code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing Data
# In this step of the project we cleaned the data. We removed punctuations, special characters, and lowercased all words first, then tokenized the words for removing of stop words. Next the words were stemmed using lemmatization. Next the words that were not longer than 4 were removed. The last step was to split the data into train, validation, and test sets.
# ### Import pyspark using Docker
import pyspark
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
# ### Start Spark Session
spark = SparkSession.builder.appName('cleaning').getOrCreate()
# ### Load Data
df = spark.read.json('Movies_and_TV.json.gz')
### View Data
df = df.select('reviewText', 'verified')
df.show(5)
### droping na values
df = df.na.drop()
df.count()
# ### Remove punctuations & special characters & lowercase words
# In this step punctuations, special characters, and numbers were removed. Also, all words were now lowercased.
### Clean Function
def clean_text(c):
c = lower(c)
c = regexp_replace(c, "^rt ", "")
c = regexp_replace(c, "[\=.]"," ")
c = regexp_replace(c, "[^a-zA-Z0-9\\s]", "")
c = regexp_replace(c, " ", " ")
c = regexp_replace(c, " ", " ")
c = regexp_replace(c, '\d+', "")
return(c)
### View Clean
df = df.withColumn("clean_text",clean_text(col('reviewText')))
df.show()
# ### Tokenize
from pyspark.ml.feature import Tokenizer
### tokenize words
tokenizer = Tokenizer(inputCol="clean_text", outputCol="token_text")
token = tokenizer.transform(df).select('verified', 'token_text')
token.show()
# ### Remove Stop words
# Stop words were removed using the pyspark ml feature “StopWordsRemover”.
from pyspark.ml.feature import StopWordsRemover
### remove stops words
remover = StopWordsRemover(inputCol='token_text', outputCol='swr_text')
swr = remover.transform(token).select('verified','swr_text')
swr.show(5)
# ### Lemmatization
# In the lemmatization step the python library “nltk” was used. A function was created to stem the words down using the “WordNetLemmatizer” from the “nltk” library.
### import nltk
import nltk
nltk.download('wordnet')
# +
### Create Function
from nltk.stem import WordNetLemmatizer
# Instantiate stemmer object
lemmer = WordNetLemmatizer()
def lem(in_vec):
out_vec = []
for t in in_vec:
t_stem = lemmer.lemmatize(t)
if len(t_stem) > 2:
out_vec.append(t_stem)
return(out_vec)
# +
### Use function
from pyspark.sql.types import *
lemmer_udf = udf(lambda x: lem(x), ArrayType(StringType()))
lem_text = swr.withColumn("lem", lemmer_udf(col("swr_text"))).select('verified', 'lem')
lem_text.show()
# -
# ### Remove Short words
# In the last cleaning step, we removed words that were not longer than four characters.
### Removing words
filter_length_udf = udf(lambda row: " ".join([x for x in row if len(x) >= 4]))
df2= lem_text.withColumn('Text', filter_length_udf(col('lem'))).select('Text', 'verified')
df2.show()
# ### Add a index to split data
# An index was added to the dataset because when we used the random split function in pyspark the text column would turn into null values. To find another way to split the data we created an index.
df2 = df2.withColumn('index', monotonically_increasing_id())
df2.show(10)
# ### Split Data to Train, Val, Test
# We split the data on a 80% train and 20% validation split with 1 row for the test set.
### 80/20 split
print('Train:',8757545 * .80 )
print('Val:',8757545 * .20 )
### Train set
train = df2.filter(df2.index <= 7006036)
train.count()
### Validation set
val = df2.filter((df2.index > 7006036) & (df2.index < 8757544))
val.count()
### Test example
test = df2.filter(df2.index == 8757544)
test.count()
# ### Export Data
# The last step was to export the data for the next three steps in the project.
### Export validation set
val.write.option("header", "true").csv('clean_val')
### Export train set
train.write.option("header", "true").csv('clean_train')
### Export test set
test.write.option("header", "true").csv('clean_test')
| Notebooks/clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# # Elasticsearch - Connect to server
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Elasticsearch/Elasticsearch_Connect_to_server.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# **Tags:** #elasticsearch #elastic #search #snippet #operations
# + [markdown] papermill={} tags=["naas", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# **Author:** [<NAME>](https://www.linkedin.com/in/ebinpaulose/)
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### 1. Prerequisites
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# - python3
# - ubuntu 18.04
# - java1.8
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### 2. Elasticsearch on local machine
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Install Linux packages
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ```sh
# $ sudo apt update
# $ sudo apt-get install apt-transport-http
# $ wget -qO - https://artifacts.elastic.co/GPG-KEY-elasticsearch | sudo apt-key add -
# $ sudo add-apt-repository "deb https://artifacts.elastic.co/packages/7.x/apt stable main"
# $ sudo apt update
# $ sudo apt install elasticsearch
# ```
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Check status of Elasticsearch server (Local)
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ```sh
# $ sudo /etc/init.d/elasticsearch status
# ```
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Start Elasticsearch server (Local)
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ```sh
# $ sudo /etc/init.d/elasticsearch start
# ```
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ##### Note : Install Java 1.8 and set Java environment variables path
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### 3. Elasticsearch on cloud
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Step 1: Login to https://www.elastic.co/ and create a deployment
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Step 2: On successful deployment get credentials
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Step 3: Create Elasticsearch credentials JSON
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# #### Credentials Json format
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ```sh
# {
# "endpoint": "< Elasticsearch endpoint from elasticsearch cloud >",
# "port": "< Port number as mentioned in elasticsearch cloud >",
# "user": "< User as mentioned in elasticsearch cloud >",
# "password": "< <PASSWORD> >",
# "protocol": "https"
# }
# ```
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### 4. Python connector for Elasticsearch
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ## Input
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### Import library
# + papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
from elasticsearchconnector import ElasticsearchConnector
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ## Model
# + papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
instance = ElasticsearchConnector("sample_credentials.json")
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### Send data to Elasticsearch server
# + papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# parameters = {'index':'< Name of the index >','type':' < Document name > '}
parameters = {'index':'students','type':'engineering'}
# data = { < Key value pairs > }
data = {"Name": "Poul", "Age":20, "address": "New york"}
result = instance.save_data(parameters,data)
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### Search data from Elasticsearch server
# + papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# parameters = {'index':'< Name of the index >','type':' < Document name > '}
parameters = {'index':'students','type':'engineering'}
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ## Output
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### Single search
# + papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# Single search
q1 = {"query": {"match": {'Name':'Poul'}}}
result = instance.search_data(parameters,[q1],search_type='search')
print(result)
# + [markdown] papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# ### Multiple search
# + papermill={} tags=["awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb", "awesome-notebooks/Elasticsearch/Elasticsearch_Connect_to_server.ipynb"]
# Multiple search
q1 = {"query": {"match": {'Name':'Poul'}}}
q2 = {"query": {"match": {'Age':27}}}
result = instance.search_data(parameters,[q1,q2],search_type='msearch')
print(result)
| Elasticsearch/Elasticsearch_Connect_to_server.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="-2aCvc1d9ezf"
# # Indexing
# To get values from a list, you use **indexing**.
#
# To index a list, you use square brackets (`[]`) with a number (or numbers) in the brackets specifying which elements of the list you want.
#
# Let's start with a list of fruit:
# + colab={} colab_type="code" id="fQJFArRj9ezg"
# command Python to make a list and store it fruit
fruit = ['apple', 'banana', 'grape', 'mango']
# command Python to get the length of your list
# + [markdown] colab_type="text" id="Nt8UDOBF9ezk"
# Let's try getting the first element in the list:
# + colab={} colab_type="code" id="O20CWJ3t9ezl"
# command Python to print one element of your list
print(fruit[1])
# + [markdown] colab_type="text" id="9ozBuT_b9ezn"
# Is this what you expected? If not, you're not alone. The trick is that Python starts counting at zero!
#
# Here's how you actually print the first element of the list:
# + colab={} colab_type="code" id="L_GS8JXH9ezo"
# command Python to print the first element of your list
# + [markdown] colab_type="text" id="TfyIKK5j9ezq"
# How do you think we can print the element mango in the list?
# + colab={} colab_type="code" id="Y8HapUry9ezr"
# command Python to print the element mango
# + [markdown] colab_type="text" id="3Hpj3r_A9ezu"
# We can also get more than one element of a list:
# + colab={} colab_type="code" id="29nPZjp99ezv"
# command Python to get the first 3 elements of the list and save it to fruit_subset
# command Python to print the length of fruit_subset
# + colab={} colab_type="code" id="4GHOM5r-9ezx"
# command Python to print the length of fruit_subset
# + [markdown] colab_type="text" id="sF63s8je9ez1"
# Note that in Python, the first number in indexing is *inclusive* and the second number is *exclusive*. In other words, you get the element corresponding to the first number (0), but not the element corresponding to the last number (3). This is like (0,3] in math.
#
# Let's print the second and third elements of the list:
# + colab={} colab_type="code" id="cQu4h5519ez2"
# command Python to get the 2nd and 3rd elements of the list
# + [markdown] colab_type="text" id="KHS0wHOF9ez5"
# Now let's get the second and fourth elment from the list:
# + colab={} colab_type="code" id="j1Oh3pDa9ez5"
# command Python to print the second and fourth elements from the list
# + [markdown] colab_type="text" id="m8yfpDIm9ez9"
# You can also get letters in a string by indexing!
# + colab={} colab_type="code" id="rYEI3WRR9ez-"
# command Python to store the first element of fruit (apple) in the variable a
# command Python to print the first 3 letters in apple
# + [markdown] colab_type="text" id="inxnEUVl9e0E"
# You just learned how to:
# * Get an element from a list or string
# * Get multiple elements from a list or string
#
# Now it's time to practice what you learned!
| Lessons/Lesson05_Indexing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Edafa on ImageNet dataset
# This notebook shows an example on how to use Edafa to obtain better results on **classification task**. We use [ImageNet](http://www.image-net.org/) dataset which has **1000 classes**. We use *pytorch* and pretrained weights of AlexNet. At the end we compare results of the same model with and without augmentations.
# #### Import dependencies
# %load_ext autoreload
# %autoreload 2
# add our package directory to the path
import sys
sys.path.append('../../')
import torchvision.models as models
import torchvision.transforms as transforms
from torch.autograd import Variable
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# #### Constants
# +
# Filename to use for comparison (3 sample files are given in 'data' folder)
FILE = '000559'
# Input size of the deeplab model
IN_SIZE = 224
# -
# #### get labels
# Let's get our class labels.
labels = []
with open('labels.txt') as f:
for line in f:
labels.append(line.split(': ')[-1][1:-3])
# #### Now we build our model (using pretrained weights)
model = models.alexnet(pretrained=True)
# #### Read and preprocess image
img_path = '../data/images/%s.jpg'%FILE
img = plt.imread(img_path)
plt.imshow(img)
transform_pipeline = transforms.Compose([ transforms.ToPILImage(),
transforms.Resize((IN_SIZE,IN_SIZE)),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])])
x = transform_pipeline(img)
x = x.unsqueeze(0)
x = Variable(x)
# ### Exp1: Predict image without augmentation
pred = model(x)
pred_without = pred.data.numpy()
# ### Exp2: Using same model with Edafa
# #### step 1: import base class `ClassPredictor`
from edafa import ClassPredictor
# #### step 2: inherit `ClassPredictor` and implement the main virtual functions: predict_patches()
class myPredictor(ClassPredictor):
def __init__(self,vgg16,pipeline,*args,**kwargs):
super().__init__(*args,**kwargs)
self.model = vgg16
self.pipe = pipeline
def predict_patches(self,patches):
preds = []
for i in range(patches.shape[0]):
processed = self.pipe(patches[i])
processed = processed.unsqueeze(0)
processed = Variable(processed)
pred = self.model(processed)
preds.append(pred.data.numpy())
return np.array(preds)
# #### step 3: make an instance of your class with the correct parameters
p = myPredictor(model,transform_pipeline,"../../conf/imagenet.json")
# #### step 4: call predict_images()
preds_with = p.predict_images([img])
# ### Compare results of Exp1 and Exp2
print('Predicted without augmentation: ', labels[pred_without.argmax()])
print('Predicted with augmentation:', labels[preds_with.argmax()])
# We can clearly see from the object image that it's a desktop computer.
# With *no augmentation* the top prediction is **Polaroid camera, Polaroid Land camera**.
# With *augmentation* the top prediction is **desktop computer**
# ### Conclusion
# Results showed that with the exact same model and by applying Edafa we can obtain better results!
| examples/pytorch/class_imagenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow_p36]
# language: python
# name: conda-env-tensorflow_p36-py
# ---
# #### variables
num_classes = 10
baseMapNum = 32
weight_decay = 1e-4
batch_size = 50
epochs=75
# #### import packages and load cifar10 data
# +
import keras
from keras.models import Sequential
from keras.utils import np_utils
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Dense, Activation, Flatten, Dropout, BatchNormalization
from keras.layers import Conv2D, MaxPooling2D
from keras.datasets import cifar10
from keras import regularizers, optimizers
import numpy as np
(train_images, train_labels), (test_images, test_labels) = cifar10.load_data()
print("Training Data Shape :","Training Data :",train_images.shape)
print("Training Label Shape :",train_labels.shape)
print("Test Data Shape :",test_images.shape)
print("Test Label Shape :",test_labels.shape)
print("Train Data Length :",len(train_images))
print("Test Data Length :",len(test_images))
# -
# #### Data Preprocessing
# +
train_images = train_images.astype('float32')
test_images = test_images.astype('float32')
#calculate z-score
mean = np.mean(train_images,axis=(0,1,2,3))
std = np.std(train_images,axis=(0,1,2,3))
train_images = (train_images-mean)/(std+1e-6)
test_images = (test_images-mean)/(std+1e-6)
#one hot encoding of labels
train_labels = np_utils.to_categorical(train_labels,num_classes)
test_labels = np_utils.to_categorical(test_labels,num_classes)
#split training data into partial_train and validation_images
#val_data = train_images[:10000]
#partial_train = train_images[10000:]
#val_labels = train_labels[:10000]
#partial_labels = train_labels[10000:]
# -
# #### generate model
# +
model = Sequential()
model.add(Conv2D(baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay), input_shape=train_images.shape[1:]))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Conv2D(baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2,2)))
#model.add(Dropout(0.2))
model.add(Conv2D(2*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Conv2D(2*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2,2)))
#model.add(Dropout(0.3))
model.add(Conv2D(4*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Conv2D(4*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2,2)))
#model.add(Dropout(0.4))
model.add(Flatten())
model.add(Dense(num_classes, activation='softmax'))
model.summary()
# -
# #### Data Augmentation
datagen = ImageDataGenerator(
featurewise_center=False,
samplewise_center=False,
featurewise_std_normalization=False,
samplewise_std_normalization=False,
zca_whitening=False,
rotation_range=15,
width_shift_range=0.1,
height_shift_range=0.1,
horizontal_flip=True,
vertical_flip=False
)
datagen.fit(train_images)
# #### fit model and evaluate
opt_rms = keras.optimizers.rmsprop(lr=0.001,decay=1e-6)
model.compile(loss='categorical_crossentropy',
optimizer=opt_rms,
metrics=['accuracy'])
history = model.fit_generator(datagen.flow(train_images, train_labels, batch_size=50),steps_per_epoch=1000,
epochs=epochs,
validation_data=datagen.flow(test_images, test_labels, batch_size=10),
validation_steps=1000)
model.save_weights('cifar10_normal_rms_ep75.h5')
# #### plot loss graph
# +
import matplotlib.pyplot as plt
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# #### plot accuracy graph
# +
plt.clf() # clear figure
acc = history.history['acc']
val_acc = history.history['val_acc']
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# #### Check output for sample
scores = model.evaluate(test_images, test_labels, batch_size=10)
print('\nTest result: %.3f loss: %.3f' % (scores[1]*100,scores[0]))
predictions = model.predict(test_data)
print("Predicted label for test sample :",np.argmax(predictions[0]))
expected_label = [i for i, j in enumerate(test_labels[0]) if j == max(test_labels[0])]
print("Actual label for test sample :", expected_label)
# #### freeze upto second last layer of model and retrain
# +
model2 = Sequential()
model2.add(Conv2D(baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay), input_shape=train_images.shape[1:]))
model2.add(Activation('relu'))
model2.add(BatchNormalization())
model2.add(Conv2D(baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model2.add(Activation('relu'))
model2.add(BatchNormalization())
model2.add(MaxPooling2D(pool_size=(2,2)))
#model.add(Dropout(0.2))
model2.add(Conv2D(2*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model2.add(Activation('relu'))
model2.add(BatchNormalization())
model2.add(Conv2D(2*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model2.add(Activation('relu'))
model2.add(BatchNormalization())
model2.add(MaxPooling2D(pool_size=(2,2)))
#model.add(Dropout(0.3))
model2.add(Conv2D(4*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model2.add(Activation('relu'))
model2.add(BatchNormalization())
model2.add(Conv2D(4*baseMapNum, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))
model2.add(Activation('relu'))
model2.add(BatchNormalization())
model2.add(MaxPooling2D(pool_size=(2,2)))
#model.add(Dropout(0.4))
model2.add(Flatten())
model2.add(Dense(num_classes, activation='softmax'))
model2.load_weights('cifar10_normal_rms_ep75.h5')
for i, layer in enumerate(model2.layers):
print(i, layer.name)
# -
for layer in model2.layers[:16]:
layer.trainable = False
for layer in model2.layers[16:]:
layer.trainable = True
opt_rms = keras.optimizers.rmsprop(lr=0.0005,decay=1e-6)
model2.compile(loss='categorical_crossentropy',
optimizer=opt_rms,
metrics=['accuracy'])
history2 = model2.fit_generator(datagen.flow(train_images, train_labels, batch_size=50),steps_per_epoch=1000,
epochs=30,
validation_data=datagen.flow(test_images, test_labels, batch_size=10),
validation_steps=1000)
model2.save('cifar10-cnn-model-compile2.h5')
# #### plot loss graph
# +
import matplotlib.pyplot as plt
loss = history2.history['loss']
val_loss = history2.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# #### plot accuracy graph
# +
plt.clf() # clear figure
acc = history2.history['acc']
val_acc = history2.history['val_acc']
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# #### Check sample output
scores = model2.evaluate(test_images, test_labels, batch_size=10)
print('\nTest result: %.3f loss: %.3f' % (scores[1]*100,scores[0]))
predictions = model2.predict(test_images)
print("Predicted label for test sample :",np.argmax(predictions[0]))
expected_label = [i for i, j in enumerate(test_labels[0]) if j == max(test_labels[0])]
print("Actual label for test sample :", expected_label)
| Assignment 2 - CNN on cifar10/cognitive-modified-assignment1-cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from tensorflow import random_uniform_initializer as rui
# -
from preprocessing import DiagramScaler, Padding
from perslay import *
# + [markdown] heading_collapsed=true
# # Input persistence diagram
# + hidden=true
diag = [np.array([[0.,4.],[1.,2.],[3.,8.],[6.,8.]])]
# + hidden=true
plt.scatter(diag[0][:,0], diag[0][:,1])
plt.plot([0.,6.],[0.,6.])
plt.show()
# + hidden=true
diag = DiagramScaler(use=True, scalers=[([0,1], MinMaxScaler())]).fit_transform(diag)
diag = Padding(use=True).fit_transform(diag)
# + hidden=true
plt.scatter(diag[0][:,0], diag[0][:,1])
plt.plot([0.,1.],[0.,1.])
plt.show()
# + hidden=true
D = np.stack(np.array(diag, dtype=np.float32), 0)
# + hidden=true
diagrams, empty_feats = [D], np.empty([1,0], dtype=np.float32)
perslayParameters = {}
# + [markdown] heading_collapsed=true
# # Persistence weight and permutation invariant operation
# + [markdown] heading_collapsed=true hidden=true
# ## Choose persistence weights
# + hidden=true
perslayParameters["pweight_train"] = False
# + hidden=true
perslayParameters["pweight"] = "power"
perslayParameters["pweight_power"] = 2
perslayParameters["pweight_init"] = 1.
# + hidden=true
perslayParameters["pweight"] = "grid"
perslayParameters["pweight_size"] = [100,100]
perslayParameters["pweight_bnds"] = ((-.001, 1.001), (-.001, 1.001))
perslayParameters["pweight_init"] = np.tile(np.arange(0.,100.,1, dtype=np.float32)[np.newaxis,:], [100,1])
# + hidden=true
perslayParameters["pweight"] = "gmix"
perslayParameters["pweight_num"] = 3
perslayParameters["pweight_init"] = np.array(np.vstack([np.random.uniform(0.,1.,[2,3]),
5.*np.ones([2,3])]), dtype=np.float32)
# + hidden=true
perslayParameters["pweight"] = None
# + [markdown] heading_collapsed=true hidden=true
# ## Choose permutation invariant operation
# + hidden=true
perslayParameters["perm_op"] = "sum"
# + hidden=true
perslayParameters["perm_op"] = "topk"
perslayParameters["keep"] = 3
# + hidden=true
perslayParameters["perm_op"] = "max"
# + hidden=true
perslayParameters["perm_op"] = "mean"
# + [markdown] heading_collapsed=true
# # Persistence representation
# + [markdown] heading_collapsed=true hidden=true
# ## Persistence image
# + hidden=true
perslayParameters["layer"] = "Image"
perslayParameters["layer_train"] = False
perslayParameters["image_size"] = (100, 100)
perslayParameters["image_bnds"] = ((-.501, 1.501), (-.501, 1.501))
perslayParameters["lvariance_init"] = .1
perslayParameters["final_model"] = tf.keras.Sequential([tf.keras.layers.Flatten()])
# + hidden=true
model = PersLayModel(name="perslay", diagdim=2, perslay_parameters=[perslayParameters], rho="identity")
vector = model([diagrams, empty_feats])
# + hidden=true
# Plot representation
V = np.flip(np.reshape(vector[0,:], [int(np.sqrt(vector[0,:].shape[0])), int(np.sqrt(vector[0,:].shape[0]))]), 0)
plt.figure()
plt.imshow(V, cmap="Purples")
cb = plt.colorbar()
cb.ax.tick_params(labelsize=14)
plt.show()
# Plot weight
if perslayParameters["pweight"] == "grid":
W = model.vars[0][0].numpy()
weights = np.flip(W, 0)
plt.figure()
plt.imshow(weights, cmap="Purples", zorder=1)
((xm,xM),(ym,yM)) = perslayParameters["pweight_bnds"]
[xs, ys] = perslayParameters["pweight_size"]
plt.scatter([int(xs*(x-xm)/(xM-xm)) for x in diag[0][:,0]],
[ys-int(ys*(y-ym)/(yM-ym)) for y in diag[0][:,1]],
s=10, color="red", zorder=2)
plt.show()
if perslayParameters["pweight"] == "gmix":
means = model.vars[0][0][:2,:].numpy()
varis = model.vars[0][0][2:,:].numpy()
x = np.arange(-.5, 1.5, .001)
y = np.arange(-.5, 1.5, .001)
xx, yy = np.meshgrid(x, y)
z = np.zeros(xx.shape)
for idx_g in range(means.shape[1]):
z += np.exp(-((xx-means[0,idx_g])**2 * (varis[0,idx_g])**2
+ (yy-means[1,idx_g])**2 * (varis[1,idx_g])**2 ))
plt.contourf(xx, yy, z)
plt.scatter(diag[0][:,0], diag[0][:,1], s=50, color="red")
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ## Persistence landscape / entropy / Betti curve
# + hidden=true
#perslayParameters["layer"] = "Landscape"
#perslayParameters["layer"] = "BettiCurve"
perslayParameters["layer"] = "Entropy"
perslayParameters["layer_train"] = False
perslayParameters["lsample_num"] = 3000
perslayParameters["lsample_init"] = np.array(np.arange(-1.,2.,.001), dtype=np.float32)
perslayParameters["theta"] = 100 # used only if layer is "BettiCurve" or "Entropy"
perslayParameters["final_model"] = "identity"
# + hidden=true
model = PersLayModel(name="perslay", diagdim=2, perslay_parameters=[perslayParameters], rho="identity")
vector = model([diagrams, empty_feats])
# + hidden=true
#Plot representation
V = vector[0,:]
plt.figure()
if perslayParameters["perm_op"] == "topk":
V = np.reshape(V, [-1, perslayParameters["keep"]])
for k in range(perslayParameters["keep"]):
plt.plot(V[:,k], linewidth=5.0)
else:
plt.plot(V, linewidth=5.0)
plt.show()
# Plot weight
if perslayParameters["pweight"] == "grid":
W = model.vars[0][0].numpy()
weights = np.flip(W, 0)
plt.figure()
plt.imshow(weights, cmap="Purples", zorder=1)
((xm,xM),(ym,yM)) = perslayParameters["pweight_bnds"]
[xs, ys] = perslayParameters["pweight_size"]
plt.scatter([int(xs*(x-xm)/(xM-xm)) for x in diag[0][:,0]],
[ys-int(ys*(y-ym)/(yM-ym)) for y in diag[0][:,1]],
s=10, color="red", zorder=2)
plt.show()
if perslayParameters["pweight"] == "gmix":
means = model.vars[0][0][:2,:].numpy()
varis = model.vars[0][0][2:,:].numpy()
x = np.arange(-.5, 1.5, .001)
y = np.arange(-.5, 1.5, .001)
xx, yy = np.meshgrid(x, y)
z = np.zeros(xx.shape)
for idx_g in range(means.shape[1]):
z += np.exp(-((xx-means[0,idx_g])**2 * (varis[0,idx_g])**2
+ (yy-means[1,idx_g])**2 * (varis[1,idx_g])**2 ))
plt.contourf(xx, yy, z)
plt.scatter(diag[0][:,0], diag[0][:,1], s=50, color="red")
plt.show()
# + hidden=true
| tutorial/visuPerslay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
import numpy as np
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
purchase_data.head()
# -
# ## Player Count
# * Display the total number of players
#
player_count = len(purchase_data["SN"].unique())
totalplayer_table = pd.DataFrame({"Total Players": player_count}, index=[0])
totalplayer_table
# ## Purchasing Analysis (Total)
# * Run basic calculations to obtain number of unique items, average price, etc.
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
#
# +
unique_items = len(purchase_data["Item ID"].unique())
average_price = round(purchase_data["Price"].mean(), 2)
total_purchases = purchase_data["Purchase ID"].count()
total_revenue = purchase_data["Price"].sum()
purch_analysis_table = pd.DataFrame({"Number of Unique Items": unique_items,
"Average Price": average_price,
"Number of Purchases": total_purchases,
"Total Revenue": total_revenue},
index=[0])
purch_analysis_table
# -
# ## Gender Demographics
# * Percentage and Count of Male Players
#
#
# * Percentage and Count of Female Players
#
#
# * Percentage and Count of Other / Non-Disclosed
#
#
#
# +
purchase_data_df = purchase_data.groupby("Gender")["SN"].nunique()
gender_pct = round(purchase_data_df / player_count, 4) * 100
demograph_table = pd.DataFrame({"Player Count": purchase_data_df, "Player Pct.": gender_pct})
demograph_table
# -
#
# ## Purchasing Analysis (Gender)
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
#
#
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
purch_analysis_df = purchase_data.groupby("Gender")["Purchase ID"].nunique()
avg_price = round(purchase_data.groupby("Gender")["Price"].mean(), 2)
tot_purch_val = round(purchase_data.groupby("Gender")["Price"].sum(), 2)
avg_tot_purch = round((tot_purch_val / purchase_data_df), 2)
analysis_table = pd.DataFrame({"Purchase Count": purch_analysis_df, "Avg Purch Price": avg_price,
"Tot Purch Val": tot_purch_val, "Avg Tot Purch per Person": avg_tot_purch})
analysis_table
# -
# ## Age Demographics
# * Establish bins for ages
#
#
# * Categorize the existing players using the age bins. Hint: use pd.cut()
#
#
# * Calculate the numbers and percentages by age group
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: round the percentage column to two decimal points
#
#
# * Display Age Demographics Table
#
# +
bins = [0, 9, 14, 19, 24, 29, 34, 39, 110]
group_labels = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
bin_data = pd.DataFrame(purchase_data)
bin_data = bin_data.groupby(["SN"], as_index=False).max()
bin_data[""] = pd.cut(bin_data["Age"], bins, labels=group_labels)
bin_data = bin_data.groupby("")
age_count = bin_data["Age"].count()
age_pct = round(age_count / player_count, 4) * 100
bin_table = pd.DataFrame({"Total Count": age_count, "Pct of Players": age_pct})
bin_table
# -
# ** PURCHASING ANALYSIS **
# * Bin the purchase_data data frame by age
#
#
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
# totpurch_per_df = pd.DataFrame(purchase_data)
# tot_purch_per = totpurch_per_df.groupby("Price")["SN"].max
# tot_purch_per
age_bins = [0, 9, 14, 19, 24, 29, 34, 39, 110]
age_labels = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
agebin_data = pd.DataFrame(purchase_data)
agebin_data["Age Ranges"] = pd.cut(agebin_data["Age"], age_bins, labels=age_labels)
agebin_data = agebin_data.groupby("Age Ranges")
newage_count = agebin_data["Purchase ID"].count()
avg_purch = round(agebin_data["Price"].mean(), 2)
tot_val = agebin_data["Price"].sum()
#avg_purch_per
newage_table = pd.DataFrame({"Purchase Count": newage_count, "Avg Purch Price": avg_purch,
"Tot Purch Val": tot_val}) #"Avg Tot Purch Per Person": avg_purch_per})
newage_table
# -
# ## Top Spenders
# * Run basic calculations to obtain the results in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the total purchase value column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
sn_purchcount = purchase_data.groupby("SN")["Purchase ID"].count()
avg_purch_price = round(purchase_data.groupby("SN")["Price"].mean(), 2)
total_purch_val = purchase_data.groupby("SN")["Price"].sum()
topspender_df = pd.DataFrame({"Purchase Count": sn_purchcount, "Avg Purch Price": avg_purch_price,
"Total Purch Val": total_purch_val})
topspender_df.sort_values("Total Purch Val", ascending=False).head()
# -
# ## Most Popular Items
# * Retrieve the Item ID, Item Name, and Item Price columns
#
#
# * Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the purchase count column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
popular_df = pd.DataFrame(purchase_data)
popular_table = popular_df.groupby("Item ID")["Item Name"]
pop_count = popular_df.groupby("Item Name")["Purchase ID"].count()
pop_price = popular_df.groupby("Item Name")["Price"].max()
pop_totval = popular_df.groupby("Item Name")["Price"].sum()
popular_table_df = pd.DataFrame({"Purchase Count": pop_count, "Item Price": pop_price, "Tot Purch Val": pop_totval})
popular_table_df.sort_values("Purchase Count", ascending=False).head()
# -
# ## Most Profitable Items
# * Sort the above table by total purchase value in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the data frame
#
#
# +
popular_df = pd.DataFrame(purchase_data)
popular_table = popular_df.groupby("Item ID")["Item Name"]
pop_count = popular_df.groupby("Item Name")["Purchase ID"].count()
pop_price = popular_df.groupby("Item Name")["Price"].max()
pop_totval = popular_df.groupby("Item Name")["Price"].sum()
popular_table_df = pd.DataFrame({"Purchase Count": pop_count, "Item Price": pop_price, "Tot Purch Val": pop_totval})
popular_table_df.sort_values("Tot Purch Val", ascending=False).head()
# -
## OBSERVABLES
# 1) Gaming is male dominated
# 2) Nearly half of gamers are in their early 20's - approx 58% of all gamers are in their 20's
# 3) Marketing dollars should focus on targeting age 15-29 males
| HeroesOfPymoli/HeroesOfPymoli_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
## Trying to find the best method of determining language of books
import pandas as pd
import numpy as np
items = pd.read_csv('items.csv', sep = '|')
# +
count = 0
item_counter = 0
for title in items['title']:
if (len(title) < 4):
print('## Title: ', title)
print("items = items.drop(", items.loc[count, 'temp'], ')')
item_counter = item_counter + 1
count = count + 1
item_counter
# +
## Here, we are removing the observations that language identifying softwares cannot handle,
## such as titles that are only numbers and titles that are too short. We can manually input
## these observations back in later with their languages
items = items.drop(124)
items = items.drop(582)
# -
## Title: 1984
items = items.drop(736)
## Title: 1984
items = items.drop(3660)
## Title: 381
items = items.drop(1282)
## Title: 2034
items = items.drop(3871)
## Title: 2021
items = items.drop(6224)
## Title: 2037
items = items.drop(7458)
## Title: 43
items = items.drop(10749)
## Title: 2069
items = items.drop(10944)
## Title: 2121
items = items.drop(13490)
## Title: 2394
items = items.drop(13888)
## Title: 2121
items = items.drop(15060)
## Title: 2048
items = items.drop(15940)
## Title: 17
items = items.drop(15555)
## Title: 2121
items = items.drop(17732)
## Title: 1632
items = items.drop(21448)
## Title: Ferris@Bruns_LLC
items = items.drop(21929)
## Title: 1814
items = items.drop(23431)
## Title: 2012
items = items.drop(25479)
## Title: 11
items = items.drop(25567)
## Title: !!
items = items.drop(31055)
## Title: Tajo@Bruns_LLC
items = items.drop(32694)
## Title: 1,2,3
items = items.drop(35645)
## Title: 110
items = items.drop(39581)
## Title: 5:55
items = items.drop(40733)
## Title: 1906
items = items.drop(41657)
## Title: 1906
items = items.drop(41664)
## Title: 2030
items = items.drop(43450)
## Title: 2047
items = items.drop(43766)
## Title: 2049
items = items.drop(77770)
## Title: 444
items = items.drop(44841)
## Title: 444
items = items.drop(44842)
## Title: 2501
items = items.drop(48254)
## Title: 2060
items = items.drop(48905)
## Title: 5028
items = items.drop(50669)
## Title: 2084
items = items.drop(55060)
## Title: 2053
items = items.drop(55370)
## Title: 1520-1522
items = items.drop(55539)
## Title: 2145
items = items.drop(61473)
## Title: 1523-1526
items = items.drop(64005)
## Title: 1712
items = items.drop(64441)
## Title: 2084
items = items.drop(64706)
## Title: 2625
items = items.drop(65362)
## Title: 2084
items = items.drop(66980)
## Title: >
items = items.drop(67092)
## Title: 2156
items = items.drop(67283)
## Title: 6984
items = items.drop(67286)
## Title: 12
items = items.drop(69842)
## Title: 71%
items = items.drop(72536)
## Title: 4
items = items.drop(73961)
## Title: Io
items = items.drop( 136 )
## Title: Ava
items = items.drop( 831 )
## Title: ABC
items = items.drop( 3677 )
## Title: EMP
items = items.drop( 4022 )
## Title: Hex
items = items.drop( 4465 )
## Title: Wir
items = items.drop( 4814 )
## Title: Boy
items = items.drop( 5054 )
## Title: Oma
items = items.drop( 5156 )
## Title: Nik
items = items.drop( 5698 )
## Title: Kim
items = items.drop( 6260 )
## Title: Pet
items = items.drop( 6407 )
## Title: M
items = items.drop( 6999 )
## Title: Äon
items = items.drop( 7000 )
## Title: Ava
items = items.drop( 7509 )
## Title: XX
items = items.drop( 7963 )
## Title: Wir
items = items.drop( 7999 )
## Title: Dry
items = items.drop( 8895 )
## Title: Ash
items = items.drop( 8896 )
## Title: FOX
items = items.drop( 11149 )
## Title: Cut
items = items.drop( 12669 )
## Title: ID
items = items.drop( 12974 )
## Title: Ehi
items = items.drop( 12987 )
## Title: ICE
items = items.drop( 13010 )
## Title: We
items = items.drop( 13142 )
## Title: Wir
items = items.drop( 13234 )
## Title: Pax
items = items.drop( 13532 )
## Title: Eve
items = items.drop( 13748 )
## Title: Ye
items = items.drop( 15053 )
## Title: Ral
items = items.drop( 15379 )
## Title: Ten
items = items.drop( 15749 )
## Title: BEX
items = items.drop( 17253 )
## Title: Äon
items = items.drop( 18532 )
## Title: Yo
items = items.drop( 18912 )
## Title: ZPG
items = items.drop( 19360 )
## Title: Liv
items = items.drop( 19716 )
## Title: Q
items = items.drop( 19982 )
## Title: Dig
items = items.drop( 20544 )
## Title: Tin
items = items.drop( 20751 )
## Title: Sea
items = items.drop( 21159 )
## Title: SHE
items = items.drop( 22397 )
## Title: Run
items = items.drop( 22675 )
## Title: Noa
items = items.drop( 22696 )
## Title: Vet
items = items.drop( 22798 )
## Title: NDI
items = items.drop( 23453 )
## Title: Mud
items = items.drop( 23766 )
## Title: COR
items = items.drop( 24594 )
## Title: Hb
items = items.drop( 25835 )
## Title: Fee
items = items.drop( 25957 )
## Title: Tao
items = items.drop( 26453 )
## Title: Lie
items = items.drop( 26609 )
## Title: KY
items = items.drop( 26723 )
## Title: Sol
items = items.drop( 27164 )
## Title: Eve
items = items.drop( 27296 )
## Title: If
items = items.drop( 28475 )
## Title: Ink
items = items.drop( 29260 )
## Title: L
items = items.drop( 29902 )
## Title: Zoe
items = items.drop( 30029 )
## Title: Tex
items = items.drop( 30314 )
## Title: Red
items = items.drop( 31667 )
## Title: MAY
items = items.drop( 32187 )
## Title: Aim
items = items.drop( 32830 )
## Title: Son
items = items.drop( 33415 )
## Title: L
items = items.drop( 33504 )
## Title: Nil
items = items.drop( 33672 )
## Title: Sie
items = items.drop( 34131 )
## Title: Eva
items = items.drop( 34908 )
## Title: Zel
items = items.drop( 35241 )
## Title: Web
items = items.drop( 35451 )
## Title: Hit
items = items.drop( 36713 )
## Title: One
items = items.drop( 36886 )
## Title: Mia
items = items.drop( 37465 )
## Title: Sun
items = items.drop( 37739 )
## Title: One
items = items.drop( 38315 )
## Title: Up
items = items.drop( 38619 )
## Title: Liv
items = items.drop( 38885 )
## Title: Max
items = items.drop( 39320 )
## Title: Run
items = items.drop( 39328 )
## Title: Spy
items = items.drop( 39776 )
## Title: Jim
items = items.drop( 40474 )
## Title: A
items = items.drop( 41040 )
## Title: Fir
items = items.drop( 41156 )
## Title: GEN
items = items.drop( 41759 )
## Title: Quo
items = items.drop( 41762 )
## Title: Sky
items = items.drop( 42360 )
## Title: Meh
items = items.drop( 42427 )
## Title: Eco
items = items.drop( 42706 )
## Title: Pug
items = items.drop( 43203 )
## Title: A
items = items.drop( 43324 )
## Title: May
items = items.drop( 45723 )
## Title: I
items = items.drop( 46217 )
## Title: Sue
items = items.drop( 46444 )
## Title: Er?
items = items.drop( 46586 )
## Title: Sin
items = items.drop( 47139 )
## Title: Sam
items = items.drop( 47752 )
## Title: Ohm
items = items.drop( 48500 )
## Title: Bob
items = items.drop( 49109 )
## Title: Aim
items = items.drop( 49184 )
## Title: Few
items = items.drop( 49591 )
## Title: A M
items = items.drop( 49704 )
## Title: War
items = items.drop( 50112 )
## Title: Z
items = items.drop( 50130 )
## Title: Kim
items = items.drop( 51251 )
## Title: Kim
items = items.drop( 52784 )
## Title: ANK
items = items.drop( 52943 )
## Title: ADA
items = items.drop( 53145 )
## Title: Ink
items = items.drop( 54822 )
## Title: KAT
items = items.drop( 55063 )
## Title: The
items = items.drop( 55499 )
## Title: ANA
items = items.drop( 55570 )
## Title: PC
items = items.drop( 57219 )
## Title: Ray
items = items.drop( 59436 )
## Title: Nyx
items = items.drop( 59743 )
## Title: Ka
items = items.drop( 59756 )
## Title: Reg
items = items.drop( 60311 )
## Title: Hex
items = items.drop( 60620 )
## Title: ABC
items = items.drop( 60681 )
## Title: TEA
items = items.drop( 61456 )
## Title: Wi
items = items.drop( 61998 )
## Title: Red
items = items.drop( 62262 )
## Title: Jo
items = items.drop( 62777 )
## Title: Tok
items = items.drop( 63018 )
## Title: Ug
items = items.drop( 64328 )
## Title: Vic
items = items.drop( 64507 )
## Title: FOX
items = items.drop( 64785 )
## Title: ISO
items = items.drop( 64919 )
## Title: Lu
items = items.drop( 65886 )
## Title: Fey
items = items.drop( 67625 )
## Title: Fey
items = items.drop( 67628 )
## Title: HPI
items = items.drop( 67743 )
## Title: Air
items = items.drop( 68029 )
## Title: Red
items = items.drop( 68476 )
## Title: Mer
items = items.drop( 68490 )
## Title: Zoo
items = items.drop( 68936 )
## Title: Bee
items = items.drop( 69689 )
## Title: PUP
items = items.drop( 70156 )
## Title: Été
items = items.drop( 71858 )
## Title: Jòn
items = items.drop( 71865 )
## Title: Ble
items = items.drop( 71876 )
## Title: Ete
items = items.drop( 71920 )
## Title: Vèt
items = items.drop( 71933 )
## Title: Buk
items = items.drop( 72131 )
## Title: Kam
items = items.drop( 72461 )
## Title: TBC
items = items.drop( 72583 )
## Title: ERA
items = items.drop( 72846 )
## Title: H2O
items = items.drop( 72850 )
## Title: Kid
items = items.drop( 72901 )
## Title: No!
items = items.drop( 73228 )
## Title: VTT
items = items.drop( 73954 )
## Title: Fix
items = items.drop( 74360 )
## Title: Bec
items = items.drop( 74598 )
## Title: Zoo
items = items.drop( 75554 )
## Title: WTF
items = items.drop( 77324 )
## Title: Kim
items = items.drop( 77674 )
## Title: Now
items = items.drop( 77983 )
# +
## Using: langdetect
from langdetect import detect
from langdetect import DetectorFactory
DetectorFactory.seed = 0
items['language'] = ''
myList = items['title']
languages = []
for text in myList:
languages.append(detect(text))
##print(text)
items['language'] = languages
# +
## Using: textblob
## Seems to be the more accurate method
## Limiting calls with the sleep function
from textblob import TextBlob
import time
items['language_textblob'] = ''
myList = items['title']
languages = []
for text in myList:
lang = TextBlob(text)
languages.append(lang.detect_language())
print(text)
time.sleep(0.5) ## To sleep for one second
items['language_textblob'] = languages
# -
items = items.drop(columns=['temp', 'language'])
items['language'] = items['language_textblob']
items = items.drop(columns = ['language_textblob'])
# +
################################
## Exporting the new data set ##
################################
items.to_csv('items_1.csv', index = False, header = True)
| Preprocessing/Creating_Language.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KaustavRaj/Text-Summarization/blob/master/Text_Summarization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Vf49qAonizPz"
# # Text Summarization
# ##### *by <NAME>, IIIT Guwahati*
# ---
# + [markdown] colab_type="text" id="_4z-8-4ei4QF"
# In this notebook, I'm building a summarizer step-by-step, based on **Amazon Fine Food Reviews** dataset, which I have stored in **/data** section but it can also be found at kaggle website.
# + [markdown] colab_type="text" id="ur_-I-_Lm7EW"
# Let's first connect google colab to google drive, where I've saved my dataset.
# + colab_type="code" id="jaZCjnAsToJn" outputId="7714c4e6-d2f5-4b76-ef62-dddaaaf242ba" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] colab_type="text" id="9d6sLE8zUAHG"
# ## Importing Libraries
# + colab_type="code" id="KWiygrk3T2O5" outputId="23d610bc-04d3-42ee-e4ad-6063a440db1e" colab={"base_uri": "https://localhost:8080/", "height": 51}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import re
import tensorflow as tf
from bs4 import BeautifulSoup
from tensorflow.keras.layers import Bidirectional, LSTM, Input, Embedding, Dense, TimeDistributed, Concatenate, Attention
from tensorflow.keras.models import Model, load_model
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
# + colab_type="code" id="m8bccQJWs5rW" outputId="df3caef6-e371-4c80-bbe7-72b68b1b81bf" colab={"base_uri": "https://localhost:8080/", "height": 326}
# !pip install contractions
# + colab_type="code" id="aVJXF2Nh3TkV" colab={}
import contractions
# + colab_type="code" id="cpWJAFCQUPtW" colab={}
dir_path = '/content/gdrive/My Drive/Colab Notebooks/Summarization/summarization v2'
# + [markdown] colab_type="text" id="_xDdll4FUVnz"
# ## Dataset Loading & Cleaning
# + colab_type="code" id="PFsOlda93m8T" colab={}
# Declaring the important variables here, which will be used later down the notebook
_NROWS = 100000
_MAX_TEXT_LEN = 0
_MAX_SUMMARY_LEN = 0
_TEXT_PADDING = 'post'
_EMBEDDING_DIM = 500
_ENCODER_DIM = 300
_DECODER_DIM = 600
_BATCH_SIZE = 64
_EPOCHS = 40
# + colab_type="code" id="W1UKq_o_tJQb" outputId="f5378bec-1631-491c-fc55-0321e3f9d0c8" colab={"base_uri": "https://localhost:8080/", "height": 527}
data = pd.read_csv(dir_path + '/data/Reviews.csv', nrows = _NROWS)
data.drop_duplicates(subset = ['Text'], inplace = True)
data.dropna(axis = 0, inplace = True)
data.head()
# + colab_type="code" id="Ku30l2oO9pbj" colab={}
def cleaner(text, remove_stopwords=True):
"""removes url's, nltk's stopwords and anything which is not an alphabet"""
stop_words = set(stopwords.words('english'))
text = re.sub(r'^https?:\/\/.*[\r\n]*', '', text.lower(), flags=re.MULTILINE)
text = re.sub(r'[^a-zA-Z]', ' ', text)
text = contractions.fix(text, slang=False)
if remove_stopwords:
text = ' '.join([word for word in text.split() if word not in stop_words]).strip()
return text
# + colab_type="code" id="JQFJOPcWtaj5" outputId="9da83c02-d7df-467a-c46c-a2d638a2b8dd" colab={"base_uri": "https://localhost:8080/", "height": 34}
from time import time
from math import ceil
t1 = time()
cleaned_text = []
for t in data['Text']:
cleaned_text.append(cleaner(t))
t2 = time()
cleaned_summary = []
for t in data['Summary']:
cleaned_summary.append(cleaner(t, remove_stopwords = False))
t3 = time()
print('Text cleaned in {} sec, and Summary in {} sec'.format(ceil(t2-t1), ceil(t3-t2)))
data = pd.DataFrame({
'text' : cleaned_text,
'summary' : cleaned_summary})
data.replace('', np.nan, inplace = True)
data.dropna(axis = 0, inplace = True)
# + colab_type="code" id="SgH33OrGNxyB" colab={}
data.reset_index(inplace=True, drop=True)
# + colab_type="code" id="TbKwk06uL4vM" outputId="667d4bbc-d867-4339-ba50-7d633de6c82e" colab={"base_uri": "https://localhost:8080/", "height": 34}
data.shape
# + colab_type="code" id="o498BXSrcuq4" colab={}
data.to_csv(dir_path + '/data/cleaned_amazon_reviews_dataset.csv', index=False)
# + [markdown] colab_type="text" id="VRC6jq-3jVCq"
# ## Visualizing the Text Data
# + colab_type="code" id="JGBzJTxPisLp" outputId="0bc18f2e-0131-4bfe-a288-27b4b303415f" colab={"base_uri": "https://localhost:8080/", "height": 607}
text_word_count = []
summary_word_count = []
for i in data['text']:
text_word_count.append(len(i.split()))
for i in data['summary']:
summary_word_count.append(len(i.split()))
visualize = pd.DataFrame({'text':text_word_count, 'summary':summary_word_count})
visualize.hist(bins = 50)
plt.rcParams['figure.figsize'] = (10, 10)
plt.show()
# + colab_type="code" id="jc0Can2OL_hG" outputId="966c982e-84e6-424c-b2e2-1c50887b0cba" colab={"base_uri": "https://localhost:8080/", "height": 34}
cnt = 0
for i in range(data.shape[0]):
if len(data['text'][i].split()) <= 60 and len(data['summary'][i].split()) <= 10:
cnt += 1
print(cnt)
# + [markdown] colab_type="text" id="lJbCyzxEsJc2"
# It's nearly 80% of our dataset, so its fairly decent in size as well.
# + colab_type="code" id="jr9KdnznisYq" colab={}
_MAX_SUMMARY_LEN = 10
_MAX_TEXT_LEN = 60
# + colab_type="code" id="g7Q0dou9URrd" colab={}
def prepare(dataset):
global _MAX_SUMMARY_LEN, _MAX_TEXT_LEN
newtext = []
newsummary = []
for i in range(len(dataset['text'])):
if len(dataset['text'][i].split()) <= _MAX_TEXT_LEN and len(dataset['summary'][i].split()) <= _MAX_SUMMARY_LEN:
newtext.append(dataset['text'][i])
newsummary.append(dataset['summary'][i])
dataset = pd.DataFrame({'text' : newtext,
'summary' : newsummary})
dataset['summary'] = dataset['summary'].apply(lambda row : 'stok ' + row + ' etok')
dataset.reset_index(drop=True, inplace=True)
return dataset
# + colab_type="code" id="ANB2RbiwLDFD" colab={}
df = prepare(data)
# + colab_type="code" id="zRJha9lPURw-" outputId="17c6cabb-0a0d-4222-c660-6671966d3cf8" colab={"base_uri": "https://localhost:8080/", "height": 34}
df.shape
# + colab_type="code" id="PSi6hKAbdC7E" colab={}
df.to_csv(dir_path + '/data/prepared_amazon_reviews_dataset.csv', index=False)
# + [markdown] colab_type="text" id="5SGi36lpYAO-"
# ## Training-Validation Data Splitting
# + [markdown] colab_type="text" id="izp5fcf7mIDh"
# Loading the data here if its already cleaned.
# + colab_type="code" id="tRgvDjlHl222" colab={}
df = pd.read_csv(dir_path + '/data/prepared_amazon_reviews_dataset.csv')[['text', 'summary']]
# + colab_type="code" id="ufhDr9KHuane" outputId="69e5d42d-13cc-4762-adcd-16496a8af3f7" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + colab_type="code" id="K8CKXSjeUR4P" colab={}
x_tr, x_val, y_tr, y_val = train_test_split(np.array(df['text']), np.array(df['summary']),
test_size=0.1, random_state=42)
# + colab_type="code" id="87HDi5dkjnm-" colab={}
# preparing 'text' for model training and validation
tok_x = Tokenizer()
tok_x.fit_on_texts(list(x_tr))
x_tr = pad_sequences(tok_x.texts_to_sequences(x_tr), maxlen=_MAX_TEXT_LEN, padding=_TEXT_PADDING)
x_val = pad_sequences(tok_x.texts_to_sequences(x_val), maxlen=_MAX_TEXT_LEN, padding=_TEXT_PADDING)
vocab_x = len(tok_x.word_counts) + 1
# + colab_type="code" id="egSUyJyOmsE7" outputId="5e8a5696-6001-43cb-ec30-c6acab0d4a99" colab={"base_uri": "https://localhost:8080/", "height": 34}
vocab_x
# + colab_type="code" id="G8OYAQzUvhqt" colab={}
# preparing 'summary' for model training and validation
tok_y = Tokenizer()
tok_y.fit_on_texts(list(y_tr))
y_tr = pad_sequences(tok_y.texts_to_sequences(y_tr), maxlen=_MAX_TEXT_LEN, padding=_TEXT_PADDING)
y_val = pad_sequences(tok_y.texts_to_sequences(y_val), maxlen=_MAX_TEXT_LEN, padding=_TEXT_PADDING)
vocab_y = len(tok_y.word_counts) + 1
# + colab_type="code" id="LNiVCaG5V2eB" colab={}
with open(dir_path + '/data/tok_x.pickle', 'wb') as f:
pickle.dump(tok_x, f)
# + colab_type="code" id="BxWLdYz-jnh4" outputId="eb5ffacc-7087-4834-e963-c957d5956e6d" colab={"base_uri": "https://localhost:8080/", "height": 34}
vocab_y
# + colab_type="code" id="sZ95yxZGjndz" outputId="c822f296-ca7f-4bb8-93a9-687d603991e9" colab={"base_uri": "https://localhost:8080/", "height": 34}
tok_y.word_counts['stok'], len(y_tr)
# + colab_type="code" id="_U_fDYsx9yoo" colab={}
index_to_word_text = tok_x.index_word
index_to_word_summary = tok_y.index_word
word_to_index_summary = tok_y.word_index
# + colab_type="code" id="RLul_vbvvuDs" colab={}
import pickle
with open(dir_path + '/data/word_indices_mapping.pickle', 'wb') as f:
store = [index_to_word_text, index_to_word_summary, word_to_index_summary]
pickle.dump(store, f)
# + [markdown] colab_type="text" id="e8vrQjknpsdm"
# ## Model Building
# + [markdown] colab_type="text" id="rdgb0IO1odXy"
# A simple Sequence to Sequence encoder-decoder model has been used, where a bidirectional lstm is used as encoder and a unidirectional lstm for decoder with attention layer.
# + colab_type="code" id="3ErjwaMmwVTS" outputId="9b3d4763-5d92-47a1-a6e4-97070baaaf66" colab={"base_uri": "https://localhost:8080/", "height": 629}
# Encoder
encoder_input = Input(shape=(_MAX_TEXT_LEN,), name='Encoder_input')
encoder_embed = Embedding(vocab_x, _EMBEDDING_DIM, trainable=True, name='Encoder_embedding')(encoder_input)
encoder = Bidirectional(LSTM(_ENCODER_DIM,
return_sequences=True,
return_state=True,
dropout=0.4, recurrent_dropout=0.4), name='Encoder_layer')
encoder_output, forward_h, forward_c, backward_h, backward_c = encoder(encoder_embed)
state_h = Concatenate()([forward_h, backward_h])
state_c = Concatenate()([forward_c, backward_c])
# Decoder
decoder_input = Input(shape=(None,), name='Decoder_input')
decoder_embed_layer = Embedding(vocab_y, _EMBEDDING_DIM, trainable=True, name='Decoder_embedding')
decoder_embed = decoder_embed_layer(decoder_input)
decoder = LSTM(_DECODER_DIM,
return_sequences=True,
return_state=True,
dropout=0.4, recurrent_dropout=0.3, name='Decoder_layer')
decoder_output, decoder_state_h, decoder_state_c = decoder(decoder_embed,initial_state=[state_h, state_c])
# Attention layer
attention_layer = Attention(name='Attention_layer')
attention_out = attention_layer([decoder_output, encoder_output])
# Concat attention layer's output and decoder's output
concat_layer = Concatenate(axis=-1, name='Concat_layer')([decoder_output, attention_out])
# Dense layer
decoder_dense = TimeDistributed(Dense(vocab_y, activation='softmax'))
decoder_output = decoder_dense(concat_layer)
# Model
model = Model([encoder_input, decoder_input], decoder_output, name='Model')
model.summary()
# + colab_type="code" id="Kdib3Z0C0LLE" outputId="a2233281-4984-4609-8bf1-ccae1b813961" colab={"base_uri": "https://localhost:8080/", "height": 289}
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy')
stopearly = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=2)
X = [x_tr,y_tr[:,:-1]]
y = y_tr.reshape(y_tr.shape[0], y_tr.shape[1], 1)[:,1:]
VAL_X = [x_val,y_val[:,:-1]]
VAL_y = y_val.reshape(y_val.shape[0], y_val.shape[1], 1)[:,1:]
history = model.fit(X, y, epochs=_EPOCHS, callbacks=[stopearly], batch_size=_BATCH_SIZE, validation_data=(VAL_X, VAL_y))
# + colab_type="code" id="0-JB5cg35_b4" outputId="c5561f52-4c6c-4c9c-b2bf-d0cccc184c28" colab={"base_uri": "https://localhost:8080/", "height": 295}
# For loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.xlabel('no. of epochs')
plt.ylabel('loss')
plt.legend(['train', 'validate'], loc='upper right')
plt.show()
# + colab_type="code" id="NK_t0ej5XQZj" colab={}
_MODEL_NAME = 'model_3'
model.save(dir_path + '/models/{}.h5'.format(_MODEL_NAME))
# + colab_type="code" id="uDrmmKkJXXec" colab={}
import json
specs = {
'model_name' : _MODEL_NAME,
'embedding_dim' : _EMBEDDING_DIM,
'encoder' : [{'type':'bi-LSTM', 'dim':_ENCODER_DIM}],
'decoder' : [{'type':'LSTM', 'dim': _DECODER_DIM}],
'attention' : {
'used' : True,
'type' : 'Luong'
},
'batch_size' : _BATCH_SIZE,
'epochs_completed' : 7,
'max_text_len' : _MAX_TEXT_LEN,
'max_summary_len' : _MAX_SUMMARY_LEN,
'nrows_dataset' : {
'total' : 100000,
'used' : 70707
},
'additional' : ""
}
with open(dir_path + '/models/specs3.json', 'a+') as f:
json.dump(specs, f)
# + [markdown] colab_type="text" id="c3cHH7789s2C"
# ## Inference
# + colab_type="code" id="M_ckN-vUjnOC" colab={}
encoder_model = Model(inputs = encoder_input,
outputs = [encoder_output, state_h, state_c])
# For storing decoder's previous states which will act as our encoder here
decoder_previous_state_h = Input(shape=(_DECODER_DIM,))
decoder_previous_state_c = Input(shape=(_DECODER_DIM,))
decoder_previous_output = Input(shape=(_MAX_TEXT_LEN, _DECODER_DIM))
# This new decoder will take the 'previous decoder' i.e, our encoder's output
decoder_embed_new = decoder_embed_layer(decoder_input)
decoder_output_new, state_h_new, state_c_new = decoder(decoder_embed_new,
initial_state = [decoder_previous_state_h, decoder_previous_state_c])
# Attention layer & concat layer for the new decoder and encoder
attention_out_new = attention_layer([decoder_output_new, decoder_previous_output])
concat_layer_final = Concatenate(axis=-1)([decoder_output_new, attention_out_new])
# A dense softmax layer to generate prob dist. over the target vocabulary
decoder_output_final = decoder_dense(concat_layer_final)
# Final decoder model
decoder_model = Model(
[decoder_input] + [decoder_previous_output, decoder_previous_state_h, decoder_previous_state_c],
[decoder_output_final] + [state_h_new, state_c_new])
# + colab_type="code" id="RW8qxy6cpK0c" colab={}
# saving encoder_model and decoder_model for later use
encoder_model.save(dir_path + '/models/encoder_model.h5')
decoder_model.save(dir_path + '/models/decoder_model.h5')
# + colab_type="code" id="pPS8uc1JjnLq" colab={}
# Summarizes an input sequence to the final sequence by making the encoder-decoder model
# predict 1 word at a time
def summarizer(input_seq):
encoder_out, encoder_h, encoder_c = encoder_model.predict(input_seq)
target_seq = np.zeros((1,1))
target_seq[0, 0] = word_to_index_summary['stok']
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict([target_seq] + [encoder_out, encoder_h, encoder_c])
# Greedy decoder
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_token = index_to_word_summary[sampled_token_index]
if sampled_token != 'etok':
decoded_sentence += sampled_token + ' '
if sampled_token == 'etok' or len(decoded_sentence.split()) >= (_MAX_SUMMARY_LEN-1):
stop_condition = True
target_seq = np.zeros((1,1))
target_seq[0, 0] = sampled_token_index
encoder_h, encoder_c = h, c
return decoded_sentence
# + colab_type="code" id="Ymmhhqz3jnIK" colab={}
def sequence_to_summary(input_seq):
s = ''
for i in input_seq:
if i != 0 and i != word_to_index_summary['stok'] and i != word_to_index_summary['etok']:
s += index_to_word_summary[i] + ' '
return s
def sequence_to_text(input_seq):
s = ''
for i in input_seq:
if i != 0:
s += index_to_word_text[i] + ' '
return s
# + [markdown] colab_type="text" id="KCzEm7BCj5RV"
# Let's test it on a few examples from our dataset itself
# + [markdown] colab_type="text" id="VaKCIdtZnNWM"
# First from the training dataset
# + colab_type="code" id="qMsmXxMWnJ2g" outputId="e1e9896a-93ca-4d09-a2f0-0af8c8488450" colab={"base_uri": "https://localhost:8080/", "height": 462}
for i in range(5):
print("Review:",sequence_to_text(x_tr[i]))
print("Original summary:",sequence_to_summary(y_tr[i]))
print("Predicted summary:",summarizer(x_tr[i].reshape(1,_MAX_TEXT_LEN)))
print("\n")
# + [markdown] colab_type="text" id="ZC7vVJC6nSvl"
# Now from the validation dataset
# + colab_type="code" id="xKsC2ch0nMGu" outputId="aa080095-2541-4b31-de24-aec903f18f81" colab={"base_uri": "https://localhost:8080/", "height": 462}
for i in range(5):
print("Review:",sequence_to_text(x_val[i]))
print("Original summary:",sequence_to_summary(y_val[i]))
print("Predicted summary:",summarizer(x_val[i].reshape(1,_MAX_TEXT_LEN)))
print("\n")
# + [markdown] colab_type="text" id="012DWqsMkEiu"
# So far so good, let's now build a wrapper function to try out any other sentence
# + colab_type="code" id="Pqbf_mIEHzmj" colab={}
def tryit(sent):
"""wrapper function to test the model"""
sent = cleaner(sent, remove_stopwords=True)
if len(sent.split()) > _MAX_TEXT_LEN:
return "make your sentence length less than {} words".format(_MAX_TEXT_LEN)
seq = tok_x.texts_to_sequences(sent.split())
seq = [[item for sublist in seq for item in sublist]]
seq = pad_sequences(seq, maxlen=_MAX_TEXT_LEN, padding=_TEXT_PADDING)
return summarizer(seq.reshape(1,_MAX_TEXT_LEN))
# + [markdown] colab_type="text" id="ZLaO9MsWkcRn"
# Next comes to trying it out on a sentence. Let's see how it performs on the below text.
# + colab_type="code" id="fMiIqP28Hzjg" colab={}
text = 'We tried the Megna for the first time last night and the food was great, freshly cooked very tasty and well presented. \
The waiters were attentive and service overall was good. The restaurant looked to have been recently decorated. \
Overall a good meal and great value we will certainly be going back.'
real_summary = 'tried megna good food attentive waiters'
# + colab_type="code" id="MAoIGlJrnsP7" outputId="cdba65f3-9d45-4f9a-e9e8-5dd282de4f18" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit(text)
# + [markdown] colab_type="text" id="bRKQzq9Lksgx"
# Not bad, since most of the amazon's dataset had similar summaries for such reviews. Now let's split each sentence and try that out.
# + colab_type="code" id="kRuwwHeCnyTZ" outputId="ed68c2b2-dc60-4c1d-fa96-2a06c4e8208a" colab={"base_uri": "https://localhost:8080/", "height": 289}
text1 = text.split('.')
for l in text1:
if len(l) > 0:
print('text :', l.strip())
print('predict:', tryit(l))
print('\n')
# + [markdown] colab_type="text" id="feQDev2GlA9H"
# It draws out the meaning atleast. Now let's try it out on more sentences.
# + colab_type="code" id="ZPYeUFD0HyzK" outputId="b23dced2-7747-4942-edbf-22343d7d2956" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('my dog was delighted to eat meat chunks')
# + colab_type="code" id="3FZQ4bb8UR1x" outputId="3957fce0-1a06-45aa-e4c1-92925800aaab" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('the food was terribly good')
# + colab_type="code" id="kczUr3uWou2m" outputId="155611a4-8836-4b35-f101-8d0396f4a604" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('the food was terrible')
# + colab_type="code" id="q1KlNY-CUR0D" outputId="e3cf63d2-e1c6-4490-deb3-3160953b06d9" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('the food was good but bad')
# + colab_type="code" id="_ifYomnvURvC" outputId="7d4516b3-b4f2-4eab-bfb0-42b8428483d0" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('the food was bad at first but finally it turned out to be otherwise')
# + colab_type="code" id="jdV1RCo8ekx7" outputId="c9544f2d-1640-4952-cebe-dc7671465c7a" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('I love maggi but couldn\'t find any')
# + colab_type="code" id="Rvw81IT6erNq" outputId="e6dbf0aa-5e1b-4034-bbee-587806332067" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('well to be honest the customer service is not good')
# + colab_type="code" id="nF74C0cOewjR" outputId="5f433c1e-398c-4f3d-be80-6df41e565b83" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('bad food')
# + colab_type="code" id="iY6WOjYFey5Y" outputId="c6dcd507-c0dc-45f0-9e39-1b41a3bb5094" colab={"base_uri": "https://localhost:8080/", "height": 34}
tryit('well the food was not that bad but certainly can be improved')
# + [markdown] colab_type="text" id="zy7bbFN6lbh1"
# Well, its working quite well except for a few sentences which has a big deeper meaning or usage of both words such as 'good' and 'bad', although it gave correct answer for second sentence despite having both 'terrible' and 'good' used together.
# + [markdown] colab_type="text" id="lwFo2G_3l7zI"
# ## Conclusion
# + [markdown] colab_type="text" id="l-ThoRKHmDel"
# The Model is working out well but it has some flaws as mentioned above. More improvements can be done to ensure better accuracy.
| Text_Summarization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# PDBe API Training
# =========
#
# This interactive Python notebook will guide you through various ways of programmatically accessing Protein Data Bank in Europe (PDBe) data using REST API
#
# The REST API is a programmatic way to obtain information from the PDB and EMDB. You can access details about:
#
# * sample
# * experiment
# * models
# * compounds
# * cross-references
# * publications
# * quality
# * assemblies
# * and more...
#
# For more information, visit http://www.ebi.ac.uk/pdbe/pdbe-rest-api
# # Notebook #2
#
# This notebook is the second in the training material series, and focuses on getting secondary structure mappings for PDB entries using the REST API of PDBe.
# ## 1) Making imports and setting variables
#
# First, we import some packages that we will use, and set some variables.
#
# Note: Full list of valid URLs is available from http://www.ebi.ac.uk/pdbe/api/doc/
# +
import re
import requests
base_url = "https://www.ebi.ac.uk/pdbe/"
api_base = base_url + "api/"
secondary_structure_url = api_base + 'pdb/entry/secondary_structure/'
# -
# ## 2) Defining request function
#
# Let's start with defining a function that can be used to GET a single PDB entry, or POST a comma-separated list of PDB entries.
#
# We will use this function to retrieve secondary structure mapping for entries.
def make_request(url, mode, pdb_id):
"""
This function can make GET and POST requests to
the PDBe API
:param url: String,
:param mode: String,
:param pdb_id: String
:return: JSON or None
"""
if mode == "get":
response = requests.get(url=url+pdb_id)
elif mode == "post":
response = requests.post(url, data=pdb_id)
if response.status_code == 200:
return response.json()
else:
print("[No data retrieved - %s] %s" % (response.status_code, response.text))
return None
# ## 3) Defining function for extracting secondary structure mapping
#
# Next, we will define a function that can be used to retrieve secondary structural element ranges for PDB entries, and extracts this information so that it can be displayed in a user-friendly way.
#
# The function will rely on the make_request() function we have defined previously.
#
# This new function should either accept a single PDB id, or a list of PDB ids, and make a GET or a POST call to the API accordingly. The data structure then has to be traversed, and the residue ranges of helices and strands have to be recorded. Since the data is in a nested JSON format, (for the sake of not touching on more advanced Python topics) we will use nested for-loops to get to the level of interest.
#
# If you are wondering how the complete JSON looks like, follow this link:
# https://www.ebi.ac.uk/pdbe/api/pdb/entry/secondary_structure/1cbs
def get_secondary_structure_ranges(pdb_id=None, pdb_list=None):
"""
This function calls the PDBe API and retrieves the residue
ranges of secondary structural elements in a single PDB entry
or in a list of PDB entries
:param pdb_id: String,
:param pdb_list: String
:return: None
"""
# If neither a single PDB id, nor a list was provided,
# exit the function
if not pdb_id and not pdb_list:
print("Either provide one PDB id, or a list of ids")
return None
if pdb_id:
# If a single PDB id was provided, call the API with GET
data = make_request(secondary_structure_url, "get", pdb_id)
else:
# If multiple PDB ids were provided, call the API with POST
# The POST API call expects PDB ids as a comma-separated lise
pdb_list_string = ", ".join(pdb_list)
data = make_request(secondary_structure_url, "post", pdb_list_string)
# When no data is returned by the API, exit the function
if not data:
print("No data available")
return None
# Loop through all the PDB entries in the retrieved data
for entry_id in data.keys():
entry = data[entry_id]
molecules = entry["molecules"]
# Loop through all the molecules of a given PDB entry
for i in range(len(molecules)):
chains = molecules[i]["chains"]
# Loop through all the chains of a given molecules
for j in range(len(chains)):
secondary_structure = chains[j]["secondary_structure"]
helices = secondary_structure["helices"]
strands = secondary_structure["strands"]
helix_list = []
strand_list = []
# Loop through all the helices of a given chain
for k in range(len(helices)):
start = helices[k]["start"]["residue_number"]
end = helices[k]["end"]["residue_number"]
helix_list.append("%s-%s" % (start, end))
# Loop through all the strands of a given chain
for l in range(len(strands)):
start = strands[l]["start"]["residue_number"]
end = strands[l]["end"]["residue_number"]
strand_list.append("%s-%s" % (start, end))
report = "%s chain %s has " % (entry_id, chains[j]["chain_id"])
if len(helix_list) > 0:
report += "helices at residue ranges %s " % str(helix_list)
else:
report += "no helices "
report += "and "
if len(strand_list) > 0:
report += "strands at %s" % str(strand_list)
else:
"no strands"
print(report)
return None
# Let's try our new function first with a single PDB entry (1cbs), and then with a list of two entries (2aqa and 2klm)
print("Example of a single PDB entry")
get_secondary_structure_ranges(pdb_id="1cbs")
print()
print("Example of multiple PDB entries")
get_secondary_structure_ranges(pdb_list=["2aqa", "2klm"])
# ## This ends the second notebook - please proceed to other notebooks of your interest
# Copyright 2018 EMBL - European Bioinformatics Institute
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| API workshop #2 - Secondary structure mapping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # 统计
# - `np.median`中位数
# + pycharm={"is_executing": false}
import numpy as np
a = np.arange(10)
np.median(a)
# + [markdown] pycharm={"name": "#%% md\n"}
# - `np.average` 加权平均值
# + pycharm={"name": "#%%\n", "is_executing": false}
np.average(a)
# + [markdown] pycharm={"name": "#%% md\n"}
# - `np.mean` 算术平均值
# + pycharm={"name": "#%%\n", "is_executing": false}
np.mean(a)
# + [markdown] pycharm={"name": "#%% md\n"}
# - `np.std`标准差
# + pycharm={"name": "#%% \n", "is_executing": false}
np.std(a)
# + [markdown] pycharm={"name": "#%% md\n"}
# - `np.var`方差
# + pycharm={"name": "#%% \n", "is_executing": false}
np.var(a)
# + [markdown] pycharm={"name": "#%% md\n"}
# - 轴向中位数
# + pycharm={"name": "#%%\n", "is_executing": false}
a = np.array([
[1,2],
[2,3],
])
np.nanmedian(a,axis=0)
# + [markdown] pycharm={"name": "#%% md\n"}
# - `np.amin`最大值(计算nan)
# - `np.amax`最小值(计算nan)
# - `np.nanmin`最大值(不计算nan)
# - `np.nanmax`最小值(不计算nan)
# + pycharm={"name": "#%%\n", "is_executing": false}
a = np.arange(4).reshape((2,2))
np.amin(a,axis=0)
# + pycharm={"name": "#%% \n", "is_executing": false}
np.amin(a)
# + pycharm={"name": "#%%\n", "is_executing": false}
np.amax(a)
# + pycharm={"name": "#%%\n", "is_executing": false}
np.amax(a,axis=1)
| doc/numpy/statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Average Daily Users
# ### Imports
from datetime import datetime
import os
import pandas as pd
import numpy as np
# ### Initializations
# Verify file path to trax data is correct
trafx_data_file_path = os.path.abspath(os.path.join(os.getcwd(), "..", "data", "raw", "TRAFx_raw.csv"))
# ### Load the trax data
trafx_df = pd.read_csv(trafx_data_file_path)
trafx_df.head(5)
# ### Clean the data
# +
# Replace any zeros with nan
trafx_df.replace(0, np.nan, inplace=True)
# Convert Day to datetime
trafx_df['Day'] = trafx_df['Day'].apply(lambda x: datetime.strptime(x, "%Y-%m-%d"))
# -
# ### Append information to main df
# *_Note: season is only Dec 1 - Apri 30, all other data excluded_*
# +
# Append season, month, and day of week on to main dataframe
trafx_df['month'] = trafx_df['Day'].apply(lambda x: x.strftime('%b'))
trafx_df['dayOfWeek'] = trafx_df['Day'].apply(lambda x: x.strftime('%A'))
def set_season(date_time):
if date_time.month == 12:
return "{}-{}".format(date_time.year, date_time.year+1)
elif date_time.month <=4:
return "{}-{}".format(date_time.year-1, date_time.year)
else:
return 'None'
trafx_df['season'] = trafx_df['Day'].apply(lambda x: set_season(x))
trafx_df = trafx_df[trafx_df['season'] != 'None']
trafx_df.head(5)
# -
# ### Group data by month and day and determine daily average users
# +
adu_df = pd.DataFrame(columns=['site', 'season', 'month', 'dayOfWeek', 'n', 'adu'])
# Get monthly averages
ms_adu_val = trafx_df.groupby(['season', 'month']).mean()
ms_adu_n = trafx_df.groupby(['season', 'month']).count()
for season, month in ms_adu_val.index.to_list():
for site in ms_adu_val.columns:
if ms_adu_n.loc[season, month][site] > 0:
adu_df.loc[len(adu_df)] = [site, season, month, 'All', ms_adu_n.loc[season, month][site], ms_adu_val.loc[season, month][site]]
adu_df.head(5)
# +
# Get daily averages
msd_adu_val = trafx_df.groupby(['season', 'month', 'dayOfWeek']).mean()
msd_adu_n = trafx_df.groupby(['season', 'month', 'dayOfWeek']).count()
for season, month, dow in msd_adu_val.index.to_list():
for site in ms_adu_val.columns:
if msd_adu_n.loc[season, month, dow][site] > 0:
adu_df.loc[len(adu_df)] = [site, season, month, dow, msd_adu_n.loc[season, month, dow][site], msd_adu_val.loc[season, month, dow][site]]
adu_df.head(5)
# -
# ### Save data to file
# This assumes we're using the cookiecutter directory structure with notebooks at the same directory
# level as data with a raw as a subdirectory of data
file_path = os.path.abspath(os.path.join(os.getcwd(), "..", "data", "processed", "daily_user_averages.csv"))
adu_df.to_csv(file_path, index=False)
| wba-trailcounting/notebooks/2.0-maf-average-daily-users.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automated Hyperparameter Optimization Training using WMLA API
#
#
#
#
#
#
#
# In this notebook, you will learn how to submit a model and dataset to the Watson Machine Learning Accelerator (WMLA) API to run Hyper Parameter Optimization (HPO). In this particular example, we will be using the Pytorch MNIST HPO model as our training model, inject hyperparameters for the sub-training during search and submit a tuning metric for better results, and then query for the best job results. This notebook runs on Python 3.6.
#
#
# 
#
#
# 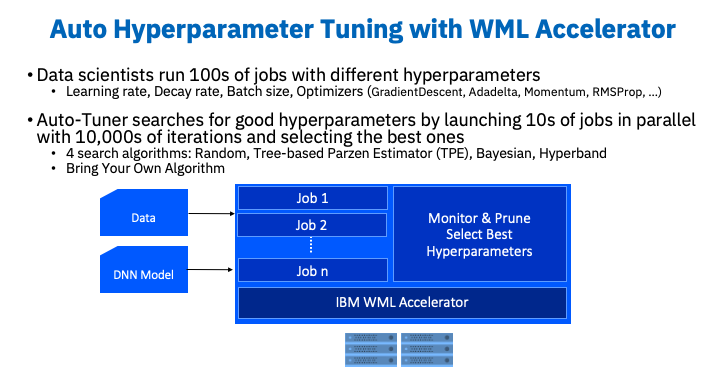
#
#
# For this notebook you will use a model and dataset that have already been set up to leverage the API. For details on the API see [API Documentation](https://www.ibm.com/support/knowledgecenter/en/SSFHA8_1.2.1/cm/deeplearning.html) in the Knowledge Center (KC).
# ## Table of contents
#
# 1. [Setup](#setup)<br>
#
# 2. [Configuring environment and project details](#configure)<br>
#
# 3. [Health Check](#health)<br>
#
# 4. [Training with the HPO API](#train)<br>
#
# 5. [Deploy the HPO task](#deploy)<br>
#
# 6. [Find best job results](#best)<br>
# <a id = "setup"></a>
# ## Step 1: Setup
#
# TODO : suggest we delete this, and ask user to open other notebook with Pytorch code ....
# First, we must import the required modules. Here we will import the Pytorch MNIST HPO model.
# !wget https://github.com/IBM/wmla-learning-path/raw/master/datasets/pytorch-mnist-hpo.modelDir.tar
# To use the WMLA API, we will be using the Python requests library.
# +
import requests
from requests.packages.urllib3.exceptions import InsecureRequestWarning
requests.packages.urllib3.disable_warnings(InsecureRequestWarning)
import json
import time
import urllib
import pandas as pd
import os,sys
import tarfile
import tempfile
from IPython.display import clear_output
import time
import pprint
# utility print function
def nprint(mystring) :
print("**{}** : {}".format(sys._getframe(1).f_code.co_name,mystring))
# utility makedir
def makeDirIfNotExist(directory) :
if not os.path.exists(directory):
nprint("Making directory {}".format(directory))
os.makedirs(directory)
else :
nprint("Directory {} already exists .. ".format(directory))
# -
# <a id = "configure"></a>
# ## Step 2: Configuring environment and project details
# Provide your credentials in this cell, including your cluster url, username and password, and instance group.
# +
def getconfig(cfg_in={}):
cfg = {}
cfg["master_host"] = '' # <=enter your host url here
cfg["dli_rest_port"] = ''
cfg["sc_rest_port"] = ''
cfg["num_images"] = {"train":200,"valid":20,"test":20}
# ==== CLASS ENTER User login details below =====
cfg["wmla_user"] = '' # <=enter your id here
cfg["wmla_pwd"] = '' # <=enter your pwd here
# ==== CLASS ENTER User login details above =====
cfg["sig_name"] = '' # <=enter instance group here
cfg["code_dir"] = "/home/wsuser/works/pytorch_hpo"
# overwrite configs if passed
for (k,v) in cfg_in.items() :
nprint("Overriding Config {}:{} with {}".format(k,cfg[k],v))
cfg[k] = v
return cfg
# cfg is used as a global variable throughout this notebook
cfg=getconfig()
# -
# Here we will get and print out the API endpoints and setup requests session. The following sections use the Watson ML Accelerator API to complete the various tasks required. We've given examples of a number of tasks but you should refer to the documentation at to see more details of what is possible and sample output you might expect.
#
# - https://www.ibm.com/support/knowledgecenter/SSFHA8_1.2.2/cm/deeplearning.html
# - https://www.ibm.com/support/knowledgecenter/SSZU2E_2.4.1/reference_s/api_references.html
#
#
# +
# REST call variables
commonHeaders = {'Accept': 'application/json'}
# Use closures for cfg for now ..
def get_tmp_dir() :
#return "/gpfs/home/s4s004/"+cfg["wmla_user"]+"/2020-05-wmla/tmp"
return "/home/wsuser/work"
def get_tar_file() :
#return get_tmp_dir() + "/" + cfg["wmla_user"]+".modelDir.tar"
return get_tmp_dir() + "/pytorch-mnist-hpo.modelDir.tar"
#get api endpoint
def get_ep(mode="sc") :
if mode=="sc" :
sc_rest_url = cfg["master_host"] +':'+ cfg["sc_rest_port"] +'/platform/rest/conductor/v1'
return sc_rest_url
elif(mode=="dl") :
dl_rest_url = cfg["master_host"] +':'+cfg["dli_rest_port"] +'/platform/rest/deeplearning/v1'
return dl_rest_url
else :
nprint("Error mode : {} not supported".format(mode))
def myauth():
return(cfg["wmla_user"],cfg["wmla_pwd"])
print ("SC API Endpoints : {}".format(get_ep("sc")))
print ("DL API Endpoints : {}".format(get_ep("dl")))
print (myauth())
print (get_tar_file())
#myauth = (wmla_user, wmla_pwd)
# Setup Requests session
req = requests.Session()
# -
# <a id = "health"></a>
# ## Step 3: Health Check
# In this step, we will check if there are any existing HPO tasks and also verify the platform health.
# Rest API: `GET platform/rest/deeplearning/v1/hypersearch`
# - `Description`: Get all the HPO tasks that the login user can access.
# - `OUTPUT`: A list of HPO tasks and each one with the same format which can be found in the api doc.
# +
def hpo_health_check():
getTuneStatusUrl = get_ep("dl") + '/hypersearch'
nprint ('getTuneStatusUrl: %s' %getTuneStatusUrl)
r = req.get(getTuneStatusUrl, headers=commonHeaders, verify=False, auth=myauth())
if not r.ok:
nprint('check hpo task status failed: code=%s, %s'%(r.status_code, r.content))
else:
if len(r.json()) == 0:
nprint('There is no hpo task been created')
for item in r.json():
nprint('Hpo task: %s, State: %s'%(item['hpoName'], item['state']))
#print('Best:%s'%json.dumps(item.get('best'), sort_keys=True, indent=4))
hpo_health_check()
# -
# <a id = "train"></a>
# ## Step 4: Training with the HPO API
#
# TODO : Now we could reference other notebook, or even put this description over there ...
#
#
# The WMLA framework requires 2 changes to your code to support the HPO API, and these are:
#
# * Inject hyperparameters for the sub-training during search
# * Retrieve sub-training result metric
#
# Note that the code sections below show a comparison between the "before" and "HPO enabled" versions of the code by using `diff`.
