイパーパラメータを指定してモデルを初期化する。
# 2. `train_input_fn`を使用してモデルにトレーニングデータを与え、`train`関数を使用してモデルをトレーニングする。
# 3. 評価セット(この例では`dfeval` DataFrame)を使用してモデルのパフォーマンスを評価する。予測値が`y_eval`配列のラベルと一致することを確認する。
#
# ブースティング木モデルをトレーニングする前に、まず線形分類器(ロジスティック回帰モデル)をトレーニングしてみましょう。ベンチマークを確立するには、より単純なモデルから始めるのがベストプラクティスです。
# + id="JPOGpmmq3rJr"
linear_est = tf.estimator.LinearClassifier(feature_columns)
# Train model.
linear_est.train(train_input_fn, max_steps=100)
# Evaluation.
result = linear_est.evaluate(eval_input_fn)
clear_output()
print(pd.Series(result))
# + [markdown] id="BarkNXwA3rJu"
# 次に、ブースティング木モデルをトレーニングしてみましょう。ブースティング木では、回帰(`BoostedTreesRegressor`)と分類(`BoostedTreesClassifier`)をサポートします。目標は、生存か非生存かのクラスを予測することなので、`BoostedTreesClassifier`を使用します。
#
# + id="tgEzMtlw3rJu"
# Since data fits into memory, use entire dataset per layer. It will be faster.
# Above one batch is defined as the entire dataset.
n_batches = 1
est = tf.estimator.BoostedTreesClassifier(feature_columns,
n_batches_per_layer=n_batches)
# The model will stop training once the specified number of trees is built, not
# based on the number of steps.
est.train(train_input_fn, max_steps=100)
# Eval.
result = est.evaluate(eval_input_fn)
clear_output()
print(pd.Series(result))
# + [markdown] id="hEflwznXvuMP"
# このトレーニングモデルを使用して、評価セットからある乗船者に予測を立てることができます。TensorFlow モデルは、バッチ、コレクション、または例に対してまとめて予測を立てられるように最適化されています。以前は、`eval_input_fn` は評価セット全体を使って定義されていました。
# + id="6zmIjTr73rJ4"
pred_dicts = list(est.predict(eval_input_fn))
probs = pd.Series([pred['probabilities'][1] for pred in pred_dicts])
probs.plot(kind='hist', bins=20, title='predicted probabilities')
plt.show()
# + [markdown] id="mBUaNN1BzJHG"
# 最後に、結果の受信者操作特性(ROC)を見てみましょう。真陽性率と偽陽性率間のトレードオフに関し、より明確な予想を得ることができます。
# + id="NzxghvVz3rJ6"
from sklearn.metrics import roc_curve
fpr, tpr, _ = roc_curve(y_eval, probs)
plt.plot(fpr, tpr)
plt.title('ROC curve')
plt.xlabel('false positive rate')
plt.ylabel('true positive rate')
plt.xlim(0,)
plt.ylim(0,)
plt.show()
| site/ja/tutorials/estimator/boosted_trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="bzRM0Gzmvaye"
# # Response function of a moving average
# [](https://github.com/eabarnes1010/course_objective_analysis/tree/main/code)
# [](https://colab.research.google.com/github/eabarnes1010/course_objective_analysis/blob/main/code/response_function_moving_avg.ipynb)
#
#
# Ever wonder what a moving average is doing to your data in frequency-space? Well - you're about to find out!
# + executionInfo={"elapsed": 118, "status": "ok", "timestamp": 1645362130160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="4OdMyhIjvayg"
import numpy as np
import matplotlib.pyplot as plt
import scipy.signal as sig
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 150
plt.rcParams['figure.figsize'] = (12.0/2, 8.0/2)
# + executionInfo={"elapsed": 109, "status": "ok", "timestamp": 1645362130498, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="86S-dBGTvayl"
#plotting commands
LW = 2 #linewidth
LFS = 6 #legend fontsize
# + [markdown] id="B3Pk9kqVvayo"
# ### Get your data together
# + colab={"base_uri": "https://localhost:8080/", "height": 629} executionInfo={"elapsed": 288, "status": "ok", "timestamp": 1645362130778, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="JgQV1HgKvayp" outputId="824d26ff-1b70-444e-e4cb-dd93bc97c503"
#t = np.arange(1,4000)
t = np.arange(1,30)
x = np.zeros(np.shape(t))
x[int(np.size(t)/2):int(np.size(t)/2+2)] = np.ones((2,))
#x = np.random.rand(np.size(t))
print(x)
plt.figure()
plt.title('boxcar smoothing');
plt.plot(t,x,'-k',linewidth = LW, label ='original data')
plt.ylim(0,1.1)
plt.xlabel('time')
plt.ylabel('units of x')
plt.legend(frameon = False, fontsize = LFS)
plt.show()
# + [markdown] id="hVjLwEaZvayt"
# First we define a simple data set x that is zeros everywhere except for two values in the middle that are 1.0.
# + [markdown] id="LkUlz3Xfvayu"
# ### Apply the filter in time-space and see what the resulting curve looks like
#
# Next, let's apply a 3-day moving window to x. We could write such a loop ourselves, or we could make use of built-in functions that do this convolution for us! In this case, I use sig.lfilter().
