code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="vPOn1ABLheho" colab_type="code" colab={}
import plotly.graph_objects as go
from collections import Counter
# + id="iF8lKtCvhKzz" colab_type="code" colab={}
text = """Pēc ideāliem cenšas lielie gari,
Bet dzīvē ieņemt vietu pirmie
Tie neiespēj, tos nomāc maizes kari,
Tos nomāc aizspriedumi sirmie.
Virs zemes nav taisnības, dūrei tik spēks,
Kas varmākām skādi dar, nosaukts tiek grēks.
Par tiesnešiem cienīti blēži sēž
Un godīgie ādu nost citiem plēš.
Un cienīgs tēvs, zaglis, teic sprediķus;
"Tik pacieties, debesīs labāki būs!"
Virs zemes nav laimes, tik zvēru pulks bļauj,
Viens otram iz mutes tie maizi sev rauj,
Un priecīgs ikkatris, kad vēders tik pilns,
Kad bērni ir veseli, dzīvoklis silts.
Un glaimojot salkušie rakstnieki sauc:
"Cik praktiska tauta! Tai cerību daudz."
Jā, cerību gan. Vēl nosirmos laiks,
Un redzams būs strādnieku moku pilns vaigs,
Vēl liekēži slinkos un godīgi krāps,
Pēc debesu viltības ļaudis vēl slāps,
Zem kājām vēl varmākas taisnību mīs,
Par kaujamiem lopiem vēl cilvēkus dzīs,
Un muļķīgas dzejas vēl dzejnieki kals
Un vaimanās paši, kad salks un sals.
Ar šīs zemes varenajiem kopā tu varēji būt
Un valdīt, un kārumu baudīt, un līgsmot ik dienas,
Bet cilvēces bēdas un trūkumus sirds tevim jūt,
Un tumsības slogus, un varmāku nastas ikvienas.
Un, tēvijas neatzīts, svešumā dodies tu tāļi
Un sludini taisnību, māci, ka cilvēki brāļi,
Ka pasaules mantas mums mierināt nespēj prāta,
Ja ticības trūkst un mīlestīb' sirdīs nav krāta.
Tu priekus tiem nesi, kas tumsībā, grūtībā smaka,
Tu vārguļu sagrauztām sirdīm bij' dzīvības aka,
Tu cilvēci sildīji garīgās verdzības salnā,
Bet atmaksu cilvēce deva tev - Golgatas kalnā."""
# + id="vLE9UcHghj_g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="79c40689-58f2-49bc-e6f1-28cbf1499764"
max_items = 45
w_count= Counter(text.split())
w_count_top = w_count.most_common(max_items)
words = [el[0] for el in w_count_top]
weights = [el[1] for el in w_count_top]
size = [len(el[0]) for el in w_count_top]
fig = go.Figure(data=[go.Scatter(
x=words,
y=weights,
mode='markers',
marker=dict(
size=size,
sizemode='area',
sizeref=2.*max(size)/(40.**2),
sizemin=1,
color=size,
showscale=True
)
)])
fig.update_layout(
title='Vārdu pielietojums Veidenbauma dzejā',
yaxis=dict(
title='Absolūtais biežums'
),
xaxis=dict(
title='Vārdi'
)
)
fig.show()
| visualization/Word_Frequency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# No notebook anterior, nós aprendemos intuitivamente como o perceptron aprende. De maneira geral, nós vamos atualizando os pesos e o bias sempre buscando diminuir uma função de custo. Nesse notebook, nós vamos ver como esse aprendizado realmente acontence, tanto na teoria quanto na prática. Também utilizaremos o Perceptron para resolver problemas de classificação e regressão.
#
# __Objetivos__:
#
# - Implementar o perceptron e seu modelo de aprendizado em Python puro e Numpy
# - Utilizar o perceptron para regressão e classificação
# + [markdown] heading_collapsed=true
# # Sumário
# + [markdown] hidden=true
# [Introdução](#Introdução)
# - [Regra de Aprendizado do Perceptron](#Regra-de-Aprendizado-do-Perceptron)
# - [Pseudo-algoritmo do Perceptron](#Pseudo-algoritmo-do-Perceptron)
#
# [Classificação](#Classificação)
# - [Porta AND/OR](#Porta-AND/OR)
# - [Exercício de Classificação](#Exerc%C3%ADcio-de-Classificação)
#
# [Regressão](#Regressão)
# - [Exercício de Regressão](#Exerc%C3%ADcio-de-Regressão)
#
# [Referências](#Referências)
# -
# # Imports e Configurações
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from random import random
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import MinMaxScaler
from sklearn.datasets.samples_generator import make_blobs
# %matplotlib inline
# + [markdown] heading_collapsed=true
# # Introdução
# + [markdown] hidden=true
# O tipo mais básico de Rede Neural Artificial é formada por apenas um neurônio, o __Perceptron__. Inicialmente, o Perceptron foi projetado para ser um __classificador binário linear__ responsável por mapear uma ou mais entradas em uma saída desejada. Porém, também podemos utilizá-lo para resolver problemas de __regressão linear__. Ele foi projetado em 1957 por <NAME>.
#
# O perceptron é formado por:
#
# <img src='images/perceptron.png' width='350'>
#
# - __entradas__ $x_1,...,x_D$: representam os atributos dos seus dados com dimensionalidade $D$. O Perceptron aceita qualquer tamanho de entrada, porém a saída é sempre apenas um valor.
# - __junção aditiva__ $\sum$: também chamada de _função agregadora_, nada mais é que a soma ponderada das entradas com os __pesos__ ($w_1,...,w_D)$. Em geral, o resultado é somado com um __bias__ $b$, responsável por deslocar o resultado do somatório. A junção aditiva é descrita pela seguinte fórmula:
#
# $$\sum_i^D{x_iw_i} + b$$
#
# - __função de ativação__ $f$: utilizada para mapear o resultado da junção aditiva em uma saída esperada. Mais detalhes abaixo.
#
# Logo, o Perceptron é representado pela seguinte fórmula matemática:
#
# $$\widehat{y}_i = f(\sum_i^D{x_iw_i} + b)$$
#
# Onde:
#
# - $D$: representa a dimensionalidade das amostras, ou seja, a quantidade de atributos de cada amostra.
# - $x_i$: representam os atributos de uma amostra que servem de entrada para o Perceptron.
# - $w_i$: representam os __pesos sinápticos__ que ponderam as entradas.
# - $b$: representa o __bias__, responsável por deslocar a fronteira de decisão além da origem e não depende de nenhum valor de entrada. Repare que o bias encontra-se fora do somatório.
# - $f$: __função de ativação__. Quando a função de ativação é linear, ou seja, nenhuma transformação é aplicada no resultado da junção aditiva, o Perceptron atua como um __Regressor Linear__. Se precisamos efetuar uma __Classificação binária__, devemos utilizar a função _step_ (também conhecida como _função degrau_) para mapear a saída em um valor discreto (0 ou 1):
#
# $$f = \begin{cases}1 & se \ wx+b > 0\\0 & caso \ contr\acute ario\end{cases}$$
#
# - $\widehat{y}$: representa a saída do Perceptron (o valor predito).
#
# __Observações importantes__:
#
# - O Perceptron não faz __Classificação Multiclasse__.
# - __A atualização dos pesos é *online*, ou seja, efetuada amostra a amostra__ utilizando uma fórmula pré-definida que veremos na seção a seguir.
# -
# ## Regra de Aprendizado do Perceptron
# O Perceptron tem sua própria forma de aprendizado conforme definido no seu artigo original. Na verdade, a fórmula para atualização dos pesos e bias é bem simples:
#
# $$w_i = w_i + \lambda(y_i - \widehat{y}_i)x_i$$
# <br>
# $$b_i = b_i + \lambda(y_i - \widehat{y}_i)$$
#
# Onde $\lambda$ é a __taxa de aprendizagem__ (___learning rate___).
#
# Repare que $y_i - \widehat{y}_i$ significa calcular a diferença entre o valor esperado ($y_i$) e o valor predito ($\widehat{y}_i$). Supondo que estamos fazendo __classificação binária__ de uma amostra $(x_i, y_i)$. Nesse caso, teremos duas possibilidades:
# - __O valor esperado é $y_i = \widehat{y}_i$__, ou seja, a saída do Perceptron (após a função de ativação _step_) é __igual__ a saída esperada. Nesse caso, __a diferença $y_i - \widehat{y}_i = 0$ e não haverá atualização de pesos__.
# - __O valor esperado é $y_i \neq \widehat{y}_i$__, ou seja, a saída do Perceptron (após a função de ativação _step_) é __diferente__ da saída esperada. Nesse caso, __a atualização dos pesos será dada pela diferença $y_i - \widehat{y}_i$__. Repare que:
# - quando essa diferença é __negativa__ (ou seja, $y_i = 0$ e $\widehat{y}_i = 1$), __os pesos tendem a diminuir__.
# - quando essa diferença é __positiva__ (ou seja, $y_i = 1$ e $\widehat{y}_i = 0$), __os pesos tendem a aumentar__.
# ## Pseudo-algoritmo do Perceptron
# 1. Inicialize os pesos $w$ e o bias $b$
# 2. Para cada amostra $(x_n, y_n)$ do nosso banco:
# 1. Calcule $\widehat{y} = f(\sum_i^D{x_iw_i} + b)$, onde $f$ é a __função _step_ para classificação__ e __linear no caso da regressão__
# 2. Calcule o $erro = y_n - \widehat{y}$
# 3. Atualize os pesos $w_i = w_i + \lambda*erro*x_i$
# 4. Atualize o bias $b_i = b_i + \lambda*erro$
# 3. Repita o passo 2 por N vezes ou até que alguma medida de custo para o $erro$ seja menor que um valor pré-determinado.
#
# Repare, como dito lá em cima, que __a atualização dos pesos e bias é feito a cada amostra__, e não somente após ver todas as amostras do banco.
# # Classificação
# ## Porta AND/OR
# +
x = np.array([[0, 0], [0, 1], [1, 0], [1, 1]])
#y = np.array([0, 1, 1, 1]) # porta OR
y = np.array([0, 0, 0, 1]).T # porta AND
print(x.shape, y.shape)
# + [markdown] heading_collapsed=true
# ### Python
# + hidden=true
D = x.shape[1] #D pesos possui a quantidade de colunas (atributos) de x == qtde entradas perceptron
w = [2*random()-1 for i in range(D)] #D pesos com valores aleatórios entre -1 e 1
b = 2*random()-1 #b fica entre -1 e 1
learning_rate = 1.0
for step in range(101):
cost = 0 #averigua se o perceptron está aprendendo
#indice n indica real
for x_n,y_n in zip(x,y):
y_pred = sum([x_i*w_i for x_i, w_i in zip(x_n,w)])+b
y_pred = 1 if y_pred > 0 else 0
error = y_n - y_pred
w = [w_i + learning_rate*error*x_i for x_i, w_i in zip(x_n,w)]
b = b + learning_rate*error
cost += error**2
if step % 10 == 0:
print('step{0}:{1}'.format(step,cost))
print('w=',w)
print('bias=',b)
print('y_pred= {0}'.format(np.dot(x,np.array(w))+b))#saída bruta, sem passar pela f. de ativação
# +
#resultado acima:
#y_pred= [-3.90047089 -1.99832769 -0.96378621 0.93835698]
#se passasse pela função de ativação, o resultado seria
#[0,0,0,1]
#OBS: cost = 3 => errou 3 de 4
# + [markdown] heading_collapsed=true
# ### Numpy
# + hidden=true
D = x.shape[1]
w = 2*np.random.random(size=D)-1
b = 2*np.random.random()-1
learning_rate = 0.1
for step in range(101):
cost = 0
for x_n,y_n in zip(x,y):
y_pred = np.dot(x_n,w)+b
y_pred = np.where(y_pred>0,1,0)
error = y_n - y_pred
w = w + learning_rate*np.dot(error,x_n)
b = b + learning_rate*error
cost += error**2
if step % 10 == 0:
print('step{0}:{1}'.format(step,cost))
print('w=',w)
print('bias=',b)
print('y_pred= {0}'.format(np.dot(x,np.array(w))+b))
# -
# ## Exercício de Classificação
# +
x, y = make_blobs(n_samples=100, n_features=2, centers=2, random_state=1234)
print(x.shape, y.shape)
plt.scatter(x[:,0], x[:,1], c=y.ravel(), cmap='bwr')
# -
def plot_linear_classifier(x, y, w, b):
x1_min, x1_max = x[:,0].min(), x[:,0].max()
x2_min, x2_max = x[:,1].min(), x[:,1].max()
x1, x2 = np.meshgrid(np.linspace(x1_min-1, x1_max+1,100), np.linspace(x2_min-1, x2_max+1, 100))
x_mesh = np.array([x1.ravel(), x2.ravel()]).T
plt.scatter(x[:,0], x[:,1], c=y.ravel(), cmap='bwr')
y_mesh = np.dot(x_mesh, np.array(w).reshape(1, -1).T) + b
y_mesh = np.where(y_mesh <= 0, 0, 1)
plt.contourf(x1, x2, y_mesh.reshape(x1.shape), cmap='bwr', alpha=0.5)
plt.xlim(x1_min-1, x1_max+1)
plt.ylim(x2_min-1, x2_max+1)
# ### Python
# +
D = x.shape[1] #D pesos possui a quantidade de colunas (atributos) de x == qtde entradas perceptron
w = [2*random()-1 for i in range(D)] #D pesos com valores aleatórios entre -1 e 1
b = 2*random()-1 #b fica entre -1 e 1
learning_rate = 0.01
for step in range(101):
cost = 0 #averigua se o perceptron está aprendendo
#indice n indica real
for x_n,y_n in zip(x,y):
y_pred = sum([x_i*w_i for x_i, w_i in zip(x_n,w)])+b
y_pred = 1 if y_pred > 0 else 0
error = y_n - y_pred
w = [w_i + learning_rate*error*x_i for x_i, w_i in zip(x_n,w)]
b = b + learning_rate*error
cost += error**2
if step % 10 == 0:
print('step{0}:{1}'.format(step,cost))
print('w=',w)
print('bias=',b)
#print('y_pred= {0}'.format(np.dot(x,np.array(w))+b))
plot_linear_classifier(x,y,w,b)
# -
# ### Numpy
# +
D = x.shape[1]
w = 2*np.random.random(size=D)-1
b = 2*np.random.random()-1
learning_rate = 0.1
for step in range(101):
cost = 0
for x_n,y_n in zip(x,y):
y_pred = np.dot(x_n,w)+b
y_pred = np.where(y_pred>0,1,0)
error = y_n - y_pred
w = w + learning_rate*np.dot(error,x_n)
b = b + learning_rate*error
cost += error**2
if step % 10 == 0:
print('step{0}:{1}'.format(step,cost))
print('w=',w)
print('bias=',b)
plot_linear_classifier(x,y,w,b)
# + [markdown] heading_collapsed=true
# # Regressão
# -
# Para transformar o Perceptron em um __regressor linear__, só o que temos de fazer é __remover a função de ativação _step___, transformando-a em uma função de ativação linear.
#
# Apesar dessa modificação, __a fórmula de atualização dos pesos não sofre nenhuma alteração__.
#
# Vamos, então, implementar nosso perceptron para classificação em Python, Numpy, Keras e TensorFlow:
# + hidden=true
df = pd.read_csv('data/medidas.csv')
print(df.shape)
df.head(10)
# + hidden=true
x = df.Altura.values
y = df.Peso.values
plt.figure()
plt.scatter(x, y)
plt.xlabel('Altura')
plt.ylabel('Peso')
# + hidden=true
print(x.shape, y.shape)
# + hidden=true
x = x.reshape(-1, 1)
print(x.shape, y.shape) #x vira matriz 100x1, y não precisa disso aqui
# + [markdown] heading_collapsed=true hidden=true
# ### Python
# + [markdown] hidden=true
# __Exercício__: tentar estimar as learning_rates de **w** e __b__. Elas são diferentes por que nossos dados não estão na mesma escala!
# + hidden=true
D = x.shape[1]
w = [2*random() - 1 for i in range(D)]
b = 2*random() - 1
for step in range(10001):
cost = 0
for x_n, y_n in zip(x, y):
# qual linha devemos remover para transformar o Perceptron num regressor?
y_pred = sum([x_i*w_i for x_i, w_i in zip(x_n, w)]) + b
#y_pred = 1 if y_pred > 0 else 0
#comentando a linha acima o perceptron vira regressor
error = y_n - y_pred
w = [w_i + 0.0000001*error*x_i for x_i, w_i in zip(x_n, w)]
b = b + 0.001*error
cost += error**2
if step%1000 == 0:
print('step {0}: {1}'.format(step, cost))
print('w: ', w)
print('b: ', b)
# + [markdown] hidden=true
# ### Numpy
# + hidden=true
D = x.shape[1]
w = 2*np.random.random(size=D)-1
b = 2*np.random.random()-1
for step in range(10001):
cost = 0
for x_n, y_n in zip(x, y):
# qual linha devemos remover para transformar o Perceptron num regressor?
y_pred = np.dot(x_n, w) + b
#y_pred = np.where(y_pred > 0, 1, 0)
error = y_n - y_pred
w = w + 1e-7*np.dot(error, x_n)
b = b + 1e-2*error
cost += error**2
if step%1000 == 0:
print('step {0}: {1}'.format(step, cost))
print('w: ', w)
print('b: ', b)
# + [markdown] hidden=true
# ### Numpy com Pré-processamento
# + hidden=true
minmax = MinMaxScaler(feature_range=(-1,1))
x = minmax.fit_transform(x.astype(np.float64))
print(x.min(), x.max())
# +
reg = LinearRegression()
reg.fit(x,y)#todas as colunas(atributos) da entrada precisam estar normalizados
#sobre os mesmos valores(a saída(y) não precisa disso)
print('w: ', reg.coef_)
print('b: ', reg.intercept_)
# + hidden=true
D = x.shape[1]
w = 2*np.random.random(size=D)-1
b = 2*np.random.random()-1
learning_rate = 0.001 # <- tente estimar a learning_rate
for step in range(1001):
cost = 0
for x_n, y_n in zip(x, y):
y_pred = np.dot(x_n, w) + b
error = y_n - y_pred
w = w + learning_rate*np.dot(error, x_n)
b = b + learning_rate*error
cost += error**2
if step%100 == 0:
print('step {0}: {1}'.format(step, cost))
print('w: ', w)
print('b: ', b)
# + [markdown] hidden=true
# ## Exercício de Regressão
# + hidden=true
df = pd.read_csv('data/notas.csv')
print(df.shape)
df.head(10)
# +
plt.figure(figsize=(20, 4))
plt.subplot(1, 3, 1)
plt.scatter(df.prova1.values, df.final.values)
plt.xlabel('Prova 1')
plt.ylabel('Final')
plt.subplot(1, 3, 2)
plt.scatter(df.prova2.values, df.final.values)
plt.xlabel('Prova 2')
plt.ylabel('Final')
plt.subplot(1, 3, 3)
plt.scatter(df.prova3.values, df.final.values)
plt.xlabel('Prova 3')
plt.ylabel('Final')
# + hidden=true
x = df[['prova1', 'prova2', 'prova3']].values
y = df['final'].values
print(x.shape, y.shape)
# + hidden=true
minmax = MinMaxScaler(feature_range=(-1,1))
x = minmax.fit_transform(x.astype(np.float64))
# + hidden=true
reg = LinearRegression()
reg.fit(x, y)
print('w: ', reg.coef_)
print('b: ', reg.intercept_)
# + [markdown] heading_collapsed=true hidden=true
# ### Python
# + hidden=true
D = x.shape[1]
w = [2*random() - 1 for i in range(D)]
b = 2*random() - 1
learning_rate = 0.01 # <- tente estimar a learning_rate
for step in range(2000): # <- tente estimar o número de passos
cost = 0
for x_n, y_n in zip(x, y):
y_pred = sum([x_i*w_i for x_i, w_i in zip(x_n, w)]) + b
error = y_n - y_pred
w = [w_i + learning_rate*error*x_i for x_i, w_i in zip(x_n, w)]
b = b + learning_rate*error
cost += error**2
if step%200 == 0:
print('step {0}: {1}'.format(step, cost))
print('w: ', w)
print('b: ', b)
# + [markdown] heading_collapsed=true hidden=true
# ### Numpy
# + hidden=true
D = x.shape[1]
w = 2*np.random.random(size=D)-1
b = 2*np.random.random()-1
learning_rate = 0.01 # <- tente estimar a learning_rate
for step in range(1001): # <- tente estimar o número de passos
cost = 0
for x_n, y_n in zip(x, y):
y_pred = np.dot(x_n, w) + b
error = y_n - y_pred
w = w + learning_rate*np.dot(error, x_n)
b = b + learning_rate*error
cost += error**2
if step%200 == 0:
print('step {0}: {1}'.format(step, cost))
print('w: ', w)
print('b: ', b)
# -
# # Referências
# - [<NAME> Perceptron](http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.335.3398&rep=rep1&type=pdf)
| Perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (tensorflow)
# language: python
# name: tensorflow
# ---
# # Introduction to Datamining
# #### Instructor : <NAME>, School of Engineering, [Azad U, South Tehran Branch](http://www/azad/ac/ir)
#
# # Modules
# 1. [Setting Up Environment](install.ipynb)
# 2. [introduction to Python](introduction_to_python.ipynb)
# 3. Lists
# 4. [Numpy](numpy_intro.ipynb)
# 5. Pandas
# 6. Statistics and Probability
# 7. Data Preprocessing
# 8. Frequent Pattern Mining
# 9. Classification
# 10. Clustering
# 11. Deep Learning and Neural Networks
#
| course_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This notebook serves as a refresher with some basic Python code and functions
# ### 1) Define a variable called x, with initial value of 5. multiply by 2 four times and print the value each time
x = 5
for i in range(4):
x = x*2
print(i, x)
# ### 2) Define a list
p = [9, 4, -5, 0, 10.9]
# Get length of list
len(p)
# index of a specific element
p.index(0)
# first element in list
p[0]
print(sum(p))
# ### 3) Create a numpy array
import numpy as np
a = np.array([5, -19, 30, 10])
# Get first element
a[0]
# Get last element
a[-1]
# Get first 3 elements
print(a[0:3])
print(a[:3])
# Get size of the array
a.shape
# ### 4) Define a dictionary that stores the age of three students.
# ### Mark: 26, Camilla: 23, Jason: 30
students = {'Mark':26,
'Camilla': 23,
'Jason':30}
students['Mark']
students.keys()
# ### 5) Create a square function
def square_number(x):
x2 = x**2
return x2
x_squared = square_number(5)
print(x_squared)
# ### 6) List comprehension
# add 2 to every element in the numpy array
numbers_array = np.arange(10, 21)
print("original array:", numbers_array)
number_array_plus_two = [x+2 for x in numbers_array]
print("array plus 2:", number_array_plus_two)
# select only even numbers
even_numbers =[x for x in numbers_array if x%2==0]
print(even_numbers)
# ### 7) Random numbers
np.random.seed(42)
rand_number = np.random.random(size =5)
print(rand_number)
np.random.seed(42)
rand_number2 = np.random.random(size =5)
print(rand_number2)
# ### 8) Matplotlib
# +
import matplotlib.pyplot as plt
# Data for plotting
t = np.arange(0.0, 5.0, 0.01)
s = 1 + np.cos(2 * np.pi * t)
fig, ax = plt.subplots()
ax.plot(t, s)
ax.set(xlabel='time (s)', ylabel='voltage (mV)',
title='Voltage vs time plot')
ax.grid()
fig.savefig("voltage_plot.png")
plt.show()
# -
# ### Plotting with seaborn
# Inspired from: https://github.com/knathanieltucker/seaborn-weird-parts
import seaborn as sns
sns.set(font_scale=1)
tips = sns.load_dataset("tips")
tips.head()
ax = sns.boxplot(x="time", y="tip", data=tips)
plt.show()
ax = sns.boxplot(x="time", y="tip", hue="smoker", data=tips)
plt.show()
fig, ax = plt.subplots(1,2, figsize=(15,7))
ax[0].scatter(tips['tip'], tips['total_bill'], color=sns.color_palette('Blues')[-1])
ax[0].set(title='Scatter plot of bills and tips', ylabel='Total bill', xlabel='Tip')
sns.boxplot(x='day', y='total_bill', data=tips, palette='Blues', ax=ax[1])
ax[1].set(title='Total bill', ylabel='Total bill', xlabel='Day of the week')
plt.show()
fig.savefig('combined_plots.png', format='png', dpi=500)
fig, ax = plt.subplots(1,1, figsize=(12,7))
ax = sns.barplot(x="day", y="total_bill", hue="sex", data=tips)
ax.set(xlabel='Day of Week', ylabel='Mean Total bill')
fig.savefig("mean_total_bill.pdf", bbox_inches='tight')
plt.show()
# +
# Plot of Gaussian PDFs
# https://en.wikipedia.org/wiki/Normal_distribution
from math import exp, pi, sqrt
def normal_pdf(x, mu=0, sigma=1.0):
'''Gaussian PDF'''
sqrt_two_pi = sqrt(2 * pi)
return exp(-(x - mu) ** 2 / 2 / sigma ** 2) / (sqrt_two_pi * sigma)
xs = [x / 10.0 for x in range(-50, 50)]
plt.plot(xs, [normal_pdf(x, sigma=1) for x in xs], '-', label='mu=0, sigma=1', linewidth=5)
plt.plot(xs, [normal_pdf(x, sigma=2) for x in xs], '--', label='mu=0, sigma=2', linewidth=5)
plt.plot(xs, [normal_pdf(x, sigma=0.5) for x in xs], ':', label='mu=0, sigma=0.5', linewidth=5)
plt.plot(xs, [normal_pdf(x, mu=-1) for x in xs], '-.', label='mu=-1, sigma=1')
plt.legend()
plt.title("Various Normal PDFs")
plt.show()
# -
# ## Exercises
# 1- Create a list with numbers from 1 to 10 (inclusive). Determine the mean of the list using two methods. Hint: numpy has a built in function
#
# +
# Code here
# -
# 2- Define a function called "expectation" that returns the mathematical expectation of variable $x$ (given as an array) and its probabilities (given as an array). Example: x = [0, 1], x_prob = [0.4, 0.6]
#
# +
# Code here
# -
# 3- Use the function "expectation" inside another function that calculates $\mathbb E(X)$, where $X\sim$ Bernoulli $(p)$. The only input of the function is $p$.
# https://en.wikipedia.org/wiki/Bernoulli_distribution
#
# +
# Code here
# -
# 4- Create a plot of two variables x and y. x ranges from 1 to 40 (step 1), and y = 8 + 3*x + $\epsilon$, where $\epsilon$ is random noise. Hint: Use a scatter plot and make sure $\epsilon$ is the right size
# +
# Code here
# -
# 5- Plot the Gaussian CDFs of the above plotted PDFs.
# +
# Code here
# -
| notebooks/1_Python_101.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature engineering
# This notebook will teach you how to extract feature values using `revscoring`'s built-in feature library as well as to build your own features.
# ## Set up the feature extractor
# This line constructs a "feature extractor" that uses Wikipedia's API. We'll need to use it later, so we'll construct it first.
import sys
sys.path.append("/usr/local/lib/python3.4/dist-packages/")
sys.path.append("/usr/local/lib/python3.4/dist-packages/revscoring/")
sys.path.append("/usr/local/lib/python3.4/dist-packages/more_itertools/")
sys.path.append("/usr/local/lib/python3.4/dist-packages/deltas/")
# !sudo pip3 install dependencies deltas
# +
from revscoring.extractors import api
import mwapi
extractor = api.Extractor(mwapi.Session("https://en.wikipedia.org",
user_agent="Revscoring feature demo <EMAIL>"))
# -
# ## Extract features
# The following line demonstrates a simple feature extraction. We'll extract two features: `wikitext.revision.chars`, the number of characters added; and `wikitext.revision.diff.chars_added`, the number of characters in the entire revision. Note that we wrap the call in a list() because it returns a generator.
# + active=""
# from revscoring.features import wikitext
# list(extractor.extract(123456789, [wikitext.revision.chars,
# wikitext.revision.diff.chars_added]))
# -
# ## Defining a custom feature
# The next block defines a new feature and sets the dependencies to be the two features we just extracted. This feature represents the proportion of characters in the current version of the page that the current edit is responsible for adding.
# +
from revscoring import Feature
chars_added_ratio_explicit = Feature(
"chars_added_ratio_explicit",
lambda a,c: a/max(c, 1), # Prevents divide by zero
depends_on=[wikitext.revision.diff.chars_added,
wikitext.revision.chars],
returns=float)
list(extractor.extract(123456789, [chars_added_ratio_explicit]))
# -
# There's easier ways that we can do this though. `revscoring.Feature` overloads simple mathematical operators to allow you to do math with features and get a feature returned. `revscoring.features.modifiers` contains a set of basic functions that do the same. This code roughly corresponds to what's going on above.
# +
from revscoring.features import modifiers
chars_added_ratio_implicit = (wikitext.revision.diff.chars_added /
modifiers.max(wikitext.revision.chars, 1))
list(extractor.extract(123456789, [chars_added_ratio_implicit]))
# -
# While the *implicit* pattern is quicker and easier than the *explicit* pattern, it's name can not be customized.
chars_added_ratio_explicit, chars_added_ratio_implicit
# ## Extracting datasources
# There's a also a set of `revscoring.Datasource`'s that are part of the dependency injection system. These "datasources" represent the data needed for feature generation. We can extract them just like `revscoring.Feature`'s.
list(extractor.extract(662953550, [wikitext.revision.diff.datasources.segments_added,
wikitext.revision.diff.datasources.segments_removed]))
# OK. Let's define a new feature for counting the number of templates added. I'll make use of mwparserfromhell to do this. See [the docs](http://mwparserfromhell.readthedocs.org/en/latest/).
# +
import mwparserfromhell as mwp
templates_added = Feature("templates_added",
lambda add_segments: sum(len(mwp.parse(s).filter_templates()) > 0 for s in add_segments),
depends_on=[wikitext.revision.diff.datasources.segments_added],
returns=int)
list(extractor.extract(662953550, [templates_added]))
# -
# ## Debugging
# There's some facilities in place to help you make sense of issues when they arise. The most important is the draw function.
from revscoring.dependencies import draw
print(draw(templates_added))
# In the tree structure above, you can see how our new feature depends on `wikitext.revision.diff.segments_added` which depends on `wikitext.revision.diff.operations` which depends (as you might imagine) on the current and parent revision. Some features can get quite complicated.
print(draw(wikitext.revision.diff.number_prop_delta_sum))
# The dependency injection system will only solve a unique dependency once for a given tree. So, even though `<revision.parent.text>` appears twice above, it will only be extracted once and then cached. This allows for multiple features to *share* large sections of their dependency trees -- and therefor minimize resource usage.
# ## Errors during extraction
#
# A `revscoring.Extractor` should be expected to throw an exception if it cannot find a missing resource during extraction. These messages are intented to clearly convey what went wrong.
try:
list(extractor.extract(2, [wikitext.revision.diff.words_added]))
except Exception as e:
print(e)
try:
list(extractor.extract(262721924, [wikitext.revision.diff.words_added]))
except Exception as e:
print(e)
from revscoring.features import revision_oriented
try:
list(extractor.extract(172665816, [revision_oriented.revision.comment_matches("foo")]))
except Exception as e:
print(e)
from revscoring.features import temporal
try:
list(extractor.extract(591839757, [revision_oriented.revision.user.text_matches("foo")]))
except Exception as e:
print(e)
| notebooks/.ipynb_checkpoints/feature_engineering-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="copyright"
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title:generic,gcp"
# # E2E ML on GCP: MLOps stage 2 : experimentation
#
# <table align="left">
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/community/ml_ops/stage2/mlops_experimentation.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# <td>
# <a href="https://console.cloud.google.com/ai/platform/notebooks/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/community/ml_ops/stage2/mlops_experimentation.ipynb">
# Open in Google Cloud Notebooks
# </a>
# </td>
# </table>
# <br/><br/><br/>
# + [markdown] id="overview:mlops"
# ## Overview
#
#
# This tutorial demonstrates how to use Vertex AI for E2E MLOps on Google Cloud in production. This tutorial covers stage 2 : experimentation.
# + [markdown] id="dataset:bq,chicago,lbn"
# ### Dataset
#
# The dataset used for this tutorial is the [Chicago Taxi](https://www.kaggle.com/chicago/chicago-taxi-trips-bq). The version of the dataset you will use in this tutorial is stored in a public BigQuery table. The trained model predicts whether someone would leave a tip for a taxi fare.
# + [markdown] id="objective:mlops,stage2,tabular"
# ### Objective
#
# In this tutorial, you create a MLOps stage 2: experimentation process.
#
# This tutorial uses the following Vertex AI:
#
# - `Vertex AI Datasets`
# - `Vertex AI Models`
# - `Vertex AI AutoML`
# - `Vertex AI Training`
# - `Vertex AI TensorBoard`
# - `Vertex AI Vizier`
# - `Vertex AI Batch Prediction`
#
# The steps performed include:
#
# - Review the `Dataset` resource created during stage 1.
# - Train an AutoML tabular binary classifier model in the background.
# - Build the experimental model architecture.
# - Construct a custom training package for the `Dataset` resource.
# - Test the custom training package locally.
# - Test the custom training package in the cloud with Vertex AI Training.
# - Hyperparameter tune the model training with Vertex AI Vizier.
# - Train the custom model with Vertex AI Training.
# - Add a serving function for online/batch prediction to the custom model.
# - Test the custom model with the serving function.
# - Evaluate the custom model using Vertex AI Batch Prediction
# - Wait for the AutoML training job to complete.
# - Evaluate the AutoML model using Vertex AI Batch Prediction with the same evaluation slices as the custom model.
# - Set the evaluation results of the AutoML model as the baseline.
# - If the evaluation of the custom model is below baseline, continue to experiment with the custom model.
# - If the evaluation of the custom model is above baseline, save the model as the first best model.
# + [markdown] id="recommendation:mlops,stage2,tabular"
# ### Recommendations
#
# When doing E2E MLOps on Google Cloud for experimentation, the following best practices with structured (tabular) data are recommended:
#
# - Determine a baseline evaluation using AutoML.
# - Design and build a model architecture.
# - Upload the untrained model architecture as a Vertex AI Model resource.
#
#
# - Construct a training package that can be ran locally and as a Vertex AI Training job.
# - Decompose the training package into: data, model, train and task Python modules.
# - Obtain the location of the transformed training data from the user metadata of the Vertex AI Dataset resource.
# - Obtain the location of the model artifacts from the Vertex AI Model resource.
# - Include in the training package initializing a Vertex AI Experiment and corresponding run.
# - Log hyperparameters and training parameters for the experiment.
# - Add callbacks for early stop, TensorBoard, and hyperparameter tuning, where hyperparameter tuning is a command-line option.
#
#
# - Test the training package locally with a small number of epochs.
# - Test the training package with Vertex AI Training.
# - Do hyperparameter tuning with Vertex AI Hyperparameter Tuning.
# - Do full training of the custom model with Vertex AI Training.
# - Log the hyperparameter values for the experiment/run.
#
#
# - Evaluate the custom model.
# - Single evaluation slice, same metrics as AutoML
# - Add evaluation to the training package and return the results in a file in the Cloud Storage bucket used for training
# - Custom evaluation slices, custom metrics
# - Evaluate custom evaluation slices as a Vertex AI Batch Prediction for both AutoML and custom model
# - Perform custom metrics on the results from the batch job
#
#
# - Compare custom model metrics against the AutoML baseline
# - If less than baseline, then continue to experiment
# - If greater then baseline, then upload model as the new baseline and save evaluation results with the model.
# + [markdown] id="install_mlops"
# ## Installations
#
# Install *one time* the packages for executing the MLOps notebooks.
# + id="install_mlops"
ONCE_ONLY = False
if ONCE_ONLY:
# ! pip3 install -U tensorflow==2.5 $USER_FLAG
# ! pip3 install -U tensorflow-data-validation==1.2 $USER_FLAG
# ! pip3 install -U tensorflow-transform==1.2 $USER_FLAG
# ! pip3 install -U tensorflow-io==0.18 $USER_FLAG
# ! pip3 install --upgrade google-cloud-aiplatform[tensorboard] $USER_FLAG
# ! pip3 install --upgrade google-cloud-pipeline-components $USER_FLAG
# ! pip3 install --upgrade google-cloud-bigquery $USER_FLAG
# ! pip3 install --upgrade google-cloud-logging $USER_FLAG
# ! pip3 install --upgrade apache-beam[gcp] $USER_FLAG
# ! pip3 install --upgrade pyarrow $USER_FLAG
# ! pip3 install --upgrade cloudml-hypertune $USER_FLAG
# ! pip3 install --upgrade kfp $USER_FLAG
# ! pip3 install --upgrade torchvision $USER_FLAG
# ! pip3 install --upgrade rpy2 $USER_FLAG
# + [markdown] id="restart"
# ### Restart the kernel
#
# Once you've installed the additional packages, you need to restart the notebook kernel so it can find the packages.
# + id="restart"
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="project_id"
# #### Set your project ID
#
# **If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="set_project_id"
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + id="autoset_project_id"
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = ! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
# + id="set_gcloud_project_id"
# ! gcloud config set project $PROJECT_ID
# + [markdown] id="region"
# #### Region
#
# You can also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
#
# - Americas: `us-central1`
# - Europe: `europe-west4`
# - Asia Pacific: `asia-east1`
#
# You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services.
#
# Learn more about [Vertex AI regions](https://cloud.google.com/vertex-ai/docs/general/locations).
# + id="region"
REGION = "us-central1" # @param {type: "string"}
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append the timestamp onto the name of resources you create in this tutorial.
# + id="timestamp"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="bucket:mbsdk"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.
#
# Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
# + id="bucket"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
# + id="autoset_bucket"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="create_bucket"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="create_bucket"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="validate_bucket"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="validate_bucket"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="set_service_account"
# #### Service Account
#
# **If you don't know your service account**, try to get your service account using `gcloud` command by executing the second cell below.
# + id="set_service_account"
SERVICE_ACCOUNT = "[your-service-account]" # @param {type:"string"}
# + id="autoset_service_account"
if (
SERVICE_ACCOUNT == ""
or SERVICE_ACCOUNT is None
or SERVICE_ACCOUNT == "[your-service-account]"
):
# Get your GCP project id from gcloud
# shell_output = !gcloud auth list 2>/dev/null
SERVICE_ACCOUNT = shell_output[2].replace("*", "").strip()
print("Service Account:", SERVICE_ACCOUNT)
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# ### Import libraries and define constants
# + id="import_aip:mbsdk"
import google.cloud.aiplatform as aip
# + [markdown] id="import_tf"
# #### Import TensorFlow
#
# Import the TensorFlow package into your Python environment.
# + id="import_tf"
import tensorflow as tf
# + [markdown] id="import_tft"
# #### Import TensorFlow Transform
#
# Import the TensorFlow Transform (TFT) package into your Python environment.
# + id="import_tft"
import tensorflow_transform as tft
# + [markdown] id="import_tfdv"
# #### Import TensorFlow Data Validation
#
# Import the TensorFlow Data Validation (TFDV) package into your Python environment.
# + id="import_tfdv"
import tensorflow_data_validation as tfdv
# + [markdown] id="init_aip:mbsdk,all"
# ### Initialize Vertex AI SDK for Python
#
# Initialize the Vertex AI SDK for Python for your project and corresponding bucket.
# + id="init_aip:mbsdk,all"
aip.init(project=PROJECT_ID, location=REGION, staging_bucket=BUCKET_NAME)
# + [markdown] id="accelerators:training,prediction,ngpu,mbsdk"
# #### Set hardware accelerators
#
# You can set hardware accelerators for training and prediction.
#
# Set the variables `TRAIN_GPU/TRAIN_NGPU` and `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
#
# (aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
#
#
# Otherwise specify `(None, None)` to use a container image to run on a CPU.
#
# Learn more about [hardware accelerator support for your region](https://cloud.google.com/vertex-ai/docs/general/locations#accelerators).
#
# *Note*: TF releases before 2.3 for GPU support will fail to load the custom model in this tutorial. It is a known issue and fixed in TF 2.3. This is caused by static graph ops that are generated in the serving function. If you encounter this issue on your own custom models, use a container image for TF 2.3 with GPU support.
# + id="accelerators:training,prediction,ngpu,mbsdk"
if os.getenv("IS_TESTING_TRAIN_GPU"):
TRAIN_GPU, TRAIN_NGPU = (
aip.gapic.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_TRAIN_GPU")),
)
else:
TRAIN_GPU, TRAIN_NGPU = (aip.gapic.AcceleratorType.NVIDIA_TESLA_K80, 4)
if os.getenv("IS_TESTING_DEPLOY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.gapic.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPLOY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (None, None)
# + [markdown] id="container:training,prediction"
# #### Set pre-built containers
#
# Set the pre-built Docker container image for training and prediction.
#
#
# For the latest list, see [Pre-built containers for training](https://cloud.google.com/ai-platform-unified/docs/training/pre-built-containers).
#
#
# For the latest list, see [Pre-built containers for prediction](https://cloud.google.com/ai-platform-unified/docs/predictions/pre-built-containers).
# + id="container:training,prediction"
if os.getenv("IS_TESTING_TF"):
TF = os.getenv("IS_TESTING_TF")
else:
TF = "2.5".replace(".", "-")
if TF[0] == "2":
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf2-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf2-cpu.{}".format(TF)
else:
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf-cpu.{}".format(TF)
TRAIN_IMAGE = "{}-docker.pkg.dev/vertex-ai/training/{}:latest".format(
REGION.split("-")[0], TRAIN_VERSION
)
DEPLOY_IMAGE = "{}-docker.pkg.dev/vertex-ai/prediction/{}:latest".format(
REGION.split("-")[0], DEPLOY_VERSION
)
print("Training:", TRAIN_IMAGE, TRAIN_GPU, TRAIN_NGPU)
print("Deployment:", DEPLOY_IMAGE, DEPLOY_GPU, DEPLOY_NGPU)
# + [markdown] id="machine:training,prediction"
# #### Set machine type
#
# Next, set the machine type to use for training and prediction.
#
# - Set the variables `TRAIN_COMPUTE` and `DEPLOY_COMPUTE` to configure the compute resources for the VMs you will use for for training and prediction.
# - `machine type`
# - `n1-standard`: 3.75GB of memory per vCPU.
# - `n1-highmem`: 6.5GB of memory per vCPU
# - `n1-highcpu`: 0.9 GB of memory per vCPU
# - `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
#
# *Note: The following is not supported for training:*
#
# - `standard`: 2 vCPUs
# - `highcpu`: 2, 4 and 8 vCPUs
#
# *Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*.
# + id="machine:training,prediction"
if os.getenv("IS_TESTING_TRAIN_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_TRAIN_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
TRAIN_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Train machine type", TRAIN_COMPUTE)
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
# + [markdown] id="find_dataset:bq"
# ### Retrieve the dataset from stage 1
#
# Next, retrieve the dataset you created during stage 1 with the helper function `find_dataset()`. This helper function finds all the datasets whose display name matches the specified prefix and import format (e.g., bq). Finally it sorts the matches by create time and returns the latest version.
# + id="find_dataset:bq"
def find_dataset(display_name_prefix, import_format):
matches = []
datasets = aip.TabularDataset.list()
for dataset in datasets:
if dataset.display_name.startswith(display_name_prefix):
try:
if (
"bq" == import_format
and dataset.to_dict()["metadata"]["inputConfig"]["bigquerySource"]
):
matches.append(dataset)
if (
"csv" == import_format
and dataset.to_dict()["metadata"]["inputConfig"]["gcsSource"]
):
matches.append(dataset)
except:
pass
create_time = None
for match in matches:
if create_time is None or match.create_time > create_time:
create_time = match.create_time
dataset = match
return dataset
dataset = find_dataset("Chicago Taxi", "bq")
print(dataset)
# + [markdown] id="load_dataset_user_metadata"
# ### Load dataset's user metadata
#
# Load the user metadata for the dataset.
# + id="load_dataset_user_metadata"
import json
try:
with tf.io.gfile.GFile(
"gs://" + dataset.labels["user_metadata"] + "/metadata.jsonl", "r"
) as f:
metadata = json.load(f)
print(metadata)
except:
print("no metadata")
# + [markdown] id="create_automl_pipeline:tabular,lbn"
# ### Create and run training pipeline
#
# To train an AutoML model, you perform two steps: 1) create a training pipeline, and 2) run the pipeline.
#
# #### Create training pipeline
#
# An AutoML training pipeline is created with the `AutoMLTabularTrainingJob` class, with the following parameters:
#
# - `display_name`: The human readable name for the `TrainingJob` resource.
# - `optimization_prediction_type`: The type task to train the model for.
# - `classification`: A tabuar classification model.
# - `regression`: A tabular regression model.
# - `column_transformations`: (Optional): Transformations to apply to the input columns
# - `optimization_objective`: The optimization objective to minimize or maximize.
# - binary classification:
# - `minimize-log-loss`
# - `maximize-au-roc`
# - `maximize-au-prc`
# - `maximize-precision-at-recall`
# - `maximize-recall-at-precision`
# - multi-class classification:
# - `minimize-log-loss`
# - regression:
# - `minimize-rmse`
# - `minimize-mae`
# - `minimize-rmsle`
#
# The instantiated object is the DAG (directed acyclic graph) for the training pipeline.
# + id="create_automl_pipeline:tabular,lbn"
dag = aip.AutoMLTabularTrainingJob(
display_name="chicago_" + TIMESTAMP,
optimization_prediction_type="classification",
optimization_objective="minimize-log-loss",
)
print(dag)
# + [markdown] id="run_automl_pipeline:async,tabular"
# #### Run the training pipeline
#
# Next, you run the DAG to start the training job by invoking the method `run`, with the following parameters:
#
# - `dataset`: The `Dataset` resource to train the model.
# - `model_display_name`: The human readable name for the trained model.
# - `training_fraction_split`: The percentage of the dataset to use for training.
# - `test_fraction_split`: The percentage of the dataset to use for test (holdout data).
# - `validation_fraction_split`: The percentage of the dataset to use for validation.
# - `target_column`: The name of the column to train as the label.
# - `budget_milli_node_hours`: (optional) Maximum training time specified in unit of millihours (1000 = hour).
# - `disable_early_stopping`: If `True`, training maybe completed before using the entire budget if the service believes it cannot further improve on the model objective measurements.
#
# The `run` method when completed returns the `Model` resource.
#
# The execution of the training pipeline will take upto 180 minutes.
# + id="run_automl_pipeline:async,tabular"
async_model = dag.run(
dataset=dataset,
model_display_name="chicago_" + TIMESTAMP,
training_fraction_split=0.8,
validation_fraction_split=0.1,
test_fraction_split=0.1,
budget_milli_node_hours=8000,
disable_early_stopping=False,
target_column="tip_bin",
sync=False,
)
# + [markdown] id="start_experiment"
# ### Create experiment for tracking training related metadata
#
# Setup tracking the parameters (configuration) and metrics (results) for each experiment:
#
# - `aip.init()` - Create an experiment instance
# - `aip.start_run()` - Track a specific run within the experiment.
#
# Learn more about [Introduction to Vertex AI ML Metadata](https://cloud.google.com/vertex-ai/docs/ml-metadata/introduction).
# + id="start_experiment"
EXPERIMENT_NAME = "chicago-" + TIMESTAMP
aip.init(experiment=EXPERIMENT_NAME)
aip.start_run("run-1")
# + [markdown] id="create_tensorboard_instance"
# ### Create a Vertex AI TensorBoard instance
#
# Create a Vertex AI TensorBoard instance to use TensorBoard in conjunction with Vertex AI Training for custom model training.
#
# Learn more about [Get started with Vertex AI TensorBoard](https://cloud.google.com/vertex-ai/docs/experiments/tensorboard-overview).
# + id="create_tensorboard_instance"
TENSORBOARD_DISPLAY_NAME = "chicago_" + TIMESTAMP
tensorboard = aip.Tensorboard.create(display_name=TENSORBOARD_DISPLAY_NAME)
tensorboard_resource_name = tensorboard.gca_resource.name
print("TensorBoard resource name:", tensorboard_resource_name)
# + [markdown] id="create_input_layer:tabular"
# ### Create the input layer for your custom model
#
# Next, you create the input layer for your custom tabular model, based on the data types of each feature.
# + id="create_input_layer:tabular"
from tensorflow.keras.layers import Input
def create_model_inputs(
numeric_features=None, categorical_features=None, embedding_features=None
):
inputs = {}
for feature_name in numeric_features:
inputs[feature_name] = Input(name=feature_name, shape=[], dtype=tf.float32)
for feature_name in categorical_features:
inputs[feature_name] = Input(name=feature_name, shape=[], dtype=tf.int64)
for feature_name in embedding_features:
inputs[feature_name] = Input(name=feature_name, shape=[], dtype=tf.int64)
return inputs
# + id="make_input_layer:tabular"
input_layers = create_model_inputs(
numeric_features=metadata["numeric_features"],
categorical_features=metadata["categorical_features"],
embedding_features=metadata["embedding_features"],
)
print(input_layers)
# + [markdown] id="create_binary_classifier:tabular"
# ### Create the binary classifier custom model
#
# Next, you create your binary classifier custom tabular model.
# + id="create_binary_classifier:tabular"
from math import sqrt
from tensorflow.keras import Model, Sequential
from tensorflow.keras.layers import (Activation, Concatenate, Dense, Embedding,
experimental)
def create_binary_classifier(
input_layers,
tft_output,
metaparams,
numeric_features,
categorical_features,
embedding_features,
):
layers = []
for feature_name in input_layers:
if feature_name in embedding_features:
vocab_size = tft_output.vocabulary_size_by_name(feature_name)
embedding_size = int(sqrt(vocab_size))
embedding_output = Embedding(
input_dim=vocab_size + 1,
output_dim=embedding_size,
name=f"{feature_name}_embedding",
)(input_layers[feature_name])
layers.append(embedding_output)
elif feature_name in categorical_features:
vocab_size = tft_output.vocabulary_size_by_name(feature_name)
onehot_layer = experimental.preprocessing.CategoryEncoding(
num_tokens=vocab_size,
output_mode="binary",
name=f"{feature_name}_onehot",
)(input_layers[feature_name])
layers.append(onehot_layer)
elif feature_name in numeric_features:
numeric_layer = tf.expand_dims(input_layers[feature_name], -1)
layers.append(numeric_layer)
else:
pass
joined = Concatenate(name="combines_inputs")(layers)
feedforward_output = Sequential(
[Dense(units, activation="relu") for units in metaparams["hidden_units"]],
name="feedforward_network",
)(joined)
logits = Dense(units=1, name="logits")(feedforward_output)
pred = Activation("sigmoid")(logits)
model = Model(inputs=input_layers, outputs=[pred])
return model
# + id="make_binary_classifier:tabular"
TRANSFORM_ARTIFACTS_DIR = metadata["transform_artifacts_dir"]
tft_output = tft.TFTransformOutput(TRANSFORM_ARTIFACTS_DIR)
metaparams = {"hidden_units": [128, 64]}
aip.log_params(metaparams)
model = create_binary_classifier(
input_layers,
tft_output,
metaparams,
numeric_features=metadata["numeric_features"],
categorical_features=metadata["categorical_features"],
embedding_features=metadata["embedding_features"],
)
model.summary()
# + [markdown] id="visualize_model"
# #### Visualize the model architecture
#
# Next, visualize the architecture of the custom model.
# + id="visualize_model"
tf.keras.utils.plot_model(model, show_shapes=True, show_dtype=True)
# + [markdown] id="save_model:gcs"
# ### Save model artifacts
#
# Next, save the model artifacts to your Cloud Storage bucket
# + id="save_model:gcs"
MODEL_DIR = f"{BUCKET_NAME}/base_model"
model.save(MODEL_DIR)
# + [markdown] id="upload_model:vertex,base_model"
# ### Upload the local model to a Vertex AI Model resource
#
# Next, you upload your local custom model artifacts to Vertex AI to convert into a managed Vertex AI Model resource.
# + id="upload_model:vertex,base_model"
vertex_custom_model = aip.Model.upload(
display_name="chicago_" + TIMESTAMP,
artifact_uri=MODEL_DIR,
serving_container_image_uri=DEPLOY_IMAGE,
labels={"base_model": "1"},
sync=True,
)
# + [markdown] id="construct_training_package"
# ### Construct the training package
#
# #### Package layout
#
# Before you start training, you will look at how a Python package is assembled for a custom training job. When unarchived, the package contains the following directory/file layout.
#
# - PKG-INFO
# - README.md
# - setup.cfg
# - setup.py
# - trainer
# - \_\_init\_\_.py
# - task.py
# - other Python scripts
#
# The files `setup.cfg` and `setup.py` are the instructions for installing the package into the operating environment of the Docker image.
#
# The file `trainer/task.py` is the Python script for executing the custom training job.
# + id="construct_training_package"
# Make folder for Python training script
# ! rm -rf custom
# ! mkdir custom
# Add package information
# ! touch custom/README.md
setup_cfg = "[egg_info]\n\ntag_build =\n\ntag_date = 0"
# ! echo "$setup_cfg" > custom/setup.cfg
setup_py = "import setuptools\n\nsetuptools.setup(\n\n install_requires=[\n\n 'google-cloud-aiplatform',\n\n 'cloudml-hypertune',\n\n 'tensorflow_datasets==1.3.0',\n\n 'tensorflow_data_validation==1.2',\n\n ],\n\n packages=setuptools.find_packages())"
# ! echo "$setup_py" > custom/setup.py
pkg_info = "Metadata-Version: 1.0\n\nName: Chicago Taxi tabular binary classifier\n\nVersion: 0.0.0\n\nSummary: Demostration training script\n\nHome-page: www.google.com\n\nAuthor: Google\n\nAuthor-email: <EMAIL>\n\nLicense: Public\n\nDescription: Demo\n\nPlatform: Vertex AI"
# ! echo "$pkg_info" > custom/PKG-INFO
# Make the training subfolder
# ! mkdir custom/trainer
# ! touch custom/trainer/__init__.py
# + [markdown] id="transform_feature_spec"
# #### Get feature specification for the preprocessed data
#
# Next, create the feature specification for the preprocessed data.
# + id="transform_feature_spec"
transform_feature_spec = tft_output.transformed_feature_spec()
print(transform_feature_spec)
# + [markdown] id="read_tfrecords_func"
# ### Load the transformed data into a tf.data.Dataset
#
# Next, you load the gzip TFRecords on Cloud Storage storage into a `tf.data.Dataset` generator. These functions are re-used when training the custom model using `Vertex Training`, so you save them to the python training package.
# + id="read_tfrecords_func"
# %%writefile custom/trainer/data.py
import tensorflow as tf
def _gzip_reader_fn(filenames):
"""Small utility returning a record reader that can read gzip'ed files."""
return tf.data.TFRecordDataset(filenames, compression_type="GZIP")
def get_dataset(file_pattern, feature_spec, label_column, batch_size=200):
"""Generates features and label for tuning/training.
Args:
file_pattern: input tfrecord file pattern.
feature_spec: a dictionary of feature specifications.
batch_size: representing the number of consecutive elements of returned
dataset to combine in a single batch
Returns:
A dataset that contains (features, indices) tuple where features is a
dictionary of Tensors, and indices is a single Tensor of label indices.
"""
dataset = tf.data.experimental.make_batched_features_dataset(
file_pattern=file_pattern,
batch_size=batch_size,
features=feature_spec,
label_key=label_column,
reader=_gzip_reader_fn,
num_epochs=1,
drop_final_batch=True,
)
return dataset
# + id="read_tfrecords"
from custom.trainer import data
TRANSFORMED_DATA_PREFIX = metadata["transformed_data_prefix"]
LABEL_COLUMN = metadata["label_column"]
train_data_file_pattern = TRANSFORMED_DATA_PREFIX + "/train/data-*.gz"
val_data_file_pattern = TRANSFORMED_DATA_PREFIX + "/val/data-*.gz"
test_data_file_pattern = TRANSFORMED_DATA_PREFIX + "/test/data-*.gz"
for input_features, target in data.get_dataset(
train_data_file_pattern, transform_feature_spec, LABEL_COLUMN, batch_size=3
).take(1):
for key in input_features:
print(
f"{key} {input_features[key].dtype}: {input_features[key].numpy().tolist()}"
)
print(f"target: {target.numpy().tolist()}")
# + [markdown] id="test_model_input"
# #### Test the model architecture with transformed input
#
# Next, test the model architecture with a sample of the transformed training input.
#
# *Note:* Since the model is untrained, the predictions should be random. Since this is a binary classifier, expect the predicted results ~0.5.
# + id="test_model_input"
model(input_features)
# + [markdown] id="train_model_func"
# ## Develop and test the training scripts
#
# When experimenting, one typically develops and tests the training package locally, before moving to training in the cloud.
#
# ### Create training script
#
# Next, you write the Python script for compiling and training the model.
# + id="train_model_func"
# %%writefile custom/trainer/train.py
from trainer import data
import tensorflow as tf
import logging
from hypertune import HyperTune
def compile(model, hyperparams):
''' Compile the model '''
optimizer = tf.keras.optimizers.Adam(learning_rate=hyperparams["learning_rate"])
loss = tf.keras.losses.BinaryCrossentropy(from_logits=False)
metrics = [tf.keras.metrics.BinaryAccuracy(name="accuracy")]
model.compile(optimizer=optimizer,loss=loss, metrics=metrics)
return model
def warmup(
model,
hyperparams,
train_data_dir,
label_column,
transformed_feature_spec
):
''' Warmup the initialized model weights '''
train_dataset = data.get_dataset(
train_data_dir,
transformed_feature_spec,
label_column,
batch_size=hyperparams["batch_size"],
)
lr_inc = (hyperparams['end_learning_rate'] - hyperparams['start_learning_rate']) / hyperparams['num_epochs']
def scheduler(epoch, lr):
if epoch == 0:
return hyperparams['start_learning_rate']
return lr + lr_inc
callbacks = [tf.keras.callbacks.LearningRateScheduler(scheduler)]
logging.info("Model warmup started...")
history = model.fit(
train_dataset,
epochs=hyperparams["num_epochs"],
steps_per_epoch=hyperparams["steps"],
callbacks=callbacks
)
logging.info("Model warmup completed.")
return history
def train(
model,
hyperparams,
train_data_dir,
val_data_dir,
label_column,
transformed_feature_spec,
log_dir,
tuning=False
):
''' Train the model '''
train_dataset = data.get_dataset(
train_data_dir,
transformed_feature_spec,
label_column,
batch_size=hyperparams["batch_size"],
)
val_dataset = data.get_dataset(
val_data_dir,
transformed_feature_spec,
label_column,
batch_size=hyperparams["batch_size"],
)
early_stop = tf.keras.callbacks.EarlyStopping(
monitor=hyperparams["early_stop"]["monitor"], patience=hyperparams["early_stop"]["patience"], restore_best_weights=True
)
callbacks = [early_stop]
if log_dir:
tensorboard = tf.keras.callbacks.TensorBoard(log_dir=log_dir)
callbacks = callbacks.append(tensorboard)
if tuning:
# Instantiate the HyperTune reporting object
hpt = HyperTune()
# Reporting callback
class HPTCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs=None):
hpt.report_hyperparameter_tuning_metric(
hyperparameter_metric_tag='val_loss',
metric_value=logs['val_loss'],
global_step=epoch
)
if not callbacks:
callbacks = []
callbacks.append(HPTCallback())
logging.info("Model training started...")
history = model.fit(
train_dataset,
epochs=hyperparams["num_epochs"],
validation_data=val_dataset,
callbacks=callbacks
)
logging.info("Model training completed.")
return history
def evaluate(
model,
hyperparams,
test_data_dir,
label_column,
transformed_feature_spec
):
logging.info("Model evaluation started...")
test_dataset = data.get_dataset(
test_data_dir,
transformed_feature_spec,
label_column,
hyperparams["batch_size"],
)
evaluation_metrics = model.evaluate(test_dataset)
logging.info("Model evaluation completed.")
return evaluation_metrics
# + [markdown] id="train_model_local"
# ### Train the model locally
#
# Next, test the training package locally, by training with just a few epochs:
#
# - `num_epochs`: The number of epochs to pass to the training package.
# - `compile()`: Compile the model for training.
# - `warmup()`: Warmup the initialized model weights.
# - `train()`: Train the model.
# + id="train_model_local"
os.chdir("custom")
import logging
from trainer import train
TENSORBOARD_LOG_DIR = "./logs"
logging.getLogger().setLevel(logging.INFO)
hyperparams = {}
hyperparams["learning_rate"] = 0.01
aip.log_params(hyperparams)
train.compile(model, hyperparams)
warmupparams = {}
warmupparams["start_learning_rate"] = 0.0001
warmupparams["end_learning_rate"] = 0.01
warmupparams["num_epochs"] = 4
warmupparams["batch_size"] = 64
warmupparams["steps"] = 50
aip.log_params(warmupparams)
train.warmup(
model, warmupparams, train_data_file_pattern, LABEL_COLUMN, transform_feature_spec
)
trainparams = {}
trainparams["num_epochs"] = 5
trainparams["batch_size"] = 64
trainparams["early_stop"] = {"monitor": "val_loss", "patience": 5}
aip.log_params(trainparams)
train.train(
model,
trainparams,
train_data_file_pattern,
val_data_file_pattern,
LABEL_COLUMN,
transform_feature_spec,
TENSORBOARD_LOG_DIR,
)
os.chdir("..")
# + [markdown] id="eval_model_local"
# ### Evaluate the model locally
#
# Next, test the evaluation portion of the training package:
#
#
# - `evaluate()`: Evaluate the model.
# + id="eval_model_local"
os.chdir("custom")
from trainer import train
evalparams = {}
evalparams["batch_size"] = 64
metrics = {}
metrics["loss"], metrics["acc"] = train.evaluate(
model, evalparams, test_data_file_pattern, LABEL_COLUMN, transform_feature_spec
)
print("ACC", metrics["acc"], "LOSS", metrics["loss"])
aip.log_metrics(metrics)
os.chdir("..")
# + [markdown] id="create_model_get"
# ### Retrieve model from Vertex AI
#
# Next, create the Python script to retrieve your experimental model from Vertex AI.
# + id="create_model_get"
# %%writefile custom/trainer/model.py
import google.cloud.aiplatform as aip
def get(model_id):
model = aip.Model(model_id)
return model
# + [markdown] id="create_task_py"
# ### Create the task script for the Python training package
#
# Next, you create the `task.py` script for driving the training package. Some noteable steps include:
#
# - Command-line arguments:
# - `model-id`: The resource ID of the `Model` resource you built during experimenting. This is the untrained model architecture.
# - `dataset-id`: The resource ID of the `Dataset` resource to use for training.
# - `experiment`: The name of the experiment.
# - `run`: The name of the run within this experiment.
# - `tensorboard-logdir`: The logging directory for Vertex AI Tensorboard.
#
#
# - `get_data()`:
# - Loads the Dataset resource into memory.
# - Obtains the user metadata from the Dataset resource.
# - From the metadata, obtain location of transformed data, transformation function and name of label column
#
#
# - `get_model()`:
# - Loads the Model resource into memory.
# - Obtains location of model artifacts of the model architecture.
# - Loads the model architecture.
# - Compiles the model.
#
#
# - `warmup_model()`:
# - Warms up the initialized model weights
#
#
# - `train_model()`:
# - Train the model.
#
#
# - `evaluate_model()`:
# - Evaluates the model.
# - Saves evaluation metrics to Cloud Storage bucket.
# + id="create_task_py"
# %%writefile custom/trainer/task.py
import os
import argparse
import logging
import json
import tensorflow as tf
import tensorflow_transform as tft
from tensorflow.python.client import device_lib
import google.cloud.aiplatform as aip
from trainer import data
from trainer import model as model_
from trainer import train
try:
from trainer import serving
except:
pass
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv('AIP_MODEL_DIR'), type=str, help='Model dir.')
parser.add_argument('--model-id', dest='model_id',
default=None, type=str, help='Vertex Model ID.')
parser.add_argument('--dataset-id', dest='dataset_id',
default=None, type=str, help='Vertex Dataset ID.')
parser.add_argument('--lr', dest='lr',
default=0.001, type=float,
help='Learning rate.')
parser.add_argument('--start_lr', dest='start_lr',
default=0.0001, type=float,
help='Starting learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=20, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=200, type=int,
help='Number of steps per epoch.')
parser.add_argument('--batch_size', dest='batch_size',
default=16, type=int,
help='Batch size.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
parser.add_argument('--tensorboard-log-dir', dest='tensorboard_log_dir',
default=os.getenv('AIP_TENSORBOARD_LOG_DIR'), type=str,
help='Output file for tensorboard logs')
parser.add_argument('--experiment', dest='experiment',
default=None, type=str,
help='Name of experiment')
parser.add_argument('--project', dest='project',
default=None, type=str,
help='Name of project')
parser.add_argument('--run', dest='run',
default=None, type=str,
help='Name of run in experiment')
parser.add_argument('--evaluate', dest='evaluate',
default=False, type=bool,
help='Whether to perform evaluation')
parser.add_argument('--serving', dest='serving',
default=False, type=bool,
help='Whether to attach the serving function')
parser.add_argument('--tuning', dest='tuning',
default=False, type=bool,
help='Whether to perform hyperparameter tuning')
parser.add_argument('--warmup', dest='warmup',
default=False, type=bool,
help='Whether to perform warmup weight initialization')
args = parser.parse_args()
logging.getLogger().setLevel(logging.INFO)
logging.info('DEVICES' + str(device_lib.list_local_devices()))
# Single Machine, single compute device
if args.distribute == 'single':
if tf.test.is_gpu_available():
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
else:
strategy = tf.distribute.OneDeviceStrategy(device="/cpu:0")
logging.info("Single device training")
# Single Machine, multiple compute device
elif args.distribute == 'mirrored':
strategy = tf.distribute.MirroredStrategy()
logging.info("Mirrored Strategy distributed training")
# Multi Machine, multiple compute device
elif args.distribute == 'multiworker':
strategy = tf.distribute.MultiWorkerMirroredStrategy()
logging.info("Multi-worker Strategy distributed training")
logging.info('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
logging.info('num_replicas_in_sync = {}'.format(strategy.num_replicas_in_sync))
# Initialize the run for this experiment
if args.experiment:
logging.info("Initialize experiment: {}".format(args.experiment))
aip.init(experiment=args.experiment, project=args.project)
aip.start_run(args.run)
metadata = {}
def get_data():
''' Get the preprocessed training data '''
global train_data_file_pattern, val_data_file_pattern, test_data_file_pattern
global label_column, transform_feature_spec, metadata
dataset = aip.TabularDataset(args.dataset_id)
METADATA = 'gs://' + dataset.labels['user_metadata'] + "/metadata.jsonl"
with tf.io.gfile.GFile(METADATA, "r") as f:
metadata = json.load(f)
TRANSFORMED_DATA_PREFIX = metadata['transformed_data_prefix']
label_column = metadata['label_column']
train_data_file_pattern = TRANSFORMED_DATA_PREFIX + '/train/data-*.gz'
val_data_file_pattern = TRANSFORMED_DATA_PREFIX + '/val/data-*.gz'
test_data_file_pattern = TRANSFORMED_DATA_PREFIX + '/test/data-*.gz'
TRANSFORM_ARTIFACTS_DIR = metadata['transform_artifacts_dir']
tft_output = tft.TFTransformOutput(TRANSFORM_ARTIFACTS_DIR)
transform_feature_spec = tft_output.transformed_feature_spec()
def get_model():
''' Get the untrained model architecture '''
global model_artifacts
vertex_model = model_.get(args.model_id)
model_artifacts = vertex_model.gca_resource.artifact_uri
model = tf.keras.models.load_model(model_artifacts)
# Compile the model
hyperparams = {}
hyperparams["learning_rate"] = args.lr
if args.experiment:
aip.log_params(hyperparams)
metadata.update(hyperparams)
with tf.io.gfile.GFile(os.path.join(args.model_dir, "metrics.txt"), "w") as f:
f.write(json.dumps(metadata))
train.compile(model, hyperparams)
return model
def warmup_model(model):
''' Warmup the initialized model weights '''
warmupparams = {}
warmupparams["num_epochs"] = args.epochs
warmupparams["batch_size"] = args.batch_size
warmupparams["steps"] = args.steps
warmupparams["start_learning_rate"] = args.start_lr
warmupparams["end_learning_rate"] = args.lr
train.warmup(model, warmupparams, train_data_file_pattern, label_column, transform_feature_spec)
return model
def train_model(model):
''' Train the model '''
trainparams = {}
trainparams["num_epochs"] = args.epochs
trainparams["batch_size"] = args.batch_size
trainparams["early_stop"] = {"monitor": "val_loss", "patience": 5}
if args.experiment:
aip.log_params(trainparams)
metadata.update(trainparams)
with tf.io.gfile.GFile(os.path.join(args.model_dir, "metrics.txt"), "w") as f:
f.write(json.dumps(metadata))
train.train(model, trainparams, train_data_file_pattern, val_data_file_pattern, label_column, transform_feature_spec, args.tensorboard_log_dir, args.tuning)
return model
def evaluate_model(model):
''' Evaluate the model '''
evalparams = {}
evalparams["batch_size"] = args.batch_size
metrics = train.evaluate(model, evalparams, test_data_file_pattern, label_column, transform_feature_spec)
metadata.update({'metrics': metrics})
with tf.io.gfile.GFile(os.path.join(args.model_dir, "metrics.txt"), "w") as f:
f.write(json.dumps(metadata))
get_data()
with strategy.scope():
model = get_model()
if args.warmup:
model = warmup_model(model)
else:
model = train_model(model)
if args.evaluate:
evaluate_model(model)
if args.serving:
logging.info('Save serving model to: ' + args.model_dir)
serving.construct_serving_model(
model=model,
serving_model_dir=args.model_dir,
metadata=metadata
)
elif args.warmup:
logging.info('Save warmed up model to: ' + model_artifacts)
model.save(model_artifacts)
else:
logging.info('Save trained model to: ' + args.model_dir)
model.save(args.model_dir)
# + [markdown] id="test_package_locally"
# ### Test training package locally
#
# Next, test your completed training package locally with just a few epochs.
# + id="test_package_locally"
DATASET_ID = dataset.resource_name
MODEL_ID = vertex_custom_model.resource_name
# !cd custom; python3 -m trainer.task --model-id={MODEL_ID} --dataset-id={DATASET_ID} --experiment='chicago' --run='test' --project={PROJECT_ID} --epochs=5 --model-dir=/tmp --evaluate=True
# + [markdown] id="warmup_base_model"
# ### Warmup training
#
# Now that you have tested the training scripts, you perform warmup training on the base model. Warmup training is used to stabilize the weight initialization. By doing so, each subsequent training and tuning of the model architecture will start with the same stabilized weight initialization.
# + id="warmup_base_model"
MODEL_DIR = f"{BUCKET_NAME}/base_model"
# !cd custom; python3 -m trainer.task --model-id={MODEL_ID} --dataset-id={DATASET_ID} --project={PROJECT_ID} --epochs=5 --steps=300 --batch_size=16 --lr=0.01 --start_lr=0.0001 --model-dir={MODEL_DIR} --warmup=True
# + [markdown] id="mirrored_intro"
# ## Mirrored Strategy
#
# When training on a single VM, one can either train was a single compute device or with multiple compute devices on the same VM. With Vertex AI Distributed Training you can specify both the number of compute devices for the VM instance and type of compute devices: CPU, GPU.
#
# Vertex AI Distributed Training supports `tf.distribute.MirroredStrategy' for TensorFlow models. To enable training across multiple compute devices on the same VM, you do the following additional steps in your Python training script:
#
# 1. Set the tf.distribute.MirrorStrategy
# 2. Compile the model within the scope of tf.distribute.MirrorStrategy. *Note:* Tells MirroredStrategy which variables to mirror across your compute devices.
# 3. Increase the batch size for each compute device to num_devices * batch size.
#
# During transitions, the distribution of batches will be synchronized as well as the updates to the model parameters.
# + [markdown] id="create_custom_pp_training_job:mbsdk"
# ### Create and run custom training job
#
#
# To train a custom model, you perform two steps: 1) create a custom training job, and 2) run the job.
#
# #### Create custom training job
#
# A custom training job is created with the `CustomTrainingJob` class, with the following parameters:
#
# - `display_name`: The human readable name for the custom training job.
# - `container_uri`: The training container image.
#
# - `python_package_gcs_uri`: The location of the Python training package as a tarball.
# - `python_module_name`: The relative path to the training script in the Python package.
# - `model_serving_container_uri`: The container image for deploying the model.
#
# *Note:* There is no requirements parameter. You specify any requirements in the `setup.py` script in your Python package.
# + id="create_custom_pp_training_job:mbsdk"
DISPLAY_NAME = "chicago_" + TIMESTAMP
job = aip.CustomPythonPackageTrainingJob(
display_name=DISPLAY_NAME,
python_package_gcs_uri=f"{BUCKET_NAME}/trainer_chicago.tar.gz",
python_module_name="trainer.task",
container_uri=TRAIN_IMAGE,
model_serving_container_image_uri=DEPLOY_IMAGE,
project=PROJECT_ID,
)
# + id="cleanup:trainer"
# ! rm -rf custom/logs
# ! rm -rf custom/trainer/__pycache__
# + [markdown] id="tarball_training_script"
# #### Store training script on your Cloud Storage bucket
#
# Next, you package the training folder into a compressed tar ball, and then store it in your Cloud Storage bucket.
# + id="tarball_training_script"
# ! rm -f custom.tar custom.tar.gz
# ! tar cvf custom.tar custom
# ! gzip custom.tar
# ! gsutil cp custom.tar.gz $BUCKET_NAME/trainer_chicago.tar.gz
# + [markdown] id="run_custom_pp_training_job:test"
# #### Run the custom Python package training job
#
# Next, you run the custom job to start the training job by invoking the method `run()`. The parameters are the same as when running a CustomTrainingJob.
#
# *Note:* The parameter service_account is set so that the initializing experiment step `aip.init(experiment="...")` has necessarily permission to access the Vertex AI Metadata Store.
# + id="run_custom_pp_training_job:test"
MODEL_DIR = BUCKET_NAME + "/testing"
CMDARGS = [
"--epochs=5",
"--batch_size=16",
"--distribute=mirrored",
"--experiment=chicago",
"--run=test",
"--project=" + PROJECT_ID,
"--model-id=" + MODEL_ID,
"--dataset-id=" + DATASET_ID,
]
model = job.run(
model_display_name="chicago_" + TIMESTAMP,
args=CMDARGS,
replica_count=1,
machine_type=TRAIN_COMPUTE,
accelerator_type=TRAIN_GPU.name,
accelerator_count=TRAIN_NGPU,
base_output_dir=MODEL_DIR,
service_account=SERVICE_ACCOUNT,
tensorboard=tensorboard_resource_name,
sync=True,
)
# + [markdown] id="delete_job"
# ### Delete a custom training job
#
# After a training job is completed, you can delete the training job with the method `delete()`. Prior to completion, a training job can be canceled with the method `cancel()`.
# + id="delete_job"
job.delete()
# + [markdown] id="model_delete:mbsdk"
# #### Delete the model
#
# The method 'delete()' will delete the model.
# + id="model_delete:mbsdk"
model.delete()
# + [markdown] id="hp_tuning"
# ## Hyperparameter tuning
#
# Next, you perform hyperparameter tuning with the training package. The training package has some additions that make the same package usable for both hyperparameter tuning, as well as local testing and full cloud training:
#
# - Command-Line:
# - `tuning`: indicates to use the HyperTune service as a callback during training.
#
#
# - `train()`: If tuning is set, creates and adds a callback to HyperTune service.
# + [markdown] id="train_custom_job_machine_specification"
# ### Prepare your machine specification
#
# Now define the machine specification for your custom training job. This tells Vertex what type of machine instance to provision for the training.
# - `machine_type`: The type of GCP instance to provision -- e.g., n1-standard-8.
# - `accelerator_type`: The type, if any, of hardware accelerator. In this tutorial if you previously set the variable `TRAIN_GPU != None`, you are using a GPU; otherwise you will use a CPU.
# - `accelerator_count`: The number of accelerators.
# + id="train_custom_job_machine_specification"
if TRAIN_GPU:
machine_spec = {
"machine_type": TRAIN_COMPUTE,
"accelerator_type": TRAIN_GPU,
"accelerator_count": TRAIN_NGPU,
}
else:
machine_spec = {"machine_type": TRAIN_COMPUTE, "accelerator_count": 0}
# + [markdown] id="train_custom_job_disk_specification"
# ### Prepare your disk specification
#
# (optional) Now define the disk specification for your custom training job. This tells Vertex what type and size of disk to provision in each machine instance for the training.
#
# - `boot_disk_type`: Either SSD or Standard. SSD is faster, and Standard is less expensive. Defaults to SSD.
# - `boot_disk_size_gb`: Size of disk in GB.
# + id="train_custom_job_disk_specification"
DISK_TYPE = "pd-ssd" # [ pd-ssd, pd-standard]
DISK_SIZE = 200 # GB
disk_spec = {"boot_disk_type": DISK_TYPE, "boot_disk_size_gb": DISK_SIZE}
# + [markdown] id="worker_pool_hpt"
# ### Define worker pool specification for hyperparameter tuning job
#
# Next, define the worker pool specification. Note that we plan to tune the learning rate and batch size, so you do not pass them as command-line arguments (omitted). The Vertex AI Hyperparameter Tuning service will pick values for both learning rate and batch size during trials, which it will pass along as command-line arguments.
# + id="worker_pool_hpt"
CMDARGS = [
"--epochs=5",
"--distribute=mirrored",
# "--experiment=chicago",
# "--run=tune",
# "--project=" + PROJECT_ID,
"--model-id=" + MODEL_ID,
"--dataset-id=" + DATASET_ID,
"--tuning=True",
]
worker_pool_spec = [
{
"replica_count": 1,
"machine_spec": machine_spec,
"disk_spec": disk_spec,
"python_package_spec": {
"executor_image_uri": TRAIN_IMAGE,
"package_uris": [BUCKET_NAME + "/trainer_chicago.tar.gz"],
"python_module": "trainer.task",
"args": CMDARGS,
},
}
]
# + [markdown] id="create_custom_job:mbsdk"
# ## Create a custom job
#
# Use the class `CustomJob` to create a custom job, such as for hyperparameter tuning, with the following parameters:
#
# - `display_name`: A human readable name for the custom job.
# - `worker_pool_specs`: The specification for the corresponding VM instances.
# + id="create_custom_job:mbsdk"
job = aip.CustomJob(
display_name="chicago_" + TIMESTAMP, worker_pool_specs=worker_pool_spec
)
# + [markdown] id="create_hpt_job:mbsdk"
# ## Create a hyperparameter tuning job
#
# Use the class `HyperparameterTuningJob` to create a hyperparameter tuning job, with the following parameters:
#
# - `display_name`: A human readable name for the custom job.
# - `custom_job`: The worker pool spec from this custom job applies to the CustomJobs created in all the trials.
# - `metrics_spec`: The metrics to optimize. The dictionary key is the metric_id, which is reported by your training job, and the dictionary value is the optimization goal of the metric('minimize' or 'maximize').
# - `parameter_spec`: The parameters to optimize. The dictionary key is the metric_id, which is passed into your training job as a command line key word argument, and the dictionary value is the parameter specification of the metric.
# - `search_algorithm`: The search algorithm to use: `grid`, `random` and `None`. If `None` is specified, the `Vizier` service (Bayesian) is used.
# - `max_trial_count`: The maximum number of trials to perform.
# + id="create_hpt_job:stage2"
from google.cloud.aiplatform import hyperparameter_tuning as hpt
hpt_job = aip.HyperparameterTuningJob(
display_name="chicago_" + TIMESTAMP,
custom_job=job,
metric_spec={
"val_loss": "minimize",
},
parameter_spec={
"lr": hpt.DoubleParameterSpec(min=0.001, max=0.1, scale="log"),
"batch_size": hpt.DiscreteParameterSpec([16, 32, 64, 128, 256], scale="linear"),
},
search_algorithm=None,
max_trial_count=8,
parallel_trial_count=1,
)
# + [markdown] id="run_hpt_job:mbsdk"
# ## Run the hyperparameter tuning job
#
# Use the `run()` method to execute the hyperparameter tuning job.
# + id="run_hpt_job:mbsdk"
hpt_job.run()
# + [markdown] id="best_trial:mbsdk"
# ### Best trial
#
# Now look at which trial was the best:
# + id="best_trial:mbsdk"
best = (None, None, None, 0.0)
for trial in hpt_job.trials:
# Keep track of the best outcome
if float(trial.final_measurement.metrics[0].value) > best[3]:
try:
best = (
trial.id,
float(trial.parameters[0].value),
float(trial.parameters[1].value),
float(trial.final_measurement.metrics[0].value),
)
except:
best = (
trial.id,
float(trial.parameters[0].value),
None,
float(trial.final_measurement.metrics[0].value),
)
print(best)
# + [markdown] id="delete_hpt_job"
# ### Delete the hyperparameter tuning job
#
# The method 'delete()' will delete the hyperparameter tuning job.
# + id="delete_hpt_job"
hpt_job.delete()
# + [markdown] id="save_hpt"
# ### Save the best hyperparameter values
# + id="save_hpt"
LR = best[2]
BATCH_SIZE = int(best[1])
# + [markdown] id="create_custom_pp_training_job:mbsdk"
# ### Create and run custom training job
#
#
# To train a custom model, you perform two steps: 1) create a custom training job, and 2) run the job.
#
# #### Create custom training job
#
# A custom training job is created with the `CustomTrainingJob` class, with the following parameters:
#
# - `display_name`: The human readable name for the custom training job.
# - `container_uri`: The training container image.
#
# - `python_package_gcs_uri`: The location of the Python training package as a tarball.
# - `python_module_name`: The relative path to the training script in the Python package.
# - `model_serving_container_uri`: The container image for deploying the model.
#
# *Note:* There is no requirements parameter. You specify any requirements in the `setup.py` script in your Python package.
# + id="create_custom_pp_training_job:mbsdk"
DISPLAY_NAME = "chicago_" + TIMESTAMP
job = aip.CustomPythonPackageTrainingJob(
display_name=DISPLAY_NAME,
python_package_gcs_uri=f"{BUCKET_NAME}/trainer_chicago.tar.gz",
python_module_name="trainer.task",
container_uri=TRAIN_IMAGE,
model_serving_container_image_uri=DEPLOY_IMAGE,
project=PROJECT_ID,
)
# + [markdown] id="run_custom_pp_training_job:full"
# #### Run the custom Python package training job
#
# Next, you run the custom job to start the training job by invoking the method `run()`. The parameters are the same as when running a CustomTrainingJob.
#
# *Note:* The parameter service_account is set so that the initializing experiment step `aip.init(experiment="...")` has necessarily permission to access the Vertex AI Metadata Store.
# + id="run_custom_pp_training_job:full"
MODEL_DIR = BUCKET_NAME + "/trained"
FULL_EPOCHS = 100
CMDARGS = [
f"--epochs={FULL_EPOCHS}",
f"--lr={LR}",
f"--batch_size={BATCH_SIZE}",
"--distribute=mirrored",
"--experiment=chicago",
"--run=full",
"--project=" + PROJECT_ID,
"--model-id=" + MODEL_ID,
"--dataset-id=" + DATASET_ID,
"--evaluate=True",
]
model = job.run(
model_display_name="chicago_" + TIMESTAMP,
args=CMDARGS,
replica_count=1,
machine_type=TRAIN_COMPUTE,
accelerator_type=TRAIN_GPU.name,
accelerator_count=TRAIN_NGPU,
base_output_dir=MODEL_DIR,
service_account=SERVICE_ACCOUNT,
tensorboard=tensorboard_resource_name,
sync=True,
)
# + [markdown] id="delete_job"
# ### Delete a custom training job
#
# After a training job is completed, you can delete the training job with the method `delete()`. Prior to completion, a training job can be canceled with the method `cancel()`.
# + id="delete_job"
job.delete()
# + [markdown] id="get_experiment"
# ### Get the experiment results
#
# Next, you use the experiment name as a parameter to the method `get_experiment_df()` to get the results of the experiment as a pandas dataframe.
# + id="get_experiment"
EXPERIMENT_NAME = "chicago"
experiment_df = aip.get_experiment_df()
experiment_df = experiment_df[experiment_df.experiment_name == EXPERIMENT_NAME]
experiment_df.T
# + [markdown] id="review_builtin_metrics"
# ## Review the custom model evaluation results
#
# Next, you review the evaluation metrics builtin into the training package.
# + id="review_builtin_metrics"
METRICS = MODEL_DIR + "/model/metrics.txt"
# ! gsutil cat $METRICS
# + [markdown] id="delete_tensorboard"
# ### Delete the TensorBoard instance
#
# Next, delete the TensorBoard instance.
# + id="delete_tensorboard"
tensorboard.delete()
# + id="reload_model"
vertex_custom_model = model
model = tf.keras.models.load_model(MODEL_DIR + "/model")
# + [markdown] id="serving_function:chicago"
# ## Add a serving function
#
# Next, you add a serving function to your model for online and batch prediction. This allows prediction requests to be sent in raw format (unpreprocessed), either as a serialized TF.Example or JSONL object. The serving function will then preprocess the prediction request into the transformed format expected by the model.
# + id="serving_function:chicago"
# %%writefile custom/trainer/serving.py
import tensorflow as tf
import tensorflow_data_validation as tfdv
import tensorflow_transform as tft
import logging
def _get_serve_features_fn(model, tft_output):
"""Returns a function that accept a dictionary of features and applies TFT."""
model.tft_layer = tft_output.transform_features_layer()
@tf.function
def serve_features_fn(raw_features):
"""Returns the output to be used in the serving signature."""
transformed_features = model.tft_layer(raw_features)
probabilities = model(transformed_features)
return {"scores": probabilities}
return serve_features_fn
def _get_serve_tf_examples_fn(model, tft_output, feature_spec):
"""Returns a function that parses a serialized tf.Example and applies TFT."""
model.tft_layer = tft_output.transform_features_layer()
@tf.function
def serve_tf_examples_fn(serialized_tf_examples):
"""Returns the output to be used in the serving signature."""
for key in list(feature_spec.keys()):
if key not in features:
feature_spec.pop(key)
parsed_features = tf.io.parse_example(serialized_tf_examples, feature_spec)
transformed_features = model.tft_layer(parsed_features)
probabilities = model(transformed_features)
return {"scores": probabilities}
return serve_tf_examples_fn
def construct_serving_model(
model, serving_model_dir, metadata
):
global features
schema_location = metadata['schema']
features = metadata['numeric_features'] + metadata['categorical_features'] + metadata['embedding_features']
print("FEATURES", features)
tft_output_dir = metadata["transform_artifacts_dir"]
schema = tfdv.load_schema_text(schema_location)
feature_spec = tft.tf_metadata.schema_utils.schema_as_feature_spec(schema).feature_spec
tft_output = tft.TFTransformOutput(tft_output_dir)
# Drop features that were not used in training
features_input_signature = {
feature_name: tf.TensorSpec(
shape=(None, 1), dtype=spec.dtype, name=feature_name
)
for feature_name, spec in feature_spec.items()
if feature_name in features
}
signatures = {
"serving_default": _get_serve_features_fn(
model, tft_output
).get_concrete_function(features_input_signature),
"serving_tf_example": _get_serve_tf_examples_fn(
model, tft_output, feature_spec
).get_concrete_function(
tf.TensorSpec(shape=[None], dtype=tf.string, name="examples")
),
}
logging.info("Model saving started...")
model.save(serving_model_dir, signatures=signatures)
logging.info("Model saving completed.")
# + [markdown] id="construct_serving_model"
# ### Construct the serving model
#
# Now construct the serving model and store the serving model to your Cloud Storage bucket.
# + id="construct_serving_model"
os.chdir("custom")
from trainer import serving
SERVING_MODEL_DIR = BUCKET_NAME + "/serving_model"
serving.construct_serving_model(
model=model, serving_model_dir=SERVING_MODEL_DIR, metadata=metadata
)
serving_model = tf.keras.models.load_model(SERVING_MODEL_DIR)
os.chdir("..")
# + [markdown] id="test_serving_model:tfrec"
# ### Test the serving model locally with tf.Example data
#
# Next, test the layer interface in the serving model for tf.Example data.
# + id="test_serving_model:tfrec"
EXPORTED_TFREC_PREFIX = metadata["exported_tfrec_prefix"]
file_names = tf.data.TFRecordDataset.list_files(
EXPORTED_TFREC_PREFIX + "/data-*.tfrecord"
)
for batch in tf.data.TFRecordDataset(file_names).batch(3).take(1):
predictions = serving_model.signatures["serving_tf_example"](batch)
for key in predictions:
print(f"{key}: {predictions[key]}")
# + [markdown] id="test_serving_model:jsonl,chicago"
# ### Test the serving model locally with JSONL data
#
# Next, test the layer interface in the serving model for JSONL data.
# + id="test_serving_model:jsonl,chicago"
schema = tfdv.load_schema_text(metadata["schema"])
feature_spec = tft.tf_metadata.schema_utils.schema_as_feature_spec(schema).feature_spec
instance = {
"dropoff_grid": "POINT(-87.6 41.9)",
"euclidean": 2064.2696,
"loc_cross": "",
"payment_type": "Credit Card",
"pickup_grid": "POINT(-87.6 41.9)",
"trip_miles": 1.37,
"trip_day": 12,
"trip_hour": 6,
"trip_month": 2,
"trip_day_of_week": 4,
"trip_seconds": 555,
}
for feature_name in instance:
dtype = feature_spec[feature_name].dtype
instance[feature_name] = tf.constant([[instance[feature_name]]], dtype)
predictions = serving_model.signatures["serving_default"](**instance)
for key in predictions:
print(f"{key}: {predictions[key].numpy()}")
# + [markdown] id="upload_serving_model:vertex,labels"
# ### Upload the serving model to a Vertex AI Model resource
#
# Next, you upload your serving custom model artifacts to Vertex AI to convert into a managed Vertex AI Model resource.
# + id="upload_serving_model:vertex,labels"
vertex_serving_model = aip.Model.upload(
display_name="chicago_" + TIMESTAMP,
artifact_uri=SERVING_MODEL_DIR,
serving_container_image_uri=DEPLOY_IMAGE,
labels={"user_metadata": BUCKET_NAME[5:]},
sync=True,
)
# + [markdown] id="evaluate_serving_model"
# ### Evaluate the serving model
#
# Next, evaluate the serving model with the evaluation (test) slices. For apples-to-apples comparison, you use the same evaluation slices for both the custom model and the AutoML model. Since your evaluation slices and metrics maybe custom, we recommend:
#
# - Send each evaluation slice as a Vertex AI Batch Prediction Job.
# - Use a custom evaluation script to evaluate the results from the batch prediction job.
# + id="evaluate_serving_model"
SERVING_OUTPUT_DATA_DIR = BUCKET_NAME + "/batch_eval"
EXPORTED_JSONL_PREFIX = metadata["exported_jsonl_prefix"]
MIN_NODES = 1
MAX_NODES = 1
job = vertex_serving_model.batch_predict(
instances_format="jsonl",
predictions_format="jsonl",
job_display_name="chicago_" + TIMESTAMP,
gcs_source=EXPORTED_JSONL_PREFIX + "*.jsonl",
gcs_destination_prefix=SERVING_OUTPUT_DATA_DIR,
model_parameters=None,
machine_type=DEPLOY_COMPUTE,
accelerator_type=DEPLOY_GPU,
accelerator_count=DEPLOY_NGPU,
starting_replica_count=MIN_NODES,
max_replica_count=MAX_NODES,
sync=True,
)
# + [markdown] id="custom_eval_script"
# ### Perform custom evaluation metrics
#
# After the batch job has completed, you input the results and target labels to your custom evaluation script. For demonstration purposes, we just display the results of the batch prediction.
# + id="custom_eval_script"
batch_dir = ! gsutil ls $SERVING_OUTPUT_DATA_DIR
batch_dir = batch_dir[0]
outputs = ! gsutil ls $batch_dir
errors = outputs[0]
results = outputs[1]
print("errors")
# ! gsutil cat $errors
print("results")
# ! gsutil cat $results | head -n10
# + id="set_model:async"
model = async_model
# + [markdown] id="automl_job_wait:mbsdk"
# ### Wait for completion of AutoML training job
#
# Next, wait for the AutoML training job to complete. Alternatively, one can set the parameter `sync` to `True` in the `run()` method to block until the AutoML training job is completed.
# + id="automl_job_wait:mbsdk"
model.wait()
# + [markdown] id="evaluate_the_model:mbsdk"
# ## Review model evaluation scores
# After your model has finished training, you can review the evaluation scores for it.
#
# First, you need to get a reference to the new model. As with datasets, you can either use the reference to the model variable you created when you deployed the model or you can list all of the models in your project.
# + id="evaluate_the_model:mbsdk"
# Get model resource ID
models = aip.Model.list(filter="display_name=chicago_" + TIMESTAMP)
# Get a reference to the Model Service client
client_options = {"api_endpoint": f"{REGION}-aiplatform.googleapis.com"}
model_service_client = aip.gapic.ModelServiceClient(client_options=client_options)
model_evaluations = model_service_client.list_model_evaluations(
parent=models[0].resource_name
)
model_evaluation = list(model_evaluations)[0]
print(model_evaluation)
# + [markdown] id="custom_vs_automl_compare"
# ## Compare metric results with AutoML baseline
#
# Finally, you make a decision if the current experiment produces a custom model that is better than the AutoML baseline, as follows:
# - Compare the evaluation results for each evaluation slice between the custom model and the AutoML model.
# - Weight the results according to your business purposes.
# - Add up the result and make a determination if the custom model is better.
# + [markdown] id="store_model_metadata"
# ### Store evaluation results for custom model
#
# Next, you use the labels field to store user metadata containing the custom metrics information.
# + id="store_model_metadata"
import json
metadata = {}
metadata["train_eval_metrics"] = METRICS
metadata["custom_eval_metrics"] = "[you-fill-this-in]"
with tf.io.gfile.GFile("gs://" + BUCKET_NAME[5:] + "/metadata.jsonl", "w") as f:
json.dump(metadata, f)
# !gsutil cat $BUCKET_NAME/metadata.jsonl
# + [markdown] id="cleanup:mbsdk"
# # Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Dataset
# - Pipeline
# - Model
# - Endpoint
# - AutoML Training Job
# - Batch Job
# - Custom Job
# - Hyperparameter Tuning Job
# - Cloud Storage Bucket
# + id="cleanup:stage2"
delete_all = False
if delete_all:
# Delete the dataset using the Vertex dataset object
try:
if "dataset" in globals():
dataset.delete()
except Exception as e:
print(e)
# Delete the model using the Vertex model object
try:
if "model" in globals():
model.delete()
except Exception as e:
print(e)
if "BUCKET_NAME" in globals():
# ! gsutil rm -r $BUCKET_NAME
| notebooks/community/ml_ops/stage2/mlops_experimentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Youth
# We learn python through a game, taken from [Code Club Projects](https://codeclubprojects.org/en-GB/python/)
#
# +
# First, let the player choose Rock, Paper or Scissors by typing the letter ‘r’, ‘p’ or ‘s’
player = input('rock (r), paper (p) or scissors (s)?')
# what did we just do? we used a built-in function in Python to prompt the user to input a letter in the console
# and we assigned the input to a variable called 'player'. the = symbol indicates that what is on the right is assigned to the variable name on the left
# Now print out what the player chose:
#print(player)
#print('you chose', player)
#print(player, 'vs')
print(player, 'vs', end=' ')
# Second: Computer's Turn
# Use 'randint' function to generate a random number to decide whether the computer has chosen rock, paper or scissors.
# we need to import it from 'random' library
from random import randint
chosen = randint(1,3)
#print(chosen)
#print('computer chose',chosen)
# we like to print the letters not numbers. let's say 1 = rock, 2 = paper, 3=scissors
# we can use if statement to assign letters to whatever number that is selected
if chosen == 1:
computer = 'r' # 'O'
elif chosen == 2:
computer = 'p' #'__'
else:
computer = 's' #'>8'
print(computer)
# notice that we are not inside the if statement because we don't use indentation
# a nicer output:
#print(player, 'vs', computer)
# let's add a code for determining the winner
# we need to compare the 'player' and 'computer' variables
if player == computer:
print('Draw!')
elif player == 'r' and computer == 's':
print('player wins!')
elif player == 'r' and computer =='p':
print('Computer wins!')
elif player == 'p' and computer == 'r':
print('Player wins!')
elif player == 'p' and computer == 's':
print('Computer wins!')
elif player == 's' and computer == 'p':
print('Player wins!')
elif player == 's' and computer == 'r':
print('Computer wins!')
# Challenge: Instead of using the letters r, p and s to represent rock, paper and scissors, can you use ASCII art?
# O for rock, __ for paper, and >8 for scissors
# so now change the lines you print the choices of the player and the computer in ASCII art
#if player == 'r':
# print('O', 'vs', end=' ')
#elif player == 'p':
# print('__', 'vs', end=' ')
#else:
# print('>8', 'vs', end =' ')
if player == 'r':
print('O', 'vs', computer)
elif player == 'p':
print('__', 'vs', computer )
else:
print('>8', 'vs', computer)
# -
# ## summary
# What we learned today:
# - built-in functions like input prompt and print
# - variables
# - numbers and strings
# - libraries and functions like randint
# - if statements
# # review:
# ## Variables
name = 'Sara'
year = 2017
# we can check the type of our variable using the type(variable_name) function
print(type(name))
#str is a string: a sequence of characters.
# ## Numbers
# You can use Python as a calculator:
7 + 8
7*8
8%7
2**4
# Python's order of operations works just like math's:
#
# - Parentheses
# - Exponents
# - Multiplication and division
# - Addition and subtraction
16 ** 0.5
16 ** (1/2)
16 ** 1/2
# # Logic
# We've talked about how Python handles numbers already. Let's now turn to how it handles inequalities – things that are either true or false.
6 > 0
4 == 6
4 <= 6
4 != 6
# you can continue here
# https://docs.trinket.io/getting-started-with-python#/logic/combining-boolean-expressions
# ## Lists
# Python has two array-like things. The first is called a "list", which can hold any data types.
fruits = ['apple', 'banana', 'mango', 'lychee']
print(fruits)
fruits.append('orange')
print(fruits)
# lists don't need to comprise of all the same type
misc = [29, 'dog', fruits]
print(misc)
print(fruits + fruits)
# The second is called a "tuple", which is an immutable list (nothing can be added or subtracted) whose elements also can't be reassigned.
tup1 = (1,2)
print(tup1)
# ## Indexing and Slicing
#
#indexing in Python starts at 0, not 1 (like in Matlab or Oracle)
print(fruits[0])
print(fruits[1])
# +
# strings are just a particular kind of list
s = 'This is a string.'