#
#
# 1. Import the dependent libararies:
#
#
#
# 
#
#
#
# 2. Get the WMLA cluster `DLI_DATA_FS`, `RESULT_DIR` and `LOG_DIR` for the HPO training job. The `DLI_DATA_FS` can be used for shared data placement, the `RESULT_DIR` can be used for final model saving, and the `LOG_DIR` can be used for user logs and monitoring.
#
#
# **Note**: `DLI_DATA_FS` is set when installing the DLI cluster; `RESULT_DIR` and `LOG_DIR` is generated by WMLA for each HPO experiment.
#
#
#
# 
#
#
#
# 3. Replace the hyperparameter definition code by reading hyperparameters from the `config.json` file. the `config.json` is generated by WMLA HPO, which contains a set of hyperparameter candidates for each tuning jobs. The hyperparameters and the search space is defined when submitting the HPO task. For example, here the hyperparameter `learning_rate` is set to tune:
#
#
#
# 
#
#
# Then you could use the hyperparameter you get from `config.json` where you want:
#
# 
#
#
#
# 4. Write the tuning result into `val_dict_list.json` under `RESULT_DIR`. WMLA HPO will read this file for each tuning job to get the metric values. Define a `test_metrics` list to store all metric values and pass the epoch parameter to the test function. Then you can add the metric values to the `test_metrics` list during the training test process. Please note that the metric names should be specified when submitting the HPO task, and be consistent with the code here.
#
# For example, at the HPO task submit request, `loss` will be used as the objective metric the tuning will try to minimize the `loss`:
#
# ```
# 'algoDef': # Define the parameters for search algorithms
# {
# # Name of the search algorithm, one of Random, Bayesian, Tpe, Hyperband
# 'algorithm': 'Random',
# # Name of the target metric that we are trying to optimize when searching hyper-parameters.
# # It is the same metric name that the model update part 2 trying to dump.
# 'objectiveMetric' : 'loss',
# # Strategy as how to optimize the hyper-parameters, minimize means to find better hyper-parameters to
# # make the above objectiveMetric as small as possible, maximize means the opposite.
# 'objective' : 'minimize',
# ...
# }
# ```
#
# The code change:
#
#
#
# 
#
#
#
# 5. After the training completes, write the metric list into the `val_dict_list.json` file.
#
#
# 
#
#
#
#
# ## NBDEV
# <a id = "deploy"></a>
# ## Step 5: Deploy the HPO task
# Here we package up our model to send to the API for HPO.
#
#
#
# REST API: `POST /platform/rest/deeplearning/v1/hypersearch`
#
# - Description: Start a new HPO task
# - Content-type: Multi-Form
# - Multi-Form Data:
# - files: Model files tar package, ending with `.modelDir.tar`
# - form-filed: {‘data’: ‘String format of input parameters to start hpo task, let’s call it as **hpo_input** and show its specification later’}
#
# #### Package model files for training
# Package the updated model files into a tar file ending with `.modelDir.tar`
#
# REST API expects a `modelDir.tar` with the model code inside ..
#
files = {'file': open(get_tar_file(), 'rb')}
print("Files : {}".format(files))
# #### Construct POST request data
# **hpo_input** will be in Python `dict` or `json` format as shown below, and will convert to string when calling REST.
# Note, this
data = {
'modelSpec': # Define the model training related parameters
{
# Spark instance group which will be used to run the HPO sub-trainings. The Spark instance group selected
# here should match the sub-training args, for example, if the sub-training args try to run a EDT job,
# then we should put a Spark instance group with capability to run EDT job here.
'sigName': cfg["sig_name"],
# These are the arguments we'll pass to the execution engine; they follow the same conventions
# of the dlicmd.py command line launcher
#
# See:
# https://www.ibm.com/support/knowledgecenter/en/SSFHA8_1.2.1/cm/dlicmd.html
# In this example, args after --model-dir are all the required parameter for the original model itself.
#
'args': '--exec-start PyTorch --cs-datastore-meta type=fs --python-version 3.6\
--gpuPerWorker 1 --model-main pytorch_mnist_HPO.py --model-dir pytorch_hpo\
--debug-level debug'
},
'algoDef': # Define the parameters for search algorithms
{
# Name of the search algorithm, one of Random, Bayesian, Tpe, Hyperband, ExperimentGridSearch
'algorithm': 'Random',
# Max running time of the hpo task in minutes, -1 means unlimited
'maxRunTime': 60,
# Max number of training job to submitted for hpo task, -1 means unlimited’,
'maxJobNum': 4,
# Max number of training job to run in parallel, default 1. It depends on both the
# avaiable resource and if the search algorithm support to run in parallel, current only Random
# fully supports to run in parallel, Hyperband and Tpe supports to to in parellel in some phase,
# Bayesian runs in sequence now.
'maxParalleJobNum': 4,
# Name of the target metric that we are trying to optimize when searching hyper-parameters.
# It is the same metric name that the model update part 2 trying to dump.
'objectiveMetric' : 'loss',
# Strategy as how to optimize the hyper-parameters, minimize means to find better hyper-parameters to
# make the above objectiveMetric as small as possible, maximize means the opposite.
'objective' : 'minimize',
},
# Define the hyper-paremeters to search and the corresponding search space.
'hyperParams':
[
{
# Hyperparameter name, which will be the hyper-parameter key in config.json
'name': 'learning_rate',
# One of Range, Discrete
'type': 'Range',
# one of int, double, str
'dataType': 'DOUBLE',
# lower bound and upper bound when type=range and dataType=double
'minDbVal': 0.001,
'maxDbVal': 0.1,
# lower bound and upper bound when type=range and dataType=int
'minIntVal': 0,
'maxIntVal': 0,
# Discrete value list when type=discrete
'discreteDbVal': [],
'discreteIntVal': [],
'discreateStrVal': []
#step size to split the Range space. ONLY valid when type is Range
#'step': '0.002',
}
]
}
mydata={'data':json.dumps(data)}
# #### Submit the Post request
# Submit the HPO task through the Post call and an HPO name/id in string format will be returned.
# <div class="alert alert-block alert-warning">Note: This cannot be submitted twice. You need to rebuild the tar file prior to resubmitting.</div>
# +
def submit_job():
startTuneUrl=get_ep('dl') + '/hypersearch'
nprint("startTuneUrl : {}".format(startTuneUrl))
nprint("files : {}".format(files))
nprint("myauth() : {}".format(myauth()))
#print("hpo_job_id : {}".format(hpo_job_id))
r = req.post(startTuneUrl, headers=commonHeaders, data=mydata, files=files, verify=False, auth=myauth())
hpo_name=None
if r.ok:
hpo_name = r.json()
print ('\nModel submitted successfully: {}'.format(hpo_name))
else:
print('\nModel submission failed with code={}, {}'. format(r.status_code, r.content))
return hpo_name
hpo_job_id = submit_job()
print("hpo_job_id : {}".format(hpo_job_id))
# -
# Print out task details here.
# +
getHpoUrl = get_ep('dl') +'/hypersearch/'+ hpo_job_id
pp = pprint.PrettyPrinter(indent=2)
keep_running=True
rr=10
res=None
while(keep_running):
res = req.get(getHpoUrl, headers=commonHeaders, verify=False, auth=myauth())
experiments=res.json()['experiments']
experiments = pd.DataFrame.from_dict(experiments)
pd.set_option('max_colwidth', 120)
clear_output()
print("Refreshing every {} seconds".format(rr))
display(experiments)
pp.pprint(res.json())
if(res.json()['state'] not in ['SUBMITTED','RUNNING']) :
keep_running=False
time.sleep(rr)
# -
# <a id = "best"></a>
# ## Step 6: See best job results
# +
# Lets query our result to see what happened during HPO training!
#res.ok
#res.json()
#print(type(res))
#print(dir(res))
#print(json.dumps(res.json(), indent=4, sort_keys=True))
print('Hpo task %s completes with state %s'%(hpo_job_id, res.json()['state']))
print("Best HPO result ...")
res.json()["best"]
# -
# #### Notebook Complete
# Congratulations, you have completed our demonstration of using WMLA for distributed hyperparameter optimization search
# Copyright © 2020 IBM. This notebook and its source code are released under the terms of the MIT License.
# <div style="background:#F5F7FA; height:110px; padding: 2em; font-size:14px;">
# <span style="font-size:18px;color:#152935;">Love this notebook? </span>
# <span style="font-size:15px;color:#152935;float:right;margin-right:40px;">Don't have an account yet?</span><br>
# <span style="color:#5A6872;">Share it with your colleagues and help them discover the power of Watson Studio!</span>
# <span style="border: 1px solid #3d70b2;padding:8px;float:right;margin-right:40px; color:#3d70b2;"><a href="https://ibm.co/wsnotebooks" target="_blank" style="color: #3d70b2;text-decoration: none;">Sign Up</a></span><br>
# </div>
| dli-learning-path/tutorials-cpd-wmla/05-wmla-api-submitting-hyperparameter-optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB
from sklearn import metrics
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix,accuracy_score
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt
import seaborn as sns
url='https://raw.githubusercontent.com/jinnatul/ml-Pro/master/data/data-diabetes.csv?token=<KEY>'
df = pd.read_csv(url)
df.head()
# -
features=['glucose','bloodpressure']
x=df[features]
y=df.diabetes
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.25, random_state=5)
model=GaussianNB()
model.fit(x_train,y_train)
predictions = model.predict(x_test)
print (predictions)
# +
# Model performance Report
print('Classification Report:\n',classification_report(y_test, predictions))
print('Confusion Matrix:\n',confusion_matrix(y_test, predictions))
print('Accuracy Score:',accuracy_score(y_test, predictions))
print("Precision:",metrics.precision_score(y_test, predictions))
print("Recall:",metrics.recall_score(y_test,predictions ))
# -
new_observation = [[40,65]]
model.predict(new_observation)
new_observation = [[35,85]]
model.predict(new_observation)
new_observation = [[42,55]]
model.predict(new_observation)
| blogPractices/Supervised-Learning/naveBeige.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print("Hello Anaconda")
# + pycharm={"is_executing": false}
"""
Please note, this script is for python3+.
If you are using python2+, please modify it accordingly.
Tutorial reference:
http://www.scipy-lectures.org/intro/matplotlib/matplotlib.html
"""
import matplotlib.pyplot as plt
import numpy as np
n = 12
X = np.arange(n)
Y1 = (1 - X / float(n)) * np.random.uniform(0.5, 1.0, n)
Y2 = (1 - X / float(n)) * np.random.uniform(0.5, 1.0, n)
plt.bar(X, +Y1, facecolor='#9999ff', edgecolor='white')
plt.bar(X, -Y2, facecolor='#ff9999', edgecolor='white')
for x, y in zip(X, Y1):
# ha: horizontal alignment
# va: vertical alignment
plt.text(x + 0.4, y + 0.05, '%.2f' % y, ha='center', va='bottom')
for x, y in zip(X, Y2):
# ha: horizontal alignment
# va: vertical alignment
plt.text(x + 0.4, -y - 0.05, '%.2f' % y, ha='center', va='top')
plt.xlim(-.5, n)
plt.xticks(())
plt.ylim(-1.25, 1.25)
plt.yticks(())
plt.show()
# +
"""
Please note, this script is for python3+.
If you are using python2+, please modify it accordingly.
Tutorial reference:
http://www.scipy-lectures.org/intro/matplotlib/matplotlib.html
"""
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(-3, 3, 50)
y1 = 2*x + 1
y2 = x**2
plt.figure(figsize=(8, 5))
plt.plot(x, y2)
# plot the second curve in this figure with certain parameters
plt.plot(x, y1, color='red', linewidth=1.0, linestyle='--')
plt.show()
# +
"""
Please note, this script is for python3+.
If you are using python2+, please modify it accordingly.
Tutorial reference:
http://www.scipy-lectures.org/intro/matplotlib/matplotlib.html
"""
import matplotlib.pyplot as plt
import numpy as np
X = np.linspace(-3, 3, 50)
Y = 2 * X + 1
T = Y # for color later on
plt.scatter(X, Y, c=T)
plt.xlim(-1.5, 1.5)
plt.xticks(()) # ignore xticks
plt.ylim(-1.5, 1.5)
plt.yticks(()) # ignore yticks
plt.show()
# +
"""
Please note, this script is for python3+.
If you are using python2+, please modify it accordingly.
"""
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(-1, 1, 50)
y = 2*x + 1
# y = x**2
plt.plot(x, y)
plt.show()
# -
| ai-engineer/jupyter/aura/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="data/photutils_banner.svg" width=500 alt="Photutils logo">
# + [markdown] slideshow={"slide_type": "slide"}
# # Photutils
#
# - Code: https://github.com/astropy/photutils
# - Documentation: https://photutils.readthedocs.io/en/stable/
# - Issue Tracker: https://github.com/astropy/photutils/issues
#
# ## Photutils can be used for:
#
# - Background and background noise estimation
# - Source Detection and Extraction
# - DAOFIND and IRAF's starfind
# - Image segmentation
# - local peak finder
# - Aperture photometry
# - PSF photometry
# - ePSF building
# - PSF matching
# - Centroids
# - Morphological properties
# - Elliptical isophote analysis
#
#
# ## In this section, we will:
#
# - Learn how to perform aperture photometry
# - Learn how to use photutils' image segmentation subpackage
#
# ---
# -
# ## Preliminaries
# + slideshow={"slide_type": "slide"}
# Initial imports
import numpy as np
import matplotlib.pyplot as plt
# Change some default plotting parameters
import matplotlib as mpl
mpl.rcParams['image.origin'] = 'lower'
mpl.rcParams['image.interpolation'] = 'nearest'
# Run the %matplotlib magic command to enable inline plotting
# in the current notebook. Choose one of these:
# %matplotlib inline
# # %matplotlib notebook
# -
# ### Load the data
# + [markdown] slideshow={"slide_type": "-"}
# We'll start by reading data and error arrays from FITS files. These are cutouts from the HST Extreme-Deep Field (XDF) taken with WFC3/IR in the F160W filter.
# -
from astropy.io import fits
# + slideshow={"slide_type": "-"}
sci_fn = 'data/xdf_hst_wfc3ir_60mas_f160w_sci.fits'
rms_fn = 'data/xdf_hst_wfc3ir_60mas_f160w_rms.fits'
sci_hdulist = fits.open(sci_fn)
rms_hdulist = fits.open(rms_fn)
sci_hdulist[0].header['BUNIT'] = 'electron/s'
# -
# Print some info about the data.
sci_hdulist.info()
# Define the data and error arrays.
# + slideshow={"slide_type": "fragment"}
data = sci_hdulist[0].data.astype(np.float)
error = rms_hdulist[0].data.astype(np.float)
# -
# Extract the data header and create a WCS object.
from astropy.wcs import WCS
# + slideshow={"slide_type": "fragment"}
hdr = sci_hdulist[0].header
wcs = WCS(hdr)
# -
# Display the data.
from astropy.visualization import simple_norm
# + slideshow={"slide_type": "fragment"}
plt.figure(figsize=(8, 8))
norm = simple_norm(data, 'sqrt', percent=99.)
plt.imshow(data, norm=norm)
plt.title('XDF F160W Cutout');
# + [markdown] slideshow={"slide_type": "slide"}
# ---
# # Part 1: Aperture Photometry
# -
# Photutils provides circular, elliptical, and rectangular aperture shapes (plus annulus versions of each). These are names of the aperture classes, defined in pixel coordinates:
#
# * `CircularAperture`
# * `CircularAnnulus`
#
# * `EllipticalAperture`
# * `EllipticalAnnulus`
#
# * `RectangularAperture`
# * `RectangularAnnulus`
#
# Along with variants of each, defined in celestial coordinates:
#
# * `SkyCircularAperture`
# * `SkyCircularAnnulus`
#
# * `SkyEllipticalAperture`
# * `SkyEllipticalAnnulus`
#
# * `SkyRectangularAperture`
# * `SkyRectangularAnnulus`
#
# These look something like this:
# <img src='data/apertures.png' alt='Figure of aperture shapes' width=700px>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Methods for handling aperture/pixel intersection
# -
# In general, the apertures will only partially overlap some of the pixels in the data.
#
# There are three methods for handling the aperture overlap with the pixel grid of the data array.
# <img src="data/photutils_aperture_methods.svg">
# NOTE: the `subpixels` keyword is ignored for the **'exact'** and **'center'** methods.
# ### Perform circular-aperture photometry on some sources in the XDF
# First, we define a circular aperture at a given position and radius (in pixels).
from photutils import CircularAperture
# + slideshow={"slide_type": "fragment"}
position = (90.73, 59.43) # (x, y) pixel position
radius = 5. # pixels
aperture = CircularAperture(position, r=radius)
# -
aperture
print(aperture)
# We can plot the aperture on the data using the aperture `plot()` method:
plt.figure(figsize=(8, 8))
plt.imshow(data, norm=norm)
aperture.plot(color='red', lw=2)
# Now let's perform photometry on the data using the `aperture_photometry()` function. **The default aperture method is 'exact'.**
#
# Also note that the input data is assumed to have zero background. If that is not the case, please see the documentation for the `photutils.background` subpackage for tools to help subtract the background.
#
# <div class="alert alert-warning alert-block">
# <h3 style="margin-top:0;">Learn More:</h3>
#
# See the [local background subtraction notebook](photutils_local_backgrounds.ipynb) for examples of local background subtraction.
# </div>
#
# The background was already subtracted for our XDF example data.
from photutils import aperture_photometry
# + slideshow={"slide_type": "fragment"}
phot = aperture_photometry(data, aperture)
phot
# -
# The output is an Astropy `QTable` (Quantity Table) with sum of data values within the aperture (using the defined pixel overlap method).
#
# The table also contains metadata, which is accessed by the `meta` attribute of the table. The metadata is stored as a python (ordered) dictionary:
phot.meta
phot.meta['version']
# Aperture photometry using the **'center'** method gives a slightly different (and less accurate) answer:
# + slideshow={"slide_type": "fragment"}
phot = aperture_photometry(data, aperture, method='center')
phot
# -
# Now perform aperture photometry using the **'subpixel'** method with `subpixels=5`:
#
# These parameters are equivalent to SExtractor aperture photometry.
# + slideshow={"slide_type": "fragment"}
phot = aperture_photometry(data, aperture, method='subpixel', subpixels=5)
phot
# -
# ## Photometric Errors
# We can also input an error array to get the photometric errors.
phot = aperture_photometry(data, aperture, error=error)
phot
# The error array in our XDF FITS file represents only the background error. If we want to include the Poisson error of the source we need to calculate the **total** error:
#
# $\sigma_{\mathrm{tot}} = \sqrt{\sigma_{\mathrm{b}}^2 +
# \frac{I}{g}}$
#
# where $\sigma_{\mathrm{b}}$ is the background-only error,
# $I$ are the data values, and $g$ is the "effective gain".
#
# The "effective gain" is the value (or an array if it's variable across an image) needed to convert the data image to count units (e.g. electrons or photons), where Poisson statistics apply.
#
# Photutils provides a `calc_total_error()` function to perform this calculation.
from photutils.utils import calc_total_error
# +
# this time include the Poisson error of the source
# our data array is in units of e-/s
# so the "effective gain" should be the exposure time
eff_gain = hdr['TEXPTIME']
tot_error = calc_total_error(data, error, eff_gain)
phot = aperture_photometry(data, aperture, error=tot_error)
phot
# -
# The total error increased only slightly because this is a small faint source.
# ## Units
# input the data units
import astropy.units as u
# `Quantity` inputs for data and error are also allowed. Note that the unit must be the same for the `data` and `error` inputs.
unit = u.electron / u.s # unit for the data and error arrays
phot = aperture_photometry(data * unit, aperture, error=tot_error * unit)
phot
# + [markdown] slideshow={"slide_type": "slide"}
# ## Performing aperture photometry at multiple positions
# -
# Now let's perform aperture photometry for three sources (all with the same aperture size). We simply define three (x, y) positions.
# + slideshow={"slide_type": "-"}
positions = [(90.73, 59.43), (73.63, 139.41), (43.62, 61.63)]
radius = 5.
apertures = CircularAperture(positions, r=radius)
# -
# An aperture object with multiple positions can be indexed or sliced to get a subset of apertures:
apertures[1] # aperture for the second position
apertures[0:2] # apertures for the first two positions
# Let's plot all three apertures on the data.
plt.figure(figsize=(8, 8))
plt.imshow(data, norm=norm)
apertures.plot(color='red', lw=2)
# Now let's perform aperture photometry.
# + slideshow={"slide_type": "-"}
phot = aperture_photometry(data * unit, apertures, error=tot_error * unit)
phot
# -
# Each source is a row in the table and is given a unique **id** (the first column).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Adding columns to the photometry table
# -
# We can add columns to the photometry table. Let's calculate the signal-to-noise (SNR) ratio of our sources and add it as a new column to the table.
# + slideshow={"slide_type": "fragment"}
snr = phot['aperture_sum'] / phot['aperture_sum_err'] # units will cancel
phot['snr'] = snr
phot
# -
# Now calculate the F160W AB magnitude and add it to the table.
# + slideshow={"slide_type": "fragment"}
f160w_zpt = 25.9463 # HST/WFC3 F160W zero point
# NOTE that the log10() function can be applied only to dimensionless quantities
# so we use the value() method to get the number value of the aperture sum
abmag = -2.5 * np.log10(phot['aperture_sum'].value) + f160w_zpt
phot['abmag'] = abmag
phot
# -
# Now, using the WCS defined above, calculate the sky coordinates for these objects and add it to the table.
# + slideshow={"slide_type": "fragment"}
# convert pixel positions to sky coordinates
x, y = np.transpose(positions)
coord = wcs.pixel_to_world(x, y)
# we can add the astropy SkyCoord object directly to the table
phot['sky_coord'] = coord
phot
# -
# We can also add separate RA and Dec columns, if preferred.
# + slideshow={"slide_type": "fragment"}
phot['ra_icrs'] = coord.icrs.ra
phot['dec_icrs'] = coord.icrs.dec
phot
# -
# If we write the table to an ASCII file using the ECSV format, we can read it back in preserving all of the units, metadata, and SkyCoord objects.
phot.write('my_photometry.txt', format='ascii.ecsv')
# view the table on disk
# !cat my_photometry.txt
# Now read the table in ECSV format.
from astropy.table import QTable
tbl = QTable.read('my_photometry.txt', format='ascii.ecsv')
tbl
tbl.meta
tbl['aperture_sum'] # Quantity array
tbl['sky_coord'] # SkyCoord array
# + [markdown] slideshow={"slide_type": "slide"}
# ## Aperture photometry using Sky apertures
# -
# First, let's define the sky coordinates by converting our pixel coordinates.
positions = [(90.73, 59.43), (73.63, 139.41), (43.62, 61.63)]
x, y = np.transpose(positions)
coord = wcs.pixel_to_world(x, y)
coord
# Now define circular apertures in sky coordinates.
#
# For sky apertures, the aperture radius must be a `Quantity`, in either pixel or angular units.
from photutils import SkyCircularAperture
# + slideshow={"slide_type": "fragment"}
radius = 5. * u.pix
sky_apers = SkyCircularAperture(coord, r=radius)
sky_apers.r
# -
radius = 0.5 * u.arcsec
sky_apers = SkyCircularAperture(coord, r=radius)
sky_apers.r
# When using a sky aperture in angular units, `aperture_photometry` needs the WCS transformation, which can be input via the `wcs` keyword:
# + slideshow={"slide_type": "fragment"}
phot = aperture_photometry(data, sky_apers, wcs=wcs)
phot
# -
# <div class="alert alert-warning alert-block">
# <h3 style='margin-top: 0;'>Learn More:</h3>
#
# Aperture Photometry in the [Extended notebook](photutils_extended.ipynb):
#
# - Bad pixel masking
# - Encircled flux
# - Aperture photometry at multiple positions using multiple apertures
#
# </div>
# + [markdown] slideshow={"slide_type": "slide"}
# ---
# # Part 2: Image Segmentation
# -
# Image segmentation is the process where sources are identified and labeled in an image.
#
# The sources are detected by using a S/N threshold level - either a per-pixel threshold image or a single value for the whole image - and defining the minimum number of pixels required within a source.
#
# First, let's define a threshold image at 2$\sigma$ (per pixel) above the background.
bkg = 0. # background level in this image
nsigma = 2.
threshold = bkg + (nsigma * error) # threshold image, this should be background-only error
# Now let's detect "8-connected" sources of minimum size 5 pixels where each pixel is 2$\sigma$ above the background.
#
# "8-connected" pixels touch along their edges or corners. "4-connected" pixels touch along their edges. For reference, SExtractor uses "8-connected" pixels.
#
# The result is a segmentation image (`SegmentationImage` object). The segmentation image is the isophotal footprint of each source above the threshold: an array in which each object is labeled with an integer. As a simple example, a segmentation map containing two distinct sources might look like this:
#
# ```
# 0 0 0 0 0 0 0 0 0 0
# 0 1 1 0 0 0 0 0 0 0
# 1 1 1 1 1 0 0 0 2 0
# 1 1 1 1 0 0 0 2 2 2
# 1 1 1 0 0 0 2 2 2 2
# 1 1 1 1 0 0 0 2 2 0
# 1 1 0 0 0 0 2 2 0 0
# 0 1 0 0 0 0 2 0 0 0
# 0 0 0 0 0 0 0 0 0 0
# ```
# where all of the pixels labeled `1` belong to the first source, all those labeled `2` belong to the second, and all `0` pixels are designated to be background.
from photutils import detect_sources
# + slideshow={"slide_type": "fragment"}
npixels = 5
segm = detect_sources(data, threshold, npixels)
print('Found {0} sources'.format(segm.nlabels))
# -
# Display the segmentation image.
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 10))
ax1.imshow(data, norm=norm)
lbl1 = ax1.set_title('Data')
ax2.imshow(segm, cmap=segm.make_cmap())
lbl2 = ax2.set_title('Segmentation Image')
# + [markdown] slideshow={"slide_type": "slide"}
# It is better to filter (smooth) the data prior to source detection.
#
# Let's use a 5x5 Gaussian kernel with a FWHM of 2 pixels.
# -
from astropy.convolution import Gaussian2DKernel
from astropy.stats import gaussian_fwhm_to_sigma
# +
sigma = 2.0 * gaussian_fwhm_to_sigma # FWHM = 2 pixels
kernel = Gaussian2DKernel(sigma, x_size=5, y_size=5)
kernel.normalize()
ssegm = detect_sources(data, threshold, npixels, filter_kernel=kernel)
print('Found {0} sources'.format(ssegm.nlabels))
# -
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 10))
ax1.imshow(segm, cmap=segm.make_cmap())
lbl1 = ax1.set_title('Original Data Segmentation')
ax2.imshow(ssegm, cmap=ssegm.make_cmap())
lbl2 = ax2.set_title('Smoothed Data Segmentation')
# ### Source deblending
# + [markdown] slideshow={"slide_type": "fragment"}
# Note above that some of our detected sources were blended. We can deblend them using the `deblend_sources()` function, which uses a combination of multi-thresholding and watershed segmentation.
#
# How the sources are deblended can be controlled with the two keywords `nlevels` and `contrast`:
# - `nlevels` is the number of multi-thresholding levels to use
# - `contrast` is the fraction of the total source flux that a local peak must have to be considered as a separate object
# -
from photutils import deblend_sources
# +
segm2 = deblend_sources(data, ssegm, npixels, filter_kernel=kernel,
contrast=0.001, nlevels=32)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(15, 8))
ax1.imshow(data, norm=norm)
ax1.set_title('Data')
ax2.imshow(ssegm, cmap=ssegm.make_cmap())
ax2.set_title('Original Segmentation Image')
ax3.imshow(segm2, cmap=segm2.make_cmap())
ax3.set_title('Deblended Segmentation Image')
print('Found {0} sources'.format(segm2.nlabels))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Measure the photometry and morphological properties of detected sources
# -
from photutils import source_properties
catalog = source_properties(data, segm2, error=error, wcs=wcs)
# `catalog` is a `SourceCatalog` object. It behaves like a list of `SourceProperties` objects, one for each source.
#
# The `to_table` method converts a subset of the `SourceCatalog` object properties (e.g., the scalar properties) to a `QTable`. We will do this so we can write the table to disk for future use, but going forward in this example we will use the `SourceCatalog` object for analysis of the detected objects.
catalog_to_tab = catalog.to_table() # convert to QTable
catalog_to_tab.remove_columns(['sky_centroid', 'sky_centroid_icrs']) # remove skycoord columns (will be useful later on)
catalog_to_tab.write('my_detected_sources.ecsv') # write to file
catalog
catalog[0] # the first source
catalog[0].xcentroid # the xcentroid of the first source
# Please go [here](http://photutils.readthedocs.io/en/latest/api/photutils.segmentation.SourceProperties.html#photutils.segmentation.SourceProperties) to see the complete list of available source properties.
# We can create a Table of isophotal photometry and morphological properties using the ``to_table()`` method of `SourceCatalog`:
tbl = catalog.to_table()
tbl
# Additional properties (not stored in the table) can be accessed directly via the `SourceCatalog` object.
# get a single object (id=12)
obj = catalog[11]
obj.id
obj
# Let's plot the cutouts of the data and error images for this source. `make_cutout` will generate a cutout of the source using the minimal bounding box of the segment.
fig, ax = plt.subplots(figsize=(12, 8), ncols=3)
ax[0].imshow(obj.make_cutout(segm2.data))
ax[0].set_title('Source id={} Segment'.format(obj.id))
ax[1].imshow(obj.data_cutout_ma)
ax[1].set_title('Source id={} Data'.format(obj.id))
ax[2].imshow(obj.error_cutout_ma)
ax[2].set_title('Source id={} Error'.format(obj.id));
# ## Exercise 1
# `data/xdf_hst_wfc3ir_60mas_f105w_sci.fits` is an F105W image of the same field used for the preceding examples. These images are aligned, so source positions detected in the F160W image should correspond to the same source if they are visible in the F105W image.
#
# In the F105W image, find the countrate in a 5-pixel aperture radius of the 3 brightest sources detected in the F160W image (hint: read in the table of detected sources that was written to disk in the previous example)
# +
# answer here
# -
# <div class="alert alert-warning alert-block">
# <h3 style="margin-top:0;">Learn More:</h3>
#
# Image Segmentation in the [Extended notebook](photutils_extended.ipynb):
#
# - Define a subset of source labels
# - Define a subset of source properties
# - Additional sources properties, such a cutout images
# - Define the approximate isophotal ellipses for each source
#
# </div>
# <div class="alert alert-warning alert-block">
# <h3 style="margin-top: 0;">PSF Photometry:</h3>
#
# See the two additional notebooks on using `photutils` for PSF-fitting photometry:
#
# - [Gaussian PSF Photometry](gaussian_psf_photometry.ipynb)
# - [Image-based PSF Photometry](image_psf_photometry_withNIRCam.ipynb)
#
# </div>
| 09-Photutils/photutils_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="eKdBA03tDnbx"
# # Teaching Physics to an AI
#
# In this Notebook, I will run simple physics simulations, and then show how neural networks can be used to "learn" or predict future states in the simulation.
# + id="nxOvYXjzZ4uz"
import time
import numpy as np
from scipy.integrate import solve_ivp
from scipy.integrate import odeint
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import pylab as py
from matplotlib.animation import FuncAnimation
from matplotlib import animation, rc
from IPython.display import HTML, Image
from matplotlib import pyplot as plt
# %config InlineBackend.figure_format = 'retina'
# + [markdown] id="o_VMgnFHDnb2"
# ## Double Pendulum Equations of Motion
#
# <img src="img/double-pendulum.png" width="100px" />
#
# $$
# F = ma = -kx
# $$
#
# with the initial conditions of $x(0) = 1$ and $v(0) = x^\prime(0) = 0$.
#
#
# ### Computational Solution
#
# Writing this as an ODE:
# $$
# x^{\prime\prime} = -\frac{k}{m}x
# $$
#
# Scipy's ODE solver can solve any system of first order ODEs, so we will rewrite this 2nd-order ODE as a system of first-order ODEs:
# $$
# \begin{align}
# x_1^\prime &= x_2 \\
# x_2^\prime &= -\frac{k}{m}x_1
# \end{align}
# $$
#
# Now let's code this up in Python.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="v72hSiz6OVsH" outputId="5bdfbcf6-34bc-47f2-b59f-ef7f91c27f16"
Image("double-pendulum.png")
# + id="dn5_xN16Dnb3"
m1 = 2 # mass of pendulum 1 (in kg)
m2 = 1 # mass of pendulum 2 (in kg)
L1 = 1.4 # length of pendulum 1 (in meter)
L2 = 1 # length of pendulum 2 (in meter)
g = 9.8 # gravitatioanl acceleration constant (m/s^2)
u0 = [-np.pi/2.2, 0, np.pi/1.8, 0] # initial conditions.
# u[0] = angle of the first pendulum
# u[1] = angular velocity of the first pendulum
# u[2] = angle of the second pendulum
# u[3] = angular velocity of the second pendulum
tfinal = 25.0 # Final time. Simulation time = 0 to tfinal.
Nt = 751
t = np.linspace(0, tfinal, Nt)
# + id="TSfqsK--F_b_"
# Differential equations describing the system
def double_pendulum(u,t,m1,m2,L1,L2,g):
# du = derivatives
# u = variables
# p = parameters
# t = time variable
du = np.zeros(4)
c = np.cos(u[0]-u[2]) # intermediate variables
s = np.sin(u[0]-u[2]) # intermediate variables
du[0] = u[1] # d(theta 1)
du[1] = ( m2*g*np.sin(u[2])*c - m2*s*(L1*c*u[1]**2 + L2*u[3]**2) - (m1+m2)*g*np.sin(u[0]) ) /( L1 *(m1+m2*s**2) )
du[2] = u[3] # d(theta 2)
du[3] = ((m1+m2)*(L1*u[1]**2*s - g*np.sin(u[2]) + g*np.sin(u[0])*c) + m2*L2*u[3]**2*s*c) / (L2 * (m1 + m2*s**2))
return du
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="6w1BTIhYGYWW" outputId="<PASSWORD>"
sol = odeint(double_pendulum, u0, t, args=(m1,m2,L1,L2,g))
#sol[:,0] = u1 = Θ_1
#sol[:,1] = u2 = ω_1
#sol[:,2] = u3 = Θ_2
#sol[:,3] = u4 = ω_2
u0 = sol[:,0] # theta_1
u1 = sol[:,1] # omega 1
u2 = sol[:,2] # theta_2
u3 = sol[:,3] # omega_2
# Mapping from polar to Cartesian
x1 = L1*np.sin(u0); # First Pendulum
y1 = -L1*np.cos(u0);
x2 = x1 + L2*np.sin(u2); # Second Pendulum
y2 = y1 - L2*np.cos(u2);
py.close('all')
py.figure(1)
#py.plot(t,x1)
#py.plot(t,y1)
py.plot(x1,y1,'.',color = '#0077BE',label = 'mass 1')
py.plot(x2,y2,'.',color = '#f66338',label = 'mass 2' )
py.legend()
py.xlabel('x (m)')
py.ylabel('y (m)')
#py.figure(2)
#py.plot(t,x2)
#py.plot(t,y2)
fig = plt.figure()
ax = plt.axes(xlim=(-L1-L2-0.5, L1+L2+0.5), ylim=(-2.5, 1.5))
#line, = ax.plot([], [], lw=2,,markersize = 9, markerfacecolor = "#FDB813",markeredgecolor ="#FD7813")
line1, = ax.plot([], [], 'o-',color = '#d2eeff',markersize = 12, markerfacecolor = '#0077BE',lw=2, markevery=10000, markeredgecolor = 'k') # line for Earth
line2, = ax.plot([], [], 'o-',color = '#ffebd8',markersize = 12, markerfacecolor = '#f66338',lw=2, markevery=10000, markeredgecolor = 'k') # line for Jupiter
line3, = ax.plot([], [], color='k', linestyle='-', linewidth=2)
line4, = ax.plot([], [], color='k', linestyle='-', linewidth=2)
line5, = ax.plot([], [], 'o', color='k', markersize = 10)
time_template = 'Time = %.1f s'
time_string = ax.text(0.05, 0.9, '', transform=ax.transAxes)
# + id="2pWgZgotGE5a"
ax.get_xaxis().set_ticks([]) # enable this to hide x axis ticks
ax.get_yaxis().set_ticks([]) # enable this to hide y axis ticks
# initialization function: plot the background of each frame
def init():
line1.set_data([], [])
line2.set_data([], [])
line3.set_data([], [])
line4.set_data([], [])
line5.set_data([], [])
time_string.set_text('')
return line3,line4, line5, line1, line2, time_string
# + id="ovAiEGd-GFFZ"
# animation function. This is called sequentially
def animate(i):
# Motion trail sizes. Defined in terms of indices. Length will vary with the time step, dt. E.g. 5 indices will span a lower distance if the time step is reduced.
trail1 = 6 # length of motion trail of weight 1
trail2 = 8 # length of motion trail of weight 2
dt = t[2]-t[1] # time step
line1.set_data(x1[i:max(1,i-trail1):-1], y1[i:max(1,i-trail1):-1]) # marker + line of first weight
line2.set_data(x2[i:max(1,i-trail2):-1], y2[i:max(1,i-trail2):-1]) # marker + line of the second weight
line3.set_data([x1[i], x2[i]], [y1[i], y2[i]]) # line connecting weight 2 to weight 1
line4.set_data([x1[i], 0], [y1[i],0]) # line connecting origin to weight 1
line5.set_data([0, 0], [0, 0])
time_string.set_text(time_template % (i*dt))
return line3, line4,line5,line1, line2, time_string
anim = animation.FuncAnimation(fig, animate, init_func=init,
frames=Nt, interval=1000*(t[2]-t[1])*0.8, blit=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="_Z-zgRrWIrRe" outputId="d7dfc8bb-a759-463d-9fa5-38466e6e9cbb"
from IPython.display import HTML
HTML(anim.to_html5_video())
# + [markdown] id="ENnpSZtADnb4"
# ### Neural Network Prediction
#
# Now let's show a neural network part of the data from this harmonic oscillator and have it try to predict the rest.
# + id="iS_fKzAIDnb-"
| notebooks/lesson-02-double-pendulum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MADE Demo
#
# Autoregressive density estimation for MNIST using masked FFNN (MADE).
# +
import sys
import argparse
import pprint
import pathlib
import numpy as np
import torch
import json
from tqdm import tqdm
from collections import OrderedDict, defaultdict
from torch.distributions import Bernoulli
import dgm
from dgm.conditional import MADEConditioner
from dgm.likelihood import FullyFactorizedLikelihood, AutoregressiveLikelihood
from dgm.opt_utils import get_optimizer, ReduceLROnPlateau
from utils import load_mnist, Batcher
# -
# Organise some hyperparameters
# +
from collections import namedtuple
Config = namedtuple(
"Config",
['seed', 'device', 'batch_size', 'data_dir',
'height', 'width', 'input_dropout',
'hidden_sizes', 'num_masks', 'resample_mask_every', 'll_samples',
'epochs', 'opt', 'lr', 'momentum', 'l2_weight', 'patience', 'early_stopping'])
args = Config(
seed=42,
device='cuda:0',
batch_size=128,
data_dir='./data',
height=28,
width=28,
input_dropout=0.,
hidden_sizes=[8000, 8000],
num_masks=1,
resample_mask_every=20,
ll_samples=10,
epochs=200,
opt='adam',
lr=1e-4,
momentum=0.,
l2_weight=1e-4,
patience=10,
early_stopping=10,
)
# -
# We like reproducibility
np.random.seed(args.seed)
torch.manual_seed(args.seed)
torch.cuda.manual_seed_all(args.seed)
# Load data
train_loader, valid_loader, test_loader = load_mnist(
args.batch_size,
save_to='{}/std/{}x{}'.format(args.data_dir, args.height, args.width),
height=args.height,
width=args.width)
# Build a model
#
# \begin{align}
# P(x) &= \prod_{i=1}^{|x|} \text{Bern}(x_i|\underbrace{f(x_{<i})}_{\text{MADE}})
# \end{align}
#
# For that we need a *conditioner*
# * the part that maps prefixes into Bernoulli parameters
#
# and an *autoregressive likelihood*
# * the part that combines the Bernoulli factors
#
#
# +
x_size = args.width * args.height
device = torch.device(args.device)
made = MADEConditioner(
input_size=x_size, # our only input to the MADE layer is the observation
output_size=x_size * 1, # number of parameters to predict
context_size=0, # we do not have any additional inputs
hidden_sizes=args.hidden_sizes,
num_masks=args.num_masks
)
model = AutoregressiveLikelihood(
event_size=x_size, # size of observation
dist_type=Bernoulli,
conditioner=made
).to(device)
print("\n# Architecture")
print(model)
# -
# Let's configure the optimiser
print("\n# Optimizer")
opt = get_optimizer(args.opt, model.parameters(), args.lr, args.l2_weight, args.momentum)
scheduler = ReduceLROnPlateau(
opt,
factor=0.5,
patience=args.patience,
early_stopping=args.early_stopping,
mode='min', threshold_mode='abs')
print(opt)
# Some helper code for batching MNIST digits
def get_batcher(data_loader):
batcher = Batcher(
data_loader,
height=args.height,
width=args.width,
device=torch.device(args.device),
binarize=True,
num_classes=10,
onehot=True
)
return batcher
# Helper code for validating a model
def validate(batcher, args, model,
optimizer=None, scheduler=None, writer=None, name='dev'):
"""
:return: stop flag, dict
NLL can be found in the dict
"""
if args.num_masks == 1:
resample_mask = False
num_samples = 1
else: # ensemble
resample_mask = True
num_samples = args.ll_samples
with torch.no_grad():
model.eval()
print_ = defaultdict(list)
nb_instances = 0.
for x_mb, y_mb in batcher:
# [B, H*W]
x_mb = x_mb.reshape(-1, args.height * args.width)
# [B, 10]
made_inputs = x_mb
# [B, H*W]
p_x = model(
inputs=None,
history=x_mb,
num_samples=num_samples, resample_mask=resample_mask
)
# [B]
nll = -p_x.log_prob(x_mb).sum(-1)
# accumulate metrics
print_['NLL'].append(nll.sum().item())
nb_instances += x_mb.size(0)
return_dict = {k: np.sum(v) / nb_instances for k, v in print_.items()}
if writer:
writer.add_scalar('%s/NLL' % name, return_dict['NLL'])
stop = False
if scheduler is not None:
stop = scheduler.step(return_dict['NLL'])
return stop, return_dict
# Main training loop
# +
print("\n# Training")
#from tensorboardX import SummaryWriter
#writer = SummaryWriter(args.logdir)
writer = None
step = 1
for epoch in range(args.epochs):
iterator = tqdm(get_batcher(train_loader))
for x_mb, y_mb in iterator:
# [B, H*W]
x_mb = x_mb.reshape(-1, args.height * args.width)
# [B, 10]
context = None
model.train()
opt.zero_grad()
if args.num_masks == 1:
resample_mask = False
else: # training with variable masks
resample_mask = args.resample_mask_every > 0 and step % args.resample_mask_every == 0
# [B, H*W]
noisy_x = torch.where(
torch.rand_like(x_mb) > args.input_dropout, x_mb, torch.zeros_like(x_mb)
)
p_x = model(
inputs=context,
history=noisy_x,
resample_mask=resample_mask
)
# [B, H*W]
ll_mb = p_x.log_prob(x_mb)
# [B]
ll = ll_mb.sum(-1)
loss = -(ll).mean()
loss.backward()
opt.step()
display = OrderedDict()
display['0s'] = '{:.2f}'.format((x_mb == 0).float().mean().item())
display['1s'] = '{:.2f}'.format((x_mb == 1).float().mean().item())
display['NLL'] = '{:.2f}'.format(-ll.mean().item())
if writer:
writer.add_scalar('training/LL', ll)
#writer.add_image('training/posterior/sample', z.mean(0).reshape(1,1,-1) * 255)
iterator.set_postfix(display, refresh=False)
step += 1
stop, dict_valid = validate(get_batcher(valid_loader), args, model, opt, scheduler,
writer=writer, name="dev")
if stop:
print('Early stopping at epoch {:3}/{}'.format(epoch + 1, args.epochs))
break
print('Epoch {:3}/{} -- '.format(epoch + 1, args.epochs) + \
', '.join(['{}: {:4.2f}'.format(k, v) for k, v in sorted(dict_valid.items())]))
# -
# Let's visualise samples
# +
from matplotlib import pyplot as plt
def visualize_made(batcher: Batcher, args, model, N=4, writer=None, name='dev'):
assert N <= args.batch_size, "N should be no bigger than a batch"
with torch.no_grad():
model.eval()
plt.figure(figsize=(2*N, 2*N))
plt.subplots_adjust(wspace=0.5, hspace=0.5)
# Some visualisations
for x_mb, y_mb in batcher:
# [B, H*W]
x_mb = x_mb.reshape(-1, args.height * args.width)
x_mb = x_mb[:N]
# [B, 10]
context = None
p_x = model(inputs=None, history=x_mb, resample_mask=False)
# [B, H*W]
ll = p_x.log_prob(x_mb)
prob = torch.exp(ll)
# reconstruct bottom half of N instances
x_rec = model.sample(inputs=None, history=x_mb, start_from=args.height * args.width // 2)
# sample N instances
x_sample = model.sample(
inputs=None,
history=torch.zeros(
[N, args.height * args.width],
device=torch.device(args.device),
dtype=torch.float32
),
start_from=0
)
for i in range(N):
plt.subplot(4, N, 0*N + i + 1)
plt.imshow(x_mb[i].reshape(args.height, args.width).cpu(), cmap='Greys')
plt.title("x%d" % (i + 1))
plt.subplot(4, N, 1*N + i + 1)
plt.imshow(prob[i].reshape(args.height, args.width).cpu(), cmap='Greys')
plt.title("prob%d" % (i + 1))
plt.subplot(4, N, 2*N + i + 1)
plt.axhline(y=args.height//2, c='red', linewidth=1, ls='--')
plt.imshow(x_rec[i].reshape(args.height, args.width).cpu(), cmap='Greys')
plt.title("rec%d" % (i + 1))
plt.subplot(4, N, 3*N + i + 1)
plt.imshow(x_sample[i].reshape(args.height, args.width).cpu(), cmap='Greys')
plt.title("sample%d" % (i + 1))
break
plt.show()
# -
# A few reconstructions for the validation set as well as samples from the autoregressive likelihood
visualize_made(get_batcher(valid_loader), args, model)
| examples/mnist/MADE demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##### Student Name: <NAME>
# ##### Student ID: 46150641
# # Sport Vouchers Program Analysis
#
# The goal of this Portfolio task is to explore data from the Federal Government Sport Vouchers program - this is a
# program that provides up to two $100 vouchers for kids to participate in organised sport. Here's the [NSW Active Kids page](https://www.service.nsw.gov.au/transaction/apply-active-kids-voucher), there are similar schemes in other states - this data is from South Australia.
#
# This is an exercise in exploring data and communicating the insights you can gain from it. The source data comes
# from the `data.gov.au` website and provides details of all Sport Vouchers that have been redeemed since February in SA 2015 as part of the Sport Voucher program: [Sports Vouchers Data](https://data.gov.au/dataset/ds-sa-14daba50-04ff-46c6-8468-9fa593b9f100/details). This download is provided for you as `sportsvouchersclaimed.csv`.
#
# To augment this data you can also make use of [ABS SEIFA data by LGA](http://stat.data.abs.gov.au/Index.aspx?DataSetCode=ABS_SEIFA_LGA#) which shows a few measures of Socioeconomic Advantage and Disadvantage for every Local Government Area. This data is provided for you as `ABS_SEIFA_LGA.csv`. This could enable you to answer questions about whether the voucher program is used equally by parents in low, middle and high socioeconomic areas. You might be interested in this if you were concerned that this kind of program might just benifit parents who are already advantaged (they might already be paying for sport so this program wouldn't be helping much).
#
# Questions:
# * Describe the distribution of vouchers by: LGA, Sport - which regions/sports stand out?
# * Are some sports more popular in different parts of the state?
# * Are any LGAs over/under represented in their use of vouchers?
# * Is there a relationship between any of the SEIFA measures and voucher use in an LGA?
#
# A challenge in this task is to display a useful summary of the data given that there are a large number of LGAs and sports involved. Try to avoid long lists and large tables. Think about what plots and tables communicate the main points of your findings.
#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Read the sports vouchers data
sa_vouchers = pd.read_csv("files/sportsvouchersclaimed.csv")
sa_vouchers.head()
# Read the socioeconomic areas data
seifa = pd.read_csv('files/ABS_SEIFA_LGA.csv')
seifa.head()
# The SEIFA data includes row for each Local Government Area (LGA) but the names of the LGAs have a letter or letters in brackets after the name. To allow us to match this up with the voucher data we remove this and convert to uppercase.
#
# For each LGA the data includes a number of measures all of which could be useful in your exploration.
# Create an LGA column by removing the letters in brackets and converting to uppercase
lga = seifa["Local Government Areas - 2011"].str.replace(' \([ACSRCDMT]+\)', '').str.upper()
seifa['LGA'] = lga
seifa.head()
# Since there are many rows per LGA we need to use `pivot_table` to create a new data frame with one row per LGA. Here
# is an example of doing this to create a table with the different SCORE measures and the population (URP) field.
# ### Question 4
#Question 4
# Group 4 measurements with LGA
LGA_scores = seifa[seifa.MEASURE == 'SCORE'].pivot_table(index="LGA", columns=["INDEX_TYPE"], values="Value")
LGA_scores.head() #Social economic level
# Create population column
LGA_pop = seifa[seifa.MEASURE == 'URP'].pivot_table(index="LGA", columns=["INDEX_TYPE"], values="Value")
LGA_scores['Population'] = LGA_pop.IEO
LGA_scores.head()
# This data frame can then be joined with the vouchers data fram to create one master data frame containing both the voucher data and the SEIFA measures.
#Join dataframe of the 4 measurements (IEO, IER, IRSAD, IRSD) and population to dataframe sa_vouchers by same LGA
sa_vouchers_scores = sa_vouchers.join(LGA_scores, on='Participant_LGA')
sa_vouchers_scores.head()
# ### Summary for question 4
# Because the 4 measurements are able to join the dataframe contain the number of vouchers use, it is possible to conclude there is a relationship between the 4 measurements and the voucher use by LGA.
sa_vouchers_scores_group = sa_vouchers_scores.groupby('Participant_LGA')
# ### Question 1
# Question 1:
#Voucher distribution by regions/ LGA
sa_LGA=pd.DataFrame({'count':sa_vouchers_scores_group.size()})
LGA_scores['count']=sa_LGA['count']
#Merge voucher by LGA to dataframe
sa_count=sa_vouchers_scores.merge(sa_LGA['count'], left_on='Participant_LGA', right_index=True)
sa_count = sa_count.rename(columns={'count': 'Count_by_LGA'})
sa_count.head()
#Scatterplot shows the voucher distribution by regions/ LGA
plt.figure(figsize = (50,40))
sns.scatterplot(data=sa_count, x="Count_by_LGA", y="Participant_LGA", hue="Participant_LGA", s=100, linewidths=.5, edgecolor="black", legend=False)
# The scatterplot shows that voucher distribution by LGA is normally distributed.
#Voucher distribution by regions/ LGA
sa_LGA=sa_LGA.join(LGA_scores['Population'], on='Participant_LGA')
sa_LGA['percent_vouchers']=100*sa_LGA['count']/sa_LGA['count'].sum() #Voucher use percentage among different LGA voucher use
sa_LGA['percent_population']=100*sa_LGA['count']/sa_LGA['Population'] #Voucher use percentage compare with total population among LGA
sa_LGA
sa_LGA['percent_vouchers'].nlargest(5) # Top 5 LGA that have largest vouchers distribution
sa_LGA['percent_population'].nlargest(5) # Top 5 LGA that have largest vouchers distribution compare to population
# Turn index to columns
sa_LGA.reset_index(inplace=True)
#Barplot for 5 LGA that have largest vouchers distribution
largest_pv=sa_LGA[sa_LGA["Participant_LGA"].isin(['ONKAPARINGA','SALISBURY', 'TEA TREE GULLY', 'PLAYFORD','CHARLES STURT'])]
largest_pv=largest_pv.sort_values(by='percent_vouchers')
plt.figure(figsize = (12,8))
ax = sns.barplot(y='percent_vouchers', x='Participant_LGA',data=largest_pv, palette="Oranges_d", linewidth=1, edgecolor=".2")
#Barplot for 5 LGA that have largest vouchers distribution
largest_pp=sa_LGA[sa_LGA["Participant_LGA"].isin(['KAROONDA EAST MURRAY','KIMBA', 'WUDINNA', 'CLEVE', 'SOUT<NAME>'])]
largest_pp=largest_pv.sort_values(by='percent_population')
plt.figure(figsize = (12,8))
ax = sns.barplot(y='percent_population', x='Participant_LGA',data=largest_pp, palette="Purples_d", linewidth=1, edgecolor=".2")
#Voucher distribution by sports
sa_vouchers_scores_sport = sa_vouchers_scores.groupby('Voucher_Sport')
sa_vouchers_scores_sport.size()
sa_sport=pd.DataFrame({'count':sa_vouchers_scores_sport.size()}) # The number of voucher use by sports
sa_sport['percent_vouchers']=100*sa_sport['count']/sa_sport['count'].sum() # Percentage of voucher use by sports
sa_sport.head()
sa_sport['percent_vouchers'].nlargest(5) # Top 5 sports that have largest vouchers distribution
# Turn index to columns
sa_sport.reset_index(inplace=True)
#Barplot for 5 sports that have largest vouchers distribution
largest_sv=sa_sport[sa_sport["Voucher_Sport"].isin(['Australian Rules','Netball', 'Football (Soccer)', 'Gymnastics', 'Basketball'])]
largest_sv=largest_sv.sort_values(by='percent_vouchers')
plt.figure(figsize = (12,8))
ax = sns.barplot(y='percent_vouchers', x='Voucher_Sport',data=largest_sv, palette="Greens_d", linewidth=1, edgecolor=".2")
# ### Summary
# * LGA that have largest vouchers distribution is **ONKAPARINGA** which take up 10% of total voucher use of all LGA. LGA that have largest vouchers distribution compare to population is **KAROONDA EAST MURRAY** which take up 37.5% of its total population.
# * The sport that have largest vouchers distribution is **Australian Rules** which take up 37.03% of total voucher use by sports and vouchers distribution for Australian Rules is much larger than other high rank sports.
# ### Question 2
# Question 2
# Voucher distribution by sport and lga
sa_vouchers_scores_VS_LGA = sa_vouchers_scores.groupby(['Participant_LGA','Voucher_Sport']) #Group data by sport and LGA
sa_vs_lga=pd.DataFrame({'Count_by_Sport':sa_vouchers_scores_VS_LGA.size()}) #Voucher use of each sport on each LGA
sa_vs_lga=sa_vs_lga.join(LGA_scores['count'], on='Participant_LGA') #Voucher use of all sports on each LGA
sa_vs_lga=sa_vs_lga.rename(columns={'count': 'Count_by_LGA'})
sa_vs_lga['percent_vouchers_by_regions']=100*sa_vs_lga['Count_by_Sport']/sa_vs_lga['Count_by_LGA'] #Percent of each sport voucher use on each LGA
sa_vs_lga=sa_vs_lga.dropna() #Drop missing values
sa_vs_lga.head()
# Turn index to columns
sa_vs_lga.reset_index(inplace=True)
sa_vs_lga.head()
#Heatmap for voucher distribution of sports in different parts of the state
vs_lga = sa_vs_lga.pivot("Voucher_Sport", "Participant_LGA", "percent_vouchers_by_regions")
plt.figure(figsize = (35,25))
ax = sns.heatmap(vs_lga, linewidths=.5, linecolor='black', cmap=sns.color_palette("flare", as_cmap=True))
# ### Summary
# * The voucher distribution by sports in different LGA varies significantly.
# * For Autralian Rules, it is more popular in COOBER PEDY and ROXBY DOWINS, and less popular in PETERBOROUGH and PORT AUGUSTA.
# * For Netball, it is more popular in KIMBA and WUDINNA, and less popular in WEST TORRENS.
# * For Swimming, it is more popular in FRANKLIN HARBOUR and less popular in WHYALLA.
# ### Question 3
# Question 3
# Voucher distribution changes by years
sa_vouchers_scores_year = sa_vouchers_scores.groupby(['Participant_LGA','Voucher_Claim_Year'])
sa_LGA_year=pd.DataFrame({'Count':sa_vouchers_scores_year.size()})
sa_LGA_year=sa_LGA_year.join(LGA_scores['Population'], on='Participant_LGA')
sa_LGA_year=sa_LGA_year.join(LGA_scores['count'], on='Participant_LGA')
sa_LGA_year=sa_LGA_year.rename(columns={'count': 'Count_by_LGA'})
sa_LGA_year['Voucher_Value_By_Year']=sa_LGA_year['Count']*50
sa_LGA_year['Total_Voucher_Value_By_LGA']=sa_LGA_year['Count_by_LGA']*50
sa_LGA_year['Percent_Value']=100*sa_LGA_year['Voucher_Value_By_Year']/sa_LGA_year['Total_Voucher_Value_By_LGA']
#Sub section data by years
pd.options.mode.chained_assignment = None
sa_LGA_2015=sa_LGA_year.xs(2015, level='Voucher_Claim_Year')
sa_LGA_2015['Percent_Value']=100*sa_LGA_2015['Voucher_Value_By_Year']/sa_LGA_2015['Voucher_Value_By_Year'].sum()
sa_LGA_2016=sa_LGA_year.xs(2016, level='Voucher_Claim_Year')
sa_LGA_2016['Percent_Value']=100*sa_LGA_2016['Voucher_Value_By_Year']/sa_LGA_2016['Voucher_Value_By_Year'].sum()
sa_LGA_2017=sa_LGA_year.xs(2017, level='Voucher_Claim_Year')
sa_LGA_2017['Percent_Value']=100*sa_LGA_2017['Voucher_Value_By_Year']/sa_LGA_2017['Voucher_Value_By_Year'].sum()
# Turn index to columns
sa_LGA_year.reset_index(inplace=True)
# sa_LGA_year
#Sum voucher values by years
sum_2015=sa_LGA_2015['Voucher_Value_By_Year'].sum()
sum_2016=sa_LGA_2016['Voucher_Value_By_Year'].sum()
sum_2017=sa_LGA_2017['Voucher_Value_By_Year'].sum()
#Create column for percentage voucher value be used by years among LGA
sa_LGA_year['Percent_Value_By_Year']=0
sa_LGA_year.head()
#Add voucher value percentage by years among LGA calculated in sub section data to original data
sa_LGA_year.loc[sa_LGA_year.Voucher_Claim_Year == 2015, "Percent_Value_By_Year"]= 100*sa_LGA_year['Voucher_Value_By_Year']/sum_2015
sa_LGA_year.loc[sa_LGA_year.Voucher_Claim_Year == 2016, "Percent_Value_By_Year"]= 100*sa_LGA_year['Voucher_Value_By_Year']/sum_2016
sa_LGA_year.loc[sa_LGA_year.Voucher_Claim_Year == 2017, "Percent_Value_By_Year"]= 100*sa_LGA_year['Voucher_Value_By_Year']/sum_2017
sa_LGA_year.head()
#Heatmap for distribution of vouchers use by LGAs that LGAs over/under represented
year_lga = sa_LGA_year.pivot("Participant_LGA", "Voucher_Claim_Year", "Percent_Value_By_Year")
plt.figure(figsize = (25,35))
ax = sns.heatmap(year_lga, linewidths=.5, linecolor='black', cmap=sns.cubehelix_palette(start=.5, rot=-.5, as_cmap=True))
# ### Summary
# * LGAs that are **over represented** their use of voucher are ONKAPARINGA, PLAYFORD, PORT ADELAIDE ENFIELD, SALISBURY, TEA TREE GULLY, CHARLES STURT, ADELAIDE HILLS, MARION and MITCHAM.
# * LGAs that are **under represented** their use of voucher are COOBER PEDY, KIMBA, KINGSTON, WUDINNA, YANKALILLA, etc.
# ### Question 4 (Cont.)
# Question 4:
#Make a copy dataframe of LGA_scores
ms_LGA=LGA_scores.copy()
# Turn index to columns
ms_LGA.reset_index(inplace=True)
ms_LGA
fig, axes = plt.subplots(4, 1, figsize=(25, 60), sharey=True)
fig.suptitle('Voucher used by different measurement')
#Scatterplot shows the voucher distribution by IEO
sns.scatterplot(ax=axes[0],data=ms_LGA, x="IEO", y="count", hue="LGA", s=100, linewidths=.5, edgecolor="black", legend=False)
axes[0].set_title('IEO')
#Scatterplot shows the voucher distribution by IER
sns.scatterplot(ax=axes[1],data=ms_LGA, x="IER", y="count", hue="LGA", s=100, linewidths=.5, edgecolor="black", legend=False)
axes[1].set_title('IER')
#Scatterplot shows the voucher distribution by IRSAD
sns.scatterplot(ax=axes[2],data=ms_LGA, x="IRSAD", y="count", hue="LGA", s=100, linewidths=.5, edgecolor="black", legend=False)
axes[2].set_title('IRSAD')
#Scatterplot shows the voucher distribution by IRSD
sns.scatterplot(ax=axes[3],data=ms_LGA, x="IRSD", y="count", hue="LGA", s=100, linewidths=.5, edgecolor="black", legend=False)
axes[3].set_title('IRSD')
# ### Summary
# As can be seen in the 4 scatterplot, there is a slight difference of the voucher use among the 4 measurements. The effects of IEO and IER to voucher use in LGA is significantly similar to each other. The effects of IRSAD and IRSD to voucher use in LGA is significantly similar to each other. However, it is quite different between (IEO, IER) and (IRSAD, IRSD).
# ## Challenge - Queensland
#
# _Note: this is an extra task that you might take on to get a better grade for your portfolio. You can get a good pass grade without doing this._
#
# Queensland has a similar program called [Get Started](https://data.gov.au/dataset/ds-qld-3118838a-d425-48fa-bfc9-bc615ddae44e/details?q=get%20started%20vouchers) and we can retrieve data from their program in a similar format.
#
# The file [round1-redeemed_get_started_vouchers.csv](files/round1-redeemed_get_started_vouchers.csv) contains records of the vouchers issued in Queensland. The date of this data is not included but the program started in 2015 so it is probably from around then.
#
# The data includes the LGA of the individual but the name of the activity is slightly different. To do a comparable analysis you would need to map the activity names onto those from South Australia.
qld_vouchers = pd.read_csv('files/round1-redeemed_get_started_vouchers.csv')
qld_vouchers.head()
# Join the QLD data with the LGA data as before to get population and SIEFA data integrated
qld_vouchers['LGA'] = qld_vouchers['Club Local Government Area Name'].str.replace(' \([RC]+\)', '').str.upper()
qld_vouchers_scores = qld_vouchers.join(LGA_scores, on='LGA')
qld_vouchers_scores.head()
sa_count['Voucher_Sport'].unique()
#unify the second table to the first one to have same Participation Activity (sport)
qld_vouchers_scores['Participation Activity'].unique()
new_qld=qld_vouchers_scores[qld_vouchers_scores['Participation Activity'].isin(['Gymnastics', 'Basketball', 'Netball', 'Golf', 'Calisthenics', 'Cricket', 'Tennis', 'Football (Soccer)', 'Taekwondo', 'Table Tennis', 'Athletics', 'Rugby League', 'Hockey', 'Ju Jitsu', 'Baseball', 'Softball', 'Swimming', 'Judo', 'Rugby Union', 'Dancesport', 'Equestrian', 'Ice Skating', 'Squash', 'Sailing', 'Roller Sports', 'Cycling', 'Archery', 'Touch', 'Karate', 'Rowing', 'Weightlifting', 'Shooting'])]
new_qld.head()
sa_count = sa_count.rename(columns={'Voucher_Sport': 'Participation Activity'})
#Merge dataframe South Australia and Queensland
sa_qld = sa_count.merge(new_qld, on='Participation Activity')
sa_qld.head()
sa_group = sa_qld.groupby(['Participation Activity', 'Participant_LGA'])
#Voucher distribution by regions/ LGA
sa_s=pd.DataFrame({'count':sa_group.size()})
sa_s.head()
qld_group = sa_qld.groupby(['Participation Activity', 'LGA'])
qld_s=pd.DataFrame({'count':qld_group.size()})
qld_s.head()
# Turn index to columns
sa_s.reset_index(inplace=True)
qld_s.reset_index(inplace=True)
qld_s=qld_s.rename(columns={'LGA': 'Participant_LGA'})
#Merge dataframe South Australia and Queensland by Participant Activity (Sport)
sa_qld_sport=sa_s.append(qld_s)
sa_qld_sport.head()
sa_qld_sport['Participant_LGA'].unique()
#Heatmap for voucher distribution of sports in different parts of the state
sa_qld_hm = sa_qld_sport.pivot("Participation Activity", "Participant_LGA", "count")
plt.figure(figsize = (35,25))
ax = sns.heatmap(sa_qld_hm, linewidths=.5, linecolor='black', cmap=sns.color_palette("crest", as_cmap=True))
# ### Summary
# * In general, the voucher distribution of South Australia and Queensland is quite similar by Participation Activity.
# * For Swimming, the voucher distribution of Queensland LGA is greater than South Australia LGA. For Netball, the voucher distribution of Queensland LGA is slightly smaller than South Australia LGA.
| Portfolio 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sys
import os
from pathlib import Path
sys.path.append(str(Path.cwd().parent))
import logging.config
from dobermann.config import settings
logging.config.dictConfig(settings.LOGGING)
# +
import datetime as dt
from dobermann.core import backtest, Timeframe, TestStrategy
report = await backtest(
strategy=TestStrategy(),
start_at=dt.datetime(2022, 1, 1),
end_at=dt.datetime(2022, 4, 1),
timeframe=Timeframe.H1,
tickers=['BTCUSDT', 'ETHUSDT', 'BNBUSDT', 'DYDXUSDT', 'NEARUSDT'],
)
# -
report.summary
report.equity_chart
| examples/backtesting_multicore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Orthogonal Functions
# > What does it mean for functions to be orthogonal?
# +
#collapse
# imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy.stats
# %matplotlib inline
plt.rcParams['figure.figsize'] = [12, 5]
plt.rcParams['figure.dpi'] = 140
π = np.pi
exp = np.exp
sin = np.sin
cos = np.cos
sqrt = np.sqrt
# -
# # Fourier Basis
# +
grid = 200
domain = [0, 2*π]
dx = (domain[1]-domain[0])/grid
grid = np.linspace(*domain, grid)
def fourier(k, x): return sin(k*x)+cos(k*x)
# +
n = 5
basis = pd.DataFrame({k: fourier(k, grid) for k in range(1,n)}, index=grid)
ax = basis.plot.line(lw=0.4, xlim=domain)
ax.axhline(0, c='black', lw='0.3')
# +
from scipy import integrate
def compare_two(i, j):
product = pd.Series(basis[i]*basis[j], name='product')
product = pd.DataFrame([basis[i], basis[j], product]).T
ax = product.plot.line(lw=0.5, color=['red', 'blue', 'purple'])
ax.fill_between(grid, product['product'], alpha=0.1)
return integrate.trapz(product['product'], x=product.index)
# -
print('integral =', np.round(compare_two(3,4), 4))
# "fourier modes as eigenfunctions of the derivative operator" What?
# # Polynomial Bases
| _notebooks/2020-12-04-orthogonal-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Factor Model of Asset Return
#
import sys
# !{sys.executable} -m pip install -r requirements.txt
import numpy as np
import pandas as pd
import time
import os
import quiz_helper
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
plt.rcParams['figure.figsize'] = (14, 8)
# ### data bundle
import os
import quiz_helper
from zipline.data import bundles
os.environ['ZIPLINE_ROOT'] = os.path.join(os.getcwd(), '..', '..','data','module_4_quizzes_eod')
ingest_func = bundles.csvdir.csvdir_equities(['daily'], quiz_helper.EOD_BUNDLE_NAME)
bundles.register(quiz_helper.EOD_BUNDLE_NAME, ingest_func)
print('Data Registered')
# ### Build pipeline engine
# +
from zipline.pipeline import Pipeline
from zipline.pipeline.factors import AverageDollarVolume
from zipline.utils.calendars import get_calendar
universe = AverageDollarVolume(window_length=120).top(500)
trading_calendar = get_calendar('NYSE')
bundle_data = bundles.load(quiz_helper.EOD_BUNDLE_NAME)
engine = quiz_helper.build_pipeline_engine(bundle_data, trading_calendar)
# -
# ### View Data¶
# With the pipeline engine built, let's get the stocks at the end of the period in the universe we're using. We'll use these tickers to generate the returns data for the our risk model.
# +
universe_end_date = pd.Timestamp('2016-01-05', tz='UTC')
universe_tickers = engine\
.run_pipeline(
Pipeline(screen=universe),
universe_end_date,
universe_end_date)\
.index.get_level_values(1)\
.values.tolist()
universe_tickers
# -
len(universe_tickers)
# +
from zipline.data.data_portal import DataPortal
data_portal = DataPortal(
bundle_data.asset_finder,
trading_calendar=trading_calendar,
first_trading_day=bundle_data.equity_daily_bar_reader.first_trading_day,
equity_minute_reader=None,
equity_daily_reader=bundle_data.equity_daily_bar_reader,
adjustment_reader=bundle_data.adjustment_reader)
# -
# ## Get pricing data helper function
from quiz_helper import get_pricing
# ## get pricing data into a dataframe
# +
returns_df = \
get_pricing(
data_portal,
trading_calendar,
universe_tickers,
universe_end_date - pd.DateOffset(years=5),
universe_end_date)\
.pct_change()[1:].fillna(0) #convert prices into returns
returns_df
# -
# ## Let's look at one stock
#
# Let's look at this for just one stock. We'll pick AAPL in this example.
# +
aapl_col = returns_df.columns[3]
asset_return = returns_df[aapl_col]
asset_return = asset_return.rename('asset_return')
# -
# ## Factor returns
# Let's make up a "factor" by taking an average of all stocks in our list. You can think of this as an equal weighted index of the 490 stocks, kind of like a measure of the "market". We'll also make another factor by calculating the median of all the stocks. These are mainly intended to help us generate some data to work with. We'll go into how some common risk factors are generated later in the lessons.
#
# Also note that we're setting axis=1 so that we calculate a value for each time period (row) instead of one value for each column (assets).
factor_return_1 = returns_df.mean(axis=1)
factor_return_2 = returns_df.median(axis=1)
factor_return_2.shape
np.array([factor_return_1.values,factor_return_2.values]).T.shape
asset_return.values.shape
# ## Factor exposures
#
# Factor exposures refer to how "exposed" a stock is to each factor. We'll get into this more later. For now, just think of this as one number for each stock, for each of the factors.
from sklearn.linear_model import LinearRegression
"""
You can run these in separate cells to see each step in detail
But for now, just assume that we're calculating a number for each
stock, for each factor, which represents how "exposed" each stock is
to each factor.
We'll discuss how factor exposure is calculated later in the lessons.
"""
lr = LinearRegression()
X = np.array([factor_return_1.values,factor_return_2.values]).T
y = np.array(asset_return.values)
lr.fit(X,y)
factor_exposure_1 = lr.coef_[0]
factor_exposure_2 = lr.coef_[1]
# ## Quiz 1 Contribution of Factors
#
# The sum of the products of factor exposure times factor return is the contribution of the factors. It's also called the "common return." calculate the common return of AAPL, given the two factor exposures and the two factor returns.
# ## Answer 1
# Calculate the contribution of the two factors to the return of this example asset
common_return = factor_exposure_1 * factor_return_1 + factor_exposure_2 * factor_return_2
common_return = common_return.rename('common_return')
# ## Quiz 2 Specific Return
# The specific return is the part of the stock return that isn't explained by the factors. So it's the actual return minus the common return.
# Calculate the specific return of the stock.
# ## Answer 2
# +
# TODO: calculate the specific return of this asset
specific_return = asset_return - common_return
specific_return = specific_return.rename('specific_return')
# -
# ## Visualize the common return and specific return
#
return_components = pd.concat([common_return,specific_return],axis=1)
return_components.head(2)
return_components.plot(title="asset return = common return + specific return");
pd.DataFrame(asset_return).plot(color='purple');
| Lesson 24 Exercises/factor_model_asset_return.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import cv2 as cv
import ipywidgets.widgets as widgets
import threading
import math
import time
import numpy as np
#Set camera display component
image_widget_src = widgets.Image(format='jpeg', width=600, height=500)
display(image_widget_src) # display camera component
image_widget_edge = widgets.Image(format='jpeg', width=600, height=500)
display(image_widget_edge) # display camera component
# +
#bgr 8 to jpeg format
import enum
import cv2
def bgr8_to_jpeg(value, quality=75):
return bytes(cv2.imencode('.jpg', value)[1])
# +
image = cv2.VideoCapture(0) # Open camera
# width=1280
# height=960
# cap.set(cv2.CAP_PROP_FRAME_WIDTH,width) # set width of image
# cap.set(cv2.CAP_PROP_FRAME_HEIGHT,height) # set height of image
image.set(3,600)
image.set(4,500)
image.set(5, 30) # set frame
image.set(cv2.CAP_PROP_FOURCC, cv2.VideoWriter.fourcc('M', 'J', 'P', 'G'))
image.set(cv2.CAP_PROP_BRIGHTNESS, 40) #set brightness -64 - 64 0.0
image.set(cv2.CAP_PROP_CONTRAST, 50) #set contrast -64 - 64 2.0
image.set(cv2.CAP_PROP_EXPOSURE, 156) #set exposure value 1.0 - 5000 156.0
ret, frame = image.read() # read camera data
image_widget_src.value = bgr8_to_jpeg(frame)
# +
# for ROI
#x = 0; y = 188
#x_ = 600; y_ = 500
while(True):
ret, src = image.read()
src = cv2.resize(src, (600,500))
#roi = src[y:y_, x:x_]
dst = cv.Canny(src, 100, 200, None, 3)
cdst = cv.cvtColor(dst, cv.COLOR_GRAY2BGR)
cdstP = np.copy(cdst)
lines = cv.HoughLines(dst, 1, np.pi / 180, 150, None, 0, 0)
if lines is not None:
for i in range(0, len(lines)):
rho = lines[i][0][0]
theta = lines[i][0][1]
a = math.cos(theta)
b = math.sin(theta)
x0 = a * rho
y0 = b * rho
pt1 = (int(x0 + 1000 * (-b)), int(y0 + 1000 * (a)))
pt2 = (int(x0 - 1000 * (-b)), int(y0 - 1000 * (a)))
cv.line(cdst, pt1, pt2, (0, 0, 255), 3, cv.LINE_AA)
linesP = cv.HoughLinesP(dst, 1, np.pi / 180, 50, None, 50, 10)
if linesP is not None:
for i in range(0, len(linesP)):
l = linesP[i][0]
cv.line(cdstP, (l[0], l[1]), (l[2], l[3]), (0, 0, 255), 3, cv.LINE_AA)
image_widget_src.value = bgr8_to_jpeg(src)
image_widget_edge.value = bgr8_to_jpeg(cdstP)
time.sleep(0.010)
# -
image.release()
| tmp_project/ipynb_test/detected_line.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Interactive mapping and analysis of geospatial big data using geemap and Google Earth Engine**
#
# This notebook was developed for the geemap workshop at the [GeoPython 2021 Conference](https://2021.geopython.net).
#
# Authors: [<NAME>](https://github.com/giswqs), [<NAME>](https://github.com/KMarkert)
#
# Link to this notebook: https://gishub.org/geopython
#
# Recorded video: https://www.youtube.com/watch?v=wGjpjh9IQ5I
#
# [](https://www.youtube.com/watch?v=wGjpjh9IQ5I)
#
#
# ## Introduction
#
# ### Description
#
# Google Earth Engine (GEE) is a cloud computing platform with a multi-petabyte catalog of satellite imagery and geospatial datasets. It enables scientists, researchers, and developers to analyze and visualize changes on the Earth’s surface. The geemap Python package provides GEE users with an intuitive interface to manipulate, analyze, and visualize geospatial big data interactively in a Jupyter-based environment. The topics to be covered in this workshop include:
#
# 1. Introducing geemap and the Earth Engine Python API
# 2. Creating interactive maps
# 3. Searching GEE data catalog
# 4. Displaying GEE datasets
# 5. Classifying images using machine learning algorithms
# 6. Computing statistics and exporting results
# 7. Producing publication-quality maps
# 8. Building and deploying interactive web apps, among others
#
# This workshop is intended for scientific programmers, data scientists, geospatial analysts, and concerned citizens of Earth. The attendees are expected to have a basic understanding of Python and the Jupyter ecosystem. Familiarity with Earth science and geospatial datasets is useful but not required.
#
# ### Useful links
# - [GeoPython 2021 Conference website](https://2021.geopython.net)
# - [Google Earth Engine](https://earthengine.google.com)
# - [geemap.org](https://geemap.org)
# - [Google Earth Engine and geemap Python Tutorials](https://www.youtube.com/playlist?list=PLAxJ4-o7ZoPccOFv1dCwvGI6TYnirRTg3) (55 videos with a total length of 15 hours)
# - [Spatial Data Management with Google Earth Engine](https://www.youtube.com/playlist?list=PLAxJ4-o7ZoPdz9LHIJIxHlZe3t-MRCn61) (19 videos with a total length of 9 hours)
# - [Ask geemap questions on GitHub](https://github.com/giswqs/geemap/discussions)
#
# ### Prerequisite
# - A Google Earth Engine account. Sigh up [here](https://earthengine.google.com) if needed.
# - [Miniconda](https://docs.conda.io/en/latest/miniconda.html) or [Anaconda](https://www.anaconda.com/products/individual)
#
#
# ### Set up a conda environment
#
# ```
# conda create -n geo python=3.8
# conda activate geo
# conda install geemap -c conda-forge
# conda install jupyter_contrib_nbextensions -c conda-forge
# jupyter contrib nbextension install --user
# ```
#
# ## geemap basics
#
# ### Import libraries
import os
import ee
import geemap
# ### Create an interactive map
Map = geemap.Map()
Map
# ### Customize the default map
#
# You can specify the center(lat, lon) and zoom for the default map. The lite mode will only show the zoom in/out tool.
Map = geemap.Map(center=(40, -100), zoom=4, lite_mode=True)
Map
# ### Add basemaps
Map = geemap.Map()
Map.add_basemap('HYBRID')
Map
from geemap.basemaps import basemaps
Map.add_basemap(basemaps.OpenTopoMap)
# ### Change basemaps without coding
# 
Map = geemap.Map()
Map
# ### Add WMS and XYZ tile layers
#
# Examples: https://viewer.nationalmap.gov/services/
#
# +
Map = geemap.Map()
url = 'https://mt1.google.com/vt/lyrs=p&x={x}&y={y}&z={z}'
Map.add_tile_layer(url, name='Google Terrain', attribution='Google')
Map
# -
naip_url = 'https://services.nationalmap.gov/arcgis/services/USGSNAIPImagery/ImageServer/WMSServer?'
Map.add_wms_layer(url=naip_url, layers='0', name='NAIP Imagery', format='image/png', shown=True)
# ### Use drawing tools
Map = geemap.Map()
Map
# +
# Map.user_roi.getInfo()
# +
# Map.user_rois.getInfo()
# -
# ### Convert GEE JavaScript to Python
#
# https://developers.google.com/earth-engine/guides/image_visualization
js_snippet = """
// Load an image.
var image = ee.Image('LANDSAT/LC08/C01/T1_TOA/LC08_044034_20140318');
// Define the visualization parameters.
var vizParams = {
bands: ['B5', 'B4', 'B3'],
min: 0,
max: 0.5,
gamma: [0.95, 1.1, 1]
};
// Center the map and display the image.