# + colab={"base_uri": "https://localhost:8080/", "height": 594} executionInfo={"elapsed": 262, "status": "ok", "timestamp": 1645362131030, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="N8Qu2jrKvayv" outputId="089ee88a-0d30-4e0e-9b81-ac074c61ff7f"
# define my moving average window
g = [1., 1., 1.] #in this case, a 3-day moving window
# apply the moving average window using a "filter" function to do the loop for me
y1 = sig.lfilter(g,np.sum(g),x)
plt.figure()
plt.title('boxcar smoothing');
plt.plot(t,x,'-k',linewidth = LW, label ='original data')
plt.plot(t,y1,'--r',linewidth = LW, label = 'smoothed with 1-1-1' )
plt.ylim(0,1.1)
plt.xlabel('time')
plt.ylabel('units of x')
plt.legend(frameon = False, fontsize = LFS)
plt.show()
# + [markdown] id="cPFRjRoAvayy"
# Notice that the data is indeed smoothed - but also phase shifted to the right! This is because of how [sig.lfilter()](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.signal.lfilter.html) is setup in python to perform a left-centered filter.
# + [markdown] id="CAgTWRQpvay0"
# Now, let's apply another 1-1-1 filter to our already smoothed data (i.e. the red curve above).
# + colab={"base_uri": "https://localhost:8080/", "height": 594} executionInfo={"elapsed": 206, "status": "ok", "timestamp": 1645362131232, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="QarFVOsBvay1" outputId="8f1900e4-9317-48c9-d2c9-12cac575ac86"
y2 = sig.filtfilt(g,np.sum(g),x) #filtfilt goes forwards and backwards to remove the phase shift
plt.figure()
plt.title('boxcar smoothing');
plt.plot(t,x,'-k',linewidth = LW, label ='original data')
plt.plot(t,y1,'--r',linewidth = LW, label = 'smoothed with 1-1-1' )
plt.plot(t,y2,'--b',linewidth = LW, label = 'smoothed with 1-1-1 twice using filtfilt' )
plt.legend(frameon = False, fontsize = LFS)
plt.ylim(0,1.1)
plt.xlabel('time')
plt.ylabel('units of x')
plt.show()
# + [markdown] id="0GnYVpkHvay5"
# A trick here is that by using the function "sig.filtfilt()" we have applied the same filter twice (in this case g = [1 1 1]) first forward and _then backward_ to remove the phase shift we got above! Just to show you what would have happened had we not used sig.filtfilt, look below. In this case, we just apply the exact same 1-1-1 filter to y1 (the red curve above) to get the orange curve.
# + colab={"base_uri": "https://localhost:8080/", "height": 594} executionInfo={"elapsed": 458, "status": "ok", "timestamp": 1645362131676, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="YZgATEdEvay6" outputId="111bfbca-9408-4f9b-8b75-3387f0eff087"
y22 = sig.lfilter(g,np.sum(g),y1) #filtfilt goes forwards and backwards to remove the phase shift
plt.figure()
plt.title('boxcar smoothing');
plt.plot(t,x,'-k',linewidth = LW, label ='original data')
plt.plot(t,y1,'--r',linewidth = LW, label = 'smoothed with 1-1-1' )
plt.plot(t,y2,'--b',linewidth = LW, label = 'smoothed with 1-1-1 twice using filtfilt' )
plt.plot(t,y22,'--',color='darkorange',linewidth = LW, label = 'smoothed with 1-1-1 twice' )
plt.legend(frameon = False, fontsize = LFS)
plt.ylim(0,1.1)
plt.xlabel('time')
plt.ylabel('units of x')
plt.show()