# -
print(s[0])
# use -1 to get the last element
print(fruits[-1])
print(fruits[-2])
# to get a slice of the string use the : symbol
print(s[0:4])
print(s[:4])
print(s[4:7])
print(s[7:])
print(s[7:len(s)])
# ## For Loops
#
nums = [23, 56, 1, 10, 15, 0]
# +
# in this case, 'n' is a dummy variable that will be used by the for loop
# you do not need to assign it ahead of time
for n in nums:
if n%2 == 0:
print('even')
else:
print('odd')
# -
# for loops can iterate over strings as well
vowels = 'aeiou'
for vowel in vowels:
print(vowel)
# ## Functions
# +
# always use descriptive naming for functions, variables, arguments etc.
def sum_of_squares(num1, num2):
"""
Input: two numbers
Output: the sum of the squares of the two numbers
"""
ss = num1**2 + num2**2
return(ss)
# The stuff inside """ """ is called the "docstring". It can be accessed by typing help(sum_of_squares)
# -
print(sum_of_squares(4,2))
# the return statement in a function allows us to store the output of a function call in a variable for later use
ss1 = sum_of_squares(5,5)
print(ss1)
# ## Useful Packages
# use a package by importing it, you can also give it a shorter alias, in this case 'np'
import numpy as np
array = np.arange(15)
lst = list(range(15))
print(array)
print(lst)
print(type(array))
print(type(lst))
# numpy arrays allow for vectorized calculations
print(array*2)
print(lst*2)
array = array.reshape([5,3])
print(array)
# we can get the mean over all rows (using axis=1)
array.mean(axis=1)
# max value in each column
array.max(axis=0)
import pandas as pd
# +
# this will read in a csv file into a pandas DataFrame
# this csv has data of country spending on healthcare
data = pd.read_csv('health.csv', header=0, index_col=0, encoding="ISO-8859-1")
# -
# the .head() function will allow us to look at first few lines of the dataframe
data.head()
# by default, rows are indicated first, followed by the column: [row, column]
data.loc['Canada', '2008']
# you can also slice a dataframe
data.loc['Canada':'Denmark', '1999':'2001']
# %matplotlib inline
import matplotlib.pyplot as plt
# the .plot() function will create a simple graph for you to quickly visualize your data
data.loc['Denmark'].plot()
data.loc['Canada'].plot()
data.loc['India'].plot()
plt.legend(loc='best')
# # Guessing game
# if we have extra time, we can play another game:
# +
import random
number = random.randint(1, 10)
tries = 0
win = False # setting a win flag to false
name = input("Hello, What is your username?")
print("Hello " + name + "." )
question = input("Would you like to play a game? [Y/N] ")
if question.lower() == "n": #in case of capital letters is entered
print("oh..okay")
exit()
elif question.lower() == "y":
print("I'm thinking of a number between 1 & 10")
while not win: # while the win is not true, run the while loop. We set win to false at the start therefore this will always run
guess = int(input("Have a guess: "))
tries = tries + 1
if guess == number:
win = True # set win to true when the user guesses correctly.
elif guess < number:
print("Guess Higher")
elif guess > number:
print("Guess Lower")
# if win is true then output message
print("Congrats, you guessed correctly. The number was indeed {}".format(number))
print("it had taken you {} tries".format(tries))
# -
| lessons/python/python-for-kids/Python-lesson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Question 2 - Build and Apply a perceptron
#
# For the same set of images and same features, build a perceptron to classify the
# images. Start the training with similar weights to both the features, and then train with the
# 40 samples to build the classifier. Apply the classifier on 5 images.
# +
# Import necessary libraries
import numpy as np
import pandas as pd
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.decomposition import PCA
# -
# ## Build a Perceptron Class
class Perceptron():
# Initialise the weights and the bias
def __init__(self, input_size, epochs=10, learning_rate=1):
self.W = np.zeros(input_size+1)
# add one ot the input shape to account for w0 which is the bias
self.b = 0
# Hyperparameters
self.epochs = epochs
self.lr = learning_rate # Fixed Learning Rate
# Create an activation function which returns 1 for positive outputs and 0 otherwise
def activation(self, x):
return 1 if x >= -0.5 else 0
# Create a prediction function which will take input features and
def get_pred(self, x):
# Get the output as Wt.X
z = np.dot(self.W.T, x)
a = self.activation(z)
return a
# Create a training function which would minimize the error
def fit(self, X, labels):
for e in range(self.epochs):
print("Training Epoch: ", e+1)
for i in range(X.shape[0]):
x = np.insert(X[i], 0, 1)
# insert a one at the beginning of the X vector to pad it
y = self.get_pred(x)
error = labels[i] - y
self.W = self.W + (self.lr * error * x)
# Create a predict function
def predict(self, X):
preds = []
for i in range(X.shape[0]):
x = np.insert(X[i], 0, 1)
y = self.get_pred(x)
preds.append(y)
return np.array(preds)
def get_weights(self):
return self.W
# ## Training Step
# Read the training and testing data
train_data = pd.read_csv('../Datasets/final_data.csv', index_col=0)
test_data = pd.read_csv('../Datasets/test_data.csv', index_col=0)
print(train_data.shape, test_data.shape)
# Assign classes as 0s and 1s
train_data['class'] = train_data['class'].map({ "cat":0, "dog":1 })
test_data['class'] = test_data['class'].map({ "cat":0, "dog":1 })
# +
# Get the training and testing data
X_train = train_data.drop(['class'], axis=1).values
X_test = test_data.drop(['class'], axis=1).values
y_train = train_data['class'].to_numpy()
y_test = test_data['class'].to_numpy()
# +
# Training Phase
perceptron = Perceptron(input_size=X_train.shape[1])
perceptron.fit(X_train, y_train)
# Collect the weights and bias
train_weights = perceptron.get_weights()
print(train_weights)
# -
# ## Testing Step
# Get the predictions
preds = perceptron.predict(X_test)
print("Accuracy of the algorithm:", accuracy_score(y_true=y_test, y_pred=preds))
# ### Testing with different learning rates
# +
learning_rate = 0.01
# Training Phase
perceptron = Perceptron(input_size=X_train.shape[1], learning_rate=learning_rate)
perceptron.fit(X_train, y_train)
# Collect the weights and bias
train_weights = perceptron.get_weights()
# Get the predictions
preds = perceptron.predict(X_test)
print("Accuracy of the algorithm:", accuracy_score(y_true=y_test, y_pred=preds))
# +
learning_rate = 0.1
# Training Phase
perceptron = Perceptron(input_size=X_train.shape[1], learning_rate=learning_rate)
perceptron.fit(X_train, y_train)
# Collect the weights and bias
train_weights = perceptron.get_weights()
# Get the predictions
preds = perceptron.predict(X_test)
print("Accuracy of the algorithm:", accuracy_score(y_true=y_test, y_pred=preds))
# +
learning_rate = 1
# Training Phase
perceptron = Perceptron(input_size=X_train.shape[1], learning_rate=learning_rate)
perceptron.fit(X_train, y_train)
# Collect the weights and bias
train_weights = perceptron.get_weights()
# Get the predictions
preds = perceptron.predict(X_test)
print("Accuracy of the algorithm:", accuracy_score(y_true=y_test, y_pred=preds))
# -
# ### Performing PCA to get image points separation (Additional Steps: Not part of the original question)
#
# Since, there are over 2000 columns, I have to apply PCA to get two components. These two components are just a representation of the whole data and will not be very accurate. To get the slope and the weights, I have done the same and tried to minimize the train weights to single weights for each dimension. I have done plotted a perceptron line for the points. The line seems to be okay and classifies dogs correctly in the data.
# +
# PCA for points
pca = PCA(n_components=2)
principalComponents = pca.fit_transform(X_test)
principalDf = pd.DataFrame(data = principalComponents, columns = ['pc1', 'pc2'])
principalDf = pd.concat([principalDf, test_data['class']], axis=1)
# +
# PCA for weights
pca1 = PCA(n_components=1)
principalComponents = pca1.fit_transform(pd.DataFrame(train_weights[1:].T))
principal_weights = pd.DataFrame(data = principalComponents, columns = ['pc1'])
# -
# Plot the points
sns.scatterplot(x='pc1', y='pc2', hue='class', data=principalDf)
# These points barely seem to be linearly separable and hence, the low accuracy seems justified.
principalDf
# Plot the line
bias = train_weights[0]
weight = principal_weights.values[1]
x = np.linspace(-50,50,100)
y = weight*x + bias
sns.scatterplot(x='pc1', y='pc2', hue='class', data=principalDf)
plt.plot(x, y, '-r')
plt.title("Perceptron Classifier Line")
plt.xlabel('x', color='#1C2833')
plt.ylabel('y', color='#1C2833')
plt.legend(loc='upper left')
plt.grid()
plt.show()
| Single Layer Perceptron/.ipynb_checkpoints/Question2_Notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ungraded Lab Part 2 - Consuming a Machine Learning Model
#
# Welcome to the second part of this ungraded lab!
# **Before going forward check that the server from part 1 is still running.**
#
# In this notebook you will code a minimal client that uses Python's `requests` library to interact with your running server.
import os
import io
import cv2
import requests
import numpy as np
from IPython.display import Image, display
# ## Understanding the URL
#
#
# ### Breaking down the URL
#
# After experimenting with the fastAPI's client you may have noticed that we made all requests by pointing to a specific URL and appending some parameters to it.
#
# More concretely:
#
# 1. The server is hosted in the URL [http://localhost:8000/](http://localhost:8000/).
# 2. The endpoint that serves your model is the `/predict` endpoint.
#
# Also you can specify the model to use: `yolov3` or`yolov3-tiny`. Let's stick to the tiny version for computational efficiency.
#
# Let's get started by putting in place all this information.
base_url = 'http://localhost:8000'
endpoint = '/predict'
model = 'yolov3-tiny'
# To consume your model, you append the endpoint to the base URL to get the full URL. Notice that the parameters are absent for now.
url_with_endpoint_no_params = base_url + endpoint
url_with_endpoint_no_params
# To set any of the expected parameters, the syntax is to add a "?" character followed by the name of the parameter and its value.
#
# Let's do it and check how the final URL looks like:
full_url = url_with_endpoint_no_params + "?model=" + model
full_url
# This endpoint expects both a model's name and an image. But since the image is more complex it is not passed within the URL. Instead we leverage the `requests` library to handle this process.
#
# # Sending a request to your server
#
# ### Coding the response_from_server function
#
# As a reminder, this endpoint expects a POST HTTP request. The `post` function is part of the requests library.
#
# To pass the file along with the request, you need to create a dictionary indicating the name of the file ('file' in this case) and the actual file.
#
# `status code` is a handy command to check the status of the response the request triggered. **A status code of 200 means that everything went well.**
def response_from_server(url, image_file, verbose=True):
"""Makes a POST request to the server and returns the response.
Args:
url (str): URL that the request is sent to.
image_file (_io.BufferedReader): File to upload, should be an image.
verbose (bool): True if the status of the response should be printed. False otherwise.
Returns:
requests.models.Response: Response from the server.
"""
files = {'file': image_file}
response = requests.post(url, files=files)
status_code = response.status_code
if verbose:
msg = "Everything went well!" if status_code == 200 else "There was an error when handling the request."
print(msg)
return response
# To test this function, open a file in your filesystem and pass it as a parameter alongside the URL:
with open("images/clock2.jpg", "rb") as image_file:
prediction = response_from_server(full_url, image_file)
# Great news! The request was successful. However, you are not getting any information about the objects in the image.
#
# To get the image with the bounding boxes and labels, you need to parse the content of the response into an appropriate format. This process looks very similar to how you read raw images into a cv2 image on the server.
#
# To handle this step, let's create a directory called `images_predicted` to save the image to:
dir_name = "images_predicted"
if not os.path.exists(dir_name):
os.mkdir(dir_name)
#
# ### Creating the display_image_from_response function
def display_image_from_response(response):
"""Display image within server's response.
Args:
response (requests.models.Response): The response from the server after object detection.
"""
image_stream = io.BytesIO(response.content)
image_stream.seek(0)
file_bytes = np.asarray(bytearray(image_stream.read()), dtype=np.uint8)
image = cv2.imdecode(file_bytes, cv2.IMREAD_COLOR)
filename = "image_with_objects.jpeg"
cv2.imwrite(f'images_predicted/{filename}', image)
display(Image(f'images_predicted/{filename}'))
display_image_from_response(prediction)
# Now you are ready to consume your object detection model through your own client!
#
# Let's test it out on some other images:
# +
image_files = [
'car2.jpg',
'clock3.jpg',
'apples.jpg'
]
for image_file in image_files:
with open(f"images/{image_file}", "rb") as image_file:
prediction = response_from_server(full_url, image_file, verbose=False)
display_image_from_response(prediction)
# -
# **Congratulations on finishing this ungraded lab!** Real life clients and servers have a lot more going on in terms of security and performance. However, the code you just experienced is close to what you see in real production environments.
# Hopefully, this lab served the purpose of increasing your familiarity with the process of deploying a Deep Learning model, and consuming from it.
#
# **Keep it up!**
# #
# ## Optional Challenge - Adding the confidence level to the request
#
# Let's expand on what you have learned so far. The next logical step is to extend the server and the client so that they can accommodate an additional parameter: the level of confidence of the prediction.
#
# **To test your extended implementation you must perform the following steps:**
#
# - Stop the server by interrupting the Kernel.
# - Extend the `prediction` function in the server.
# - Re run the cell containing your server code.
# - Re launch the server.
# - Extend your client.
# - Test it with some images (either with your client or fastAPI's one).
#
# Here are some hints that can help you out throughout the process:
#
# #### Server side:
# - The `prediction` function that handles the `/predict` endpoint needs an additional parameter to accept the confidence level. Add this new parameter before the `File` parameter. This is necessary because `File` has a default value and must be specified last.
#
#
# - `cv.detect_common_objects` accepts the `confidence` parameter, which is a floating point number (type `float`in Python).
#
#
# #### Client side:
# - You can add a new parameter to the URL by extending it with an `&` followed by the name of the parameter and its value. The name of this new parameter must be equal to the name used within the `prediction` function in the server. An example would look like this: `myawesomemodel.com/predict?model=yolov3-tiny&newParam=value`
#
# ##### Sample Solution:
# - Once you're done with this optional task or if you got stuck while doing it, you can see a sample solution by one of your course mentors [here](https://community.deeplearning.ai/t/c1-w1-optional-challenge-confidence-level/67619). Just make sure you've already joined our Discourse community as shown in an earlier reading item. This is posted in the [MLEP Learner Projects](https://community.deeplearning.ai/c/machine-learning-engineering-for-production/mlep-learner-projects/224) category and feel free to post your own solution (and other content-related projects) there as well. Just remember **not to post any graded material** so as not to violate the Honor Code. You can instead take one of the tools/concepts taught in the lectures or labs then apply it to a mini-project. [Here](https://community.deeplearning.ai/t/fastapi-for-text-classification-problem-in-arabic/56857) is an example. We encourage you to explore your fellow learners' projects and comment on the ones you find interesting.
#
#
# **You can do it!**
| course1/week1-ungraded-lab/client.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from data_cleaning import clean_defender_data
from flask import *
import pandas as pd
app = Flask(__name__)
@app.route("/tables")
def show_tables():
DefenderDF = clean_defender_data()
DefenderDF.set_index(['player_id'], inplace=True)
DefenderDF.index.name=None
return render_template('defenderDF.html',tables=[DefenderDF.to_html(classes='defenders')],
titles = ['na', 'Defenders'])
if __name__ == "__main__":
app.run(debug=True)
# -
| FantasyPL-Flask/Untitled1.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Classification on Iris dataset with sklearn and DJL
//
// In this notebook, you will try to use a pre-trained sklearn model to run on DJL for a general classification task. The model was trained with [Iris flower dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set).
//
// ## Background
//
// ### Iris Dataset
//
// The dataset contains a set of 150 records under five attributes - sepal length, sepal width, petal length, petal width and species.
//
// Iris setosa | Iris versicolor | Iris virginica
// :-------------------------:|:-------------------------:|:-------------------------:
//  |  | 
//
// The chart above shows three different kinds of the Iris flowers.
//
// We will use sepal length, sepal width, petal length, petal width as the feature and species as the label to train the model.
//
// ### Sklearn Model
//
// You can find more information [here](http://onnx.ai/sklearn-onnx/). You can use the sklearn built-in iris dataset to load the data. Then we defined a [RandomForestClassifer](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html) to train the model. After that, we convert the model to onnx format for DJL to run inference. The following code is a sample classification setup using sklearn:
//
// ```python
// # Train a model.
// from sklearn.datasets import load_iris
// from sklearn.model_selection import train_test_split
// from sklearn.ensemble import RandomForestClassifier
// iris = load_iris()
// X, y = iris.data, iris.target
// X_train, X_test, y_train, y_test = train_test_split(X, y)
// clr = RandomForestClassifier()
// clr.fit(X_train, y_train)
// ```
//
//
// ## Preparation
//
// This tutorial requires the installation of Java Kernel. To install the Java Kernel, see the [README](https://github.com/deepjavalibrary/djl/blob/master/jupyter/README.md).
//
// These are dependencies we will use. To enhance the NDArray operation capability, we are importing ONNX Runtime and PyTorch Engine at the same time. Please find more information [here](https://github.com/deepjavalibrary/djl/blob/master/docs/onnxruntime/hybrid_engine.md#hybrid-engine-for-onnx-runtime).
// +
// // %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// %maven ai.djl:api:0.12.0
// %maven ai.djl.onnxruntime:onnxruntime-engine:0.12.0
// %maven ai.djl.pytorch:pytorch-engine:0.12.0
// %maven org.slf4j:slf4j-api:1.7.26
// %maven org.slf4j:slf4j-simple:1.7.26
// %maven com.microsoft.onnxruntime:onnxruntime:1.4.0
// %maven ai.djl.pytorch:pytorch-native-auto:1.8.1
// -
import ai.djl.inference.*;
import ai.djl.modality.*;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.*;
import ai.djl.repository.zoo.*;
import ai.djl.translate.*;
import java.util.*;
// ## Step 1 create a Translator
//
// Inference in machine learning is the process of predicting the output for a given input based on a pre-defined model.
// DJL abstracts away the whole process for ease of use. It can load the model, perform inference on the input, and provide
// output. DJL also allows you to provide user-defined inputs. The workflow looks like the following:
//
// 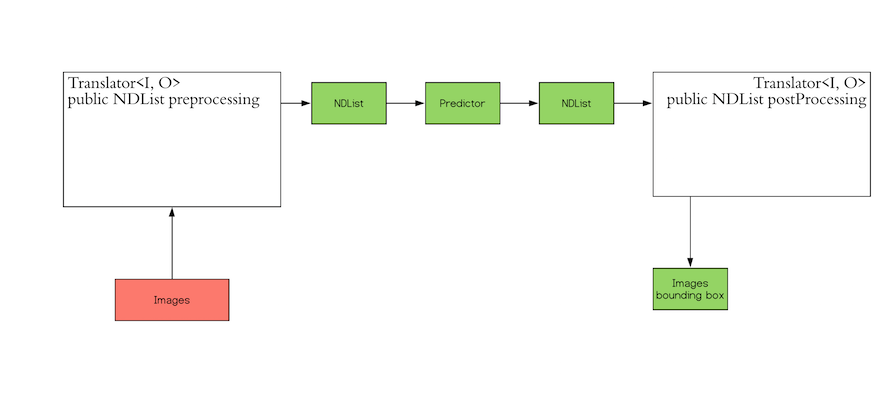
//
// The `Translator` interface encompasses the two white blocks: Pre-processing and Post-processing. The pre-processing
// component converts the user-defined input objects into an NDList, so that the `Predictor` in DJL can understand the
// input and make its prediction. Similarly, the post-processing block receives an NDList as the output from the
// `Predictor`. The post-processing block allows you to convert the output from the `Predictor` to the desired output
// format.
//
// In our use case, we use a class namely `IrisFlower` as our input class type. We will use [`Classifications`](https://javadoc.io/doc/ai.djl/api/latest/ai/djl/modality/Classifications.html) as our output class type.
public static class IrisFlower {
public float sepalLength;
public float sepalWidth;
public float petalLength;
public float petalWidth;
public IrisFlower(float sepalLength, float sepalWidth, float petalLength, float petalWidth) {
this.sepalLength = sepalLength;
this.sepalWidth = sepalWidth;
this.petalLength = petalLength;
this.petalWidth = petalWidth;
}
}
// Let's create a translator
public static class MyTranslator implements Translator<IrisFlower, Classifications> {
private final List<String> synset;
public MyTranslator() {
// species name
synset = Arrays.asList("setosa", "versicolor", "virginica");
}
@Override
public NDList processInput(TranslatorContext ctx, IrisFlower input) {
float[] data = {input.sepalLength, input.sepalWidth, input.petalLength, input.petalWidth};
NDArray array = ctx.getNDManager().create(data, new Shape(1, 4));
return new NDList(array);
}
@Override
public Classifications processOutput(TranslatorContext ctx, NDList list) {
return new Classifications(synset, list.get(1));
}
@Override
public Batchifier getBatchifier() {
return null;
}
}
// ## Step 2 Prepare your model
//
// We will load a pretrained sklearn model into DJL. We defined a [`ModelZoo`](https://javadoc.io/doc/ai.djl/api/latest/ai/djl/repository/zoo/ModelZoo.html) concept to allow user load model from varity of locations, such as remote URL, local files or DJL pretrained model zoo. We need to define `Criteria` class to help the modelzoo locate the model and attach translator. In this example, we download a compressed ONNX model from S3.
String modelUrl = "https://mlrepo.djl.ai/model/tabular/softmax_regression/ai/djl/onnxruntime/iris_flowers/0.0.1/iris_flowers.zip";
Criteria<IrisFlower, Classifications> criteria = Criteria.builder()
.setTypes(IrisFlower.class, Classifications.class)
.optModelUrls(modelUrl)
.optTranslator(new MyTranslator())
.optEngine("OnnxRuntime") // use OnnxRuntime engine by default
.build();
ZooModel<IrisFlower, Classifications> model = criteria.loadModel();
// ## Step 3 Run inference
//
// User will just need to create a `Predictor` from model to run the inference.
Predictor<IrisFlower, Classifications> predictor = model.newPredictor();
IrisFlower info = new IrisFlower(1.0f, 2.0f, 3.0f, 4.0f);
predictor.predict(info);
| jupyter/onnxruntime/machine_learning_with_ONNXRuntime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# !head -n 1000 violations.csv > small-violations.csv
# # 1. I want to make sure my Plate ID is a string. Can't lose the leading zeroes!
plate_info = {'Plate ID': 'str'}
df = pd.read_csv("small-violations.csv", dtype=plate_info)
df
df.head()
df.head(10)
df.tail()
# # 2. I don't think anyone's car was built in 0AD. Discard the '0's as NaN.
plate_info = {'Plate ID': 'str'}
df = pd.read_csv("small-violations.csv", dtype=plate_info, na_values={'Vehicle Year': '0', 'Date First Observed': '0'})
df.head()
# # 3. I want the dates to be dates! Read the read_csv documentation to find out how to make pandas automatically parse dates.
import dateutil
def date_to_date(date):
date = str(date)
parsed_date = dateutil.parser.parse(date)
return parsed_date
df.columns
df['New Issue Date']= df['Issue Date'].apply(date_to_date)
import datetime
def convert_to_time(time):
try:
str_time = str(time)
return datetime.datetime.strptime(str_time, "%Y%m%d")
except:
return None
other_df = df[df['Vehicle Expiration Date'] != 0]
other_df.head()
other_df['New Vehicle Expiration Date']= other_df['Vehicle Expiration Date'].apply(convert_to_time)
other_df.head()
# # 4. "Date first observed" is a pretty weird column, but it seems like it has a date hiding inside. Using a function with .apply, transform the string (e.g. "20140324") into a Python date. Make the 0's show up as NaN.
other_df.columns
other_df['Date First Observed'].dtypes
other_df['Date First Observed'].tail()
import dateutil
other_df['Date First Observed']
other_df['Violation Time'].head()
other_df['Violation Time'].tail()
def int_to_date(integer):
if not pd.isnull(integer):
date = str(int(integer))
parsed_date = dateutil.parser.parse(date)
return parsed_date.strftime("%Y-%-m-%d")
other_df['Date First Observed'].apply(int_to_date)
# # 5. "Violation time" is... not a time. Make it a time.
def violation_time_to_time(time):
try:
hour = time[0:2]
minutes = time[2:4]
am_pm= time[4]
regular_time= hour + ":" + minutes + " " + am_pm + 'm'
violation_time_fixed = dateutil.parser.parse(regular_time)
return violation_time_fixed.strftime("%H:%M%p")
except:
return None
other_df['Violation Time'].apply(violation_time_to_time)
# # 6. There sure are a lot of colors of cars, too bad so many of them are the same. Make "BLK" and "BLACK", "WT" and "WHITE", and any other combinations that you notice.
other_df['Vehicle Color'].value_counts()
# +
def color_rename(color):
if (color == 'BLACK') or (color == 'BLK') or (color == 'BK'):
return 'BLACK'
elif (color == 'WHITE') or (color == 'WHT') or (color == 'WH') or (color == 'W'):
return 'WHITE'
other_df['Vehicle Color'].apply(color_rename)
# -
# # 7. Join the data with the Parking Violations Code dataset from the NYC Open Data site.
parking_violations_df = pd.read_csv("DOF_Parking_Violation_Codes.csv", encoding="mac_roman", error_bad_lines=False)
parking_violations_df.head()
parking_violations_df['CODE'].describe()
other_df['Violation Code'].describe()
def convert_to_str(n):
return str(n)
parking_violations_df['Code'] = parking_violations_df['CODE'].apply(convert_to_str)
other_df['Violation code'] = other_df['Violation Code'].apply(convert_to_str)
parking_violations_df.head()
updated_parking_violations_df = parking_violations_df.rename(columns={'Manhattan  96th St. & below': 'Manhattan 96th & below', 'All Other Areas': 'All other areas'})
updated_parking_violations_df.head()
other_df.head()
diff_violations_df = pd.merge(other_df, updated_parking_violations_df, left_on='Violation code', right_on='Code')
diff_violations_df.head()
# # 8. How much money did NYC make off of parking violations?
diff_violations_df['Manhattan 96th & below'].describe()
diff_violations_df['All other areas'].describe()
diff_violations_df['Manhattan 96th & below'].apply(convert_to_str).head()
diff_violations_df['All other areas'].apply(convert_to_str).head()
diff_violations_df = new_violations_df[new_violations_df['Manhattan 96th & below'] != 'vary']
diff_violations_df.head()
import re
def strip_and_convert_to_int(string):
match = re.findall(r"^\$?\d*", string)
if match:
new_string = string.replace("$", "").split()
new_int = int(new_string[0])
return new_int
else:
return None
diff_violations_df['Manhattan 96th and below'] = diff_violations_df['Manhattan 96th & below'].apply(strip_and_convert_to_int)
diff_violations_df.head()
diff_violations_df['All Other Areas'] = diff_violations_df['All other areas'].apply(strip_and_convert_to_int)
diff_violations_df.tail()
diff_violations_df['All Other Areas'].value_counts().head()
manhattan_violations = diff_violations_df.groupby('Violation code')['All Other Areas'].sum()
manhattan_violations.sum()
violations_not_man = diff_violations_df.groupby('Violation code')['Manhattan 96th and below'].sum()
violations_not_man.sum()
violations_revenue = violations_not_man.sum() + manhattan_violations.sum()
violations_revenue
# # 9. What's the most lucrative kind of parking violation? The most frequent?
manhattan_violations.sort_values(ascending=False)
violations_not_man.sort_values(ascending=False)
new_violations_df['Violation code'].value_counts()
# # 10. New Jersey has bad drivers, but does it have bad parkers, too? How much money does NYC make off of all non-New York vehicles?
out_of_staters_df = diff_violations_df[diff_violations_df['Registration State'] != 'NY']
out_of_staters_df.head()
out_of_staters_other = out_of_staters_df.groupby('Violation code')['All Other Areas'].sum()
out_of_staters_other.sum()
out_of_staters_manhattan= out_of_staters_df.groupby('Violation code')['Manhattan 96th and below'].sum()
out_of_staters_manhattan.sum()
total_out_of_staters_violations = out_of_staters_other.sum()+ out_of_staters_manhattan.sum()
total_out_of_staters_violations
# # 11. Make a chart of the top few.
# %matplotlib inline
out_of_staters_other.sort_values(ascending=False).plot(kind='bar', x='Violation code')
out_of_staters_manhattan.sort_values(ascending=False).plot(kind='bar', x='Violation code')
# # 12. What time of day do people usually get their tickets? You can break the day up into several blocks - for example 12am-6am, 6am-12pm, 12pm-6pm, 6pm-12am.
# # 13. What's the average ticket cost in NYC?
average_tix_price = total_out_of_staters_violations / diff_violations_df['Violation code'].value_counts().sum()
average_tix_price
# # 14. Make a graph of the number of tickets per day.
diff_violations_df['Issue Date'].value_counts().head(10).plot(kind='barh')
# # 15. Make a graph of the amount of revenue collected per day.
daily_revenue = total_out_of_staters_violations / new_violations_df['New Issue Date'].value_counts()
daily_revenue.sort_values(ascending=False).head(20).plot(kind='bar')
#
# # 16. Manually construct a dataframe out of https://dmv.ny.gov/statistic/2015licinforce-web.pdf (only NYC boroughts - bronx, queens, manhattan, staten island, brooklyn), having columns for borough name, abbreviation, and number of licensed drivers.
nyc_licenses = pd.read_excel("NYC.xlsx")
nyc_licenses
# # 17. What's the parking-ticket-$-per-licensed-driver in each borough of NYC? Do this with pandas and the dataframe you just made, not with your head!
diff_violations_df.columns
diff_violations_df['Violation County'].value_counts()
bronx_violations = diff_violations_df[diff_violations_df['Violation County'] == 'BX']
bronx_licenses = nyc_licenses['Total'][nyc_licenses['Abbreviation'] == 'BX']
bronx_tix = bronx_violations.groupby('Violation code')['All Other Areas'].sum()
driver_bronx_tix = bronx_licenses / bronx_tix.sum()
driver_bronx_tix
queens_violations = diff_violations_df[diff_violations_df['Violation County'] == 'Q']
queens_licenses = nyc_licenses['Total'][nyc_licenses['Abbreviation'] == 'Q']
queens_tix = queens_violations.groupby('Violation code')['All Other Areas'].sum()
driver_queens_tix = queens_licenses / queens_tix.sum()
driver_queens_tix
ny_violations = diff_violations_df[diff_violations_df['Violation County'] == 'NY']
ny_licenses = nyc_licenses['Total'][nyc_licenses['Abbreviation'] == 'NY']
ny_tix = ny_violations.groupby('Violation code')['All Other Areas'].sum()
driver_ny_tix = ny_licenses / ny_tix.sum()
driver_ny_tix
brooklyn_violations = diff_violations_df[diff_violations_df['Violation County'] == 'R']
brooklyn_licenses = nyc_licenses['Total'][nyc_licenses['Abbreviation'] == 'R']
brooklyn_tix = brooklyn_violations.groupby('Violation code')['All Other Areas'].sum()
driver_brooklyn_tix = brooklyn_licenses / brooklyn_tix.sum()
driver_brooklyn_tix
staten_is_violations = diff_violations_df[diff_violations_df['Violation County'] == 'K']
staten_is_licenses = nyc_licenses['Total'][nyc_licenses['Abbreviation'] == 'K']
staten_is_tix = violations_kings.groupby('Violation code')['All Other Areas'].sum()
driver_staten_is_tix = staten_is_licenses / staten_is_tix.sum()
driver_staten_is_tix
| Homework 11 Soma.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pymc3 as pm
import tqdm
import spacepy.toolbox as tb
# %matplotlib inline
# -
# # Setup a convolution data set and try and doconvolve it
#
# +
np.random.seed(8675309)
dat_len = 100
xval = np.arange(dat_len)
realdat = np.zeros(dat_len, dtype=int)
realdat[40:60] = 50
noisemean = 2
real_n = np.zeros_like(realdat)
for i in range(len(realdat)):
real_n[i] = np.random.poisson(realdat[i]+noisemean)
# make a detector
# triangular with FWFM 5 and is square
det = np.array([1,1,1,1,1])
# the numpy convolve I don't understand the normalization
obs = np.convolve(real_n, det, mode='same')
obs = tb.normalize(obs)
obs *= real_n.max()
# Two subplots, the axes array is 1-d
f, axarr = plt.subplots(4, sharex=True)
axarr[0].plot(xval, realdat)
axarr[0].set_ylabel('Truth')
axarr[0].set_ylim((0,60))
axarr[1].plot(xval, real_n)
axarr[1].set_ylabel('T+N')
axarr[2].plot(np.arange(len(det)), det)
axarr[2].set_ylabel('Det')
axarr[3].plot(xval, obs)
axarr[3].set_ylabel('Obs')
# -
# So the det provides a point spread function that is U(0,5)
# +
# generate some data
with pm.Model() as model:
truth_mc = pm.Uniform('truth', 0, 100, shape=dat_len)
noisemean_mc = pm.Uniform('noisemean', 0, 100)
noise_mc = pm.Poisson('noise', noisemean_mc, observed=obs[1:20])
real_n_mc = pm.Poisson('real_n', truth_mc+noisemean_mc, shape=dat_len)
psf = pm.Uniform('psf', 0, 5, observed=det)
obs_mc = pm.Normal('obs', (truth_mc+noisemean_mc)*psf.max(), 1/5**2, observed=obs, shape=dat_len)
trace = pm.sample(5000)
# -
pm.traceplot(trace)
pm.summary(trace)
# +
# plt.plot(trace['truth'][0:5,:].T)
trace['truth'].shape
iqr = np.zeros((dat_len,3))
for i in range(dat_len):
iqr[i] = np.percentile(trace['truth'].T[i], (25,50,75), axis=0)
plt.plot(xval, iqr[:,1], label='recovered')
plt.fill_between(xval, iqr[:,0], iqr[:,2], alpha=0.2)
plt.plot(xval, real_n, c='r', label='pre psf')
plt.plot(xval, realdat, c='g', label='truth')
plt.plot(xval, obs, c='k', label='observed', lw=3)
plt.legend()
plt.figure()
snr = iqr[:,1]/(iqr[:,2], iqr[:,0])
perixval.shape, snr.shape
plt.plot(xval, snr)
# -
print(np.percentile(trace['noisemean'], (25,50,75)), noisemean)
obs[1:20]
| Deconvolution/convolution1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem 250 - 250250
#
# Find the number of non-empty subsets of $\{1^1, 2^2, 3^3, \ldots, 250250^{250250}\}$, the sum of whose elements is divisible by 250. Enter the rightmost 16 digits as your answer.
a = [1] + [0]*249
for i in range(1, 250251):
n = pow(i, i, 250)
b = [0] * 250
for j in range(250):
b[j] = (a[j] + a[j-n]) % 10**16
a = b
print(a[0] - 1)
| Euler 250 - 250250.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # ESTUDIO DE CASO: PREDICCIÓN DE ENFERMEDAD DEL CORAZÓN
# 1- Demostrar conocimiento organizacional o de contexto
# 2- Demostrar conocimiento detallado de los conjuntos de datos correspondientes
# 3- Demostrar capacidad en la preparación previa de los datos
#
#
# 4- Algoritmos y modelos
#
# En cuanto al algoritmo de aprendizaje para generar el modelo de predicción de enfermedades del corazón, claramente debemos considerar los algoritmos supervisados de clasificación. Dentro de estos, se podría justificar no utilizar knn por un consumo alto de memoria debido a que dataset de entrenamiento es uno grande, pero con los bajos costos de memoria de hoy en día, este no debería ser un impedimento para un conjunto de datos que pese un par de megas. Utilizaría Naive Bayes al principio como estándar o benchmark para comparar con otros modelos más complejos. Luego de obtener las métricas para el modelo de Naive Bayes, utilizaré difentes tipos de ensambles inicialmente Random Forest y alguna variante de Boosted Trees como XGBoost debido a su alta performance para casos de la industria estudiada. Además, como otra justificación para utilizar estos métodos, no existe un requerimiento de que las predicciones tengan que ser extremadamente rápidas, por lo que a utilización de un modelo de Random Forest con miles de árboles no sería un problema. Sería interesante también estudiar la performance de SVMs con distintos kernels para este caso particular.
# Utilizando los estudios Citados por el paper "Heart Failure: Diagnosis, Severity Estimation and Prediction of Adverse
# Events Through Machine Learning Techniques" (<NAME>, <NAME>, <NAME>, <NAME>, <NAME>), Diversos grupos de investigadores utilizaron diferentes algoritmos de ML, por lo que elegir uno a priori sin probar una gran variedad de ellos sería un enfoque bastante ingenuo y sesgado en mi opinión. Debido a esto, utilizaré una variedad de algoritmos y elegiré el que mejor se adecúe al dataset elegido.
#
| Caso de estudio 3/caso_de_estudio_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pickle
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from importlib import reload
import sys
from scipy.stats import binned_statistic
import warnings
from scipy.optimize import minimize
warnings.filterwarnings("ignore")
# +
sys.path.insert(1,'../../snowmicropyn')
import snowmicropyn
print(snowmicropyn.__file__)
from snowmicropyn import density_ssa
# -
# # Read in mCT data
# +
def get_mct_frame(site):
frames = pickle.load(open('../data/microCT/processed_mCT.p', 'rb'))
mct_df = frames[site]
mct_df = mct_df.drop_duplicates(subset='height_ave (cm)',keep="first")
mct_df.set_index('height_ave (cm)', inplace=True)
return(mct_df)
site = '2N13'
mct_df = get_mct_frame(site)
mct_df
# +
fig,ax = plt.subplots()
ax.plot(mct_df['Equiv. Diam (mm)'],
mct_df.index)
ax.set_xlabel('Equiv. Diam (mm)')
ax.set_ylabel('Height above snow-soil interface [cm]')
ax.set_title(f'Site {site.upper()}')
plt.show()
# return df to get a look at it
mct_df
# -
# # Read SMP data
# +
p = snowmicropyn.Profile.load(f'../data/SMP/SNEX20_SMP_S19M1172_2N13_20200206.PNT')
p.detect_ground()
# -
# # Sensitivity analysis to window size
# +
C20_ssa_coeffs = [0.57, -18.56, -3.66]
C20_coeffs = {'density': [295.8, 65.1, -43.2, 47.1],
'ssa':C20_ssa_coeffs,
'equation':'ssa'}
P15_ssa_coeffs = [0.131, 0.355, 0.0291]
P15_coeffs = {'density':[420.47, 102.47, -121.15, -169.96],
'ssa':P15_ssa_coeffs,
'equation':'l_ex'}
C20_window = 1
P15_window = 2.5
# +
def get_P15_C20(p):
df_C20 = snowmicropyn.density_ssa.calc(p.samples,
coeff_model=C20_coeffs,
window=C20_window,
overlap=50)
df_P15 = snowmicropyn.density_ssa.calc(p.samples,
coeff_model=P15_coeffs,
window=P15_window,
overlap=50)
###################################################
df_C20.columns = ['distance', 'Cdensity', 'Cssa']
df_P15.columns = ['distance', 'Pdensity', 'Pssa']
###################################################
df_combined = pd.merge(left=df_P15, right=df_C20)
df_combined = df_combined[(df_combined['distance'] < p.ground)]
df_combined['distance_up'] = (p.ground - df_combined['distance'])/10
df_combined.set_index('distance_up',inplace=True)
df_combined.sort_index()
df_combined.dropna(how='any')
return df_combined
PC_df = get_P15_C20(p)
PC_df
# +
def resample_PC_to_mCT(mct_df, PC_df):
for code in ['Cssa', 'Pssa']:
mct_df[code] = [np.nanmean(PC_df[code][(PC_df.index < u) & (PC_df.index > l)])
for l, u in zip(mct_df['height_min (cm)'], mct_df['height_max (cm)']) ]
return mct_df
com_df = resample_PC_to_mCT(mct_df, PC_df)
com_df
# +
fig,ax = plt.subplots(figsize=(8,5))
ax.plot(com_df['Pssa'], com_df.index,label='P2015')
ax.plot(com_df['Cssa'], com_df.index,label='C2020')
ax.plot(com_df['SSA (m2/kg)'],
com_df.index,
label='mCT',
color='k',
zorder=5)
ax.set_xlabel('SSA (m2/kg)', fontsize='x-large')
ax.set_ylabel('Height above snow-soil interface [cm]', fontsize='x-large')
ax.set_title(f'Site {site.upper()}', fontsize='x-large')
ax.legend(loc='lower right', fontsize='x-large')
plt.show()
# +
def get_RMSE(mct_df):
CRMSE = np.sqrt(np.nanmean(np.square(mct_df['SSA (m2/kg)']-mct_df['Cssa'])))
PRMSE = np.sqrt(np.nanmean(np.square(mct_df['SSA (m2/kg)']-mct_df['Pssa'])))
return {'CRMSE':CRMSE,
'PRMSE':PRMSE}
get_RMSE(com_df)
# -
# # Now iterate coefficients to find best fit
# +
def get_ssa(p, coeffs_dict):
df = snowmicropyn.density_ssa.calc(p.samples,
coeff_model=coeffs_dict,
window=C20_window,
overlap=50)
df = df[(df['distance'] < p.ground)]
df['distance_up'] = (p.ground - df['distance'])/10
df.set_index('distance_up',inplace=True)
df.sort_index()
df.dropna(how='any')
return df
def resample_SMP_to_mCT(mct_df, df):
com_df = mct_df.copy()
com_df['ssa'] = [np.nanmean(df['ssa'][(df.index < u) & (df.index > l)])
for l, u in zip(mct_df['height_min (cm)'], mct_df['height_max (cm)']) ]
return com_df
# +
def compare_smp_to_mct(coeffs_list,mct_df,p,eqn='ssa'):
coeffs_dict = {'density': [295.8, 65.1, -43.2, 47.1],
'ssa':coeffs_list,
'equation':eqn}
df = get_ssa(p, coeffs_dict)
# print(df)
com_df = resample_SMP_to_mCT(mct_df, df)
# print(com_df)
return(com_df)
def calc_RMSE(coeffs_list,mct_df,p,eqn='ssa'):
com_df = compare_smp_to_mct(coeffs_list,mct_df,p,eqn)
RMSE = np.sqrt(np.nanmean(np.square(com_df['SSA (m2/kg)']-com_df['ssa'])))
print(RMSE)
return RMSE
# +
site = '2N13'
mct_df = get_mct_frame(site)
RMSE = calc_RMSE(C20_ssa_coeffs,mct_df,p)
# +
initial_guess = C20_ssa_coeffs
min_output = minimize(fun=calc_RMSE,
x0=initial_guess,
args=(mct_df, p))
# -
# # Plot the results of the minimization exercise
# +
ssa_coeffs_min = list(min_output.x)
min_df = compare_smp_to_mct(ssa_coeffs_min,mct_df,p)
#############
C20_coeffs = [0.57, -18.56, -3.66]
C20_df = compare_smp_to_mct(C20_coeffs,mct_df,p)
# ################
P15_coeffs = [0.131, 0.355, 0.0291]
P15_df = compare_smp_to_mct(P15_coeffs,mct_df,p,eqn='l_ex')
# +
plt.plot(min_df['ssa'],min_df.index,label='minimized coeffs')
plt.plot(P15_df['ssa'],min_df.index,label='P2015')
plt.plot(C20_df['ssa'],min_df.index,label='C2020')
plt.plot(min_df['SSA (m2/kg)'],min_df.index,label='micro-CT')
plt.xlabel('SSA (m2/kg)', fontsize='x-large')
plt.ylabel('Height above snow-\nsoil interface [cm]', fontsize='x-large')
plt.legend()
| notebooks/Compare_mCT_vs_SMP_SSA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # [Species Name] Records from GBIF and/or eBird
# +
# Specify some paths and names in this cell.
# Path to folder where you saved your wildlifeconfig file.
config_path = ""
# Path to an existing filter set
filter_set_json = ""
# Path to taxa concept JSON
taxon_json = ""
# Your unique, informative name for the query
query_name = ""
# Path to R install (e.g,"C:/Miniconda3/envs/wrangler040/lib/R" or "~/Miniconda3/envs/wrangler040/lib/R")
R_home = ""
ask_eBird = True
ask_GBIF = True
# +
# Nothing to fill out in this cell
# %matplotlib inline
import sys
import os
import sqlite3
import pprint
import json
import pandas as pd
import geopandas as gpd
import numpy as np
import matplotlib.pyplot as plt
from datetime import datetime
sys.path.append(config_path)
if os.path.exists(config_path + "wranglerconfig.py") is True:
import wranglerconfig as config
else:
print("!!! wranglerconfig.ply was not found")
sys.path.append(config.codeDir)
import wrangler_functions as functions
# Define some variables
t1 = datetime.now()
working_directory = config.workDir
temp_directory = config.tempDir
username = config.gbif_username
password = <PASSWORD>
email = config.gbif_email
EBD_file = config.EBD_file
output_database = working_directory + query_name + '.sqlite'
os.environ["TMP"] = temp_directory
os.environ["TEMP"] = temp_directory
# Set display options
pd.set_option('display.max_colwidth', 45)
pd.set_option('display.max_rows', 150)
print("Notebook run " + str(t1))
print("Results were saved in " + output_database)
# -
# ## Taxon Concept
your_taxon_id = None
gbif_id = None
ebird_id = None
detection_distance_m = None
taxon_polygon = None
# +
# If a json was provided, use it, otherwise create a new one with info that was provided.
if taxon_json is None:
# Build a species dictionary
taxon_info = {"ID": your_taxon_id, "GBIF_ID": gbif_id, "EBIRD_ID": ebird_id, "detection_distance_m": detection_distance_m,
"TAXON_EOO": taxon_polygon}
# Save as json object
out_file = open(working_directory + your_taxon_id + ".json", "w")
json.dump(taxon_info, out_file)
out_file.close()
if taxon_json is not None:
with open(taxon_json, "r") as f:
taxon_info = json.load(f)
f.close()
pprint.pprint(taxon_info)
# -
# ## Filter Set
filter_set_name = "" # What to name your filter set
print("Filter set name: " + str(filter_set_name))
# #### GBIF Request Method
"""
Whether to get records from GBIF in a darwin core archive.
"False" uses the GBIF API, which has limitations that may be important.
"True" requests results be emailed in a darwin core archive.
"""
get_dwca = True
print("Request a Darwin Core Archive? " + str(get_dwca))
# #### Date Limits
# Notes:
'''
Enter year and month ranges. For example, years_range = 2015,2017 and months_range = 3,6
'''
years_range = "2000,2021"
months_range = "1,12"
print("Years: " + str(years_range))
print("Months: " + str(months_range))
# #### Country
# Notes:
country = None
print("Country: " + str(country))
# #### Bounding Box
# Notes:
'''
Coordinates should correspond to WGS84 (EPSG:4326). Don't use this option if you specify a query polygon below.
'''
lat_range = None
lon_range = None
print("Latitude range: " + str(lat_range))
print("Longitude range: " + str(lon_range))
# #### Area of Interest
# Notes:
'''
Specify polygons to use for spatial filtering.
Records with coordinates outside of the polygons will be removed.
You can specify a geometry for the query and one for the species.
The species geometry is included to facilitate better handling of taxonomic issues.
If both are provided, the intersection is calculated and used as the filter.
The format should be well known text in WGS84 (EPSG 4326), and very importantly, vertices need to be listed counter-clockwise.
See the ccw_wkt_from_shp() function in wrangler functions for help.
'''
query_polygon = None
print(query_polygon)
# #### Taxon EOO
# Notes:
'''
True or False whether you want to apply the taxon EOO to the filtering.
If True, removes records with centroids outside of the extent of occurrence geometry you provided in taxon_info.
'''
use_taxon_geometry = False
print("Use taxon extent of occurrence? " + str(use_taxon_geometry))
# #### Geoissue
# Notes:
'''
Get only records with geospatial issues? GBIF only.
'''
geoissue = False
print("Records with geoissues OK? " + str(geoissue))
# #### Collections
# Notes:
'''
List collection codes that you'd like to omit. GBIF only.
'''
collection_codes_omit = None
print("Omit: " + str(collection_codes_omit))
# #### Institutions
# Notes:
'''
List institution codes that you'd like to omit. GBIF only
'''
institutions_omit = None
print("Omit: " + str(institutions_omit))
# #### Datasets
# Notes:
'''
List datasets that you'd like to omit.
'''
datasets_omit = None
print("Omit: " + str(datasets_omit))
# #### Coordinate Uncertainty
# Notes:
'''
has_coordinate_uncertainty -- Do you want to remove records without coordinate uncertainty (True) or leave them in the data set
(False)? Note that eBird records in GBIF (EOD) do not have this and nor do EBD data.
default_coordUncertainty -- coordinateUncertaintyInMeters is often not provided.
Here is an option to use a default. If you don't want anything entered, set this as None. Note that when the value is
None, georeferences will be approximated with information from other field. See the User's Guide for more information.
max_coordinate_uncertainty -- A maximum for coordinate uncertainty can also be set in meters. It should be an integer
greater than zero.
'''
has_coordinate_uncertainty = False
default_coordUncertainty = None
max_coordinate_uncertainty = 100000
print("Coordinate uncertainty required? " + str(has_coordinate_uncertainty))
print("Default coordinate uncertainty to use: " + str(default_coordUncertainty))
print("Maximum allowable coordinate uncertainty: " + str(max_coordinate_uncertainty))
# #### Bases
# Notes:
'''
List bases of records that you want to omit. GBIF only.
'''
bases_omit = None
print("Omit: " + str(bases_omit))
# #### Sampling Protocols
# Notes:
'''
List sampling protocols that you would like to omit.
'''
sampling_protocols_omit = None
print("Omit: " + str(sampling_protocols_omit))
# #### Issues
# Notes:
'''
List issues that you want to omit. GBIF only.
'''
issues_omit = None
print("Omit: " + str(issues_omit))
# #### Duplicates
# Notes:
'''
Specify whether duplicates on latitude, longitude, and date should be included.
'''
duplicate_coord_date_OK = True
print("Allow duplicates? " + str(duplicate_coord_date_OK))
# #### Filter Set Summary
# +
if filter_set_json is None:
# Build a filter set dictionary
filter_set = {"name": filter_set_name, "query_polygon": query_polygon, "issues_omit": issues_omit,
"sampling_protocols_omit": sampling_protocols_omit, "bases_omit": bases_omit,
"has_coordinate_uncertainty": has_coordinate_uncertainty, "geoissue": geoissue,
"default_coordUncertainty": default_coordUncertainty,
"max_coordinate_uncertainty": max_coordinate_uncertainty,
"datasets_omit": datasets_omit, "collection_codes_omit": collection_codes_omit,
"institutions_omit": institutions_omit, "geoissue": geoissue, "use_taxon_geometry": use_taxon_geometry,
"lat_range": lat_range, "lon_range": lon_range, "country": country,
"years_range": years_range, "months_range": months_range, "duplicate_coord_date_OK": duplicate_coord_date_OK, "get_dwca": get_dwca}
# Replace empty strings with None
for x in filter_set.keys():
if filter_set[x] == "":
filter_set[x] = None
# Save as json object
with open(working_directory + filter_set_name + ".json", "w") as f:
json.dump(filter_set, f)
f.close()
if filter_set_json is not None:
with open(filter_set_json, "r") as f:
filter_set = json.load(f)
f.close()
# Replace empty strings with None
for x in filter_set.keys():
if filter_set[x] == "":
filter_set[x] = None
pprint.pprint(filter_set)
# -
# ## Processing
# +
# Create an output database
functions.build_output_database(output_database)
# Save taxon and filter set info into database
output_db_conn= sqlite3.connect(output_database)
cursor = output_db_conn.cursor()
pd.DataFrame(taxon_info.values(), taxon_info.keys()).applymap(str).to_sql(name='taxon_concept', con=output_db_conn, if_exists='replace')
pd.DataFrame(filter_set.values(), filter_set.keys()).applymap(str).to_sql(name='filter_set', con=output_db_conn, if_exists='replace')
output_db_conn.close()
# +
# Run the appropriate queries
if ask_eBird == True and ask_GBIF == True:
# Run GBIF query
gbif_data = functions.get_GBIF_records(taxon_info, filter_set, query_name, working_directory, username, password, email)
# Run eBird query
ebird_data = functions.get_EBD_records(taxon_info, filter_set, working_directory, EBD_file, query_name, R_home)
elif ask_eBird == True and ask_GBIF == False:
# Run eBird query
ebird_data = functions.get_EBD_records(taxon_info, filter_set, working_directory, EBD_file, query_name, R_home)
gbif_data = None
elif ask_eBird == False and ask_GBIF == True:
# Run GBIF query
gbif_data = functions.get_GBIF_records(taxon_info, filter_set, query_name, working_directory, username, password, email)
ebird_data = None
# -
# Filter out records with undesirable values, locations, and/or duplication.
functions.process_records(ebird_data=ebird_data, gbif_data=gbif_data, filter_set=filter_set,
taxon_info=taxon_info, working_directory=working_directory,
query_name=query_name)
output_db_conn= sqlite3.connect(output_database)
cursor = output_db_conn.cursor()
record_count = cursor.execute("SELECT COUNT(record_id) FROM occurrence_records;").fetchone()
print(str(record_count[0]) + " records were saved in the output database")
# ## Results of the Filtering
# #### Sources
pd.set_option('display.max_colwidth', 31)
sources = pd.read_sql(sql="SELECT * FROM sources;", con=output_db_conn)
print(sources[['institutionID', 'collectionCode', 'datasetName', 'acquired', 'removed', 'retained']])
# #### Bases
bases = pd.read_sql(sql="SELECT * FROM attribute_value_counts WHERE attribute = 'basisOfRecord';", con=output_db_conn)
print(bases[['value', 'acquired', 'removed', 'retained']])
# #### Protocols
pd.set_option('display.max_colwidth', 90)
protocols = pd.read_sql(sql="SELECT * FROM attribute_value_counts WHERE attribute = 'samplingProtocol';", con=output_db_conn)
if protocols.empty == True:
print("No protocols were documented.")
if protocols.empty == False:
print(protocols[['value', 'acquired', 'removed', 'retained']])
# #### Issues
pd.set_option('display.max_colwidth', 90)
issues = pd.read_sql(sql="SELECT * FROM attribute_value_counts WHERE attribute = 'issues';", con=output_db_conn)
if issues.empty == True:
print("No issues were documented.")
if issues.empty == False:
print(issues[['value', 'acquired', 'removed', 'retained']])
# #### Establishment Means
establishment = pd.read_sql(sql="SELECT * FROM attribute_value_counts WHERE attribute = 'establishmentMeans';", con=output_db_conn)
if establishment.empty == True:
print("No establishment means were reported.")
if establishment.empty == False:
print(establishment[['value', 'acquired', 'removed', 'retained']])
# #### Identification Qualifiers
qualifications = pd.read_sql(sql="SELECT * FROM attribute_value_counts WHERE attribute = 'identificationQualifers';", con=output_db_conn)
if qualifications.empty == True:
print("No identification qualifiers were reported.")
if qualifications.empty == False:
print(establishment[['value', 'acquired', 'removed', 'retained']])
# ## Descriptions of Retained Records
# #### Locations
# +
# Get the record coordinates as a data frame
record_coordinates = (pd.read_sql("""SELECT decimalLatitude, decimalLongitude, radius_m
FROM occurrence_records""",
con=output_db_conn)
.astype({'decimalLongitude': 'float', 'decimalLatitude': 'float',
'radius_m': 'float'}))
# Make the data frame spatial
gdf = gpd.GeoDataFrame(record_coordinates, geometry=gpd.points_from_xy(record_coordinates['decimalLongitude'],
record_coordinates['decimalLatitude']))
# Set the coordinate reference system
gdf.crs={'init' :'epsg:4326'}
# Create world map <<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
base = world.plot(figsize=(12,12), color = 'darkkhaki')
gdf.plot(ax=base, marker='o', color='k', markersize=5)
plt.show()
# Create USA map <<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
if filter_set["country"] == "US":
usa_bbox = np.array([-124.725839, 24.498131, -66.949895, 49.384358])
fig, ax = plt.subplots(figsize=(12,12))
ax.set_xlim(([usa_bbox[0], usa_bbox[2]]))
ax.set_ylim(([usa_bbox[1], usa_bbox[3]]))
world.plot(ax=ax, color='darkkhaki')
gdf.plot(ax=ax, marker='o', color='k', markersize=5)
plt.show()
# Create coordinate extent map <<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
os.chdir(config.codeDir)
states = gpd.read_file(os.getcwd() + '/data/us_states.shp')
# Reproject states and record coordinates to facilitate buffering
states = states.to_crs(epsg=5070)
footprints = gdf.to_crs(epsg=5070)
# Buffer points for record footprints
footprints['footprint']=footprints.apply(lambda x: x.geometry.buffer(x.radius_m), axis=1)
footprints.set_geometry(col='footprint', inplace=True, drop=True)
# Map the buffered points/footprints
fig, ax = plt.subplots(figsize=(12,12))
coordinate_bbox = footprints.geometry.total_bounds
ax.set_xlim(([coordinate_bbox[0], coordinate_bbox[2]]))
ax.set_ylim(([coordinate_bbox[1], coordinate_bbox[3]]))
states.plot(ax=ax, color = 'darkkhaki')
footprints.boundary.plot(ax=ax, color='k')
plt.show()
# Cleanup
del footprints, gdf
# -
# #### Years Represented
occ_years = [int(x[0]) for x in cursor.execute("SELECT strftime('%Y', eventDate) FROM occurrence_records").fetchall()]
years = filter_set['years_range']
years = years.split(',')
yearsrng = list(range(int(years[0]), int(years[1]), 1))
binsnum = int(years[1]) - int(years[0])
plt.hist(occ_years, bins=binsnum)
plt.ylabel("number of records")
plt.xlabel("year")
plt.xticks(yearsrng, rotation=90)
plt.title("Occurrences per Year")
# #### Months Represented
occ_months = [int(x[0]) for x in cursor.execute("SELECT strftime('%m', eventDate) FROM occurrence_records").fetchall()]
plt.hist(occ_months, bins=range(1, 14), color="g")
plt.ylabel("number of records")
plt.xlabel("month")
plt.xticks(range(1, 13))
plt.title("Occurrences per Month")
# #### Distribution of Coordinate Uncertainty Values for Retained Records
try:
occ_cert = [float(x[0]) for x in cursor.execute("SELECT coordinateUncertaintyInMeters FROM occurrence_records WHERE coordinateUncertaintyInMeters > 0.0;").fetchall()]
maxi = max(occ_cert)
plt.figure(figsize=(16,4))
plt.hist(occ_cert, bins=50, color="r")
plt.xticks(range(0, int(maxi), int(maxi/50)), rotation=90)
plt.ylabel("number of records")
plt.xlabel("meters")
plt.title("Reported Coordinate Uncertainties")
except:
print("No coordinate uncertanties values were reported.")
try:
rng_max = 2000
occ_cert2 = [x for x in occ_cert if x <= rng_max]
plt.figure(figsize=(12,4))
plt.hist(occ_cert2, bins=30, color="m", align='mid')
plt.xticks(range(0, rng_max + 100, int(rng_max/30.)), rotation=90)
plt.ylabel("number of records")
plt.xlabel("meters")
plt.title("Reported Coordinate Uncertainties Below 2km")
except:
print("No coordinate uncertanties values were reported.")
# #### Distribution of Point-radius Values for Retained Records
occ_cert = [float(x[0]) for x in cursor.execute("SELECT radius_m FROM occurrence_records;").fetchall()]
plt.figure(figsize=(11,4))
plt.boxplot(occ_cert, vert=False)
plt.xlabel("meters")
plt.title("Compiled Point-radius Buffer Lengths")
# #### Distribution of Nominal Coordinate Precisions for Retained Records
occ_cert = [float(x[0]) for x in cursor.execute("SELECT nominal_xy_precision FROM occurrence_records;").fetchall()]
plt.figure(figsize=(11,4))
plt.boxplot(occ_cert, vert=False)
plt.xlabel("meters")
plt.title("Nominal Precisions of Coordinates")
# #### Remarks
print("General remarks:")
remarks = output_db_conn.execute("SELECT DISTINCT general_remarks FROM occurrence_records;").fetchall()
if len(remarks) <= 20:
try:
for rem in remarks:
if rem[0][0:1] == ';':
print(rem[0][2:])
else:
print(rem[0])
except:
pass
else:
print("More than 20 remarks, consult the occurrence database.")
print("Event remarks:")
remarks = output_db_conn.execute("SELECT DISTINCT eventRemarks FROM occurrence_records;").fetchall()
if len(remarks) <= 20:
try:
for rem in remarks:
if rem[0][0:1] == ';':
print(rem[0][2:])
else:
print(rem[0])
except:
pass
else:
print("More than 20 remarks, consult the occurrence database.")
print("Occurrence remarks:")
remarks = output_db_conn.execute("SELECT DISTINCT occurrenceRemarks FROM occurrence_records;").fetchall()
if len(remarks) <= 20:
try:
for rem in remarks:
if rem[0][0:1] == ';':
print(rem[0][2:])
else:
print(rem[0])
except:
pass
else:
print("More than 20 remarks, consult the occurrence database.")
print("Location remarks:")
remarks = output_db_conn.execute("SELECT DISTINCT locationRemarks FROM occurrence_records;").fetchall()
if len(remarks) <= 20:
try:
for rem in remarks:
if rem[0][0:1] == ';':
print(rem[0][2:])
else:
print(rem[0])
except:
pass
else:
print("More than 20 remarks, consult the occurrence database.")
print("Identified remarks:")
remarks = output_db_conn.execute("SELECT DISTINCT general_remarks FROM occurrence_records;").fetchall()
if len(remarks) <= 20:
try:
for rem in remarks:
if rem[0][0:1] == ';':
print(rem[0][2:])
else:
print(rem[0])
except:
pass
else:
print("More than 20 remarks, consult the occurrence database.")
print("Georeference remarks:")
remarks = output_db_conn.execute("SELECT DISTINCT georeferenceRemarks FROM occurrence_records;").fetchall()
if len(remarks) <= 20:
try:
for rem in remarks:
if rem[0][0:1] == ';':
print(rem[0][2:])
else:
print(rem[0])
except:
pass
else:
print("More than 20 remarks, consult the occurrence database.")
# #### Attributes Returned for GBIF Records
# This count was made before filters were applied
pd.set_option('display.max_colwidth', 100)
if ask_GBIF == True:
fields_summary = pd.read_sql("SELECT * FROM gbif_fields_returned", output_db_conn)
fields_summary.index.name = 'Field'
pd.set_option('display.max_rows', 250)
print(fields_summary.sort_values(by="attribute"))
if ask_GBIF == False:
print("GBIF was not queried.")
# #### Attributes Returned for eBird Records
# This count was made before filters were applied
if ask_eBird == True:
fields_summary = pd.read_sql("SELECT * FROM ebird_fields_returned", output_db_conn)
fields_summary.index.name = 'Field'
pd.set_option('display.max_rows', 250)
print(fields_summary)
if ask_eBird == False:
print("No eBird Basic Dataset was queried.")
# ## Citations
# #### eBird
if ask_eBird == True:
with open(config.EBD_file[:-22] + "recommended_citation.txt", "r") as reference:
print(reference.readlines()[0])
else:
print("No eBird Basic Dataset was queried")
# #### GBIF
if ask_GBIF == True:
if filter_set["get_dwca"] == True:
print("Citations-- ")
print(cursor.execute("SELECT citations FROM GBIF_download_info").fetchall()[0][0])
else:
print("Set 'get_dwca' to True to acquire a list of citations")
if ask_GBIF == False:
print("GBIF was not queried.")
if ask_GBIF == True:
if filter_set["get_dwca"] == True:
print("Rights-- ")
print(output_db_conn.execute("SELECT rights FROM GBIF_download_info").fetchall()[0][0])
else:
print("Set 'get_dwca' to True to see the rights")
if ask_GBIF == False:
print("GBIF was not queried.")
if ask_GBIF == True:
if filter_set["get_dwca"] == True:
print("DOI-- ")
doi = output_db_conn.execute("SELECT doi FROM GBIF_download_info").fetchall()[0][0]
print("https://doi.org/" + doi)
else:
print("Set 'get_dwca' to True to perform a search with a doi assigned")
if ask_GBIF == False:
print("GBIF was not queried.")
if ask_GBIF == True:
if filter_set["get_dwca"] == True:
print("GBIF download key-- ")
print(output_db_conn.execute("SELECT download_key FROM GBIF_download_info").fetchall()[0][0])
else:
print("Set 'get_dwca' to True to perform a search with a download key assigned")
if ask_GBIF == False:
print("GBIF was not queried")
# ## Runtime
output_db_conn.close()
del cursor
t2 = datetime.now()
print(t2 - t1)
# Check for violations of filter set and print a message if an error is found.
functions.verify_results(output_database)
# ## Optional Output
# Remove "#" to activate desired statements.
# +
# Make a shapefile of record footprints
#shp1 = functions.spatial_output(database=output_database, make_file=True, output_file=working_directory + your_taxon_id + "_footprints.shp", epsg=5070, mode="footprints")
# Make a shapefile of record coordinates
#shp2 = functions.spatial_output(database=output_database, make_file=True, output_file=working_directory + your_taxon_id + "_points.shp", epsg=5070, mode="points")
# Make a shapefile of random points
#shp3 = functions.spatial_output(database=output_database, make_file=True, output_file=working_directory + your_taxon_id + "_random.shp", epsg=5070, mode="random")
# +
# Export this notebook as an html file
#os.chdir(working_directory)
# #!jupyter nbconvert --to html --TemplateExporter.exclude_input=True --output-dir="C:/PATH/WHERE/TO/SAVE/" ENTER_QUERY_NAME_HERE.ipynb
| Query_TEMPLATE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_qwukc5b"
# # Exercise: Heterogenous Plugin and the DevCloud
#
# Now that you've learned about the **Heterogenous plugin**, you will have the opportunity to load a model on all three hardware devices, CPU, GPU, and FPGA, using 3 device combinations and run inference on an image.
#
# In this exercise, you will do the following:
# 1. Write a Python script to load a model and run inference 100 times on a device on Intel's DevCloud.
# * Calculate the time it takes to load the model.
# * Calculate the time it takes to run inference 100 times.
# 2. Write a shell script to submit a job to Intel's DevCloud.
# 3. Submit a job using `qsub` on an **IEI Tank-870** edge node using `HETERO`, run `liveQStat` to view the status of your submitted jobs, and then retrieve and view the results from your job.
# * One job using `FPGA/CPU` as the device.
# * One job using `GPU/CPU` as the device.
# * One job using `FPGA/GPU/CPU` as the device.
# 4. Plot and compare the results using bar graphs with `matplotlib` for the following metrics:
# * Model Loading Time
# * Inference Time
# * Frames Per Second (FPS)
#
# Click the **Exercise Overview** button below for a demonstration.
# + [markdown] graffitiCellId="id_z8bfs11"
# <span class="graffiti-highlight graffiti-id_z8bfs11-id_d97ox8f"><i></i><button>Exercise Overview</button></span>
# + [markdown] graffitiCellId="id_0untint"
# #### IMPORTANT: Set up paths so we can run Dev Cloud utilities
# You *must* run this every time you enter a Workspace session.
# + graffitiCellId="id_axn1sb2"
# %env PATH=/opt/conda/bin:/opt/spark-2.4.3-bin-hadoop2.7/bin:/opt/conda/bin:/usr/local/sbin:/usr/local/bin:/usr/sbin:/usr/bin:/sbin:/bin:/opt/intel_devcloud_support
import os
import sys
sys.path.insert(0, os.path.abspath('/opt/intel_devcloud_support'))
sys.path.insert(0, os.path.abspath('/opt/intel'))
# + [markdown] graffitiCellId="id_mhiayyz"
# ## The Model
#
# We will be using the `vehicle-license-plate-detection-barrier-0106` model for this exercise.
#
# Remember to use the appropriate model precisions for each device:
#
# * FPGA - `FP16`
# * GPU - `FP16`
# * CPU - It is prefered to use `FP32`, but we have to use `FP16` since **GPU** and **FPGA** use `FP16`
#
# The model has already been downloaded for you in the `/data/models/intel` directory on Intel's DevCloud.
#
# We will be running inference on an image of a car. The path to the image is `/data/resources/car.png`.
# + [markdown] graffitiCellId="id_ltf95ei"
# # Step 1: Creating a Python Script
#
# The first step is to create a Python script that you can use to load the model and perform inference. We'll use the `%%writefile` magic to create a Python file called `inference_on_device.py`. In the next cell, you will need to complete the `TODO` items for this Python script.
#
# `TODO` items:
#
# 1. Load the model
#
# 2. Get the name of the input node
#
# 3. Prepare the model for inference (create an input dictionary)
#
# 4. Run inference 100 times in a loop
#
# If you get stuck, you can click on the **Show Solution** button below for a walkthrough with the solution code.
# + graffitiCellId="id_bpywo8s"
# %%writefile inference_on_device.py
import time
import numpy as np
import cv2
from openvino.inference_engine import IENetwork
from openvino.inference_engine import IECore
import argparse
def main(args):
model=args.model_path
model_weights=model+'.bin'
model_structure=model+'.xml'
start=time.time()
# TODO: Load the model
load_time=time.time()-start
print(f"Time taken to load model = {load_time} seconds")
# Get the name of the input node
# Reading and Preprocessing Image
input_img=cv2.imread('car.png')
input_img=cv2.resize(input_img, (300,300), interpolation = cv2.INTER_AREA)
input_img=np.moveaxis(input_img, -1, 0)
# TODO: Prepare the model for inference (create input dict etc.)
start=time.time()
for _ in range(100):
# TODO: Run Inference in a Loop
inference_time=time.time()-start
fps=100/inference_time
print(f"Time Taken to run 100 Inference is = {inference_time} seconds")
with open(f"/output/{args.path}.txt", "w") as f:
f.write(str(load_time)+'\n')
f.write(str(inference_time)+'\n')
f.write(str(fps)+'\n')
if __name__=='__main__':
parser=argparse.ArgumentParser()
parser.add_argument('--model_path', required=True)
parser.add_argument('--device', default=None)
parser.add_argument('--path', default=None)
args=parser.parse_args()
main(args)
# + [markdown] graffitiCellId="id_1rnmf5g"
# <span class="graffiti-highlight graffiti-id_1rnmf5g-id_nmeqj1a"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_ufbi2ll"
# ## Step 2: Creating a Job Submission Script
#
# To submit a job to the DevCloud, you'll need to create a shell script. Similar to the Python script above, we'll use the `%%writefile` magic command to create a shell script called `inference_model_job.sh`. In the next cell, you will need to complete the `TODO` items for this shell script.
#
# `TODO` items:
# 1. Create three variables:
# * `DEVICE` - Assign the value as the first argument passed into the shell script.
# * `MODELPATH` - Assign the value as the second argument passed into the shell script.
# * `SAVEPATH` - Assign the value as the third argument passed into the shell script.
# 2. Call the Python script using the three variable values as the command line argument
#
# If you get stuck, you can click on the **Show Solution** button below for a walkthrough with the solution code.
# + graffitiCellId="id_5r13clu"
# %%writefile inference_model_job.sh
# #!/bin/bash
exec 1>/output/stdout.log 2>/output/stderr.log
# mkdir -p /output
# TODO: Create DEVICE variable
# TODO: Create MODELPATH variable
# TODO: Create SAVEPATH variable
if echo "$DEVICE" | grep -q "FPGA"; then # if device passed in is FPGA, load bitstream to program FPGA
export AOCL_BOARD_PACKAGE_ROOT=/opt/intel/openvino/bitstreams/a10_vision_design_sg2_bitstreams/BSP/a10_1150_sg2
source /opt/altera/aocl-pro-rte/aclrte-linux64/init_opencl.sh
aocl program acl0 /opt/intel/openvino/bitstreams/a10_vision_design_sg2_bitstreams/2020-2_PL2_FP16_MobileNet_Clamp.aocx
export CL_CONTEXT_COMPILER_MODE_INTELFPGA=3
fi
# TODO: Call the Python script
# cd /output
tar zcvf output.tgz * # compresses all files in the current directory (output)
# + [markdown] graffitiCellId="id_f1nbmn9"
# <span class="graffiti-highlight graffiti-id_f1nbmn9-id_ia7yjlq"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_2wqvzne"
# ## Step 3: Submitting a Job to Intel's DevCloud
#
# In the next three sub-steps, you will write your `!qsub` commands to submit your jobs to Intel's DevCloud to load your model and run inference on the **IEI Tank-870** edge node with an **Intel Core i5** CPU and an **Intel Arria 10 FPGA**. You will use the **HETERO** device plugin to run inference on three device combinations.
#
# Your `!qsub` command should take the following flags and arguments:
# 1. The first argument should be the shell script filename
# 2. `-d` flag - This argument should be `.`
# 3. `-l` flag - This argument should request an edge node with an **IEI Tank-870**. The default quantity is 1, so the **1** after `nodes` is optional.
# * **Intel Core i5 6500TE** for your `CPU`.
# * **Intel HD Graphics 530** for your `IGPU`.
# * **Intel Arria 10** for your `FPGA`.
#
# To get the queue labels for these devices, you can go to [this link](https://devcloud.intel.com/edge/get_started/devcloud/)
#
# 4. `-F` flag - This argument should contain the three values to assign to the variables of the shell script:
# * **DEVICE** - Device type for the job: You will have to use `HETERO` with three different combinations of `CPU`,`GPU` or `FPGA`.
# - `FPGA,CPU`
# - `GPU,CPU`
# - `FPGA,GPU,CPU`
# * **MODELPATH** - Full path to the model for the job. As a reminder, the model is located in `/data/models/intel`.
# * **SAVEPATH** - Name of the file you want to save the performance metrics as. These should be named as the following:
# - `fpga_cpu_stats` for the `FPGA/CPU` job
# - `gpu_cpu_stats` for the `GPU/CPU` job
# - `fpga_gpu_cpu_stats` for the `FPGA/GPU/CPU` job
#
# **Note**: There is an optional flag, `-N`, you may see in a few exercises. This is an argument that only works on Intel's DevCloud that allows you to name your job submission. This argument doesn't work in Udacity's workspace integration with Intel's DevCloud.
# + [markdown] graffitiCellId="id_28fed2h"
# ## Step 3a: Running on the FPGA and CPU
#
# In the cell below, write the qsub command that will submit your job to both the FPGA and CPU.
#
# If you get stuck, you can click on the **Show Solution** button below for a walkthrough with the solution code.
# + graffitiCellId="id_6awpacu"
fpga_cpu_job = # TODO: Write qsub command
print(fpga_cpu_job[0])
# + [markdown] graffitiCellId="id_cvp3lyi"
# <span class="graffiti-highlight graffiti-id_cvp3lyi-id_chmeh50"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_dfm31r5"
# ### Check Job Status
#
# To check on the job that was submitted, use `liveQStat` to check the status of the job. The cell is locked until this finishes polling 10 times or you can interrupt the kernel to stop it by pressing the stop button at the top: 
#
# Column `S` shows the state of your running jobs.
#
# For example:
# - If `JOB ID`is in Q state, it is in the queue waiting for available resources.
# - If `JOB ID` is in R state, it is running.
# + graffitiCellId="id_0756ni1"
import liveQStat
liveQStat.liveQStat()
# + [markdown] graffitiCellId="id_t7nm9be"
# ###### Get Results
#
# Run the next cell to retrieve your job's results.
# + graffitiCellId="id_zbykzwd"
import get_results
get_results.getResults(fpga_cpu_job[0], filename="output.tgz", blocking=True)
# + [markdown] graffitiCellId="id_e8klgeq"
# ###### Unpack your output files and view stdout.log
# + graffitiCellId="id_3hp5mib"
# !tar zxf output.tgz
# + graffitiCellId="id_c1lmzsp"
# !cat stdout.log
# + [markdown] graffitiCellId="id_0k0k9jp"
# ###### View stderr.log
# This can be used for debugging
# + graffitiCellId="id_m33x45r"
# !cat stderr.log
# + [markdown] graffitiCellId="id_io25c53"
# ## Step 3a: Running on the GPU and CPU
#
# In the cell below, write the qsub command that will submit your job to both the GPU and CPU.
#
# If you get stuck, you can click on the **Show Solution** button below for a walkthrough with the solution code.
# + graffitiCellId="id_v5klpi1"
gpu_cpu_job = # TODO: Write qsub command
print(gpu_cpu_job[0])
# + [markdown] graffitiCellId="id_7k34s6u"
# <span class="graffiti-highlight graffiti-id_7k34s6u-id_022l4bj"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_dfm31r5"
# ### Check Job Status
#
# To check on the job that was submitted, use `liveQStat` to check the status of the job. The cell is locked until this finishes polling 10 times or you can interrupt the kernel to stop it by pressing the stop button at the top: 
#
# Column `S` shows the state of your running jobs.
#
# For example:
# - If `JOB ID`is in Q state, it is in the queue waiting for available resources.
# - If `JOB ID` is in R state, it is running.
# + graffitiCellId="id_0756ni1"
import liveQStat
liveQStat.liveQStat()
# + [markdown] graffitiCellId="id_t7nm9be"
# ###### Get Results
#
# Run the next cell to retrieve your job's results.
# + graffitiCellId="id_zbykzwd"
import get_results
get_results.getResults(gpu_cpu_job[0], filename="output.tgz", blocking=True)
# + [markdown] graffitiCellId="id_e8klgeq"
# ###### Unpack your output files and view stdout.log
# + graffitiCellId="id_3hp5mib"
# !tar zxf output.tgz
# + graffitiCellId="id_c1lmzsp"
# !cat stdout.log
# + [markdown] graffitiCellId="id_0k0k9jp"
# ###### View stderr.log
# This can be used for debugging
# + graffitiCellId="id_m33x45r"
# !cat stderr.log
# + [markdown] graffitiCellId="id_io25c53"
# ## Step 3c: Running on the FPGA, GPU, and CPU
#
# In the cell below, write the qsub command that will submit your job to all three devices, FPGA, GPU, and CPU.
#
# If you get stuck, you can click on the **Show Solution** button below for a walkthrough with the solution code.
# + graffitiCellId="id_v5klpi1"
fpga_gpu_cpu_job = # TODO: Write qsub command
print(fpga_gpu_cpu_job[0])
# + [markdown] graffitiCellId="id_mxh5ozv"
# <span class="graffiti-highlight graffiti-id_mxh5ozv-id_qicoukm"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_dfm31r5"
# ### Check Job Status
#
# To check on the job that was submitted, use `liveQStat` to check the status of the job. The cell is locked until this finishes polling 10 times or you can interrupt the kernel to stop it by pressing the stop button at the top: 
#
# Column `S` shows the state of your running jobs.
#
# For example:
# - If `JOB ID`is in Q state, it is in the queue waiting for available resources.
# - If `JOB ID` is in R state, it is running.
# + graffitiCellId="id_0756ni1"
import liveQStat
liveQStat.liveQStat()
# + [markdown] graffitiCellId="id_t7nm9be"
# ###### Get Results
#
# Run the next cell to retrieve your job's results.
# + graffitiCellId="id_zbykzwd"
import get_results
get_results.getResults(fpga_gpu_cpu_job[0], filename="output.tgz", blocking=True)
# + [markdown] graffitiCellId="id_e8klgeq"
# ###### Unpack your output files and view stdout.log
# + graffitiCellId="id_3hp5mib"
# !tar zxf output.tgz
# + graffitiCellId="id_c1lmzsp"
# !cat stdout.log
# + [markdown] graffitiCellId="id_0k0k9jp"
# ###### View stderr.log
# This can be used for debugging
# + graffitiCellId="id_m33x45r"
# !cat stderr.log
# + [markdown] graffitiCellId="id_4rf323l"
# ## Step 4: Plot and Compare Results
#
# Run the cells below to plot and compare the results.
# + graffitiCellId="id_bkny5ta"
import matplotlib.pyplot as plt
# + graffitiCellId="id_njkbzfn"
def plot(labels, data, title, label):
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.set_ylabel(label)
ax.set_title(title)
ax.bar(labels, data)
def read_files(paths, labels):
load_time=[]
inference_time=[]
fps=[]
for path in paths:
if os.path.isfile(path):
f=open(path, 'r')
load_time.append(float(f.readline()))
inference_time.append(float(f.readline()))
fps.append(float(f.readline()))
plot(labels, load_time, 'Model Load Time', 'seconds')
plot(labels, inference_time, 'Inference Time', 'seconds')
plot(labels, fps, 'Frames per Second', 'Frames')
paths=['fpga_cpu_stats.txt', 'gpu_cpu_stats.txt', 'fpga_gpu_cpu_stats.txt']
read_files(paths, ['FPGA/CPU', 'GPU/CPU', 'FPGA/GPU/CPU'])
| Jupyter Tutorial/Exercise_Heterogenous_plugin_and_the_DevCloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.0.0b0 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# # 人脸关键点检测
#
# **作者:** [ssz95](https://github.com/zzs95) <br>
# **日期:** 2021.05 <br>
# **摘要:** 本示例教程将会演示如何使用飞桨实现人脸关键点检测。
# ## 一、简介
# 在图像处理中,关键点本质上是一种特征。它是对一个固定区域或者空间物理关系的抽象描述,描述的是一定邻域范围内的组合或上下文关系。它不仅仅是一个点信息,或代表一个位置,更代表着上下文与周围邻域的组合关系。关键点检测的目标就是通过计算机从图像中找出这些点的坐标,作为计算机视觉领域的一个基础任务,关键点的检测对于高级别任务,例如识别和分类具有至关重要的意义。
#
# 关键点检测方法总体上可以分成两个类型,一个种是用坐标回归的方式来解决,另一种是将关键点建模成热力图,通过像素分类任务,回归热力图分布得到关键点位置。这两个方法,都是一种手段或者是途径,解决的问题就是要找出这个点在图像当中的位置与关系。
#
# 其中人脸关键点检测是关键点检测方法的一个成功实践,本示例简要介绍如何通过飞桨开源框架,实现人脸关键点检测的功能。这个案例用到的是第一种关键点检测方法——坐标回归。将使用到 Paddle 2.1的API,集成式的训练接口,能够很方便对模型进行训练和预测。
#
# ## 二、环境设置
#
# 本教程基于Paddle 2.1 编写,如果你的环境不是本版本,请先参考官网[安装](https://www.paddlepaddle.org.cn/install/quick) Paddle 2.1 。
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import os
import paddle
from paddle.io import Dataset
from paddle.vision.transforms import transforms
from paddle.vision.models import resnet18
from paddle.nn import functional as F
print(paddle.__version__)
# -
# ## 三、数据集
# ### 3.1 数据集下载
# 本案例使用了Kaggle官方举办的人脸关键点检测challenge数据集,官网:[https://www.kaggle.com/c/facial-keypoints-detection](https://www.kaggle.com/c/facial-keypoints-detection)
#
# 官方数据集将人脸图像和标注数据打包成了csv文件,使用panda来读取。其中数据集中的文件:<br>
# training.csv: 包含了用于训练的人脸关键点坐标和图像。<br>
# test.csv: 包含了用于测试的人脸关键点图像, 没有标注关键点坐标。<br>
# IdLookupTable.csv: 测试集关键点的位置的对应名称。<br>
#
# 图像的长和宽都为96像素,所需要检测的一共有15个关键点。
# !unzip -o ./test.zip -d data/data60
# !unzip -o ./training.zip -d data/data60
# ### 3.2 数据集定义
# 飞桨(PaddlePaddle)数据集加载方案是统一使用Dataset(数据集定义) + DataLoader(多进程数据集加载)。
#
# 首先进行数据集的定义,数据集定义主要是实现一个新的Dataset类,继承父类paddle.io.Dataset,并实现父类中以下两个抽象方法,__getitem__和__len__:
# +
Train_Dir = './data/data60/training.csv'
Test_Dir = './data/data60/test.csv'
lookid_dir = './data/data60/IdLookupTable.csv'
class ImgTransforms(object):
"""
图像预处理工具,用于将图像进行升维(96, 96) => (96, 96, 3),
并对图像的维度进行转换从HWC变为CHW
"""
def __init__(self, fmt):
self.format = fmt
def __call__(self, img):
if len(img.shape) == 2:
img = np.expand_dims(img, axis=2)
img = img.transpose(self.format)
if img.shape[0] == 1:
img = np.repeat(img, 3, axis=0)
return img
class FaceDataset(Dataset):
def __init__(self, data_path, mode='train', val_split=0.2):
self.mode = mode
assert self.mode in ['train', 'val', 'test'], \
"mode should be 'train' or 'test', but got {}".format(self.mode)
self.data_source = pd.read_csv(data_path)
# 清洗数据, 数据集中有很多样本只标注了部分关键点, 这里有两种策略
# 第一种, 将未标注的位置从上一个样本对应的关键点复制过来
# self.data_source.fillna(method = 'ffill',inplace = True)
# 第二种, 将包含有未标注的样本从数据集中移除
self.data_source.dropna(how="any", inplace=True)
self.data_label_all = self.data_source.drop('Image', axis = 1)
# 划分训练集和验证集合
if self.mode in ['train', 'val']:
np.random.seed(43)
data_len = len(self.data_source)
# 随机划分
shuffled_indices = np.random.permutation(data_len)
# 顺序划分
# shuffled_indices = np.arange(data_len)
self.shuffled_indices = shuffled_indices
val_set_size = int(data_len*val_split)
if self.mode == 'val':
val_indices = shuffled_indices[:val_set_size]
self.data_img = self.data_source.reindex().iloc[val_indices]
self.data_label = self.data_label_all.reindex().iloc[val_indices]
elif self.mode == 'train':
train_indices = shuffled_indices[val_set_size:]
self.data_img = self.data_source.reindex().iloc[train_indices]
self.data_label = self.data_label_all.reindex().iloc[train_indices]
elif self.mode == 'test':
self.data_img = self.data_source
self.data_label = self.data_label_all
self.transforms = transforms.Compose([
ImgTransforms((2, 0, 1))
])
# 每次迭代时返回数据和对应的标签
def __getitem__(self, idx):
img = self.data_img['Image'].iloc[idx].split(' ')
img = ['0' if x == '' else x for x in img]
img = np.array(img, dtype = 'float32').reshape(96, 96)
img = self.transforms(img)
label = np.array(self.data_label.iloc[idx,:],dtype = 'float32')/96
return img, label
# 返回整个数据集的总数
def __len__(self):
return len(self.data_img)
# 训练数据集和验证数据集
train_dataset = FaceDataset(Train_Dir, mode='train')
val_dataset = FaceDataset(Train_Dir, mode='val')
# 测试数据集
test_dataset = FaceDataset(Test_Dir, mode='test')
# -
# ### 3.3 数据集抽样展示
# 实现好Dataset数据集后,来测试一下数据集是否符合预期,因为Dataset是一个可以被迭代的Class,通过for循环从里面读取数据来用matplotlib进行展示。关键点的坐标在数据集中进行了归一化处理,这里乘以图像的大小恢复到原始尺度,并用scatter函数将点画在输出的图像上。
# +
def plot_sample(x, y, axis):
img = x.reshape(96, 96)
axis.imshow(img, cmap='gray')
axis.scatter(y[0::2], y[1::2], marker='x', s=10, color='b')
fig = plt.figure(figsize=(10, 7))
fig.subplots_adjust(
left=0, right=1, bottom=0, top=1, hspace=0.05, wspace=0.05)
# 随机取16个样本展示
for i in range(16):
axis = fig.add_subplot(4, 4, i+1, xticks=[], yticks=[])
idx = np.random.randint(train_dataset.__len__())
# print(idx)
img, label = train_dataset[idx]
label = label*96
plot_sample(img[0], label, axis)
plt.show()
# -
# ## 四、定义模型
# 这里使用到 ``paddle.vision.models`` 中定义的 ``resnet18`` 网络模型。在ImageNet分类任务中,图像分成1000类,在模型后接一个全连接层,将输出的1000维向量映射成30维,对应15个关键点的横纵坐标。
class FaceNet(paddle.nn.Layer):
def __init__(self, num_keypoints, pretrained=False):
super(FaceNet, self).__init__()
self.backbone = resnet18(pretrained)
self.outLayer1 = paddle.nn.Sequential(
paddle.nn.Linear(1000, 512),
paddle.nn.ReLU(),
paddle.nn.Dropout(0.1))
self.outLayer2 = paddle.nn.Linear(512, num_keypoints*2)
def forward(self, inputs):
out = self.backbone(inputs)
out = self.outLayer1(out)
out = self.outLayer2(out)
return out
# ### 4.1 模型可视化
# 调用飞桨提供的summary接口对组建好的模型进行可视化,方便进行模型结构和参数信息的查看和确认。
# +
from paddle.static import InputSpec
num_keypoints = 15
model = paddle.Model(FaceNet(num_keypoints))
model.summary((1,3, 96, 96))
# -
# ## 五、训练模型
# 在这个任务是对坐标进行回归,使用均方误差(Mean Square error )损失函数`paddle.nn.MSELoss()`来做计算,飞桨2.1中,在nn下将损失函数封装成可调用类。这里使用paddle.Model相关的API直接进行训练,只需要定义好数据集、网络模型和损失函数即可。
#
# 使用模型代码进行Model实例生成,使用prepare接口定义优化器、损失函数和评价指标等信息,用于后续训练使用。在所有初步配置完成后,调用fit接口开启训练执行过程,调用fit时只需要将前面定义好的训练数据集、测试数据集、训练轮次(Epoch)和批次大小(batch_size)配置好即可。
model = paddle.Model(FaceNet(num_keypoints=15))
optim = paddle.optimizer.Adam(learning_rate=1e-3,
parameters=model.parameters())
model.prepare(optim, paddle.nn.MSELoss())
model.fit(train_dataset, val_dataset, epochs=60, batch_size=256)
# ## 六、模型预测
# 为了更好的观察预测结果,分别可视化验证集结果与标注点的对比,和在未标注的测试集的预测结果。
# ### 6.1 验证集结果可视化
# 红色的关键点为网络预测的结果, 绿色的关键点为标注的groundtrue。
result = model.predict(val_dataset, batch_size=1)
# +
def plot_sample(x, y, axis, gt=[]):
img = x.reshape(96, 96)
axis.imshow(img, cmap='gray')
axis.scatter(y[0::2], y[1::2], marker='x', s=10, color='r')
if gt!=[]:
axis.scatter(gt[0::2], gt[1::2], marker='x', s=10, color='lime')
fig = plt.figure(figsize=(10, 7))
fig.subplots_adjust(
left=0, right=1, bottom=0, top=1, hspace=0.05, wspace=0.05)
for i in range(16):
axis = fig.add_subplot(4, 4, i+1, xticks=[], yticks=[])
idx = np.random.randint(val_dataset.__len__())
img, gt_label = val_dataset[idx]
gt_label = gt_label*96
label_pred = result[0][idx].reshape(-1)
label_pred = label_pred*96
plot_sample(img[0], label_pred, axis, gt_label)
plt.show()
# -
# ### 6.2 测试集结果可视化
result = model.predict(test_dataset, batch_size=1)
# +
fig = plt.figure(figsize=(10, 7))
fig.subplots_adjust(
left=0, right=1, bottom=0, top=1, hspace=0.05, wspace=0.05)
for i in range(16):
axis = fig.add_subplot(4, 4, i+1, xticks=[], yticks=[])
idx = np.random.randint(test_dataset.__len__())
img, _ = test_dataset[idx]
label_pred = result[0][idx].reshape(-1)
label_pred = label_pred*96
plot_sample(img[0], label_pred, axis)
plt.show()
# -
| docs/practices/landmark_detection/landmark_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="glA3NV9MRl63" colab_type="text"
# # Kalman Filter - Lab 7.2
#
# ## Recap
# This is the Lab on using a Kalman Filter in CE6003's Object Tracking. You should complete the tasks in this lab as part of the Kalman Filter section of the lesson.
#
# Please remember this lab must be completed before taking the quiz at the end of this lesson.
#
# First, if we haven't already done so, we need to clone the various images and resources needed to run these labs into our workspace.
# + id="md1qGUkYR6H-" colab_type="code" colab={}
# !git clone https://github.com/EmdaloTechnologies/CE6003.git
# + [markdown] id="Nvp_ecbWSB6V" colab_type="text"
#
# **Program Description**
#
# This program demonstrates a very simple 'tracking' mechanism - derived from a Kalman filter. We're going to use our Kalman filter to track a single object, namely a person.
#
# + id="qWvUXFH1iaoJ" colab_type="code" colab={}
import os
import re
import io
import cv2
import time
import numpy as np
import base64
from IPython.display import clear_output, Image, display
# + [markdown] id="_GSU76keSNr7" colab_type="text"
# #The Story So Far
#
# To illustrate how to track something in a video stream, we have used the following technique to generate a set of images for you to work on.
#
# What we did was we generated a short video - just recording one person walking around, on an iPhone.
#
# Then we used ```ffmpeg``` to decompose about 7 seconds of that video down into still images.
#
# ```ffmpeg -i $vid_in -vf fps=30 imgs/daire%03d.png```
#
# We saved those frames as ```imgs/daire%03d.png``` in the git repository in the single-detections directory
#
# We've run yolo3 over those frames to generate bounding boxes and saved those bounding boxes into the same directory.
#
# The file format is comma-separated values and the values are as shown here:
#
# frame index | object-type | centre-x | centre-y | width | height | confidence
# --- | --- | --- | --- | --- | --- | ---
# int | -1 | float | float | float | float | float
#
# * The object type is always a person - that's all we inferred for.
# * The centre-x and width are fractions of the image's width
# * The centre-y and height are fractions of the image's height
# * The confidence is supplied by Yolo3
# + [markdown] id="GYWphuuLX6hr" colab_type="text"
# *What Happens Now*
#
# For each image in the directory, in order,
# * we'll find the centre of the detection in that image (if any)
# * we'll build a bounding box for the detection in that image
# * we'' derive a variance term (crudely) from the Yolo confidence for that image
# * and we'll supply the centre of that bounding box along with the variance term to a Kalman Filter implementation
#
# Then, we'll explore how a Kalman filter tracks the object in the image stream.
# + [markdown] id="XsQyMnFpZBUP" colab_type="text"
# **Get File Handles**
#
# This function gets the filenames of all the files in the directory, in a reproducible order, and loads in the bounding boxes from file.
# + id="I1lGqvyGZBjj" colab_type="code" colab={}
def get_pngs_and_boxes():
pngdir = "/content/CE6003/images/lab7/single-objects/"
bbdir = "/content/CE6003/images/lab7/single-objects/"
pngfolder = os.fsencode(pngdir)
bbfolder = os.fsencode(bbdir)
pngfiles = []
for filename in os.listdir(pngfolder):
if filename.decode().endswith(".png"):
pngfiles.append(pngdir + filename.decode())
pngfiles.sort()
for filename in os.listdir(bbfolder):
if filename.decode().endswith(".boxes"):
bbfilename = bbdir + filename.decode()
bb = open(bbfilename, "r")
bb_lines = bb.readlines()
bb.close()
return bb_lines, pngfiles
# + [markdown] id="qDJASmq8ZrwJ" colab_type="text"
# **Parse Detections**
#
# We'll use this function in the main loop to wrangle the detections into the format we want to supply to our Kalman Filter.
#
# Essentially it takes the name of png file, an img object and the list of bounding boxes as inputs.
#
# It then finds the correct record (if any) for that image in the bounding boxes list and converts the bounding box parameters into a format which we'll use for the rest of the program (it converts back to absolute pixel values).
#
# It returns a centre and a confidence value for the image supplied to it.
# + id="HQw8QFkqZxMD" colab_type="code" colab={}
def parse_detections(bboxes, pngfile, img):
# Sample Line: 400,-1,0.285417,0.241667,0.094792,0.483333,0.999797,-1,-1,-1
# Index, object type,
# x - centre of bounding box (as fraction of image width
# y - centre of bounding box (as fraction of image height
# w - width of bounding box (as fraction of image width)
# h - height of bounding box (as fraction of image height
# prob, _,_,_
# extract the frame index of the png file -
# use it to find the detections for that frame
index = int(re.findall(r'\d+', pngfile)[-1])
imgh, imgw = img.shape[:2]
centre = np.zeros(shape=(2, 1))
P = 0.000001 # hack to avoid div by zero
for line in bboxes:
np_array = np.genfromtxt(io.StringIO(line), delimiter=",")
lineindex = int(np_array[0])
if lineindex == index:
centre = np_array[2:4]
P += np_array[6]
centre[0] *= imgw
centre[1] *= imgh
return centre, P
return centre, P
# + [markdown] id="iuM6UO7Jan5V" colab_type="text"
# **Kalman 2D**
#
# This function specialises the kalman() routine below for a particular model.
#
# In this example, we are going to create a 2D Kalman, and we'll use a constant velocity model - so we'll use a term for x (expressed as position and velocity) and a term for y (also expressed as position and velocity).
#
# ### State Estimate Term
# Therefore our state estimate will be represented by 4 state terms
#
# $x_{state} = \begin{bmatrix}
# x_{pos} \\
# y_{pos} \\
# \dot{x} \\
# \dot{y}
# \end{bmatrix}$
#
# Note: the $x$ in $x_{state}$ is separate from $x_{pos}$.
#
#
# ### State Estimate Error Covariance
# Our state estimate error covariance matrix will have a $4 \times 4$ shape.
# We'll supply something like:
#
# $P_{0} = \begin{bmatrix}
# 100 && 0 && 0 && 0 \\
# 0 && 100 && 0 && 0 \\
# 0 && 0 && 100 && 0 \\
# 0 && 0 && 0 && 100 \\
# \end{bmatrix}$
#
# to this matrix later on to initialize our state estimate error covariance, indicating a high initial degree of uncertainty around each of our state parameters and indicating an initial belief that the uncertainty in each state term is not linked to the uncertainty in the other state terms.
#
# This is a measure of the initialial estimated accuracy of the state estimate $x_{state}$ above.
#
# Its typical to initialize $P$ by setting the diagonal elements to the uncertainty in your initial values of the state and normally to quite a large value.
#
# The measurements will normally bring the state estimate error covariances down when running the filter and in almost all cases, the off diagonal elements will become non-zero as the filter works.
#
# For the case of the simple 2 state position-velocity filter, the off diagonal matrix elements make sure that if you do have a velocity measurement, that the position is also corrected. So, the state estimate error covariance matrix 'learns' how variance in one state term affects the variance in the other state terms.
#
# ### Process Noise Covariance
#
# We'll initialise the process noise covariance as an Identity matrix as shown here. We're setting it up along the diagonal because this is a reasonable way to set up when we have no good information about the process. You can better model this term (particularly for more advanced filters) but for our purposes it leads to satisfactory tracking. Essentially we are saying that we believe each state term is effectively independent of each other state term. It turns out that even in cases where this is patently not true, it leads to reasonable behaviour and obviates the need to a detailed process model - which suits our purposes in this demonstration.
#
# $Q_{0} = \begin{bmatrix}
# 1 && 0 && 0 && 0 \\
# 0 && 1 && 0 && 0 \\
# 0 && 0 && 1 && 0 \\
# 0 && 0 && 0 && 1
# \end{bmatrix}$
#
#
# ### Measurement Terms
#
# We'll supply a measurement term (aka an observation term) later on dynamically to the filter for each frame we process. In terms of the shape of the measurement term its going to represent what we can measure - i.e. $x_{pos}$ and $y_{pos}$. So, our measurement term will be of this form:
#
# $measurement = \begin{bmatrix}
# x_{pos} \\
# y_{pos} \\
# \end{bmatrix}$
#
# ### Measurement uncertainty covariance
# Again, we'll supply our measurement noise co-variance values (aka our observation nose covariance) later on dynamically, but, in terms of shape it will look like this. We're assuming empirically a certain measurement noise in the two terms we're measuring $x_{pos}$ and $y_{pos}$.
#
# $R = \begin{bmatrix}
# x_{noise} && 0 \\
# 0 && y_{noise}
# \end{bmatrix}$
#
# We'll derive empirically $x_{noise}$ and $y_{noise}$ from yolo later for each frame we process.
#
# ### State-Transition Model and Observation Model
# Finally, we'll use two shape terms to tell a generic Kalman Filter algorithm what terms to multiply at each stage.
#
# $F$, the state transition model, will effectively enforce the constant velocity model on the process.
#
# $F =\begin{bmatrix}
# 1 && 0 && 1 && 0 \\
# 0 && 1 && 0 && 1 \\
# 0 && 0 && 1 && 0 \\
# 0 && 0 && 0 && 1
# \end{bmatrix}$
#
# $H$, the observation model matrix, effectively informs the other matrices in the ```kalman()``` function below that we are only measuring $x_{pos}$ and $y_{pos}$
#
# $H=\begin{bmatrix}
# 1 && 0 && 0 && 0 \\
# 0 && 1 && 0 && 0
# \end{bmatrix}$
#
# ### A note on 'Tuning' the Kalman Filter
# Setting each term along the diagonal of the $Q$ matrix (the process error covariance term) to 1 works as a 'bucket chemistry' tuning technique for filter responsiveness when working in conjunction with estimates for the $R$ matrix (the observation covariance). For some projects we have the time and resources to derive statistical models to accurately model $Q$ and $R$ but for some projects - such as this introductory example - its relatively common to tune them quickly by hand.
#
# Some tuning tips include:
# * Trial and error tuning is common;
# * Setting large values of $Q$ relative to $R$ tells the Kalman Filter to trust the observations more than the model;
# * setting large values of $R$ relative to $Q$ tells the Kalman Filter to trust the model more than the observations;
# * setting $Q$ to all ones along the diagonal is reasonably common as its relatively neutral and means you can tune by only adjusting $R$.
#
# ### A note on types
#
# Finally, we initialise everything using floats, to get a self-consistent set of data types - everything is floating point.
#
# ### Summary
#
# With these matrices shaped and initialized, we have specialised a generic ```kalman()``` routine to perform Kalman Filtering using a constant velocity model in two dimensions; x and y. For simplicity, we are not using $B_{k}$, the control-input model, in this example.
# + id="TZAHf_dQau78" colab_type="code" colab={}
def kalman_2d(x, P, observation, R, Q=np.matrix(np.eye(4))):
x, P = kalman(x, P, observation, R, Q,
# State Transition Model matrix, assuming constant velocity model (x, y, x_dot, y_dot)
F=np.matrix('''
1. 0. 1. 0.;
0. 1. 0. 1.;
0. 0. 1. 0.;
0. 0. 0. 1.
'''),
# Observation Model matrix, assuming we can only measure
# the co-ordinates x, y
H=np.matrix('''
1. 0. 0. 0.;
0. 1. 0. 0.