Map.setCenter(-122.1899, 37.5010, 10); // San Francisco Bay
Map.addLayer(image, vizParams, 'false color composite');
"""
geemap.js_snippet_to_py(js_snippet, add_new_cell=True, import_ee=True, import_geemap=True, show_map=True)
# You can also convert GEE JavaScript to Python without coding.
#
# 
Map = geemap.Map()
Map
# ## Earth Engine datasets
#
# ### Load Earth Engine datasets
# +
Map = geemap.Map()
# Add Earth Engine datasets
dem = ee.Image('USGS/SRTMGL1_003')
landcover = ee.Image("ESA/GLOBCOVER_L4_200901_200912_V2_3").select('landcover')
landsat7 = ee.Image('LE7_TOA_5YEAR/1999_2003')
states = ee.FeatureCollection("TIGER/2018/States")
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Add Earth Eninge layers to Map
Map.addLayer(dem, vis_params, 'SRTM DEM', True, 0.5)
Map.addLayer(landcover, {}, 'Land cover')
Map.addLayer(landsat7, {'bands': ['B4', 'B3', 'B2'], 'min': 20, 'max': 200, 'gamma': 1.5}, 'Landsat 7')
Map.addLayer(states, {}, "US States")
Map
# -
# ### Search the Earth Engine Data Catalog
Map = geemap.Map()
Map
dem = ee.Image('CGIAR/SRTM90_V4')
Map.addLayer(dem, {}, "CGIAR/SRTM90_V4")
# +
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
Map.addLayer(dem, vis_params, "DEM")
# -
# ### Use the datasets module
from geemap.datasets import DATA
# +
Map = geemap.Map()
dem = ee.Image(DATA.USGS_SRTMGL1_003)
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
Map.addLayer(dem, vis_params, 'SRTM DEM')
Map
# -
# ### Use the Inspector tool
# 
# +
Map = geemap.Map()
# Add Earth Engine datasets
dem = ee.Image('USGS/SRTMGL1_003')
landcover = ee.Image("ESA/GLOBCOVER_L4_200901_200912_V2_3").select('landcover')
landsat7 = ee.Image('LE7_TOA_5YEAR/1999_2003').select(['B1', 'B2', 'B3', 'B4', 'B5', 'B7'])
states = ee.FeatureCollection("TIGER/2018/States")
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Add Earth Eninge layers to Map
Map.addLayer(dem, vis_params, 'SRTM DEM', True, 0.5)
Map.addLayer(landcover, {}, 'Land cover')
Map.addLayer(landsat7, {'bands': ['B4', 'B3', 'B2'], 'min': 20, 'max': 200, 'gamma': 1.5}, 'Landsat 7')
Map.addLayer(states, {}, "US States")
Map
# -
# ## Data visualization
#
# ### Use the Plotting tool
# 
# +
Map = geemap.Map()
landsat7 = ee.Image('LE7_TOA_5YEAR/1999_2003').select(['B1', 'B2', 'B3', 'B4', 'B5', 'B7'])
landsat_vis = {
'bands': ['B4', 'B3', 'B2'],
'gamma': 1.4
}
Map.addLayer(landsat7, landsat_vis, "Landsat")
hyperion = ee.ImageCollection('EO1/HYPERION') \
.filter(ee.Filter.date('2016-01-01', '2017-03-01'))
hyperion_vis = {
'min': 1000.0,
'max': 14000.0,
'gamma': 2.5,
}
Map.addLayer(hyperion, hyperion_vis, 'Hyperion')
Map
# -
# ### Change layer opacity
# +
Map = geemap.Map(center=(40, -100), zoom=4)
dem = ee.Image('USGS/SRTMGL1_003')
states = ee.FeatureCollection("TIGER/2018/States")
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
Map.addLayer(dem, vis_params, 'SRTM DEM', True, 1)
Map.addLayer(states, {}, "US States", True)
Map
# -
# ### Visualize raster data
# +
Map = geemap.Map(center=(40, -100), zoom=4)
# Add Earth Engine dataset
dem = ee.Image('USGS/SRTMGL1_003')
landsat7 = ee.Image('LE7_TOA_5YEAR/1999_2003').select(['B1', 'B2', 'B3', 'B4', 'B5', 'B7'])
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
Map.addLayer(dem, vis_params, 'SRTM DEM', True, 1)
Map.addLayer(landsat7, {'bands': ['B4', 'B3', 'B2'], 'min': 20, 'max': 200, 'gamma': 2}, 'Landsat 7')
Map
# -
# ### Visualize vector data
# +
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
Map.addLayer(states, {}, "US States")
Map
# +
vis_params = {
'color': '000000',
'colorOpacity': 1,
'pointSize': 3,
'pointShape': 'circle',
'width': 2,
'lineType': 'solid',
'fillColorOpacity': 0.66
}
palette = ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']
Map.add_styled_vector(states, column="NAME", palette=palette, layer_name="Styled vector", **vis_params)
# -
# ### Add a legend
legends = geemap.builtin_legends
for legend in legends:
print(legend)
Map = geemap.Map()
Map.add_basemap('HYBRID')
landcover = ee.Image('USGS/NLCD/NLCD2016').select('landcover')
Map.addLayer(landcover, {}, 'NLCD Land Cover')
Map.add_legend(builtin_legend='NLCD')
Map
# +
Map = geemap.Map()
legend_dict = {
'11 Open Water': '466b9f',
'12 Perennial Ice/Snow': 'd1def8',
'21 Developed, Open Space': 'dec5c5',
'22 Developed, Low Intensity': 'd99282',
'23 Developed, Medium Intensity': 'eb0000',
'24 Developed High Intensity': 'ab0000',
'31 Barren Land (Rock/Sand/Clay)': 'b3ac9f',
'41 Deciduous Forest': '68ab5f',
'42 Evergreen Forest': '1c5f2c',
'43 Mixed Forest': 'b5c58f',
'51 Dwarf Scrub': 'af963c',
'52 Shrub/Scrub': 'ccb879',
'71 Grassland/Herbaceous': 'dfdfc2',
'72 Sedge/Herbaceous': 'd1d182',
'73 Lichens': 'a3cc51',
'74 Moss': '82ba9e',
'81 Pasture/Hay': 'dcd939',
'82 Cultivated Crops': 'ab6c28',
'90 Woody Wetlands': 'b8d9eb',
'95 Emergent Herbaceous Wetlands': '6c9fb8'
}
landcover = ee.Image('USGS/NLCD/NLCD2016').select('landcover')
Map.addLayer(landcover, {}, 'NLCD Land Cover')
Map.add_legend(legend_title="NLCD Land Cover Classification", legend_dict=legend_dict)
Map
# -
# ### Add a colorbar
# +
Map = geemap.Map()
dem = ee.Image('USGS/SRTMGL1_003')
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
Map.addLayer(dem, vis_params, 'SRTM DEM')
colors = vis_params['palette']
vmin = vis_params['min']
vmax = vis_params['max']
Map.add_colorbar(vis_params, label="Elevation (m)", layer_name="SRTM DEM")
Map
# -
Map.add_colorbar(vis_params, label="Elevation (m)", layer_name="SRTM DEM", orientation="vertical")
Map.add_colorbar(vis_params, label="Elevation (m)", layer_name="SRTM DEM", orientation="vertical", transparent_bg=True)
Map.add_colorbar(vis_params, label="Elevation (m)", layer_name="SRTM DEM", orientation="vertical", transparent_bg=True, discrete=True)
# ### Create a split-panel map
Map = geemap.Map()
Map.split_map(left_layer='HYBRID', right_layer='TERRAIN')
Map
Map = geemap.Map()
Map.split_map(left_layer='NLCD 2016 CONUS Land Cover', right_layer='NLCD 2001 CONUS Land Cover')
Map
# +
nlcd_2001 = ee.Image('USGS/NLCD/NLCD2001').select('landcover')
nlcd_2016 = ee.Image('USGS/NLCD/NLCD2016').select('landcover')
left_layer = geemap.ee_tile_layer(nlcd_2001, {}, 'NLCD 2001')
right_layer = geemap.ee_tile_layer(nlcd_2016, {}, 'NLCD 2016')
Map = geemap.Map()
Map.split_map(left_layer, right_layer)
Map
# -
# ### Create linked maps
# +
image = ee.ImageCollection('COPERNICUS/S2') \
.filterDate('2018-09-01', '2018-09-30') \
.map(lambda img: img.divide(10000)) \
.median()
vis_params = [
{'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.3},
{'bands': ['B8', 'B11', 'B4'], 'min': 0, 'max': 0.3, 'gamma': 1.3},
{'bands': ['B8', 'B4', 'B3'], 'min': 0, 'max': 0.3, 'gamma': 1.3},
{'bands': ['B12', 'B12', 'B4'], 'min': 0, 'max': 0.3, 'gamma': 1.3}
]
labels = [
'Natural Color (B4/B3/B2)',
'Land/Water (B8/B11/B4)',
'Color Infrared (B8/B4/B3)',
'Vegetation (B12/B11/B4)'
]
geemap.linked_maps(rows=2, cols=2, height="400px", center=[38.4151, 21.2712], zoom=12,
ee_objects=[image], vis_params=vis_params, labels=labels, label_position="topright")
# -
# ### Create timelapse animations
geemap.show_youtube('https://youtu.be/mA21Us_3m28')
# ### Create time-series composites
geemap.show_youtube('https://youtu.be/kEltQkNia6o')
# ## Data analysis
# ### Descriptive statistics
# +
Map = geemap.Map()
centroid = ee.Geometry.Point([-122.4439, 37.7538])
image = ee.ImageCollection('LANDSAT/LC08/C01/T1_SR') \
.filterBounds(centroid) \
.first()
vis = {
'min': 0,
'max': 3000,
'bands': ['B5', 'B4', 'B3']
}
Map.centerObject(centroid, 8)
Map.addLayer(image, vis, "Landsat-8")
Map
# -
image.propertyNames().getInfo()
image.get('CLOUD_COVER').getInfo()
props = geemap.image_props(image)
props.getInfo()
stats = geemap.image_stats(image, scale=90)
stats.getInfo()
# ### Zonal statistics
# +
Map = geemap.Map()
# Add Earth Engine dataset
dem = ee.Image('USGS/SRTMGL1_003')
# Set visualization parameters.
dem_vis = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Add Earth Engine DEM to map
Map.addLayer(dem, dem_vis, 'SRTM DEM')
# Add Landsat data to map
landsat = ee.Image('LE7_TOA_5YEAR/1999_2003')
landsat_vis = {
'bands': ['B4', 'B3', 'B2'],
'gamma': 1.4
}
Map.addLayer(landsat, landsat_vis, "LE7_TOA_5YEAR/1999_2003")
states = ee.FeatureCollection("TIGER/2018/States")
Map.addLayer(states, {}, 'US States')
Map
# +
out_dir = os.path.expanduser('~/Downloads')
out_dem_stats = os.path.join(out_dir, 'dem_stats.csv')
if not os.path.exists(out_dir):
os.makedirs(out_dir)
# Allowed output formats: csv, shp, json, kml, kmz
# Allowed statistics type: MEAN, MAXIMUM, MINIMUM, MEDIAN, STD, MIN_MAX, VARIANCE, SUM
geemap.zonal_statistics(dem, states, out_dem_stats, statistics_type='MEAN', scale=1000)
# -
out_landsat_stats = os.path.join(out_dir, 'landsat_stats.csv')
geemap.zonal_statistics(landsat, states, out_landsat_stats, statistics_type='SUM', scale=1000)
# ### Zonal statistics by group
# +
Map = geemap.Map()
dataset = ee.Image('USGS/NLCD/NLCD2016')
landcover = ee.Image(dataset.select('landcover'))
Map.addLayer(landcover, {}, 'NLCD 2016')
states = ee.FeatureCollection("TIGER/2018/States")
Map.addLayer(states, {}, 'US States')
Map.add_legend(builtin_legend='NLCD')
Map
# +
out_dir = os.path.expanduser('~/Downloads')
nlcd_stats = os.path.join(out_dir, 'nlcd_stats.csv')
if not os.path.exists(out_dir):
os.makedirs(out_dir)
# statistics_type can be either 'SUM' or 'PERCENTAGE'
# denominator can be used to convert square meters to other areal units, such as square kilimeters
geemap.zonal_statistics_by_group(landcover, states, nlcd_stats, statistics_type='SUM', denominator=1000000, decimal_places=2)
# -
# ### Unsupervised classification
#
# Source: https://developers.google.com/earth-engine/guides/clustering
#
# The `ee.Clusterer` package handles unsupervised classification (or clustering) in Earth Engine. These algorithms are currently based on the algorithms with the same name in [Weka](http://www.cs.waikato.ac.nz/ml/weka/). More details about each Clusterer are available in the reference docs in the Code Editor.
#
# Clusterers are used in the same manner as classifiers in Earth Engine. The general workflow for clustering is:
#
# 1. Assemble features with numeric properties in which to find clusters.
# 2. Instantiate a clusterer. Set its parameters if necessary.
# 3. Train the clusterer using the training data.
# 4. Apply the clusterer to an image or feature collection.
# 5. Label the clusters.
#
# The training data is a `FeatureCollection` with properties that will be input to the clusterer. Unlike classifiers, there is no input class value for an `Clusterer`. Like classifiers, the data for the train and apply steps are expected to have the same number of values. When a trained clusterer is applied to an image or table, it assigns an integer cluster ID to each pixel or feature.
#
# Here is a simple example of building and using an ee.Clusterer:
#
# 
# **Add data to the map**
# +
Map = geemap.Map()
point = ee.Geometry.Point([-87.7719, 41.8799])
image = ee.ImageCollection('LANDSAT/LC08/C01/T1_SR') \
.filterBounds(point) \
.filterDate('2019-01-01', '2019-12-31') \
.sort('CLOUD_COVER') \
.first() \
.select('B[1-7]')
vis_params = {
'min': 0,
'max': 3000,
'bands': ['B5', 'B4', 'B3']
}
Map.centerObject(point, 8)
Map.addLayer(image, vis_params, "Landsat-8")
Map
# -
# **Make training dataset**
#
# There are several ways you can create a region for generating the training dataset.
#
# - Draw a shape (e.g., rectangle) on the map and the use `region = Map.user_roi`
# - Define a geometry, such as `region = ee.Geometry.Rectangle([-122.6003, 37.4831, -121.8036, 37.8288])`
# - Create a buffer zone around a point, such as `region = ee.Geometry.Point([-122.4439, 37.7538]).buffer(10000)`
# - If you don't define a region, it will use the image footprint by default
# +
training = image.sample(**{
# 'region': region,
'scale': 30,
'numPixels': 5000,
'seed': 0,
'geometries': True # Set this to False to ignore geometries
})
Map.addLayer(training, {}, 'training', False)
# -
# **Train the clusterer**
# Instantiate the clusterer and train it.
n_clusters = 5
clusterer = ee.Clusterer.wekaKMeans(n_clusters).train(training)
# **Classify the image**
# +
# Cluster the input using the trained clusterer.
result = image.cluster(clusterer)
# # Display the clusters with random colors.
Map.addLayer(result.randomVisualizer(), {}, 'clusters')
Map
# -
# **Label the clusters**
# +
legend_keys = ['One', 'Two', 'Three', 'Four', 'ect']
legend_colors = ['#8DD3C7', '#FFFFB3', '#BEBADA', '#FB8072', '#80B1D3']
# Reclassify the map
result = result.remap([0, 1, 2, 3, 4], [1, 2, 3, 4, 5])
Map.addLayer(result, {'min': 1, 'max': 5, 'palette': legend_colors}, 'Labelled clusters')
Map.add_legend(legend_keys=legend_keys, legend_colors=legend_colors, position='bottomright')
# -
# **Visualize the result**
print('Change layer opacity:')
cluster_layer = Map.layers[-1]
cluster_layer.interact(opacity=(0, 1, 0.1))
Map
# **Export the result**
out_dir = os.path.expanduser('~/Downloads')
out_file = os.path.join(out_dir, 'cluster.tif')
geemap.ee_export_image(result, filename=out_file, scale=90)
# +
# geemap.ee_export_image_to_drive(result, description='clusters', folder='export', scale=90)
# -
# ### Supervised classification
# Source: https://developers.google.com/earth-engine/guides/classification
#
# The `Classifier` package handles supervised classification by traditional ML algorithms running in Earth Engine. These classifiers include CART, RandomForest, NaiveBayes and SVM. The general workflow for classification is:
#
# 1. Collect training data. Assemble features which have a property that stores the known class label and properties storing numeric values for the predictors.
# 2. Instantiate a classifier. Set its parameters if necessary.
# 3. Train the classifier using the training data.
# 4. Classify an image or feature collection.
# 5. Estimate classification error with independent validation data.
#
# The training data is a `FeatureCollection` with a property storing the class label and properties storing predictor variables. Class labels should be consecutive, integers starting from 0. If necessary, use remap() to convert class values to consecutive integers. The predictors should be numeric.
#
# 
# **Add data to the map**
# +
Map = geemap.Map()
point = ee.Geometry.Point([-122.4439, 37.7538])
image = ee.ImageCollection('LANDSAT/LC08/C01/T1_SR') \
.filterBounds(point) \
.filterDate('2016-01-01', '2016-12-31') \
.sort('CLOUD_COVER') \
.first() \
.select('B[1-7]')
vis_params = {
'min': 0,
'max': 3000,
'bands': ['B5', 'B4', 'B3']
}
Map.centerObject(point, 8)
Map.addLayer(image, vis_params, "Landsat-8")
Map
# -
# **Make training dataset**
#
# There are several ways you can create a region for generating the training dataset.
#
# - Draw a shape (e.g., rectangle) on the map and the use `region = Map.user_roi`
# - Define a geometry, such as `region = ee.Geometry.Rectangle([-122.6003, 37.4831, -121.8036, 37.8288])`
# - Create a buffer zone around a point, such as `region = ee.Geometry.Point([-122.4439, 37.7538]).buffer(10000)`
# - If you don't define a region, it will use the image footprint by default
# +
# region = Map.user_roi
# region = ee.Geometry.Rectangle([-122.6003, 37.4831, -121.8036, 37.8288])
# region = ee.Geometry.Point([-122.4439, 37.7538]).buffer(10000)
# -
# In this example, we are going to use the [USGS National Land Cover Database (NLCD)](https://developers.google.com/earth-engine/datasets/catalog/USGS_NLCD) to create label dataset for training
#
#
# 
nlcd = ee.Image('USGS/NLCD/NLCD2016').select('landcover').clip(image.geometry())
Map.addLayer(nlcd, {}, 'NLCD')
Map
# +
# Make the training dataset.
points = nlcd.sample(**{
'region': image.geometry(),
'scale': 30,
'numPixels': 5000,
'seed': 0,
'geometries': True # Set this to False to ignore geometries
})
Map.addLayer(points, {}, 'training', False)
# -
# **Train the classifier**
# +
# Use these bands for prediction.
bands = ['B1', 'B2', 'B3', 'B4', 'B5', 'B6', 'B7']
# This property of the table stores the land cover labels.
label = 'landcover'
# Overlay the points on the imagery to get training.
training = image.select(bands).sampleRegions(**{
'collection': points,
'properties': [label],
'scale': 30
})
# Train a CART classifier with default parameters.
trained = ee.Classifier.smileCart().train(training, label, bands)
# -
# **Classify the image**
# +
# Classify the image with the same bands used for training.
result = image.select(bands).classify(trained)
# # Display the clusters with random colors.
Map.addLayer(result.randomVisualizer(), {}, 'classfied')
Map
# -
# **Render categorical map**
#
# To render a categorical map, we can set two image properties: `landcover_class_values` and `landcover_class_palette`. We can use the same style as the NLCD so that it is easy to compare the two maps.
class_values = nlcd.get('landcover_class_values').getInfo()
class_palette = nlcd.get('landcover_class_palette').getInfo()
landcover = result.set('classification_class_values', class_values)
landcover = landcover.set('classification_class_palette', class_palette)
Map.addLayer(landcover, {}, 'Land cover')
Map.add_legend(builtin_legend='NLCD')
Map
# **Visualize the result**
print('Change layer opacity:')
cluster_layer = Map.layers[-1]
cluster_layer.interact(opacity=(0, 1, 0.1))
# **Export the result**
out_dir = os.path.expanduser('~/Downloads')
out_file = os.path.join(out_dir, 'landcover.tif')
geemap.ee_export_image(landcover, filename=out_file, scale=900)
# +
# geemap.ee_export_image_to_drive(landcover, description='landcover', folder='export', scale=900)
# -
# ### Training sample creation
#
# 
geemap.show_youtube('https://youtu.be/VWh5PxXPZw0')
Map = geemap.Map()
Map
# ### WhiteboxTools
import whiteboxgui
whiteboxgui.show()
whiteboxgui.show(tree=True)
# 
Map = geemap.Map()
Map
# ## Map making
# ### Plot a single band image
import matplotlib.pyplot as plt
from geemap import cartoee
geemap.ee_initialize()
srtm = ee.Image("CGIAR/SRTM90_V4")
region = [-180, -60, 180, 85] # define bounding box to request data
vis = {'min':0, 'max':3000} # define visualization parameters for image
# +
fig = plt.figure(figsize=(15, 10))
cmap = "gist_earth" # colormap we want to use
# cmap = "terrain"
# use cartoee to get a map
ax = cartoee.get_map(srtm, region=region, vis_params=vis, cmap=cmap)
# add a colorbar to the map using the visualization params we passed to the map
cartoee.add_colorbar(ax, vis, cmap=cmap, loc="right", label="Elevation", orientation="vertical")
# add gridlines to the map at a specified interval
cartoee.add_gridlines(ax, interval=[60,30], linestyle="--")
# add coastlines using the cartopy api
ax.coastlines(color="red")
ax.set_title(label = 'Global Elevation Map', fontsize=15)
plt.show()
# -
# ### Plot an RGB image
# +
# get a landsat image to visualize
image = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_044034_20140318')
# define the visualization parameters to view
vis ={"bands": ['B5', 'B4', 'B3'], "min": 0, "max":5000, "gamma":1.3}
# +
fig = plt.figure(figsize=(15, 10))
# here is the bounding box of the map extent we want to use
# formatted a [W,S,E,N]
zoom_region = [-122.6265, 37.3458, -121.8025, 37.9178]
# plot the map over the region of interest
ax = cartoee.get_map(image, vis_params=vis, region=zoom_region)
# add the gridlines and specify that the xtick labels be rotated 45 degrees
cartoee.add_gridlines(ax, interval=0.15, xtick_rotation=45, linestyle=":")
# add coastline
ax.coastlines(color="yellow")
# add north arrow
cartoee.add_north_arrow(ax, text="N", xy=(0.05, 0.25), text_color="white", arrow_color="white", fontsize=20)
# add scale bar
cartoee.add_scale_bar_lite(ax, length=10, xy=(0.1, 0.05), fontsize=20, color="white", unit="km")
ax.set_title(label = 'Landsat False Color Composite (Band 5/4/3)', fontsize=15)
plt.show()
# -
# ### Add map elements
from matplotlib.lines import Line2D
# +
# get a landsat image to visualize
image = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_044034_20140318')
# define the visualization parameters to view
vis ={"bands": ['B5', 'B4', 'B3'], "min": 0, "max":5000, "gamma":1.3}
# +
fig = plt.figure(figsize=(15, 10))
# here is the bounding box of the map extent we want to use
# formatted a [W,S,E,N]
zoom_region = [-122.6265, 37.3458, -121.8025, 37.9178]
# plot the map over the region of interest
ax = cartoee.get_map(image, vis_params=vis, region=zoom_region)
# add the gridlines and specify that the xtick labels be rotated 45 degrees
cartoee.add_gridlines(ax, interval=0.15, xtick_rotation=0, linestyle=":")
# add coastline
ax.coastlines(color="cyan")
# add north arrow
cartoee.add_north_arrow(ax, text="N", xy=(0.05, 0.25), text_color="white", arrow_color="white", fontsize=20)
# add scale bar
cartoee.add_scale_bar_lite(ax, length=10, xy=(0.1, 0.05), fontsize=20, color="white", unit="km")
ax.set_title(label = 'Landsat False Color Composite (Band 5/4/3)', fontsize=15)
# add legend
legend_elements = [Line2D([], [], color='#00ffff', lw=2, label='Coastline'),
Line2D([], [], marker='o', color='#A8321D', label='City', markerfacecolor='#A8321D', markersize=10, ls ='')]
cartoee.add_legend(ax, legend_elements, loc='lower right')
plt.show()
# -
# ### Plot multiple layers
# +
Map = geemap.Map()
image = ee.ImageCollection('MODIS/MCD43A4_006_NDVI') \
.filter(ee.Filter.date('2018-04-01', '2018-05-01')) \
.select("NDVI")\
.first()
vis_params = {
'min': 0.0,
'max': 1.0,
'palette': [
'FFFFFF', 'CE7E45', 'DF923D', 'F1B555', 'FCD163', '99B718', '74A901',
'66A000', '529400', '3E8601', '207401', '056201', '004C00', '023B01',
'012E01', '011D01', '011301'
],
}
Map.setCenter(-7.03125, 31.0529339857, 2)
Map.addLayer(image, vis_params, 'MODIS NDVI')
countries = geemap.shp_to_ee("../data/countries.shp")
style = {
"color": "00000088",
"width": 1,
"fillColor": "00000000"}
Map.addLayer(countries.style(**style), {}, "Countries")
ndvi = image.visualize(**vis_params)
blend = ndvi.blend(countries.style(**style))
Map.addLayer(blend, {}, "Blend")
Map
# -
# specify region to focus on
bbox = [-180, -88, 180, 88]
# +
fig = plt.figure(figsize=(15,10))
# plot the result with cartoee using a PlateCarre projection (default)
ax = cartoee.get_map(blend, region=bbox)
cb = cartoee.add_colorbar(ax, vis_params=vis_params, loc='right')
ax.set_title(label = 'MODIS NDVI', fontsize = 15)
# ax.coastlines()
plt.show()
# -
import cartopy.crs as ccrs
# +
fig = plt.figure(figsize=(15,10))
projection = ccrs.EqualEarth(central_longitude=-180)
# plot the result with cartoee using a PlateCarre projection (default)
ax = cartoee.get_map(blend, region=bbox, proj=projection)
cb = cartoee.add_colorbar(ax, vis_params=vis_params, loc='right')
ax.set_title(label = 'MODIS NDVI', fontsize = 15)
# ax.coastlines()
plt.show()
# -
# ### Use custom projections
import cartopy.crs as ccrs
# get an earth engine image of ocean data for Jan-Mar 2018
ocean = (
ee.ImageCollection('NASA/OCEANDATA/MODIS-Terra/L3SMI')
.filter(ee.Filter.date('2018-01-01', '2018-03-01'))
.median()
.select(["sst"], ["SST"])
)
# set parameters for plotting
# will plot the Sea Surface Temp with specific range and colormap
visualization = {'bands':"SST", 'min':-2, 'max':30}
# specify region to focus on
bbox = [-180, -88, 180, 88]
# +
fig = plt.figure(figsize=(15,10))
# plot the result with cartoee using a PlateCarre projection (default)
ax = cartoee.get_map(ocean, cmap='plasma', vis_params=visualization, region=bbox)
cb = cartoee.add_colorbar(ax, vis_params=visualization, loc='right', cmap='plasma')
ax.set_title(label = 'Sea Surface Temperature', fontsize = 15)
ax.coastlines()
plt.show()
# +
fig = plt.figure(figsize=(15,10))
# create a new Mollweide projection centered on the Pacific
projection = ccrs.Mollweide(central_longitude=-180)
# plot the result with cartoee using the Mollweide projection
ax = cartoee.get_map(ocean, vis_params=visualization, region=bbox,
cmap='plasma', proj=projection)
cb = cartoee.add_colorbar(ax,vis_params=visualization, loc='bottom', cmap='plasma',
orientation='horizontal')
ax.set_title("Mollweide projection")
ax.coastlines()
plt.show()
# +
fig = plt.figure(figsize=(15,10))
# create a new Robinson projection centered on the Pacific
projection = ccrs.Robinson(central_longitude=-180)
# plot the result with cartoee using the Goode homolosine projection
ax = cartoee.get_map(ocean, vis_params=visualization, region=bbox,
cmap='plasma', proj=projection)
cb = cartoee.add_colorbar(ax, vis_params=visualization, loc='bottom', cmap='plasma',
orientation='horizontal')
ax.set_title("Robinson projection")
ax.coastlines()
plt.show()
# +
fig = plt.figure(figsize=(15,10))
# create a new equal Earth projection focused on the Pacific
projection = ccrs.EqualEarth(central_longitude=-180)
# plot the result with cartoee using the orographic projection
ax = cartoee.get_map(ocean, vis_params=visualization, region=bbox,
cmap='plasma', proj=projection)
cb = cartoee.add_colorbar(ax, vis_params=visualization, loc='right', cmap='plasma',
orientation='vertical')
ax.set_title("Equal Earth projection")
ax.coastlines()
plt.show()
# +
fig = plt.figure(figsize=(15,10))
# create a new orographic projection focused on the Pacific
projection = ccrs.Orthographic(-130,-10)
# plot the result with cartoee using the orographic projection
ax = cartoee.get_map(ocean, vis_params=visualization, region=bbox,
cmap='plasma', proj=projection)
cb = cartoee.add_colorbar(ax, vis_params=visualization, loc='right', cmap='plasma',
orientation='vertical')
ax.set_title("Orographic projection")
ax.coastlines()
plt.show()
# -
# ### Create timelapse animations
# +
Map = geemap.Map()
lon = -115.1585
lat = 36.1500
start_year = 1984
end_year = 2000
point = ee.Geometry.Point(lon, lat)
years = ee.List.sequence(start_year, end_year)
def get_best_image(year):
start_date = ee.Date.fromYMD(year, 1, 1)
end_date = ee.Date.fromYMD(year, 12, 31)
image = ee.ImageCollection("LANDSAT/LT05/C01/T1_SR") \
.filterBounds(point) \
.filterDate(start_date, end_date) \
.sort("CLOUD_COVER") \
.first()
return ee.Image(image)
collection = ee.ImageCollection(years.map(get_best_image))
vis_params = {
"bands": ['B4', 'B3', 'B2'],
"min": 0,
"max": 5000
}
image = ee.Image(collection.first())
Map.addLayer(image, vis_params, 'First image')
Map.setCenter(lon, lat, 8)
Map
# +
w = 0.4
h = 0.3
region = [lon-w, lat-h, lon+w, lat+h]
fig = plt.figure(figsize=(10, 8))
# use cartoee to get a map
ax = cartoee.get_map(image, region=region, vis_params=vis_params)
# add gridlines to the map at a specified interval
cartoee.add_gridlines(ax, interval=[0.2, 0.2], linestyle=":")
# add north arrow
north_arrow_dict = {
"text": "N",
"xy": (0.1, 0.3),
"arrow_length": 0.15,
"text_color": "white",
"arrow_color": "white",
"fontsize": 20,
"width": 5,
"headwidth": 15,
"ha": "center",
"va": "center"
}
cartoee.add_north_arrow(ax, **north_arrow_dict)
# add scale bar
scale_bar_dict = {
"length": 10,
"xy": (0.1, 0.05),
"linewidth": 3,
"fontsize": 20,
"color": "white",
"unit": "km",
"ha": "center",
"va": "bottom"
}
cartoee.add_scale_bar_lite(ax, **scale_bar_dict)
ax.set_title(label = 'Las Vegas, NV', fontsize=15)
plt.show()
# -
cartoee.get_image_collection_gif(
ee_ic = collection,
out_dir = os.path.expanduser("~/Downloads/timelapse"),
out_gif = "animation.gif",
vis_params = vis_params,
region = region,
fps = 5,
mp4 = True,
grid_interval = (0.2, 0.2),
plot_title = "Las Vegas, NV",
date_format = 'YYYY-MM-dd',
fig_size = (10, 8),
dpi_plot = 100,
file_format = "png",
north_arrow_dict = north_arrow_dict,
scale_bar_dict = scale_bar_dict,
verbose = True
)
# ## Data export
# ### Export ee.Image
# +
Map = geemap.Map()
image = ee.Image('LE7_TOA_5YEAR/1999_2003')
landsat_vis = {
'bands': ['B4', 'B3', 'B2'],
'gamma': 1.4
}
Map.addLayer(image, landsat_vis, "LE7_TOA_5YEAR/1999_2003", True, 1)
Map
# +
# Draw any shapes on the map using the Drawing tools before executing this code block
roi = Map.user_roi
if roi is None:
roi = ee.Geometry.Polygon([[[-115.413031, 35.889467],
[-115.413031, 36.543157],
[-114.034328, 36.543157],
[-114.034328, 35.889467],
[-115.413031, 35.889467]]])
# +
# Set output directory
out_dir = os.path.expanduser('~/Downloads')
if not os.path.exists(out_dir):
os.makedirs(out_dir)
filename = os.path.join(out_dir, 'landsat.tif')
# -
# Exporting all bands as one single image
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=90, region=roi, file_per_band=False)
# Exporting each band as one image
geemap.ee_export_image(image, filename=filename, scale=90, region=roi, file_per_band=True)
# Export an image to Google Drive¶
# +
# geemap.ee_export_image_to_drive(image, description='landsat', folder='export', region=roi, scale=30)
# -
# ### Export ee.ImageCollection
loc = ee.Geometry.Point(-99.2222, 46.7816)
collection = ee.ImageCollection('USDA/NAIP/DOQQ') \
.filterBounds(loc) \
.filterDate('2008-01-01', '2020-01-01') \
.filter(ee.Filter.listContains("system:band_names", "N"))
collection.aggregate_array('system:index').getInfo()
geemap.ee_export_image_collection(collection, out_dir=out_dir)
# +
# geemap.ee_export_image_collection_to_drive(collection, folder='export', scale=10)
# -
# ### Extract pixels as a numpy array
# +
import matplotlib.pyplot as plt
img = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_038029_20180810') \
.select(['B4', 'B5', 'B6'])
aoi = ee.Geometry.Polygon(
[[[-110.8, 44.7],
[-110.8, 44.6],
[-110.6, 44.6],
[-110.6, 44.7]]], None, False)
rgb_img = geemap.ee_to_numpy(img, region=aoi)
print(rgb_img.shape)
# -
rgb_img_test = (255*((rgb_img[:, :, 0:3] - 100)/3500)).astype('uint8')
plt.imshow(rgb_img_test)
plt.show()
# ### Export pixel values to points
# +
Map = geemap.Map()
# Add Earth Engine dataset
dem = ee.Image('USGS/SRTMGL1_003')
landsat7 = ee.Image('LE7_TOA_5YEAR/1999_2003')
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Add Earth Eninge layers to Map
Map.addLayer(landsat7, {'bands': ['B4', 'B3', 'B2'], 'min': 20, 'max': 200}, 'Landsat 7')
Map.addLayer(dem, vis_params, 'SRTM DEM', True, 1)
Map
# -
# **Download sample data**
work_dir = os.path.expanduser('~/Downloads')
in_shp = os.path.join(work_dir, 'us_cities.shp')
if not os.path.exists(in_shp):
data_url = 'https://github.com/giswqs/data/raw/main/us/us_cities.zip'
geemap.download_from_url(data_url, out_dir=work_dir )
in_fc = geemap.shp_to_ee(in_shp)
Map.addLayer(in_fc, {}, 'Cities')
# **Export pixel values as a shapefile**
out_shp = os.path.join(work_dir, 'dem.shp')
geemap.extract_values_to_points(in_fc, dem, out_shp)
# **Export pixel values as a csv**
out_csv = os.path.join(work_dir, 'landsat.csv')
geemap.extract_values_to_points(in_fc, landsat7, out_csv)
# ### Export ee.FeatureCollection
# +
Map = geemap.Map()
fc = ee.FeatureCollection('users/giswqs/public/countries')
Map.addLayer(fc, {}, "Countries")
Map
# -
out_dir = os.path.expanduser('~/Downloads')
out_shp = os.path.join(out_dir, 'countries.shp')
geemap.ee_to_shp(fc, filename=out_shp)
out_csv = os.path.join(out_dir, 'countries.csv')
geemap.ee_export_vector(fc, filename=out_csv)
out_kml = os.path.join(out_dir, 'countries.kml')
geemap.ee_export_vector(fc, filename=out_kml)
# +
# geemap.ee_export_vector_to_drive(fc, description="countries", folder="export", file_format="shp")
# -
# ## Web apps
# ### Deploy web apps using ngrok
# **Steps to deploy an Earth Engine App:**
# 1. Install ngrok by following the [instruction](https://ngrok.com/download)
# 3. Download the notebook [71_timelapse.ipynb](https://geemap.org/notebooks/71_timelapse/71_timelapse.ipynb)
# 4. Run this from the command line: `voila --no-browser 71_timelapse.ipynb`
# 5. Run this from the command line: `ngrok http 8866`
# 6. Copy the link from the ngrok terminal window. The links looks like the following: https://randomstring.ngrok.io
# 7. Share the link with anyone.
#
# **Optional steps:**
# * To show code cells from you app, run this from the command line: `voila --no-browser --strip_sources=False 71_timelapse.ipynb`
# * To protect your app with a password, run this: `ngrok http -auth="username:password" 8866`
# * To run python simple http server in the directory, run this:`sudo python -m http.server 80`
geemap.show_youtube("https://youtu.be/eRDZBVJcNCk")
# ### Deploy web apps using Heroku
# **Steps to deploy an Earth Engine App:**
#
# - [Sign up](https://signup.heroku.com/) for a free heroku account.
# - Follow the [instructions](https://devcenter.heroku.com/articles/getting-started-with-python#set-up) to install [Git](https://git-scm.com/book/en/v2/Getting-Started-Installing-Git) and Heroku Command Line Interface (CLI).
# - Authenticate heroku using the `heroku login` command.
# - Clone this repository: <https://github.com/giswqs/geemap-heroku>
# - Create your own Earth Engine notebook and put it under the `notebooks` directory.
# - Add Python dependencies in the `requirements.txt` file if needed.
# - Edit the `Procfile` file by replacing `notebooks/geemap.ipynb` with the path to your own notebook.
# - Commit changes to the repository by using `git add . && git commit -am "message"`.
# - Create a heroku app: `heroku create`
# - Run the `config_vars.py` script to extract Earth Engine token from your computer and set it as an environment variable on heroku: `python config_vars.py`
# - Deploy your code to heroku: `git push heroku master`
# - Open your heroku app: `heroku open`
#
# **Optional steps:**
#
# - To specify a name for your app, use `heroku apps:create example`
# - To preview your app locally, use `heroku local web`
# - To hide code cells from your app, you can edit the `Procfile` file and set `--strip_sources=True`
# - To periodically check for idle kernels, you can edit the `Procfile` file and set `--MappingKernelManager.cull_interval=60 --MappingKernelManager.cull_idle_timeout=120`
# - To view information about your running app, use `heroku logs --tail`
# - To set an environment variable on heroku, use `heroku config:set NAME=VALUE`
# - To view environment variables for your app, use `heroku config`
geemap.show_youtube("https://youtu.be/nsIjfD83ggA")
| examples/workshops/GeoPython_2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nlp_env
# language: python
# name: nlp_env
# ---
# # Using `BertNLU`
# +
# imports
from dialognlu import BertNLU, AutoNLU
from dialognlu.readers.goo_format_reader import Reader
import ipywidgets as widgets
import json
# -
# ## Reading Training and Validation Datasets
# +
train_path = "../data/snips/train" # please, specify the path to your dataset
val_path = "../data/snips/valid" # please, specify the path to your dataset
train_dataset = Reader.read(train_path)
val_dataset = Reader.read(val_path)
# -
# ## Initialize `BertNLU` model
# Choose `model_type`
model_type_wid = widgets.Dropdown(options=['bert', 'albert'], value='bert', description='model_type:',
disabled=False)
display(model_type_wid)
# +
model_type = model_type_wid.value
config = {
"model_type": model_type #"bert" #"albert"
}
print(config)
# initialize BertNLU from config
nlu = BertNLU.from_config(config)
# -
# ## Train `BertNLU` model
# start model training
nlu.train(train_dataset, val_dataset, epochs=3, batch_size=32)
# ## Save the trained model
# +
save_path = f"../saved_models/joint_{model_type}_model"
print(f"Saving model to '{save_path}' ...")
nlu.save(save_path)
print("Saved!")
# -
# ## [Optional] Load the model from disk
print("Loading model ...")
nlu = AutoNLU.load(save_path)
# ## Reading Testing Dataset
test_path = "../data/snips/test" # please, specify the path to the testing dataset
test_dataset = Reader.read(test_path)
# ## Evaluating model
# +
token_f1_score, tag_f1_score, report, acc = nlu.evaluate(test_dataset)
print('Slot Classification Report:', report)
print('Slot token f1_score = %f' % token_f1_score)
print('Slot tag f1_score = %f' % tag_f1_score)
print('Intent accuracy = %f' % acc)
# -
# ## Make Predictions
# +
l = widgets.Layout(flex='0 1 auto', height='40px', min_height='30px', width='auto')
utterance_txt = widgets.Textarea(
value='add <NAME> to the grime instrumentals playlist',
placeholder='Type utterance here!',
description='Utterance:',
layout=l,
disabled=False
)
predict_btn = widgets.Button(
description='Predict'
)
result_layout = widgets.Layout(flex='0 1 auto', height='400px', min_height='30px', width='auto')
result_txt = widgets.Textarea(
value='',
placeholder='Result goes here',
description='Result:',
layout=result_layout,
disabled=True
)
def btn_click_event(x):
result = nlu.predict(utterance_txt.value)
result = json.dumps(result, indent=4, sort_keys=True)
result_txt.value = result
predict_btn.on_click(btn_click_event)
display(utterance_txt, predict_btn, result_txt)
# -
| notebooks/using_bert_nlu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
import pandas as pd
from scipy.optimize import minimize
import geopandas as gpd
# ### Definition of the model
# The SIR model differential equations.
def deriv(y, t, N, beta,gamma):
S,I,R = y
dSdt = -(beta*I/N)*S
dIdt = (beta*S/N)*I - gamma*I
dRdt = gamma*I
return dSdt, dIdt, dRdt
# ### Integration of the differential equations
def time_evo(N,beta,gamma,I0=1,R0=0,t=np.arange(0,365)):
# Definition of the initial conditions
# I0 and R0 denotes the number of initial infected people (I0)
# and the number of people that recovered and are immunized (R0)
# t ise the timegrid
S0=N-I0-R0 # number of people that can still contract the virus
# Initial conditions vector
y0 = S0, I0, R0
# Integrate the SIR equations over the time grid, t.
ret = odeint(deriv, y0, t, args=(N,beta,gamma))
S, I, R = np.transpose(ret)
return (t,S,I,R)
# # All-in-one
# +
popolation_regions = np.array([ 1304970, 559084, 533050, 1947131, 5801692, 4459477, 1215220,5879082, 1550640, 10060574, 1525271, 305617, 4356406, 4029053, 1639591, 4999891, 3729641, 541380, 882015, 125666, 4905854])
name_regions = np.array(['Abruzzo','Basilicata','<NAME>','Calabria','Campania','Emilia-Romagna','<NAME>','Lazio','Liguria','Lombardia','Marche','Molise','Piemonte','Puglia','Sardegna','Sicilia','Toscana','<NAME>','Umbria','Valle d\'Aosta','Veneto'])
data = pd.read_csv('https://raw.githubusercontent.com/pcm-dpc/COVID-19/master/dati-regioni/dpc-covid19-ita-regioni.csv')
df_r0=pd.DataFrame(data['data'].tail(1))
for region in name_regions:
N = popolation_regions[name_regions == region]
ydata = np.array(data.loc[data['denominazione_regione'] == region, "totale_casi"])
ydata_death = np.array(data.loc[data['denominazione_regione'] == region, "deceduti"])
ydata_rec = np.array(data.loc[data['denominazione_regione'] == region, "dimessi_guariti"])
ydata_inf = ydata-ydata_rec-ydata_death
xdata = pd.to_numeric(range(ydata.shape[0]))
today = len(xdata)
def minimizer(R0,t1=today-7,t2=today):
#true data
ydata_inf_2=np.array(ydata_inf[t1:t2])
xdata_2=np.arange(0,len(ydata_inf_2))
#model
fin_result=time_evo(N,0.1*R0,0.1,I0=ydata_inf_2[0])
i_vec=fin_result[2]
i_vec_2=i_vec[0:len(xdata_2)]
#average error
error=np.sum(np.abs(ydata_inf_2-i_vec_2)/ydata_inf_2)*100
return error
minimizer_vec=np.vectorize(minimizer)
xgrid = np.arange(1,1.3,0.01)
ygrid = minimizer_vec(xgrid)
r0_ideal = round(xgrid[np.argmin(ygrid)],2)
print('r0_ideal for the '+region+': ',r0_ideal)
ydata_inf_2 = np.array(ydata_inf[today-7:today])
xdata_2 = np.arange(0,len(ydata_inf_2))
print('ydata_inf.shape '+region+': ',ydata_inf.shape)
print('ydata_inf for the '+region+': ',ydata_inf)
print('ydata_inf_2 for the '+region+': ',ydata_inf_2)
fin_result = time_evo(N,1/14*r0_ideal,1/14,I0=ydata_inf_2[0])
t=fin_result[0]
s_vec=fin_result[1]
i_vec=fin_result[2]
r_vec=fin_result[3]
def minimizer_gen(t1,t2):
xgrid=np.arange(0.1,7.2,0.01)
ygrid=minimizer_vec(xgrid,t1=t1,t2=t2)
r0_ideal=round(xgrid[np.argmin(ygrid)],2)
return r0_ideal
r0_time=[]
# for i in range(today-4):
# min_val=minimizer_gen(i,i+5)
# r0_time.append(min_val)
# print(i,min_val)
min_val=minimizer_gen(today-7,today)
df_r0[region] = min_val
r0_time.clear()
# -
df = df_r0.T
df['description'] = df.index
df.rename(columns={ df.columns[0]: "R0" }, inplace = True)
df = df.iloc[1:]
df['description'][df.description == "<NAME>"] = "<NAME>"
trentino = round(float((sum(df.R0[df.description == "P.A. Trento"], df.R0[df.description == "P.A. Bolzano"])/2)), 2)
row_df = pd.DataFrame([{'R0':trentino, "description":"Trentino-Alto Adige"}])
df = pd.concat([df, row_df], ignore_index=True)
df
map = gpd.read_file("regioni_italiane.geojson")
map = map.merge(df, on='description', how='left')
map.to_file("export/r0_regioni.geojson", driver='GeoJSON')
classificazione = pd.read_excel('classificazione_regioni.xlsx')
map = map.merge(classificazione, on='description', how='left')
map[["description", "R0", "Area"]].to_csv("export/r0_regioni.csv")
map
| regioni/R0_map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Merge Datasets
#
# This recipe demonstrates a simple pattern for merging FiftyOne Datasets via [Dataset.merge_samples()](https://voxel51.com/docs/fiftyone/api/fiftyone.core.dataset.html?highlight=merge_samples#fiftyone.core.dataset.Dataset.merge_samples).
#
# Merging datasets is an easy way to:
#
# - Combine multiple datasets with information about the same underlying raw media (images and videos)
# - Add model predictions to a FiftyOne dataset, to compare with ground truth annotations and/or other models
# ## Setup
#
# In this recipe, we'll work with a dataset downloaded from the [FiftyOne Dataset Zoo](https://voxel51.com/docs/fiftyone/user_guide/dataset_creation/zoo.html).
#
# To access the dataset, install `torch` and `torchvision`, if necessary:
# Modify as necessary (e.g., GPU install). See https://pytorch.org for options
# !pip install torch
# !pip install torchvision
# Then download the test split of [CIFAR-10](https://www.cs.toronto.edu/~kriz/cifar.html):
# Download the validation split of COCO-2017
# !fiftyone zoo download cifar10 --splits test
# ## Merging model predictions
#
# Load the test split of CIFAR-10 into FiftyOne:
# +
import random
import os
import fiftyone as fo
import fiftyone.zoo as foz
# Load test split of CIFAR-10
dataset = foz.load_zoo_dataset("cifar10", split="test", dataset_name="merge-example")
classes = dataset.info["classes"]
print(dataset)
# -
# The dataset contains ground truth labels in its `ground_truth` field:
# Print a sample from the dataset
print(dataset.first())
# Suppose you would like to add model predictions to some samples from the dataset.
#
# The usual way to do this is to just iterate over the dataset and add your predictions directly to the samples:
def run_inference(filepath):
# Run inference on `filepath` here.
# For simplicity, we'll just generate a random label
label = random.choice(classes)
return fo.Classification(label=label)
# +
# Choose 100 samples at random
random_samples = dataset.take(100)
# Add model predictions to dataset
for sample in random_samples:
sample["prediction"] = run_inference(sample.filepath)
sample.save()
print(dataset)
# -
# However, suppose you store the predictions in a separate dataset:
# +
# Filepaths of images to proces
filepaths = [s.filepath for s in dataset.take(100)]
# Run inference
predictions = fo.Dataset()
for filepath in filepaths:
sample = fo.Sample(filepath=filepath)
sample["prediction"] = run_inference(filepath)
predictions.add_sample(sample)
print(predictions)
# -
# You can easily merge the `predictions` dataset into the main dataset via [Dataset.merge_samples()](https://voxel51.com/docs/fiftyone/api/fiftyone.core.dataset.html?highlight=merge_samples#fiftyone.core.dataset.Dataset.merge_samples).
#
# Let's start by loading a fresh copy of CIFAR-10 that doesn't have predictions:
dataset2 = foz.load_zoo_dataset("cifar10", split="test", dataset_name="merge-example2")
# Now merge the predictions:
# +
# Merge predictions
dataset2.merge_samples(predictions)
# Verify that 100 samples in `dataset2` now have predictions
print(dataset2.exists("prediction"))
# -
# By default, samples with the same absolute `filepath` are merged. However, you can customize this as desired via various keyword arguments of [Dataset.merge_samples()](https://voxel51.com/docs/fiftyone/api/fiftyone.core.dataset.html?highlight=merge_samples#fiftyone.core.dataset.Dataset.merge_samples).
#
# For example, the command below will merge samples with the same base filename, ignoring the directory:
# Merge predictions, using the base filename of the samples to decide which samples to merge
# In this case, we've already performed the merge, so the existing data is overwritten
dataset.merge_samples(predictions, key_fcn=lambda p: os.path.basename(p))
# Let's print a sample with predictions to verify that the merge happened as expected:
# Print a sample with predictions
print(dataset2.exists("prediction").first())
| docs/source/recipes/merge_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
from pandas import Series, DataFrame
import pandas as pd
# +
#Now we'll learn DataFrames
#Let's get some data to play with. How about the NFL?
import webbrowser
website = 'http://en.wikipedia.org/wiki/NFL_win-loss_records'
webbrowser.open(website)
# -
#Copy and read to get data
nfl_frame = pd.read_clipboard()
#Show
nfl_frame
# We can grab the oclumn names with .columns
nfl_frame.columns
#Lets see some specific data columns
DataFrame(nfl_frame,columns=['Team','First Season','Total Games'])
#What happens if we ask for a column that doesn't exist?
DataFrame(nfl_frame,columns=['Team','First Season','Total Games','Stadium'])
# Call columns
nfl_frame.columns
#We can retrieve individual columns
nfl_frame.Team
# Or try this method for multiple word columns
nfl_frame['Total Games']
#We can retrieve rows through indexing
nfl_frame.ix[3]
#We can also assign value sto entire columns
nfl_frame['Stadium']="Levi's Stadium" #Careful with the ' here
nfl_frame
# +
#Putting numbers for stadiums
nfl_frame["Stadium"] = np.arange(5)
#Show
nfl_frame
# -
# Call columns
nfl_frame.columns
#Adding a Series to a DataFrame
stadiums = Series(["Levi's Stadium","AT&T Stadium"],index=[4,0])
# +
#Now input into the nfl DataFrame
nfl_frame['Stadium']=stadiums
#Show
nfl_frame
# +
#We can also delete columns
del nfl_frame['Stadium']
nfl_frame
# +
#DataFrames can be constructed many ways. Another way is from a dictionary of equal length lists
data = {'City':['SF','LA','NYC'],
'Population':[837000,3880000,8400000]}
city_frame = DataFrame(data)
#Show
city_frame
# -
#For full list of ways to create DataFrames from various sources go to teh documentation for pandas:
website = 'http://pandas.pydata.org/pandas-docs/dev/generated/pandas.DataFrame.html'
webbrowser.open(website)
| Lec 15 - DataFrames.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="84163f3ca19c0b7c9fda47121b3bc4cadfaf1fcc"
# # Gensim Fasttext for Wikipedia corpus
# + _uuid="cc7b3e6ca62670ff13626705402f626778487204"
import re # For preprocessing
import pandas as pd # For data handling
from time import time # To time our operations
from collections import defaultdict # For word frequency
import spacy # For preprocessing
import nltk
# -
import os
cwd = os.getcwd()
cwd
# + _uuid="6453b9c3f797e51923e030090ead659253f4e459"
corpus = open("../output/wikipedia2008_fi_lemmatized.txt").read()
# + [markdown] _uuid="7f07dca2a2656dcd9e0c315afa36af32a992eef7"
# ## Alternative cleaning:
# -
# Since Wiki data is already lemmatized, I will split the data at periods to get sentences and then split sentences at whitespace
#
# Resulting data: list of lists
# +
# speeches is a list of list of sentences
sentences = re.split(r"[.!?]", corpus)
# sentences as list of words
sents = [sent.strip().split() for sent in sentences if sent != ""]
# -
sents[0:3]
# + [markdown] _uuid="31b4a744059df490ddb47ab6cdec008dc929ede3"
# ## Bigrams:
# + _uuid="af6d420284a0ff7a7407d4c526754ffe850d6170"
from gensim.models.phrases import Phrases, Phraser
# + [markdown] _uuid="bb7766b322cbc1d3381912b890585eb249ac5304"
# Creates the relevant phrases from the list of sentences:
# + _uuid="8befad8c76c54bd2b831b0942a2f626f7d8a6dac"
phrases = Phrases(sents, min_count=30, progress_per=10000)
# + [markdown] _uuid="45bae4a953f2ad8951e4efb234e1e357857a33b3"
# The goal of Phraser() is to cut down memory consumption of Phrases(), by discarding model state not strictly needed for the bigram detection task:
# + _kg_hide-input=true _uuid="b8ae81ba230013aefe7c584338de7376fedf6294"
bigram = Phraser(phrases)
# + [markdown] _uuid="4a58380f19d159688aeee665d1afb96289fdd4b8"
# Transform the corpus based on the bigrams detected:
# + _uuid="8051b56890c147119db3df529d3cfd3cf675fdca"
sentences = bigram[sents]
# + [markdown] _uuid="a4f81e8bb2c09a67b00cd24db28353eca8ae188c"
# ## Most Frequent Words:
# Mainly a sanity check of the effectiveness of the lemmatization, removal of stopwords, and addition of bigrams.
# + _uuid="eeb8afe1cfcb7ba65bd14d657455600acacf39ba"
word_freq = defaultdict(int)
for sent in sentences:
for i in sent:
word_freq[i] += 1
len(word_freq)
# + _uuid="5b010149150b2b2eaf332d79bcde0649b8a3c2b5"
sorted(word_freq, key=word_freq.get, reverse=True)[:10]
# + [markdown] _uuid="500ab7b5c84dc006d7945f339c40725a82856fdf"
# # Training the model
# ## Gensim Word2Vec Implementation:
# + _uuid="3269be205cadbad499aa87890893d92da6adc796"
import multiprocessing
from gensim.models import FastText
# + [markdown] _uuid="7c524bc49c41a6c37f9e754a38797c9501202090"
# ## Why I seperate the training of the model in 3 steps:
# I prefer to separate the training in 3 distinctive steps for clarity and monitoring.
# 1. `Word2Vec()`:
# >In this first step, I set up the parameters of the model one-by-one. <br>I do not supply the parameter `sentences`, and therefore leave the model uninitialized, purposefully.
# 2. `.build_vocab()`:
# >Here it builds the vocabulary from a sequence of sentences and thus initialized the model. <br>With the loggings, I can follow the progress and even more important, the effect of `min_count` and `sample` on the word corpus. I noticed that these two parameters, and in particular `sample`, have a great influence over the performance of a model. Displaying both allows for a more accurate and an easier management of their influence.
# 3. `.train()`:
# >Finally, trains the model.<br>
# The loggings here are mainly useful for monitoring, making sure that no threads are executed instantaneously.
# + _uuid="03488d9b68963579c96094aca88a302c9f2753a7"
cores = multiprocessing.cpu_count() # Count the number of cores in a computer
# + [markdown] _uuid="89c305fcd163488441ac2ac6133678bd973b4419"
# ## The parameters:
#
# * `min_count` <font color='purple'>=</font> <font color='green'>int</font> - Ignores all words with total absolute frequency lower than this - (2, 100)
#
#
# * `window` <font color='purple'>=</font> <font color='green'>int</font> - The maximum distance between the current and predicted word within a sentence. E.g. `window` words on the left and `window` words on the left of our target - (2, 10)
#
#
# * `size` <font color='purple'>=</font> <font color='green'>int</font> - Dimensionality of the feature vectors. - (50, 300)
#
#
# * `sample` <font color='purple'>=</font> <font color='green'>float</font> - The threshold for configuring which higher-frequency words are randomly downsampled. Highly influencial. - (0, 1e-5)
#
#
# * `alpha` <font color='purple'>=</font> <font color='green'>float</font> - The initial learning rate - (0.01, 0.05)
#
#
# * `min_alpha` <font color='purple'>=</font> <font color='green'>float</font> - Learning rate will linearly drop to `min_alpha` as training progresses. To set it: alpha - (min_alpha * epochs) ~ 0.00
#
#
# * `negative` <font color='purple'>=</font> <font color='green'>int</font> - If > 0, negative sampling will be used, the int for negative specifies how many "noise words" should be drown. If set to 0, no negative sampling is used. - (5, 20)
#
#
# * `workers` <font color='purple'>=</font> <font color='green'>int</font> - Use these many worker threads to train the model (=faster training with multicore machines)
# + _uuid="ad619db82c219d6cb81fad516563feb0c4d474cd"
fasttext_model = FastText(min_count=20,
window=2,
size=300,
sample=6e-5,
alpha=0.03,
min_alpha=0.0007,
negative=20,
workers=cores-1)
# + [markdown] _uuid="d7e9f1bd338f9e15647b5209ffd8fbb131cd7ee5"
# ## Building the Vocabulary Table:
# Word2Vec requires us to build the vocabulary table (simply digesting all the words and filtering out the unique words, and doing some basic counts on them):
# + _uuid="66358ad743e05e17dfbed3899af9c41056143daa"
t = time()
fasttext_model.build_vocab(sentences, progress_per=10000)
print('Time to build vocab: {} mins'.format(round((time() - t) / 60, 2)))
# + [markdown] _uuid="63260d82061abb47db7f2f8b23e07ec629adf5a9"
# ## Training of the model:
# _Parameters of the training:_
# * `total_examples` <font color='purple'>=</font> <font color='green'>int</font> - Count of sentences;
# * `epochs` <font color='purple'>=</font> <font color='green'>int</font> - Number of iterations (epochs) over the corpus - [10, 20, 30]
# + _uuid="07a2a047e701e512fd758edff186daadaeea6461"
t = time()
fasttext_model.train(sentences, total_examples=fasttext_model.corpus_count, epochs=30, report_delay=1)
print('Time to train the model: {} mins'.format(round((time() - t) / 60, 2)))
# -
fasttext_model.save("fasttext.model.alt_preprocessing.wiki")
# + _uuid="34dd51c7f2f39d016b982ef81e4df576f6b31bcb"
# Make model more memory-efficient
fasttext_model.init_sims(replace=True)
# + [markdown] _uuid="a420d5a98eb860cff1f4bbac8cbe2054459b6200"
# # Exploring the model
# ## Most similar to:
#
# + _uuid="339207a733a1ac42fe60e32a29f9e5d5ca0a9275"
fasttext_model.wv.most_similar(positive=["keskustapuolue"])
# + [markdown] _uuid="3b6686e6fa956a98450259b063b4cf51019a6d0b"
# _A small precision here:_<br>
# The dataset is the Simpsons' lines of dialogue; therefore, when we look at the most similar words from "homer" we do **not** necessary get his family members, personality traits, or even his most quotable words. No, we get what other characters (as Homer does not often refers to himself at the 3rd person) said along with "homer", such as how he feels or looks ("depressed"), where he is ("hammock"), or with whom ("marge").
#
# Let's see what the bigram "homer_simpson" gives us by comparison:
# + _uuid="22595f98c675a9697243b7e826b2840e5fc3e5f5"
fasttext_model.wv.most_similar(positive=["Matti_Vanhanen"])
# + _uuid="ac9ba47738e596dce6552099e76f303f28577943"
fasttext_model.wv.most_similar(positive=["Paavo_Väyrynen"])
# -
fasttext_model.wv.most_similar(positive=["Timo_Soini"])
# + [markdown] _uuid="d8b5937dfd7584f168a33060c435036cad5b390b"
# ## Similarities:
# + _uuid="367755f5c9e00de4bc5056c978f5b50a38c1368b"
fasttext_model.wv.similarity("keskusta", 'kokoomus')
# + _uuid="349828078b5a438d93e5494478e88095913dc58e"
fasttext_model.wv.similarity("<NAME>", "<NAME>")
# + [markdown] _uuid="08a999d758ac687d626b631a8ce393eaa26f41e7"
# ## Odd-One-Out:
#
# Here, we ask our model to give us the word that does not belong to the list!
# + _uuid="d982e44d9c212b5ee09bcaebd050a725ab5e508e"
fasttext_model.wv.doesnt_match(["Paavo_Väyrynen", "Matti_Vanhanen", "Timo_Soini"])
# -
fasttext_model.wv.doesnt_match(["Vihreät", "Vasemmistoliitto", "Kokoomus"])
# + [markdown] _uuid="df95cdff693e843ab4b4c174fea24029447573cd"
# ## Analogy difference:
# + _uuid="812961e79dde9f2032f708755ca287c0aef838d0"
fasttext_model.wv.most_similar(positive=["keskusta|puolue", "Jan_Vapaavuori"], negative=["Matti_Vanhanen"], topn=3)
# + [markdown] _uuid="773c0acc8750ba8e728ff261f2e9ec39694c245c"
# ### t-SNE visualizations:
# t-SNE is a non-linear dimensionality reduction algorithm that attempts to represent high-dimensional data and the underlying relationships between vectors in a lower-dimensional space.<br>
# Here is a good tutorial on it: https://medium.com/@luckylwk/visualising-high-dimensional-datasets-using-pca-and-t-sne-in-python-8ef87e7915b
# + _uuid="27ec46110042fc28da900b1b344ae4e0692d5dc2"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
# + [markdown] _uuid="22693eaa25253b38cee3c5cd5db6b6fdddb575a4"
# Our goal in this section is to plot our 300 dimensions vectors into 2 dimensional graphs, and see if we can spot interesting patterns.<br>
# For that we are going to use t-SNE implementation from scikit-learn.
#
# To make the visualizations more relevant, we will look at the relationships between a query word (in <font color='red'>**red**</font>), its most similar words in the model (in <font color="blue">**blue**</font>), and other words from the vocabulary (in <font color='green'>**green**</font>).
# + _uuid="489a7d160dcd92da0ce42a3b5b461368c9ffe5f1"
def tsnescatterplot(model, word, list_names):
""" Plot in seaborn the results from the t-SNE dimensionality reduction algorithm of the vectors of a query word,
its list of most similar words, and a list of words.
"""
arrays = np.empty((0, 300), dtype='f')
word_labels = [word]
color_list = ['red']
# adds the vector of the query word
arrays = np.append(arrays, model.wv.__getitem__([word]), axis=0)
# gets list of most similar words
close_words = model.wv.most_similar([word])
# adds the vector for each of the closest words to the array
for wrd_score in close_words:
wrd_vector = model.wv.__getitem__([wrd_score[0]])
word_labels.append(wrd_score[0])
color_list.append('blue')
arrays = np.append(arrays, wrd_vector, axis=0)
# adds the vector for each of the words from list_names to the array
for wrd in list_names:
wrd_vector = model.wv.__getitem__([wrd])
word_labels.append(wrd)
color_list.append('green')
arrays = np.append(arrays, wrd_vector, axis=0)
# Reduces the dimensionality from 300 to 50 dimensions with PCA
reduc = PCA(n_components=21).fit_transform(arrays)
# Finds t-SNE coordinates for 2 dimensions
np.set_printoptions(suppress=True)
Y = TSNE(n_components=2, random_state=0, perplexity=15).fit_transform(reduc)
# Sets everything up to plot
df = pd.DataFrame({'x': [x for x in Y[:, 0]],
'y': [y for y in Y[:, 1]],
'words': word_labels,
'color': color_list})
fig, _ = plt.subplots()
fig.set_size_inches(9, 9)
# Basic plot
p1 = sns.regplot(data=df,
x="x",
y="y",
fit_reg=False,
marker="o",
scatter_kws={'s': 40,
'facecolors': df['color']
}
)
# Adds annotations one by one with a loop
for line in range(0, df.shape[0]):
p1.text(df["x"][line],
df['y'][line],
' ' + df["words"][line].title(),
horizontalalignment='left',
verticalalignment='bottom', size='medium',
color=df['color'][line],
weight='normal'
).set_size(15)
plt.xlim(Y[:, 0].min()-50, Y[:, 0].max()+50)
plt.ylim(Y[:, 1].min()-50, Y[:, 1].max()+50)
plt.title('t-SNE visualization for {}'.format(word.title()))
# + [markdown] _uuid="c73fc2faaf0baecc84f02a97b50cb9ccefa48686"
# ## 10 Most similar words vs. 10 Most dissimilar
#
# + _uuid="10c77b072f7c281f2be919341be116565c20d8a8"
tsnescatterplot(fasttext_model, 'Matti_Vanhanen', [i[0] for i in fasttext_model.wv.most_similar(negative=["Matti_Vanhanen"])])
# + [markdown] _uuid="87315bfbaceb3733bd7af035db6c59cfc4b1ba7f"
# ## 10 Most similar words vs. 11th to 20th Most similar words:
#
# + _uuid="e6f0bc598922f4f2cd17d2511560242a3c35fdd9" pycharm={"name": "#%%\n"}
tsnescatterplot(fasttext_model, "Matti_Vanhanen", [t[0] for t in fasttext_model.wv.most_similar(positive=["Matti_Vanhanen"], topn=20)][10:])
# -
| notebooks/gensim_fasttext_wiki_corpus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extract authors from PMC-OAI frontmatter `<article>` records
# +
import pathlib
import pandas
from pubmedpy.xml import yield_etrees_from_zip
from pubmedpy.pmc_oai import extract_authors_from_article
# -
zip_paths = sorted(pathlib.Path('data/pmc/oai/pmc_fm').glob('*.zip'))
zip_paths
authors = list()
for zip_path in zip_paths:
for name, article in yield_etrees_from_zip(zip_path):
authors.extend(extract_authors_from_article(article))
author_df = pandas.DataFrame(authors)
author_df = author_df.sort_values(['pmcid', 'position'])
affiliation_df = author_df[["pmcid", "position", "affiliations"]]
author_df = author_df.drop(columns=['affiliations'])
author_df.tail()
# create affiliations table
affiliation_df = (
affiliation_df
.explode('affiliations')
.rename(columns={"affiliations": "affiliation"})
[["pmcid", "position", "affiliation"]]
.dropna(subset=["affiliation"])
)
affiliation_df.head(2)
# Show 10 random affiliations
print(*affiliation_df.sample(10, random_state=0).affiliation, sep='\n')
# number of unique affiliations
affiliation_df.affiliation.nunique()
# Total number of articles
author_df.pmcid.nunique()
# number of corresponding authors per paper
n_corresponding = author_df.groupby("pmcid").corresponding.sum()
pmcids_without_corresponding = set(n_corresponding[n_corresponding == 0].index)
# Probability of author position being corresponding,
# given that there's at least one corresponding author
# and the author is not the last author
(
author_df
.query("pmcid not in @pmcids_without_corresponding")
.query("reverse_position > 1")
.groupby("position")
.corresponding
.mean()
.map("{:.1%}".format)
.head()
)
# Probability of author reverse position being corresponding,
# given that there's at least one corresponding author
# and the author is not the first author
(
author_df
.query("pmcid not in @pmcids_without_corresponding")
.query("position > 1")
.groupby("reverse_position")
.corresponding
.mean()
.map("{:.1%}".format)
.head()
)
# Corresponding author counts
n_corresponding.value_counts().sort_index()
# Testing: show some articles without any corresponding authors
n_corresponding.reset_index().query("corresponding == 0").head()
# Testing: show some articles without >10 corresponding authors
n_corresponding.reset_index().query("corresponding >= 10")
# +
# Write author dataframe to a TSV
author_df.to_csv('data/pmc/authors.tsv.xz', index=False, sep='\t')
# Write affiliation dataframe to a TSV
affiliation_df.to_csv('data/pmc/affiliations.tsv.xz', index=False, sep='\t')
| 04.process-pmc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## File I/O (Input / Output)
# ## Jupyter only writing to text file
# * %%writefile filename.ext
#
# Writes in the working directory (first run pwd)
# %%writefile somefile.txt
Oh my text first line
Super easy second line
OH and third line is
# +
# %%writefile myFunctions.py
def myCoolFun(a,b):
return a+b
def mySecondFun(a,b):
return a*b
# -
import myFunctions
myFunctions.myCoolFun(5,6)
import myFunctions as mf
mf.myCoolFun(5,6)
mf.mySecondFun(6,7)
import mylib
mylib.nb_year(100,2,0,200)
from mylib import nb_year
nb_year(100,20,20,200)
from mylib import nb_year as mynb
mynb(555,55,100,10000)
# filein is our own name for file
filein = open('somefile.txt')
type(filein)
filein.
# +
mytext = filein.read() # usually not that useful
print(mytext)
# -
filein.read()
# there can be many pointers to file stream
fin = filein
fin.name
fin.read() # what will happen ?
fin.seek(2)
fin.read()
fin.seek(6)
text6= fin.read()
print(text6)
# we seek the very beginning of the file
fin.seek(0)
fulltext = fin.read()
print(fulltext)
fulltext[6:]
fin.seek(0)
mylines=fin.readlines()
mylines
type(mylines)
len(mylines)
# list comprehension to generate line lengths
linelengths = [len(line) for line in mylines]
linelengths
mylines[0]
print(mylines[0])
# if we need the representation of string then we use built in repr function
print(mylines[0])
print(repr(mylines[0])) # same as mylines[0] as the first last item
mylines[0]
# we go trough each line and do some work
for line in mylines:
print(f'My Line Is:{line}')
# Do more work on lines
"He said 'nice weather' , really"
# We can go throught the file directly (without reading the whole file into memory)
fin.seek(0)
# we can go through very large files here
for line in fin:
print(line, end='')
for line in mylines:
print(line, end='')
#close the file if you .open() it!!
fin.close()
# ## Use with open always!
#
# * closes automatically!
# * throws exceptions on errors
with open('somefile.txt') as fin:
for line in fin:
# do something with each line
print(line)
# +
# Idiom on how to open AND close a file for reading and doing work
with open('somefile.txt') as fin:
results = []
for line in fin:
results.append(line.count('a'))
print(line)
# do wo with each line here,save into a list or other structure
# we can do more work with file here
# maybe fin.seek(0) to read it again for some reason
# File will be closed once this line ends
print("file is closed already here")
print(results)
#closes here!
#closes automatically!!!
# -
fin.read()
# ## For MacOS and Linux
# * use pwd to see where you are
# ### myfile = open("/Users/MyUserName/SomeFolder/MaybeAnotherFolder/myfile.txt")
# ## For Windows
# * use pwd to see where you are
# ### myfile = open("C:\\Users\\MyUserName\\SomeFolder\\MaybeAnotherFolder\\myfile.txt")
# Jupyter Magic !someOScommand for example !dir or !ls
# !dir
pwd
# !dir
pwd
# importing OS specific library for system work
import os
cwd = os.getcwd()
cwd
with open(cwd+'\\somefile.txt') as f:
for line in f:
print(line)
# join path no mater what OS we have
fullpath = os.path.join(cwd, 'somefile.txt')
fullpath
os.path.join(os.getcwd(), "myimages", "catpictures", "mycat.png")
# i get the current working directory and join the file path
with open(os.path.join(os.getcwd(), 'somefile.txt')) as f:
for line in f:
print(line)
with open('../data/Veidenbaums.txt', encoding='UTF-8') as f:
mytext = f.read()
len(mytext)
mytext[:100]
mytext[-100:]
with open('../data/Veidenbaums.txt', encoding='UTF-8') as f:
mylines = f.readlines()
len(mylines)
cleanlines = [line.strip() for line in mylines]
len(cleanlines)
textlines = [line for line in cleanlines if len(line) > 0]
len(textlines)
textlines[100:105]
'***' in textlines[100]
'***' in textlines[101]
noheadlines = [line for line in textlines if not '***' in line]
len(noheadlines)
noheadlines[:10]
"# #".join(["Valdis", "likes", "food"])
poemtext= " ".join(noheadlines)
len(poemtext), poemtext[:50]
words = poemtext.split(" ")
len(words)
words[:20]
badchars = ".,:'\"!?"
for c in badchars:
print(c)
print(len(poemtext))
for c in badchars:
poemtext = poemtext.replace(c, "")
len(poemtext)
words = poemtext.split()
len(words)
lowercase = [word.lower() for word in words]
len(lowercase), lowercase[:10]
from collections import Counter
wordcount = Counter(lowercase)
wordcount.most_common(20)
specialwords = ['alu', 'ala', 'opa']
# +
longwords = [word for word in lowercase if len(word) > 3 or word in specialwords]
len(lowercase),len(longwords)
# -
longcount = Counter(longwords)
longcount.most_common(20)
longcount['alus']
longcount['alu']
#mode=w is write only and it deletes OLD file!!
with open('numbers.txt', mode='w') as fwriter:
for n in range(1,10):
fwriter.write(f'The number is {n*2} \n')
#fout.write(f'The number is {n}') # diferences between \n and no \n
#file is closed is here
len('The number is {n*2} \n')*9
with open('numbers.txt') as freader:
mytext = freader.read()
len(mytext)
with open('numbers.txt') as freader:
sum = 0
for line in freader:
print(line)
print(len(line))
sum += len(line)
print(sum)
from datetime import datetime
now = datetime.now()
now
today = datetime.today()
print(today)
#mode=w is write only and it deletes OLD file!!
today = datetime.today()
with open(f'numbers{today.hour}_{today.minute}_{today.second}.txt', mode='w') as fwriter:
for n in range(1,10):
fwriter.write(f'The number is {n*2} \n')
#fout.write(f'The number is {n}') # diferences between \n and no \n
#file is closed is here
today
today.hour
today.minute
datetime.today()
timestamp = datetime.timestamp(now)
timestamp
str(datetime.now())
with open('numbers.txt') as fin:
print(fin.readlines())
with open('numbers.txt', mode='r') as fin:
for line in fin:
print(line, end="")
# We can append to the files without overwriting
with open('numbers.txt', mode='a') as fin:
fin.write("This might not be the end\nThis is really the end")
with open('numbers.txt', mode='a') as fin:
fin.seek(4) # seek will not work here for writing
fin.write("This is not the end")
with open('numbers.txt') as f:
print(f.read())
# Jupyter magic for reading files into notebook
# %%readfile numbers.txt
The number is 2
The number is 4
The number is 6
The number is 8
The number is 10
The number is 12
The number is 14
The number is 16
The number is 18
This is the end!
# +
# Generally Preferably to read and write separately - VS
# -
with open('somefile.txt', mode="r+") as f:
print(f.readlines())
f.write('moreinfo\tmore\n')
with open('sometext.txt') as f:
print(f.readlines())
# ### Modes:
# * mode='r' - Read Only
# * 'w' - Write Only (and will overwrite existing files!!!)
# * 'a' - Apend Only (stream is at the end of file!)
# * 'r+' - Read and Write
# * 'w+' - Write and Read with Overwriting existing/make new files
#
# From C (fopen)
# * ``r+'' Open for reading and writing. The stream is positioned at the
# beginning of the file.
#
# * ``w+'' Open for reading and writing. The file is created if it does not
# exist, otherwise it is truncated(**destroyed!**). The stream is positioned at
# the beginning of the file.
with open('numbers.txt', mode='a') as f:
f.write("New Line\n")
# !cd data/
# !dir
mylist = list(range(1,30))
mytextlist = [str(x)+'\n' for x in range(1,30)]
with open('list.txt', mode='w') as fwriter:
fwriter.writelines(mytextlist)
with open('biglist.txt', mode='w') as fwriter:
for n in range(1,1_000_000):
fwriter.write('Line:'+ str(n)+'\n')
# +
# Optimal writing for large files will be between how much you can hold in memory and how big the file will be
# -
# ## More on String Formatting
# ## How do you convert values to strings?
#
# ### In Python: pass it to the repr() or str() functions.
#
# The str() function is meant to return representations of values which are fairly human-readable, while repr() is meant to generate representations which can be read by the interpreter (or will force a SyntaxError if there is no equivalent syntax). For objects which don’t have a particular representation for human consumption, str() will return the same value as repr().
print(str(34341235421))
print(repr(54))
print(str(54))
str(54)==repr(54)
hi = "Hello\n"
hir = repr(hi)
print(hi)
print(hir)
hir==hi
# ### The goal of __repr__ is to be unambiguous:
# ### The goal of __str__ is to be readable
#
# ## In other words: __repr__ is for developers, __str__ is for customers (end users)
for x in range(1,12):
print(f'{x:2d} {x*x:3d} {x**3:4f}')
'-3.14'.zfill(7) # pads numeric string on the left with zeros
# write to a text file squares.txt first 10 squares
# 1 squared is 1
# 2 squared is 4
for n in range(1,11):
print(f'{n} squared is {n*n}')
# write to a text file squares.txt first 10 squares
# 1 squared is 1
# 2 squared is 4
with open('squares.txt', mode='w') as f:
for n in range(1,11):
myline = f'{n} squared is {n*n}\n'
print(myline)
f.write(myline)
# write to a text file squares.txt first 10 squares
# 1 squared is 1
# 2 squared is 4
with open('squares2.txt', mode='w') as f:
mylines = [f'{n} squared is {n*n}\n' for n in range(1,11)]
f.writelines(mylines)
with open('squares2.txt') as f:
for line in f:
# do something with line for example print it
print(line, end="")
type(f)
dir(f)
f.close()
f.readlines()
# +
## Homework
## Write function which writes Fizzbuzz 1 to 100 (5,7) to file fizzbuzz.txt
## Format example:
## 1: 1
## 2: 2
## 5: Fizz
## 6: 6
## 7: Buzz
#... 35: FizzBuzz
# -
len(mytext)
with open('../data/Veidenbaums.txt', encoding='UTF-8') as f:
veidtext = [line for line in f]
len(veidtext)
veidtext[:10]
| Python_Core/Python File IO_in_class_01_with_Veidenbaums.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
import ipywidgets as widgets
from IPython.display import display, clear_output
# +
# #!jupyter nbextension enable --py widgetsnbextension --sys-prefix
# #!jupyter serverextension enable voila --sys-prefix
# +
# Image Widget
file = open("grandma.jpg", "rb")
image = file.read()
image_headline = widgets.Image(
value=image,
format='jpg',
width='300'
)
label_headline = widgets.Label(
value='Photo by CDC on Unsplash',
style={'description_width': 'initial'}
)
vbox_headline = widgets.VBox([image_headline, label_headline])
# +
# grandson/granddaughter
grand = widgets.ToggleButtons(
options=['grandson', 'granddaughter']
)
# +
# name
name = widgets.Text(placeholder='Your name here')
# -
date = widgets.DatePicker(description='Pick a Date')
# +
# number of friends
friends = widgets.IntSlider(
value=3, # default value
min=0,
max=10,
step=1,
style={'description_width': 'initial', 'handle_color': '#16a085'}
)
# +
# button send
button_send = widgets.Button(
description='Send to grandma',
tooltip='Send',
style={'description_width': 'initial'}
)
output = widgets.Output()
def on_button_clicked(event):
with output:
clear_output()
print("Sent message: ")
print(f"Dear Grandma! This is your favourite {grand.value}, {name.value}.")
print(f"I would love to come over on {date.value} for dinner, if that's okay for you!")
print(f"Also, if you don't mind, I'll bring along {friends.value} hungry ghosts for your delicious food!")
button_send.on_click(on_button_clicked)
vbox_result = widgets.VBox([button_send, output])
# +
# stacked right hand side
text_0 = widgets.HTML(value="<h1>Dear Grandma!</h1>")
text_1 = widgets.HTML(value="<h2>This is your favourite</h2>")
text_2= widgets.HTML(value="<h2>I would love to come over on </h2>")
text_3= widgets.HTML(value="<h2>for dinner, if that's okay for you!</h2>")
text_4= widgets.HTML(value="<h2>Also, if you don't mind, I'll bring along </h2>")
text_5= widgets.HTML(value="<h2>hungry ghosts for dinner, if that's okay for you!</h2>")
vbox_text = widgets.VBox([text_0, text_1, grand, name, text_2, date, text_3, text_4, friends, text_5, vbox_result])
# -
page = widgets.HBox([vbox_headline, vbox_text])
display(page)
# !pip freeze > requirements.txt
| Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IPC Alerts of Acute Food Insecurity
# Capture current IPC alerts as displayed on this ipcinfo.org [page](http://www.ipcinfo.org/ipcinfo-website/resources/alerts-archive/en/).