# + [markdown] id="0MmlTQMUvay8"
# We now see that this curve is even more phase shifted. This should teach you two things:
# * know what your software is doing!
# * filtfilt (or forward-backward applications of filters) can be very handy
# + [markdown] id="YJ3BE7fgvay9"
# ### Compute the response functions from the data and from theory
#
# The question is now - what does this smoothing with a 1-1-1 filter do to our data in frequency space? This is known as the "response function". We are going to calculate these response functions in two ways:
# * from theory (see lecture notes)
# * from the ratio of the power spectrum of the smoothed data to the power spectrum of the original raw data
# + executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1645362131676, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="J719CTztvay-"
# calculate the FFTs of the original data, the 1-1-1 smoothed data, and the data smoothed with a 1-1-1 twice
Z_x = np.fft.fft(x)/np.size(x)
Z_y1 = np.fft.fft(y1)/np.size(y1)
Z_y2 = np.fft.fft(y2)/np.size(y2)
# compute the power spectrum by squaring the FFT (and taking only first half)
Ck2_x = np.abs(Z_x[0:int(np.size(Z_x)/2 + 1)])**2
Ck2_y1 = np.abs(Z_y1[0:int(np.size(Z_y1)/2 + 1)])**2
Ck2_y2 = np.abs(Z_y2[0:int(np.size(Z_y2)/2 + 1)])**2
# compute the response function from theory
freq = np.arange(0,np.size(x)/2)/float(np.size(x))
Rg_y1 = 1./3 + (2./3)*np.cos(freq*2.*np.pi)
Rg2_y1 = Rg_y1**2
Rg_y2 = (1./3 + (2./3)*np.cos(freq*2*np.pi))**2
Rg2_y2 = Rg_y2**2
# + [markdown] id="rXusnUv0vazB"
# ### Plot the spectrum after filtering
# + colab={"base_uri": "https://localhost:8080/", "height": 594} executionInfo={"elapsed": 297, "status": "ok", "timestamp": 1645362131968, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="I2912WISvazC" outputId="cfda5318-0ed7-4e30-ef91-cf5b992ef84a"
#%% plot normalized spectrum of the raw data and smoothed data
maxval = np.max(Ck2_x)
plt.figure()
plt.title('Power Spectra of Raw and Smoothed Data')
plt.plot(freq,Ck2_x/maxval,'-k',linewidth = LW, label = 'original data')
plt.plot(freq,Ck2_y1/maxval,'-',color='darkorange',linewidth = LW, \
label = 'data after applying forward 1-1-1')
plt.plot(freq,Ck2_y2/maxval,'-',color='cornflowerblue',linewidth = LW, \
label = 'data after applying forward/backward 1-1-1')
plt.legend(fontsize = LFS, frameon = False)
plt.ylim(0,1)
plt.xlim(0,.5)
plt.xlabel('frequency')
plt.ylabel('normalized power')
plt.show()
# + [markdown] id="nHAwYv6PvazG"
# ### Plot the theoretical squared response functions and compare to our data
# + colab={"base_uri": "https://localhost:8080/", "height": 594} executionInfo={"elapsed": 362, "status": "ok", "timestamp": 1645362132320, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="LvSHNqX4vazH" outputId="6685ef61-e210-4f01-d8ff-8746e9de9dc0"
plt.figure()
plt.title('Squared Response Functions')
plt.plot(freq,Rg2_y1,'-k',linewidth = LW, label = '1-1-1 theoretical response')
plt.plot(freq,Ck2_y1/Ck2_x,'--',color='fuchsia',linewidth = LW, label = '1-1-1 $Ck^2_{output}/Ck^2_{orig}$')
plt.plot(freq,Rg2_y2,'-k',linewidth = LW, label = '1-1-1 x 2 theoretical response')
plt.plot(freq,Ck2_y2/Ck2_x,'--',color = 'cornflowerblue',linewidth = LW, label = '1-1-1 x 2 $Ck^2_{output}/Ck^2_{orig}$')
plt.ylim(0,1)
plt.xlim(0,.5)
plt.legend(fontsize = LFS*1.5, frameon = False)
plt.ylabel('filter power factor')
plt.xlabel('frequency')
plt.show()
# + [markdown] id="Rgeqb862vazJ"
# We see that indeed, the theory and data agree perfectly! Wow! In addition, we see that the additional smoothing moves the power to lower frequencies - as expected from the fact that we are _smoothing_ our data!
# -
# ## Example for lecture
# + executionInfo={"elapsed": 133, "status": "ok", "timestamp": 1645362132442, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiNPVVIWP6XAkP_hwu-8rAxoeeNuk2BMkX5-yuA=s64", "userId": "07585723222468022011"}, "user_tz": 420} id="CwHq_9ApvazL"
fig, axs = plt.subplots(1,2,figsize=(10,3))
ax = axs[0]
ax.plot(freq,np.sqrt(Ck2_x),'-',color='cornflowerblue',linewidth = LW, label='original data')
ax.plot(freq,np.sqrt(Ck2_y2),'-',color='orange',linewidth = LW, label='filtered')
ax.set_ylabel('|C_k|')
ax.set_xlabel('frequency')
ax.set_title('C_k')
ax.legend()
ax.set_ylim(0,.07)
ax.set_xlim(0,.5)
ax = axs[1]
ax.plot(freq,Rg_y2,'-k',linewidth = LW, label='response function')
ax.plot(freq,np.sqrt(Ck2_y2)/np.sqrt(Ck2_x),'--',color='tab:pink',linewidth = LW, label='response function')
ax.set_title('Response Function')
ax.set_xlabel('frequency')
ax.set_ylim(0,1.)