'''))
return x, P
# + [markdown] id="x4RRg22Hitb5" colab_type="text"
# #Kalman Filter
#
# Derived from Wikipedia
#
# See http://en.wikipedia.org/wiki/Kalman_filter
#
# Look back over Kalman Introduction and Kalman Maths for an insight into how Kalman is operating.
#
# The concept is:
# * For a low computational cost
# * Generate a state update/prediction
# * Generate a measurement prediction from that state
# * Calculate the difference between the predicted measurement and the actual measurement
# * Adjust the state update/prediction, and repeat....
#
# All done on normal probabilities - i.e. means and co-variances. Effectively each key term is held as two parameters - a mean term and a covariance term for that mean.
#
# To this filter, we supply the old state / variance and the new measurement value / variance and we take away a new state / variance.
#
# Two key terms to watch are the process error estimate covariance term and the observation error estimate covariance term as their interaction influences the filter responsiveness.
# + id="Cn7nKwdli4e9" colab_type="code" colab={}
def kalman(x, P, z, R, Q, F, H):
'''
Dynamic Parameters
x: state estimate
P: state estimate uncertainty covariance
z: observation
R: the covariance of the observation noise
Additionally
F: the state-transition model
H: the observation model - maps true state space into the observed space
Q: the covariance of the process noise
return:
updated and predicted new values for (x, P)
'''
# Update Step
# Update x and P based on measurement m
# distance between measured and current position-belief
# Innovation (or measurement) pre-fit residual
y = np.matrix(z).T - H * x
# Innovation (or pre-fit residual) co-variance
S = H * P * H.T + R
# Optimal Kalman Gain
try:
inv = S.I
except:
inv = np.linalg.pinv(S)
K = P * H.T * inv
# Updated (a posteriori state estimate)
x = x + K * y
# Updated a posteriori estimate co-variance
I = np.matrix(np.eye(F.shape[0]))
P = (I - K*H)*P
# Predict Step
# Predict x and P
x = F*x
P = F*P*F.T + Q
return x, P
# + [markdown] id="d7dlyv0Nk7EU" colab_type="text"
# #Demo
#
# ## Program Execution
# For each file:
# * get centre of detection (if any) and confidence from Yolo
# * feed Kalman with these values
# * Extract $x_{pos}$ and $y_{pos}$ from Kalman state term.
# * Print original centre of detection
# * Print filtered centre of detection
#
# ## Initialisation
#
# ### State Estimate Initialisation
# We need to initialize the state estimate vector to something. We'll initialise it to all zeros for reproducibility.
#
#
# $x_{0} = \begin{bmatrix}
# 0 \\
# 0 \\
# 0 \\
# 0 \\
# \end{bmatrix}$
#
# ### State Estimate Error Covariance Initialisation
# We need to initialize the state estimate error covariance matrix (estimated accuracy of state estimate). We'll initialise it along the diagonal as being highly uncertain as shown here.
#
# $P_{0} = \begin{bmatrix}
# 100 && 0 && 0 && 0 \\
# 0 && 100 && 0 && 0 \\
# 0 && 0 && 100 && 0 \\
# 0 && 0 && 0 && 100
# \end{bmatrix}$
#
# Typically we initialize the $P$ matrix on the diagonal, with each component corresponding to the expected variance in the corresponding state term, i.e. how much deviation you might expect in the initialization of that state term. If you have no insight into the behaviour of your model, this is a fairly typical way to get started while figuring out a better $P$ matrix initialisation.
#
# ### Observation Estimate Noise Covariance
# And we'll set up a matrix to hold R, the measurement uncertainty (or more formally the estimated covariance of the observation noise.
#
# $R_{0} = \begin{bmatrix}
# 0 && 0 \\
# 0 && 0
# \end{bmatrix}$
#
# As we process each frame, we're going to adjust this term based on the confidence yolo has in its detection.
# + id="56tX2yoKlFiS" colab_type="code" colab={}
writer = None
def demo_kalman_2d():
global writer
# Initialise state estimate (x, y, x_dot, y_dot) to no position
x = np.matrix('0. 0. 0. 0.').T
# Initialise uncertainty to all highly uncertain
P = np.matrix(np.eye(4))*100
raw_centres = [] # a list of unfiltered centres
filtered_centres = [] # a list of filtered centres
R = np.zeros(shape=(2, 2)) # a shape to hold measurement uncertainty
bb_lines, pngfiles = get_pngs_and_boxes()
for pngfile in pngfiles:
#print("handling .." + os.path.basename(pngfile))
img = cv2.imread(pngfile)
# Derive R from yolo confidence level in detection
raw_centre, conf = parse_detections(bb_lines, pngfile, img)
# Crudely derive R (the covariance of the observation noise).
# If yolo is confident we want a small
# uncertainty. If yolo isn't confident, translate to
# a large uncertainty.
R *= 1/conf
# Keep track of unfiltered bounding box centres - these will be
# the basis of our Kalman
raw_centres.append(raw_centre.astype(int))
# reshape observation for Kalman - it expects
# [x]
# [y]
# not [x,y]
observed = raw_centre.reshape(1, 2)
# Update the Kalman with state estimate, state covariance
# observation and observation noise estimate
x, P = kalman_2d(x, P, observed, R)
# just track x and y values from Kalman state
# (we just want to visualise positions)
filtered_centre = x[:2]
# Keep track of filtered x & y values
filtered_centres.append(filtered_centre)
# Pretty print a track of original centres
# and filtered centres
for i in range(1, len(raw_centres)):
# print unfiltered
cv2.line(img, tuple(raw_centres[i-1]),
tuple(raw_centres[i]), (0, 255, 0), 20)
# print filtered
cv2.line(img, tuple(filtered_centres[i-1]),
tuple(filtered_centres[i]), (255, 0, 0), 20)
# Resize and show the image
img = cv2.resize(img, (int(img.shape[1]/4), int(img.shape[0]/4)))
# Build a frame of our output video
if writer is None:
# Initialize our video writer
fourcc = cv2.VideoWriter_fourcc(*'VP80')
writer = cv2.VideoWriter('video.webm', fourcc, 30, (img.shape[1], img.shape[0]), True)
# Write the output frame to disk
writer.write(img)
# Release the file pointers
writer.release()
demo_kalman_2d()
# + [markdown] id="8fZJvZXeY15y" colab_type="text"
# **Video**
#
# Thia code plays the video we just made.
#
# The Kalman Filtered track plays in green, the unfilterered track plays in blue.
#
# As you can see, the Kalman Filtering has a role to play in predicting a reasonable guess for where the object might be while it is off-camera.
# + id="mxvG8It3RlWy" colab_type="code" colab={}
# Set this to 1 if video display
# is not working - works with chrome and firefox, not with safari
videoBodge = 0
def arrayShow (imageArray):
ret, png = cv2.imencode('.png', imageArray)
encoded = base64.b64encode(png)
return Image(data=encoded.decode('ascii'))
if(videoBodge == 0):
from IPython.display import HTML
from base64 import b64encode
webm = open('video.webm','rb').read()
data_url = "data:video/webm;base64," + b64encode(webm).decode()
else:
video = cv2.VideoCapture("video.webm")
while(video.isOpened()):
clear_output(wait=True)
ret, frame = video.read()
if(ret == False):
break
lines, columns, _ = frame.shape
img = arrayShow(frame)
display(img)
time.sleep(1)
# + id="DewKk12799jm" colab_type="code" colab={}
# Display Video
HTML("""
<video width=200 controls>
<source src="%s" type="video/webm">
</video>
""" % data_url)
# + [markdown] id="OH4IGJkZlO_u" colab_type="text"
# # Conclusion
#
# ## Exercises
# **Exercise 1**
# Simulate occluding the object being detected - for example, only supply every second measurement update to the Kalman algorithm and observe the Kalman predictions.
#
# **Exercise 2**
# Again, similarly to the Bayes lab, vary the initial state covariance, process covariance and the measurement covariance relative to each other and observe how that affects the Kalman Filter's predictions.
#
# **Advanced Exercise**
# Think about how you might extend the model to account for acceleration in x and y.
#
# ## Takeaways
# 1. You've seen a Kalman Filter used for single object tracking
# 2. You've seen that a Kalman Filter can help deal with occlusions - i.e. in this example the object being tracked disappeared for a few frames and the Kalman continued to predict motion for it based on its model.
# 3. You've seen that a typical approach to writing a Kalman is to develop the core algorithm independent of the number of terms in the state variable and to specialise it for a particular model.
#
# ## Next Steps
# 1. We'll see the Kalman's strengths lie with predictable behaviour (typically referred to as a linear model) and we'll look at a derivitive technique (the Particle Filter) that attempts to improve the Kalman in the presence of less predictable behaviour (aka a non-linear model).
| Lab_7_2_KalmanFilter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # 机器学习工程师纳米学位
# ## 强化学习
# ## 项目 4: 训练智能出租车学会驾驶
#
# 欢迎来到机器学习工程师纳米学位的第四个项目!在这个notebook文件中,模板代码已经提供给你,有助于你对*智能出租车*的分析和实现学习算法。你无须改动已包含的代码,除非另有要求。 你需要回答notebook文件中给出的与项目或可视化相关的问题。每一个你要回答的问题前都会冠以**'问题 X'**。仔细阅读每个问题,并在后面**'回答'**文本框内给出完整的回答。你提交的项目会根据你对于每个问题的回答以及提交的`agent.py`的实现来进行评分。
#
# >**提示:** Code 和 Markdown 单元格可通过 **Shift + Enter** 快捷键来执行。此外,Markdown可以通过双击进入编辑模式。
# + [markdown] deletable=true editable=true
# -----
#
# ## 开始
# 在这个项目中,你将构建一个优化的Q-Learning驾驶代理程序,它会操纵*智能出租车* 通过它的周边环境到达目的地。因为人们期望*智能出租车*要将乘客从一个地方载到另一个地方,驾驶代理程序会以两个非常重要的指标来评价:**安全性**和**可靠性**。驾驶代理程序在红灯亮时仍然让*智能出租车*行驶往目的地或者勉强避开事故会被认为是**不安全**的。类似的,驾驶代理程序频繁地不能适时地到达目的地会被认为**不可靠**。最大化驾驶代理程序的**安全性**和**可靠性**保证了*智能出租车*会在交通行业获得长期的地位。
#
# **安全性**和**可靠性**用字母等级来评估,如下:
#
# | 等级 | 安全性 | 可靠性 |
# |:-----: |:------: |:-----------: |
# | A+ | 代理程序没有任何妨害交通的行为,<br/>并且总是能选择正确的行动。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的100%。 |
# | A | 代理程序有很少的轻微妨害交通的行为,<br/>如绿灯时未能移动。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的90%。 |
# | B | 代理程序频繁地有轻微妨害交通行为,<br/>如绿灯时未能移动。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的80%。 |
# | C | 代理程序有至少一次重大的妨害交通行为,<br/>如闯红灯。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的70%。 |
# | D | 代理程序造成了至少一次轻微事故,<br/>如绿灯时在对面有车辆情况下左转。 | 代理程序在合理时间内到达目的地的次数<br />占行驶次数的60%。 |
# | F | 代理程序造成了至少一次重大事故,<br/>如有交叉车流时闯红灯。 | 代理程序在合理时间内到达目的地的次数<br />未能达到行驶次数的60%。 |
#
# 为了协助评估这些重要的指标,你会需要加载可视化模块的代码,会在之后的项目中用到。运行下面的代码格来导入这个代码,你的分析中会需要它。
# + deletable=true editable=true
# Import the visualization code
import visuals as vs
# Pretty display for notebooks
# %matplotlib inline
# + [markdown] deletable=true editable=true
# ### 了解世界
# 在开始实现你的驾驶代理程序前,首先需要了解*智能出租车*和驾驶代理程序运行的这个世界(环境)。构建自我学习的代理程序重要的组成部分之一就是了解代理程序的特征,包括代理程序如何运作。原样直接运行`agent.py`代理程序的代码,不需要做任何额外的修改。让结果模拟运行一段时间,以观察各个不同的工作模块。注意在可视化模拟程序(如果启用了),**白色车辆**就是*智能出租车*。
# + [markdown] deletable=true editable=true
# ### 问题 1
# 用几句话,描述在运行默认的`agent.py`代理程序中,你在模拟程序里观察到了什么。一些你可以考虑的情况:
# - *在模拟过程中,智能出租车究竟移动了吗?*
# - *驾驶代理程序获得了什么样的奖励?*
# - *交通灯的颜色改变是如何影响奖励的?*
#
# **提示:** 从顶层的`/smartcab/`目录(这个notebook所在的地方),运行命令
# ```bash
# 'python smartcab/agent.py'
# ```
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# ### 理解代码
# 除了要了解世界之外,还需要理解掌管世界、模拟程序等等如何运作的代码本身。如果一点也不去探索一下*“隐藏”*的器件,就试着去创建一个驾驶代理程序会很难。在顶层的`/smartcab/`的目录下,有两个文件夹:`/logs/` (之后会用到)和`/smartcab/`。打开`/smartcab/`文件夹,探索每个下面的Python文件,然后回答下面的问题。
# + [markdown] deletable=true editable=true
# ### 问题 2
# - *在*`agent.py`* Python文件里,选择 3 个可以设定的 flag,并描述他们如何改变模拟程序的。*
# - *在*`environment.py`* Python文件里,当代理程序执行一个行动时,调用哪个Environment类的函数?*
# - *在*`simulator.py`* Python 文件里,*`'render_text()'`*函数和*`'render()'`*函数之间的区别是什么?*
# - *在*`planner.py`* Python文件里,*`'next_waypoint()`* 函数会先考虑南北方向还是东西方向?*
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# -----
# ## 实现一个基本的驾驶代理程序
#
# 创建一个优化Q-Learning的驾驶代理程序的第一步,是让代理程序确实地执行有效的行动。在这个情况下,一个有效的行动是`None`(不做任何行动)、`'Left'`(左转)、`'Right'`(右转)或者`'Forward'`(前进)。作为你的第一个实现,到`'choose_action()'`代理程序函数,使驾驶代理程序随机选择其中的一个动作。注意你会访问到几个类的成员变量,它们有助于你编写这个功能,比如`'self.learning'`和`'self.valid_actions'`。实现后,运行几次代理程序文件和模拟程序来确认你的驾驶代理程序每步都执行随机的动作。
# + [markdown] deletable=true editable=true
# ### 基本代理程序模拟结果
# 要从最初的模拟程序获得结果,你需要调整下面的标志:
# - `'enforce_deadline'` - 将此标志设定为`True`来强制驾驶代理程序捕获它是否在合理时间内到达目的地。
# - `'update_delay'` - 将此标志设定为较小数值(比如`0.01`)来减少每次试验中每步之间的时间。
# - `'log_metrics'` - 将此标志设定为`True`将模拟结果记录为在`/logs/`目录下的`.csv`文件。
# - `'n_test'` - 将此标志设定为`'10'`则执行10次测试试验。
#
# 可选的,你还可以通过将`'display'`标志设定为`False`来禁用可视化模拟(可以使得试验跑得更快)。调试时,设定的标志会返回到他们的默认设定。重要的是要理解每个标志以及它们如何影响到模拟。
#
# 你成功完成了最初的模拟后(有20个训练试验和10个测试试验),运行下面的代码单元格来使结果可视化。注意运行同样的模拟时,日志文件会被覆写,所以留意被载入的日志文件!在 projects/smartcab 下运行 agent.py 文件。
# + deletable=true editable=true
# Load the 'sim_no-learning' log file from the initial simulation results
vs.plot_trials('sim_no-learning.csv')
# + [markdown] deletable=true editable=true
# ### 问题 3
# 利用上面的从你初始模拟中得到的可视化结果,给出关于驾驶代理程序的分析和若干观察。确保对于可视化结果上的每个面板你至少给出一条观察结果。你可以考虑的一些情况:
# - *驾驶代理程序多频繁地做出不良决策?有多少不良决策造成了事故?*
# - *假定代理程序是随机驾驶,那么可靠率是否合理?*
# - *代理程序对于它的行动会获得什么样的奖励?奖励是否表明了它收到严重的惩罚?*
# - *随着试验数增加,结果输出是否有重大变化?*
# - *这个智能出租车对于乘客来说,会被人为是安全的且/或可靠的吗?为什么或者为什么不?*
# + [markdown] deletable=true editable=true
# **答案:**
# + [markdown] deletable=true editable=true
# -----
# ## 通知驾驶代理程序
# 创建一个优化Q-Learning的驾驶代理程序的第二步,是定义一系列代理程序会在环境中发生的状态。根据输入、感知数据和驾驶代理程序可用的变量,可以为代理程序定义一系列状态,使它最终可以*学习*在一个状态下它需要执行哪个动作。对于每个状态的`'如果这个处于这个状态就那个行动'`的状况称为**策略**,就是最终驾驶代理程序要学习的。没有定义状态,驾驶代理程序就不会明白哪个动作是最优的——或者甚至不会明白它要关注哪个环境变量和条件!
# + [markdown] deletable=true editable=true
# ### 识别状态
# 查看`'build_state()'`代理程序函数,它显示驾驶代理函数可以从环境中获得下列数据:
# - `'waypoint'`,*智能出租车*去向目的地应该行驶的方向,它是*智能出租车*车头方向的相对值。
# - `'inputs'`,*智能出租车*的感知器数据。它包括
# - `'light'`,交通灯颜色。
# - `'left'`,*智能出租车*左侧车辆的目的方向。如果没有车辆,则返回`None`。
# - `'right'`,*智能出租车*右侧车辆的目的方向。如果没有车辆,则返回`None`。
# - `'oncoming'`,*智能出租车*交叉方向车辆的目的方向。如果没有车辆,则返回`None`。
# - `'deadline'`,*智能出租车*在时间之内到达目的地还所需的剩余动作数目。
# + [markdown] deletable=true editable=true
# ### 问题 4
# *代理程序的哪些可用特征与学习**安全性**和**效率**相关性最高?你为什么认为这些特征适合在环境中对**智能出租车**建模?如果你没有选择某些特征,放弃他们的原因是什么?*
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# ### 定义状态空间
# 当定义一系列代理程序会处于的状态,必需考虑状态空间的*大小*。就是说,如果你期望驾驶代理程序针对每个状态都学习一个**策略**,你会需要对于每一个代理状态都有一个最优的动作。如果所有可能状态的数量非常大,最后会变成这样的状况,驾驶代理程序对于某些状态学不到如何行动,会导致未学习过的决策。例如,考虑用下面的特征定义*智能出租车*的状态的情况:
#
# `('is_raining', 'is_foggy', 'is_red_light', 'turn_left', 'no_traffic', 'previous_turn_left', 'time_of_day')`.
# 发生如`(False, True, True, True, False, False, '3AM')`的状态的频次如何?没有近乎无限数量的训练,很怀疑代理程序会学到一个合适的动作!
# + [markdown] deletable=true editable=true
# ### 问题 5
# *如果用你在**问题4**中选择的特征来定义一个状态,状态空间的大小是多少?假定你了解环境以及它是如何模拟的,你觉得经过合理数量的训练之后,代理驾驶能学到一个较好的策略吗?(遇见绝大部分状态都能作出正确决策。)*
# **提示:** 考虑特征*组合*来计算状态的总数!
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# ### 更新驾驶代理程序的状态
# 要完成你的第二个实现,去到`'build_state()'`代理程序函数。根据你在**问题4**给出的判断,你现在要将`'state'`变量设定为包含所有Q-Learning所需特征的元组。确认你的驾驶代理程序通过运行代理程序文件和模拟会更新它的状态,注意状态是否显示了。如果用了可视化模拟,确认更新的状态和在模拟程序里看到的一致。
#
# **注意:** 观察时记住重置模拟程序的标志到默认设定!
# + [markdown] deletable=true editable=true
# -----
# ## 实现Q-Learning驾驶代理程序
# 创建一个优化Q-Learning的驾驶代理程序的第三步,是开始实现Q-Learning自身的功能。Q-Learning的概念相当直接:每个访问的状态,为所有可用的状态-行动配对在Q-table里创建一条记录。然后,当代理程序遇到一个状态并执行了一个动作,基于获得的奖励和设定的相互的更新规则,来更新关联的状态-动作配对的Q-value。当然,Q-Learning还带来其他的收益,如此我们可以让代理程序根据每个可能的状态-动作配对的Q-values,来为每个状态选择*最佳*动作。在这个项目里,你会实现一个*衰减* $\epsilon$ *-贪心* 的Q-learning算法,不含折扣因子。遵从每个代理程序函数的**TODO**下的实现指导。
#
# 注意代理程序的属性`self.Q`是一个字典:这就是Q-table的构成。每个状态是`self.Q`字典的键,每个值是另一个字典,包含了*action*和*Q-value*。这里是个样例:
#
# ```
# { 'state-1': {
# 'action-1' : Qvalue-1,
# 'action-2' : Qvalue-2,
# ...
# },
# 'state-2': {
# 'action-1' : Qvalue-1,
# ...
# },
# ...
# }
# ```
#
# 此外,注意你要求利用一个*衰减*$\epsilon$*(探索)因子*。因此,随着试验的增加,$\epsilon$会向0减小。这是因为,代理程序会从它的行为中学习,然后根据习得的行为行动。而且当$\epsilon$达到特定阈值后(默认阈值为0.01),代理程序被以它所学到的东西来作检测。作为初始的Q-Learning实现,你将实现一个线性衰减$\epsilon$的函数。
# + [markdown] deletable=true editable=true
# ### Q-Learning模拟结果
# 要从最初的Q-learning程序获得结果,你需要调整下面的标志和设置:
# - `'enforce_deadline'` - 将此标志设定为`True`来强制驾驶代理程序捕获它是否在合理时间内到达目的地。
# - `'update_delay'` - 将此标志设定为较小数值(比如`0.01`)来减少每次试验中每步之间的时间。
# - `'log_metrics'` - 将此标志设定为`True`将模拟结果记录为在`/logs/`目录下的`.csv`文件,Q-table存为`.txt`文件。
# - `'n_test'` - 将此标志设定为`'10'`则执行10次测试试验。
# - `'learning'` - 将此标志设定为`'True'`来告诉驾驶代理使用你的Q-Learning实现。
#
# 此外,使用下面的$\epsilon$衰减函数:
#
# $$ \epsilon_{t+1} = \epsilon_{t} - 0.05, \hspace{10px}\textrm{for trial number } t$$
#
# 如果你在实施时遇到困难,尝试把`'verbose'`标志设为`True`来调试。调试时,在这里设定的标志会返回到它们的默认设定。重要的是你要理解每个标志做什么并且解释它们怎么影响模拟!
#
# 当你成功完成初始的Q-Learning模拟程序后,运行下面代码单元格来使结果可视化。注意当相同的模拟运行时,log文件会被覆写,所以要留意载入的log文件!
# + deletable=true editable=true
# Load the 'sim_default-learning' file from the default Q-Learning simulation
vs.plot_trials('sim_default-learning.csv')
# + [markdown] deletable=true editable=true
# ### 问题 6
# 利用上面的从你默认的Q-Learning模拟中得到的可视化结果,像在**问题3**那样,给出关于驾驶代理程序的分析和若干观察。注意模拟程序应该也产生了Q-table存在一个文本文件中,可以帮到你观察代理程序的算法。你可以考虑的一些情况:
# - *有没有观察到基本驾驶代理程序和默认的Q-Learning代理程序的相似之处?*
# - *在测试之前驾驶代理大约需要做多少训练试验?在给定的$\epsilon$ 容忍度下,这个数字是否合理?*
# - *你实现的$\epsilon$(探索因子)衰减函数是否准确地在参数面板中显示?*
# - *随着试验数增加,不良动作的数目是否减少?平均奖励是否增加?*
# - *与初始的驾驶代理程序相比,安全性和可靠性评分怎样?*
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# -----
# ## 改进Q-Learning驾驶代理程序
# 创建一个优化Q-Learning的驾驶代理程序的第三步,是执行优化!现在Q-Learning算法已经实现并且驾驶代理程序已经成功学习了,需要调整设定、调节参数让驾驶代理程序学习**安全性**和**效率**。通常这一步需要很多试验和错误,因为某些设定必定会造成更糟糕的学习。要记住的一件事是学习的行为本身和需要的时间:理论上,我们可以允许代理程序用非常非常长的时间来学习;然而,Q-Learning另一个目的是*将没有习得行为的试验试验变为有习得行为的行动*。例如,训练中总让代理程序执行随机动作(如果$\epsilon = 1$并且永不衰减)当然可以使它*学习*,但是不会让它*行动*。当改进你的Q-Learning实现时,要考虑做一个特定的调整的意义,以及它是否逻辑上是否合理。
# + [markdown] deletable=true editable=true
# ### 改进Q-Learning的模拟结果
# 要从最初的Q-learning程序获得结果,你需要调整下面的标志和设置:
# - `'enforce_deadline'` - 将此标志设定为`True`来强制驾驶代理程序捕获它是否在合理时间内到达目的地。
# - `'update_delay'` - 将此标志设定为较小数值(比如`0.01`)来减少每次试验中每步之间的时间。
# - `'log_metrics'` - 将此标志设定为`True`将模拟结果记录为在`/logs/`目录下的`.csv`文件,Q-table存为`.txt`文件。
# - `'learning'` - 将此标志设定为`'True'`来告诉驾驶代理使用你的Q-Learning实现。
# - `'optimized'` - 将此标志设定为`'True'`来告诉驾驶代理你在执行一个优化版本的Q-Learning实现。
#
# 优化Q-Learning代理程序可以调整的额外的标志:
# - `'n_test'` - 将此标志设定为某个正数(之前是10)来执行那么多次测试试验。
# - `'alpha'` - 将此标志设定为0 - 1之间的实数来调整Q-Learning算法的学习率。
# - `'epsilon'` - 将此标志设定为0 - 1之间的实数来调整Q-Learning算法的起始探索因子。
# - `'tolerance'` - 将此标志设定为某个较小的大于0的值(默认是0.05)来设定测试的epsilon阈值。
#
# 此外,使用一个你选择的$\epsilon$ (探索因子)衰减函数。注意无论你用哪个函数,**一定要以合理的速率衰减**到`'tolerance'`。Q-Learning代理程序到此才可以开始测试。某个衰减函数的例子($t$是试验的数目):
#
# $$ \epsilon = a^t, \textrm{for } 0 < a < 1 \hspace{50px}\epsilon = \frac{1}{t^2}\hspace{50px}\epsilon = e^{-at}, \textrm{for } 0 < a < 1 \hspace{50px} \epsilon = \cos(at), \textrm{for } 0 < a < 1$$
#
# 如果你想的话,你也可以使用$\alpha$ (学习率) 的衰减函数,当然这通常比较少见。如果你这么做了,确保它满足不等式$0 \leq \alpha \leq 1$。
# 如果你在实施时遇到困难,尝试把`'verbose'`标志设为`True`来调试。调试时,在这里设定的标志会返回到它们的默认设定。重要的是你要理解每个标志做什么并且解释它们怎么影响模拟!
#
# 当你成功完成初始的Q-Learning模拟程序后,运行下面代码单元格来使结果可视化,请注意为了达到项目要求你需要在安全性和可靠性上获得至少都为A的评分。注意当相同的模拟运行时,log文件会被覆写,所以要留意载入的log文件!
# + deletable=true editable=true
# Load the 'sim_improved-learning' file from the improved Q-Learning simulation
vs.plot_trials('sim_improved-learning.csv')
# + [markdown] deletable=true editable=true
# ### 问题7
# 利用上面的从你改进的Q-Learning模拟中得到的可视化结果,像在**问题6**那样,给出关于改进的驾驶代理程序的最终分析和观察。你需要回答的问题:
# - *使用了什么epsilon(探索因子)的衰减函数?*
# - *在测试之前驾驶代理大约需要做多少训练试验?*
# - *你用了什么epsilon-tolerance和alpha(学习率)值?为什么?*
# - *与之前的默认Q-Learning学习器相比,这个Q-Learning学习器有多少改进? *
# - *你会说Q-Learning学习器的结果表明了你的驾驶代理程序成功地学习了一个合适的策略吗?*
# - *你对*智能出租车*的安全性和可靠性评分满意吗?*
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# ### 定义一个最优策略
#
# 有时,对于重要的问题*“我要让我的代理程序学习什么?”*的答案,只是理论性的,无法具体描述。然而这里,你可以具体定义代理程序要学什么,就是美国通行权交通法案。这些法律是已知信息,你可以基于这些法律,为*智能出租车*进一步定义每一个状态所做的最优动作。在那种情况下,我们称这一系列最优状态-动作配对为**最优策略**。因此,不像那些理论性的回答,不仅通过收到的奖励(惩罚),而且纯观察,代理程序是否在“错误”地行动能很清晰地得知。如果代理程序闯了红灯,我们既看见它获得了一个负面奖励,也知道这是一个错误的行为。这可以用来帮你验证驾驶代理程序习得的**策略**是否正确,或只是个**次优策略**。
# + [markdown] deletable=true editable=true
# ### 问题 8
# 给出几个关于最优策略是什么样子的例子(用你已定义的状态)。之后,查看`'sim_improved-learning.txt'`文本文件,看你的改进的Q-Learning算法的结果。_每个从模拟中纪录的状态,对于给定的状态,**策略**(得分最高的动作)是否正确?是否对于有些状态,有策略不同于预期的最优策略?_给出一个状态和记录的状态-动作的奖励,解释为什么正是个正确的策略。
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# -----
# ### 选做:未来奖励 - 折扣因子 `'gamma'`
# 也许你会好奇,作为Q-Learning算法的一部分,之前要求你在实现中**不要**使用折扣引子`'gamma'`。在算法中包含未来奖励能有助于在未来状态回溯到当前状态时的反向正面奖励。本质上,如果给予驾驶代理程序执行若干动作到达不同状态的选择,包含未来奖励会是代理程序偏向可以得到更多奖励的状态。一个例子是驶向目的的驾驶代理程序:所有行动和奖励都相等,那么理论上如果到达目的地会有额外奖励,驶向目的会获得更好的奖励。然而,即使在这个项目里,驾驶代理程序也要在规定的时间里到达目的地,包含未来奖励不会有益于代理程序。实际上,如果代理程序给予多次试验学习,它甚至会给Q-value带来负面影响!
# + [markdown] deletable=true editable=true
# ### 可选问题 9
# *在项目中有两个特点使得未来奖励在这个Q-Learning算法无效。一个特点是关于*智能出租车*本身,另一个是关于环境。你能指出它们是什么以及为什么未来奖励不会在这个项目中起效?*
# + [markdown] deletable=true editable=true
# **回答:**
# + [markdown] deletable=true editable=true
# > **注意**:当你写完了所有的代码,并且回答了所有的问题。你就可以把你的 iPython Notebook 导出成 HTML 文件。你可以在菜单栏,这样导出**File -> Download as -> HTML (.html)**把这个 HTML 和这个 iPython notebook 一起做为你的作业提交。
| Notes/P3 SmartCab/smartcab/.ipynb_checkpoints/smartcabzh-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HighLvRiver/HighLvRiver.github.io/blob/master/AIC_Project1_Jayden_v_Surprise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="akL3g19N8fmp" colab_type="code" outputId="6c2e7c3c-54af-4d77-c372-4750dfcbd1f6" colab={"base_uri": "https://localhost:8080/", "height": 98}
# !pip3 install scikit-surprise
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import surprise
import time
from surprise.model_selection import cross_validate, train_test_split
from surprise import accuracy
from surprise.accuracy import rmse
from surprise import NormalPredictor
from surprise import KNNBasic
from surprise import KNNWithMeans
from surprise import KNNWithZScore
from surprise import KNNBaseline
from surprise import SVD
from surprise import BaselineOnly
from surprise import SVDpp
from surprise import NMF
from surprise import SlopeOne
from surprise import CoClustering
# + id="sfyGlG0-8h9M" colab_type="code" outputId="4a87292d-9f97-45fe-8b86-4320e8213f6b" colab={"base_uri": "https://localhost:8080/", "height": 367}
# %time train = pd.read_csv("https://www.dropbox.com/s/hp2dn6l6op4n4mv/watcha_aic_train_data_for_problem_1.csv?dl=1")
train.head(10)
# + id="ogx2CJTC8iXo" colab_type="code" outputId="c482296f-0bd4-4cff-b72b-0fee6163c63a" colab={"base_uri": "https://localhost:8080/", "height": 367}
# %time test = pd.read_csv("https://www.dropbox.com/s/0g70q9k7pm5so1n/watcha_aic_test_data_for_problem_1.csv?dl=1")
test.head(10)
# + id="9ByaUtED8j0r" colab_type="code" outputId="b3717c4a-9ad9-435f-94db-86695e0dd632" colab={"base_uri": "https://localhost:8080/", "height": 33}
test.shape
# + id="SYXK0tX18lMS" colab_type="code" outputId="d145ee41-8fbb-4f3a-eee5-9adb25fc78fc" colab={"base_uri": "https://localhost:8080/", "height": 33}
train.shape
# + id="nOsWt8oU8mW5" colab_type="code" colab={}
# 최종 점수 산정 함수
def score(ans, pred):
if ans == 0:
return 0
elif ans == pred:
return 1
else:
return -1
# + id="IItj-tLb8n0s" colab_type="code" colab={}
# 최대 점수 체크하기 : 2,973,129
# # %time test.apply(lambda s: score(s['preference'],s['preference']), axis=1).sum() # 2973129
# + [markdown] id="OtfApz-n1_fK" colab_type="text"
#
# + id="uYnkhvy98pnc" colab_type="code" outputId="d82b2a7f-15c7-41d3-c51f-4b9305910532" colab={"base_uri": "https://localhost:8080/", "height": 330}
plt.hist(train['preference'])
# + id="vxF9YkVC9PKY" colab_type="code" outputId="111b07a7-f880-4333-a3ef-fc9e43f2da90" colab={"base_uri": "https://localhost:8080/", "height": 191}
train['preference'] = train['preference'] + 2
train.head()
# + id="iBMN6xt49jyK" colab_type="code" colab={}
test['preference'] = test['preference'] + 2
# + id="_ZyFhmw1_uYu" colab_type="code" colab={}
# sample = train.sample(n=10000, random_state=1)
# + id="fhKrJpxXEThg" colab_type="code" colab={}
train = train[train['preference'] != 2]
# + id="eViJXzXRCdHx" colab_type="code" outputId="bf5f26e0-28b8-41c7-979e-b1e429dec3e7" colab={"base_uri": "https://localhost:8080/", "height": 330}
plt.hist(train['preference'])
# + id="qQqFV5mnJ7xa" colab_type="code" colab={}
reader = surprise.Reader(rating_scale=(1,3))
# + id="MLPzmdvz-m6b" colab_type="code" colab={}
sample_data = surprise.Dataset.load_from_df(sample, reader)
# + id="CSrrmTJc_GzC" colab_type="code" outputId="bc403433-6b9a-4639-cec4-115a3766716d" colab={"base_uri": "https://localhost:8080/", "height": 822}
benchmark = []
for algorithm in [SVD(), NMF(), NormalPredictor()
, KNNBaseline(), KNNBasic(), KNNWithMeans(), KNNWithZScore()
, BaselineOnly(), CoClustering()]:
results = cross_validate(algorithm, sample_data, measures=['RMSE'], cv=3, verbose=False)
tmp = pd.DataFrame.from_dict(results).mean(axis=0)
tmp = tmp.append(pd.Series([str(algorithm).split(' ')[0].split('.')[-1]], index=['Algorithm']))
benchmark.append(tmp)
pd.DataFrame(benchmark).set_index('Algorithm').sort_values('test_rmse')
# + id="i8ZbdNbfBRoq" colab_type="code" colab={}
# bsl_option = {'method':'als',
# 'n_epochs':5,
# 'reg_u':12,
# 'reg_i':5
# }
# train_data = surprise.Dataset.load_from_df(train, reader)
train_data = surprise.Dataset.load_from_df(sample, reader)
# + id="J27oOC8pEIAF" colab_type="code" outputId="6aa1e847-4ba4-4f97-ec2e-2677aa4d8809" colab={"base_uri": "https://localhost:8080/", "height": 33}
start = time.time()
algo = SVD()
algo.fit(train_data.build_full_trainset())
print('Training time(s) :'+str(int(time.time() - start)))
# + id="RDNpD5U1EKwr" colab_type="code" outputId="1071c559-cbd1-4afb-d8d5-e1ec435af01c" colab={"base_uri": "https://localhost:8080/", "height": 33}
print(algo.__class__.__name__)
# + id="ENC9bbFhIVrQ" colab_type="code" colab={}
# # %time test = pd.read_csv("https://www.dropbox.com/s/0g70q9k7pm5so1n/watcha_aic_test_data_for_problem_1.csv?dl=1")
# test.head(10)
# + id="yE0QgS4BEVh0" colab_type="code" colab={}
predictions = algo.test(np.array(test))
pred_ratings = np.array([pred.est for pred in predictions])
# + id="QfyLcgMnFTbp" colab_type="code" colab={}
plt.hist(pred_ratings)
# + id="JzF2E2KpFXg7" colab_type="code" colab={}
def pred_rat(x):
if x>2: return 1
else: return -1
# + id="3N15fv0bFY1f" colab_type="code" colab={}
test['pred']=pd.Series(map(pred_rat, pred_ratings))
# + id="0EiSh25HFbUb" colab_type="code" colab={}
test['ans']= test.apply(lambda x:score(x['preference'], x['pred']), axis=1)
# + id="YiJtudNsFeRr" colab_type="code" outputId="790cff04-7375-42b7-b1dd-88f75085ca8e" colab={"base_uri": "https://localhost:8080/", "height": 335}
test.head(10)
# + id="6znWLZO8Ffzu" colab_type="code" outputId="5473ceb5-fe82-4a51-8<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 33}
np.sum(test['ans'])
# + id="EaO10rowFEJM" colab_type="code" colab={}
sample = train.sample(frac=0.1, random_state=1)
# + id="9Jf4tVCsD4D4" colab_type="code" colab={}
prediction_test = algo.test(sample)
| AIC_Project1_Jayden_v_Surprise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spartan Data Science
#
# # Imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
train = pd.read_csv('G:/Titanic/train.csv')
test = pd.read_csv('G:/Titanic/test.csv')
print(train.columns.values)
print(test.columns.values)
train.info()
test.info()
# +
# As we can see from the above information of the training dataset
# that there are a lot of missing values in the Cabin column.
# +
# While the Age column contains a very few missing values which can be
# substituted by the average age value within each class i.e imputation.
# +
# Moreover, only two values missing in the Embarked column.
# -
# # Data Visualization
# The following Heatmap will reveal the missing values.
# White lines indicate the missing values.
sns.heatmap(train.isnull(), yticklabels=False, cbar=False, cmap="Reds")
# Checking how many survived vs. how many did not with respect to gender.
sns.set_style('whitegrid')
sns.countplot(x='Survived',hue='Sex',data=train,palette='Set2')
# Checking how many survived vs. how many did not with respect to class.
sns.set_style('whitegrid')
sns.countplot(x='Survived',hue='Pclass',data=train,palette='Accent_r',edgecolor=sns.color_palette("dark", 3))
# Checking the distribution of age
sns.distplot(train['Age'].dropna(),kde=False,color='red',bins=30)
# Checking the age groups of the people within each class.
# Grouped into classes
plt.figure(figsize=(12, 7))
sns.boxplot(x='Pclass',y='Age',data=train,palette='winter_r')
# Plotting people who came in groups or alone
sns.countplot(x = 'SibSp', data = train, edgecolor=sns.color_palette("dark", 3))
# Plotting the Fare column
sns.countplot(x = 'Fare', data = train)
# A better representation for the above distribution using pandas
train['Fare'].hist(bins=30,figsize=(10,4))
# And lastly, distribution for Parch
sns.countplot(x = 'Parch', data = train, edgecolor=sns.color_palette("dark", 3))
# # Data Preprocessing
# We'll perform the following tasks:
# 1. Take care of all the missing values
# 2. Convert Categorical Values into Dummy Variables so that the Machine Learning Model can interpret them.
# 3. Take care of the Multicolinearity issue by dropping one column of the dummy variables from each set of dummy variables.
#
# +
# Imputing the Age Column
def AgeImputation(column):
Age = column[0]
Pclass = column[1]
if pd.isnull(Age):
if Pclass == 1:
return 37
elif Pclass == 2:
return 29
else:
return 24
else:
return Age
train['Age'] = train[['Age','Pclass']].apply(AgeImputation,axis=1)
test['Age'] = test[['Age','Pclass']].apply(AgeImputation,axis=1)
# -
sns.heatmap(test.isnull(),yticklabels=False,cbar=False,cmap="Reds")
sns.heatmap(train.isnull(),yticklabels=False,cbar=False,cmap="Reds")
# Dropping the Cabin column because it has too many missing values. Imputing wont give accurate representation for the data.
train.drop('Cabin',axis=1,inplace=True)
test.drop('Cabin',axis=1,inplace=True)
sns.heatmap(train.isnull(),yticklabels=False,cbar=False,cmap="Reds")
# +
# Lastly, dealing with the Embarked Column.
# We're dropping the rows containing null values for any column column in the Training Set
train.dropna(inplace=True)
# fill (instead of drop) the missing value of Fare with the mean of Fares
# so that there are exactly 418 rows (required for submission)
mean = test['Fare'].mean()
test['Fare'].fillna(mean, inplace=True)
# -
# All missing values have been taken care of.
sns.heatmap(train.isnull(),yticklabels=False,cbar=False,cmap="Reds")
# All missing values have been taken care of.
sns.heatmap(test.isnull(),yticklabels=False,cbar=False,cmap="Reds")
train.info()
test.info()
# Let's see what are the current columns
train.head()
test.head()
# convert categorical variables into dummy/indicator variables
# drop_first drops one column to remove multi-colinearity i.e one or more columns predicting the other
sex = pd.get_dummies(train['Sex'],drop_first=True)
embark = pd.get_dummies(train['Embarked'],drop_first=True)
# dropping the Name and Ticket columns because they have no role in the model training and prediction
# dropping the Sex and Embarked columns to replace them with the new columns with dummy variables
train.drop(['Sex','Embarked','Name','Ticket'],axis=1,inplace=True)
train = pd.concat([train,sex,embark],axis=1)
train.head()
# Since passenger id wont give any information about their survival
train.drop(['PassengerId'],axis=1,inplace=True)
train.head()
# Repeating the above process for test
sex = pd.get_dummies(test['Sex'],drop_first=True)
embark = pd.get_dummies(test['Embarked'],drop_first=True)
test.drop(['Sex','Embarked','Name','Ticket'],axis=1,inplace=True)
test = pd.concat([test,sex,embark],axis=1)
test.head()
# Since passenger id wont give any information about their survival
P_ID = test['PassengerId'] # Saving for later
test.drop(['PassengerId'],axis=1,inplace=True)
test.head()
train.head()
P_ID.head()
train.info()
test.info()
P_ID
# # Preparing the Dataset for Machine Learning
# +
from sklearn.model_selection import train_test_split
X = train.drop('Survived', axis = 1)
y = train['Survived']
X_train, X_test, y_train, y_test = train_test_split(train.drop('Survived',axis=1),
train['Survived'], test_size = 0.30,
random_state=101)
# -
# # Random Forest Classifier
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(n_estimators = 10, criterion = 'entropy', random_state = 0)
classifier.fit(X_train, y_train)
survived = classifier.predict(test)
test['Survived'] = survived
test['PassengerId'] = P_ID
test.info()
test[['PassengerId', 'Survived']].to_csv('G:/Titanic/First_Random_Forest_Classifier.csv', index=False)
| Titanic - Random Forest (0.73205).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# requests is a library that helps get web pages
# urllib2 is an alternative
import requests
# ### Getting the content of a simple web page ###
# Get the webpage
response = requests.get('https://www.bloomberg.com/asia')
# Get the attributes
dir(response)
response.status_code
# * 200 : Success
# * 401 : Unauthorised
# * 400 : Bad request
# * 403 : Forbidden
# * 404 : Not found
# * 500 : Internal server error
response.headers # examine the nature of content returned
response.text # looks garbled because this is the actual text
# ### Getting actual JSON data ###
api_response = requests.get('https://eservices.mas.gov.sg/api/action/datastore/search.json?resource_id=d3aedcad-ad7a-4172-90eb-407ab4c50096&limit=5')
# json allows us to access the json data as a dict
data = api_response.json()
data.keys()
# We can continue to drill down
data['result']
len(data['result'])
data['result'].keys()
len(data['result']['records'])
for i in range(len(data['result']['records'])):
print(data['result']['records'][i])
| notebook/Requests_Stub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Rating Movies IMDB
# ## Importing the libraries
# +
from plotnine import *
from sklearn import metrics
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import itertools
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pickle
import seaborn as sns
# -
# ## Reading the data
data_frame = pd.read_csv('dataset/movie_metadata.csv')
data_frame.head(10)
data_frame.shape
data_frame.dtypes
list(data_frame.columns)
# ## Data analysis
data_frame.drop('movie_imdb_link', axis=1, inplace=True)
data_frame['color'].value_counts()
data_frame.drop('color', axis=1, inplace=True)
data_frame.isna().any()
data_frame.isna().sum()
data_frame.dropna(axis=0, subset=['director_name', 'num_critic_for_reviews', 'duration',
'director_facebook_likes', 'actor_3_facebook_likes', 'actor_2_name',
'actor_1_facebook_likes', 'actor_1_name', 'actor_3_name',
'facenumber_in_poster', 'num_user_for_reviews', 'language',
'country', 'plot_keywords', 'title_year',
'actor_2_facebook_likes'],
inplace=True)
data_frame.shape
data_frame['content_rating'].value_counts()
data_frame['content_rating'].fillna('R', inplace=True)
data_frame['aspect_ratio'].value_counts()
data_frame['aspect_ratio'].fillna(data_frame['aspect_ratio'].median(), inplace=True)
data_frame['budget'].fillna(data_frame['budget'].median(), inplace=True)
data_frame['gross'].fillna(data_frame['gross'].median(), inplace=True)
data_frame.isna().sum()
data_frame.duplicated().sum()
data_frame.drop_duplicates(inplace=True)
data_frame.shape
data_frame['language'].value_counts()
data_frame.drop('language', axis=1, inplace=True)
data_frame['country'].value_counts()
data_frame.drop('country', axis=1, inplace=True)
data_frame['profit'] = data_frame['budget'].sub(data_frame['gross'], axis=0)
data_frame.head(10)
data_frame['profit_percentage'] = (data_frame['profit'] / data_frame['gross']) * 100
data_frame.head(10)
data_frame.to_csv('exploratory_data_analysis_movies_imdb_.csv', index=False)
# ## Data visualization
ggplot(aes(x='imdb_score', y='profit'), data=data_frame) +\
geom_line() +\
stat_smooth(colour='blue', span=1)
ggplot(data_frame) +\
aes(x='imdb_score', y='movie_facebook_likes') +\
geom_line() +\
labs(title='IMDB Score vs Movie Facebook Likes', x='IMDB Score', y='Movie Facebook Likes')
# +
plt.figure(figsize=(10, 8))
data_frame = data_frame.sort_values(by='imdb_score', ascending=False)
data_frame_2 = data_frame.head(20)
ax = sns.pointplot(data_frame_2['actor_1_name'], data_frame_2['imdb_score'], hue=data_frame_2['movie_title'])
ax.set_xticklabels(ax.get_xticklabels(), rotation=40, ha="right")
plt.tight_layout()
plt.show()
# -
# ## Data preparation
data_frame.drop(columns=['director_name', 'actor_1_name', 'actor_2_name',
'actor_3_name', 'plot_keywords', 'movie_title'], axis=1, inplace=True)
data_frame['genres'].value_counts()
data_frame.drop('genres', axis=1, inplace=True)
data_frame.drop(columns=['profit', 'profit_percentage'], axis=1, inplace=True)
# +
corr = data_frame.corr()
sns.set_context("notebook", font_scale=1.0, rc={"lines.linewidth": 2.5})
plt.figure(figsize=(13, 7))
mask = np.zeros_like(corr)
mask[np.triu_indices_from(mask, 1)] = True
a = sns.heatmap(corr, mask=mask, annot=True, fmt='.2f')
rotx = a.set_xticklabels(a.get_xticklabels(), rotation=90)
roty = a.set_yticklabels(a.get_yticklabels(), rotation=30)
# -
data_frame['other_actors_facebook_likes'] = data_frame['actor_2_facebook_likes'] + data_frame['actor_3_facebook_likes']
data_frame.drop(columns=['actor_2_facebook_likes', 'actor_3_facebook_likes', 'cast_total_facebook_likes'],
axis=1,
inplace=True)
data_frame['critic_review_ratio'] = data_frame['num_critic_for_reviews'] / data_frame['num_user_for_reviews']
data_frame.drop(columns=['num_critic_for_reviews', 'num_user_for_reviews'], axis=1, inplace=True)
# +
corr = data_frame.corr()
sns.set_context("notebook", font_scale=1.0, rc={"lines.linewidth": 2.5})
plt.figure(figsize=(13, 7))
mask = np.zeros_like(corr)
mask[np.triu_indices_from(mask, 1)] = True
a = sns.heatmap(corr, mask=mask, annot=True, fmt='.2f')
rotx = a.set_xticklabels(a.get_xticklabels(), rotation=90)
roty = a.set_yticklabels(a.get_yticklabels(), rotation=30)
# -
data_frame['imdb_binned_score'] = pd.cut(data_frame['imdb_score'], bins=[0, 4, 6, 8, 10], right=True, labels=False) + 1
data_frame.head(10)
data_frame = pd.get_dummies(data=data_frame, columns=['content_rating'], prefix=['content_rating'], drop_first=True)
data_frame.head(10)
data_frame.to_csv('data_imdb_scored.csv', index=False)
X = pd.DataFrame(columns=['duration', 'director_facebook_likes', 'actor_1_facebook_likes', 'gross',
'num_voted_users', 'facenumber_in_poster', 'budget', 'title_year',
'aspect_ratio', 'movie_facebook_likes', 'other_actors_facebook_likes', 'critic_review_ratio',
'content_rating_G', 'content_rating_GP', 'content_rating_M', 'content_rating_NC-17',
'content_rating_Not Rated', 'content_rating_PG', 'content_rating_PG-13', 'content_rating_Passed',
'content_rating_R', 'content_rating_TV-14', 'content_rating_TV-G', 'content_rating_TV-PG',
'content_rating_Unrated', 'content_rating_X'],
data=data_frame)
y = pd.DataFrame(columns=['imdb_binned_score'], data=data_frame)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# +
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
# -
X.isna().sum()
# +
logit = LogisticRegression(verbose=1, max_iter=1000)
logit.fit(X_train, np.ravel(y_train, order='C'))
y_pred = logit.predict(X_test)
# -
y_pred
cnf_matrix = metrics.confusion_matrix(y_test, y_pred)
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
plot_confusion_matrix(cnf_matrix, classes=['1','2', '3', '4'],
title='Matriz de confusão não normalizada', normalize=False)
data_frame['imdb_binned_score'].value_counts()
print(metrics.classification_report(y_test, y_pred, target_names=['1','2', '3', '4']))
trained_model = 'imdb_movies_model.sav'
pickle.dump(logit, open(trained_model, 'wb'))
loaded_model = pickle.load(open(trained_model, 'rb'))
X_test[0]
loaded_model.predict([X_test[0]])
| rating-movies-imdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Two stream numerical code and tests
import numpy as np
import matplotlib.pylab as plt
# %matplotlib inline
tau,omega,xi,omegaSoil = 5.,0.5,0.0,0.0
nsd=5
theta=45
dtau=0.01
correct=False
plot=False
diffuse=True
# +
# how many layers in canopy: nz_canopy
nz_canopy = int(np.ceil(tau/(dtau)))
dt = 1./float(nz_canopy)
# need to know filter width : nz_filter
# value of tau for the filter : tauFilt
tauFilt = np.arange(-nsd,nsd,dt)
nz_filter = len(tauFilt)
# need to know total extent to store: nz
nz = int(nz_canopy+2*nz_filter)
nz = 2*int(nz/2)
# create canopy array of zeros : canopyI0
tauCanopy = np.arange(nz)*dt - nz_filter*dt
canopyI0 = np.zeros_like(tauCanopy)
# source term: put unity at top of
# canopy: index nz_filter-1
canopyI0[tauCanopy==0] = 1.0
canopyExtent = np.logical_and(tauCanopy>0,tauCanopy<=tau)
print(canopyExtent.sum()*dt)
# -
plt.plot(tauCanopy,canopyExtent)
# +
'''
Calculate Int(exp(-(t / mu) mu,dmu)/Int(exp(-(t / mu) mu,dmu)
scipy.special.expi
integral(exp(t)/t,t=-inf..x)
'''
n = 10001
dmu = 1./(n-1)
mu = np.arange(n)*dmu
mask = mu==0
m = np.zeros_like(mu)
m[~mask] = 1./mu[~mask]
m[mask] = 1e200
dtau = 10./(n-1)
tau = np.arange(n)*dtau
ee = []
for t in tau:
ee.append((np.exp(-t *m) * mu * dmu).sum()/( np.atleast_1d(mu * dmu)).sum())
# -
plt.plot(ee)
| 001_Overview.ipynb |
# # Trying the Julia programming language in the Jupyter Notebook
println("Hello world!")
f(z, c) = z.*z .+ c
f(2.0 + 1.0im, 1.0)
z = [-1.0 - 1.0im 1.0 - 1.0im;
-1.0 + 1.0im 1.0 + 1.0im]
z[1,end]
f(z, 0)
function julia(z, c; maxiter=200)
for n = 1:maxiter
if abs2(z) > 4.0
return n-1
end
z = f(z, c)
end
return maxiter
end
Pkg.add("PyCall")
using PyCall
@pyimport numpy as np
z = np.linspace(-1., 1., 100)
m = [julia(z[i], 0.5) for i=1:100]
Pkg.add("Gadfly")
using Gadfly
# + podoc={"output_text": "<Plot(...)>"}
plot(x=1:100, y=m, Geom.point, Geom.line)
# -
@time m = [julia(complex(r, i), complex(-0.06, 0.67))
for i = 1:-.001:-1,
r = -1.5:.001:1.5];
Pkg.add("PyPlot")
using PyPlot
# + podoc={"output_text": "<Julia output>"}
imshow(m, cmap="RdGy",
extent=[-1.5, 1.5, -1, 1]);
| Chapter05/12_julia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # ADM Quantities in terms of BSSN Quantities
#
# ## Author: <NAME>
# ### Formatting improvements courtesy <NAME>
#
# [comment]: <> (Abstract: TODO)
#
# **Notebook Status:** <font color='orange'><b> Self-Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). **Additional validation tests may have been performed, but are as yet, undocumented. (TODO)**
#
# ### NRPy+ Source Code for this module: [ADM_in_terms_of_BSSN.py](../edit/BSSN/ADM_in_terms_of_BSSN.py)
#
# ## Introduction:
# This tutorial notebook constructs all quantities in the [ADM formalism](https://en.wikipedia.org/wiki/ADM_formalism) (see also Chapter 2 in Baumgarte & Shapiro's book *Numerical Relativity*) in terms of quantities in our adopted (covariant, tensor-rescaled) BSSN formalism. That is to say, we will write the ADM quantities $\left\{\gamma_{ij},K_{ij},\alpha,\beta^i\right\}$ and their derivatives in terms of the BSSN quantities $\left\{\bar{\gamma}_{ij},\text{cf},\bar{A}_{ij},\text{tr}K,\alpha,\beta^i\right\}$ and their derivatives.
#
# ### A Note on Notation:
#
# As is standard in NRPy+,
#
# * Greek indices refer to four-dimensional quantities where the zeroth component indicates temporal (time) component.
# * Latin indices refer to three-dimensional quantities. This is somewhat counterintuitive since Python always indexes its lists starting from 0. As a result, the zeroth component of three-dimensional quantities will necessarily indicate the first *spatial* direction.
#
# As a corollary, any expressions in NRPy+ involving mixed Greek and Latin indices will need to offset one set of indices by one; a Latin index in a four-vector will be incremented and a Greek index in a three-vector will be decremented (however, the latter case does not occur in this tutorial notebook).
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Initialize core Python/NRPy+ modules
# 1. [Step 2](#threemetric): The ADM three-metric $\gamma_{ij}$ and its derivatives in terms of rescaled BSSN quantities
# 1. [Step 2.a](#derivatives_e4phi): Derivatives of $e^{4\phi}$
# 1. [Step 2.b](#derivatives_adm_3metric): Derivatives of the ADM three-metric: $\gamma_{ij,k}$ and $\gamma_{ij,kl}$
# 1. [Step 2.c](#christoffel): Christoffel symbols $\Gamma^i_{jk}$ associated with the ADM 3-metric $\gamma_{ij}$
# 1. [Step 3](#extrinsiccurvature): The ADM extrinsic curvature $K_{ij}$ and its derivatives in terms of rescaled BSSN quantities
# 1. [Step 4](#code_validation): Code Validation against `BSSN.ADM_in_terms_of_BSSN` NRPy+ module
# 1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize core Python/NRPy+ modules \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# Let's start by importing all the needed modules from Python/NRPy+:
# +
# Step 1.a: Import all needed modules from NRPy+
import NRPy_param_funcs as par # NRPy+: parameter interface
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import sys # Standard Python module for multiplatform OS-level functions
# Step 1.b: Set the coordinate system for the numerical grid
par.set_parval_from_str("reference_metric::CoordSystem","Spherical")
# Step 1.c: Given the chosen coordinate system, set up
# corresponding reference metric and needed
# reference metric quantities
# The following function call sets up the reference metric
# and related quantities, including rescaling matrices ReDD,
# ReU, and hatted quantities.
rfm.reference_metric()
# Step 1.d: Set spatial dimension (must be 3 for BSSN, as BSSN is
# a 3+1-dimensional decomposition of the general
# relativistic field equations)
DIM = 3
# Step 1.e: Import all basic (unrescaled) BSSN scalars & tensors
import BSSN.BSSN_quantities as Bq
Bq.BSSN_basic_tensors()
gammabarDD = Bq.gammabarDD
cf = Bq.cf
AbarDD = Bq.AbarDD
trK = Bq.trK
Bq.gammabar__inverse_and_derivs()
gammabarDD_dD = Bq.gammabarDD_dD
gammabarDD_dDD = Bq.gammabarDD_dDD
Bq.AbarUU_AbarUD_trAbar_AbarDD_dD()
AbarDD_dD = Bq.AbarDD_dD
# -
# <a id='threemetric'></a>
#
# # Step 2: The ADM three-metric $\gamma_{ij}$ and its derivatives in terms of rescaled BSSN quantities. \[Back to [top](#toc)\]
# $$\label{threemetric}$$
#
# The ADM three-metric is written in terms of the covariant BSSN three-metric tensor as (Eqs. 2 and 3 of [Ruchlin *et al.*](https://arxiv.org/pdf/1712.07658.pdf)):
# $$
# \gamma_{ij} = \left(\frac{\gamma}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{i j},
# $$
# where $\gamma=\det{\gamma_{ij}}$ and $\bar{\gamma}=\det{\bar{\gamma}_{ij}}$.
#
# The "standard" BSSN conformal factor $\phi$ is given by (Eq. 3 of [Ruchlin *et al.*](https://arxiv.org/pdf/1712.07658.pdf)):
#
# \begin{align}
# \phi &= \frac{1}{12} \log\left(\frac{\gamma}{\bar{\gamma}}\right) \\
# \implies e^{\phi} &= \left(\frac{\gamma}{\bar{\gamma}}\right)^{1/12} \\
# \implies e^{4 \phi} &= \left(\frac{\gamma}{\bar{\gamma}}\right)^{1/3}
# \end{align}
#
# Thus the ADM three-metric may be written in terms of the BSSN three-metric and conformal factor $\phi$ as
#
# $$
# \gamma_{ij} = e^{4 \phi} \bar{\gamma}_{i j}.
# $$
#
# NRPy+'s implementation of BSSN allows for $\phi$ and two other alternative conformal factors to be defined:
#
# \begin{align}
# \chi &= e^{-4\phi} \\
# W &= e^{-2\phi},
# \end{align}
#
# Thus if `"BSSN_quantities::EvolvedConformalFactor_cf"` is set to `"chi"`, then
#
# \begin{align}
# \gamma_{ij} &= \frac{1}{\chi} \bar{\gamma}_{i j} \\
# &= \frac{1}{\text{cf}} \bar{\gamma}_{i j},
# \end{align}
#
# and if `"BSSN_quantities::EvolvedConformalFactor_cf"` is set to `"W"`, then
# \begin{align}
# \gamma_{ij} &= \frac{1}{W^2} \bar{\gamma}_{i j} \\
# &= \frac{1}{\text{cf}^2} \bar{\gamma}_{i j}.
# \end{align}
# +
# Step 2: The ADM three-metric gammaDD and its
# derivatives in terms of BSSN quantities.
gammaDD = ixp.zerorank2()
exp4phi = sp.sympify(0)
if par.parval_from_str("EvolvedConformalFactor_cf") == "phi":
exp4phi = sp.exp(4*cf)
elif par.parval_from_str("EvolvedConformalFactor_cf") == "chi":
exp4phi = (1 / cf)
elif par.parval_from_str("EvolvedConformalFactor_cf") == "W":
exp4phi = (1 / cf**2)
else:
print("Error EvolvedConformalFactor_cf type = \""+par.parval_from_str("EvolvedConformalFactor_cf")+"\" unknown.")
sys.exit(1)
for i in range(DIM):
for j in range(DIM):
gammaDD[i][j] = exp4phi*gammabarDD[i][j]
# -
# <a id='derivatives_e4phi'></a>
#
# ## Step 2.a: Derivatives of $e^{4\phi}$ \[Back to [top](#toc)\]
# $$\label{derivatives_e4phi}$$
#
# To compute derivatives of $\gamma_{ij}$ in terms of BSSN variables and their derivatives, we will first need derivatives of $e^{4\phi}$ in terms of the conformal BSSN variable `cf`.
#
# \begin{align}
# \frac{\partial}{\partial x^i} e^{4\phi} &= 4 e^{4\phi} \phi_{,i} \\
# \implies \frac{\partial}{\partial x^j} \frac{\partial}{\partial x^i} e^{4\phi} &= \frac{\partial}{\partial x^j} \left(4 e^{4\phi} \phi_{,i}\right) \\
# &= 16 e^{4\phi} \phi_{,i} \phi_{,j} + 4 e^{4\phi} \phi_{,ij}
# \end{align}
#
# Thus computing first and second derivatives of $e^{4\phi}$ in terms of the BSSN quantity `cf` requires only that we evaluate $\phi_{,i}$ and $\phi_{,ij}$ in terms of $e^{4\phi}$ (computed above in terms of `cf`) and derivatives of `cf`:
#
# If `"BSSN_quantities::EvolvedConformalFactor_cf"` is set to `"phi"`, then
# \begin{align}
# \phi_{,i} &= \text{cf}_{,i} \\
# \phi_{,ij} &= \text{cf}_{,ij}
# \end{align}
#
# If `"BSSN_quantities::EvolvedConformalFactor_cf"` is set to `"chi"`, then
# \begin{align}
# \text{cf} = e^{-4\phi} \implies \text{cf}_{,i} &= -4 e^{-4\phi} \phi_{,i} \\
# \implies \phi_{,i} &= -\frac{e^{4\phi}}{4} \text{cf}_{,i} \\
# \implies \phi_{,ij} &= -e^{4\phi} \phi_{,j} \text{cf}_{,i} -\frac{e^{4\phi}}{4} \text{cf}_{,ij}\\
# &= -e^{4\phi} \left(-\frac{e^{4\phi}}{4} \text{cf}_{,j}\right) \text{cf}_{,i} -\frac{e^{4\phi}}{4} \text{cf}_{,ij} \\
# &= \frac{1}{4} \left[\left(e^{4\phi}\right)^2 \text{cf}_{,i} \text{cf}_{,j} -e^{4\phi} \text{cf}_{,ij}\right] \\
# \end{align}
#
# If `"BSSN_quantities::EvolvedConformalFactor_cf"` is set to `"W"`, then
# \begin{align}
# \text{cf} = e^{-2\phi} \implies \text{cf}_{,i} &= -2 e^{-2\phi} \phi_{,i} \\
# \implies \phi_{,i} &= -\frac{e^{2\phi}}{2} \text{cf}_{,i} \\
# \implies \phi_{,ij} &= -e^{2\phi} \phi_{,j} \text{cf}_{,i} -\frac{e^{2\phi}}{2} \text{cf}_{,ij}\\
# &= -e^{2\phi} \left(-\frac{e^{2\phi}}{2} \text{cf}_{,j}\right) \text{cf}_{,i} -\frac{e^{2\phi}}{2} \text{cf}_{,ij} \\
# &= \frac{1}{2} \left[e^{4\phi} \text{cf}_{,i} \text{cf}_{,j} -e^{2\phi} \text{cf}_{,ij}\right] \\
# \end{align}
# +
# Step 2.a: Derivatives of $e^{4\phi}$
phidD = ixp.zerorank1()
phidDD = ixp.zerorank2()
cf_dD = ixp.declarerank1("cf_dD")
cf_dDD = ixp.declarerank2("cf_dDD","sym01")
if par.parval_from_str("EvolvedConformalFactor_cf") == "phi":
for i in range(DIM):
phidD[i] = cf_dD[i]
for j in range(DIM):
phidDD[i][j] = cf_dDD[i][j]
elif par.parval_from_str("EvolvedConformalFactor_cf") == "chi":
for i in range(DIM):
phidD[i] = -sp.Rational(1,4)*exp4phi*cf_dD[i]
for j in range(DIM):
phidDD[i][j] = sp.Rational(1,4)*( exp4phi**2*cf_dD[i]*cf_dD[j] - exp4phi*cf_dDD[i][j] )
elif par.parval_from_str("EvolvedConformalFactor_cf") == "W":
exp2phi = (1 / cf)
for i in range(DIM):
phidD[i] = -sp.Rational(1,2)*exp2phi*cf_dD[i]
for j in range(DIM):
phidDD[i][j] = sp.Rational(1,2)*( exp4phi*cf_dD[i]*cf_dD[j] - exp2phi*cf_dDD[i][j] )
else:
print("Error EvolvedConformalFactor_cf type = \""+par.parval_from_str("EvolvedConformalFactor_cf")+"\" unknown.")
sys.exit(1)
exp4phidD = ixp.zerorank1()
exp4phidDD = ixp.zerorank2()
for i in range(DIM):
exp4phidD[i] = 4*exp4phi*phidD[i]
for j in range(DIM):
exp4phidDD[i][j] = 16*exp4phi*phidD[i]*phidD[j] + 4*exp4phi*phidDD[i][j]
# -
# <a id='derivatives_adm_3metric'></a>
#
# ## Step 2.b: Derivatives of the ADM three-metric: $\gamma_{ij,k}$ and $\gamma_{ij,kl}$ \[Back to [top](#toc)\]
# $$\label{derivatives_adm_3metric}$$
#
# Recall the relation between the ADM three-metric $\gamma_{ij}$, the BSSN conformal three-metric $\bar{\gamma}_{i j}$, and the BSSN conformal factor $\phi$:
#
# $$
# \gamma_{ij} = e^{4 \phi} \bar{\gamma}_{i j}.
# $$
#
# Now that we have constructed derivatives of $e^{4 \phi}$ in terms of the chosen BSSN conformal factor `cf`, and the [BSSN.BSSN_quantities module](../edit/BSSN/BSSN_quantities.py) ([**tutorial**](Tutorial-BSSN_quantities.ipynb)) defines derivatives of $\bar{\gamma}_{ij}$ in terms of rescaled BSSN variables, derivatives of $\gamma_{ij}$ can be immediately constructed using the product rule:
#
# \begin{align}
# \gamma_{ij,k} &= \left(e^{4 \phi}\right)_{,k} \bar{\gamma}_{i j} + e^{4 \phi} \bar{\gamma}_{ij,k} \\
# \gamma_{ij,kl} &= \left(e^{4 \phi}\right)_{,kl} \bar{\gamma}_{i j} + \left(e^{4 \phi}\right)_{,k} \bar{\gamma}_{i j,l} + \left(e^{4 \phi}\right)_{,l} \bar{\gamma}_{ij,k} + e^{4 \phi} \bar{\gamma}_{ij,kl}
# \end{align}
# +
# Step 2.b: Derivatives of gammaDD, the ADM three-metric
gammaDDdD = ixp.zerorank3()
gammaDDdDD = ixp.zerorank4()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
gammaDDdD[i][j][k] = exp4phidD[k]*gammabarDD[i][j] + exp4phi*gammabarDD_dD[i][j][k]
for l in range(DIM):
gammaDDdDD[i][j][k][l] = exp4phidDD[k][l]*gammabarDD[i][j] + \
exp4phidD[k]*gammabarDD_dD[i][j][l] + \
exp4phidD[l]*gammabarDD_dD[i][j][k] + \
exp4phi*gammabarDD_dDD[i][j][k][l]
# -
# <a id='christoffel'></a>
#
# ## Step 2.c: Christoffel symbols $\Gamma^i_{jk}$ associated with the ADM 3-metric $\gamma_{ij}$ \[Back to [top](#toc)\]
# $$\label{christoffel}$$
#
# The 3-metric analog to the definition of Christoffel symbol (Eq. 1.18) in Baumgarte & Shapiro's *Numerical Relativity* is given by
# $$
# \Gamma^i_{jk} = \frac{1}{2} \gamma^{il} \left(\gamma_{lj,k} + \gamma_{lk,j} - \gamma_{jk,l} \right),
# $$
# which we implement here:
# +
# Step 2.c: 3-Christoffel symbols associated with ADM 3-metric gammaDD
# Step 2.c.i: First compute the inverse 3-metric gammaUU:
gammaUU, detgamma = ixp.symm_matrix_inverter3x3(gammaDD)
GammaUDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
for l in range(DIM):
GammaUDD[i][j][k] += sp.Rational(1,2)*gammaUU[i][l]* \
(gammaDDdD[l][j][k] + gammaDDdD[l][k][j] - gammaDDdD[j][k][l])
# -
# <a id='extrinsiccurvature'></a>
#
# # Step 3: The ADM extrinsic curvature $K_{ij}$ and its derivatives in terms of rescaled BSSN quantities. \[Back to [top](#toc)\]
# $$\label{extrinsiccurvature}$$
#
# The ADM extrinsic curvature may be written in terms of the BSSN trace-free extrinsic curvature tensor $\bar{A}_{ij}$ and the trace of the ADM extrinsic curvature $K$:
#
# \begin{align}
# K_{ij} &= \left(\frac{\gamma}{\bar{\gamma}}\right)^{1/3} \bar{A}_{ij} + \frac{1}{3} \gamma_{ij} K \\
# &= e^{4\phi} \bar{A}_{ij} + \frac{1}{3} \gamma_{ij} K \\
# \end{align}
#
# We only compute first spatial derivatives of $K_{ij}$, as higher-derivatives are generally not needed:
# $$
# K_{ij,k} = \left(e^{4\phi}\right)_{,k} \bar{A}_{ij} + e^{4\phi} \bar{A}_{ij,k} + \frac{1}{3} \left(\gamma_{ij,k} K + \gamma_{ij} K_{,k}\right)
# $$
# which is expressed in terms of quantities already defined.
# +
# Step 3: Define ADM extrinsic curvature KDD and
# its first spatial derivatives KDDdD
# in terms of BSSN quantities
KDD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
KDD[i][j] = exp4phi*AbarDD[i][j] + sp.Rational(1,3)*gammaDD[i][j]*trK
KDDdD = ixp.zerorank3()
trK_dD = ixp.declarerank1("trK_dD")
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
KDDdD[i][j][k] = exp4phidD[k]*AbarDD[i][j] + exp4phi*AbarDD_dD[i][j][k] + \
sp.Rational(1,3)*(gammaDDdD[i][j][k]*trK + gammaDD[i][j]*trK_dD[k])
# -
# <a id='code_validation'></a>
#
# # Step 4: Code Validation against `BSSN.ADM_in_terms_of_BSSN` NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions between
#
# 1. this tutorial and
# 2. the NRPy+ [BSSN.ADM_in_terms_of_BSSN](../edit/BSSN/ADM_in_terms_of_BSSN.py) module.
#
# +
all_passed=True
def comp_func(expr1,expr2,basename,prefixname2="Bq."):
if str(expr1-expr2)!="0":
print(basename+" - "+prefixname2+basename+" = "+ str(expr1-expr2))
all_passed=False
def gfnm(basename,idx1,idx2=None,idx3=None,idx4=None):
if idx2 is None:
return basename+"["+str(idx1)+"]"
if idx3 is None:
return basename+"["+str(idx1)+"]["+str(idx2)+"]"
if idx4 is None:
return basename+"["+str(idx1)+"]["+str(idx2)+"]["+str(idx3)+"]"
return basename+"["+str(idx1)+"]["+str(idx2)+"]["+str(idx3)+"]["+str(idx4)+"]"
expr_list = []
exprcheck_list = []
namecheck_list = []
import BSSN.ADM_in_terms_of_BSSN as AB
AB.ADM_in_terms_of_BSSN()
namecheck_list.extend(["detgamma"])
exprcheck_list.extend([AB.detgamma])
expr_list.extend([detgamma])
for i in range(DIM):
for j in range(DIM):
namecheck_list.extend([gfnm("gammaDD",i,j),gfnm("gammaUU",i,j),gfnm("KDD",i,j)])
exprcheck_list.extend([AB.gammaDD[i][j],AB.gammaUU[i][j],AB.KDD[i][j]])
expr_list.extend([gammaDD[i][j],gammaUU[i][j],KDD[i][j]])
for k in range(DIM):
namecheck_list.extend([gfnm("gammaDDdD",i,j,k),gfnm("GammaUDD",i,j,k),gfnm("KDDdD",i,j,k)])
exprcheck_list.extend([AB.gammaDDdD[i][j][k],AB.GammaUDD[i][j][k],AB.KDDdD[i][j][k]])
expr_list.extend([gammaDDdD[i][j][k],GammaUDD[i][j][k],KDDdD[i][j][k]])
for l in range(DIM):
namecheck_list.extend([gfnm("gammaDDdDD",i,j,k,l)])
exprcheck_list.extend([AB.gammaDDdDD[i][j][k][l]])
expr_list.extend([gammaDDdDD[i][j][k][l]])
for i in range(len(expr_list)):
comp_func(expr_list[i],exprcheck_list[i],namecheck_list[i])
if all_passed:
print("ALL TESTS PASSED!")
else:
print("ERROR. ONE OR MORE TESTS FAILED")
sys.exit(1)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 5: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ADM_in_terms_of_BSSN.pdf](Tutorial-ADM_in_terms_of_BSSN.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ADM_in_terms_of_BSSN")
| Tutorial-ADM_in_terms_of_BSSN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# For more information about Monte Carlo Tree Search check out our web site at www.mcts.ai
# http://mcts.ai/about/index.html
from math import *
import random
# -
def upper_confidence_bounds(node_value, num_parent_visits, num_node_visits):
""" the UCB1 formula """
return node_value + sqrt(2 * log(num_parent_visits) / num_node_visits)
class OXOEnv:
""" A state of the game, i.e. the game board.
Squares in the board are in this arrangement
012
345
678
where 0 = empty, 1 = player 1 (X), 2 = player 2 (O)
"""
def __init__(self):
self.current_player = 1 # At the root pretend the current player 'Player 1'
self.board = [0, 0, 0, 0, 0, 0, 0, 0, 0] # 0 = empty, 1 = player 1, 2 = player 2
def clone(self):
""" Create a deep clone of this game state.
"""
env = OXOEnv()
env.current_player = self.current_player
env.board = self.board[:]
return env
def do_move(self, location):
""" Update a state by carrying out the given move.
Must update playerToMove.
"""
assert 8 >= location >= 0 == self.board[location] and location == int(location)
self.board[location] = self.current_player
if self.current_player == 1:
self.current_player = 2
elif self.current_player == 2:
self.current_player = 1
else:
assert False
def get_possible_locations(self):
""" Get all possible moves from this state.
"""
return [i for i in range(9) if self.board[i] == 0]
def get_result(self, player_just_moved):
""" Get the game result from the viewpoint of playerjm.
Case == 1.0: player 1 win,
Case == 2.0: player 2 win,
Case == 3.0: both player 1 and 2 win (draw)
"""
case = 0.0
for (x, y, z) in [(0, 1, 2), (3, 4, 5), (6, 7, 8), (0, 3, 6), (1, 4, 7), (2, 5, 8), (0, 4, 8), (2, 4, 6)]:
if self.board[x] == self.board[y] == self.board[z] == player_just_moved:
case += 1
elif self.board[x] == self.board[y] == self.board[z] == (3 - player_just_moved):
case += 2
if case == 1:
return 1 # player 1 win
elif case == 2:
return 2 # player 2 win
elif case == 3 or not self.get_possible_locations():
print("draw - ", case)
return 0 # draw
else:
return -1.0 # continue
assert False # Should not be possible to get here
def __repr__(self):
s = ""
for i in range(9):
s += ".XO"[self.board[i]]
if i % 3 == 2:
s += "\n"
return s[0:-1]
class Node:
""" A node in the game tree.
Note wins is always from the viewpoint of current_player.
Crashes if state not specified.
"""
def __init__(self, location=None, parent=None, env=None):
self.location = location # "None" for the root node
self.parent_node = parent # "None" for the root node
self.child_nodes = []
self.wins = 0
self.visits = 0
self.unvisited_locations = env.get_possible_locations() # future child nodes
def uct_select_child(self):
""" Use the UCB1 formula to select a child node.
Often a constant UCTK is applied so we have UCB1 to vary the amount of exploration versus exploitation.
"""
s = sorted(
self.child_nodes,
key=lambda c: upper_confidence_bounds(c.wins / c.visits, self.visits, c.visits)
)[-1]
return s
def add_child(self, loc, env):
""" Remove loc from untried_locations and add a new child node for the location loc.
Return the added child node
"""
n = Node(location=loc, parent=self, env=env)
self.unvisited_locations.remove(loc)
self.child_nodes.append(n)
return n
def update(self, result):
""" Update this node - one additional visit and result additional wins.
result must be from the viewpoint of current_player.
"""
self.visits += 1
self.wins += result
def to_tree_string(self, indent):
s = self.indent_string(indent) + str(self)
for c in self.child_nodes:
s += c.to_tree_string(indent + 1)
return s
@staticmethod
def indent_string(indent):
s = "\n"
for i in range(1, indent + 1):
s += "| "
return s
def to_children_string(self):
s = ""
for c in self.child_nodes:
s += str(c) + "\n"
return s
def __repr__(self):
return "[Location: {0}, W/V: {1}/{2}, U: {3}, Child Nodes: {4}]".format(
self.location,
self.wins,
self.visits,
str(self.unvisited_locations),
str([x.location for x in self.child_nodes])
)
def search_by_uct(env, iter_max, verbose=False):
""" Conduct a UCT (Upper Confidence Bounds for Trees) search for itermax iterations starting from rootstate.
Return the best move from the root_node.
Assumes 2 alternating players (player 1 starts), with game results in the range [1, 2, 0, -1].
"""
root_node = Node(location=None, parent=None, env=env)
print("[Search By UCT]")
for i in range(iter_max):
node = root_node
env2 = env.clone()
# Select
while node.unvisited_locations == [] and node.child_nodes != []: # node is fully expanded and non-terminal
node = node.uct_select_child()
env2.do_move(node.location)
print("Iter: {0}, Player {1} selects the best child node {2}".format(
i,
env2.current_player,
node.location
))
# Expand
if node.unvisited_locations: # if we can expand (i.e. state/node is non-terminal)
m = random.choice(node.unvisited_locations)
env2.do_move(m)
print("Iter: {0}, Player {1} expands to an arbitrary location {2}".format(
i,
env2.current_player,
m
))
node = node.add_child(m, env2) # add child and descend tree
# Rollout - this can often be made orders of magnitude quicker using a state.GetRandomMove() function
j = 0
while env2.get_possible_locations(): # while state is non-terminal
m = random.choice(env2.get_possible_locations())
env2.do_move(m)
print("Iter: {0} and {1}, Player {2} rolls out to the location {3}".format(
i,
j,
env2.current_player,
m
))
j += 1
# Backpropagate
j = 0
print("{0} - Cloned Env:\n{1}".format(3 - env.current_player, env2))
while node: # backpropagate from the expanded node and work back to the root node
node.update(env2.get_result(3 - env.current_player)) # state is terminal. Update node with result from
# point of view of 3 - env.current_player
print("Iter: {0}, {1}, Evaluate the node {2}: Wins/Visits - {3}/{4}".format(
i,
j,
node.location,
node.wins,
node.visits
))
node = node.parent_node
j += 1
print()
# Output some information about the tree - can be omitted
if verbose:
print(root_node.to_tree_string(0))
else:
print(root_node.to_children_string())
return sorted(root_node.child_nodes, key=lambda c: c.visits)[-1].location # return the move that was most visited
def play_game(verbose=True):
""" Play a sample game between two UCT players where each player gets a different number
of UCT iterations (= simulations = tree nodes).
"""
# state = OthelloState(4) # uncomment to play Othello on a square board of the given size
env = OXOEnv() # uncomment to play OXO
# state = NimState(15) # uncomment to play Nim with the given number of starting chips
while env.get_possible_locations():
print("Original Env:\n{0}".format(env))
if env.current_player == 1:
m = search_by_uct(env=env, iter_max=2, verbose=verbose) # Player 1
else:
m = search_by_uct(env=env, iter_max=2, verbose=verbose) # Player 2
print("Best Move: " + str(m) + "\n")
env.do_move(m)
print("Original Env:\n{0}".format(env))
if env.get_result(env.current_player) == 1:
print("Player " + str(env.current_player) + " wins!")
break
elif env.get_result(env.current_player) == 2:
print("Player " + str(3 - env.current_player) + " wins!")
break
elif env.get_result(env.current_player) == 0:
print("Nobody wins!")
break
elif env.get_result(env.current_player) == -1:
print("Continue...\n")
else:
assert False
if __name__ == "__main__":
""" Play a single game to the end using UCT for both players.
"""
play_game()
| 2.ReinforcementLearning/MCTS/mcts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mushroom classification using catboost
# +
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# +
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
# %matplotlib inline
# -
df = pd.read_csv("dataset/mushrooms.csv")
df.shape
df.head(5)
df.columns
df.info()
df.nunique()
# cast all columns to categorical
df = df.apply(lambda col: col.astype("category"))
df.info()
# +
from sklearn.model_selection import train_test_split
X = df.iloc[:, 1:]
y = df.iloc[:, 0]
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, stratify=y, random_state=42
)
X_train.shape, X_test.shape
# -
# Exploratory analysis on the data
df_train = pd.concat((X_train, y_train), axis=1)
df_train.columns
df_train.head(1)
figure, axes = plt.subplots(nrows=8, ncols=3, figsize=(20, 30))
for col_no in range(len(df_train.columns) - 1):
sns.countplot(
x=df_train.columns[col_no], hue="class", data=df_train, ax=axes[col_no // 3][col_no % 3]
);
# data distribution by class
df_train.groupby(by="class")["cap-shape"].count()
# +
from sklearn.preprocessing import LabelEncoder
# From the column description, https://www.kaggle.com/uciml/mushroom-classification
# all features seem to be nominal features.
feature_cols = df_train.columns[:-1]
target_col = df_train.columns[-1]
target_encoder = LabelEncoder()
y_train_trans = target_encoder.fit_transform(y_train)
X_train.shape, y_train_trans.shape
# -
# ## Exploring catboost
# +
from catboost import CatBoostClassifier
from sklearn.model_selection import RepeatedStratifiedKFold, cross_val_score
catboost_clf = CatBoostClassifier(iterations=10, cat_features=feature_cols)
# cross validation
# catboost can handle categorical variables and no preprocessing is required
cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=42)
scores = cross_val_score(catboost_clf, X_train, y_train_trans, scoring="accuracy", cv=cv, n_jobs=-1)
scores.mean(), scores.std()
# +
from sklearn.model_selection import RandomizedSearchCV
catboost_params = {
"iterations": range(10, 100, 5),
"learning_rate": [0.01, 0.1, 1],
"depth": [2, 5, 7, 9],
}
rnd_cv = RandomizedSearchCV(catboost_clf, catboost_params, n_iter=100, n_jobs=-1, cv=3, verbose=2)
rnd_cv.fit(X_train, y_train_trans)
rnd_cv.best_score_, rnd_cv.best_params_
# -
# training catboost using GPU
# best params after hyperparameter tuning
best_params = {"iterations": 65, "learning_rate": 0.1, "depth": 7}
catboost_clf_gpu = CatBoostClassifier(**best_params, task_type="GPU", cat_features=feature_cols)
catboost_clf_gpu.fit(X_train, y_train_trans)
y_hat = catboost_clf_gpu.predict(X_test)
# +
# evaluate the model on the test set
from sklearn.metrics import (
accuracy_score,
confusion_matrix,
f1_score,
precision_score,
recall_score,
roc_auc_score,
)
y_test_trans = target_encoder.transform(y_test)
confusion_matrix(y_test_trans, y_hat)
accuracy_score(y_test_trans, y_hat)
precision_score(y_test_trans, y_hat)
recall_score(y_test_trans, y_hat)
f1_score(y_test_trans, y_hat)
roc_auc_score(y_test_trans, y_hat)
| mushroom_classification/mushroom_classification_catboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Import most generic modules
import importlib
import pathlib
import os
import sys
from datetime import datetime, timedelta
import pandas as pd
from IPython.display import display, Markdown
import warnings
warnings.filterwarnings("ignore")
module_path = os.path.abspath(os.path.join("../.."))
if module_path not in sys.path:
sys.path.append(module_path)
# + tags=["parameters"]
# Parameters that will be replaced when calling this notebook
ticker = "TSLA"
report_name = f"{datetime.now().strftime('%Y%m%d_%H%M%S')}_comparison_{ticker.upper()}"
# -
display(
Markdown(
f"# {ticker.upper()} - Comparison - {datetime.now().strftime('%Y/%m/%d %H:%M:%S')}"
)
)
# +
from gamestonk_terminal.stocks.comparison_analysis import (
finviz_compare_model,
yahoo_finance_model,
)
ticker = ticker.upper()
finviz_similar = finviz_compare_model.get_similar_companies(
ticker, ["Sector", "Industry", "Country"]
)[0]
# -
# ### Similar companies and descriptions
# +
from gamestonk_terminal.stocks.fundamental_analysis import yahoo_finance_model
df_info = yahoo_finance_model.get_info(ticker)
if "Long business summary" in df_info.index:
summary = df_info.loc["Long business summary"].values[0]
if "Website" in df_info.index:
website = df_info.loc["Website"].values[0]
if finviz_similar:
print(f"{ticker}: {website}")
print(summary)
for symbol in finviz_similar:
df_info = yahoo_finance_model.get_info(symbol)
if "Long business summary" in df_info.index:
summary = df_info.loc["Long business summary"].values[0]
if "Website" in df_info.index:
website = df_info.loc["Website"].values[0]
print("")
print(f"{symbol}: {website}")
print(summary)
# + [markdown] tags=[]
# ### Historical prices
# +
import math
from gamestonk_terminal.stocks.comparison_analysis import yahoo_finance_view
if finviz_similar and finviz_similar != [""]:
for i in range(math.ceil(len(finviz_similar) / 4)):
yahoo_finance_view.display_historical(
similar_tickers=finviz_similar[4 * (i) : 4 * (i + 1)],
)
else:
print("Ticker not found in CoinGeckoAPI")
# -
# ### Historical correlation
# +
from matplotlib import pyplot as plt
if finviz_similar and finviz_similar != [""]:
plt.figure(figsize=(25, 10))
yahoo_finance_view.display_correlation(
similar_tickers=finviz_similar,
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Historical volumes
# -
if finviz_similar and finviz_similar != [""]:
for i in range(math.ceil(len(finviz_similar) / 4)):
yahoo_finance_view.display_volume(
similar_tickers=finviz_similar[4 * (i) : 4 * (i + 1)],
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Overview
# +
from gamestonk_terminal.stocks.comparison_analysis import finviz_compare_view
if finviz_similar and finviz_similar != [""]:
finviz_compare_view.screener(
similar=finviz_similar,
data_type="overview",
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Valuation
# +
from gamestonk_terminal.stocks.comparison_analysis import finviz_compare_view
if finviz_similar and finviz_similar != [""]:
finviz_compare_view.screener(
similar=finviz_similar,
data_type="valuation",
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Financial
# +
from gamestonk_terminal.stocks.comparison_analysis import finviz_compare_view
if finviz_similar and finviz_similar != [""]:
finviz_compare_view.screener(
similar=finviz_similar,
data_type="financial",
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Ownership
# +
from gamestonk_terminal.stocks.comparison_analysis import finviz_compare_view
if finviz_similar and finviz_similar != [""]:
finviz_compare_view.screener(
similar=finviz_similar,
data_type="ownership",
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Performance
# +
from gamestonk_terminal.stocks.comparison_analysis import finviz_compare_view
if finviz_similar and finviz_similar != [""]:
finviz_compare_view.screener(
similar=finviz_similar,
data_type="performance",
)
else:
print("Ticker not found in CoinGeckoAPI")
# + [markdown] tags=[]
# ### Technical
# +
from gamestonk_terminal.stocks.comparison_analysis import finviz_compare_view
if finviz_similar and finviz_similar != [""]:
finviz_compare_view.screener(
similar=finviz_similar,
data_type="technical",
)
else:
print("Ticker not found in CoinGeckoAPI")
# -
# !jupyter nbconvert {report_name + ".ipynb"} --to html --no-input
| gamestonk_terminal/jupyter/reports/similar_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !jupyter nbconvert --to html DnnMath.ipynb
# # AI 융합교육 (수학교육용 데이터 수집 + 선처리 + AI 코딩)
#
#
# <p>
#
#
# # 스마트 수학교육 특강과 이산수학 공통 학습자료
# # =====================================
#
# <p>
# <font color=blue>
#
# # +++++++++++++++++++++++++++++++++++++
#
# </font>
# <p>
#
# # ++++++++++++++++++++++++++++++++++++++
#
# # Data 기반 딥러닝 분석 및 사용 (1) (2) (3) (4) (5) 실습
#
# ### (1) XOR 뉴럴네트워크와 인공지능
#
# ### (2) CNN 신경망과 Mnist 데이터
#
# ### (3) 자연어 처리와 chatbot 데이터
#
# ### (4) RNN 신경망과 RNN 실험 (Mnist)
#
# ### (5) RNN+CNN 텍스트 분석 (imdb)
#
# # ++++++++++++++++++++++++++++++
#
# <p>
#
#
# 동영상 자료 :
#
# https://www.youtube.com/playlist?list=PLZbbT5o_s2xq7LwI2y8_QtvuXZedL6tQU
#
#
# https://end-to-end-machine-learning.teachable.com/courses/how-deep-neural-networks-work/lectures/9485279
# # ==================================
#
# <p>
#
# # Deep Neural Network 역사와 행렬 합성함수
#
# # (1) XOR 뉴럴네트워크와 인공지능
#
# <p>
#
#
# ## XOR 문제 : 1969년 패펄트와 민스키
#
# ### 이 문제가 일으킨 인공지능 겨울과 거북이 LOGO 언어
#
#
#
# <p>
# 
#
# <p>
#
#
# 
#
# <p>
#
#
# 
#
# <p>
#
# 
#
# <p>
#
# 
# +
print( 3+2)
# 2와 3을 더하는 명령
# -
# # ++++++++++++++++++++++++++++++++++
#
#
# <p>
#
# # numpy 행렬 만들기와 텐서플로우 그래프와 변수
#
# <p>
#
# ### 참고자료 : https://medium.com/ai-india/hello-world-tensorflow-6ce3f5bcbb6b
#
# <p> 텐서플로우는 그래프 위에 변수를 연결하고 나중에 sess.run 시킨다
#
#
#
# <p>
#
#
# ### placeholder
#
# 생성될 때 값을 가지지 않고, 자리(place)를 유지(hold)하는 개념입니다. 함수 f(x)의 x
#
# placeholder(
# dtype,
# shape=None,
# name=None
# )
#
# dtype : 데이터 타입을 의미하며 반드시 적어주어야 한다.
#
# shape : 입력 데이터의 형태를 의미한다. 상수 값이 될 수도 있고 다차원 배열의 정보가 들어올 수도 있다.
# ( 디폴트 파라미터로 None 지정 )
#
# name : 해당 placeholder의 이름을 부여하는 것으로 적지 않아도 된다. ( 디폴트 파라미터로 None 지정 )
#
# ### variable
#
# 생성될때 값을 갖는 상수와 같은 것 (일차함수 ax+b 의 기울기 및 절편과 같은 것)
#
# ### constant
#
# 콘스탄트 (잔짜 상수)
a=2
b=3
c=a+b
print(c)
# # Part 1: building the GRAPH,
# ### it represents the data flow of the computations
#
# <p>
#
# # Part 2: running a SESSION,
# ### it executes the operations in the graph
#
# <p>
# <p>
#
# 
#
# <p>
#
#
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
x=2
y=3
add_op = tf.add(x, y)
print( add_op )
# 주석을 달아주세요
print('텐서플로우 버젼 : ' , tf.__version__)
mul_op = tf.multiply(x, y)
pow_op = tf.pow( add_op, mul_op)
useless_op = tf.multiply(x, add_op)
with tf.Session() as sess:
pow_op, useless_op = sess.run([pow_op, useless_op])
print( 'pow_op 의 값 : ', pow_op)
print(useless_op)
# -
#
#
#
# # [탐구] 다음을 스토리텔링
#
# 
#
# <p>
#
# # +++++++++++++++
#
# <p>
# # 학교수학의 일차함수 및 일차변환
# +
import numpy as np
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
# X 와 Y 의 상관관계를 분석하는 기초적인 선형 회귀 모델을 만들고 실행해봅니다.
# y = w*x ===> w=1 찾기
x_data = [1, 2, 3]
y_data = [1, 2, 3]
# x의 값이 y 에 대응된다
# -
# # 일차함수 = weight 기울기=> y = x * W + b
# +
W = tf.Variable(tf.random_uniform([1], -1.0, 1.0))
b = tf.Variable(tf.random_uniform([1], -1.0, 1.0))
# name: 나중에 텐서보드등으로 값의 변화를 추적하거나 살펴보기 쉽게 하기 위해 이름을 붙여줍니다.
X = tf.placeholder(tf.float32, name="X")
Y = tf.placeholder(tf.float32, name="Y")
print("X = ", X)
print('Y = ', Y)
# X 와 Y 의 상관 관계를 분석하기 위한 가설 수식을 작성합니다.
# y = W * x + b
# W 와 X 가 행렬이 아니므로 tf.matmul 이 아니라 기본 곱셈 기호를 사용했습니다.
hypothesis = W * X + b
# 손실 함수를 작성합니다.
# mean(h - Y)^2 : 예측값과 실제값의 거리를 비용(손실) 함수로 정합니다.
cost = tf.reduce_mean(tf.square(hypothesis - Y))
# 텐서플로우에 기본적으로 포함되어 있는 함수를 이용해 경사 하강법 최적화를 수행합니다.
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.1)
# 비용을 최소화 하는 것이 최종 목표
train_op = optimizer.minimize(cost)
#########
# 세션을 생성하고 초기화합니다.
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
# 최적화를 100번 수행합니다.
for step in range(100):
# sess.run 을 통해 train_op 와 cost 그래프를 계산합니다.
# 이 때, 가설 수식에 넣어야 할 실제값을 feed_dict 을 통해 전달합니다.
_, cost_val = sess.run([train_op, cost], feed_dict={X: x_data, Y: y_data})
if step % 10 ==0 :
print(step, '=> ' , cost_val, ' :: y = ', sess.run(W), ' * x + ', sess.run(b))
# 최적화가 완료된 모델에 테스트 값을 넣고 결과가 잘 나오는지 확인해봅니다.
print("\n=== Test ===")
print("X: 5, Y:", sess.run(hypothesis, feed_dict={X: 5}))
print("X: 2.5, Y:", sess.run(hypothesis, feed_dict={X: 2.5}))
# -
#
# # ==============================
#
# <p>
#
#
# # XOR 문제와 인공지능의 겨울 그리고 봄
# # 일차변환 : 벡터를 벡터로 대응시키는 규칙
# # true AND true = true : true AND false = false
#
# # 중간에 hiden layer 들어간 example
# 
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
# 장의역 nput X vector
X = [[0, 0], [0, 1], [1, 0], [1, 1]]
# 치역 output Y vector
Y = [[0], [1], [1], [0]]
# 이 것에 대응하는 일차 아핀변환은 무엇 ??
# Placeholders (앞에서의 X,Y 를 input and output 들어가게 하는 곳)
x = tf.placeholder(tf.float32, shape=[4,2])
y = tf.placeholder(tf.float32, shape=[4,1])
# W matrix ( 이 행렬을 구하는 것이 목표 : 모델이라 부른다)
W1 = tf.Variable([[1.0, 0.0], [1.0, 0.0]], shape=[2,2])
W2 = tf.Variable([[0.0], [1.0]], shape=[2,1])
# Biases (아핀변환의 절편 : 행렬과 절편 즉 모델을 구함이 목표)
B1 = tf.Variable([0.0, 0.0], shape=[2] )
B2 = tf.Variable([0.0], shape=[1] )
# 앞의 example 에서는 하나의 아핀변환 (시작을 이렇게 랜덤하게)
# default range is [0, 1). 이 것을 -1 이상에서 1 미만 사이로
# W = tf.Variable(tf.random_uniform([1], -1.0, 1.0))
# b = tf.Variable(tf.random_uniform([1], -1.0, 1.0))
########################################################
# 아핀변환 2개 이상의 합성으로 된다는 것이 1969 XOR
# Hidden layer and outout layer : 일차변환(아핀변환) 합성
output =tf.sigmoid(tf.matmul(tf.sigmoid(tf.matmul(x, W1) + B1), W2) + B2)
# 시그모이드 함수는 실수 x 를 1/ 1+(e)^x 에 대응: 함수 0과 1 사이가 치역
# error estimation (tf.matmu : matrix multiplicaiton)
# tf.squared_difference, tf.reduced_mean 찾아보라 !!
cost = tf.reduce_mean( tf.squared_difference( y, output) )
train_op = tf.train.GradientDescentOptimizer(0.1).minimize( cost )
# 다른 옵티마이제이션 실습 (시험문제로 좋음 : 찾아서 스토리텔링)
#train_op = tf.train.AdamOptimizer(0.1).minimize( cost )
# 다른 optimizer 사용
# Adam’s optimizer was presented by <NAME> from OpenAI
# and <NAME> from the University of Toronto in their 2015
######### 변수를 computation graph 에 생성시키고 실행 #########
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
for i in range (50001):
error = sess.run( train_op , feed_dict={x: X, y: Y})
if i % 5000 == 0:
print('Epoch: ' + str(i))
print('Error: ' + str( sess.run( cost, feed_dict={x: X, y: Y})))
for answer in sess.run( output, feed_dict={x: X, y: Y}):
print(' ', answer)
print ("실행 끝 : 아직 sess 는 살아있음")
# -
# ### 러닝 rate 와 옵티마이져 구하기 ?
#
# https://medium.com/octavian-ai/which-optimizer-and-learning-rate-should-i-use-for-deep-learning-5acb418f9b2
#
#
# 
# # 위에서 adam optimization 더 좋다
# <p>
#
# # 컴퓨터가 찾아낸 행렬 규칙 2개를 보자
# +
# 우리의 목표는 행렬과 절편(bias) 을 구하는 것
print( '첫번째 행렬 \n', sess.run(W1), '\n', sess.run(B1))
print( '두번째 행렬 \n', sess.run(W2), '\n', sess.run(B2))
print( ' 아직 sess 살아있음 ')
"""
손실 함수(Loss function) 값 cost 값 -훈련 하는 동안 모델의 오차를 측정.
행렬 절편 모델의 학습이 올바른 방향으로 향하도록 이 함수를 최소화해야 한다.
옵티마이저(Optimizer)-데이터와 손실 함수를 바탕으로 모델의 업데이트 방법을 결정.
따라서 딥러닝에서는 optimizer 가 핵심적인 파라메터 .....
"""
# +
# reduce_mean 탐구
x = tf.constant([[1. , 3.], [2. , 6.]])
print(sess.run(x))
print( '\n' )
print(sess.run(tf.reduce_mean(x)))
print(sess.run(tf.reduce_mean(x, 0)))
print(sess.run(tf.reduce_mean(x, 1)))
"""
tf.reduce_mean(x)처럼 두번째 인자를 적지 않은 경우
변수 x가 가리키는 배열 전체 원소의 합을 원소 개수로 나누어 계산합니다.
( 1 + 3 + 2 + 6 ) / 4 = 3 으로 계산
tf.reduce_mean(x, 0)는 열 단위로 평균을 냅니다.
첫번째 열의 원소의 평균은 1.5이고 두번째 열의 원소의 평균은 4.5입니다.
그래서 계산 결과가 [1.5 4.5] 입니다.
tf.reduce_mean(x, 1) 은 ??