# +
# General imports
# helpers
from pathlib import Path
import urllib.request
from datetime import datetime
from tqdm import tqdm
import copy
# data processing
import pandas as pd
import requests
from urllib.request import Request, urlopen
from bs4 import BeautifulSoup
# conf
import sys
sys.path.insert(0,'../../..') # TODO: be more elegant / robust here
from config.config import config
sources = config.sources
# -
# ## Finding all alerts, then finding all related documents
def get_all_alerts_hyperlinks(url):
# Find all hyperlinks for alerts on the page,
headers = {"User-Agent": "Mozilla/5.0"}
req = Request(url=url, headers=headers)
with urllib.request.urlopen(req) as page:
# Parse .xml file
xml = BeautifulSoup(page.read(), "lxml")
# Get all dataset info
links = []
for link in xml.findAll("a"):
title = link.get("title")
url = link.get("href")
if (
(title is not None)
and (title != "image thumbnail")
and (url.startswith("/ipcinfo-website/alerts-archive/issue-"))
):
url = "http://www.ipcinfo.org" + url
links.append({"title": title, "url": url})
# TODO: add the alerts paragraph to the metadata
return links
# +
def get_pdf_from_url(url, output_path=None):
"""Download a pdf from a given url.
:param url: web url from where to download the pdf
:param output_path: a pathlib.Path object to which to write the downloaded pdf
"""
# create the folder if not existing
output_path.parent.mkdir(parents=True, exist_ok=True)
# download
r = requests.get(url)
with open(output_path, "wb") as outfile:
outfile.write(r.content)
def get_all_documents_from_alerts_references(links):
documents = []
for link in tqdm(links):
headers = {"User-Agent": "Mozilla/5.0"}
req = Request(url=link["url"], headers=headers)
with urllib.request.urlopen(req) as page:
xml = BeautifulSoup(page.read(), "lxml")
alert_page_title = xml.find("div", class_="csc-header csc-header-n2").text
# TODO : Using spacy entity recognition, find the country/region + map it to ISO code
for new_link in xml.find_all("a"):
href = new_link.get("href")
if href.endswith(".pdf"):
if href.startswith("/"):
href = "http://www.ipcinfo.org" + href
# For each download, based on the pdf's name (report/annexes/snapshots, etc...), indicate the type of file in a metadata file
document_type = "unknown_type"
pdf_filename = href.split("/")[-1]
for potential_document_type in ["snapshot", "report", "annexes"]:
if potential_document_type in pdf_filename.lower():
document_type = potential_document_type
output_path = output_folder_path / pdf_filename
link.update(
{
"alert_page_title": alert_page_title,
"pdf_url": href,
"document_type": document_type,
"pdf_local_path": str(output_path),
}
)
# Download the items
get_pdf_from_url(href, output_path=output_path)
documents.append(copy.copy(link))
# save the resulting csv of metadata
df = pd.DataFrame.from_dict(documents)
df.to_csv(output_metadata_path, sep=";", index=False)
return df
# -
# ## Scrape them all
# +
ipc_alerts_hyperlink = sources.ipc_alerts_url
output_folder_path = sources.ipc_alerts_folder_path
output_metadata_path = sources.ipc_alerts_metadata_path
# Find all hyperlinks for alerts on the page,
links = get_all_alerts_hyperlinks(ipc_alerts_hyperlink)
df = get_all_documents_from_alerts_references(links)
# + tags=["dev"]
df.head()
# -
| src/notebooks/collection/IPC Alerts of Acute Food Insecurity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/julianovale/PO240_Meta_heuristica/blob/main/PO240_Semana07_NSGA2_Portfolio.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="jd453rmkY-as" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 73} outputId="eac7db5e-cd6c-4293-84c8-673a8d4a1b1b"
from google.colab import files
uploaded = files.upload() # subir arquivo cotacoes.xlsx
# + id="kQ1GqtOdW7MF"
# Program Name: NSGA-II.py
# Description: This is a python implementation of Prof. Kalyanmoy Deb's popular NSGA-II algorithm
# Author: <NAME>
# Supervisor: Prof. <NAME>
#Importing required modules
import math
import random
import matplotlib.pyplot as plt
#Function to find index of list
def index_of(a,list):
for i in range(0,len(list)):
if list[i] == a:
return i
return -1
#Function to sort by values
def sort_by_values(list1, values):
sorted_list = []
while(len(sorted_list)!=len(list1)):
if index_of(min(values),values) in list1:
sorted_list.append(index_of(min(values),values))
values[index_of(min(values),values)] = math.inf
return sorted_list
#Function to carry out NSGA-II's fast non dominated sort
def fast_non_dominated_sort(values1, values2): # dois objetivos de maximização
S=[[] for i in range(0,len(values1))]
front = [[]]
n=[0 for i in range(0,len(values1))]
rank = [0 for i in range(0, len(values1))]
for p in range(0,len(values1)):
S[p]=[]
n[p]=0
for q in range(0, len(values1)):
if (values1[p] > values1[q] and values2[p] > values2[q]) or (values1[p] >= values1[q] and values2[p] > values2[q]) or (values1[p] > values1[q] and values2[p] >= values2[q]):
if q not in S[p]:
S[p].append(q) # p domina q
elif (values1[q] > values1[p] and values2[q] > values2[p]) or (values1[q] >= values1[p] and values2[q] > values2[p]) or (values1[q] > values1[p] and values2[q] >= values2[p]):
n[p] = n[p] + 1 # q domina p
if n[p]==0:
rank[p] = 0
if p not in front[0]:
front[0].append(p)
i = 0
while(front[i] != []):
Q=[]
for p in front[i]:
for q in S[p]:
n[q] =n[q] - 1
if( n[q]==0):
rank[q]=i+1
if q not in Q:
Q.append(q)
i = i+1
front.append(Q)
del front[len(front)-1]
return front
#Function to calculate crowding distance
def crowding_distance(values1, values2, front):
distance = [0 for i in range(0,len(front))]
sorted1 = sort_by_values(front, values1[:])
sorted2 = sort_by_values(front, values2[:])
distance[0] = 4444444444444444
distance[len(front) - 1] = 4444444444444444
for k in range(1,len(front)-1):
distance[k] = distance[k]+ (values1[sorted1[k+1]] - values2[sorted1[k-1]])/(max(values1)-min(values1))
for k in range(1,len(front)-1):
distance[k] = distance[k]+ (values1[sorted2[k+1]] - values2[sorted2[k-1]])/(max(values2)-min(values2))
return distance
# + id="RmmpsisAU7zr"
import numpy as np
# from nsgaII import fast_non_dominated_sort
# from nsgaII import crowding_distance
# from nsgaII import sort_by_values
# from nsgaII import index_of
import random
import matplotlib.pyplot as plt
import pandas as pd
# sobre esta função: Author: <NAME>
def geneticoMultiobjetivo(populacao, TAM_POP, GERACOES, retATIVOS, covATIVOS, ATIVOS):
gen_no = 0 # geração número 0
while(gen_no < GERACOES):
# function1_values é uma lista com todos os valores de f1 (retorno) de todos os indivíduos
function1_values = [function1(populacao[i], retATIVOS, ATIVOS) for i in range(TAM_POP)]
# function2_values é uma lista com todos os valores de f2 (-risco) de todos os indivíduos
function2_values = [function2(populacao[i], covATIVOS, ATIVOS) for i in range(TAM_POP)]
non_dominated_sorted_solution = fast_non_dominated_sort(function1_values[:],function2_values[:])
crowding_distance_values=[]
for i in range(0,len(non_dominated_sorted_solution)):
crowding_distance_values.append(crowding_distance(function1_values[:],function2_values[:],non_dominated_sorted_solution[i][:]))
# montando a população mista (solution2): população atual + filhos
solution2 = populacao[:] # copia população atual toda
# gerando os filhos
while(len(solution2) != 2*TAM_POP):
pai1 = random.randint(0,TAM_POP-1) # seleciona um índice
pai2 = random.randint(0,TAM_POP-1)
# coloca na mista os filhos gerados no crossover
filho1, filho2 = crossoverMutacao(populacao[pai1], populacao[pai2], ATIVOS)
solution2.append(filho1)
solution2.append(filho2)
# avalia a população mista toda
function1_values2 = [function1(solution2[i], retATIVOS, ATIVOS)for i in range(0,2*TAM_POP)]
function2_values2 = [function2(solution2[i], covATIVOS, ATIVOS)for i in range(0,2*TAM_POP)]
non_dominated_sorted_solution2 = fast_non_dominated_sort(function1_values2[:],function2_values2[:])
crowding_distance_values2=[]
for i in range(0,len(non_dominated_sorted_solution2)):
crowding_distance_values2.append(crowding_distance(function1_values2[:],function2_values2[:],non_dominated_sorted_solution2[i][:]))
new_solution= []
for i in range(0,len(non_dominated_sorted_solution2)):
non_dominated_sorted_solution2_1 = [index_of(non_dominated_sorted_solution2[i][j],non_dominated_sorted_solution2[i] ) for j in range(0,len(non_dominated_sorted_solution2[i]))]
front22 = sort_by_values(non_dominated_sorted_solution2_1[:], crowding_distance_values2[i][:])
front = [non_dominated_sorted_solution2[i][front22[j]] for j in range(0,len(non_dominated_sorted_solution2[i]))]
front.reverse()
for value in front:
new_solution.append(value)
if(len(new_solution)==TAM_POP):
break
if (len(new_solution) == TAM_POP):
break
populacao = [solution2[i] for i in new_solution]
gen_no = gen_no + 1
#Lets plot the final front now
# function1_values é uma lista com todos os valores de f1 (retorno) de todos os indivíduos
function1_values = [function1(populacao[i], retATIVOS, ATIVOS) for i in range(TAM_POP)]
# function2_values é uma lista com todos os valores de f2 (-risco) de todos os indivíduos
function2_values = [function2(populacao[i], covATIVOS, ATIVOS) for i in range(TAM_POP)]
non_dominated_sorted_solution = fast_non_dominated_sort(function1_values[:],function2_values[:])
pontos_function1 = []
pontos_function2 = []
for indiceIndiv in non_dominated_sorted_solution[0]: # somente primeira fronteira
pontos_function1.append(function1_values[indiceIndiv])
pontos_function2.append(-function2_values[indiceIndiv])
return pontos_function1, pontos_function2
# coletar tabela de preços, conforme o exemplo
def leitura(arquivo):
ARQ = pd.read_excel(arquivo, sheet_name='Valores')#abre o arquivo para leitura, na aba Valores
ATIVOS = ARQ.at[0,'Qtde']
PERIODOS = ARQ.at[1,'Qtde']
ARQ = pd.read_excel(arquivo, sheet_name='Planilha')
Preco = []
for i in range(ATIVOS): # coluna
nomecoluna = "Acao"+str(i+1)
PrecoI = [] # uma coluna de inputs
for t in range(PERIODOS): # linha
PrecoIK = ARQ.at[t, nomecoluna] # um número de input
PrecoI.append(PrecoIK) # coloco um número de input na coluna
Preco.append(PrecoI) # coloco a coluna em x
return ATIVOS, PERIODOS, Preco
# calcula o retorno (maximização)
def function1(x, retATIVOS, ATIVOS):
total = 0
for i in range(ATIVOS):
total = total + retATIVOS[i]*x[i]
return total
# calcula o risco (minimização risco = - maximização (- risco))
def function2(x, covATIVOS, ATIVOS):
total = 0
for i in range(ATIVOS):
for j in range(ATIVOS):
total = total + covATIVOS[i][j]*x[i]*x[j]
return -total # trocar sinal para problema ter apenas objetivos de max
def preProcessamento(ATIVOS, PERIODOS, Preco):
# criar a matriz r de retornos
r = []
for i in range (ATIVOS): # para cada linha
crialinha = []
for t in range (PERIODOS): # para cada coluna
crialinha.append(0) # preencho a linha com 0s em todas as colunas
r.append(crialinha)
# calculo do r
for i in range (ATIVOS):
for t in range (1, PERIODOS):
r[i][t-1] = (Preco[i][t]/Preco[i][t-1]) - 1
# criar o vetor retATIVOS
retATIVOS = []
for i in range (ATIVOS):
retATIVOS.append(0)
# calculo
for i in range (ATIVOS):
for t in range (PERIODOS-1):
retATIVOS[i] = retATIVOS[i] + r[i][t]
retATIVOS[i] = retATIVOS[i]/(PERIODOS - 1)
# criar a matriz covATIVOS
covATIVOS = []
for i in range (ATIVOS): # para cada linha
crialinha = []
for j in range (ATIVOS): # para cada coluna
crialinha.append(0) # preencho a linha com 0s em todas as colunas
covATIVOS.append(crialinha)
# calculo
for i in range(ATIVOS):
for j in range (ATIVOS):
for t in range (PERIODOS):
covATIVOS[i][j] = covATIVOS[i][j] + (r[i][t] - retATIVOS[i])*(r[j][t] - retATIVOS[j])
return retATIVOS, covATIVOS
def inicializaPopulacao(TAM_POP, ATIVOS):
populacao = []
for j in range (TAM_POP):
x = []
for i in range (ATIVOS):
aleatorio = random.random()
if aleatorio <= 0.1:
aleatorio = 0
x.append(aleatorio) # preenchendo a i-ésima posição com o aletorio
populacao.append(x)
normalizarPopulacao(populacao, TAM_POP, ATIVOS) # somatório x[i] = 1
return populacao
def normalizarPopulacao(populacao, TAM_POP, ATIVOS):
for j in range (TAM_POP):
populacao[j] = normalizarSolucao(populacao[j], ATIVOS)
def normalizarSolucao(x, ATIVOS):
soma = 0
for i in range (ATIVOS):
if x[i] <= 0.01:
x[i] = 0
soma = soma + x[i] #está somando todos os valores de x
if soma < 1 or soma > 1:
for i in range (ATIVOS):
x[i] = x[i]/soma
return x
def crossoverMutacao(pai1, pai2, ATIVOS):
PC = 0.9
prob = random.random()
if prob < PC:
alpha = random.random()
filho1 = []
filho2 = []
for i in range (ATIVOS):
filho1.append(alpha*pai1[i] + (1-alpha)*pai2[i])
filho2.append(alpha*pai2[i] + (1-alpha)*pai1[i])
else:
filho1 = pai1.copy()
filho2 = pai2.copy()
# mutação
filho1m = mutacao(filho1, ATIVOS)
filho2m = mutacao(filho2, ATIVOS)
filho1mn = normalizarSolucao(filho1m, ATIVOS)
filho2mn = normalizarSolucao(filho2m, ATIVOS)
return filho1mn, filho2mn
def mutacao(x, ATIVOS):
PM = 0.1
for i in range (ATIVOS):
prob = random.random()
if prob < PM:
x[i] = np.random.normal(x[i],0.1)
return x
def imprimeRodadas(matriz3D):
plt.xlabel('Retorno', fontsize=15)
plt.ylabel('Risco', fontsize=15)
xmin = matriz3D[0][0][0]
xmax = matriz3D[0][0][0]
ymin = matriz3D[0][1][0]
ymax = matriz3D[0][1][0]
for fronteira in matriz3D:
pontos_function1 = fronteira[0]
pontos_function2 = fronteira[1]
# para encontrar limites do gráfico
if min(pontos_function1) < xmin:
xmin = min(pontos_function1)
if max(pontos_function1) > xmax:
xmax = max(pontos_function1)
if min(pontos_function2) < ymin:
ymin = min(pontos_function2)
if max(pontos_function2) > ymax:
ymax = max(pontos_function2)
plt.scatter(pontos_function1, pontos_function2)
plt.xlim(xmin, xmax)
plt.ylim(ymin, ymax)
plt.show()
# função principal: chama as demais funções
def main(TAM_POP, GERACOES, RODADAS):
random.seed() # inicia a semente dos número pseudo randômicos
# leitura e preparação
ATIVOS, PERIODOS, Preco = leitura("cotacoes.xlsx")
retATIVOS, covATIVOS = preProcessamento(ATIVOS, PERIODOS, Preco)
matriz3D = [] # matriz com todas as fronteiras
for r in range(RODADAS):
# Genético
populacao = inicializaPopulacao(TAM_POP, ATIVOS)
pontos_function1, pontos_function2 = geneticoMultiobjetivo(populacao, TAM_POP, GERACOES, retATIVOS, covATIVOS, ATIVOS)
matriz3D.append([pontos_function1, pontos_function2])
imprimeRodadas(matriz3D)
# + id="g_qhy8VYW6De" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="706ff975-a8a1-4fb1-9dd8-f7e89be76a34"
# TAM_POP, GERACOES, RODADAS
main(100, 10, 4)
# + id="tbkquZWLU70L" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="68a3ac25-e2c8-4540-bbcb-32c83d27bed2"
# TAM_POP, GERACOES, RODADAS
main(100, 200, 4)
| PO240_Semana07_NSGA2_Portfolio.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p><font size="6"><b> CASE - Observation data - data cleaning and enrichment</b></font></p>
#
# > *© 2021, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
# -
# **Scenario**:<br>
#
# Observation data of species (when and where is a given species observed) is typical in biodiversity studies. Large international initiatives support the collection of this data by volunteers, e.g. [iNaturalist](https://www.inaturalist.org/). Thanks to initiatives like [GBIF](https://www.gbif.org/), a lot of these data is also openly available.
# You decide to share data of a field campaign, but the data set still requires some cleaning and standardization. For example, the coordinates, can be named `x`/`y`, `decimalLatitude`/`decimalLongitude`, `lat`/`long`... Luckily, you know of an international **open data standard** to describe occurrence/observation data, i.e. [Darwin Core (DwC)](http://rs.tdwg.org/dwc/terms). Instead of inventing your own data model, you decide to comply to this international standard. The latter will enhance communication and will also make your data compliant with GBIF.
# In short, the DwC describes a flat table (cfr. `CSV`) with an agreed name convention on the header names and conventions on how certain data types need to be represented (as a reference, an in depth description is given [here](https://www.tdwg.org/standards/dwc/)). For this tutorial, we will focus on a few of the existing terms to learn some elements about data cleaning:
# * `eventDate`: ISO 6801 format of dates
# * `scientificName`: the accepted scientific name of the species
# * `decimalLatitude`/`decimalLongitude`: coordinates of the occurrence in WGS84 format
# * `sex`: either `male` or `female` to characterize the sex of the occurrence
# * `occurrenceID`: an identifier within the data set to identify the individual records
# * `datasetName`: a static string defining the source of the data
#
# Furthermore, additional information concerning the taxonomy will be added using an external API service
# **Dataset to work on:**
# For this data set, the data is split up in the following main data files:
# * `surveys.csv` the data with the surveys in the individual plots
# * `species.csv` the overview list of the species short-names
# * `plot_location.xlsx` the overview of coordinates of the individual locations
#
# The data originates from a [study](http://esapubs.org/archive/ecol/E090/118/metadata.htm) of a Chihuahuan desert ecosystem near Portal, Arizona.
#
# 
# ## 1. Survey-data
# Reading in the data of the individual surveys:
survey_data = pd.read_csv("data/surveys.csv")
survey_data.head()
# <div class="alert alert-success">
#
# **EXERCISE 1**
#
# - How many individual records (occurrences) does the survey data set contain?
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing1.py
# -
# ### Adding the data source information as static column
# For convenience when this data-set will be combined with other datasets, we first add a column of static values, defining the `datasetName` of this particular data:
datasetname = "Ecological Archives E090-118-D1."
# Adding this static value as a new column `datasetName`:
# <div class="alert alert-success">
#
# **EXERCISE 2**
#
# Add a new column, `datasetName`, to the survey data set with `datasetname` as value for all of the records (static value for the entire data set)
#
# <details><summary>Hints</summary>
#
# - When a column does not exist, a new `df["a_new_column"]` can be created by assigning a value to it.
# - No `for`-loop is required, as Pandas will automatically broadcast a single string value to each of the rows in the `DataFrame`.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing2.py
# -
# ### Cleaning the `sex_char` column into a DwC called [sex](http://rs.tdwg.org/dwc/terms/#sex) column
# <div class="alert alert-success">
#
# **EXERCISE 3**
#
# - Get a list of the unique values for the column `sex_char`.
#
# <details><summary>Hints</summary>
#
# - To find the unique values, look for a function called `unique` (remember `SHIFT`+`TAB` combination to explore the available methods/attributes?)
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing3.py
# -
# So, apparently, more information is provided in this column, whereas according to the [metadata](http://esapubs.org/archive/ecol/E090/118/Portal_rodent_metadata.htm) information, the sex information should be either `M` (male) or `F` (female). We will create a column, named `sex` and convert the symbols to the corresponding sex, taking into account the following mapping of the values (see [metadata](http://esapubs.org/archive/ecol/E090/118/Portal_rodent_metadata.htm) for more details):
# * `M` -> `male`
# * `F` -> `female`
# * `R` -> `male`
# * `P` -> `female`
# * `Z` -> nan
#
# At the same time, we will save the original information of the `sex_char` in a separate column, called `verbatimSex`, as a reference in case we need the original data later.
# In summary, we have to:
# * rename the `sex_char` column to `verbatimSex`
# * create a new column with the name `sex`
# * map the original values of the `sex_char` to the values `male` and `female` according to the mapping above
# First, let's convert the name of the column header `sex_char` to `verbatimSex` with the `rename` function:
survey_data = survey_data.rename(columns={'sex_char': 'verbatimSex'})
# <div class="alert alert-success">
#
# **EXERCISE 4**
#
# - Express the mapping of the values (e.g. `M` -> `male`) into a Python dictionary object with the variable name `sex_dict`. `Z` values correspond to _Not a Number_, which can be defined as `np.nan`.
# - Use the `sex_dict` dictionary to replace the values in the `verbatimSex` column to the new values and save the mapped values in a new column 'sex' of the DataFrame.
#
# <details><summary>Hints</summary>
#
# - A dictionary is a Python standard library data structure, see https://docs.python.org/3/tutorial/datastructures.html#dictionaries - no Pandas magic involved when you need a key/value mapping.
# - When you need to replace values, look for the Pandas method `replace`.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing4.py
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing5.py
# -
# Checking the current frequency of values of the resulting `sex` column (this should result in the values `male`, `female` and `nan`):
survey_data["sex"].unique()
# To check what the frequency of occurrences is for male/female of the categories, a bar chart is a possible representation:
# <div class="alert alert-success">
#
# **EXERCISE 5**
#
# - Make a horizontal bar chart comparing the number of male, female and unknown (`NaN`) records in the data set.
#
# <details><summary>Hints</summary>
#
# - Pandas provides a shortcut method `value_counts` which works on Pandas `Series` to count unique values. Explore the documentation of the `value_counts` method to include the `NaN` values as well.
# - Check in the help of the Pandas plot function for the `kind` parameter.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing6.py
# -
# <div class="alert alert-warning">
#
# <b>NOTE</b>: The usage of `groupby` combined with the `size` of each group would be an option as well. However, the latter does not support to count the `NaN` values as well. The `value_counts` method does support this with the `dropna=False` argument.
#
# </div>
# ### Solving double entry field by decoupling
# When checking the species unique information:
survey_data["species"].unique()
survey_data.head(10)
# There apparently exists a double entry: `'DM and SH'`, which basically defines two records and should be decoupled to two individual records (i.e. rows). Hence, we should be able to create an additional row based on this split. To do so, Pandas provides a dedicated function since version 0.25, called `explode`. Starting from a small subset example:
example = survey_data.loc[7:10, "species"]
example
# Using the `split` method on strings, we can split the string using a given character, in this case the word `and`:
example.str.split("and")
# The `explode` method will create a row for each element in the list:
example_split = example.str.split("and").explode()
example_split
# Hence, the `DM` and `SH` are now enlisted in separate rows. Other rows remain unchanged. The only remaining issue is the spaces around the characters:
example_split.iloc[1], example_split.iloc[2]
# Which we can solve again using the string method `strip`, removing the spaces before and after the characters:
example_split.str.strip()
# To make this reusable, let's create a dedicated function to combine these steps, called `solve_double_field_entry`:
def solve_double_field_entry(df, keyword="and", column="verbatimEventDate"):
"""Split on keyword in column for an enumeration and create extra record
Parameters
----------
df: pd.DataFrame
DataFrame with a double field entry in one or more values
keyword: str
word/character to split the double records on
column: str
column name to use for the decoupling of the records
"""
df = df.copy() # copy the input DataFrame to avoid editing the original
df[column] = df[column].str.split(keyword)
df = df.explode(column)
df[column] = df[column].str.strip() # remove white space around the words
return df
# The function takes a `DataFrame` as input, splits the record into separate rows and returns an updated `DataFrame`. We can use this function to get an update of the `DataFrame`, with an additional row (observation) added by decoupling the specific field. Let's apply this new function.
# <div class="alert alert-success">
#
# **EXERCISE 6**
#
# - Use the function `solve_double_field_entry` to update the `survey_data` by decoupling the double entries. Save the result as a variable `survey_data_decoupled`.
#
# <details><summary>Hints</summary>
#
# - As we added a 'docstring' to the function, we can check our own documentation to know how to use the function and which inputs we should provide. You can use `SHIFT` + `TAB` to explore the documentation just like any other function.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing7.py
# -
survey_data_decoupled["species"].unique()
survey_data_decoupled.head(11)
# ### Create new occurrence identifier
# The `record_id` is no longer a unique identifier for each observation after the decoupling of this data set. We will make a new data set specific identifier, by adding a column called `occurrenceID` that takes a new counter as identifier. As a simple and straightforward approach, we will use a new counter for the whole dataset, starting with 1:
np.arange(1, len(survey_data_decoupled) + 1, 1)
# To create a new column with header `occurrenceID` with the values 1 -> 35550 as field values:
survey_data_decoupled["occurrenceID"] = np.arange(1, len(survey_data_decoupled) + 1, 1)
# To overcome the confusion on having both a `record_id` and `occurrenceID` field, we will remove the `record_id` term:
survey_data_decoupled = survey_data_decoupled.drop(columns="record_id")
# Hence, columns can be `drop`-ped out of a DataFrame
survey_data_decoupled.head(10)
# ### Converting the date values
# In the survey data set we received, the `month`, `day`, and `year` columns are containing the information about the date, i.e. `eventDate` in DarwinCore terms. We want this data in a ISO format `YYYY-MM-DD`. A convenient Pandas function is the usage of `to_datetime`, which provides multiple options to interpret dates. One of the options is the automatic interpretation of some 'typical' columns, like `year`, `month` and `day`, when passing a `DataFrame`.
# +
# pd.to_datetime(survey_data_decoupled[["year", "month", "day"]]) # uncomment the line and test this statement
# -
# This is not working, not all dates can be interpreted... We should get some more information on the reason of the errors. By using the option `coerce`, the problem makers will be labeled as a missing value `NaT`. We can count the number of dates that can not be interpreted:
sum(pd.to_datetime(survey_data_decoupled[["year", "month", "day"]], errors='coerce').isna())
# <div class="alert alert-success">
#
# **EXERCISE 7**
#
# - Make a selection of `survey_data_decoupled` containing those records that can not correctly be interpreted as date values and save the resulting `DataFrame` as a new variable `trouble_makers`
#
# <details><summary>Hints</summary>
#
# - The result of the `.isna()` method is a `Series` of boolean values, which can be used to make a selection (so called boolean indexing or filtering)
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing8.py
# -
# Checking some charactersitics of the trouble_makers:
trouble_makers.head()
trouble_makers["day"].unique()
trouble_makers["month"].unique()
trouble_makers["year"].unique()
# The issue is the presence of day `31` during the months April and September of the year 2000. At this moment, we would have to recheck the original data in order to know how the issue could be solved. Apparently, - for this specific case - there has been a data-entry problem in 2000, making the `31` days during this period should actually be `30`. It would be optimal to correct this in the source data set, but for the exercise, we will correct it here.
# <div class="alert alert-success">
#
# **EXERCISE 8**
#
# - Assign in the `DataFrame` `survey_data_decoupled` all of the troublemakers `day` values the value 30 instead of 31.
#
# <details><summary>Hints</summary>
#
# - No `for`-loop is required, but use the same boolean mask to assign the new value to the correct rows.
# - Check `pandas_03b_indexing.ipynb` for the usage of `loc` and `iloc` to assign new values.
# - With `loc`, specify both the selecting for the rows and for the columns (`df.loc[row_indexer, column_indexer] = ..`).
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing9.py
# -
# Now, we do the parsing again to create a proper `eventDate` field, containing the dates:
survey_data_decoupled["eventDate"] = \
pd.to_datetime(survey_data_decoupled[["year", "month", "day"]])
# <div class="alert alert-success">
#
# **EXERCISE 9**
#
# - Check the number of observations for each year. Create a horizontal bar chart with the number of rows/observations for each year.
#
# <details><summary>Hints</summary>
#
# - To get the total number of observations, both the usage of `value_counts` as using `groupby` + `size` will work. `value_counts` is a convenient function when all you need to do is counting rows.
# - When using `value_counts`, the years in the index will no longer be in ascending order. You can chain methods and include a `sort_index()` method to sort these again.
#
# </details>
#
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing10.py
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing11.py
# -
survey_data_decoupled.head()
# Currently, the dates are stored in a python specific date format:
survey_data_decoupled["eventDate"].dtype
# This is great, because it allows for many functionalities using the `.dt` accessor:
survey_data_decoupled.eventDate.dt #add a dot (.) and press TAB to explore the date options it provides
# <div class="alert alert-success">
#
# **EXERCISE 10**
#
# - Create a horizontal bar chart with the number of records for each year (cfr. supra), but without using the column `year`, using the `eventDate` column directly.
#
# <details><summary>Hints</summary>
#
# - Check the `groupby` + `size` solution of the previous exercise and use this to start with. Replace the `year` inside the `groupby` method...
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing12.py
# -
# We actually do not need the `day`, `month`, `year` columns anymore, but feel free to use what suits you best.
# <div class="alert alert-success">
#
# **EXERCISE 11**
#
# - Create a bar chart with the number of records for each day of the week (`dayofweek`)
#
# <details><summary>Hints</summary>
#
# - Pandas has an accessor for `dayofweek` as well.
# - You can specify the days of the week yourself to improve the plot, or use the Python standard library `calendar.day_name` (import the calendar module first) to get the names.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing13.py
# -
# When saving the information to a file (e.g. `CSV`-file), this data type will be automatically converted to a string representation. However, we could also decide to explicitly provide the string format the dates are stored (losing the date type functionalities), in order to have full control on the way these dates are formatted:
survey_data_decoupled["eventDate"] = survey_data_decoupled["eventDate"].dt.strftime('%Y-%m-%d')
survey_data_decoupled["eventDate"].head()
# For the remainder, let's remove the day/year/month columns.
survey_data_decoupled = survey_data_decoupled.drop(columns=["day", "month", "year"])
# ## 2. Add species names to dataset
# The column `species` only provides a short identifier in the survey overview. The name information is stored in a separate file `species.csv`. We want our data set to include this information, read in the data and add it to our survey data set:
# <div class="alert alert-success">
#
# **EXERCISE 12**
#
# - Read in the 'species.csv' file and save the resulting `DataFrame` as variable `species_data`.
#
# <details><summary>Hints</summary>
#
# - Check the delimiter (`sep`) parameter of the `read_csv` function.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing14.py
# -
species_data.head()
# ### Fix a wrong acronym naming
# When reviewing the metadata, you see that in the data-file the acronym `NE` is used to describe `Neotoma albigula`, whereas in the [metadata description](http://esapubs.org/archive/ecol/E090/118/Portal_rodent_metadata.htm), the acronym `NA` is used.
# <div class="alert alert-success">
#
# **EXERCISE 13**
#
# - Convert the value of 'NE' to 'NA' by using Boolean indexing/Filtering for the `species_id` column.
#
# <details><summary>Hints</summary>
#
# - To assign a new value, use the `loc` operator.
# - With `loc`, specify both the selecting for the rows and for the columns (`df.loc[row_indexer, column_indexer] = ..`).
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing15.py
# -
# ### Merging surveys and species
# As we now prepared the two series, we can combine the data, using again the `pd.merge` operation.
# We want to add the data of the species to the survey data, in order to see the full species names in the combined data table.
# <div class="alert alert-success">
#
# **EXERCISE 14**
#
# Combine the DataFrames `survey_data_plots` and the `DataFrame` `species_data` by adding the corresponding species information (name, class, kingdom,..) to the individual observations. Assign the output to a new variable `survey_data_species`.
#
# <details><summary>Hints</summary>
#
# - This is an example of a database JOIN operation. Pandas provides the `pd.merge` function to join two data sets using a common identifier.
# - Take into account that our key-column is different for `species_data` and `survey_data_plots`, respectively `species` and `species_id`. The `pd.merge()` function has `left_on` and `right_on` keywords to specify the name of the column in the left and right `DataFrame` to merge on.
#
# </details>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing16.py
# -
len(survey_data_species) # check length after join operation
# The join is ok, but we are left with some redundant columns and wrong naming:
survey_data_species.head()
# We do not need the columns `species_x` and `species_id` column anymore, as we will use the scientific names from now on:
survey_data_species = survey_data_species.drop(["species_x", "species_id"], axis=1)
# The column `species_y` could just be named `species`:
survey_data_species = survey_data_species.rename(columns={"species_y": "species"})
survey_data_species.head()
# + tags=[]
len(survey_data_species)
# -
# ## 3. Add coordinates from the plot locations
# ### Loading the coordinate data
# The individual plots are only identified by a `plot` identification number. In order to provide sufficient information to external users, additional information about the coordinates should be added. The coordinates of the individual plots are saved in another file: `plot_location.xlsx`. We will use this information to further enrich our data set and add the Darwin Core Terms `decimalLongitude` and `decimalLatitude`.
# <div class="alert alert-success">
#
# **EXERCISE 15**
#
# - Read the excel file 'plot_location.xlsx' and store the data as the variable `plot_data`, with 3 columns: plot, xutm, yutm.
#
# <details><summary>Hints</summary>
#
# - Pandas read methods all have a similar name, `read_...`.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing17.py
# -
plot_data.head()
# ### Transforming to other coordinate reference system
# These coordinates are in meters, more specifically in the [UTM 12 N](https://en.wikipedia.org/wiki/Universal_Transverse_Mercator_coordinate_system) coordinate system. However, the agreed coordinate representation for Darwin Core is the [World Geodetic System 1984 (WGS84)](http://spatialreference.org/ref/epsg/wgs-84/).
#
# As this is not a GIS course, we will shortcut the discussion about different projection systems, but provide an example on how such a conversion from `UTM12N` to `WGS84` can be performed with the projection toolkit `pyproj` and by relying on the existing EPSG codes (a registry originally setup by the association of oil & gas producers).
# First, we define out two projection systems, using their corresponding EPSG codes:
from pyproj import Transformer
transformer = Transformer.from_crs("EPSG:32612", "epsg:4326")
# The reprojection can be done by the function `transform` of the projection toolkit, providing the coordinate systems and a set of x, y coordinates. For example, for a single coordinate, this can be applied as follows:
transformer.transform(681222.131658, 3.535262e+06)
# Such a transformation is a function not supported by Pandas itself (it is in https://geopandas.org/). In such an situation, we want to _apply_ a custom function to _each row of the DataFrame_. Instead of writing a `for` loop to do this for each of the coordinates in the list, we can `.apply()` this function with Pandas.
# <div class="alert alert-success">
#
# **EXERCISE 16**
#
# Apply the pyproj function `transform` to plot_data, using the columns `xutm` and `yutm` and save the resulting output in 2 new columns, called `decimalLongitude` and `decimalLatitude`:
#
# - Create a function `transform_utm_to_wgs` that takes a row of a `DataFrame` and returns a `Series` of two elements with the longitude and latitude.
# - Test this function on the first row of `plot_data`
# - Now `apply` this function on all rows (use the `axis` parameter correct)
# - Assign the result of the previous step to `decimalLongitude` and `decimalLatitude` columns
#
# <details><summary>Hints</summary>
#
# - Convert the output of the transformer to a Series before returning (`pd.Series(....)`)
# - A convenient way to select a single row is using the `.loc[0]` operator.
# - `apply` can be used for both rows (`axis` 1) as columns (`axis` 0).
# - To assign two columns at once, you can use a similar syntax as for selecting multiple columns with a list of column names (`df[['col1', 'col2']]`).
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing18.py
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing19.py
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing20.py
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing21.py
# -
plot_data.head()
# The above function `transform_utm_to_wgs` you have created is a very specific function that knows the structure of the `DataFrame` you will apply it to (it assumes the 'xutm' and 'yutm' column names). We could also make a more generic function that just takes a X and Y coordinate and returns the `Series` of converted coordinates (`transform_utm_to_wgs2(X, Y)`).
#
# An alternative to apply such a custom function to the `plot_data` `DataFrame` is the usage of the `lambda` construct, which lets you specify a function on one line as an argument:
#
# transformer = Transformer.from_crs("EPSG:32612", "epsg:4326")
# plot_data.apply(lambda row : transformer.transform(row['xutm'], row['yutm']), axis=1)
#
#
# <div class="alert alert-warning">
#
# __WARNING__
#
# Do not abuse the usage of the `apply` method, but always look for an existing Pandas function first as these are - in general - faster!
#
# </div>
# ### Join the coordinate information to the survey data set
# We can extend our survey data set with this coordinate information. Making the combination of two data sets based on a common identifier is completely similar to the usage of `JOIN` operations in databases. In Pandas, this functionality is provided by [`pd.merge`](http://pandas.pydata.org/pandas-docs/stable/merging.html#database-style-DataFrame-joining-merging).
#
# In practice, we have to add the columns `decimalLongitude`/`decimalLatitude` to the current data set `survey_data_decoupled`, by using the plot identification number as key to join.
# <div class="alert alert-success">
#
# **EXERCISE 17**
#
# - Extract only the columns to join to our survey dataset: the `plot` identifiers, `decimalLatitude` and `decimalLongitude` into a new variable named `plot_data_selection`
#
# <details><summary>Hints</summary>
#
# - To select multiple columns, use a `list` of column names, e.g. `df[["my_col1", "my_col2"]]`
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing22.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 18**
#
# Combine the `DataFrame` `plot_data_selection` and the `DataFrame` `survey_data_decoupled` by adding the corresponding coordinate information to the individual observations using the `pd.merge()` function. Assign the output to a new variable `survey_data_plots`.
#
# <details><summary>Hints</summary>
#
# - This is an example of a database JOIN operation. Pandas provides the `pd.merge` function to join two data sets using a common identifier.
# - The key-column is the `plot`.
#
# </details>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing23.py
# -
survey_data_plots.head()
# The plot locations need to be stored with the variable name `verbatimLocality` indicating the identifier as integer value of the plot:
survey_data_plots = survey_data_plots.rename(columns={'plot': 'verbatimLocality'})
# Let's now save our clean data to a `csv` file, so we can further analyze the data in a following notebook:
survey_data_plots.to_csv("interim_survey_data_species.csv", index=False)
# ## (OPTIONAL SECTION) 4. Using a API service to match the scientific names
# As the current species names are rather short and could eventually lead to confusion when shared with other users, retrieving additional information about the different species in our dataset would be useful to integrate our work with other research. An option is to match our names with an external service to request additional information about the different species.
#
# One of these services is [GBIF API](http://www.gbif.org/developer/species). The service can most easily be illustrated with a small example:<br><br>
# In a new tab blad of the browser, go to the URL [http://www.gbif.org/species/2475532](http://www.gbif.org/species/2475532), which corresponds to the page of `Alcedo atthis` (*ijsvogel* in dutch). One could for each of the species in the list we have do a search on the website of GBIF to find the corresponding page of the different species, from which more information can be extracted manually. However, this would take a lot of time...
#
# Therefore, GBIF (as many other organizations!) provides a service (or API) to extract the same information in a machine-readable way, in order to automate these searches. As an example, let's search for the information of `Alcedo atthis`, using the GBIF API: Go to the URL: [http://api.gbif.org/v1/species/match?name=Alcedo atthis](http://api.gbif.org/v1/species/match?name=%22Alcedo%20atthis%22) and check the output. What we did is a machine-based search on the GBIF website for information about `Alcedo atthis`.
#
# The same can be done using Python. The main library we need to this kind of automated searches is the [`requests` package](http://docs.python-requests.org/en/master/), which can be used to do request to any kind of API out there.
import requests
# ### Example matching with Alcedo Atthis
# For the example of `Alcedo atthis`:
species_name = 'Alcedo atthis'
base_string = 'http://api.gbif.org/v1/species/match?'
request_parameters = {'verbose': False, 'strict': True, 'name': species_name}
message = requests.get(base_string, params=request_parameters).json()
message
# From which we get a dictionary containing more information about the taxonomy of the `Alcedo atthis`.
# In the species data set available, the name to match is provided in the combination of two columns, so we have to combine those to in order to execute the name matching:
genus_name = "Callipepla"
species_name = "squamata"
name_to_match = '{} {}'.format(genus_name, species_name)
base_string = 'http://api.gbif.org/v1/species/match?'
request_parameters = {'strict': True, 'name': name_to_match} # use strict matching(!)
message = requests.get(base_string, params=request_parameters).json()
message
# To apply this on our species data set, we will have to do this request for each of the individual species/genus combination. As, this is a returning functionality, we will write a small function to do this:
# ### Writing a custom matching function
# <div class="alert alert-success">
#
# **EXERCISE 19**
#
# - Write a function, called `name_match` that takes the `genus`, the `species` and the option to perform a strict matching or not as inputs, performs a matching with the GBIF name matching API and return the received message as a dictionary.
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing24.py
# -
# <div class="alert alert-info">
#
# **NOTE**
#
# For many of these API request handling, dedicated packages do exist, e.g. <a href="https://github.com/sckott/pygbif">pygbif</a> provides different functions to do requests to the GBIF API, basically wrapping the request possibilities. For any kind of service, just ask yourself: is the dedicated library providing sufficient additional advantage, or can I easily setup the request myself. (or sometimes: for which the documentation is the best...)<br><br>Many services do exist for a wide range of applications, e.g. scientific name matching, matching of addresses, downloading of data,...
#
# </div>
# Testing our custom matching function:
genus_name = "Callipepla"
species_name = "squamata"
name_match(genus_name, species_name, strict=True)
# However, the matching won't provide an answer for every search:
genus_name = "Lizard"
species_name = "sp."
name_match(genus_name, species_name, strict=True)
# ### Match each of the species names of the survey data set
# Hence, in order to add this information to our survey DataFrame, we need to perform the following steps:
# 1. extract the unique genus/species combinations in our dataset and combine them in single column
# 2. match each of these names to the GBIF API service
# 3. process the returned message:
# * if a match is found, add the information of the columns 'class', 'kingdom', 'order', 'phylum', 'scientificName', 'status' and 'usageKey'
# * if no match was found: nan-values
# 4. Join the DataFrame of unique genus/species information with the enriched GBIF info to the `survey_data_species` data set
# <div class="alert alert-success">
#
# **EXERCISE 20**
#
# - Extract the unique combinations of genus and species in the `survey_data_species` using the function `drop_duplicates()`. Save the result as the variable `unique_species` and remove the `NaN` values using `.dropna()`.
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing25.py
# -
len(unique_species)
# <div class="alert alert-success">
#
# **EXERCISE 21**
#
# - Extract the unique combinations of genus and species in the `survey_data_species` using `groupby`. Save the result as the variable `unique_species`.
#
# <details><summary>Hints</summary>
#
# - As `groupby` needs an aggregation function, this can be `first()` (the first of each group) as well.
# - Do not forget to `reset_index` after the `groupby`.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing26.py
# -
len(unique_species)
# <div class="alert alert-success">
#
# **EXERCISE 22**
#
# - Combine the columns genus and species to a single column with the complete name, save it in a new column named 'name'
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing27.py
# -
unique_species.head()
# To perform the matching for each of the combination, different options do exist (remember `apply`?)
#
# Just to showcase the possibility of using `for` loops in such a situation, let's do the addition of the matched information with a `for` loop. First, we will store everything in one dictionary, where the keys of the dictionary are the index values of `unique_species` (in order to later merge them again) and the values are the entire messages (which are dictionaries on itself). The format will look as following:
#
# ```
# species_annotated = {O: {'canonicalName': 'Squamata', 'class': 'Reptilia', 'classKey': 358, ...},
# 1: {'canonicalName':...},
# 2:...}
# ```
# this will take a bit as we do a request to gbif for each individual species
species_annotated = {}
for key, row in unique_species.iterrows():
species_annotated[key] = name_match(row["genus"], row["species"], strict=True)
# +
#species_annotated # uncomment to see output
# -
# We can now transform this to a pandas `DataFrame`:
# <div class="alert alert-success">
#
# **EXERCISE 23**
#
# - Convert the dictionary `species_annotated` into a pandas DataFrame with the row index the key-values corresponding to `unique_species` and the column headers the output columns of the API response. Save the result as the variable `df_species_annotated`.
#
# <details><summary>Hints</summary>
#
# - The documentation of `pd.DataFrame` says the input van be 'ndarray (structured or homogeneous), Iterable, dict, or DataFrame'.
# - `transpose` can be used to flip rows and columns.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing28.py
# -
df_species_annotated.head()
# ### Select relevant information and add this to the survey data
# <div class="alert alert-success">
#
# **EXERCISE 24**
#
# - Subselect the columns 'class', 'kingdom', 'order', 'phylum', 'scientificName', 'status' and 'usageKey' from the DataFrame `df_species_annotated`. Save it as the variable `df_species_annotated_subset`
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing29.py
# -
df_species_annotated_subset.head()
# <div class="alert alert-success">
#
# **EXERCISE 25**
#
# - Join the `df_species_annotated_subset` information to the `unique_species` overview of species. Save the result as variable `unique_species_annotated`.
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing30.py
# -
unique_species_annotated.head()
# <div class="alert alert-success">
#
# **EXERCISE 26**
#
# - Join the `unique_species_annotated` data to the `survey_data_species` data set, using both the genus and species column as keys. Save the result as the variable `survey_data_completed`.
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/case2_observations_processing31.py
# -
len(survey_data_completed)
survey_data_completed.head()
# Congratulations! You did a great cleaning job, save your result:
survey_data_completed.to_csv("survey_data_completed_.csv", index=False)
# ## Acknowledgements
# * `species.csv` and `survey.csv` are used from the [data carpentry workshop](https://github.com/datacarpentry/python-ecology-lesson) This data is from the paper <NAME>, <NAME>, and <NAME>.
# Brown. 2009. Long-term monitoring and experimental manipulation of a Chihuahuan Desert ecosystem near Portal, Arizona, USA. Ecology 90:1708. http://esapubs.org/archive/ecol/E090/118/
# * The `plot_location.xlsx` is a dummy created location file purely created for this exercise, using the plots location on google maps
# * [GBIF API](http://www.gbif.org/developer/summary)
| notebooks/case2_observations_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.transforms as trans
# Here we plot the results from the normalized average final collisions of all scenarios
data = pd.read_csv('normalized_average_collisions2.csv')
data.head()
# +
# Group scenarios by vehicle number
veh50 = data[data['VehicleNumber']==50]
veh100 = data[data['VehicleNumber']==100]
veh200 = data[data['VehicleNumber']==200]
# +
z50 = np.polyfit(100-veh50['Adherence'], veh50['normave'], 1)
p50 = np.poly1d(z50)
z100 = np.polyfit(100-veh100['Adherence'], veh100['normave'], 1)
p100 = np.poly1d(z100)
z200 = np.polyfit(100-veh200['Adherence'], veh200['normave'], 1)
p200 = np.poly1d(z200)
from matplotlib.legend_handler import HandlerTuple
fig, axs = plt.subplots(1, 1, figsize=(6.0, 4.5))
plt.rc('axes', axisbelow=True)
plt.grid()
fifty1 = plt.scatter(100-veh50['Adherence'], veh50['normave'], color='k', label='50 vehicles', marker='o', s=150.0, edgecolor='k')
fifty2 = plt.plot(100-veh50['Adherence'], p50(100-veh50['Adherence']), color='k', linestyle='--', alpha=0.7)
hundred1 = plt.scatter(100-veh100['Adherence'], veh100['normave'], color='b', label='100 vehicles', marker='^', s=150.0, edgecolor='k')
hundred2 = plt.plot(100-veh100['Adherence'], p100(100-veh100['Adherence']), color='b', linestyle='--', alpha=0.7)
thundred1 = plt.scatter(100-veh200['Adherence'], veh200['normave'], color='r', label='200 vehicles', marker='P', s=150.0, edgecolor='k')
thundred2 = plt.plot(100-veh200['Adherence'], p200(100-veh200['Adherence']), color='r', linestyle='--', alpha=0.7)
plt.legend(fontsize=15.0)
plt.errorbar(100-veh50['Adherence'], veh50['normave'], yerr=veh50['std'], linestyle='None', color='k', capsize=5.0, alpha=0.6, label='50 vehicles')
plt.errorbar(100-veh100['Adherence'], veh100['normave'], yerr=veh100['std'], linestyle='None', color='b', capsize=5.0, alpha=0.6, label='100 vehicles')
plt.errorbar(100-veh200['Adherence'], veh200['normave'], yerr=veh200['std'], linestyle='None', color='r', capsize=5.0, alpha=0.6, label='200 vehicles')
plt.ylabel('Normalized collisions', color='k', fontsize=15.0)
plt.xlabel('Percentage of non-adherence', color='k', fontsize=15.0)
plt.xticks(fontsize=15.0)
plt.yticks(fontsize=15.0)
plt.savefig('Collisions_vs_adherence.png', dpi=300)
plt.show()
# -
| normalized_collisions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Principal Componenet Analysis (PCA)
# The PCA algorithm is a dimensionality reduction algorithm which works really well for datasets which have correlated columns. It combines the features of X in linear combination such that the new components capture the most information of the data.
# The PCA model is implemented in the cuML library and can accept the following parameters:
# 1. svd_solver: selects the type of algorithm used: Jacobi or full (default = full)
# 2. n_components: the number of top K vectors to be present in the output (default = 1)
# 3. random_state: select a random state if the results should be reproducible across multiple runs (default = None)
# 4. copy: if 'True' then it copies the data and removes mean from it else the data will be overwritten with its mean centered version (default = True)
# 5. whiten: if True, de-correlates the components (default = False)
# 6. tol: if the svd_solver = 'Jacobi' then this variable is used to set the tolerance (default = 1e-7)
# 7. iterated_power: if the svd_solver = 'Jacobi' then this variable decides the number of iterations (default = 15)
#
# The cuml implementation of the PCA model has the following functions that one can run:
# 1. Fit: it fits the model with the dataset
# 2. Fit_transform: fits the PCA model with the dataset and performs dimensionality reduction on it
# 3. Inverse_transform: returns the original dataset when the transformed dataset is passed as the input
# 4. Transform: performs dimensionality reduction on the dataset
# 5. Get_params: returns the value of the parameters of the PCA model
# 6. Set_params: allows the user to set the value of the parameters of the PCA model
#
# The model accepts only numpy arrays or cudf dataframes as the input. In order to convert your dataset to cudf format please read the cudf documentation on https://rapidsai.github.io/projects/cudf/en/latest/. For additional information on the PCA model please refer to the documentation on https://rapidsai.github.io/projects/cuml/en/latest/index.html
import numpy as np
import pandas as pd
from sklearn.decomposition import PCA as skPCA
from cuml import PCA as cumlPCA
import cudf
import os
# # Helper Functions
# check if the mortgage dataset is present and then extract the data from it, else do not run
import gzip
def load_data(nrows, ncols, cached = 'data/mortgage.npy.gz'):
if os.path.exists(cached):
print('use mortgage data')
with gzip.open(cached) as f:
X = np.load(f)
X = X[np.random.randint(0,X.shape[0]-1,nrows),:ncols]
else:
# throws FileNotFoundError error if mortgage dataset is not present
raise FileNotFoundError('Please download the required dataset or check the path')
df = pd.DataFrame({'fea%d'%i:X[:,i] for i in range(X.shape[1])})
return df
# +
# this function checks if the results obtained from two different methods (sklearn and cuml) are the equal
from sklearn.metrics import mean_squared_error
def array_equal(a,b,threshold=2e-3,with_sign=True):
a = to_nparray(a)
b = to_nparray(b)
if with_sign == False:
a,b = np.abs(a),np.abs(b)
error = mean_squared_error(a,b)
res = error<threshold
return res
# the function converts a variable from ndarray or dataframe format to numpy array
def to_nparray(x):
if isinstance(x,np.ndarray) or isinstance(x,pd.DataFrame):
return np.array(x)
elif isinstance(x,np.float64):
return np.array([x])
elif isinstance(x,cudf.DataFrame) or isinstance(x,cudf.Series):
return x.to_pandas().values
return x
# -
# # Run tests
# +
# %%time
# nrows = number of samples
# ncols = number of features of each sample
nrows = 2**15
nrows = int(nrows * 1.5)
ncols = 400
X = load_data(nrows,ncols)
print('data',X.shape)
# -
# set parameters for the PCA model
n_components = 10
whiten = False
random_state = 42
svd_solver="full"
# %%time
# use the sklearn PCA on the dataset
pca_sk = skPCA(n_components=n_components,svd_solver=svd_solver,
whiten=whiten, random_state=random_state)
# creates an embedding
result_sk = pca_sk.fit_transform(X)
# %%time
# convert the pandas dataframe to cudf dataframe
X = cudf.DataFrame.from_pandas(X)
# %%time
# use the cuml PCA model on the dataset
pca_cuml = cumlPCA(n_components=n_components,svd_solver=svd_solver,
whiten=whiten, random_state=random_state)
# obtain the embedding of the model
result_cuml = pca_cuml.fit_transform(X)
# calculate the attributes of the two models and compare them
for attr in ['singular_values_','components_','explained_variance_',
'explained_variance_ratio_']:
passed = array_equal(getattr(pca_sk,attr),getattr(pca_cuml,attr))
message = 'compare pca: cuml vs sklearn {:>25} {}'.format(attr,'equal' if passed else 'NOT equal')
print(message)
# compare the results of the two models
passed = array_equal(result_sk,result_cuml)
message = 'compare pca: cuml vs sklearn transformed results %s'%('equal'if passed else 'NOT equal')
print(message)
| cuml/pca_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### 降维技术
# 对数据进行简化降维的原因:
#
# - 使得数据更易使用
# - 使得数据更易可视化
# - 降低很多算法的开销
# - 去除噪声
# - 使得结果易懂
# 数据降维方法主要有三种:
#
# - **主成分分析(Principal Component Analysis, PCA)**
# 在PCA中,数据从原来的坐标系转换到了新的坐标系,新坐标系的选择由原始数据本身决定。第一个新坐标轴选择的是原始数据中方差最大的方向,第二个新坐标轴的选择是和第一个坐标轴正交而且具有最大方差的方向。此过程一直反复,重复次数为原始数据中的特征数目。
# 大部分方差都包含在最前面的几个新坐标轴中。
# - **因子分析(Factor Analysis)**
# 在因子分析中,假设在观察数据的生成中有一些观察不到的隐变量(latent variable)。假设观察数据是这些**隐变量和某些噪声**的线性组合。那么隐变量的数据可能比观察数据的数目少,即通过找到隐变量就可以实现数据降维。
# - **独立成分分析(Independent Component Analysis, ICA)**
# ICA假设数据是从N个数据源生成的,这一点和因子分析有些类似。假设数据为多个数据源的混合观察结果,这些数据源之间在统计上是**相互独立**的,而在PCA中**只假设数据是不相关**的。同因子分析一样,数据源数目少于观察数据数目时,即可实现降维。
# ### PCA
# 将数据转换成前N个主成分的PCA伪代码如下:
#
# 去除平均值
# 计算协方差矩阵
# 计算协方差矩阵的特征值和特征向量
# 将特征值从大到小排序
# 保留最上面的N个特征向量
# 将数据转换到上述N个特征向量构建的空间中
#
import pca
import numpy as np
dataMat = pca.loadDataSet('testSet.txt')
lowDMat, reconMat = pca.pca(dataMat, 1)
np.shape(lowDMat)
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_subplot(111)
ax.scatter(dataMat[:,0].flatten().A[0], dataMat[:,1].flatten().A[0], marker='^', s=20)
ax.scatter(reconMat[:,0].flatten().A[0], reconMat[:,1].flatten().A[0], marker='o', s=10, c='red')
plt.show()
lowDMat, reconMat = pca.pca(dataMat, 2)
fig = plt.figure()
ax = fig.add_subplot(111)
ax.scatter(dataMat[:,0].flatten().A[0], dataMat[:,1].flatten().A[0], marker='^', s=20)
ax.scatter(reconMat[:,0].flatten().A[0], reconMat[:,1].flatten().A[0], marker='o', s=10, c='red')
plt.show()
reload(pca)
dataMat = pca.replaceNanWithMean()
meanVals = np.mean(dataMat, axis=0)
meanRemoved = dataMat - meanVals
covMat = np.cov(meanRemoved, rowvar=0)
print type(covMat)
eigVals, eigVects = np.linalg.eig(np.mat(covMat))
eigVals[:20]
| Ch13_PCA/pca.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # Uranium - Basic Usage
// + active=""
// Welcome to Uranium's Basic Usage page. On this page you can find everything you need to know about how to install, test, build, and quick build using the command line tools provided with the repository.
// -
// ## Windows
// + active=""
// All of the windows commands, except quickbuild, will bring you through the installation process of Yarn and NPM if you do not have them already.
// + active=""
// For windows, all 4 commands are supported (which are):
// -
// - install
// - test
// - build
// - quickbuild
// + active=""
// This allows for some great usage on windows, as this project was originally made for windows devices.
// + [markdown] tags=[]
// #### Install
// + active=""
// The install command is a shorthand tool that allows to quickly install all of the required packages for the repository at once.
// + active=""
// To use the command, all you have to do is simply run:
// -
// ```cmd
// install
// ```
// #### Test
// + active=""
// The test command is a shorthand tool that allows to test your edits to the repository easily.
// + active=""
// To use the command, all you have to do is simply run:
// -
// ```cmd
// test
// ```
// #### Build
// + active=""
// The build command is a shorthand tool that allows to build your modified Uranium repository easily.
// + active=""
// To use the command, all you have to do is simply run:
// -
// ```cmd
// build
// ```
// #### Quickbuild
// + active=""
// The quickbuild command is a shorthand tool that allows to build your modified Uranium repository as a single command, as with the normal build it requires a double-quit and two command true command calls to run.
// + active=""
// To use the command, all you have to do is simply run:
// -
// ```cmd
// quickbuild
// ```
// ## Linux & MacOS
// + active=""
// All of the Linux/MacOS commands, except quickbuild, will tell you how you can install Yarn and NPM.
// + active=""
// For Linux/MacOS, all 4 commands are supported (which are):
// -
// - install
// - test
// - build
// - quickbuild
// + active=""
// This allows for some great usage on Linux and MacOS, even though this project was originally made for windows devices.
// + [markdown] tags=[]
// #### Linstall
// + active=""
// The linstall command is a shorthand tool that allows to quickly install all of the required packages for the repository at once.
// + active=""
// To use the command, all you have to do is run the sh file.
// + active=""
// Linux:
// -
// ```cmd
// ./linstall.sh
// ```
// + active=""
// MacOS:
// -
// ```cmd
// sh linstall.sh
// ```
// #### Ltest
// + active=""
// The ltest command is a shorthand tool that allows to test your edits to the repository easily.
// + active=""
// To use the command, all you have to do is run the sh file.
// + active=""
// Linux:
// -
// ```cmd
// ./ltest.sh
// ```
// + active=""
// MacOS:
// -
// ```cmd
// sh ltest.sh
// ```
// #### Lbuild
// + active=""
// The lbuild command is a shorthand tool that allows to build your modified Uranium repository easily.
// + active=""
// Linux:
// -
// ```cmd
// ./lbuild.sh
// ```
// + active=""
// MacOS:
// -
// ```cmd
// sh lbuild.sh
// ```
// #### Quickbuild
// + active=""
// The quickbuild command is a shorthand tool that allows to build your modified Uranium repository as a single command, as with the normal build it requires a double-quit and two command true command calls to run.
// + active=""
// Linux:
// -
// ```cmd
// ./quickbuild.sh
// ```
// + active=""
// MacOS:
// -
// ```cmd
// sh quickbuild.sh
// ```
// ## Future Support
// + active=""
// Unfortunately, we do not plan on supporting any other operating systems in the future. This does not mean other operating systems aren't supported though. Any form of Linux should be supported along with most versions of MacOS. None of the commands for Windows should work on versions prior to Windows 10 (ex: Windows 7 or Windows XP). We may bring support to other Unix-Like operating systems but that is not gauranteed.
| cli-info.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import datetime
import matplotlib.pyplot as plt
import matplotlib.style as style
import statsmodels.api as sm
import statsmodels.formula.api as smf
import patsy
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score, mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn.linear_model import RidgeCV, Ridge, Lasso
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import KFold
from sklearn.linear_model import lars_path
style.use('fivethirtyeight')
# -
# ## EDA
car_sales = pd.read_csv('Car_Sales_Numbers.csv', index_col=0, parse_dates=True)
car_sales = car_sales.rename(columns={'ALTSALES':'Cars_Sold_e6'})
car_sales.head()
car_sales_diff1 = car_sales.diff().fillna(car_sales)
car_sales_diff2 = car_sales_diff1.diff().fillna(car_sales_diff1)
#car_sales_diff2.isnull().values.any()
fig, ax= plt.subplots(figsize=(8,5))
fig.tight_layout()
ax.plot(car_sales, label='Car_Sales_x10^6')
ax.plot(car_sales_diff2, label='Car_Sales_Diff2', color='red')
ax.legend(loc='best', frameon=False)
plt.xticks(rotation=90)
plt.show()
# +
population = pd.read_csv('Population.csv', parse_dates=True)
population.iloc[:,1] = population.iloc[:,1]/(10**6)
population = population.rename(columns={'LFWA64TTUSM647S':'Population_e6'})
population = population.set_index('DATE')
population.head()
population_diff1 = population.diff().fillna(population)
population_diff2 = population_diff1.diff().fillna(population_diff1)
population_diff2.isnull().values.any()
# +
fig, ax1= plt.subplots(figsize=(8,5))
fig.tight_layout()
color = 'tab:blue'
ax1.plot(population, label='population_x10^6')
ax1.legend(loc='best', frameon=False)
ax1.tick_params(axis='x', rotation=90)
color = 'tab:red'
ax2 = ax1.twinx()
ax2.plot(population_diff2, linestyle='dashed', color='red', label='population_Diff2')
ax2.legend(loc='best', frameon=False)
ax2.set_ylim(-2, 2)
ax1.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
# +
interest_rate = pd.read_csv('Auto_Interest_Rates.csv', parse_dates=True)
#population.iloc[:,1] = population.iloc[:,1]/(10**6)
population = population.rename(columns={'LFWA64TTUSM647S':'Population_e6'})
population = population.set_index('DATE')
population.head()
population_diff1 = population.diff().fillna(population)
population_diff2 = population_diff1.diff().fillna(population_diff1)
population_diff2.isnull().values.any()
# -
djia = pd.read_csv('DJIA.csv', parse_dates=True)
djia = djia.rename(columns={'Adj Close':'DJIA', 'Date':'DATE'})
cols = ['DATE', 'DJIA']
djia = djia[cols]
djia.head()
djia = djia.set_index('DATE')
djia_diff1 = djia.diff().fillna(djia)
djia_diff2 = djia_diff1.diff().fillna(djia_diff1)
djia_diff2.isnull().values.any()
fig, ax= plt.subplots(figsize=(8,5))
fig.tight_layout()
ax.plot(djia, label='DJIA_Data')
ax.plot(djia_diff2, label='DJIA_Diff2')
ax.legend(loc='best', frameon=False)
ax.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
gold = pd.read_csv('Gold_prices.csv', parse_dates=True)
gold = gold.rename(columns={'GOLDAMGBD228NLBM':'Gold_Price_Index'})
gold.fillna(0)
gold = gold.set_index('DATE')
#print(type(gold.Gold_Price_Index[2]))
gold['Gold_Price_Index'] = gold['Gold_Price_Index'].apply(pd.to_numeric, errors='coerce')
gold.head()
gold=gold.fillna(0)
gold_diff1 = gold.diff().fillna(gold)
gold_diff2 = gold_diff1.diff().fillna(gold_diff1)
gold_diff2.isnull().values.any()
fig, ax= plt.subplots(figsize=(8,5))
fig.tight_layout()
ax.plot(gold, label='gold_Data')
ax.plot(gold_diff2, label='gold_Diff1')
ax.legend(loc='best', frameon=False)
ax.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
gdp = pd.read_excel('GDP.xlsx', sheet_name='Monthly Data Sheet', header=1)
gdp = gdp.rename(columns={'Month-Year':'DATE', 'Stock-Watson Real GDP':'GDP'})
gdp['DATE']=gdp['DATE'].astype(str)
cols = ['DATE', 'GDP']
gdp = gdp[cols]
#,'States':'United_States','east':'Northeast','West':'Midwest','Unnamed: 4':'South','Unnamed: 5':'West'})
gdp=gdp.set_index('DATE')
gdp=gdp.fillna(0)
gdp.head()
gdp_diff1 = gdp.diff().fillna(gdp)
gdp_diff2 = gdp_diff1.diff().fillna(gdp_diff1)
gdp_diff2.isnull().values.any()
# +
fig, ax1= plt.subplots(figsize=(8,5))
fig.tight_layout()
color = 'tab:blue'
ax1.plot(gdp, label='GDP')
ax1.legend(loc='best', frameon=False)
ax1.tick_params(axis='x', rotation=90)
color = 'tab:red'
ax2 = ax1.twinx()
ax2.plot(gdp_diff2, linestyle='dashed', color='red', label='GDP_Diff1')
ax2.legend(loc='best', frameon=False)
ax2.set_ylim(-1000, 1000)
ax1.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
# -
wti = pd.read_csv('WTI_Crude_Oil.csv', parse_dates=True)
wti = wti.rename(columns={'DCOILWTICO':'WTI'})#,'States':'United_States','east':'Northeast','West':'Midwest','Unnamed: 4':'South','Unnamed: 5':'West'})
#wti=wti.fillna(0)
wti=wti.set_index('DATE')
wti.head()
wti['WTI']=wti['WTI'].apply(pd.to_numeric, errors='coerce')
wti=wti.fillna(0)
wti_diff1 = wti.diff().fillna(wti)
wti_diff2 = wti_diff1.diff().fillna(wti_diff1)
wti_diff2.isnull().values.any()
# +
fig, ax1= plt.subplots(figsize=(8,5))
fig.tight_layout()
color = 'tab:blue'
ax1.plot(wti, label='WTI')
ax1.legend(loc='upper left', frameon=False)
ax1.tick_params(axis='x', rotation=90)
color = 'tab:red'
ax2 = ax1.twinx()
ax2.plot(wti_diff2, linestyle='dashed', color='red', label='WTI_Diff1')
ax2.legend(loc='best', frameon=False)
ax2.set_ylim(-250, 250)
ax1.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
# -
unemployment = pd.read_csv('Unemployment.csv', parse_dates=True)
unemployment = unemployment.rename(columns={'UNRATE':'Unemp_rate'})#,'States':'United_States','east':'Northeast','West':'Midwest','Unnamed: 4':'South','Unnamed: 5':'West'})
unemployment=unemployment.set_index('DATE')
unemployment.head()
unemployment['Unemp_rate']=unemployment['Unemp_rate'].apply(pd.to_numeric, errors='coerce')
unemployment_diff1 = unemployment.diff().fillna(unemployment)
unemployment_diff2 = unemployment_diff1.diff().fillna(unemployment_diff1)
unemployment_diff2.isnull().values.any()
# +
fig, ax1= plt.subplots(figsize=(8,5))
fig.tight_layout()
color = 'tab:blue'
ax1.plot(unemployment, label='Unemp_rate')
ax1.legend(loc='upper left', frameon=False)
ax1.tick_params(axis='x', rotation=90)
color = 'tab:red'
ax2 = ax1.twinx()
ax2.plot(unemployment_diff2, linestyle='dashed', color='red', label='Unemp_rate_Diff1')
ax2.legend(loc='best', frameon=False)
#ax2.set_ylim(-250, 250)
ax1.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
# -
house = pd.read_csv('house.csv', parse_dates=True)
house = house.rename(columns={'HSN1F':'Houses_Sold_e3'})#:'United_States','east':'Northeast','West':'Midwest','Unnamed: 4':'South','Unnamed: 5':'West'})
house=house.set_index('DATE')
house.head()
house['Houses_Sold_e3']=house['Houses_Sold_e3'].apply(pd.to_numeric, errors='coerce')
house_diff1 = house.diff().fillna(house)
house_diff2 = house_diff1.diff().fillna(house_diff1)
house_diff2.isnull().values.any()
# +
fig, ax1= plt.subplots(figsize=(8,5))
fig.tight_layout()
color = 'tab:blue'
ax1.plot(house, label='Houses_sold_x10^3')
ax1.legend(loc='upper left', frameon=False)
ax1.tick_params(axis='x', rotation=90)
color = 'tab:red'
ax2 = ax1.twinx()
ax2.plot(house_diff2, linestyle='dashed', color='red', label='Houses_sold_Diff1')
ax2.legend(loc='best', frameon=False)
#ax2.set_ylim(-250, 250)
ax1.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
# -
result = pd.merge(car_sales, right=population, left_index=True, right_index=True)
result_diff = pd.merge(car_sales_diff2, right=population_diff2, left_index=True, right_index=True)
result_diff
result = pd.merge(result, right=unemployment, left_index=True, right_index=True)
result_diff = pd.merge(result_diff, right=unemployment_diff2, left_index=True, right_index=True)
result_diff
result = pd.merge(result, right=gold, left_index=True, right_index=True)
result_diff = pd.merge(result_diff, right=gold_diff2, left_index=True, right_index=True)
result_diff
result = pd.merge(result, right=wti, left_index=True, right_index=True)
result_diff = pd.merge(result_diff, right=wti_diff2, left_index=True, right_index=True)
result_diff
result = pd.merge(result, right=djia, left_index=True, right_index=True)
result_diff = pd.merge(result_diff, right=djia_diff2, left_index=True, right_index=True)
result_diff
result = pd.merge(result, right=gdp, left_index=True, right_index=True)
result_diff = pd.merge(result_diff, right=gdp_diff2, left_index=True, right_index=True)
result_diff
result = pd.merge(result, right=house, left_index=True, right_index=True)
result_diff = pd.merge(result_diff, right=house_diff2, left_index=True, right_index=True)
result_diff
result_diff = result_diff.reset_index()
result_diff['DATE']=pd.to_datetime(result_diff.DATE).dt.date
result_diff = result.sort_values('DATE', ascending=True)
#result_diff[result_diff.isnull()]
result.head()
result1 = result.squeeze()
result_diff.info()
#cols = ['Cars_Sold_x10^6','Population_x10^8','Unemp_rate','Gold_Price_Index','WTI','DJIA','GDP','Houses_Sold_x10^3']
#result[cols]=result[cols].apply(pd.to_numeric, errors='coerce')
sns.heatmap(result_diff.corr(), cmap="coolwarm", annot=True, vmin=-1, vmax=1);
# +
#sns.pairplot(result_diff, height=1.2, aspect=1.5);
# -
result_diff.columns
# ## Base Model - LR with Stats Model - Before Feature Engg / Regularization
# +
# Create your feature matrix (X) and target vector (y)
y, X = patsy.dmatrices('Cars_Sold_e6 ~ Population_e6 + Unemp_rate + Gold_Price_Index + WTI + DJIA + GDP + Houses_Sold_e3', data=result_diff, return_type="dataframe")
# Create your model
model = sm.OLS(y, X)
# Fit your model to your training set
fit2 = model.fit()
# Print summary statistics of the model's performance
fit2.summary()
# -
X = result_diff.iloc[:, 1:]
y = result_diff.iloc[:, 0]
X.shape
# +
def split_and_validate(X, y):
'''
For a set of features and target X, y, perform a 80/20 train/val split,
fit and validate a linear regression model, and report results
'''
# perform train/val split
X_train, X_val, y_train, y_val = \
train_test_split(X, y, test_size=0.2, random_state=50)
# fit linear regression to training data
lr_model = LinearRegression()
lr_model.fit(X_train, y_train)
# score fit model on validation data
val_score = lr_model.score(X_val, y_val)
# report results
print('\nValidation R^2 score was:', val_score)
print('Feature coefficient results: \n')
for feature, coef in zip(X.columns, lr_model.coef_):
print(feature, ':', f'{coef:.2f}')
split_and_validate(X, y)
# -
# ## Cross-Validation
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, random_state=10) #hold out 20% of the data for final testing
#this helps with the way kf will generate indices below
#X, y = np.array(X), np.array(y)
kf = KFold(n_splits=4,shuffle=True, random_state=1)
#print(X)
# +
lr_r2 = []
l1_r2 = []
l2_r2 = []
lr_rmse = []
l1_rmse = []
l2_rmse = []
Y_lr_predicted = []
Y_ridge_predicted = []
Y_lasso_predicted =[]
for tr,te in kf.split(X,y):
# print("Fold")
lr = LinearRegression()
l1 = Ridge(alpha=0.1)
l2 = Lasso(alpha=0.01)
X_tr,X_te = X.iloc[tr] ,X.iloc[te]
y_tr,y_te = y.iloc[tr] ,y.iloc[te]
scale = StandardScaler()
#scale.fit(X_tr)
X_tr_sc = scale.fit_transform(X_tr)
X_te_sc = scale.transform(X_te)
lr.fit(X_tr_sc,y_tr)
# print("L1")
l1.fit(X_tr_sc,y_tr)
# print("L2")
l2.fit(X_tr_sc,y_tr)
lr_r2.append(r2_score(y_te,lr.predict(X_te_sc)))
l1_r2.append(r2_score(y_te,l1.predict(X_te_sc)))
l2_r2.append(r2_score(y_te,l2.predict(X_te_sc)))
Y_lr_predicted = lr.predict(X)
Y_ridge_predicted = l1.predict(X)
Y_lasso_predicted = l2.predict(X)
lr_rmse.append(np.sqrt(mean_squared_error(y_te,lr.predict(X_te_sc))))
l1_rmse.append(np.sqrt(mean_squared_error(y_te,l1.predict(X_te_sc))))
l2_rmse.append(np.sqrt(mean_squared_error(y_te,l2.predict(X_te_sc))))
print(f'Lin_Reg Rsquared Value:{np.mean(lr_r2)}, RMSE: {np.mean(lr_rmse)}')
print('\r\n')
print(f'Ridge Rsquared Value:{np.mean(l1_r2)}, RMSE: {np.mean(l1_rmse)}')
print('\r\n')
print(f'Lasso Rsquared Value:{np.mean(l2_r2)}, RMSE: {np.mean(l2_rmse)}')
for feature, coef in zip(X.columns, lr.coef_):
print(feature, ':', f'{coef:.2f}')
#print(X)
#print(Y_lr_predicted)
# -
plt.hist(X_tr_sc[:,6])
# +
## Tuning Regularization
# -
#Mean Absolute Error (MAE)
def mae(y_true, y_pred):
return np.mean(np.abs(y_pred - y_true))
# +
alphalist = 10**(np.linspace(-4,2,400))
err_vec_val = np.zeros(len(alphalist))
err_vec_train = np.zeros(len(alphalist))
for i,curr_alpha in enumerate(alphalist):
# note the use of a new sklearn utility: Pipeline to pack
# multiple modeling steps into one fitting process
steps = [('standardize', StandardScaler()),
('lasso', Lasso(alpha = curr_alpha))]
pipe = Pipeline(steps)
pipe.fit(X_tr, y_tr)
val_set_pred = pipe.predict(X_te)
err_vec_val[i] = mae(y_te, val_set_pred)
plt.plot(np.log10(alphalist), err_vec_val)
# -
# +
result_diff_predict = result_diff.copy()
result_diff_predict['Y_ridge_predicted']=Y_ridge_predicted
result_diff_predict.head()
fig, ax1= plt.subplots(figsize=(8,5))
fig.tight_layout()
color = 'tab:blue'
ax1.plot('Y_ridge_predicted', data=result_diff_predict)#label='Y_ridge_predicted',
ax1.legend(loc='upper left', frameon=False)
ax1.tick_params(axis='x', rotation=90)
color = 'tab:red'
ax2 = ax1.twinx()
ax2.plot(car_sales, linestyle='dashed', color='red', label='Cars_Sold_Actual')#, data=result_diff)
ax2.legend(loc='best', frameon=False)
#ax2.set_ylim(-250, 250)
ax1.xaxis.set_major_locator(plt.MaxNLocator(10))
plt.xticks(rotation=90)
plt.show()
# -
std = StandardScaler()
std.fit(X_tr)
X_tr = std.transform(X_tr)
print("Computing regularization path using the LARS ...")
alphas, _, coefs = lars_path(X_tr, y_tr, method='lasso')
# +
# plotting the LARS path
xx = np.sum(np.abs(coefs.T), axis=1)
xx /= xx[-1]
plt.figure(figsize=(10,10))
plt.plot(xx, coefs.T)
ymin, ymax = plt.ylim()
plt.vlines(xx, ymin, ymax, linestyle='dashed')
plt.xlabel('|coef| / max|coef|')
plt.ylabel('Coefficients')
plt.title('LASSO Path')
plt.axis('tight')
plt.legend(X_train.columns)
plt.show()
# -
| code/Dataset/MacroEconDatasets/.ipynb_checkpoints/Sales_Linear_Model_2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Importing Data with NumPy
import numpy as np
# ### np.loadtxt() vs np.genfromtxt()
lendingCoDataNumeric=np.loadtxt("LendingCompanyNumericData.csv",delimiter=',')
lendingCoDataNumeric
lendingCoDataNumeric1=np.genfromtxt("LendingCompanyNumericData.csv",delimiter=',')
lendingCoDataNumeric1
np.array_equal(lendingCoDataNumeric,lendingCoDataNumeric1)
# looks like so far loadtxt and genfromtxt are the same, lets find out from the following code.The main difference is that, the loadtxt is the faster of the two but it breaks when we feed it incomplete or ill formatted data. On the other hand, genfromtxt is slightly slower but can handle missing values. Now lets import a dataset with missing values in it and see what happens
lendingCoDataNumericNAN=np.genfromtxt("LendingCoNumericDataWithNAN.csv",delimiter=';')
lendingCoDataNumericNAN
lendingCoDataNumericNAN=np.loadtxt("LendingCoNumericDataWithNAN.csv",delimiter=';',dtype=str)
lendingCoDataNumericNAN
#we can use this method only when we want to observe the data values and not to perform any
# mathematical operations
# In practice, we often deal with incomplete data. Sometimes we do not create this data ourselves. This is why genfromtxt is a great choice when loading files with numpy
# ### Partial Cleaning While Importing
lendingCoDataNumericNAN=np.genfromtxt("LendingCoNumericDataWithNAN.csv",delimiter=';')
lendingCoDataNumericNAN
# We will use a function called skip_header which is used to remove the first line of our dataset. To remove the first two lines, we use skip_header=2
# +
lendingCoDataNumericNAN=np.genfromtxt("LendingCoNumericDataWithNAN.csv",
delimiter=';',
skip_header=2)
lendingCoDataNumericNAN
#Now comparing the output of this code with that of the previous code, we realize that, the first two
#rows of the first code have been removed
#we can use skip_header=10 if we want to remove the first ten rows from the top
# -
# What if we want to remove columns and not rows? what do we do? let's do this
lendingCoDataNumericNAN=np.genfromtxt("LendingCoNumericDataWithNAN.csv",
delimiter=';',
usecols=(0,1,5))
lendingCoDataNumericNAN
#the above code lets us use only the first, second and the sixt columns
# One interesting thing is that, we can use only the columns we want and also arrange them in the order that we want. Lets do this
lendingCoDataNumericNAN=np.genfromtxt("LendingCoNumericDataWithNAN.csv",
delimiter=';',
usecols=(5,1,0))
lendingCoDataNumericNAN
#the above code rearranges our columns as we instructed
lendingCoDataNumericNAN=np.genfromtxt("LendingCoNumericDataWithNAN.csv",
delimiter=';',
skip_header=2,
skip_footer=2,
usecols=(5,1,0))
lendingCoDataNumericNAN
# Now we can set each of these columns to some variables. This is how we do that.Check the code below
rowOne,rowTwo,rowThree=np.genfromtxt("LendingCoNumericDataWithNAN.csv",
delimiter=';',
skip_header=2,
skip_footer=2,
usecols=(5,1,0))
lendingCoDataNumericNAN
# Oh! Too many values to upack? alright, we can do this. Check the code below
rowOne,rowTwo,rowThree=np.genfromtxt("LendingCoNumericDataWithNAN.csv",
delimiter=';',
skip_header=2,
skip_footer=2,
usecols=(5,1,0),
unpack=True)
lendingCoDataNumericNAN
rowOne
rowThree
# ### String vs Object vs Numbers
lendingCoLT=np.genfromtxt('lendingCoLT.csv',delimiter=',')
lendingCoLT
#we first have to open the file outside this program to see the delimiter whether a colon or a comma
# To get a neater output, we can print the variable. Let's do this
lendingCoLT=np.genfromtxt('lendingCoLT.csv',delimiter=',')
print(lendingCoLT)
#it looks more organized to print out our values than to just call them directly
| Notebooks and Data/Importing text data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="UW1pYG8_r9_d"
# # Lab 2 (Part 2): Regularizing MLPs
#
# **<NAME> - 100438045**
#
# ------------------------------------------------------
# *Deep Learning. Master in Big Data Analytics*
#
# *<NAME> <EMAIL>*
#
# ------------------------------------------------------
#
#
# In this second part of the lab, we'll be using the [Fashion-MNIST dataset](https://github.com/zalandoresearch/fashion-mnist), a drop-in replacement for the MNIST dataset. MNIST is actually quite trivial with neural networks where you can easily achieve better than 97% accuracy. Fashion-MNIST is a set of 28x28 greyscale images of clothes. It's more complex than MNIST, so it's a better representation of the actual performance of your network, and a better representation of datasets you'll use in the real world. You can see a sample below.