ax.set_xlim(0,.5)
plt.show()
# -
| code/response_function_moving_avg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # Math
#
# Inline maths with inline role: $ x^3+\frac{1+\sqrt{2}}{\pi} $
#
# Inline maths using dollar signs (not supported yet): $x^3+frac{1+sqrt{2}}{pi}$ as the
# backslashes are removed.
#
# $$
# x^3+\frac{1+\sqrt{2}}{\pi}
# $$
#
# check math with some more advanced LaTeX, previously reported as an issue.
#
# $$
# \mathbb P\{z = v \mid x \}
# = \begin{cases}
# f_0(v) & \mbox{if } x = x_0, \\
# f_1(v) & \mbox{if } x = x_1
# \end{cases}
# $$
#
# and labeled test cases
#
#
# <a id='equation-firsteq'></a>
# $$
# \mathbb P\{z = v \mid x \}
# = \begin{cases}
# f_0(v) & \mbox{if } x = x_0, \\
# f_1(v) & \mbox{if } x = x_1
# \end{cases} \tag{1}
# $$
# # Further Inline
#
# A continuation Ramsey planner at $ t \geq 1 $ takes
# $ (x_{t-1}, s_{t-1}) = (x_-, s_-) $ as given and before
# $ s $ is realized chooses
# $ (n_t(s_t), x_t(s_t)) = (n(s), x(s)) $ for $ s \in {\cal S} $
# # Referenced Math
#
# Simple test case with reference in text
#
#
# <a id='equation-test'></a>
# $$
# v = p + \beta v \tag{2}
# $$
#
# this is a reference to [(2)](#equation-test) which is the above equation
| tests/ipynb/math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cupy as cp
import cusignal
from scipy import signal
import numpy as np
# ### Cross Power Spectral Density
cx = np.random.rand(int(1e8))
cy = np.random.rand(int(1e8))
fs = int(1e6)
# %%time
ccsd = signal.csd(cx, cy, fs, nperseg=1024)
gx = cp.random.rand(int(1e8))
gy = cp.random.rand(int(1e8))
fs = int(1e6)
# %%time
gcsd = cusignal.csd(gx, gy, fs, nperseg=1024)
# ### Periodogram
csig = np.random.rand(int(1e8))
fs = int(1e6)
# %%time
f, Pxx_spec = signal.periodogram(csig, fs, 'flattop', scaling='spectrum')
gsig = cp.random.rand(int(1e8))
fs = int(1e6)
# %%time
gf, gPxx_spec = cusignal.periodogram(gsig, fs, 'flattop', scaling='spectrum')
# ### Welch PSD
csig = np.random.rand(int(1e8))
fs = int(1e6)
# %%time
cf, cPxx_spec = signal.welch(csig, fs, nperseg=1024)
gsig = cp.random.rand(int(1e8))
fs = int(1e6)
# %%time
gf, gPxx_spec = cusignal.welch(gsig, fs, nperseg=1024)
# ### Spectrogram
csig = np.random.rand(int(1e8))
fs = int(1e6)
# %%time
cf, ct, cPxx_spec = signal.spectrogram(csig, fs)
gsig = cp.random.rand(int(1e8))
fs = int(1e6)
# %%time
gf, gt, gPxx_spec = cusignal.spectrogram(gsig, fs)
# ### Coherence
cx = np.random.rand(int(1e8))
cy = np.random.rand(int(1e8))
fs = int(1e6)
# %%time
cf, cCxy = signal.coherence(cx, cy, fs, nperseg=1024)
gx = cp.random.rand(int(1e8))
gy = cp.random.rand(int(1e8))
fs = int(1e6)
# %%time
gf, gCxy = cusignal.coherence(gx, gy, fs, nperseg=1024)
# ### Short Time Fourier Transform
cx = np.random.rand(int(1e8))
fs = int(1e6)
# %%time
cf, ct, cZxx = signal.stft(cx, fs, nperseg=1000)
gx = cp.random.rand(int(1e8))
fs = int(1e6)
# %%time
gf, gt, gZxx = cusignal.stft(gx, fs, nperseg=1024)
| notebooks/spectral_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Google maps distances
# Get an API key from Google [here](https://console.developers.google.com/apis/).
now = datetime.now()
gmaps = googlemaps.Client(key='')
gmaps.distance_matrix("1 n ogden ave chicago illinois usa",
"10 s. kedzie chicago illinois usa",
mode="transit",
departure_time=now)['rows'][0]['elements'][0]['distance']['value']
# After building a loop to get the distances between schools on a map, I realized I was attempting to hammer Google's API server 7 million times. That'll cost ya. So I deleted the logic and I'll simply use a competely fake distance measure based on the row indices and the cluster indices.
| resolver/google_maps_distances.ipynb |