"""
# +
a = tf.constant(10)
b = tf.constant(32)
print(a)
print( a+ b )
print('=======')
print('a의 값 : ', sess.run(a))
print('a+b 의 값 : ', sess.run(a+b))
# +
sess.close()
# 여기서 sess 실행 상태를 종료
# -
# # [실습 탐구문제 ]
#
#
#
# <p>
#
# # 2차원에서 1차원 가는 상황 만들어 탐구
# ## (예) 고등학교 and , or 논리 연산
#
# ## y = x W + b
#
# ## x: 2차원 row vector,
#
# ## y는 1차원, W는 2x1 행렬, b 는 숫자
#
# <p>
#
# ### ++++++++++++++++++++++++++++++
#
#
# # Hidden layer 합성함수의 경우에는 좀더
# # backpropagation, optimization 알아야 함
# +
x_data = np.array([[0, 0],
[0, 1],
[1, 0],
[1, 1]], dtype=np.float32)
y_data = np.array([[0],
[0],
[0],
[1]], dtype=np.float32)
print( y_data.T.dot( x_data ))
print( tf.matmul( y_data.T , x_data ) )
print( y_data.shape )
print( y_data.T.shape , '===:===' , x_data.shape )
a = tf.constant(10)
b = tf.constant(32)
# print(sess.run(a+b))
print( a+ b )
# -
# [[1. 1.]]
# tf.Tensor([[1. 1.]], shape=(1, 2), dtype=float32)
# (4, 1)
# (1, 4) ===:=== (4, 2)
# tf.Tensor(42, shape=(), dtype=int32)
# ## Kernel 메뉴의 restart 하면 새로 시작
# +
# tf.disable_eager_execution()
############################
c = tf.constant(10)
d = tf.constant(32)
# print(sess.run(a+b))
print( c+d )
# -
# ## tf.disable_eager_execution() 하면 sess로 run
# +
#tf.disable_eager_execution()
###################################
# initialize the variables
# init = tf.global_variables_initializer()
# sess.run(init)
####################################
hello = tf.constant('Hello, TensorFlow!')
e = tf.constant(10)
f = tf.constant(32)
sess = tf.Session()
###################
print( sess.run(hello) )
print( hello )
print( sess.run( e+f))
print( e+f )
print( sess.run( tf.matmul( y_data.T , x_data ) ) )
print( tf.matmul( y_data.T , x_data ) )
# -
A = tf.constant(10)
B = tf.constant(32)
print(sess.run(A+B))
print( A+ B )
# ### 텐서플로우 그래프를 먼저 만들고 sess.run
# +
# 텐서플로우 2.0 을 1.* 버젼으로
# 텐서플로우 2.0 이상에는 없다는 뜻
X = tf.placeholder(tf.float32, [None, 2], name='x-input')
Y = tf.placeholder(tf.float32, [None, 1], name='y-input')
# -
# # 일차변환 (아핀) = weighted matrix => y = x* W + b
# +
W = tf.Variable(tf.random_normal([2, 1]), name='weight')
b = tf.Variable(tf.random_normal([1]), name='bias')
# Ws = tf.Variable(tf.random_normal([2, 1]), name='weight')
hypothesis = tf.sigmoid(tf.matmul(X, W) + b)
# sum = tf.sigmoid(tf.matmul(X, Ws) + b)
print( hypothesis)
# print( sum )
# -
# ## 텐서플로우 sess.run 시키는 다음 코드를 설명하자
#
# ### 인터넷 자료를 참고해서, tensorflow 1.* 의 sess.run 이해하자
# +
# cost/loss function
cost = -tf.reduce_mean(Y * tf.log(hypothesis) + (1 - Y) * tf.log(1 - hypothesis))
train = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(cost)
# Accuracy computation
# True if hypothesis>0.5 else False
predicted = tf.cast(hypothesis > 0.5, dtype=tf.float32)
accuracy = tf.reduce_mean(tf.cast(tf.equal(predicted, Y), dtype=tf.float32))
# +
# initialize the variables
init = tf.global_variables_initializer()
# Launch graph
with tf.Session() as sess:
# Initialize TensorFlow variables
sess.run( init )
for step in range(201):
sess.run(train, feed_dict={X: x_data, Y: y_data})
if step % 100 == 0:
print(step, ' ==> ', sess.run(cost, feed_dict={X: x_data, Y: y_data}), sess.run(W))
# Accuracy report
h, c, a = sess.run([hypothesis, predicted, accuracy], feed_dict={X: x_data, Y: y_data})
print("\nHypothesis: ", h, "\nCorrect: ", c, "\nAccuracy: ", a)
print( '행렬 \n', sess.run(W) )
# -
# # ====================================
#
#
# # ===================================
#
# <p>
# # (4) RNN 순환신경망과 RNN 실험
#
# <p>
#
# # 학부 이산수학의 점화식 및 recurrence equation 관련
#
# # a_1, a_2 ... , a_n 에서 a_n 은 그 전의 항들과 관계
# # +++++++++++++++++++++++++++++++++++
# <p>
#
# # 먼저 baby 순환신경망 RNN을 만들고 실험한다
#
# <p>
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
import numpy as np
# +
# input data
X0_data = np.array([
# steps 1st 2nd 3rd
[1, 2], [3, 4], [5, 6] # first batch
]) # shape: [batch_size, n_steps, n_inputs]
# input data
X1_data = np.array([
# steps 1st 2nd 3rd
[7, 8], [9, 10], [11, 12] # second batch
]) # shape: [batch_size, n_steps, n_inputs]
# input data
X2_data = np.array([
# steps 1st 2nd 3rd
[13, 14], [15, 16], [17, 18] # third batch
]) # shape: [batch_size, n_steps, n_inputs]
print( X0_data.shape )
# -
# <img src="images/RNN11.png" style="width:500;height:300px;">
# <caption><center> **Figure 1**: Basic RNN model </center></caption>
#
# <img src="images/rnn_step_forward.png" style="width:700px;height:300px;">
# +
#tf.reset_default_graph()
########################
# placeholder 등 그래프 리셋
# hyparameters
n_neurons = 8
# parameters
n_inputs = 2
# build a sequence to sequence rnn model
X0 = tf.placeholder(tf.float32, [None, n_inputs])
# shape = [batch_size, n_inputs]
X1 = tf.placeholder(tf.float32, [None, n_inputs])
X2 = tf.placeholder(tf.float32, [None, n_inputs])
Wx = tf.Variable(tf.random_normal([n_inputs, n_neurons]))
b = tf.Variable(tf.zeros([1, n_neurons]))
Wy = tf.Variable(tf.random_normal([n_neurons, n_neurons]))
y0 = tf.tanh(tf.matmul(X0, Wx) + b) # shape: [batch_size, n_neurons]
y1 = tf.tanh(tf.matmul(y0, Wy) + tf.matmul(X1, Wx) + b)
y2 = tf.tanh(tf.matmul(y1, Wy) + tf.matmul(X2, Wx) + b)
output0 = tf.layers.dense(y0, 1) # shape: [batch_size, 1]
output1 = tf.layers.dense(y1, 1)
output2 = tf.layers.dense(y2, 1)
# -
# # ================================
#
# <p>
# +
# input data
X_data = np.array([
# steps 1st 2nd 3rd
[[1, 2], [7, 8], [13, 14]], # first batch
[[3, 4], [9, 10], [15, 16]], # second batch
[[5, 6], [11, 12], [17, 18]] # third batch
]) # shape: [batch_size, n_steps, n_inputs]
# +
tf.reset_default_graph()
# 이 것이 있어야 에러가 안난다.
##############################
# hyperparameters
n_neurons = 8
# parameters
n_inputs = X_data.shape[2]
n_steps = X_data.shape[1]
print( X_data.shape )
print( n_inputs)
print( n_steps )
# rnn model
X = tf.placeholder(tf.float32, [None, n_steps, n_inputs])
cell = tf.nn.rnn_cell.BasicRNNCell(num_units=n_neurons)
output, state = tf.nn.dynamic_rnn(cell, X, dtype=tf.float32)
# +
# initialize the variables
init = tf.global_variables_initializer()
# train
with tf.Session() as sess:
sess.run(init)
feed_dict = {X: X_data}
output_shape = sess.run(tf.shape(output), feed_dict=feed_dict)
state_shape = sess.run(tf.shape(state), feed_dict=feed_dict)
print('output shape [batch_size, n_steps, n_neurons]: ', output_shape)
print('state shape [batch_size, n_neurons]: ', state_shape)
# -
# ## 학생들이 자체적으로 앞의 코드 작동원리를 스토리텔링
#
# ## 인터넷 서치를 하면 어딘가에는 이런 질문과 답이 있다
#
# https://stackoverflow.com/questions/48384434/what-is-the-structure-of-the-network-based-on-basicrnncell
#
# # +++++++++++++++++++++++++++++++++
# # [탐구] tf.nn.rnn_cell 함수 탐구
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
import numpy as np
# -
X_data = np.array([
# steps 1st 2nd 3rd
[[1, 2], [7, 8], [13, 14]], # first batch
[[3, 4], [9, 10], [15, 16]], # second batch
[[5, 6], [11, 12], [17, 18]] # third batch
]) # shape: [batch_size, n_steps, n_inputs]
# 
# +
tf.reset_default_graph()
# hyperparameters
n_neurons = 8
# parameters
n_steps = X_data.shape[1]
n_inputs = X_data.shape[2]
n_layers = 5 # 5 hidden layers
# rnn model
X = tf.placeholder(tf.float32, [None, n_steps, n_inputs])
layers = [tf.nn.rnn_cell.BasicRNNCell(num_units=n_neurons) for _ in range(n_layers)]
multi_rnn = tf.nn.rnn_cell.MultiRNNCell(layers)
output, state = tf.nn.dynamic_rnn(multi_rnn, X, dtype=tf.float32)
# +
# initializer the variables
init = tf.global_variables_initializer()
# train
with tf.Session() as sess:
sess.run(init)
feed_dict = {X: X_data}
output_shape = sess.run(tf.shape(output), feed_dict=feed_dict)
state_shape = sess.run(tf.shape(state), feed_dict=feed_dict)
print('output shape [batch_size, n_steps, n_neurons]: ', output_shape)
print('state shape [n_layers, batch_size, n_neurons]: ' ,state_shape)
# -
# # +++++++++++++++++++++++++++++++++
# +
import tensorflow.compat.v1 as tf
import numpy as np
tf.disable_eager_execution()
batch_size = 4
vector_size = 3
inputs = tf.placeholder(tf.float32, [batch_size, vector_size])
# print( inputs.shape)
print( inputs ) # Tensor("Placeholder_3:0", shape=(4, 3), dtype=float32)
num_units = 2
state = tf.zeros([batch_size, num_units], tf.float32)
print( state )
cell = tf.nn.rnn_cell.BasicRNNCell(num_units=num_units)
output, newstate = cell(inputs=inputs, state=state)
print("Output of cell.variables is a list of Tensors:")
print(cell.variables)
# [<tf.Variable 'basic_rnn_cell_5/kernel:0' shape=(5, 2) dtype=float32>,
# <tf.Variable 'basic_rnn_cell_5/bias:0' shape=(2,) dtype=float32>]
kernel, bias = cell.variables
print( kernel )
print( bias )
X = np.zeros([batch_size, vector_size])
print(X)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
output_, newstate_, k_, b_ = sess.run(
[output, newstate, kernel, bias], feed_dict = {inputs: X})
print("Output:\n" , output_)
print("New State == Output:\n", newstate_)
print("\nKernel: \n", k_)
print("\nBias:\n", b_)
sess.close()
# -
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
import numpy as np
tf.reset_default_graph()
# Values is data batch_size=2, sequence_length = 3, num_features = 1
values = tf.constant(np.array([
[[1], [2], [3]],
[[2], [3], [4]]
]), dtype=tf.float32)
lstm_cell = tf.nn.rnn_cell.LSTMCell(100)
outputs, state = tf.nn.dynamic_rnn(cell=lstm_cell, dtype=tf.float32, inputs=values)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
output_run, state_run = sess.run([outputs, state])
# +
print( outputs )
print( state.c )
print( state.h )
np.all(output_run[:,-1] == state_run.h)
# -
# <img src="images/LSTM.png" style="width:500;height:400px;">
# # +++++++++++++++++++++++++++
# +
tf.reset_default_graph()
# Values is data batch_size=2, sequence_length = 3, num_features = 1
values = tf.constant(np.array([
[[1], [2], [3]],
[[2], [3], [4]]
]), dtype=tf.float32)
lstm_cell_fw = tf.nn.rnn_cell.LSTMCell(100)
lstm_cell_bw = tf.nn.rnn_cell.LSTMCell(105) # change to 105 just so can see the effect in output
(output_fw, output_bw), (output_state_fw, output_state_bw) = tf.nn.bidirectional_dynamic_rnn(
cell_fw=lstm_cell_fw,
cell_bw=lstm_cell_bw,
inputs=values,
dtype=tf.float32)
# -
output_state_bw.c
# +
tf.reset_default_graph()
# Values is data batch_size=2, sequence_length = 3, num_features = 1
values = tf.constant(np.array([
[[1], [2], [3]],
[[2], [3], [4]]
]), dtype=tf.float32)
lstm_cell = lambda: tf.nn.rnn_cell.LSTMCell(100)
multi_cell = tf.nn.rnn_cell.MultiRNNCell([lstm_cell() for _ in range(3)])
outputs, state = tf.nn.dynamic_rnn(cell=multi_cell, dtype=tf.float32, inputs=values)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
output_run, state_run = sess.run([outputs, state])
# -
state
# # +++++++++++++++++++++++++++++++++
# +
# http://www.wildml.com/2016/08/rnns-in-tensorflow-a-practical-guide-and-undocumented-features/
# http://learningtensorflow.com/index.html
# http://suriyadeepan.github.io/2016-12-31-practical-seq2seq/
import tensorflow.compat.v1 as tf
import numpy as np
tf.disable_eager_execution()
# One hot encoding for each char in 'hello'
h = [1, 0, 0, 0]
e = [0, 1, 0, 0]
l = [0, 0, 1, 0]
o = [0, 0, 0, 1]
# -
# 
#
# +
import pprint
pp = pprint.PrettyPrinter(indent=4)
sess = tf.InteractiveSession()
with tf.variable_scope('one_cell') as scope:
# One cell RNN input_dim (4) -> output_dim (2)
hidden_size = 2
cell = tf.keras.layers.SimpleRNNCell(units=hidden_size)
print(cell.output_size, cell.state_size)
x_data = np.array([[h]], dtype=np.float32) # x_data = [[[1,0,0,0]]]
pp.pprint(x_data)
outputs, _states = tf.nn.dynamic_rnn(cell, x_data, dtype=tf.float32)
sess.run(tf.global_variables_initializer())
pp.pprint(outputs.eval())
# -
# 
#
#
with tf.variable_scope('two_sequances') as scope:
# One cell RNN input_dim (4) -> output_dim (2). sequence: 5
hidden_size = 2
cell = tf.keras.layers.SimpleRNNCell(units=hidden_size)
x_data = np.array([[h, e, l, l, o]], dtype=np.float32)
print(x_data.shape)
pp.pprint(x_data)
outputs, _states = tf.nn.dynamic_rnn(cell, x_data, dtype=tf.float32)
sess.run(tf.global_variables_initializer())
pp.pprint(outputs.eval())
# 
with tf.variable_scope('3_batches') as scope:
# One cell RNN input_dim (4) -> output_dim (2). sequence: 5, batch 3
# 3 batches 'hello', 'eolll', 'lleel'
x_data = np.array([[h, e, l, l, o],
[e, o, l, l, l],
[l, l, e, e, l]], dtype=np.float32)
pp.pprint(x_data)
hidden_size = 2
cell = tf.nn.rnn_cell.LSTMCell(num_units=hidden_size, state_is_tuple=True)
outputs, _states = tf.nn.dynamic_rnn(
cell, x_data, dtype=tf.float32)
sess.run(tf.global_variables_initializer())
pp.pprint(outputs.eval())
with tf.variable_scope('3_batches_dynamic_length') as scope:
# One cell RNN input_dim (4) -> output_dim (5). sequence: 5, batch 3
# 3 batches 'hello', 'eolll', 'lleel'
x_data = np.array([[h, e, l, l, o],
[e, o, l, l, l],
[l, l, e, e, l]], dtype=np.float32)
pp.pprint(x_data)
hidden_size = 2
cell = tf.nn.rnn_cell.LSTMCell(num_units=hidden_size, state_is_tuple=True)
outputs, _states = tf.nn.dynamic_rnn(
cell, x_data, sequence_length=[5,3,4], dtype=tf.float32)
sess.run(tf.global_variables_initializer())
pp.pprint(outputs.eval())
# # ==================================
#
# <p>
#
# # 참고 : 학부 이산수학 U2.TF-DNN_MinMax.ipynb
#
# # [뉴럴네트워크와 이산수학 최적화이론](./U2.TF-DNN_MinMax.ipynb)
# <p>
#
#
# # +++++++++++++++++++++++++++++++++++
#
#
# <p>
#
# #### 참고 사이트 : http://www.easy-tensorflow.com/tf-tutorials/recurrent-neural-networks
# # 텐서플로우와 일차함수, 일차변환 최적화
# +
import tensorflow.compat.v1 as tf
print( tf.__version__)
tf.disable_eager_execution()
# tf.disable_eager_execution() 하여 텐서플로우 2.0 기능을 disable
# -
# ## 강의용 WinPython37F 는 텐서플로우 2.0 버젼이지만
#
# ## 1.* 버젼으로 낮추어 다룬다 (예전 코드와의 호환성 고려)
#
# <p>
#
# ## ++++++++++++++++++++++++++++++++++++++++++
# <p>
#
# ## 텐서플로우 그래프 위에 3종류의 변수를 배치시킨 후 실행 !!
#
# <p>
#
# ## 참고 자료 : https://medium.com/ai-india/hello-world-tensorflow-6ce3f5bcbb6b
#
# <p>
# +
print( tf.add(5,3) )
# + language="javascript"
#
# element.text( 3+5 )
#
# -
#
#
# <h2 id="1.1.-Computational-Graph">1.1. Computational Graph<a class="anchor-link" href="http://i-systems.github.io/HSE545/iAI/DL/topics/02_optimization/01_Optimization_TensorFlow.html#5.1.-Computational-Graph">¶</a></h2><ul>
# <li><code>tf.constant</code></li>
# <li><code>tf.Variable</code></li>
# <li><code>tf.placeholder</code></li>
# </ul>
#
#
# ### 텐서플로우는 먼저 변수를 그래프 구조에 배치를 시킨 후,
#
# ### 그래프를 initiallize 하며 session 을 만들어 run 시킨다.
#
# <h3> tf.constant (텐서플로우 상수)</h3>
# <p>
# <code>tf.constant</code> creates a constant tensor specified by value, dtype, shape and so on.</p>
#
# +
a = tf.constant([1,2,3])
b = tf.constant(4, shape=[1.3])
A = a + b
B = a*b
# -
#
# <p>The result of the lines of code is an abstract tensor in the computation graph. However, contrary to what you might expect, the result doesn’t actually get calculated. It just defined the model, but no process ran to calculate the result.</p>
#
# +
print(A)
# +
B
# -
# ## 이처럼 print 시켜도 아직은 변수의 형태만 써준다 !
#
# <p>To run any of the three defined operations, we need to create a <code>session</code> for that graph. The session will also allocate memory to store the current value of the variable.</p>
# <p>When you think of doing things in TensorFlow, you might want to think of creating tensors (like matrices), adding operations (that output other tensors), and then executing the computation (running the computational graph). In particular, it's important to realize that when you add an operation on tensors, it doesn't execute immediately. Rather, TensorFlow waits for you to define all the operations you want to perform. Then, TensorFlow optimizes the computation graph, deciding how to execute the computation, before generating the data. Because of this, a tensor in TensorFlow isn't so much holding the data as a placeholder for holding the data, waiting for the data to arrive when a computation is executed.</p>
# <p><br/></p>
# <p></p><center><img src="./images/tf_session.png" width="500"/><p></p>
# </center>
# ## 변수들을 tf.Session() 만들어 run 시킨다
# +
sess = tf.Session()
sess.run(A)
# +
sess.run(B)
# -
#
# <p>
#
# ### 아래와 같이 상수 a, b 를 곱하는 연산을 result 라 하고, sess.run 실행시킨 결과 </p>
#
# constant 는 variable 과는 다르게 initialize 시킬 필요는 없다.
#
# +
a = tf.constant([1,2,3])
b = tf.constant([4,5,6])
result = tf.multiply(a, b)
with tf.Session() as sess:
output = sess.run(result)
print(output)
# -
#
# <h3>tf.Variable (텐서플로우 변수)</h3><p><code>tf.Variable</code> is regarded as the decision variable in optimization. We should initialize variables to use <code>tf.Variable</code>.</p>
#
# +
x1 = tf.Variable([1, 1])
x2 = tf.Variable([2, 2])
y = x1 + x2
# +
print(y)
# -
# ## 변수는 먼저 initialize 시킨 후, run 시킨다
# +
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
print( sess.run(y) )
# x1 = tf.Variable([10, 10])
sess.run(y)
# -
#
# <h3>tf.placeholder (값을 담을 메모리 상자)</h3><p>The value of <code>tf.placeholder</code> 는 <code>feed_dict</code> 명령으로 값을 공급하고, 아래와 같이 <code>Session.run()</code> 시킨다.</p>
#
# +
sess = tf.Session()
x = tf.placeholder(tf.float32)
sess.run(x, feed_dict = {x : [[1,2],[3,4]]})
# +
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
sum = a + b
print(sess.run(sum, feed_dict = {a : 1, b : 2}))
print(sess.run(sum, feed_dict = {a : [1,2], b : [3,4]}))
# -
# ## 텐서플로우 그래프에 배치된 변수의 연산을 run 하는 방법
#
#
# <h2 id="1.2.-Tensor-Manipulation">1.2. 텐서프로우 연산</h2><ul>
# <li><code>Adding Matrices</code></li>
# <li><code>Multiplying Matrices</code></li>
# <li><code>Reshape</code></li>
# </ul>
#
#
# <h3 id="Adding-Matrices">Adding Matrices</a></h3>
#
# 두 가지의 sess.run 표현
#
# +
x1 = tf.constant(1, shape = [3])
x2 = tf.constant(2, shape = [3])
output = tf.add(x1, x2)
# method1 use session
sess = tf.Session()
result = sess.run(output)
print(result)
sess.close()
# method2 use session
with tf.Session() as sess:
result = sess.run(output)
print(result)
# +
x1 = tf.constant(1, shape = [2, 3])
x2 = tf.constant(2, shape = [2, 3])
output = tf.add(x1, x2)
with tf.Session() as sess:
result = sess.run(output)
print(result)
# -
#
# <h3 id="Multiplying-Matrices">Multiplying Matrices</a></h3>
#
# 두 가지 표현 방법
#
# +
x1 = tf.constant([[1, 2],
[3, 4]])
x2 = tf.constant([[2],[3]])
# +
output1 = tf.matmul(x1, x2)
with tf.Session() as sess:
result = sess.run(output1)
print(result)
# +
output2 = x1*x2
with tf.Session() as sess:
result = sess.run(output2)
print(result)
# -
#
# <h3 id="Reshape">행렬 모양 바꾸기</h3>
#
# +
x = [1, 2, 3, 4, 5, 6, 7, 8]
x_re = tf.reshape(x, [4,2])
sess = tf.Session()
sess.run(x_re)
# +
x_re = tf.reshape(x, [2,-1])
sess = tf.Session()
sess.run(x_re)
# -
# ### ++++++++++++++++++++++++++++++++++++++
#
#
# # 텐서플로우 코딩을 활용한 Min-Max
#
#
# <h2 id="1.3.-TensorFlow-as-Optimization-Solver">1.3. 텐서플로우로 함수의 최대최소 접근하기</h2><p><br/>
# <span class="MathJax_Preview" style="color: inherit;"></span></p><div class="MathJax_Display" style="text-align: center;"><span class="MathJax" data-mathml='<math xmlns="http://www.w3.org/1998/Math/MathML" display="block"><munder><mo movablelimits="true" form="prefix">min</mo><mrow class="MJX-TeXAtom-ORD"><mi>&#x03C9;</mi></mrow></munder><mspace width="thickmathspace" /><mspace width="thickmathspace" /><mo stretchy="false">(</mo><mi>&#x03C9;</mi><mo>&#x2212;</mo><mn>4</mn><msup><mo stretchy="false">)</mo><mn>2</mn></msup></math>' id="MathJax-Element-61-Frame" role="presentation" style="text-align: center; position: relative;" tabindex="0"><nobr aria-hidden="true"><span class="math" id="MathJax-Span-2183" style="width: 7.086em; display: inline-block;"><span style="display: inline-block; position: relative; width: 5.896em; height: 0px; font-size: 120%;"><span style="position: absolute; clip: rect(1.134em, 1005.9em, 3.098em, -999.997em); top: -2.199em; left: 0em;"><span class="mrow" id="MathJax-Span-2184"><span class="munderover" id="MathJax-Span-2185"><span style="display: inline-block; position: relative; width: 1.67em; height: 0px;"><span style="position: absolute; clip: rect(3.158em, 1001.67em, 4.17em, -999.997em); top: -3.985em; left: 0em;"><span class="mo" id="MathJax-Span-2186" style="font-family: MathJax_Main;">min</span><span style="display: inline-block; width: 0px; height: 3.991em;"></span></span><span style="position: absolute; clip: rect(3.515em, 1000.42em, 4.289em, -999.997em); top: -3.39em; left: 0.598em;"><span class="texatom" id="MathJax-Span-2187"><span class="mrow" id="MathJax-Span-2188"><span class="mi" id="MathJax-Span-2189" style="font-size: 70.7%; font-family: MathJax_Math-italic;">ω</span></span></span><span style="display: inline-block; width: 0px; height: 3.991em;"></span></span></span></span><span class="mspace" id="MathJax-Span-2190" style="height: 0em; vertical-align: 0em; width: 0.301em; display: inline-block; overflow: hidden;"></span><span class="mspace" id="MathJax-Span-2191" style="height: 0em; vertical-align: 0em; width: 0.301em; display: inline-block; overflow: hidden;"></span><span class="mo" id="MathJax-Span-2192" style="font-family: MathJax_Main;">(</span><span class="mi" id="MathJax-Span-2193" style="font-family: MathJax_Math-italic;">ω</span><span class="mo" id="MathJax-Span-2194" style="font-family: MathJax_Main; padding-left: 0.241em;">−</span><span class="mn" id="MathJax-Span-2195" style="font-family: MathJax_Main; padding-left: 0.241em;">4</span><span class="msubsup" id="MathJax-Span-2196"><span style="display: inline-block; position: relative; width: 0.836em; height: 0px;"><span style="position: absolute; clip: rect(3.039em, 1000.3em, 4.408em, -999.997em); top: -3.985em; left: 0em;"><span class="mo" id="MathJax-Span-2197" style="font-family: MathJax_Main;">)</span><span style="display: inline-block; width: 0px; height: 3.991em;"></span></span><span style="position: absolute; top: -4.402em; left: 0.42em;"><span class="mn" id="MathJax-Span-2198" style="font-size: 70.7%; font-family: MathJax_Main;">2</span><span style="display: inline-block; width: 0px; height: 3.991em;"></span></span></span></span></span><span style="display: inline-block; width: 0px; height: 2.205em;"></span></span></span><span style="display: inline-block; overflow: hidden; vertical-align: -0.925em; border-left: 0px solid; width: 0px; height: 2.075em;"></span></span></nobr>
#
# <span class="MJX_Assistive_MathML MJX_Assistive_MathML_Block" role="presentation">
#
# <math display="block" xmlns="http://www.w3.org/1998/Math/MathML"><munder><mo form="prefix" movablelimits="true">min</mo><mrow class="MJX-TeXAtom-ORD"><mi>ω</mi></mrow></munder><mspace width="thickmathspace"></mspace><mspace width="thickmathspace"></mspace><mo stretchy="false">(</mo><mi>ω</mi><mo>−</mo><mn>4</mn><msup><mo stretchy="false">)</mo><mn>2</mn></msup>
# </math>
#
# </span>
#
# </span></div>
#
# <script id="MathJax-Element-61" type="math/tex; mode=display"> \min_{\omega}\;\;(\omega - 4)^2</script>
#
# ## TF 와 numpy 그리고 matplotlib 로 코딩을 해보자 !!
#
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
print(tf.__version__)
# -
#
# <p>
#
# ## y = f(x) = x* x -8* x + 16 이차함수의 최솟값을 구하자 !!
# +
# input weight w 에 대한 cost 줄이기
w = tf.Variable(0, dtype = tf.float32)
# 함수의 변수값 : float
cost = w*w - 8*w + 16
# 함숫값 y=f(x) 의 값
#########################
# Learning Rate 의 준말 LR
LR = 0.05
optm = tf.train.GradientDescentOptimizer(LR).minimize(cost)
######## optm 실행 ++> cost 최소화 ++> w 의 값을 찾음 ###
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
sess.run( w )
# -
#
# ### 이 결과는 w 의 초기값 0
#
# LR = 0.05
#
# optm = tf.train.GradientDescentOptimizer(LR).minimize(cost)
#
#
# ### sess.run( optm ) 시키면 어떻게 되는가 ??
# # GradientDescentOptimizer(LearningRate) ??
#
#
# ## WinPython37F 의 python-3.7.2 안의 이 코드를 불러서 쓴다
#
# https://github.com/tensorflow/tensorflow/blob/v2.1.0/tensorflow/python/training/gradient_descent.py#L30-L82
# +
######## optm 실행 ++> cost 최소화 ++> w 의 값을 찾음 ###
# runs one step of gradient descent
sess.run(optm)
print(sess.run(w))
# cost 최소화를 위해 w 를 바꿈
# runs two step of gradient descent
sess.run(optm)
print(sess.run(w))
# -
#
# <p>Then after just one step of gradient decent the variable was <span class="MathJax_Preview" style="color: inherit;"></span><span class="MathJax" data-mathml='<math xmlns="http://www.w3.org/1998/Math/MathML"><mn>0.4</mn></math>' id="MathJax-Element-62-Frame" role="presentation" style="position: relative;" tabindex="0"><nobr aria-hidden="true"><span class="math" id="MathJax-Span-2199" style="width: 1.551em; display: inline-block;"><span style="display: inline-block; position: relative; width: 1.253em; height: 0px; font-size: 120%;"><span style="position: absolute; clip: rect(1.372em, 1001.25em, 2.384em, -999.997em); top: -2.199em; left: 0em;"><span class="mrow" id="MathJax-Span-2200"><span class="mn" id="MathJax-Span-2201" style="font-family: MathJax_Main;">0.4</span></span><span style="display: inline-block; width: 0px; height: 2.205em;"></span></span></span><span style="display: inline-block; overflow: hidden; vertical-align: -0.068em; border-left: 0px solid; width: 0px; height: 1.004em;"></span></span></nobr><span class="MJX_Assistive_MathML" role="presentation"><math xmlns="http://www.w3.org/1998/Math/MathML"><mn>0.4</mn></math></span></span><script id="MathJax-Element-62" type="math/tex">0.4</script>, and <span class="MathJax_Preview" style="color: inherit;"></span><span class="MathJax" data-mathml='<math xmlns="http://www.w3.org/1998/Math/MathML"><mi>&#x03C9;</mi></math>' id="MathJax-Element-63-Frame" role="presentation" style="position: relative;" tabindex="0"><nobr aria-hidden="true"><span class="math" id="MathJax-Span-2202" style="width: 0.836em; display: inline-block;"><span style="display: inline-block; position: relative; width: 0.658em; height: 0px; font-size: 120%;"><span style="position: absolute; clip: rect(1.551em, 1000.66em, 2.324em, -999.997em); top: -2.14em; left: 0em;"><span class="mrow" id="MathJax-Span-2203"><span class="mi" id="MathJax-Span-2204" style="font-family: MathJax_Math-italic;">ω</span></span><span style="display: inline-block; width: 0px; height: 2.146em;"></span></span></span><span style="display: inline-block; overflow: hidden; vertical-align: -0.068em; border-left: 0px solid; width: 0px; height: 0.718em;"></span></span></nobr><span class="MJX_Assistive_MathML" role="presentation"><math xmlns="http://www.w3.org/1998/Math/MathML"><mi>ω</mi></math></span></span><script id="MathJax-Element-63" type="math/tex">\omega</script> reaches <span class="MathJax_Preview" style="color: inherit;"></span><span class="MathJax" data-mathml='<math xmlns="http://www.w3.org/1998/Math/MathML"><mn>0.76</mn></math>' id="MathJax-Element-64-Frame" role="presentation" style="position: relative;" tabindex="0"><nobr aria-hidden="true"><span class="math" id="MathJax-Span-2205" style="width: 2.146em; display: inline-block;"><span style="display: inline-block; position: relative; width: 1.789em; height: 0px; font-size: 120%;"><span style="position: absolute; clip: rect(1.372em, 1001.73em, 2.384em, -999.997em); top: -2.199em; left: 0em;"><span class="mrow" id="MathJax-Span-2206"><span class="mn" id="MathJax-Span-2207" style="font-family: MathJax_Main;">0.76</span></span><span style="display: inline-block; width: 0px; height: 2.205em;"></span></span></span><span style="display: inline-block; overflow: hidden; vertical-align: -0.068em; border-left: 0px solid; width: 0px; height: 1.004em;"></span></span></nobr><span class="MJX_Assistive_MathML" role="presentation"><math xmlns="http://www.w3.org/1998/Math/MathML"><mn>0.76</mn></math></span></span><script id="MathJax-Element-64" type="math/tex">0.76</script> after two steps of gradient decent.</p>
#
# +
for _ in range(100):
sess.run(optm)
print(sess.run(w))
sess.close()
# -
#
# <p>After 100 more steps we are reaching 3.999914.</p>
#
# ## 텐서플로우 실행으로 최솟값 찾기 !
# +
w = tf.Variable(0, dtype = tf.float32)
cost = w*w - 8*w + 16
LR = 0.05
with tf.Session() as sess:
tf.global_variables_initializer().run()
optm = tf.train.GradientDescentOptimizer(LR).minimize(cost)
print( 'w의 시작값 : ', sess.run(w) )
for _ in range(30):
sess.run(optm)
print('w의 값 : ', sess.run(w))
print( ' w = ', sess.run(w) , ' 에서의 cost 값 = ' , sess.run( cost ))
# -
# ## matplotlib 의 plot 으로 최솟값의 그래프를 그리며 계산 !
# +
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
import matplotlib.pyplot as plt
# %matplotlib inline
############# Graph 만들기 ##########
w = tf.Variable(0, dtype = tf.float32)
cost = w*w - 8*w +16
LR = 0.05
optm = tf.train.GradientDescentOptimizer(LR).minimize(cost)
############ session 실행하기 ##########
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
cost_record = []
print( 'cost의 시작값 : ', sess.run(cost) )
for _ in range(50):
sess.run(optm)
print('cost 값 : ', sess.run(cost), ' at w = ', sess.run(w))
cost_record.append(sess.run(cost))
print("\n 최적값 optimal w =", sess.run(w) , ' 에서의 최솟값은 ' , sess.run(cost) )
########### 그래프 그리기 #############
plt.figure(figsize = (10,8))
plt.plot(cost_record)
plt.xlabel('iteration', fontsize = 15)
plt.ylabel('cost', fontsize = 15)
plt.show()
# -
# # [연습문제] 다양한 중학교 함수로 최대최소를 찾아본다
# # [발전문제] 고등학교와 대학교 함수의 경우로 확대한다
# # GradientDescentOptimizer(LearningRate) ??
#
#
# ## 텐서플로우 기본과 경사하강법 그리고 house 가격에 응용
#
# 참고 자료 : https://medium.com/ai-india/hello-world-tensorflow-6ce3f5bcbb6b
#
# # [응용] 다음과 같은 house 크기와 가격의 관계를 함수로 나타내자
#
# <p>
#
#
# ## (1 단계) data_in/houses.csv 파일의 데이터를 전처리 preprocessing !!
#
# +
import pandas as pd
# import library to split the data
from sklearn.model_selection import train_test_split
# %matplotlib inline
import matplotlib.pyplot as plt
def read_data():
"""
read the CSV data
"""
data = pd.read_csv("./images/houses.csv")
print(data.head())
size_data = data["Size"].values
price_data = data["Price"].values
return size_data, price_data
# read the data from the csv file.
size, price = read_data()
print( len(size) )
plt.scatter(size , price, label='house prices')
plt.draw()
# -
# ## 데이터를 train, test 용으로 나누고 normalize !!
# +
def split_test_train(size, price):
# split the data, test size = 33%
size_train, size_test, price_train, price_test = train_test_split(size, price, test_size=0.33)
return size_train, size_test, price_train, price_test
# Normalize a data set
def normalize(array):
return (array - array.mean()) / array.std()
# split the data into testing and training set.
X_train, X_test, Y_train,Y_test = split_test_train(size, price)
# 779 개의 데이터 print( len(size))
# print the length of test and train dataset.
print("Length of the training data: ", len(X_train))
print("Length of the testing data: ", len(X_test))
# normalize the data
X_train = normalize(X_train)
Y_train = normalize(Y_train)
X_test = normalize(X_test)
Y_test = normalize(Y_test)
# plot the normalized data
plt.scatter(X_train, Y_train, label='Samples data')
plt.draw()
# -
# ## ( 2 단계) 텐서플로우 변수를 그래프 위에 만들자 !!
# +
import tensorflow.compat.v1 as tf
import numpy as np
def get_model_tensors():
"""
function obtain model tensor
"""
# X is the placeholder for size of the house from the dataset.
# Y is the placeholder for size of the house from the dataset.
X = tf.placeholder("float")
Y = tf.placeholder("float")
# The parameters theta0 and theta1
theta1 = tf.Variable(np.random.randn(), name="weight")
theta0 = tf.Variable(np.random.randn(), name="bias")
# Hypothesis = theta0 + theta1 * X
x_theta1 = tf.multiply(X, theta1)
model = tf.add(x_theta1 , theta0)
return X, Y, theta1, theta0, model
def get_cost_optimizer_tensor(Y, model, size, learning_rate):
"""
Tensor for calculating the cost function and the the optimizer for
minimizing the cost.
"""
# Cost function tensor.
cost_function = tf.reduce_sum(tf.pow(model - Y, 2))/(2 * size)
# gradient descent tensor.
gradient_descent = tf.train.GradientDescentOptimizer(learning_rate)
# optimization tensor.
optimizer = gradient_descent.minimize(cost_function)
return optimizer, cost_function
# Set parameters
learning_rate = 0.1
training_iteration = 200
X, Y, theta1, theta0, model = get_model_tensors()
optimizer, cost_function = get_cost_optimizer_tensor(Y, model,len(X_train), learning_rate )
# -
# ## (3 단계) session 을 만들어 실행을 시키자 !!
# +
# Initialize variables
init = tf.initialize_all_variables()
# Launch a graph
with tf.Session() as sess:
sess.run(init)
display_step = 20
# Fit all training data
for iteration in range(training_iteration):
# Run the gradient descent optimizer
sess.run(optimizer, feed_dict={X: X_train, Y: Y_train})
# Display logs per iteration step
if iteration % display_step == 0:
training_cost = sess.run(cost_function, feed_dict={X: X_train, Y: Y_train})
print( "Cost function during training: ", "cost=", "{:.9f}\n".format(training_cost),
"a=", sess.run(theta1), "b=", sess.run(theta0), "\n" )
tuning_cost = sess.run(cost_function, feed_dict={X: X_train, Y: Y_train})
print( "Tuning completed:", "cost=", "{:.9f}\n".format(tuning_cost),
"a=", sess.run(theta1), "b=", sess.run(theta0), "\n" )
# Validate a tuning model
testing_cost = sess.run(cost_function, feed_dict={X: X_test, Y: Y_test})
print( "Testing data cost:" , testing_cost )
# Display a plot
plt.figure()
plt.plot(X_train, Y_train, 'ro', label='Normalized samples')
plt.plot(X_test, Y_test, 'go', label='Normalized testing samples')
plt.plot(X_train, sess.run(theta1) * X_train + sess.run(theta0), label='Fitted line')
plt.legend()
plt.show()
print( "일차함수는 : ", sess.run(theta1) ," * x + ", sess.run(theta0) )
# -
# # [연습문제] 이와 같은 regression 예를 찾아 코딩한다
# # +++++++++++++++++++++++++++++++++++++++
#
#
# <p>
#
# # 파트 C : AI 융합수학교육 참고자료
#
# <p>
#
# ## 고등학교에서 텐서플로우로 대학교 최적화 문제를 다루려면 ?
#
#
#
# <p>
#
# # +++++++++++++++++++++++++++++++++++++++
# 
#
# ### 서울대학교 1학년 문과수학 내용 중에 나오는 최적화 문제
#
# ### 파이썬 코딩으로 해당하는 함수의 등고선 그래프를 그림
# <p>
# <p>
#
# # z=f(x,y) 삼차원 그래프 (자바스크립트 코딩)
#
# <p>
#
# ### 아래를 [run] 시키면 3차원 터틀크래프트 환경으로 연결됩니다.
#
# <p>
# + language="html"
#
#
# <iframe src="http://www.javamath.com/snucode/" width=900 height=500>
# </iframe>
# -
# ## 바로 위를 [run] 하여 터틀크래프트가 나오게 하고
#
# ## 아래의 코드를 복사하고 [beginxyz] 단추를 누르자.
# # +++++++++++++++++++++++++++++++
#
# beginxyz
#
# window( 0.05 )
#
# if( abs( x* x* x + y* y* y - 3* x* y - z ) < 0.2 ) return 7
#
# return 0
# ### ++++++++++++++++++++++
#
#
# beginxyz
#
# x=0.05*x
# y=0.05*y
# z=0.05*z
# // window(0.05) 로 대치 가능
#
# function f(x,y) {
# return y*y*y + x*x*x - 3*x*y
# }
#
# if( z<=f(x,y) && z>=f(x,y)-0.5 ) {
# if(z>=0) {return 6}
# else if(z<=-1) { return 5}
# else return 7
# }
#
# return 0
#
# # z=f(x,y) 그래프 파이썬 코딩
# +
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
sz = 100
x,y = np.meshgrid(np.linspace(-2,2,sz),np.linspace(2,-2,sz))
z = x*x*x + y*y*y - 3*x*y
#plt.contourf(x,y,z, vmin=-7, vmax=7, cmap='jet')
level = np.arange(-7,7,0.5)
plt.contourf(x,y,z, vmin=-2, vmax=2, cmap='jet', levels=level)
plt.contour(x,y,z,colors='black', levels=level)
#plt.show()
# -
# ## 다음 파이썬 코드는 진화전략을 사용한 애니메이션
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
sz = 100
x,y = np.meshgrid(np.linspace(-2,2,sz),np.linspace(2,-2,sz))
# 넘파이 매쉬그리드는 엑스축으로 먼저, 와이축으로 다음
# 행렬의 크기 순서와 mashgrid랑 차이남
Z = x*x*x + y*y*y - 3*x*y
# plt.imshow( z, vmin=-1, vmax=1, cmap='jet')
from IPython.display import clear_output
import time
np.random.seed(3)
nn = 7 # number of steps to take (and plot horizontally)
alpha = 0.03 # learning rate
sigma = 3 # standard deviation of the samples around current parameter vector
w = np.array([70.0, 30.0])
# start point : size=100 이므로 가로x 먼저 세로y 다음
#plt.ion() # something about plotting
#plt.figure(figsize=(10,7))
#plt.figure(figsize=(20,15))
prevx, prevy = [], []
for q in range(nn):
plt.figure(figsize=(10,10))
# draw the optimization landscape
#ax1 = plt.subplot(1,nn,q+1)
plt.imshow(Z, vmin=-1, vmax=1, cmap='jet')
# draw a population of samples in black
noise = np.random.randn(200, 2)
wp = np.expand_dims(w, 0) + sigma*noise
x,y = zip(*wp)
plt.scatter(x,y,4,'k', edgecolors='face')
# draw the current parameter vector in white
plt.scatter( [w[0]],[w[1]], 40,'w', edgecolors='face')
# draw estimated gradient as white arrow
R = np.array( [ Z[ int(wi[1]), int(wi[0]) ] for wi in wp ] )
R -= R.mean()
R /= R.std() # standardize the rewards to be N(0,1) gaussian
g = np.dot(R, noise)
u = alpha * g
plt.arrow( w[0], w[1], u[0], u[1], head_width=3, head_length=5, fc='w', ec='w')
plt.axis('off')
plt.title('iteration %d, reward %.2f' % (q+1, Z[int(w[0]), int(w[1])]))
# draw the history of optimization as a white line
prevx.append(w[0])
prevy.append(w[1])
if len(prevx) > 0:
plt.plot(prevx, prevy, 'wo-')
w += u
#plt.axis('tight')
plt.pause(1.0)
#time.sleep(1.0)
clear_output(wait=True)
# -
# # [문제] 문제 만들고 탐구하기
#
#
# 
# beginxyz
#
# window(0.05)
#
# function f(x,y) {
# return 3*(-x*x*x-y*y*y)*exp(-x*x-y*y)
# }
#
# if( z<=f(x,y) && z>=f(x,y)-0.1 ) {
# if(z>1) {return 6}
# else if(z<-1) { return 5}
# else return 7
# }
#
# return 0
#
# ### 자신이 만든 z=f(x,y) 함수의 그래프를 그리고 최대 최소를 탐구하자.
# 
#
# 
# # Rosenbrock 함수를 찾아보고, 최솟값을 구해보자
#
# beginxyz
#
# window(0.05)
# a=1; b=100; c=0.001
#
# function f(x,y) {
# return c*((a-x)*(a-x)+b*pow(y-x*x,2))
# }
#
# if( z<=f(x,y) && z>=f(x,y)-c*250 ) {
# if(z==0) { return 5 }
# else return 7
# }
#
# return 0
# +
# %matplotlib inline
import matplotlib.pyplot as plt
#plt.style.use('seaborn-white')
import numpy as np
#from mpl_toolkits import mplot3d
def rosen(x):
"""Generalized n-dimensional version of the Rosenbrock function"""
return sum(100*(x[1:]-x[:-1]**2.0)**2.0 +(1-x[:-1])**2.0)
x = np.linspace(-5, 5, 100)
y = np.linspace(-5, 5, 100)
X, Y = np.meshgrid(x, y)
Z = rosen(np.vstack([X.ravel(), Y.ravel()])).reshape((100,100))
# Note: the global minimum is at (1,1) in a tiny contour island
plt.contour(X, Y, Z, 1000, cmap = 'jet')
plt.text(1, 1, 'x', va='center', ha='center', color='red', fontsize=20);
# +
def Rosenbrock(x,y):
return (1 + x)**2 + 100*(y - x**2)**2
def Grad_Rosenbrock(x,y):
g1 = -400*x*y + 400*x**3 + 2*x -2
g2 = 200*y -200*x**2
return np.array([g1,g2])
def Hessian_Rosenbrock(x,y):
h11 = -400*y + 1200*x**2 + 2
h12 = -400 * x
h21 = -400 * x
h22 = 200
return np.array([[h11,h12],[h21,h22]])
# +
def Gradient_Descent(Grad,x,y, gamma = 0.00125, epsilon=0.0001, nMax = 10000 ):
#Initialization
i = 0
iter_x, iter_y, iter_count = np.empty(0),np.empty(0), np.empty(0)
error = 10
X = np.array([x,y])
#Looping as long as error is greater than epsilon
while np.linalg.norm(error) > epsilon and i < nMax:
i +=1
iter_x = np.append(iter_x,x)
iter_y = np.append(iter_y,y)
iter_count = np.append(iter_count ,i)
#print(X)
X_prev = X
X = X - gamma * Grad(x,y)
error = X - X_prev
x,y = X[0], X[1]
print(X)
return X, iter_x,iter_y, iter_count
root,iter_x,iter_y, iter_count = Gradient_Descent(Grad_Rosenbrock,-2,2)
# +
x = np.linspace(-2,2,250)
y = np.linspace(-1,3,250)
X, Y = np.meshgrid(x, y)
Z = Rosenbrock(X, Y)
#Angles needed for quiver plot
anglesx = iter_x[1:] - iter_x[:-1]
anglesy = iter_y[1:] - iter_y[:-1]
# %matplotlib inline
fig = plt.figure(figsize = (16,8))
#Surface plot
ax = fig.add_subplot(1, 2, 1, projection='3d')
ax.plot_surface(X,Y,Z,rstride = 5, cstride = 5, cmap = 'jet', alpha = .4, edgecolor = 'none' )
ax.plot(iter_x,iter_y, Rosenbrock(iter_x,iter_y),color = 'r', marker = '*', alpha = .4)
ax.view_init(45, 280)
ax.set_xlabel('x')
ax.set_ylabel('y')
#Contour plot
ax = fig.add_subplot(1, 2, 2)
ax.contour(X,Y,Z, 50, cmap = 'jet')
#Plotting the iterations and intermediate values
ax.scatter(iter_x,iter_y,color = 'r', marker = '*')
ax.quiver(iter_x[:-1], iter_y[:-1], anglesx, anglesy, scale_units = 'xy', angles = 'xy', scale = 1, color = 'r', alpha = .3)
ax.set_title('Gradient Descent with {} iterations'.format(len(iter_count)))
plt.show()
# -
# # +++++++++++++++++++++++++++
#
#
# # 텐서플로 2.0 버젼으로 바꾸기 ????
#
#
#
# # +++++++++++++++++++++++++++
| TF-Keras_CHATBOT/.ipynb_checkpoints/DnnMath-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib as mat
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import statsmodels.api as stats
import sklearn as sklearn
from sklearn import preprocessing
print("pandas version:", pd.__version__)
print("numpy version:", np.__version__)
print("matplotlib version:", mat.__version__)
print("statsmodels version:", stats.__version__)
print("sklearn version:", sklearn.__version__)
# +
# Read the data from the csv file
dataset = pd.read_csv("train.csv")
# print(dataset.describe())
# print(dataset.shape) # (1460, 81)
# The dataset is huge and has so many independent variables.
# But we will be taking into account only the Living-Area to predict the Sales-Price of the houses.
# X corresponds to the gross living area (feature)
# Y corresponds to the predicted sales price (label)
X = dataset['GrLivArea']
Y = dataset['SalePrice']
X_mean = X.mean()
Y_mean = Y.mean()
print("Mean for X:", X_mean)
print("Mean for Y:", Y_mean)
print()
X_std = X.std()
Y_std = Y.std()
print("Standard Deviation for X:", X_std)
print("Standard Deviation for Y:", Y_std)
print()
X_min = X.min()
X_max = X.max()
X_diff = (X_max - X_min)
print("X_min: {} and X_max: {} for X".format(X_min, X_max))
print("Diff of X_min and X_max for X is {}".format(X_diff))
print()
plt.scatter(X, Y, color = "red")
plt.title("Scatter Plot", fontsize = 20)
plt.xlabel("Living-Area", fontsize = 20)
plt.ylabel("Sales-Price", fontsize = 20)
# This shows the point at which both X and Y values are equal to X_mean and Y_mean respectively.
plt.scatter(X_mean, Y_mean, color = "black")
plt.show()
print()
# +
# Normalize the dataset
X_normalized = ((X - X_min)/X_diff) # not being here
# Standardize the dataset
X_standardized = ((X - X_mean)/X_std)
# Add a column of 1s for the gradient descent
X_modified = np.c_[np.ones(X.shape[0]), X_standardized]
print(X_modified)
# NOTE: We can also make use of de-standardize and de-normalize preocesses if the situation requires it.
# +
# Gradient Descent
alpha = 0.01 #Step size
iterations = 2000 #No. of iterations
m = Y.size #No. of data points
np.random.seed(8496) #Set the seed
theta = np.random.rand(2) #Pick some random values to start with
# print(theta) # [0.76779152 0.1691429 ]
#GRADIENT DESCENT
def gradient_descent(x, y, theta, iterations, alpha):
past_costs = []
past_thetas = [theta]
for i in range(iterations):
prediction = np.dot(x, theta)
error = prediction - y
cost = 1/(2*m) * np.dot(error.T, error)
past_costs.append(cost)
theta = theta - (alpha * (1/m) * np.dot(x.T, error))
past_thetas.append(theta)
return past_thetas, past_costs
#Pass the relevant variables to the function and get the new values back...
past_thetas, past_costs = gradient_descent(X_modified, Y, theta, iterations, alpha)
theta = past_thetas[-1]
#Print the results...
print("Gradient Descent: {:.2f}, {:.2f}".format(theta[0], theta[1]))
print()
# print(past_thetas)
# -
# # Plot the cost function v/s number of iterations
plt.title('Cost Function J', fontsize = 20)
plt.xlabel('No. of iterations', fontsize = 20)
plt.ylabel('Cost', fontsize = 20)
plt.plot(past_costs)
plt.show()
print()
# +
# Show the model
print("theta[0]: {} and theta[1]: {}".format(theta[0], theta[1]))
X_retrieved = X_modified[:,1]
Y_predicted = theta[1]*X_retrieved + theta[0]
X_retrieved_mean = X_retrieved.mean()
Y_predicted_mean = Y_predicted.mean()
plt.scatter(X_retrieved, Y, color = "red")
plt.gca().set_title("Gradient Descent Linear Regressor", fontsize = 15)
plt.xlabel("Living-Area", fontsize = 20)
plt.ylabel("Sales-Price", fontsize = 20)
# This shows the point at which both X and Y values are equal to X_mean and Y_mean respectively.
plt.plot(X_retrieved, Y_predicted, lw = 4, c = 'orange', label = 'regression line')
plt.scatter(X_retrieved_mean, Y_predicted_mean, color = "black")
plt.show()
print()
| regression-problems/univariate-linear-regression-with-gradient-descent/linear-regression-with-gradient-descent-impl-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Traduction automatique
# Attention: ce notebook ne fonctionne pas dans l'environnement Docker. Vous pouvez le tester à travers Google Colab: https://colab.research.google.com/
# <img src="colab.png" width="1000"/>
# ## Exemple d'utilisation d'un modèle transformers avec Hugging Face
# Librairies supplémentaires à installer :
# !pip install transformers torch sentencepiece
# Commençons par importer deux modules :
from transformers import AutoTokenizer, AutoModelForSeq2SeqLM
# Ensuite téléchargeons un modèle :
model_name = "Helsinki-NLP/opus-mt-fr-en"
_ = AutoTokenizer.from_pretrained(model_name)
_ = AutoModelForSeq2SeqLM.from_pretrained(model_name)
# Chargeons en mémoire le tokenizer et le modèle :
tokenizer = AutoTokenizer.from_pretrained(model_name, local_files_only=True)
model = AutoModelForSeq2SeqLM.from_pretrained(model_name, local_files_only=True)
# Nous pouvons à présent traduire...
# +
text = "Mon tailleur est riche."
tokenized_text = tokenizer(text, return_tensors='pt', padding=True)
translation = model.generate(**tokenized_text)
translated = tokenizer.batch_decode(translation, skip_special_tokens=True)
print(translated)
# -
# Pour plus de facilité, vous pouvez aussi utiliser <https://github.com/Teuze/translate>
| module5/s2_machine_translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# language: python
# name: python37464bita6c5c53d7b16474cae5bc645c5b09138
# ---
# # Getting started with modeling and analysis of biological systems
# + [markdown] slideshow={"slide_type": "slide"}
# 
# 
#
# ***
# ***
# + [markdown] slideshow={"slide_type": "slide"}
# # What is this "guide"?
#
# I am writing this article as a guide for anyone who wants to start working on modeling and analysis of biological systems. The topics include but are not limited to creating mathematical models for biological systems, running simulations, accessing public repositories of biological models, and some advanced mathematical modeling discussions.
#
# # # Who?
#
# If you are a mathematician, control theorist, engineer interested in applying your skills to biological systems that inspire you - this is definitely an article for you.
#
# If you are a bioengineer/biologist working on experiments but want to use mathematical models to improve analysis and understanding of your systems - this article could be helpful to you as well.
#
# # # Where?
#
# The article is right here - you are reading it, but this guide is supposed to be an interactive guide. So, you can download a Jupyter notebook [here] and play around with it.
#
# More information is available on a course webpage that I helped teach at Caltech in Spring 2020 - [BE240](https://www.cds.caltech.edu/~murray/wiki/BE_240,_Spring_2020)
#
# # Background
#
# Before jumping right ahead with creating and analyzing models, it is important to get a grasp of a "language" in which biological models are written. This standardized language is called Systems Biology Markup Language (SBML).
#
# ## Introduction to SBML
#
# As shown in the Figure above, SBML is a language which can be used to exchange your biological models in. If a lab or a set of users write models or develop tools compatible with SBML, then all of their tools and models can be easily accessed and used by the community all around the world. A one-line definition of SBML from the SBML official webpage concisely describes this:
#
# ### [SBML](http://sbml.org) is a _"free and open interchange format for computer models of biological processes"_
#
# ### 1. SBML uses the language of XML
# Even the simplest SBML model file can contain hundreds of lines, full of various XML tags. The header looks like this:
# ```
# <sbml xmlns="http://www.sbml.org/sbml/level2/version3" level="2" metaid="_153818" version="3">
# <model id="BIOMD0000000012" metaid="_000001" name="Elowitz2000 - Repressilator">
# ```
# **You don't have to write your own SBML files by hand! - That's where various tools help you. **
# + [markdown] slideshow={"slide_type": "subslide"}
# ## A quick peek into the SBML model:
# Model header:
# Species: 
# + [markdown] slideshow={"slide_type": "slide"}
#
# ## 2. Software that support SBML : More than 300!
#
# * Model building tools: [Tellurium](http://tellurium.analogmachine.org/) (Python), [Sub-SBML](https://github.com/BuildACell/subsbml) (Python), [iBioSim](https://async.ece.utah.edu/tools/ibiosim/) (GUI), [MATLAB SimBiology](https://www.mathworks.com/products/simbiology.html)
# * Model simulation tools: [bioscrape](https://github.com/biocircuits/bioscrape/) (Python), [COPASI](http://copasi.org/) (GUI), [LibRoadRunner](http://libroadrunner.org/) (Python), [MATLAB SimBiology](https://www.mathworks.com/products/simbiology.html)
# * Analysis tools: [bioscrape inference](https://github.com/biocircuits/bioscrape/) (Python), [COPASI](http://copasi.org/) (GUI), [ABC-SysBio](http://www.theosysbio.bio.ic.ac.uk/resources/abc-sysbio/) (Python)
#
# Follow [this link](http://sbml.org/SBML_Software_Guide/SBML_Software_Summary) for detailed descriptions of various software tools that support SBML.
#
#
# Models in SBML : A big curated model database : [BioModels](https://www.ebi.ac.uk/biomodels/). An example model from this database at the end of this notebook.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 3. For software developers : SBML API for Python : python-libsbml.
#
# `python-libsbml` is the API to write/read SBML models using a script. Unless you are developing your own new software, you wouldn't need to learn about it in detail. Just knowing that the API exists and the software tools that are compatible with SBML use it is enough for a non-developer user. However, in case you are planning to develop your own software or change an existing one, you might need to learn about how this API functions. Here are a couple lines of introduction of this API. Assume that you have an object called `model`. This is an instance of the Model class in libsbml, each model has an `SBMLDocument` object associated to it as well that functions as a "holder" of the `Model` object. For example: To
#
# * set/change parameter values: `model.getParameter(6).setValue(1e3)` => Changes the value of the 6th parameter in the list of parameters to 1e3.
# * set initial conditions: `model.getSpecies('id').setInitialAmount(50)` => Changes the initial condition of the species with identifier "id" to 50.
#
# and so on...
#
# In my opinion, a good place to start working on your own SBML development project is by looking at the documentation for the `Model` class in python-libsbml. Once you get a hang of how the objects work here, you should feel much more comfortable. The [Documentation](http://sbml.org/Software/libSBML/5.18.0/docs/python-api/) is available on this link. It is also worth mentioning that the SBML and the libsbml community is one of the most helpful and active software development community. So, if you face any issues with SBML or related tools, feel free to jump on their [Google groups discussion forums](https://groups.google.com/forum/#!forum/libsbml-development)!
#
# ***
# ---
# -
# # Let's take an example : The famous "repressilator" system:
#
# Here's what we will do
# * Find the SBML model for the "repressilator" as implemented in their paper [[1]](https://www.nature.com/articles/35002125).
# * Choose a simulator to run the SBML model.
# * Run the simulations, study the results.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Simulating a repressilator circuit (using its SBML model)
# From [biomodels](https://www.ebi.ac.uk/biomodels/) SBML model repository, we can get the SBML model that accompanies the original repressilator paper [[1]](https://www.nature.com/articles/35002125).
# -
# ## Simulator options:
#
# 1. COPASI - http://copasi.org/ - A commonly used GUI based simulator and modeling engine for biological models. Plenty of online resources available to learn this tool. Recommended for users who like a graphical user interface to create and analyze their models.
#
# 2. RoadRunner - http://libroadrunner.org/ - A deterministic simulation engine specifically designed for SBML models.
#
# 3. Bioscrape - https://github.com/biocircuits/bioscrape/ - (Personal bias warning) - A fast stochastic and deterministic simulator tool based on Python. I am a developer of this tool so I am biased towards using it ;). Next, I demonstrate how this can be used to simulate SBML models:
# # Using [bioscrape](https://github.com/biocircuits/bioscrape/)
# We import the SBML file obtained into bioscrape to simulate it. For more information on how to simulate a bioscrape model, refer to [this](http://www.cds.caltech.edu/%7Emurray/courses/be240/sp2020/W2_bioscrape.ipynb) notebook.
# + [markdown] slideshow={"slide_type": "slide"}
# ## The ODE model (from [[1]](https://www.nature.com/articles/35002125.pdf)):
#
# 
# In the SBML model: PX: lacI protein, PY: TetR protein, PZ: cI protein.
# + slideshow={"slide_type": "slide"}
# Import bioscrape simulator and import_sbml from sbmlutil to import SBML files into bioscrape.
from bioscrape.sbmlutil import import_sbml
from bioscrape.simulator import py_simulate_model
# Import the SBML file : Usage : import_sbml('sbml_filename.xml'), returns bioscrape Model object
# (Make sure that the file path is correctly specified wherever the SBML file ending in .xml is present in your directory)
M_represillator = import_sbml('repressilator_sbml.xml', sbml_warnings = False)
#Simulate Deterministically and Stochastically
import numpy as np
timepoints = np.linspace(0,700,10000)
result_det = py_simulate_model(timepoints, Model = M_represillator)
result_stoch = py_simulate_model(timepoints, Model = M_represillator, stochastic = True)
# + slideshow={"slide_type": "skip"}
# Import relevant settings and packages to create plots
import matplotlib.pyplot as plt
import matplotlib as mpl
color_list = ['r', 'k', 'b','g','y','m','c']
mpl.rc('axes', prop_cycle=(mpl.cycler('color', color_list) ))
mpl.rc('xtick', labelsize=16)
mpl.rc('ytick', labelsize=16)
plt.figure(figsize = (10, 4))
# + slideshow={"slide_type": "subslide"}
#Plot Results
for i in range(len(M_represillator.get_species_list())):
s = M_represillator.get_species_list()[i]
plt.plot(timepoints, result_det[s], color = color_list[i], label = "Deterministic "+s)
plt.plot(timepoints, result_stoch[s], ":", color = color_list[i], label = "Stochastic "+s)
plt.title('Repressilator Model')
plt.xlabel('Time', FontSize = 16)
plt.ylabel('Amount', FontSize = 16)
plt.legend()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## More examples (try these out yourself! I will be happy to chat more about this):
#
# 1. **Try your own SBML model!**
# * Similar to the repressilator example above, try creating your own SBML model and run a simulation.
# * If you are interested, use [Tellurium](http://tellurium.analogmachine.org/) or [COPASI](http://copasi.org/) to create a SBML model, then simulate using bioscrape or compare simulations with other libraries.
# * Get a SBML model from one of the repositories online of your favorite paper, then simulate using bioscrape. For example, similar to the repressilator example above, BioModels repository consists SBML models of a
# * Toggle Switch
# * Influenza Viral Dynamics Spread
# * Circadian Oscillator
# * MAPK/ERK pathway
# * Other interesting examples available [here](https://www.ebi.ac.uk/biomodels/content/model-of-the-month?all=yes).
# * Pro Tip : For bioscrape users, you can directly load an SBML model using the `Model`constructor. Usage : `M = Model(sbml_filename = "sbml_filename.xml")`
#
| getting-started/getting_started_biology_modeling_tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day and Night Image Classifier
# ---
#
# The day/night image dataset consists of 200 RGB color images in two categories: day and night. There are equal numbers of each example: 100 day images and 100 night images.
#
# We'd like to build a classifier that can accurately label these images as day or night, and that relies on finding distinguishing features between the two types of images!
#
# *Note: All images come from the [AMOS dataset](http://cs.uky.edu/~jacobs/datasets/amos/) (Archive of Many Outdoor Scenes).*
#
# ### Import resources
#
# Before you get started on the project code, import the libraries and resources that you'll need.
# +
import cv2 # computer vision library
import helpers
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# -
# ## Training and Testing Data
# The 200 day/night images are separated into training and testing datasets.
#
# * 60% of these images are training images, for you to use as you create a classifier.
# * 40% are test images, which will be used to test the accuracy of your classifier.
#
# First, we set some variables to keep track of some where our images are stored:
#
# image_dir_training: the directory where our training image data is stored
# image_dir_test: the directory where our test image data is stored
# Image data directories
image_dir_training = "day_night_images/training/"
image_dir_test = "day_night_images/test/"
# ## Load the datasets
#
# These first few lines of code will load the training day/night images and store all of them in a variable, `IMAGE_LIST`. This list contains the images and their associated label ("day" or "night").
#
# For example, the first image-label pair in `IMAGE_LIST` can be accessed by index:
# ``` IMAGE_LIST[0][:]```.
#
# Using the load_dataset function in helpers.py
# Load training data
IMAGE_LIST = helpers.load_dataset(image_dir_training)
# ---
# # 1. Visualize the input images
#
# +
# Select an image and its label by list index
image_index = 0
selected_image = IMAGE_LIST[image_index][0]
selected_label = IMAGE_LIST[image_index][1]
## TODO: Print out 1. The shape of the image and 2. The image's label `selected_label`
print('Shape of image:', selected_image.shape)
print('Label of image:', selected_label)
plt.imshow(selected_image)
plt.show()
## TODO: Display a night image
# Note the differences between the day and night images
# Any measurable differences can be used to classify these images
# Zoek de eerste nachtfoto en laat deze ook zien
for pair in IMAGE_LIST:
if pair[1] == "night":
plt.imshow(pair[0])
plt.show()
break
# -
| 1_1_Image_Representation/6_1. Visualizing the Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Welcome to the SETI Institute Code Challenge!
#
# This first tutorial will explain a little bit on what the data is and where to get it.
#
# # Update 23 Januar 2018
#
# This project has been a huge success and we'd like to thank all of the participants, the winning team `Effsubsee` and the scientific members of the SETI Insitute and IBM.
#
# We are beginning to decommision this project. However, it will still be useful as a learning tool. The only real change is that the primary full data set will be removed. The `basic`, `primary small` and `primary medium` data sets will remain.
#
#
# # Update 21 June 2017
#
# We learned a lot at the hackathon on June 10-11th and decided to regenerate the primary data set. This is called the `v3` primary data set. The changes, compared to `v2` are: the noise background is gaussian white noise instead of noise from the Sun, the signal amplitudes are higher and the characteristics should make them more distinguishable, and there are only 140k in the full set (20k per signal type), compared with 350k previously (50k per signal type).
#
# The `basic` data set remains unchanged from before.
#
# # Introduction
#
# For the Code Challenge, you will be using the **"primary" data set**, as we've called it. The primary data set is
#
# * labeled data set of 35000 simulated signals
# * 7 different labels, or "signal classifications"
# * total of about 10 GB of data
#
# This data set should be used to train your models.
#
# As stated above, we no longer have the full 140,000 data set (51 GB). All of the data are found in the `primary medium` data set below. Additionally , there is the `basic4` data set and the and `primay small` sub set. They are explained below.
#
#
# ## Simple Data Format
#
# Each data file has a simple format:
#
# * file name = <UUID>.dat
# * a JSON header in the first line that contains:
# * UUID
# * signal_classification (label)
# * followed by stream complex-valued time-series data.
#
# The `ibmseti` Python package is available to assist in reading this data and performing some basic operations for you.
#
# ## Basic Warmup Data Set.
#
# There is also a second, simple and clean data set that you may use for warmup, which we call the **"basic" data set**. This basic set should be used as a sanity check and for very early-stage prototyping. We recommend that everybody starts with this.
#
# * Only 4 different signal classifications
# * 1000 simulation files for each class: 4000 files total
# * Available as single zip file
# * ~1 GB in total.
#
# ### Basic Set versus Primary Set
#
# > The difference between the `basic` and `primary` data sets is that the signals simulated in the `basic` set have, on average, much higher signal to noise ratio (they are larger amplitude signals). They also have other characteristics that will make the different signal classes very distinguishable. **You should be able to get very high signal classification accuracy with the basic data set.** The primary data set has smaller amplitude signals and can look more similar to each other, making classification accuracy more difficult with this data set. There are also only 4 classes in the basic data set and 7 classes in the primary set.
#
#
# ## Primary Data Sets
#
# ### Primary Small
#
# The `primary small` is a subset of the full primary data set. Use for early-stage prototyping.
#
# * All 7 signal classifications
# * 1000 simulations / class (7 classes = 7000 files)
# * Available as single zip file
# * ~2 GB in total
#
# ### Primary Medium
#
# The `primary medium` was a subset of the full primary data set but it not constitutes the entire data set. You may want to consider ways to augment this data set in order to create more training samples. Additionally, you could consider splitting each file up into 4 or 5 smaller files and simply build models that accept smaller files. You wouldn't be able to use this for post scores to the Scoreboards, but it would be one way to generate more data. Finally, we hope to one day release the simulation code, which would allow you to generate your own data sets.
#
# * All 7 signal classifications
# * 5000 simulations / class (7 classes = 35000 files)
# * Large enough for relatively robust model construction
# * Available in 5 separate zip files
# * ~10 GB in total
#
# ### Primary Full
#
# #### **THIS DATA SET IS NO LONGER AVAILABLE.**
#
#
#
# ## Index Files
#
# For all data sets, there exists an **index** file. That file is a CSV file. Each row holds the UUID, signal_classification (label) for a simulation file in the data set. You can use these index files in a few different ways (from using to keep track of your downloads, to facilitate parallelization of your analysis on Spark).
#
#
#
# ## Direct Data URLs if you are working from outside of IBM Data Science Experience
#
# ### Basic4
# [Data (1.1 GB)](https://ibm.box.com/shared/static/1zhbd1yvblmkt42485y1r7qlqo5ny6jm.zip)
#
# [Index File](https://ibm.box.com/shared/static/lv7w6lxxpleeqpqpqg3p7e96kpe6nbmi.csv)
#
#
# ### Primary Small
#
# [Data (1.9 GB)](https://ibm.box.com/shared/static/8rh54gtz6wqnzas849r1vkjewtf68mvp.zip)
#
# [Index File](https://ibm.box.com/shared/static/6y66vklx1mnx1ozn1zww58mrdsfzjc7h.csv)
#
# ### Primary Medium
#
# [Data Zip File 1 (1.9 GB)](https://ibm.box.com/shared/static/n7zexohsj0eil18bg1ubv2exlmw3hia1.zip)
#
# [Data Zip File 2 (1.9 GB)](https://ibm.box.com/shared/static/bdp01bj3vzzxeq0ta7d53ekyx71w2n7q.zip)
#
# [Data Zip File 3 (1.9 GB)](https://ibm.box.com/shared/static/dh2bab97hqqztwrcihpe0jqmcg6s6755.zip)
#
# [Data Zip File 4 (1.9 GB)](https://ibm.box.com/shared/static/ndldz1sh1jr3zfb990wi8fkqz1kk1wot.zip)
#
# [Data Zip File 5 (1.9 GB)](https://ibm.box.com/shared/static/wwhg8wz85rjynygpnyvwc6eey8yspsnn.zip)
#
# [Index File](https://ibm.box.com/shared/static/mdzp9yxjvp3ljpcjyw380mvm4d7haavm.csv)
#
#
# It's probably easiest to download these zip files, unzip them separately, then move the contents of to a single folder.
#
#
#
# # Test Data Sets
#
# Once you've trained your model, done all of your cross-validation testing, and are ready to submit an entry to the contest, you'll need to download the test data set and score the test set data with your model.
#
#
# The test data files are nearly the same as the training sets. The only difference is the JSON header in each file does not contain the signal class. You can use `ibmseti` python package to read each file, just like you would the training data. See [Step_2_reading_SETI_code_challenge_data.ipynb](https://github.com/setiQuest/ML4SETI/blob/master/tutorials/Step_2_reading_SETI_code_challenge_data.ipynb) for examples.
#
# ### Note:
# ### July 1 - July 21: Only the "Preview" test set is available.
# ### July 21 - July 31: The final test set is now available.
#
# <br>
#
# ## Preview Test Set
# The `primary_testset_preview_v3` data set contains 2414 test simulation files. Each data file is the same as the above training data except the JSON header does NOT contain the 'signal_classification' key.
#
# * All 7 classes
# * Roughly 340 simulations per class
# * JSON header with UUID only
# * Available as single zip file
# * 665 MB in total
#
#
# ### Direct Download Link
#
# [Preview Test Set Zip File](https://ibm.box.com/shared/static/91z783n1ysyrzomcvj4o89f4b8ss76ct.zip)
#
# [Preview Test Set Index File](https://ibm.box.com/shared/static/mvlhxesth3gsd5b6gkj91me7noydr3fl.csv)
#
#
#
# ## Final Test Set
# The `primary_testset_final_v3` data set contains 2496 test simulation files. Each data file is the same as the above training data except the JSON header does NOT contain the 'signal_classification' key.
#
# * All 7 classes
# * Roughly 350 simulations per class
# * JSON header with UUID only
# * Available as single zip file
# * 687 MB in total
#
#
#
# ### Direct Download Link
#
# [Final Test Set Zip File](https://ibm.box.com/shared/static/kg50txjb1j3yawoys8gfyoxsedx84lvv.zip)
#
# [Final Test Set Index File](https://ibm.box.com/shared/static/lmsgynqttidmcitx1uqwnyrz9utpl27x.csv)
#
#
#
#
#
# ### Submitting Classification Results
#
# See the [Judging Criteria](https://github.com/setiQuest/ML4SETI/blob/master/Judging_Criteria.ipynb) notebook for information on submitting your test-set classifications.
# # Getting Data from IBM Spark service
#
# If you're working with IBM Watson Data Platform (or Data Science Experience), you can use either `wget` or `curl` from a Jupyter notebook cell. Or you can use the `requests` library, or similar, to download the files programmatically. (This should work for both the IBM Spark service backend or the IBM Analytics Engine backend.) Simply call `wget` command-line from the shell using the appropriate shell command syntax. The shell command syntax is different for Python kernels versus Scala kernels. Below we show you the Python kernel way, asssuming that the vast majority will use Python.
# +
# #copy link from above.
#make sure to use the -O <filename.zip> to redirect the output
# !wget https://ibm.box.com/shared/static/91z783n1ysyrzomcvj4o89f4b8ss76ct.zip -O primary_testset_preview_v3.zip
# -
# !ls -al primary_testset_preview_v3.zip
import zipfile
zz = zipfile.ZipFile('primary_testset_preview_v3.zip')
zz.namelist()[:10]
| tutorials/Step_1_Get_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3HTm3uxaAblr"
# # GC-SAN
# > Graph Contextualized Self-Attention Network for Session-based Recommendation.
# + [markdown] id="hRoFlQ80AiEr"
# Session-based recommendation, which aims to predict the user’s immediate next action based on anonymous sessions, is a key task in many online services (e.g. e-commerce, media streaming). Recently, Self-Attention Network (SAN) has achieved significant success in various sequence modeling tasks without using either recurrent or convolutional network. However, SAN lacks local dependencies that exist over adjacent items and limits its capacity for learning contextualized representations of items in sequences. Graph contextualized self-attention model (GC-SAN) utilizes both graph neural network and self-attention mechanism, for session-based recommendation. In GC-SAN, we dynamically construct a graph structure for session sequences and capture rich local dependencies via graph neural network (GNN). Then each session learns long-range dependencies by applying the self-attention mechanism. Finally, each session is represented as a linear combination of the global preference and the current interest of that session.
# + [markdown] id="OFlDQ7etBFC1"
# ## Architecture
#
# <p><center><img src='_images/C699874_1.png'></p></center>
#
# We first construct a directed graph of all session sequences. Based on the graph, we apply graph neural network to obtain all node vectors involved in the session graph. After that, we use a multi-layer self-attention network to capture long-range dependencies between items in the session. In prediction layer, we represent each session as a linear of the global preference and the current interest of that session. Finally, we compute the ranking scores of each candidate item for recommendation.
# + [markdown] id="5gHzeQCPBKff"
# ### Performance
#
# <p><center><img src='_images/C699874_2.png'></p></center>
| docs/C699874_GC_SAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Physics laws commonly used in mathematical modeling of biological systems
# This notebook lists physics laws which are commonly used in modeling biological systems. It is intended for biology students who want to have an idea of the content and meaning of physical laws relevant to their systems, in order to improve their communication skills with quantitative scientists in collaborative modeling projects. It describes the laws, the meaning of the variables and parameters, the range of applicability and the biological context(s) in which the laws are useful/used, and the conservation laws it is generally associated with. When possible, simplified versions of the law and/or partially solved problems that are very common in biology will be presented. This is a "forever-work-in-progress" notebook, that will be enriched by your suggestions!
#
#
#
#
# ## 1. Fick's law(s) of diffusion
# Many biomolecular systems of interest in biomedicine are inhomogeneous, and experiments measure spatial gradients of molecule concentration, activity, or interactions... etc. When inhomogeneity arises in a continuous system, there is a universal "force" that tends to drive the system back to homogeneity over time. This trend is quantified by the Fick's law:
#
# $$ \vec{J} = -D * \vec\nabla{C}$$
#
# This law represents the flux vector $\vec{J}$ of a molecule in a solution, that is generated by the existence of a spatial gradient $\vec\nabla{C}$ in the concentration $C$ of the molecule. In this law, the concentration flux is quantified per unit time, and across a unit surface orthogonal to the gradient. $D$ is the diffusion coefficient (always positive), it has the units (in physics, we call this "dimension") of a surface per unit time, and depends on the type of molecule, the existence or not of a physical barrier orthogonal to the gradient (e.g. porous membrane...), temperature... etc. This first Fick's law expresses the fact that a flux is generated by a concentration gradient. It is a consequence of a much more general law of Nature: systems tend to become homogeneous over time. Hence, try to have Fick's law in mind everytime you know there is some inhomogeneity in any feature of the system you model: there will be a flux, along the feature gradient, from regions where the feature is "high" to regions where it is "low".
#
# The equation for the flux is often coupled to a balance equation for the concentration C(x,y,z,t) of molecule (or more generally the inhomogenous feature) that depends on time and the 3 space coordinates x, y, z:
#
#
# $$ \frac{\partial C}{\partial t} + \vec\nabla . \vec J = R$$
#
# where $R$ is the net rate of molecule production (production - destruction/conversion to something else) at this particular location of the system. The term $ \vec\nabla . \vec J = \frac{\partial J_x}{\partial x} + \frac{\partial J_y}{\partial y} + \frac{\partial J_z}{\partial z}$ (where x, y z are the 3 directions of space and $J_x,y,z$ the components of the flux vector along these directions) is called the <b> divergence </b> of the flux vector $\vec J$ and expresses the balance of inward and outward molecule fluxes at the particular location where you are looking at (local) concentration changes. Indeed, in a continuous system, the flux vector might be different at different positions in space because spatial gradients might <b> themselves </b> be inhomogenous. The total net amount of molecules transported per unit time at your particular location is the difference between the outward flux at our location and the inward flux from <b> neighbor locations </b>. Hence, when computing this net flux, the derivatives of the flux vector $\vec J$ components along the three directions of space naturally appear when writing the "spatial balance equation".
#
# But long story short:
#
# $$ \frac{\partial C}{\partial t} + \vec\nabla . \vec J = \frac{\partial C}{\partial t} + \frac{\partial J_x}{\partial x} + \frac{\partial J_y}{\partial y} + \frac{\partial J_z}{\partial z} =R$$
#
# is a fundamental "conservation equation" that expresses the fact that, locally, within a short amount of time $dt$ there is no other changes in the local amount of molecules than the net production rate $R$: molecules are not instantaneously transported to or from distant places in the system. <b> This local conservation equation shall be written for any space-dependent model </b>.
#
# An immediate consequence is that, when plugging the first Fick's law in this conservation equation (and assuming the diffusion coefficient $D$ is homogeneous in space, one obtains the <b> diffusion equation </b>:
#
#
# $$ \frac{\partial C}{\partial t} = D* \Delta C + R = D*(\frac{\partial^2 C}{\partial x^2}+\frac{\partial^2 C}{\partial y^2}+\frac{\partial^2 C}{\partial z^2}) + R$$
#
#
# We can remark here that this balance equation for the concentration $C$ is very similar to the ODEs developed in the notebook Section 2 - Animating the structure, with the addition of one extra term: the diffusion term $D*(\frac{\partial^2 C}{\partial x^2}+\frac{\partial^2 C}{\partial y^2}+\frac{\partial^2 C}{\partial z^2})$.
#
# The Fick's law(s) shall be used in modeling any interacting biomolecular systems (in the continuum approximation) that is not homogeneous in space, so typically with existing gradients of molecules concentrations or activity within a given organelle. It can also be used to model the passive transfer of biomolecules through porous membranes (such as blood vessels) due to concentration gradients, osmotic fluxes ... any situation where an inhomogenous system has the potential to become homogenous over time.
#
#
# ## 2. Solid dynamics and the Newton equations
# Many fields of biomedicine require the description/modeling of processes that involve more than just biochemical signaling. Once cells or tissues are not just anymore the "physical supports" of biochemical reactions, but are themselves moving, growing or changing their shape, they are subject to universal physical laws. Depending on the situation and the degree of complexity of the model, biological objects can be modeled as rigid solids, fluids, or deformable (elastic/nonelastic) solids. These 3 situations are summarized in sections 2, 3 and 4 of this notebook.
#
# Non-deformable solids can have 2 types of motion: an overall motion of their center of mass, and a rotational motion around the center of mass (that leads the position of the center of mass unaffected). Real dynamics often involve complex motion patterns, but patterns can always be decomposed as a sum of center of mass motion + rotations.
#
# ### Motion of the center of mass
# The motion of the center of mass of an object A is governed by the second Newton law:
#
# $$\frac{d}{dt}(M*\vec V_A) = \Large\Sigma \vec F_{all->A} $$
#
# where $M$ is the total mass, $V_A$ is the velocity of the center of mass, and $\Large\Sigma \vec F_{all->A}$ is the sum of all forces acting on the object.
#
# In other words, if there is no net total force acting on an object, the motion will continue indefinitely along a straight line with constant velocity. If the object is immobile, it will remain immobile. And if you want to initiate an overall motion, you have to apply a net force.
#
#
# ### Rotational motion
# But having a null net force does not mean no force at all is applied on the object. You could apply a force at one point of the object, and an opposite force at another point. The net force is 0 and the center of mass won't move. And, if the forces exerted are along the line joining the two points where they are exerted, nothing will happen. Think about a Hockey puck you would equally press on two diametrally opposed points. However, if now those opposite forces are exerted with some angle relative to the puck diameter joining the two points, then the puck will spin. You have exerted what is called a <b> Torque </b>, denoted $\tau$. Specifically, you are applying a torque along the direction which is orthogonal to both the direction of the force, and the line joining the two points. This is the rotational equivalent of the net force for overall motions, and it obeys a very similar equation:
#
# $$\frac{d}{dt}(I*\vec\Omega_A) = \Large\Sigma \vec \tau_{all->A},$$
#
# where $\vec\Omega_A$ represent the angular velocity vector (rotations with respect to x, y and z axes), $I$ is called the <b> moment of inertia </b> and accounts for how the mass of the solid is distributed with respect to the axes, and as above $\Large\Sigma \vec \tau_{all->A}$ is the total net torque acting on the solid A.
#
#
# ### Action-reaction principle
# Forces between object are a consequence of microscopic <b> interactions. </b>. As a consequence, is we write or say that an object A exerts a force $\vec F_{A->B}$ on an object B, then we implicitly write or say that the object B exerts an opposite force, because the "direction" of the interaction is a matter of point of view:
#
# $$\vec F_{B->A}= -\vec F_{A->B}.$$
#
# Both cannot be dissociated, if $\vec F_{A->B}$ exists, $\vec F_{B->A}$ exists as well and is opposite. However, generally we are interested in the dynamics of A, or of B, so we only use one of the two forces.
#
#
# ### Example of some common forces in biomechanics
# Friction (resists motion of two object relative to each other; solid friction, fluid friction or viscous force, lubricated friction... etc); Adhesion forces, like cell-cell junctions, originate in actin-supported transmembrane protein anchoring to a support or another cell); Traction forces, have many origins including retrograde actin flow in crawling cells; Tensile/compression forces generated by myosin motion along cytoskeleton fibers; Elastic forces, that oppose the deformation of an object...
#
# Cells generate forces using their cytoskeleton and other molecular tools, hence they use biochemistry. In return, external force sensing by cells is also based on biochemistry. How the different forces mentioned above are mathematically linked to the biochemical signals defines a biomechanical model.
#
#
# ## 3. Fluid dynamics and the Navier-Stokes equations
# The mechanical laws given above apply to non-deformable solids. Similar laws apply to the mechanics of deformable materials. The most deformable materials are fluids, which immediately flow (and thus, irreversibly deform) when a stress is exerted on them. They immediately dissipate the mechanical energy due to the stress. Less deformable are the deformable solids (see 4.), which have the ability to reversibly deform (elastic deformation), and then regain their original shape. We say that elastic solids have the ability to transiently store the mechanical emergy of the stress. Under physiologically relevant timescales cells are intrinsically viscoelastic, meaning they display a combination of both elastic and viscous reponses to mechanical stress. So it is good to know a bit about the equations that govern the dynamics of (viscous) fluids (section 3 here), and the equations that govern elastic deformations (section 4).
#
# Let's start with (viscous) fluids. Their dynamics are governed by the Navier Stokes equation:
#
#
# $$\rho \frac{\partial \vec u}{\partial t}+ \rho (\vec u. \vec \nabla) \vec u = - \vec \nabla P + \vec \nabla . T + \vec f $$
#
# It seems complicated... It is not. $\rho$ is the volumetric mass, $\vec u$ the velocity field (see below), $P$ the pressure field, and $T$ the stress tensor that characterizes the mechanical properties of the fluid (see below).
#
# It is the equivalent of the second Newton law, but adapted to fluids. The velocity of a fluid does not make any sense: think about a river flow, where should we measure the velocity? The answer is: everywhere. For fluids, we do not talk of a unique velocity, but of a "velocity field", which is no more no less than the water flow vector (direction+intensity) at any point within the fluid. Idem for the pressure, which is a defined "locally" at any point within the fluid.
#
# And because we're talking of a <b> local </b> velocity (field), rather than being weighted by the entire fluid mass like in Newton's law, it is weighted by the volumetric mass $\rho$, i.e. the mass per unit volume of fluid. So we understand that $\rho \frac{\partial \vec u}{\partial t}$ is the equivalent of $M \frac{d V_A}{dt}$. So what about the second term on the left hand side?
#
# We have to take a step back and remember the balance equations. To establish a balance equation on a variable Y, we list sources of the rate of change of Y at time t, and write Y(t+dt)=Y(t)+dt*Rate_of_change_of_Y. What if Y also depends on space, e.g. the coordinate x? Then the equation takes a form of Y(t+dt,x)=Y(t,x)+dt*Rate_of_change_of_Y(x)+dx*Y_flux_from_x-dx - dx*Y_flux_to_x+dx. But now what happens if, in our system, there is an <b> intrinsic flow </b> that transports small sections of the system further away along the x axis with a velocity $u_x$ ? Then it does not make sense to compare Y(t+dt,x) with Y(t,x) because the in the meantime, the initial position x in our system has been transported to a new location $x+u_x*dt$, so when calculating the balance on Y, we should compare $Y(t+dt,x+u_x*dt)$ with $Y(t,x)$. Because we don't like implementing 2 small changes at the same time (i.e., a small time shift dt and a space shift), we compensate exactly for the effect of the internal flow on "displacing" system elements by adding this second term. Thinking in more mathematical way, if we have to derivate overtime a function $Y(t,x)$ where $x(t)$ also depends on time (which is the case in presence of the flow), then $dY/dt=\partial Y / \partial t + \partial x/ \partial t * \partial Y / \partial x$.
#
# In 3 dimensions, where the velocity field has components along the y and z axis as well, it becomes $$ \frac{\partial Y}{\partial t}+ (\vec u. \vec \nabla) Y$$. This is called the <b> material derivative </b>. This expression for the time derivative of any function of space and time $Y(t,x,y,z)$ needs always to be used when there is a flow. In particular, we need to take this into account when modeling cellular actin density, and related biochemistry, in the presence of a retrograde flow. In it a very common situation in biology. And in the context of the Navier Stokes equation, the variable $Y$ we are interested in is ... the velocity vector $\vec u$ itself. Hence the left hand side of the NS equation.
#
# What about the right hand side? As in Newton's formula, it contains the forces. Because the equation is written locally, since the velocity field is different everywhere, the right hand side includes the local pressure gradient (analogous to Fick's law, a pressure gradient generates material motion to "equilibrate" pressures), the local force field $\vec f$ if any (including, for instance, gravity, electromagnetic forces if the fluid has electromagnetic properties... anything that acts "remotely" on the local fluid elements), and finally the divergence of the stress tensor $\vec \nabla T$ that represents the local forces that are due to elements of the fluid acting on their neighbor.
#
# The stress tensor contains all the information on the mechanical properties of our system, and how it dissipates emergy. Typically, in biological problems, the stress tensor accounts for shear stresses (fluid analogous of torques for solids), and fluid viscosity (the viscous force being due to the friction of neighboring layers of fluids when you want to move on piece of fluid further...). If you drag your finger through a liquid, your finger will pull some liquid due to viscosity, and this might also cause some turbulence in the fluid due to shear stresses.
#
# If the fluid is incompressible, or nearly, which is the case of water and dilute acqueous solutions, or even lipids that make cell membranes, in physiological temperature and pressure conditions, the stress tensor simplifies and the Navier Stokes equation becomes:
#
#
# $$\rho \frac{\partial \vec u}{\partial t}+ \rho (\vec u. \vec \nabla) \vec u = - \vec \nabla P + \vec f + \mu \Delta \vec u ,$$
#
# where the $\Delta$ operator sums up all second derivatives relative to spatial coordinates (see diffusion equation above). The constant coefficient $\mu$ is called viscosity, and determines how much force is locally generated by a flux of material at the velocity $\vec u$. Similarly, the viscosity defines the rate at which a fluid can flow under a given external stress ($\vec f$ and $-\vec\nabla P$).
#
# So, in short, without solving the full (or the reduced) Navier Stokes equation, in a viscous fluid the local stress is proportional to the gradient of the velocity, i.e. to the <b> rate of strain </b>, the proportionality constant being the viscosity:
#
# $$ \sigma = \mu * \frac{d \epsilon}{dt} $$.
#
# This is why the divergence of the stress tensor produces the term $ \mu \Delta \vec u $ in the full equation, And this is what we will remember here.
#
# ## 4. In between solids and liquids: notion of elasticity in biology
#
# While fluid dynamics theories based on Navier Stokes-like equations might prove efficient to model what happens in the cytosol, or within organells which are essentially aqueous media, such theories become insufficient to explain cellular properties where the cell, or part of it (like a rigid organelle, the plasma membrane...etc) opposes a resistance to the deformation. In this situation, an approach accounting for the elastic properties of the cell or its components is required.
#
# What are the physics equations governing elasticity ?
#
# When a spring is stretched from its equilibrium by an extra length $\Delta l$, there is a restoring force actin on whatever is stretching the spring, that opposes the direction of the stretch and has an amplitude of $k*\Delta l$ where $k$ is called the spring constant and depends on what the spring is made of and how it's made. If we stop applying the force, the spring will oscillate back to its initial size. This is called an elastic deformation. If we pull too much, we might irreversibly deform the spring: we have been outside of the "elastic response" range of the material.
#
# Similarly, for any deformable material there is a range of "strain" $\epsilon$ (i.e., transcient deformation) in which the stress $\sigma$ (force) produced by the solid to recover its initial shape is linearly proportional to the strain. The constant $E$ is called the Young modulus and we have:
#
# $$ \sigma = E*\epsilon $$.
#
# If the strain is applied along one axis, there is also a resulting deformation along the transverse axis that results from the microscopic interactions that hold the solid together, and this transverse strain is also proportional to the initial strain:
#
# $$ \epsilon_{transv.} = - \epsilon / \nu $$
#
# where $\nu$ is called the Poisson ratio and the minus sign stands there because if you try to elongate the solid along one direction, it will naturally respond by "compressing" along the transverse direction to minimize volume changes. You have experienced this a thousand times in your everyday life.
#
# Because both the transverse strain and the stress that follow a given strain are both proportional to the initial strain, we talk of <b> linear elasticity theory </b>.
#
# Why is this useful when modeling biochemical systems?
#
#
# ## 5. Viscous and elastic at the same time: the viscoelastic cellular medium
# The interior of the cell is a very complex medium where elastic organelles and the visco-elastic cytoskeleton are immersed in a viscous fluid and wrapped in viscous fluid-like membranes.
# Hence, the local stress tensor $\sigma$ has components both proportional to the strain tensor, and to the rate of strain tensor (see above).
#
# In this view, the cellular medium can be modeled as a assembly of springs and dashpots representing respectively the elastic components and the viscous components. How springs and dashpots are combined will produce different force/velocity (or stress/strain) relationships, that will need to be compared with experiments.
#
#
# More information available in this article, written in a very pedagogical way yet exposing up-to-date models:
#
# https://doi.org/10.1063/1.4900941
#
#
#
#
#
#
# ## 6. Other possibilities (temp, work in progress, depending on what's covered in 3 and 4 already)
#
#
#
# Young - Laplace law:
# MT = (P * r) / 2 (bubble/spherical),
# MT = wall tension,
# p = pressure,
# r = radius
#
#
# Flow:
#
# F = deltaP / R,
# deltaP = pressure change between two points,
# R = peripheral resistance,
# F = volumetric flow,
#
#
# Laminar flow:
#
# Q = (Pi r^4 deltaP) / (8 n L),
# Q = volumetric flow rate,
# r = radius,
# deltaP = pressure gradient,
# n= dynamic viscosity,
# L = length,
#
#
# Turbulent flow speed:
#
# Vc = (R n) / (p r),
# R = Reynolds nr,
# n = dynamic viscosity,
# p = density,
# r = radius,
#
#
# Reynolds number:
#
# R = (Vc p r) / n
#
#
# Laplace's law:
#
# T = P*r (cyllindrical vessel),
# T = wall tension,
# p = pressure,
# r = radius,
#
#
# Hydrostatic pressure:
# HSP = p g l,
# p = density,
# g = gravity,
# l = height
#
#
# Osmotic pressure:
# Pi = RTci,
# R = gas constant,
# T = absolute temperature,
# c = concentration,
# i = Hoff's coefficient,
#
#
#
#
# Internal force friction:
#
# F = (n deltaV s) / d,
# n = dynamic viscocity,
# deltaV = difference in speed,
# s = surface area,
# d = distance,
#
#
# Flow viscosimeter:
#
# n = (n0 (pt)) / (p0 * t0),
# n = dynamic viscocity,
# 0 = known value,
# p = density,
# t = time,
#
#
# Poiseuilles law:
#
# deltaP = (8 n L Q) / (Pi r^4),
# deltaP = pressure change,
# n = dynamic viscocity,
# L = length,
# Q = flow rate,
# r = radius,
#
#
#
# More reading on cell mechanics:
#
# https://doi.org/10.1002/wsbm.1275
#
| J_Mathematical_Modeling/Section 2/SuppNoteBook-PhysicsLawsInBiology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem Solving Approaches
# ## Greedy Algorithms
#
# A greedy algorithm is an algorithm that works by selecting the best choice at each step.
#
# An algorithm can be solved with a greedy algorithm if it has the following properties.
#
# __Greedy Choice Property__
#
# An optimal solution can be found by choosing the best choice at each step without reconsidering the previous steps.
#
# __Optimal Substructure Property__
#
# The optimal solution to the problem is also the optimal solution to all of the subproblems.
#
# ### Greedy Approach
#
# 1. The solution step starts as empty
# 2. At each step, an answer is added until the solution is complete
# 3. If the solution set is feasible, the current item is kept
# 4. Otherwise, the item is rejected and never looked at again.
#
# ### Example Greedy Problem
#
# ```
# Q. Make $18 using $1, $5, and $10
# A. We can take the biggest at each step
# ```
#
# ### Example Non-Greedy Problem
# Greedy algorithms will not always given the best approach.
#
# For example: Find the max path in a graph will not work, since certain weights at different paths may be larger than the max weight at each step.
# ## Dynamic Algorithms
#
# Dynamic programming problems cache (memoization) the result of subproblems to find the optimal result.
#
# Dynamic programming is similar to greedy algorithms, but greedy algorithms look for a locally optimum solutions, while dynamic programming combines the results of subprograms to find the optimum solution.
#
# We can tell if a program can be solved by dynamic programming if it satisfies the following properties.
#
# __Optimal Substructure__
#
# The optimal solution can be found by using the optimal solution of the subproblems.
#
# For example, the shortest path would have optimal substructure, but the longest path would not.
#
# The knapsack problem has an optimal substructure since the max value for items[i-1] gives the max value without item[i].
#
# __Overlapping Subproblems__
#
# To find an answer to a problem i, we have to use solutions that were computed when finding i-1. For example, when computing fib(9), we need to use answers we found when computing fib(8).