# + colab={"base_uri": "https://localhost:8080/", "height": 221} executionInfo={"elapsed": 1638, "status": "ok", "timestamp": 1614601591119, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="TiHYLJaCr9_i" outputId="41c1ad0f-8954-4d2e-bdcb-84072d48fcef"
from IPython.display import Image
from IPython.core.display import HTML
Image(url= "https://miro.medium.com/max/3200/1*QQVbuP2SEasB0XAmvjW0AA.jpeg", width=400, height=200)
# + [markdown] id="XUTNj0umr9_m"
# Our goal is to build a neural network that can take one of these images and predict the digit in the image. Unlike the MNIST case, for this problem you will notice that the model **easily overfits**, so addressing this issue is an important problem here. To do so, we will experiment with early stopping, dropout, and L2 weight regularization.
#
# Note: a big part of the following material is a personal wrap-up of [Facebook's Deep Learning Course in Udacity](https://www.udacity.com/course/deep-learning-pytorch--ud188). So all credit goes for them!!
# + id="_ZPWyYdXr9_n"
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina' #To get figures with high quality!
import numpy as np
import torch
from torch import nn
from torch import optim
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="q_-PTTwwr9_o"
# ## Part I. Download FMNIST with `torchvision`
#
# The code below will download the MNIST dataset, then create training and test datasets for us. It is mostly the same code we used to download MNIST in the previous part of the Lab.
# + id="a7iJsMPur9_p"
import torch
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
# Download and load the training data
trainset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
# + [markdown] id="ma5XLalQr9_p"
# Lets take a look to the mini-batch size and plot a sample.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1606, "status": "ok", "timestamp": 1614601591124, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="z536aiwbr9_q" outputId="d28802f9-9f51-49b7-c72e-f333583a73d5"
dataiter = iter(trainloader) #To iterate through the dataset
images, labels = dataiter.next()
print(type(images))
print(images.shape)
print(labels.shape)
# + [markdown] id="05kLNsV8r9_q"
# This is what one of the images looks like.
# + colab={"base_uri": "https://localhost:8080/", "height": 282} executionInfo={"elapsed": 1835, "status": "ok", "timestamp": 1614601591365, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="AbEEDcDAr9_r" outputId="b2f51d8d-54dd-4dd4-9349-f2254bd7de00"
plt.imshow(images[1].numpy().reshape([28,28]), cmap='Greys_r')
print(f'Digit associated to this is: {labels[1].item()}')
# + [markdown] id="SxH4lrgsr9_r"
# ## Part II. Visualize overfiting
#
# > **Exercise**: Train a Neural Network with four layers, hidden dimmensions 256, 128 and 64 neurons. Use ReLU activation functions, and a log-Softmax output layer. To do so, complete the following steps:
# > - Create a class defining the NN model
# > - Extend the class to incorporate a training method. **Call it trainloop instead of train**. The reason will be clear later.
# > - Train the model for 30 epochs and evaluate train/test performance
# + id="Vjzib0zWr9_r"
# Creating the NN class
class MLP(nn.Module):
def __init__(self, dimx, nlabels, hidden1=256, hidden2=128, hidden3=64):
super().__init__()
# first layer
self.output1 = nn.Linear(dimx,hidden1)
# second layer
self.output2 = nn.Linear(hidden1, hidden2)
# third layer
self.output3 = nn.Linear(hidden2, hidden3)
# fourth layer
self.output4 = nn.Linear(hidden3, nlabels)
# ReLu activation function
self.relu = nn.ReLU()
# log-softmax activation fun.
self.logsoft = nn.LogSoftmax(dim=1)
def forward(self, x):
# first layer
x = self.relu(self.output1(x))
# second layer
x = self.relu(self.output2(x))
# third layer
x = self.relu(self.output3(x))
# fourth layer
x = self.logsoft(self.output4(x))
return x
# + id="ByGpsK0Hr9_s"
# Now we create the class that we will use for training and testing
class MLP_extended(MLP):
def __init__(self, dimx, nlabels, hidden1=256, hidden2=128, hidden3=64, epochs=30, lr = 1e-3):
super().__init__(dimx, nlabels, hidden1=256, hidden2=128, hidden3=64)
# attributes:
self.lr = lr # learning rate
self.optim = optim.Adam(self.parameters(), self.lr) #optimizer
self.epochs = epochs # epochs
self.criterion = nn.NLLLoss()
# create a list to store the loss at each epoch:
self.loss_during_training = []
def trainloop(self, loader):
# optimization loop
for j in range(int(self.epochs)): # looping for all the epochs
# making the error equal to zero:
running_loss = 0
for images, labels in loader:
self.optim.zero_grad()
# get output of value:
out = self.forward(images.view(images.shape[0], -1))
# computing the loss
loss = self.criterion(out, labels)
# adding up to the running loss
running_loss += loss.item()
loss.backward()
# changing the parameters with the optimizer:
self.optim.step()
# appending the avg. running_loss to the epoch error list:
self.loss_during_training.append(running_loss/len(trainloader))
# printing the errors:
if(j % 10 == 0):
print(f'Training Loss after {j} epochs: {self.loss_during_training[-1]}')
def accuracy(self, loader, part):
accuracy = 0
with torch.no_grad():
for images, labels in loader:
logprobs = self.forward(images.view(images.shape[0], -1))
top_p, top_class = logprobs.topk(1, dim=1)
equals = (top_class == labels.view(images.shape[0], -1))
accuracy += torch.mean(equals.type(torch.FloatTensor))
# printing the accuracy:
print(f'The accuracy in the {part} dataset is: {accuracy/len(loader)}')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 146285, "status": "ok", "timestamp": 1614602125402, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="26NvPpGRr9_t" outputId="aca6bf76-bc24-4395-e292-fea25475e4a3"
#YOUR CODE HERE
# training and evaluating the model:
dimx = 28*28
my_MLP = MLP_extended(dimx, 10)
# train the model
my_MLP.trainloop(trainloader)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 14529, "status": "ok", "timestamp": 1614602139915, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="thH9TLCfoy2h" outputId="681dc6e3-6502-4be0-bfc0-af69706ebf38"
# training accuracy
my_MLP.accuracy(trainloader, 'training')
# testing accuracy
my_MLP.accuracy(testloader, 'testing')
# + [markdown] id="6zgNQPxMr9_u"
# In light of the train/test performance, certainly the model is performing significantly better in the train set than in the test set. This is a sign of overfitting. For an early detection of overfitting, we will make use of a **validation set** that we will use to visualize the evolution of the loss function during training.
#
# With the following code we split the train set into one training set (45k images) and a validation set (15k images). We do a naive splitting assuming that the data is randomized. **Keep in mind that in general you can do something smarter than this like K-Fold cross validation**, but here we keep it simple.
#
#
# + id="6stascs7r9_u"
import copy
validloader = copy.deepcopy(trainloader) # Creates a copy of the object
#We take the first 45k images for training
trainloader.dataset.data = trainloader.dataset.data[:45000,:,:]
trainloader.dataset.targets = trainloader.dataset.targets[:45000]
#And the rest for validation
validloader.dataset.data = validloader.dataset.data[45000:,:,:]
validloader.dataset.targets = validloader.dataset.targets[45000:]
# + [markdown] id="hpqmK-PPr9_v"
# > **Exercise**: Modify your code class above so that, during training, everytime an epoch is finished you compute the loss function over the validation set. You must store these values into a list name as `valid_loss_during_training`. When performing this step, do not forget to turn off gradients by using `with torch.no_grad()`.
# >
# >Then, repeat the training (30 epochs) and plot the train/validation loss along epochs. Compute the final train/validation/test performance.
# + id="iv5GAViwr9_w"
#YOUR CODE HERE
# Now add the computation of the loss with respect to the validation set
class MLP_extended2(MLP):
def __init__(self, dimx, nlabels, hidden1=256, hidden2=128, hidden3=64, epochs=30, lr = 1e-3):
super().__init__(dimx, nlabels, hidden1=256, hidden2=128, hidden3=64)
# attributes:
self.lr = lr # learning rate
self.optim = optim.Adam(self.parameters(), self.lr) #optimizer
self.epochs = epochs # epochs
self.criterion = nn.NLLLoss()
# create a list to store the loss at each epoch:
self.loss_during_training = []
# create a list to store the loss over validation:
self.valid_loss_during_training = []
def trainloop(self, trainloader, validloader):
# optimization loop
for j in range(int(self.epochs)): # looping for all the epochs
# making the error equal to zero:
running_loss = 0
valid_loss = 0
for images, labels in trainloader:
self.optim.zero_grad()
# get output of value:
out = self.forward(images.view(images.shape[0], -1))
# computing the loss
loss = self.criterion(out, labels)
# adding up to the running loss
running_loss += loss.item()
loss.backward()
# changing the parameters with the optimizer:
self.optim.step()
# computation of the loss w.r.t the validation:
with torch.no_grad():
for images_valid, labels_valid in validloader:
out_valid = self.forward(images_valid.view(images_valid.shape[0], -1))
loss_valid = self.criterion(out_valid, labels_valid)
valid_loss += loss_valid
# appending the avg. running_loss to the epoch error list:
self.loss_during_training.append(running_loss/len(trainloader))
# appending the validation loss
self.valid_loss_during_training.append(valid_loss/len(validloader))
# printing the errors for both training and validation datasets:
if(j % 5 == 0):
print(f'Training Loss after {j} epochs: {self.loss_during_training[-1]}')
print(f'Validation Loss after {j} epochs: {self.valid_loss_during_training[-1]}')
def accuracy(self, loader, part='Training'):
accuracy = 0
with torch.no_grad():
for images, labels in loader:
logprobs = self.forward(images.view(images.shape[0], -1))
top_p, top_class = logprobs.topk(1, dim=1)
equals = (top_class == labels.view(images.shape[0], -1))
accuracy += torch.mean(equals.type(torch.FloatTensor))
# printing the accuracy:
print(f'The accuracy in the {part} dataset is: {accuracy/len(loader)}')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 474121, "status": "ok", "timestamp": 1614602614030, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="nLHkDspPpFqq" outputId="1de98008-8785-46ae-d0f8-0c043ecfc24d"
# now we train and implement the new method:
my_mlp = MLP_extended2(dimx, 10)
my_mlp.trainloop(trainloader, validloader)
# + id="YpB068VCrRqX"
plt.plot(my_mlp.loss_during_training)
plt.plot(my_mlp.valid_loss_during_training)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Training and Validation Errors during training')
plt.show()
# + id="h0cVo1HisRIS"
# now we get the performances of all datasets:
my_mlp.accuracy(trainloader, 'training')
my_mlp.accuracy(validloader, 'validation')
my_mlp.accuracy(testloader, 'testing')
# + [markdown] id="2R4Gwy8zr9_w"
# If we look at the training and validation losses as we train the network, we can see a phenomenon known as overfitting.
#
# The network learns the training set better and better, resulting in lower training losses. However, it starts having problems generalizing to data outside the training set leading to the validation loss increasing. The ultimate goal of any deep learning model is to make predictions on new data, so we should strive to get the lowest validation loss possible. One option is to use the version of the model with the lowest validation loss, here the one around 8-10 training epochs. This strategy is called *early-stopping*. In practice, you'd save the model frequently as you're training then later choose the model with the lowest validation loss. **Note that with early stopping we are using the validation set to select the appropiate number of epochs.**
#
# > **Exercise:** According to your results, re-train the model again for the right number of epochs (just before the validation loss starts to grow). Compare the train, validation and test performance.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 121867, "status": "ok", "timestamp": 1614602938187, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="sDt3iYGZr9_x" outputId="1e8da216-e232-4e4c-b389-34dc3c287080"
#YOUR CODE HERE
# get the epoch that has the lowest validation loss:
epoch = my_mlp.valid_loss_during_training.index(min(my_mlp.valid_loss_during_training)) + 1 # subtract 1 because of the index
# now we train the model with only that number of epochs:
my_mlp2 = MLP_extended2(dimx, 10, epochs=epoch)
my_mlp2.trainloop(trainloader, validloader)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 135728, "status": "ok", "timestamp": 1614602952054, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="EZ_9MkVsvyF_" outputId="d53dd9f0-b043-4b19-b201-4524726c118b"
# now we see the performance of each case:
my_mlp2.accuracy(trainloader, 'training')
my_mlp2.accuracy(validloader, 'validation')
my_mlp2.accuracy(testloader, 'testing')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 135726, "status": "ok", "timestamp": 1614602952056, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="w2mg43b_wL0U" outputId="3789b28c-be77-4ec9-f51a-af5566828d42"
plt.plot(my_mlp2.loss_during_training)
plt.plot(my_mlp2.valid_loss_during_training)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Training and Validation Errors during training')
plt.show()
# + [markdown] id="9ov2_9cbr9_y"
# ## Part III. Using Dropout Regularization
#
# The most common method to reduce overfitting (outside of early-stopping) is *dropout*, where we randomly drop input units. This forces the network to share information between weights, increasing it's ability to generalize to new data. Adding dropout in PyTorch is straightforward using the [`nn.Dropout`](https://pytorch.org/docs/stable/nn.html#torch.nn.Dropout) module.
#
# The following code corresponds to a 2 layer NN where we use dropout in the intermediate hidden space:
#
#
# ```python
# class MLP_dropout(nn.Module):
# def __init__(self,dimx,hidden1,nlabels): #Nlabels will be 10 in our case
# super().__init__()
#
# self.output1 = nn.Linear(dimx,hidden1)
#
# self.output2 = nn.Linear(hidden1,nlabels)
#
# self.relu = nn.ReLU()
#
# self.logsoftmax = nn.LogSoftmax(dim=1)
#
# # Dropout module with 0.2 drop probability
# self.dropout = nn.Dropout(p=0.2)
#
# def forward(self, x):
# # Pass the input tensor through each of our operations
# x = self.output1(x)
# x = self.relu(x)
# x = self.dropout(x)
# x = self.output2(x)
# x = self.logsoftmax(x) #YOUR CODE HERE
# return x
#
# ```
#
# During training we want to use dropout to prevent overfitting, but during inference we want to use the entire network. So, we need to turn off dropout during validation, testing, and whenever we're using the network to make predictions. To do this, you use `self.eval()`. This sets the model to evaluation mode where the dropout probability is 0. You can turn dropout back on by setting the model to train mode with `self.train()` (**This is why we cannot call our training method `train` anymore**). In general, the pattern for the validation loop will look like this, where you turn off gradients, set the model to evaluation mode, calculate the validation loss and metric, then set the model back to train mode.
#
# ```python
# # turn off gradients
# with torch.no_grad():
#
# # set model to evaluation mode
# self.eval()
#
# # validation pass here
# for images, labels in testloader:
# ...
#
# # set model back to train mode
# self.train()
# ```
#
# > **Exercise:**
# > - Create a new NN class that modifies the previous one by incorporating a dropout step with `p=0.2` after every ReLU non-linearity is applied.
# > - Modified the extended clases to set `model.eval()` when appropiate (do not forget to go back to `model.train()`)
# > - For this new model, plot the evolution of the training and validation losses. Compare with the case with no dropout. Discuss the results. Is early stopping still required? If so, when you should stop training? Compare the train, validation and test performance.
# > - Repeat the experiments for a dropout probability of `p=0.1` and `p=0.3`. Which value provides the best validation performance?
# + executionInfo={"elapsed": 464, "status": "ok", "timestamp": 1614607374298, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="w_ErYlTFr9_z"
# new model applying the dropout option to avoid overfitting
class MLPdrop(nn.Module):
def __init__(self, dimx, nlabels, hidden1=256, hidden2=128, hidden3=64, pr=0.2):
super().__init__()
# first layer
self.output1 = nn.Linear(dimx,hidden1)
# second layer
self.output2 = nn.Linear(hidden1, hidden2)
# third layer
self.output3 = nn.Linear(hidden2, hidden3)
# fourth layer
self.output4 = nn.Linear(hidden3, nlabels)
# ReLu activation function
self.relu = nn.ReLU()
# log-softmax activation fun.
self.logsoft = nn.LogSoftmax(dim=1)
# Dropout method
self.dropout = nn.Dropout(p=pr)
def forward(self, x):
# first layer
x = self.relu(self.output1(x))
# apply dropout method
x = self.dropout(x)
# second layer
x = self.relu(self.output2(x))
# apply dropout method
x = self.dropout(x)
# third layer
x = self.relu(self.output3(x))
# apply dropout method
x = self.dropout(x)
# fourth layer
x = self.logsoft(self.output4(x))
return x
# + executionInfo={"elapsed": 487, "status": "ok", "timestamp": 1614608375302, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="Wsp8O0q8r9_1"
class MLPdrop_extended(MLPdrop):
def __init__(self, dimx, nlabels, hidden1=256, hidden2=128, hidden3=64, pr = 0.2, epochs=30, lr = 1e-3):
super().__init__(dimx, nlabels, hidden1=hidden1, hidden2=hidden2, hidden3=hidden3, pr=pr)
# attributes:
self.lr = lr # learning rate
self.optim = optim.Adam(self.parameters(), self.lr) #optimizer
self.epochs = epochs # epochs
self.criterion = nn.NLLLoss()
# create a list to store the loss at each epoch:
self.loss_during_training = []
# create a list to store the loss over validation:
self.valid_loss_during_training = []
def trainloop(self, trainloader, validloader):
# setting training mode on
# optimization loop
for j in range(int(self.epochs)): # looping for all the epochs
# making the error equal to zero:
running_loss = 0.
valid_loss = 0.
for images, labels in trainloader:
self.optim.zero_grad()
self.train()
# get output of value:
out = self.forward(images.view(images.shape[0], -1))
# computing the loss
loss = self.criterion(out, labels)
# adding up to the running loss
running_loss += loss.item()
loss.backward()
# changing the parameters with the optimizer:
self.optim.step()
# computation of the loss w.r.t the validation:
with torch.no_grad():
self.eval()
for images_valid, labels_valid in validloader:
out_valid = self.forward(images_valid.view(images_valid.shape[0], -1))
loss_valid = self.criterion(out_valid, labels_valid)
valid_loss += loss_valid
self.train()
# appending the avg. running_loss to the epoch error list:
self.loss_during_training.append(running_loss/len(trainloader))
# appending the validation loss
self.valid_loss_during_training.append(valid_loss/len(validloader))
# printing the errors for both training and validation datasets:
if(j % 5 == 0):
print(f'Training Loss after {j} epochs: {self.loss_during_training[-1]}')
print(f'Validation Loss after {j} epochs: {self.valid_loss_during_training[-1]}')
def accuracy(self, loader, part='Training'):
# setting evaluation mode on:
self.eval()
accuracy = 0
with torch.no_grad():
for images, labels in loader:
logprobs = self.forward(images.view(images.shape[0], -1))
top_p, top_class = logprobs.topk(1, dim=1)
equals = (top_class == labels.view(images.shape[0], -1))
accuracy += torch.mean(equals.type(torch.FloatTensor))
self.accu = accuracy/len(loader)
# printing the accuracy:
print(f'The accuracy in the {part} dataset is: {self.accu}')
return self.accu
# + [markdown] id="qSglOX6qfIAM"
# Now we will analyize this method plotting the training and validation loss scores during training
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 468797, "status": "ok", "timestamp": 1614608051713, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="DkVKi5Lxr9_2" outputId="1c6bde20-72c4-434b-af60-72eea1d5cceb"
mlp_drop = MLPdrop_extended(dimx, 10, pr = 0.2)
mlp_drop.trainloop(trainloader, validloader)
# + colab={"base_uri": "https://localhost:8080/", "height": 294} executionInfo={"elapsed": 468306, "status": "ok", "timestamp": 1614608052058, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="uYiR5Z-Pfmjr" outputId="8ee30ed5-899b-4214-a2fe-ec354f4f84bc"
# plotting the training and validation training_losses
plt.plot(mlp_drop.loss_during_training)
plt.plot(mlp_drop.valid_loss_during_training)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Training and Validation Errors with 0.2 Dropout prob.')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 479278, "status": "ok", "timestamp": 1614608066533, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="FQuv4Z27fzy9" outputId="88ea8806-a447-4416-902d-330bfb1bcb2b"
# getting the accuracies of each of the datasets:
acc_02_training = mlp_drop.accuracy(trainloader, 'training')
acc_02_validation = mlp_drop.accuracy(validloader, 'validation')
acc_02_testing = mlp_drop.accuracy(testloader, 'Testing')
# + executionInfo={"elapsed": 446, "status": "ok", "timestamp": 1614608388457, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="0pRHZv6ug6XG"
Performance_02 = {'Training': acc_02_training.item(), 'Validating': acc_02_validation.item(), 'Testing': acc_02_testing.item()}
# + [markdown] id="lACxQcZ0gLaz"
# Now we repeat the process but with dropout probablity of 0.1 and 0.3
# + [markdown] id="WbQrhrewgSbQ"
# **Dropout Probability 0.1**
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 491002, "status": "ok", "timestamp": 1614608888628, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="ictj4eXlr9_3" outputId="9f462972-1634-4bb5-8dad-7398189657f8"
# instantiate and train the model
mlp_drop1 = MLPdrop_extended(dimx, 10, pr = 0.1)
mlp_drop1.trainloop(trainloader, validloader)
# + colab={"base_uri": "https://localhost:8080/", "height": 294} executionInfo={"elapsed": 491466, "status": "ok", "timestamp": 1614608889111, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="uAxs_8czr9_3" outputId="c51fcabe-b128-4f6e-d639-cfc3ec4dbdaa"
# plotting the training and validation training_losses
plt.plot(mlp_drop1.loss_during_training)
plt.plot(mlp_drop1.valid_loss_during_training)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Training and Validation Errors with 0.1 Dropout prob.')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 505631, "status": "ok", "timestamp": 1614608903289, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="fIz8Hu7rr9_4" outputId="eb0e04a4-fd19-4030-d7ca-6fdc90a37bbb"
acc_01_training = mlp_drop1.accuracy(trainloader, 'training')
acc_01_validation = mlp_drop1.accuracy(validloader, 'validation')
acc_01_testing = mlp_drop1.accuracy(testloader, 'testing')
# + executionInfo={"elapsed": 505624, "status": "ok", "timestamp": 1614608903290, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="eHRlA5EwkZPf"
Performance_01 = {'Training': acc_01_training.item(), 'Validating': acc_01_validation.item(), 'Testing': acc_01_testing.item()}
# + [markdown] id="ZGXwFqdmgm37"
# **Dropout Probability 0.3**
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1027035, "status": "ok", "timestamp": 1614609424706, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="2Rp1nPPXgqeM" outputId="dbc66c76-0886-4276-ccfc-fd94cecc8ea0"
# instantiate and train the model
mlp_drop3 = MLPdrop_extended(dimx, 10, pr = 0.3)
mlp_drop3.trainloop(trainloader, validloader)
# + colab={"base_uri": "https://localhost:8080/", "height": 294} executionInfo={"elapsed": 1027375, "status": "ok", "timestamp": 1614609425056, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="nEZ0Buulgvj4" outputId="fc558e56-75bb-4133-c7f1-766fc3f99f2d"
# plotting the training and validation training_losses
plt.plot(mlp_drop3.loss_during_training)
plt.plot(mlp_drop3.valid_loss_during_training)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Training and Validation Errors with 0.3 Dropout prob.')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1042843, "status": "ok", "timestamp": 1614609440542, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="bzANPgsjgvdo" outputId="5f75ebdb-af62-418a-ec04-11b8f3e85b1e"
acc_03_training = mlp_drop3.accuracy(trainloader, 'training')
acc_03_validation = mlp_drop3.accuracy(validloader, 'validation')
acc_03_testing = mlp_drop3.accuracy(testloader, 'testing')
# + executionInfo={"elapsed": 1042843, "status": "ok", "timestamp": 1614609440544, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="GdpUHAWFkdgG"
Performance_03 = {'Training': acc_03_training.item(), 'Validating': acc_03_validation.item(), 'Testing': acc_03_testing.item()}
# + colab={"base_uri": "https://localhost:8080/", "height": 142} executionInfo={"elapsed": 1042834, "status": "ok", "timestamp": 1614609440547, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="f3_wGn_RgvVs" outputId="3875285f-4977-4d80-c79b-5623a2aa7976"
Performance = {'Dropout Prob: 0.1': Performance_01, 'Dropout Prob: 0.2': Performance_02, 'Dropout Prob: 0.3': Performance_03}
pd.DataFrame.from_dict(Performance)
# + [markdown] executionInfo={"elapsed": 1042836, "status": "ok", "timestamp": 1614609440553, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12531941878311717446"}, "user_tz": -60} id="vLCkuxpAmwS-"
# We see in the summary table above that the dropout probability affects the traininset performance, but not really the validation and testing sets. Since we see that there is no significant change between the performances of the three cases. And actually it could be also said that it decays in a small amount.
#
# In general, the dropout functionality can help decrease the overfitting of the training step, but at least in this example it does not really chanage much the testing performance of the model, once it has been trained.
# -
| Deep_Learning/Dense_Neural_Networks/JijonVorbeck_Lab2DNN/JijonVorbeck_Lab_2_Part_2.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:annorxiver]
# language: python
# name: conda-env-annorxiver-py
# ---
# # Calculate Odds Ratios for each Square Bin
# +
# %load_ext autoreload
# %autoreload 2
import csv
from collections import Counter, defaultdict
import json
import lzma
from multiprocessing import Process, Manager
from pathlib import Path
import pickle
import re
import sys
from threading import Thread
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook
from annorxiver_modules.corpora_comparison_helper import (
aggregate_word_counts,
get_term_statistics,
)
from annorxiver_modules.word_bin_helper import lemmatize_tokens, write_lemma_counts
# -
# # Gather Paper Bins Dataframe
pmc_df = pd.read_csv("output/paper_dataset/paper_dataset_tsne_square.tsv", sep="\t")
print(pmc_df.shape)
pmc_df.head()
word_count_folder = Path("../pmc_corpus/pmc_word_counts/")
word_counter_file = "output/app_plots/global_doc_word_counter.tsv.xz"
field_names = ["document", "lemma", "count"]
n_jobs = 3
QUEUE_SIZE = 75000 # Queue Size if too big then will need to make smaller
doc_xpath = "//abstract/sec/*|//abstract/p|//body/sec/*|//body/p"
with Manager() as m:
# Set up the Queue
doc_path_queue = m.JoinableQueue(QUEUE_SIZE)
lemma_queue = m.JoinableQueue(QUEUE_SIZE)
# Start the document object feeder
t = Thread(
target=write_lemma_counts,
args=(word_counter_file, field_names, lemma_queue, n_jobs),
)
t.start()
running_jobs = []
# Start the jobs
for job in range(n_jobs):
p = Process(
target=lemmatize_tokens, args=(doc_xpath, doc_path_queue, lemma_queue)
)
running_jobs.append(p)
p.start()
for idx, row in tqdm_notebook(pmc_df.iterrows()):
doc_path = f"../journals/{row['journal']}/{row['document']}.nxml"
doc_path_queue.put(doc_path)
# Poison pill to end running jobs
for job in running_jobs:
doc_path_queue.put(None)
# Wait for jobs to finish
for job in running_jobs:
job.join()
# Wait until thread is done running
t.join()
with lzma.open(word_counter_file, "rt") as infile:
reader = csv.DictReader(infile, delimiter="\t")
background_bin_dictionaries = defaultdict(Counter)
word_bin_dictionaries = {
squarebin_id: defaultdict(Counter)
for squarebin_id in pmc_df.squarebin_id.unique()
}
document_mapper = dict(zip(pmc_df.document.tolist(), pmc_df.squarebin_id.tolist()))
for line in tqdm_notebook(reader):
squarebin_id = document_mapper[line["document"]]
background_bin_dictionaries.update({line["lemma"]: int(line["count"])})
word_bin_dictionaries[squarebin_id].update({line["lemma"]: int(line["count"])})
# +
cutoff_score = 20
background_sum = sum(background_bin_dictionaries.values())
bin_ratios = {}
for squarebin in tqdm_notebook(word_bin_dictionaries):
bin_dict = word_bin_dictionaries[squarebin]
bin_sum = sum(word_bin_dictionaries[squarebin].values())
# Try and filter out low count tokens to speed function up
filtered_bin_dict = {
lemma: bin_dict[lemma] for lemma in bin_dict if bin_dict[lemma] > cutoff_score
}
if len(filtered_bin_dict) > 0:
bin_dict = filtered_bin_dict
# Calculate odds ratio
bin_words = set(bin_dict.keys())
background_words = set(background_bin_dictionaries.keys())
words_to_compute = bin_words & background_words
word_odd_ratio_records = []
for idx, word in enumerate(words_to_compute):
top = float(bin_dict[word] * background_sum)
bottom = float(background_bin_dictionaries[word] * bin_sum)
word_odd_ratio_records.append(
{"lemma": word, "odds_ratio": np.log(top / bottom)}
)
sorted(word_odd_ratio_records, key=lambda x: x["odds_ratio"], reverse=True)
bin_ratios[squarebin] = word_odd_ratio_records[0:20]
# -
# # Insert Bin Word Associations in JSON File
square_bin_plot_df = pd.read_json(
open(Path("output") / Path("app_plots") / Path("pmc_square_plot.json"))
)
square_bin_plot_df.head()
bin_odds_df = pd.DataFrame.from_records(
[{"bin_id": key, "bin_odds": bin_ratios[key]} for key in bin_ratios]
)
bin_odds_df.head()
(
square_bin_plot_df.merge(bin_odds_df, on=["bin_id"]).to_json(
Path("output") / Path("app_plots") / Path("pmc_square_plot.json"),
orient="records",
lines=False,
)
)
| pmc/journal_recommendation/04_square_bins_word_ratios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #1. Import modules
# + colab={} colab_type="code" executionInfo={"elapsed": 1389, "status": "ok", "timestamp": 1593417151055, "user": {"displayName": "<NAME>.", "photoUrl": "", "userId": "13008728338773453082"}, "user_tz": -180} id="Y9PwncccuAVk"
from torchvision import transforms
from torchvision import datasets
import numpy as np
import PIL
import torch
import torch.nn as nn
import torchvision.datasets as dset
import torch.utils.data
import torchvision
# -
# #2. Choose the torch device
if torch.cuda.is_available():
torch_device = 'cuda:0'
else:
torch_device = 'cpu'
# # 3. Prepare converter
class CnnInfer(torch.nn.Module):
'''
accepts RGB uint8 image as tensor
'''
def __init__(self, cnn_model):
super(CnnInfer, self).__init__()
self.mean = [0.5, 0.5, 0.5]
self.std = [0.5, 0.5, 0.5]
self.cnn = cnn_model
self.cnn.eval()
def forward(self, img):
x = img.permute(2,0,1).to(torch.float) / 255
x = torchvision.transforms.functional.normalize(x, self.mean, self.std).unsqueeze(0) # add batch dimension
y = self.cnn(x)
return y.squeeze(0) # remove batch dimension
# #3. Convert loaded model to script for C++
# +
import json
import utils
# available architectures:
# - densenet121, se_densenet121
# - resnet18, resnet34, resnet50, se_resnet18, se_resnet34, se_resnet50, resnext50, se_resnext50
# - inception_v3, se_inception_v3, xception, se_xception, inception_resnet
cnn_model, frame_size, classes_header = utils.load_model("se_xception", 5, "models/se_xception.ckpt", device=torch_device);
cnn_infer = CnnInfer(cnn_model)
# создаем скрипт
sample_frame_size = (frame_size[0], frame_size[1], 3)
sample = torch.randint(low=0, high=255, size=sample_frame_size, dtype=torch.uint8)
# scripted_model = torch.jit.script(cnn_infer, sample)
scripted_model = torch.jit.trace(cnn_infer, sample)
scripted_model.save('scripted_model.pth')
# метаданные (заголовки, размер изображений и т.п.)
metadata = {
"classes_header": ",".join(classes_header),
"input_size": {
"width": frame_size[1],
"height": frame_size[0],
"depth": 3
}}
with open('metadata.json', 'w') as f:
f.write(json.dumps(metadata, indent=4))
print("please, check metadata:")
print(json.dumps(metadata, indent=4))
# +
import os
import zipfile
output_filename = "script.zip"
with zipfile.ZipFile(output_filename, 'w') as myzip:
myzip.write('scripted_model.pth')
myzip.write('metadata.json')
| cnn_converter/converter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python(tensorflow-gpu)
# language: python
# name: tensorflow-gpu
# ---
# +
import os,time,cv2, sys, math
import tensorflow as tf
import argparse
import numpy as np
import os
from utils import utils, helpers
from builders import model_builder
# sepcify the gpu
os.environ['CUDA_VISIBLE_DEVICES'] = '/device:GPU:0'
# hyper-parameters setting
parser = argparse.ArgumentParser()
parser.add_argument('--image', type=str, default="./images/0016E5_08147_L.png", help='The image you want to predict on.')
parser.add_argument('--checkpoint_path', type=str, default="./checkpoints/latest_model_BiSeNet_CamVid.ckpt", help='The path to the latest checkpoint weights for your model.')
parser.add_argument('--crop_height', type=int, default=640, help='Height of cropped input image to network')
parser.add_argument('--crop_width', type=int, default=800, help='Width of cropped input image to network')
parser.add_argument('--model', type=str, default="BiSeNet", help='The model you are using')
parser.add_argument('--dataset', type=str, default="CamVid", help='The dataset you are using')
args = parser.parse_args(args=[])
class_names_list, label_values = helpers.get_label_info(os.path.join(args.dataset, "class_dict.csv"))
num_classes = len(label_values)
infer_size = (800, 640)
print("\n***** Begin prediction *****")
print("Dataset -->", args.dataset)
print("Model -->", args.model)
print("Crop Height -->", args.crop_height)
print("Crop Width -->", args.crop_width)
print("Num Classes -->", num_classes)
print("Image -->", args.image)
# allow the use of gpu
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess=tf.Session(config=config)
# Use placeholders as formal parameters to reduce the op of the training process
net_input = tf.placeholder(tf.float32,shape=[None,None,None,3])
# load the network (BiseNet, depth-wise BiseNet, depth-wise AAFF, depth-wise AAFF2)
network, _ = model_builder.build_model(args.model, net_input=net_input,
num_classes=num_classes,
crop_width=args.crop_width,
crop_height=args.crop_height,
is_training=False)
sess.run(tf.global_variables_initializer())
print('Loading model checkpoint weights')
saver=tf.train.Saver(max_to_keep=500)
# load the model weights
saver.restore(sess, args.checkpoint_path)
# load the target image for prediction
loaded_image = utils.load_image(args.image)
resized_image =cv2.resize(loaded_image, (args.crop_width, args.crop_height))
input_image = np.expand_dims(np.float32(resized_image[:args.crop_height, :args.crop_width]),axis=0)/255.0
# perform prediction
output_image = sess.run(network, feed_dict={net_input:input_image})
# compute the inference speed
elapse = []
for i in range(50):
start = time.time()
output_image = sess.run(network, feed_dict={net_input:input_image})
duration = time.time() - start
print('time: {:.4f}, about {:.6f} fps'.format(duration, 1 / duration))
elapse.append(duration)
print('Average time: {:.4f}, about {:.6f} fps'.format(np.mean(elapse), 1 / np.mean(elapse)))
# output a colored predicted image
output_image = np.array(output_image[0,:,:,:])
output_image = helpers.reverse_one_hot(output_image)
out_vis_image = helpers.colour_code_segmentation(output_image, label_values)
file_name = utils.filepath_to_name(args.image)
cv2.imwrite("%s_pred.png"%(file_name),cv2.cvtColor(np.uint8(out_vis_image), cv2.COLOR_RGB2BGR))
print("")
print("Finished!")
print("Wrote image " + "%s_pred.png"%(file_name))
# -
import tensorflow as tf
print(tf.__version__)
| predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
pip install kafka-python
from kafka import KafkaConsumer
import matplotlib.pyplot as plt
import json
import threading
consumer = KafkaConsumer('powergrid1',
group_id='powergrid1',
bootstrap_servers=['localhost:9092'],
)
print("consumer started ...")
x = {}
def plot():
global x
for message in consumer:
x[json.loads((message.value).decode("utf-8"))["date"]] = (json.loads((message.value).decode("utf-8"))["volume"])
print(x)
plot_thread = threading.Thread(target=plot)
plot_thread.start()
fig = plt.figure(figsize = (12, 6))
x = dict(sorted(x.items(), key=lambda item: item[0], reverse=False))
plt.bar([*x.keys()][:30], [*x.values()][:30], width=0.5)
plt.xlabel("Date", fontsize=15)
plt.ylabel("Volume", fontsize=15)
plt.xticks(rotation=90)
plt.title("POWERGRIDE SHARES", fontsize=15)
consumer = KafkaConsumer('ntpc1',
group_id='ntpc1',
bootstrap_servers=['localhost:9092'],
)
print("consumer started ...")
x = {}
def plot():
global x
for message in consumer:
x[json.loads((message.value).decode("utf-8"))["date"]] = (json.loads((message.value).decode("utf-8"))["volume"])
print(x)
plot_thread = threading.Thread(target=plot)
plot_thread.start()
fig = plt.figure(figsize = (12, 6))
x = dict(sorted(x.items(), key=lambda item: item[0], reverse=False))
plt.bar([*x.keys()][:30], [*x.values()][:30], width=0.5)
plt.xlabel("Date", fontsize=15)
plt.ylabel("Volume", fontsize=15)
plt.xticks(rotation=90)
plt.title("NTPC SHARES", fontsize=15)
| EV/Date vs Volume/charging point/output/Charging_Point.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import opendatablend as odb
import pandas as pd
# +
# Set the path for the dataset metadata. Find this using the 'Get metadata' button on a dataset e.g. https://www.opendatablend.io/dataset?name=open-data-blend-road-safety
dataset_path = 'https://packages.opendatablend.io/v1/open-data-blend-road-safety/datapackage.json'
# Set your acess key. Note: leaving this blank will result in anonymous/public calls and will consume your free API call allowance.
access_key = ''
# We want Pandas to display up to 100 rows for dataframes
pd.options.display.max_rows = 100
# -
# Download the date dimension and load it into a DataFrame
resource_name = 'date-parquet'
output = odb.get_data(dataset_path, resource_name, access_key=access_key)
df_date = pd.read_parquet(output.data_file_name, columns=['drv_date_key', 'drv_date', 'drv_month_name', 'drv_month_number', 'drv_quarter_name', 'drv_quarter_number', 'drv_year'])
df_date.head()
# Download the road safety-accident-info dimension and load it into a DataFrame
resource_name = 'road-safety-accident-info-parquet'
output = odb.get_data(dataset_path, resource_name, access_key=access_key)
df_accident_info = pd.read_parquet(output.data_file_name, columns=['drv_road_safety_accident_info_key', 'src_road_surface_condition', 'src_speed_limit', 'src_weather_condition', 'src_police_force'])
df_accident_info.head()
# Download the road safety accident fact data for 2017 and load it into a DataFrame
resource_name = 'road-safety-accident-2017-parquet'
output = odb.get_data(dataset_path, resource_name, access_key=access_key)
df_accidents_2017 = pd.read_parquet(output.data_file_name, columns=['drv_accident_date_key', 'drv_road_safety_accident_info_key', 'src_number_of_casualties', 'src_number_of_vehicles'])
# Download the road safety accident fact data for 2018 and load it into a DataFrame
resource_name = 'road-safety-accident-2018-parquet'
output = odb.get_data(dataset_path, resource_name, access_key=access_key)
df_accidents_2018 = pd.read_parquet(output.data_file_name, columns=['drv_accident_date_key', 'drv_road_safety_accident_info_key', 'src_number_of_casualties', 'src_number_of_vehicles'])
# Download the road safety accident fact data for 2019 and load it into a DataFrame
resource_name = 'road-safety-accident-2019-parquet'
output = odb.get_data(dataset_path, resource_name, access_key=access_key)
df_accidents_2019 = pd.read_parquet(output.data_file_name, columns=['drv_accident_date_key', 'drv_road_safety_accident_info_key', 'src_number_of_casualties', 'src_number_of_vehicles'])
df_accidents_combined = pd.concat([df_accidents_2017, df_accidents_2018, df_accidents_2019])
df_accidents_combined
# Extend the accident with the data and road safety accident into dimensions
df_accidents = df_date.merge(df_accidents_combined, left_on='drv_date_key', right_on='drv_accident_date_key').merge(df_accident_info, on='drv_road_safety_accident_info_key')
df_accidents
# Total number of accidents per year
df_accidents.groupby('drv_year').agg({'drv_year': np.size}).rename(columns={"drv_year": "total_accidents"}).sort_index()
df_accidents['number_of_accidents'] = 1
# High-level stats by year
df_accidents[['drv_year', 'number_of_accidents', 'src_number_of_casualties', 'src_number_of_vehicles']].groupby(by=['drv_year']).sum().sort_index()
# High-level stats by year and police force
df_accidents[['drv_year', 'src_police_force', 'number_of_accidents', 'src_number_of_casualties', 'src_number_of_vehicles']].groupby(by=['drv_year', 'src_police_force']).sum().sort_index()
# High-level stats by year and speed limit
df_accidents[['drv_year', 'src_speed_limit', 'number_of_accidents', 'src_number_of_casualties', 'src_number_of_vehicles']].groupby(by=['drv_year', 'src_speed_limit']).sum().sort_index()
| examples/open_data_blend_pandas_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow2]
# language: python
# name: conda-env-tensorflow2-py
# ---
import pandas as pd
import numpy as np
datFile = 'data/foodNetwork_count.json'
tmp1 = pd.read_json(datFile)
print(tmp.drop_duplicates().numLinks.sum())
print(tmp1.drop_duplicates().numLinks.sum())
tmp.sort_values(by = 'numLinks', ascending=False)
| get_foodnetwork_counts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# | [Contents](toc.ipynb) | [Index](index.ipynb) | [PyRosetta Google Drive Setup](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.01-PyRosetta-Google-Drive-Setup.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.00-How-to-Get-Started.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # How to Get Started
# ## Students
# **Step 1:** Sign in to Google Drive.
#
# **Step 2:** Make a new folder called “PyRosetta” in your top level directory on Google Drive.
#
from IPython.display import Image
Image('./Media/PyRosetta-folder.png',width='600')
# **Step 3:** Get your PyRosetta license.
#
# **Step 4:** Download the Linux [PyRosetta package](http://www.pyrosetta.org/dow) that is called "Python-3.7.MinSizeRel".
#
Image('./Media/linux-download.png',width='400')
# **Step 5:** Upload the Linux PyRosetta package (.tar.bz2 file) to the PyRosetta folder in your Google Drive.
#
# **Step 6:** Upload the PyRosetta notebooks folder to the top-level directory of your Google Drive.
Image('./Media/google-drive.png',width='600')
# **Step 7:** Open up “01.01-PyRosetta-Google-Drive-Setup.ipynb” in Google Colab by right-clicking, going under “Open With…” and clicking Google Colaboratory. If you have never used Google Colab before, you may need to first install the Google Colab [app](https://workspace.google.com/u/2/marketplace/app/colaboratory/1014160490159?hl=en&pann=drive_app_widget).
#
# **Step 8:** Follow steps to install PyRosetta. Note: You only need to do this once since this installation will be associated with your Google Drive account.
#
# **Step 9:** In “01.01-PyRosetta-Google-Drive-Setup.ipynb”, follow the steps to install third-party external packages, using `pip`:
#
# `pip install biopython blosc dask dask-jobqueue distributed graphviz jupyter matplotlib numpy pandas py3Dmol scipy seaborn traitlets --user`
#
# **Step 9.1:**
#
# **For Chapter 16, Running PyRosetta in Parallel (and a few other notebooks), you will need to use a specific version of PyRosetta that is built for parallelization.** This is the serialization build. Besides manually building it from the Rosetta C++ source code, the general way to obtain this is through the use of a `conda` environment.
#
# A `conda` environment is a way to run code that has specific versions of required packages, that instead of being installed globally, will be installed as a local virtual environment that you may run whenever you wish. This is extremely useful when some packages require specific versions of other packages, as is the case for some rosetta distributed code.
#
# You will need to pass the username and password of PyRosetta to conda.
# In order to do this, we will create a file in your home directory called `.condarc`. If you already have this file, please edit it instead of overriding it below.
#
# **Here, instead of USERNAME and PASSWORD, enter the USERNAME and PASSWORD you were given while gaining access to PyRosetta.**
#
# ```
# # echo "channels:" >> $HOME/.condarc
# # echo " - https://USERNAME:PASSWORD@conda.graylab.jhu.edu" >> $HOME/.condarc
# # echo " - defaults" >> $HOME/.condarc
# conda env create -f environment.yml
# conda activate PyRosetta.notebooks
# ```
#
# Each time you wish to run this environment, use `conda activate PyRosetta.notebooks` to create the local virtual environment BEFORE running the jupyter notebook. You may wish to put this in your system configuration on startup. It is recommended to activate conda before you run the notebooks in general. The correct kernel is set for them.
#
# **NOTE:**
# When using a notebook with this environment - the python **Kernel** must be set to this env. *In Chapter 16 and other parellel notebooks, this is done for you*, but if you wish to use this environment in other notebooks, make sure to run conda manually change this!
#
# You can do this by looking at the jupyter menu - `Kernel` is after `Cell` and before `Widgets`. The option is 'Change Kernel`. This is how you would run python2 vs python3 or run a kernal with other conda environments you have installed on your computer.
#
# **Step 10:** Open up “01.02-PyRosetta-Google-Drive-Usage-Example.ipynb” in Colab to make sure that PyRosetta was installed properly. You are now ready to move on to the other notebooks!
# **Known Limitations:**
#
# - If you receive the error `AttributeError: module 'pyrosetta.rosetta' has no attribute '__path__'` or `TypeError: int returned non-int (type str)` then serialized functions need to locally import libraries (specifically, add `import pyrosetta`, `import pyrosetta.distributed` and other `pyrosetta` imports to the serialized functions, i.e. any functions passed to `client.submit` or `dask.delayed`).
#
# Example:
#
# ```
# def my_function(packed_pose):
# import pyrosetta
# pyrosetta.MoveMap()
# return packed_pose
# client.submit(my_function, packed_pose)
# ```
#
#
# - If running dask's `LocalCluster(n_workers=2, threads_per_worker=2)`, you may need to adjust to `n_workers=1` and `threads_per_worker=1` if you encounter errors on some systems.
#
#
# - If running dask's `LocalCluster(n_workers=2, threads_per_worker=2)` in a standalone python script, the `LocalCluster` instance must be called within the `if __name__ == "__main__"` block.
#
#
# - If using `Jupyter Lab`, the `pyrosetta.distributed.viewer` visualizer with `py3Dmol` bindings may not function correctly (i.e. the interactive scrollbar widget may not work or the visualizer doesn't appear). To fix this, you may have to install the following Jupyter Lab extensions:
#
# `jupyter labextension install jupyterlab_3dmol`
#
# `jupyter labextension install @jupyter-widgets/jupyterlab-manager`
#
# ## Instructors
# **Step 1:** Please install extensions nbgrader and nbpages.
#
# **Step 2:** Make any changes to the workshops in the "notebooks" folder. (You can follow the directions for students and edit them in Google Colab or on your own computer using a Jupyter Notebooks browser.)
#
# **Step 3:** When you are done making changes, run make-student-nb.bash to automatically generate the student notebooks in a separate folder, the table of contents, and the keyword index. The student notebooks are copies of the instructor's version that have omitted solutions.
# **Chapter contributors:**
#
# - <NAME> (Johns Hopkins University)
# - <NAME>kov (Johns Hopkins University)
# - <NAME> (University of Washington; Lyell Immunopharma)
# <!--NAVIGATION-->
# | [Contents](toc.ipynb) | [Index](index.ipynb) | [PyRosetta Google Drive Setup](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.01-PyRosetta-Google-Drive-Setup.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.00-How-to-Get-Started.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
| notebooks/01.00-How-to-Get-Started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import talib
plt.style.use('fivethirtyeight')
df = pd.read_csv('3665.csv')
df = df.set_index(pd.DatetimeIndex(df['Date'].values))
ShortEMA = df.Close.ewm(span=12, adjust=False).mean() # Fast Moving Average
LongEMA = df.Close.ewm(span=26, adjust=False).mean() # Slow Moving Average
MACD = ShortEMA - LongEMA
signal = MACD.ewm(span=9, adjust=False).mean()
df['MACD'] = MACD
df['Signal Line'] = signal
flag = 0
def InvestStrategy(signal):
BuyStock = []
SellStock = []
global flag
# If MACD > Signal Line, Buy The Stock. If Not, Sell The Stock
for i in range(0, len(signal)):
if signal['MACD'][i] > signal['Signal Line'][i]:
if flag == 0:
BuyStock.append(signal['Close'][i])
SellStock.append(np.nan)
flag = 1
else:
BuyStock.append(np.nan)
SellStock.append(np.nan)
elif signal['MACD'][i] < signal['Signal Line'][i]:
if flag == 1:
SellStock.append(signal['Close'][i])
BuyStock.append(np.nan)
flag = 0
else:
BuyStock.append(np.nan)
SellStock.append(np.nan)
else: # Handling nan values
BuyStock.append(np.nan)
SellStock.append(np.nan)
return (BuyStock, SellStock)
temp = InvestStrategy(df)
df['BuyStockPrice'] = temp[0]
df['SellStockPrice'] = temp[1]
TotalCost = np.sum(df['BuyStockPrice'])
TotalEarn = np.sum(df['SellStockPrice'])
if flag == 1:
TotalEarn = TotalEarn + df['Close'].tail(1)
NetValue = round(TotalEarn - TotalCost, 2)
ReturnRate = round(100 * (TotalEarn - TotalCost) / TotalCost, 2)
# Visualize The Strategy
plt.figure(figsize=(12.2,4.5)) #width = 12.2in, height = 4.5
plt.plot(df.index, MACD, label='STOCK', color = 'red')
plt.plot(df.index, signal, label='SIGNAL LINE', color='blue')
plt.xticks(rotation=45)
plt.legend(loc='upper left')
plt.show()
title = 'Close Price History & Buy / Sell Signals '
my_stocks = df
plt.figure(figsize=(12.2,4.5))
plt.scatter(my_stocks.index, my_stocks['BuyStockPrice'], color = 'green', label='Buy Signal', marker = '^', alpha = 1)
plt.scatter(my_stocks.index, my_stocks['SellStockPrice'], color = 'red', label='Sell Signal', marker = 'v', alpha = 1)
plt.plot(my_stocks['Close'], label='Close Price', alpha = 0.35)
plt.xticks(rotation=45)
plt.title(title)
plt.ylabel('Close Price NTD ($)',fontsize=18)
plt.legend(loc='upper left')
plt.show()
print("Total Cost:", TotalCost, "NTD")
print("Total Earn:", TotalEarn, "NTD")
print("Net Value:", NetValue, "NTD")
print("Rate Of Return:", ReturnRate, "%")
# +
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import talib
plt.style.use('fivethirtyeight')
df = pd.read_csv('4171.csv')
df = df.set_index(pd.DatetimeIndex(df['Date'].values))
ShortEMA = df.Close.ewm(span=12, adjust=False).mean() # Fast Moving Average
LongEMA = df.Close.ewm(span=26, adjust=False).mean() # Slow Moving Average
MACD = ShortEMA - LongEMA
signal = MACD.ewm(span=9, adjust=False).mean()
df['MACD'] = MACD
df['Signal Line'] = signal
flag = 0
def InvestStrategy(signal):
BuyStock = []
SellStock = []
global flag
# If MACD > Signal Line, Buy The Stock. If Not, Sell The Stock
for i in range(0, len(signal)):
if signal['MACD'][i] > signal['Signal Line'][i]:
if flag == 0:
BuyStock.append(signal['Close'][i])
SellStock.append(np.nan)
flag = 1
else:
BuyStock.append(np.nan)
SellStock.append(np.nan)
elif signal['MACD'][i] < signal['Signal Line'][i]:
if flag == 1:
SellStock.append(signal['Close'][i])
BuyStock.append(np.nan)
flag = 0
else:
BuyStock.append(np.nan)
SellStock.append(np.nan)
else: # Handling nan values
BuyStock.append(np.nan)
SellStock.append(np.nan)
return (BuyStock, SellStock)
temp = InvestStrategy(df)
df['BuyStockPrice'] = temp[0]
df['SellStockPrice'] = temp[1]
TotalCost = np.sum(df['BuyStockPrice'])
TotalEarn = np.sum(df['SellStockPrice'])
if flag == 1:
TotalEarn = TotalEarn + df['Close'].tail(1)
NetValue = round(TotalEarn - TotalCost, 2)
ReturnRate = round(100 * (TotalEarn - TotalCost) / TotalCost, 2)
# Visualize The Strategy
plt.figure(figsize=(12.2,4.5)) #width = 12.2in, height = 4.5
plt.plot(df.index, MACD, label='STOCK', color = 'red')
plt.plot(df.index, signal, label='SIGNAL LINE', color='blue')
plt.xticks(rotation=45)
plt.legend(loc='upper left')
plt.show()
title = 'Close Price History & Buy / Sell Signals '
my_stocks = df
plt.figure(figsize=(12.2,4.5))
plt.scatter(my_stocks.index, my_stocks['BuyStockPrice'], color = 'green', label='Buy Signal', marker = '^', alpha = 1)
plt.scatter(my_stocks.index, my_stocks['SellStockPrice'], color = 'red', label='Sell Signal', marker = 'v', alpha = 1)
plt.plot(my_stocks['Close'], label='Close Price', alpha = 0.35)
plt.xticks(rotation=45)
plt.title(title)
plt.ylabel('Close Price NTD ($)',fontsize=18)
plt.legend(loc='upper left')
plt.show()
print("Total Cost:", TotalCost, "NTD")
print("Total Earn:", TotalEarn, "NTD")
print("Net Value:", NetValue, "NTD")
print("Rate Of Return:", ReturnRate, "%")
# +
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import talib
plt.style.use('fivethirtyeight')
df = pd.read_csv('4743.csv')
df = df.set_index(pd.DatetimeIndex(df['Date'].values))
ShortEMA = df.Close.ewm(span=12, adjust=False).mean() # Fast Moving Average
LongEMA = df.Close.ewm(span=26, adjust=False).mean() # Slow Moving Average
MACD = ShortEMA - LongEMA
signal = MACD.ewm(span=9, adjust=False).mean()
df['MACD'] = MACD
df['Signal Line'] = signal
flag = 0
def InvestStrategy(signal):
BuyStock = []
SellStock = []
global flag
# If MACD > Signal Line, Buy The Stock. If Not, Sell The Stock
for i in range(0, len(signal)):
if signal['MACD'][i] > signal['Signal Line'][i]:
if flag == 0:
BuyStock.append(signal['Close'][i])
SellStock.append(np.nan)
flag = 1
else:
BuyStock.append(np.nan)
SellStock.append(np.nan)
elif signal['MACD'][i] < signal['Signal Line'][i]:
if flag == 1:
SellStock.append(signal['Close'][i])
BuyStock.append(np.nan)
flag = 0
else:
BuyStock.append(np.nan)
SellStock.append(np.nan)
else: # Handling nan values
BuyStock.append(np.nan)
SellStock.append(np.nan)
return (BuyStock, SellStock)
temp = InvestStrategy(df)
df['BuyStockPrice'] = temp[0]
df['SellStockPrice'] = temp[1]
TotalCost = np.sum(df['BuyStockPrice'])
TotalEarn = np.sum(df['SellStockPrice'])
if flag == 1:
TotalEarn = TotalEarn + df['Close'].tail(1)
NetValue = round(TotalEarn - TotalCost, 2)
ReturnRate = round(100 * (TotalEarn - TotalCost) / TotalCost, 2)
# Visualize The Strategy
plt.figure(figsize=(12.2,4.5)) #width = 12.2in, height = 4.5
plt.plot(df.index, MACD, label='STOCK', color = 'red')
plt.plot(df.index, signal, label='SIGNAL LINE', color='blue')
plt.xticks(rotation=45)
plt.legend(loc='upper left')
plt.show()
title = 'Close Price History & Buy / Sell Signals '
my_stocks = df
plt.figure(figsize=(12.2,4.5))
plt.scatter(my_stocks.index, my_stocks['BuyStockPrice'], color = 'green', label='Buy Signal', marker = '^', alpha = 1)
plt.scatter(my_stocks.index, my_stocks['SellStockPrice'], color = 'red', label='Sell Signal', marker = 'v', alpha = 1)
plt.plot(my_stocks['Close'], label='Close Price', alpha = 0.35)
plt.xticks(rotation=45)
plt.title(title)
plt.ylabel('Close Price NTD ($)',fontsize=18)
plt.legend(loc='upper left')
plt.show()
print("Total Cost:", TotalCost, "NTD")
print("Total Earn:", TotalEarn, "NTD")
print("Net Value:", NetValue, "NTD")
print("Rate Of Return:", ReturnRate, "%")
# -
| Best Performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from scipy import stats
import itertools
from sklearn import linear_model
from numpy import ones,vstack
from numpy.linalg import lstsq
pd.set_option('display.max_rows', 500)
# +
df = pd.read_csv("../data/Ames_Housing_Price_Data_v2.csv", index_col=0)
df_re=pd.read_csv("../data/Ames_Real_Estate_Data_raw.csv", index_col=0)
df=df.reset_index(drop=False)
df_re=df_re.reset_index(drop=False)
# -
df_re
df_re[df_re['MapRefNo']=='531477050']
merged=pd.merge(df,df_re,how='left', left_on='PID',right_on='MapRefNo')
merged.head(5).T
# merged['Address']=merged['PA-Nmbr'].fillna(' ').astype(str) \
# # + " " + merged['PA-PreD'].fillna(' ').astype(str) \
# # + " " + merged['PA-Strt'].fillna(' ').astype(str) \
# # + " " + merged['PA-StSfx'].fillna(' ').astype(str) \
# # + " " + merged['PA-PostD'].fillna(' ').astype(str)
merged['Address']=merged['Prop_Addr'].fillna(' ').astype(str) + ", Ames, Iowa, USA"
merged[merged['Address']==' , Ames, Iowa, USA'].T
# +
from geopy.geocoders import Nominatim
geocoder = Nominatim(user_agent = 'operation_goldfish')
# adding 1 second padding between calls
from geopy.extra.rate_limiter import RateLimiter
geocode = RateLimiter(geocoder.geocode, min_delay_seconds = 1, return_value_on_exception = None)
def coord_finder(address):
try:
return geocode(address, exactly_one=True, limit=None, addressdetails=False, language=False, geometry=None, extratags=False, country_codes=None, viewbox=None, bounded=False, featuretype=None, namedetails=False)
except:
return "No coordinates found"
# -
merged['Coords']=merged['Address'].apply(lambda row: coord_finder(row))
# +
import geopy
geolocator = geopy.geocoders.OpenMapQuest(api_key='<KEY>')
# adding 1 second padding between calls
from geopy.extra.rate_limiter import RateLimiter
geocode = RateLimiter(geocoder.geocode, min_delay_seconds = 1, return_value_on_exception = None)
def coord_finder2(address):
try:
return geolocator.geocode(address, exactly_one=True, limit=None, addressdetails=False, language=False, geometry=None, extratags=False, country_codes=None, viewbox=None, bounded=False, featuretype=None, namedetails=False)
except:
return "No coordinates found"
# -
merged['Coords2']=merged['Address'].apply(lambda row: coord_finder2(row))
# +
import geopy
geolocator = geopy.geocoders.GoogleV3(api_key='<KEY>', domain='maps.googleapis.com')
# adding 1 second padding between calls
from geopy.extra.rate_limiter import RateLimiter
geocode = RateLimiter(geocoder.geocode, min_delay_seconds = 1, return_value_on_exception = None)
def coord_finder3(address):
try:
return geolocator.geocode(address, exactly_one=True, language=False)
except:
return "No coordinates found"
# -
merged['Coords4']=merged['Address'].apply(lambda row: coord_finder3(row))
merged['Coords4'][1].latitude
merged['latitude']=merged['Coords4'].apply(lambda x: x.latitude)
merged['longitude']=merged['Coords4'].apply(lambda x: x.longitude)
house_coords=merged[['PID','Address','Coords4','latitude','longitude']]
# +
#house_coords.to_csv('../data/house_coordinates.csv')
# -
house_coords=pd.read_csv('../data/house_coordinates.csv')
house_coords=house_coords.drop('Unnamed: 0',axis=1).drop_duplicates()
def group_vincinity(df_house, df_place, colname="POI", mileage = 0.25):
'''
Input variables:
df_house: Dataframe of the houses. Need to contain the columns ["longitude", "latitude"].
df_place: Dataframe of places of interests. Need to contain the columns ["longitude", "latitude"].
colname: A string, indicates the column name which will be attached to df_house.
mileage: A float. Will look for all pairs of df_house and df_place within th mileage miles.
Output variables:
An updated dataframe of df_house which has a new column "POI".
The column consists integers which indicates the number of 'df_place' within 'mileage' in each 'df_house'.
'''
#for each line:
#sum
df_place=house_coords.drop('')
house_coords["gps"] = house_coords[["latitude", "longitude"]].apply(lambda x: (x["latitude"], x["longitude"]), axis = 1);
house_coords[colname] = house_coords[["gps"]].apply(lambda x: np.sum(df_place[["latitude", "longitude"]].\
apply(lambda y: geopy.distance.geodesic((y["latitude"],y["longitude"]), x).miles < mileage, axis = 1)), axis = 1);
house_coords.drop("gps", axis = 1, inplace = True);
return df_house;
# +
df = pd.read_csv("../data/Ames_Housing_Price_Data_v2.csv", index_col=0)
df=df.reset_index()
house_coords["gps"] = house_coords[["latitude", "longitude"]].apply(lambda x: (x["latitude"], x["longitude"]), axis = 1);
house_coords=house_coords.merge(df[['PID','SalePrice']],how='left')
# +
import geopy
from geopy.distance import geodesic
def group_average(index, mileage = 0.25):
coords=house_coords.loc[index,'gps']
df_place=house_coords.drop(index,axis=0).copy()
df_place['in_distance']=df_place['gps'].apply(lambda x: geopy.distance.geodesic(x, coords).miles < mileage)
return df_place.loc[df_place['in_distance']==True,'SalePrice'].mean()
# +
house_coords2=house_coords.copy()
for i in range(0,len(house_coords2)):
house_coords2.loc[i,'AvgPrice-0.25']=group_average(i,mileage=0.25)
house_coords2.loc[i,'AvgPrice-0.5']=group_average(i,mileage=0.5)
house_coords2.to_csv('../data/house_surrounding_avg_prices.csv')
# -
house_coords2
| Moritz/House_coordinate_identification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_python3)
# language: python
# name: conda_python3
# ---
# + [markdown] nbpresent={"id": "42b5e80b-ad1d-4335-a1f7-10a91127e3dc"}
# # Time Series Forecasting with Linear Learner
# _**Using Linear Regression to Forecast Monthly Demand**_
#
# ---
#
# ---
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Host](#Host)
# 1. [Forecast](#Forecast)
# 1. [Extensions](#Extensions)
#
# ---
#
# ## Background
#
# Forecasting is potentially the most broadly relevant machine learning topic there is. Whether predicting future sales in retail, housing prices in real estate, traffic in cities, or patient visits in healthcare, almost every industry could benefit from improvements in their forecasts. There are numerous statistical methodologies that have been developed to forecast time-series data, but still, the process for developing forecasts tends to be a mix of objective statistics and subjective interpretations.
#
# Properly modeling time-series data takes a great deal of care. What's the right level of aggregation to model at? Too granular and the signal gets lost in the noise, too aggregate and importent variation is missed. Also, what is the right cyclicality? Daily, weekly, monthly? Are there holiday peaks? How should we weight recent versus overall trends?
#
# Linear regression with appropriate controls for trend, seasonality, and recent behavior, remains a common method for forecasting stable time-series with reasonable volatility. This notebook will build a linear model to forecast weekly output for US gasoline products starting in 1991 to 2005. It will focus almost exclusively on the application. For a more in-depth treatment on forecasting in general, see [Forecasting: Principles & Practice](https://robjhyndman.com/uwafiles/fpp-notes.pdf).
#
# ---
#
# ## Setup
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
bucket = '<your_s3_bucket_name_here>'
prefix = 'sagemaker/linear_time_series_forecast'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
# + [markdown] nbpresent={"id": "b2548d66-6f8f-426f-9cda-7a3cd1459abd"}
# Now we'll import the Python libraries we'll need.
# + nbpresent={"id": "bb88eea9-27f3-4e47-9133-663911ea09a9"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
import os
import time
import json
import sagemaker.amazon.common as smac
import sagemaker
from sagemaker.predictor import csv_serializer, json_deserializer
# + [markdown] nbpresent={"id": "142777ae-c072-448e-b941-72bc75735d01"}
# ---
# ## Data
#
# Let's download the data. More information about this dataset can be found [here](https://rdrr.io/github/robjhyndman/fpp/man/gasoline.html).
# + nbpresent={"id": "78105bc7-ce5d-4003-84f6-4dc5700c5945"}
# !wget http://robjhyndman.com/data/gasoline.csv
# + [markdown] nbpresent={"id": "b472326f-3584-4b61-aecc-04b35486a1ab"}
# And take a look at it.
# + nbpresent={"id": "f8976dad-6897-4c7e-8c95-ae2f53070ef5"}
gas = pd.read_csv('gasoline.csv', header=None, names=['thousands_barrels'])
display(gas.head())
plt.plot(gas)
plt.show()
# + [markdown] nbpresent={"id": "1c44e72e-1b0d-4dcb-91b9-9b9f28a697b0"}
# As we can see, there's a definitive upward trend, some yearly seasonality, but sufficient volatility to make the problem non-trivial. There are several unexpected dips and years with more or less pronounced seasonality. These same characteristics are common in many topline time-series.
#
# Next we'll transform the dataset to make it look a bit more like a standard prediction model. Our target variable is `thousands_barrels`. Let's create explanatory features, like:
# - `thousands_barrels` for each of the 4 preceeding weeks.
# - Trend. The chart above suggests the trend is simply linear, but we'll create log and quadratic trends in case.
# - Indicator variables {0 or 1} that will help capture seasonality and key holiday weeks.
# + nbpresent={"id": "6af8d66e-2ef6-4e8d-bb23-d2bd3dbb0b20"}
gas['thousands_barrels_lag1'] = gas['thousands_barrels'].shift(1)
gas['thousands_barrels_lag2'] = gas['thousands_barrels'].shift(2)
gas['thousands_barrels_lag3'] = gas['thousands_barrels'].shift(3)
gas['thousands_barrels_lag4'] = gas['thousands_barrels'].shift(4)
gas['trend'] = np.arange(len(gas))
gas['log_trend'] = np.log1p(np.arange(len(gas)))
gas['sq_trend'] = np.arange(len(gas)) ** 2
weeks = pd.get_dummies(np.array(list(range(52)) * 15)[:len(gas)], prefix='week')
gas = pd.concat([gas, weeks], axis=1)
# + [markdown] nbpresent={"id": "1c77ea86-256b-4601-a5d5-2f875c0649c9"}
# Now, we'll:
# - Clear out the first four rows where we don't have lagged information.
# - Split the target off from the explanatory features.
# - Split the data into training, validation, and test groups so that we can tune our model and then evaluate its accuracy on data it hasn't seen yet. Since this is time-series data, we'll use the first 60% for training, the second 20% for validation, and the final 20% for final test evaluation.
# + nbpresent={"id": "80c0adca-5db2-4152-a9f4-42cbc1dbde84"}
gas = gas.iloc[4:, ]
split_train = int(len(gas) * 0.6)
split_test = int(len(gas) * 0.8)
train_y = gas['thousands_barrels'][:split_train]
train_X = gas.drop('thousands_barrels', axis=1).iloc[:split_train, ].as_matrix()
validation_y = gas['thousands_barrels'][split_train:split_test]
validation_X = gas.drop('thousands_barrels', axis=1).iloc[split_train:split_test, ].as_matrix()
test_y = gas['thousands_barrels'][split_test:]
test_X = gas.drop('thousands_barrels', axis=1).iloc[split_test:, ].as_matrix()
# + [markdown] nbpresent={"id": "ff9d10f9-b611-423b-80da-6dcdafd1c8b9"}
# Now, we'll convert the datasets to the recordIO-wrapped protobuf format used by the Amazon SageMaker algorithms and upload this data to S3. We'll start with training data.
# -
buf = io.BytesIO()
smac.write_numpy_to_dense_tensor(buf, np.array(train_X).astype('float32'), np.array(train_y).astype('float32'))
buf.seek(0)
key = 'linear_train.data'
boto3.resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train', key)).upload_fileobj(buf)
s3_train_data = 's3://{}/{}/train/{}'.format(bucket, prefix, key)
print('uploaded training data location: {}'.format(s3_train_data))
# Next we'll convert and upload the validation dataset.
buf = io.BytesIO()
smac.write_numpy_to_dense_tensor(buf, np.array(validation_X).astype('float32'), np.array(validation_y).astype('float32'))
buf.seek(0)
key = 'linear_validation.data'
boto3.resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation', key)).upload_fileobj(buf)
s3_validation_data = 's3://{}/{}/validation/{}'.format(bucket, prefix, key)
print('uploaded validation data location: {}'.format(s3_validation_data))
# + [markdown] nbpresent={"id": "f3b125ad-a2d5-464c-8cfa-bd203034eee4"}
# ---
# ## Train
#
# Now we can begin to specify our linear model. First, let's specify the containers for the Linear Learner algorithm. Since we want this notebook to run in all 4 of Amazon SageMaker's regions, we'll create a small lookup. More details on algorithm containers can be found in [AWS documentation](https://docs-aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html).
# -
containers = {'us-west-2': '174872318107.dkr.ecr.us-west-2.amazonaws.com/linear-learner:latest',
'us-east-1': '382416733822.dkr.ecr.us-east-1.amazonaws.com/linear-learner:latest',
'us-east-2': '404615174143.dkr.ecr.us-east-2.amazonaws.com/linear-learner:latest',
'eu-west-1': '438346466558.dkr.ecr.eu-west-1.amazonaws.com/linear-learner:latest'}
# Amazon SageMaker's Linear Learner actually fits many models in parallel, each with slightly different hyperparameters, and then returns the one with the best fit. This functionality is automatically enabled. We can influence this using parameters like:
#
# - `num_models` to increase to total number of models run. The specified parameters will always be one of those models, but the algorithm also chooses models with nearby parameter values in order to find a solution nearby that may be more optimal. In this case, we're going to use the max of 32.
# - `loss` which controls how we penalize mistakes in our model estimates. For this case, let's use absolute loss as we haven't spent much time cleaning the data, and absolute loss will adjust less to accomodate outliers.
# - `wd` or `l1` which control regularization. Regularization can prevent model overfitting by preventing our estimates from becoming too finely tuned to the training data, which can actually hurt generalizability. In this case, we'll leave these parameters as their default "auto" though.
#
# Let'd kick off our training job in SageMaker's distributed, managed training. Because training is managed (AWS handles spinning up and spinning down hardware), we don't have to wait for our job to finish to continue, but for this case, we'll use the Python SDK to track to wait and track our progress.
# +
sess = sagemaker.Session()
linear = sagemaker.estimator.Estimator(containers[boto3.Session().region_name],
role,
train_instance_count=1,
train_instance_type='ml.c4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
linear.set_hyperparameters(feature_dim=59,
mini_batch_size=100,
predictor_type='regressor',
epochs=10,
num_models=32,
loss='absolute_loss')
linear.fit({'train': s3_train_data, 'validation': s3_validation_data})
# + [markdown] nbpresent={"id": "2adcc348-9ab5-4a8a-8139-d0ecd740208a"}
# ---
# ## Host
#
# Now that we've trained the linear algorithm on our data, let's create a model and deploy that to a hosted endpoint.
# -
linear_predictor = linear.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
# ### Forecast
#
# Now that we have our hosted endpoint, we can generate statistical forecasts from it. Let's forecast on our test dataset to understand how accurate our model may be.
#
# There are many metrics to measure forecast error. Common examples include include:
# - Root Mean Square Error (RMSE)
# - Mean Absolute Percent Error (MAPE)
# - Geometric Mean of the Relative Absolute Error (GMRAE)
# - Quantile forecast errors
# - Errors that account for asymmetric loss in over or under-prediction
#
# For our example we'll keep things simple and use Median Absolute Percent Error (MdAPE), but we'll also compare it to a naive benchmark forecast (that week last year's demand * that week last year / that week two year's ago).
#
# There are also multiple ways to generate forecasts.
# - One-step-ahead forecasts: When predicting for multiple data points, one-step-ahead forecasts update the history with the correct known value. These are common, easy to produce, and can give us some intuition of whether out model is performing as expected. However, they can also present an unnecessarily optimistic evaluation of the forecast. In most real-life cases, we want to predict out well into the future, because the actions we may take based on that forecast are not immediate. In these cases, we want know what the time-periods in between will bring, so generating a forecast based on the knowledge that we do, can be misleading.
# - Multi-step-ahead (or horizon) forecasts: In this case, when forecasting out of sample, each forecast builds off of the forecasted periods that precede it. So, errors early on in the test data can compound to create large deviations for observations late in the test data. Although this is more realistic, it can be difficult to create the forecasts, particularly as model complexity increases.
#
# For our example, we'll calculate both, but focus on the multi-step forecast accuracy.
#
# Let's start by generating the naive forecast.
gas['thousands_barrels_lag52'] = gas['thousands_barrels'].shift(52)
gas['thousands_barrels_lag104'] = gas['thousands_barrels'].shift(104)
gas['thousands_barrels_naive_forecast'] = gas['thousands_barrels_lag52'] ** 2 / gas['thousands_barrels_lag104']
naive = gas[split_test:]['thousands_barrels_naive_forecast'].as_matrix()
# And investigating it's accuracy.
print('Naive MdAPE =', np.median(np.abs(test_y - naive) / test_y))
plt.plot(np.array(test_y), label='actual')
plt.plot(naive, label='naive')
plt.legend()
plt.show()
# Now we'll generate the one-step-ahead forecast. First we need a function to convert our numpy arrays into a format that can be handled by the HTTP POST request we pass to the inference container. In this case that's a simple CSV string. The results will be published back as JSON. For these common formats we can use the Amazon SageMaker Python SDK's built in `csv_serializer` and `json_deserializer` functions.
linear_predictor.content_type = 'text/csv'
linear_predictor.serializer = csv_serializer
linear_predictor.deserializer = json_deserializer
# Next, we'll invoke the endpoint to get predictions.
result = linear_predictor.predict(test_X)
one_step = np.array([r['score'] for r in result['predictions']])
# Let's compare forecast errors.
print('One-step-ahead MdAPE = ', np.median(np.abs(test_y - one_step) / test_y))
plt.plot(np.array(test_y), label='actual')
plt.plot(one_step, label='forecast')
plt.legend()
plt.show()
# As we can see our MdAPE is substantially better than the naive, and we actually swing from a forecasts that's too volatile to one that under-represents the noise in our data. However, the overall shape of the statistical forecast does appear to better represent the actual data.
#
# Next, let's generate multi-step-ahead forecast. To do this, we'll need to loop over invoking the endpoint one row at a time and make sure the lags in our model are updated appropriately.
# +
multi_step = []
lags = test_X[0, 0:4]
for row in test_X:
row[0:4] = lags
result = linear_predictor.predict(row)
prediction = result['predictions'][0]['score']
multi_step.append(prediction)
lags[1:4] = lags[0:3]
lags[0] = prediction
multi_step = np.array(multi_step)
# -
# And now calculate the accuracy of these predictions.
print('Multi-step-ahead MdAPE =', np.median(np.abs(test_y - multi_step) / test_y))
plt.plot(np.array(test_y), label='actual')
plt.plot(one_step, label='forecast')
plt.legend()
plt.show()
# As we can see our multi-step ahead error performs worse than our one-step ahead forecast, but nevertheless remains substantially stronger than the naive benchmark forecast. This 1.5 percentage point difference may not seem particularly meaningful, but at the large scale of many topline forecasts can mean millions of dollars in excess supply or lost sales.
# ---
# ## Extensions
#
# Our linear model does a good job of predicting gasoline demand, but of course, improvements could be made. The fact that statistical forecast actually underrepresents some of the volatility in the data could suggest that we have actually over-regularized the data. Or, perhaps our choice of absolute loss was incorrect. Rerunning the model with further tweaks to these hyperparameters may provide more accurate out of sample forecasts. We also did not do a large amount of feature engineering. Occasionally, the lagging time-periods have complex interrelationships with one another that should be explored. Finally, alternative forecasting algorithms could be explored. Less interpretable methods like ARIMA, and black-box methods like LSTM Recurrent Neural Networks have been shown to predict time-series very well. Balancing the simplicity of a linear model with predictive accuracy is an important subjective question where the right answer depends on the problem being solved, and its implications to the business.