#
# We can cache these answers using memoization.
| Python/Problem Solving Approach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.039647, "end_time": "2021-12-05T05:22:18.496611", "exception": false, "start_time": "2021-12-05T05:22:18.456964", "status": "completed"} tags=[]
# # **Introduction**
# Predictive models are often used by investors to decide whether a budding business would be profitable in their domain. The success of a new restaurant can be similarly predicted based on past data relating to the location and services offered. We will develop a predictor model to find out the popularity and success a new restaurant can generate with an extensive study. This study aims to offer huge insights on which factors determine the success of a new restaurant and predict ratings for newer restaurants
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 1.404123, "end_time": "2021-12-05T05:22:19.939620", "exception": false, "start_time": "2021-12-05T05:22:18.535497", "status": "completed"} tags=[]
#Importing Libraries
import numpy as np #NumPy is a general-purpose array-processing package.
import pandas as pd #It contains high-level data structures and manipulation tools designed to make data analysis fast and easy.
import matplotlib.pyplot as plt #It is a Plotting Library
import seaborn as sns #Seaborn is a Python data visualization library based on matplotlib.
from sklearn.linear_model import LogisticRegression #Logistic Regression is a Machine Learning classification algorithm
from sklearn.linear_model import LinearRegression #Linear Regression is a Machine Learning classification algorithm
from sklearn.model_selection import train_test_split #Splitting of Dataset
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import r2_score
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + papermill={"duration": 11.597494, "end_time": "2021-12-05T05:22:31.576122", "exception": false, "start_time": "2021-12-05T05:22:19.978628", "status": "completed"} tags=[]
df = pd.read_csv('../input/zomato-bangalore-restaurants/zomato.csv')
df.head()
# + papermill={"duration": 0.050804, "end_time": "2021-12-05T05:22:31.668629", "exception": false, "start_time": "2021-12-05T05:22:31.617825", "status": "completed"} tags=[]
df.shape
# + [markdown] papermill={"duration": 0.039348, "end_time": "2021-12-05T05:22:31.747718", "exception": false, "start_time": "2021-12-05T05:22:31.708370", "status": "completed"} tags=[]
# # **EDA and Cleaning**
# + papermill={"duration": 0.169728, "end_time": "2021-12-05T05:22:31.959076", "exception": false, "start_time": "2021-12-05T05:22:31.789348", "status": "completed"} tags=[]
df.info()
# + papermill={"duration": 0.147168, "end_time": "2021-12-05T05:22:32.147218", "exception": false, "start_time": "2021-12-05T05:22:32.000050", "status": "completed"} tags=[]
df.isna().sum()
# + papermill={"duration": 0.055269, "end_time": "2021-12-05T05:22:32.243653", "exception": false, "start_time": "2021-12-05T05:22:32.188384", "status": "completed"} tags=[]
#dropping irrelevant columns
zomato=df.drop(['url','address','phone','dish_liked', 'menu_item', 'reviews_list'],axis=1)
# + papermill={"duration": 0.180424, "end_time": "2021-12-05T05:22:32.463902", "exception": false, "start_time": "2021-12-05T05:22:32.283478", "status": "completed"} tags=[]
#Removing the Duplicates
zomato.duplicated().sum()
zomato.drop_duplicates(inplace=True)
# + papermill={"duration": 0.225961, "end_time": "2021-12-05T05:22:32.729708", "exception": false, "start_time": "2021-12-05T05:22:32.503747", "status": "completed"} tags=[]
#Remove the NaN values from the dataset
zomato.isnull().sum()
zomato.dropna(how='any',inplace=True)
zomato.info()
# + papermill={"duration": 0.056654, "end_time": "2021-12-05T05:22:32.829311", "exception": false, "start_time": "2021-12-05T05:22:32.772657", "status": "completed"} tags=[]
#Changing the Columns Names
zomato.columns
zomato = zomato.rename(columns={'approx_cost(for two people)':'cost','listed_in(type)':'type',
'listed_in(city)':'city', 'rate':'rating'})
zomato.columns
# + papermill={"duration": 0.13473, "end_time": "2021-12-05T05:22:33.005042", "exception": false, "start_time": "2021-12-05T05:22:32.870312", "status": "completed"} tags=[]
#Some Transformations
zomato['cost'] = zomato['cost'].astype(str)
zomato['cost'] = zomato['cost'].apply(lambda x: x.replace(',','.'))
zomato['cost'] = zomato['cost'].astype(float)
zomato.info()
# + papermill={"duration": 0.148383, "end_time": "2021-12-05T05:22:33.194732", "exception": false, "start_time": "2021-12-05T05:22:33.046349", "status": "completed"} tags=[]
#Removing '/5' from Rates
zomato['rating'].unique()
zomato = zomato.loc[zomato.rating !='NEW']
zomato = zomato.loc[zomato.rating !='-'].reset_index(drop=True)
remove_slash = lambda x: x.replace('/5', '') if type(x) == np.str else x
zomato.rating = zomato.rating.apply(remove_slash).str.strip().astype('float')
zomato['rating'].head()
# + papermill={"duration": 0.120594, "end_time": "2021-12-05T05:22:33.356359", "exception": false, "start_time": "2021-12-05T05:22:33.235765", "status": "completed"} tags=[]
# Adjust the column names
zomato.name = zomato.name.apply(lambda x:x.title())
zomato.online_order.replace(('Yes','No'),(True, False),inplace=True)
zomato.book_table.replace(('Yes','No'),(True, False),inplace=True)
zomato.head()
# + [markdown] papermill={"duration": 0.042736, "end_time": "2021-12-05T05:22:33.442108", "exception": false, "start_time": "2021-12-05T05:22:33.399372", "status": "completed"} tags=[]
# # **Visualization**
# + [markdown] papermill={"duration": 0.041826, "end_time": "2021-12-05T05:22:33.526271", "exception": false, "start_time": "2021-12-05T05:22:33.484445", "status": "completed"} tags=[]
# **Top 10 most popular restaurant types**
# + papermill={"duration": 0.39546, "end_time": "2021-12-05T05:22:33.964325", "exception": false, "start_time": "2021-12-05T05:22:33.568865", "status": "completed"} tags=[]
from collections import Counter
r_type=zomato['rest_type'].value_counts()[:10]
sns.barplot(x=r_type,y=r_type.index)
plt.title("Most In-Demand Restaurant Types")
plt.xlabel("count")
# + [markdown] papermill={"duration": 0.044494, "end_time": "2021-12-05T05:22:34.054832", "exception": false, "start_time": "2021-12-05T05:22:34.010338", "status": "completed"} tags=[]
# **Proportion of Restaurants that provide online service**
# + papermill={"duration": 0.163913, "end_time": "2021-12-05T05:22:34.261831", "exception": false, "start_time": "2021-12-05T05:22:34.097918", "status": "completed"} tags=[]
zomato.online_order.value_counts().plot(kind='pie')
w_circle=plt.Circle((0,0),0.7,color='white')
p=plt.gcf()
p.gca().add_artist(w_circle)
# + [markdown] papermill={"duration": 0.044457, "end_time": "2021-12-05T05:22:34.350926", "exception": false, "start_time": "2021-12-05T05:22:34.306469", "status": "completed"} tags=[]
# **Proportion of restaurants that allow booking tables**
# + papermill={"duration": 0.144284, "end_time": "2021-12-05T05:22:34.540388", "exception": false, "start_time": "2021-12-05T05:22:34.396104", "status": "completed"} tags=[]
zomato.book_table.value_counts().plot(kind='pie')
w_circle=plt.Circle((0,0),0.7,color='white')
p=plt.gcf()
p.gca().add_artist(w_circle)
# + [markdown] papermill={"duration": 0.045271, "end_time": "2021-12-05T05:22:34.631453", "exception": false, "start_time": "2021-12-05T05:22:34.586182", "status": "completed"} tags=[]
# **Most popular restaurant chains**
# + papermill={"duration": 0.312259, "end_time": "2021-12-05T05:22:34.991121", "exception": false, "start_time": "2021-12-05T05:22:34.678862", "status": "completed"} tags=[]
ax=df.name.value_counts()[:10].plot(kind='bar')
plt.xlabel("Restaurant Name")
plt.ylabel("No. of branches")
plt.title("Top 10 Chain Restaurants")
# + [markdown] papermill={"duration": 0.047268, "end_time": "2021-12-05T05:22:35.085322", "exception": false, "start_time": "2021-12-05T05:22:35.038054", "status": "completed"} tags=[]
# **Impact of Online Ordering on Ratings**
#
# It can be seen from the chart below that restaurants that allow ordering online tend to have higher ratings
# + papermill={"duration": 0.738599, "end_time": "2021-12-05T05:22:35.871371", "exception": false, "start_time": "2021-12-05T05:22:35.132772", "status": "completed"} tags=[]
sns.set_style('white')
y=pd.crosstab(zomato.rating,zomato.online_order)
y.plot(kind='bar',stacked=True)
# + [markdown] papermill={"duration": 0.047834, "end_time": "2021-12-05T05:22:35.967491", "exception": false, "start_time": "2021-12-05T05:22:35.919657", "status": "completed"} tags=[]
# **Impact of Table Booking on Ratings**
#
# It can be seen from the chart below that restaurants that allow table bookings tend to have higher ratings.
# + papermill={"duration": 0.900435, "end_time": "2021-12-05T05:22:36.915538", "exception": false, "start_time": "2021-12-05T05:22:36.015103", "status": "completed"} tags=[]
sns.set_style('white')
y=pd.crosstab(zomato.rating,zomato.book_table)
y.plot(kind='bar',stacked=True)
# + [markdown] papermill={"duration": 0.049741, "end_time": "2021-12-05T05:22:37.014670", "exception": false, "start_time": "2021-12-05T05:22:36.964929", "status": "completed"} tags=[]
# **Top 10 retaurant locations**
# + papermill={"duration": 0.357128, "end_time": "2021-12-05T05:22:37.421072", "exception": false, "start_time": "2021-12-05T05:22:37.063944", "status": "completed"} tags=[]
sns.set_style('darkgrid')
ch=zomato['location'].value_counts()[:10]
sns.barplot(x=ch,y=ch.index,palette='viridis')
plt.xlabel('Number of Outlets')
plt.ylabel('Location')
plt.show()
# + [markdown] papermill={"duration": 0.049225, "end_time": "2021-12-05T05:22:37.519675", "exception": false, "start_time": "2021-12-05T05:22:37.470450", "status": "completed"} tags=[]
# # **Predictive Models**
#
# We now proceed to build a model that can predict ratings for new restaurants on the market to enable informed decision making by the stakeholders. The first step for this is to encode the columns containing string types into some sort of code.
# + papermill={"duration": 0.097007, "end_time": "2021-12-05T05:22:37.666147", "exception": false, "start_time": "2021-12-05T05:22:37.569140", "status": "completed"} tags=[]
def Encode(zomato):
for column in zomato.columns[~zomato.columns.isin(['rating', 'cost', 'votes'])]:
zomato[column] = zomato[column].factorize()[0]
return zomato
encodedZomato = Encode(zomato.copy())
# + [markdown] papermill={"duration": 0.050282, "end_time": "2021-12-05T05:22:37.765980", "exception": false, "start_time": "2021-12-05T05:22:37.715698", "status": "completed"} tags=[]
# Next we try to plot a heatmap to get the correlation between the various available variables
# + papermill={"duration": 1.457853, "end_time": "2021-12-05T05:22:39.273908", "exception": false, "start_time": "2021-12-05T05:22:37.816055", "status": "completed"} tags=[]
#Get Correlation between different variables
corr = encodedZomato.corr(method='kendall')
plt.figure(figsize=(15,8))
sns.heatmap(corr, annot=True)
# + [markdown] papermill={"duration": 0.05301, "end_time": "2021-12-05T05:22:39.381033", "exception": false, "start_time": "2021-12-05T05:22:39.328023", "status": "completed"} tags=[]
# ## **Splitting the Dataset**
#
# Next up is the very important step of the deciding the train-test split for the different models. In this, after careful consideration, a train-test split of 4:1 is chosen. Also, the dependent and independent variables are declared for the models. The target variable here is rating which is what we aim to predict for future restaurants.
# + papermill={"duration": 0.080299, "end_time": "2021-12-05T05:22:39.514902", "exception": false, "start_time": "2021-12-05T05:22:39.434603", "status": "completed"} tags=[]
#Defining X and Y for model, Train-test split
x = encodedZomato[["online_order","book_table","votes","location","rest_type","cuisines","cost","type"]]
y = encodedZomato['rating']
#Getting Test and Training Set
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=.2,random_state=353)
x_train.shape, y_train.shape
# + [markdown] papermill={"duration": 0.053063, "end_time": "2021-12-05T05:22:39.621175", "exception": false, "start_time": "2021-12-05T05:22:39.568112", "status": "completed"} tags=[]
# ## **Extra Tree Regressor**
#
# Extra Trees Regressor is an ensemble learning method fundamentally based on decision trees. Extra Trees Regressor, like RandomForest, randomizes certain decisions and subsets of data to minimize over-learning from the data and overfitting
#
# For more details, visit https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.ExtraTreesRegressor
# + papermill={"duration": 10.783061, "end_time": "2021-12-05T05:22:50.457874", "exception": false, "start_time": "2021-12-05T05:22:39.674813", "status": "completed"} tags=[]
#Extra Tree Regression
from sklearn.ensemble import ExtraTreesRegressor
ETree=ExtraTreesRegressor(n_estimators = 200)
ETree.fit(x_train,y_train)
y_predict=ETree.predict(x_test)
r2_score(y_test,y_predict)
# + [markdown] papermill={"duration": 0.055239, "end_time": "2021-12-05T05:22:50.567311", "exception": false, "start_time": "2021-12-05T05:22:50.512072", "status": "completed"} tags=[]
# ## **Decision Tree Regressor**
#
# Decision Trees (DTs) are a non-parametric supervised learning method used for classification and regression. The goal is to create a model that predicts the value of a target variable by learning simple decision rules inferred from the data features. A tree can be seen as a piecewise constant approximation.
#
# For more details, visit https://scikit-learn.org/stable/modules/tree.html
#
# also, https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeRegressor
# + papermill={"duration": 0.214442, "end_time": "2021-12-05T05:22:50.836374", "exception": false, "start_time": "2021-12-05T05:22:50.621932", "status": "completed"} tags=[]
#DTree Regression
from sklearn.tree import DecisionTreeRegressor
DTree=DecisionTreeRegressor(min_samples_leaf=.00001)
DTree.fit(x_train,y_train)
y_predict=DTree.predict(x_test)
r2_score(y_test,y_predict)
# + [markdown] papermill={"duration": 0.055056, "end_time": "2021-12-05T05:22:50.949740", "exception": false, "start_time": "2021-12-05T05:22:50.894684", "status": "completed"} tags=[]
# ## **Linear Regression Model**
#
# In statistics, linear regression is a linear approach for modelling the relationship between a scalar response and one or more explanatory variables. The case of one explanatory variable is called simple linear regression.
#
# Visit: https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression
# + papermill={"duration": 0.08989, "end_time": "2021-12-05T05:22:51.096239", "exception": false, "start_time": "2021-12-05T05:22:51.006349", "status": "completed"} tags=[]
#Some LinReg Model
reg=LinearRegression()
reg.fit(x_train,y_train)
y_pred=reg.predict(x_test)
r2_score(y_test,y_pred)
# + [markdown] papermill={"duration": 0.057461, "end_time": "2021-12-05T05:22:51.257723", "exception": false, "start_time": "2021-12-05T05:22:51.200262", "status": "completed"} tags=[]
# ## **Gradient Boosting Regressor**
#
# GB builds an additive model in a forward stage-wise fashion; it allows for the optimization of arbitrary differentiable loss functions. In each stage a regression tree is fit on the negative gradient of the given loss function.
#
# For more details, visit https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingRegressor
# + papermill={"duration": 3.011628, "end_time": "2021-12-05T05:22:54.323669", "exception": false, "start_time": "2021-12-05T05:22:51.312041", "status": "completed"} tags=[]
from sklearn.ensemble import GradientBoostingRegressor
gbr = GradientBoostingRegressor()
gbr.fit(x_train, y_train)
y_predict=gbr.predict(x_test)
r2_score(y_test,y_predict)
# + [markdown] papermill={"duration": 0.053679, "end_time": "2021-12-05T05:22:54.432101", "exception": false, "start_time": "2021-12-05T05:22:54.378422", "status": "completed"} tags=[]
# ## **XGBoost Regressor**
#
# XGBoost is a decision-tree-based ensemble Machine Learning algorithm that uses a gradient boosting framework. In prediction problems involving unstructured data (images, text, etc.) artificial neural networks tend to outperform all other algorithms or frameworks. However, when it comes to small-to-medium structured/tabular data, decision tree based algorithms are considered best-in-class right now.
# Visit https://xgboost.readthedocs.io/en/latest/python/python_api.html?highlight=xgbregressor#xgboost.XGBRegressor to learn more
# + papermill={"duration": 13.193953, "end_time": "2021-12-05T05:23:07.679946", "exception": false, "start_time": "2021-12-05T05:22:54.485993", "status": "completed"} tags=[]
from xgboost import XGBRegressor
xgb = XGBRegressor(booster = 'gbtree', learning_rate = 0.1, max_depth = 15, n_estimators = 200)
xgb.fit(x_train, y_train)
y_predict=xgb.predict(x_test)
r2_score(y_test,y_predict)
# + [markdown] papermill={"duration": 0.05412, "end_time": "2021-12-05T05:23:07.789939", "exception": false, "start_time": "2021-12-05T05:23:07.735819", "status": "completed"} tags=[]
# ## **Random Forest Regressor**
#
# A random forest is a meta estimator that fits a number of classifying decision trees on various sub-samples of the dataset and uses averaging to improve the predictive accuracy and control over-fitting.
#
# Visit https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestRegressor to learn more
# + papermill={"duration": 19.52206, "end_time": "2021-12-05T05:23:27.367566", "exception": false, "start_time": "2021-12-05T05:23:07.845506", "status": "completed"} tags=[]
from sklearn.ensemble import RandomForestRegressor
# tuning=dict()
rf=RandomForestRegressor(n_estimators=200,random_state=50,min_samples_leaf=.00001)
rf.fit(x_train,y_train)
y_pred=rf.predict(x_test)
r2_score(y_test,y_pred)
# + [markdown] papermill={"duration": 0.054154, "end_time": "2021-12-05T05:23:27.476917", "exception": false, "start_time": "2021-12-05T05:23:27.422763", "status": "completed"} tags=[]
# Now let us compare and contrast the performance of each of our models by arranging their r-squared scores in a tabulated format
# + papermill={"duration": 1.305617, "end_time": "2021-12-05T05:23:28.837350", "exception": false, "start_time": "2021-12-05T05:23:27.531733", "status": "completed"} tags=[]
models = pd.DataFrame({
'Model' : ['Linear Regression', 'Decision Tree', 'Random Forest','Extra Tree Regressor', 'Gradient Boost', 'XgBoost'],
'Score' : [reg.score(x_test, y_test), DTree.score(x_test, y_test), rf.score(x_test, y_test), ETree.score(x_test, y_test),
gbr.score(x_test, y_test), xgb.score(x_test, y_test)]
})
models.sort_values(by = 'Score', ascending = False)
# + [markdown] papermill={"duration": 0.055638, "end_time": "2021-12-05T05:23:28.949609", "exception": false, "start_time": "2021-12-05T05:23:28.893971", "status": "completed"} tags=[]
# # **Inferences**
#
# It is clearly visible that the XGBoost regressor works the best with our dataset, which means that we'll be using this model to predict the ratings of the our future dataset.
# + [markdown] papermill={"duration": 0.05592, "end_time": "2021-12-05T05:23:29.063796", "exception": false, "start_time": "2021-12-05T05:23:29.007876", "status": "completed"} tags=[]
# Now we need to use our model, the XGBoost regressor, to predict the ratings for new restaurants
# + [markdown] papermill={"duration": 0.056962, "end_time": "2021-12-05T05:23:29.177894", "exception": false, "start_time": "2021-12-05T05:23:29.120932", "status": "completed"} tags=[]
# # **Predicting Ratings for New Restaurants**
# + papermill={"duration": 0.095239, "end_time": "2021-12-05T05:23:29.329421", "exception": false, "start_time": "2021-12-05T05:23:29.234182", "status": "completed"} tags=[]
df_new = pd.read_csv('../input/zomato-prediction/Zomato_predictions.csv')
#Converting cost to float
df_new['cost'] = df_new['cost'].astype(float)
df_new
# + papermill={"duration": 0.080185, "end_time": "2021-12-05T05:23:29.466661", "exception": false, "start_time": "2021-12-05T05:23:29.386476", "status": "completed"} tags=[]
zomatoNew = Encode(df_new.copy())
zomatoNew
# + papermill={"duration": 0.084532, "end_time": "2021-12-05T05:23:29.608111", "exception": false, "start_time": "2021-12-05T05:23:29.523579", "status": "completed"} tags=[]
x_pred = zomatoNew[["online_order","book_table","votes","location","rest_type","cuisines","cost","type"]]
y_predict=xgb.predict(x_pred)
y_pred_list = [ round(elem, 1) for elem in y_predict ]
y_pred_list
# + papermill={"duration": 0.08558, "end_time": "2021-12-05T05:23:29.752451", "exception": false, "start_time": "2021-12-05T05:23:29.666871", "status": "completed"} tags=[]
df_new["rating"] = y_pred_list
df_new
# + papermill={"duration": 0.073273, "end_time": "2021-12-05T05:23:29.884640", "exception": false, "start_time": "2021-12-05T05:23:29.811367", "status": "completed"} tags=[]
df_new.to_csv("../working/zomato_predicted_ratings.csv", index = False)
# + [markdown] papermill={"duration": 0.061174, "end_time": "2021-12-05T05:23:30.006408", "exception": false, "start_time": "2021-12-05T05:23:29.945234", "status": "completed"} tags=[]
# # **Conclusions**
#
# We have successfully trained and built multiple models on the dataset. We also found that the XGBoost regressor works incredibly well for this data. This analysis shows that once the model has been trained and tested on the data, we can actually predict the ratings for new restaurants as well with the independent variables being available to us.
| Source/zomato-success-factors-rating-predictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Beamprofile (Figure 7)
# +
# Import modules
import sys, h5py
import numpy as np
import scipy as sp
# %matplotlib inline
# Import modules from src directory
sys.path.append("../src")
import plotting
# -
# Physical constants
h = 6.62606957e-34 #Js
c = 299792458 #m/s
hc = h*c #Jm
# Lorentzian
lorentz1d = lambda x, I, gamma: (I * gamma**2) / (x**2 + gamma**2)
# ### 1. Loading data from file
# Load size/intensity distribution
results_file = '../meta/results.h5'
with h5py.File(results_file, 'r') as f:
diameter = f['diameter'][:] # nm
intensity = f['intensity'][:] # mJ/um2
# Rescale intensity
wavelength = 0.2262 #nm
mJtoPh = lambda i: ((i / 1000.) * wavelength*1e-9) / (hc) * 1e-12
# ### 2. Invert intensity data (35 - 45 nm) to get beam profile in the center
# +
# Select size range
size_min, size_max = (35,45)
intensity_profile = intensity[(diameter > size_min) & (diameter < size_max)]
diameter_profile = diameter[(diameter > size_min) & (diameter < size_max)]
# Model for beam profile
Ebefore = 3.29 # mJ
transmission = 0.15
s = np.argsort(intensity_profile)[::-1]
pintensity_center = intensity_profile[s]
r_center = np.sqrt((float(Ebefore*transmission) / np.sum(intensity_profile) / np.pi)*np.arange(0,intensity_profile.shape[0]+1))
fwhm_center = r_center[np.argmin(np.abs(pintensity_center - (pintensity_center[0]/2)))]*2 * 1000. # in nm
# Mirror the profile
xmirrored_center = np.hstack([-r_center[:-1][::-1],r_center[:-1]])
ymirrored_center = mJtoPh(np.hstack([pintensity_center[::-1], pintensity_center])) # in Nr. Photons per um2
# Intensity histogram
hist,edges = np.histogram(mJtoPh(intensity_profile), range=(0,0.5), bins=50)
# -
# ### 3. Invert intensity data (235 - 300 nm) to get beam profile in the tails
# +
# Select size range
size_min, size_max = (235,300)
intensity_profile = intensity[(diameter > size_min) & (diameter < size_max)]
diameter_profile = diameter[(diameter > size_min) & (diameter < size_max)]
# Model for beam profile
Ebefore = 3.29 # mJ
transmission = 0.15
s = np.argsort(intensity_profile)[::-1]
pintensity_tails = intensity_profile[s]
r_tails = np.sqrt((float(Ebefore*transmission) / np.sum(intensity_profile) / np.pi)*np.arange(0,intensity_profile.shape[0]+1))
fwhm_tails = r_tails[np.argmin(np.abs(pintensity_tails - (pintensity_tails[0]/2)))]*2 * 1000. # in nm
# Mirror the profile
xmirrored_tails = np.hstack([-r_tails[:-1][::-1],r_tails[:-1]])
ymirrored_tails = mJtoPh(np.hstack([pintensity_tails[::-1], pintensity_tails])) # in Nr. Photons per um2
# Intensity histogram
hist,edges = np.histogram(mJtoPh(intensity_profile), range=(0,0.5), bins=50)
# +
# Combined profile
select_tails = r_tails[:-1] > 0.68
select_center = r_center[:-1] < 0.7
pintensity_combined = np.hstack([pintensity_center[select_center], pintensity_tails[select_tails]])
# Do normalization on the combined profile
Ebefore = 3.29 # mJ
transmission = 0.2
r_combined = np.sqrt((float(Ebefore*transmission) / np.sum(pintensity_combined) / np.pi)*np.arange(pintensity_combined.shape[0]+1))
fwhm_combined = r_combined[np.argmin(np.abs(pintensity_combined - (pintensity_combined[0]/2)))]*2 * 1000. # in nm
#print fwhm_combined, 2*r_combined.max()
# Mirror the profile
xmirrored_combined = np.hstack([-r_combined[:-1][::-1],r_combined[:-1]])
ymirrored_combined = mJtoPh(np.hstack([pintensity_combined[::-1], pintensity_combined])) # in Nr. Photons per um2
# +
# Resample the combined profile on a regular grid
dx = 1e-3
x = np.arange(xmirrored_combined.min(), xmirrored_combined.max(), dx)
f = sp.interpolate.interp1d(xmirrored_combined, ymirrored_combined)
y = f(x)
# Fit single lorentzian to initial beam reconstruction
popt, pcov = sp.optimize.curve_fit(lorentz1d, x, y)
Ifit ,gammafit = popt
lfit = lorentz1d(x, Ifit, gammafit)
# -
# ### 3. Plotting
# +
plot = plotting.Plot(rows=1, cols=2, aspect=1, fontsize=10, legend=True, legend_frameon=False, legend_location=2,save_pdf=True)
plot.xlabel = [r'Distance to beam axis [arb. units]', r'Distance to beam axis [$\mu$m]']
plot.ylabel = 2*[r'Photon intensity [$10^{12}$ photons /$\mu$m$^2$]']
plot.title_label = ['', '']
# Plotting the beam profile
plot.plotting_traces(0, [xmirrored_center, xmirrored_tails],
[ymirrored_center, ymirrored_tails],
[' 35 - 45 nm particles', '235 - 300 nm particles'],
colors=['g', 'b'], mecs=['g', 'b'], mfcs=['g', 'b'], linestyles=['None', 'None'],
linewidths=[0,0,], markers=['.','.'], markersize=3, logy=False,
ylim=[0,1.3], xlim=[-2.5, 2.5])
plot.axes[0].tick_params(axis='y', which='both', right='off', left='on', labelright='off', labelleft='on')
#direction='inout', pad=20)
plot.axes[0].tick_params(axis='x', which='both', bottom='on', top='off', labelbottom='on', labeltop='off')
plot.axes[0].yaxis.set_label_position("left")
plot.axes[0].xaxis.labelpad = 7
plot.axes[0].spines['top'].set_visible(False)
plot.axes[0].spines['right'].set_visible(False)
plot.axes[0].text(-2.5,1.45, '(a)', va='top', ha='center', fontsize=10, color='k')
plot.axes[0].set_yticks([0,0.2,0.4,0.6,0.8,1.,1.2])
# Plotting the beam profile
plot.plotting_traces(1, [xmirrored_combined], [ymirrored_combined],
['Combined reconstruction'],
colors=['r'], mecs=['r'], mfcs=['r'], linestyles=['None'],
linewidths=[0], markers=['.'], markersize=3, logy=False,
ylim=[0,1.3], xlim=[-2., 2.])
plot.axes[1].tick_params(axis='y', which='both', right='on', left='off', labelright='on', labelleft='off')
plot.axes[1].tick_params(axis='x', which='both', bottom='on', top='off', labelbottom='on', labeltop='off')
plot.axes[1].yaxis.set_label_position("right")
plot.axes[1].xaxis.labelpad = 7
plot.axes[1].spines['top'].set_visible(False)
plot.axes[1].spines['left'].set_visible(False)
plot.axes[1].text(-2.,1.45, '(b)', va='top', ha='left', fontsize=10, color='k')
plot.axes[1].set_xticks([-2,-1,0,1,2])
plot.axes[1].set_yticks([0,0.2,0.4,0.6,0.8,1.,1.2])
plot.plotting_traces(1, [x], [lfit],
['Lorentzian fit'],
colors=['0.5'], mecs=['0.5'], mfcs=['0.5'], linestyles=['-'],
linewidths=[1], markersize=1, logy=False,
ylim=[0,1.3], xlim=[-2., 2])
plot.show()
# -
# **Figure 7.**
# Reconstruction of the average X-ray beam profile in the focus. Injected particles are assumed to sample this profile uniformly.
# (a) Independent reconstructions using intensities that correspond to the smallest detectable particle sizes ($35$--$45$~nm)
# and the largest observed particle sizes ($235$--$300$~nm).
# (b) Combined reconstruction using the blue tails and the green center from (a) with the x-axis being rescaled such that the
# integrated profile equals a pulse energy in the focus of $0.66$~mJ (based on $3.29$~mJ measured upstream of the optics and assuming 20 \% transmission).
# With this scale, the reconstructed profile has a full width at half the maximum (FWHM) of $522$ nm. A Lorentzian fit to the profile is shown in gray.
| ipynb/fig07_beamprofile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CS207 Project Group 9
# # Bambanta: Forward and Reverse Automatic Differentiation
#
# *****
# # I. Introduction
# The software implements **‘Automatic Differentiation’ (AD)**. This is a technique to computationally evaluate the derivative of a specified function. Importantly, AD is not the same as symbolic differentiation or numerical differentiation, and holds important advantages over both. Symbolic differentiation, which is equivalent to analytically solving differential equations, can find the exact solution (with machine precision), but is very computationally expensive, and so with very large functions can be infeasible. Numerical differentiation, which uses the finite-difference approximation method, is computationally efficient, but is ultimately only approximate, and can be subject to both rounding error and discretisation error, meaning that it cannot be perfectly accurate. Both of these ‘traditional’ methods of differentiation run into problems and inefficiencies when calculating higher derivatives and partial derivatives with respect to many inputs (which is an important component of gradient-based optimisation methods).
#
#
# Automatic differentiation solves all these problems as it is able to solve derivatives to machine precision with comparative computational efficiency. As a result, automatic differentiation has incredibly important applications; in its ‘reverse-mode’ (discussed below), it is the basis of back-propagation, a fundamental process in neural network machine learning algorithms - as such this technique is leveraged by open-source machine learning libraries such as TensorFlow. A result of its efficient accuracy and iterative method, AD is capable of algorithmic differentiation: Because of the fact that every computer program, from mathematical algorithms to web-pages, can be expressed as a sequence and combination of arithmetic operations and elementary functions, the derivative of any computer program can be found using automatic differentiation.
# # II. Background
# Automatic differentiation is essentially the iterative application of the chain-rule. As mentioned above, any function can be considered a sequence of basic arithmetic operations or elementary functions (addition, multiplication, division, subtraction, trigonometric functions, exponential, logarithms etc.) and so any function can be interpreted in the following way (albeit often less simply):
# $$y = f(x) = f(g(h((x)))$$
#
# This can be rewritten as:
# $$y = f(g(h(x0))) = f(g(x1)) = f(x2) = x3$$
#
# Often, this decomposition is represented as an acyclic, directed computational graph that illustrates the route from the base function x0 to y, as illustrated by the example below:
#
# $ x_0\rightarrow^{h(x)}x_1\rightarrow^{g(x)}x_2\rightarrow^{f(x)}x_3\rightarrow y $
#
#
#
# In forward mode, automatic differentiation works by decomposing the function into this structure, and working through each component function finding the derivative using the chain rule ‘inside out’. That is to say, dx0/dx is found first, following by dx1/dx and so on until dy/dx itself is found. All this requires initial values to be set for x0, and x0’.
#
#
# Reverse mode, however, works in the opposite direction; rather than finding the derivative of the most fundamental component, and then finding the derivative of parent expressions in terms of these children components recursively until the final gradient is found, reverse mode goes the other way. It finds the derivative of each ‘child’ function in terms of its parent function recursively until the basic level derivative is found, at which point the final gradient can be found.
#
#
# One way of achieving forward mode AD is to use dual numbers. These are an extension of real numbers, somewhat analogous to imaginary numbers, such that every number additionally contains a dual component, $\epsilon$, where $\epsilon^2$ = 0. Given any polynomial function (or, in fact, any analytic real function via its Taylor series), if we replace x with (x+x'$\epsilon$), we find that the function will become: f(x) + f'(x)$\epsilon$. This provides a routine to automatically compute the derivative of the function f(x), and so is used in forward AD.
#
# Sources: https://en.wikipedia.org/wiki/Automatic_differentiation,
# http://www.columbia.edu/~ahd2125/post/2015/12/5/
#
# # III. How to Use
# ## a) Installation
# We recommend installing our package in a virtual environment. Please ensure that a version of virtualenv is already installed in your machine, and follow the steps:
#
# ```
# virtualenv env
#
# source env/bin/activate
#
# pip install Bambanta
# ```
#
# Alternatively, `pip install Bambanta` can be called alone.
# ## b) Usage
# ### i) Importing
# Once installed, the package can be imported simply through:
# ```python
# from Bambanta import AutoDiff
# ```
from Bambanta import AutoDiff
# ### ii) Forward Mode
# ###### Univariate
# For a simple, univariate example, let's find the value of the derivate of $y=3x^2-4x$ at $x=3$.
#
# we create an instance of the AutoDiff object as the basic building block for the equation - in other words, a single value of the independent variable. This object can then be used with binary and unary mathematical operators to construct the full function being evaluated. For each operation, a new function value (fAD.val) and derivative value (fAD.der) are calculated, such that once all operations are complete for the function, the function object's 'der' attribute will be that function's derivative at the point specificied at AutoDiff object creation.
#
# **N.B.**: It is important to note that AutoDiff assumes that the object being initialised is elementary and as such will have a derivative value of 1. Users can pass different der values as an argument for the function if required.
# +
a = 3.0
x = AutoDiff.fAD(a)
y = 3*x**2 - 4*x
print(y.der)
print("- - - - - - - - - -")
# better way to return values:
print("val: ",y.get_val(),"der: ", y.get_jac())
# -
# More complex cases can be handled, for example:
#
# $$y = \frac{e^{2x}|1-\log{x}|}{\sin{x} - \cos{x}}$$
# +
x = AutoDiff.fAD(a)
y = (AutoDiff.exp(2*x)*abs(1-AutoDiff.log(x)))/(AutoDiff.sin(x) - AutoDiff.cos(x))
y.get_jac()
# -
# ##### Multivariate
# AutoDiff.fAD is designed to handle multivariate calculus as well. For this, the additional functions **create_f** and **stack_f** must be used.
#
# Create_f is used to set up a system of fAD objects, in the form of a list, for construction of a multivariable function, whilst stack_f can be used to take in multiple independent fAD objects and construct a new fAD object that combines these objects such that the values of all objects can be returned at once and the objects can be altered simultaneously.
#
# **AutoDiff.create_f - for creating multiple variables for a single function**
#
# Multiple variables can be passed to create_f **as a list** and it will return an object for each variable with the necessary set of partial derivative values. When finding the derivative of a function with more than one variable, these variables should _always_ be created together using create_f, as this will ensure each variable has the correct vector of partial derivative values. Moreover, variables from unrelated functions should not be created together with create. Passing both $a$ and $b$ into create_f will create partial derivatives of $a$ with respect to $b$ and $b$ with respect to $a$ - if this is not desired then $a$ and $b$ should be created separately. Note that just like the single-variable case, create_f will assume that each object being created is elementary and will assign a derivative value of 1 to each unless otherwise instructed by the user.
#
# **AutoDiff.stack_f - for creating a vector of independent functions**
#
# One can pass multiple fAD objects into stack_f **as a list** and it will return a single fAD object with each of the objects values and derivatives stored as a vector. This allows one to return the derivative of all objects at once, or to manipulate each function in the same way all at once. Calling the `val` attribute of the stacked AutoDiff object will return the value of each function at the specified point. Calling the `der` attribute will return the Jacobian of the derivatives.
#
# **N.B.** Creating a stack_f object for vector value functions allows each function within the vector to be altered uniformly and simultaneously (see below) - but only operations with scalars (i.e. not other fAD objects) are currently supported.
# For example, let's deal with the following example of a vector-valued function:
#
# $\begin{bmatrix} f_1 \\ f_2 \\ f_3 \end{bmatrix} = \begin{bmatrix} x^2+2y^2 \\ |x-\cos{y}| \\ x-\frac{y}{x-y} \end{bmatrix}$
#
# at the points:
#
# $x = 4$, $y = 6$
#
# Once we've found the values and derivatives for this vector valued function $F(x,y)$, let's find the values and derivatives of another vector valued function, $G$:
#
# $G(x,y) = 2F(x,y)+5$
# +
x, y = AutoDiff.create_f([4,6])
f1 = x**2 + 2*y**2
f2 = abs(x - AutoDiff.cos(y))
f3 = x - y/(x-y)
F = AutoDiff.stack_f([f1,f2,f3])
print('F function values: ', F.get_val())
print('F derivative values:\n', F.get_jac())
G = F*2+5
print("---------------------------")
print('G Function values: ', G.get_val())
print('G Derivative values:\n', G.get_jac())
# -
# ## iii) Usage: Reverse Mode
# AutoDiff can also implement reverse mode automatic differentiation through the rAD class. The usage is slightly different. Variables can be initialised in a similar way, however this time, multiple variables within one equation do not necessarily need to be defined at once using create_r (although this function still exists should one want to use it). Once the function has been defined, the user must explicitly state which variable is the 'outer' variable of the function using the method outer(). then, to find the gradient of the outer variable with respect to, for example, the variable x, one must call **x**.grad().
#
# This is because, unlike forward mode which determines values and derivatives in the same directions whilst implicitly traversing the function's computational graph, reverse mode traverses the computational graph in one direction, storing the connections between nodes as it does so, and in this first traversal determine function values. Then, to find function derivatives it must traverse the computational graph in the reverse direction. To travel in the reverse direction from outside inward, grad() starts at the variable with respect to which the function is being differentiated, and recursively travels through the computational tree until it reaches the variable declared as the outer variable. At this point, grad recursively applies the chain rule to find the gradient of the function.
#
#
# This is done by recursing through the from x (inner object) to y (outer object) and then calculating the derivative through the recursion.
#
# Stack_r does exist, but behaves differently to stack_f, and requires the functions (as Python functions) are passed in as arguments to allow for construction of vector-valued function. because objects can only be defined in the context of one function at a time, this function returns the values and derivatives for each function rather than a rAD object storing all functions themselves.
#
# Let's again find the value of the derivate of $y=3x^2-4x$ at $x=3$.
# +
a = 3
x = AutoDiff.rAD(a)
y = 3*x**2-4*x
y.outer()
x.grad()
x.get_grad()
# -
# More complex cases can be handled, for example:
#
# $$y = \frac{e^{2x}|1-\log{x}|}{\sin{x} - \cos{x}}$$
# +
x = AutoDiff.rAD(a)
y = AutoDiff.exp(2*x)*abs(1-AutoDiff.log(x))/(AutoDiff.sin(x) - AutoDiff.cos(x))
y.outer()
x.grad()
x.get_grad()
# -
# #### _Multivariate_
# Multivariate cases can also be handled. Again, let's find the derivative of:
#
# $\begin{bmatrix} f_1 \\ f_2 \\ f_3 \end{bmatrix} = \begin{bmatrix} x^2+2y^2 \\ |x-\cos{y}| \\ x-\frac{y}{x-y} \end{bmatrix}$
#
# at the points:
#
# $x = 4$, $y = 6$
#
# This time round, things are a bit more complicated, as gradients must be reset between the functions; the variables can only be defined in the context of one outer function at a time.
# +
x, y = AutoDiff.create_f([4,6])
x = AutoDiff.rAD(4)
y = AutoDiff.rAD(6)
f1 = x**2 + 2*y**2
f1.outer()
x.grad()
y.grad()
print(x.get_grad(),y.get_grad())
AutoDiff.reset_der((x,y))
f2 = abs(x - AutoDiff.cos(y))
f2.outer()
x.grad()
y.grad()
print(x.get_grad(),y.get_grad())
AutoDiff.reset_der((x,y))
f3 = x - y/(x-y)
f3.outer()
x.grad()
y.grad()
print(x.get_grad(),y.get_grad())
# -
# # IV. Software Organisation
# ## _a) Directory Structure_
# ```
# cs207-FinalProject\
# AutoDiff/
# __init__.py
# __pycache__.py
# AutoDiff.py
# test_AutoDiff.py
# docs/
# milestone1.ipynb
# milestone2.ipynb
# final.ipynb
# README.md
# requirement.txt
# setup.cfg
# ```
# ## _b) Modules_
# Within Bambanta are two Python modules (other than `__init__.py`):
#
# 1. AutoDiff.py: This contains all functional code for automatic differentiation. within this file there are two classes, each with two complementary functions for equation construction along with numerous mathematical functions:
# * fAD - Class object that is the basic building block for forward-mode differentiation. Stores function and derivative values and has overloaded mathematical methods to allow construction of mathematical functions.
# * create_f - Function that is used to create multiple fAD objects that are within the same function and thus contain partial derivatives with respect to each other.
# * stack_f - stacks multiple fAD objects, given as a list, into a single fAD object that represents a vector-valued function.
# * rAD - Class object that is the basic building block for reverse-mode differentiation. Stores function value and children lists that, when the grad() method is called, are defined iteratively through reverse mode. Also contains a function outer() which is required to define the final parent node of the function's computational graph.
# * create_r - analogous function to its forward-mode counterpart, though not strictly necessary for multivariate functions as create_f is for forward mode.
# * stack_r - analogous to forward mode counterpart, creates rAD object for multiple functions - unlike forward mode, the functions (as python functions or lambda functions) must be passed to stack alongside the rAD objects. returns function values and jacobian matrix of derivatives.
# * mathematical functions - designed to handle forward and reverse mode objects as well as standard numerics: includes sin, cos, tan, arcsin, arccos, arctan, sinh, cosh, tanh, exp, log, and sqrt.
# * mul_by_row - Allows multiplication of forward-mode autodiff object with 2-dimensional derivatives, used for generalised overloading of multiplicative magic methods.
# * reset_ders - reset derivative values of reverse mode objects so that they can be reused in new functions.
# 2. test_AutoDiff.py: This is the test-suite for AutoDiff.py. Tests for correct function of above classes and functions. Tests are run using pytest, with the assistance of numpy.testing functions `assert_array_equal` and `assert_array_almost_equal` for dealing with cases where values are subject to a degree of rounding error. The tests are linked with Travis CI, which provides continuous integration software testing, and Coveralls, which provides a code coverage service to ensure that all of the code is being tested.
# ## _c) Implementation_
# ### Class: fAD
# This class encapsulates the fundamental machinery of forward mode automatic differentiation, and is capable of dealing with both single and multi-variable cases.
#
# #### Dependencies
#
# - numpy (imported as np): used for numerous mathematical (e.g. trigonometric, logarithmic operations)
# - numbers: used to ensure user passes numerical values.
#
# #### Attributes (Data Structures)
# - val: array of floats (of size 1 or more)
# - Numeric values, indicating the value of each entry in the current AutoDiff instance. For cases with only one function, val will size 1. For vector-valued functions, val can be longer.
#
# - der: 2D array of floats
# - Values representing the derivative value(s) in the current AutoDiff instance. The returned 2D array can be thought of as the Jacobian of all functions and variables for the AutoDiff instance. Suppose there are m elementary variables constructing n functions, all stored within the AutoDiff instance, 'der' would be a n\*m array of elements, with the (i,j) entry representing the derivative value of the i-th AutoDiff with respect to the j-th elementary variable.
#
# #### Methods
# (The following demonstrations are for the case when there is only one value in self.val. When the fAD object is in higher dimension, storing the values in an array allows us to simply apply the computation to each entry.)
#
# 0. `__init__`:
# - arguments:
# - a list/array of fAD instances or numerics
# - sets self.val as a list of 'val' attributes of the input AD instances
# - combines the 'der' attributes of the input fAD instances as a 2D array and save as self.der
#
#
# 1. `__add__` & `__radd__`:
# - arguments:
# - self
# - other: a float, int, or fAD
# - returns:
# - if other is an AD -> a new fAD instance with new.val = self.val + other.val, new.der = self.der + other.der
# - if other is a numeric value -> a new fAD instance with new.val = self.val + other, new.der = self.der
#
#
# 2. `__sub__`
# - arguments:
# - self
# - other: a float, int, or fAD
# - returns:
# - if other is a fAD -> a new fAD instance with new.val = self.val - other.val, new.der = self.der - other.der
# - if other is a numeric value -> a new fAD instance with new.val = self.val - other, new.der = self.der
#
#
# 3. `__rsub__`
# - arguments:
# - self
# - other: a float, int, or fAD
# - returns:
# - if other is a fAD -> a new fAD instance with new.val = other.val - self.val, new.der = other.der - self.der
# - if other is a numeric value -> a new fAD instance with new.val = other - self.val, new.der = -self.der
#
#
# 4. `__mul__` & `__rmul__`
# - arguments:
# - self
# - other: a float, int, or fAD
# - returns:
# - if other is a fAD -> a new fAD instance with new.val = self.val \* other.val, new.der = self.val \* other.der + self.der \* other.val
# - if other is a numeric value -> a new fAD instance with new.val = self.val \* other, new.der = self.der \* other
#
#
# 5. `__truediv__`
# - arguments:
# - self
# - other: a float, int, or fAD
# - returns:
# - if other is a fAD -> a new fAD instance with new.val = self.val / other.val, new.der = self.der\/other.val-self.val\*other.der\/(other.val\*\*2)
# - if other is a numeric value -> a new fAD instance with new.val = self.val / other, new.der = self.der / other
# - raises:
# - ZeroDivisionError when other.val = 0 or other = 0
#
#
# 6. `__rtruediv__`
# - arguments:
# - self
# - other: a float, int, or fAD
# - returns:
# - if other is a fAD -> a new fAD instance with new.val = other.val / self.val, new.der = other.val / self.der-other.val\*self.der / (self.val\*\*2)
# - if other is a numeric value -> a new fAD instance with new.der = -other \* self.der / (self.val \*\*2)
# - raises:
# - ZeroDivisionError when self.val = 0
#
#
# 7. `__pow__`
# - arguments:
# - self
# - exp: a float, int, or fAD
# - returns:
# - if exp is a fAD -> a new fAD instance with new.val = self.val \*\* exp.val, new.der = (self.val\*\*exp.val) * (self.der\*exp.val /self.val + exp.der\*np.log(self.val))
# - if other is a numeric value -> a new fAD instance with new.val = self.val \*\* exp, new.der = exp*(self.val\*\*(exp-1))\*self.der
#
#
# 8. `__rpow__`
# - arguments:
# - self
# - base: a float, int, or fAD
# - returns:
# - if base is a fAD -> a new fAD instance with new.val = base.val\*\*self.val, new.der = (base.val\*\*self.val) * (base.der\*self.val /base.val + self.der\*np.log(base.val))
# - if base is a numeric value -> a new fAD instance with new.val = base\*\*self.val, new.der = np.log(base)\*(base\*\*self.val)\*self.der
#
#
# 9. `__neg__`
# - arguments:
# - self
# - returns:
# - a new fAD instance with new.val = -self.val, new.der = -self.der
#
#
# 10. `__abs__`
# - arguments:
# - self
# - returns:
# - a new fAD instance with new.val = abs(self.val), new.der = (self.val / abs(self.val)) \* self.der
#
#
#
# 11. `__eq__`
# - arguments:
# - self
# - other: a fAD instance
# - returns:
# - 'True' if self.val==other.val and self.der==other.der, 'False' otherwise
#
# 12. `__ne__`
# - arguments:
# - self
# - other: a fAD instance
# - returns:
# - 'False' if self.val==other.val and self.der==other.der, 'True' otherwise
#
# 12. `__str__`
# - arguments:
# - self
# - returns:
# - a string describing the value and derivatives of the current instance
#
#
# 13. `__len__`
# - arguments:
# - self
# - returns:
# - len(self.val); number of function values stored within the fAD object.
#
#
# 14. `__repr__`
# - arguments:
# - self
# - returns:
# - string describing fAD(self.val,self.der)
#
#
# 15. `get_val()`
# - arguments:
# - self
# - returns:
# - self.val formatted correctly
#
#
# 16. `get_jac()`
# - arguments:
# - self
# - returns:
# - self.der formatted correctly
#
# ### Class: rAD
# This class encapsulates the fundamental machinery of forward mode automatic differentiation, and is capable of dealing with both single and multi-variable cases.
#
# #### Dependencies
#
# - numpy (imported as np): used for numerous mathematical (e.g. trigonometric, logarithmic operations)
# - numbers: used to ensure user passes numerical values.
#
# #### Attributes (Data Structures)
# - val: array of floats (of size 1 or more)
# - Numeric values, indicating the value of each entry in the current AutoDiff instance. For cases with only one function, val will size 1. For vector-valued functions, val can be longer.
#
# #### Methods
# (The following demonstrations are for the case when there is only one value in self.val. When the AD object is in higher dimension, storing the values in an array allows us to simply apply the computation to each entry.)
#
# 0. `__init__`:
# - arguments:
# - a list/array of AD instances
# - sets self.val as a list of 'val' attributes of the input AD instances
# - creates empty list for children and sets derivative value to None
#
#
# 1. `grad`:
# - arguments:
# - self
# - returns:
# - gradient of outer object with respect to this object; calling this before variable.der / variable.get_der() will update derivatives for outer variable from None to its gradient with respect to this object.
#
# 2. `__add__` & `__radd__`:
# - arguments:
# - self
# - other: a float, int, or rAD object
# - returns:
# - if other is a rAD -> a new rAD instance with new.val = self.val + other.val
# - if other is a numeric value -> a new AD instance with new.val = self.val + other
# - appends to the children of self and other (if other is rAD) a tuple of weight = self.val, and the new rAD object.
#
#
# 3. `__sub__`
# - arguments:
# - self
# - other: a float, int, or rAD object
# - returns:
# - if other is a rAD -> a new rAD instance with new.val = self.val - other.val
# - if other is a numeric value -> a new AD instance with new.val = self.val - other
# - appends to the children of self and other (if other is rAD) a tuple of weight = self.val (times 1 and -1 respectively), and the new rAD object.
#
#
# 4. `__rsub__`
# - arguments:
# - self
# - other: a float, int, or rAD object
# - returns:
# - if other is a rAD -> a new rAD instance with new.val= other.val - self.val
# - if other is a numeric value -> a new AD instance with new.val = other - self.val
# - appends to the children of self and other (if other is rAD) a tuple of weight = self.val (times -1 and 1 respectively), and the new rAD object.
#
#
#
# 5. `__mul__` & `__rmul__`
# - arguments:
# - self
# - other: a float, int, or rAD
# - returns:
# - if other is a rAD -> a new rAD instance with new.val = self.val \* other.val,
# - appends (other.val, new value) to self's children and (self.val, new value) to other's children
# - if other is a numeric value -> a new rAD instance with new.val = self.val \* other, new.der = self.der \* other
# - appends (other\*self.value, new value) to self's children
#
#
# 6. `__truediv__`
# - arguments:
# - self
# - other: a float, int, or rAD
# - returns:
# - if other is a rAD -> a new rAD instance with new.val = self.val / other.val
# - appends (1/other.val, new value) to self's children and ((-self.val / other.val\*\*2),new value) to other's children
# - if other is a numeric value -> a new rAD instance with new.val = self.val / other, and (1/other, new value) is appended to self's children.
# - raises:
# - ZeroDivisionError when other.val = 0 or other = 0
#
#
# 7. `__rtruediv__`
# - arguments:
# - self
# - other: a float, int, or rAD
# - returns:
# - if other is a rAD -> a new rAD instance with new.val = other.val / self.val,
# - appends (-other.val / self.val\*\*2, new value) to self's children and (1/self.val, new value) to other's children
# - if other is a numeric value -> a new rAD instance and appends (-other/self.val\*\*2, new value) to self's children
# - raises:
# - ZeroDivisionError when self.val = 0
#
#
# 8. `__pow__`
# - arguments:
# - self
# - exp: a float, int, or rAD
# - returns:
# - if exp is a rAD -> a new rAD instance with new.val = self.val \*\* exp.val
# - appends (self.val\*\*(other.val-1)\*other.val, new value) to self's children and (self.val\*\*other.val\*np.log(self.val), new value) to other's children
# - if other is a numeric value -> a new rAD instance with new.val = self.val \*\* exp, and appends self's children with (self.val\*\*(other-1)\*other, new value)
#
#
# 9. `__rpow__`
# - arguments:
# - self
# - base: a float, int, or rAD
# - returns:
# - if base is a rAD -> a new rAD instance with new.val = base.val\*\*self.val
# - appends self's children with (other.val\*\*self.val\*np.log(other.val), new value) and other's children is appended with (other.val\*\*(self.val-1)\*self.val, new value)
# - if base is a numeric value -> a new rAD instance with new.val = base\*\*self.val, and self' children is appended with (other\*\*self.val\*np.log(other), new value)
#
#
# 10. `__neg__`
# - arguments:
# - self
# - returns:
# - a new AD instance with new.val = -self.val, and (-1,new value) is appended to self's children
#
# 11. `__abs__`
# - arguments:
# - self
# - returns:
# - a new AD instance with new.val = abs(self.val), self's children is appended with (self.val / abs(self.val), new value)
#
#
#
# 12. `__eq__`
# - arguments:
# - self
# - other: a rAD instance
# - returns:
# - 'True' if self.val==other.val and self.der==other.der, 'False' otherwise
#
# 13. `__ne__`
# - arguments:
# - self
# - other: a rAD instance
# - returns:
# - 'False' if self.val==other.val and self.der==other.der, 'True' otherwise
#
# 14. `__str__`
# - arguments:
# - self
# - returns:
# - a string describing the value and derivatives of the current instance
#
# 15. `get_val()`
# - arguments:
# - self
# - returns:
# - self.val formatted correctly
#
#
# 16. `get_grad()`
# - arguments:
# - self
# - returns:
# - gradient of self with respect to outer object
#
# 17. `outer`
# - arguments:
# - self
# - returns:
# - nothing
# - sets this object to be the outer variable for a function by setting self.der = 1 (derivative of self with respect to self = 1).
#
# ### Functions: create, stack, sin, cos, log, exp.
# #### Dependencies
#
# - numpy (imported as np): used for numerous mathematical (e.g. trigonometric operations)
# - numbers: used to ensure user passes numerical values.
# - math: used for logarithms with variable bases.
#
# The following functions exist outside the AutoDiff class:
#
# 1. `create_f`
# - allows the users to quickly create multiple fAD instances
# - arguments:
# - val: a list of values
# - der: optional, assigned derivative values
# - returns:
# - a list of fAD objects
#
# 1. `create_r`
# - allows the users to quickly create multiple fAD instances
# - arguments:
# - val: a list of values
# - returns:
# - a list of rAD objects
#
# 2. `stack_f`
# - allows users to stack multiple fAD instances into one high-dimentional fAD instance
# - arguments:
# - vals: a list of multiple fAD instances
# - returns:
# - one fAD object, with *val* = an array of *val*'s of the fAD instances in the argument fADs, and *der* = an array of *der*'s of the fAD instances in the argument
#
# 3. `stack_r`
# - allows users to stack multiple rAD instances into one high-dimentional rAD instance
# - arguments:
# - vals: a list of multiple rAD instances
# - functions: list of multiple functions to be combined
# - returns:
# - function values and jacobian matrix
#
# 3. `sin`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns sin(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `cos`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns cos(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
#
# 3. `tan`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns tan(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `arcsin`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns arcsin(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `arccos`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns arccos(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `arctan`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns arctan(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `sinh`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns sinh(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `cosh`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns cosh(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 3. `tanh`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns tanh(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 5. `exp`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns exp(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 6. `log`
# - arguments:
# - x (AutoDiff or numeric)
# - returns:
# - returns log(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 5. `sqrt`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns sqrt(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 5. `logistic`
# - arguments:
# - x (fAD, rAD or numeric)
# - returns:
# - returns logistic(x) as the appropriate object (fAD, rAD or numeric) with correct derivative/children as appropriate.
#
# 5. `mul_by_row`
# - arguments:
# - val: array of values
# - der: array of derivatives
# - returns:
# - performs row-wise multiplication for forward-mode autodiff objects, facilitating calculations with 2-dimensional derivatives.
#
# 5. `reset_der`
# - arguments:
# - rADs: single instance or array of rAD objects
# - returns:
# - nothing
# - resets children and derivative values for all rAD objects given.
# # _d) Future_
# The main improvement that could be made is to improve the user-friendliness and intuitiveness of the reverse mode. Due to the fact that the values and computational tree are created in one direction and the derivatives are determined by traversing the other direction, the usage is slightly counter-intuitive; the derivative of y with respect to x is an attribute of x, not y. An ideal usage might be that the derivative of y with respect to x is an attribute of x could be attained through calling something like y.grad(x).
#
# Exactly how this could be achieved is not immediately clear - whatever method used must allow the outer function, y, to access all associated variable objects. For example, consider the equation $y=x\cdot z$.
#
# One possibility is to augment all mathematical functions to also return a list of the involved objects. The `__mul__` method, for example, currently returns `rAD(val = self.val * other.val)`. The rAD object could accept an additional argument, say `object_list`, that contained pointers to the self and other rAD objects and objects with which they themselves have been linked (something like `rAD(val = self.val * other.val, object_list = [self,other, selfs_objects, others_objects])`, then, in the new rAD object's `__init__` constructor, these objects could be put into a container stored with the y object such that methods of y can now access both x and z objects (and any object to which they themselves have been linked). Calling y.grad(x) would lead grad to access x and its computational graph, and calculate the gradient from there.
| docs/Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # QCoDeS Example with Keysight 344xxA
# `344xxA` models of Keysight digital multimeters have similar QCoDeS drivers. In this tutorial, `Keysight_34465A` is chosen for showcasing the usage of the instrument.
#
# Note however that not every feature/parameter is available on all `344xxA` models. This, when possible, is reflected in the instantiated driver object. Also note that models like `34465A` have options (like `DIG` and `MEM`) that can either be enabled or disabled on a particular instrument; this also has impact on availability of some features/parameters and/or their settings. In general, refer to the instrument's manual for detailed information.
#
# __NOTE__: Beginning with firmware revision 3.0, the digitizing and advanced triggering option, referred to as "DIG", for models 34465A-DIG/34470A-DIG, is now standard.
#
# The driver does not cover all the features of the instrument. __At present, the driver only supports being used to measure DC Voltage__
#
# +
import time
import numpy
import qcodes
from qcodes.dataset.measurements import Measurement
from qcodes.dataset.plotting import plot_dataset
from qcodes.instrument_drivers.Keysight.Keysight_34465A_submodules import Keysight_34465A
# -
dmm = Keysight_34465A('dmm', 'TCPIP0::172.20.2.182::inst0::INSTR')
dmm.reset()
# ## Parameters and methods overview
#
# Here is an overview (not exhaustive) of the parameters and methods that are available in the driver.
#
# The driver is not only comprised of root-level parameters and methods but also contains submodules which logically group some functionality.
#
# * Measurements
# * `dmm.init_measurement()`
# * `dmm.fetch()`
# * `dmm.read()`
# * `dmm.volt` - immediately measure one (the present) voltage value
# * Time trace measurements
# * `dmm.timetrace` - covered in more detail below
# * `dmm.time_axis`
# * `dmm.timetrace_dt`
# * `dmm.timetrace_npts`
# * Range settings
# * `dmm.range`
# * `dmm.autorange`
# * `dmm.autorange_once()`
# * Triggering
# * `dmm.trigger.source`
# * `dmm.trigger.delay`
# * `dmm.trigger.auto_delay_enabled`
# * `dmm.trigger.count`
# * `dmm.trigger.slope`
# * `dmm.trigger.level`
# * `dmm.trigger.force()`
# * Sample settings
# * `dmm.sample.count`
# * `dmm.sample.source`
# * `dmm.sample.timer`
# * `dmm.sample.pretrigger_count`
# * Display control
# * `dmm.display.text`
# * `dmm.display.clear()`
# * `dmm.display.enabled`
# * Measurement type
# * `dmm.sense_function`
# * Measurement accuracy (NB: DC Voltage mode only)
# * `dmm.NPLC`
# * `dmm.resolution`
# * `dmm.line_frequency`
# * `dmm.aperture_mode`
# * `dmm.aperture_time`
# * `dmm.autozero`
#
# As an extra, let's print the readable snapshot of the instrument.
dmm.print_readable_snapshot(update=True)
# ## Single value reading
#
# If one simpy wants to measure a single voltage value right now, the convenient `volt` parameter can be used. This parameter will always return a voltage reading, even if the instrument is set up to measure something else than voltage. The instrument state is left untouched by getting this parameter.
#
# This is a convenience parameter that does not utilize the full power of the instrument. In the next two sections, we will present more prowerful and robust ways of performing measurements.
dmm.volt()
# ## Time trace type measurements
#
# For the commonly occuring case were one would like to measure N voltage points equidistantly spaced in time, the Keysight DMM driver offers the convenient `timetrace` parameter. It comes with three associated (helper) parameters:
#
# - `timetrace_dt` - the time spacing between the acquired points
# - `timetrace_npts` - the number of points
# - `time_axis` - the corresponding time axis (np.array, starts at 0)
# - `timetrace` - the array of voltage values
#
# Note that changing `timetrace_dt` and `timetrace_npts` does not change any corresponding setting on the instrument before `timetrace.get` is called. Once that happens, the state of the instrument is __temporarily__ switched into one compatible with performing a time trace measurement. This specifically means that we set
#
# - `trigger.count` to 1
# - `trigger.source` to "BUS" (internal trigger - __no other trigger can be used__)
# - `sample.timer` to `timetrace_dt`
# - `sample.count` to `timetrace_npts`
#
# After the acquisition has completed, the previous instrument settings are restored. This behaviour guarantees that calling `timetrace.get` always works, irrespective of instrument state. The only exception is if the specified `timetrace_dt` can not be realized with the present `dmm.NPLC` and/or `dmm.aperture_time` settings. If that is the case, the `timetrace.get` method will raise a `RuntimeError` with an instructive error message. The user may manually compare the value of `sample.timer_minimum` to the value of `timetrace_dt`.
# The usage is straightforward.
# +
meas = Measurement()
meas.register_parameter(dmm.timetrace)
dmm.NPLC(0.006)
dmm.timetrace_dt(0.02)
dmm.timetrace_npts(500)
print(f'Minimal allowable dt: {dmm.sample.timer_minimum()} s')
with meas.run() as datasaver:
datasaver.add_result((dmm.timetrace, dmm.timetrace()),
(dmm.time_axis, dmm.time_axis()))
time_trace_ds = datasaver.dataset
# -
axs, cbs = plot_dataset(time_trace_ds)
# ## General guide: Multivalue triggered measurements
# __NOTE__: Refer to the instrument manual for more information on how to perform measurements with the instrument; here, only the most basic and frequently used ones are demonstated.
#
# Measurements with the instrument are performed in the following way: the instrument's settings are set for a particular kind of measurement, then the measurement is started/initialized, then after all the data has been acquired, it is retrieved from the instrument. Below is an example of such a measurement.
# Use `range`, `autorange` parameters or `autorange_once` method to set the measurement range. Disabling autorange is recommended by the instrument's manual for speeding up the measurement.
dmm.autorange_once()
# In order to set up the accuracy of the measurements and related settings, set up `NPLC` or `aperture_*` parameters (if available).
dmm.aperture_mode('ON')
dmm.aperture_time(2e-5)
# Set up triggering mechanism. Note that trigger settings and methods are inside `trigger` submodule of the instrument driver. Here, we will use immediate triggering (the measurement is triggered uppon measurement initialization, that is when `init_measurement` is called) with 1 trigger without any delays. Consulm the instrument's manual for more information on various triggering options.
dmm.trigger.source('IMM')
dmm.trigger.count(1)
dmm.trigger.delay(0.0)
# Set up sampling settings. Note that sampling parameters and method are inside `sample` submodule of the instrument driver. Here, we set to measure 15 samples.
dmm.sample.count(15)
dmm.sample.pretrigger_count(0)
# We are also going to set the sample source to timer (not avaliable in all models) so that the instrument ensures that the samples are taken with fixed periods between them. The `timer` parameter allows to set the value of that fixed period. For simplicity, we are going to let the instrument deduce the minimum value of it according to the current instrument configuration (!) and set it.
dmm.sample.source('TIM')
dmm.sample.timer('MIN')
# It turns out that commands are executed faster when the display of the instrument is disabled or is displaying text. One of the further section expands on it. Here, we will just set the display to some text.
dmm.display.text('Example with 15 samples')
# In order to initiate the measurement, call `init_measurement` method of the driver. In the case of this example, the instrument will get into "wait-for-trigger" mode but because the trigger source is "immediate", the instrument will immediately start measuring the 15 samples.
dmm.init_measurement()
# The instrument is going to measure 15 samples and save them to its memory. Once the measurement is completed, we can call `fetch` method of the driver to retreive the acquired data.
# While the measurement is going, there are two things we can do. One is to `sleep` until the end of the measurement, and then call `fetch`. The other one is to call `fetch` immediately after the measurement has been initiated - this way the instrument will return the acquired data right when the measurement is finished. This sounds pretty useful, however there are two considerations to keep in mind: the instrument manual hints that calling fetching immediately after initiation may be slower than waiting for the measurement to complete; if the measurement takes longer than the VISA command timeout, the code may raise a VISA timeout exception while the measurement is properly running (e.g. waiting for a trigger). To overcome the latter problem, one can temporarily change the VISA timeout value during the data fetching:
# ```python
# new_timeout = old_timeout + n_samples * time_per_sample
# # where, n_samples == dmm.sample.count(),
# # and time_per_sample == dmm.sample.timer()
# with dmm.timeout.set_to(new_timeout):
# data = dmm.fetch()
# ```
# Assuming that we've just slept or waited enough for the measurement to complete, let's `fetch` the data from the instrument. Note that due to the nature of the `fetch` command of the instrument, one can fetch the same measured data more than once (until, for example, a new measurement has been initiated; refer to the instrument's manual for more information on this).
data = dmm.fetch()
data
# Note that there is also a `read` method. It's difference from `fetch` is that it also initiates the new measurement. Using `read` might be convenient for some cases while `init_measurement` + `fetch` definitely allow for more control.
# Since the measurement is finished, let's bring back the display to life.
dmm.display.clear()
# If needed, it is straightforward to calculate a vector of times when the acquired data points were measured; for example, like this:
n = dmm.sample.count()
t = dmm.sample.timer()
setpoints = numpy.linspace(0, n*t, n)
setpoints # in seconds
# Note that for this simple type of measurement with measurement times being equidistantly spaced, the `timetrace` parameter is preferred.
# ## Special values of some parameters
# Some parameters can be set to special values like `MIN`/`MAX`/`DEF` which usually mean minimum/maximum/default, respectively.
#
# In order to obtain the actual value of the parameter that gets set when setting it to one of these special values, just call the get method of the parameter.
# Find out what the maximum value of `sample_timer` can be
dmm.sample.timer('MAX')
dmm.sample.timer()
# Find out what the default value of `sample_timer` is
dmm.sample.timer('DEF')
dmm.sample.timer()
# Find out what the recommended minumum value of `sample_timer` is
dmm.sample.timer('MIN')
dmm.sample.timer()
# Alternatively, if available, use a conveniently implemented
# get-only parameter to find out the actual value,
# for example, for MIN value of `sample_timer` there is such
# a convenient get-only parameter:
dmm.sample.timer_minimum()
# ## Display state impacts command execution speed
# To improve the execution speed of commands on the instrument, the user may disable the display's updates. When the display is not updating, it may show a static message of up to 40 characters. The `timetrace` parameter (see above) makes use of this functionality to execute faster and at the same time display a message that a measurement is in progress.
#
# The driver provides `display` submodule with `text` parameter that displays a given text on the insrturment, and a `clear` method that clears the text from display.
#
# The driver's `display` submodule also provides `enabled` parameter. When it is set to `False`, the state of the display is such that it does not show anything. Note, however, that displaying text is still possible when the `display.enabled` is `False` (when `display.enabled` is `False`, `display.clear` clears the text from the screen but does not enable it).
# Displays the text
dmm.display.text('Hello, World!')
# Returns display to its normal state
dmm.display.clear()
# Note that a call to `display_clear` also updates
# the value of the `display_text` parameter:
assert dmm.display.text() == ''
# Display can also be cleared by setting
# `display_text` to an empty string
dmm.display.text('some text') # Displays some text
time.sleep(0.5)
dmm.display.text('') # Returns display to its normal state
# Disables the display, which makes it turn black
dmm.display.enabled(False)
# Shows some text on a disabled display
dmm.display.text("I'm disabled but still showing text")
# Enabling display in this state
# won't change what's being displayed
dmm.display.enabled(True)
# ... but since the display is now enabled,
# clearing the display will not only remove the text
# but also show all the normal display indicators.
dmm.display.clear()
# ## Error handling
# Use the following methods to read the error queue of the instrument. The instrument has an error queue of length up to 20 messages. The queue message retrieval is first-in-first-out.
# Retrieve the first (i.e. oldest) error message
# in the queue (and thereby remove from the queue)
dmm.error()
# +
# The entire queue can be flushed out
# using `flush_error_queue` method.
# Printing the messages in enabled by default
# and can be disables with the `verbose` kwarg.
# generate a few errors
for _ in range(3):
dmm.write('produce an error!')
# -
dmm.flush_error_queue(verbose=True)
| docs/examples/driver_examples/Qcodes example with Keysight 344xxA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Handwritten Digits recognization using Deep Learning
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
(X_train, y_train) , (X_test, y_test) = keras.datasets.mnist.load_data()
len(X_train)
len(X_test)
X_train[0].shape
X_train[0]
plt.matshow(X_train[0])
y_train[0]
X_train = X_train / 255
X_test = X_test / 255
X_train[0]
X_train_flattened = X_train.reshape(len(X_train), 28*28)
X_test_flattened = X_test.reshape(len(X_test), 28*28)
X_train_flattened.shape
X_train_flattened[0]
# +
model = keras.Sequential([
keras.layers.Dense(10, input_shape=(784,), activation='sigmoid')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(X_train_flattened, y_train, epochs=10)
# -
model.evaluate(X_test_flattened, y_test)
# <img src=digits_neural_network.jpg>
y_predicted = model.predict(X_test_flattened)
y_predicted[0]
plt.matshow(X_test[0])
np.argmax(y_predicted[0]) # return index of max
y_predicted_labels = [np.argmax(i) for i in y_predicted]
y_predicted_labels[:5]
cm = tf.math.confusion_matrix(labels=y_test,predictions=y_predicted_labels)
cm
import seaborn as sn
plt.figure(figsize = (10,7))
sn.heatmap(cm, annot=True, fmt='d')
plt.xlabel('Predicted')
plt.ylabel('Truth')
# ## Using hidden layer
# +
model = keras.Sequential([
keras.layers.Dense(100, input_shape=(784,), activation='relu'),
keras.layers.Dense(10, activation='sigmoid')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(X_train_flattened, y_train, epochs=7)
# -
model.evaluate(X_test_flattened,y_test)
# +
y_predicted = model.predict(X_test_flattened)
y_predicted_labels = [np.argmax(i) for i in y_predicted]
cm = tf.math.confusion_matrix(labels=y_test,predictions=y_predicted_labels)
plt.figure(figsize = (10,7))
sn.heatmap(cm, annot=True, fmt='d')
plt.xlabel('Predicted')
plt.ylabel('Truth')
# -
# ## Using Flatten layer so that we don't have to call .reshape on input dataset
#
# +
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(100, activation='relu'),
keras.layers.Dense(10, activation='sigmoid')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(X_train, y_train, epochs=10)
# -
model.evaluate(X_test,y_test)
| Deep Learning/4. Handwritten Digits recognization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # All decoders (except KF, NB, and ensemble) run with varying amounts of training data
# ## User Options
# Define what folder you're saving to
# save_folder=''
save_folder='/home/jglaser/Files/Neural_Decoding/Results/'
# Define what folder you're loading the files from
# load_folder=''
load_folder='/home/jglaser/Data/DecData/'
# Define what dataset you are using
# dataset='s1'
# dataset='m1'
dataset='hc'
# Define which decoder to run
run_wf=1
run_wc=0
run_rnn=0
run_dnn=0
run_gru=0
run_lstm=0
run_xgb=0
run_svr=0
# ## 1. Import Packages
#
# We import both standard packages, and functions from the accompanying .py files
# +
#Import standard packages
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import io
from scipy import stats
import pickle
import time
import sys
#Add the main folder to the path, so we have access to the files there.
#Note that if your working directory is not the Paper_code folder, you may need to manually specify the path to the main folder. For example: sys.path.append('/home/jglaser/GitProj/Neural_Decoding')
sys.path.append('..')
#Import function to get the covariate matrix that includes spike history from previous bins
from preprocessing_funcs import get_spikes_with_history
#Import metrics
from metrics import get_R2
from metrics import get_rho
#Import decoder functions
from decoders import WienerCascadeDecoder
from decoders import WienerFilterDecoder
from decoders import DenseNNDecoder
from decoders import SimpleRNNDecoder
from decoders import GRUDecoder
from decoders import LSTMDecoder
from decoders import XGBoostDecoder
from decoders import SVRDecoder
#Import Bayesian Optimization package
from bayes_opt import BayesianOptimization
# +
#Turn off deprecation warnings
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# -
# ## 2. Load Data
#
# The data that we load is in the format described below. We have another example script, "neural_preprocessing.py" that may be helpful towards putting the data in this format.
#
# Neural data should be a matrix of size "number of time bins" x "number of neurons", where each entry is the firing rate of a given neuron in a given time bin
#
# The output you are decoding should be a matrix of size "number of time bins" x "number of features you are decoding"
# +
if dataset=='s1':
with open(load_folder+'s1_test_data.pickle','rb') as f:
# neural_data,vels_binned,pos_binned,acc_binned=pickle.load(f,encoding='latin1')
neural_data,vels_binned,pos_binned,acc_binned=pickle.load(f)
if dataset=='m1':
with open(load_folder+'m1_test_data.pickle','rb') as f:
# neural_data,vels_binned,pos_binned,acc_binned=pickle.load(f,encoding='latin1')
neural_data,vels_binned,pos_binned,acc_binned=pickle.load(f)
if dataset=='hc':
with open(load_folder+'hc_test_data.pickle','rb') as f:
# neural_data,vels_binned,pos_binned,acc_binned=pickle.load(f,encoding='latin1')
neural_data,pos_binned=pickle.load(f)
# -
# ## 3. Preprocess Data
# ### 3A. User Inputs
# The user can define what time period to use spikes from (with respect to the output).
# +
if dataset=='s1':
bins_before=6 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=6 #How many bins of neural data after (and including) the output are used for decoding
if dataset=='m1':
bins_before=13 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=0 #How many bins of neural data after (and including) the output are used for decoding
if dataset=='hc':
bins_before=4 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=5 #How many bins of neural data after (and including) the output are used for decoding
# -
# ### 3B. Format Covariates
# #### Format Input Covariates
#Remove neurons with too few spikes in HC dataset
if dataset=='hc':
nd_sum=np.nansum(neural_data,axis=0)
rmv_nrn=np.where(nd_sum<100)
neural_data=np.delete(neural_data,rmv_nrn,1)
# +
# Format for recurrent neural networks (SimpleRNN, GRU, LSTM)
# Function to get the covariate matrix that includes spike history from previous bins
X=get_spikes_with_history(neural_data,bins_before,bins_after,bins_current)
# Format for Wiener Filter, Wiener Cascade, XGBoost, and Dense Neural Network
#Put in "flat" format, so each "neuron / time" is a single feature
X_flat=X.reshape(X.shape[0],(X.shape[1]*X.shape[2]))
# -
# #### Format Output Covariates
#Set decoding output
if dataset=='s1' or dataset=='m1':
y=vels_binned
if dataset=='hc':
y=pos_binned
# #### In HC dataset, remove time bins with no output (y value)
if dataset=='hc':
#Remove time bins with no output (y value)
rmv_time=np.where(np.isnan(y[:,0]) | np.isnan(y[:,1]))
X=np.delete(X,rmv_time,0)
X_flat=np.delete(X_flat,rmv_time,0)
y=np.delete(y,rmv_time,0)
# ### 3C. Define training/testing/validation sets
# The testing and validation sets stay consistent, and we vary the length of the training set before them.
# +
#Bin size
if dataset=='s1' or dataset=='m1':
dt=.05
if dataset=='hc':
dt=.2
if dataset=='hc':
#Size of sets
test_size=int(450/dt) #7.5 min
valid_size=test_size #validation size is the same as the test size
train_size_min=test_size #The minimum training size
#End indices
end_idx=np.int(X.shape[0]*.8) #End of test set
tr_end_idx=end_idx-test_size-valid_size #End of training set
#Range of sets
testing_range=[end_idx-test_size,end_idx] #Testing set (length of test_size, goes up until end_idx)
valid_range=[end_idx-test_size-valid_size,end_idx-test_size] #Validation set (length of valid_size, goes up until beginning of test set)
#Below is a list with the range of all training sets tested
training_range_all=[[tr_end_idx-1*train_size_min,tr_end_idx],[tr_end_idx-2*train_size_min,tr_end_idx],[tr_end_idx-3*train_size_min,tr_end_idx],[tr_end_idx-4*train_size_min,tr_end_idx],[tr_end_idx-5*train_size_min,tr_end_idx]]
if dataset=='s1':
#Size of sets
test_size=int(300/dt) #5 min
valid_size=test_size
train_size_min=int(60/dt) # 1 min, the minimum training size
#End indices
end_idx=np.int(X.shape[0]*.9)
tr_end_idx=end_idx-test_size-valid_size
#Range of sets
testing_range=[end_idx-test_size,end_idx] #Testing set (length of test_size, goes up until end_idx)
valid_range=[end_idx-test_size-valid_size,end_idx-test_size] #Validation set (length of valid_size, goes up until beginning of test set)
#Below is a list with the range of all training sets tested
training_range_all=[[tr_end_idx-1*train_size_min,tr_end_idx],[tr_end_idx-2*train_size_min,tr_end_idx],[tr_end_idx-3*train_size_min,tr_end_idx],[tr_end_idx-4*train_size_min,tr_end_idx],[tr_end_idx-5*train_size_min,tr_end_idx],[tr_end_idx-10*train_size_min,tr_end_idx],[tr_end_idx-20*train_size_min,tr_end_idx]]
if dataset=='m1':
#Size of sets
test_size=int(300/dt) #5 min
valid_size=test_size
train_size_min=int(60/dt) # 1 min, the minimum training size
#End indices
end_idx=np.int(X.shape[0]*1)
tr_end_idx=end_idx-test_size-valid_size
#Range of sets
testing_range=[end_idx-test_size,end_idx] #Testing set (length of test_size, goes up until end_idx)
valid_range=[end_idx-test_size-valid_size,end_idx-test_size] #Validation set (length of valid_size, goes up until beginning of test set)
#Below is a list with the range of all training sets tested
training_range_all=[[tr_end_idx-1*train_size_min,tr_end_idx],[tr_end_idx-2*train_size_min,tr_end_idx],[tr_end_idx-3*train_size_min,tr_end_idx],[tr_end_idx-4*train_size_min,tr_end_idx],[tr_end_idx-5*train_size_min,tr_end_idx],[tr_end_idx-10*train_size_min,tr_end_idx]]
num_folds=len(training_range_all) #Number of loops we'll do (I'm just calling it "folds" so I can keep old code that used CV folds)
# -
# ## 4. Run Decoders
# **Initialize lists of results**
# +
#R2 values
mean_r2_wf=np.empty(num_folds)
mean_r2_wc=np.empty(num_folds)
mean_r2_xgb=np.empty(num_folds)
mean_r2_dnn=np.empty(num_folds)
mean_r2_rnn=np.empty(num_folds)
mean_r2_gru=np.empty(num_folds)
mean_r2_lstm=np.empty(num_folds)
mean_r2_svr=np.empty(num_folds)
#Actual Data
y_test_all=[]
y_train_all=[]
y_valid_all=[]
#Test predictions
y_pred_wf_all=[]
y_pred_wc_all=[]
y_pred_xgb_all=[]
y_pred_dnn_all=[]
y_pred_rnn_all=[]
y_pred_gru_all=[]
y_pred_lstm_all=[]
y_pred_svr_all=[]
# -
# **In the following section, we**
# 1. Loop over varying amounts of training data
# 2. Extract the training/validation/testing data
# 3. Preprocess the data
# 4. Run the individual decoders (whichever have been specified in user options). This includes the hyperparameter optimization
# 5. Save the results
#
# +
t1=time.time()
num_examples=X.shape[0] #number of examples (rows in the X matrix)
for i in range(num_folds): #Loop through different amounts of training data
######### SPLIT DATA INTO TRAINING/TESTING/VALIDATION #########
#Note that all sets have a buffer of"bins_before" bins at the beginning, and "bins_after" bins at the end
#This makes it so that the different sets don't include overlapping neural data
#Testing set
testing_set=np.arange(testing_range[0]+bins_before,testing_range[1]-bins_after)
#Validation set
valid_set=np.arange(valid_range[0]+bins_before,valid_range[1]-bins_after)
#Training_set
training_range=training_range_all[i] #Get the training range for this loop
training_set=np.arange(training_range[0]+bins_before,training_range[1]-bins_after)
#Get training data
X_train=X[training_set,:,:]
X_flat_train=X_flat[training_set,:]
y_train=y[training_set,:]
#Get testing data
X_test=X[testing_set,:,:]
X_flat_test=X_flat[testing_set,:]
y_test=y[testing_set,:]
#Get validation data
X_valid=X[valid_set,:,:]
X_flat_valid=X_flat[valid_set,:]
y_valid=y[valid_set,:]
##### PREPROCESS DATA #####
#Z-score "X" inputs.
X_train_mean=np.nanmean(X_train,axis=0)
X_train_std=np.nanstd(X_train,axis=0)
X_train=(X_train-X_train_mean)/X_train_std
X_test=(X_test-X_train_mean)/X_train_std
X_valid=(X_valid-X_train_mean)/X_train_std
#Z-score "X_flat" inputs.
X_flat_train_mean=np.nanmean(X_flat_train,axis=0)
X_flat_train_std=np.nanstd(X_flat_train,axis=0)
X_flat_train=(X_flat_train-X_flat_train_mean)/X_flat_train_std
X_flat_test=(X_flat_test-X_flat_train_mean)/X_flat_train_std
X_flat_valid=(X_flat_valid-X_flat_train_mean)/X_flat_train_std
#Zero-center outputs
y_train_mean=np.nanmean(y_train,axis=0)
y_train=y_train-y_train_mean
y_test=y_test-y_train_mean
y_valid=y_valid-y_train_mean
#Z-score outputs (for SVR)
y_train_std=np.nanstd(y_train,axis=0)
y_zscore_train=y_train/y_train_std
y_zscore_test=y_test/y_train_std
y_zscore_valid=y_valid/y_train_std
################# DECODING #################
#Add actual train/valid/test data to lists (for saving)
y_test_all.append(y_test)
y_train_all.append(y_train)
y_valid_all.append(y_valid)
###### WIENER FILTER ###
if run_wf:
#Declare model
model_wf=WienerFilterDecoder()
#Fit model
model_wf.fit(X_flat_train,y_train)
#Get predictions
y_test_predicted_wf=model_wf.predict(X_flat_test)
#Get metric of fit
mean_r2_wf[i]=np.mean(get_R2(y_test,y_test_predicted_wf))
R2s_wf=get_R2(y_test,y_test_predicted_wf)
print('R2s_wf:', R2s_wf)
y_pred_wf_all.append(y_test_predicted_wf)
##### WIENER CASCADE #####
if run_wc:
#Get hyperparameters using validation set
def wc_evaluate(degree):
model_wc=WienerCascadeDecoder(degree)
model_wc.fit(X_flat_train,y_train)
y_valid_predicted_wc=model_wc.predict(X_flat_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_wc))
wcBO = BayesianOptimization(wc_evaluate, {'degree': (1, 5.01)}, verbose=0)
wcBO.maximize(init_points=3, n_iter=3)
best_params=wcBO.res['max']['max_params']
degree=best_params['degree']
print("degree=", degree)
# Run model w/ above hyperparameters
model_wc=WienerCascadeDecoder(degree)
model_wc.fit(X_flat_train,y_train)
y_test_predicted_wc=model_wc.predict(X_flat_test)
mean_r2_wc[i]=np.mean(get_R2(y_test,y_test_predicted_wc))
R2s_wc=get_R2(y_test,y_test_predicted_wc)
print('R2s_wc:', R2s_wc)
y_pred_wc_all.append(y_test_predicted_wc)
##### SIMPLE RNN ######
if run_rnn:
#Get hyperparameters using validation set
def rnn_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_rnn=SimpleRNNDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_rnn.fit(X_train,y_train)
y_valid_predicted_rnn=model_rnn.predict(X_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_rnn))
rnnBO = BayesianOptimization(rnn_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
rnnBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=rnnBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_rnn=SimpleRNNDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_rnn.fit(X_train,y_train)
y_test_predicted_rnn=model_rnn.predict(X_test)
mean_r2_rnn[i]=np.mean(get_R2(y_test,y_test_predicted_rnn))
R2s_rnn=get_R2(y_test,y_test_predicted_rnn)
print('R2s:', R2s_rnn)
y_pred_rnn_all.append(y_test_predicted_rnn)
##### GRU ######
if run_gru:
#Get hyperparameters using validation set
def gru_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_gru=GRUDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_gru.fit(X_train,y_train)
y_valid_predicted_gru=model_gru.predict(X_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_gru))
gruBO = BayesianOptimization(gru_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
gruBO.maximize(init_points=20, n_iter=20,kappa=10)
best_params=gruBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_gru=GRUDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_gru.fit(X_train,y_train)
y_test_predicted_gru=model_gru.predict(X_test)
mean_r2_gru[i]=np.mean(get_R2(y_test,y_test_predicted_gru))
R2s_gru=get_R2(y_test,y_test_predicted_gru)
print('R2s:', R2s_gru)
y_pred_gru_all.append(y_test_predicted_gru)
##### LSTM ######
if run_lstm:
#Get hyperparameters using validation set
def lstm_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_lstm=LSTMDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_lstm.fit(X_train,y_train)
y_valid_predicted_lstm=model_lstm.predict(X_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_lstm))
lstmBO = BayesianOptimization(lstm_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
lstmBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=lstmBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_lstm=LSTMDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_lstm.fit(X_train,y_train)
y_test_predicted_lstm=model_lstm.predict(X_test)
mean_r2_lstm[i]=np.mean(get_R2(y_test,y_test_predicted_lstm))
R2s_lstm=get_R2(y_test,y_test_predicted_lstm)
print('R2s:', R2s_lstm)
y_pred_lstm_all.append(y_test_predicted_lstm)
##### Dense (Feedforward) NN ######
if run_dnn:
#Get hyperparameters using validation set
def dnn_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs)
model_dnn.fit(X_flat_train,y_train)
y_valid_predicted_dnn=model_dnn.predict(X_flat_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_dnn))
dnnBO = BayesianOptimization(dnn_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
dnnBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=dnnBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs)
model_dnn.fit(X_flat_train,y_train)
y_test_predicted_dnn=model_dnn.predict(X_flat_test)
mean_r2_dnn[i]=np.mean(get_R2(y_test,y_test_predicted_dnn))
R2s_dnn=get_R2(y_test,y_test_predicted_dnn)
print('R2s:', R2s_dnn)
y_pred_dnn_all.append(y_test_predicted_dnn)
##### SVR #####
if run_svr:
#Get hyperparameters using validation set
max_iter=4000 #2000 for M1, 4000 for HC
def svr_evaluate(C):
model_svr=SVRDecoder(C=C, max_iter=max_iter)
model_svr.fit(X_flat_train,y_zscore_train)
y_valid_predicted_svr=model_svr.predict(X_flat_valid)
return np.mean(get_R2(y_zscore_valid,y_valid_predicted_svr))
svrBO = BayesianOptimization(svr_evaluate, {'C': (.5, 10)}, verbose=0)
svrBO.maximize(init_points=5, n_iter=5)
best_params=svrBO.res['max']['max_params']
C=best_params['C']
print("C=", C)
# Run model w/ above hyperparameters
model_svr=SVRDecoder(C=C, max_iter=max_iter)
model_svr.fit(X_flat_train,y_zscore_train)
y_test_predicted_svr=model_svr.predict(X_flat_test)
mean_r2_svr[i]=np.mean(get_R2(y_zscore_test,y_test_predicted_svr))
R2s_svr=get_R2(y_zscore_test,y_test_predicted_svr)
print('R2s_svr:', R2s_svr)
y_pred_svr_all.append(y_test_predicted_svr)
##### XGBOOST ######
if run_xgb:
#Get hyperparameters using validation set
def xgb_evaluate(max_depth,num_round,eta):
max_depth=int(max_depth)
num_round=int(num_round)
eta=float(eta)
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta)
model_xgb.fit(X_flat_train,y_train)
y_valid_predicted_xgb=model_xgb.predict(X_flat_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_xgb))
xgbBO = BayesianOptimization(xgb_evaluate, {'max_depth': (2, 10.01), 'num_round': (100,700), 'eta': (0, 1)})
xgbBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=xgbBO.res['max']['max_params']
num_round=np.int(best_params['num_round'])
max_depth=np.int(best_params['max_depth'])
eta=best_params['eta']
# Run model w/ above hyperparameters
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta)
model_xgb.fit(X_flat_train,y_train)
y_test_predicted_xgb=model_xgb.predict(X_flat_test)
mean_r2_xgb[i]=np.mean(get_R2(y_test,y_test_predicted_xgb))
R2s_xgb=get_R2(y_test,y_test_predicted_xgb)
print('R2s:', R2s_xgb)
y_pred_xgb_all.append(y_test_predicted_xgb)
print ("\n")
time_elapsed=time.time()-t1
###### SAVE RESULTS #####
#Note that I save them after every cross-validation fold rather than at the end in case the code/computer crashes for some reason while running
#Only save results for the decoder we chose to run
if run_wf:
with open(save_folder+dataset+'_results_amt_wf.pickle','wb') as f:
pickle.dump([mean_r2_wf,y_pred_wf_all],f)
if run_wc:
with open(save_folder+dataset+'_results_amt_wc.pickle','wb') as f:
pickle.dump([mean_r2_wc,y_pred_wc_all],f)
if run_xgb:
with open(save_folder+dataset+'_results_amt_xgb.pickle','wb') as f:
pickle.dump([mean_r2_xgb,y_pred_xgb_all],f)
if run_dnn:
with open(save_folder+dataset+'_results_amt_dnn.pickle','wb') as f:
pickle.dump([mean_r2_dnn,y_pred_dnn_all,time_elapsed],f)
if run_rnn:
with open(save_folder+dataset+'_results_amt_rnn.pickle','wb') as f:
pickle.dump([mean_r2_rnn,y_pred_rnn_all,time_elapsed],f)
if run_gru:
with open(save_folder+dataset+'_results_amt_gru.pickle','wb') as f:
pickle.dump([mean_r2_gru,y_pred_gru_all,time_elapsed],f)
if run_lstm:
with open(save_folder+dataset+'_results_amt_lstm.pickle','wb') as f:
pickle.dump([mean_r2_lstm,y_pred_lstm_all,time_elapsed],f)
if run_svr:
with open(save_folder+dataset+'_results_amt_svr.pickle','wb') as f:
pickle.dump([mean_r2_svr,y_pred_svr_all,time_elapsed],f)
print("time_elapsed:",time_elapsed)
#Save ground truth results
with open(save_folder+dataset+'_ground_truth_amt.pickle','wb') as f:
pickle.dump([y_test_all,y_train_all,y_valid_all],f)
# -
mean_r2_wf
| Paper_code/ManyDecoders_DataAmt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# reload packages
# %load_ext autoreload
# %autoreload 2
# ### Choose GPU (this may not be needed on your computer)
# %env CUDA_DEVICE_ORDER=PCI_BUS_ID
# %env CUDA_VISIBLE_DEVICES=0
import tensorflow as tf
gpu_devices = tf.config.experimental.list_physical_devices('GPU')
if len(gpu_devices)>0:
tf.config.experimental.set_memory_growth(gpu_devices[0], True)
print(gpu_devices)
# ### load packages
from tfumap.umap import tfUMAP
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
import umap
import pandas as pd
# ### Load dataset
dataset = 'fmnist'
dims = (28,28,1)
from tensorflow.keras.datasets import fashion_mnist
# +
# load dataset
(train_images, Y_train), (test_images, Y_test) = fashion_mnist.load_data()
X_train = (train_images/255.).astype('float32')
X_test = (test_images/255.).astype('float32')
X_train = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
# subset a validation set
n_valid = 10000
X_valid = X_train[-n_valid:]
Y_valid = Y_train[-n_valid:]
X_train = X_train[:-n_valid]
Y_train = Y_train[:-n_valid]
# flatten X
X_train_flat = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test_flat = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
X_valid_flat= X_valid.reshape((len(X_valid), np.product(np.shape(X_valid)[1:])))
print(len(X_train), len(X_valid), len(X_test))
# -
# ### define networks
dims = (28,28,1)
n_components = 2
encoder = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=dims),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation="relu"
),
tf.keras.layers.Conv2D(
filters=128, kernel_size=3, strides=(2, 2), activation="relu"
),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=n_components),
])
# ### Create model and train
batch_size = 5000
from tfumap.paths import ensure_dir, MODEL_DIR, DATA_DIR
from tfumap.parametric_tsne import compute_joint_probabilities, tsne_loss
save_loc = DATA_DIR/ 'parametric_tsne'/ dataset / 'P.npy'
if save_loc.exists():
P = np.load(save_loc)
else:
P = compute_joint_probabilities(X_train_flat, batch_size=batch_size, perplexity=30, verbose=2)
ensure_dir(save_loc)
np.save(save_loc, P)
# Joint probabilities of data
Y_train_tsne = P.reshape(X_train.shape[0], -1)
opt = tf.keras.optimizers.Adam(lr=0.01)
encoder.compile(loss=tsne_loss(d=n_components, batch_size=batch_size), optimizer=opt)
X_train = np.reshape(X_train, ([len(X_train)]+ list(dims)))
X_test = np.reshape(X_test, ([len(X_test)]+ list(dims)))
# because shuffle == False, the same batches are used each time...
history = encoder.fit(X_train, Y_train_tsne, batch_size=batch_size, shuffle=False, nb_epoch=1000)
# ### get z for training and test
z = encoder.predict(X_train)
z_test = encoder.predict(X_test)
# ### Test plot
fig, axs = plt.subplots(ncols = 2, figsize=(10, 5))
axs[0].scatter(z[:, 0], z[:, 1], s=0.1, alpha=0.5, c=Y_train, cmap=plt.cm.tab10)
axs[1].scatter(z_test[:, 0], z_test[:, 1], s=1, alpha=0.5, c=Y_test, cmap=plt.cm.tab10)
# ### Save models + projections
import os
output_dir = MODEL_DIR/'projections'/ dataset / 'parametric-tsne'
encoder.save(os.path.join(output_dir, "encoder"))
np.save(output_dir / 'z.npy', z)
np.save(output_dir / 'z_test.npy', z_test)
# ### compute metrics
# #### silhouette
from tfumap.silhouette import silhouette_score_block
ss, sil_samp = silhouette_score_block(z, Y_train, n_jobs = -1)
ss
ss_test, sil_samp_test = silhouette_score_block(z_test, Y_test, n_jobs = -1)
ss_test
fig, axs = plt.subplots(ncols = 2, figsize=(10, 5))
axs[0].scatter(z[:, 0], z[:, 1], s=0.1, alpha=0.5, c=sil_samp, cmap=plt.cm.viridis)
axs[1].scatter(z_test[:, 0], z_test[:, 1], s=1, alpha=0.5, c=sil_samp_test, cmap=plt.cm.viridis)
# #### KNN
from sklearn.neighbors import KNeighborsClassifier
neigh5 = KNeighborsClassifier(n_neighbors=5)
neigh5.fit(z, Y_train)
score_5nn = neigh5.score(z_test, Y_test)
score_5nn
neigh1 = KNeighborsClassifier(n_neighbors=1)
neigh1.fit(z, Y_train)
score_1nn = neigh1.score(z_test, Y_test)
score_1nn
# #### Trustworthiness
from sklearn.manifold import trustworthiness
tw = trustworthiness(X_train_flat[:10000], z[:10000])
tw_test = trustworthiness(X_test_flat[:10000], z_test[:10000])
tw, tw_test
# #### save output metrics
metrics_df = pd.DataFrame(
columns=[
"dataset",
"class_",
"dim",
"trustworthiness",
"silhouette_score",
"silhouette_samples",
]
)
metrics_df.loc[len(metrics_df)] = [dataset, 'parametric-tsne', n_components, tw, ss, sil_samp]
metrics_df
save_loc = DATA_DIR / 'projection_metrics' / 'train' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
metrics_df.to_pickle(save_loc)
metrics_df_test = pd.DataFrame(
columns=[
"dataset",
"class_",
"dim",
"trustworthiness",
"silhouette_score",
"silhouette_samples",
]
)
metrics_df_test.loc[len(metrics_df)] = [dataset, 'parametric-tsne', n_components, tw_test, ss_test, sil_samp_test]
metrics_df_test
save_loc = DATA_DIR / 'projection_metrics' / 'test' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
metrics_df.to_pickle(save_loc)
nn_acc_df = pd.DataFrame(columns = ["method_","dimensions","dataset","1NN_acc","5NN_acc"])
nn_acc_df.loc[len(nn_acc_df)] = ['parametric-tsne', n_components, dataset, score_1nn, score_5nn]
nn_acc_df
save_loc = DATA_DIR / 'knn_classifier' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
nn_acc_df.to_pickle(save_loc)
| notebooks/dataset-projections/fmnist/fashion-mnist-parametric-tsne.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manejo de Ficheros
# Python nos permite trabajar en dos niveles diferentes con respecto al sistema de archivos y directorios. Uno de ellos, es a través del módulo os, que como su nombre lo indica, nos facilita el trabajo con todo el sistema de archivos y directorios, a nivel del propios Sistema Operativo. El segundo nivel -más simple-, es el que nos permite trabajar con archivos, manipulando su lectura y escritura a nivel de la aplicación y tratando a cada archivo como un objeto.
#
# ## Objeto File
# ------------------------------
# Al igual que sucede con otras variables, manipular una de ellas como un objeto File, es posible, cuando a ésta, se le asigna como valor un archivo.
# Para asignar a una variable un valor de tipo file, solo es necesario recurrir a la función integrada open(), la cuál está destinada a la apertura de un archivo.
#
# - La función integrada open(), recibe dos parámetros: – El primero de ellos, es la ruta hacia el archivo que se desea abrir
#
# - Y el segundo, el modo en el cual abrirlo
# +
# Forma 1 de apertura a partir del método with
with open('./src/texto.txt',mode='r') as f:
data= f.readlines()
print(data[::200])
# Archivo se cierra de forma automática
# +
# Forma 2 de apertura
f = open('./src/texto.txt')
data = f.readlines() # readlines -> permite leer todas las líneas del archivo, en forma de lista
print(data[::200])
f.close()
# -
# Valido si archivo se encuentra efectivamente cerrado
f.closed
# ### Modos de Apertura
# Por defecto un archivo se apertura en modo **lectura (r)** pero en la práctica existen muchos más modos los cuales nos permitiran escribir (w) o modificar un archivo ya existente
# <img src='./img/modo_apertura.PNG'>
# <img src='./img/modo_apertura2.PNG'>
# ### Métodos de Archivos
# El objeto file, entre sus métodos más frecuentes, dispone de los siguientes
# <img src='./img/metodos_files.PNG'>
# ## Lectura
# +
# Ruta donde leeremos el fichero, r indica lectura (por defecto ya es r)
fichero = open('./src/texto.txt','r')
# Lectura completa
texto = fichero.read() # read -> Metodo que permite la lectura de todo el archivo
# Cerramos el fichero
fichero.close()
print(texto)
# -
# También se puede leer un fichero utilizando la instrucción estándar with de la siguiente forma:
with open('./src/texto.txt', "r") as fichero:
for linea in fichero:
print(linea)
# ## Escritura
# +
texto = "Una línea con texto\nOtra línea con texto"
# Ruta donde crearemos el fichero, w indica escritura (puntero al principio)
# De existir el archivo, este será eliminado y creado uno nuevo
with open('./src/fichero.txt','w') as f:
# Escribimos el texto
f.write(texto)
# -
with open('./src/fichero.txt','r') as f:
# Escribimos el texto
data = f.readlines()
data
# Este modo nos permite añadir datos al final de un fichero:
# Ruta donde leeremos el fichero, a indica extensión (puntero al final)
with open('./src/fichero.txt','a') as f:
f.write('\nOtra línea más abajo del todo')
# La variante 'a+' permite crear el fichero si no existe:
# Ruta archivo del tipo a+
with open('fichero_inventado.txt','a+') as f:
f.write('\nhola mundo')
pass
# ## Lectura con escritura
# Se puede abrir un fichero en modo lectura con escritura, pero éste debe existir préviamente. Además por defecto el puntero estará al principio y si escribimos algo sobreescribiremos el contenido actual, así que prestad atención a los saltos de línea y caracteres especiales:
#
# Creamos un fichero de prueba con 4 líneas
fichero = open('fichero2.txt','w')
texto = "Línea 1\nLínea 2\nLínea 3\nLínea 4"
fichero.write(texto)
fichero.close()
# Lo abrimos en lectura con escritura y escribimos algo
fichero = open('fichero2.txt','r+')
fichero.write("0123456")
fichero.close()
# Volvemos a ponter el puntero al inicio y leemos hasta el final
fichero.seek(0)
fichero.read()
fichero.close()
# ### Modificar una línea
# Para lograr este fin lo mejor es leer todas las líneas en una lista, modificar la línea en la lista, posicionar el puntero al principio y reescribir de nuevo todas las líneas:
#
# +
fichero = open('fichero2.txt','r+')
texto = fichero.readlines()
# Modificamos la línea que queramos a partir del índice
texto[2] = "Esta es la línea 3 modificada\n"
# Volvemos a ponter el puntero al inicio y reescribimos
fichero.seek(0)
fichero.writelines(texto)
fichero.close()
# Leemos el fichero de nuevo
with open("fichero2.txt", "r") as fichero:
print(fichero.read())
# -
# ### <a href='https://docs.python.org/2.4/lib/standard-encodings.html'>Encoding</a>
# **encoding** es el nombre de la codificación utilizada para decodificar o codificar el archivo. Esto solo debe usarse en modo texto. La codificación predeterminada es dependiente de la plataforma, pero cualquier codificación compatible con Python se puede utilizar.
# Más conocidos:
#
# - utf-8
# - latin1
# - ascii
#
# #### Lectura sin encoding con texto español
#
#
#
with open('./src/fichero_esp.txt') as f:
texto = f.read()
print(texto)
# #### Lectura con encoding utf-8
with open('./src/fichero_esp.txt',encoding='utf-8') as f:
texto = f.read()
print(texto)
# # EJERCICIOS
# ----------------------------
# <h3>1.</h3>
#
# En este ejercicio deberás crear un script llamado <code>personas.py</code> que lea los datos de un fichero de texto, que transforme cada fila en un diccionario y lo añada a una lista llamada personas. Luego rocorre las personas de la lista y paracada una muestra de forma amigable todos sus campos.
#
# El fichero de texto se denominará <code>personas.txt</code> y tendrá el siguiente contenido en texto plano (créalo previamente):
# <code>1;Carlos;Pérez;05/01/1989
# 2;Manuel;Heredia;26/12/1973
# 3;Rosa;Campos;12/06/1961
# 4;David;García;25/07/2006</code>
# +
from io import open
fichero = open('personas.txt','r', encoding="utf8")
lineas = fichero.readlines()
fichero.close()
personas = []
for linea in lineas:
# Borramos los saltos de línea y separamos
campos = linea.replace("\n", "").split(";")
persona = {"id":campos[0], "nombre":campos[1],
"apellido":campos[2], "nacimiento":campos[3]}
personas.append(persona)
for p in personas:
print("(id={}) {} {} => {} ".format( p['id'], p['nombre'],
p['apellido'], p['nacimiento']) )
# -
| Modulo4/1. Manejo de Ficheros.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
'''Deep Dreaming in Keras.
Run the script with:
```
python deep_dream.py path_to_your_base_image.jpg prefix_for_results
```
e.g.:
```
python deep_dream.py img/mypic.jpg results/dream
```
It is preferable to run this script on GPU, for speed.
If running on CPU, prefer the TensorFlow backend (much faster).
Example results: http://i.imgur.com/FX6ROg9.jpg
'''
from __future__ import print_function
from keras.preprocessing.image import load_img, img_to_array
import numpy as np
from scipy.misc import imsave
from scipy.optimize import fmin_l_bfgs_b
import time
import argparse
from keras.applications import vgg16
from keras import backend as K
from keras.layers import Input
# +
#parser = argparse.ArgumentParser(description='Deep Dreams with Keras.')
#parser.add_argument('base_image_path', metavar='base', type=str,
# help='Path to the image to transform.')
#parser.add_argument('result_prefix', metavar='res_prefix', type=str,
# help='Prefix for the saved results.')
#args = parser.parse_args()
base_image_path = 'base_image.jpg'#args.base_image_path
result_prefix = 'results'#args.result_prefix
# dimensions of the generated picture.
img_height = 2233
img_width = 4613
# path to the model weights file.
weights_path = 'vgg16_weights.h5'
# some settings we found interesting
saved_settings = {
'bad_trip': {'features': {'block4_conv1': 0.05,
'block4_conv2': 0.01,
'block4_conv3': 0.01},
'continuity': 0.1,
'dream_l2': 0.8,
'jitter': 5},
'dreamy': {'features': {'block5_conv1': 0.05,
'block5_conv2': 0.02},
'continuity': 0.1,
'dream_l2': 0.02,
'jitter': 0},
}
# the settings we will use in this experiment
settings = saved_settings['bad_trip']
# +
# util function to open, resize and format pictures into appropriate tensors
def preprocess_image(image_path):
img = load_img(image_path, target_size=(img_height, img_width))
img = img_to_array(img)
img = np.expand_dims(img, axis=0)
img = vgg16.preprocess_input(img)
return img
# util function to convert a tensor into a valid image
def deprocess_image(x):
if K.image_dim_ordering() == 'th':
x = x.reshape((3, img_height, img_width))
x = x.transpose((1, 2, 0))
else:
x = x.reshape((img_height, img_width, 3))
# Remove zero-center by mean pixel
x[:, :, 0] += 103.939
x[:, :, 1] += 116.779
x[:, :, 2] += 123.68
# 'BGR'->'RGB'
x = x[:, :, ::-1]
x = np.clip(x, 0, 255).astype('uint8')
return x
# +
if K.image_dim_ordering() == 'th':
img_size = (3, img_height, img_width)
else:
img_size = (img_height, img_width, 3)
# this will contain our generated image
dream = Input(batch_shape=(1,) + img_size)
# build the VGG16 network with our placeholder
# the model will be loaded with pre-trained ImageNet weights
model = vgg16.VGG16(input_tensor=dream,
weights='imagenet', include_top=False)
print('Model loaded.')
# get the symbolic outputs of each "key" layer (we gave them unique names).
layer_dict = dict([(layer.name, layer) for layer in model.layers])
# +
# continuity loss util function
def continuity_loss(x):
assert K.ndim(x) == 4
if K.image_dim_ordering() == 'th':
a = K.square(x[:, :, :img_height - 1, :img_width - 1] -
x[:, :, 1:, :img_width - 1])
b = K.square(x[:, :, :img_height - 1, :img_width - 1] -
x[:, :, :img_height - 1, 1:])
else:
a = K.square(x[:, :img_height - 1, :img_width - 1, :] -
x[:, 1:, :img_width - 1, :])
b = K.square(x[:, :img_height - 1, :img_width - 1, :] -
x[:, :img_height - 1, 1:, :])
return K.sum(K.pow(a + b, 1.25))
# -
# define the loss
loss = K.variable(0.)
for layer_name in settings['features']:
# add the L2 norm of the features of a layer to the loss
assert layer_name in layer_dict.keys(), 'Layer ' + layer_name + ' not found in model.'
coeff = settings['features'][layer_name]
x = layer_dict[layer_name].output
shape = layer_dict[layer_name].output_shape
# we avoid border artifacts by only involving non-border pixels in the loss
if K.image_dim_ordering() == 'th':
loss -= coeff * K.sum(K.square(x[:, :, 2: shape[2] - 2, 2: shape[3] - 2])) / np.prod(shape[1:])
else:
loss -= coeff * K.sum(K.square(x[:, 2: shape[1] - 2, 2: shape[2] - 2, :])) / np.prod(shape[1:])
# +
# add continuity loss (gives image local coherence, can result in an artful blur)
loss += settings['continuity'] * continuity_loss(dream) / np.prod(img_size)
# add image L2 norm to loss (prevents pixels from taking very high values, makes image darker)
loss += settings['dream_l2'] * K.sum(K.square(dream)) / np.prod(img_size)
# feel free to further modify the loss as you see fit, to achieve new effects...
# compute the gradients of the dream wrt the loss
grads = K.gradients(loss, dream)
outputs = [loss]
if isinstance(grads, (list, tuple)):
outputs += grads
else:
outputs.append(grads)
f_outputs = K.function([dream], outputs)
def eval_loss_and_grads(x):
x = x.reshape((1,) + img_size)
outs = f_outputs([x])
loss_value = outs[0]
if len(outs[1:]) == 1:
grad_values = outs[1].flatten().astype('float64')
else:
grad_values = np.array(outs[1:]).flatten().astype('float64')
return loss_value, grad_values
# this Evaluator class makes it possible
# to compute loss and gradients in one pass
# while retrieving them via two separate functions,
# "loss" and "grads". This is done because scipy.optimize
# requires separate functions for loss and gradients,
# but computing them separately would be inefficient.
class Evaluator(object):
def __init__(self):
self.loss_value = None
self.grad_values = None
def loss(self, x):
assert self.loss_value is None
loss_value, grad_values = eval_loss_and_grads(x)
self.loss_value = loss_value
self.grad_values = grad_values
return self.loss_value
def grads(self, x):
assert self.loss_value is not None
grad_values = np.copy(self.grad_values)
self.loss_value = None
self.grad_values = None
return grad_values
evaluator = Evaluator()
# -
# run scipy-based optimization (L-BFGS) over the pixels of the generated image
# so as to minimize the loss
x = preprocess_image(base_image_path)
for i in range(15):
print('Start of iteration', i)
start_time = time.time()
# add a random jitter to the initial image. This will be reverted at decoding time
random_jitter = (settings['jitter'] * 2) * (np.random.random(img_size) - 0.5)
x += random_jitter
# run L-BFGS for 7 steps
x, min_val, info = fmin_l_bfgs_b(evaluator.loss, x.flatten(),
fprime=evaluator.grads, maxfun=7)
print('Current loss value:', min_val)
# decode the dream and save it
x = x.reshape(img_size)
x -= random_jitter
img = deprocess_image(np.copy(x))
fname = result_prefix + '_at_iteration_%d.png' % i
imsave(fname, img)
end_time = time.time()
print('Image saved as', fname)
print('Iteration %d completed in %ds' % (i, end_time - start_time))
| Deep Dreaming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # T-Student regression
# ---
# In this notebook we analyze how to adjust a *bayesian linear regression* code when the outcome has heavy tail and the hypotesis of *normal distribution* are not vadid. We use a synthetic dataset in order to see which is the impact of the right model on the results. The code used is *Python* with a specific bayesian framework **PyMC3**.
import warnings
warnings.filterwarnings("ignore")
from scipy import stats
import arviz as az
import numpy as np
import matplotlib.pyplot as plt
import pymc3 as pm
import seaborn as sns
import pandas as pd
import random
from theano import shared
from sklearn import preprocessing
print('Running on PyMC3 v{}'.format(pm.__version__))
# Let's simulate a riproducible synthetic dataset. By default we don't have mean and variance for *T-Student distribution* but we can extend to the usual three parameter **location-scale** family as follow:
# $X = \mu + \sigma T$, see [here](https://en.wikipedia.org/wiki/Student%27s_t-distribution#Generalized_Student's_t-distribution)
# +
random.seed(1990)
N = 359 # Sample size
b0 = 3.8 # b0 coefficient (intercept)
b1 = 2.4 # b1 coefficient (slope)
# Simulate indipendent variable
x = np.linspace(-3,4,N)
# Simulate the output
y = b0 + b1*x + np.random.normal(0,3,N) + [*np.repeat([0], 300, axis=0), *20*np.random.binomial(1,0.3,59)]
# -
plt.hist(y)
plt.title('Histogram Outcome')
plt.show()
# ## Gaussian Model
with pm.Model() as model_gaussian:
# Prior distribution
sigma = pm.HalfCauchy("sigma", beta=10, testval=1.0)
beta0 = pm.Normal("beta0", 0, sigma=20)
beta1 = pm.Normal("beta1", 0, sigma=20)
# Likelihood
lik = pm.Normal('y', mu = beta0 + beta1 * x, sigma=sigma, observed=y)
# draw 1000 posterior samples using NUTS sampling + 1000 of warmup on 4 chain
trace_gaussian = pm.sample(1000, tune=1000)
az.plot_trace(trace_gaussian);
# ## T-Student Model
with pm.Model() as model_tstudent:
# Prior distribution
sigma = pm.HalfCauchy("sigma", beta=10, testval=1.0)
beta0 = pm.Normal("beta0", 0, sigma=20)
beta1 = pm.Normal("beta1", 0, sigma=20)
n = pm.Gamma('n', alpha = 2, beta = 0.1)
# Likelihood
lik = pm.StudentT('y', nu=n, mu = beta0 + beta1 * x, sigma=sigma, observed=y)
# draw 1000 posterior samples using NUTS sampling + 1000 of warmup on 4 chain
trace_tstudent = pm.sample(1000, tune=1000)
az.plot_trace(trace_tstudent);
# ## Results
az.summary(trace_gaussian)
az.summary(trace_tstudent)
fig = plt.figure(figsize=(7, 7))
ax = fig.add_subplot(111, xlabel="x", ylabel="y", title="Generated data and underlying model")
ax.plot(x, y, "x", label="sampled data")
ax.plot(x, b0 + b1*x, label="true regression line", lw=2.0)
ax.plot(x, trace_gaussian["beta0"].mean() + trace_gaussian["beta1"].mean()*x, label="regression line", lw=2.0, color = "r")
ax.plot(x, trace_tstudent["beta0"].mean() + trace_tstudent["beta1"].mean()*x, label="t-regression line", lw=2.0, color = "m")
plt.legend(loc=0);
| t_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 6: Web Applications
#
# For this homework, you're going to write a web API for the lake data in the MONDIAL database. (Make sure you've imported the data as originally outlined in [our week 1 tutorial](https://github.com/ledeprogram/data-and-databases/blob/master/SQL_notes.md).)
#
# The API should perform the following tasks:
#
# * A request to `/lakes` should return a JSON list of dictionaries, with the information from the `name`, `elevation`, `area` and `type` fields from the `lake` table in MONDIAL.
# * The API should recognize the query string parameter `sort`. When left blank or set to `name`, the results should be sorted by the name of the lake (in alphabetical order). When set to `area` or `elevation`, the results should be sorted by the requested field, in *descending* order.
# * The API should recognize the query string parameter `type`. When specified, the results should *only* include rows that have the specified value in the `type` field.
# * You should be able to use *both* the `sort` and `type` parameters in any request.
#
# This notebook contains only *test requests to your API*. Write the API as a standalone Python program, start the program and then run the code in the cells below to ensure that your API produces the expected output. When you're done, paste the source code in the final cell (so we can check your work, if needed).
#
# Hints when writing your API code:
#
# * You'll need to construct the SQL query as a string, piece by piece. This will likely involve a somewhat messy tangle of `if` statements. Lean into the messy tangle.
# * Make sure to use parameter placeholders (%s) in the query.
# * If you're getting SQL errors, print out your SQL statement in the request handler function so you can debug it. (When you use `print()` in Flask, the results will display in your terminal window.)
# * When in doubt, return to the test code. Examine it carefully and make sure you know exactly what it's trying to do.
#
# ## Problem set #1: A list of lakes
#
# Your API should return a JSON list of dictionaries (objects). Use the code below to determine what the keys of the dictionaries should be. (For brevity, this example only prints out the first ten records, but of course your API should return all of them.)
#
# Expected output:
#
# 143 lakes
# Ammersee - elevation: 533 m / area: 46 km^2 / type: None
# Arresoe - elevation: None m / area: 40 km^2 / type: None
# Atlin Lake - elevation: 668 m / area: 798 km^2 / type: None
# Balaton - elevation: 104 m / area: 594 km^2 / type: None
# Barrage de Mbakaou - elevation: None m / area: None km^2 / type: dam
# Bodensee - elevation: 395 m / area: 538 km^2 / type: None
# Brienzersee - elevation: 564 m / area: 29 km^2 / type: None
# Caspian Sea - elevation: -28 m / area: 386400 km^2 / type: salt
# Chad Lake - elevation: 250 m / area: 23000 km^2 / type: salt
# Chew Bahir - elevation: 520 m / area: 800 km^2 / type: salt
import requests
data = requests.get('http://localhost:5000/lakes').json()
print(len(data), "lakes")
for item in data[:10]:
print(item['name'], "- altitude:", item['altitude'], "m / area:", item['area'], "km^2 / type:", item['type'])
# ## Problem set #2: Lakes of a certain type
#
# The following code fetches all lakes of type `salt` and finds their average area and elevation.
#
# Expected output:
#
# average area: 18880
# average elevation: 970
import requests
data = requests.get('http://localhost:5000/lakes?type=salt').json()
avg_area = sum([x['area'] for x in data if x['area'] is not None]) / len(data)
avg_elev = sum([x['altitude'] for x in data if x['altitude'] is not None]) / len(data)
print("average area:", int(avg_area))
print("average altitude:", int(avg_elev))
# ## Problem set #3: Lakes in order
#
# The following code fetches lakes in reverse order by their elevation and prints out the name of the first fifteen, excluding lakes with an empty elevation field.
#
# Expected output:
#
# * Licancabur Crater Lake
# * Nam Co
# * Lago Junin
# * Lake Titicaca
# * Poopo
# * Salar de Uyuni
# * Koli Sarez
# * Lake Irazu
# * Qinghai Lake
# * Segara Anak
# * Lake Tahoe
# * Crater Lake
# * Lake Tana
# * Lake Van
# * Issyk-Kul
import requests
data = requests.get('http://localhost:5000/lakes?sort=elevation').json()
for item in [x['name'] for x in data if x['elevation'] is not None][:15]:
print("*", item)
# ## Problem set #4: Order and type
#
# The following code prints the names of the largest caldera lakes, ordered in reverse order by area.
#
# Expected output:
#
# * Lake Nyos
# * Lake Toba
# * Lago Trasimeno
# * <NAME>
# * <NAME>
# * Crater Lake
# * <NAME>
# * <NAME>
import requests
data = requests.get('http://localhost:5000/lakes?sort=area&type=caldera').json()
for item in data:
print("*", item['name'])
# ## Problem set #5: Error handling
#
# Your API should work fine even when faced with potential error-causing inputs. For example, the expected output for this statement is an empty list (`[]`), *not* every row in the table.
import requests
data = requests.get('http://localhost:5000/lakes', params={'type': "' OR true; --"}).json()
data
# Specifying a field other than `name`, `area` or `elevation` for the `sort` parameter should fail silently, defaulting to sorting alphabetically. Expected output: `['Ammersee', 'Arresoe', 'Atlin Lake', 'Balaton', 'Barrage de Mbakaou']
# `
import requests
data = requests.get('http://localhost:5000/lakes', params={'sort': "florb"}).json()
[x['name'] for x in data[:5]]
# ## Paste your code
#
# Please paste the code for your entire Flask application in the cell below, in case we want to take a look when grading or debugging your assignment.
# +
# paste code here
| homework6/Homework_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import os
from sklearn import preprocessing
import tensorflow as tf
from tensorflow.keras import backend as K
from tqdm import tqdm
#loading_data
data_dir='dataset'
model_dir='pretrained'
eol_data = np.load('%s/battery_EoL.npy'%(data_dir),allow_pickle='TRUE')
battery_id = np.load('%s/index_battery.npy'%(data_dir),allow_pickle='TRUE')
charge_data=np.load('%s/charge_data.npy'%(data_dir),allow_pickle='TRUE').tolist()
discharge_data=np.load('%s/discharge_data.npy'%(data_dir),allow_pickle='TRUE').tolist()
summary_data=np.load('%s/summary_data.npy'%(data_dir),allow_pickle='TRUE').tolist()
charge_norm=np.load('%s/charge_norm.npy'%(data_dir),allow_pickle='TRUE').tolist()
discharge_norm=np.load('%s/discharge_norm.npy'%(data_dir),allow_pickle='TRUE').tolist()
summary_norm=np.load('%s/summary_norm.npy'%(data_dir),allow_pickle='TRUE').tolist()
# +
# visualize charging data
cell_number=50
cycle=100
fig,axes=plt.subplots(nrows=4,ncols=1,figsize=(6,6),dpi=200)
for i in range(4):
if i==0:
axes[i].plot(charge_data[cell_number][cycle,i],c='b',label='cycle:100')
else:
axes[i].plot(charge_data[cell_number][cycle,i],c='b')
cycle=900
for i in range(4):
if i==0:
axes[i].plot(charge_data[cell_number][cycle,i],c='r',label='cycle:900')
else:
axes[i].plot(charge_data[cell_number][cycle,i],c='r')
axes[0].legend()
axes[0].set_ylabel('Capacity(Ah)')
axes[1].set_ylabel('Voltage(V)')
axes[2].set_ylabel('Current(C)')
axes[3].set_ylabel('Temperature(°C)')
fig.suptitle('Charging info', fontsize=16)
# -
# visualize discharging data
cycle=100
fig,axes=plt.subplots(nrows=4,ncols=1,figsize=(6,6),dpi=200)
for i in range(4):
if i==0:
axes[i].plot(discharge_data[cell_number][cycle,i],c='b',label='cycle:100')
else:
axes[i].plot(discharge_data[cell_number][cycle,i],c='b')
cycle=900
for i in range(4):
if i==0:
axes[i].plot(discharge_data[cell_number][cycle,i],c='r',label='cycle:900')
else:
axes[i].plot(discharge_data[cell_number][cycle,i],c='r')
axes[0].legend()
axes[0].set_ylabel('Capacity(Ah)')
axes[1].set_ylabel('Voltage(V)')
axes[2].set_ylabel('Current(C)')
axes[3].set_ylabel('Temperature(°C)')
fig.suptitle('Discharging info', fontsize=16)
# +
# visualize Summary data
fig,axes=plt.subplots(nrows=3,ncols=1,figsize=(6,10),dpi=200)
axes[0].plot(summary_data[cell_number][:,0],label='charge capacity')
axes[0].plot(summary_data[cell_number][:,1],label='discharge capacity')
axes[0].legend()
axes[1].plot(summary_data[cell_number][:,2],label='temperature mean')
axes[1].plot(summary_data[cell_number][:,3],label='temperature min')
axes[1].plot(summary_data[cell_number][:,4],label='temperature max')
axes[1].legend()
axes[2].plot(summary_data[cell_number][:,5],label='charge time')
axes[2].legend()
axes[0].set_ylabel('Capacity(Ah)')
axes[1].set_ylabel('Temperature(°C)')
axes[2].set_ylabel('Time(minute)')
axes[2].set_xlabel('Timestep')
fig.suptitle('Summary info', fontsize=16)
# -
| 1_Predicting/data_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import make_blobs
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score, mean_squared_error
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import BaggingClassifier
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
RANDOM_SEED = 139
# -
train_data, train_labels = make_blobs(n_samples=200, centers=[(0,1),(-3,-3),(4,2)],
n_features=2, random_state=RANDOM_SEED,
cluster_std=(1.2,1.5,1,))
train_data.shape
# Let’s write an auxiliary function that will return grid for further visualization.
def get_grid(data):
x_min, x_max = data[:, 0].min() - 1, data[:, 0].max() + 1
y_min, y_max = data[:, 1].min() - 1, data[:, 1].max() + 1
return np.meshgrid(np.arange(x_min, x_max, 0.01), np.arange(y_min, y_max, 0.01))
# +
clf_tree = DecisionTreeClassifier(criterion='entropy', max_depth=3,
random_state=RANDOM_SEED)
# training the tree
clf_tree.fit(train_data, train_labels)
# -
# some code to depict separating surface
xx, yy = get_grid(train_data)
predicted = clf_tree.predict(np.c_[xx.ravel(),
yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, predicted, cmap='coolwarm')
plt.scatter(train_data[:, 0], train_data[:, 1], c=train_labels, s=100,
cmap='coolwarm', edgecolors='black', linewidth=1.5);
df = pd.read_csv('../Data/bill_authentication.xls')
df
X = df.drop('Class', axis=1).values
y = df.Class.values
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=17, test_size=0.2)
# +
clf_tree = DecisionTreeClassifier(
max_depth=3,
random_state=17,
max_features=2
)
# training the tree
clf_tree.fit(X_train, y_train)
# -
y_pred = clf_tree.predict(X_test)
round(f1_score(y_test, y_pred), 3)
X_test.shape
print(f"Class is {clf_tree.predict(np.array([2.04378,-0.38422,1.437292,0.76421]).reshape([1, 4]))}.")
# ## Overfitting
# +
RANDOM_SEED = 139
train_data, train_labels = make_blobs(n_samples=100, centers=[(-3,-3),(4,2)],
n_features=2, random_state=RANDOM_SEED,
cluster_std=(5,5))
clf_tree = DecisionTreeClassifier(criterion='entropy', max_depth=None,
random_state=RANDOM_SEED)
# training the tree
clf_tree.fit(train_data, train_labels)
# some code to depict separating surface
xx, yy = get_grid(train_data)
predicted = clf_tree.predict(np.c_[xx.ravel(),
yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, predicted, cmap='coolwarm')
plt.scatter(train_data[:, 0], train_data[:, 1], c=train_labels, s=100,
cmap='coolwarm', edgecolors='black', linewidth=1.5);
# -
# ## Regression
# +
from sklearn.tree import DecisionTreeRegressor
from sklearn.datasets import make_blobs
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
RANDOM_SEED = 139
n_train = 150
n_test = 1000
noise = 0.1
def f(x):
x = x.ravel()
return np.exp(-x ** 2) + 1.5 * np.exp(-(x - 5) ** 2)
def generate(n_samples, noise):
X = np.random.rand(n_samples) * 10 - 5
X = np.sort(X).ravel()
y = np.exp(-X ** 2) + 1.5 * np.exp(-(X - 5) ** 2) + \
np.random.normal(0.0, noise, n_samples)
X = X.reshape((n_samples, 1))
return X, y
X_train, y_train = generate(n_samples=n_train, noise=noise)
X_test, y_test = generate(n_samples=n_test, noise=noise)
from sklearn.tree import DecisionTreeRegressor
reg_tree = DecisionTreeRegressor(max_depth=4, random_state=RANDOM_SEED)
reg_tree.fit(X_train, y_train)
reg_tree_pred = reg_tree.predict(X_test)
plt.figure(figsize=(10, 6))
plt.plot(X_test, f(X_test), "b")
plt.scatter(X_train, y_train, c="b", s=20)
plt.plot(X_test, reg_tree_pred, "g", lw=2)
plt.xlim([-5, 5])
plt.title("Decision tree regressor, MSE = %.2f" % np.sum((y_test - reg_tree_pred) ** 2))
plt.show()
# -
data = pd.read_csv('../Data/petrol_consumption.xls')
data.head()
X = data.drop('Petrol_Consumption', axis=1).values
y = data.Petrol_Consumption.values
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, test_size=0.3)
# +
reg_tree = DecisionTreeRegressor(random_state=42)
# training the tree
reg_tree.fit(X_train, y_train)
y_pred = reg_tree.predict(X_test)
round(np.sqrt(mean_squared_error(y_test, y_pred)), 2)
# -
reg_tree.get_depth()
wine = pd.read_csv('../Data/winequality-red.xls', sep=';')
wine.head()
wine['is_good'] = wine.quality.apply(lambda x: 1 if x >= 6 else 0)
wine.head()
X = wine.drop(['quality', 'is_good'], axis=1).values
y = wine.is_good.values
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, test_size=0.3)
# +
log_reg = LogisticRegression(random_state=42)
# training the reg
log_reg.fit(X_train, y_train)
y_pred_log = log_reg.predict(X_test)
print(f"Decision tree f1-score: {f1_score(y_test, y_pred_log)}.")
reg_tree = DecisionTreeClassifier(random_state=42, max_depth=10)
# training the tree
reg_tree.fit(X_train, y_train)
y_pred_tree = reg_tree.predict(X_test)
print(f"Decision tree f1-score: {f1_score(y_test, y_pred_tree)}.")
# +
ens_trees = BaggingClassifier(random_state=42, n_estimators=1500)
# training the tree
ens_trees.fit(X_train, y_train)
y_pred_ens = ens_trees.predict(X_test)
print(f"Bagging f1-score: {f1_score(y_test, y_pred_ens)}.")
# +
weather = pd.read_csv('../Data/temps_extended.xls')
y = weather['actual']
X = weather.drop(['actual','weekday','month','day','year'],axis =1)
X_train, X_val, y_train, y_val=train_test_split(X,y,test_size=0.3, random_state=42)
# -
weather.head()
from sklearn.ensemble import RandomForestRegressor
from pprint import pprint
rf = RandomForestRegressor(random_state = 42)
rf.fit(X_train, y_train)
y_pred = rf.predict(X_val)
orig_mse = mean_squared_error(y_val, y_pred)
# Look at parameters used by our current forest
print('Параметры по умолчанию:\n')
pprint(rf.get_params())
from sklearn.model_selection import RandomizedSearchCV
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 2000, num = 10)]
max_features = ['auto', 'sqrt']
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
min_samples_split = [2, 5, 10]
min_samples_leaf = [1, 2, 4]
bootstrap = [True, False]
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
rf = RandomForestRegressor(random_state=42)
rf_random = RandomizedSearchCV(estimator=rf, param_distributions=random_grid, n_iter=100,
cv=3, verbose=2, random_state=42, n_jobs=-1)
rf_random.fit(X_train, Y_train)
rf_random.best_params_
# +
rf = RandomForestRegressor(random_state = 42,
n_estimators = 1000,
min_samples_split = 5,
min_samples_leaf = 2,
max_features = 'sqrt',
max_depth = 10,
bootstrap = True)
rf.fit(X_train, y_train)
y_pred = rf.predict(X_val)
changed_mse = mean_squared_error(y_val, y_pred)
# -
round(orig_mse - changed_mse, 1)
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import cross_val_score, train_test_split
from sklearn.ensemble import BaggingClassifier, RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
df = pd.read_csv('../Data/HR-dataset.csv')
np.random.seed(42)
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
target = 'left'
features = df.columns.drop(target)
features = features.drop('empid') # Удалим идентификатор пользователя как нерепрезентативный признак
print(features)
X, y = df[features].copy(), df[target]
# +
salary_ordinals = {'low': 1, 'medium': 2, 'high': 3}
X['dept'] = X['dept'].apply(X['dept'].value_counts().get)
X['salary'] = X['salary'].apply(salary_ordinals.get)
# -
X
scaler = StandardScaler()
X = pd.DataFrame(data=scaler.fit_transform(X), columns=X.columns)
def estimate_accuracy(clf, X, y, cv=5):
return cross_val_score(clf, X, y, cv=5, scoring='accuracy').mean()
tree = DecisionTreeClassifier(max_depth=30)
print("Decision tree:", estimate_accuracy(tree, X, y))
bagging_trees = BaggingClassifier(tree)
print("Decision tree bagging:", estimate_accuracy(bagging_trees, X, y))
# +
random_tree = DecisionTreeClassifier(max_features=int(np.sqrt(len(features))), max_depth=30)
print("Random tree:", estimate_accuracy(random_tree, X, y))
bagging_random_trees = BaggingClassifier(random_tree)
print("Random tree bagging:", estimate_accuracy(bagging_random_trees, X, y))
# -
random_forest = RandomForestClassifier(
n_estimators=100,
max_features=int(np.sqrt(len(features))),
max_depth=30,
oob_score=True,
n_jobs=-1
)
random_forest.fit(X, y)
random_forest.oob_score_.mean()
# +
lr = LogisticRegression(solver='saga', max_iter=200)
lr.fit(X, y)
print("LR:", estimate_accuracy(lr, X, y))
random_logreg = BaggingClassifier(
lr,
n_estimators=10,
n_jobs=-1,
random_state=42
)
print("Bagging for LR:", estimate_accuracy(random_logreg, X, y))
random_logreg = BaggingClassifier(
lr,
n_estimators=10,
n_jobs=-1,
max_features=0.5,
random_state=42
)
print("Bagging for LR:", estimate_accuracy(random_logreg, X, y))
# -
def plot_predictions(X, y, clf, proba=False, points_size=7, xlabel='x', ylabel='y'):
"""Fits the classifier on the data (X, y) and plots the result on a 2-D plane."""
def get_grid(data):
x_std, y_std = data.std(axis=0)
x_min, x_max = data[:, 0].min() - x_std / 2, data[:, 0].max() + x_std / 2
y_min, y_max = data[:, 1].min() - y_std / 2, data[:, 1].max() + y_std / 2
return np.meshgrid(np.linspace(x_min, x_max, num=200),
np.linspace(y_min, y_max, num=200))
clf.fit(X, y)
xx, yy = get_grid(X)
if proba:
predicted = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1].reshape(xx.shape)
else:
predicted = clf.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.figure(figsize=(10.0, 10.0))
plt.pcolormesh(xx, yy, predicted, cmap=plt.cm.coolwarm, alpha=0.1)
plt.scatter(X[:, 0], X[:, 1], c=y, s=points_size, cmap=plt.cm.coolwarm, alpha=0.90)
plt.ylim([yy.min(),yy.max()])
plt.xlim([xx.min(),xx.max()])
plt.xlabel(xlabel)
plt.ylabel(ylabel)
return clf
from sklearn.datasets import load_digits
data = load_digits()
X = data['data']
y = data['target']
from sklearn.model_selection import cross_val_score
# +
tree = DecisionTreeClassifier()
cross_val_score(tree, X, y, cv=10, n_jobs = -1).mean()
# -
tree_bag = BaggingClassifier(tree, n_estimators=100)
cross_val_score(tree_bag, X, y, cv=10, n_jobs = -1).mean()
tree_bag = BaggingClassifier(tree, n_estimators=100, max_features=int(np.sqrt(X.shape[1])))
cross_val_score(tree_bag, X, y, cv=10, n_jobs = -1).mean()
# +
tree = DecisionTreeClassifier(max_features=int(np.sqrt(X.shape[1])))
tree_bag = BaggingClassifier(tree, n_estimators=100)
cross_val_score(tree_bag, X, y, cv=10, n_jobs = -1).mean()
# -
random_forest = RandomForestClassifier(
n_estimators=100,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
random_forest.oob_score_.mean()
random_forest = RandomForestClassifier(
n_estimators=500,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
random_forest.oob_score_.mean()
random_forest = RandomForestClassifier(
n_estimators=800,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
random_forest.oob_score_.mean()
random_forest = RandomForestClassifier(
n_estimators=1000,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
random_forest.oob_score_.mean()
random_forest = RandomForestClassifier(
n_estimators=10,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
random_forest.oob_score_.mean()
# +
random_forest = RandomForestClassifier(
n_estimators=100,
max_depth=5,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
print(random_forest.oob_score_.mean())
random_forest = RandomForestClassifier(
n_estimators=100,
max_features=int(np.sqrt(X.shape[1])),
oob_score=True,
)
random_forest.fit(X, y)
print(random_forest.oob_score_.mean())
# -
-0.5*np.log2(0.5) -0.5*np.log2(0.5)
| module_6/Module_suppl_notebooks/ML-5.Decision_trees + RF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import unittest
class MyTestCase(unittest.TestCase):
def test_1(self):
with self.assertRaises(Exception):
1 + '1'
MyTestCase().test_1()
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm
import pandas as pd
import seaborn as sns
from sklearn import svm
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn import neighbors, datasets
from sklearn.model_selection import cross_val_score
from sklearn.datasets import make_blobs
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import GradientBoostingClassifier
from scipy.spatial import ConvexHull
from tqdm import tqdm
import random
plt.style.use('ggplot')
import pickle
from sklearn import tree
from sklearn.tree import export_graphviz
from joblib import dump, load
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from scipy.interpolate import interp1d
# %matplotlib inline
# +
def getAuc(X,y,test_size=0.25,max_depth=None,n_estimators=100,
minsplit=4,FPR=[],TPR=[],VERBOSE=False, USE_ONLY=None):
'''
get AUC given training data X, with target labels y
'''
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size)
CLASSIFIERS=[DecisionTreeClassifier(max_depth=max_depth, min_samples_split=minsplit,class_weight='balanced'),
RandomForestClassifier(n_estimators=n_estimators,
max_depth=max_depth,min_samples_split=minsplit,class_weight='balanced'),
ExtraTreesClassifier(n_estimators=n_estimators,
max_depth=max_depth,min_samples_split=minsplit,class_weight='balanced'),
AdaBoostClassifier(n_estimators=n_estimators),
GradientBoostingClassifier(n_estimators=n_estimators,max_depth=max_depth),
svm.SVC(kernel='rbf',gamma='scale',class_weight='balanced',probability=True)]
if USE_ONLY is not None:
if isinstance(USE_ONLY, (list,)):
CLASSIFIERS=[CLASSIFIERS[i] for i in USE_ONLY]
if isinstance(USE_ONLY, (int,)):
CLASSIFIERS=CLASSIFIERS[USE_ONLY]
for clf in CLASSIFIERS:
clf.fit(X_train,y_train)
y_pred=clf.predict_proba(X_test)
fpr, tpr, thresholds = metrics.roc_curve(y_test,y_pred[:,1], pos_label=1)
auc=metrics.auc(fpr, tpr)
if VERBOSE:
print(auc)
FPR=np.append(FPR,fpr)
TPR=np.append(TPR,tpr)
points=np.array([[a[0],a[1]] for a in zip(FPR,TPR)])
hull = ConvexHull(points)
x=np.argsort(points[hull.vertices,:][:,0])
auc=metrics.auc(points[hull.vertices,:][x,0],points[hull.vertices,:][x,1])
return auc,CLASSIFIERS
def saveFIG(filename='tmp.pdf',AXIS=False):
'''
save fig for publication
'''
import pylab as plt
plt.subplots_adjust(top = 1, bottom = 0, right = 1, left = 0,
hspace = 0, wspace = 0)
plt.margins(0,0)
if not AXIS:
plt.gca().xaxis.set_major_locator(plt.NullLocator())
plt.gca().yaxis.set_major_locator(plt.NullLocator())
plt.savefig(filename,dpi=300, bbox_inches = 'tight',
pad_inches = 0,transparent=False)
return
# -
NUM=3
df=pd.read_csv('psychoByDiag.csv',index_col=0,sep=',')
# +
df=df[df['DX']>0]
#df=df[df.DX.between(1,2)]
X=df.iloc[:,1:].values
y=df.iloc[:,0].values.astype(str)
#y=[(x=='1')+0 for x in y]
y=[(int(x)<3)+0 for x in y ]
Xdiag=X
# -
Xdiag.shape
ACC=[]
CLFdiag=None
for run in tqdm(np.arange(500)):
auc,CLFS=getAuc(X,y,test_size=0.2,max_depth=NUM,n_estimators=2,
minsplit=2,VERBOSE=False, USE_ONLY=[2])
ACC=np.append(ACC,auc)
if auc > 0.85:
CLFdiag=CLFS
sns.distplot(ACC)
np.median(ACC)
# +
df=pd.read_csv('PSYCHO.DAT',header=None,index_col=0,sep='\s+')
df=df[df[1]>0]
#df=df[df[1].between(1,2)]
X=df.loc[:,2:].values
#y=df.loc[:,1].values.astype(str)
#y=(df.loc[:,1]==1)+0
y=[(df.loc[:,1]<3)+0 for x in y ]
Xpsy=X
# +
df=pd.read_csv('/home/ishanu/Dropbox/scratch_/Qfeatures.csv')
df=df[df.labels>0]
#df=df[df.labels.between(1,2)]
Xq=df.drop('labels',axis=1).values
#y=df.labels.values.astype(str)
X=np.c_[Xpsy,Xq]
#X=Xpsy
#X=np.c_[X,Xdiag]
#X=np.c_[Xpsy,Xdiag]
#X=X1
#X=np.c_[Xpsy,Xdiag]
# -
df.labels.value_counts()
y=(df.labels<3)+0
X.shape
qACC=[]
CLF={}
for run in tqdm(np.arange(2000)):
auc,CLFS=getAuc(X,y,test_size=0.6,max_depth=NUM,n_estimators=2,
minsplit=2,VERBOSE=False, USE_ONLY=[2])
qACC=np.append(qACC,auc)
if auc > 0.8:
CLF[auc]=CLFS
#print('.')
ax=sns.distplot(ACC,label='noq')
sns.distplot(qACC,ax=ax,label='Q')
ax.legend()
np.median(qACC)
#CLF
# +
CLFstar=CLF[np.array([k for k in CLF.keys()]).max()][0]
# -
auc_=[]
ROC={}
fpr_ = np.linspace(0, 1, num=20, endpoint=True)
for run in np.arange(1000):
clf=CLFstar
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5)
y_pred=clf.predict_proba(X_test)
fpr, tpr, thresholds = metrics.roc_curve(y_test,y_pred[:,1], pos_label=1)
f = interp1d(fpr, tpr)
auc_=np.append(auc_,metrics.auc(fpr_, f(fpr_)))
ROC[metrics.auc(fpr, tpr)]={'fpr':fpr_,'tpr':f(fpr_)}
sns.distplot(auc_)
auc_.mean()
# +
# confidence bound calculations
from scipy import interpolate
import subprocess
from sklearn import metrics
xnew = np.arange(0.01, 1, 0.01)
Y=[]
for a in ROC.keys():
#print(a)
#break
x=ROC[a]['fpr']
y=ROC[a]['tpr']
f = interpolate.interp1d(x, y)
ynew = f(xnew)
Y=np.append(Y,ynew)
#plt.plot(x, y, 'o', xnew, ynew, '-')
#break
Y=pd.DataFrame(Y.reshape(int(len(Y)/len(xnew)),len(xnew))).sample(20).transpose()
Y.to_csv('Y.csv',index=None,header=None,sep=' ')
T=0.99
CNFBD="~/ZED/Research/data_science_/bin/cnfbd "
subprocess.call(CNFBD+" -N 5 -f Y.csv -a "+str(T)+" > Y.dat ", shell=True)
Yb=pd.read_csv('Y.dat',header=None,sep=' ',names=['lb','mn','ub'])
Yb['fpr']=xnew
Yb.head()
BND=[metrics.auc(Yb.fpr, Yb.lb),metrics.auc(Yb.fpr, Yb.mn),metrics.auc(Yb.fpr, Yb.ub)]
BND
print(T, '% cnfbnd', BND[0],BND[2], ' mean:', BND[1])
# -
str(BND[1]*100)[:5]
# +
TPR=[]
plt.figure(figsize=[6,5])
for a in ROC.keys():
#print(a)
#break
plt.plot(ROC[a]['fpr'],ROC[a]['tpr'],'-k',alpha=.05)
TPR=np.append(TPR,ROC[a]['tpr'])
TPR=TPR.reshape(int(len(TPR)/len(fpr_)),len(fpr_))
plt.plot(fpr_,np.median(TPR,axis=0),'-r')
metrics.auc(fpr_,np.median(TPR,axis=0))
#plt.gca().set_title('schizophrenia + scheff vs others')
#plt.text(.6,.65,'AUC: '+str(metrics.auc(fpr_,np.median(TPR,axis=0)))[:5],color='r')
#plt.text(.6,.65,'AUC: '+str(93.0)+'%',color='r')
#plt.text(.6,.31,'AUC: '+str(metrics.auc(fpr_,np.median(tprA,axis=0)))[:5],color='b')
#plt.text(.6,.19,'AUC: '+str(metrics.auc(fpr_,np.median(tprB,axis=0)))[:5],color='g')
#plt.gca().set_xlabel('1-specificity')
#plt.gca().set_ylabel('sensitivity')
FS=18
#plt.gca().set_title('schizophrenia or schizoaffective vs others',fontsize=18,y=1.02)
#plt.text(.6,.65,'AUC: '+str(BND[1]*100)[:5]+'%',color='r',fontsize=FS)
#plt.text(.6,.25,'AUC: '+str(metrics.auc(fpr_,np.median(TPR,axis=0)))[:5],color='r')
#plt.text(.6,.31,'AUC: '+str(metrics.auc(fpr_,np.median(tprA,axis=0)))[:5],color='b')
#plt.text(.6,.19,'AUC: '+str(metrics.auc(fpr_,np.median(tprB,axis=0)))[:5],color='g')
FS=18
plt.gca().set_title('AUC: '+str(BND[1]*100)[:5]+'%')
plt.gca().set_ylabel('sensitivity',fontsize=FS,labelpad=10,color='.5')
plt.gca().set_xlabel('1-specificity',fontsize=FS,labelpad=10,color='.5')
plt.gca().tick_params(axis='x', labelsize=FS,labelcolor='.5' )
plt.gca().tick_params(axis='y', labelsize=FS ,labelcolor='.5')
saveFIG(str(NUM)+'sczsceff.pdf',AXIS=True)
# -
#6 0.88795924 0.89931849 mean: 0.89363888
# +
def pickleModel(models,threshold=0.87,filename='model.pkl',verbose=True):
'''
save trained model set
'''
MODELS=[]
for key,mds in models.items():
if key >= threshold:
mds_=mds
MODELS.extend(mds_)
if verbose:
print("number of models (tests):", len(MODELS))
FS=getCoverage(MODELS,verbose=True)
print("Item Use Fraction:", FS.size/(len(MODELS)+0.0))
dump(MODELS, filename)
return MODELS
def loadModel(filename):
'''
load models
'''
return load(filename)
def drawTrees(model):
'''
draw the estimators (trees)
in a single model
'''
N=len(model.estimators_)
for count in range(N):
estimator = model.estimators_[count]
export_graphviz(estimator, out_file=str(NUM)+'_X4X_PSYtree.dot',
#feature_names = iris.feature_names,
#class_names = iris.target_names,
rounded = True, proportion = False,
precision = 2, filled = True)
from subprocess import call
call(['dot', '-Tpng', str(NUM)+'_X4X_PSYtree.dot', '-o', str(NUM)+'_X4X_PSYtree'+str(count)+'.png', '-Gdpi=600'])
#from IPython.display import Image
#Image(filename = 'PSYtree'+str(count)+'.png')
def getCoverage(model,verbose=True):
'''
return how many distinct items (questions)
are used in the model set.
This includes the set of questions being
covered by all forms that may be
generated by the model set
'''
FS=[]
for m in model:
for count in range(len(m.estimators_)):
clf=m.estimators_[count]
fs=clf.tree_.feature[clf.tree_.feature>0]
FS=np.array(list(set(np.append(FS,fs))))
if verbose:
print("Number of items used: ", FS.size)
return FS
# -
models=pickleModel(CLF,threshold=.81,filename='SCH+scffvxmodel_3_2.pkl',verbose=True)
models
drawTrees(models[13])
models[13].estimators_[0].tree_.feature
features=[x for x in models[13].estimators_[0].tree_.feature if x != -2]
models[13].feature_importances_
F=pd.DataFrame([x.feature_importances_[:73] for x in models]).mean()
F=F/F.sum()
Fc=pd.DataFrame([x.feature_importances_[:73] for x in models]).astype(bool).sum()
Fc=Fc/Fc.sum()
dF=pd.DataFrame(F).join(pd.DataFrame(Fc),lsuffix='imp',rsuffix='cnt')
dF=dF.reset_index()
dF.columns=['Item','Importance','Usage']
dF.Item = dF.Item+1
texTable(dF,str(NUM)+'_4_Item_Importance_Usage.tex',INDEX=False)
#F.plot(kind='bar')
dF
# +
FS=18
font = {'size' : FS}
plt.rc('font', **font)
fig=plt.figure(figsize=[8,7])
ax=fig.gca()
ax=sns.scatterplot(ax=ax,x='Usage',y='Importance',data=dF,hue='Item')
FS=18
#plt.gca().set_title('schizophrenia or schizoaffective vs others',fontsize=18,y=1.02)
#plt.text(.6,.65,'AUC: '+str(93.0)+'%',color='r',fontsize=FS)
#plt.text(.6,.25,'AUC: '+str(metrics.auc(fpr_,np.median(TPR,axis=0)))[:5],color='r')
#plt.text(.6,.31,'AUC: '+str(metrics.auc(fpr_,np.median(tprA,axis=0)))[:5],color='b')
#plt.text(.6,.19,'AUC: '+str(metrics.auc(fpr_,np.median(tprB,axis=0)))[:5],color='g')
plt.gca().set_ylabel('Item Importance',fontsize=FS,labelpad=10,color='.5')
plt.gca().set_xlabel('Item Usage Fraction',fontsize=FS,labelpad=10,color='.5')
plt.gca().tick_params(axis='x', labelsize=FS,labelcolor='.5' )
plt.gca().tick_params(axis='y', labelsize=FS ,labelcolor='.5')
plt.gca().set_xlim(0.005,0.035)
plt.gca().set_ylim(0.0,0.045);
saveFIG(str(NUM)+'IMPFRACsczsceff.pdf',AXIS=True)
# +
L=3
features0=[model.estimators_[0].tree_.feature[:L] for model in models]
features1=[model.estimators_[1].tree_.feature[:L] for model in models]
F0=pd.DataFrame(features0)
F0.columns=['x'+str(i) for i in np.arange(L)]
#print(F0.x0.value_counts())
#print(F0.x1.value_counts())
#print(F0.x2.value_counts())
xF0=F0.x0.value_counts()
xF0=xF0[(xF0.index<73) & (xF0.index>=0)].head(10)
xF1=F0.x1.value_counts()
xF1=xF1[(xF1.index<73) & (xF1.index>=0)].head(10)
xF2=F0.x2.value_counts()
xF2=xF2[(xF2.index<73) & (xF2.index>=0)].head(10)
#pd.DataFrame(xF1).merge(pd.DataFrame(xF0))#.join(pd.DataFrame(xF2))
xF0=xF0.reset_index()
xF0.columns=['Item_0','Frequency_0']
xF0.Item_0=xF0.Item_0+1
xF1=xF1.reset_index()
xF1.columns=['Item_1','Frequency_1']
xF1.Item_1=xF1.Item_1+1
xF2=xF2.reset_index()
xF2.columns=['Item_2','Frequency_2']
xF2.Item_2=xF2.Item_2+1
xF0=xF0.join(xF1).join(xF2)
texTable(xF0,str(NUM)+'_4_tab1.tex',INDEX=False)
# +
import pandas as pd
import numpy as np
DUMMY=False
STRA='L{1in}|L{1.25in}|L{1.25in}|L{1.5in}|L{.3in}|L{.3in}'
def texTable(df,tabname='tmp.tex',FORMAT='%1.2f',INDEX=True,DUMMY=DUMMY,USE_l=False):
'''
write latex table
'''
if DUMMY:
return
if INDEX:
df=df.reset_index()
columns=df.columns
df.columns=[x.replace('_','\\_') for x in columns]
for col in df.columns:
if df[col].dtype == 'object':
df[col]=df[col].str.replace('_','\\_')
if USE_l:
TABFORMAT='l'*len(df.columns)
else:
TABFORMAT='L{1in}|'*len(df.columns)
TABFORMAT=TABFORMAT[:-1]
STR='\\begin{tabular}{'+TABFORMAT+'}\n'
with open(tabname,'w') as f:
f.write(STR)
df.to_csv(tabname,float_format=FORMAT,
line_terminator='\\\\\\hline\n',
sep='&',quotechar=' ',index=None,mode='a')
with open(tabname,'a') as f:
f.write('\\end{tabular}\n')
# -
xF0
| code/notebooks/CAD-psych-paper/SCZSAFF/psychosis-Copy4.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,.md//md
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 An introduction to remote services and multi-agent systems
#
# In the previous session, you had an opportunity to experiment hands-on with some neural networks. You may have finished that lab session wondering how neural networks can be built into real robots, particularly low-cost robots rather than expensive cutting-edge robots found only in research labs. In this session, you will find out.
#
# The robot simulator we're using was originally designed to simulate the behaviour of a Lego Mindstorms EV3 controlled robot. The EV3 brick is excellent for introductory robotics, but it has limitations in terms of processor speed and memory capacity. This makes it impractical to train anything other than a small neural network on a Lego EV3 robot, although we may be able to use pre-trained models to perform "on-device" classification tasks.
#
# In general, we are often faced with the problem that we may want to run powerful programs on low-cost hardware that really isn't up to the job. Upgrading a robot with a more powerful processor might not be a solution because it adds cost and may demand extra electrical power or create heat management issues: driving a processor hard can generate a lot of heat, and you need to remove that heat somehow. Heatsinks are heavy and take up physical space, and cooling fans are heavy, take up space, and need access to power. As you add more mass, you need more powerful motors, which themselves add mass and require more power. Bigger batteries add more mass, so you can see where this argument leads...
#
# A possible alternative is to think about using remote services or *multi-agent* systems approaches. In either case, we might use a low-cost robot as a mobile agent to gather data and send that back to a more-powerful computer for processing.
#
# In the first case, we might think of the robot as a remote data collector, collecting data on our behalf and then returning that data to us in response to a request for it. In the RoboLab environment we might think of the notebook Python environment as our local computing environment and the robot as a remote service. Every so often, we might *pull* data from the robot by making a request for a copy of it from the notebook so that we can then analyse the data at our leisure. Alternatively, each time the robot collects a dataset, it might *push* a copy of the data to the notebook's Python environment. In each case, we might think of this as "uploading" data from the simulated robot back to the notebook.
#
# The model is a bit like asking a research librarian for some specific information, the research librarian researching the topic, perhaps using resources you don't have direct access to, and then the research librarian providing you with the information you requested.
#
# In a more dynamic multi-agent case, we might consider the robot and the notebook environment to be acting as peers sending messages as and when they can between each other. For example, we might have two agents: a Lego mobile robot and a personal computer (PC), or the simulated robot and the notebook. In computational terms, *agents* are long-lived computational systems that can deliberate on the actions they may take in pursuit of their own goals based on their own internal state (often referred to as "beliefs") and sensory inputs. Their actions are then performed by means of some sort of effector system that can act to change the state of the environment within which they reside.
#
# In a multi-agent system, two or more agents may work together to combine to perform some task that not only meets the (sub)goals of each individual agent, but that might also strive to attain some goal agreed on by each member of the multi-agent system. Agents may communicate by making changes to the environment, for example, by leaving a trail that other agents may follow (an effect known as *stigmergy*), or by passing messages between themselves directly.
#
# To a limited extent, we may view our simulated robot / Python system as a model for a simple multi-agent system where two agents — the robot, and the classifying neural network or deliberative rule-based system, for example — work together to perform the task of classifying and identifying patterns in some environment that the individual agents could not achieve by themselves. We let the robot do what it does best – move around while logging data – and then it actively sends the data back to the notebook Python environment for processing. The Python environment processes the data using a trained neural network, or perhaps a complex rule-based system, and sends back a message to the robot giving an appropriate response. In each case, the two agents act independently, sending messages to the other party when they have data or a classification to share, rather than just responding to requests for data or a particular service as they occur.
#
# In this session, we will explore how we might use the simulated robot as a remote data collector. Every so often, we will grab a copy of the logged data into the notebook Python environment and analyse it within the notebook.
# + [markdown] tags=["alert-success"]
# *There is quite a lot of provided code in this week's notebooks. You are not necessarily expected to be able to create this sort of code yourself. Instead, try to focus on the process of how various tasks are broken down into smaller discrete steps, as well as how small code fragments can be combined to create "higher-level" functions that perform ever more powerful tasks.*
# -
# ## 1.1 Using a pre-trained MLP to categorise light sensor array data logged from the simulator
#
# The *MNIST_Digits* simulator background includes various digit images from the MNIST dataset, arranged in a grid.
#
# Alongside each digit is a grey square, where the grey level is used to encode the actual label associated with the image. (You can see how the background was created in the `Background Image Generator.ipynb` notebook in the top-level `backgrounds` folder.)
#
# Typically, we use the light sensor to return a single value, such as the reflected light intensity value. However, in this notebook, you will use the light sensor as a simple low-resolution camera. Rather than returning a single value, the sensor returns an array of data containing the values associated individual pixel values from a sampled image. We can then use this square array of pixel data collected by the robot inside the simulator, rather than a single reflected-light value, as the basis for trying to detect what the robot can actually see.
# + [markdown] tags=["alert-danger"]
# *Note that this low resolution camera-like functionality is not supported by the real Lego light sensor.*
# -
# Let's start by loading in the simulator:
# +
from nbev3devsim.load_nbev3devwidget import roboSim, eds
# %load_ext nbev3devsim
# -
# In order to collect the sensor image data, if the simulated robot program `print()` message starts with the word `image_data`, then we can send light sensor array data from the left, right or both light sensors to a datalog in the notebook Python environment.
#
# The `-R` (`--autorun`) switch in the magic at the start of the following code cell will run the program in the simulator once it has been downloaded.
# +
# %%sim_magic_preloaded -b MNIST_Digits -OA -R -x 400 -y 50
# Configure a light sensor
colorLeft = ColorSensor(INPUT_2)
# This is a command invocation rather than a print statement
print("image_data left")
# The command is responded to by
# the "Image data logged..." message display
# -
# As we're going to be collecting data from the simulator into the notebook Python environment, we should take the precaution of clearing the notebook datalog before we start using it:
# %sim_data --clear
# ### 1.1.1 Pushing the sensor array datalog from the simulator to the notebook
#
# We can now run the data collection routine by calling a simple line magic that teleports the robot to a specific location, runs the data collection program (`-R`) and pushes the light sensor array data to the notebook Python environment:
# +
# %sim_magic -RA -x 400 -y 850
# Wait a moment to give data time to synchronise
import time
time.sleep(1)
# -
# We may need to wait a few moments for the program to execute and the data to be sent to the notebook Python environment.
#
# In the current example, the simulator is *pushing* the light sensor array data to the notebook each time the robot sends a particular message to the simulator output window.
#
# With the data pushed from the simulator to the notebook Python environment, we should be able to see a dataframe containing the retrieved data:
roboSim.image_data()
# Each row of the dataframe represents a single captured image from one of the light sensors.
# ### 1.1.2 Previewing the sampled sensor array data (optional)
#
# Having grabbed the data, we can explore the data as rendered images.
# The data representing the image is a long list of RGB (red green, blue) values. We can generate an image from a
# specific row of the dataframe, given the row index:
# +
from nn_tools.sensor_data import generate_image, zoom_img
index = -1 # Get the last image in the dataframe
img = generate_image(roboSim.image_data(),
index, mode='rgb')
zoom_img(img)
# + [markdown] tags=["alert-warning"]
# If you don't see a figure image displayed, check that the robot is placed over a figure by reviewing the sensor array display in the simulator. If the image is there, rerun the previous code cell to see if the data is now available. If it isn't, rerun the data-collecting magic cell, wait a view seconds, and then try to view the zoomed image display.
# -
# We can check the color depth of the image by calling the `.getbands()` method on it:
img.getbands()
# As we might expect from the robot colour sensor, this is a tri-band, RGB image.
#
# Alternatively, we can generate an image directly as a greyscale image, either by setting the mode explicitly or by omitting it (`mode=greyscale` is the default setting):
# +
img = generate_image(roboSim.image_data(), index)
zoom_img(img)
img.getbands()
# -
# The images we trained the network on were size 28 × 28 pixels. The raw images retrieved from the simulator sensor are slightly smaller, coming in at 20 × 20 pixels.
img.size
# The collected image also presents a square profile around the "circular" sensor view. We might thus reasonably decide that we are going to focus our attention on the 14 × 14 square area in the centre of the collected image, with top left pixel `(3, 3)`.
zoom_img(img)
# One of the advantages of using the Python `PIL` package is that a range of *methods* (that is, *functions*) are defined on each image object that allow us to manipulate it *as an image*. (We can then also access the data defining the transformed image *as data* if we need it in that format.)
#
# We can preview the area in our sampled image by cropping the image to the area of interest:
# +
img = generate_image(roboSim.image_data(), index,
crop=(3, 3, 17, 17))
display(img.size)
zoom_img( img )
# + [markdown] tags=["alert-success"]
# If required, we can resize the image by passing the desired size to the `generate_image()` function via the `resize` parameter, setting it either to a specified size, such as `resize=(28, 28)` (that is, 28 × 28 pixels) or back to the original, uncropped image size (`resize=('auto')`)
#
# ```python
#
# img = generate_image(roboSim.image_data(), index,
# crop=(3, 3, 17, 17),
# resize = (28, 28))
# ```
# -
# ### 1.1.3 Collecting multiple sample images
#
# The *MINIST_Digits* simulator background contains a selection of handwritten digit images arranged in a sparse grid on the background which we shall refer to as image sampling point locations. These image locations within the background can be found at the following co-ordinates:
#
# - along rows `100` pixels apart, starting at `x=100` and ending at `x=2000`
# - along columns `100` pixels apart, starting at `y=50` and ending at `y=1050`.
#
# We can collect images from this grid by using magic to teleport the robot to each sampling location and then automatically run the robot program to log the sensor data. For example, to collect images from one column of the background arrangement — that is, images with a particular *x* co-ordinate — we need to calculate the required *y*-values for each sampling point.
#
# To start, let's just check we can generate the required *y*-values:
# +
# Generate a list of integers with desired range and gap
min_value = 50
max_value = 1050
step = 100
list(range(min_value, max_value+1, step))
# -
# To help us keep track of where we are in the sample collection, we can use a visual indicator such as a progress bar.
#
# The `tqdm` Python package provides a wide range of tools for displaying progress bars in Python programs. For example the `tqdm.notebook.trange` function enhances the range iterator with an interactive progress bar that allows us to follow the progress of the iterator:
# +
# Provide a progress bar when iterating through the range
from tqdm.notebook import trange
import time
for i in trange(min_value, max_value, step):
#Wait a moment
time.sleep(0.5)
# -
# We can now create a simple script that will:
#
# - clear the datalog;
# - iterate through the desired *y* locations with a visual indicator of how much progress we have made;
# - use line magic to locate the robot at each step and run the already downloaded image-sampling program.
#
# To access the value of the iterated *y* value in the magic, we need to prefix it with a `$` when we refer to it.
# +
# We need to add a short delay between iterations to give
# the data time to synchronise
import time
# Clear the datalog so we know it's empty
# %sim_data --clear
for _y in trange(min_value, max_value+1, step):
# %sim_magic -R -x 100 -y $_y
# Give the data time to synchronise
time.sleep(1)
# -
# We can view the collected samples via a *pandas* dataframe:
image_data_df = roboSim.image_data()
image_data_df
# We can access a centrally cropped black and white version of an image extracted from the retrieved data by index number (`--index / -i`) by calling the `%sim_bw_image_data` magic, optionally setting the `--threshold / -t` value away from its default value of `127`. Using the `--nocrop / -n` flag will prevent the autocropping behaviour.
# We can convert the image to a black and white image by setting pixels above a specified threshold value to white (`255`), otherwise colouring the pixel black (`0`) using the `generate_bw_image()` function. This will select a row from the datalog at a specific location and optionally crop it to a specific area. Pixel values greater than the threshold will be set to white (`255`) and pixel values equal to or below the threshold will be set to `0`.
# +
from nn_tools.sensor_data import generate_bw_image
index = -1
xx = generate_bw_image(image_data_df,
index,
threshold=100,
crop=(3, 3, 17, 17))
zoom_img(xx)
# -
# The `%sim_bw_image_data` magic performs a similar operation and can also retrieve a random image from the collected data using the `--random / -r` flag. (The crop limits are also assumed by the magic.)
# cropped_bw_image = %sim_bw_image_data --random --threshold 100
zoom_img(cropped_bw_image)
# + [markdown] tags=["alert-success"]
# *End-user applications are simple applications created by users themselves to simplify the performance of certain tasks. Such applications may be brittle and only work in specific situations or circumstances. The code may not be as elegant, well engineered or maintainable as "production code" used in applications made available to other users. One of the advantages of learning to code is the ability to create your own end-user applications.*
# + [markdown] activity=true
# ### 1.1.4 Activity — Observing the effect of changing threshold value when converting the image from a greyscale to a black and white image (optional)
#
# Use the following end-user application to observe the effects of setting different threshold values when creating the black and white binarised version of the image from the original greyscale image data.
# + activity=true
from nn_tools.sensor_data import generate_bw_image
from ipywidgets import interact_manual
@interact_manual(threshold=(0, 255),
index=(0, len(image_data_df)-1))
def bw_preview(index=0, threshold=200,
crop=False):
# Optionally crop to the centre of the image
_crop = (3, 3, 17, 17) if crop else None
_original_img = generate_image(image_data_df, index)
# Generate a black and white image
_demo_img = generate_bw_image(image_data_df, index,
threshold=threshold,
crop=_crop)
# # %sim_bw_image_data --index -1 --threshold 100 --crop 3.3,17,17
zoom_img( _original_img)
zoom_img( _demo_img )
# Preview the actual sized image
# display(_original_img, _demo_img)
# + [markdown] tags=["alert-warning"]
# *The `sensor_image_focus()` function is another convenience function for returning the image in the centre of the sensor array.*
#
# ```python
# from nn_tools.sensor_data import sensor_image_focus
#
# original_image = generate_image(image_data_df, index)
# focal_image = sensor_image_focus( original_image )
# zoom_img( focal_image )
# ```
# -
# ## 1.2 Testing the robot sampled images using a pre-retrained MLP
#
# Having grabbed the image data, we can pre-process it as required and then present it to an appropriately trained neural network to see if the network can identify the digit it represents.
# ### 1.2.1 Loading in a previously saved MLP model
#
# Rather than train a new model, we can load in an MLP we have trained previously. Remember that when using a neural network model, we need to make sure that we know how many inputs it expects, which in our case matches the size of presented images.
#
# You can either use the pre-trained model that is provided in the same directory as this notebook (`mlp_mnist14x14.joblib`), or use your own model created in an earlier notebook.
# +
# Load model
from joblib import load
MLP = load('mlp_mnist14x14.joblib')
# -
# Check the configuration of the MLP:
from nn_tools.network_views import network_structure
network_structure(MLP)
# The 196 input features correspond to an input grid of 14 × 14 pixels.
#
# *For a square array, we get the side length as the square root of the number of features.*
# ### 1.2.2 Using the pre-trained classifier to recognise sampled images
#
# What happens if we now try to recognise images sampled from the simulator light sensor array using our previously trained MLP classifier?
# +
import random
from nn_tools.network_views import image_class_predictor
# Get a random image index value
index = random.randint(0, len(image_data_df)-1)
# Generate the test image as a black and white image
test_image = generate_bw_image(image_data_df, index,
threshold=127,
crop=(3, 3, 17, 17))
# Display a zoomed version of the test image
zoom_img(test_image)
# Print the class prediction report
image_class_predictor(MLP, test_image);
# +
# test_image2 = %sim_bw_image_data --index -1
# Display a zoomed version of the test image
zoom_img(test_image2)
# Print the class prediction report
image_class_predictor(MLP, test_image2);
# -
# How well did the classifier perform?
# + [markdown] student=true
# *Make your own notes and observations about the MLP's performance here. If anything strikes you as unusual, why do you think the MLP is performing the way it is?*
# -
# We can create a simple interactive application to test the other images more easily:
@interact_manual(threshold=(0, 255),
index=(0, len(image_data_df)-1))
def test_image(index=0, threshold=200, show_image=True):
# Create the test image
test_image = generate_bw_image(image_data_df, index,
threshold=threshold,
crop=(3, 3, 17, 17))
# Generate class prediction chart
image_class_predictor(MLP, test_image)
if show_image:
zoom_img(test_image)
# In general, how well does the classifier appear to perform?
# + [markdown] student=true
# *Record your own notes and observations about the behaviour and performance of the MLP here.*
# + [markdown] activity=true
# ### 1.2.3 Activity — collecting image sample data at a specific location
#
# Write a simple line magic command to collect the image data for the handwritten digit centred on the location `(600, 750)`.
#
# Note that you may need to wait a short time between running the data collection program and trying to view it.
#
# Display a zoomed version of the image in the notebook. By observation, what digit does it represent?
#
# Using the `image_class_predictor()` function, how does the trained MLP classify the image? Does this match your observation?
#
# Increase the light sensor noise in the simulator to its maximum value and collect and test the data again. How well does the network perform this time?
#
#
# *Hint: data is collected into a dataframe returned by calling `roboSim.image_data()`.*
#
# *Hint: remember that you need to crop the image to a 14 × 14 array.*
# + student=true
# Your image-sampling code here
# + student=true
# Your image-viewing code here
# + [markdown] student=true
# *From your own observation, record which digit is represented by the image here.*
# + student=true
# How does the trained MLP classify the image?
# + [markdown] student=true
# *How well does the prediction match your observation? Is the MLP confident in its prediction?*
# + [markdown] student=true
# Increase the level of light sensor noise to its maximum value and re-run the experiment:
# + student=true
# Collect data with noise
# + student=true
# Preview image with noise
# + student=true
# Classify image with noise
# + [markdown] student=true
# *Add your own notes and observations on how well the network performed the classification task in the presence of sensor noise here.*
# + [markdown] activity=true heading_collapsed=true
# #### Example discussion
#
# *Click on the arrow in the sidebar or run this cell to reveal an example discussion.*
# + [markdown] activity=true hidden=true
# We can collect the image data by calling the `%sim_magic` with the `-R` switch so that it runs the current program directly. We also need to set the location using the `-x` and `-y` parameters.
# + activity=true hidden=true
# %sim_magic -R -x 600 -y 750
# + [markdown] activity=true hidden=true
# The data is available in a dataframe returned by calling `roboSim.image_data()`.
# + [markdown] activity=true hidden=true
# To view the result, we can zoom the display of the last-collected image in the notebook synched datalog.
# + activity=true hidden=true
# Get data for the last image in the dataframe
index = -1
my_img = generate_bw_image(roboSim.image_data(), index,
crop=(3, 3, 17, 17))
zoom_img(my_img)
# + [markdown] activity=true hidden=true
# By my observation, the digit represented by the image at the specified location is a figure `3`.
#
# The trained MLP classifies the object as follows:
# + activity=true hidden=true
image_class_predictor(MLP, my_img)
# + [markdown] activity=true hidden=true
# This appears to match my prediction.
# -
# ## 1.3 Summary
#
# In this notebook, you have seen how we can use the robot's light sensor as a simple low-resolution camera to sample handwritten digit images from the background. Collecting the data from the robot, we can then convert it to an image and pre-process it before testing it with a pre-trained multi-layer perceptron.
#
# Using captured images that are slightly offset from the centre of the image array essentially provides us with a "jiggled" image, which tends to increase the classification error.
#
# You have also seen how we can automate the way the robot collects image data by "teleporting" the robot to a particular location and then sampling the data there.
#
# In the next notebook, you will see how we can use this automation approach to collect image and class data "in bulk" from the simulator.
| content/08. Remote services and multi-agent systems/08.1 Introducing remote services and multi-agent systems.ipynb |
from traj_py import XPStraj
import bluesky.preprocessors as bpp
import bluesky.plan_stubs as bps
# +
class ScanningExperimentalModule2():
""" the zero for Ry must be set correctly so that Rx is pointing in the x direction
once homed, this position i1s at -6.0
"""
x = EpicsMotor('XF:16IDC-ES:Scan2{Ax:sX}Mtr', name='ss2_x')
x1 = EpicsMotor('XF:16IDC-ES:Scan2{Ax:X}Mtr', name='ss2_x1')
y = EpicsMotor('XF:16IDC-ES:Scan2{Ax:sY}Mtr', name='ss2_y')
z = EpicsMotor('XF:16IDC-ES:InAir{Mscp:1-Ax:F}Mtr', name='focus')
# this is the Standa stepper stage
rx = EpicsMotor('XF:16IDC-ES:Scan2{Ax:RX1}Mtr', name='ss2_rx')
ry = EpicsMotor('XF:16IDC-ES:Scan2{Ax:RY}Mtr', name='ss2_ry')
ss2 = ScanningExperimentalModule2()
xps_trj = XPStraj('10.16.2.100', 'scan', 'test')
# +
xps_trj_motors = {'scan.rY': ss2.ry,
'scan.Y': ss2.y,
'scan.X': ss2.x}
DETS = [pil1M_ext, pilW1_ext, pilW2_ext, em1, em2]
# -
def raster(detectors, exp_time, fast_axis, f_start, f_end, Nfast,
slow_axis=None, s_start=0, s_end=0, Nslow=1, monitors=[em1, em2], md=None):
""" raster scan in fly mode using detectors with exposure time of exp_time
detectors must be a member of pilatus_detectors_ext
fly on the fast_axis, step on the slow_axis, both specified as Ophyd motors
the fast_axis must be one of member of xps_trj.motors, for now this is hard-coded
the specified positions are relative to the current position
for the fast_axis are the average positions during detector exposure
use it within the run engine: RE(raster(...))
"""
if not set(detectors).issubset(pilatus_detectors_ext):
raise Exception("only pilatus_detectors_ext can be used in this raster scan.")
if fast_axis not in xps_trj_motors.values():
raise Exception("the fast_axis is not supported in this raster scan: ", fast_axis.name)
fast_axis_name = list(xps_trj_motors.keys())[list(xps_trj_motors.values()).index(fast_axis)]
#
step_size = (f_end-f_start)/(Nfast-1)
dt = exp_time + 0.005 # exposure_period is 5ms longer than exposure_time, as defined in Pilatus
xps_trj.define_traj(fast_axis_name, Nfast-1, step_size, dt)
p0_fast = fast_axis.position
ready_pos = {}
ready_pos[True] = p0_fast+f_start-xps_trj.traj_par['rampup_distance']-step_size/2
ready_pos[False] = p0_fast+f_end+xps_trj.traj_par['rampup_distance']+step_size/2
if slow_axis is not None:
p0_slow = slow_axis.position
pos_s = p0_slow+np.linspace(s_start, s_end, Nslow)
else:
Nslow = 1
pilatus_ct_time(exp_time)
set_pil_num_images(Nfast*Nslow)
print('setting up to collect %d exposures of %.2f sec ...' % (Nfast*Nslow, exp_time))
motor_names = [fast_axis.name, slow_axis.name]
motors = [fast_axis, slow_axis]
scan_shape = [Nfast, Nslow]
_md = {'shape': tuple(scan_shape),
'plan_args': {'detectors': list(map(repr, detectors))},
'plan_name': 'raster',
'plan_pattern': 'outer_product',
'motors': tuple(motor_names),
'hints': {},
}
_md.update(md or {})
_md['hints'].setdefault('dimensions', [(('time',), 'primary')])
@bpp.stage_decorator([xps_trj] + detectors)
@bpp.run_decorator(md=_md)
@fast_shutter_decorator()
def inner(fast_axis, ready_pos, slow_axis, Nslow, pos_s):
running_forward = True
for i in range(Nslow):
if slow_axis is not None:
yield from mov(fast_axis, ready_pos[running_forward], slow_axis, pos_s[i])
else:
yield from mov(fast_axis, ready_pos[running_forward])
yield from wait()
xps_trj.select_forward_traj(running_forward)
yield from bps.kickoff(xps_trj, wait=True)
yield from bps.complete(xps_trj, wait=True)
yield from bps.collect(xps_trj)
running_forward = not running_forward
#yield from bps.open_run()
#for mo in monitors:
# yield from bps.monitor(mo)
yield from inner(fast_axis, ready_pos, slow_axis, Nslow, pos_s)
#for mo in monitors:
# yield from bps.unmonitor(mo)
#yield from bps.close_run()
if slow_axis is not None:
yield from mov(fast_axis, p0_fast, slow_axis, p0_slow)
else:
yield from mov(fast_axis, p0_fast)
for m in raster([pil1M_ext], 0.2, ss2.x, -0.1, 0.1, 21, ss2.y, -0.1, 0.1, 11):
print(m)
login('test', 'test', 'test')
RE(raster([pil1M_ext, pilW1_ext, pilW2_ext], 0.2, ss2.x, -0.1, 0.1, 21, ss2.y, -0.1, 0.1, 11))
hdr = db[-1]
hdr.fields()
hdr.table(stream_name='test')
hdr.descriptors
| tests/flyscaning_2018Nov14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Anonymizing data with Faker
from pprint import pprint
from faker import Faker
from faker.providers import internet
# "Faker is a Python package that generates fake data for you. Whether you need to bootstrap your database, create good-looking XML documents, fill-in your persistence to stress test it, or anonymize data taken from a production service, Faker is for you."
#
# https://faker.readthedocs.io/en/master/
# "Use `faker.Faker()` to create and initialize a faker generator, which can generate data by accessing properties named after the type of data you want."
fake = Faker()
# Pour créer un faux nom.
print(fake.name())
# Pour créer une fausse adresse.
print(fake.address())
# Il est possible de créer plusieurs faux noms, ici en français, avec une boucle.
fake_fr = Faker('fr_FR')
for _ in range(10):
print(fake_fr.name())
# Même chose en néerlandais.
fake_nl = Faker('nl_NL')
for _ in range(10):
print(fake_nl.name())
# Nous pouvons créer une fausse adresse IP.
fake.add_provider(internet)
print(fake.ipv4_public())
# Nous pouvons créer un faux profil : adresse, date de naissance, e-mail, nom, genre, et username.
profile = fake_fr.simple_profile()
pprint(profile)
| module5/s2_anonymization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import gdxpds
import os
south_share = 0.75; #65
freq = '150Min'; #125
variance = 0.25; #0.35
# +
# scale renewable profiles
res = pd.read_excel('in_48h.xlsx', sheet_name = 'timeseries')
res = res.iloc[2:27, 1:9]
res.columns = ['int', 'load_north', 'load_south', 'int_2', 'Wind_north', 'Solar_north', 'Wind_south', 'Solar_south']
res.index = pd.to_timedelta(res.index-2, 'hours')
for c in res.columns:
res[c] = res[c].astype(float, errors = 'raise')
renewables = res.resample(freq).mean().round(2)
combined = renewables[['load_north', 'load_south', 'Wind_north', 'Solar_north', 'Wind_south', 'Solar_south']]
#load profile
load = pd.read_pickle('2015-2020_load_generation.pkl')
df = load['DE']
df = df.loc[df.index.year == 2020]
load_curve = df.groupby(df.index.hour).mean()
load_curve.index = pd.to_timedelta(load_curve.index, 'hours')
series = load_curve.resample(freq).mean()
#generic load profile
#series = pd.Series(np.ones(series.shape), series.index, name=series.name) * round(series.mean(),-3)
load = pd.DataFrame(data = series.values, index = series.index, columns = ['total load'])
np.random.seed([3,14])
load['north'] = load['total load'] *((1-south_share) + np.random.uniform(-variance, variance, len(load)))
load['north'] = [max(0,i) for i in load['north']]
load['south'] = load['total load'] - load['north']
load = load.round(-1)
# --- SAVE LOAD DATA ---
load = load.reset_index()[['north','south']]
load.index = load.index.rename('t')
load.index = load.index+1
load.to_excel('load.xlsx')
load = load.stack().reset_index();
load.columns = ['t','n','value'];
data_ready_for_GAMS = { 'i_load': load }
gdx_file = '../Two node models/load.gdx'
gdx = gdxpds.to_gdx(data_ready_for_GAMS, gdx_file)
# --- SAVE RES DATA
north = renewables[['Wind_north', 'Solar_north']]
north.columns = ['wind', 'solar']
north.loc[:,'n'] = 'north'
north.loc[:,'t'] = range(1,len(north)+1)
south = renewables[['Wind_south', 'Solar_south']]
south.columns = ['wind', 'solar']
south.loc[:,'n'] = 'south'
south.loc[:,'t'] = range(1,len(north)+1)
re = north.append(south);
re = re.iloc[:, [3, 2, 0, 1]]
re = re.set_index(['t','n'])
re = re.unstack()
re.to_excel('avail.xlsx')
tmp = pd.DataFrame(re.stack().stack())
tmp.index.names = ['t','n','tec']
tmp = tmp.reset_index()
tmp = tmp.iloc[:, [0, 2, 1, 3]]
tmp = tmp.rename(columns = {0:'value'})
data_ready_for_GAMS = { 'i_avail': tmp }
gdx_file = '../Two node models/avail.gdx'
gdx = gdxpds.to_gdx(data_ready_for_GAMS, gdx_file)
# +
#os.system(str('cmd /k "gams ../Two node models/nodal.gms"'))
# -
load
tmp
load.loc[load['n']=='north']['value'].std()
# # OLD
new_frequency['south'] / new_frequency['total load']
#RES data
res = pd.read_excel('in.xlsx', sheet_name = 'timeseries')
res = res.iloc[2:27, 5:9,]
res.index = pd.to_timedelta((res.index-2), 'hours')
res.columns = [['Wind_north', 'Solar_north', 'Wind_south', 'Solar_south']]
res
res = res.resample(freq).mean().iloc[:-1,:]
new_frequency[['Wind_north', 'Solar_north', 'Wind_south', 'Solar_south']] = res
new_frequency
new_frequency.to_csv('12_load.csv')
new_frequency.plot()
# +
import numpy as np
import matplotlib.pyplot as plt
# Create some mock data
fig, ax1 = plt.subplots()
ax1.set_xlabel('time')
ax1.set_ylabel('Availability (in %)')
ax1.plot(new_frequency.index, new_frequency[['Wind_north', 'Solar_north', 'Wind_south', 'Solar_south']])
ax1.tick_params(axis='y')
ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
ax2.set_ylabel('Load (in MW)')
ax2.plot(new_frequency.index, new_frequency[['north','south']], color = 'black')
ax2.tick_params(axis='y')
ax2.set_ylim([0, 65000])
#ax2.ylim[0,max(new_frequency['south'])+ 1000]
fig.tight_layout() # otherwise the right y-label is slightly clipped
plt.show()
# -
| Input data/.ipynb_checkpoints/Input_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python36
# language: python
# name: python36
# ---
# +
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
import matplotlib
matplotlib.use("Agg") #Needed to save figures
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
import xgboost as xgb
from sklearn.metrics import roc_auc_score
training = pd.read_csv("/Users/szkfzx/datasets/santander-customer-satisfaction/train.csv")
test = pd.read_csv("/Users/szkfzx/datasets/santander-customer-satisfaction/test.csv")
print(training.shape)
print(test.shape)
# Replace -999999 in var3 column with most common value 2
# See https://www.kaggle.com/cast42/santander-customer-satisfaction/debugging-var3-999999
# for details
training = training.replace(-999999,2)
# Replace 9999999999 with NaN
# See https://www.kaggle.com/c/santander-customer-satisfaction/forums/t/19291/data-dictionary/111360#post111360
# training = training.replace(9999999999, np.nan)
# training.dropna(inplace=True)
# Leads to validation_0-auc:0.839577
X = training.iloc[:,:-1]
y = training.TARGET
# Add zeros per row as extra feature
X['n0'] = (X == 0).sum(axis=1)
# # Add log of var38
# X['logvar38'] = X['var38'].map(np.log1p)
# # Encode var36 as category
# X['var36'] = X['var36'].astype('category')
# X = pd.get_dummies(X)
# Add PCA components as features
from sklearn.preprocessing import normalize
from sklearn.decomposition import PCA
X_normalized = normalize(X, axis=0)
pca = PCA(n_components=2)
X_pca = pca.fit_transform(X_normalized)
X['PCA1'] = X_pca[:,0]
X['PCA2'] = X_pca[:,1]
from sklearn.feature_selection import SelectPercentile
from sklearn.feature_selection import f_classif,chi2
from sklearn.preprocessing import Binarizer, scale
p = 86 # 308 features validation_1-auc:0.848039
p = 80 # 284 features validation_1-auc:0.848414
p = 77 # 267 features validation_1-auc:0.848000
p = 75 # 261 features validation_1-auc:0.848642
# p = 73 # 257 features validation_1-auc:0.848338
# p = 70 # 259 features validation_1-auc:0.848588
# p = 69 # 238 features validation_1-auc:0.848547
# p = 67 # 247 features validation_1-auc:0.847925
# p = 65 # 240 features validation_1-auc:0.846769
# p = 60 # 222 features validation_1-auc:0.848581
X_bin = Binarizer().fit_transform(scale(X))
selectChi2 = SelectPercentile(chi2, percentile=p).fit(X_bin, y)
selectF_classif = SelectPercentile(f_classif, percentile=p).fit(X, y)
chi2_selected = selectChi2.get_support()
chi2_selected_features = [ f for i,f in enumerate(X.columns) if chi2_selected[i]]
print('Chi2 selected {} features {}.'.format(chi2_selected.sum(),
chi2_selected_features))
f_classif_selected = selectF_classif.get_support()
f_classif_selected_features = [ f for i,f in enumerate(X.columns) if f_classif_selected[i]]
print('F_classif selected {} features {}.'.format(f_classif_selected.sum(),
f_classif_selected_features))
selected = chi2_selected & f_classif_selected
print('Chi2 & F_classif selected {} features'.format(selected.sum()))
features = [ f for f,s in zip(X.columns, selected) if s]
print (features)
X_sel = X[features]
X_train, X_test, y_train, y_test = train_test_split(X_sel, y, random_state=1301, stratify=y, test_size=0.4)
# xgboost parameter tuning with p = 75
# recipe: https://www.kaggle.com/c/bnp-paribas-cardif-claims-management/forums/t/19083/best-practices-for-parameter-tuning-on-models/108783#post108783
ratio = float(np.sum(y == 1)) / np.sum(y==0)
# Initial parameters for the parameter exploration
# clf = xgb.XGBClassifier(missing=9999999999,
# max_depth = 10,
# n_estimators=1000,
# learning_rate=0.1,
# nthread=4,
# subsample=1.0,
# colsample_bytree=0.5,
# min_child_weight = 5,
# scale_pos_weight = ratio,
# seed=4242)
# gives : validation_1-auc:0.845644
# max_depth=8 -> validation_1-auc:0.846341
# max_depth=6 -> validation_1-auc:0.845738
# max_depth=7 -> validation_1-auc:0.846504
# subsample=0.8 -> validation_1-auc:0.844440
# subsample=0.9 -> validation_1-auc:0.844746
# subsample=1.0, min_child_weight=8 -> validation_1-auc:0.843393
# min_child_weight=3 -> validation_1-auc:0.848534
# min_child_weight=1 -> validation_1-auc:0.846311
# min_child_weight=4 -> validation_1-auc:0.847994
# min_child_weight=2 -> validation_1-auc:0.847934
# min_child_weight=3, colsample_bytree=0.3 -> validation_1-auc:0.847498
# colsample_bytree=0.7 -> validation_1-auc:0.846984
# colsample_bytree=0.6 -> validation_1-auc:0.847856
# colsample_bytree=0.5, learning_rate=0.05 -> validation_1-auc:0.847347
# max_depth=8 -> validation_1-auc:0.847352
# learning_rate = 0.07 -> validation_1-auc:0.847432
# learning_rate = 0.2 -> validation_1-auc:0.846444
# learning_rate = 0.15 -> validation_1-auc:0.846889
# learning_rate = 0.09 -> validation_1-auc:0.846680
# learning_rate = 0.1 -> validation_1-auc:0.847432
# max_depth=7 -> validation_1-auc:0.848534
# learning_rate = 0.05 -> validation_1-auc:0.847347
#
clf = xgb.XGBClassifier(missing=9999999999,
max_depth = 5,
n_estimators=1000,
learning_rate=0.1,
nthread=4,
subsample=1.0,
colsample_bytree=0.5,
min_child_weight = 3,
scale_pos_weight = ratio,
reg_alpha=0.03,
seed=1301)
clf.fit(X_train, y_train, early_stopping_rounds=50, eval_metric="auc",
eval_set=[(X_train, y_train), (X_test, y_test)])
print('Overall AUC:', roc_auc_score(y, clf.predict_proba(X_sel, ntree_limit=clf.best_iteration)[:,1]))
test['n0'] = (test == 0).sum(axis=1)
# test['logvar38'] = test['var38'].map(np.log1p)
# # Encode var36 as category
# test['var36'] = test['var36'].astype('category')
# test = pd.get_dummies(test)
test_normalized = normalize(test, axis=0)
pca = PCA(n_components=2)
test_pca = pca.fit_transform(test_normalized)
test['PCA1'] = test_pca[:,0]
test['PCA2'] = test_pca[:,1]
sel_test = test[features]
y_pred = clf.predict_proba(sel_test, ntree_limit=clf.best_iteration)
submission = pd.DataFrame({"ID":test.index, "TARGET":y_pred[:,1]})
submission.to_csv("submission.csv", index=False)
submission
| Kaggle/Playgroud/Satisfaction/0-84-score-with-36-features-only.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SIR Modeling in Python, based on Numberphile
#
# This notebook is designed to be a bare-bones introudction to SIR modeling in Python.
#
# I am using the example from Numberphile video: https://www.youtube.com/watch?v=k6nLfCbAzgo where <NAME> discusses simple ODE modeling in Geogebra using SIR models as the example.
# There is a Geogebra workbook demonstrating this, as in the video, here: https://www.geogebra.org/classic/aqpv5df7
#
# also taking examples from https://scipython.com/book/chapter-8-scipy/additional-examples/the-sir-epidemic-model/
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
import pandas as pd
# In the numberphile example, Ben uses transm and recov for beta and gamma, so I have replaced them here. If you want to set it back, just change all transm to beta and all recov to gamma.