# ### (Optional) Clean-up
#
# If you're ready to be done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
sagemaker.Session().delete_endpoint(linear_predictor.endpoint)
| introduction_to_applying_machine_learning/linear_time_series_forecast/linear_time_series_forecast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/custom-hyper/CoinGeckoAPI/blob/main/watchlist_report.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UzdbdQAquXzi"
#
#
# * Spikes
# * % Rankings
# * Medium website for newsfeed scraper
# * Twitter account for newsfeed scrape
# * Forum website for newsfeed scraper
# * Homepage website for HTML download/entity recognition/NLP
# * Loop for a list of exchanges
# * Proprietary score
# * Price action tool
#
# + [markdown] id="3_rFSz85Oj3u"
# #Project Setup
# + [markdown] id="tbB5g3qzEzGh"
#
# + id="5m4_0fgAPIA4" colab={"base_uri": "https://localhost:8080/"} outputId="169ec255-7caa-475f-8289-3fb862c4f2ae"
#@title Install Libraries { display-mode: "code" }
# !pip install plotly
# !pip install pytrends
# !pip install pycoingecko
# !pip install pandas
# !pip install tweepy
# !pip install requests
# !pip install pycoingecko
# + [markdown] id="sDymunfdxzyI"
# # CoinGecko API Statistics Report
# + id="aG_Ot5aN_OMT" colab={"base_uri": "https://localhost:8080/"} outputId="eb8c9001-1777-47de-c0f9-bea83ef47762"
#@title Available Currencies { display-mode: "code" }
import requests
import datetime
import time
import json
from google.colab import data_table
import plotly.graph_objects as go
import pandas as pd
import matplotlib.pyplot as plt
from pandas.io.json import json_normalize
from pycoingecko import CoinGeckoAPI
#General Information table
def currency_stats(url):
"""
Generates a list of the currencies on CoinGecko
Args: url of the CoinGecko website
"""
frames = []
response = requests.get(url)
data = response.json()
df = pd.DataFrame (data)
print('Stats: ')
print(df.info(verbose=True))
print(df.isnull().sum())
print(data)
print(df.describe())
def main():
url = "https://api.coingecko.com/api/v3/simple/supported_vs_currencies"
currency_stats(url)
if __name__ == "__main__":
main()
# + id="CpfyE_IK70Aj" colab={"base_uri": "https://localhost:8080/"} outputId="72c82d39-44ae-4b63-85cb-038a1f3d605e"
#@title Available Coins { display-mode: "code" }
import requests
import datetime
import time
import json
from google.colab import data_table
import plotly.graph_objects as go
import pandas as pd
import matplotlib.pyplot as plt
from pandas.io.json import json_normalize
#General Information table
def coins_stats(url):
"""
Generates a list of the coins on CoinGecko
Args: url of the CoinGecko website
"""
frames = []
response = requests.get(url)
data = response.json()
token_list = pd.DataFrame (data)
print('Stats: ')
print(token_list.info(verbose=True))
print(token_list.isnull().sum())
print(token_list.describe())
return token_list
def main():
url = "https://api.coingecko.com/api/v3/coins/list"
print(coins_stats(url))
if __name__ == "__main__":
main()
# + [markdown] id="YHPqTPJ-MeAM"
#
# + [markdown] id="XVcNrRwDAIvd"
# # CoinGecko API Extraction
# + id="d5ttw5sPWAq_" colab={"base_uri": "https://localhost:8080/"} outputId="216f1603-c18a-4bef-fb2a-4aea353d58e1"
from google.colab import drive
drive.mount('/content/drive')
# + id="omzIJ-K6lijc"
#@title Download Coins Data Reports { display-mode: "code" }
import requests
import datetime
import time
import json
from google.colab import data_table
import plotly.graph_objects as go
import pandas as pd
import matplotlib.pyplot as plt
from pandas.io.json import json_normalize
import logging
import sys
from tqdm.notebook import tqdm
currency = 'usd'
pages = range(1, 135) #can find the number of pages by dividing the total number of coins by 100
frames=[]
for page_num in tqdm(pages):
url = "https://api.coingecko.com/api/v3/coins/markets?vs_currency={}&order=market_cap_desc&per_page=100&page={}&sparkline=false&price_change_percentage=1h%2C%2024h%2C%207d%2C14d%2C30d%2C200d%2C1y".format(currency, page_num)
#Obtain JSON formatted data
response = requests.get(url)
data = response.json()
#Convert data to list of dataframes
full_table = pd.json_normalize(data)
results = [full_table]
frames.append(full_table)
#Slow down requests
#Status report
logging.info('Page '+ str(page_num) + ' – Rows ' + str(len(full_table)) + ' – source: '+ str(url))
single = pd.concat(frames) #Careful, this line must be outside of the for loop
#Create ID
id = single['id']
symbol = single['symbol'].str.upper()
name = single['name']
single['Airtable_ID'] = symbol + '.' + name.str.replace(" ", "")
"""
x = single['fully_diluted_valuation']
y = single['market_cap']
try:
return single['ratiofully_diluted_valuation'] = single['fully_diluted_valuation']/single['market_cap']
except ZeroDivisionError:
return 0
"""
single['CoinGecko_website'] = 'https://www.coingecko.com/en/coins/'+ id
path = "/content/drive/MyDrive/workspace/pipelines/datasets/coingecko/coingecko-coin-list/output"
single.to_csv(path+'/CoinGecko_coins_markets_{}.csv'.format(currency))
single.info()
# + id="zDbHILx-MnfE"
#General Information table
i = 'helium'
cur = 'usd'
response = requests.get("https://api.coingecko.com/api/v3/coins/{}".format(i))
output = response.text
json_parsed = json.loads(output)
response_2 = requests.get("https://api.coingecko.com/api/v3/simple/price?ids={}&vs_currencies={}".format(i,cur))
output_2 = response_2.text
json_parsed_2 = json.loads(output_2)
print('')
print('MARKET DATA')
print('')
#Indicators
# Basics
id = json_parsed['id']
symbol = json_parsed['symbol'].upper()
name = json_parsed['name']
ID = symbol + '.' + name.replace(" ", "")
asset_platform_id = json_parsed['asset_platform_id']
platforms = json_parsed['platforms']
localization = json_parsed['localization']
public_notice = json_parsed['public_notice']
description = json_parsed['description']['en']
short_desc = json_parsed['ico_data']['short_desc']
# price Volume
price = json_parsed_2[i][cur]
# Categories
categories = json_parsed['links']['categories']
categories_1 = json_parsed['links']['categories'][0]
categories_2 = json_parsed['links']['categories'][1]
categories_3 = json_parsed['links']['categories'][2]
# forum
official_forum_url = json_parsed['links']['official_forum_url'].str.contains("hello")]
official_forum_url_1 = json_parsed['links']['official_forum_url'][0]
official_forum_url_2 = json_parsed['links']['official_forum_url'][1]
official_forum_url_3 = json_parsed['links']['official_forum_url'][2] Backed up the
links = json_parsed['links'].str.contains("medium")]
link_CoinGecko = 'https://www.coingecko.com/en/coins/{}'.format(i)
# Medium
links_announcement_url_medium = json_parsed['links']['announcement_url'][0]
# forum
links_announcement_url_forum = json_parsed['links']['announcement_url'][1]
# Homepage
links_homepage_1 = json_parsed['links']['homepage'][0]b Tectonic Doctrine
links_homepage_2 = json_parsed['links']['homepage'][1]
links_homepage_3 = json_parsed['links']['homepage'][2]
# Twitter
links_twitter_screen_name = json_parsed['links']['twitter_screen_name']
account_twitter = '@{}'.format(json_parsed['links']['twitter_screen_name'])
links_twitter = 'https://twitter.com/{}'.format(json_parsed['links']['twitter_screen_name'])
market_cap = json_parsed['market_data']['market_cap']['usd']
fdv_to_tvl_rati = json_parsed['market_data']['fdv_to_tvl_ratio']
mcap_to_tvl_ratio = json_parsed['market_data']['mcap_to_tvl_ratio']
fully_diluted_valuation = json_parsed['fully_diluted_valuation']
dilution_ratio = fully_diluted_valuation / market_cap
total_volume = json_parsed['market_data']['total_volume']['usd']
total_supply_rotation = total_volume / circulating_supply
# Tokenomics
total_value_locked = json_parsed['market_data']['total_value_locked']
total_supply = json_parsed['total_supply']
max_supply = json_parsed['max_supply']
circulating_supply = json_parsed['circulating_supply']
supply_expansion = max_supply / circulating_supply
# Adjusted capitalization
total_supply_mcap = price * total_supply
max_supply_mcap = price * max_supply
circulating_supply_mcap = price * circulating_supply
# volume
total_volume = json_parsed['market_data']['total_volume']['usd']
total_volume_rotation = total_volume / circulating_supply
# multiple price change percentages
price_change_percentage_7d = json_parsed['market_data']['price_change_percentage_7d']
price_change_percentage_14d = json_parsed['market_data']['price_change_percentage_14d']
price_change_percentage_30d = json_parsed['market_data']['price_change_percentage_30d']
price_change_percentage_60d = json_parsed['market_data']['price_change_percentage_60d']
price_change_percentage_200d = json_parsed['market_data']['price_change_percentage_200d']
price_change_percentage_1y = json_parsed['market_data']['price_change_percentage_1y']
# Community followers
community_data = json_parsed['community_data']
community_data_reddit_subscribers = json_parsed['community_data']['reddit_subscribers']
community_data_telegram_channel_user_count = json_parsed['community_data']['telegram_channel_user_count']
community_data_twitter_followers = json_parsed['community_data']['twitter_followers']
#Ratio market CAP to social media followers
market_cap_twitter_followers_ratio = market_cap / community_data_twitter_followers
market_cap_reddit_subscribers_ratio = market_cap / community_data_reddit_subscribers
market_cap_telegram_channel_user_count_ratio = market_cap / community_data_telegram_channel_user_count
# CoinGecko Score
CoinGecko_market_cap_rank = json_parsed['market_cap_rank']
CoinGecko_coingecko_rank = json_parsed['coingecko_rank']
CoinGecko_coingecko_score = json_parsed['coingecko_score']
CoinGecko_developer_score = json_parsed['developer_score']
CoinGecko_community_score = json_parsed['community_score']
CoinGecko_liquidity_score = json_parsed['liquidity_score']
# Ratio market CAP versus core
market_cap_CoinGecko_coingecko_score_ratio = market_cap / CoinGecko_coingecko_score
market_cap_CoinGecko_developer_score_ratio = market_cap / CoinGecko_developer_score
market_cap_CoinGecko_community_score_ratio = market_cap / CoinGecko_community_score
market_cap_CoinGecko_liquidity_score_ratio = market_cap / CoinGecko_liquidity_score
developer_data = json_parsed['developer_data']
sentiment_votes_down_percentage = json_parsed['sentiment_votes_down_percentage']
sentiment_votes_percentageup_ = json_parsed['sentiment_votes_percentageup_']
df_community_data = pd.DataFrame(data=community_data, index=[0])
# + id="Q8_m0sRZRHpc"
#General Information table
The
i = 'helium'
cur = 'usd'
period = 'max'
response = requests.get("https://api.coingecko.com/api/v3/coins/{}/market_chart?vs_currency={}&days={}".format(i,cur,period))
output = response.text
json_parsed = json.loads(output)
json_parsed .json_normalize
# + id="TFrCK_v9yq1w"
#Indicators
df = {}
df['asset_platform_id'] = asset_platform_id
df['id'] = asset_platform_id
df['link_CoinGecko'] = 'https://www.coingecko.com/en/coins/{}'.format(i)
# + id="6sFo92TQOwJP"
'https://api.coingecko.com/api/v3/coins/{}/market_chart/range?vs_currency={}&from={}&to={}'.format('solana','usd',,)
# + id="QROQCR-UNs83"
df_community_data
# + id="g1ev1MPwOAXY"
df_developer_data
# + id="6J5fuRFuQImF"
from pycoingecko import CoinGeckoAPI
cg = CoinGeckoAPI()
a = cg.get_coin_ohlc_by_id(id=i, vs_currency='usd', days = 'max')
df = pd.DataFrame(data=a, columns = ['Date','Open','High', 'Low', 'Close'])
fig2 = go.Figure(data=[go.Candlestick(x=df['Date'],open=df['Open'], high=df['High'],low=df['Low'], close=df['Close'])])
fig2.show()
# + [markdown] id="T3crwZswAEnZ"
# # Crypto fees website
| watchlist_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.3.7
# language: julia
# name: julia-0.3
# ---
using PyCall
unshift!(PyVector(pyimport("sys")["path"]), "");
@pyimport GLS_Apr_weekend
xi_list = GLS_Apr_weekend.xi_list
P = GLS_Apr_weekend.P
L = GLS_Apr_weekend.L # dimension of xi
size(P, 1), size(P, 2)
# +
using JuMP
mGLSJulia = Model()
@defVar(mGLSJulia, lam[1:size(P,1)] >= 0)
@defVar(mGLSJulia, p[1:size(P,1), 1:size(P,2)] >= 0)
for i = 1:size(P,1)
for j = 1:size(P,2)
if P[i,j] == 0
@addConstraint(mGLSJulia, p[i,j] == 0)
end
end
end
for i = 1:size(P,1)
@addNLConstraint(mGLSJulia, sum{p[i,j], j = 1:size(P,2)} == 1)
end
for l = 1:L
@addNLConstraint(mGLSJulia, sum{p[i,l] * lam[i], i = 1:size(P,1)} == xi_list[l])
end
@setNLObjective(mGLSJulia, Min, sum{p[1,j], j = 1:size(P,2)}) # play no actual role, but could not use zero objective
solve(mGLSJulia)
# -
getValue(lam)
getObjectiveValue(mGLSJulia)
GLS_Apr_weekend.saveDemandVec(getValue(lam))
| 03_OD_matrix_estimation_journal18/OD_matrix_estimation_GLS_julia_Apr_weekend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Natural language Preprocessing & Feature Engineering
#
# * LABELS:
# - Sentiment Analysis, Classification
# - regex, TFIDF, NLTK
# ## Imports & read in data
import pandas as pd
import numpy as np
import seaborn as sns
from nltk.tokenize import word_tokenize, sent_tokenize, regexp_tokenize
from nltk import pos_tag
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from sklearn.preprocessing import LabelEncoder
from collections import defaultdict
from nltk.corpus import wordnet as wn
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn import model_selection, naive_bayes, svm
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt
np.random.seed(500)
#Corpus = pd.read_csv("corpus.csv",encoding='latin-1')
Corpus_all = pd.read_csv("corpus.csv",encoding='latin-1')
Corpus=Corpus_all[:50]
# ### Data pre-processing
# * This involves transforming raw data into an understandable format for NLP models.
#
#
# * **Tokenization**:
# This is a process of breaking a stream of text up into words, phrases, symbols, or other meaningful elements called tokens. The list of tokens becomes input for further processing.
#
# **NLTK** Library has word_tokenize and sent_tokenize to easily break a stream of text into a list of words or sentences, respectively. Its TweetTokenizer separates out hashtags, mentions & exclamation marks <.-)!! #NLPisfun
#
#
# * **Word Stemming/Lemmatization**:
# The aim of both processes is the same, reducing the inflectional forms of each word into a common base or root. Lemmatization is closely related to stemming. The difference is that a stemmer operates on a single word without knowledge of the context, and therefore cannot discriminate between words which have different meanings depending on part of speech. However, stemmers are typically easier to implement and run faster, and the reduced accuracy may not matter for some applications.
#
# * **Bag of Words**:
# Basic method for finding topics in text. Based on the theory - more frequent words will be the important words
#
#
# * **Stop-Words**:
# Words like 'and', 'the' etc. which don't carry much meaning, are removed from text.
#
#
# * **Word Vector**:
# Multi-dimentional representation of a word,with sparse features, trained from a reference or corpora. It contains information about the relation of the word with other words in text.
#
#
# * A complete list of steps may involve (add or remove based on task):
#
# 1. Remove Blank rows in Data, if any
# 2. Change all the text to lower case
# 3. Word Tokenization
# 4. Remove Stop words
# 5. Remove Non-alpha text
# 6. Word Lemmatization
# +
# Step - a : Remove blank rows if any.
print(Corpus.isnull().any()) # gives False.
# Step - b : Change all the text to lower case. This is required as python interprets 'dog' and 'DOG' differently
Corpus['text_lower']=[doc.lower() for doc in Corpus.text]
# Step - c : Tokenization : In this each entry in the corpus will be broken into set of words
Corpus['text_final']=[word_tokenize(doc) for doc in Corpus.text_lower]
# Step - d : Remove Stop words, Non-Numeric and perfom Word Stemming/Lemmenting.
# WordNetLemmatizer requires Pos tags to understand if the word is noun or verb or adjective etc.
# By default it is set to Noun.
lemmatizer = WordNetLemmatizer()
def nltk_tag_to_wordnet_tag(nltk_tag):
if nltk_tag.startswith('J'):
return wn.ADJ
elif nltk_tag.startswith('V'):
return wn.VERB
elif nltk_tag.startswith('N'):
return wn.NOUN
elif nltk_tag.startswith('R'):
return wn.ADV
else:
return wn.NOUN
for i in range(len(Corpus.text_final)):
Corpus.text_final[i]=[t for t in Corpus.text_final[i] if t.isalpha() and t not in stopwords.words('english')]
Corpus.text_final[i]=[lemmatizer.lemmatize(word, nltk_tag_to_wordnet_tag(tag)) for word,tag in pos_tag(Corpus.text_final[i])]
Corpus.text_final[i]=str(Corpus.text_final[i])
# -
#Number of sentences per text
line_num = [len(sent_tokenize(t, r"\w+")) for t in Corpus['text_lower']]
sns.distplot(line_num)
# ### Prepare Test/ Train/ Datasets
# +
# Import Counter
from collections import Counter
# Tokenize the article: tokens
tokens = word_tokenize(holy_grail)
# Convert the tokens into lowercase: lower_tokens
lower_tokens = [t.lower() for t in tokens]
# Create a Counter with the lowercase tokens: bow_simple
bow_simple =[bag for t in Corpus['text_final']]Counter(lower_tokens)
# Print the 10 most common tokens
print(bow_simple.most_common(10))
# -
X=Corpus['text_final']
y=Corpus['label']
X_train, X_test, y_train, y_test = model_selection.train_test_split(X,y,test_size=0.3, stratify=y)
# ### Encoding
#
# * Label encode the target variable — This is done to transform Categorical data of string type in the data set into numerical values which the model can understand.
#
#
# * **Gensim** package: a great package for processing texts, working with word vector models (such as Word2Vec, FastText etc) and for building topic models.
# * Significant advantage with gensim is: it lets you handle large text files without having to load the entire file in memory.
# * Gensim models can be saved updated and reused.
#
#
# * **TFIDF**: Term Frequency – Inverse Document Frequency(TF-IDF): document specific common words are highly weighted. For example, many astonomy documents may have the word 'Sky'. Sky will be downweighted for this corpus.
#
#
# * **NER (Names Entity Recognition)**: People, places, dates, organizations can be recognized & tagged.
Encoder = LabelEncoder()
y_train = Encoder.fit_transform(y_train)
y_test = Encoder.transform(y_test)
Tfidf_vect = TfidfVectorizer(max_features=5000)
Tfidf_vect.fit(X)
Train_X_Tfidf = Tfidf_vect.transform(X_train)
Test_X_Tfidf = Tfidf_vect.transform(X_test)
print(Tfidf_vect.vocabulary_)
print(Train_X_Tfidf)
# ##### STEP -7: Use the ML Algorithms
# Classifier - Algorithm - SVM
# fit the training dataset on the classifier
SVM = svm.SVC(C=1.0, kernel='linear', degree=3, gamma='auto')
SVM.fit(Train_X_Tfidf,y_train)
# predict the labels on validation dataset
predictions_SVM = SVM.predict(Test_X_Tfidf)
# Use accuracy_score function to get the accuracy
print("SVM Accuracy Score -> ",accuracy_score(predictions_SVM, y_test)*100)
# ***STEP -8: Cross-validation***
from sklearn.model_selection import cross_val_score
cv_scores =cross_val_score(SVM, Train_X_Tfidf, y_train, cv=5)
cv_scores
# ## Trying out other classification models
#
# - LinearSVC (faster SVM than SVC, but no kernel support)
# - Trying different kernels using SVC
# - SGDClassifier (Can handle large amounts of data, for 'online' model, out-of-core support.)
# - Logistic Regression
# - Desion Tree
# - Random Forest
# - Naive Bayes
# - KNN
# ### LinearSVC
# +
from sklearn.svm import LinearSVC
linear_svc = LinearSVC(C=1, loss="hinge") #use dual=False when n_samples>n_features
linear_svc.fit(Train_X_Tfidf,y_train)
predictions_lin = linear_svc.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(predictions_lin, y_test)*100)
cv_scores =cross_val_score(linear_svc, Train_X_Tfidf, y_train, cv=5)
cv_scores
# -
# ### switching out kernels
# +
poly = svm.SVC(kernel='poly')
poly.fit(Train_X_Tfidf,y_train)
predictions_poly = poly.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(predictions_poly, y_test)*100)
cv_scores =cross_val_score(poly, Train_X_Tfidf, y_train, cv=5)
cv_scores
# +
rbf = svm.SVC() #default is 'rbf'
rbf.fit(Train_X_Tfidf,y_train)
predictions_rbf = rbf.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(predictions_rbf, y_test)*100)
cv_scores =cross_val_score(rbf, Train_X_Tfidf, y_train, cv=5)
cv_scores
# +
sig = svm.SVC(kernel='sigmoid')
sig.fit(Train_X_Tfidf,y_train)
predictions_sig = sig.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(predictions_sig, y_test)*100)
cv_scores =cross_val_score(sig, Train_X_Tfidf, y_train, cv=5)
cv_scores
# -
# ### SGDClassifier : Binary Classifier
# +
"""
SGDClassifier(loss="hinge",alpha=1/(m*C)).
This applies regular Stochastic Gradient Descent to train a linear SVM classifier. It does not converge as fast as the LinearSVC class, but it
can be useful to handle huge datasets that do not fit in memory (out-of-core training), or to handle online classification tasks.
"""
from sklearn.linear_model import SGDClassifier
sgd_clf = SGDClassifier(max_iter=1000, tol=1e-3, random_state=42)
sgd_clf.fit(Train_X_Tfidf,y_train)
y_pred_sgd=sgd_clf.predict(Test_X_Tfidf)
print("SGD Accuracy Score -> ",accuracy_score(y_pred_sgd, y_test)*100)
from sklearn.model_selection import cross_val_score
cv_scores =cross_val_score(sgd_clf, Train_X_Tfidf, y_train, cv=5)
cv_scores
# -
# ### Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression()
log_reg.fit(Train_X_Tfidf,y_train)
y_pred_lr=log_reg.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(y_pred_lr, y_test)*100)
cv_scores =cross_val_score(log_reg, Train_X_Tfidf, y_train, cv=5)
cv_scores
# +
print('model = {} @ features + {}'.format(log_reg.coef_,log_reg.intercept_))
print("\nFirst Instance: ",X_test.iloc[3])
print("Actual label: ", y_test[3])
print('\nPrediction for first instance: ', log_reg.predict(Test_X_Tfidf[3]))
print('Prediction porbability for first instance: ', log_reg.predict_proba(Test_X_Tfidf[3]))
print("\nAnother Instance: ",X_test.iloc[4])
print("Actual label: ", y_test[4])
print('\nPrediction for first instance: ', log_reg.predict(Test_X_Tfidf[4]))
print('Prediction porbability for first instance: ', log_reg.predict_proba(Test_X_Tfidf[4]))
# -
# Sort the coefficients of features in ascending and desceding order.
# Extract the indexes of these features into inds_ascending and inds_descending
inds_ascending = np.argsort(log_reg.coef_.flatten())
inds_descending = inds_ascending[::-1]
# find the words corresponding to lowest coefficients
for i in range(10):
for key, value in Tfidf_vect.vocabulary_.items():
if value==inds_ascending[i]:
print(key)
# find the words corresponding to highest coefficients
for i in range(10):
for key, value in Tfidf_vect.vocabulary_.items():
if value==inds_descending[i]:
print(key)
# ### Logistic regression with L1 penalty
# +
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression(solver='liblinear',penalty='l1')
log_reg.fit(Train_X_Tfidf,y_train)
y_pred_lr=log_reg.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(y_pred_lr, y_test)*100)
cv_scores =cross_val_score(log_reg, Train_X_Tfidf, y_train, cv=5)
cv_scores
# -
# Sort the coefficients of features in ascending and desceding order.
# Extract the indexes of these features into inds_ascending and inds_descending
inds_ascending = np.argsort(log_reg.coef_.flatten())
inds_descending = inds_ascending[::-1]
# find the words corresponding to lowest coefficients
for i in range(10):
for key, value in Tfidf_vect.vocabulary_.items():
if value==inds_ascending[i]:
print(key)
# find the words corresponding to highest coefficients
for i in range(10):
for key, value in Tfidf_vect.vocabulary_.items():
if value==inds_descending[i]:
print(key)
# ### Decision tree, Random Forest
# +
from sklearn.tree import DecisionTreeClassifier
tree_clf = DecisionTreeClassifier(max_features='auto')
tree_clf.fit(Train_X_Tfidf,y_train)
y_pred=tree_clf.predict(Test_X_Tfidf)
print("Accuracy Score -> ",accuracy_score(y_pred, y_test)*100)
cv_scores =cross_val_score(tree_clf, Train_X_Tfidf, y_train, cv=5)
cv_scores, cv_scores.mean()
# -
tree_clf.get_depth(),tree_clf.get_n_leaves(), tree_clf.n_features_
inds_ascending=np.argsort(tree_clf.feature_importances_.flatten())
inds_descending = inds_ascending[::-1]
# find the words corresponding to lowest coefficients
for i in range(10):
for key, value in Tfidf_vect.vocabulary_.items():
if value==inds_ascending[i]:
print(key)
# find the words corresponding to highest coefficients
for i in range(20):
for key, value in Tfidf_vect.vocabulary_.items():
if value==inds_descending[i]:
print(key)
from sklearn.ensemble import RandomForestClassifier
rf=RandomForestClassifier()
rf.fit(Train_X_Tfidf,y_train)
y_pred=rf.predict(Test_X_Tfidf)
print("Accuracy Score -> ",accuracy_score(y_pred, y_test)*100)
cv_scores =cross_val_score(rf, Train_X_Tfidf, y_train, cv=5)
cv_scores
# ### Naive Bays
# +
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
gnb.fit(Train_X_Tfidf.toarray(),y_train)
y_pred=gnb.predict(Test_X_Tfidf.toarray())
print("Accuracy Score -> ",accuracy_score(y_pred, y_test)*100)
cv_scores =cross_val_score(gnb, Train_X_Tfidf.toarray(), y_train, cv=5)
cv_scores
# -
# ### KNN
# ### String kernels?
# ### Parameter tuning for SVC
# +
from sklearn.model_selection import GridSearchCV
param_grid = dict(kernel = [‘linear’, ‘rbf’, ‘poly’], gamma = [0.1, 1, 10, 100], c = [0.1, 1, 10, 100, 1000],
degree = [0, 1, 2, 3, 4, 5, 6])
non.fit(Train_X_Tfidf,y_train)
predictions_non = non.predict(Test_X_Tfidf)
print(" Accuracy Score -> ",accuracy_score(predictions_non, y_test)*100)
cv_scores =cross_val_score(non, Train_X_Tfidf, y_train, cv=5)
cv_scores
svm= svm.SVC()
svm_cv = GridSearchCV(svm, param_grid, cv=5)
svm_cv.fit(X, y)
svm_cv.best_params_
svm_cv.best_score
_
| NLP Experimentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Leaflet cluster map of talk locations
#
# Run this from the _talks/ directory, which contains .md files of all your talks. This scrapes the location YAML field from each .md file, geolocates it with geopy/Nominatim, and uses the getorg library to output data, HTML, and Javascript for a standalone cluster map.
# !pip install getorg python-frontmatter
import glob
import getorg
from geopy import Nominatim
import frontmatter
g = glob.glob("_talks/*.md")
geocoder = Nominatim()
location_dict = {}
location = ""
permalink = ""
title = ""
a = frontmatter.load(g[0])
a.to_dict()
suff_dict = {", CA": " California", ", WA": " Washington", ", TX": " Texas", ", MT": " Montana "}
for file in g:
data = frontmatter.load(file)
data_dict = data.to_dict()
location = data_dict['location']
if location[-4:] in suff_dict:
location = location[:-3] + suff_dict[location[-4:]]
descript_name = location + " | " + data_dict['venue'] + " | " + data_dict['title']
location_dict[descript_name] = geocoder.geocode(location)
print(descript_name, "\n", location, "\n", location_dict[descript_name], "\n\n")
m = getorg.orgmap.create_map_obj()
getorg.orgmap.output_html_cluster_map(location_dict, folder_name="talk_map", hashed_usernames=False)
location_dict
location
| talkmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Identifying Clusters
#
# Before we get too far along, let's take a look at some different sets of data to practice identifying clusters.
#
# Start by running the cell below.
# +
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
import helper_functions as h
import test_file as t
from IPython import display
# %matplotlib inline
# Make the images larger
plt.rcParams['figure.figsize'] = (16, 9)
# -
# `1.` Run the cell below to generate a set of data. Then enter an integer next to **`question_1_clusters`** that identifies the number of clusters you think appear in the plot.
#
# If you think that there are 2 clusters in the plot, you should enter:
#
# ```
# question_1_clusters = 2
# ```
h.plot_q1_data()
# +
question_1_clusters = 4 # Enter the number of clusters you see here as an integer
#Then this will test your number against what we used to generate the data
t.test_question_1(question_1_clusters)
# -
# `2.` Run the cell below to generate a set of data. Then, similar to the first question, enter the number of clusters you think appear in the plot.
h.plot_q2_data()
# +
question_2_clusters = 2 # Enter the number of clusters you see here as an integer
#Then this will test your number against what we used to generate the data
t.test_question_2(question_2_clusters)
# -
# `3.` Run the cell below to generate a set of data. Then, similar to the previous questions, enter the number of clusters you think appear in the plot.
h.plot_q3_data()
# +
question_3_clusters = 6 # Enter the number of clusters you see here as an integer
#Then this will test your number against what we used to generate the data
t.test_question_3(question_3_clusters)
# -
# `4.` Now one final time, run the cell below, and identify the number of clusters you think are in the plot created.
h.plot_q4_data()
# +
question_4_clusters = 7 # Enter the number of clusters you see here as an integer
#Then this will test your number against what we used to generate the data
display.HTML(t.test_question_4(question_4_clusters))
# -
# **You can find a solution to this by clicking the orange jupyter image at the top of this notebook.**
| Unsupervised_Learning/1. Clustering/Identifying_Clusters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import bar_chart_race as bcr
import pandas as pd
import numpy as np
import yfinance as yf
import warnings
warnings.filterwarnings('ignore')
import os
import pickle
COLUMNS=['zip', 'sector', 'fullTimeEmployees', 'longBusinessSummary', 'city', 'phone', 'state', 'country', 'companyOfficers',
'website', 'maxAge', 'address1', 'fax', 'industry', 'previousClose', 'regularMarketOpen', 'twoHundredDayAverage',
'trailingAnnualDividendYield', 'payoutRatio', 'volume24Hr', 'regularMarketDayHigh', 'navPrice', 'averageDailyVolume10Day',
'totalAssets', 'regularMarketPreviousClose', 'fiftyDayAverage', 'trailingAnnualDividendRate', 'open', 'averageVolume10days',
'expireDate', 'yield', 'algorithm', 'dividendRate', 'exDividendDate', 'beta', 'circulatingSupply', 'startDate',
'regularMarketDayLow', 'priceHint', 'currency', 'trailingPE', 'regularMarketVolume', 'lastMarket', 'maxSupply', 'openInterest',
'marketCap', 'volumeAllCurrencies', 'strikePrice', 'averageVolume', 'priceToSalesTrailing12Months', 'dayLow', 'ask', 'ytdReturn',
'askSize', 'volume', 'fiftyTwoWeekHigh', 'forwardPE', 'fromCurrency', 'fiveYearAvgDividendYield', 'fiftyTwoWeekLow', 'bid',
'tradeable', 'dividendYield', 'bidSize', 'dayHigh', 'exchange', 'shortName', 'longName', 'exchangeTimezoneName',
'exchangeTimezoneShortName', 'isEsgPopulated', 'gmtOffSetMilliseconds', 'underlyingSymbol', 'quoteType', 'symbol',
'underlyingExchangeSymbol', 'headSymbol', 'messageBoardId', 'uuid', 'market', 'annualHoldingsTurnover', 'enterpriseToRevenue',
'beta3Year', 'profitMargins', 'enterpriseToEbitda', '52WeekChange', 'morningStarRiskRating', 'forwardEps', 'revenueQuarterlyGrowth',
'sharesOutstanding', 'fundInceptionDate', 'annualReportExpenseRatio', 'bookValue', 'sharesShort', 'sharesPercentSharesOut',
'fundFamily', 'lastFiscalYearEnd', 'heldPercentInstitutions', 'netIncomeToCommon', 'trailingEps', 'lastDividendValue',
'SandP52WeekChange', 'priceToBook', 'heldPercentInsiders', 'nextFiscalYearEnd', 'mostRecentQuarter', 'shortRatio',
'sharesShortPreviousMonthDate', 'floatShares', 'enterpriseValue', 'threeYearAverageReturn', 'lastSplitDate', 'lastSplitFactor',
'legalType', 'morningStarOverallRating', 'earningsQuarterlyGrowth', 'dateShortInterest', 'pegRatio', 'lastCapGain',
'shortPercentOfFloat', 'sharesShortPriorMonth', 'category', 'fiveYearAverageReturn', 'regularMarketPrice', 'logo_url']
# +
def get_all_stock_info(name='all_stock_info'):
all_df = pd.DataFrame(columns=COLUMNS)
filename = 'fixtures/{}.parquet.gzip'.format(name)
if not os.path.exists(filename):
with open('fixtures/tickers.pickle', 'rb') as f:
tickers = pickle.load(f)
for ticker in sorted(set(tickers)):
yahoo_ticker = ticker.replace('.', '-')
info_file = 'fixtures/stocks/{}.pickle'.format(yahoo_ticker)
if not os.path.exists(info_file):
print('Processing {}'.format(yahoo_ticker))
t = yf.Ticker(yahoo_ticker)
try:
info = dict(t.info)
except:
info = None
with open(info_file, 'wb') as f:
pickle.dump(info, f)
else:
with open(info_file, 'rb') as f:
info = pickle.load(f)
try:
if info:
all_df.loc[yahoo_ticker] = info
else:
print('Skipped ticker: {}'.format(yahoo_ticker))
except:
continue
all_df.fillna(value=np.nan, inplace=True)
all_df.to_parquet(filename, compression='gzip')
print('All stock data processed.')
else:
all_df = pd.read_parquet(filename)
print('All stock data loaded.')
return all_df
def get_all_stocks(start='', end=''):
dfs = {}
with open('fixtures/tickers.pickle', 'rb') as f:
tickers = pickle.load(f)
for ticker in sorted(set(tickers)):
yahoo_ticker = ticker.replace('.', '-')
stock_file = 'fixtures/stocks/{}.parquet.gzip'.format(yahoo_ticker)
if not os.path.exists(stock_file):
print('Processing {}'.format(yahoo_ticker))
t = yf.Ticker(yahoo_ticker)
df = t.history(period='max')
df.to_parquet(stock_file, compression='gzip')
else:
df = pd.read_parquet(stock_file)
if start == '' and end == '':
dfs[yahoo_ticker] = df
elif start == '':
dfs[yahoo_ticker] = df[:end]
elif end == '':
dfs[yahoo_ticker] = df[start:]
else:
dfs[yahoo_ticker] = df[start:end]
return dfs
# -
df = get_all_stock_info()
df[(df['country'] == 'China') & (df['city'] == 'Beijing')]['marketCap'].sum() / 1000000000
df[(df['country'] == 'China') & (df['city'] == 'Shanghai')]['marketCap'].sum() / 1000000000
df[(df['country'] == 'China') & (df['city'] == 'Shenzhen')]['marketCap'].sum() / 1000000000
df[(df['country'] == 'China') & (df['city'] == 'Guangzhou')]['marketCap'].sum() / 1000000000
df[(df['country'] == 'China')]['marketCap'].sum() / 1000000000
df1 = df.groupby('city').sum()
s1 = df1['marketCap'] / 1000000000
s2 = df1['fullTimeEmployees']
df2 = pd.DataFrame({'marketCap': s1, 'Employees': s2})
df2.sort_values(by=['marketCap'], ascending=False)[:30]
df['E']
| temp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison of Uncertainty Estimation on Toy Example
# +
import numpy as np
import numpy.matlib
import seaborn as sns
import matplotlib.pyplot as plt
import tensorflow as tf
from spinup.algos.uncertainty_estimate.core import MLP, BeroulliDropoutMLP, BootstrappedEnsemble, get_vars, ReplayBuffer
# -
# # Generate Training Data
# +
# Target from "Deep Exploration via Bootstrapped DQN"
# y = x + sin(alpha*(x+w)) + sin(beta*(x+w)) + w
# w ~ N(mean=0, var=0.03**2)
# Training set: x in (0, 0.6) or (0.8, 1), alpha=4, beta=13
def generate_label(x, noisy=True):
num = len(x)
alpha, beta = 4, 13
if noisy:
sigma = 0.03
else:
sigma = 0
omega = np.random.normal(0, sigma, num)
y = x + np.sin(alpha*(x+omega)) + np.sin(beta*(x+omega)) + omega
return y
def plot_training_data_and_underlying_function(train_size=20, train_s1=0, train_e1=0.6, train_s2=0.8, train_e2=1.4):
x_f = np.arange(-1, 2, 0.005)
# True function
y_f = generate_label(x_f, noisy=False)
# Noisy data
y_noisy = generate_label(x_f, noisy=True)
# Training data
x_train = np.concatenate((np.random.uniform(train_s1, train_e1, int(train_size/2)), np.random.uniform(train_s2, train_e2, int(train_size/2))))
y_train = generate_label(x_train)
plt.figure()
plt.plot(x_f, y_f, color='k')
plt.plot(x_f, y_noisy, '.', color='r', alpha=0.3)
plt.plot(x_train, y_train, '.', color='b')
plt.legend(['underlying function', 'noisy data', '{} training data'.format(train_size)])
plt.tight_layout()
plt.savefig('./underlying_function_for_generating_data.jpg', dpi=300)
plt.show()
return x_train, y_train, x_f, y_f
# -
# sns.set(style="darkgrid", font_scale=1.5)
training_data_size = 200#20#50
x_train, y_train, x_f, y_f = plot_training_data_and_underlying_function(train_size=training_data_size,
train_s1=0, train_e1=0.6, train_s2=0.8, train_e2=1.4)
x_train = x_train.reshape(-1,1)
# X_train = np.concatenate([x_train, x_train**2, x_train**3], axis=1)
# X_train = x_train
X_train = np.concatenate([x_train, x_train, x_train], axis=1)
X_train.shape
# # Build Neural Networks
seed=0
x_dim=X_train.shape[1]
y_dim = 1
hidden_sizes = [300, 300]
x_low = -10
x_high = 10
max_steps=int(1e6)
learning_rate=1e-3
batch_size=100
replay_size=int(1e6)
BerDrop_n_post=50#100
dropout_rate = 0.05
bootstrapp_p = 0.75
# +
tf.set_random_seed(seed)
np.random.seed(seed)
# Define input placeholder
x_ph = tf.placeholder(dtype=tf.float32, shape=(None, x_dim))
y_ph = tf.placeholder(dtype=tf.float32, shape=(None, y_dim))
layer_sizes = hidden_sizes + [y_dim]
hidden_activation=tf.keras.activations.relu
output_activation = tf.keras.activations.linear
# +
# 1. Create MLP to learn RTN:
# which is only used for generating target value.
mlp_replay_buffer = ReplayBuffer(x_dim=x_dim, y_dim=y_dim, size=replay_size)
with tf.variable_scope('MLP'):
mlp = MLP(layer_sizes, hidden_activation=hidden_activation, output_activation=output_activation)
mlp_y = mlp(x_ph)
mlp_loss = tf.reduce_mean((y_ph - mlp_y)**2) # mean-square-error
mlp_optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
mlp_train_op = mlp_optimizer.minimize(mlp_loss, var_list=mlp.variables)
# 2. Create BernoulliDropoutMLP:
# which is trained with dropout masks and regularization term
with tf.variable_scope('BernoulliDropoutUncertaintyTrain'):
bernoulli_dropout_mlp = BeroulliDropoutMLP(layer_sizes, weight_regularizer=1e-6, dropout_rate=dropout_rate,
hidden_activation = hidden_activation,
output_activation = output_activation)
ber_drop_mlp_y = bernoulli_dropout_mlp(x_ph, training=True) # Must set training=True to use dropout mask
ber_drop_mlp_reg_losses = tf.reduce_sum(
tf.losses.get_regularization_losses(scope='BernoulliDropoutUncertaintyTrain'))
ber_drop_mlp_loss = tf.reduce_sum(
(y_ph - ber_drop_mlp_y) ** 2 + ber_drop_mlp_reg_losses) # TODO: heteroscedastic loss
ber_drop_mlp_optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
ber_drop_mlp_train_op = ber_drop_mlp_optimizer.minimize(ber_drop_mlp_loss,
var_list=bernoulli_dropout_mlp.variables)
# 3. Create lazy BernoulliDropoutMLP:
# which copys weights from MLP by:
# lazy_bernoulli_dropout_mlp_sample.set_weights(mlp.get_weights())
# then post sample predictions with dropout masks.
with tf.variable_scope('LazyBernoulliDropoutUncertaintySample'):
lazy_bernoulli_dropout_mlp = BeroulliDropoutMLP(layer_sizes, weight_regularizer=1e-6, dropout_rate=dropout_rate,
hidden_activation=hidden_activation,
output_activation=output_activation)
lazy_ber_drop_mlp_y = lazy_bernoulli_dropout_mlp(x_ph, training=True) # Set training=True to sample with dropout masks
lazy_ber_drop_mlp_update = tf.group([tf.assign(v_lazy_ber_drop_mlp, v_mlp)
for v_mlp, v_lazy_ber_drop_mlp in zip(mlp.variables, lazy_bernoulli_dropout_mlp.variables)])
# Create BootstrappedEnsembleNN
with tf.variable_scope('BootstrappedEnsembleUncertainty'):
boots_ensemble = BootstrappedEnsemble(ensemble_size=BerDrop_n_post, x_dim=x_dim, y_dim=y_dim, replay_size=replay_size,
x_ph=x_ph, y_ph=y_ph, layer_sizes=layer_sizes,
hidden_activation=hidden_activation,
output_activation=output_activation,
learning_rate=learning_rate)
# -
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# # Training
# Add training set to bootstrapped_ensemble
for i in range(X_train.shape[0]):
boots_ensemble.add_to_replay_buffer(X_train[i], y_train[i], bootstrapp_p=bootstrapp_p)
# +
training_epoches = 500#1000#500
ber_drop_mlp_train_std = np.zeros((training_epoches,))
ber_drop_mlp_train_loss = np.zeros((training_epoches,))
lazy_ber_drop_mlp_train_std = np.zeros((training_epoches,))
lazy_ber_drop_mlp_train_loss = np.zeros((training_epoches,))
boots_ensemble_train_std = np.zeros((training_epoches,))
boots_ensemble_train_loss = np.zeros((training_epoches,))
for ep_i in range(training_epoches):
if ep_i%100==0:
print('epoch {}'.format(ep_i))
# TODO: uncertainty on training set
# repmat X_train for post sampling: N x BerDrop_n_post x x_dim
ber_drop_mlp_post = np.zeros([X_train.shape[0], BerDrop_n_post, y_dim])
lazy_ber_drop_mlp_post = np.zeros([X_train.shape[0], BerDrop_n_post, y_dim])
boots_ensemble_post = np.zeros([X_train.shape[0], BerDrop_n_post, y_dim])
for x_i in range(X_train.shape[0]):
x_post = np.matlib.repmat(X_train[x_i,:], BerDrop_n_post, 1) # repmat x for post sampling
# BernoulliDropoutMLP
ber_drop_mlp_post[x_i,:,:] = sess.run(ber_drop_mlp_y, feed_dict={x_ph: x_post})
# LazyBernoulliDropoutMLP
lazy_ber_drop_mlp_post[x_i,:,:] = sess.run(lazy_ber_drop_mlp_y, feed_dict={x_ph: x_post})
# BootstrappedEnsemble
boots_ensemble_post[x_i,:,:] = boots_ensemble.prediction(sess, X_train[x_i,:])
# Everage std on training set
ber_drop_mlp_train_std[ep_i] = np.mean(np.std(ber_drop_mlp_post,axis=1))
lazy_ber_drop_mlp_train_std[ep_i] = np.mean(np.std(lazy_ber_drop_mlp_post,axis=1))
boots_ensemble_train_std[ep_i] = np.mean(np.std(boots_ensemble_post,axis=1))
# Train MLP
mlp_outs = sess.run([mlp_loss, mlp_train_op], feed_dict={x_ph: X_train, y_ph: y_train.reshape(-1,y_dim)})
lazy_ber_drop_mlp_train_loss[ep_i] = mlp_outs[0]
sess.run(lazy_ber_drop_mlp_update) # copy weights
# Train BernoulliDropoutMLP on the same batch with MLP
ber_drop_outs = sess.run([ber_drop_mlp_loss, ber_drop_mlp_train_op], feed_dict={x_ph:X_train, y_ph: y_train.reshape(-1,y_dim)})
ber_drop_mlp_train_loss[ep_i] = ber_drop_outs[0]
# Train BootstrappedEnsemble
boots_ensemble_loss = boots_ensemble.train(sess, batch_size)
boots_ensemble_train_loss[ep_i] = np.mean(boots_ensemble_loss)
# +
marker = '.'
markersize = 1
# Loss
f, axes = plt.subplots(1, 3)
f.set_figwidth(18)
f.set_figheight(3.5)
axes[0].plot(lazy_ber_drop_mlp_train_loss, marker, markersize=markersize)
axes[0].set_title('LazyBernoulliDropout (MLP) Average Training Loss')
axes[0].set_xlabel('Training Epochs')
axes[0].set_ylabel('Loss Value on Training Data')
axes[1].plot(ber_drop_mlp_train_loss, marker, markersize=markersize)
axes[1].set_title('BernoulliDropout Average Training Loss')
axes[1].set_xlabel('Training Epochs')
axes[2].plot(boots_ensemble_train_loss, marker, markersize=markersize)
axes[2].set_title('BootsEnsemble Average Training Loss')
axes[2].set_xlabel('Training Epochs')
f.savefig('./toy_example_loss_on_training_data.jpg', dpi=300)
# Uncertainty
f, axes = plt.subplots(1, 3, sharey=True)
f.set_figwidth(18)
f.set_figheight(3.5)
axes[0].plot(lazy_ber_drop_mlp_train_std, markersize=markersize)
axes[0].set_title('Lazy Bernoulli Dropout Average Uncertainty')
axes[0].set_xlabel('Training Epochs')
axes[0].set_ylabel('Average Uncertainty on Trainig Data')
axes[1].plot(ber_drop_mlp_train_std,marker, markersize=markersize)
axes[1].set_title('Bernoulli Dropout Average Uncertainty')
axes[1].set_xlabel('Training Epochs')
axes[2].plot(boots_ensemble_train_std, marker, markersize=markersize)
axes[2].set_title('Bootstrapped Ensemble Average Uncertainty')
axes[2].set_xlabel('Training Epochs')
f.savefig('./toy_example_uncertainty_on_training_data.jpg', dpi=300)
# -
# # Post Sampling to Estimate Uncertainty
x_test = np.arange(-1, 2, 0.005)
x_test = x_test.reshape(-1,1)
# X_test = np.concatenate([x_test, x_test**2, x_test**3], axis=1)
# X_test = x_test
X_test = np.concatenate([x_test, x_test, x_test], axis=1)
X_test.shape
# +
# post sampling
mlp_postSamples = np.zeros([X_test.shape[0], BerDrop_n_post, y_dim])
ber_drop_mlp_postSamples = np.zeros([X_test.shape[0], BerDrop_n_post, y_dim])
lazy_ber_drop_mlp_postSamples = np.zeros([X_test.shape[0], BerDrop_n_post, y_dim])
boots_ensemble_postSamples = np.zeros([X_test.shape[0], BerDrop_n_post, y_dim])
for i in range(X_test.shape[0]):
x = X_test[i,:]
x_postSampling = np.matlib.repmat(x, BerDrop_n_post, 1) # repmat x for post sampling
# MLP
mlp_postSamples[i,:,:] = sess.run(mlp_y, feed_dict={x_ph: x_postSampling})
# BernoulliDropoutMLP
ber_drop_mlp_postSamples[i,:,:] = sess.run(ber_drop_mlp_y, feed_dict={x_ph: x_postSampling})
# LazyBernoulliDropoutMLP
sess.run(lazy_ber_drop_mlp_update) # copy weights
lazy_ber_drop_mlp_postSamples[i,:,:] = sess.run(lazy_ber_drop_mlp_y, feed_dict={x_ph: x_postSampling})
# BootstrappedEnsemble
boots_ensemble_postSamples[i,:,:] = boots_ensemble.prediction(sess, x)
# +
mlp_mean = np.mean(mlp_postSamples,axis=1)
mlp_std = np.std(mlp_postSamples,axis=1)
ber_drop_mlp_mean = np.mean(ber_drop_mlp_postSamples,axis=1)
ber_drop_mlp_std = np.std(ber_drop_mlp_postSamples,axis=1)
lazy_ber_drop_mlp_mean = np.mean(lazy_ber_drop_mlp_postSamples,axis=1)
lazy_ber_drop_mlp_std = np.std(lazy_ber_drop_mlp_postSamples,axis=1)
boots_ensemble_mean = np.mean(boots_ensemble_postSamples,axis=1)
boots_ensemble_std = np.std(boots_ensemble_postSamples,axis=1)
# +
markersize = 5
f, axes = plt.subplots(1,4,sharey=True)
# f.suptitle('n_training_data={}, n_post_samples={}, dropout_rate={}, n_trainig_epochs={}, bootstrapp_p={}'.format(training_data_size,
# BerDrop_n_post,
# dropout_rate,
# training_epoches,
# bootstrapp_p),
# fontsize=20)
f.set_figwidth(20)
f.set_figheight(4)
axes[0].plot(x_test, mlp_mean, 'k')
axes[0].plot(x_train, y_train, 'r.', markersize=markersize)
axes[0].plot(x_f, y_f,'m', alpha=0.5)
axes[0].fill_between(x_test.flatten(),
(mlp_mean+mlp_std).flatten(),
(mlp_mean-mlp_std).flatten())
axes[0].set_title('MLP', fontsize=15)
axes[1].plot(x_test, lazy_ber_drop_mlp_mean, 'k')
axes[1].plot(x_train, y_train, 'r.', markersize=markersize)
axes[1].plot(x_f, y_f,'m', alpha=0.5)
axes[1].fill_between(x_test.flatten(),
(lazy_ber_drop_mlp_mean+lazy_ber_drop_mlp_std).flatten(),
(lazy_ber_drop_mlp_mean-lazy_ber_drop_mlp_std).flatten())
axes[1].set_title('LazyBernoulliDropoutMLP', fontsize=15)
axes[2].plot(x_test, ber_drop_mlp_mean, 'k')
axes[2].plot(x_train, y_train, 'r.', markersize=markersize)
axes[2].plot(x_f, y_f,'m', alpha=0.5)
axes[2].fill_between(x_test.flatten(),
(ber_drop_mlp_mean+ber_drop_mlp_std).flatten(),
(ber_drop_mlp_mean-ber_drop_mlp_std).flatten())
axes[2].set_title('BernoulliDropoutMLP', fontsize=15)
prediction_mean_h, = axes[3].plot(x_test, boots_ensemble_mean, 'k')
training_data_h, = axes[3].plot(x_train, y_train, 'r.', markersize=markersize)
underlying_function_h, = axes[3].plot(x_f, y_f,'m', alpha=0.5)
prediction_std_h = axes[3].fill_between(x_test.flatten(),
(boots_ensemble_mean+boots_ensemble_std).flatten(),
(boots_ensemble_mean-boots_ensemble_std).flatten())
axes[3].set_title('BootstrappedEnsemble', fontsize=15)
axes[3].set_ylim(-6, 9)
axes[0].legend(handles=[underlying_function_h, training_data_h, prediction_mean_h, prediction_std_h],
labels=['underlying function', '{} training data'.format(training_data_size), 'prediction mean', 'prediction mean $\pm$ standard deviation'])
plt.tight_layout()
f.subplots_adjust(top=0.8)
plt.savefig('./toy_example_comparison_of_uncertainty_estimation.jpg', dpi=300)
# -
| spinup/algos/uncertainty_estimate/Comparison_of_Uncertainty_Estimation_on_Toy_Example-Copy2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# !pip install citipy
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
# Import API key
from config import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/world_cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# -
#
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
count = 1
# instantiate the lists to hold the data for each city
city_list = []
cloudiness = []
country = []
date = []
humidity = []
lat = []
lng = []
max_temp = []
wind_speed = []
for city in cities:
# assemble url and make API request
print(f"Processing Record {count} of Set 1 | {city}.")
count = count + 1
target_url = ("http://api.openweathermap.org/data/2.5/weather?"
'appid={0}&q={1}').format(weather_api_key,city)
try:
# extract results
response = requests.get(target_url).json()
# add the city info to the lists
city_list.append(response['name'])
cloudiness.append(response['clouds']['all'])
country.append(response['sys']['country'])
date.append(response['dt'])
humidity.append(response['main']['humidity'])
lat.append(response['coord']['lat'])
lng.append(response['coord']['lon'])
max_temp.append(response['main']['temp_max'])
wind_speed.append(response['wind']['speed'])
except:
print("City not found. Skipping...")
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# +
# convert the max temp to F from Kelvin
max_temp = [(9/5)*(x - 273) + 32 for x in max_temp]
# populate the dataframe with the city info
city_df = pd.DataFrame({
"City":city_list,
"Cloudiness":cloudiness,
"Country":country,
"Date":date,
"Humidity":humidity,
"Lat":lat,
"Lng":lng,
"Max Temp":max_temp,
"Wind Speed":wind_speed
})
# write the dataframe to a csv
city_df.to_csv("world_cities.csv")
city_df.head()
# -
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# +
#No Cities with humidity > 100%
# -
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
fig, ax = plt.subplots()
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (F)')
ax.set_title('City Latitude vs. Max Temperature')
ax.scatter(city_df['Lat'],city_df['Max Temp'],edgecolors='black')
plt.grid()
fig.savefig('lat_vs_max_temp.png')
# ## Latitude vs. Humidity Plot
fig, ax = plt.subplots()
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
ax.set_title('City Latitude vs. Humidity')
ax.scatter(city_df['Lat'],city_df['Humidity'],edgecolors='black')
plt.grid()
fig.savefig('lat_vs_humidity.png')
# ## Latitude vs. Cloudiness Plot
fig, ax = plt.subplots()
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
ax.set_title('City Latitude vs. Cloudiness')
ax.scatter(city_df['Lat'],city_df['Cloudiness'],edgecolors='black')
plt.grid()
fig.savefig('lat_vs_cloudiness.png')
# ## Latitude vs. Wind Speed Plot
fig, ax = plt.subplots()
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
ax.set_title('City Latitude vs. Wind Speed')
ax.scatter(city_df['Lat'],city_df['Wind Speed'],edgecolors='black')
plt.grid()
fig.savefig('lat_vs_wind_speed.png')
# ## Linear Regression
x_val=north_hem["Latitude"]
y_val=north_hem["Temperature"]
(slope, intercept, rvalue, pvalue, stderr)=linregress(x_val,y_val)
regress_val=x_val*slope+intercept
line_eq="y="+str(round(slope,2))+"x+"+str(round(intercept,2))
plt.scatter(x_val,y_val)
plt.plot(x_val,regress_val,"r-")
plt.annotate(line_eq,(-50,80),color="red")
plt.title("North Hemisphere: Latitude v Temperature")
plt.xlabel("Latitude")
plt.ylabel("Temperature")
print(f"rvalue={round(rvalue,2)}")
plt.savefig("../Images/north_hem_temp.png")
plt.show()
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
| WeatherPy/WeatherPy_Starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="h5oP7zLQACZR"
# # Intro to Digital Agriculture
# ## Week 6
# Instructors: <NAME>, <NAME>
#
# TAs for this week: <NAME>, <NAME>
#
# On this practical we will work with the satellite images, look for agricultural fields and visualize them
# + [markdown] id="QD9-AvLKCEQJ"
# # Part 0
# ### Data download and preparation
# Lets first download the data (can take several minutes)
# + id="6_yz9jPtAB6q"
# !wget --no-check-certificate "https://onedrive.live.com/download?cid=1E2DE865E90D4259&resid=1E2DE865E90D4259%21258622&authkey=<KEY>" -O Farmpins.zip
# !wget --no-check-certificate "https://onedrive.live.com/download?cid=1E2DE865E90D4259&resid=1E2DE865E90D4259%21195470&authkey=AFPw5W-8uzm5vpM" -O unet_parts.py
# + id="ywXB6zwlSlaB"
# !unzip ./Farmpins.zip
# + [markdown] id="gX0DUzrPFPQc"
# And import necessary libraries
# + id="X-xXliNb-QhN"
import numpy as np
import imageio as io
import pandas as pd
import skimage.transform as transforms
import skimage.util as utils
import os
from matplotlib import pyplot as plt
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from torch.utils.tensorboard import SummaryWriter
from torch.optim import Adam, lr_scheduler
import gdal
from unet_parts import *
import torchvision
from tqdm import tnrange, tqdm
from IPython.display import clear_output
# + id="xpihfoyP-QhW"
from sklearn.metrics import accuracy_score, confusion_matrix, recall_score, f1_score, precision_score, classification_report, SCORERS
# + [markdown] id="CA5dTlCoGZlp"
# # Part 1. EDA
#
# Lets load the images and look what we have inside
# + id="2oqFprb7GlFq"
data_f = gdal.Open('./20170101_mosaic_cropped.tif')
data_mask_train = gdal.Open('./train_crops.tif')
# + [markdown] id="hrUGLAEnUUjB"
# `fields` - data tensor (satellite image) with surface reflectance values
#
# `train_mask` - crop labels for field (used for training)
# + id="QOxtI5WzGuuF"
fields = np.array(data_f.ReadAsArray())
train_mask = np.array(data_mask_train.ReadAsArray())
# + id="aezLtcIqHEK3"
train_id = gdal.Open('./train_field_id.tif')
train_field_id_map = np.array(train_id.ReadAsArray()).astype(np.uint16)
train_field_id_map = np.hstack([train_field_id_map, np.zeros((train_field_id_map.shape[0], 1))])
train_field_id_map = train_field_id_map*(train_mask > 0)
# train_field_id = np.unique(train_field_id_map)
# train_field_id = train_field_id[train_field_id > 0]
# + [markdown] id="hKcQKTwDUrw_"
# Lets plot `train_field_id_map`
# + id="1fVbzbaOUIde"
plt.figure(figsize=(15, 15))
plt.imshow(train_field_id_map)
# plt.colorbar()
# + [markdown] id="ZV313q5TU434"
# As we can see, `train_field_id_map` contains masks with field ids (pixels corresponding to some particular field #$n$ will have all value $n$)
# + colab={"base_uri": "https://localhost:8080/"} id="juCo_lK3VXpB" outputId="34db42e3-700f-4d70-d611-9f4ab0871643"
train_mask[train_mask > 15] = 0
np.unique(train_mask)
# + [markdown] id="b1dZ7EYdVqpB"
# We can also see, that it has only 10 unique values, thus it is most probably (and really is) crop types (out labels for field). Lets visualize them.
# + id="p9TaQQkzVPqW"
plt.figure(figsize=(15, 15))
plt.imshow(train_mask)
# + [markdown] id="K3wOkKPEWKYo"
# Now it is time to look at satellite image itself. Look at image shape. It should have 13 channels.
# + colab={"base_uri": "https://localhost:8080/"} id="eKRuw4kaWax4" outputId="4b430561-42f2-4ee8-c7e8-806b064cc19a"
fields.shape
# + [markdown] id="KOv9_rpMWeWV"
# Since we are working with the Sentinel - it captures multispectral images: in our case it has 13 channels. In order to plot the RGB or any other image of the planet surface we should Google tech sheet for Sentinel 2B satellite and search for channels description :[Sentinel 2 Wiki Page](https://en.wikipedia.org/wiki/Sentinel-2#Instruments)
#
# Lets plot the NIR channel (7)
# + id="3X1mlv07ahCO"
plt.figure(figsize=(15, 15))
plt.imshow(fields[7, :2000, :3000], cmap='gray')
# + [markdown] id="37SvP2PLcUcg"
# One can play and plot different channels and convince yourself that different channels have different spatial resolution. Sentinel have 3 types of resolutions (60m, 20m and 10m per pix). They all have been upsampled up to 10m per pixel, but visually one can notice the difference
# + [markdown] id="ADUo4E4lc-PY"
# Lets visualize agro fields on the image
# + id="UY9zeQZZc-vh"
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
#here we define the color scheme for our filed, such that every crop should have
#its own constant color
viridis = cm.get_cmap('viridis', 256)
train_colors = viridis(np.linspace(0, 1, 256))
transp = np.array([0, 0, 0, 0])
train_colors[:25, :] = transp
train_cmp = ListedColormap(train_colors)
plt.figure(figsize=(15, 15))
plt.imshow(fields[12, :2000, :3000], cmap='gray')
plt.imshow(train_mask[:2000, :3000], alpha=0.5, cmap=train_cmp)
# + [markdown] id="Nz_FidMOXWxg"
# **Task 1 (1 point for each index)**. Plot the NDVI, EVI, NDRE indices for patch of original image for coordinates(pixels) in range for x in [0, 3000] and y in [500:1500]. Just Google the indices, if you do not know exact formulae.
#
# *Hint: mind the proper order when slicing the tensor*
# + [markdown] id="TaP27XSaYxw_"
# Since our dataset is imbalanced, training of even simplest segmentation model could be quite difficult. One of the simplest ways to fight with this issue is to assign the weight for each class in a loss function.
#
# **Task 2 (2 points)**. Calculate the number of *pixels* (1 pt) and *fields* (1 pt) for each crop and print them out. This information will help us further in training loop
# + [markdown] id="TyDqhsp-dxQG"
# # Part 2. Download data for training
# + [markdown] id="Jlh7ZQoOT6hg"
# In order to train the network, we need to split our big satellite image into patches in a such way, that on the one hand, we could process it using neural networks, but on the other hand we could split it on training/validation set with equal contribution of all classes.
#
# One way to do this is to crop each field from the original image and resize them (for classification) or not (for segmentation). Another way is to just use patches with constant size. In this work we will use second approach, even though it is much harder to properly split it with equal class proportions. One can try to do that for additional points, but I suggest you to download prepared already split dataset with the link below.
#
# + id="efW4k3VGT535"
# !wget --no-check-certificate "https://onedrive.live.com/download?cid=1E2DE865E90D4259&resid=1E2DE865E90D4259%21258624&authkey=<KEY>" -O Patches.zip
# !unzip ./Patches.zip
# + [markdown] id="ExS53xOS1MlV"
# The main task is to train networks for classification using fully convolutional network like UNet with Squeeze and Excitation blocks.
#
# Downloaded data have the following structure: it has the `train` and `val` folder with the patches for *train* and *validation* purpouses. Each folder contains 2 folders: `images` with the multispectral image patches and `labels` of corresponding crops.
# + [markdown] id="yyEYzwvh2Oo3"
# Bellow the structure of a network is defined. Ones who are interested can look through it and investigate it. It is basically the UNet with channel-wise attention mechanism called Squeeze and Excitation (SE_block here)
# + id="zCxqgk2P-QhX"
class SE_block(nn.Module):
def __init__(self, channels, squeese_rate=1):
super(SE_block, self).__init__()
self.fc1 = nn.Linear(channels, channels//squeese_rate)
self.fc2 = nn.Linear(channels//squeese_rate, channels)
def forward(self, input):
g_avg = torch.mean(input, [-1, -2])
x = self.fc1(g_avg).relu()
x = self.fc2(x).sigmoid()
return torch.unsqueeze(torch.unsqueeze(x, 2), 3)*input
# + id="VR02P7Yj-QhX"
class UNet(nn.Module):
def __init__(self, n_channels, n_classes):
super(UNet, self).__init__()
self.inc = inconv(n_channels, 64)
self.down1 = down(64, 128)
self.se1 = SE_block(128)
self.down2 = down(128, 256)
self.se2 = SE_block(256)
self.down3 = down(256, 256) #was 512
self.se3 = SE_block(256) #was 512
self.down4 = down(512, 512)
self.up1 = up(1024, 256, bilinear=False)
self.up2 = up(512, 128, bilinear=False)
self.up3 = up(256, 64, bilinear=False)
self.up4 = up(128, 64, bilinear=False)
self.outc = outconv(64, n_classes)
def forward(self, x):
x1 = self.inc(x)
x2 = self.down1(x1)
x3 = self.down2(self.se1(x2))
x4 = self.down3(self.se2(x3))
# x5 = self.down4(self.se3(x4))
# x = self.up1(x5, x4)
x = self.up2(x4, x3)
x = self.up3(x, x2)
x = self.up4(x, x1)
x = self.outc(x)
return x
# + [markdown] id="YgiShdCJ2y7k"
# Since we have relatively low amount of data, we should use augmentation in order to prevent overfitting and increase generalization a little bit.
#
# Below are the parameters for augmentation
# + id="_RFyphL1-QhY"
img_size = 100
pad_size = img_size//2
shift = img_size//5
rot_angle = 60
# + [markdown] id="epyH1Eps7tue"
# It is also important to normalize the data, so do it.
#
#
# **Task 3 (1 point)** Calculate the *mean* and *standart deviation* for each channel in the input image.
# + id="iAnc6Nbh-QhY"
'''
mean, std: torch.tensor or nd.array of shape (channels)
'''
# Your code here
# mean = ...
# std = ...
# + [markdown] id="WfOvktGQ88rD"
# **Task 4 (2 points)** Below you can see the bodies for the functions to additional features represented by different vegetation indices: we suggest you to fill in these functions with the code for generation features, but you can also find some additional features and implement them here in a similar fashion
# + id="Jr4-Tjo--QhY"
def generate_NDVI(features):
'''
Arguments:
features: torch.tensor of shape (Chanels, H, W)
Return:
ndvi: torch.tensor of shape (H, W)
'''
# Your code here
# ndvi = ...
return ndvi
# + id="S82GTXfK-QhZ"
def generate_EVI(features):
'''
Arguments:
features: torch.tensor of shape (Chanels, H, W)
Return:
evi: torch.tensor of shape (H, W)
'''
# Your code here
# evi = ...
return evi
# + id="g6cSJjth-QhZ"
def generate_NDRE(features):
'''
Arguments:
features: torch.tensor of shape (Chanels, H, W)
Return:
ndre: torch.tensor of shape (H, W)
'''
# Your code here
# ndre = ...
return ndre
# + id="IlPAMTm9-QhZ"
def generate_MSAVI(features):
'''
Arguments:
features: torch.tensor of shape (Chanels, H, W)
Return:
msavi: torch.tensor of shape (H, W)
'''
# Your code here
# msavi = ...
return msavi
# + [markdown] id="BsFqEam40vom"
# Here we gather all generated features in one tensor
# + id="ctC1nCI0-QhZ"
def generate_all_indices(features, u2b4evi=True):
ndvi = generate_NDVI(features)
evi = generate_EVI(features, u2b4evi)
ndre = generate_NDRE(features)
msavi = generate_MSAVI(features)
return torch.cat([features, ndvi.unsqueeze(0), evi.unsqueeze(0), ndre.unsqueeze(0), msavi.unsqueeze(0)], dim=0)
# + [markdown] id="vmyM9OlL0_Z7"
# This is the class for `torch` `Dataset` which loads the data from folders and do the preprocessing after loading
# + [markdown] id="FPUvCEwT1SYF"
# We have computed `mean` and `std` in previous task, thus we should apply it. Keep in mind that one should apply normalization after all the features are generated and concatenated to the original features.
#
# **Task 4 (1 point)** Using function `generate_all_features()` you are suggested to calculate all indices and normalize the satellite image.
#
# *Hint: It is recommended to normalize only satellite image part and leave generated indices as it is*
# + id="uvyCD_hh-Qha"
class CropFieldsDataset(Dataset):
def __init__(self, images_dir, labels_dir, transform=None):
self.images_dir = images_dir
self.labels_dir = labels_dir
self.transform = transform
i_list = os.listdir(self.images_dir)
self.im_list = []
for image in i_list:
if (image[-3:] == 'npy'):
self.im_list.append(image)
self.mean = mean[:, None, None]
self.std = std[:, None, None]
def __len__(self):
return len(self.im_list)
def __getitem__(self, idx):
img_name = os.path.join(self.images_dir, self.im_list[idx])
image = np.load(img_name)#*255
labels_name = os.path.join(self.labels_dir, self.im_list[idx])
labels = np.load(labels_name)
if(self.transform != None):
randangle = np.random.randint(-rot_angle, rot_angle)
sc = tuple(np.random.uniform(0.75, 1, 2))
tf = transforms.AffineTransform(scale = sc)
image = transforms.rotate(image, randangle, mode='reflect')
image = utils.pad(image, ((pad_size, pad_size), (pad_size, pad_size), (0, 0)), mode='reflect')
image = transforms.warp(image, tf, mode='reflect')
labels = transforms.rotate(labels, randangle, mode='reflect', order=0, preserve_range=True)
labels = utils.pad(labels, ((pad_size, pad_size), (pad_size, pad_size)), mode='reflect')
labels = transforms.warp(labels, tf, mode='reflect', order=0)
M, N, D = image.shape
randshiftx = np.random.randint(-shift, shift)
randshifty = np.random.randint(-shift, shift)
image = utils.crop(image, (((M + randshiftx - img_size)//2, (M - randshiftx + 1 - img_size)//2),
((N + randshifty - img_size)//2, (N - randshifty + 1 - img_size)//2), (0, 0)))
labels= utils.crop(labels,(((M + randshiftx - img_size)//2, (M - randshiftx + 1 - img_size)//2),
((N + randshifty - img_size)//2, (N - randshifty + 1 - img_size)//2)))
image = torchvision.transforms.ToTensor()(image)
labels = torch.tensor(labels).to(torch.long)
## Your code here
# image = ...
# norm_image = ...
## ------------------
return norm_image, labels
# + [markdown] id="o-cjbbsKB8dU"
# Lets make the instance for `Dataset` and make the dataloade (one could try different batch sizes which is appropriate for GPU you use)
# + id="BtwHK6_e-Qha"
train_set = CropFieldsDataset("./train/images", "./train/labels", transform=True)
train_loader = DataLoader(train_set, batch_size=8, shuffle=True)
# + [markdown] id="2uLqImLUCTrR"
# And lets visualize the how augmentation works
# + id="4T7gHkAm-Qhb"
image, labels = train_set[10]
_, ax = plt.subplots(ncols=2)
ax[0].imshow(image[12, :, :], cmap='gray')
ax[1].imshow(labels[:, :])
image.shape
# + [markdown] id="z6c9Y8yVCcxD"
# Here comes the validation dataset also
# + id="rWFmm-fY-Qhd"
val_set = CropFieldsDataset("./val/images", "./val/labels")
val_loader = DataLoader(val_set, batch_size=1)
# + [markdown] id="xn0rRbjGCjJk"
# One could look for the expected result, which is generated from validation dataset
# + id="vQWWd9Ty-Qhe"
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
viridis = cm.get_cmap('viridis', 256)
train_colors = viridis(np.linspace(0, 1, 256))
test_colors = viridis(np.linspace(0, 1, 256))
transp = np.array([0, 0, 0, 0])
train_colors[:25, :] = transp
train_cmp = ListedColormap(train_colors)
# + id="6_kHRt0w-Qhe"
# len(train_set)
image, labels = val_set[18]
print(image.shape)
_, ax = plt.subplots(ncols=2, figsize=(10, 10), dpi=100)
ax[0].imshow(image[2, :, :], cmap='gray')
ax[1].imshow(image[2, :, :], cmap='gray')
ax[1].imshow(labels[:, :], cmap=train_cmp)
ax[0].set_axis_off()
ax[1].set_axis_off()
ax[0].set_title('Band 3')
ax[1].set_title('Crop map')
# + [markdown] id="WMfnDODUCrQb"
# Now we are going to train the network
# + id="EHV_Y6ey-Qhe"
if(torch.cuda.is_available()):
device = torch.device('cuda:0')
else:
device = torch.device('cpu')
# Here we initialize the network: UNet(input_features, n_classes)
# If you generate additional features, you should put here number of channels
# of your final input (13 + number_of_generated_indices)
model = UNet(17, 10).to(device)
# + id="munZdXfM-Qhf"
def get_lr(optimizer):
for param_group in optimizer.param_groups:
return param_group['lr']
# + [markdown] id="SaAmFnXtDUzh"
# **Task 5 (1 point)** Complete training loop with missing parts (look at the comments for guidance) and
#
# **Task 6 (2 points)** Train your network. You will get full score (**2 points**) if the $accuracy$ on validation $\geq 63\%$, **1 point** if it will be in range $50\% \leq accuracy < 63\%$, and **no points** if $accuracy < 50\%$
# + id="Af57Az-8-Qhf"
train_weights = torch.tensor([0, 0.14852941, 0, 0.28836312, 0.04097108, 0.07013557, 0.10067904, 0.07735483, 0.02633347, 0]).to(device)
val_weights = torch.tensor([0, 0.20184716, 0, 0.23679423, 0.03584126, 0.06965259, 0.09625471, 0.07470775, 0.02500757, 0]).to(device)
optimizer = Adam(model.parameters(), lr = 0.01, weight_decay=0.0001)
train_criterion = nn.CrossEntropyLoss(ignore_index=0, weight=train_weights, reduction='mean').to(device)
val_criterion = nn.CrossEntropyLoss(ignore_index=0, weight=val_weights, reduction='mean').to(device)
lr_sch = lr_scheduler.StepLR(optimizer, 1, gamma=0.977)
epochs_num = 350
loss_list = []
acc_list = []
best_acc = 0.0 #0.628
for epoch in range(epochs_num): # loop over the dataset multiple times
running_train_loss = 0.0
train_accuracy_pix = 0
train_sum_pix = 0
model.train()
for data in tqdm(train_loader):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# transfer data to the device
# inputs = ...
# labels = ...
# zero the parameter gradients
# ...
# make forward pass, compute the loss, make backward pass and optimization step
# outputs = ...
# loss = ...
# ...
# ...
# print statistics
running_train_loss += loss.item()
result = F.softmax(outputs, dim=1).detach().cpu().numpy()
pred = np.argmax(result, axis=1)
labels = labels.cpu().numpy()
fl_labels = labels.flatten()
train_accuracy_pix += np.sum(pred.flatten()[fl_labels > 0] == fl_labels[fl_labels > 0])
train_sum_pix += np.sum(labels > 0)
# train_writer.add_scalar('Loss', running_train_loss, global_step=epoch)
running_train_loss /= len(train_set)
loss_list.append(running_train_loss)
acc_list.append(train_accuracy_pix/train_sum_pix)
model.eval()
running_val_loss = 0.0
val_accuracy_pix = 0
val_sum_pix = 0
for data in tqdm(val_loader):
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
with torch.no_grad():
outputs = model(inputs)
loss = val_criterion(outputs, labels)
running_val_loss += loss.item()
result = F.softmax(outputs, dim=1).detach().cpu().numpy()
pred = np.argmax(result, axis=1)
labels = labels.cpu().numpy()
fl_labels = labels.flatten()
val_accuracy_pix += np.sum(pred.flatten()[fl_labels > 0] == fl_labels[fl_labels > 0])
val_sum_pix += np.sum(labels > 0)
running_val_loss /= len(val_set)
# train_writer.add_scalar('Accuracy', train_accuracy_pix/train_sum_pix, global_step=epoch)
# train_writer.add_scalar("Loss", running_train_loss, global_step=epoch)
# val_writer.add_scalar('Accuracy', val_accuracy_pix/val_sum_pix, global_step=epoch)
# val_writer.add_scalar("Loss", running_val_loss, global_step=epoch)
print("Epoch: {0:3d} | LR: {1}\n Train Loss: {2:4f} \n Train Acc: \t {3:4.1f}%\n Val Loss: {4:4f} \n Val Acc: \t {5:4.1f}%".format(
epoch+1, get_lr(optimizer),
running_train_loss,
100*train_accuracy_pix/train_sum_pix,
running_val_loss,
100*val_accuracy_pix/val_sum_pix))
if (val_accuracy_pix/val_sum_pix > best_acc):
best_acc = val_accuracy_pix/val_sum_pix
torch.save(model.state_dict(), 'best_model.pth')
lr_sch.step()
print('Finished Training')
# + [markdown] id="_86qYlsypUem"
# Next cells are just for validation
#
#
#
# + id="4Srzo8TJ-Qhh"
model.eval()
val_accuracy_pix = 0
val_sum_pix = 0
correct_class_pix = np.zeros((10), dtype=np.int)
class_pix = np.zeros((10), dtype=np.int)
preds_list = []
labels_list = []
for data in tqdm(val_loader):
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
with torch.no_grad():
outputs = model(inputs)
result = F.softmax(outputs, dim=1).detach().cpu().numpy()
pred = np.argmax(result, axis=1)
preds_list.append(pred.flatten())
labels = labels.cpu().numpy()
fl_labels = labels.flatten()
labels_list.append(fl_labels)
val_accuracy_pix += np.sum(pred.flatten()[fl_labels > 0] == fl_labels[fl_labels > 0])
val_sum_pix += np.sum(labels > 0)
for i in range(10):
class_pix[i] += np.sum(labels == i)
correct_class_pix[i] += np.sum((pred == i)*(labels == i))
class_acc = correct_class_pix/class_pix
pix_df = pd.DataFrame(data=np.vstack([correct_class_pix, class_pix]),
index=['correct_pix', 'pix'],
columns=np.arange(10))
acc_df = pd.DataFrame(data=np.round(class_acc[None, :], 3), index=['accuracy'], columns=np.arange(10))
print('Accuracy', val_accuracy_pix/val_sum_pix)
display(pix_df)
display(acc_df)
# + id="sNp8Aivp-Qhh"
all_preds = np.hstack(preds_list)
all_labels = np.hstack(labels_list)
# + [markdown] id="AmF942STqzDx"
# Cell below are for visualization of correctness of the models prediction (green means correct, red - incorrect). Just launch all remaining cells
# + id="rs86Fpj3-Qhi"
images = []
results = []
llabels = []
with torch.no_grad():
for i, data in tqdm(enumerate(val_loader)):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
images.append(inputs.numpy())
llabels.append(labels.numpy())
inputs = inputs.to(device)
# forward
outputs = model(inputs)
result = F.softmax(outputs, dim=1).detach().cpu().numpy()
pred = np.argmax(result, axis=1)
results.append(pred)
# + id="vxTa72ke-Qhj"
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
viridis = cm.get_cmap('viridis', 256)
train_colors = viridis(np.linspace(0, 1, 256))
test_colors = viridis(np.linspace(0, 1, 256))
transp = np.array([0, 0, 0, 0])
train_colors[:24, :] = transp
train_cmp = ListedColormap(train_colors)
pink = np.array([248/256, 24/256, 148/256, 1])
test_colors[:128, :] = transp
test_colors[128:, :] = pink
test_cmp = ListedColormap(test_colors)
# + id="Os5aSTDA-Qhj"
test_scale = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
# + id="-nUlm167-Qhj"
for i in range(len(val_set)):
right = (results[i][0, ...] == llabels[i][0, ...]).astype(int) - (results[i][0, ...] != llabels[i][0, ...]).astype(int)
pic1 = results[i][0, ...]*(llabels[i][0, ...] > 0)
pic2 = llabels[i][0, ...]#.numpy()
pic1[0, :10] = test_scale
pic2[0, :10] = test_scale
_, ax = plt.subplots(ncols=3, figsize=(12, 4))
ax[0].imshow(images[i][0, 3, ...], cmap='gray')
fig1 = ax[0].imshow(pic1, alpha=1, cmap=train_cmp)
ax[0].set_title('Model Prediction')
fig2 = ax[1].imshow(pic2, cmap=viridis)
ax[1].set_title('Ground Truth')
ax[2].imshow(right*(llabels[i][0, ...] > 0), cmap='RdYlGn')
ax[2].set_title('Correctness')
# plt.colorbar(fig1, ax=ax[0])
# plt.colorbar(fig2, ax=ax[1])
ax[0].xaxis.set_visible(False)
ax[0].yaxis.set_visible(False)
ax[1].xaxis.set_visible(False)
ax[1].yaxis.set_visible(False)
ax[2].xaxis.set_visible(False)
ax[2].yaxis.set_visible(False)
# plt.savefig('Figures/fig_{}.jpg'.format(i), dpi=100)
# + id="g6ETVnG-pKVu"
| Satellite_Imagery/Satellite_Images_Hometask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="RCsEqTmRFIlC"
# ## Environment Setup
# + colab={"base_uri": "https://localhost:8080/"} id="OSJsyCG7GMyM" executionInfo={"status": "ok", "timestamp": 1648015138981, "user_tz": -480, "elapsed": 6569, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}} outputId="2469db45-dc35-4a82-a478-29bebcbf190d"
# !pip install albumentations==0.4.6
# + id="bQ261RzvFDSh" executionInfo={"status": "ok", "timestamp": 1648015153103, "user_tz": -480, "elapsed": 9709, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
import torch
import os
import albumentations as A
import cv2
import numpy as np
from albumentations.pytorch import ToTensorV2
import glob as glob
from xml.etree import ElementTree as et
from torch.utils.data import Dataset, DataLoader
import torchvision
from torchvision.models.detection.faster_rcnn import FastRCNNPredictor
from tqdm.auto import tqdm
import matplotlib.pyplot as plt
import time
# + colab={"base_uri": "https://localhost:8080/"} id="9UUHeuEwFLuT" executionInfo={"status": "ok", "timestamp": 1648015181580, "user_tz": -480, "elapsed": 28495, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}} outputId="31ee5af7-6d11-46e0-ec67-4d728ff85b1d"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="w62DaROVFX-D" executionInfo={"status": "ok", "timestamp": 1648015182873, "user_tz": -480, "elapsed": 1298, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}} outputId="385fa7f2-f865-4b5d-de72-3357d8b6681a"
path = '/content/drive/My Drive/IS5451 Project/'
os.listdir(path)
# + [markdown] id="YbXr03iwF4_2"
# ### Configuration
# + id="FR62C7pSEeCb" executionInfo={"status": "ok", "timestamp": 1648015280048, "user_tz": -480, "elapsed": 911, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
BATCH_SIZE = 128 # increase / decrease according to GPU memeory
RESIZE_TO = 512 # resize the image for training and transforms
NUM_EPOCHS = 5 # number of epochs to train for
DEVICE = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# training images and XML files directory
TRAIN_DIR = path + 'data/train'
# validation images and XML files directory
VALID_DIR = path + 'data/test'
# classes: 0 index is reserved for background
CLASSES = [
'background', 'Beer', 'Butter', 'Bread', 'Yoghurt', 'Egg(s)', 'Cheese', 'Milk', 'Banana'
]
NUM_CLASSES = 3
# whether to visualize images after crearing the data loaders
VISUALIZE_TRANSFORMED_IMAGES = False
# location to save model and plots
OUT_DIR = path + 'outputs'
SAVE_PLOTS_EPOCH = 2 # save loss plots after these many epochs
SAVE_MODEL_EPOCH = 2 # save model after these many epochs
# + [markdown] id="DmjOk6pLF_qY"
# ### Utilities
# + id="vKtlR7A2FzrM" executionInfo={"status": "ok", "timestamp": 1648015283313, "user_tz": -480, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
# this class keeps track of the training and validation loss values...
# ... and helps to get the average for each epoch as well
class Averager:
def __init__(self):
self.current_total = 0.0
self.iterations = 0.0
def send(self, value):
self.current_total += value
self.iterations += 1
@property
def value(self):
if self.iterations == 0:
return 0
else:
return 1.0 * self.current_total / self.iterations
def reset(self):
self.current_total = 0.0
self.iterations = 0.0
# + id="XcSrzkNrGGJU" executionInfo={"status": "ok", "timestamp": 1648015283314, "user_tz": -480, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
def collate_fn(batch):
"""
To handle the data loading as different images may have different number
of objects and to handle varying size tensors as well.
"""
return tuple(zip(*batch))
# define the training tranforms
def get_train_transform():
return A.Compose([
A.Flip(0.5),
A.RandomRotate90(0.5),
A.MotionBlur(p=0.2),
A.MedianBlur(blur_limit=3, p=0.1),
A.Blur(blur_limit=3, p=0.1),
ToTensorV2(p=1.0),
], bbox_params={
'format': 'pascal_voc',
'label_fields': ['labels']
})
# define the validation transforms
def get_valid_transform():
return A.Compose([
ToTensorV2(p=1.0),
], bbox_params={
'format': 'pascal_voc',
'label_fields': ['labels']
})
def show_tranformed_image(train_loader):
"""
This function shows the transformed images from the `train_loader`.
Helps to check whether the tranformed images along with the corresponding
labels are correct or not.
Only runs if `VISUALIZE_TRANSFORMED_IMAGES = True` in config.py.