# +
N = 1
Istart = 0.01
Sstart = N - Istart
Rstart = 0
transm = 3.25
recov = 0.30
maxT = 20
t = np.linspace(0, maxT, maxT)
# The SIR model differential equations.
def deriv(SIR, t, N, transm, recov):
S, I, R = SIR
dSdt = -transm * S * I
dIdt = transm * S * I - recov * I
dRdt = recov * I
return dSdt, dIdt, dRdt
# Initial conditions vector
SIR0 = Sstart, Istart, Rstart
# Integrate the SIR equations over the time grid, t.
ret = odeint(deriv, SIR0, t, args=(N, transm, recov))
S, I, R = ret.T
# -
plotData = pd.DataFrame(ret.T)
plotData = plotData.transpose()
plotData.columns = ['S','I','R']
with pd.plotting.plot_params.use('x_compat',True):
plotData['S'].plot(color='g')
plotData['I'].plot(color='r')
plotData['R'].plot(color='b')
| SIRModeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# name: python3
# ---
# ## Importing Libraries & getting Data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('dataset/fake_job_postings.csv')
data.head()
data.info()
data.describe()
data.shape
# ## Feature Selection
data.columns
data = data[['title', 'location', 'company_profile', 'requirements', 'telecommuting', 'has_company_logo', 'has_questions', 'employment_type',
'required_experience', 'required_education', 'industry', 'function', 'salary_range','fraudulent']]
# ## Handling missing values & Outliers
# checking for duplicates
data.duplicated().sum()
data.drop_duplicates(inplace=True)
data.duplicated().sum()
# +
data_num = data[['telecommuting','has_company_logo','has_questions','fraudulent','salary_range']]
data_categ = data[['title', 'location', 'company_profile', 'requirements', 'employment_type','required_experience', 'required_education', 'industry', 'function']]
# -
plt.figure(figsize=(16,8))
sns.boxplot(data=data)
# plt.xticks(rotation=30)
plt.show()
# +
# removing outliers
data_num = data_num[data_num['telecommuting'] < 0.9]
data_num = data_num[data_num['has_company_logo'] > 0.1]
data_num = data_num[data_num['fraudulent'] < 0.9]
data_num
# -
data.isnull().sum()
data.dropna(axis=0, how='any', inplace=True)
data.isnull().sum()
data.shape
# ## EDA
data_categ.columns, data_num.columns
plt.hist(data.employment_type, color='red', edgecolor='black', alpha=0.7)
plt.xlabel('\nEmployment Type')
plt.show()
plt.hist(data.required_experience, color='blue', edgecolor='black', alpha=0.7)
plt.xlabel('\nRequired Experience')
plt.xticks(rotation=90)
plt.show()
plt.hist(data.fraudulent, color='green', edgecolor='black', alpha=0.7)
plt.xlabel('\nFrauds')
plt.show()
plt.figure(figsize=(10,5))
sns.set_style('darkgrid')
sns.countplot(x='function' ,data=data)
plt.xticks(rotation=90)
plt.show()
data_jobs_fulltime = data[(data['employment_type'] == 'Full-time') & (data['fraudulent'] == 0)]
data_jobs_fulltime.shape
# +
data_jobs_fulltime['title'].value_counts()
# Agent-Inbound Sales Position has the most opportunities (12) of a full-time job with no frauds.
# -
data_industry_mostfake = data[data['fraudulent'] == 1]
data_industry_mostfake.shape
# +
data_industry_mostfake['industry'].value_counts()
# Oil & Energy is the industry with the most fake jobs(19), followed by Real Estate(10).
# -
# ## Getting Balanced Dataset
data['fraudulent'].value_counts()
# +
fraud_jobs = data[data['fraudulent'] == 1]
real_jobs = data[data['fraudulent'] == 0]
fraud_jobs.shape ,real_jobs.shape
# +
# oversampling 'fraud' dataframe for getting balanced dataset
fraud_jobs = fraud_jobs.sample(1403 ,replace=True)
fraud_jobs.shape, real_jobs.shape
# -
data = fraud_jobs.append(real_jobs)
data.reset_index()
# ## Encoding
data_categ.columns
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
data['title'] = le.fit_transform(data['title'])
data['location'] = le.fit_transform(data['location'])
data['company_profile'] = le.fit_transform(data['company_profile'])
data['requirements'] = le.fit_transform(data['requirements'])
data['employment_type'] = le.fit_transform(data['employment_type'])
data['required_experience'] = le.fit_transform(data['required_experience'])
data['required_education'] = le.fit_transform(data['required_education'])
data['industry'] = le.fit_transform(data['industry'])
data['function'] = le.fit_transform(data['function'])
data['salary_range'] = le.fit_transform(data['salary_range'])
data.reset_index()
data.head()
data.shape
# # Model Building
# +
X = data[['title', 'location', 'company_profile', 'requirements','telecommuting', 'has_company_logo', 'has_questions', 'employment_type',
'required_experience', 'required_education', 'industry', 'function',
'salary_range']].values
y = data['fraudulent'].values
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.33)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# -
# ## Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression(solver='lbfgs', max_iter=3000)
lr.fit(X_train ,y_train)
y_predict_lr = lr.predict(X_test)
# -
from sklearn.metrics import accuracy_score
print('Accuracy score using Logistic Regression :',accuracy_score(y_test ,y_predict_lr))
# ## KNN
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
# +
# choosing best number of neighbors for our model
neighbors_with_accuracies = {}
for i in range(1,20):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train ,y_train)
y_pred_knn = knn.predict(X_test)
acc = accuracy_score(y_test ,y_pred_knn)
neighbors_with_accuracies[i] = acc
# -
neighbors_with_accuracies
max(neighbors_with_accuracies.values())
knn = KNeighborsClassifier(n_neighbors=2)
knn.fit(X_train, y_train)
y_predict_knn = knn.predict(X_test)
print("Accuracy Score using KNN:", accuracy_score(y_test, y_predict_knn))
# ## Random Forest
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier()
rf.fit(X_train ,y_train)
y_predict_rf = rf.predict(X_test)
print("Accuracy Score using Random Forest:", accuracy_score(y_test, y_predict_rf))
# # Model Evaluation
# +
# creating Dataframe to check which model was the best
models = pd.DataFrame({
'Model': ['Logistic Regression', 'KNN', 'Random Forest'],
'Score': [accuracy_score(y_test, y_predict_lr),
accuracy_score(y_test, y_predict_knn),
accuracy_score(y_test, y_predict_rf),
]
})
models.sort_values(by='Score', ascending=False)
| Kaggle Datasets/ML/Real or Fake Jobs/fakejobs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python List Iteration
# A variety of ways to iterate Lists, including for loop, while loop, enumerate.
# ----
# The standard for loop works well if it is used inside the loop you only need the item and not its index.
# +
letters = ['a', 'b', 'c', 'd', 'e']
for letter in letters:
print(letter)
# -
# ----
# If you need the index inside the loop you can use range(len(list)).
# Then you can always get the list item if needed by using the index.
for index in range(len(letters)):
print('letters', index, '=', letters[index])
# ----
# Best option if you need both index and item inside the loop is to use Python's **enumerate** function.
# Enumerate works in both Python 2.x and 3.x
for index, item in enumerate(letters):
print('letters', index, '=', item)
# Enumerate actually returns an iterable enumerate object,
# which is a sequence of tuples of (index, item).
enum_obj = enumerate(letters)
print(next(enum_obj))
print(next(enum_obj))
print(type(enum_obj))
# ----
# Probably the clumsiest way to iterate a list in Python -- the **while loop**.
# Requires index initialization before list, and incrementation inside loop.
index = 0
while index < len(letters):
print('letters', index, '=', letters[index])
index += 1
| Python Programming/10. Extras/Python List Iteration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import _pickle as pkl
import pandas as pd
import os,sys,glob, h5py
from dlab import reach_align_pipeline as rf
from dlab.generalephys import placeAxesOnGrid, cleanAxes
from dlab.continuous_traces import gaussian_filter1d
from scipy.signal import find_peaks
from scipy.stats import pearsonr, spearmanr, zscore
from itertools import combinations
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import seaborn as sns
sns.set_style('white')
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings('ignore')
# -
# # import ephys and timestamp info
# +
#import dataframe with information for all mice (dailey- should be on your desktop in kimdata named dfexcept28_ypos.json)
data_path = '/Users/kimberlygagnon/Documents/CU/Welle Lab/experiments/daileydata/dfexcept28_ypos.json'
df = pd.read_json(data_path)
ts_path = '/Users/kimberlygagnon/Documents/CU/Welle Lab/experiments/daileydata/finalized/df_ts.json'
df_ts = pd.read_json(data_path)
p_start = '/Users/kimberlygagnon/Documents/CU/Welle Lab/experiments/daileydata/finalized/Neuropixels_ProcessorStartTimes.csv'
df_start = pd.read_csv(p_start)
# -
# # initialize dataframe with behavior alignments
df_align = pd.DataFrame(df,columns = ['times','ypos','mouse','cohort','cell','overall_rate','baseline_fr','reach_fr','waveform_class'])
# # import curated behavior file
#import curated behavior for specific mouse (save xls as csv)
reach_path = '/Users/kimberlygagnon/Documents/CU/Welle Lab/experiments/daileydata/finalized/Curated_Video_Timestamps/20200804_unit00_session001_NPCE_026.csv'
df_reaches = pd.read_csv(reach_path)
#remove random nans
df_reaches = df_reaches.dropna(axis = 1 , how = 'all')
df_reaches = df_reaches.dropna(axis = 0 , how = 'all')
df_reaches.head()
# # synch behavior and create relevant dataframes
for mouse in df.mouse.unique():
df_ = df[df.mouse==mouse]
#align software with ephys
soft_start = df_start[df_start.Mouse_number==mouse].Processor_Start_time
frame_ts = df_ts[df_ts.mouse==mouse].timestamps
#WRITE CODE THAT LOADS RELEVANT CURATED BEHAVIOR EXCEL SPREADSHEET SAVED AS CSV
reach_times = synchphys(soft_start,frame_ts) #gets reach times
df_reaches['rMax_t']= reach_times
df_reaches = df_reaches[df_reaches.behaviors!='none'][df_reaches.behaviors!='arm_movement']
#in below code can look at success vs. failure reaches by inputing df_reaches[df_reaches=='success'] or 'x_failure'
reaches = rf.epochfr(df_,df_reaches)
df_align['epoch_fr'] = reaches
baseline = rf.epochfr(df_,df_reaches,start = 1.0,end = -0.5,binsize)
df_align['baseline_fr'] = baseline
ave_reach,normedbins,ave_reach_ = rf.reachave_tensor(df_,df_reaches) #gets -4 to +2 epoch for plotting heatmap as well as tensor for PCA
df_align['bin_ave'] = ave_reach
df_align['norm_bin_ave'] = normedbins
traj = rf.center(reach_ave_)
p,ev = rf.pca(traj)
df_align['expl_var'] = ev
df_align['pcs'] = p
smooth_pcs = gaussian_filter1d(p, sigma=8)
df_align['smooth_pcs'] = smooth_pcs
#need to concat df_align with larger dataframe in for loop?? to make complete
| reach_align_batch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.api as sm
import scipy.stats as stats
import sys
sys.path.append("../")
import vuong_tests3
# +
def compute_loglike(resid):
sigma = np.sqrt(np.sum(resid**2)/resid.shape[0])
ll = np.log(stats.norm.pdf(resid,loc=0,scale=sigma))
return ll
def compute_score(yn,xn,params):
xn = sm.add_constant(xn)
resid = yn - np.matmul(xn,params)
k = len(params)
scale = (resid**2).mean()
tile_resid = np.tile( resid, k)
tile_resid = np.reshape(tile_resid, (k,xn.shape[0]) ).transpose()
grad = tile_resid*xn/scale
return grad
def compute_hess(yn,xn,params):
pass
def setup_shi(yn,xn,return_model=False,num_params=15):
x1n,x2n = xn[:,0],xn[:,1:num_params]
# model 1 grad, etc.
model1 = sm.OLS(yn,sm.add_constant(x1n))
model1_fit = model1.fit(disp=False)
params1 = (model1_fit.params)
ll1 = compute_loglike(model1_fit.resid)
grad1 = compute_score(yn,x1n,model1_fit.params)
hess1 = model1.hessian(model1_fit.params)
#model 2 grad, etc.
model2 = sm.OLS(yn,sm.add_constant(x2n))
model2_fit = model2.fit(disp=False)
params2 = (model2_fit.params)
ll2 = compute_loglike(model2_fit.resid)
grad2 = compute_score(yn,x2n,model2_fit.params)
hess2 = model2.hessian(model2_fit.params)
if return_model:
return ll1,grad1,hess1,params1,model1,ll2,grad2,hess2,params2,model2
return ll1,grad1,hess1,params1,ll2,grad2,hess2,params2
# -
# # a = .25
# +
def gen_data(nobs=1000, a=0.25, num_params=15):
x = np.random.normal(scale=3., size=(nobs,1+num_params))
e = np.random.normal(loc=0.0, scale=1.0, size=nobs)
y = 1 + a*x[:,0] + a/np.sqrt(num_params)*x[:,1:num_params+1].sum(axis=1) + e
return y,x,nobs
gen_data_ex = lambda : gen_data(nobs=1000, a=0.25, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
# -
gen_data_ex = lambda : gen_data(nobs=500, a=0.25, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
gen_data_ex = lambda : gen_data(nobs=250, a=0.25, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
# # a = .125
gen_data_ex = lambda : gen_data(nobs=1000, a=0.125, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
gen_data_ex = lambda : gen_data(nobs=500, a=0.125, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
gen_data_ex = lambda : gen_data(nobs=250, a=0.125, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
# # a = 0
gen_data_ex = lambda : gen_data(nobs=1000, a=0.0, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
gen_data_ex = lambda : gen_data(nobs=500, a=0.0, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
gen_data_ex = lambda : gen_data(nobs=250, a=0.0, num_params=15)
mc_out = vuong_tests3.monte_carlo(1000,gen_data_ex,setup_shi)
vuong_tests3.print_mc(mc_out)
print(mc_out)
# # other weird DGPs
# +
def gen_data(beta=0):
nobs = 250
#x = np.random.normal(low=-3., high=3., size=(nobs,3))
x = np.random.normal(scale=3., size=(nobs,4))
e = np.random.normal(loc=0.0, scale=1.0, size=nobs)
y = 1 + 1/np.sqrt(2)*x[:,3] + 1/np.sqrt(2)*x[:,2] + 1*x[:,0] + beta * x[:,1] + e
return y,x,nobs
yn,xn,nobs = gen_data()
ll1,grad1,hess1,ll2,k1, grad2,hess2,k2 = setup_shi(yn,xn)
mc_out = vuong_tests3.monte_carlo(100,gen_data,setup_shi)
vuong_tests3.print_mc(mc_out)
# -
| overlapping_reg/pre_summer2021/overlap_shi_ex_pt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
from __future__ import division
import logging
import os
import xml.etree.ElementTree as ET
from senpy.plugins import SenpyPlugin, EmotionPlugin
from senpy.models import Results, Entry, Error
logger = logging.getLogger(__name__)
import numpy as np
import math, itertools
from collections import defaultdict
import gzip
from datetime import datetime
import subprocess
class machineTranslation(EmotionPlugin):
def __init__(self, info, *args, **kwargs):
super(machineTranslation, self).__init__(info, *args, **kwargs)
self.name = info['name']
self.id = info['module']
self._info = info
local_path = os.path.dirname(os.path.abspath(__file__))
def activate(self, *args, **kwargs):
st = datetime.now()
logger.info("{} {}".format(datetime.now() - st, "active"))
st = datetime.now()
subprocess.run(['rm','-f', 'translate.perl'])
subprocess.run( ['wget','http://server1.nlp.insight-centre.org/docker/translate.perl','-O','translate.perl'] )
subprocess.run( ['chmod','+x','translate.perl'] )
logger.info("{} {}".format(datetime.now() - st, "translation script downloaded"))
logger.info("%s plugin is ready to go!" % self.name)
def deactivate(self, *args, **kwargs):
try:
logger.info("%s plugin is being deactivated..." % self.name)
except Exception:
print("Exception in logger while reporting deactivation of %s" % self.name)
## CUSTOM METHODS
def _translate(self, source_language_code, target_language_code, text_input):
st = datetime.now()
command = './translate.perl %s %s "%s"' % (source_language_code, target_language_code, text_input)
logger.info("executing '%s'" % command)
# command = './translate.perl£££%s£££%s£££"%s"' % (source_language_code, target_language_code, text_input)
command = ['./translate.perl', str(source_language_code), str(target_language_code), str(text_input)]
result = subprocess.run( command, stdout=subprocess.PIPE )
logger.info("{} {}".format(datetime.now() - st, "translation is complete"))
result = result.stdout.decode("utf-8")
return result
def analyse(self, **params):
logger.debug("machine translation with params {}".format(params))
text_input = params.get("input", None)
source_language_code = params.get("sourcelanguage", None)
target_language_code = params.get("targetlanguage", None)
if source_language_code == target_language_code:
text_output = text_input
elif 'en' in [source_language_code, target_language_code]:
text_output = str(self._translate(source_language_code, target_language_code, text_input))
else:
raise Error("Unavailable language pair")
response = Results()
entry = Entry()
entry.nif__isString = text_input
entry['nif:predLang'] = source_language_code
translation = {}
translation['nif:isString'] = text_output
translation['nif:predLang'] = target_language_code
translation['nif:wasTranslatedFrom'] = entry.id
entry['nif:translation'] = [translation]
response.entries.append(entry)
return response
| machineTranlsation/machineTranslation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><font size=12>
# Weather Derivatites </h1>
# <h1> Transition LSTM <br></h1>
#
# Developed by [<NAME>](mailto:<EMAIL>) <br>
# 22 September 2018
#
# Import needed libraries.
import numpy as np
import pandas as pd
import random as rand
import matplotlib.pyplot as plt
import time
from io import StringIO
import datetime
import pickle
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
# # Import Datasets
# +
# Import total dataset.
# Configure path to read txts.
path = '../datasets/'
# Download the update dataset.
import os
if not os.path.exists(path+'/fullDataset/completeDailyDataset.pickle'):
# ! wget https://github.com/jesugome/WeatherDerivates/raw/master/datasets/fullDataset/completeDailyDataset.pickle -P path
allDataDataframe = pickle.load(open(path+'/fullDataset/completeDailyDataset.pickle','rb'))
# -
allDataDataframe.head()
# Normalize DataFrame
scaler = MinMaxScaler()
scaled_values = scaler.fit_transform(allDataDataframe)
allDataDataframe.loc[:,:] = scaled_values
allDataDataframe.head()
# # Preparing Datasets
# +
# Split original data into 0's and 1's (Dry and wet days).
dryDays = allDataDataframe[allDataDataframe['state']==0]
wetDays = allDataDataframe[allDataDataframe['state']==1]
# +
# Create X and Y training matrices(dry).
xDry = dryDays.drop(['nextState','state','Prep','probNeutral','nino34'],axis=1)
xtrain = xDry.values
test = pd.DataFrame(xtrain)
xtrainD = np.reshape(xtrain, ( xtrain.shape[0], 1 , xtrain.shape[1]))
yDry = dryDays['nextState']
ytrain = yDry.values
ytrainD = np.reshape(ytrain, (ytrain.shape[0] , ) )
print(xtrainD.shape, ' -- ' ,ytrainD.shape)
# +
# Create X and Y training matrices(Wet).
xWet = wetDays.drop(['nextState','state','probNeutral','nino34'],axis=1)
xtrain = xWet.values
test = pd.DataFrame(xtrain)
xtrainW = np.reshape(xtrain, ( xtrain.shape[0], 1 , xtrain.shape[1]))
yWet = wetDays['nextState']
ytrain = yWet.values
ytrainW = np.reshape(ytrain, (ytrain.shape[0] , ) )
print(xtrainW.shape, ' -- ' ,ytrainW.shape)
# -
# Split dataset.
xTrainD, xValidationD, yTrainD, yValidationD = train_test_split(xtrainD, ytrainD, test_size=0.25, random_state=22)
xTrainW, xValidationW, yTrainW, yValidationW = train_test_split(xtrainW, ytrainW, test_size=0.25, random_state=22)
# # Neural Network
# Import neural networks libraries.
from keras.models import Sequential
from keras.utils import np_utils
from keras.layers import Dense, Flatten, Dropout, Activation, BatchNormalization, LSTM
from keras import optimizers as opts
from keras.callbacks import History
from keras.layers import Conv1D, GlobalMaxPooling1D
from livelossplot import PlotLossesKeras
# ### Dry
# +
from keras import backend as K
K.clear_session()
model = Sequential()
model.add(LSTM(100, input_shape= (xtrainD.shape[1],xtrainD.shape[2])))
model.add(Dense(1, activation='sigmoid'))
#Defines optimizer.
op = opts.adam(lr=0.01)
model.compile(loss='binary_crossentropy', optimizer=op, metrics=['accuracy'])
model.summary()
batch_size = 365
epochs = 200
#model.fit(xtrain, ytrain, validation_split=0.1, batch_size=2000, epochs=50, verbose=1, callbacks=[PlotLossesKeras()])
model.fit(xTrainD, yTrainD,
batch_size=batch_size,
epochs=epochs,
verbose=1,
shuffle = False,
validation_data = [xValidationD,yValidationD],
#validation_split = 0.1,
callbacks=[PlotLossesKeras()])
model.save('../results/visibleMarkov/rainfall_lstmDry_LSTM.h5')
# +
from sklearn.metrics import roc_curve, auc, roc_auc_score
yPred = model.predict(xValidationD)
AUC = roc_auc_score(yValidationD, yPred)
# Perform the neural network over the test set.
print('Evaluating the test set...')
score = model.evaluate(xValidationD, yValidationD)
#roc_curve(yValidationTxt,yPredGenre)
print('\nTest score:', score[0])
print('Test accuracy:', score[1])
print('Model AUC is: ', AUC )
# -
# ### Wet
# +
from keras import backend as K
K.clear_session()
model = Sequential()
model.add(LSTM(100, input_shape= (xtrainW.shape[1],xtrainW.shape[2])))
model.add(Dense(1, activation='sigmoid'))
#Defines optimizer.
op = opts.adam(lr=0.01)
model.compile(loss='binary_crossentropy', optimizer=op, metrics=['accuracy'])
model.summary()
batch_size = 365
epochs = 200
#model.fit(xtrain, ytrain, validation_split=0.1, batch_size=2000, epochs=50, verbose=1, callbacks=[PlotLossesKeras()])
model.fit(xTrainW, yTrainW,
batch_size=batch_size,
epochs=epochs,
verbose=1,
shuffle = False,
validation_data = [xValidationW,yValidationW],
#validation_split = 0.1,
callbacks=[PlotLossesKeras()])
model.save('../results/visibleMarkov/rainfall_lstmWet_LSTM.h5')
# +
from sklearn.metrics import roc_curve, auc, roc_auc_score
yPred = model.predict(xValidationW)
AUC = roc_auc_score(yValidationW, yPred)
# Perform the neural network over the test set.
print('Evaluating the test set...')
score = model.evaluate(xValidationW, yValidationW)
#roc_curve(yValidationTxt,yPredGenre)
print('\nTest score:', score[0])
print('Test accuracy:', score[1])
print('Model AUC is: ', AUC )
# -
xValidationD.shape
val = np.array([0.1, 0.85, 0.14])
val.shape
xt = np.reshape(val, ( 1, 1 , val.shape[0]))
xt.shape
xWet
| code/nnLSTMtransition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="gAlLjDLPDGC4"
# # COCOデータセットダウンロード
# + id="-OUkCFdBa_4N"
# !wget http://images.cocodataset.org/zips/train2017.zip
# !unzip -n train2017.zip
# !rm -f train2017.zip
# !wget http://images.cocodataset.org/zips/val2017.zip
# !unzip -n val2017.zip
# !rm -f val2017.zip
# !wget http://images.cocodataset.org/annotations/annotations_trainval2017.zip
# !unzip -n annotations_trainval2017.zip
# !rm -f annotations_trainval2017.zip
# + [markdown] id="iWG6w8-ADOMa"
# # COCO minitrainダウンロード
# + id="bJkV6uAB4kLz"
# !pip install -U gdown
# + id="xU6Oxd313cLr"
import gdown
gdown.download('https://drive.google.com/uc?id=1lezhgY4M_Ag13w0dEzQ7x_zQ_w0ohjin', 'instances_minitrain2017.json', quiet=False)
# + [markdown] id="FwvZHQb3DRzf"
# # YOLOX環境準備
# + id="osoPIFsGbHAI"
# !git clone https://github.com/Megvii-BaseDetection/YOLOX
# %cd YOLOX
# !pip install -U pip && pip install -r requirements.txt
# !pip install -v -e .
# + [markdown] id="yGpyYsMFDoTe"
# # 学習データセットコピー
# + id="YyZER5PJc01v"
# !mkdir dataset
# !mkdir dataset/images
# !mkdir dataset/annotations
# !cp -rf /content/train2017 dataset/images
# !cp -rf /content/val2017 dataset/images
# !cp -rf /content/instances_minitrain2017.json dataset/annotations
# !cp -rf /content/annotations/instances_train2017.json dataset/annotations
# !cp -rf /content/annotations/instances_val2017.json dataset/annotations
# + [markdown] id="3kYqH0LsDqwI"
# # YOLOX-Nano学習用ファイル準備
# + id="PcUJTQ10ie5o"
# %%writefile nano.py
# #!/usr/bin/env python3
# -*- coding:utf-8 -*-
# Copyright (c) Megvii, Inc. and its affiliates.
import os
import torch.nn as nn
from yolox.exp import Exp as MyExp
class Exp(MyExp):
def __init__(self):
super(Exp, self).__init__()
self.depth = 0.33
self.width = 0.25
self.input_size = (416, 416)
self.random_size = (10, 20)
self.mosaic_scale = (0.5, 1.5)
self.test_size = (416, 416)
self.mosaic_prob = 0.5
self.enable_mixup = False
self.exp_name = os.path.split(os.path.realpath(__file__))[1].split(".")[0]
self.num_classes = 80
# Define yourself dataset path
self.data_dir = "/content/YOLOX/dataset/images"
# self.train_ann = "/content/YOLOX/dataset/annotations/instances_train2017.json"
self.train_ann = "/content/YOLOX/dataset/annotations/instances_minitrain2017.json"
self.val_ann = "/content/YOLOX/dataset/annotations/instances_val2017.json"
self.max_epoch = 30
self.data_num_workers = 4
self.eval_interval = 1
def get_model(self, sublinear=False):
def init_yolo(M):
for m in M.modules():
if isinstance(m, nn.BatchNorm2d):
m.eps = 1e-3
m.momentum = 0.03
if "model" not in self.__dict__:
from yolox.models import YOLOX, YOLOPAFPN, YOLOXHead
in_channels = [256, 512, 1024]
# NANO model use depthwise = True, which is main difference.
backbone = YOLOPAFPN(self.depth,
self.width,
in_channels=in_channels,
depthwise=True)
head = YOLOXHead(self.num_classes,
self.width,
in_channels=in_channels,
depthwise=True)
self.model = YOLOX(backbone, head)
self.model.apply(init_yolo)
self.model.head.initialize_biases(1e-2)
return self.model
# + [markdown] id="hExwDSqHDvW1"
# # 訓練済みモデルダウンロード
# + id="O0YLPYqnjWp_"
# !wget https://github.com/Megvii-BaseDetection/storage/releases/download/0.0.1/yolox_nano.pth
# + [markdown] id="gVNOvnRrDycK"
# # 訓練
# + id="hWLvBYzwjYm1"
# !python tools/train.py \
# -f nano.py \
# -d 1 \
# -b 64 \
# --fp16 \
# -o \
# -c yolox_nano.pth
# + [markdown] id="U1MBD8YCD1AN"
# # ONNX変換
# + id="Mj1yC0-zjb1i"
# !python tools/export_onnx.py \
# --output-name yolox_nano.onnx \
# -n yolox-nano \
# -f nano.py \
# -c '/content/YOLOX/YOLOX_outputs/nano/best_ckpt.pth'
# + id="rqPbGHuttk2B"
from google.colab import files
files.download('yolox_nano.onnx')
| YOLOX-Colaboratory-coco-minitrain.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
from IPython.display import Image
HTML("""
<style>
.output_png {
display: table-cell;
text-align: center;
vertical-align: middle;
}
</style>
""")
# # *Circuitos Elétricos I*
# ## Semana 1 - Convenções para aplicação das Leis de Kirchhoff na análise de circuitos
#
#
#
# ### Caso 1
Image("./figures/J1C1.png", width=500)
# #### Lei de Kirchhoff das tensões (LKT)
#
# Em qualquer malha frechada do circuito $\sum_k v_k = 0$
#
# `Convenção arbitrária (1): ao percorrer a malha, escolha um sinal (+ ou -) para indicar aumentos de tensão e o sinal oposto para indicar quedas de tensão no somatório da LKT.`
#
# Logo, atribuindo o sinal (-) para aumentos de tensão e o sinal (+) para quedas de tensão, ao aplicar a LKT no circuito mostrado acima, temos:
#
# $$
# \begin{align}
# -10 + v_1 + v_2 &= 0\\
# -v_2 + v_3 + v_4 &= 0
# \end{align}
# $$
# #### Lei de Kirchhoff das correntes (LKC)
#
# Em qualquer nó do circuito $\sum_k i_k = 0$
#
# `Convenção arbitrária (2): para o nó em questão, escolha um sinal (+ ou -) para indicar correntes chegando ao nó e o sinal oposto para indicar correntes deixando o nó no somatório da LKT.`
#
# ou, para evitar erros com troca de sinais, simplesmente faça
#
# `Somatório das correntes chegando ao nó igual ao somatório das correntes deixando o nó.`
#
# $$
# \begin{align}
# i_1 &= i_2 + i_3\\
# i_3 &= -0.5~A
# \end{align}
# $$
# #### Lei de Ohm (+convenção passiva)
#
# `Convenção passiva (3): qualquer expressão que relacione as grandezas de tensão e corrente num elemento ideal de dois terminais deve ser escrita de acordo com a convenção passiva.`
#
# A convenção passiva estabelece que:
#
# 1. Se o sentido de referência adotado para corrente coincide com a queda de tensão na polaridade de referência ($+ \rightarrow -$), *qualquer expressão envolvendo $v$ e $i$* para o elemento em questão deve ser escrita com **sinal positivo**.
#
#
# 2. Se o sentido de referência adotado para corrente coincide com o aumento de tensão na polaridade de referência ($+ \leftarrow -$), *qualquer expressão envolvendo $v$ e $i$* para o elemento em questão deve ser escrita com **sinal negativo**.
#
# A Lei de Ohm expressa a relação entre tensão, corrente e resistência num resistor ideal. Logo, as expressões da Lei de Ohm devem obedecer a convenção passiva.
#
# Desse modo, podemos escrever as seguintes equações para o circuito acima.
#
# $$
# \begin{align}
# v_1 &= 10i_1\\
# v_2 &= 50i_2\\
# v_3 &= 20i_3
# \end{align}
# $$
# Logo:
#
# $$
# \begin{align}
# -10 + 10i_1 + 50i_2 &= 0\\
# -50i_2 -10 + v_4 &= 0\\
# i_1 - i_2 &= -0.5
# \end{align}
# $$
#
# Rearranjando as equações:
#
# $$
# \begin{align}
# 10i_1 + 50i_2 &= 10\\
# -50i_2 + v_4 &= 10\\
# i_1 - i_2 &= -0.5
# \end{align}
# $$
# ### Solução das equações
import sympy as sp
import numpy as np
# +
# define as N variáveis desconhecidas
i1, i2, v4 = sp.symbols('i1, i2, v4')
# define os sistema de N equações
eq1 = sp.Eq(10*i1+50*i2, 10)
eq2 = sp.Eq(-50*i2 + v4, 10)
eq3 = sp.Eq(i1 - i2, -.5)
# resolve o sistema
soluc = sp.solve((eq1, eq2, eq3), dict=True)
i1 = np.array([sol[i1] for sol in soluc])
i2 = np.array([sol[i2] for sol in soluc])
v4 = np.array([sol[v4] for sol in soluc])
i3 = -0.5
print('Solução do sistema:\n\n i1 = %.2f A,\n i2 = %.2f A,\n i3 = %.2f A,\n v4 = %.2f V.' %(i1, i2, i3, v4))
# -
# #### Cálculo das potências
# +
# expressões para a Lei de Ohm (convenção passiva)
v1 = 10 * i1
v2 = 50 * i2
v3 = 20 * i3
# expressões para as potências (convenção passiva)
p10V = -10 * i1
p1 = v1 * i1
p2 = v2 * i2
p3 = v3 * i3
p4 = -v4 * 0.5
print('Potências:\n\n p10V = %.2f W\n p1 = %.2f W,\n p2 = %.2f W,\n p3 = %.2f W,\n p4 = %.2f W\n' %(p10V, p1, p2, p3, p4))
print('P: ', p10V + p1 + p2 + p3 + p4);
# -
# calcula somatório das potências
print('Somatório das potências : %.2f W\n' %(p10V+p1+p2+p3+p4))
# Simulação do circuito: https://tinyurl.com/yfbwd4vz
# ### Caso 2
Image("./figures/J1C2.png", width=500)
# +
# define as N variáveis desconhecidas
i1, i2, v4 = sp.symbols('i1, i2, v4')
# define os sistema de N equações
eq1 = sp.Eq(-10*i1 + 50*i2, -10)
eq2 = sp.Eq(50*i2 + v4, 10)
eq3 = sp.Eq(i1 + i2, -.5)
# resolve o sistema
soluc = sp.solve((eq1, eq2, eq3), dict=True)
i1 = np.array([sol[i1] for sol in soluc])
i2 = np.array([sol[i2] for sol in soluc])
v4 = np.array([sol[v4] for sol in soluc])
i3 = 0.5
print('Solução do sistema:\n\n i1 = %.2f A,\n i2 = %.2f A,\n i3 = %.2f A,\n v4 = %.2f V.' %(i1, i2, i3, v4))
# +
# expressões para a Lei de Ohm (convenção passiva)
v1 = 10*i1
v2 = -50*i2
v3 = -20*i3
# expressões para as potências (convenção passiva)
p10V = -10*i1
p1 = v1*i1
p2 = -v2*i2
p3 = -v3*i3
p4 = -v4*i3
print('Potências:\n\n p10V = %.2f W\n p1 = %.2f W,\n p2 = %.2f W,\n p3 = %.2f W,\n p4 = %.2f W\n' %(p10V, p1, p2, p3, p4))
print('P: ', p10V + p1 + p2 + p3 + p4);
# -
# ### Caso 3
Image("./figures/J1C3.png", width=500)
# +
# define as N variáveis desconhecidas
i1, i2, v4 = sp.symbols('i1, i2, v4')
# define os sistema de N equações
eq1 = sp.Eq( )
eq2 = sp.Eq( )
eq3 = sp.Eq( )
# resolve o sistema
soluc = sp.solve((eq1, eq2, eq3), dict=True)
i1 = np.array([sol[i1] for sol in soluc])
i2 = np.array([sol[i2] for sol in soluc])
v4 = np.array([sol[v4] for sol in soluc])
i3 = 0.5
print('Solução do sistema:\n\n i1 = %.2f A,\n i2 = %.2f A,\n i3 = %.2f A,\n v4 = %.2f V.' %(i1, i2, i3, v4))
# +
# expressões para a Lei de Ohm (convenção passiva)
v1 =
v2 =
v3 =
# expressões para as potências (convenção passiva)
p10V =
p1 =
p2 =
p3 =
p4 =
print('Potências:\n\n p10V = %.2f W\n p1 = %.2f W,\n p2 = %.2f W,\n p3 = %.2f W,\n p4 = %.2f W\n' %(p10V, p1, p2, p3, p4))
| Jupyter notebooks/Circuitos Eletricos I - Semana 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="gghLdT_L86m3" outputId="057f8fce-505f-4f1b-ecd3-c81c091dd8db"
# !pip install deepface
# + id="g3HdH8vW89J2"
from deepface import DeepFace
# + colab={"base_uri": "https://localhost:8080/"} id="eXHqgctf9ACL" outputId="a503c00c-bd03-4973-93f5-2ef1d56e947a"
DeepFace.verify(img1_path= "img1.jpg", img2_path = "img2.jpg", model_name = "ArcFace", detector_backend = 'retinaface')
# + colab={"base_uri": "https://localhost:8080/"} id="4hpJ8hpj9EXL" outputId="12806770-9278-4401-95bc-9b61f14194f6"
obj
# + id="MTCAVagU9T8O"
| DeepFace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 给定一棵二叉树,确定它是否高度平衡。
# 针对这个问题,将高度平衡二叉树定义为:每个节点的左右子树高度相差不超过1的二叉树。
# -
# <img src='110.jpg' width=600>
# +
# Definition for a binary tree node.
class TreeNode:
def __init__(self, val=0, left=None, right=None):
self.val = val
self.left = left
self.right = right
class Solution:
def isBalanced(self, root: TreeNode) -> bool:
if not root: return True
nodes = [root]
while nodes:
temp_node = []
for n in nodes:
l_depth = self.helper(n.left) + 1
r_depth = self.helper(n.right) + 1
if abs(l_depth - r_depth) > 1:
return False
if n.left:
temp_node.append(n.left)
if n.right:
temp_node.append(n.right)
nodes = temp_node
return True
def helper(self, node):
if not node: return 0
depth = max(self.helper(node.left), self.helper(node.right)) + 1
return depth
# +
t1 = TreeNode(3)
t2 = TreeNode(9)
t3 = TreeNode(20)
t4 = TreeNode(15)
t5 = TreeNode(7)
t1.left = t2
t1.right = t3
t3.left= t4
t3.right = t5
root_ = t1
solution = Solution()
solution.isBalanced(root_)
# -
| Tree/0827/110. Balanced Binary Tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import os
from PIL import Image
import matplotlib.pyplot as plt
import matplotlib as mpl
PIC = Image.open("timg.jpeg", 'r')
piv_value = np.array(PIC)
PIC
piv_value.shape
## svd
u_r, sigma_R, v_r = np.linalg.svd(piv_value[:, :, 0])
u_g, sigma_G, v_g = np.linalg.svd(piv_value[:, :, 1])
u_b, sigma_B, v_b = np.linalg.svd(piv_value[:, :, 2])
sigma_r=np.diag(sigma_R)
sigma_g=np.diag(sigma_G)
sigma_b=np.diag(sigma_B)
plt.plot(np.log(sigma_G))
plt.show()
K=200
R=np.rint(np.dot(np.dot(u_r[:,0:K],sigma_r[0:K,0:K]),v_r[0:K,:])).astype('uint8')
R[R<0]=0;R[R>255]=255
G=np.rint(np.dot(np.dot(u_g[:,0:K],sigma_g[0:K,0:K]),v_g[0:K,:])).astype('uint8')
G[G<0]=0;G[G>255]=255
B=np.rint(np.dot(np.dot(u_b[:,0:K],sigma_b[0:K,0:K]),v_b[0:K,:])).astype('uint8')
B[B<0]=0;B[B>255]=255
I= np.stack((R, G, B), axis=2)
plt.imshow(I)
plt.show()
from sklearn.decomposition import PCA
n_components=200
pca=PCA(n_components=n_components)
R=piv_value[:, :,0]
G=piv_value[:, :,1]
B=piv_value[:, :,2]
R_pca=np.rint(pca.inverse_transform(pca.fit_transform(R))).astype('uint8')
G_pca=np.rint(pca.inverse_transform(pca.fit_transform(G))).astype('uint8')
B_pca=np.rint(pca.inverse_transform(pca.fit_transform(B))).astype('uint8')
I= np.stack((R_pca, G_pca, B_pca), axis=2)
plt.imshow(I)
plt.show()
| 1104-linearalgebra/PCA_SVD_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import numpy as np
import pandas as pd
def generate_embedding(dataset,
iterations,
perplexity,
pca_dim,
learning_rate,
verbose=1):
path = f'../demo_embeddings/{dataset}/iterations_{iterations}/perplexity_{perplexity}/pca_{pca_dim}/learning_rate_{learning_rate}'
def display(string):
if verbose:
print(string)
if os.path.exists(path):
if os.path.exists(path + '/data.csv'):
display(f'{dataset} already exists.')
return
else:
os.makedirs(path)
data = pd.read_csv(f"../data/{dataset}_input.csv")
labels = pd.read_csv(f"../data/{dataset}_labels.csv")
nb_col = data.shape[1]
pca = PCA(n_components=min(nb_col, pca_dim))
data_pca = pca.fit_transform(data.values)
tsne = TSNE(n_components=3,
n_iter=iterations,
learning_rate=learning_rate,
perplexity=perplexity,
random_state=1131)
embedding = tsne.fit_transform(data_pca)
embedding_df = pd.DataFrame(embedding, columns=['x', 'y', 'z'])
embedding_df.index = labels.values
embedding_df.to_csv(path + f'/data.csv')
display(f'{path} has been generated.')
generate_embedding('mnist_3000', 250, 3, 25, 100, verbose=1)
# +
dataset = 'mnist_3000'
iterations = 250
perplexity = 3
pca_dim = 25
learning_rate = 10
path = f'../demo_embeddings/{dataset}/iterations_{iterations}/perplexity_{perplexity}/pca_{pca_dim}/learning_rate_{learning_rate}'
pd.read_csv(path+f'/data.csv', index_col=0)
# -
# %timeit df = pd.read_csv("mnist_3000_input.csv")
# %timeit df2 = pd.read_csv('../demo_embeddings/mnist_3000/iterations_250/perplexity_3/pca_25/learning_rate_10/data.csv')
# %timeit combined_df = df.join(df2.loc[:,'x':'z']).set_index(['x','y','z'])
df2 = pd.read_csv('../demo_embeddings/mnist_3000/iterations_250/perplexity_3/pca_25/learning_rate_10/data.csv')
df2.head()
image = df.iloc[0]
matrix = image.values.reshape(28,28)
# +
di = {'x': -3.510562, 'y': 1.200590}
def compare(coord):
return df2[coord] == di[coord]
print(di['y'])
df2[compare('y')]
# -
hoverData = { "points": [ { "x": 0.86785585, "y": 2.3639283, "z": 1.0667368, "curveNumber": 4, "pointNumber": 171, "text": "Digit 4" } ] }
hover_point_np = np.array([hoverData['points'][0][i] for i in ['x', 'y', 'z']]).astype(np.float64)
hover_point_np
# +
mask = df2.loc[:,'x':'z'].eq(hover_point_np).all(axis=1)
df2[mask].index[0]
image = df.iloc[df2[mask].index[0]].values.reshape(28,28).astype(np.float64)
Image.fromarray(np.uint8(255 * image))
# +
import base64
from PIL import Image
from io import BytesIO
def numpy_to_b64(array, scalar=True):
if scalar:
array = np.uint8(255 * array)
im_pil = Image.fromarray(array)
buff = BytesIO()
im_pil.save(buff, format="png")
im_b64 = base64.b64encode(buff.getvalue()).decode("utf-8")
return im_b64
numpy_to_b64(image)
| apps/dash-tsne/notebooks/Pandas Tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# +
def get_returns(file):
"""Reads data file from the disk and returns percentage change"""
return pd.read_csv(file + '.csv', index_col=0, parse_dates=True).pct_change()
def get_data(file):
"""Fetch data from the disk"""
data = pd.read_csv(file + '.csv', index_col='Date', parse_dates=['Date'])
return data
def calc_corr(ser1, ser2, window):
"""Calculates correlation of two series"""
ret1 = ser1.pct_change()
ret2 = ser2.pct_change()
corr = ret1.rolling(window).corr(ret2)
return corr
# -
df = get_returns('sp500')
df['NDX'] = get_returns('NDX')
df.tail()
df['SP500'].rolling(50).corr(df['NDX'])[-200:].plot();
df['SP500'].rolling(100).corr(df['NDX'])[-200:].plot();
df['SP500'].rolling(10).corr(df['NDX'])[-200:].plot();
# +
points_to_plot = 300
data = get_data('indexes')
# -
for ind in data:
data[ind + '_rebased'] = (data[-points_to_plot:][ind].pct_change() + 1).cumprod()
data.tail()
# +
# Relative strength NDX to SP500
data['RSI'] = data['NDX'] / data['SP500']
# Calculate 50 day rolling correlation
data['corr'] = calc_corr(data['NDX'], data['SP500'], 50)
# -
data.tail(20)
# Slice the data - cut the points we don't intend to plot
plot_data = data[-points_to_plot :]
# +
fig = plt.figure(figsize=(12,8))
# The first sublot, planning for 3 plots high, 1 plot wide, this being the first
ax = fig.add_subplot(311)
ax.set_title('Index Comparison')
ax.semilogy(plot_data['SP500_rebased'], linestyle='-', label='SP500', linewidth=3)
ax.semilogy(plot_data['NDX_rebased'], linestyle='--', label='NDX', linewidth=3)
ax.legend()
ax.grid(False);
# Second sub plot
ax2 = fig.add_subplot(312)
ax2.plot(plot_data['RSI'], label='Relative Strength, NDX to SP500', linestyle=':', linewidth=3)
ax2.legend()
ax2.grid(True)
#Thrird sub plot
ax3 = fig.add_subplot(313)
ax3.plot(plot_data['corr'], label='Correlation between NDX and SP500', linestyle='-', linewidth=3)
ax3.legend()
ax3.grid(True)
# -
| before_zipline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pref
# language: python
# name: pref
# ---
# +
# %load_ext autoreload
# %autoreload 2
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import torch
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib as mpl
from test_functions import problem_setup
from colour import Color
from sim_helpers import (
fit_outcome_model,
gen_random_candidates,
fit_pref_model,
gen_rand_X,
PosteriorMeanDummySampler,
gen_comps,
gen_expected_util_candidates
)
from botorch.optim.optimize import optimize_acqf
from helper_classes import LearnedPrefereceObjective
from botorch.sampling.samplers import MCSampler, SobolQMCNormalSampler
from botorch.acquisition.utils import prune_inferior_points
from botorch.acquisition.monte_carlo import (
qNoisyExpectedImprovement,
qSimpleRegret,
)
import numpy as np
import pandas as pd
from botorch.test_functions.multi_objective import DTLZ2
from torch.distributions import MultivariateNormal
mpl.rcParams["figure.dpi"]= 120
mpl.rcParams['pdf.fonttype'] = 42
mpl.rcParams['ps.fonttype'] = 42
mpl.rcParams['text.usetex'] = False
import warnings
warnings.filterwarnings(
"ignore",
message="Could not update `train_inputs` with transformed inputs",
)
warnings.filterwarnings(
"ignore",
message="NumericalWarning: A not p.d., added jitter of 1.0e-08 to the diagonal",
)
# -
X_dim = 3
Y_dim = 2
problem = DTLZ2(dim=X_dim, num_objectives=Y_dim, noise_std=0)
def util_func(Y):
mean = torch.tensor([0.5, 1]).double()
diag_mat = torch.diag(torch.tensor([1, 1.5]))
theta = torch.tensor(0.4)
R = torch.tensor([[torch.cos(theta), -torch.sin(theta)], [torch.sin(theta), torch.cos(theta)]])
covar = (R @ diag_mat @ R.T).double()
mn = MultivariateNormal(loc=mean, covariance_matrix=covar)
return mn.log_prob(Y)
# +
# Find out feasible Y
n_mc = 1024
mc_X = gen_rand_X(n_mc, problem)
mc_Y = problem(mc_X)
mc_util = mc_Y.sum(dim=-1)
# Simulate preference heatmap
repeated_Ys = []
ymin, ymax = -1, 2.5
util_min = util_func(mc_Y).min()
for y1 in np.arange(ymin, ymax, 0.1):
for y2 in np.arange(ymin, ymax, 0.1):
util_val = util_func(torch.tensor([y1, y2]))
int_util_val = ((util_val - util_min) * 20).clamp(min=0).round().int()
single_Y = torch.tensor([y1, y2])
repeated_Ys.append(single_Y.repeat(int_util_val, 1))
repeated_Ys = torch.cat(repeated_Ys)
print(repeated_Ys.shape)
# +
torch.manual_seed(2)
EUBO_X = []
EUBO_Y = []
feasible_sample_Y = []
# Outcome points
outcome_X = gen_rand_X(8, problem)
outcome_Y = problem(outcome_X)
outcome_util = util_func(outcome_Y)
outcome_comps = gen_comps(utility=outcome_util, comp_noise_type="constant", comp_noise=0)
outcome_model = fit_outcome_model(outcome_X, outcome_Y, X_bounds=problem.bounds)
train_Y = outcome_model.posterior(gen_rand_X(2, problem)).mean.detach()
train_util = util_func(train_Y)
train_comps = gen_comps(utility=train_util, comp_noise_type="constant", comp_noise=0)
post = outcome_model.posterior(mc_X)
for i in range(10):
print(i)
pref_model = fit_pref_model(
train_Y,
train_comps,
kernel="default",
transform_input=True,
Y_bounds=torch.stack([torch.zeros(Y_dim), mc_Y.max(dim=-2).values * 1.2])
)
cand_X, cand_Y, acqf_val, acqf = gen_expected_util_candidates(
outcome_model=outcome_model,
pref_model=pref_model,
problem=problem,
previous_winner=None,
search_space_type="rff",
return_acqf=True
)
EUBO_X.append(cand_X)
EUBO_Y.append(cand_Y)
# EUBO-f
feasible_sample_Y.append(acqf.gp_samples.posterior(mc_X).mean.squeeze(0).clone().detach())
train_Y = torch.cat((train_Y, cand_Y))
train_util = util_func(train_Y)
train_comps = gen_comps(utility=train_util, comp_noise_type="constant", comp_noise=0)
# +
plot_EUBO_Y = torch.cat(EUBO_Y).detach().numpy()
plot_every_n = 4
n_iter_to_plot = plot_EUBO_Y.shape[0] // (plot_every_n * 2) + 1
plot_idx = [i * plot_every_n for i in range(n_iter_to_plot)]
pair_plot_idx = np.array([(2 * plot_every_n * i, 2 * plot_every_n * i + 1) for i in range(n_iter_to_plot)]).flatten()
plot_EUBO_Y = plot_EUBO_Y[pair_plot_idx, :]
cmap = "Reds"
ftrue_color = "#0096FF"
init_color = Color("#000000")
final_color = Color("#eeeeee")
colors = [c.hex_l for c in init_color.range_to(final_color, plot_EUBO_Y.shape[0]//2)]
pair_colors = np.repeat(colors, 2)
plot_ymin, plot_ymax = ymin, ymax
fig, ax = plt.subplots(figsize=(5, 4.5))
sns.kdeplot(x=repeated_Ys[:,0], y=repeated_Ys[:,1], cmap=cmap, shade=True, levels=100, ax=ax, thresh=0.001)
sns.kdeplot(x=mc_Y[:,0], y=mc_Y[:,1], levels=1, ax=ax, color=ftrue_color, thresh=0.005, linewidths=2)
for i, idx in enumerate(plot_idx):
sns.kdeplot(x=feasible_sample_Y[idx][:,0], y=feasible_sample_Y[idx][:,1], levels=1, ax=ax, color=colors[i], thresh=0.005, alpha=0.8, linewidths=1.5, linestyles="--", zorder=i + 10)
ax.scatter(x=plot_EUBO_Y[[i*2, i*2+1], 0], y=plot_EUBO_Y[[i*2, i*2+1], 1], marker="x", s=70, linewidths=2.5, color=colors[i], alpha=1, zorder=i + 10, label=idx + 1)
ax.set_xlim(-1, 2.0)
ax.set_ylim(-0.75, 2.75)
ax.set_xlabel("y1 (outcome 1)")
ax.set_ylabel("y2 (outcome 2)")
plt.legend(title="Iteration")
plt.savefig("../plots/example_eubo.pdf", bbox_inches='tight')
| notebooks/illustrative_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Safety of a Buckling Plate
import grama as gr
X = gr.Intention()
# # Plate in danger of buckling
#
# ---
#
# 
#
# **Question**: Will the plate buckle?
# ## Modeling Safety
#
# ---
#
# $$\text{Safety} = \text{Stress}_{\text{Critical}} - \text{Stress}_{\text{Applied}}$$
#
# Important values!
#
# | Value | Means |
# |---|---|
# | $\text{Safety} > 0$ | Safe |
# | $\text{Safety} \leq 0$ | Unsafe! |
#
# $\text{Stress}_{\text{Critical}}$ and $\text{Stress}_{\text{Applied}}$ come from engineering analysis
## Model building == Engineering analysis
from plate_analysis import md_plate
## Show model structure
md_plate.show_dag()
## Simulate the model 1000 times and plot
df_mc = \
md_plate >> \
gr.ev_monte_carlo(n=1e3, df_det="nom")
df_mc >> \
gr.pt_hists(out=["E", "t", "mu", "h"])
# ## Study: Ignoring Uncertainty
#
# ---
#
df_conservative = \
md_plate >> \
gr.ev_nominal(df_det="nom")
df_conservative[["safety"]]
# **Conclusion**: Plate will not buckle
# ## Study: Considering Uncertainty
#
# ---
#
## Analyze the model
df_mc >> gr.pt_hists(out=["safety"])
df_mc >> gr.tf_summarize(safety=gr.quant(X.safety, p=0.01))
# **Conclusion**: Plate has a *small probability* of buckling
| examples/demo/buckling_plate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mleyvaz/Data-science-python/blob/master/ExplainModel.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="VqezIXgnfiXd" outputId="1184d6e3-7129-4a54-9c29-e1c7f8ca97fd"
# !pip install explainerdashboard
# + colab={"base_uri": "https://localhost:8080/", "height": 462} id="75yyUE2Bfb1q" outputId="07f58fbe-1bb8-41b3-ef57-55abfb33a91c"
from sklearn.ensemble import RandomForestClassifier
from explainerdashboard import ClassifierExplainer, ExplainerDashboard
from explainerdashboard.datasets import titanic_survive, titanic_names
feature_descriptions = {
"Sex": "Gender of passenger",
"Gender": "Gender of passenger",
"Deck": "The deck the passenger had their cabin on",
"PassengerClass": "The class of the ticket: 1st, 2nd or 3rd class",
"Fare": "The amount of money people paid",
"Embarked": "the port where the passenger boarded the Titanic. Either Southampton, Cherbourg or Queenstown",
"Age": "Age of the passenger",
"No_of_siblings_plus_spouses_on_board": "The sum of the number of siblings plus the number of spouses on board",
"No_of_parents_plus_children_on_board" : "The sum of the number of parents plus the number of children on board",
}
X_train, y_train, X_test, y_test = titanic_survive()
train_names, test_names = titanic_names()
model = RandomForestClassifier(n_estimators=50, max_depth=5)
model.fit(X_train, y_train)
explainer = ClassifierExplainer(model, X_test, y_test,
cats=['Deck', 'Embarked',
{'Gender': ['Sex_male', 'Sex_female', 'Sex_nan']}],
descriptions=feature_descriptions, # defaults to None
labels=['Not survived', 'Survived'], # defaults to ['0', '1', etc]
idxs = test_names, # defaults to X.index
index_name = "Passenger", # defaults to X.index.name
target = "Survival", # defaults to y.name
)
db = ExplainerDashboard(explainer,
title="Titanic Explainer", # defaults to "Model Explainer"
whatif=False, # you can switch off tabs with bools
)
db.run(port=8050)
# + colab={"base_uri": "https://localhost:8080/", "height": 821} id="bihzZK7vgPoh" outputId="870015eb-6507-405e-e21e-196bc98fe96e"
from explainerdashboard import InlineExplainer
InlineExplainer(explainer).tab.whatif()
| ExplainModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python для анализа данных
#
# *<NAME>, НИУ ВШЭ*
#
# дополнения: *<NAME>, НИУ ВШЭ*
# Посмотрим на другие примеры использования `selenium`.
#
# **Пример.** Зайдем на сайт книжного магазина и найдем все книги про Python. Загрузим библиотеку, веб-драйвер и откроем страницу в браузере через Python.
# +
# from selenium import webdriver as wb
# br = wb.Firefox()
from selenium import webdriver
from webdriver_manager.chrome import ChromeDriverManager
br = webdriver.Chrome(ChromeDriverManager().install())
# открываем страницу в Chrome в автоматическом режиме
br.get("http://www.biblio-globus.ru/")
# -
# Найдем с помощью CSS Selector'а (*SelectorGadget*) поле для ввода названия книги или автора.
field = br.find_element_by_css_selector("input")
# Сохраним запрос:
author = "Python" # переменная author - условность
# Введем запрос в поле для поиска (`.send_keys`) и подождем чуть-чуть:
field.send_keys(author)
br.implicitly_wait(2) # подождем пару секунд
# Теперь найдем кнопку для поиска (значок *лупа* рядом со строкой поиска) через CSS Selector:
submit = br.find_element_by_css_selector("#search_submit")
# Кликнем на нее:
submit.click()
# Сохраним первую страницу с результатами в переменную `page1`.
page1 = br.page_source
# +
# page1
# -
# Теперь обработаем эту страницу через `BeautifulSoup`:
from bs4 import BeautifulSoup
soup1 = BeautifulSoup(page1, 'lxml')
# Найдем все названия книг на этой странице. По исходному коду можно увидеть, что они имеют тэг `a` с атрибутом `class`, равным `name`:
soup1.find_all('a', {'class':'name'})
# С помощью списковых включений выберем из ссылок с тэгом `<a>` текст (так мы уже делали, и не раз).
books1 = [b.text for b in soup1.find_all('a', {'class':'name'})]
books1
# Теперь аналогичным образом сгрузим информацию об авторах:
# [a.text for a in soup1.find_all('div', {'class': 'author'})]
# то же самое что и
authors1 = []
for a in soup1.find_all('div', {'class': 'author'}):
authors1.append(a.text)
authors1
soup1.find_all('div', {'class': 'author'})
authors1 = [a.text for a in soup1.find_all('div', {'class': 'author'})]
# Сгрузим расположение:
place1 = [p.text for p in soup1.find_all('div', {'class':'placement'})]
place1
# И, конечно, цену:
price1 = [p.text for p in soup1.find_all('div',
{'class':'title_data price'})]
price1
# Осталось пройтись по всем страницам, которые были выданы в результате поиска. Для примера перейдем на страницу 2 и на этом остановимся.
next_p = br.find_element_by_css_selector('.next_page')
next_p.click()
# Проделаем то же самое, что и с первой страницей. По-хорошему нужно написать функцию, которая будет искать на странице названия книг, их расположение и цену. Но оставим это в качестве задания читателю :)
page2 = br.page_source
soup2 = BeautifulSoup(page2, 'lxml')
books2 = [b.text for b in soup2.find_all('a', {'class':'name'})]
author2 = [a.text for a in soup2.find_all('div', {'class': 'author'})]
place2 = [p.text for p in soup2.find_all('div', {'class':'placement'})]
price2 = [p.text for p in soup2.find_all('div', {'class':'title_data price'})]
# Расширим списки результатов с первой страницы данными, полученными со второй страницы, используя метод `.extend()`.
books1.extend(books2) # books1 + books2
authors1.extend(books2)
place1.extend(place2)
price1.extend(price2)
# Осталось импортировать библиотеку `pandas` и создать датафрейм.
import pandas as pd
# Для разнообразия создадим датафрейм не из списка списков, а из словаря. Ключами словаря будут названия столбцов в таблице, а значениями – списки с сохраненной информацией (названия книг, цены и проч.).
df = pd.DataFrame({'book': books1, 'author': authors1,
'placement': place1, 'price': price1})
df.head()
# Давайте приведем столбец с ценой к числовому типу. Уберем слова *Цена* и *руб*, а потом сконвертируем строки в числа с плавающей точкой. Напишем функцию `get_price()`,
df.iloc[1, 3]
float(df.iloc[1, 3].split()[1].replace(',', '.'))
float('.'.join(re.findall(r'\d+', df.iloc[1, 3])))
def get_price(price):
book_price = price.split(' ')[1] # разобьем строку по пробелу и возьмем второй элемент
book_price = book_price.replace(',', '.') # заменим запятую на точку
price_num = float(book_price) # сконвертируем в float
return price_num
import re
def preis(x):
return float('.'.join(re.findall(r'\d+',x)))
# проверка
get_price(df.price[0])
preis(df.price[0])
# Всё отлично работает! Применим функцию к столбцу *price* и создадим новый столбец *nprice*.
df['nprice'] = df.price.apply(preis)
df.head()
# Теперь можем расположить книги по цене в порядке возрастания:
df.sort_values('nprice')
# И сохраним всю таблицу в csv-файл:
df.to_csv("books.csv", index=False)
br.close()
# +
from selenium import webdriver
from webdriver_manager.chrome import ChromeDriverManager
br = webdriver.Chrome(ChromeDriverManager().install())
# открываем страницу в Chrome в автоматическом режиме
br.get("http://www.biblio-globus.ru/")
# -
knigi = '//*[@id="TableMRight"]/tbody/tr/td/table/tbody/tr[2]/td[2]/a'
knigi_el = br.find_element_by_xpath(knigi)
knigi_el.click()
det_css = 'body > table > tbody > tr:nth-child(2) > td.column_right > div > div.card-columns > div:nth-child(1) > div > ul > li > ul > li:nth-child(2) > ul > li:nth-child(2) > a'
det_el = br.find_element_by_css_selector(det_css)
det_el.click()
page1 = BeautifulSoup(br.page_source)
books_p1 = page1.find_all('div', {'class': 'details_1'})
len(books_p1)
books_p1[0]
books_p1[0].divv is None
books_p1[0].find('div', {'class': 'author'}).text
books_p1[0].a.text
books_p1[0].find('div', {'class': 'placement'}).text
books_p1[0].find('div', {'class': 'title_data price'}).text
books_p1[0].find('div', {'class': 'title_data pricee'}) is None
titles = []
authors = []
places = []
prices = []
for book in books_p1:
if book.find('div', {'class': 'author'}) is not None:
authors.append(book.find('div', {'class': 'author'}).text)
else:
authors.append('')
if book.a is not None:
titles.append(book.a.text)
else:
titles.append('')
if book.find('div', {'class': 'placement'}) is not None:
places.append(book.find('div', {'class': 'placement'}).text)
else:
places.append('')
if book.find('div', {'class': 'title_data price'}) is not None:
prices.append(book.find('div', {'class': 'title_data price'}).text)
else:
prices.append('')
titles
def get_page_info(books_p):
titles = []
authors = []
places = []
prices = []
for book in books_p:
if book.div is not None:
authors.append(book.div.text)
else:
authors.append('')
if book.a is not None:
titles.append(book.a.text)
else:
titles.append('')
if book.find('div', {'class': 'placement'}) is not None:
places.append(book.find('div', {'class': 'placement'}).text)
else:
places.append('')
if book.find('div', {'class': 'title_data price'}) is not None:
prices.append(book.find('div', {'class': 'title_data price'}).text)
else:
prices.append('')
return titles, authors, places, prices
next_page = '//*[@id="main_wrapper"]/ul/li[4]/a'
page2 = br.find_element_by_xpath(next_page)
page2.click()
# +
# next_page_2 = '//*[@id="main_wrapper"]/ul/li[8]/a'
# next_page_2 = br.find_element_by_xpath(next_page_2)
# next_page_2.click()
# -
from time import sleep
np_xpath = '//*[@id="main_wrapper"]/ul/li[8]/a'
for _ in range(1000):
try:
page = BeautifulSoup(br.page_source)
books = page.find_all('div', {'class': 'details_1'})
t, a, pl, pr = get_page_info(books)
titles.extend(t)
authors.extend(a)
places.extend(pl)
prices.extend(pr)
np = br.find_element_by_xpath(np_xpath)
sleep(3)
np.click()
except:
print('all pages parsed')
break
df = pd.DataFrame({'book': titles, 'author': authors,
'placement': places, 'price': prices})
df.head()
# +
# br.close()
| lect11_Selenium_API/2021_DPO_11_3_selenium-books.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas 3: Shaping data
#
# The second in a series of notebooks that describe Pandas' powerful data management tools. This one covers shaping methods: switching rows and columns, pivoting, and stacking. We'll see that this is all about the indexes: the row and column labels.
#
# Outline:
#
# * [Example: WEO debt and deficits](#wants). Something to work with.
# * [Indexing](#index). Setting and resetting the index. Multi-indexes.
# * [Switching rows and columns](#pivot). Transpose. Referring to variables with multi-indexes.
# * [Stack and unstack](#stack). Managing column structure and labels.
# * [Pivot](#pivot). Unstack shortcut if we start with wide data.
# * [Review](#review). Apply what we've learned.
#
# More data management topics coming.
#
# **Note: requires internet access to run.**
#
# <!--
# internal links http://sebastianraschka.com/Articles/2014_ipython_internal_links.html
# -->
#
# This IPython notebook was created by <NAME>, <NAME>, and <NAME> for the NYU Stern course [Data Bootcamp](http://databootcamp.nyuecon.com/).
# <a id=prelims></a>
# ## Preliminaries
#
# Import packages, etc.
# +
import sys # system module
import pandas as pd # data package
import matplotlib.pyplot as plt # graphics module
import datetime as dt # date and time module
import numpy as np # foundation for Pandas
# %matplotlib inline
# check versions (overkill, but why not?)
print('Python version:', sys.version)
print('Pandas version: ', pd.__version__)
print('Today: ', dt.date.today())
# -
# <a id=wants></a>
# ## Example: WEO debt and deficits
#
# We spend most of our time on one of the examples from the previous notebook. The problem in this example is that variables run across rows, rather than down columns. Our **want** is to flip some of the rows and columns so that we can plot the data against time. The question is how.
#
# We use a small subset of the IMF's [World Economic Outlook database](https://www.imf.org/external/ns/cs.aspx?id=28) that contains two variables and three countries.
url1 = 'http://www.imf.org/external/pubs/ft/weo/2015/02/weodata/'
url2 = 'WEOOct2015all.xls'
url = url1 + url2
weo = pd.read_csv(url, sep='\t',
usecols=[1,2,3,4,6,40,41,42],
thousands=',',
na_values=['n/a', '--'])
print('Variable dtypes:\n', weo.dtypes, sep='')
# +
# create debt and deficits dataframe: two variables and three countries
variables = ['GGXWDG_NGDP', 'GGXCNL_NGDP']
countries = ['ARG', 'DEU', 'GRC']
dd = weo[weo['WEO Subject Code'].isin(variables) & weo['ISO'].isin(countries)]
# change column labels to something more intuitive
dd = dd.rename(columns={'WEO Subject Code': 'Variable',
'Subject Descriptor': 'Description'})
# rename variables
dd['Variable'] = dd['Variable'].replace(to_replace=['GGXWDG_NGDP', 'GGXCNL_NGDP'],
value=['Debt', 'Surplus'])
dd
# -
# ### Reminders
#
# What kind of object does each of the following produce?
dd.index
dd.columns
dd['ISO']
dd[['ISO', 'Variable']]
dd[dd['ISO'] == 'ARG']
# ### Wants
#
# We might imagine doing several different things with this data:
#
# * Plot a specific variable (debt or surplus) for a given date.
# * Time series plots for a specific country.
# * Time series plots for a specific variable.
#
# Depending on which we want, we might organize the data differently. We'll focus on the last two.
#
# Here's a brute force approach to the problem: simply transpose the data. This is where that leads:
dd.T
# **Comments.** The problem here is that the columns include both the numbers (which we want to plot) and some descriptive information (which we don't).
# <a id='index'></a>
# ## Setting and resetting the index
#
# We start by setting and resetting the index. That may sound like a step backwards -- haven't we done this already? -- but it reminds us of some things that will be handy later.
#
# Take the dataframe `dd`. What would we like in the index? Evenutally we'd like the dates `[2011, 2012, 2013]`, but right now the row labels are more naturally the variable or country. Here are some varriants.
# ### Setting the index
dd.set_index('Country')
# we can do the same thing with a list, which will be meaningful soon...
dd.set_index(['Country'])
# **Exercise.** Set `Variable` as the index.
# **Comment.** Note that the new index brought its **name** along: `Country` in the two examples, `Variable` in the exercise. That's incredibly useful because we can refer to index levels by name. If we happen to have an index without a name, we can set it with
#
# ```python
# df.index.name = 'Whatever name we like'
# ```
# ### Multi-indexes
#
# We can put more than one variable in an index, which gives us a **multi-index**. This is sometimes called a **hierarchical index** because the **levels** of the index (as they're called) are ordered.
#
# Multi-indexes are more common than you might think. One reason is that data itself is often multi-dimensional. A typical spreadsheet has two dimensions: the variable and the observation. The WEO data is naturally three dimensional: the variable, the year, and the country. (Think about that for a minute, it's deeper than it sounds.)
#
# The problem we're having is fitting this nicely into two dimensions. A multi-index allows us to manage that. A two-dimensional index would work here -- the country and the variable code -- but right now we have some redundancy.
# **Example.** We push all the descriptive, non-numerical columns into the index, leaving the dataframe itself with only numbers, which seems like a step in thee right direction.
ddi = dd.set_index(['Variable', 'Country', 'ISO', 'Description', 'Units'])
ddi
# Let's take a closer look at the index
ddi.index
# That's a lot to process, so we break it into pieces.
#
# * `ddi.index.names` contains a list of level names. (Remind yourself that lists are ordered, so this tracks levels.)
# * `ddi.index.levels` contains the values in each level.
#
# Here's what they like like here:
# Chase and Spencer like double quotes
print("The level names are:\n", ddi.index.names, "\n", sep="")
print("The levels (aka level values) are:\n", ddi.index.levels, sep="")
# Knowing the order of the index components and being able to inspect their values and names is fundamental to working with a multi-index.
# ### Resetting the index
#
# We've seen that `set_index` pushes columns into the index. Here we see that `reset_index` does the reverse: it pushes components of the index back to the columns.
#
# **Example.**
ddi.head(2)
ddi.reset_index()
# or we can reset the index by level
ddi.reset_index(level=1).head(2)
# or by name
ddi.reset_index(level='Country').head(2)
# or do more than one at a time
ddi.reset_index(level=[1,3]).head(2)
# **Comment.** By default, `reset_index` pushes one or more index levels into columns. If we want to discard that level of the index altogether, we use the parameter `drop=True`.
ddi.reset_index(level=[1,3], drop=True).head(2)
# **Exercise.** For the dataframe `ddi`:
#
# * Use the `reset_index` method to move the `Units` level of the index to a column of the dataframe.
# * Use the `drop` parameter of `reset_index` to delete `Units` from the dataframe.
# ## Switching rows and columns
#
# If we take the dataframe `ddi`, we see that the everything's been put into the index but the data itself. Perhaps we can get what we want if we just flip the rows and columns. Roughly speaking, we refer to this as **pivoting**.
# ### First look at switching rows and columns
#
# The simplest way to flip rows and columns is to use the `T` or transpose property. When we do that, we end up with a lot of stuff in the column labels, as the multi-index for the rows gets rotated into the columns. Other than that, we're good. We can even do a plot. The only problem is all the stuff we've pushed into the column labels -- it's kind of a mess.
ddt = ddi.T
ddt
# **Comment.** We see here that the multi-index for the rows has been turned into a multi-index for the columns. Works the same way.
#
# The only problem here is that the column labels are more complicated than we might want. Here, for example, is what we get with the plot method. As usual, `.plot()` plots all the columns of the dataframe, but here that means we're mixing variables. And the legend contains all the levels of the column labels.
ddt.plot()
# **Comment.** Ooooh, that's ugly! We're on the right track, but evidently not there yet.
# ### Referring to variables with a multi-index
#
# Can we refer to variables in the same way? Sort of, as long as we refer to the top level of the column index. It gives us a dataframe that's a subset of the original one.
#
# Let's try each of these:
#
# * `ddt['Debt']`
# * `ddt['Debt']['Argentina']`
# * `ddt['Debt', 'Argentina']`
# * `ddt['ARG']`
#
# What do you see? What's going on? The theme is that we can reference the top level, which in `ddi` is the `Variable`. If we try to access a lower level, it bombs.
# indexing by variable
debt = ddt['Debt']
# **Exercise.** With the dataframe `ddt`:
#
# * What type of object is `Debt`?
# * Construct a line plot of `Debt` over time with one line for each country.
# **Example.** Let's do this together. How would we fix up the legend? What approaches cross your mind? (No code, just the general approach.)
# ### Swapping levels
#
# Since variables refer to the first level of the column index, it's not clear how we would group data by country. Suppose, for example, we wanted to plot `Debt` and `Surplus` for a specific country. What would we do?
#
# One way to do that is to make the country the top level with the `swaplevel` method. Note the `axis` parameter. With `axis=1` we swap column levels, with `axis=0` (the default) we swap row levels.
ddts = ddt.swaplevel(0,1, axis=1)
ddts
# **Exercise.** Use the dataframe `ddts` to plot `Debt` and `Surplus` across time for Argentina. *Hint:* In the `plot` method, set `subplots=True` so that each variable is in a separate subplot.
# ### The `xs` method
#
# Another approach to extracting data that cuts across levels of the row or column index: the `xs` method. This is recent addition tpo Pandas and an extremely good method once you get the hang of it.
#
# The basic syntax is `df.xs(item, axis=X, level=N)`, where `N` is the name or number of an index level and `X` describes if we are extracting from the index or column names. Setting `X=0` (so `axis=0`) will slice up the data along the index, `X=1` extracts data for column labels.
#
# Here's how we could use `xs` to get the Argentina data without swapping the level of the column labels
# +
# #ddt.xs?
# -
ddt.xs("Argentina", axis=1, level="Country")
ddt.xs("Argentina", axis=1, level="Country")["Debt"]
# **Exercise.** Use a combination of `xs` and standard slicing with `[...]` to extract the variable `Debt` for Greece.
# **Exercise.** Use the dataframe `ddt` -- and the `xs` method -- to plot `Debt` and `Surplus` across time for Argentina.
# <a id='stack'></a>
# ## Stacking and unstacking
#
# The `set_index` and `reset_index` methods work on the row labels -- the index. They move columns to the index and the reverse. The `stack` and `unstack` methods move index levels to and from column levels:
#
# * `stack` stacks the data up, moving the columns to the index and creating a **long** dataframe.
# * `unstack` does the reverse, moving columns or index levels into the column labels and creating a **wide** dataframe.
#
# We use both to shape (or reshape) our data. We use `set_index` to push things into the index. And then use `reset_index` to push some of them back to the columns. That gives us pretty fine-grainded control over the shape of our data.
# We start by simplifying our initial dataframe.
# drop some of the index levels (think s for small)
dds = ddi.reset_index(level=[1,3,4], drop=True)
# give a name to the column labels
dds.columns.name = 'Year'
dds
# Let's remind ourselves **what we want.** We want to
#
# * move the column index (Year) into the row index
# * move the `Variable` and `ISO` levels the other way, into the column labels.
#
# The first one uses `stack`, the second one `unstack`.
# ### Stacking
#
# We stack our data, one variable on top of another, with a multi-index to keep track of what's what. In simple terms, we change the data from a **wide** format to a **long** format. The `stack` method takes the lowest column level and makes it the lowest row level.
# convert to long format. Notice printing is different... what `type` is ds?
ds = dds.stack()
ds
# same thing with explicit reference to column name
dds.stack(level='Year').head(8)
# or with level number
dds.stack(level=0).head(8)
# ### Unstacking
#
# Stacking moves columns into the index, "stacking" the data up into longer columns. Unstacking does the reverse, taking levels of the row index and turning them into column labels. Roughly speaking we're rotating or **pivoting** the data.
# now go long to wide
ds.unstack() # default is lowest value level='ISO'
# different level
ds.unstack(level='Variable')
# or two at once
ds.unstack(level=['Variable', 'ISO'])
# **Exercise.** Run the code below and explain what each line of code does.
# stacked dataframe
ds.head(8)
du1 = ds.unstack()
du2 = du1.unstack()
# **Exercise (challenging).** Take the unstacked dataframe `dds`. Use some combination of `stack`, `unstack`, and `plot` to plot the variable `Surplus` against `Year` for all three countries. Challenging mostly because you need to work out the steps by yourself.
# <a id='pivot'></a>
# ## Pivoting
#
# The `pivot` method: a short cut to some kinds of unstacking. In rough terms, it takes a wide dataframe and constructs a long one. The inputs are columns, not index levels.
# ### Example: BDS data
#
# The Census's [Business Dynamnics Statistics](http://www.census.gov/ces/dataproducts/bds/data.html) collects annual information about the hiring decisions of firms by size and age. This table list the number of firms and total employment by employment size categories: 1 to 4 employees, 5 to 9, and so on.
#
# **Apply want operator.** Our **want** is to plot total employment (the variable `Emp`) against size (variable `fsize`). Both are columns in the original data.
#
# Here we construct a subset of the data, where we look at two years rather than the whole 1976-2013 period.
# +
url = 'http://www2.census.gov/ces/bds/firm/bds_f_sz_release.csv'
raw = pd.read_csv(url)
raw.head()
sizes = ['a) 1 to 4', 'b) 5 to 9', 'c) 10 to 19', 'd) 20 to 49']
bds = raw[(raw['year2']>=2012) & raw['fsize'].isin(sizes)][['year2', 'fsize', 'Firms', 'Emp']]
bds
# -
# ### Pivoting the data
#
# Let's think specifically about what we **want**. We want to graph `Emp` against `fsize` for (say) 2013. This calls for:
#
# * The index should be the size categories `fsize`.
# * The column labels should be the entries of `year2`, namely `2012` and `2013`.
# * The data should come from the variable `Emp`.
#
# These inputs translate directly into the following `pivot` method:
# pivot and divide by a million (dividing so bars aren't too long)
bdsp = bds.pivot(index='fsize', columns='year2', values='Emp')/10**6
bdsp
# **Comment.** Note that all the parameters here are columns. That's not a choice, it's the way the the `pivot` method is written.
#
# We do a plot for fun:
# plot 2013 as bar chart
fig, ax = plt.subplots()
bdsp[2013].plot.barh(ax=ax)
ax.set_ylabel('')
ax.set_xlabel('Number of Employees (millions)')
# <a id='review'></a>
# ## Review
#
# We return to the OECD's healthcare data, specifically a subset of their table on the number of doctors per one thousand population. This loads and cleans the data:
url1 = 'http://www.oecd.org/health/health-systems/'
url2 = 'OECD-Health-Statistics-2015-Frequently-Requested-Data.xls'
docs = pd.read_excel(url1+url2,
skiprows=3,
usecols=[0, 51, 52, 53, 54, 55, 57],
sheetname='Physicians',
na_values=['..'],
skip_footer=21)
# rename country variable
names = list(docs)
docs = docs.rename(columns={names[0]: 'Country'})
# strip footnote numbers from country names
docs['Country'] = docs['Country'].str.rsplit(n=1).str.get(0)
docs = docs.head()
docs
# Use this data to:
#
# * Set the index as `Country`.
# * Construct a horizontal bar chart of the number of doctors in each country in "2013 (or nearest year)".
# * Apply the `drop` method to `docs` to create a dataframe `new` that's missing the last column.
# * *Challenging.* Use `stack` and `unstack` to "pivot" the data so that columns are labeled by country names and rows are labeled by year. This is challenging because we have left out the intermediate steps.
# * Plot the number of doctors over time in each country as a line in the same plot.
#
# *Comment.* In the last plot, the x axis labels are non-intuitive. Ignore that.
# ## Resources
#
# Far and away the best material on this subject is <NAME>' 2015 Pycon presentation. 2 hours and 25 minutes and worth every second.
#
# * Video: https://youtu.be/5JnMutdy6Fw
# * Materials: https://github.com/brandon-rhodes/pycon-pandas-tutorial
# * Outline: https://github.com/brandon-rhodes/pycon-pandas-tutorial/blob/master/script.txt
| Code/IPython/bootcamp_pandas-shape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd07945e9a82d7512fbf96246d9bbc29cd2f106c1a4a9cf54c9563dadf10f2237d4
# ---
def funcion ():
#VARIABLE LOCAL DE LA VARIABLE
x=2
return 2
print(x)
x=99
def funcion():
#variable global
global x
x=2 # se asigna a una variable local una global que se llama igual
return x
funcion()
print(x)
x=99
def funcion():
#variable global
x=7
global x
x=2 # error porque se ha asignado una variable x antes que una global dentro de la funcion
return x
funcion()
print(x)
x=99
l1=[0,1,2]
def funcion(lista):
#variable global
global x
x=2
lista.append("AS")
funcion(lista=l1)
print("l1:",l1)
print("x:",x)
x=99
l1=[0,1,2]
def funcion(lista):
#variable global
global x
x=2
lista.append("AS")
l1=1 # le da igual a python xq ws una variable local no global
funcion(lista=l1)
print("l1:",l1)
print("x:",x)
x=99
l1=[0,1,2]
def funcion(lista):
#variable global
global x
global l1
x=2
lista.append("AS")
l1=1 #asi cambio la variable que he metido dentro de global
funcion(lista=l1)
print("l1:",l1)
print("x:",x)
x=99
l1=[0,1,2]
def funcion(lista):
#variable global
global x
global l1
x=2
l1= lista.append("AS") #asi te devuelve el return, asi que no se puede usar asi
funcion(lista=l1)
print("l1:",l1)
print("x:",x)
x=99
l1=[0,1,2]
def funcion(lista):
#variable global
global x
x=2
l1= lista.append("AS") #auqnue guardes lo que te devuelve esa funcion a l1, como haces el print l1 que es una funcion global te muestra la lista appendada
print("l1 local:",l1)
funcion(lista=l1)
print("l1:",l1)
print("x:",x)
x=99
l1=[0,1,2]
def funcion(lista):
#variable global
global x, l1
x=2
lista.append("AS")
print("l1 dentro de funcion1:", l1)
print("lista dentro de la funcion 1:", lista)
l1= lista.append("AS") #auqnue guardes lo que te devuelve esa funcion a l1, como haces el print l1 que es una funcion global te muestra la lista appendada
print("l1 dentro de la funcion2:", l1)
print("lista dentro de funcion2:", lista)
print("l1 local:",l1)
funcion(lista=l1)
print("l1:",l1)
print("x:",x)
# +
x = 2
def funcion(x):
x = 7 + x
print(x)
print(x)
funcion(x=7)
print(x)
# +
x = 2
a = 10
def funcion(x):
x = 7 + x + a
print(x)
print(x)
funcion(x=7)
print(x)
# +
x = 2
a = 10
def funcion(x):
b = a + 4
print("b:", b)
x = 7 + x + a
print(x)
print(x)
funcion(x=7)
print(x)
print(a)
# +
x = 2
a = 10
def funcion(x):
global a
a = a + 2
b = a + 4
print("a:", a)
print("b:", b)
x = 7 + x + a
print(x)
print(x)
funcion(x=7)
print(x)
print(a)
# -
| week4_EDA_np_pd_json_apis_regex/day1_numpy_pandas_I/theory/python/global_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Using the RelevantFeatureAugmenter with separate datasets for train and test data
#
# This notebook illustrates the RelevantFeatureAugmenter in pipelines where you have first train on samples from dataset `df_train` but then want to test using samples from another `df_test`.
# (Here `df_train` and `df_test` refer to the dataframes that contain the time series data)
#
# Due to limitations in the sklearn pipeline API one has to use the `ppl.set_params(fresh__timeseries_container=df)` method for those two dataframes between train and test run.
# + deletable=true editable=true
import pandas as pd
from sklearn.pipeline import Pipeline
from sklearn.cross_validation import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report
from tsfresh.examples.robot_execution_failures import download_robot_execution_failures
from tsfresh.examples import load_robot_execution_failures
from tsfresh.transformers import RelevantFeatureAugmenter
# + deletable=true editable=true
download_robot_execution_failures
df, y = load_robot_execution_failures()
df.shape
# -
# Here, df contains the time series of both train and test set.
# We will split it into a train df_train and a test set df_test:
y_train, y_test = train_test_split(y)
df_train = df.loc[df.id.isin(y_train.index)]
df_test = df.loc[df.id.isin(y_test.index)]
X_train = pd.DataFrame(index=y_train.index)
X_test = pd.DataFrame(index=y_test.index)
df_train.shape, df_test.shape
# + deletable=true editable=true
ppl = Pipeline([('fresh', RelevantFeatureAugmenter(column_id='id', column_sort='time')),
('clf', RandomForestClassifier())])
# + deletable=true editable=true
# for the fit on the train test set, we set the fresh__timeseries_container to `df_train`
ppl.set_params(fresh__timeseries_container=df_train)
ppl.fit(X_train, y_train)
# + deletable=true editable=true
# for the predict on the test test set, we set the fresh__timeseries_container to `df_test`
ppl.set_params(fresh__timeseries_container=df_test)
y_pred = ppl.predict(X_test)
# + deletable=true editable=true
print(classification_report(y_test, y_pred))
| notebooks/pipeline_with_two_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="fMpWRG9LdF8r"
# I need to Install Tensorlow using command
# !pip install -q tensorflow-gpu==2.7.0
# + [markdown] id="x_Rb5zcXLGzJ"
# Get the version
# + colab={"base_uri": "https://localhost:8080/"} id="2EjLsFxyXwXL" outputId="d3ac7702-dfe7-4060-9fe2-4133633e78b1"
try :
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
print(tf.__version__)
# + [markdown] id="4-zAoU-HNcu2"
# All my imports
# + id="ZuRKqiXNW5-J"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# here i import the layer that i need to create 1 demesion neural netword
from tensorflow.keras.layers import Dense, Input, GlobalMaxPool1D
from tensorflow.keras.layers import Conv1D, MaxPool1D, Embedding
from tensorflow.keras.models import Model
from keras.models import Sequential
from keras import layers
# here a import Tokenizer and pad_seqeunces in order to preprocess text
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
# + [markdown] id="ruYNVlpZDjZ_"
#
#
# ```
# # I import ny dataset that i have (train, val, test)
#
# # I use pandas to read the Data and also renames the columns
# + id="jC8985nHdNPL"
# Train Data
data_train = pd.read_csv('/content/train.csv', names=['sentence', 'label'])
# Validation Data
data_val = pd.read_csv('/content/val.csv', names=['sentence', 'label'])
# Test Data
data_test = pd.read_csv('/content/test.csv', names=['sentence', 'label'])
# + [markdown] id="xKalZMA5OrNy"
# Here i check the Data using .head
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="nzlkZO1xdjTT" outputId="c2c2544a-7dae-4f5b-d34f-9f3ac3e9ba64"
data_train.head()
# + id="91J5fF0admzz"
# I store the reviews and the labels in arrays
reviews = data_train['sentence'].values
label = data_train['label'].values
reviewstest = data_test['sentence'].values
labeltest = data_test['label'].values
reviewsval = data_val['sentence'].values
labelval = data_val['label'].values
# + id="sRY-3LcRdoWY"
reviews_train, y_train = (reviews, label)
reviewsVal, y_Val = (reviewsval, labelval)
reviewstest, y_test = (reviewstest, labeltest)
# + id="5P8M5Wm4dr-p"
#print(reviews_train)
# + [markdown] id="e_rGkxwCPkMM"
# Here I Convert sentences to sequences using the Tokenizer , first i called the fit_on_text on a trainset and i called text_to_sequebces for train, validation, text, this give me a sequences train and sequennces validation and also for test
#
# + id="7iNJDOZ2dthd"
MAX_VOCAB_S = 20000
tokenizer = Tokenizer(num_words = MAX_VOCAB_S)
tokenizer.fit_on_texts(reviews_train)
# new variable X_train and y_train and X_test
seq_train = tokenizer.texts_to_sequences(reviews_train)
seq_val = tokenizer.texts_to_sequences(reviewsVal)
seq_test = tokenizer.texts_to_sequences(reviewstest)
# + [markdown] id="KFAwirOOQSXa"
# I get the word index mapping and also vocabulary size V, 37252 tokens
# + colab={"base_uri": "https://localhost:8080/"} id="iAwhAq5edvzT" outputId="c2a2ba18-deb0-4d84-dccf-73a3a7e3d86a"
wordToIndex = tokenizer.word_index
V = len(wordToIndex) + 1
print(reviews_train[0])
print(seq_train[0])
V
# + [markdown] id="1fea6oy-ooZo"
#
#
# ```
# # Ce texte est au format code
# ```
#
# I called the pad_sequences to pad trainset
# + colab={"base_uri": "https://localhost:8080/"} id="ZMaxxmNhgii_" outputId="140eed57-9e59-469b-e188-c574b6b88bef"
data_train = pad_sequences(seq_train)
print('Shape of the data train : ', data_train.shape )
T = data_train.shape[1]
# + [markdown] id="ZCrJydWMo8fR"
# I called the pad_sequences to pad valtest using 2299 as maxlen
# + colab={"base_uri": "https://localhost:8080/"} id="xCNo57pShLqE" outputId="ded6c2a4-e18e-4dce-f37c-e85816fb0452"
data_val = pad_sequences(seq_val, maxlen=T)
print('Shape of the data val : ', data_val.shape )
# + [markdown] id="7YQjqiCCpGhS"
# I i called the pad_sequences to pad the testset using 2299 as **maxlen**
# + colab={"base_uri": "https://localhost:8080/"} id="Far5AB2chd6J" outputId="8cd05773-dd27-4a8a-fc4e-171650894500"
data_test = pad_sequences(seq_test,maxlen=T)
print('Shape of the data test : ', data_test.shape )
# + [markdown] id="_cvlHZHFgEQJ"
# **Creating of the model**
#
# I stared by Embedding layer dimensionality D = 50
# than i will have convolutional layer follow by Globalmaxpooling
#
# + id="eSb6L1qOZL-f"
D = 50
i = Input(shape=(T,))
x = Embedding(V, D)(i)
x = Conv1D(32, 3, activation='relu')(x)
x = GlobalMaxPool1D()(x)
x = Dense(10, activation='relu')(x)
x = Dense(1, activation='sigmoid')(x)
model = Model(i, x)
# + id="iUzAJcuYdbLT"
#D = 50
#i = Input(shape=(T,))
#x = Embedding(V, D)(i)
#x = Conv1D(32, 3, activation='relu')(x)
#x = MaxPool1D(3)(x)
#x = Conv1D(64, 3, activation='relu')(x)
#x = MaxPool1D(3)(x)
#x = Conv1D(128, 3, activation='relu')(x)
#x = GlobalMaxPool1D()(x)
#x = Dense(1, activation='sigmoid')(x)
#model = Model(i, x)
# + [markdown] id="ymQS_KsgoJtM"
# **for Compile and fitting the**
# + [markdown] id="OlPGH0ODqLrh"
# i called model.compile and model.fil
# + colab={"base_uri": "https://localhost:8080/"} id="EDEaBhPJd9il" outputId="448030ef-48ad-4e0e-a870-8b125e7efc5b"
model.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
print('Training model**')
Training = model.fit(data_train, y_train, epochs = 10, validation_data=(data_val, y_Val))
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="Ic_MyTxreawj" outputId="d957cd09-28f3-4bc2-f04a-ed41116bacb5"
loss, accuracy = model.evaluate(data_train, y_train, verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
loss, accuracy = model.evaluate(data_val, y_Val, verbose=False)
print("Validation Accuracy: {:.4f}".format(accuracy))
# + colab={"base_uri": "https://localhost:8080/", "height": 164} id="S96kE-F0edy3" outputId="ffe66217-ad8d-45ea-90f5-129c57b93ffa"
# Plot loss per iteration
plt.subplot(211)
plt.plot(Training.history['loss'], label='loss')
plt.plot(Training.history['val_loss'], label='val_loss')
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 164} id="8J20jNTLefnn" outputId="642c1b0d-155e-4bb4-8a67-1ecb12e4552d"
# Plot accuracy per iteration
plt.subplot(212)
plt.plot(Training.history['accuracy'], label='acc')
plt.plot(Training.history['val_accuracy'], label='val_acc')
plt.legend()
# + id="epeQWIHNgAbm" colab={"base_uri": "https://localhost:8080/"} outputId="bf2fdaf4-2e1f-4a5b-dc22-eb7b860c0b3d"
loss, accuracy = model.evaluate(data_test, y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
| Artificial Intelligence Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### <center>Update TCGA Workspace Data Model with Compact DRS URLs/Identifier</center>
#
# ##### Description
# This notebook allows a user to update single entity (ex. participant, sample, and pair) data model tables that currently have columns with "drs://dataguids.org" file paths, denoting a DRS URI (location) of a file, to the newer compact DRS URL - drs://dg.4DFC:UUID - format. This notebook will isolate eligible columns that point to a file with the dataguids.org pointer, create a new updated tsv file, and update the data model.
#
#
# ##### Options
# The dry_run option (default = True) will print out the changes that will be made to each table ahead of modifying the data tables. The stdout will show the data table name, the individual columns, and the path to the workspace bucket location of the updated .tsv. Users can examine and verify the changes before setting dry_run = False and re-running the cells to make real updates to the data model table.
#
#
# ##### Execution
# 1. Set dry_run = True or dry_run = False and execute cell (Shift + Enter).
# 2. Run each following cell once the preeceding cell has completed.
# [*] denotes a cell that is not finished executing.
#
# ##### Notes
# The set entity (participant_set, sample_set, and pair_set) data model tables are not modified in this script. The set tables point to the unique IDs of the set constituents - a value that is not modified - thus, not requiring any updates.
#
# + code_folding=[]
# variable that allows user to run script and look at the updated .tsv files before updating data model
# set dry_run to "False" and re-run script to perform actual update of data model with DRS URLs
# DEFAULT: dry_run is set to True and will list the columns in each table that will be updated
# it will also provide the location of the .tsv files with the DRS url updates for inspection
dry_run = True
# + code_folding=[0]
# Imports relevant packages. (Shift + Enter) to execute.
import os
import json
import re
from firecloud import api as fapi
import pandas as pd
from io import StringIO
import csv
import pprint
from collections import OrderedDict
# + code_folding=[0]
# Sets up workspace environment variables. (Shift + Enter) to execute.
ws_project = os.environ['WORKSPACE_NAMESPACE']
ws_name = os.environ['WORKSPACE_NAME']
ws_bucket = os.environ['WORKSPACE_BUCKET']
# print(ws_project + "\n" + ws_name + "\n" + "bucket: " + ws_bucket)
# + code_folding=[0]
# Gets list of single entity types in workspace that need DRS URL updates. (Shift + Enter) to execute.
# API call to get all entity types in workspace
res_etypes = fapi.list_entity_types(ws_project, ws_name)
dict_all_etypes = json.loads(res_etypes.text)
# get non-set entities and add to list
# "set" entities do not need to be updated because they only reference the unique ID of each single entity
# the unique ID of any single entity is not modified so sets should remain the same
single_etypes_list = []
single_etypes_list = [key for key in dict_all_etypes.keys() if not key.endswith("_set")]
print(f"List of entity types that will be updated, if applicable:")
print('\n'.join(['\t' * 7 + c for c in single_etypes_list]))
# + code_folding=[0]
# Updates the data model, for single entity types, with DRS URLs. (Shift + Enter) to execute.
# set guid pattern for guid validation
guid_pattern = re.compile(r'^[\da-f]{8}-([\da-f]{4}-){3}[\da-f]{12}$', re.IGNORECASE)
for etype in single_etypes_list:
print(f'Starting TCGA DRS updates for entity: {etype}')
# get entity table response for API call
res_etype = fapi.get_entities_tsv(ws_project, ws_name, etype, model="flexible")
# Save current/original data model tsv files to the bucket for provenance
print(f'Saving original {etype} TSV to {ws_bucket}')
original_tsv_name = "original_" + etype + "_table.tsv"
with open(original_tsv_name, "w") as f:
f.write(res_etype.text)
# copy files to workspace bucket
# !gsutil cp $original_tsv_name $ws_bucket 2> stdout
# read entity table response into dictionary to perform DRS URL updates
dict_etype = list(csv.DictReader(StringIO(res_etype.text), delimiter='\t'))
# create empty list to add updated rows and list to capture list of columns that were modified
drs_dict_table = []
modified_cols = set()
# for "row" (each row is [list] of column:values)
for row in dict_etype:
drs_row = row.copy()
# for each column in row
for col in row:
# check if the col values are dataguids.org URLs and parse out guid
if row[col].startswith("drs://dataguids.org"):
guid = row[col].split("/")[3] #[0]
# only modify col if guid is valid and exists
if guid and guid_pattern.match(guid):
drs_url = "drs://dg.4DFC:" + guid
drs_row[col] = drs_url
modified_cols.add(col)
# append new "row" with updated drs values to new list
drs_dict_table.append(drs_row)
# set output file name and write tsv files
updated_tsv_name = "updated_" + etype + "_table.tsv"
tsv_headers = drs_dict_table[0].keys()
with open(updated_tsv_name, 'w') as outfile:
# get keys from OrderedDictionary and write rows, separate with tabs
writer = csv.DictWriter(outfile, tsv_headers, delimiter="\t")
writer.writeheader()
writer.writerows(drs_dict_table)
print(f'Saving DRS URL updated {etype} TSV to {ws_bucket}')
# !gsutil cp $updated_tsv_name $ws_bucket 2> stdout
# list of the columns that are scoped to be updated if re-run with dry_run = False
modified_cols = list(modified_cols)
if dry_run:
print(f'Columns in the {etype} table that *will be* be updated when notebook is re-run with `dry_run = False`:')
if not modified_cols:
print('\t' * 4 + f"No columns to update in the {etype} table." + "\n\n")
else:
print('\n'.join(['\t' * 4 + c for c in modified_cols]))
print(f'To view in detail what will be updated, inspect the {updated_tsv_name} file in the workspace bucket, {ws_bucket}.' + "\n\n")
else:
# upload newly created tsv file containing drs urls
print(f"Starting update of the {etype} table with compact DRS identifiers (drs://df.4DFC:GUID).")
res_update = fapi.upload_entities_tsv(ws_project, ws_name, updated_tsv_name, model="flexible")
if res_update.status_code != 200:
print(f"Could not update existing {etype} table. Error message: {res_update.text}")
print(f'Finished uploading TCGA DRS updated .tsv for entity: {etype}' + "\n")
# -
| notebooks/Update_Data_Model_to_Compact_DRS_Identifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/krissydolor/Linear-Algebra-_-2nd-Sem/blob/main/Activity_1_Python_Fundamentals.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="EDQuZvw-sLys"
# # Welcome to Python Fundamentals
# In this module, we are going to establish or review our skills in Python programming. In this notebook we are going to cover:
# * Variables and Data Types
# * Operations
# * Input and Output Operations
# * Logic Control
# * Iterables
# * Functions
# + [markdown] id="s_6-KVCKtFeX"
# ## Variable and Data Types
# + [markdown] id="8E6dQdCnbZKR"
# ###### Variables and data types in Python are, as the name implies, values that change. A variable is a memory region in a computer language to store a value. The value you've saved may change in the future if the specs change.
# + id="2GoBp1jUtOb_"
x = 1
a,b = 0, -1
# + colab={"base_uri": "https://localhost:8080/"} id="H20uvU1_tVfs" outputId="a693ecd9-f8bd-4087-e064-f00ced478e56"
type(x)
# + colab={"base_uri": "https://localhost:8080/"} id="QjkU0tf1tvpf" outputId="fd9d0bcf-027b-4b24-a2e1-138364fdce09"
y = 1.0
type(y)
# + colab={"base_uri": "https://localhost:8080/"} id="pOwxhCvZt9Kn" outputId="89577745-7d3d-4e06-8f68-b80ff8013b81"
x = float(x)
type(x)
# + colab={"base_uri": "https://localhost:8080/"} id="ZQ711S6zwrNL" outputId="c6e9e6e2-98ce-4dbc-e158-4b200a8c5a70"
s, t, u = "0", "1", 'one'
type(s)
# + colab={"base_uri": "https://localhost:8080/"} id="6YJNLtGaxQFP" outputId="eae42a72-7ef1-431d-8e53-0067bdc2dada"
s_int = int(s)
s_int
# + [markdown] id="OdEau0nYyApB"
# ## Operations
# + [markdown] id="kbArVmbOBF1i"
# ###### Operators are Python's particular characters for solving math or logic computation. The operand is the value that the operator performs on—for instance, 2+10=12. Here the + sign is the operator that carries out the given equation. Additionally, 2 and 10 are known as the operands, and 12 is the operation's output.
#
# + [markdown] id="oBNuygbJyHY6"
# ### Arithmetic
# + [markdown] id="95YQX8g6iUtQ"
# ###### In Python, the arithmetic operators execute mathematical operations, including addition, subtraction, multiplication, and division.
# + [markdown] id="9qU34HyNBiYX"
# ###### The addition operator in Python is +. It's used to combine two numbers.
# ###### The subtraction operator in Python is -. It takes the second number and subtracts it from the first.
# ###### The multiplication operator in Python is *. It's used to calculate the product of two numbers.
# ###### The division operator in Python is /. When the first operand is divided by the second, it is utilized to get the quotient.
# ###### The exponentiation operator in Python is **. It elevates the first operand to the second operand's power.
# ###### The modulus operator in Python is %. When the first operand is divided by the second, it gets the remaining.
#
# + id="gAb9cfcPyPdq"
a,b,c,d = 2.0, -0.5, 0, -32
# + colab={"base_uri": "https://localhost:8080/"} id="OCilWW12ye5E" outputId="b44ed68a-8c64-4d1f-af59-a5e9698d7d3f"
### Addition
S = a+b
S
# + colab={"base_uri": "https://localhost:8080/"} id="fnS6fMZCyrZS" outputId="9e2b4e44-46b6-44b6-fb19-975d98e049c6"
### Subtraction
D = b-d
D
# + colab={"base_uri": "https://localhost:8080/"} id="f6KCAAI_yyij" outputId="143c5d52-5471-43e3-f33e-c113b2b1f876"
### Multiplication
P = a*d
P
# + colab={"base_uri": "https://localhost:8080/"} id="ChjhteXyzcww" outputId="672ddc04-dab4-45e6-d63d-9ebc97b6b7b0"
### Divison
Q = c/a
Q
# + colab={"base_uri": "https://localhost:8080/"} id="JZv6MJqwzoDd" outputId="7a7c7315-4a25-4725-eef7-488c9ae49778"
### Floor Division
Fq = a//b
Fq
# + colab={"base_uri": "https://localhost:8080/"} id="byAb-uXMz6-l" outputId="7be5ce8f-1e39-4042-f58b-413732e9871b"
### Exponentiation
E = a**b
E
# + colab={"base_uri": "https://localhost:8080/"} id="PtmPVcJ90p5V" outputId="2a8ce376-41c3-4e15-a403-e179da6777c1"
### Modulo
mod = d%a
mod
# + [markdown] id="2-w5y9Pp1A4B"
# ### Assignment Operations
# + [markdown] id="HUwR-QGIjxdH"
# ###### Assignment Operators functions to assign values to variables. The following are some examples of assignment operators that this activity utilized:
#
# + [markdown] id="uX4_UdtXCntE"
# ###### 1. =, this functions to assign the value of ride of expression to the operand on the left side
# ###### 2. +=, this will add the operand on the right side alongside the operand on the left side operand
# ###### 3. -=, this assignment operator subtracts the right operand from the left operand then assign it to the left operand
# ###### 4. *=, it multiplies right operand with left operand then assign to left operand
# ###### 5. **=, this will calculate the exponent value using operands and assign value to left operand
# + id="QVFJEw0L1Ett"
G, H, J, K = 0, 100, 2, 2
# + colab={"base_uri": "https://localhost:8080/"} id="G4rMNSFa1sj5" outputId="6bf6f86e-480a-401f-976d-cae9b79e3d4f"
G += a
a
# + colab={"base_uri": "https://localhost:8080/"} id="gF-TrNv72rnl" outputId="ad5d8579-9abd-4f49-9ca7-443399894463"
H -= d
H
# + colab={"base_uri": "https://localhost:8080/"} id="FgH-GE491380" outputId="0bde117b-c486-41a2-9c58-18fa71b6d2a6"
J *= 2
J
# + colab={"base_uri": "https://localhost:8080/"} id="NvDygy4G2SiD" outputId="f0b8bdbb-87b1-4655-d960-a122b1c4bb5c"
K **= 2
K
# + [markdown] id="sJcUaEQ13Zzm"
# ### Comparators
# + [markdown] id="F9sv7dwDlNrb"
# ###### Comparators are used to compare two values. It performs comparisons and generates boolean values. Although it is often used with numerical values, other data types may also be utilized.
# + [markdown] id="guHWtUy_C6Ry"
# ###### == symbol represents the equal to operator. It compares the values on the left and right sides. It returns True if the left-side value equals the right-side value. Otherwise, False is returned.
# ###### != symbol represents the not equal to operator. It compares the values on the left and right sides. It returns True if the left-side value is not equal to the right-side value. Otherwise, False is returned.
# ###### == symbol represents the equal to operator. It compares the values on the left and right sides.It returns True if the left-side value equals the right-side value. Otherwise, False is returned.
# ###### != symbol represents the not equal to operator. It compares the values on the left and right sides. It returns True if the left-side value is not equal to the right-side value. Otherwise, False is returned.
# ###### < symbol represents the less-than operator. It compares the values on the left and right sides. It returns True if the left-side value is less. Otherwise, False is returned.
# ###### <= symbol represents the operator less than or equal to. It compares the values on the left and right sides. It returns True if the left-side value is less than or equal to the right-side value. Otherwise, False is returned.
# + id="_DX8McaI3enz"
res_1, res_2, res_3 = 1, 2.0, "1"
true_val = 1.0
# + colab={"base_uri": "https://localhost:8080/"} id="lR0rh1N03sRZ" outputId="f19b7235-512e-4e51-e3bb-73f6347363ac"
## Equality
res_1 == true_val
# + colab={"base_uri": "https://localhost:8080/"} id="7R3LDDNz34HI" outputId="b05135b1-b015-4489-d533-52dbe2c7c9f4"
## Non-equality
res_3 != true_val
# + colab={"base_uri": "https://localhost:8080/"} id="04u8XuDH4C0P" outputId="582148c8-a38a-4e83-879f-1894e0e05b5f"
## Inequality
t1 = res_1 > res_2
t2 = res_1 < res_2/2
t3 = res_1 >= res_2/2
t4 = res_1 <= res_2
t3
# + [markdown] id="ogV1uC_W5LQs"
# ### Logical
# + [markdown] id="iqtZP8GXDdR6"
# ###### A logical operator is a character or term that joins various expressions. The value of the originalstatements entirely influences the validity of the compound expression generated and the operator's significance. This activity used logical operators such as AND, OR, and NOT
# + colab={"base_uri": "https://localhost:8080/"} id="lpEsFcvd5Qs4" outputId="8a711fa3-2fe1-471f-c72d-1b405630ae0f"
res_1 == true_val
# + colab={"base_uri": "https://localhost:8080/"} id="EDNdnQDa5crP" outputId="467972db-c896-4683-a48d-4a7ab560a355"
res_1 is true_val
# + colab={"base_uri": "https://localhost:8080/"} id="7CVYR6MX5hXP" outputId="8aafd9d2-8068-48ba-a4be-44fd2ca7b462"
res_1 is not true_val
# + colab={"base_uri": "https://localhost:8080/"} id="1Gcfoi-l5mMy" outputId="bd935d4c-cd30-4e52-b67b-6ede5abcba93"
p,q = True, False
conj = p and q
conj
# + colab={"base_uri": "https://localhost:8080/"} id="-emtrjeh7ICh" outputId="63ddb0da-d95f-4e12-f650-b5766f0f460d"
p,q = True, False
disj = p or q
disj
# + colab={"base_uri": "https://localhost:8080/"} id="avJNToSa7Y6U" outputId="561b5ed8-2ae2-4d91-ff7f-4e6c3fb72c80"
p,q = True, False
e = not(p and q)
e
# + colab={"base_uri": "https://localhost:8080/"} id="mo1XSlwC7qLM" outputId="84115a7c-bdaa-4c80-fbda-5f5d0b06312b"
p,q = True, False
xor = (not p and q) or (p and not q)
xor
# + [markdown] id="u30LBoHv8C-1"
# ### I/O
# + [markdown] id="nPSkCQXdejhV"
# ###### Input and output are concepts that refer to the way a computer software communicates with its user. The user provides input to the program, while the software provides output to the user.
# + [markdown] id="m3vwxFTwDuzv"
# ###### The print() method sends a message to the screen or another standard output device.
# ###### A counter tool “cnt” is included to help with quick and easy tallying.
# ###### In Python, single quotes, double quotes, and even triple quotes may generate strings. Unicode characters are represented as strings, which are arrays of bytes.
# ###### You need to give the names of the variables within a set of curly brackets when using f-Strings to show variables. All variable names will be replaced with their relevant values during runtime.
#
# + colab={"base_uri": "https://localhost:8080/"} id="lQjq_WFV8FlV" outputId="0eb3e836-67df-4656-d008-99079e8a04a5"
print("Hello World")
# + id="2SmKetBm8X8-"
cnt = 1
# + colab={"base_uri": "https://localhost:8080/"} id="RwbkCUyi8afK" outputId="7cbd4ef1-c2fe-4f8b-9c8f-128dff3634dd"
string = "Hello World"
print(string, ", Current run count is: ", cnt)
cnt += 1
# + colab={"base_uri": "https://localhost:8080/"} id="k3PYwbvN8t00" outputId="d748208e-b49d-427e-ea07-3d36c5607561"
print(f"{string}, Current count is: {cnt}")
# + colab={"base_uri": "https://localhost:8080/"} id="9XUzVsem9mwe" outputId="2a8e60c9-15ff-4803-ba1e-a23675f29667"
sem_grade = 82.243564657461234
name = "qtpie"
print("Hello {}, your semestral grade is: {}". format(name, sem_grade))
# + colab={"base_uri": "https://localhost:8080/"} id="Gm0ch5do-oEc" outputId="5eda83da-d629-4779-cfbf-2e7aefa0d068"
sem_grade = 82.243564657461234
name = ""
print("Hello {}, your semestral grade is: {}". format(name, sem_grade))
# + colab={"base_uri": "https://localhost:8080/"} id="9zdQrzp1ty4g" outputId="c377f828-e12b-48d1-f9d0-9d88f12fe77f"
w_pg, w_mg, w_fg = 0.3, 0.3, 0.4
print("The weights of your semestral grade are: \
\n\t{: .2%} for Prelims\
\n\t{: .2%} for Midterms, and\
\n\t{: .2%} for Finals.".format(w_pg, w_mg, w_fg))
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="bg--pwxnuibp" outputId="27a5c30c-5d34-402c-913d-34423c4be68d"
x = input("enter a number: ")
x
# + colab={"base_uri": "https://localhost:8080/"} id="x9oOs1usuqvU" outputId="f793f146-bcb0-411e-85c0-0f3a7221fd74"
name= input("K<NAME>: ")
pg = input("Enter prelim grade: ")
mg = input("Entermidterm grade ")
fg = input("Entermidterm grade ")
sem_grade = None
print("Hello {}, your semestral grade is: {}" .format(name, sem_grade))
# + [markdown] id="xOQ7_clnwHUv"
# ### Looping Statement
# + [markdown] id="NTYRO4bKECeQ"
# ###### It comprises a series of operations that are repeated numerous times until the desired result is reached. Python includes two looping statements: for and while, which are employed to identify them in this exercise.
#
# + [markdown] id="PIvyRe8wwOFR"
# ### While
# + [markdown] id="bTmbYUXupvuI"
# ###### Python While Loop is used to repeatedly execute a block of statements until a given condition is satisfied. And when the situation becomes false, the line immediately after the loop in the program is executed.
# + colab={"base_uri": "https://localhost:8080/"} id="2cP9z6_hwRRw" outputId="8da60426-3f68-4972-a7df-67b005d4b36b"
## while loops
i, j = 0, 10
while(i<=j):
print(f"{i}\t|\t{j}")
i+=1
# + [markdown] id="ru_nZCfExaio"
# ### For
# + [markdown] id="AfwtCevKENKc"
# ###### A "For" Loop is used to loop a code block a predetermined amount of times. In the "For" loop, the number of iterations to be performed is already known. It is employed to achieve the desired outcome. On the contrary, in a while loop, the command runs until a particular condition is satisfied and the statement is proven to be wrong.
#
# + colab={"base_uri": "https://localhost:8080/"} id="0tTI6E99xdDA" outputId="7a28d278-de3a-42d5-b78b-1e5a3addd9dd"
# for(int i=0; i<10; i++){
# printf(i)
# }
i=0
for i in range(10):
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="VBYv2CPDyBo7" outputId="ca5572e7-3d20-4145-f4b9-49db6815f3fc"
playlist = ["Crazier", "Happier", "Bahay-Kubo"]
print('Now Playing:\n')
for song in playlist:
print(song)
# + [markdown] id="MPGGRZSmy1D4"
# ### Flow Control
# + [markdown] id="ofeAfXhWteW8"
# ###### Flow control statements may be used to determine which Python instructions should be executed when such criteria are met.
# + [markdown] id="7w7Zw6ZmiqMy"
# ###### The if-elif-else statement is used to conditionally execute a statement or a block of statements. Conditions can be true or false, execute one thing when the condition is true, something else when the condition is false.
# + [markdown] id="T0bCdbxFEr9g"
# ### Condition Statements
# + [markdown] id="RzbAdN7eE1du"
# ###### In a Python programming language, the if statement governs the program flow. The "elif" and "else" functions are seen in most if statements. This indicates that while there may be various "elif", there is only one "else". The if statement must end with the "else"; thus, no other "elif" branches can be added.
#
# + colab={"base_uri": "https://localhost:8080/"} id="wd7GD0n0yzsK" outputId="6e5da050-d68a-411e-b335-df17b65df084"
numeral1, numeral2 = 13, 14
if(numeral1 == numeral2) :
print("Yey")
elif(numeral1>numeral2):
print("Hoho")
else:
print("Awww")
# + [markdown] id="FVKdciK41tc0"
# ### Functions
# + id="fBys9uQg1gq7"
# void DeleteUser(int usedid){
# delete(userid);
# }
def delete_user (userid):
print("Successfully deleted user: {}". format(userid))
def delete_all_users ():
print("All of us are dead")
# + colab={"base_uri": "https://localhost:8080/"} id="QDe1P5Ky2ow1" outputId="5bf91622-91d6-4607-b48d-be310b920758"
userid = "Krissy"
delete_user("Krissy")
delete_all_users()
# + id="QkwuXG8v2_ET"
def add(addend1,addend2):
return addend1 + addend2
def power_of_base2(exponent):
return 2**exponent
# + colab={"base_uri": "https://localhost:8080/"} id="FKbeL9Gj3plM" outputId="ee7050ea-4d3d-489b-c5a3-fceceb7bc5ef"
addend1, addend2 = 24, 25
add(addend1, addend2)
exponent = 8
power_of_base2(exponent)
# + [markdown] id="lJyOrmlpa64U"
# ### Assignment
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="VCOai1s3a_aR" outputId="605627cb-d507-4edb-e930-8e60e2e59669"
w_pg, w_mg, w_fg = 0.30, 0.30, 0.40
name = input("Enter your name: ")
course = input("Enter your course: ")
yearlevel = input("Enter your year level: ")
pg = float(input("Enter Prelim Grade: "))
mg = float(input("Enter Midterm Grade: "))
fg = float(input("Enter Finals Grade: "))
sem_grade = (pg*w_pg) + (mg*w_mg) + (fg*w_fg)
print("Your Sem Grade is: {: .2f} ".format (sem_grade))
if(sem_grade > 70): print("\U0001F600")
elif(sem_grade == 70): print("\U0001F606")
else: print("\U0001F62D")
| Activity_1_Python_Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="KK-Ovk_OL8gb"
# Assignment 5
#
#
#
#
#
#
#
#
# + [markdown] id="D8T7g_TzC3S5"
# Konzo is a distinct upper motor neuron disease that is prevalent in sub-Saharan Africa. As part of a pilot study to investigate the relationship of the gutmicrobiome and konzo, individuals with a heavy reliance on cassava, whose consumption without proper detoxification is implicated in konzo, were assessed from regions with varying prevalence of konzo.