"""
if len(train_loader) > 0:
for i in range(1):
images, targets = next(iter(train_loader))
images = list(image.to(DEVICE) for image in images)
targets = [{k: v.to(DEVICE) for k, v in t.items()} for t in targets]
boxes = targets[i]['boxes'].cpu().numpy().astype(np.int32)
sample = images[i].permute(1, 2, 0).cpu().numpy()
for box in boxes:
cv2.rectangle(sample,
(box[0], box[1]),
(box[2], box[3]),
(0, 0, 255), 2)
cv2.imshow('Transformed image', sample)
cv2.waitKey(0)
cv2.destroyAllWindows()
# + [markdown] id="J-FEjbTjHIbi"
# ## Data preparation
# + id="o6PhqP1JHRGQ" executionInfo={"status": "ok", "timestamp": 1648015283815, "user_tz": -480, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
# the dataset class
class Groceries(Dataset):
def __init__(self, dir_path, width, height, classes, transforms=None):
self.transforms = transforms
self.dir_path = dir_path
self.height = height
self.width = width
self.classes = classes
# get all the image paths in sorted order
self.image_paths = glob.glob(f"{self.dir_path}/*.jpeg")
self.all_images = [image_path.split('/')[-1] for image_path in self.image_paths]
self.all_images = sorted(self.all_images)
def __getitem__(self, idx):
# capture the image name and the full image path
image_name = self.all_images[idx]
image_path = os.path.join(self.dir_path, image_name)
# read the image
image = cv2.imread(image_path)
# convert BGR to RGB color format
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB).astype(np.float32)
image_resized = cv2.resize(image, (self.width, self.height))
image_resized /= 255.0
# capture the corresponding XML file for getting the annotations
annot_filename = image_name[:-4] + 'xml'
annot_file_path = os.path.join(self.dir_path, annot_filename)
boxes = []
labels = []
tree = et.parse(annot_file_path)
root = tree.getroot()
# get the height and width of the image
image_width = image.shape[1]
image_height = image.shape[0]
# box coordinates for xml files are extracted and corrected for image size given
for member in root.findall('object'):
# map the current object name to `classes` list to get...
# ... the label index and append to `labels` list
labels.append(self.classes.index(member.find('name').text))
# xmin = left corner x-coordinates
xmin = int(member.find('bndbox').find('xmin').text)
# xmax = right corner x-coordinates
xmax = int(member.find('bndbox').find('xmax').text)
# ymin = left corner y-coordinates
ymin = int(member.find('bndbox').find('ymin').text)
# ymax = right corner y-coordinates
ymax = int(member.find('bndbox').find('ymax').text)
# resize the bounding boxes according to the...
# ... desired `width`, `height`
xmin_final = xmin#(xmin/image_width)*self.width
xmax_final = xmax#(xmax/image_width)*self.width
ymin_final = ymin#(ymin/image_height)*self.height
yamx_final = ymax#(ymax/image_height)*self.height
clipped = np.array([xmin_final, ymin_final, xmax_final, yamx_final])
# clipped = np.clip(clipped, a_min=0, a_max=1)
boxes.append(list(clipped))
# bounding box to tensor
boxes = torch.as_tensor(boxes, dtype=torch.float32)
# area of the bounding boxes
area = (boxes[:, 3] - boxes[:, 1]) * (boxes[:, 2] - boxes[:, 0])
# no crowd instances
iscrowd = torch.zeros((boxes.shape[0],), dtype=torch.int64)
# labels to tensor
labels = torch.as_tensor(labels, dtype=torch.int64)
# prepare the final `target` dictionary
target = {}
target["boxes"] = boxes
target["labels"] = labels
target["area"] = area
target["iscrowd"] = iscrowd
image_id = torch.tensor([idx])
target["image_id"] = image_id
# apply the image transforms
if self.transforms:
sample = self.transforms(image = image_resized,
bboxes = target['boxes'],
labels = labels)
image_resized = sample['image']
target['boxes'] = torch.Tensor(sample['bboxes'])
return image_resized, target
def __len__(self):
return len(self.all_images)
# + colab={"base_uri": "https://localhost:8080/"} id="-LEO51qnHpQG" executionInfo={"status": "ok", "timestamp": 1648015283816, "user_tz": -480, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}} outputId="a9be59e8-e8be-4f33-c56a-387c683fd296"
# prepare the final datasets and data loaders
train_dataset = Groceries(TRAIN_DIR, RESIZE_TO, RESIZE_TO, CLASSES, get_train_transform())
valid_dataset = Groceries(VALID_DIR, RESIZE_TO, RESIZE_TO, CLASSES, get_valid_transform())
train_loader = DataLoader(
train_dataset,
batch_size=BATCH_SIZE,
shuffle=True,
num_workers=0,
collate_fn=collate_fn
)
valid_loader = DataLoader(
valid_dataset,
batch_size=BATCH_SIZE,
shuffle=False,
num_workers=0,
collate_fn=collate_fn
)
print(f"Number of training samples: {len(train_dataset)}")
print(f"Number of validation samples: {len(valid_dataset)}\n")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Pkzr0Fw7Hssy" executionInfo={"status": "ok", "timestamp": 1648015289662, "user_tz": -480, "elapsed": 1037, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}} outputId="e03053d2-15ea-4bbc-db67-e7a6449ae340"
# execute datasets.py using Python command from Terminal...
# ... to visualize sample images
# USAGE: python datasets.py
# if __name__ == '__main__':
from google.colab.patches import cv2_imshow
# sanity check of the Dataset pipeline with sample visualization
dataset = Groceries(
TRAIN_DIR, RESIZE_TO, RESIZE_TO, CLASSES
)
print(f"Number of training images: {len(dataset)}")
# function to visualize a single sample
def visualize_sample(image, target):
box = target['boxes'][0]
label = '0'
# print(target['labels'])
label = CLASSES[target['labels'][0]]
cv2.rectangle(
image,
(int(box[0]), int(box[1])), (int(box[2]), int(box[3])),
(0, 255, 0), 2
)
cv2.putText(
image, label, (int(box[0]), int(box[1]-5)),
cv2.FONT_HERSHEY_SIMPLEX, 0.7, (0, 0, 255), 2
)
cv2_imshow(image)
cv2.waitKey(0)
NUM_SAMPLES_TO_VISUALIZE = 5
for i in range(NUM_SAMPLES_TO_VISUALIZE):
image, target = dataset[i]
visualize_sample(image, target)
# + [markdown] id="ktvrxbNdIqqF"
# ## Training
# + id="iyHaoxc4Iug3" executionInfo={"status": "ok", "timestamp": 1648015293814, "user_tz": -480, "elapsed": 854, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
def create_model(num_classes):
# load Faster RCNN pre-trained model
model = torchvision.models.detection.fasterrcnn_resnet50_fpn(pretrained=True)
# get the number of input features
in_features = model.roi_heads.box_predictor.cls_score.in_features
# define a new head for the detector with required number of classes
model.roi_heads.box_predictor = FastRCNNPredictor(in_features, num_classes)
return model
# + id="49_q3zYmI8jb" executionInfo={"status": "ok", "timestamp": 1648015294248, "user_tz": -480, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
plt.style.use('ggplot')
# + id="EJzotQPcJIZD" executionInfo={"status": "ok", "timestamp": 1648015296242, "user_tz": -480, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
# function for running training iterations
def train(train_data_loader, model):
print('Training')
global train_itr
global train_loss_list
# initialize tqdm progress bar
prog_bar = tqdm(train_data_loader, total=len(train_data_loader))
for i, data in enumerate(prog_bar):
optimizer.zero_grad()
images, targets = data
print(i)
images = list(image.to(DEVICE) for image in images)
targets = [{k: v.to(DEVICE) for k, v in t.items()} for t in targets]
loss_dict = model(images, targets)
losses = sum(loss for loss in loss_dict.values())
loss_value = losses.item()
train_loss_list.append(loss_value)
train_loss_hist.send(loss_value)
losses.backward()
optimizer.step()
train_itr += 1
# update the loss value beside the progress bar for each iteration
prog_bar.set_description(desc=f"Loss: {loss_value:.4f}")
return train_loss_list
# + id="HYEiQAsVJNUN" executionInfo={"status": "ok", "timestamp": 1648015298021, "user_tz": -480, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}}
# function for running validation iterations
def validate(valid_data_loader, model):
print('Validating')
global val_itr
global val_loss_list
# initialize tqdm progress bar
prog_bar = tqdm(valid_data_loader, total=len(valid_data_loader))
for i, data in enumerate(prog_bar):
images, targets = data
images = list(image.to(DEVICE) for image in images)
targets = [{k: v.to(DEVICE) for k, v in t.items()} for t in targets]
with torch.no_grad():
loss_dict = model(images, targets)
losses = sum(loss for loss in loss_dict.values())
loss_value = losses.item()
val_loss_list.append(loss_value)
val_loss_hist.send(loss_value)
val_itr += 1
# update the loss value beside the progress bar for each iteration
prog_bar.set_description(desc=f"Loss: {loss_value:.4f}")
return val_loss_list
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["445f60593de449278d38259bda128f24", "b168c91ccc104d42a85f882fe5e8cbf5", "fb5ce8cf271d4859bdf1bc504f147b6e", "4c859f04f06d418caed4cc60b21a3dc7", "6bf5397a11aa44718c3b8ac5c90d6055", "aed744a993d042fa9d980dc606ff290d", "b8c2a21ec73c4abaa6d3a85b03db9c0b", "425212021ed34716a7916dd62b0df399", "<KEY>", "577111dd1fdf4811883dae30364e53fa", "05c8d1cd97f34808927fcc35e442de7b", "e395a5cf54704ada8eaf7863501f5ee6", "<KEY>", "<KEY>", "8eed87c8d4ad44faa899801bbca355ac", "c37ac6deb86d4fe7a5c5d3f5ac9e2233", "fb7e1d93006a4fc8bed7f6a2c512af77", "5011a225c5ee49c6a33167035ebb6dab", "<KEY>", "<KEY>", "4ed129d56b2d47868a0940dd4210ba53", "<KEY>"]} id="on3T_fUPJSED" executionInfo={"status": "error", "timestamp": 1648015312957, "user_tz": -480, "elapsed": 14244, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04915486098817437213"}} outputId="3e06808d-98c7-442b-9884-99c0f2ae5d68"
# initialize the model and move to the computation device
model = create_model(num_classes=NUM_CLASSES)
model = model.to(DEVICE)
# get the model parameters
params = [p for p in model.parameters() if p.requires_grad]
# define the optimizer
optimizer = torch.optim.SGD(params, lr=0.001, momentum=0.9, weight_decay=0.0005)
# initialize the Averager class
train_loss_hist = Averager()
val_loss_hist = Averager()
train_itr = 1
val_itr = 1
# train and validation loss lists to store loss values of all...
# ... iterations till ena and plot graphs for all iterations
train_loss_list = []
val_loss_list = []
# name to save the trained model with
MODEL_NAME = 'model'
# whether to show transformed images from data loader or not
if VISUALIZE_TRANSFORMED_IMAGES:
show_tranformed_image(train_loader)
# start the training epochs
for epoch in range(NUM_EPOCHS):
print(f"\nEPOCH {epoch+1} of {NUM_EPOCHS}")
# reset the training and validation loss histories for the current epoch
train_loss_hist.reset()
val_loss_hist.reset()
# create two subplots, one for each, training and validation
figure_1, train_ax = plt.subplots()
figure_2, valid_ax = plt.subplots()
# start timer and carry out training and validation
start = time.time()
train_loss = train(train_loader, model)
val_loss = validate(valid_loader, model)
print(f"Epoch #{epoch} train loss: {train_loss_hist.value:.3f}")
print(f"Epoch #{epoch} validation loss: {val_loss_hist.value:.3f}")
end = time.time()
print(f"Took {((end - start) / 60):.3f} minutes for epoch {epoch}")
if (epoch+1) % SAVE_MODEL_EPOCH == 0: # save model after every n epochs
torch.save(model.state_dict(), f"{OUT_DIR}/model{epoch+1}.pth")
print('SAVING MODEL COMPLETE...\n')
if (epoch+1) % SAVE_PLOTS_EPOCH == 0: # save loss plots after n epochs
train_ax.plot(train_loss, color='blue')
train_ax.set_xlabel('iterations')
train_ax.set_ylabel('train loss')
valid_ax.plot(val_loss, color='red')
valid_ax.set_xlabel('iterations')
valid_ax.set_ylabel('validation loss')
figure_1.savefig(f"{OUT_DIR}/train_loss_{epoch+1}.png")
figure_2.savefig(f"{OUT_DIR}/valid_loss_{epoch+1}.png")
print('SAVING PLOTS COMPLETE...')
if (epoch+1) == NUM_EPOCHS: # save loss plots and model once at the end
train_ax.plot(train_loss, color='blue')
train_ax.set_xlabel('iterations')
train_ax.set_ylabel('train loss')
valid_ax.plot(val_loss, color='red')
valid_ax.set_xlabel('iterations')
valid_ax.set_ylabel('validation loss')
figure_1.savefig(f"{OUT_DIR}/train_loss_{epoch+1}.png")
figure_2.savefig(f"{OUT_DIR}/valid_loss_{epoch+1}.png")
torch.save(model.state_dict(), f"{OUT_DIR}/model{epoch+1}.pth")
plt.close('all')
# sleep for 5 seconds after each epoch
time.sleep(5)
# + id="Py4el73-Jkko"
| source/Machine Learning/model/Object Detection Fast R-CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
a = np.array([1,2,3,4])
print(a)
# +
import time
a = np.random.rand(1000000) # one million
b = np.random.rand(1000000)
tic = time.time()
c = np.dot(a,b)
toc = time.time()
print(c)
print("Vectorized version:", str(1000*(toc-tic)) + "ms")# 逗号连接
c =0
tic = time.time()
for i in range(1000000):
c+=a[i]*b[i]
toc=time.time()
print(c)
print("For loop" + str(1000*(toc-tic))+"ms")# 加号连接
# 差别300倍
# -
| 01神经网络和深度学习/2.11 Vectorization demo向量化.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Open-Loop Evaluation
#
# In this notebook you are going to evaluate a CNN-based policy to control the SDV with a protocol named *open-loop* evaluation.
#
# **Note: this notebook assumes you've already run the [training notebook](./train.ipynb) and stored your model successfully.**
#
# ## What is open-loop evaluation?
# In open-loop evaluation we evaluate our model prediction as we follow the annotated ground truth.
#
# In each frame, we compare the predictions of our model against the annotated ground truth. This can be done with different metrics, and we will see a few of them in the following.
#
# **Regardless of the metric used, this evaluation protocol doesn't modify the future locations according to our predictions.**
#
# 
#
#
# ## What can we use open-loop evaluation for?
# Open-loop evaluation can be used for a frame by frame comparison between the expert and the policy. This is extremely useful for debugging the model behaviours and investigate outlier predictions in specific situations (e.g. at crossings or unprotected turn).
#
# ## Is open-loop evaluation enough?
# Regardless of the quality of the open-loop results, **this evaluation is not enough** to ensure your model will be able to actually drive on the road (that's where we all want to go in the end). If your model is not in full control of the SDV, you can't really say it will work once the annotated trajectory won't be available anymore.
#
# Before drawing conclusions on our model we must test it when it is in full control of the SDV, in a setting called **closed-loop**. You can try just that in our [dedicated closed-loop evaluation notebook](./closed_loop_test.ipynb)
# +
from tempfile import gettempdir
import matplotlib.pyplot as plt
import numpy as np
import torch
from torch import nn, optim
from torch.utils.data import DataLoader
from torch.utils.data.dataloader import default_collate
from tqdm import tqdm
from l5kit.configs import load_config_data
from l5kit.data import LocalDataManager, ChunkedDataset
from l5kit.dataset import EgoDataset
from l5kit.rasterization import build_rasterizer
from l5kit.geometry import transform_points, angular_distance
from l5kit.visualization import TARGET_POINTS_COLOR, PREDICTED_POINTS_COLOR, draw_trajectory
from l5kit.kinematic import AckermanPerturbation
from l5kit.random import GaussianRandomGenerator
import os
# -
# ## Prepare data path and load cfg
#
# By setting the `L5KIT_DATA_FOLDER` variable, we can point the script to the folder where the data lies.
#
# Then, we load our config file with relative paths and other configurations (rasteriser, training params...).
# set env variable for data
os.environ["L5KIT_DATA_FOLDER"] = "/tmp/l5kit_data"
dm = LocalDataManager(None)
# get config
cfg = load_config_data("./config.yaml")
# ## Load the model
model_path = "/tmp/planning_model.pt"
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model = torch.load(model_path).to(device)
model = model.eval()
# ## Load the evaluation data
# This is almost the exact same code you've already seen in the [training notebook](./train.ipynb). Apart from the different dataset we load, the biggest difference is that **we don't perturb our data here**.
#
# When performing evaluation we're interested in knowing the performance on the annotated data, not on perturbed one.
# ===== INIT DATASET
eval_cfg = cfg["val_data_loader"]
rasterizer = build_rasterizer(cfg, dm)
eval_zarr = ChunkedDataset(dm.require(eval_cfg["key"])).open()
eval_dataset = EgoDataset(cfg, eval_zarr, rasterizer)
eval_dataloader = DataLoader(eval_dataset, shuffle=eval_cfg["shuffle"], batch_size=eval_cfg["batch_size"],
num_workers=eval_cfg["num_workers"])
print(eval_dataset)
# # Evaluation loop
#
# Here, we loop through the data and store predicted and annotated trajectories (positions + yaws).
#
# Note: we're not taking into account availability here. We acknowledge this can reflect in a lower score.
# +
# ==== EVAL LOOP
position_preds = []
yaw_preds = []
position_gts = []
yaw_gts = []
torch.set_grad_enabled(False)
for idx_data, data in enumerate(tqdm(eval_dataloader)):
data = {k: v.to(device) for k, v in data.items()}
result = model(data)
position_preds.append(result["positions"].detach().cpu().numpy())
yaw_preds.append(result["yaws"].detach().cpu().numpy())
position_gts.append(data["target_positions"].detach().cpu().numpy())
yaw_gts.append(data["target_yaws"].detach().cpu().numpy())
if idx_data == 10:
break
position_preds = np.concatenate(position_preds)
yaw_preds = np.concatenate(yaw_preds)
position_gts = np.concatenate(position_gts)
yaw_gts = np.concatenate(yaw_gts)
# -
# # Quantitative evaluation
# ## ADE, FDE and angle distance
#
# ### Positional displacement
# [Average Displacement Error (ADE) and Final Displacement Error (FDE)](https://en.wikipedia.org/wiki/Mean_squared_displacement) are standard metrics used to evaluate future predictions for AVs.
#
# We can compute them by comparing predicted and annotated positions, which we have stored in the previous cell.
# Additionally, we can plot histograms of their distributions across samples to better capture the variance of our error.
#
# ### Angle displacement
#
# For the yaw, we can use the Minimum Angle Distance to check the error. Again, we can plot a histogram to inspect the error distribution.
#
# Although yaw may seem redundant here, it's actually crucial to fully control the SDV. We'll use it extensively in the closed-loop evaluation notebook.
# +
pos_errors = np.linalg.norm(position_preds - position_gts, axis=-1)
# DISPLACEMENT AT T
plt.plot(np.arange(pos_errors.shape[1]), pos_errors.mean(0), label="Displacement error at T")
plt.legend()
plt.show()
# ADE HIST
plt.hist(pos_errors.mean(-1), bins=100, label="ADE Histogram")
plt.legend()
plt.show()
# FDE HIST
plt.hist(pos_errors[:,-1], bins=100, label="FDE Histogram")
plt.legend()
plt.show()
angle_errors = angular_distance(yaw_preds, yaw_gts).squeeze()
# ANGLE ERROR AT T
plt.plot(np.arange(angle_errors.shape[1]), angle_errors.mean(0), label="Angle error at T")
plt.legend()
plt.show()
# ANGLE ERROR HIST
plt.hist(angle_errors.mean(-1), bins=100, label="Angle Error Histogram")
plt.legend()
plt.show()
# -
# # Qualitative evaluation
# ## Visualise results
# We can also visualise some images with predicted and annotated trajectories using L5Kit visualisation features.
#
# In this example, we draw 20 images from our dataset and we visualise predicted and annotated trajectories on top of them.
for frame_number in range(0, len(eval_dataset), len(eval_dataset) // 20):
data = eval_dataloader.dataset[frame_number]
data_batch = default_collate([data])
data_batch = {k: v.to(device) for k, v in data_batch.items()}
result = model(data_batch)
predicted_positions = result["positions"].detach().cpu().numpy().squeeze()
im_ego = rasterizer.to_rgb(data["image"].transpose(1, 2, 0))
target_positions = data["target_positions"]
predicted_positions = transform_points(predicted_positions, data["raster_from_agent"])
target_positions = transform_points(target_positions, data["raster_from_agent"])
draw_trajectory(im_ego, predicted_positions, PREDICTED_POINTS_COLOR)
draw_trajectory(im_ego, target_positions, TARGET_POINTS_COLOR)
plt.imshow(im_ego)
plt.axis("off")
plt.show()
# ## Visualise the open-loop
#
# To visualise the open loop we can just repeat the same operations for the consecutive frames.
#
# In this example, we show the first 200 frames for our dataset, plotting predicted and annotated trajectories.
#
# **We want to stress this out again: this is an open loop evaluation, we are NOT controlling the AV with our predictions**
# +
from IPython.display import display, clear_output
import PIL
for frame_number in range(200):
data = eval_dataloader.dataset[frame_number]
data_batch = default_collate([data])
data_batch = {k: v.to(device) for k, v in data_batch.items()}
result = model(data_batch)
predicted_positions = result["positions"].detach().cpu().numpy().squeeze()
predicted_positions = transform_points(predicted_positions, data["raster_from_agent"])
target_positions = transform_points(data["target_positions"], data["raster_from_agent"])
im_ego = rasterizer.to_rgb(data["image"].transpose(1, 2, 0))
draw_trajectory(im_ego, target_positions, TARGET_POINTS_COLOR)
draw_trajectory(im_ego, predicted_positions, PREDICTED_POINTS_COLOR)
clear_output(wait=True)
display(PIL.Image.fromarray(im_ego))
# -
# # Pre-trained model results
#
# We include here the open-loop results of one scene using one of our pre-trained model. The predicted trajectory is well overlapped with the annotated one.
#
# 
| examples/planning/open_loop_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/avinsit123/Tweetopedia_Collab_Notebooks/blob/master/Query_based_Convolutional_Sentiment_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dtYwHmWtsT96" colab_type="text"
# # Convolutional Sentiment Analysis
# CNNs are type of neural architecture which are used to do image classification.They involve passing a convolutional filter through the image and rapidly downscaling these images until the size decreases to a portion where we can connect it a fully connected network.
#
# The intuitive idea behind learning the weights is that your convolutional layers act like *feature extractors*, extracting parts of the image that are most important for your CNN's goal, e.g. if using a CNN to detect faces in an image, the CNN may be looking for features such as the existance of a nose, mouth or a pair of eyes in the image.
#
# So why use CNNs on text? In the same way that a 3x3 filter can look over a patch of an image, a 1x2 filter can look over a 2 sequential words in a piece of text, i.e. a bi-gram. In the previous tutorial we looked at the FastText model which used bi-grams by explicitly adding them to the end of a text, in this CNN model we will instead use multiple filters of different sizes which will look at the bi-grams (a 1x2 filter), tri-grams (a 1x3 filter) and n-grams (a 1x$n$ filter) within the text.
#
# The intuition here is that the appearance of certain bi-grams, tri-grams and n-grams within the review will be a good indication of the final sentiment.First let us install the required libraries.
# + id="kPipOzGIsdMY" colab_type="code" outputId="65d1b821-8ab6-49c0-81c1-3216469a93fc" colab={"base_uri": "https://localhost:8080/", "height": 441}
# !pip install torch
# !pip install torchtext
# + [markdown] id="vIIXpmBgsT9_" colab_type="text"
# ## Preparing Data
#
# Let us first prepare the data.
# + id="ZeO7kR-osT-D" colab_type="code" outputId="a849f01c-b3cb-4378-efb8-9e6087e01eff" colab={"base_uri": "https://localhost:8080/", "height": 613}
import torch
from torchtext import data
from torchtext import datasets
import random
SEED = 1234
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
TEXT = data.Field(tokenize='spacy')
LABEL = data.LabelField(dtype=torch.float)
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
train_data, valid_data = train_data.split(random_state=random.seed(SEED))
# + [markdown] id="AuWGPo87sT-K" colab_type="text"
# Build the vocab and load the pre-trained word embeddings.
#
# Download Glove Embeddings where each word is representated by 100 Degree Vectors.
# + id="vPL_mGSbsT-M" colab_type="code" outputId="439c1b1f-5d35-4b13-89bd-3ce18b516e95" colab={"base_uri": "https://localhost:8080/", "height": 1020}
TEXT.build_vocab(train_data, max_size=25000, vectors="glove.6B.100d")
LABEL.build_vocab(train_data)
# + [markdown] id="eZlV2a9QsT-T" colab_type="text"
# As before, we create the iterators.
# + id="Aq2pHcQgsT-W" colab_type="code" colab={}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size=BATCH_SIZE,
device=device)
# + [markdown] id="lHoJs_qMsT-e" colab_type="text"
# ## Build the Model
#
# Now to build our model.
#
# The first major hurdle is visualizing how CNNs are used for text. Images are typically 2 dimensional (we'll ignore the fact that there is a third "colour" dimension for now) whereas text is 1 dimensional. However, we know that the first step in almost all of our previous tutorials (and pretty much all NLP pipelines) is converting the words into word embeddings. This is how we can visualize our words in 2 dimensions, each word along one axis and the elements of vectors aross the other dimension. Consider the 2 dimensional representation of the embedded sentence below:
#
# 
#
# We can then use a filter that is **[n x emb_dim]**. This will cover $n$ sequential words entirely, as their width will be `emb_dim` dimensions. Consider the image below, with our word vectors are represented in green. Here we have 4 words with 5 dimensional embeddings, creating a [4x5] "image" tensor. A filter that covers two words at a time (i.e. bi-grams) will be **[2x5]** filter, shown in yellow, and each element of the filter with have a _weight_ associated with it. The output of this filter (shown in red) will be a single real number that is the weighted sum of all elements covered by the filter.
#
# 
#
# The filter then moves "down" the image (or across the sentence) to cover the next bi-gram and another output (weighted sum) is calculated.
#
# 
#
# Finally, the filter moves down again and the final output for this filter is calculated.
#
# 
#
# In our case (and in the general case where the width of the filter equals the width of the "image"), our output will be a vector with number of elements equal to the height of the image (or lenth of the word) minus the height of the filter plus one, $4-2+1=3$ in this case.
#
# This example showed how to calculate the output of one filter. Our model (and pretty much all CNNs) will have lots of these filters. The idea is that each filter will learn a different feature to extract. In the scenario of analysing text, we are hoping each of the **[2 x emb_dim]** filters will be looking for the occurence of different bi-grams.
#
# In our model, we will also have different sizes of filters, heights of 3, 4 and 5, with 100 of each of them. The intuition is that we will be looking for the occurence of different tri-grams, 4-grams and 5-grams that are relevant for analysing sentiment of movie reviews.
#
# The next step in our model is to use *pooling* (specifically *max pooling*) on the output of the convolutional layers. This is similar to the FastText model where we performed the average over each of the word vectors, implemented by the `F.avg_pool2d` function, however instead of taking the average over a dimension, we are taking the maximum value over a dimension. Below an example of taking the maximum value (0.9) from the output of the convolutional layer on the example sentence (not shown in the activation function applied to the output of the convolutions).
#
# 
#
# The idea here is that the maximum value is the "most important" feature for determining the sentiment of the review, which corresponds to the "most important" n-gram within the review. How do we know what the "most important" n-gram is? Luckily, we don't have to! Through backpropagation, the weights of the filters are changed so that whenever certain n-grams that are highly indicative of the sentiment are seen, the output of the filter is a "high" value. This "high" value then passes through the max pooling layer if it is the maximum value in the output.
#
# As our model has 100 filters of 3 different sizes, that means we have 300 different n-grams the model thinks are important. We concatenate these together into a single vector and pass them through a linear layer to predict the sentiment. We can think of the weights of this linear layer as "weighting up the evidence" from each of the 300 n-grams and making a final decision.
#
# ### Implementation Details
#
# We implement the convolutional layers with `nn.Conv2d`. The `in_channels` argument is the number of "channels" in your image going into the convolutional layer. In actual images this is usually 3 (one channel for each of the red, blue and green channels), however when using text we only have a single channel, the text itself. The `out_channels` is the number of filters and the `kernel_size` is the size of the filters. Each of our `kernel_size`s is going to be **[n x emb_dim]** where $n$ is the size of the n-grams.
#
# In PyTorch, RNNs want the input with the batch dimension second, whereas CNNs want the batch dimension first. Thus, the first thing we do to our input is `permute` it to make it the correct shape. We then pass the sentence through an embedding layer to get our embeddings. The second dimension of the input into a `nn.Conv2d` layer must be the channel dimension. As text technically does not have a channel dimension, we `unsqueeze` our tensor to create one. This matches with our `in_channels=1` in the initialization of our convolutional layers.
#
# We then pass the tensors through the convolutional and pooling layers, using the `ReLU` activation function after the convolutional layers. Another nice feature of the pooling layers is that they handle sentences of different lengths. The size of the output of the convolutional layer is dependent on the size of the input to it, and different batches contain sentences of different lengths. Without the max pooling layer the input to our linear layer would depend on the size of the input sentence (not what we want). One option to rectify this would be to trim/pad all sentences to the same length, however with the max pooling layer we always know the input to the linear layer will be the total number of filters. **Note**: there an exception to this if your sentence(s) are shorter than the largest filter used. You will then have to pad your sentences to the length of the largest filter. In the IMDb data there are no reviews only 5 words long so we don't have to worry about that, but you will if you are using your own data.
#
# Finally, we perform dropout on the concatenated filter outputs and then pass them through a linear layer to make our predictions.
# + id="wB40Qc6GsT-h" colab_type="code" colab={}
import torch.nn as nn
import torch.nn.functional as F
class CNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, n_filters, filter_sizes, output_dim, dropout):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim)
self.conv_0 = nn.Conv2d(in_channels=1, out_channels=n_filters, kernel_size=(filter_sizes[0],embedding_dim))
self.conv_1 = nn.Conv2d(in_channels=1, out_channels=n_filters, kernel_size=(filter_sizes[1],embedding_dim))
self.conv_2 = nn.Conv2d(in_channels=1, out_channels=n_filters, kernel_size=(filter_sizes[2],embedding_dim))
self.fc = nn.Linear(len(filter_sizes)*n_filters, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
#x = [sent len, batch size]
x = x.permute(1, 0)
#x = [batch size, sent len]
embedded = self.embedding(x)
#embedded = [batch size, sent len, emb dim]
embedded = embedded.unsqueeze(1)
#embedded = [batch size, 1, sent len, emb dim]
conved_0 = F.relu(self.conv_0(embedded).squeeze(3))
conved_1 = F.relu(self.conv_1(embedded).squeeze(3))
conved_2 = F.relu(self.conv_2(embedded).squeeze(3))
#conv_n = [batch size, n_filters, sent len - filter_sizes[n]]
pooled_0 = F.max_pool1d(conved_0, conved_0.shape[2]).squeeze(2)
pooled_1 = F.max_pool1d(conved_1, conved_1.shape[2]).squeeze(2)
pooled_2 = F.max_pool1d(conved_2, conved_2.shape[2]).squeeze(2)
#pooled_n = [batch size, n_filters]
cat = self.dropout(torch.cat((pooled_0, pooled_1, pooled_2), dim=1))
#cat = [batch size, n_filters * len(filter_sizes)]
return self.fc(cat)
# + [markdown] id="OOR6f-F8sT-q" colab_type="text"
# Currently the `CNN` model can only use 3 different sized filters, but we can actually improve the code of our model to make it more generic and take any number of filters.
#
# We do this by placing all of our convolutional layers in a `nn.ModuleList`, a function used to hold a list of PyTorch `nn.Module`s. If we simply used a standard Python list, the modules within the list cannot be "seen" by any modules outside the list which will cause us some errors.
#
# We can now pass an arbitrary sized list of filter sizes and the list comprehension will create a convolutional layer for each of them. Then, in the `forward` method we iterate through the list applying each convolutional layer to get a list of convolutional outputs, which we also feed through the max pooling in a list comprehension before concatenating together and passing through the dropout and linear layers.
# + id="6cXcDMtKsT-t" colab_type="code" colab={}
class CNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, n_filters, filter_sizes, output_dim, dropout):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim)
self.convs = nn.ModuleList([nn.Conv2d(in_channels=1, out_channels=n_filters, kernel_size=(fs,embedding_dim)) for fs in filter_sizes])
self.fc = nn.Linear(len(filter_sizes)*n_filters, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
#x = [sent len, batch size]
x = x.permute(1, 0)
#x = [batch size, sent len]
embedded = self.embedding(x)
#embedded = [batch size, sent len, emb dim]
embedded = embedded.unsqueeze(1)
#embedded = [batch size, 1, sent len, emb dim]
conved = [F.relu(conv(embedded)).squeeze(3) for conv in self.convs]
#conv_n = [batch size, n_filters, sent len - filter_sizes[n]]
pooled = [F.max_pool1d(conv, conv.shape[2]).squeeze(2) for conv in conved]
#pooled_n = [batch size, n_filters]
cat = self.dropout(torch.cat(pooled, dim=1))
#cat = [batch size, n_filters * len(filter_sizes)]
return self.fc(cat)
# + [markdown] id="aImBe7Br_4GQ" colab_type="text"
# Below are listed some of the parameters which you can change .Their names are actually implied.Change these parameters and run the training cell.
# + id="66rHuY5I_3h0" colab_type="code" colab={}
BATCH_SIZE = 64
N_EPOCHS = 5
INPUT_DIM = len(TEXT.vocab) #Do not change this
EMBEDDING_DIM = 100 #Embedding Dims for Glove vectors
N_FILTERS = 100
FILTER_SIZES = [3,4,5] #Size of Convolutional Filter
OUTPUT_DIM = 1 #Output Dimensions = 1
DROPOUT = 0.5
device = [ "cuda" if torch.cuda.is_available() else "cpu"]
model_name = "hateorlove" #Name of check point file
# + [markdown] id="sJfLOpx_sT-z" colab_type="text"
# We create an instance of our `CNN` class.
# + id="fs7muZRHsT-1" colab_type="code" outputId="e32bc4e6-1eaf-4d87-db60-417a1f86ac3e" colab={"base_uri": "https://localhost:8080/", "height": 94}
model = CNN(INPUT_DIM, EMBEDDING_DIM, N_FILTERS, FILTER_SIZES, OUTPUT_DIM, DROPOUT)
# + [markdown] id="QhnCINB7sT-4" colab_type="text"
# And load the pre-trained embeddings
# + id="UOfxgtfjsT-6" colab_type="code" outputId="acab8200-f4d9-4b91-d56d-f9e0dff35976" colab={"base_uri": "https://localhost:8080/", "height": 193}
pretrained_embeddings = TEXT.vocab.vectors
model.embedding.weight.data.copy_(pretrained_embeddings)
# + [markdown] id="A4nKo_U_sT_B" colab_type="text"
# ## Train the Model
# + [markdown] id="TyvYfhO9sT_D" colab_type="text"
# We initialize the optimizer, loss function (criterion) and place the model and criterion on the GPU (if available)
# + id="9nzqeSbFsT_F" colab_type="code" colab={}
import torch.optim as optim
optimizer = optim.Adam(model.parameters())
criterion = nn.BCEWithLogitsLoss()
model = model.to(device)
criterion = criterion.to(device)
# + [markdown] id="-xXtWsQusT_I" colab_type="text"
# We implement the function to calculate accuracy...
# + id="7wvO43CasT_J" colab_type="code" colab={}
def binary_accuracy(preds, y):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
#round predictions to the closest integer
rounded_preds = torch.round(torch.sigmoid(preds))
correct = (rounded_preds == y).float() #convert into float for division
acc = correct.sum()/len(correct)
return acc
# + [markdown] id="HtP5iRRSsT_M" colab_type="text"
# We define a function for training our model...
#
# **Note**: as we are using dropout again, we must remember to use `model.train()` to ensure the dropout is "turned on" while training.
# + id="plKlzMNosT_N" colab_type="code" colab={}
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + [markdown] id="Ew3fFD4jsT_Q" colab_type="text"
# We define a function for testing our model...
#
# **Note**: again, as we are now using dropout, we must remember to use `model.eval()` to ensure the dropout is "turned off" while evaluating.
# + id="7LGqRQTssT_Q" colab_type="code" colab={}
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + [markdown] id="A3qcg1VnsT_T" colab_type="text"
# Finally, we train our model...
# + id="Q3q6J2PDsT_U" colab_type="code" outputId="618de873-5313-4dcb-fa2d-455a1a8f7d4b" colab={"base_uri": "https://localhost:8080/", "height": 214}
for epoch in range(N_EPOCHS):
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
print(f'| Epoch: {epoch+1:02} | Train Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}% | Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}% |')
# + [markdown] id="Jy_yIr5CsT_Y" colab_type="text"
# ...and get our best test accuracy yet!
# + id="jDrUFScZsT_Z" colab_type="code" outputId="ba58d843-f55a-474f-eb15-77c562e55ff1" colab={"base_uri": "https://localhost:8080/", "height": 114}
test_loss, test_acc = evaluate(model, test_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}% |')
# + [markdown] id="ugE5RXiMsT_f" colab_type="text"
# ## User Input
#
# And again, as a sanity check we can check some input sentences
#
# **Note**: As mentioned in the implementation details, the input sentence has to be at least as long as the largest filter height used. We modify our `predict_sentiment` function to also accept a minimum length argument. If the tokenized input sentence is less than `min_len` tokens, we append padding tokens (`<pad>`) to make it `min_len` tokens.
# + id="rtk2IRh2sT_g" colab_type="code" colab={}
import spacy
nlp = spacy.load('en')
def predict_sentiment(sentence, min_len=5):
tokenized = [tok.text for tok in nlp.tokenizer(sentence)]
print(tokenized)
if len(tokenized) < min_len:
tokenized += ['<pad>'] * (min_len - len(tokenized))
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(1)
prediction = torch.sigmoid(model(tensor))
return prediction.item()
# + [markdown] id="J3oA-tRtsT_i" colab_type="text"
# An example negative review...
# + [markdown] id="XXS-TjddsT_n" colab_type="text"
# An example positive review...
# + id="PdoRTTaVsT_o" colab_type="code" outputId="92782257-3bd7-4c8b-c7bd-98e224e983ee" colab={"base_uri": "https://localhost:8080/", "height": 114}
predict_sentiment("I love how the socially retarded, AKA Trump fanatics, praise this sub-human entity as it plunges our once great nation into utter ruin. I'm not sure if I should laugh or cringe. ")
# + id="y0EM7Ry3yKSq" colab_type="code" colab={}
#Do not Make Alterations to this cell
checkpoint ={
'state_dict' : model.state_dict(),
'stoi' : TEXT.vocab.stoi
}
torch.save(checkpoint,model_name + '.pth')
#Go and Download Checkpoint file from the Side Window
| Query_based_Convolutional_Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment #02: using the standard library
# Today's exercises have to be done with help of the python standard library alone! No external module can be used.
# ## Exercise #02-01: cryptography for <s>dummies</s> Roman Emperors
# A very simple encryption technique is called the [Caesar cipher](http://en.wikipedia.org/wiki/Caesar_cipher). The basic idea is that each letter is replaced by a letter that is a certain number of letters away, so for example if the shift was 2, then A would become C, B would become D, and Z will become B.
#
# **A. Write a function that given a string and a shift, will produce the encrypted string for that shift**. The rules are:
# - you should accept lowercase and uppercase letters, and return letters of the same case
# - spaces or other punctuation characters should not be changed.
#
# There are several ways to reach this result. Just pick the one which makes more sense to you. Then, decrypt the following message, which was encrypted with a shift of 13:
#
# Pbatenghyngvbaf, lbh unir fhpprrqrq va qrpelcgvat gur fgevat.
#
# **B. Now write a decoding script** which, when run in the linux or windows command line, prints the decoded phrase after asking the user to enter a phrase and a shift with which it was encrypted. Test your script on the sentence above.
#
#
# **C. Now try to decrypt this sentence, for which the shift is unknown**:
#
# Gwc uivioml bw nqvl bpm zqopb apqnb.
#
# One way to solve this problem involves human decision, but another way can be fully automated (and implies more work). Pick the one you want!
# ## Exercise #02-02: automated data download
# [SRTM](https://en.wikipedia.org/wiki/Shuttle_Radar_Topography_Mission) is a digital elevation model at ~90 m resolution covering almost the entire globe (up to $\pm$ 60° latitude). The data is organized in 5°x5° tiles. To see a map of the tiles have a look at [this download page](http://srtm.csi.cgiar.org/SELECTION/inputCoord.asp). This tool is nice to use if you know which tile you want, but not very useful if you want more than one tile at different places of the globe.
#
# Fortunately, the entire dataset is available on this server: http://droppr.org/srtm/v4.1/6_5x5_TIFs (click on the link to display the data). The file naming convention is very simple:
# - the first two digits number gives the location of the tile in the longitudes (starting at 180° West)
# - the last two digits gives the location of the tile in the latitudes (starting at 60° North and going southwards)
#
# For example, here are some examples of locations and their associated tile:
# - (-179, 59) -> 'srtm_01_01.zip'
# - (-179, 51) -> 'srtm_01_02.zip'
# - (-174, 54) -> 'srtm_02_02.zip'
#
# And so forth.
#
# **A. Write a script which, given a longitude and a latitude as arguments, downloads the corresponding file** in the current directory. The function should raise an error when the given location is not valid.
#
# *Hints: define "valid" locations first: some are easy to catch, some cannot be caught automatically. Do we really have to deal with those?*
#
# **B.** Extend this script to be a bit more clever: **download the data file only if the file isn't already available in the current directory**.
#
# **C. Optional**: extend this script to be even more clever: given a range of longitudes and latitudes, it should download all the files convering this area. For example, the range 9°W to 18°W and 44°N to 47°N would download 6 files.
# *Note 1: we will have a look at the data later in the course. But If you want to display them you can open them with the [qgis](https://www.qgis.org) software for example.*
#
# *Note 2: at the edge of the tiles (i.e. coordinates -175, 55) the problem might be not well defined because of accuracy errors (we will get back to these). You shouldn't care about this for now and try to get things right for everywhere but at the exact boundaries.*
# ## Tips
# Both scripts are asking the user for an input, but in a different way:
#
# - **for #02-01**, I am asking you to make use of python's [input()](https://docs.python.org/3/library/functions.html#input) function, which asks the user to type in the command line.
# - **for #02-02**, I am asking you to use the script with [command line arguments](https://docs.python.org/3/tutorial/stdlib.html?highlight=sys%20argv#command-line-arguments).
#
# You'll find numerous examples online for both procedures. Both work fine with ipython and/or spyder. For example, here is my script call for the optional exercise #02-02-C:
#
# ```
# # %run download_srtm.py 9 44 18 47
# ```
#
# *Back to the [table of contents](00-Introduction.ipynb#ctoc)*
| notebooks/08-Assignment-02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import cmath
import scipy.integrate as spint
import matplotlib.pyplot as plt
# +
def mthm_vpi(v):
coeff = FYFG *(3/2)*n*(1-e**2)**2
term1 = (np.cos(v - x0)/(e*(-1+e**2)*np.sqrt(1-e**2)))
term2 = (3/4)*J2*(Rp/a)**2* n/(1-e**2)**2 * (2 - (5/2)*(np.sin(i)**2))
period = 11.86 * (a / 5.2)**(3/2) * (M_sol)**(1/2)
term3 = 2*np.pi / period
return coeff*term1 + term2 - term3
def mthm_e(e):
coeff = FYFG *(3/2)*n*(1-e**2)**2
term1 = np.sin(v-x0)/((-1+e**2)*np.sqrt(1-e**2))
return coeff*term1
def yark_acc(x, y, z):
L = L_sol*0.000235 # solar luminosity in au^2 M_sol/yr^3
c = 63197.8 # speed of light in au/yr
r = np.sqrt(x**2 + y**2 + z**2)
A = (R**2 * L_sol)/(4*mast*c)
ax = (A/r**3) * x
ay = (A/r**3) * (0.25*x + y)
az = (A/r**3) * z
return mast*np.sqrt(ax**2 + ay**2 + az**2), np.arctan2(ay,ax)
es = np.linspace(0,1,100)
v_ = np.linspace(0,2*np.pi,100)
rhill = 0.35
G = 4*np.pi**2
Mj = 9.55e-4
J2 = 1.47e-2
Rp = 4.78e-4
a = 0.07
i = 0
L_sol = 1
M_sol = 1
rkm = 10 # moon radius in km
R = rkm*6.68e-9 # converts moon radius to AU
v_ast = (rkm**5)**3 * (4/3) * np.pi # volume of moon in cm^3
m_ast = v_ast * (3/1000) # mass of moon in kg if density ~ 3 g/cm^3
mast = m_ast / 1.989e30 # converts moon mass to M_sol
B = 0
FY, x0 = yark_acc(7.5,0,0)
n = np.sqrt(G*Mj / a**3)
FG = (G*Mj*mast)/(a**2)
FYFG = FY/FG
edata = np.zeros((len(es),len(v_)))
vdata = np.zeros((len(es),len(v_)))
for et in range(len(es)):
e = es[et]
for it in range(len(v_)):
v = v_[it]
m = v_[it]
evals = mthm_e(e)
vvals = mthm_vpi(v)
edata[et,it] = evals
vdata[et,it] = vvals
# -
plt.plot(es,(vdata[40]),'b')
plt.plot(es,(edata[:,0]),'r')
plt.plot(es,np.zeros_like(es),'k',ls='--',alpha=0.5)
#plt.fill_between(es, (idata) - ierrs, (idata) + ierrs, facecolor='b', alpha=0.3,zorder=0)
#plt.fill_between(es, edata - eerrs, edata + eerrs, facecolor='r', alpha=0.3,zorder=0)
plt.legend(labels=[r"<d$\varpi$/dt>","<de/dt>"], fontsize=12, frameon=False,loc='lower right')
#plt.ylim(-1e-7,1.2e-6)
plt.xlabel("e", fontsize=16)
plt.ylabel("Rate of change", fontsize=12)
#plt.text(0,6e-7,r"i = %2.f$^\circ$" %(i*180/np.pi), fontsize=16)
plt.show()
print("Constants:")
print(r"FY / FG = %.5f" %FYFG)
print("n = %1.1f AU / yr" %n)
print("Theta_0 = %1.1f" %x0)
print("i = %1.1f" %i)
print("B = %1.1f" %B)
varpi_dot = (3/2)*J2*(Rp/a)**2* n
ef = (3/2) * n/varpi_dot * FYFG * np.cos(B)
print(ef)
# +
import matplotlib.colors as colors
# Example of making your own norm. Also see matplotlib.colors.
# From <NAME>: This one gives two different linear ramps:
class MidpointNormalize(colors.Normalize):
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
self.midpoint = midpoint
colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# I'm ignoring masked values and all kinds of edge cases to make a
# simple example...
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return np.ma.masked_array(np.interp(value, x, y))
class MidpointNorm(colors.Normalize):
def __init__(self, vmin=-.004, vmax=.004, midpoint=None, clip=False):
self.midpoint = midpoint
colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# I'm ignoring masked values and all kinds of edge cases to make a
# simple example...
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return np.ma.masked_array(np.interp(value, x, y))
#####
# +
fig, ax = plt.subplots(1,2,figsize=(16,7))
speed = np.sqrt(vdata**2 + edata**2)
for i in range(2):
ax[i].set_xlabel(r"$\varpi$",fontsize=16)
ax[i].set_ylabel("e",fontsize=16)
ax[i].set_xticks([0,np.pi,2*np.pi])
ax[i].set_xticklabels([0,r"$\pi$",r"2$\pi$"])
ax[i].set_xlim(0,2*np.pi)
ax[i].streamplot(v_,es,vdata,edata,color='k')
#ax[i].hlines(ef,0,2*np.pi,color='k',ls="--")
ax[0].set_title(r"Colormap shows <d$\varpi$/dt>",fontsize=18)
ax[1].set_title(r"Colormap shows <de/dt>",fontsize=18)
ax0 = ax[0].pcolor(v_,es,vdata,cmap='seismic',norm=MidpointNormalize(midpoint=0))
ax1 = ax[1].pcolor(v_,es,edata,cmap='seismic',norm=MidpointNormalize(midpoint=0))
fig.colorbar(ax0,ax=ax[0])
fig.colorbar(ax1,ax=ax[1])
fig.show()
fig.tight_layout()
# -
| 5-Irregs/Integral Check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Theory of design
# ## Dynamic modeling and design of basic pole-placement controller
# Begin with the Lagrangian.
# $$
# L=T-V=K_e -P_e
# \tag{1}
# $$
# Determine the kinetic energies of the cart and pendulum pop, which is treated as a point mass at the end of a mass-less rod. The pendulum bob's position can be treated as such:
# $$
# p_x=x+lsin\theta \\
# p_y=lcos\theta
# \tag{2}
# $$
# This has corresponding velocities:
# $$
# v_px=\dot x+l\dot\theta cos\theta \\
# v_py=-l\dot\theta sin\theta
# \tag{3}
# $$
# Their squared magnitude:
# $$
# v_p^2=(\dot x+l\dot\theta cos\theta)^2+(-l\dot\theta sin\theta)^2 \\
# = \dot x^2 + 2 \dot xl\dot\theta cos\theta+l^2\dot\theta^2cos^2\theta+l^2 \dot\theta^2sin^2\theta \\
# =\dot x^2+2\dot xl\dot\theta cos\theta +l^2\dot\theta^2
# \tag{4}
# $$
# Using this for the pendulum's kinetic energy
# $$
# KE_p=\frac{1}{2}m\dot (\dot x^2+2\dot xl\dot\theta cos\theta +l^2\dot\theta^2) \\
# KE_c=\frac{1}{2}M\dot x^2
# $$
# Only the pendulum contributes to the potential energy of the system, and is defined as:
# $$
# PE_p=mgl cos\theta
# $$
# Returning to the Lagrangian:
# $$
# L = \frac{1}{2}M\dot x^2+\frac{1}{2}m\dot (\dot x^2+2\dot xl\dot\theta cos\theta +l^2\dot\theta^2)-mglcos\theta \\
# =\frac{1}{2}(M+m)\dot x^2+m\dot xl\dot\theta cos\theta + \frac{1}{2}ml^2\dot\theta^2-mglcos\theta
# \tag{5}
# $$
# From here we determine the Euler-Lagrange equations of motion (EOM), which are of the form:
# $$
# \frac{d}{dt}\left\lgroup \frac{\partial L}{\partial \dot x}\right\rgroup-\frac{\partial L}{\partial x}=f \\
# \frac{d}{dt}\left\lgroup \frac{\partial L}{\partial \dot \theta}\right\rgroup-\frac{\partial L}{\partial \theta}=0
# $$
# For the four partial derivative terms, we have:
# $$
# \frac{\partial L}{\partial \dot x}=(M+m)\dot x+ml\dot\theta cos\theta \\
# \frac{\partial L}{\partial \dot \theta}=ml\dot x cos\theta+ml^2\dot\theta \\
# \frac{\partial L}{\partial x}=0 \\
# \frac{\partial L}{\partial \theta}=-mlsin\theta\dot x\dot\theta+mglsin\theta
# $$
# And their time derivatives:
# $$
# \frac{d}{dt}\left\lgroup \frac{\partial L}{\partial \dot x}\right\rgroup=(M+m)\ddot x+ml\ddot\theta cos\theta-ml\dot\theta^2 sin\theta \\
# \frac{d}{dt}\left\lgroup \frac{\partial L}{\partial \dot \theta}\right\rgroup=ml^2\ddot\theta+ml\ddot x cos\theta-ml\dot x \dot\theta sin\theta
# $$
# Substitute everything in, and at last we have a model for the system in the form of two nonlinear second-order ODEs:
# $$
# (M+m)\ddot x+ml\ddot\theta cos\theta-ml\dot\theta^2 sin\theta=f \\
# ml^2\ddot\theta+ml\ddot xcos\theta - mglsin\theta=0
# \tag{6}
# $$
# To make the transition to state-space, first we linearize the above using small-angle approximations:
# $$
# cos\theta \approxeq1 \\ sin\theta\approxeq\theta \\ \dot\theta^2\approxeq0
# $$
# Which give the linearized dynamic model:
# $$
# (M+m)\ddot x + ml\ddot\theta=f \\
# m\ddot x +ml\ddot\theta - mg\theta=0
# \tag{7}
# $$
# We use substitutions to solve for each second-order term:
# $$
# \ddot x=\frac{-mg}{M}\theta + \frac{1}{M}f \\
# \ddot\theta=\frac{M+m}{Ml}g\theta - \frac{-1}{Ml}f
# $$
# With these linearly separable terms, we can represent the system in canonical state-space form:
# $$
# \dot \bold x=\bold A \bold x+\bold Bu
# $$
# Where the state vector is defined as:
# $$
# \bold x = [x, \dot x,\theta, \dot\theta]^T
# $$
# The system and input matrices follow:
# $$
# \bold A= \left[
# \begin{matrix}
# 0&1&0&0 \\
# 0&0&\frac{-mg}{M}&0 \\
# 0&0&0&1 \\
# 0&0&\frac{M+m}{Ml}g&0 \\
# \end{matrix}
# \right]
# \hspace{1cm}
# \bold B=\left[
# \begin{matrix}
# 0 \\
# \frac{1}{M} \\
# 0 \\
# \frac{-1}{Ml} \\
# \end{matrix}
# \right]
# \tag{8}
# $$
# Here we introduce the feeback gain matrix **K** and apply it to our input, which we now modify to include an added setpoint. This produces a corresponding closed-loop system:
# $$
# \bold K=\begin{matrix}[k1&k2&k3&k4]\end{matrix} \\
# \bold u=-\bold K \bold y+\bold r\\
# \dot \bold x=(\bold A-\bold B \bold K) \bold x+\bold B \bold r \\
# \bold y=\bold C\bold x+\bold D \bold u
# $$
# This closed-loop system has a characteristic equation:
# $$
# |s\bold I-(\bold A-\bold B \bold K)|=0 \\
# s^4+(a_4+k_4)s^3+(a_3+k_3)s^2+(a_2+k_2)s+(a_1+k_1)=0
# \tag{9}
# $$
# To find our gain values we must choose some performance criteria for the transient response, namely the maximum settling time and percent overshoot. This produces a 2nd-order characteristic polynomial to which we add two more terms to make the overall 'desired' polynomial a 4th-order. The added terms simply place then two remaining roots (poles) at 5x the distance of the other two on the real axis. We can then set this equal to our actual characteristic and solve for our gains by matching like-terms.
# $$
# T_s=\frac{4}{\zeta\omega_n} \\
# \zeta=\frac{-\ln(\frac{\%OS}{100})}{\sqrt{\pi^2+\ln^2(\frac{\%OS}{100})}}
# \\
# (s^2+2\zeta\omega_ns+\omega_n^2)(s+5\zeta\omega_n)(s+5\zeta\omega_n)=s^4+(a_4+k_4)s^3+(a_3+k_3)s^2+(a_2+k_2)s+(a_1+k_1)
# $$
# This manual pole-placement can be improved and the system optimally controlled by instead designing a Linear Quadratic Regulator (LQR). This method minimizes a cost function J:
# $$
# J=\int\limits_0^\infin [\bold x^T \bold Q \bold x+\bold u^T \bold R \bold u]dt
# $$
# Here **Q** and **R** are symmetric and diagonal and act as weightings for the state and input, respectively. For this application we will assume a negligible cost of control (i.e. assume a wall-powered and sufficiently sized motor). We also treat angular movement of the pendulum with higher relative cost penalty than linear movement of the cart. Our weighting terms could then look similar to the following:
# $$
# \bold Q= \left[
# \begin{matrix}
# 1&0&0&0 \\
# 0&1&0&0 \\
# 0&0&10&0 \\
# 0&0&0&100 \\
# \end{matrix}
# \right]
# \hspace{1cm}
# \bold R=[0.01]
# \tag{10}
# $$
# With weights chosen, gains are determined by solving the algebraic Ricatti equation. With gains chosen for state variable feedback and applying them to the difference of the current state from a desired setpoint, the input signal becomes:
# $$
# u = -\bold K(\bold x - \bold x_d)
# \tag{11}
# $$
| simulation/abstract.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Processamento de consulta
# +
import math
import json
import operator
from collections import Counter
class QueryProcessing():
def load(self, path): # carregar o indice invertido no formato json
f = open(path)
return json.load(f)
def init_vectors(self, spl): # inicia as tres estruturas base, recebe o indice invertido como paramentro (sample)
docs = set() # lista de documentos reconhecidos pelo indice invertido
length = {} # a quantidade de vezes que um documento foi referenciado
scores = {} # estrutura base de score, cada chave é um documento que tem valor 0 (zero)
for key in spl.keys(): # processa cada token do indice invertido
dc = spl[key][1] # lista de documentos onde o token ocorre
for item in dc.keys():
docs.add(item) # adiciona o documento
if item not in length:
length[item] = 0
length[item] += dc[item] # contabiliza um para esse documento
scores[item] = 0 # inicializa o valor 0 (zero) para esse documento
return (docs, length, scores)
def __init__(self, index_path):
self.sample = self.load(index_path)
self.docs, self.length, self.scores = self.init_vectors(self.sample)
def rank(self, query, attr="", idf=False): # ranqueamento dos documentos para uma query
query_terms = set([attr + "." + q if attr != "" else q for q in query.split()]) # processa de acordo com o attr informado (para consultas estruturadas)
query_tf = Counter(query_terms) # tf da consulta, contagem de termos
vocabulary = set(self.sample.keys()) # vocabulario do indice invertido (lista de tokens)
terms = query_terms.intersection(vocabulary) # filtro de tokens que existem no vocabulario
docs_score = dict(self.scores) # uma copia da estrutura de scores para os documentos que serao ranqueados
for term in terms: # metodo Term-at-a-time
term_docs = self.sample[term][1] # lista de documentos que o termo ocorre
wq = query_tf[term] # peso da query (tf puro)
if idf: # caso considere o idf
widf = math.log10(len(self.docs)/len(term_docs))
for doc in term_docs:
if idf:
docs_score[doc] += term_docs[doc] * wq * widf
else:
docs_score[doc] += term_docs[doc] * wq
for s in self.scores.keys(): # normalizacao
docs_score[s] = docs_score[s]/self.length[s]
return sorted(docs_score.items(), key=operator.itemgetter(1), reverse=True) # ordenar pelo maior ranking
# -
# ### Como usar
# Primeiramente deve ser instanciado a classe QueryProcessing, informando o índice invertido a ser utilizado
qp = QueryProcessing(index_path="inverted_index.json")
# Com isso, serão construídos o conjunto de documentos reconhecidos pelo índice, e algumas estruturas base
# O próximo passo é chamar o método ```rank```, passando como parametro a query a ser processada, obtendo a lista ranqueada de documentos que correspondem a essa query. Esse método tem dois parametros opcionais: ```attr```, serve para consultas estruturadas ```[direcao, titulo, elenco]```, seu valor padrão é "", caso não especificado; ```idf```, um valor booleano que corresponde a utilização, ou não, do idf no momento do ranqueamento, seu valor padrão é ```False```
qp.rank(query="roman polanski", attr="direcao", idf=True)
| query_processor/.ipynb_checkpoints/Query processing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Detecting Cotton Plant Disease Using ConvNets
# ### By: <NAME>
# ## Importing the libraries
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
import tensorflow
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import Adam
import matplotlib.pyplot as plt
# -
# ## Defining the Train & Test Paths
train_data_path = '../input/cotton-disease-dataset/Cotton Disease/train'
test_data_path = '../input/cotton-disease-dataset/Cotton Disease/test'
training_datagen = ImageDataGenerator(rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
training_data = training_datagen.flow_from_directory(train_data_path,
target_size=(150, 150),
batch_size=32,
class_mode='categorical')
test_datagen = ImageDataGenerator(rescale =1./255)
test_data = test_datagen.flow_from_directory(test_data_path,
target_size=(150,150),
batch_size=32,
class_mode='categorical')
# ## Building the CNN Model
# +
model = keras.models.Sequential([
keras.layers.Conv2D(filters=32, kernel_size=3, input_shape=[150, 150, 3]),
keras.layers.MaxPooling2D(pool_size=(2,2)),
keras.layers.Conv2D(filters=64, kernel_size=3),
keras.layers.MaxPooling2D(pool_size=(2,2)),
keras.layers.Conv2D(filters=128, kernel_size=3),
keras.layers.MaxPooling2D(pool_size=(2,2)),
keras.layers.Conv2D(filters=256, kernel_size=3),
keras.layers.MaxPooling2D(pool_size=(2,2)),
keras.layers.Conv2D(filters=512, kernel_size=3),
keras.layers.MaxPooling2D(pool_size=(2,2)),
keras.layers.Dropout(0.5),
keras.layers.Flatten(),
keras.layers.Dense(units=128, activation='relu'),
keras.layers.Dropout(0.1),
keras.layers.Dense(units=256, activation='relu'),
keras.layers.Dropout(0.25),
keras.layers.Dense(units=4, activation='softmax')
])
# -
model.compile(optimizer=Adam(learning_rate=0.0001),loss='categorical_crossentropy',metrics=['accuracy'])
history = model.fit(training_data,
epochs=50,
verbose=1,
validation_data=test_data,
)
# ### Plotting the Accuracy & Loss Plots
# +
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('CNN Model\'s Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('CNN Model\'s Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# ## Defining the Image Classification Function
# +
from tensorflow.keras.preprocessing import image
import numpy as np
IMAGE_SIZE =(150,150)
def predictor(image_location):
test_image=image.load_img(image_location, target_size = IMAGE_SIZE)
plt.imshow(test_image)
test_image=image.img_to_array(test_image)
test_image=test_image/255
test_image = np.expand_dims(test_image, axis = 0)
preds=np.argmax(model.predict(test_image))
if preds==0:
print("The leaf is diseased cotton leaf")
elif preds==1:
print("The leaf is diseased cotton plant")
elif preds==2:
print("The leaf is fresh cotton leaf")
else:
print("The leaf is fresh cotton plant")
# -
# #### TEST 1
predictor('../input/cotton-disease-dataset/Cotton Disease/val/diseased cotton leaf/dis_leaf (100)_iaip.jpg')
# #### TEST 2
predictor('../input/cotton-disease-dataset/Cotton Disease/val/diseased cotton plant/dd (10)_iaip.jpg')
# #### TEST 3
predictor('../input/cotton-disease-dataset/Cotton Disease/val/fresh cotton leaf/d (106)_iaip.jpg')
# #### TEST 4
predictor('../input/cotton-disease-dataset/Cotton Disease/val/fresh cotton plant/dsd (141)_iaip.jpg')
# ## Validating the Test Set
validation_path = '../input/cotton-disease-dataset/Cotton Disease/val'
valid_data = test_datagen.flow_from_directory(validation_path, target_size=(150, 150),
batch_size=32,
class_mode='categorical')
predicted = model.predict(valid_data[0][0])
# +
actual=[]
for i in valid_data[0][1]:
actual.append(np.argmax(i))
predict=[]
for i in predicted:
predict.append(np.argmax(i))
# -
from sklearn.metrics import accuracy_score
print( "CNN Model's Accuracy", accuracy_score(actual, predict))
# ## Saving the CNN Model
model.save("Cotton-CNN-Model.h5")
# ## THE END
| Cotton Disease Detection using ConvNets/Spp-DL-CNN-3-CottonDisease-Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Ndo4ERqnwQOU"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="MTKwbguKwT4R"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="xfNT-mlFwxVM"
# # 卷积变分自编码器
# + [markdown] colab_type="text" id="0TD5ZrvEMbhZ"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/tutorials/generative/cvae">
# <img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />
# 在 tensorFlow.google.cn 上查看</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/generative/cvae.ipynb">
# <img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />
# 在 Google Colab 中运行</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/generative/cvae.ipynb">
# <img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />
# 在 GitHub 上查看源代码</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/generative/cvae.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载 notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="fKsm6LhC7TAw"
# Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
# [官方英文文档](https://www.tensorflow.org/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
# [tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
# [<EMAIL> Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
# + [markdown] colab_type="text" id="ITZuApL56Mny"
# 
#
# 本笔记演示了如何通过训练变分自编码器([1](https://arxiv.org/abs/1312.6114), [2](https://arxiv.org/abs/1401.4082))来生成手写数字图片。
#
# + colab={} colab_type="code" id="P-JuIu2N_SQf"
# 用于生成 gif
# !pip install imageio
# + [markdown] colab_type="text" id="e1_Y75QXJS6h"
# ## 导入 Tensorflow 与其他库
# + colab={} colab_type="code" id="YfIk2es3hJEd"
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version 仅应用于 Colab。
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
import os
import time
import numpy as np
import glob
import matplotlib.pyplot as plt
import PIL
import imageio
from IPython import display
# + [markdown] colab_type="text" id="iYn4MdZnKCey"
# ## 加载 MNIST 数据集
#
# 每个 MNIST 图片最初都是包含 784 个整数的向量,每个整数取值都在 0-255 之间,表示像素的强度。我们在模型中使用伯努利分布对每个像素进行建模,并对数据集进行静态二值化。
# + colab={} colab_type="code" id="a4fYMGxGhrna"
(train_images, _), (test_images, _) = tf.keras.datasets.mnist.load_data()
# + colab={} colab_type="code" id="NFC2ghIdiZYE"
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1).astype('float32')
test_images = test_images.reshape(test_images.shape[0], 28, 28, 1).astype('float32')
# 标准化图片到区间 [0., 1.] 内
train_images /= 255.
test_images /= 255.
# 二值化
train_images[train_images >= .5] = 1.
train_images[train_images < .5] = 0.
test_images[test_images >= .5] = 1.
test_images[test_images < .5] = 0.
# + colab={} colab_type="code" id="S4PIDhoDLbsZ"
TRAIN_BUF = 60000
BATCH_SIZE = 100
TEST_BUF = 10000
# + [markdown] colab_type="text" id="PIGN6ouoQxt3"
# ## 使用 *tf.data* 来将数据分批和打乱
# + colab={} colab_type="code" id="-yKCCQOoJ7cn"
train_dataset = tf.data.Dataset.from_tensor_slices(train_images).shuffle(TRAIN_BUF).batch(BATCH_SIZE)
test_dataset = tf.data.Dataset.from_tensor_slices(test_images).shuffle(TEST_BUF).batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="THY-sZMiQ4UV"
# ## 通过 *tf.keras.Sequential* 连接生成网络与推理网络
#
#
# 在我们的 VAE 示例中,我们将两个小型的 ConvNet 用于生成和推断网络。由于这些神经网络较小,我们使用 `tf.keras.Sequential` 来简化代码。在下面的描述中,令 $x$ 和 $z$ 分别表示观测值和潜在变量
#
# ### 生成网络
#
#
# 这里定义了生成模型,该模型将潜在编码作为输入,并输出用于观测条件分布的参数,即 $p(x|z)$。另外,我们对潜在变量使用单位高斯先验 $p(z)$。
#
# ### 推理网络
#
#
# 这里定义了近似后验分布 $q(z|x)$,该后验分布以观测值作为输入,并输出用于潜在表示的条件分布的一组参数。在本示例中,我们仅将此分布建模为对角高斯模型。在这种情况下,推断网络将输出因式分解的高斯均值和对数方差参数(为了数值稳定性使用对数方差而不是直接使用方差)。
#
# ### 重参数化技巧
#
# 在优化过程中,我们可以从 $q(z|x)$ 中采样,方法是首先从单位高斯采样,然后乘以标准偏差并加平均值。这样可以确保梯度能够通过样本传递到推理网络参数。
#
# ### 网络架构
#
# 对于推理网络,我们使用两个卷积层,后接一个全连接层。在生成网络中,我们通过使用全连接层,后接三个卷积转置层(在某些情况下也称为反卷积层)来镜像词体系结构。请注意,在训练 VAE 时避免使用批归一化(batch normalization)是一种常见的做法,因为使用小批量处理会导致额外的随机性,从而加剧随机抽样的不稳定性。
# + colab={} colab_type="code" id="VGLbvBEmjK0a"
class CVAE(tf.keras.Model):
def __init__(self, latent_dim):
super(CVAE, self).__init__()
self.latent_dim = latent_dim
self.inference_net = tf.keras.Sequential(
[
tf.keras.layers.InputLayer(input_shape=(28, 28, 1)),
tf.keras.layers.Conv2D(
filters=32, kernel_size=3, strides=(2, 2), activation='relu'),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation='relu'),
tf.keras.layers.Flatten(),
# No activation
tf.keras.layers.Dense(latent_dim + latent_dim),
]
)
self.generative_net = tf.keras.Sequential(
[
tf.keras.layers.InputLayer(input_shape=(latent_dim,)),
tf.keras.layers.Dense(units=7*7*32, activation=tf.nn.relu),
tf.keras.layers.Reshape(target_shape=(7, 7, 32)),
tf.keras.layers.Conv2DTranspose(
filters=64,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation='relu'),
tf.keras.layers.Conv2DTranspose(
filters=32,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation='relu'),
# No activation
tf.keras.layers.Conv2DTranspose(
filters=1, kernel_size=3, strides=(1, 1), padding="SAME"),
]
)
@tf.function
def sample(self, eps=None):
if eps is None:
eps = tf.random.normal(shape=(100, self.latent_dim))
return self.decode(eps, apply_sigmoid=True)
def encode(self, x):
mean, logvar = tf.split(self.inference_net(x), num_or_size_splits=2, axis=1)
return mean, logvar
def reparameterize(self, mean, logvar):
eps = tf.random.normal(shape=mean.shape)
return eps * tf.exp(logvar * .5) + mean
def decode(self, z, apply_sigmoid=False):
logits = self.generative_net(z)
if apply_sigmoid:
probs = tf.sigmoid(logits)
return probs
return logits
# + [markdown] colab_type="text" id="0FMYgY_mPfTi"
# ## 定义损失函数和优化器
#
# VAE 通过最大化边际对数似然的证据下界(ELBO)进行训练:
#
# $$\log p(x) \ge \text{ELBO} = \mathbb{E}_{q(z|x)}\left[\log \frac{p(x, z)}{q(z|x)}\right].$$
#
# 实际上,我们优化了此期望的单样本蒙卡特罗估计:
#
# $$\log p(x| z) + \log p(z) - \log q(z|x),$$
# 其中 $z$ 从 $q(z|x)$ 中采样。
#
# **注意**:我们也可以分析性地计算 KL 项,但简单起见,这里我们将所有三个项合并到蒙卡特罗估计器中。
# + colab={} colab_type="code" id="iWCn_PVdEJZ7"
optimizer = tf.keras.optimizers.Adam(1e-4)
def log_normal_pdf(sample, mean, logvar, raxis=1):
log2pi = tf.math.log(2. * np.pi)
return tf.reduce_sum(
-.5 * ((sample - mean) ** 2. * tf.exp(-logvar) + logvar + log2pi),
axis=raxis)
@tf.function
def compute_loss(model, x):
mean, logvar = model.encode(x)
z = model.reparameterize(mean, logvar)
x_logit = model.decode(z)
cross_ent = tf.nn.sigmoid_cross_entropy_with_logits(logits=x_logit, labels=x)
logpx_z = -tf.reduce_sum(cross_ent, axis=[1, 2, 3])
logpz = log_normal_pdf(z, 0., 0.)
logqz_x = log_normal_pdf(z, mean, logvar)
return -tf.reduce_mean(logpx_z + logpz - logqz_x)
@tf.function
def compute_apply_gradients(model, x, optimizer):
with tf.GradientTape() as tape:
loss = compute_loss(model, x)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
# + [markdown] colab_type="text" id="Rw1fkAczTQYh"
# ## 训练
#
# * 我们从迭代数据集开始
# * 在每次迭代期间,我们将图像传递给编码器,以获得近似后验 $q(z|x)$ 的一组均值和对数方差参数
# * 然后,我们应用 *重参数化技巧* 从 $q(z|x)$ 中采样
# * 最后,我们将重新参数化的样本传递给解码器,以获取生成分布 $p(x|z)$ 的 logit
# * **注意:**由于我们使用的是由 keras 加载的数据集,其中训练集中有 6 万个数据点,测试集中有 1 万个数据点,因此我们在测试集上的最终 ELBO 略高于对 Larochelle 版 MNIST 使用动态二值化的文献中的报告结果。
#
# ## 生成图片
#
# * 进行训练后,可以生成一些图片了
# * 我们首先从单位高斯先验分布 $p(z)$ 中采样一组潜在向量
# * 随后生成器将潜在样本 $z$ 转换为观测值的 logit,得到分布 $p(x|z)$
# * 这里我们画出伯努利分布的概率
# + colab={} colab_type="code" id="NS2GWywBbAWo"
epochs = 100
latent_dim = 50
num_examples_to_generate = 16
# 保持随机向量恒定以进行生成(预测),以便更易于看到改进。
random_vector_for_generation = tf.random.normal(
shape=[num_examples_to_generate, latent_dim])
model = CVAE(latent_dim)
# + colab={} colab_type="code" id="RmdVsmvhPxyy"
def generate_and_save_images(model, epoch, test_input):
predictions = model.sample(test_input)
fig = plt.figure(figsize=(4,4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0], cmap='gray')
plt.axis('off')
# tight_layout 最小化两个子图之间的重叠
plt.savefig('image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
# + colab={} colab_type="code" id="2M7LmLtGEMQJ"
generate_and_save_images(model, 0, random_vector_for_generation)
for epoch in range(1, epochs + 1):
start_time = time.time()
for train_x in train_dataset:
compute_apply_gradients(model, train_x, optimizer)
end_time = time.time()
if epoch % 1 == 0:
loss = tf.keras.metrics.Mean()
for test_x in test_dataset:
loss(compute_loss(model, test_x))
elbo = -loss.result()
display.clear_output(wait=False)
print('Epoch: {}, Test set ELBO: {}, '
'time elapse for current epoch {}'.format(epoch,
elbo,
end_time - start_time))
generate_and_save_images(
model, epoch, random_vector_for_generation)
# + [markdown] colab_type="text" id="P4M_vIbUi7c0"
# ### 使用 epoch 编号显示图片
# + colab={} colab_type="code" id="WfO5wCdclHGL"
def display_image(epoch_no):
return PIL.Image.open('image_at_epoch_{:04d}.png'.format(epoch_no))
# + colab={} colab_type="code" id="5x3q9_Oe5q0A"
plt.imshow(display_image(epochs))
plt.axis('off')# 显示图片
# + [markdown] colab_type="text" id="NywiH3nL8guF"
# ### 生成所有保存图片的 GIF
# + colab={} colab_type="code" id="IGKQgENQ8lEI"
anim_file = 'cvae.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info >= (6,2,0,''):
display.Image(filename=anim_file)
# + [markdown] colab_type="text" id="yQXO_dlXkKsT"
# 如果您正使用 Colab,您可以使用以下代码下载动画。
# + colab={} colab_type="code" id="4fSJS3m5HLFM"
try:
from google.colab import files
except ImportError:
pass
else:
files.download(anim_file)
| site/zh-cn/tutorials/generative/cvae.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''PythonDataV2'': conda)'
# name: python3
# ---
# + [markdown] id="0G6gj1vRR48a"
# # random_forest_model
# ----
#
# Written in the Python 3.7.9 Environment with the following package versions
#
# * joblib 1.0.1
# * numpy 1.19.5
# * pandas 1.3.1
# * scikit-learn 0.24.2
# * tensorflow 2.5.0
#
# By <NAME>
#
# This workbook builds a model from 2017 flight performance and predicts 2018 performance.
# + id="byo9Yx__R48d"
# Import Dependencies
# Plotting
# %matplotlib inline
import matplotlib.pyplot as plt
# Data manipulation
import numpy as np
import pandas as pd
import math
import datetime
from statistics import mean
from operator import itemgetter
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder, MinMaxScaler, OneHotEncoder
from tensorflow.keras.utils import to_categorical
# Parameter Selection
from sklearn import tree
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
# Model Development
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Dropout
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
# Model Metrics
from sklearn.metrics import classification_report
# Save/load files
from tensorflow.keras.models import load_model
import joblib
# from google.colab import files
# # Ignore deprecation warnings
# import warnings
# warnings.simplefilter('ignore', FutureWarning)
# + id="8un601m2R48e"
# Set the seed value for the notebook, so the results are reproducible
from numpy.random import seed
seed(1)
# + [markdown] id="h2XPthEaR48e"
# # Read the clean CSV
# + colab={"base_uri": "https://localhost:8080/", "height": 213} id="6sAlS5XWR48e" outputId="2a4a0b18-7f5b-4a50-b90a-67135cf4dc7b"
# Import data
# Read the CSV file from Github to Pandas Dataframe
url = "https://raw.githubusercontent.com/NicoleLund/flight_delay_prediction/main/data_manipulation_modeling/data_clean/2017_TUS.csv"
df = pd.read_csv(url)
df.head(3)
# + [markdown] id="4xVuJisFR48g"
# # Additional Data Formatting
# + id="M07O7sX_esdt"
# Convert the time fields into decimal hour
def time_conv(mil_time):
hours = []
for time in mil_time:
if time == 2400:
time = 0
if math.isnan(time):
hours.append(time)
elif time < 59:
hour = int(datetime.datetime.strptime(str(int(time)), '%M').strftime('%H'))
minute = int(datetime.datetime.strptime(str(int(time)), '%M').strftime('%M'))
hours.append(hour + minute/60)
else:
hour = int(datetime.datetime.strptime(str(int(time)), '%H%M').strftime('%H'))
minute = int(datetime.datetime.strptime(str(int(time)), '%H%M').strftime('%M'))
hours.append(hour + minute/60)
return hours
# + id="V6Sp0zw3esdt"
df['CRS_DEP_hours'] = time_conv(df['CRS_DEP_TIME'])
df['CRS_ARR_hours'] = time_conv(df['CRS_ARR_TIME'])
# + colab={"base_uri": "https://localhost:8080/"} id="2mvarRxcR48g" outputId="0630b131-4915-41e1-9b8e-eaff9991dba3"
# Split dataframe into X and y
# Define model variables
# Model input
X_categorical_df = df[['OP_CARRIER', 'day_of_week', 'DEST']]
X_numeric_df = df[['OP_CARRIER_FL_NUM', 'CRS_DEP_hours', 'CRS_ARR_hours', 'DISTANCE']]
X_numeric_df.DISTANCE = X_numeric_df.DISTANCE.astype(int)
# Model output
y_df = df[['CANCELLED', 'DIVERTED', 'DELAY']]
y_df.CANCELLED = y_df.CANCELLED.astype(int)
y_df.DIVERTED = y_df.DIVERTED.astype(int)
# + colab={"base_uri": "https://localhost:8080/", "height": 329} id="CdNAGF5XVwQ6" outputId="7f76b4f5-f8ec-4de3-9a92-9887c0ee8ffc"
# Review model output
print(y_df.info())
y_df.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 346} id="lQJv7IovUTMx" outputId="f56919bd-3ec6-4221-e516-34077ddc3420"
# Review model numeric input
print(X_numeric_df.info())
X_numeric_df.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 329} id="4s_93gLRdNWi" outputId="252d3402-544c-4947-a509-ddbce9540665"
# Review model categorical input
print(X_categorical_df.info())
X_categorical_df.head(3)
# + [markdown] id="0z2q7RuAesdv"
# # One-hot encode categorical values
# + colab={"base_uri": "https://localhost:8080/"} id="cgEJEqyQbY9X" outputId="9997284f-8674-42d9-a813-1b9819ed3910"
# View categories for one hot encoder
carriers = pd.unique(X_categorical_df.OP_CARRIER)
day = pd.unique(X_categorical_df.day_of_week)
destination = pd.unique(X_categorical_df.DEST)
print(carriers)
print(day)
print(destination)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="UX5RQGxpesdv" outputId="85d5cdf4-2404-4eb7-f4f4-4bf9b46acedf"
# Force 2017 Categorical transformation to include Carriers 'YV', 'F9', 'G4'
# and Destinations 'PVU', 'CLT', 'BLI'
X_categorical_df.loc[len(X_categorical_df.index)] = ['YV','Sunday','PVU']
X_categorical_df.loc[len(X_categorical_df.index)] = ['F9','Sunday','CLT']
X_categorical_df.loc[len(X_categorical_df.index)] = ['G4','Sunday','BLI']
X_categorical_df.tail(5)
# + colab={"base_uri": "https://localhost:8080/"} id="o2Ui_6FEesdv" outputId="64efb2b8-6632-49b1-919a-4733b3a1ff48"
# Verify categories for one hot encoder including 'YV', 'F9', 'G4', 'PVU', 'CLT', 'BLI'
carriers = pd.unique(X_categorical_df.OP_CARRIER)
day = pd.unique(X_categorical_df.day_of_week)
destination = pd.unique(X_categorical_df.DEST)
print(carriers)
print(day)
print(destination)
# + id="q83aKPc4v4ti"
# Define categorical column names
# Note: 'YV', 'F9', 'G4' are present in the 2018 data and needs 0-filled columns in the model
column_names = ['DL', 'OO', 'UA', 'WN', 'AA', 'EV', 'AS', 'YV', 'F9', 'G4', \
'Sunday', 'Monday', 'Tuesday', 'Wednesday', \
'Thursday', 'Friday', 'Saturday', 'ATL', 'DEN', \
'DFW', 'HOU', 'IAH', 'JFK', 'LAS', 'LAX', 'MDW', \
'MSP', 'OAK', 'ORD', 'PDX', 'PHX', 'SAN', 'SEA', 'SFO', 'SJC', 'SLC','PVU', 'CLT', 'BLI']
# + colab={"base_uri": "https://localhost:8080/"} id="Yvwq5SFIbvKd" outputId="79ae3a30-7c27-41af-9005-3ea296370338"
# One-hot-encode categorical fields
encoder = OneHotEncoder(categories=[carriers,day,destination])
encoder.fit(X_categorical_df)
X_encoded = encoder.transform(X_categorical_df)
X_encoded.toarray()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="9GMmBu_ug2zR" outputId="dfe263a4-50b6-4960-ae66-7b26e4c5e16c"
# Create encoded DataFrame of Categorical Fields
X_encoded_df = pd.DataFrame(X_encoded.toarray(), columns=column_names).astype(int)
X_encoded_df.tail(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="EKVOjFODesdx" outputId="a03d0057-97e8-43e8-e8b4-94f67af9da8f"
# Remove last 3 rows for forced data
X_encoded_df = X_encoded_df.drop([len(X_encoded_df.index)-1], axis=0)
X_encoded_df = X_encoded_df.drop([len(X_encoded_df.index)-1], axis=0)
X_encoded_df = X_encoded_df.drop([len(X_encoded_df.index)-1], axis=0)
X_encoded_df.tail(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 213} id="e_nlkauJesdx" outputId="56d9d1bb-9cc7-4b2c-d449-16578984656e"
# Join encoded data to original DataFrame
encoded_df = df
encoded_df = encoded_df.join(X_encoded_df)
encoded_df.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="zQ1s538wk09F" outputId="23fbe287-d369-446d-e9f8-633b99e1566e"
# Join encoded data to X
X_df = X_numeric_df
X_df = X_df.join(X_encoded_df)
X_df.head(3)
# + [markdown] id="ytW5uyKkesdy"
# # Model Pre-Processing
# + id="tgUzpL5CR48g"
# Split X and y into training and testing groups
X_train, X_test, y_train, y_test = train_test_split(
X_df, y_df, test_size=0.3, random_state=42)
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="WJo3R8jMR48h" outputId="51b38d35-f44f-4a34-ffee-f60374ff51ea"
# Display training data
X_train.head(3)
# + id="ZVZTjs6RR48h"
# Scale the data with MinMaxScaler
X_scaler = MinMaxScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# + [markdown] id="fo4BJNnLesdy"
# # Random Forest Modeling
# + colab={"base_uri": "https://localhost:8080/"} id="COHMmyCBesdy" outputId="960b2962-87af-4239-ef46-4027c3849af5"
# Hyperparamter tuning
model = RandomForestClassifier(n_jobs=-1)
param_grid = {
'n_estimators': [100, 1000, 5000],
'max_features': ['auto', 'sqrt', 'log2']
}
grid = GridSearchCV(estimator=model, param_grid=param_grid, scoring='accuracy')
grid.fit(X_train, y_train)
grid.best_params_
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="b5BBABt4esdz" outputId="af0646a3-b52f-42d4-f3d3-1978c06b3e2f"
# Save the grid
joblib.dump(grid, 'random_forest_grid_v1.sav')
# files.download('random_forest_grid_v1.sav')
# + colab={"base_uri": "https://localhost:8080/"} id="7m8EOXm8esdz" outputId="5a118e3d-6cb1-4a90-feee-9582c31e0f11"
# Build final model
best_model = RandomForestClassifier(n_estimators=5000, max_features= 'log2', n_jobs=-1)
best_model.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="3dzT9tmaesd0" outputId="519daf9d-6e10-4082-d765-d5897cbd7686"
# Save the model
joblib.dump(best_model, 'random_forest_model_v1.sav')
# files.download('random_forest_model_v1.sav')
# + [markdown] id="JDQmf-yMesd0"
# # Score the model
# + colab={"base_uri": "https://localhost:8080/"} id="GC1bazZVesd0" outputId="13dc7155-d382-4088-a6f2-58a375a534f3"
print(f'Random Forest Score: {best_model.score(X_test, y_test)}')
# + id="iWsEhg5qesd0"
predictions = best_model.predict(X_test)
results_cancelled = pd.DataFrame({ \
"CANCELLED": y_test.CANCELLED,"CANCELLED_PREDICT": predictions[:,0]})
results_diverted = pd.DataFrame({ \
"DIVERTED": y_test.DIVERTED,"DIVERTED_PREDICT": predictions[:,1]})
results_delayed = pd.DataFrame({ \
"DELAY": y_test.DELAY, "DELAY_PREDICT": predictions[:,2]})
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="0cFKBgQ_esd0" outputId="ce12fd5f-2ae1-49ea-a5b8-c9b3c3a19095"
print(classification_report(y_test.CANCELLED, predictions[:,0]))
results_cancelled.apply(pd.value_counts)
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="CG9uKo-nesd0" outputId="9b6e5304-7ac8-4b69-f42f-bf7aa61f1734"
print(classification_report(y_test.DIVERTED, predictions[:,1]))
results_diverted.apply(pd.value_counts)
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="SwbgS-3resd0" outputId="0907ff2e-edfc-4f5d-f19f-3b00c1a97fdf"
print(classification_report(y_test.DELAY, predictions[:,2]))
results_delayed.apply(pd.value_counts)
# + [markdown] id="Bc8YfxB2esd1"
# # Review Feature Importance
# + id="9Ow_YoTOesd1"
# Sort the features by their importance
rf_feature_sort = sorted(zip(X_train.columns,best_model.feature_importances_),key=itemgetter(1), reverse=False)
# rf_feature_sort
# + colab={"base_uri": "https://localhost:8080/", "height": 719} id="HzpKjz24esd1" outputId="6b93164b-2114-4afe-a45f-c59de962e8e3"
# Plot Random Forest Feature Importance
fig = plt.figure(figsize=[12,12])
plt.barh(*zip(* (rf_feature_sort)))
plt.xlabel('Feature Importance')
plt.ylabel('Feature Name')
plt.title('Random Forest Features')
plt.show()
# + [markdown] id="PwIPsvwQesd1"
# # Format 2018 data for prediction
# + colab={"base_uri": "https://localhost:8080/", "height": 213} id="PepgmmVKesd1" outputId="d559446c-5887-4cd4-e7f4-4d9abd95f511"
# Import data
# Read the CSV file from Github to Pandas Dataframe
df_2018 = pd.read_csv('https://raw.githubusercontent.com/NicoleLund/flight_delay_prediction/main/data_manipulation_modeling/data_clean/2018_TUS.csv')
df_2018.head(3)
# + id="WGQAc_Lxesd1"
# Convert the time fields into decimal hour for 2018 Prediction
df_2018['CRS_DEP_hours'] = time_conv(df_2018['CRS_DEP_TIME'])
df_2018['CRS_ARR_hours'] = time_conv(df_2018['CRS_ARR_TIME'])
# + colab={"base_uri": "https://localhost:8080/"} id="CLxjyyuiesd1" outputId="1ab53e17-3e29-4ef9-92d6-d35b84835d3c"
# Define Model input for 2018 Prediction
X_categorical_2018_df = df_2018[['OP_CARRIER', 'day_of_week', 'DEST']]
X_numeric_2018_df = df_2018[['OP_CARRIER_FL_NUM', 'CRS_DEP_hours', 'CRS_ARR_hours', 'DISTANCE']]
X_numeric_2018_df.DISTANCE = X_numeric_2018_df.DISTANCE.astype(int)
# + colab={"base_uri": "https://localhost:8080/"} id="YzZH51IJesd2" outputId="81792bc3-dd18-4e86-a18d-9ef8a5dae5ba"
# One-hot-encode categorical fields for 2018 Prediction
X_encoded_2018 = encoder.transform(X_categorical_2018_df)
X_encoded_2018.toarray()
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="jdJp2chc9_fS" outputId="c1d659b2-aa1c-4ecd-9854-5751167181a8"
# Create encoded DataFrame of Categorical Fields
X_encoded_2018_df = pd.DataFrame(X_encoded_2018.toarray(), columns=column_names).astype(int)
X_encoded_2018_df.tail(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="EY9YqpeH-Eod" outputId="6aa3ca2e-0319-449b-bfb2-ab07fed63dcc"
# Join encoded data to X predicted
X_2018_df = X_numeric_2018_df
X_2018_df = X_2018_df.join(X_encoded_2018_df)
X_2018_df.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 69} id="Yytz_BTK-S3V" outputId="a90b07e6-1f67-4b3e-9c23-0203b69a0635"
X_2018_df[X_2018_df.isna().any(axis=1)]
# + colab={"base_uri": "https://localhost:8080/"} id="XdcQNErQ-XNp" outputId="25af6dda-72ff-4af2-c7b2-fe00229d153d"
# Verify resulting dataframe shapes
print(X_numeric_2018_df.shape)
print(X_encoded_2018_df.shape)
print(X_2018_df.shape)
# + id="uCvNzWFb-hm8"
# Scale the prediction data with MinMaxScaler
X_2018_scaled = X_scaler.transform(X_2018_df)
# + [markdown] id="WPJ2Oykc-y0D"
# # Make Predictions
# + colab={"base_uri": "https://localhost:8080/"} id="oMhtIX3b-1k9" outputId="7a3a7959-0bcc-45c7-ff79-8c6748d5b35a"
# Predictions
predictions_2018 = best_model.predict(X_2018_scaled)
predictions_2018
# + colab={"base_uri": "https://localhost:8080/"} id="1LiOng-y_Beb" outputId="620b6c2f-8cf2-4bca-d31f-d17fb7b85f14"
# Verify array shapes
print(predictions_2018.shape)
print(df_2018.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="ad-s-tldAKDh" outputId="8bafa462-04e5-4233-f764-c1895087899f"
# Create DataFrame of predictions
predictions_2018_df = pd.DataFrame({ \
"CANCELLED_PREDICT": predictions_2018[:,0], \
"DIVERTED_PREDICT": predictions_2018[:,1], \
"DELAY_PREDICT": predictions_2018[:,2]})
predictions_2018_df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 213} id="n7fESjz9_aaq" outputId="e0e08f2b-fd2a-41bd-87bb-3103eade06b9"
# Join predicted data to full file
df_2018_predictions = df_2018
df_2018_predictions = df_2018_predictions.join(predictions_2018_df)
df_2018_predictions.head(3)
# + colab={"base_uri": "https://localhost:8080/"} id="QtkAPYCHAtoD" outputId="3c23136f-0777-41c6-96e5-0bb2e61a174c"
print(df_2018_predictions.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="gCDp9aLVAw2k" outputId="3a416e87-5ba5-4021-9134-9ac3f5a6dfa0"
# Save output to csv
df_2018_predictions.to_csv('2018_TUS_wPredictions_v1.csv', index = False)
# files.download('2018_TUS_wPredictions_v1.csv')
| data_manipulation_modeling/model_best_fit/Archive/random_forest_model_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 001_basics
# [Source](https://github.com/send2manoo/Python-TheNoTheoryGuide/)
# The Zen of Python
import this
# Comments
# This is a python tutorial and a single line comment
''' This is a multiline comment
pretty awesome!!