#
# Samples were taken from the urban capital of Kinshasa (Kin) where no outbreaks of konzo are documented. Additional samples from a rural control, Masimanimba (Mas), where no outbreaks of konzo are historically reported were also taken. Individuals from two regions of high (HPZ) and low prevalence (LPZ) of konzo from the Kahemba region were taken with unaffected (U) and konzo (K) individuals from each.
#
#
# Bacteroides and Prevotella are genus that have known associations with urban and rural lifestyles, respectively. Here we assess using the Kruskal-Wallis test where there is a significant difference in the relative abundance of these genus in the six groups, and the data is visualized using box plots.
# + id="-5LcQaiSLiVJ"
import pandas as pd
import numpy as np
import scipy
import scipy.stats
import plotly.express as px
# + colab={"base_uri": "https://localhost:8080/"} id="KvGQD8EoYJte" outputId="ecc77f19-1d67-4298-9ad7-2456204093f2"
from google.colab import drive
drive.mount('/content/drive')
# + id="tAxzzMGdhIiQ" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="805e4a0f-9d78-4aff-fd25-342d18cd452e"
#get csv file that contains read counts for genus for different samples. Header is true
#1st column contains genus names. 2nd to 4th column is additional info. Starting at colum 5 is samples read counts for wach genus
genus = pd.read_csv("/content/drive/My Drive/KinshasaControl_Konzo3_Bacteria_Genus_ReadCounts.csv")
genus
# + id="CqINnA5eheXq" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="2d5826fa-8591-47c1-a4cb-1df9cdbfe4ea"
#meta file contains sample data (such as which geographical location were the samples collected from)
meta = pd.read_csv("/content/drive/My Drive/KinshasaControl_Konzo3_sampleData.csv")
meta
# + id="2ARfL38mhs_d" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="9fbfa1ef-c884-46c2-c061-c352941f6f85"
#set NA's to 0. Remove the unnecessary colums
genus = genus.replace(np.nan,0)
genus.pop("taxRank")
genus.pop("taxID")
genus.pop("Max")
genus
# + id="ljRpXdR0cC9h"
#Make the column with genus into row names so data structure is only readcounts.
genus = genus.set_index('name')
# + colab={"base_uri": "https://localhost:8080/", "height": 640} id="R7lXUX90yRTb" outputId="adcd120f-0280-49fb-ddb8-366b7d202342"
#Conver read counts to relative abundance (done with each column since each sample is one column)
genus = genus.apply(lambda x : x / x.sum())
genus
#genus
# + id="C7NdfsMn1efx"
#transpose data frame so samples are rows.
#Also remove NA's since those got introduces for genus whose sum was 0 (making denominator 0 for relative abundance calculation)
genus_t = genus.transpose()
genus_t = genus_t.replace(np.nan,0)
# + id="w1C_ygL02WYN"
#might be a better way to do this, but convert rownames back to column so we can merge the meta file with sample name
genus_t.index.name = 'name'
genus_t.reset_index(inplace=True)
# + id="RpC6HouT3f-w"
#name column Sample to match meta file
genus_t = genus_t.rename(columns=str).rename(columns={'name':'Sample'})
# + id="GRTiufY84N7-"
#Merge meta data with genus_t
genus_tj = pd.merge(genus_t, meta, on=['Sample'])
genus_tj
# + id="cH0Se-WGiM3a"
genus_tj = genus_tj.set_index('Sample')
# + colab={"base_uri": "https://localhost:8080/"} id="vY5Y_9VA40mY" outputId="44644011-88fd-4661-9b5c-9787db7f1994"
#Do Kruskal Wallis test to see if there is a sig difference in the relative abundance of Prevotella genus between the six groups
#microbiome data tends to not be normally distributed so a non-parametric test is appropriate
#Bacteroides has been previously shown to be enriched in urban populations
bact_kw = scipy.stats.kruskal(*[group["Bacteroides"].values for name, group in genus_tj.groupby("Status")])
bact_kw
#KruskalResult(statistic=2.0190546347452027, pvalue=0.8465022320762265)
# + colab={"base_uri": "https://localhost:8080/"} id="fvhtwyFk-dYu" outputId="addaa87b-6275-4521-ee9a-d8c35cf983b2"
#Prevotella genus has previously been shown to be enriched in in rural populations
prev_kw = scipy.stats.kruskal(*[group["Prevotella"].values for name, group in genus_tj.groupby("Status")])
prev_kw
#KruskalResult(statistic=39.928496009821856, pvalue=1.5437782911043988e-07)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="z_hDba1i91nU" outputId="f9ab057b-041c-4798-ce2f-14d637dff781"
Bact = px.box(genus_tj, x="Status", y="Bacteroides", color = "Status", category_orders={ "Status" :
["Kinshasa",
"Masimanimba",
"Unaffected_Low_Prevalence_Zone",
"Konzo_Low_Prevalence_Zone",
"Unaffected_High_Prevalence_Zone",
"Konzo_High_Prevalence_Zone"]}, boxmode="overlay")
Bact.update_layout(
xaxis = dict(
tickvals = ["Kinshasa",
"Masimanimba",
"Unaffected_Low_Prevalence_Zone",
"Konzo_Low_Prevalence_Zone",
"Unaffected_High_Prevalence_Zone",
"Konzo_High_Prevalence_Zone"],
ticktext = ["Kin", "Mas", "ULPZ", "KLPZ", "UHPZ", "KHPZ"]
),
showlegend=False
)
Bact.show()
#Although Kruskal-Wallis test resulted in a p-value > 0.05, a post-hoc test may be considered to see if there is an enrichment of Bacteroides in urban population in this dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="HdM3mwq0-Qz8" outputId="7469d536-cdfb-486e-a5a1-0d05b670f3fa"
Prev = px.box(genus_tj, x="Status", y="Prevotella", color = "Status", category_orders={ "Status" :
["Kinshasa",
"Masimanimba",
"Unaffected_Low_Prevalence_Zone",
"Konzo_Low_Prevalence_Zone",
"Unaffected_High_Prevalence_Zone",
"Konzo_High_Prevalence_Zone"]}, boxmode="overlay")
Prev.update_layout(
xaxis = dict(
tickvals = ["Kinshasa",
"Masimanimba",
"Unaffected_Low_Prevalence_Zone",
"Konzo_Low_Prevalence_Zone",
"Unaffected_High_Prevalence_Zone",
"Konzo_High_Prevalence_Zone"],
ticktext = ["Kin", "Mas", "ULPZ", "KLPZ", "UHPZ", "KHPZ"]
),
showlegend=False
)
Prev.show()
#The Kruskal-Wallis test resulted in a p-value < 0.01, a post-hoc test is necessary to see if there is an enrichment of Prevotella in rural population in specific pairwise comparisons
| assignmet5_Neerja.ipynb |
# + [markdown] colab_type="text" id="copyright-notice"
# #### Copyright 2017 Google LLC.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="copyright-notice2"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="g4T-_IsVbweU"
# # 稀疏性和 L1 正则化
# + [markdown] colab_type="text" id="g8ue2FyFIjnQ"
# **学习目标:**
# * 计算模型大小
# * 通过应用 L1 正则化来增加稀疏性,以减小模型大小
# + [markdown] colab_type="text" id="ME_WXE7cIjnS"
# 降低复杂性的一种方法是使用正则化函数,它会使权重正好为零。对于线性模型(例如线性回归),权重为零就相当于完全没有使用相应特征。除了可避免过拟合之外,生成的模型还会更加有效。
#
# L1 正则化是一种增加稀疏性的好方法。
#
#
# + [markdown] colab_type="text" id="fHRzeWkRLrHF"
# ## 设置
#
# 运行以下单元格,以加载数据并创建特征定义。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="pb7rSrLKIjnS"
from __future__ import print_function
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.cn/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="3V7q8jk0IjnW"
def preprocess_features(california_housing_dataframe):
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
["latitude",
"longitude",
"housing_median_age",
"total_rooms",
"total_bedrooms",
"population",
"households",
"median_income"]]
processed_features = selected_features.copy()
# Create a synthetic feature.
processed_features["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] /
california_housing_dataframe["population"])
return processed_features
def preprocess_targets(california_housing_dataframe):
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = pd.DataFrame()
# Create a boolean categorical feature representing whether the
# median_house_value is above a set threshold.
output_targets["median_house_value_is_high"] = (
california_housing_dataframe["median_house_value"] > 265000).astype(float)
return output_targets
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="pAG3tmgwIjnY"
# Choose the first 12000 (out of 17000) examples for training.
training_examples = preprocess_features(california_housing_dataframe.head(12000))
training_targets = preprocess_targets(california_housing_dataframe.head(12000))
# Choose the last 5000 (out of 17000) examples for validation.
validation_examples = preprocess_features(california_housing_dataframe.tail(5000))
validation_targets = preprocess_targets(california_housing_dataframe.tail(5000))
# Double-check that we've done the right thing.
print("Training examples summary:")
display.display(training_examples.describe())
print("Validation examples summary:")
display.display(validation_examples.describe())
print("Training targets summary:")
display.display(training_targets.describe())
print("Validation targets summary:")
display.display(validation_targets.describe())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="gHkniRI1Ijna"
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="bLzK72jkNJPf"
def get_quantile_based_buckets(feature_values, num_buckets):
quantiles = feature_values.quantile(
[(i+1.)/(num_buckets + 1.) for i in range(num_buckets)])
return [quantiles[q] for q in quantiles.keys()]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="al2YQpKyIjnd"
def construct_feature_columns():
"""Construct the TensorFlow Feature Columns.
Returns:
A set of feature columns
"""
bucketized_households = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("households"),
boundaries=get_quantile_based_buckets(training_examples["households"], 10))
bucketized_longitude = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("longitude"),
boundaries=get_quantile_based_buckets(training_examples["longitude"], 50))
bucketized_latitude = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("latitude"),
boundaries=get_quantile_based_buckets(training_examples["latitude"], 50))
bucketized_housing_median_age = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("housing_median_age"),
boundaries=get_quantile_based_buckets(
training_examples["housing_median_age"], 10))
bucketized_total_rooms = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("total_rooms"),
boundaries=get_quantile_based_buckets(training_examples["total_rooms"], 10))
bucketized_total_bedrooms = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("total_bedrooms"),
boundaries=get_quantile_based_buckets(training_examples["total_bedrooms"], 10))
bucketized_population = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("population"),
boundaries=get_quantile_based_buckets(training_examples["population"], 10))
bucketized_median_income = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("median_income"),
boundaries=get_quantile_based_buckets(training_examples["median_income"], 10))
bucketized_rooms_per_person = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("rooms_per_person"),
boundaries=get_quantile_based_buckets(
training_examples["rooms_per_person"], 10))
long_x_lat = tf.feature_column.crossed_column(
set([bucketized_longitude, bucketized_latitude]), hash_bucket_size=1000)
feature_columns = set([
long_x_lat,
bucketized_longitude,
bucketized_latitude,
bucketized_housing_median_age,
bucketized_total_rooms,
bucketized_total_bedrooms,
bucketized_population,
bucketized_households,
bucketized_median_income,
bucketized_rooms_per_person])
return feature_columns
# + [markdown] colab_type="text" id="hSBwMrsrE21n"
# ## 计算模型大小
#
# 要计算模型大小,只需计算非零参数的数量即可。为此,我们在下面提供了一个辅助函数。该函数深入使用了 Estimator API,如果不了解它的工作原理,也不用担心。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="e6GfTI0CFhB8"
def model_size(estimator):
variables = estimator.get_variable_names()
size = 0
for variable in variables:
if not any(x in variable
for x in ['global_step',
'centered_bias_weight',
'bias_weight',
'Ftrl']
):
size += np.count_nonzero(estimator.get_variable_value(variable))
return size
# + [markdown] colab_type="text" id="XabdAaj67GfF"
# ## 减小模型大小
#
# 您的团队需要针对 *SmartRing* 构建一个准确度高的逻辑回归模型,这种指环非常智能,可以感应城市街区的人口统计特征(`median_income`、`avg_rooms`、`households` 等等),并告诉您指定城市街区的住房成本是否高昂。
#
# 由于 SmartRing 很小,因此工程团队已确定它只能处理**参数数量不超过 600 个**的模型。另一方面,产品管理团队也已确定,除非所保留测试集的**对数损失函数低于 0.35**,否则该模型不能发布。
#
# 您可以使用秘密武器“L1 正则化”调整模型,使其同时满足大小和准确率限制条件吗?
# + [markdown] colab_type="text" id="G79hGRe7qqej"
# ### 任务 1:查找合适的正则化系数。
#
# **查找可同时满足以下两种限制条件的 L1 正则化强度参数:模型的参数数量不超过 600 个且验证集的对数损失函数低于 0.35。**
#
# 以下代码可帮助您快速开始。您可以通过多种方法向您的模型应用正则化。在此练习中,我们选择使用 `FtrlOptimizer` 来应用正则化。`FtrlOptimizer` 是一种设计成使用 L1 正则化比标准梯度下降法得到更好结果的方法。
#
# 重申一次,我们会使用整个数据集来训练该模型,因此预计其运行速度会比通常要慢。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="1Fcdm0hpIjnl"
def train_linear_classifier_model(
learning_rate,
regularization_strength,
steps,
batch_size,
feature_columns,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a linear regression model.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
regularization_strength: A `float` that indicates the strength of the L1
regularization. A value of `0.0` means no regularization.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
feature_columns: A `set` specifying the input feature columns to use.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `LinearClassifier` object trained on the training data.
"""
periods = 7
steps_per_period = steps / periods
# Create a linear classifier object.
my_optimizer = tf.train.FtrlOptimizer(learning_rate=learning_rate, l1_regularization_strength=regularization_strength)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
linear_classifier = tf.estimator.LinearClassifier(
feature_columns=feature_columns,
optimizer=my_optimizer
)
# Create input functions.
training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value_is_high"],
batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value_is_high"],
num_epochs=1,
shuffle=False)
predict_validation_input_fn = lambda: my_input_fn(validation_examples,
validation_targets["median_house_value_is_high"],
num_epochs=1,
shuffle=False)
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print("Training model...")
print("LogLoss (on validation data):")
training_log_losses = []
validation_log_losses = []
for period in range (0, periods):
# Train the model, starting from the prior state.
linear_classifier.train(
input_fn=training_input_fn,
steps=steps_per_period
)
# Take a break and compute predictions.
training_probabilities = linear_classifier.predict(input_fn=predict_training_input_fn)
training_probabilities = np.array([item['probabilities'] for item in training_probabilities])
validation_probabilities = linear_classifier.predict(input_fn=predict_validation_input_fn)
validation_probabilities = np.array([item['probabilities'] for item in validation_probabilities])
# Compute training and validation loss.
training_log_loss = metrics.log_loss(training_targets, training_probabilities)
validation_log_loss = metrics.log_loss(validation_targets, validation_probabilities)
# Occasionally print the current loss.
print(" period %02d : %0.2f" % (period, validation_log_loss))
# Add the loss metrics from this period to our list.
training_log_losses.append(training_log_loss)
validation_log_losses.append(validation_log_loss)
print("Model training finished.")
# Output a graph of loss metrics over periods.
plt.ylabel("LogLoss")
plt.xlabel("Periods")
plt.title("LogLoss vs. Periods")
plt.tight_layout()
plt.plot(training_log_losses, label="training")
plt.plot(validation_log_losses, label="validation")
plt.legend()
return linear_classifier
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="9H1CKHSzIjno"
linear_classifier = train_linear_classifier_model(
learning_rate=0.1,
# TWEAK THE REGULARIZATION VALUE BELOW
regularization_strength=0.0,
steps=300,
batch_size=100,
feature_columns=construct_feature_columns(),
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
print("Model size:", model_size(linear_classifier))
# + [markdown] colab_type="text" id="yjUCX5LAkxAX"
# ### 解决方案
#
# 点击下方即可查看可能的解决方案。
# + [markdown] colab_type="text" id="hgGhy-okmkWL"
# 正则化强度为 0.1 应该就足够了。请注意,有一个需要做出折中选择的地方:正则化越强,我们获得的模型就越小,但会影响分类损失。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="_rV8YQWZIjns"
linear_classifier = train_linear_classifier_model(
learning_rate=0.1,
regularization_strength=0.1,
steps=300,
batch_size=100,
feature_columns=construct_feature_columns(),
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
print("Model size:", model_size(linear_classifier))
| ml/cc/exercises/zh-CN/sparsity_and_l1_regularization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Run fidimag inside a Docker Container
# ## Setup Docker
# Install Docker, follow instructions at https://www.docker.com/products/docker
#
#
# ## Setup fidimag docker container with Jupyter Notebook
#
# Pull the fidimag notebook container:
#
# ```
# docker pull fidimag/notebook
# ```
# ## Start Notebook on container
# ```
# docker run -p 30000:8888 fidimag/notebook
# ```
#
# This command starts a [Jupyter Notebook](http://jupyter.org) in which [fidimag]() can be used. The Jupyter notebook inside the container is listening on port 8888.
#
# The parameter `-p 30000:8888` says that the port 8888 inside the container is exposed on the host system as port 30000.
#
# On a Linux host machine, you can connect to `http://localhost:30000` to see the notebook.
#
# On a Mac, you need to find out the right IP address to which to connect. The information is provided by
# ```
# docker-machine ip
# ```
# For example, if `docker-machine ip` returns `192.168.99.100`, then the right URL to paste into the browser on the host system is `http://192.168.99.100:30000`.
#
# [*How does this work on Windows? Pull requests welcome.*]
# ## Detach the docker image (to run in the background)
#
# You can add the `-d` switch to the `docker run` command to *detach* the process:
# ```
# docker run -d -p 30000:8888 fidimag/notebook
# ```
# ## Show active Docker containers
# ```docker ps``` lists all running containers.
#
# To only show the `id`s, we can use
#
# docker ps -q
#
# To only show the containers that was last started, we can use the `-l` flag:
#
# docker ps -l
# ## Stop a docker container
#
# To stop the last container started, we can use the ```docker stop ID``` command, where we need to find the `ID` first. We can do this using ```docker ps -l -q```. Putting the commands together, we have
# ```
# docker stop $(docker ps -l -q)
# ```
# to stop the last container we started.
# ## Explore the docker container / run Fidimag from (I)Python
#
# We can start the docker container with the `-ti` switch, and we can provide `bash` as the command to execute:
# ```
# docker run -ti fidimag/notebook bash
# ```
#
# A bash prompt appears (and we are now inside the container):
# ```
# jovyan@4df962d27520:~/work$
# ```
#
# and can start Python inside the container, and import fidimag:
#
# ```
# jovyan@4df962d27520:~/work$ python
# Python 3.5.1 |Continuum Analytics, Inc.| (default, Jun 15 2016, 15:32:45)
# [GCC 4.4.7 20120313 (Red Hat 4.4.7-1)] on linux
# Type "help", "copyright", "credits" or "license" for more information.
# >>> import fidimag
# ```
#
# We could also start IPython:
# ```
# jovyan@4df962d27520:~/work$ ipython
# Python 3.5.1 |Continuum Analytics, Inc.| (default, Jun 15 2016, 15:32:45)
# Type "copyright", "credits" or "license" for more information.
#
# IPython 4.2.0 -- An enhanced Interactive Python.
# ? -> Introduction and overview of IPython's features.
# # %quickref -> Quick reference.
# help -> Python's own help system.
# object? -> Details about 'object', use 'object??' for extra details.
#
# In [1]:
# ```
#
# [The switch `-t` stands for `Allocate a pseudo-TTY` and `-i` for `Keep STDIN open even if not attached`.]
# ## Mount the local file system to exchange files and data with container
# Often, we may have a fidimag Python script `run.py` we want to execute in our current working directory in the host machine. We want output files from that command to be written to the same working directory on the host machine.
#
# We can use the container like this to achieve that:
#
# ```
# docker run -v `pwd`:/io -ti fidimag/notebook python run.py```
#
# The ```-v `pwd`:/io``` tells the docker container to take the current working directory on the host (``` `pwd` ```) and mount it to the path `/io` on the container. The container is set up so that the default working directory is `/io`.
# Here is an example file `run.py` that reads
# ```
# import fidimag # to proof we can import it
# print("Hello from the container")
# # and write to a file
# open("data.txt", "w").write("Data from the container.\n")
# ```
#
# This can be executed with
# ```
# docker run -v `pwd`:/io -ti fidimag/notebook python hello.py
# ```
# and will create a data file `data.txt` that is visible from the host's working directory.
#
#
# ## Explore Jupyter notebook examples with the Docker container
# ```
# git clone https://github.com/computationalmodelling/fidimag.git
# # cd fidimag/doc/ipynb/
# docker run -v `pwd`:/io -p 30000:8888 -d fidimag/notebook
# ```
#
#
# ## Use smaller docker containers
# Two alternative docker containers are available that provide only Fidimag, but not the Jupyter Notebook, nor scipy. They are available under the names `fidimag/minimal-py2` and `fidimag/minimal-py3`. Use these names instead of `fidimag/notebook` in the examples above.
#
# The `fidimag/minimal-py2` version uses Python2, the `fidimag/minimal-py3` version uses Python 3.
| doc/user_guide/ipynb/tutorial-docker-container.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cmath
z = complex(input())
print(*cmath.polar(z), sep='\n')
# +
# another way
import cmath
c = complex(input())
r, phi = cmath.polar(c)
print(r, phi, sep='\n')
| Python/6. Math/31. polar coordinates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # 数值稳定性和模型初始化
#
# 理解了正向传播与反向传播以后,我们来讨论一下深度学习模型的数值稳定性问题以及模型参数的初始化方法。深度模型有关数值稳定性的典型问题是衰减(vanishing)和爆炸(explosion)。
#
#
# ## 衰减和爆炸
#
# 当神经网络的层数较多时,模型的数值稳定性容易变差。假设一个层数为$L$的多层感知机的第$l$层$\boldsymbol{H}^{(l)}$的权重参数为$\boldsymbol{W}^{(l)}$,输出层$\boldsymbol{H}^{(L)}$的权重参数为$\boldsymbol{W}^{(L)}$。为了便于讨论,不考虑偏差参数,且设所有隐藏层的激活函数为恒等映射(identity mapping)$\phi(x) = x$。给定输入$\boldsymbol{X}$,多层感知机的第$l$层的输出$\boldsymbol{H}^{(l)} = \boldsymbol{X} \boldsymbol{W}^{(1)} \boldsymbol{W}^{(2)} \ldots \boldsymbol{W}^{(l)}$。此时,如果层数$l$较大,$\boldsymbol{H}^{(l)}$的计算可能会出现衰减或爆炸。举个例子,假设输入和所有层的权重参数都是标量,比如权重参数为0.2和5,多层感知机的第30层输出为输入$\boldsymbol{X}$分别与$0.2^{30} \approx 1 \times 10^{-21}$(衰减)和$5^{30} \approx 9 \times 10^{20}$(爆炸)的乘积。类似地,当层数较多时,梯度的计算也更容易出现衰减或爆炸。
#
# 随着内容的不断深入,我们会在后面的章节进一步介绍深度学习的数值稳定性问题以及解决方法。
#
#
# ## 随机初始化模型参数
#
# 在神经网络中,我们通常需要随机初始化模型参数。下面我们来解释这样做的原因。
#
# 回顾[“多层感知机”](mlp.md)一节图3.3描述的多层感知机。为了方便解释,假设输出层只保留一个输出单元$o_1$(删去$o_2, o_3$和指向它们的箭头),且隐藏层使用相同的激活函数。如果将每个隐藏单元的参数都初始化为相等的值,那么在正向传播时每个隐藏单元将根据相同的输入计算出相同的值,并传递至输出层。在反向传播中,每个隐藏单元的参数梯度值相等。因此,这些参数在使用基于梯度的优化算法迭代后值依然相等。之后的迭代也是如此。这种情况下,无论隐藏单元有多少,隐藏层本质上只有1个隐藏单元在发挥作用。因此,正如我们在前面的实验中所做的那样,我们通常将神经网络的模型参数,特别是权重参数,进行随机初始化。
#
#
# ### MXNet的默认随机初始化
#
# 随机初始化模型参数的方法有很多。在[“线性回归的简洁实现”](linear-regression-gluon.md)一节中,我们使用`net.initialize(init.Normal(sigma=0.01))`使模型`net`的权重参数采用正态分布的随机初始化方式。如果不指定初始化方法,例如`net.initialize()`,MXNet将使用默认的随机初始化方法:权重参数每个元素随机采样于-0.07到0.07之间的均匀分布,偏差参数全部清零。
#
#
# ### Xavier随机初始化
#
# 还有一种比较常用的随机初始化方法叫做Xavier随机初始化 [1]。
# 假设某全连接层的输入个数为$a$,输出个数为$b$,Xavier随机初始化将使得该层中权重参数的每个元素都随机采样于均匀分布
#
# $$U\left(-\sqrt{\frac{6}{a+b}}, \sqrt{\frac{6}{a+b}}\right).$$
#
# 它的设计主要考虑到,模型参数初始化后,每层输出的方差不该受该层输入个数影响,且每层梯度的方差也不该受该层输出个数影响。
#
# ## 小结
#
# * 深度模型有关数值稳定性的典型问题是衰减和爆炸。当神经网络的层数较多时,模型的数值稳定性容易变差。
# * 我们通常需要随机初始化神经网络的模型参数。
#
#
# ## 练习
#
# * 有人说随机初始化模型参数是为了“打破对称性”。这里的“对称”应如何理解?
# * 我们是否可以将线性回归或softmax回归中所有的权重参数都初始化为相同值?
#
#
# ## 扫码直达[讨论区](https://discuss.gluon.ai/t/topic/8052)
#
# 
#
# ## 参考文献
#
# [1] <NAME>., & <NAME>. (2010, March). Understanding the difficulty of training deep feedforward neural networks. In Proceedings of the thirteenth international conference on artificial intelligence and statistics (pp. 249-256).
| chapter_deep-learning-basics/numerical-stability-and-init.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
d = [0,1,2,3,4,5,6,7,8,9]
df = pd.DataFrame(d)
df
df.columns = ['Rev']
df
df['NewCol'] = 5
df
del df['NewCol']
df
df['test'] = 3
df['col'] = df['Rev']
df
i = ['a','b','c','d','e','f','g','h','i','j']
df.index = i
df
df.loc['a']
df.loc['a':'d']
df.iloc[0:3]
df['Rev']
df[['Rev', 'test']]
df.loc[df.index[5:],'col']
df.loc[df.index[:3],['col', 'test']]
#показывает первые пять записей (настройка по умолчанию)
df.head()
#показывает последние пять записей (настройка по умолчанию)
df.tail()
| learning python/learning pandas/pandas_lec_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## LeNet
#
# 
# + slideshow={"slide_type": "slide"}
import d2l
import mxnet as mx
from mxnet import autograd, gluon, init, nd
from mxnet.gluon import loss as gloss, nn
import time
net = nn.Sequential()
net.add(nn.Conv2D(channels=6, kernel_size=5, padding=2, activation='sigmoid'),
nn.AvgPool2D(pool_size=2, strides=2),
nn.Conv2D(channels=16, kernel_size=5, activation='sigmoid'),
nn.AvgPool2D(pool_size=2, strides=2),
# Dense will transform the input of the shape (batch size, channel, height, width)
# into the input of the shape (batch size, channel *height * width) automatically
# by default.
nn.Dense(120, activation='sigmoid'),
nn.Dense(84, activation='sigmoid'),
nn.Dense(10))
# + [markdown] slideshow={"slide_type": "slide"}
# Feeding a single observation through the network
# -
X = nd.random.uniform(shape=(1, 1, 28, 28))
net.initialize()
for layer in net:
X = layer(X)
print(layer.name, 'output shape:\t', X.shape)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data and training
# -
batch_size = 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size=batch_size)
# +
# use a GPU if we have it
def try_gpu():
try:
ctx = mx.gpu()
_ = nd.zeros((1,), ctx=ctx)
except mx.base.MXNetError:
ctx = mx.cpu()
return ctx
ctx = try_gpu()
ctx
# + [markdown] slideshow={"slide_type": "slide"}
# ## Accuracy
# -
def evaluate_accuracy(data_iter, net, ctx):
acc_sum, n = nd.array([0], ctx=ctx), 0
for X, y in data_iter:
# If ctx is the GPU, copy the data to the GPU.
X, y = X.as_in_context(ctx), y.as_in_context(ctx).astype('float32')
acc_sum += (net(X).argmax(axis=1) == y).sum()
n += y.size
return acc_sum.asscalar() / n
# + [markdown] slideshow={"slide_type": "slide"}
# ## Training loop
# -
# This function has been saved in the d2l package for future use.
def train(net, train_iter, test_iter, batch_size, trainer, ctx,
num_epochs):
print('training on', ctx)
loss = gloss.SoftmaxCrossEntropyLoss()
for epoch in range(num_epochs):
train_l_sum, train_acc_sum, n, start = 0.0, 0.0, 0, time.time()
for X, y in train_iter:
X, y = X.as_in_context(ctx), y.as_in_context(ctx)
with autograd.record():
y_hat = net(X)
l = loss(y_hat, y).sum()
l.backward()
trainer.step(batch_size)
y = y.astype('float32')
train_l_sum += l.asscalar()
train_acc_sum += (y_hat.argmax(axis=1) == y).sum().asscalar()
n += y.size
test_acc = evaluate_accuracy(test_iter, net, ctx)
print('epoch %d, loss %.4f, train acc %.3f, test acc %.3f, '
'time %.1f sec'
% (epoch + 1, train_l_sum / n, train_acc_sum / n, test_acc,
time.time() - start))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Network initialization and training
# -
lr, num_epochs = 0.9, 5
net.initialize(force_reinit=True, ctx=ctx, init=init.Xavier())
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': lr})
train(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)
| slides/2_28/lenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Cycle through each year and extract the major topics
import os
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.externals import joblib
from sklearn.feature_extraction.text import TfidfVectorizer
import operator
from sklearn.externals import joblib
from sklearn import decomposition
# %matplotlib inline
total_bldgsim = pd.read_pickle("/Users/nus/twenty-years-of-bldgsim-textmining/processed_data/total_email_data.pkl")
total_bldgsim.info()
# # Loop through each year and calculate the tf-idf and the topics for each year
def rank_terms( A, terms ):
# get the sums over each column
sums = A.sum(axis=0)
# map weights to the terms
weights = {}
for col, term in enumerate(terms):
weights[term] = sums[0,col]
# rank the terms by their weight over all documents
return sorted(weights.items(), key=operator.itemgetter(1), reverse=True)
def get_descriptor( terms, H, topic_index, top ):
# reverse sort the values to sort the indices
top_indices = np.argsort( H[topic_index,:] )[::-1]
# now get the terms corresponding to the top-ranked indices
top_terms = []
for term_index in top_indices[0:top]:
top_terms.append( terms[term_index] )
top_terms = [item for item in top_terms if not item.isdigit()]
return top_terms
k = 6
# # Run through each year, get the top 10 topics and their top 10 words
#
# Put them in a dataframe with the year
all_topics = []
condensed_topics = []
for year in total_bldgsim.index.year.unique():
bldg_sim_peryear = total_bldgsim[total_bldgsim.index.year == year]
out_array = np.array(bldg_sim_peryear.fillna("x").Body)
custom_stop_words = []
with open( "/Users/nus/twenty-years-of-bldgsim-textmining/processed_data/stopwords_annual_analysis_3.txt", "r" ) as fin:
for line in fin.readlines():
custom_stop_words.append( line.strip() )
print(year)
# use a custom stopwords list, set the minimum term-document frequency to 20
vectorizer = CountVectorizer(stop_words = custom_stop_words, min_df = 20, encoding='latin-1')
A = vectorizer.fit_transform(out_array)
print( "Created %d X %d document-term matrix" % (A.shape[0], A.shape[1]) )
# extract the resulting vocabulary
terms = vectorizer.get_feature_names()
print("Vocabulary has %d distinct terms" % len(terms))
# we can pass in the same preprocessing parameters
vectorizer = TfidfVectorizer(stop_words=custom_stop_words, min_df = 20, encoding='latin-1')
A = vectorizer.fit_transform(out_array)
print("Created %d X %d TF-IDF-normalized document-term matrix" % (A.shape[0], A.shape[1]) )
joblib.dump((A,terms), "/Users/nus/twenty-years-of-bldgsim-textmining/pkl_files/emails-raw"+str(year)+".pkl")
(A,terms) = joblib.load( "/Users/nus/twenty-years-of-bldgsim-textmining/pkl_files/emails-raw"+str(year)+".pkl" )
print( "Loaded %d X %d document-term matrix" % (A.shape[0], A.shape[1]) )
model = decomposition.NMF( init="nndsvd", n_components=k )
# apply the model and extract the two factor matrices
W = model.fit_transform( A )
H = model.components_
descriptors = []
condensed = []
for topic_index in range(k):
descriptors.append( get_descriptor( terms, H, topic_index, 5) )
str_descriptor = ", ".join( descriptors[topic_index] )
condensed.append(str_descriptor)
print("Topic %02d: %s" % ( topic_index+1, str_descriptor ) )
condensed_topics.append(condensed)
descriptors_df = pd.DataFrame(descriptors).T
descriptors_df['year'] = year
all_topics.append(descriptors_df)
condensed_topics = pd.DataFrame(condensed_topics)
condensed_topics.to_excel("/Users/nus/twenty-years-of-bldgsim-textmining/processed_data/condensed_topics.xlsx")
all_topics_df = pd.concat(all_topics).reset_index(drop=True)
all_topics_df.info()
all_topics_df
all_topics_df_melted = all_topics_df.melt(id_vars='year')
all_topics_df_melted.head()
all_topics_df_melted.info()
wordfreq = all_topics_df_melted.value.value_counts()
wordfreq.head(40)
wordfreq
wordfreq.to_csv("/Users/nus/twenty-years-of-bldgsim-textmining/processed_data/word_frequency.csv")
| .ipynb_checkpoints/07_tf-idf extraction and processing for each year - 6 topics per year of 5 words-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CRC Make-A-Thon Crash Course - API Number Exercise
# Lets pick a API_14 number and assign it to a string variable.
API_14 = '04030565740000'
print(API_14)
# Lets then use API_14 to split the sting and find API_12, API_10, and API_8
# +
API_12 = API_14[:12]
print(API_12)
API_10 = API_14[:10]
print(API_10)
API_08 = API_14[2:-4]
print(API_08)
# -
# ## Exercise 1:
# ## Lets present the results in a few ways
# ### 1. Try presenting the results in a simple way by inserting the results into strings.
# '\t' creates a tab seporator within the string.
#example
value = '####'
print('Name:\t' + value)
#Answer
print('API14:\t' + API_14)
print('API12:\t' + API_12)
print('API10:\t' + API_10)
print('API08:\t' + API_08)
# ### 2. Try presenting the results by passing the API strings to one string statement.
# '\n' creates a new line within the string.
#example
str1 = '{}:\t{}'.format('Name1', value)
str2 = '{}:\t{}'.format('Name2', value)
statement = str1 + '\n' + str2
print(statement)
#Answer
str1 = '{}:\t{}'.format('API14', API_14)
str2 = '{}:\t{}'.format('API12', API_12)
str3 = '{}:\t{}'.format('API10', API_10)
str4 = '{}:\t{}'.format('API08', API_08)
statement = str1 + '\n' + str2 + '\n' + str3 + '\n' + str4
print(statement)
print(type(statement))
# ### 3. Try presenting the results by iterating over a list of the results.
# First create two lists for our API varible names and values. Then use a for loop to print API_no and API_vlaue iterating over a range from 0 to length of API names.
# +
#example
Names = ['Name1','Name2']
print(Names)
Values = ['####','####']
print(Values)
# +
#example
r = range(0, len(Names))
for i in r:
print('{}:\t{}'.format(Names[i],Values[i]))
# +
#Answer
names = ['API14','API12','API10','API08']
print(names)
values = [API_14, API_12, API_10, API_08]
print(values)
# +
#Answer
numbers = range(0, len(names))
for i in numbers:
print('{}:\t{}'.format(names[i],values[i]))
# -
# ### 4. Try presenting the results by iterating over a dictionary of the results
# First create a dictionary of our API variables. Then use a for loop to print API variables iterating through the dict.
#example
Name_dict = {'Name1':'#####', 'Name2':'#####'}
print(Name_dict)
#example
for key, value in Name_dict.items():
print('{}:\t{}'.format(key,value))
#Answer
name_dict = {'API14':API_14,'API12':API_12,'API10':API_10,'API08':API_08}
print(name_dict)
#Answer
for key, value in name_dict.items():
print('{}:\t{}'.format(key,value))
# # Exercise 2:
# # Lets assume our well was redrilled and will need an updated API_14.
# For redrilled wells the API_12 value updates by 1. Therefore 04030565740000 becomes 04030565740100.
# ### 1. Try updating the API_14 using a sting replace.
# example
Name = 'James'
new_Name = Name.replace('es','ie')
print('new_Name:\t' + new_Name)
#Answer
name = API_14
new_name = name.replace('0000','0100')
print('old_name:\t' + name)
print('new_name:\t' + new_name)
# ### 2. Try updating the API_14 by finding the API_10 and add the updated last four values.
# +
#example
Name = '012345'
Name = Name[:2]
new_Name = Name + '678'
print('new_Name:\t' + new_Name)
# -
#Answer
name = API_14
new_name2 = name[:-4] + '0100'
print('new_name2:\t' + new_name2)
# ### 3. Try updating the API_14 by splitingthe API_14 before and after the 12 value.
# +
#example
Name = '012345'
new_Name = Name[:2] + '1' + Name[3:]
print('new_Name:\t' + new_Name)
# +
#Answer
name = API_14
new_name3 = name[:11] + '1' + name[12:]
print('new_name3:\t' + new_name3)
# -
# ### 4. FYI. We can also change the API_12 directly by first turning the string to a list and replace the 12th number then joining the list back to one string.
# +
#Answer
l = list(API_14)
l[11] = '1'
new_api_14 = ''.join(l)
print(l)
print(new_api_14)
# -
| CRC Make A Thon Crash Course API Number Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CSC413 - Tutorial 2
# ## Tutorial overview
#
# - Introduction to interactive notebooks
#
# - Introduction to Autograd and PyTorch
#
# - Walk-through several PyTorch models (with increasing levels of complexity and abstraction
# ## Interactive notebooks
#
# ### Overview
#
# - Keep documentation, code, and outputs in the same file.
#
# - Many different implementations (e.g. Jupyter, Google Colab, Zeppelin, etc.)
#
# - Many different languages (e.g. Python, C++ (xeus), Scala, etc.)
#
# ### Jupyter notebooks / lab
#
# -
#
# ### Google Colab
#
# - Hosted interactive notebooks (similar to Jupyter notebooks).
#
# - Can get a free GPU or TPU (for at most 12 hours...).
#
# - (Note to self:) Make sure to select a runtime with a GPU.
# + [markdown] colab_type="text" id="oW6E9VvcmYpd"
# ## Automatic differentiation
#
# References:
#
# - <NAME>' talk: https://www.youtube.com/watch?v=sq2gPzlrM0g
# - Backpropagation notes from Stanford's CS231n: http://cs231n.github.io/optimization-2/
# - Autograd Github Repository (contains a tutorial and examples): https://github.com/HIPS/autograd
#
# ### Approaches for Computing Derivatives
#
# - **Symbolic differentiation:** automatic manipulation of mathematical expressions to get derivatives
# - Takes a math expression and returns a math expression: $f(x) = x^2 \rightarrow \frac{df(x)}{dx} = 2x$
# - Used in Mathematica, Maple, SymPy, etc.
#
# - **Numeric differentiation:** Approximating derivatives by finite differences:
# $$
# \frac{\partial}{\partial x_i} f(x_1, \dots, x_N) = \lim_{h \to 0} \frac{f(x_1, \dots, x_i + h, \dots, x_N) - f(x_1, \dots, x_i - h, \dots, x_N)}{2h}
# $$
#
# - **Automatic differentiation:** Takes code that computes a function and returns code that computes the derivative of that function.
# - Reverse Mode AD: A method to get exact derivatives efficiently, by storing information as you go forward that you can reuse as you go backwards
# - _"The goal isn't to obtain closed-form solutions, but to be able to wirte a program that efficiently computes the derivatives."_ - Lecture 6 Slides (Backpropagation)
# - **PyTorch**, **Autograd**, **JAX**
# + [markdown] colab_type="text" id="oW6E9VvcmYpd"
# ### Reverse mode automatic differentiation
#
# In machine learning, we have functions that have large fan-in, e.g. a neural net can have millions of parameters, that all squeeze down to one scalar that tells you how well it predicts something.
#
# #### General idea for implementation
#
# * Create a "tape" data structure that tracks the operations performed in computing a function.
# * Overload primitives to:
# - Add themselves to the tape when called.
# - Compute gradients with respect to their local inputs.
# * _Forward pass_ computes the function, and adds operations to the tape.
# * _Reverse pass_ accumulates the local gradients using the chain rule.
# * This is efficient for graphs with large fan-in, like most loss functions in ML.
# + [markdown] colab_type="text" id="0JxunitHmYpf"
# ### Autograd
#
# * [Autograd](https://github.com/HIPS/autograd) is a Python package for automatic differentiation.
# * To install Autograd:
# pip install autograd
# * There are a lot of great [examples](https://github.com/HIPS/autograd/tree/master/examples) provided with the source code.
#
# ### What can Autograd do?
#
# From the Autograd Github repository:
#
# * Autograd can automatically differentiate native Python and Numpy code.
# * It can handle a large subset of Python's features, including loops, conditional statements (if/else), recursion and closures
# * It can also compute higher-order derivatives
# * It uses reverse-mode differentiation (a.k.a. backpropagation) so it can efficiently take gradients of scalar-valued functions with respect to array-valued arguments.
#
#
# ### Autograd vs Tensorflow, Theano, etc.
#
# Many deep learning packages implement automatic differentiation using small _domain-specific languages_ within Python:
# - Theano
# - Caffe
# - Vanilla Torch (as compared to Autograd for Torch)
# - Tensorflow
#
# Most of these alternatives require you to _explicitly_ construct a computation graph; Autograd constructs a computation graph _implicitly_, by tracking the sequence of operations that have been performed during the execution of a program.
#
# + [markdown] colab_type="text" id="1kDjpkTqmYph"
# ### Autograd basic usage
# -
import autograd.numpy as jnp # Import thinly-wrapped numpy
from autograd import grad # Basicallly the only autograd function you need
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="szDksM6wmYpn" outputId="f0f05894-4e66-4679-8db0-c8a2424c2237"
# Define a function like normal, using Python and (autograd's) NumPy
def tanh(x):
y = jnp.exp(-x)
return (1.0 - y) / (1.0 + y)
# Create a *function* that computes the gradient of tanh
grad_tanh = grad(tanh)
# Evaluate the gradient at x = 1.0
print(grad_tanh(1.0))
# Compare to numeric gradient computed using finite differences
print((tanh(1.0001) - tanh(0.9999)) / 0.0002)
# + [markdown] colab_type="text" id="SDGKdlknJMuK"
# ## PyTorch
#
# - Makes it possible to work with arrays and tensors efficiently in Python (wraps NumPy for CPU tensors).
#
# - Adds GPU support.
#
# - Adds automatic differentiation.
#
# - Provides a high-level abstractions for working with neural networks.
# +
import matplotlib.pyplot as plt
import numpy as np
import numpy.random as npr
import torch
# %matplotlib inline
# -
# ### PyTorch — API
#
# See: https://pytorch.org/docs/stable/index.html (especially `torch`, `torch.nn`, `torch.nn.functional`, and `torch.Tensor`).
# +
x = torch.tensor([1.0, 2.0, 3.0, 4.0])
x
# +
x_np = np.array([1.0, 2.0, 3.0, 4.0], dtype=np.float32)
x = torch.from_numpy(x_np)
# Torch abstracts over NumPy but uses a NumPy-compatible representation under the hood
x_np[0] = 100.0
x[1] = 200.0
x.data.numpy()[2] = 300
x
# -
# Broadcasting
x.reshape(-1, 1) + x
# Dot product
x @ x.T
# ### PyTorch — GPU support
#
# We can move PyTorch tensors to the GPU, which allows us to perform some computations much faster.
# +
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
# +
x = x.to(device=device)
x
# -
x @ x.T
# + [markdown] colab_type="text" id="SDGKdlknJMuK"
# ### PyTorch — Automatic differentiation
#
# PyTorch allows us to dynamically define computational graphs that can be evaluated efficently on GPUs.
# -
data = torch.tensor([1.0])
data
param = torch.tensor([1.0], requires_grad=True)
param
(data + param)
# For a more concrete example, let's work with the function:
#
# $$f(x) = x^2 + 2x + 6$$
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="lKC3Lzwqs07E" outputId="ef425156-f89a-43ac-aa1d-1671544c6065"
def f(x):
return x ** 2 + 2 * x + 6
np_x = np.array([4.0])
x = torch.from_numpy(np_x).requires_grad_(True)
y = f(x)
print(y)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="kWQJmBlytIOV" outputId="2075ad1e-e13a-4f3b-98b7-4991060cf467"
y.backward()
x.grad
# + colab={} colab_type="code" id="nCdN3mtXtX-N"
np_x = np.array([5.0])
x = torch.from_numpy(np_x).requires_grad_(True)
y = f(x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="2TWXc5gmtdrA" outputId="2e058699-1e92-470b-8128-749cb5938c45"
y.backward()
x.grad
# + [markdown] colab_type="text" id="v60TtmvumYps"
# ### PyTorch autodiff vs. manual gradients via staged computation
#
# In this example, we will see how a complicated computation can be written as a composition of simpler functions, and how this provides a scalable strategy for computing gradients using the chain rule.
#
# Say we want to write a function to compute the gradient of the *sigmoid function*:
# $$
# \sigma(x) = \frac{1}{1 + e^{-x}}
# $$
# We can write $\sigma(x)$ as a composition of several elementary functions, as $\sigma(x) = s(c(b(a(x))))$, where:
#
# $$
# a(x) = -x
# $$
#
# $$
# b(a) = e^a
# $$
#
# $$
# c(b) = 1 + b
# $$
#
# $$
# s(c) = \frac{1}{c}
# $$
#
# Here, we have "staged" the computation such that it contains several intermediate variables, each of which are basic expressions for which we can easily compute the local gradients.
#
# The computation graph for this expression is shown in the figure below.
#
# 
#
# The input to this function is $x$, and the output is represented by node $s$. We wish compute the gradient of $s$ with respect to $x$, $\frac{\partial s}{\partial x}$. In order to make use of our intermediate computations, we can use the chain rule as follows:
# $$
# \frac{\partial s}{\partial x} = \frac{\partial s}{\partial c} \frac{\partial c}{\partial b} \frac{\partial b}{\partial a} \frac{\partial a}{\partial x}
# $$
#
# <!--
# Given a vector-to-scalar function, $\mathbb{R}^D \to \mathbb{R}$, composed of a set of primitive functions
# $\mathbb{R}^M \to \mathbb{R}^N$ (for various $M$, $N$), the gradient of the composition is given by the product of the gradients of the primitive functions, according to the chain rule. But the chain rule doesn’t prescribe the order in which to multiply the gradients. From the perspective of computational complexity, the order makes all the
# difference.
# -->
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="D5RUMyRsmYpt" outputId="60bb0e8a-e8da-4ea3-dd80-483fa8c50dd0"
def sigmoid(x):
"""Sigmoid function reimplemented for clarity. Use `torch.sigmoid` in real life!"""
y = 1.0 / (1.0 + torch.exp(-x))
return y
def grad_sigmoid_pytorch(x):
x = x.clone().requires_grad_(True)
y = sigmoid(x)
y.backward()
return x.grad
def grad_sigmoid_manual(x):
"""Implements the gradient of the logistic sigmoid function
$\sigma(x) = 1 / (1 + e^{-x})$ using staged computation
"""
# Forward pass, keeping track of intermediate values for use in the
# backward pass
a = -x # -x in denominator
b = np.exp(a) # e^{-x} in denominator
c = 1 + b # 1 + e^{-x} in denominator
s = 1.0 / c # Final result, 1.0 / (1 + e^{-x})
# Backward pass
dsdc = -1.0 / (c ** 2)
dsdb = dsdc * 1
dsda = dsdb * torch.exp(a)
dsdx = dsda * (-1)
return dsdx
def grad_sigmoid_symbolic(x):
# Since d sigmoid(x) / dx = sigmoid(x) * (1 - sigmoid(x))
s = sigmoid(x)
dsdx = s * (1 - s)
return dsdx
input_x = torch.tensor([2.0])
# Compare the results of manual and automatic gradient functions:
print("autograd:", grad_sigmoid_pytorch(input_x).item())
print("manual:", grad_sigmoid_manual(input_x).item())
print("symbolic:", grad_sigmoid_symbolic(input_x).item())
# + [markdown] colab_type="text" id="YTHQYM2ImYq4"
# ### Implementing custom gradients
#
# One thing you can do is define custom gradients for your own functions. There are several reasons you might want to do this, including:
#
# 1. **Speed:** You may know a faster way to compute the gradient for a specific function.
# 2. **Numerical Stability**
# 3. When your code depends on **external library calls**
#
# The `@primitive` decorator wraps a function so that its gradient can be specified manually and its invocation can be recorded.
# -
class MySigmoid(torch.autograd.Function):
"""
We can implement our own custom autograd Functions by subclassing
torch.autograd.Function and implementing the forward and backward passes
which operate on Tensors.
"""
@staticmethod
def forward(ctx, input):
"""
In the forward pass we receive a Tensor containing the input and return
a Tensor containing the output. ctx is a context object that can be used
to stash information for backward computation. You can cache arbitrary
objects for use in the backward pass using the ctx.save_for_backward method.
"""
ans = 1.0 / (1.0 + torch.exp(-input))
ctx.save_for_backward(input, ans)
return ans
@staticmethod
def backward(ctx, grad_output):
"""
In the backward pass we receive a Tensor containing the gradient of the loss
with respect to the output, and we need to compute the gradient of the loss
with respect to the input.
"""
input, ans = ctx.saved_tensors
return grad_output * ans * (1 - ans)
# +
my_sigmoid = MySigmoid.apply
x = input_x.clone().requires_grad_(True)
y = my_sigmoid(x)
y.backward()
x.grad.item()
# + [markdown] colab_type="text" id="I-ePOja_mYrA"
# ## Basic models
#
# The next three sections of the notebook show examples of using pytorch in the context of three problems:
#
# 1. **1-D linear regression**, where we try to fit a model to a function $y = wx + b$
# 2. **Linear regression using a polynomial feature map**, to fit a function of the form $y = w_0 + w_1 x + w_2 x^2 + \dots + w_M x^M$
# 3. **Nonlinear regression using a neural network**
# + [markdown] colab_type="text" id="JBs8UkXfmYrC"
# ### Linear Regression
# + [markdown] colab_type="text" id="cLHB3U0BmYrD"
# #### Review
#
# We are given a set of data points $\{ (x_1, t_1), (x_2, t_2), \dots, (x_N, t_N) \}$, where each point $(x_i, t_i)$ consists of an *input value* $x_i$ and a *target value* $t_i$.
#
# The **model** we use is:
# $$
# y_i = wx_i + b
# $$
#
# We want each predicted value $y_i$ to be close to the ground truth value $t_i$. In linear regression, we use squared error to quantify the disagreement between $y_i$ and $t_i$. The **loss function** for a single example is:
# $$
# \mathcal{L}(y_i,t_i) = \frac{1}{2} (y_i - t_i)^2
# $$
#
# The **cost function** is the loss averaged over all the training examples:
# $$
# \mathcal{E}(w,b) = \frac{1}{N} \sum_{i=1}^N \mathcal{L}(y_i, t_i) = \frac{1}{N} \sum_{i=1}^N \frac{1}{2} \left(wx_i + b - t_i \right)^2
# $$
# + [markdown] colab={} colab_type="code" id="dOjH3W31ITDU"
# #### Data generation
# + [markdown] colab_type="text" id="b-BhEYZOmYrJ"
# We generate a synthetic dataset $\{ (x_i, t_i) \}$ by first taking the $x_i$ to be linearly spaced in the range $[0, 10]$ and generating the corresponding value of $t_i$ using the following equation (where $w = 4$ and $b=10$):
# $$
# t_i = 4 x_i + 10 + \epsilon
# $$
#
# Here, $\epsilon \sim \mathcal{N}(0, 2)$ (that is, $\epsilon$ is drawn from a Gaussian distribution with mean 0 and variance 2). This introduces some random fluctuation in the data, to mimic real data that has an underlying regularity, but for which individual observations are corrupted by random noise.
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" id="mOLDubBYmYrK" outputId="5c85ac5d-bb0e-4ec9-f0cb-74c2d707006f"
# In our synthetic data, we have w = 4 and b = 10
N = 100 # Number of training data points
x = np.linspace(0, 10, N)
t = 4 * x + 10 + npr.normal(0, 2, x.shape[0])
plt.plot(x, t, "r.")
x = torch.from_numpy(x)
t = torch.from_numpy(t)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="DWKVaOrimYrO" outputId="1a1e3f1d-5389-45c7-9da2-91472cdf95a7"
# Initialize random parameters
params = {
"w": torch.randn(1).requires_grad_(True),
"b": torch.randn(1).requires_grad_(True),
}
def cost(params):
y = params["w"] * x + params["b"]
return (1 / N) * torch.sum(0.5 * (y - t) ** 2)
# Find the gradient of the cost function using pytorch
num_epochs = 1000 # Number of epochs of training
alpha = 0.01 # Learning rate
for i in range(num_epochs):
# Evaluate the gradient of the current parameters stored in params
loss = cost(params)
loss.backward()
if i % 100 == 0:
print(f"i: {i:<5d} loss: {loss.item():.4f}")
# Update parameters w and b
with torch.no_grad():
params["w"].data = params["w"] - alpha * params["w"].grad
params["b"].data = params["b"] - alpha * params["b"].grad
params["w"].grad.zero_()
params["b"].grad.zero_()
print(params)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="Q6JKtls8mYrY" outputId="154d136d-5e78-49cd-8cbf-46fa2029ff08"
# Plot the training data again, together with the line defined by y = wx + b
# where w and b are our final learned parameters
plt.plot(x, t, "r.")
plt.plot([0, 10], [params["b"], params["w"] * 10 + params["b"]], "b-")
# + [markdown] colab_type="text" id="UMf3GbOhmYrc"
# ### Linear regression with a feature mapping
# + [markdown] colab_type="text" id="I6sur0MDmYrd"
# In this example we will fit a polynomial using linear regression with a polynomial feature mapping.
# The target function is:
#
# $$
# t = x^4 - 10 x^2 + 10 x + \epsilon
# $$
#
# where $\epsilon \sim \mathcal{N}(0, 4)$.
#
# This is an example of a _generalized linear model_, in which we perform a fixed nonlinear transformation of the inputs $\mathbf{x} = (x_1, x_2, \dots, x_D)$, and the model is still linear in the _parameters_. We can define a set of _feature mappings_ (also called feature functions or basis functions) $\phi$ to implement the fixed transformations.
#
# In this case, we have $x \in \mathbb{R}$, and we define the feature mapping:
# $$
# \mathbf{\phi}(x) = \begin{pmatrix}\phi_1(x) \\ \phi_2(x) \\ \phi_3(x) \\ \phi_4(x) \end{pmatrix} = \begin{pmatrix}1\\x\\x^2\\x^3\end{pmatrix}
# $$
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" id="D-XKvKC4mYre" outputId="47d5582c-ef14-4989-d23d-826b94d3a762"
# Generate synthetic data
N = 100 # Number of data points
x = np.linspace(-3, 3, N) # Generate N values linearly-spaced between -3 and 3
t = x ** 4 - 10 * x ** 2 + 10 * x + npr.normal(0, 4, x.shape[0]) # Generate corresponding targets
plt.plot(x, t, "r.") # Plot data points
t = torch.from_numpy(t).view(-1, 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="oGgROsxlmYrk" outputId="c47b640f-3343-413b-f453-82825c30219a"
M = 4 # Degree of polynomial to fit to the data (this is a hyperparameter)
feature_matrix = torch.tensor(
[[item ** i for i in range(M + 1)] for item in x], dtype=torch.float32
)
params = {
"w": torch.randn(M + 1, 1).requires_grad_(True),
}
print(params["w"].shape)
def cost(params):
y = torch.mm(feature_matrix, params["w"])
return (1.0 / N) * torch.sum(0.5 * (y - t) ** 2)
# Compute the gradient of the cost function using Autograd
num_epochs = 1000
learning_rate = 0.001
# Manually implement gradient descent
for i in range(num_epochs):
loss = cost(params)
loss.backward()
if i % 100 == 0:
print(f"i: {i:<5d} loss: {loss.item():.4f}")
with torch.no_grad():
params["w"].data = params["w"] - learning_rate * params["w"].grad
params["w"].grad.zero_()
# Print the final learned parameters.
print(params["w"])
w = params["w"].detach().cpu().numpy()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="CY3XajWcmYrp" outputId="2d6298d8-472a-472c-c970-60989157078b"
# Plot the original training data again, together with the polynomial we fit
plt.plot(x, t, "r.")
plt.plot(x, np.dot(feature_matrix, w), "b-")
# + [markdown] colab_type="text" id="OWYEe11YmYrs"
# ### Neural net regression
# + [markdown] colab_type="text" id="C7bnv5LemYru"
# In this example we will implement a (nonlinear) regression model using a neural network. To implement and train a neural net using Autograd, you only have to define the forward pass of the network and the loss function you wish to use; you do _not_ need to implement the _backward pass_ of the network. When you take the gradient of the loss function using `grad`, Autograd automatically computes computes the backward pass. It essentially executes the backpropagation algorithm implicitly.
#
# 
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" id="0TuIWDkCmYr5" outputId="7a7ead0c-353a-47ba-e020-568c9b652000"
# Generate synthetic data
x = np.linspace(-5, 5, 1000)
t = x ** 3 - 20 * x + 10 + npr.normal(0, 4, x.shape[0])
plt.plot(x, t, "r.")
x = torch.from_numpy(x).float()
t = torch.from_numpy(t)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="yphlCjMmmYr9" outputId="13396351-9c29-425b-85ac-65692a823a4c"
inputs = x.reshape(x.shape[-1], 1)
params = {
"W1": torch.randn(1, 4).requires_grad_(True),
"b1": torch.randn(4).requires_grad_(True),
"W2": torch.randn(4, 4).requires_grad_(True),
"b2": torch.randn(4).requires_grad_(True),
"W3": torch.randn(4, 1).requires_grad_(True),
"b3": torch.randn(1).requires_grad_(True),
}
# We can define an optimizer which takes care of updating parameters based on their gradient. We can use more complex optimizers like SGD+Momntum or Adam.
optimizer = torch.optim.SGD(params.values(), lr=0.0001, weight_decay=0.0001, momentum=0.9)
# Pytorch also has implementation of wide range of activation functions such as: Tanh, ReLU, LeakyReLU, ...
nonlinearity = torch.nn.ReLU()
def predict(params, inputs):
h1 = nonlinearity(torch.mm(inputs, params["W1"]) + params["b1"])
h2 = nonlinearity(torch.mm(h1, params["W2"]) + params["b2"])
output = torch.mm(h2, params["W3"]) + params["b3"]
return output
def cost(params):
output = predict(params, inputs)
return (1.0 / inputs.shape[0]) * torch.sum(0.5 * (output.reshape(output.shape[0]) - t) ** 2)
print(cost(params))
num_epochs = 10000
for i in range(num_epochs):
# Evaluate the gradient of the current parameters stored in params
loss = cost(params)
if i % 500 == 0:
print(f"i: {i:<5d} loss: {loss.item():.4f}")
optimizer.zero_grad()
loss.backward()
optimizer.step()
with torch.no_grad():
final_y = predict(params, inputs)
plt.plot(x, t, "r.")
plt.plot(x, final_y, "b-")
# -
# ## Neural network models
# + [markdown] colab_type="text" id="QPKpJ3UyanuH"
# Unlike Tensorflow, we can define the graph on the fly. That is why it is more convenient to define a function in Python: we call the function as part of constructing the graph.
#
# Let's now create a simple model for classifiying MNIST digits.
# + [markdown] colab_type="text" id="lkdDq61Ea38-"
# ### MNIST classification
# + [markdown] colab_type="text" id="hiJiOPA5a-lu"
# MNIST is a famous dataset containing hand-written digits. The training set contains 60k and the test set contains 10k images. Pytorch has built-in functions for downloading well-known datasets like MNIST.
# + colab={"base_uri": "https://localhost:8080/", "height": 513} colab_type="code" id="eeeIOIMha5da" outputId="7855870d-92d3-419a-e635-b9172ab46e34"
from torchvision import datasets, transforms
mnist_train = datasets.MNIST("data", train=True, download=True, transform=transforms.ToTensor())
mnist_test = datasets.MNIST("../data", train=False, download=True, transform=transforms.ToTensor())
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="nTdP7vjzb35H" outputId="37165ce6-13ed-43fb-adda-d1b73fbbad65"
print(mnist_train)
print(mnist_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="1dWTw-Rgbh5O" outputId="b19feac0-8842-45a4-cf4e-b10354777392"
i = npr.randint(1, 50000)
example = mnist_train[i]
print("Label: ", example[1])
plt.imshow(example[0].reshape((28, 28)), cmap=plt.cm.gray)
plt.grid(None)
# + [markdown] colab_type="text" id="F5cYaF7PcB7v"
# Pytorch's DataLoader is responsible for creating an iterator over the dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 231} colab_type="code" id="4WDyu63wcV3Z" outputId="2160a2b1-c814-47de-fc8d-88ce272bebf2"
import torchvision
from torch.utils.data import DataLoader
mnist_train = datasets.MNIST("data", train=True, download=True, transform=transforms.ToTensor())
mnist_test = datasets.MNIST("data", train=False, download=True, transform=transforms.ToTensor())
bs = 32
train_dl = DataLoader(mnist_train, batch_size=bs)
test_dl = DataLoader(mnist_test, batch_size=100)
dataiter = iter(train_dl)
images, labels = dataiter.next()
viz = torchvision.utils.make_grid(images, nrow=10, padding=2).numpy()
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(np.transpose(viz, (1, 2, 0)))
# ax.grid(None)
# + [markdown] colab_type="text" id="779GU7DKdVbn"
# Using pytorch built-in function we can easily define any model like multi-layer perceptrons. After training, we just care about the average test accuracy, so let's write a function to compute the accuracy over test set.
# + colab={} colab_type="code" id="2Led8bErdZrw"
def get_test_stat(model, dl, device):
model.eval()
cum_loss, cum_acc = 0.0, 0.0
for i, (xb, yb) in enumerate(dl):
xb = xb.to(device)
yb = yb.to(device)
xb = xb.view(xb.size(0), -1)
y_pred = model(xb)
loss = loss_fn(y_pred, yb)
acc = (torch.max(y_pred.data, 1)[1] == yb).sum() # accuracy(y_pred, yb)
cum_loss += loss.item() * len(yb)
cum_acc += acc.item() * len(yb)
cum_loss /= 10000
cum_acc /= 10000
model.train()
return cum_loss, cum_acc
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="ohxuCpR9eLGt" outputId="bf710e4d-25eb-4595-c66a-8f340947cde2"
dim_x = 784
dim_h = 100
dim_out = 10
model = torch.nn.Sequential(
torch.nn.Linear(dim_x, dim_h),
torch.nn.ReLU(),
torch.nn.Linear(dim_h, dim_out),
)
learning_rate = 1e-2
epochs = 2
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# Using GPUs in PyTorch is pretty straightforward
if torch.cuda.is_available():
print("Using cuda")
use_cuda = True
device = torch.device("cuda")
else:
device = "cpu"
# we need to tell pytorch to move the model to gpu
model.to(device)
loss_fn = torch.nn.CrossEntropyLoss()
model.train()
for epoch in range(epochs):
print(epoch)
for i, (xb, yb) in enumerate(train_dl):
# We also need to transfer the data to the target device
xb = xb.to(device)
yb = yb.to(device)
xb = xb.view(xb.size(0), -1)
# Forward pass
y_pred = model(xb)
loss = loss_fn(y_pred, yb)
# Backward pass
model.zero_grad() # Zero out the previous gradient computation
loss.backward() # Compute the gradient
optimizer.step() # Use the gradient information to make a step
test_loss, test_acc = get_test_stat(model, test_dl, device)
print("Test loss: {} Test acc: {}".format(test_loss, test_acc))
# + [markdown] colab_type="text" id="nSTuUZlRgeUD"
# ### Dynamic network
# + [markdown] colab_type="text" id="Ji_qMWsOgbbj"
# To showcase the power of PyTorch dynamic graphs, we will implement a very strange model: a fully-connected ReLU network that on each forward pass randomly chooses a number between 1 and 4 and has that many hidden layers, reusing the same weights multiple times to compute the innermost hidden layers.
#
# By <NAME> https://github.com/jcjohnson/pytorch-examples/blob/master/nn/dynamic_net.py
# + colab={"base_uri": "https://localhost:8080/", "height": 901} colab_type="code" id="D5dRsCN9gnvU" outputId="53d5f259-33e5-471a-c77c-3a4e311e3202"
import random
class DynamicNet(torch.nn.Module):
def __init__(self, D_in, H, D_out):
"""
In the constructor we construct three nn.Linear instances that we will use
in the forward pass.
"""
super(DynamicNet, self).__init__()
self.input_linear = torch.nn.Linear(D_in, H)
self.middle_linear = torch.nn.Linear(H, H)
self.output_linear = torch.nn.Linear(H, D_out)
def forward(self, x, verbose=False):
"""
For the forward pass of the model, we randomly choose either 0, 1, 2, or 3
and reuse the middle_linear Module that many times to compute hidden layer
representations.
Since each forward pass builds a dynamic computation graph, we can use normal
Python control-flow operators like loops or conditional statements when
defining the forward pass of the model.
Here we also see that it is perfectly safe to reuse the same Module many
times when defining a computational graph. This is a big improvement from Lua
Torch, where each Module could be used only once.
"""
h_relu = self.input_linear(x).clamp(min=0)
n_layers = random.randint(0, 3)
if verbose:
print("The number of layers for this run is", n_layers)
# print(h_relu)
for _ in range(n_layers):
h_relu = self.middle_linear(h_relu).clamp(min=0)
if verbose:
pass
# print(h_relu)
y_pred = self.output_linear(h_relu)
return y_pred
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 64, 1000, 10, 1
# Create random Tensors to hold inputs and outputs, and wrap them in Variables
x = torch.randn(N, D_in)
y = torch.randn(N, D_out).requires_grad_(False)
# Construct our model by instantiating the class defined above
model = DynamicNet(D_in, H, D_out)
# Construct our loss function and an Optimizer. Training this strange model with
# vanilla stochastic gradient descent is tough, so we use momentum
criterion = torch.nn.MSELoss(reduction="sum")
optimizer = torch.optim.SGD(model.parameters(), lr=1e-4, momentum=0.9)
for t in range(50):
# Forward pass: Compute predicted y by passing x to the model
y_pred = model(x)
# Compute and print loss
loss = criterion(y_pred, y)
if t % 10 == 0:
print(t, loss.data.item())
# Zero gradients, perform a backward pass, and update the weights.
optimizer.zero_grad()
loss.backward()
optimizer.step()
# + [markdown] colab_type="text" id="FX-mRC_5iNvs"
# ### CIFAR10 classification
# + [markdown] colab_type="text" id="KVftofphh-ng"
# We will finish with an example on CIFAR10, highlighting the importance of applying transformations to your inputs. Example is lifted from:
#
# https://github.com/uoguelph-mlrg/Cutout/blob/master/train.py
# + colab={} colab_type="code" id="9iAw2caKiW97"
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 120)
self.fc3 = nn.Linear(120, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
# + colab={} colab_type="code" id="A3p2SQViibIc"
def get_data(data_normalize=False, data_augment=False):
train_transform = transforms.Compose([])
test_transform = transforms.Compose([])
if data_augment:
train_transform.transforms.append(transforms.RandomCrop(32, padding=4))
train_transform.transforms.append(transforms.RandomHorizontalFlip())
train_transform.transforms.append(transforms.ToTensor())
test_transform.transforms.append(transforms.ToTensor())
if data_normalize:
normalize = transforms.Normalize(
mean=[x / 255.0 for x in [125.3, 123.0, 113.9]],
std=[x / 255.0 for x in [63.0, 62.1, 66.7]],
)
train_transform.transforms.append(normalize)
test_transform.transforms.append(normalize)
train_dataset = datasets.CIFAR10(
root="data/", train=True, transform=train_transform, download=True
)
test_dataset = datasets.CIFAR10(
root="data/", train=False, transform=test_transform, download=True
)
train_loader = torch.utils.data.DataLoader(
dataset=train_dataset, batch_size=128, shuffle=True, num_workers=2
)
test_loader = torch.utils.data.DataLoader(
dataset=test_dataset, batch_size=128, shuffle=False, num_workers=2
)
return train_loader, test_loader
def test(net, loader):
net.eval() # Change model to 'eval' mode (BN uses moving mean/var).
correct = 0.0
total = 0.0
for images, labels in loader:
with torch.no_grad():
pred = net(images)
pred = torch.max(pred.data, 1)[1]
total += labels.size(0)
correct += (pred == labels).sum().item()
val_acc = correct / total
net.train()
return val_acc
# + colab={} colab_type="code" id="-GR395vyihIU"
def train_model(train_loader, test_loader, epochs=5):
net = Net()
optimizer = torch.optim.SGD(net.parameters(), lr=0.1, momentum=0.9)
criterion = nn.CrossEntropyLoss()
train_accs = []
test_accs = []
net.train()
for epoch in range(epochs):
print(epoch)
xentropy_loss_avg = 0.0
correct = 0.0
total = 0.0
for i, (images, labels) in enumerate(train_loader):
net.zero_grad()
pred = net(images)
xentropy_loss = criterion(pred, labels)
xentropy_loss.backward()
optimizer.step()
xentropy_loss_avg += xentropy_loss.item()
# Calculate running average of accuracy
pred = torch.max(pred.data, 1)[1]
total += labels.size(0)
correct += (pred == labels.data).sum().item()
accuracy = correct / total
test_acc = test(net, test_loader)
print("Test acc: ", test_acc)
train_accs.append(accuracy)
test_accs.append(test_acc)
return train_accs, test_accs
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="MADQb81UijK7" outputId="de18d442-5354-4099-95c1-6ab26a4cba3e"
train_loader, test_loader = get_data(data_augment=False, data_normalize=False)
train_accs, test_accs = train_model(train_loader, test_loader, epochs=3)
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="UnYCP-REikzs" outputId="661de97c-cc2f-4c88-9dc9-49fcf1db96a2"
train_loader, test_loader = get_data(data_augment=False, data_normalize=True)
normalize_train_accs, normalize_test_accs = train_model(train_loader, test_loader, epochs=3)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="aRwFghLzimSJ" outputId="adf16f14-c354-4bfd-a9d7-f97b4d8bc759"
fig, ax = plt.subplots()
epochs = 3
ax.plot(range(epochs), train_accs, c="blue", label="no input normalization")
ax.plot(range(epochs), normalize_train_accs, c="red", label="input normalization")
ax.legend()
ax.set_xlabel("Epochs")
ax.set_ylabel("Accuracy")
ax.set_title("Train Accuracy")
fig.show()
# -
| assets/tutorials/tut02_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''sdc-dmri_venv'': pyenv)'
# metadata:
# interpreter:
# hash: df9a1fb9ddadb6ac6ff99c8fe9ccf963050f17681aeb0dab008e78c4d51aff8c
# name: python3
# ---
# # Introduction to Diffusion MRI data
#
# _*Much of the material in both the current notebook and the upcoming notebooks are adapted from the lessons on [neuroimaging analysis with Python](https://github.com/datacarpentry/python-neuroimaging-lesson)._
#
# ## Diffusion Weighted Imaging (DWI)
#
# Diffusion imaging probes the random, microscopic motion of water protons by employing MRI sequences which are sensitive to the geometry and environmental organization surrounding the water protons. This is a popular technique for studying the white matter of the brain. The diffusion within biological structures, such as the brain, are often restricted due to barriers (eg. cell membranes), resulting in a preferred direction of diffusion (anisotropy). A typical diffusion MRI scan will acquire multiple volumes that are sensitive to a particular diffusion direction and result in diffusion-weighted images (DWI). Diffusion that exhibits directionality in the same direction result in an attenuated signal. With further processing (to be discussed later in the lesson), the acquired images can provide measurements which are related to the microscopic changes and estimate white matter trajectories. Images with no diffusion weighting are also acquired as part of the acquisition protocol.
#
#  \
# Diffusion along X, Y, and Z directions
# ## b-values & b-vectors
#
# In addition to the acquired diffusion images, two files are collected as part of the diffusion dataset. These files correspond to the gradient amplitude (b-values) and directions (b-vectors) of the diffusion measurement and are named with the extensiosn `.bval` and `.bvec` respectively. The b-value is the diffusion-sensitizing factor, and reflects the timing & strength of the gradients used to acquire the diffusion-weighted images. The b-vector corresponds to the direction of the diffusion sensitivity. Together these two files define the diffusion MRI measurement as a set of gradient directions and corresponding amplitudes.
# ## Dataset
#
# For the rest of this tutorial, we will make use of a subset of publicly available dataset, ds000030, from [openneuro.org](https://openneuro.org/datasets/ds000030) The dataset is structured according to the Brain Imaging Data Structure ([BIDS](https://bids-specification.readthedocs.io/en/etable/)).
# Below is a tree diagram showing the folder structure of a single MR subject and session within ds000221. This was obtained by using the bash command `tree`.
#
# `!tree ../../data/ds000221`
#
# ```
# ../../data/ds000221
# ├── .bidsignore
# ├── CHANGES
# ├── dataset_description.json
# ├── participants.tsv
# ├── README
# ├── derivatives/
# ├── sub-010001/
# └── sub-010002/
# ├── ses-01/
# │ ├── anat
# │ │ ├── sub-010002_ses-01_acq-lowres_FLAIR.json
# │ │ ├── sub-010002_ses-01_acq-lowres_FLAIR.nii.gz
# │ │ ├── sub-010002_ses-01_acq-mp2rage_defacemask.nii.gz
# │ │ ├── sub-010002_ses-01_acq-mp2rage_T1map.nii.gz
# │ │ ├── sub-010002_ses-01_acq-mp2rage_T1w.nii.gz
# │ │ ├── sub-010002_ses-01_inv-1_mp2rage.json
# │ │ ├── sub-010002_ses-01_inv-1_mp2rage.nii.gz
# │ │ ├── sub-010002_ses-01_inv-2_mp2rage.json
# │ │ ├── sub-010002_ses-01_inv-2_mp2rage.nii.gz
# │ │ ├── sub-010002_ses-01_T2w.json
# │ │ └── sub-010002_ses-01_T2w.nii.gz
# │ ├── dwi
# │ │ ├── sub-010002_ses-01_dwi.bval
# │ │ │── sub-010002_ses-01_dwi.bvec
# │ │ │── sub-010002_ses-01_dwi.json
# │ │ └── sub-010002_ses-01_dwi.nii.gz
# │ ├── fmap
# │ │ ├── sub-010002_ses-01_acq-GEfmap_run-01_magnitude1.json
# │ │ ├── sub-010002_ses-01_acq-GEfmap_run-01_magnitude1.nii.gz
# │ │ ├── sub-010002_ses-01_acq-GEfmap_run-01_magnitude2.json
# │ │ ├── sub-010002_ses-01_acq-GEfmap_run-01_magnitude2.nii.gz
# │ │ ├── sub-010002_ses-01_acq-GEfmap_run-01_phasediff.json
# │ │ ├── sub-010002_ses-01_acq-GEfmap_run-01_phasediff.nii.gz
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-AP_epi.json
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-AP_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-PA_epi.json
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-PA_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-sefmapBOLDpre_dir-AP_epi.json
# │ │ ├── sub-010002_ses-01_acq-sefmapBOLDpre_dir-AP_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-sefmapBOLDpre_dir-PA_epi.json
# │ │ ├── sub-010002_ses-01_acq-sefmapBOLDpre_dir-PA_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-AP_epi.json
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-AP_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-PA_epi.json
# │ │ ├── sub-010002_ses-01_acq-SEfmapBOLDpost_dir-PA_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-SEfmapDWI_dir-AP_epi.json
# │ │ ├── sub-010002_ses-01_acq-SEfmapDWI_dir-AP_epi.nii.gz
# │ │ ├── sub-010002_ses-01_acq-SEfmapDWI_dir-PA_epi.json
# │ │ └── sub-010002_ses-01_acq-SEfmapDWI_dir-PA_epi.nii.gz
# │ └── fmap
# │ │ ├── sub-010002_ses-01_task-rest_acq-AP_run-01_bold.json
# │ │ └── sub-010002_ses-01_task-rest_acq-AP_run-01_bold.nii.gz
# └── ses-02/
# ```
# ## Querying a BIDS Dataset
#
# [`pybids`](https://bids-standard.github.io/pybids/) is a Python API for querying, summarizing and manipulating the BIDS folder structure. We will make use of `pybids` to query the necessary files.
#
# Lets first pull the metadata from its associated JSON file using the `get_metadata()` function for the first run.
# ?BIDSLayout
# +
from bids.layout import BIDSLayout
layout = BIDSLayout("../../../data/ds000221", validate=False)
# + jupyter={"outputs_hidden": true}
dwi = layout.get(subject='010006', suffix='dwi', extension='nii.gz', return_type='file')[0]
layout.get_metadata(dwi)
# -
# ## [`dipy`](http://dipy.org)
#
# For this lesson, we will use the `Dipy` (Diffusion Imaging in Python) package for processing and analysing diffusion MRI.
#
# ### Why `dipy`?
#
# - Fully free and open source
# - Implemented in Python. Easy to understand, and easy to use.
# - Implementations of many state-of-the art algorithms
# - High performance. Many algorithms implemented in [`cython`](http://cython.org/)
#
# ### Installing `dipy`
#
# The easiest way to install `Dipy` is to use `pip`! Additionally, `Dipy` makes use of the FURY library for visualization. We will also install this using `pip`!
#
# We can install it by entering the following in a terminal `pip install dipy`. We will do so using Jupyter Magic in the following cell!
# ! pip install dipy
# ! pip install fury
# ### Defining a measurement: `GradientTable`
#
# `Dipy` has a built-in function that allows us to read in `bval` and `bvec` files named `read_bvals_bvecs` under the `dipy.io.gradients` module. Let's first grab the path to our gradient directions and amplitude files and load them into memory.
# + jupyter={"outputs_hidden": true}
bvec = layout.get_bvec(dwi)
bval = layout.get_bval(dwi)
# +
from dipy.io.gradients import read_bvals_bvecs
from dipy.core.gradients import gradient_table
gt_bvals, gt_bvecs = read_bvals_bvecs(bval, bvec)
# + [markdown] jupyter={"outputs_hidden": true}
# There is a also simple `GradientTable` object implemented in the `dipy.core.gradients` module. The input to the `GradientTable` should be our the values for our gradient directions and amplitudes we just read.
# -
gtab = gradient_table(gt_bvals, gt_bvecs)
# We will need this gradient table later on to process our data and generate diffusion tensor images (DTI)!
#
# There is also a built in function for gradient tables, `b0s_mask` that can be used to separate difussion weighted measurements from non-diffusion weighted measurements (b=0s/mm^2). Try to extract the vector corresponding to diffusion weighted measurements in the following cell!
gtab.bvecs[~gtab.b0s_mask]
# It is also important to know where our diffusion weighting free measurements are as we need them for registration in our preprocessing, (our next notebook). The `b0s_mask` shows that this is the first volume of our dataset.
gtab.b0s_mask
# In the next few notebooks, we will talk more about preprocessing the diffusion weighted images and reconstructing the Tensor model
| code/introduction/solutions/introduction_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load the dataset.
with open("./datasets/football_key.tsv") as f:
# Each line is of form: <country_id> <country_name>
def fmt(line):
return (int(line[0])-1, line[1].strip('"'))
data_key = [fmt(line.strip().split()) for line in f if line[0] != '*']
with open("./datasets/football_pairs.tsv") as f:
# Each line is of form: <country_a_id> <country_b_id> <number_of_players>
def fmt(pair):
return (int(pair[0])-1, int(pair[1])-1, 1)
data_pairs = [fmt(line.strip().split()) for line in f if line[0] != '*']
# # Turn into useful format
#
# Edit the `neighbours[]` below and `similarity` func below that to create a new metric.
neighbours = [set() for _ in range(len(data_key))]
for p in data_pairs:
neighbours[p[0]].add(p[1])
neighbours[p[1]].add(p[0])
def similarity_CN(x, y, ignore_set=None):
# Common neighbours
# x, y are indices to neighbours[]
if ignore_set is None:
ignore_set = set()
return len((neighbours[x] & neighbours[y]) - set(t[1] for t in ignore_set))
# # Compute similarity matrix.
def compute_similarities(ignore_set=None):
# S_CN[x][y] contains the similarity of nodes x and y using the Common Neighbours (CN) metric.
S_CN = [[0 for _ in range(len(data_key))] for _ in range(len(data_key))]
for i in range(len(data_key)-1):
for j in range(0, len(data_key)):
S_CN[i][j] = similarity_CN(i, j, ignore_set=ignore_set)
return S_CN
# A quick eyeball check of a subset of the data.
S_CN = compute_similarities()
num_to_print = len(data_key)//2
print(' '*4 + ' '.join(d[1] for d in data_key[:num_to_print]))
print('\n'.join(data_key[i][1] + ' ' + ','.join('{:>3}'.format(c) for c in S_CN[i][:num_to_print]) for i in range(num_to_print)))
# # Create test sets.
#
# Split the list of links into 10 random partitions, as the paper does, to get comparable measurements. Also create a set of all links which are not in the dataset.
# +
def chunks(l, n):
"""Yield successive n-sized chunks from l."""
for it in range(0, len(l), n):
yield l[it:it + n]
e = []
predict = []
for i in range(len(data_key)):
for j in range(i+1, len(data_key)):
if i in neighbours[j]:
e.append((i, j))
else:
predict.append((i, j))
# e now contains all link pairs
# predict contains all non-existing links from the original data
# each pair is a tuple (a, b), where a < b
# We now randomly shuffle this list
import random
random.shuffle(e)
print('len(e)', len(e))
print('len(e)//10 =', len(e)//10)
# Create e_prime, a list of 10 partitions
e_prime = list(chunks(e, len(e)//10 + 1))
# The following is a quick eyeball test to make sure the partitions look ok.
print('10 subsets:')
for i in range(len(e_prime)):
entry = e_prime[i]
print(entry)
# +
aucs = []
n1s = []
n2s = []
n3s = []
ns = []
# Column headings.
print('\t\tn1 \tn2 \tn3 \tAUC')
# Iterate across the 10 folds.
for i in range(10):
test = e_prime[i]
S_CN = compute_similarities(ignore_set=None)
n1 = 0 # missing_link > nonexistant_link
n2 = 0 # missing_link = nonexistant_link
n3 = 0 # missing_link < nonexistant_link
n = 0 # total link comparisons
for missing_link in test:
a_score = S_CN[missing_link[0]][missing_link[1]]
for nonexistant_link in predict:
b_score = S_CN[nonexistant_link[0]][nonexistant_link[1]]
if abs(a_score-b_score) < 0.0005:
n2 += 1
elif a_score > b_score:
n1 += 1
else:
n3 += 1
n += 1
auc = (n1 + 0.5*n2)/(n)
aucs.append(auc)
n1s.append(n1)
n2s.append(n2)
n3s.append(n3)
ns.append(n)
print('Fold {:<2}:\t{:<5}\t{:<5}\t{:<5}\t{:<.6f}'.format(i+1, n1, n2, n3, auc))
def avg(seq):
return sum(seq)/len(seq)
print('Average:\t{:<5}\t{:<5}\t{:<5}\t{:<.6f}'.format(int(round(avg(n1s))), int(round(avg(n2s))), int(round(avg(n3s))), avg(aucs)))
# -
| research/common_neighbours_football.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sugatoray/stackoverflow/blob/master/src/answers/Q_69321580/Q_69321580.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="FgCpNh4TLjfI"
# # Stackoverflow Question: [69321580][#Q]
#
# [#Q]: https://stackoverflow.com/questions/69321580/jinja-table-when-column-value-then-add-div-to-specific-column/69321665#69321665
# + id="BgcozP01A3ck"
src/notebooks/jinja2_python_notebook.ipynb
# + id="4ockw8cDPaOp"
import string
from typing import List
import numpy as np
import pandas as pd
from jinja2 import Template
def make_row(letter:str, ncols: int=5) -> List[str]:
return [ f'{letter}{str(i).zfill(len(str(ncols)))}' for i in range(ncols) ]
def make_data(nrows: int=5, ncols: int=6) -> pd.DataFrame:
data = []
# nrows = 5
# ncols = 6
columns = list(string.ascii_uppercase[0:ncols])
rowlabels = list(string.ascii_uppercase[0:nrows])
for letter in rowlabels:
data.append(make_row(letter=letter, ncols=ncols-1))
data = np.array(data)
data = np.hstack([data, np.random.randint(0,2,nrows).reshape((nrows,1))])
df = pd.DataFrame(data, columns=columns)
return df
# + colab={"base_uri": "https://localhost:8080/"} id="lIYdn1woOS1n" outputId="87525d9d-ee5a-441a-fae2-9cbe3b9011c0"
# %%writefile template.html
<table>
<thead>
<tr>
{% for col in column_names %}
<th>
{{col}}
</th>
{% endfor %}
</tr>
</thead>
<tbody>
{% for row in row_data %}
<tr>
{% for col, row_ in zip(column_names, row) %}
{% if loop.index == 1 %}
<td>
{{ '<div class="circle" style="float: left;">LB</div>' if row[target_col_num] == 1 else '' }} {{ row_ }}
</td>
{% else %}
<td>{{ row_ }}</td>
{% endif %}
{% endfor %}
</tr>
{% endfor %}
</tbody>
</table>
# + id="_LOTXlqVaz4j"
df = make_data(nrows=5, ncols=6)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="cFPDYSA1R0g4" outputId="813fd4b7-97fc-4ce6-fac8-51e9ad6b5914"
df
# + colab={"base_uri": "https://localhost:8080/"} id="XjNmUIx0SeeS" outputId="17dc1911-889e-46d3-cab7-fcf9f9810974"
df.columns
# + id="CiER3zyKW0mm"
with open('template.html', 'r') as f:
template = f.read()
# + id="owgVc_CaBFI7"
t = Template(template)
output = t.render(
column_names=df.columns,
row_data=df.loc[0].tolist(),
target_col_num=5,
zip=zip,
)
# + id="M2xJGFGmUN8n"
t = Template("This is {{ fname }}.")
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="k1gSYhu8V6L6" outputId="1feb19ca-31f0-48ec-9d37-23ada431cca8"
t.render(fname="Joe")
# + id="GKflAGmdWeX2"
from IPython.display import HTML
# + colab={"base_uri": "https://localhost:8080/", "height": 169} id="eDQ_rW9PbPt8" outputId="338aade6-25d1-4a4d-bc19-aa9506e65712"
HTML(output)
# + id="waGX6K9ZbXTR"
| src/answers/Q_69321580/Q_69321580.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Bin data with logarithmically spaced bins notebook
# Copyright 2020 <NAME>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# +
import matplotlib
matplotlib.use('module://ipympl.backend_nbagg')
from matplotlib import pyplot as plt
import numpy as np
import os
import sys
# Load packages
software_path = os.path.join('..')
sys.path.append(software_path)
from functions.binning import calculate_bin_means
# +
# Simulate noisy data: a line of 500 data points with noise between x = [1:1000] (f(x) = x/10)
start_x = 1
stop_x = 1000
number_of_datapoints = 500
number_of_bins = 10
noise_level = 10
x = np.linspace(start_x, stop_x, number_of_datapoints)
y = x/10 + np.random.randn(number_of_datapoints) * noise_level
# Calculate binned data with log spaced bins
bin_edges = np.logspace(np.log10(start_x), np.log10(stop_x), number_of_bins+1)
print('Calculate binned data with {} log spaced bins from x = {} to {}.'.format(number_of_bins, start_x, stop_x))
# Concatenate x and y to one 2 dimensional array
data = np.c_[x, y]
# Bin data along x (sortcolumn=0)
edges, centers, width, bin_means, bin_stds, bin_Ns = calculate_bin_means(data, edges=bin_edges, sortcolumn=0)
x_binned = bin_means[:,0]
y_binned = bin_means[:,1]
x_errs = bin_stds[:,1] / np.sqrt(bin_Ns)
y_errs = bin_stds[:,1] / np.sqrt(bin_Ns)
# Plot the data and binned data in a semilog plot
fig, ax = plt.subplots()
ax.scatter(x, y, alpha=0.15, color='black', marker='.', lw=0, label="raw data")
ax.errorbar(centers, y_binned, yerr=y_errs, fmt='o', label="bin centers")
ax.errorbar(x_binned, y_binned, xerr=x_errs, yerr=y_errs, fmt='.', label="bin means")
ax.set_xscale('log')
ax.legend()
fig.show()
| functions/notebooks/Bin data with logarithmically spaced bins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#TEST 01
#trying to write to csv file
#training the above code
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import csv
data=pd.read_csv("dr_features.csv")
data_new=pd.read_csv("dr_features.csv")
predictions=data_new['count']
#data_new
threshold = 30
data_new['pred_value'] = predictions.apply(lambda x: 1 if x > threshold else 0)
features_raw = data_new[[ "count", "area"]]
predict_class = data_new['pred_value']
#writing to csv file
count=0
with open('dr_features.csv','r') as csvinput:
with open('dr_features_output.csv', 'w') as csvoutput:
writer = csv.writer(csvoutput, lineterminator='\n')
reader = csv.reader(csvinput)
all = []
row = next(reader)
row.append('pred_values')
all.append(row)
for row in reader:
row.append(predict_class[count])
count = count + 1
all.append(row)
writer.writerows(all)
| test061_training00.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 2. Solving a 1D diffusion equation
# +
# Document Author: Dr. <NAME>
# Author email: <EMAIL>
# License: MIT
# This tutorial is applicable for NAnPack version 1.0.0-alpha4
# -
# ### I. Background
#
# The objective of this tutorial is to present the step-by-step solution of a 1D diffusion equation using NAnPack such that users can follow the instructions to learn using this package. The numerical solution is obtained using the Forward Time Central Spacing (FTCS) method. The detailed description of the FTCS method is presented in Section IV of this tutorial.
#
# ### II. Case Description
#
# We will be solving a classical probkem of a suddenly accelerated plate in fluid mechanicas which has the known exact solution. In this problem, the fluid is
# bounded between two parallel plates. The upper plate remains stationary and the lower plate is suddenly accelerated in *y*-direction at velocity $U_o$. It is
# required to find the velocity profile between the plates for the given initial and boundary conditions.
#
# (For the sake of simplicity in setting up numerical variables, let's assume that the *x*-axis is pointed in the upward direction and *y*-axis is pointed along the horizontal direction as shown in the schematic below:
# 
# **Initial conditions**
# $$u(t=0.0, 0.0<x\leq H) = 0.0 \;m/s$$
# $$u(t=0.0, x=0.0) = 40.0 \;m/s$$
#
# **Boundary conditions**
# $$u(t\geq0.0, x=0.0) = 40.0 \;m/s$$
# $$u(t\geq0.0, x=H) = 0.0 \;m/s$$
#
# Viscosity of fluid, $\;\;\nu = 2.17*10^{-4} \;m^2/s$
# Distance between plates, $\;\;H = 0.04 \;m$
# Grid step size, $\;\;dx = 0.001 \;m$
# Simulation time, $\;\;T = 1.08 \;sec$
#
# Specify the required simulation inputs based on our setup in the configuration file provided with this package. You may choose to save the configuration file with any other filename. I have saved the configuration file in the "input" folder of my project directory such that the relative path is `./input/config.ini`.
# ### III. Governing Equation
#
# The governing equation for the given application is the simplified for the the Navies-Stokes equation which is given as:
#
# $$\frac{\partial u} {\partial t} = \nu\frac{\partial^2 u}{\partial x^2}$$
#
# This is the diffusion equation model and is classified as the parabolic PDE.
#
# ### IV. FTCS method
#
# The forward time central spacing approximation equation in 1D is presented here. This is a time explicit method which means that one unknown is calculated using the known neighbouring values from the previous time step. Here *i* represents grid point location, *n*+1 is the future time step, and *n* is the current time step.
#
# $$u_{i}^{n+1} = u_{i}^{n} + \frac{\nu\Delta t}{(\Delta x)^2}(u_{i+1}^{n} - 2u_{i}^{n} + u_{i-1}^{n})$$
#
# The order of this approximation is $[(\Delta t), (\Delta x)^2]$
#
# The diffusion number is given as $d_{x} = \nu\frac{\Delta t}{(\Delta x)^2}$ and for one-dimensional applications the stability criteria is $d_{x}\leq\frac{1}{2}$
#
# The solution presented here is obtained using a diffusion number = 0.5 (CFL = 0.5 in configuration file). Time step size will be computed using the expression of diffusion number. Beginners are encouraged to try diffusion numbers greater than 0.5 as an exercise after running this script.
#
# Users are encouraged to read my blogs on numerical methods - [link here](https://www.linkedin.com/in/vishalsharmaofficial/detail/recent-activity/posts/).
#
# ### V. Script Development
#
# *Please note that this code script is provided in file `./examples/tutorial-02-diffusion-1D-solvers-FTCS.py`.*
#
# As per the Python established coding guidelines [PEP 8](https://www.python.org/dev/peps/pep-0008/#imports), all package imports must be done at the top part of the script in the following sequence --
# 1. import standard library
# 2. import third party modules
# 3. import local application/library specific
#
# Accordingly, in our code we will importing the following required modules (in alphabetical order). If you are using Jupyter notebook, hit `Shift + Enter` on each cell after typing the code.
import matplotlib.pyplot as plt
from nanpack.benchmark import ParallelPlateFlow
import nanpack.preprocess as pre
from nanpack.grid import RectangularGrid
from nanpack.parabolicsolvers import FTCS
import nanpack.postprocess as post
# As the first step in simulation, we have to tell our script to read the inputs and assign those inputs to the variables/objects that we will use in our entire code. For this purpose, there is a class `RunConfig` in `nanpack.preprocess` module. We will call this class and assign an object (instance) to it so that we can use its member variables. The `RunConfig` class is written in such a manner that its methods get executed as soon as it's instance is created. The users must provide the configuration file path as a parameter to `RunConfig` class.
FileName = "D:/MyProjects/projectroot/nanpack/input/config.ini" # specify the correct file path
cfg = pre.RunConfig(FileName) # cfg is an instance of RunConfig class which can be used to access class variables. You may choose any variable in place of cfg.
# You will obtain several configuration messages on your output screen so that you can verify that your inputs are correct and that the configuration is successfully completed. Next step is the assignment of initial conditions and the boundary conditions. For assigning boundary conditions, I have created a function `BC()` which we will be calling in the next cell. I have included this function at the bottom of this tutorial for your reference. It is to be noted that U is the dependent variable that was initialized when we executed the configuration, and thus we will be using `cfg.U` to access the initialized U. In a similar manner, all the inputs provided in the configuration file can be obtained by using configuration class object `cfg.` as the prefix to the variable names. Users are allowed to use any object of their choice.
#
# *If you are using Jupyter Notebook, the function BC must be executed before referencing to it, otherwise, you will get an error. Jump to the bottom of this notebook where you see code cell # 1 containing the `BC()` function*
# +
# Assign initial conditions
cfg.U[0] = 40.0
cfg.U[1:] = 0.0
# Assign boundary conditions
U = BC(cfg.U)
# -
# Next, we will be calculating location of all grid points within the domain using the function `RectangularGrid()` and save values into X. We will also require to calculate diffusion number in X direction. In nanpack, the program treats the diffusion number = CFL for 1D applications that we entered in the configuration file, and therefore this step may be skipped, however, it is not the same in two-dimensional applications and therefore to stay consistent and to avoid confusion we will be using the function `DiffusionNumbers()` to compute the term `diffX`.
X, _ = RectangularGrid(cfg.dX, cfg.iMax)
diffX,_ = pre.DiffusionNumbers(cfg.Dimension, cfg.diff, cfg.dT, cfg.dX)
# Next, we will initialize some local variables before start the time stepping:
Error = 1.0 # variable to keep track of error
n = 0 # variable to advance in time
# Start time loop using while loop such that if one of the condition returns False, the time stepping will be stopped. For explanation of each line, see the comments. Please note the identation of the codes within the while loop. Take extra care with indentation as Python is very particular about it.
while n <= cfg.nMax and Error > cfg.ConvCrit: # start loop
Error = 0.0 # reset error to 0.0 at the beginning of each step
n += 1 # advance the value of n at each step
Uold = U.copy() # store solution at time level, n
U = FTCS(Uold, diffX) # solve for U using FTCS method at time level n+1
Error = post.AbsoluteError(U, Uold) # calculate errors
U = BC(U) # Update BC
post.MonitorConvergence(cfg, n, Error) # Use this function to monitor convergence
post.WriteSolutionToFile(U, n, cfg.nWrite, cfg.nMax,\
cfg.OutFileName, cfg.dX) # Write output to file
post.WriteConvHistToFile(cfg, n, Error) # Write convergence log to history file
# In the above convergence monitor, it is worth noting that the solution error is gradually moving towards zero which is what we need to confirm stability in the solution. If the solution becomes unstable, the errors will rise, probably upto the point where your code will crash. As you know that the solution obtained is a time-dependent solution and therefore, we didn't allow the code to run until the convergence is observed. If a steady-state solution is desired, change the STATE key in the configuration file equals to "STEADY" and specify a much larger value of nMax key, say nMax = 5000. This is left as an exercise for the users to obtain a stead-state solution. Also, try running the solution with the larger grid step size, $\Delta x$ or a larger time step size, $\Delta t$.
#
# After the time stepping is completed, save the final results to the output files.
# Write output to file
post.WriteSolutionToFile(U, n, cfg.nWrite, cfg.nMax,
cfg.OutFileName, cfg.dX)
# Write convergence history log to a file
post.WriteConvHistToFile(cfg, n, Error)
# Verify that the files are saved in the target directory.
# Now let us obtain analytical solution of this flow that will help us in validating our codes.
# Obtain analytical solution
Uana = ParallelPlateFlow(40.0, X, cfg.diff, cfg.totTime, 20)
# Next, we will validate our results by plotting the results using the matplotlib package that we have imported above. Type the following lines of codes:
plt.rc("font", family="serif", size=8) # Assign fonts in the plot
fig, ax = plt.subplots(dpi=150) # Create axis for plotting
plt.plot(U, X, ">-.b", linewidth=0.5, label="FTCS",\
markersize=5, markevery=5) # Plot data with required labels and markers, customize the plot however you may like
plt.plot(Uana, X, "o:r", linewidth=0.5, label="Analytical",\
markersize=5, markevery=5) # Plot analytical solution on the same plot
plt.xlabel('Velocity (m/s)') # X-axis labelling
plt.ylabel('Plate distance (m)') # Y-axis labelling
plt.title(f"Velocity profile\nat t={cfg.totTime} sec", fontsize=8) # Plot title
plt.legend()
plt.show() # Show plot- this command is very important
# Function for the boundary conditions.
def BC(U):
"""Return the dependent variable with the updated values at the boundaries."""
U[0] = 40.0
U[-1] = 0.0
return U
# Congratulations, you have completed the first coding tutoria using nanpack package and verified that your codes produced correct results. If you solve some other similar diffusion-1D model example, share it with the nanpack community. I will be excited to see your projects.
| docs/build/html/examples/tutorial-02-diffusion-1D-solvers-FTCS.ipynb |