Let me introduce you to jennifer!'''
# Simple imports
import math
import random
# +
# importing specific functions from modules
# imports just the factorial function from math
from math import factorial
# imports all the functions from math
from math import *
# +
# Giving aliases
# The Module name is alaised
import math as m
# The function name is alaised
from math import factorial as fact
# +
# Calling imported functions
# If you import the module you have to call the functions from the module
import math
print (math.factorial(12))
# If you import the functions you can call the function as if it is in your program
from random import randrange as rg
print (rg(23, 1000))
# -
# Variables
msg = "Python!" # String
v2 = 'Python!' # Also String works same
v1 = 2 # Numbers
v3 = 3.564 # Floats / Doubles
v4 = True # Boolean (True / False)
# print()
# automatically adds a newline
print (msg)
print (v2)
print (v1)
print (v3)
print (v4)
print ("Hello Python!")
# +
# Note: Both " and ' can be used to make strings. And this flexibility allows for the following:
msg2 = 'Jennifer said, "I love Python!"'
msg3 = "After that Jennifer's Python Interpreter said it back to her!"
msg4 = 'Of Course she used the command `print("I love Jennifer")`'
print (msg2)
print (msg3)
print (msg4)
# -
# input()
msg = input()
# input() with message
msg = input ("Provide some input: ")
print (msg)
# Check for specific input without storing it
if input("Enter something: ") == "something":
print ("Something something")
else: print ("Not Something")
# +
# Python takes every input as a string
# So, if required you can convert to the required type
msg = input("Enter a number: ")
print (type(msg))
msg = int(input ("Enter a number again, if not a number this will throw an error: "))
print (type(msg))
# +
# Basic Arithmetic operations
# Add
print (3 + 2)
print (3.4565 + 56.232)
print ('------------')
# Subtract
print (3 - 4)
print (34.56 - 3.78)
print ('------------')
# Multiply
print (4 * 3)
print (7.56 * 34)
print ('------------')
# Division
print (5 / 2)
print (5.0 / 2)
print (5 / 2.0)
print (25.0 / 5)
print ('------------')
# Exponents
print (4 ** 4)
print (5.67 ** 3)
print ('------------')
# Modulo
print (10%3)
print (10%11)
| 001_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DATA 531 - Lab 2: Practice with Python - Solving problems, docstrings, exception handling, conditions and loops
# ****
# **Name**: FirstName LastName
#
# **Date**: September 15, 2020
# ****
# ## Objectives
#
# 1. Practice solving problems with programs that use variables, expressions, decisions, and looping/iteration.
# 2. Perform input and output using Python.
# 3. Experience with string manipulation including data splitting, cleaning, and analysis.
# 4. Practice writing docstrings for your functions
# 5. Practice writing assert statements for your functions
# 6. Fluency in reading and writing text (CSV) files including using try-except to handle exceptions.
# 7. Experience analyzing real-world, open data sets.
# 8. Retrieve files from web sites using Python code.
# 9. Using modules `pandas`, `numpy`, and `scipy` for executing a data analysis workflow including data loading, cleaning, and filtering
# 10. Practice using python functions from external modules by reading the docstrings, manual
# ## Question #1 - Calculating Taxes (5 marks)
#
# Write a Python function that calculates and prints the tax on a purchased item. The input of this function should be a string, and the pre-tax price of the item.
#
# Details:
#
# - Add a docstring to this function so users know how to use it. (1 mark)
# - Calculate the provincial tax (5%) federal tax (7%), and store them as separate variables. (1 mark)
# - Choose one: A) Add at least two assert statements as tests for your code OR B) Use try/except statements to test for inputs to the function (1 mark)
# - Print the item amount, the provincial tax, the federal tax, and the total with all taxes included. (2 mark)
# - **Bonus:** Round tax amounts to the nearest cent and display with exactly two decimal points. (1 mark)
# ## Question #2 - Data Cleaning (15 marks)
#
# Create a Python program that cleans data in string format. Data set (copy as string into Python code):
#
# data = """5:Joe:35000:1970-08-09
# 4:Steve:49999:1955-01-02
# 1:Leah:154000:1999-06-12
# 3:Sheyanne:255555:1987-05-14
# 2:Matt:24000:1972-11-03
# 7:Kyla:1000000:1950-02-01
# 8:Dave:15000:2000-09-05
# """
#
# Details:
#
# - Use `split()` to separate data into rows (one per line). (1 mark)
# - Use a for loop to process each line: (1 mark)
# - Use split() to divide data into four fields (id, name, salary, birthdate). Output the fields. Hint: Use "\t" to add tab in output. (2 marks)
# - Calculate the age using the birthdate and the current date. Print the age. [Reference to convert string to date using strptime](https://docs.python.org/3/library/datetime.html#datetime.datetime.strptime) Note: May also use date.fromisoformat(). (3 marks)
# - Calculate and print the total number of people, average salary, highest salary, and youngest employee. (4 marks)
# - Use a for loop to process the data set again: (1 mark)
# - Increase the salary by 20% for any employee whose salary < 40000 or has a name that is less than 5 characters long. Print out new and previous salary. (2 marks)
# - **Bonus:** Create a list to store data in after convert from string in first pass so on second loop processing data use the list rather than parsing the string again. (2 marks)
# - **Bonus:** Update the salary directly in the list and print out the list. (1 mark)
#
# ### Sample Output
#
# Id: 5 Name: Joe Salary: 35000 Birthdate: 1970-08-09
# Age: 49
# Id: 4 Name: Steve Salary: 49999 Birthdate: 1955-01-02
# Age: 64
# Id: 1 Name: Leah Salary: 154000 Birthdate: 1999-06-12
# Age: 20
# Id: 3 Name: Sheyanne Salary: 255555 Birthdate: 1987-05-14
# Age: 32
# Id: 2 Name: Matt Salary: 24000 Birthdate: 1972-11-03
# Age: 46
# Id: 7 Name: Kyla Salary: 1000000 Birthdate: 1950-02-01
# Age: 69
# Id: 8 Name: Dave Salary: 15000 Birthdate: 2000-09-05
# Age: 19
#
# Number of people: 7
# Total salary: 1533554 Average salary: 219079.14285714287 Max salary: 1000000
# Youngest employee: Dave
#
# Joe Old salary: 35000 New salary: 42000.0
# Leah Old salary: 154000 New salary: 184800.0
# Matt Old salary: 24000 New salary: 28800.0
# Kyla Old salary: 1000000 New salary: 1200000.0
# Dave Old salary: 15000 New salary: 18000.0
#
# Bonus:
# Joe Old salary: 35000 New salary: 42000.0
# Leah Old salary: 154000 New salary: 184800.0
# Matt Old salary: 24000 New salary: 28800.0
# Kyla Old salary: 1000000 New salary: 1200000.0
# Dave Old salary: 15000 New salary: 18000.0
# [['5', 'Joe', 42000.0, '1970-08-09'], ['4', 'Steve', 49999, '1955-01-02'], ['1', 'Leah', 184800.0, '1999-06-12'], ['3', 'Sheyanne', 255555, '1987-05-14'], ['2', 'Matt', 28800.0, '1972-11-03'], ['7', 'Kyla', 1200000.0, '1950-02-01'], ['8', 'Dave', 18000.0, '2000-09-05']]
# +
### Your answer here
# -
# ## Question #3 - Data Analysis of GDP Data Stored as a CSV File (5 marks)
#
# Create a Python program that uses the open data set on GDP from the Canadian government [36100293.csv](data/36100293.csv) ([Original data source](https://open.canada.ca/data/en/dataset/b0c4a8e1-bb44-4ea8-a389-a3a6b87573aa)) to answer some GDP questions and produce a graph. Details:
#
# - Ensure the data file [36100293.csv](data/36100293.csv) is in your code directory for input. There is also a metadata file [36100293_MetaData.csv](data/36100293_MetaData.csv) providing info on the data file (not required to use).
# - Create an empty dictionary that will eventually have the year as a key and the GDP for that year as a value. Create two empty lists to store year and GDP data. (1 marks)
# - Read the data file using the ``csv`` module and store the GDP for each year in the dictionary and the year and GDP in each list. Code must use try-except to handle any errors. Note that this data set has lots of data besides GDP so you will need to determine which rows store GDP values. (1 mark)
# - Prompt the user for a year and output the GDP for that year from the dictonary. (1 mark)
# - Calculate and print the GDP change from the user year and the GDP in the 1970. (1 mark)
# - Create a line chart using matplotlib showing the GDP from 1961 to 2011. Here is [matplotlib line chart example code](https://matplotlib.org/3.1.1/gallery/lines_bars_and_markers/simple_plot.html) Use the GDP and year lists created previously for the chart data. (1 mark)
#
# ### Sample Output
#
# Enter a year to lookup GDP: 1965
# GDP in 1965 was 17.2 billion dollars which is -20.000000000000004 % different than 1970.
#
# 
# +
### Helpful hint
"""You can prompt the user for a value like this:"""
value_returned = input('Value')
# +
### Your answer here
# -
# ## Optional question (for practice; not for marks)
# ### Question #2 - Data Analysis Pipeline using Python Modules (15 marks)
#
# Create a Python program that reads data on Canadian household incomes from open data web site ([Data source](https://open.canada.ca/data/en/dataset/a932e23b-6480-404d-9d15-708dc9cf9b81)) and performs analysis and reporting. Details:
#
# - Put a comment at the top of the Python file called ``lab4q2`` with your name.
# - The data source URL is [https://www150.statcan.gc.ca/n1/tbl/csv/11100032-eng.zip](https://www150.statcan.gc.ca/n1/tbl/csv/11100032-eng.zip). Download zip file using ``urllib.request`` A method called ``urlretrieve`` may be deprecated in the future but is very easy to use for this. (1 mark) [Reference example for urllib](https://stackoverflow.com/questions/7243750/download-file-from-web-in-python-3)
# - Unzip the file using ``zipfile`` module and ``extractall`` method. (1 mark) [zipfile reference](https://docs.python.org/3/library/zipfile.html) and [zipfile extractall example](https://thispointer.com/python-how-to-unzip-a-file-extract-single-multiple-or-all-files-from-a-zip-archive/) Note: Once you have the data downloaded and unzipped, comment out this code until submission so you do not keep downloading the data!
# - Use ``pandas`` module and ``read_csv`` method to read CSV file. (1 mark)
# - Filter the data to include only ``Canada`` data and print the number of rows. (2 marks)
# - Produce two subsets of the data:
# 1. All data rows with ``Average total income``.
# 2. All data rows with ``Average income tax``.
# - Print out number of rows in each data set and use ``head(4)`` to print out first 4 rows in tax data set (showing only ``Income concept`` and ``VALUE`` columns.) [Pandas reference](https://pandas.pydata.org/pandas-docs/stable/indexing.html) (3 marks)
# - Perform and graph the output for a linear regression ``(tax, income)`` on the ``VALUE`` column for these two data subsets. (2 marks)
# - Create a ``matplotlib`` bar chart with bars for ``tax`` and ``income`` and x-axis labels the family categories (``Economic family type``). Note: To resize figure larger use: ``plt.figure(figsize=(6,6))`` (2 marks)
# - Perform a k-means clustering with 3 clusters and graph results. Use sample code in notes with slight modification. (3 marks)
#
# ### Sample Output
#
# Number of data rows for Canada: 216
# Number of data rows: 18
# Number of data rows: 18
# Income concept VALUE
# 108 Average income tax 12400.0
# 109 Average income tax 16000.0
# 110 Average income tax 9700.0
# 111 Average income tax 9000.0
#
# 
# 
# 
| labs/lab2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
rm(list= ls())
library(dplyr)
library(tidyr)
library(ggplot2)
library(BayesFactor)
library(lme4)
library(lmerTest)
data.folder <- 'Experiment 3. Visibility of the illusory Kanizsa triangle'
# +
### loading raw results and computing duration of individual event
results= read.csv(file.path(data.folder, 'Experiment3.csv'), sep=';')
compute.duration <- function(times, events){
duration <- rep(NA, length(times))
duration[events==0] <- times[events==1]-times[events==0]
return(duration)
}
results <- results %>%
# removing early static masks condition
filter(RotationSpeed!=0) %>%
# converting time to seconds
mutate(Time= Time / 1000) %>%
# computing duration of disappearances for each target
group_by(ID, Block, Target) %>%
mutate(Duration= compute.duration(Time, Event)) %>%
# making sure that factors as indeed factors
ungroup() %>%
mutate(MasksN = factor(MasksN, levels= c(3, 1)))
# -
# ## Proportion of time participant report that at least one target was invisible
# +
disappearance <- results %>%
# computing proportion for each block
group_by(ID, MasksN, Block) %>%
summarise(time.prop= sum(Duration[Target>0], na.rm= TRUE)/BlockDuration[1]) %>%
# averaging across blocks for each observer and condition
group_by(ID, MasksN) %>%
summarise(time.prop= 100*mean(time.prop))
disappearance <- data.frame(disappearance)
# -
# Looking at the effect of the mask using linear-mixed models and Bayesian ANOVA
# +
lm.null <- lme4::lmer(time.prop ~ 1 + (1|ID), data= disappearance, REML= FALSE)
lm.masks <- update(lm.null, .~. + MasksN)
anova(lm.null, lm.masks)
set.seed(211122017)
sort(anovaBF(time.prop ~ MasksN + ID, data= disappearance, whichRandom = 'ID'))
# -
# Plotting
# +
# averages per group
time.per.condition.plot <- disappearance %>%
# Adjusting observers' means following Loftus & Masson (1994)
group_by(ID) %>%
mutate(ID.avg= mean(time.prop)) %>%
ungroup() %>%
mutate(overall.avg= mean(time.prop)) %>%
mutate(time.adjusted= time.prop - ID.avg + overall.avg) %>%
# computing group averages per condition
group_by(MasksN) %>%
summarise(time.avg= mean(time.adjusted),
time.serr= sd(time.adjusted)/sqrt(n()))
# comparison to the baseline condition (M3)
duration.lmer <- summary(lmerTest::lmer(time.prop ~ MasksN + (1|ID), data= disappearance))
duration.lmer
rcontrast<-function(t, df) {
return (sqrt(t^2/(t^2 + df)))
}
coefficients.only <- data.frame(duration.lmer$coefficients)
colnames(coefficients.only) <- c('Estimate', 'Std.Error', 'df', 't.value', 'p.value')
dplyr::mutate(coefficients.only, R.sqr= rcontrast(t.value, df))
time.plot <- ggplot(data= time.per.condition.plot, aes(x= MasksN, y= time.avg, ymin= time.avg-time.serr, ymax= time.avg+time.serr))+
geom_errorbar(color= 'darkgreen', width=0.3)+
geom_point(color= 'darkgreen', size= 3) +
ylab('Disappearance time [%]')
# ggtitle('Average disappearance time across all targets')
print(time.plot)
ggsave('Figures/Exp3 - Kanitza visibility - disappearance duration.pdf', time.plot, width= 4, height= 5, units = 'cm', useDingbats = FALSE)
# -
# ## Proportion of time participant report that at the illusory Kanizsa triangle is visible
# +
visibility <- results %>%
# computing proportion for each block
group_by(ID, MasksN, Block) %>%
summarise(time.prop= sum(Duration[Target==0], na.rm= TRUE)/BlockDuration[1]) %>%
# averaging across blocks for each observer and condition
group_by(ID, MasksN) %>%
summarise(time.prop= 100*mean(time.prop))
visibility <- data.frame(visibility)
# -
# Looking at the effect of the mask using linear-mixed models and Bayesian ANOVA
# +
lm.null <- lme4::lmer(time.prop ~ 1 + (1|ID), data= visibility, REML= FALSE)
lm.masks <- update(lm.null, .~. + MasksN)
anova(lm.null, lm.masks)
set.seed(111122017)
sort(anovaBF(time.prop ~ MasksN + ID, data= visibility, whichRandom = 'ID'))
# -
# Plotting
# +
# averages per group
time.per.condition.plot <- visibility %>%
# Adjusting observers' means following Loftus & Masson (1994)
group_by(ID) %>%
mutate(ID.avg= mean(time.prop)) %>%
ungroup() %>%
mutate(overall.avg= mean(time.prop)) %>%
mutate(time.adjusted= time.prop - ID.avg + overall.avg) %>%
# computing group averages per condition
group_by(MasksN) %>%
summarise(time.avg= mean(time.adjusted),
time.serr= sd(time.adjusted)/sqrt(n()))
# comparison to the baseline condition (M3)
duration.lmer <- summary(lmerTest::lmer(time.prop ~ MasksN + (1|ID), data= visibility))
duration.lmer
rcontrast<-function(t, df) {
return (sqrt(t^2/(t^2 + df)))
}
coefficients.only <- data.frame(duration.lmer$coefficients)
colnames(coefficients.only) <- c('Estimate', 'Std.Error', 'df', 't.value', 'p.value')
dplyr::mutate(coefficients.only, R.sqr= rcontrast(t.value, df))
time.plot <- ggplot(data= time.per.condition.plot, aes(x= MasksN, y= time.avg, ymin= time.avg-time.serr, ymax= time.avg+time.serr))+
geom_errorbar(color= 'orange', width=0.3)+
geom_point(color= 'orange', size= 3) +
ylab('visibility time [%]')
# ggtitle('Average visibility time across all targets')
print(time.plot)
ggsave('Figures/Exp3 - Kanitza visibility - visibility duration.pdf', time.plot, width= 4, height= 5, units = 'cm', useDingbats = FALSE)
| Experiment 3. Analysis.ipynb |
# +
# Control variates demo
# https://en.wikipedia.org/wiki/Control_variates
import numpy as np
np.random.seed(0)
N = 1500
u = np.random.uniform(size=N)
f = 1 / (1 + u)
mu_naive = np.mean(f)
se_naive = np.sqrt(np.var(f) / N)
print("naive {:0.4f}, se {:0.4f}".format(mu_naive, se_naive))
# control variate version
c = 0.4773
g = 1 + u
baseline = 3.0 / 2
cv = f + c * (g - baseline)
mu_cv = np.mean(cv)
se_cv = np.sqrt(np.var(cv) / N)
print("cv {:0.4f}, se {:0.4f}".format(mu_cv, se_cv))
| notebooks/book2/10/control_variates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# # Kepler Hack: Q1–Q17 Occurrence Rate Calculation
#
# By: <NAME>
#
# This is a version of [a blog post I wrote](http://dan.iel.fm/posts/exopop/) updated for the most recent Kepler data release. The main change from Q1–Q16 is that the completeness model has changed. The main changes are:
#
# 1. the MES threshold should be set to 15
# 2. the matched filter was no longer a box. Eherefore the "depth" relevant for the completeness should be the *minimum* not the *mean*.
#
# First, a helper function for downloading data from the Exoplanet Archive:
# +
import os
import requests
import numpy as np
import pandas as pd
from io import BytesIO # Python 3 only!
import matplotlib.pyplot as pl
def get_catalog(name, basepath="data"):
"""
Download a catalog from the Exoplanet Archive by name and save it as a
Pandas HDF5 file.
:param name: the table name
:param basepath: the directory where the downloaded files should be saved
(default: ``data`` in the current working directory)
"""
fn = os.path.join(basepath, "{0}.h5".format(name))
if os.path.exists(fn):
return pd.read_hdf(fn, name)
if not os.path.exists(basepath):
os.makedirs(basepath)
print("Downloading {0}...".format(name))
url = ("http://exoplanetarchive.ipac.caltech.edu/cgi-bin/nstedAPI/"
"nph-nstedAPI?table={0}&select=*").format(name)
r = requests.get(url)
if r.status_code != requests.codes.ok:
r.raise_for_status()
fh = BytesIO(r.content)
df = pd.read_csv(fh)
df.to_hdf(fn, name, format="t")
return df
# -
# Here's the completeness model to apply to Q1—Q17 catalog:
# +
def get_duration(period, aor, e):
"""
Equation (1) from Burke et al. This estimates the transit
duration in the same units as the input period. There is a
typo in the paper (24/4 = 6 != 4).
:param period: the period in any units of your choosing
:param aor: the dimensionless semi-major axis (scaled
by the stellar radius)
:param e: the eccentricity of the orbit
"""
return 0.25 * period * np.sqrt(1 - e**2) / aor
def get_a(period, mstar, Go4pi=2945.4625385377644/(4*np.pi*np.pi)):
"""
Compute the semi-major axis of an orbit in Solar radii.
:param period: the period in days
:param mstar: the stellar mass in Solar masses
"""
return (Go4pi*period*period*mstar) ** (1./3)
def get_delta(k, c=1.0874, s=1.0187, mean=False):
"""
Estimate the approximate expected transit depth as a function
of radius ratio. There might be a typo here. In the paper it
uses c + s*k but in the public code, it is c - s*k:
https://github.com/christopherburke/KeplerPORTs
:param k: the dimensionless radius ratio between the planet and
the star
"""
delta_max = k*k * (c + s*k)
if mean:
return 0.84 * delta_max
return delta_max
cdpp_cols = [k for k in get_catalog("q1_q17_dr24_stellar").keys() if k.startswith("rrmscdpp")]
cdpp_vals = np.array([k[-4:].replace("p", ".") for k in cdpp_cols], dtype=float)
def get_mes(star, period, rp, tau, re=0.009171, mean=False):
"""
Estimate the multiple event statistic value for a transit.
:param star: a pandas row giving the stellar properties
:param period: the period in days
:param rp: the planet radius in Earth radii
:param tau: the transit duration in hours
"""
# Interpolate to the correct CDPP for the duration.
cdpp = np.array(star[cdpp_cols], dtype=float)
sigma = np.interp(tau, cdpp_vals, cdpp)
# Compute the radius ratio and estimate the S/N.
k = rp * re / star.radius
snr = get_delta(k, mean=mean) * 1e6 / sigma
# Scale by the estimated number of transits.
ntrn = star.dataspan * star.dutycycle / period
return snr * np.sqrt(ntrn)
# Pre-compute and freeze the gamma function from Equation (5) in
# Burke et al.
mesthresh = 15
def get_pdet(star, aor, period, rp, e, comp_p, mean=False):
"""
Equation (5) from Burke et al. Estimate the detection efficiency
for a transit.
:param star: a pandas row giving the stellar properties
:param aor: the dimensionless semi-major axis (scaled
by the stellar radius)
:param period: the period in days
:param rp: the planet radius in Earth radii
:param e: the orbital eccentricity
"""
tau = get_duration(period, aor, e) * 24.
mes = get_mes(star, period, rp, tau, mean=mean)
y = np.polyval(comp_p, mes) / (1 + np.exp(-2.0*(mes-mesthresh)))
return y * (y <= 1.0) + 1.0 * (y > 1.0)
def get_pwin(star, period):
"""
Equation (6) from Burke et al. Estimates the window function
using a binomial distribution.
:param star: a pandas row giving the stellar properties
:param period: the period in days
"""
M = star.dataspan / period
f = star.dutycycle
omf = 1.0 - f
pw = 1 - omf**M - M*f*omf**(M-1) - 0.5*M*(M-1)*f*f*omf**(M-2)
msk = (pw >= 0.0) * (M >= 2.0)
return pw * msk
def get_pgeom(aor, e):
"""
The geometric transit probability.
See e.g. Kipping (2014) for the eccentricity factor
http://arxiv.org/abs/1408.1393
:param aor: the dimensionless semi-major axis (scaled
by the stellar radius)
:param e: the orbital eccentricity
"""
return 1. / (aor * (1 - e*e)) * (aor > 1.0)
def get_completeness(star, period, rp, e, comp_p, with_geom=True, mean=False):
"""
A helper function to combine all the completeness effects.
:param star: a pandas row giving the stellar properties
:param period: the period in days
:param rp: the planet radius in Earth radii
:param e: the orbital eccentricity
:param with_geom: include the geometric transit probability?
"""
aor = get_a(period, star.mass) / star.radius
pdet = get_pdet(star, aor, period, rp, e, comp_p, mean=mean)
pwin = get_pwin(star, period)
if not with_geom:
return pdet * pwin
pgeom = get_pgeom(aor, e)
return pdet * pwin * pgeom
# -
# And a function for estimating the occurrence rate (assumed constant) in a bin in $T_\mathrm{eff}$ and period:
def run_analysis(trng, period_rng):
stlr = get_catalog("q1_q17_dr24_stellar")
# Select the stars.
m = np.isfinite(stlr.teff) & (trng[0] <= stlr.teff) & (stlr.teff < trng[1])
m &= np.isfinite(stlr.logg) & (4.0 <= stlr.logg)
# Only include stars with sufficient data coverage.
m &= (stlr.dutycycle * stlr.dataspan) > 2*365.25
m &= stlr.dutycycle > 0.33
# Only select stars with mass estimates.
m &= np.isfinite(stlr.mass)
stlr = pd.DataFrame(stlr[m])
print("Selected {0} targets after cuts".format(len(stlr)))
# KOI catalog.
kois = get_catalog("q1_q17_dr24_koi")
# Select candidates.
rp_rng = (1.5, 2.3)
# Join on the stellar list.
kois = pd.merge(kois, stlr[["kepid", "teff", "radius"]], on="kepid", how="inner")
# Only select the KOIs in the relevant part of parameter space.
m = kois.koi_pdisposition == "CANDIDATE"
base_kois = pd.DataFrame(kois[m])
m &= (period_rng[0] <= kois.koi_period) & (kois.koi_period < period_rng[1])
m &= np.isfinite(kois.koi_prad) & (rp_rng[0] <= kois.koi_prad) & (kois.koi_prad < rp_rng[1])
m &= np.isfinite(kois.koi_max_mult_ev) & (kois.koi_max_mult_ev > 15.0)
kois = pd.DataFrame(kois[m])
print("Selected {0} KOIs after cuts".format(len(kois)))
# Calibrate the completeness.
inj = pd.read_csv("data/DR24-Pipeline-Detection-Efficiency-Table.txt", delim_whitespace=True,
skiprows=4, header=None, names=[
"kepid", "sky", "period", "epoch", "t_depth", "t_dur", "t_b", "t_ror", "t_aor",
"offset_from_source", "offset_distance", "expect_mes", "recovered", "meas_mes",
"r_period", "r_epoch", "r_depth", "r_dur", "r_b", "r_ror", "r_aor"
], na_values="null")
# Join on the stellar list.
inj = pd.merge(inj, stlr[["kepid"]], on="kepid", how="inner")
# Estimate the linear trend above 15 MES.
bins = np.linspace(mesthresh, 80, 20)
n_tot, _ = np.histogram(inj.expect_mes, bins)
m = inj.meas_mes > mesthresh
# m = inj.expect_mes > mesthresh
m &= inj.recovered
n_rec, _ = np.histogram(inj.expect_mes[m], bins)
x = 0.5 * (bins[:-1] + bins[1:])
y = n_rec / n_tot
m = np.isfinite(y)
x, y = x[m], y[m]
pl.figure()
comp_p = np.polyfit(x, y, 1)
pl.plot(x, y)
x0 = np.linspace(0, 80, 500)
pl.plot(x0, np.polyval(comp_p, x0) / (1 + np.exp(-2*(x0-mesthresh))))
pl.xlabel("expected MES");
# Compute the mean completeness.
print("Computing mean completeness...")
p = np.exp(np.random.uniform(np.log(period_rng[0]), np.log(period_rng[1]), 5000))
r = np.exp(np.random.uniform(np.log(rp_rng[0]), np.log(rp_rng[1]), len(p)))
c = np.zeros(len(p))
for _, star in stlr.iterrows():
c += get_completeness(star, p, r, 0.0, comp_p, with_geom=True)
# Compute occurrence rate.
Q = np.mean(c)
N = len(kois)
occ = N / Q
sig = occ / np.sqrt(N)
print("{0:.3} ± {1:.3}".format(occ, sig))
return occ, sig, N, Q, comp_p
# G-dwarfs:
run_analysis((5300.0, 6000.0), (40, 80))
# M-dwarfs:
run_analysis((2400.0, 3900.0), (20, 40))
| keplerhack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(IRdisplay)
library(devtools)
dev_mode(on=T)
install_github("mathisonian/lightning-rstat/LightningR")
library(LightningR)
vizserver <- Lightning$new("http://public.lightning-viz.org/", notebook=T)
vizserver$createsession()
vizserver$line(c(1,2,3,4,5,6,7,8))
vizserver$scatter(c(1, 2, 3), c(1, 2, 3), )
x <- seq(-3, 7, by = 1/8)
vizserver$line(cos(x))
vizserver$scatter(x, cos(x))
# ## excercise use the scatter3 function to plot a sin wave in webgl
#
# ### try to modify the color of each point
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import tensorflow as tf
import os
import random
from collections import defaultdict
import pandas as pd
import time
def load_data_train():
user_movie = defaultdict(set)
data=pd.read_csv('BRP_datas\\BRP_common_user_book\\common_user_book_19_1VS2.csv')
num_user=len(pd.unique(data['user_id']))
num_book=len(pd.unique(data['book_id']))
print('训练集借阅记录数:{}'.format(data.shape[0]))
for row,val in data.iterrows():
u = int(val['user_id'])
i = int(val['book_id'])
user_movie[u].add(i) #
print("num_user:", num_user)
print("num_book", num_book)
return num_user, num_book, user_movie
def load_data_test():
user_movie = defaultdict(set)
data=pd.read_csv('BRP_datas\\BRP_common_user_book\\common_user_book_19_2VS1.csv')
num_user=len(pd.unique(data['user_id']))
num_book=len(pd.unique(data['book_id']))
print('测试集借阅记录数:{}'.format(data.shape[0]))
for row,val in data.iterrows():
u = int(val['user_id'])
i = int(val['book_id'])
user_movie[u].add(i)
print("num_user:", num_user)
print("num_book", num_book)
return num_user, num_book, user_movie
def generate_test(user_movie_pair_test):
"""
对每一个用户u,在user_movie_pair_test中随机找到他借阅过的一本书,保存在user_ratings_test,
后面构造训练集和测试集需要用到。
"""
user_test = dict()
for u,i_list in user_movie_pair_test.items():
user_test[u] = random.sample(user_movie_pair_test[u],1)[0]
return user_test
def generate_train_batch(user_movie_pair_train,item_count,batch_size=50):
t = []
for b in range(batch_size):
u = random.sample(user_movie_pair_train.keys(),1)[0]
i = random.sample(user_movie_pair_train[u],1)[0]
j = random.randint(0,item_count)
while j in user_movie_pair_train[u]:
j = random.randint(0,item_count)
t.append([u,i,j])
return np.asarray(t)
def generate_test_batch(user_ratings_test,user_movie_pair_test,item_count):
"""
对于每个用户u,它的评分图书i是我们在user_ratings_test中随机抽取的,它的j是用户u所有没有借阅过的图书集合,
比如用户u有1000本书没有借阅,那么这里该用户的测试集样本就有1000个
"""
for u in user_movie_pair_test.keys():
t = []
i = user_ratings_test[u]
for j in range(0,item_count):
if not(j in user_movie_pair_test[u]):
t.append([u,i,j])
yield np.asarray(t)
def bpr_mf(user_count,item_count,hidden_dim):
u = tf.placeholder(tf.int32,[None])
i = tf.placeholder(tf.int32,[None])
j = tf.placeholder(tf.int32,[None])
user_emb_w = tf.get_variable("user_emb_w", [user_count+1, hidden_dim],
initializer=tf.random_normal_initializer(0, 0.1))
item_emb_w = tf.get_variable("item_emb_w", [item_count+1, hidden_dim],
initializer=tf.random_normal_initializer(0, 0.1))
u_emb = tf.nn.embedding_lookup(user_emb_w, u)
i_emb = tf.nn.embedding_lookup(item_emb_w, i)
j_emb = tf.nn.embedding_lookup(item_emb_w, j)
x = tf.reduce_sum(tf.multiply(u_emb,(i_emb-j_emb)),1,keep_dims=True)
mf_auc = tf.reduce_mean(tf.to_float(x>0))
l2_norm = tf.add_n([
tf.reduce_sum(tf.multiply(u_emb, u_emb)),
tf.reduce_sum(tf.multiply(i_emb, i_emb)),
tf.reduce_sum(tf.multiply(j_emb, j_emb))
])
regulation_rate = 0.0001
bprloss = regulation_rate * l2_norm - tf.reduce_mean(tf.log(tf.sigmoid(x)))
train_op = tf.train.GradientDescentOptimizer(0.01).minimize(bprloss)
return u, i, j, mf_auc, bprloss, train_op
start=time.clock()
user_count,item_count,user_movie_pair_train = load_data_train()
test_user_count,test_item_count,user_movie_pair_test = load_data_test()
user_ratings_test = generate_test(user_movie_pair_test)
print('user_ratings_test的值为:{}'.format(user_ratings_test))
with tf.Session() as sess:
u,i,j,mf_auc,bprloss,train_op = bpr_mf(user_count,item_count,20)
sess.run(tf.global_variables_initializer())
for epoch in range(1,6):
print('epoch的值为{}'.format(epoch))
_batch_bprloss = 0
for k in range(1,5000):
uij = generate_train_batch(user_movie_pair_train,item_count)
_bprloss,_train_op = sess.run([bprloss,train_op],
feed_dict={u:uij[:,0],i:uij[:,1],j:uij[:,2]})
_batch_bprloss += _bprloss
print("epoch:",epoch)
print("bpr_loss:",_batch_bprloss / k)
print("_train_op")
user_count = 0
_auc_sum = 0.0
for t_uij in generate_test_batch(user_ratings_test,user_movie_pair_test,item_count):
_auc, _test_bprloss = sess.run([mf_auc, bprloss],
feed_dict={u: t_uij[:, 0], i: t_uij[:, 1], j: t_uij[:, 2]}
)
user_count += 1
_auc_sum += _auc
print("test_loss: ", _test_bprloss, "test_auc: ", _auc_sum / user_count)
print("")
variable_names = [v.name for v in tf.trainable_variables()]
values = sess.run(variable_names)
for k, v in zip(variable_names, values):
print("Variable: ", k)
print("Shape: ", v.shape)
print(v)
session1 = tf.Session()
u1_all = tf.matmul(values[0], values[1],transpose_b=True)
result_1 = session1.run(u1_all)
print (result_1)
p = np.squeeze(result_1)
# np.argsort(p),将元素从小到大排列,提取对应的索引。找到了索引就是找到了书
ind = np.argsort(p)[:,-5:]
print('top5对应的索引为{}'.format(ind))
num=0
all_num_user_item=0
for ii in range(len(user_movie_pair_test)):
num_user_item=0
for jj in user_movie_pair_test[ii]:
num_user_item+=1
if jj in (ind[ii]):
num+=1
all_num_user_item+=num_user_item
print('num的值为:{}'.format(num))
print('用户的数目为{}'.format(len(user_movie_pair_test)))
print('用户喜欢的物品的数目为:{}'.format(all_num_user_item))
print('召回率为{}'.format(num/all_num_user_item))
print('准确率为{}'.format(num/(len(user_movie_pair_test)*5)))
duration=time.clock()-start
print('耗费时间:{}'.format(duration))
| campus_data_mining/reference/5/BRP_1VS2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
def haldane_honeycomb(kx, ky, m=0.5, phi=np.pi/2):
k = np.array([kx / np.sqrt(3.), ky * 2. / 3.])
t1 = t2 = 1.
a1 = np.array([np.sqrt(3) * 0.5, 0.5])
a2 = np.array([0, -1])
a3 = np.array([-np.sqrt(3) * 0.5, 0.5])
b1 = a2 - a3
b2 = a3 - a1
b3 = a1 - a2
pauli0 = np.eye(2)
pauli1 = np.array([[0, 1], [1, 0]])
pauli2 = np.array([[0, -1j], [1j, 0]])
pauli3 = np.array([[1, 0], [0, -1]])
hk = 2 * t2 * np.cos(phi) * (
np.cos(k @ b1) + np.cos(k @ b2) + np.cos(k @ b3)
) * pauli0 + t1 * (
(np.cos(k @ a1) + np.cos(k @ a2) + np.cos(k @ a3)) * pauli1 +
(np.sin(k @ a1) + np.sin(k @ a2) + np.sin(k @ a3)) * pauli2
) + (m - 2 * t2 * np.sin(phi) * (
np.sin(k @ b1) + np.sin(k @ b2) + np.sin(k @ b3)
)) * pauli3
return hk
# -
from chern import Hamiltonian, Chern
hk = Hamiltonian(haldane_honeycomb, "haldane")
hk
cn = Chern(hk)
cn
cn.chern
| examples/haldane.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.3
# language: julia
# name: julia-1.6
# ---
# https://twitter.com/MathSorcerer/status/1302576992965984261
# +
module Goma # 再編集しやすいように仮想モジュールを設定
abstract type AbstractEdge{T<:Integer} end
struct Edge{T} <: AbstractEdge{T}
data::NTuple{2, T}
Edge(x::NTuple{2, T}) where T = new{T}(x)
# To define Edge{Int}(1, 2)
Edge{T}(x::Vararg{T2, 2}) where {T, T2} = new{T}(NTuple{2, T}(convert.(T, x)))
# To define Edge{Int}([1, 2])
Edge{T}(x::AbstractVector{T2}) where {T,T2} = new{T}(NTuple{2, T2}(x))
# To define Edge{Int}((1,2))
Edge{T}(x::Tuple) where T = new{T}(NTuple{2, T}(x))
end
function Edge(x::Tuple)
nt = NTuple(promote(x...))
if eltype(nt) <: Integer
Edge(nt)
else
try
Edge(Int.(nt))
catch
error("Fail to convert $(nt) to NTuple{$(length(nt)), Int}")
end
end
end
Edge(v::AbstractVector) = Edge(promote(v...))
Edge(vargs...) = Edge(vargs)
const Right = Edge(1, 0)
const Left = Edge(-1, 0)
const Up = Edge(0, 1)
const Down = Edge(0, -1)
# override functions
Base.getindex(e::AbstractEdge, i::Int) = e.data[i]
Base.iterate(e::AbstractEdge, state=1) = iterate(e.data, state)
Base.length(e::AbstractEdge) = length(e.data)
Base.size(e::AbstractEdge) = (length(e.data),)
function Base.getproperty(e::Edge, sym::Symbol)
if sym in fieldnames(Edge)
return getfield(e, sym)
elseif sym === :x
return e[1]
elseif sym === :y
return e[2]
else
error("invalid symbol $sym")
end
end
function Base.show(io::IO, mime::MIME"text/plain", e::AbstractEdge{T}) where {T}
println(io, e |> typeof)
for (i, (s, d)) in enumerate(zip([:x,:y], e))
println(io, "$s = $(e[i])")
end
end
# Define conversion and promotion
Base.promote_rule(a::Type{Edge{T1}}, b::Type{Edge{T2}}) where {T1, T2} = Edge{promote_type(T1, T2)}
Edge{T}(x::Edge{T2}) where {T, T2} = Edge{T}(x.data)
Edge(x::Edge) = x
Base.convert(::Type{Edge{T}}, x::Edge{T2}) where {T, T2} = Edge{T}(x)
# Operation
for op in [:+, :-, :*]
@eval function Base.$op(e1::AbstractEdge{T1}, e2::AbstractEdge{T2}) where {T1, T2}
promote_type(typeof(e1), typeof(e2))(broadcast($op, e1.data, e2.data))
end
end
abstract type AbstractPath{T<:Integer} end
struct Path{T} <: AbstractPath{T}
edges::Vector{Edge{T}}
origin::NTuple{2, T}
Path(edges::Vector{Edge{T}}) where {T} = new{T}(Edge{T}.(edges), NTuple{2, T}(zeros(T , 2)))
Path{T}(edges::Vector{Edge{T1}}) where {T, T1} = new{T}(Edge{T}.(edges), NTuple{2, T}(zeros(T, 2)))
Path(edges::Vector{Edge{T1}}, origin::NTuple{2, T2}) where {T1, T2} = new{promote_type(T1, T2)}(edges, origin)
Path{T}(edges::Vector{Edge{T1}}, origin) where {T, T1} = new{T}(Edge{T}.(edges), NTuple{2, T}(origin))
end
end
Edge = Goma.Edge
Path = Goma.Path
R = Right = Goma.Right
L = Left = Goma.Left
U = Up = Goma.Up
D = Down = Goma.Down;
# -
using Plots
# + tags=[]
edges = Edge{Int32}.([
[1,0],
R,
U,
L,
U,
[1,1],
R,
R,
[0,-1],
R,
])
path = Path(edges)
x, y = path.origin
xmin = xmax = x
ymin = ymax = y
p = plot(xlabel="x", ylabel="y")
for e in path.edges
quiver!(p, [x], [y], quiver=([e.x], [e.y]))
x += e.x
y += e.y
xmin = min(xmin, x + e.x)
ymin = min(ymin, y + e.y)
xmax = max(xmax, x + e.x)
ymax = max(ymax, y + e.y)
end
plot!(p, xticks = xmin:xmax, yticks = ymin:ymax)
p
# -
methods(Goma.Edge)
# +
module Kuma # 再編集しやすいように仮想モジュールを設定
abstract type AbstractEdge{T<:Integer} end
struct Edge{T} <: AbstractEdge{T}
data::NTuple{2, T}
# Automatically defined are
# Edge(data::Tuple{T, T}) where T
# Edge{T}(data)
# Edge(x::NTuple{2, T}) where T = new{T}(x)
# # To define Edge{Int}(1, 2)
# Edge{T}(x::Vararg{T2, 2}) where {T, T2} = new{T}(NTuple{2, T}(convert.(T, x)))
# # To define Edge{Int}([1, 2])
# Edge{T}(x::AbstractVector{T2}) where {T,T2} = new{T}(NTuple{2, T2}(x))
# # To define Edge{Int}((1,2))
# Edge{T}(x::Tuple) where T = new{T}(NTuple{2, T}(x))
end
Edge{T}(x, y) where T = Edge{T}((x, y))
Edge{T}(v::AbstractVector) where T = Edge{T}(v...)
function Edge(x::Tuple)
nt = NTuple(promote(x...))
if eltype(nt) <: Integer
Edge(nt)
else
try
Edge(Int.(nt))
catch
error("Fail to convert $(nt) to NTuple{$(length(nt)), Int}")
end
end
end
Edge(x, y) = Edge((x, y))
Edge(v::AbstractVector) = Edge(v...)
const Right = Edge(1, 0)
const Left = Edge(-1, 0)
const Up = Edge(0, 1)
const Down = Edge(0, -1)
# override functions
Base.getindex(e::AbstractEdge, i::Int) = e.data[i]
Base.iterate(e::AbstractEdge, state=1) = iterate(e.data, state)
Base.length(e::AbstractEdge) = length(e.data)
Base.size(e::AbstractEdge) = (length(e.data),)
function Base.getproperty(e::Edge, sym::Symbol)
if sym in fieldnames(Edge)
return getfield(e, sym)
elseif sym === :x
return e[1]
elseif sym === :y
return e[2]
else
error("invalid symbol $sym")
end
end
function Base.show(io::IO, mime::MIME"text/plain", e::AbstractEdge{T}) where {T}
println(io, e |> typeof)
for (i, (s, d)) in enumerate(zip([:x,:y], e))
println(io, "$s = $(e[i])")
end
end
# Define conversion and promotion
Base.promote_rule(a::Type{Edge{T1}}, b::Type{Edge{T2}}) where {T1, T2} = Edge{promote_type(T1, T2)}
Edge{T}(x::Edge) where T = Edge{T}(x.data)
Edge(x::Edge) = x
Base.convert(::Type{Edge{T}}, x::Edge{T2}) where {T, T2} = Edge{T}(x)
# Operation
for op in [:+, :-, :*]
@eval function Base.$op(e1::AbstractEdge{T1}, e2::AbstractEdge{T2}) where {T1, T2}
promote_type(typeof(e1), typeof(e2))(broadcast($op, e1.data, e2.data))
end
end
abstract type AbstractPath{T<:Integer} end
struct Path{T} <: AbstractPath{T}
edges::Vector{Edge{T}}
origin::NTuple{2, T}
Path(edges::Vector{Edge{T}}) where {T} = new{T}(Edge{T}.(edges), NTuple{2, T}(zeros(T , 2)))
Path{T}(edges::Vector{Edge{T1}}) where {T, T1} = new{T}(Edge{T}.(edges), NTuple{2, T}(zeros(T, 2)))
Path(edges::Vector{Edge{T1}}, origin::NTuple{2, T2}) where {T1, T2} = new{promote_type(T1, T2)}(edges, origin)
Path{T}(edges::Vector{Edge{T1}}, origin) where {T, T1} = new{T}(Edge{T}.(edges), NTuple{2, T}(origin))
end
end
Edge = Kuma.Edge
Path = Kuma.Path
R = Right = Kuma.Right
L = Left = Kuma.Left
U = Up = Kuma.Up
D = Down = Kuma.Down;
# +
edges = Edge{Int32}.([
[1,0],
R,
U,
L,
U,
[1,1],
R,
R,
[0,-1],
R,
])
path = Path(edges)
x, y = path.origin
xmin = xmax = x
ymin = ymax = y
p = plot(xlabel="x", ylabel="y")
for e in path.edges
quiver!(p, [x], [y], quiver=([e.x], [e.y]))
x += e.x
y += e.y
xmin = min(xmin, x + e.x)
ymin = min(ymin, y + e.y)
xmax = max(xmax, x + e.x)
ymax = max(ymax, y + e.y)
end
plot!(p, xticks = xmin:xmax, yticks = ymin:ymax)
p
# -
methods(Kuma.Edge) |> display
methods(Kuma.Edge{Int}) |> display
methods(Goma.Edge) |> display
methods(Goma.Edge{Int}) |> display
Kuma.Edge{Int}((1.0, 2.0))
@which Kuma.Edge{Int}((1.0, 2.0))
Goma.Edge{Int}((1.0, 2.0))
@which Goma.Edge{Int}((1, 2))
Kuma.Edge{Int}([1.0, 2.0])
@which Kuma.Edge{Int}([1.0, 2.0])
Goma.Edge{Int}([1.0, 2.0])
@which Goma.Edge{Int}([1.0, 2.0])
Kuma.Edge{Int}([1.0, 2.0]...)
@which Kuma.Edge{Int}([1.0, 2.0]...)
Goma.Edge{Int}([1.0, 2.0]...)
@which Goma.Edge{Int}([1.0, 2.0]...)
Kuma.Edge{Int}((1.0f0, 2.0)...)
@which Kuma.Edge{Int}((1.0f0, 2.0)...)
Goma.Edge{Int}((1.0f0, 2.0)...)
| 0024/draw_edge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
@author: mdigi14
"""
import pandas as pd
"""
Update Parameters Here
"""
FILE = "quaks"
PATH = "../metadata/rarity_data/{}_raritytools.csv".format(FILE)
MINT_PATH = "../minting_data/{}_minting.csv".format(FILE)
GRIFTER_ADDRESS = "0x111c26a02ca4050684d4083d72e2a7c1dcba853f"
TOP_N = 150
LOWER_BOUND = 0
UPPER_BOUND = 10000
# +
"""
Generate Report
"""
RARITY_DB = pd.read_csv(PATH)
RARITY_DB = RARITY_DB[RARITY_DB['TOKEN_ID'].duplicated()==False]
MINT_DB = pd.read_csv(MINT_PATH)
MINT_DB = MINT_DB[MINT_DB['to_account'] == GRIFTER_ADDRESS]
RARE_MINT_DB = MINT_DB[MINT_DB['rank'] < TOP_N]
ALT_DB = RARITY_DB[RARITY_DB['TOKEN_ID'] > LOWER_BOUND]
ALT_DB = ALT_DB[ALT_DB['TOKEN_ID'] < UPPER_BOUND]
RARE_DB = ALT_DB[ALT_DB['Rank'] < TOP_N]
rare_tokens_in_range = len(RARE_DB)
tokens_minted_by_grifter = len(list(MINT_DB['TOKEN_ID']))
rare_tokens_minted_by_grifter = len(RARE_MINT_DB)
grifter_mint_share = tokens_minted_by_grifter / (UPPER_BOUND - LOWER_BOUND)
grifter_rare_mint_share = rare_tokens_minted_by_grifter / rare_tokens_in_range
print("Project: ", FILE)
print("Grifter: ", GRIFTER_ADDRESS)
print("\n")
print("Total tokens in range: ", UPPER_BOUND - LOWER_BOUND)
print("Total rares in range: ", rare_tokens_in_range)
print("\n")
print("Total tokens minted by grifter: ", tokens_minted_by_grifter)
print("Rare tokens minted by grifter: ", rare_tokens_minted_by_grifter)
print("\n")
print("Rare tokens minted by grifter (by ID): ", list(RARE_MINT_DB['TOKEN_ID']))
print("\n")
print("Percentage of tokens in range minted by grifter: ", "{:.2%}".format(grifter_mint_share))
print("Percentage of rare tokens in range minted by grifter: ", "{:.2%}".format(grifter_rare_mint_share))
print("Multiple of rare token pct vs total token pct: ", round(grifter_rare_mint_share / grifter_mint_share))
| fair_drop/grifter_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.5.3
# language: ruby
# name: ruby
# ---
require 'daru/view'
Daru::View.plotting_library = :highcharts
# +
# scatter chart : basic
opts = {
chart: {
type: 'scatter',
zoomType: 'xy'
},
title: {
text: 'Height Versus Weight of 507 Individuals by Gender'
},
subtitle: {
text: 'Source: Heinz 2003'
},
xAxis: {
title: {
enabled: true,
text: 'Height (cm)'
},
startOnTick: true,
endOnTick: true,
showLastLabel: true
},
yAxis: {
title: {
text: 'Weight (kg)'
}
},
legend: {
layout: 'vertical',
align: 'left',
verticalAlign: 'top',
x: 100,
y: 70,
floating: true,
backgroundColor:" (Highcharts.theme && Highcharts.theme.legendBackgroundColor) || '#FFFFFF'".js_code,
borderWidth: 1
},
plotOptions: {
scatter: {
marker: {
radius: 5,
states: {
hover: {
enabled: true,
lineColor: 'rgb(100,100,100)'
}
}
},
states: {
hover: {
marker: {
enabled: false
}
}
},
tooltip: {
headerFormat: '<b>{series.name}</b><br>',
pointFormat: '{point.x} cm, {point.y} kg'
}
}
},
}
series_dt = [
{
name: 'Female',
color: 'rgba(223, 83, 83, .5)',
data: [[161.2, 51.6], [167.5, 59.0], [159.5, 49.2], [157.0, 63.0], [155.8, 53.6],
[170.0, 59.0], [159.1, 47.6], [166.0, 69.8], [176.2, 66.8], [160.2, 75.2],
[172.5, 55.2], [170.9, 54.2], [172.9, 62.5], [153.4, 42.0], [160.0, 50.0],
[147.2, 49.8], [168.2, 49.2], [175.0, 73.2], [157.0, 47.8], [167.6, 68.8],
[159.5, 50.6], [175.0, 82.5], [166.8, 57.2], [176.5, 87.8], [170.2, 72.8],
[174.0, 54.5], [173.0, 59.8], [179.9, 67.3], [170.5, 67.8], [160.0, 47.0],
[154.4, 46.2], [162.0, 55.0], [176.5, 83.0], [160.0, 54.4], [152.0, 45.8],
[162.1, 53.6], [170.0, 73.2], [160.2, 52.1], [161.3, 67.9], [166.4, 56.6],
[168.9, 62.3], [163.8, 58.5], [167.6, 54.5], [160.0, 50.2], [161.3, 60.3],
[167.6, 58.3], [165.1, 56.2], [160.0, 50.2], [170.0, 72.9], [157.5, 59.8],
[167.6, 61.0], [160.7, 69.1], [163.2, 55.9], [152.4, 46.5], [157.5, 54.3],
[168.3, 54.8], [180.3, 60.7], [165.5, 60.0], [165.0, 62.0], [164.5, 60.3],
[156.0, 52.7], [160.0, 74.3], [163.0, 62.0], [165.7, 73.1], [161.0, 80.0],
[162.0, 54.7], [166.0, 53.2], [174.0, 75.7], [172.7, 61.1], [167.6, 55.7],
[151.1, 48.7], [164.5, 52.3], [163.5, 50.0], [152.0, 59.3], [169.0, 62.5],
[164.0, 55.7], [161.2, 54.8], [155.0, 45.9], [170.0, 70.6], [176.2, 67.2],
[170.0, 69.4], [162.5, 58.2], [170.3, 64.8], [164.1, 71.6], [169.5, 52.8],
[163.2, 59.8], [154.5, 49.0], [159.8, 50.0], [173.2, 69.2], [170.0, 55.9],
[161.4, 63.4], [169.0, 58.2], [166.2, 58.6], [159.4, 45.7], [162.5, 52.2],
[159.0, 48.6], [162.8, 57.8], [159.0, 55.6], [179.8, 66.8], [162.9, 59.4],
[161.0, 53.6], [151.1, 73.2], [168.2, 53.4], [168.9, 69.0], [173.2, 58.4],
[171.8, 56.2], [178.0, 70.6], [164.3, 59.8], [163.0, 72.0], [168.5, 65.2],
[166.8, 56.6], [172.7, 105.2], [163.5, 51.8], [169.4, 63.4], [167.8, 59.0],
[159.5, 47.6], [167.6, 63.0], [161.2, 55.2], [160.0, 45.0], [163.2, 54.0],
[162.2, 50.2], [161.3, 60.2], [149.5, 44.8], [157.5, 58.8], [163.2, 56.4],
[172.7, 62.0], [155.0, 49.2], [156.5, 67.2], [164.0, 53.8], [160.9, 54.4],
[162.8, 58.0], [167.0, 59.8], [160.0, 54.8], [160.0, 43.2], [168.9, 60.5],
[158.2, 46.4], [156.0, 64.4], [160.0, 48.8], [167.1, 62.2], [158.0, 55.5],
[167.6, 57.8], [156.0, 54.6], [162.1, 59.2], [173.4, 52.7], [159.8, 53.2],
[170.5, 64.5], [159.2, 51.8], [157.5, 56.0], [161.3, 63.6], [162.6, 63.2],
[160.0, 59.5], [168.9, 56.8], [165.1, 64.1], [162.6, 50.0], [165.1, 72.3],
[166.4, 55.0], [160.0, 55.9], [152.4, 60.4], [170.2, 69.1], [162.6, 84.5],
[170.2, 55.9], [158.8, 55.5], [172.7, 69.5], [167.6, 76.4], [162.6, 61.4],
[167.6, 65.9], [156.2, 58.6], [175.2, 66.8], [172.1, 56.6], [162.6, 58.6],
[160.0, 55.9], [165.1, 59.1], [182.9, 81.8], [166.4, 70.7], [165.1, 56.8],
[177.8, 60.0], [165.1, 58.2], [175.3, 72.7], [154.9, 54.1], [158.8, 49.1],
[172.7, 75.9], [168.9, 55.0], [161.3, 57.3], [167.6, 55.0], [165.1, 65.5],
[175.3, 65.5], [157.5, 48.6], [163.8, 58.6], [167.6, 63.6], [165.1, 55.2],
[165.1, 62.7], [168.9, 56.6], [162.6, 53.9], [164.5, 63.2], [176.5, 73.6],
[168.9, 62.0], [175.3, 63.6], [159.4, 53.2], [160.0, 53.4], [170.2, 55.0],
[162.6, 70.5], [167.6, 54.5], [162.6, 54.5], [160.7, 55.9], [160.0, 59.0],
[157.5, 63.6], [162.6, 54.5], [152.4, 47.3], [170.2, 67.7], [165.1, 80.9],
[172.7, 70.5], [165.1, 60.9], [170.2, 63.6], [170.2, 54.5], [170.2, 59.1],
[161.3, 70.5], [167.6, 52.7], [167.6, 62.7], [165.1, 86.3], [162.6, 66.4],
[152.4, 67.3], [168.9, 63.0], [170.2, 73.6], [175.2, 62.3], [175.2, 57.7],
[160.0, 55.4], [165.1, 104.1], [174.0, 55.5], [170.2, 77.3], [160.0, 80.5],
[167.6, 64.5], [167.6, 72.3], [167.6, 61.4], [154.9, 58.2], [162.6, 81.8],
[175.3, 63.6], [171.4, 53.4], [157.5, 54.5], [165.1, 53.6], [160.0, 60.0],
[174.0, 73.6], [162.6, 61.4], [174.0, 55.5], [162.6, 63.6], [161.3, 60.9],
[156.2, 60.0], [149.9, 46.8], [169.5, 57.3], [160.0, 64.1], [175.3, 63.6],
[169.5, 67.3], [160.0, 75.5], [172.7, 68.2], [162.6, 61.4], [157.5, 76.8],
[176.5, 71.8], [164.4, 55.5], [160.7, 48.6], [174.0, 66.4], [163.8, 67.3]]
}, {
name: 'Male',
color: 'rgba(119, 152, 191, .5)',
data: [[174.0, 65.6], [175.3, 71.8], [193.5, 80.7], [186.5, 72.6], [187.2, 78.8],
[181.5, 74.8], [184.0, 86.4], [184.5, 78.4], [175.0, 62.0], [184.0, 81.6],
[180.0, 76.6], [177.8, 83.6], [192.0, 90.0], [176.0, 74.6], [174.0, 71.0],
[184.0, 79.6], [192.7, 93.8], [171.5, 70.0], [173.0, 72.4], [176.0, 85.9],
[176.0, 78.8], [180.5, 77.8], [172.7, 66.2], [176.0, 86.4], [173.5, 81.8],
[178.0, 89.6], [180.3, 82.8], [180.3, 76.4], [164.5, 63.2], [173.0, 60.9],
[183.5, 74.8], [175.5, 70.0], [188.0, 72.4], [189.2, 84.1], [172.8, 69.1],
[170.0, 59.5], [182.0, 67.2], [170.0, 61.3], [177.8, 68.6], [184.2, 80.1],
[186.7, 87.8], [171.4, 84.7], [172.7, 73.4], [175.3, 72.1], [180.3, 82.6],
[182.9, 88.7], [188.0, 84.1], [177.2, 94.1], [172.1, 74.9], [167.0, 59.1],
[169.5, 75.6], [174.0, 86.2], [172.7, 75.3], [182.2, 87.1], [164.1, 55.2],
[163.0, 57.0], [171.5, 61.4], [184.2, 76.8], [174.0, 86.8], [174.0, 72.2],
[177.0, 71.6], [186.0, 84.8], [167.0, 68.2], [171.8, 66.1], [182.0, 72.0],
[167.0, 64.6], [177.8, 74.8], [164.5, 70.0], [192.0, 101.6], [175.5, 63.2],
[171.2, 79.1], [181.6, 78.9], [167.4, 67.7], [181.1, 66.0], [177.0, 68.2],
[174.5, 63.9], [177.5, 72.0], [170.5, 56.8], [182.4, 74.5], [197.1, 90.9],
[180.1, 93.0], [175.5, 80.9], [180.6, 72.7], [184.4, 68.0], [175.5, 70.9],
[180.6, 72.5], [177.0, 72.5], [177.1, 83.4], [181.6, 75.5], [176.5, 73.0],
[175.0, 70.2], [174.0, 73.4], [165.1, 70.5], [177.0, 68.9], [192.0, 102.3],
[176.5, 68.4], [169.4, 65.9], [182.1, 75.7], [179.8, 84.5], [175.3, 87.7],
[184.9, 86.4], [177.3, 73.2], [167.4, 53.9], [178.1, 72.0], [168.9, 55.5],
[157.2, 58.4], [180.3, 83.2], [170.2, 72.7], [177.8, 64.1], [172.7, 72.3],
[165.1, 65.0], [186.7, 86.4], [165.1, 65.0], [174.0, 88.6], [175.3, 84.1],
[185.4, 66.8], [177.8, 75.5], [180.3, 93.2], [180.3, 82.7], [177.8, 58.0],
[177.8, 79.5], [177.8, 78.6], [177.8, 71.8], [177.8, 116.4], [163.8, 72.2],
[188.0, 83.6], [198.1, 85.5], [175.3, 90.9], [166.4, 85.9], [190.5, 89.1],
[166.4, 75.0], [177.8, 77.7], [179.7, 86.4], [172.7, 90.9], [190.5, 73.6],
[185.4, 76.4], [168.9, 69.1], [167.6, 84.5], [175.3, 64.5], [170.2, 69.1],
[190.5, 108.6], [177.8, 86.4], [190.5, 80.9], [177.8, 87.7], [184.2, 94.5],
[176.5, 80.2], [177.8, 72.0], [180.3, 71.4], [171.4, 72.7], [172.7, 84.1],
[172.7, 76.8], [177.8, 63.6], [177.8, 80.9], [182.9, 80.9], [170.2, 85.5],
[167.6, 68.6], [175.3, 67.7], [165.1, 66.4], [185.4, 102.3], [181.6, 70.5],
[172.7, 95.9], [190.5, 84.1], [179.1, 87.3], [175.3, 71.8], [170.2, 65.9],
[193.0, 95.9], [171.4, 91.4], [177.8, 81.8], [177.8, 96.8], [167.6, 69.1],
[167.6, 82.7], [180.3, 75.5], [182.9, 79.5], [176.5, 73.6], [186.7, 91.8],
[188.0, 84.1], [188.0, 85.9], [177.8, 81.8], [174.0, 82.5], [177.8, 80.5],
[171.4, 70.0], [185.4, 81.8], [185.4, 84.1], [188.0, 90.5], [188.0, 91.4],
[182.9, 89.1], [176.5, 85.0], [175.3, 69.1], [175.3, 73.6], [188.0, 80.5],
[188.0, 82.7], [175.3, 86.4], [170.5, 67.7], [179.1, 92.7], [177.8, 93.6],
[175.3, 70.9], [182.9, 75.0], [170.8, 93.2], [188.0, 93.2], [180.3, 77.7],
[177.8, 61.4], [185.4, 94.1], [168.9, 75.0], [185.4, 83.6], [180.3, 85.5],
[174.0, 73.9], [167.6, 66.8], [182.9, 87.3], [160.0, 72.3], [180.3, 88.6],
[167.6, 75.5], [186.7, 101.4], [175.3, 91.1], [175.3, 67.3], [175.9, 77.7],
[175.3, 81.8], [179.1, 75.5], [181.6, 84.5], [177.8, 76.6], [182.9, 85.0],
[177.8, 102.5], [184.2, 77.3], [179.1, 71.8], [176.5, 87.9], [188.0, 94.3],
[174.0, 70.9], [167.6, 64.5], [170.2, 77.3], [167.6, 72.3], [188.0, 87.3],
[174.0, 80.0], [176.5, 82.3], [180.3, 73.6], [167.6, 74.1], [188.0, 85.9],
[180.3, 73.2], [167.6, 76.3], [183.0, 65.9], [183.0, 90.9], [179.1, 89.1],
[170.2, 62.3], [177.8, 82.7], [179.1, 79.1], [190.5, 98.2], [177.8, 84.1],
[180.3, 83.2], [180.3, 83.2]]
}
]
scatter_basic = Daru::View::Plot.new(series_dt, opts)
scatter_basic.show_in_iruby
# +
# note : dependent js is highcharts-more.js not included in IRuby right now
# bubble chart : basic
opts = {
chart: {
type: 'bubble',
plotBorderWidth: 1,
zoomType: 'xy'
},
modules: ['highcharts-more'],
legend: {
enabled: false
},
title: {
text: 'Sugar and fat intake per country'
},
subtitle: {
text: 'Source: <a href="http://www.euromonitor.com/">Euromonitor</a> and <a href="https://data.oecd.org/">OECD</a>'
},
xAxis: {
gridLineWidth: 1,
title: {
text: 'Daily fat intake'
},
labels: {
format: '{value} gr'
},
plotLines: [{
color: 'black',
dashStyle: 'dot',
width: 2,
value: 65,
label: {
rotation: 0,
y: 15,
style: {
fontStyle: 'italic'
},
text: 'Safe fat intake 65g/day'
},
zIndex: 3
}]
},
yAxis: {
startOnTick: false,
endOnTick: false,
title: {
text: 'Daily sugar intake'
},
labels: {
format: '{value} gr'
},
maxPadding: 0.2,
plotLines: [{
color: 'black',
dashStyle: 'dot',
width: 2,
value: 50,
label: {
align: 'right',
style: {
fontStyle: 'italic'
},
text: 'Safe sugar intake 50g/day',
x: -10
},
zIndex: 3
}]
},
tooltip: {
useHTML: true,
headerFormat: '<table>',
pointFormat: '<tr><th colspan="2"><h3>{point.country}</h3></th></tr>' +
'<tr><th>Fat intake:</th><td>{point.x}g</td></tr>' +
'<tr><th>Sugar intake:</th><td>{point.y}g</td></tr>' +
'<tr><th>Obesity (adults):</th><td>{point.z}%</td></tr>',
footerFormat: '</table>',
followPointer: true
},
plotOptions: {
series: {
dataLabels: {
enabled: true,
format: '{point.name}'
}
}
},
}
series_dt = [
{
data: [
{ x: 95, y: 95, z: 13.8, name: 'BE', country: 'Belgium' },
{ x: 86.5, y: 102.9, z: 14.7, name: 'DE', country: 'Germany' },
{ x: 80.8, y: 91.5, z: 15.8, name: 'FI', country: 'Finland' },
{ x: 80.4, y: 102.5, z: 12, name: 'NL', country: 'Netherlands' },
{ x: 80.3, y: 86.1, z: 11.8, name: 'SE', country: 'Sweden' },
{ x: 78.4, y: 70.1, z: 16.6, name: 'ES', country: 'Spain' },
{ x: 74.2, y: 68.5, z: 14.5, name: 'FR', country: 'France' },
{ x: 73.5, y: 83.1, z: 10, name: 'NO', country: 'Norway' },
{ x: 71, y: 93.2, z: 24.7, name: 'UK', country: 'United Kingdom' },
{ x: 69.2, y: 57.6, z: 10.4, name: 'IT', country: 'Italy' },
{ x: 68.6, y: 20, z: 16, name: 'RU', country: 'Russia' },
{ x: 65.5, y: 126.4, z: 35.3, name: 'US', country: 'United States' },
{ x: 65.4, y: 50.8, z: 28.5, name: 'HU', country: 'Hungary' },
{ x: 63.4, y: 51.8, z: 15.4, name: 'PT', country: 'Portugal' },
{ x: 64, y: 82.9, z: 31.3, name: 'NZ', country: 'New Zealand' }
]
}
]
bubble_basic = Daru::View::Plot.new(series_dt, opts)
bubble_basic.show_in_iruby
# highcharts-more.js needed, otherwise chart will not be displayed
# +
# highcharts-more.js needed, otherwise chart will not be displayed
# buuble chart : bubble 3d
opts = {
chart: {
type: 'bubble',
plotBorderWidth: 1,
zoomType: 'xy'
},
title: {
text: 'Highcharts bubbles with radial gradient fill'
},
xAxis: {
gridLineWidth: 1
},
yAxis: {
startOnTick: false,
endOnTick: false
},
}
series_dt = [
{
data: [
[9, 81, 63],
[98, 5, 89],
[51, 50, 73],
[41, 22, 14],
[58, 24, 20],
[78, 37, 34],
[55, 56, 53],
[18, 45, 70],
[42, 44, 28],
[3, 52, 59],
[31, 18, 97],
[79, 91, 63],
[93, 23, 23],
[44, 83, 22]
],
# beloe code not working
marker: {
fillColor: {
# radialGradient: { cx: 0.4, cy: 0.3, r: 0.7 },
# stops: [
# [0, 'rgba(255,255,255,0.5)'],
# [1, "Highcharts.Color(Highcharts.getOptions().colors[0]).setOpacity(0.5).get('rgba')".js_code]
# ]
}
}
}, {
data: [
[42, 38, 20],
[6, 18, 1],
[1, 93, 55],
[57, 2, 90],
[80, 76, 22],
[11, 74, 96],
[88, 56, 10],
[30, 47, 49],
[57, 62, 98],
[4, 16, 16],
[46, 10, 11],
[22, 87, 89],
[57, 91, 82],
[45, 15, 98]
],
marker: {
fillColor: {
# radialGradient: { cx: 0.4, cy: 0.3, r: 0.7 },
# stops: [
# [0, 'rgba(255,255,255,0.5)'],
# [1, "Highcharts.Color(Highcharts.getOptions().colors[1]).setOpacity(0.5).get('rgba')".js_code]
# ]
}
}
}
]
bubble_3d= Daru::View::Plot.new(series_dt, opts)
bubble_3d.show_in_iruby
# -
| spec/dummy_iruby/Highcharts - scatter -bubble charts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Poi of Seattle
# +
import pandas as pd
from searchpoi import SearchPoi
import numpy as np
import matplotlib.pyplot as plt
df = pd.read_csv('./listings.csv')
# -
# We import Poi of Seattle using [Overpass API](https://wiki.openstreetmap.org/wiki/Overpass_API).
p = SearchPoi("Seattle")
# For each host of Airbnb, we find a summary of the closest points (a buffer of 5km) regrouped by [tag](https://wiki.openstreetmap.org/wiki/Map_Features). This operation is complex for CPU and takes much time.
r = []
for index,row in df.iterrows():
r.append(p.closest(float(row['longitude']),float(row['latitude'])))
# In the precedent row, we had used a list to reduce the overhead of data frame concatenation. Now we are proceeding to create a data frame with the correct index
r = pd.concat(r, axis = 0)
r.set_index(df.index, inplace = True)
# We save the data frame
r.to_csv('./seattle_poi.csv')
| ImportSeattlePoi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TODO:
# - Clean (see other notebook)
# - Create "distance from Seattle" feature
# +
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import scipy.stats as st
# import pymc3 as pm
import seaborn as sns
# enables inline plots, without it plots don't show up in the notebook
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
# # %config InlineBackend.figure_format = 'png'
# mpl.rcParams['figure.dpi']= 300
# -
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', 25)
pd.set_option('display.precision', 3)
pd.set_option('display.float_format', lambda x: '%.3f' % x)
import pickle
def load_pd_pkl(file):
import pandas as pd
try:
with open(f"{file}.pkl",'rb') as picklefile:
return pickle.load(picklefile)
except FileNotFoundError:
df = pd.read_csv(f"{file}.csv")
with open(f"{file}.pkl", 'wb') as picklefile:
pickle.dump(df, picklefile)
return df
df = load_pd_pkl('data/raw_wta_df')
# +
#df.to_csv('data/raw_wta_df.csv')
# -
df.shape
df.reset_index(drop=True, inplace=True)
#rename columns
df['votes'] = df['votes'].astype(int)
df['countreports'] = df['countreports'].astype(int)
df['rating'] = df['rating'].astype(float)
df['length'] = df['length'].astype(float)
df['gain'] = df['gain'].astype(float)
df['hpoint'] = df['hpoint'].astype(float)
df['lat'] = df['lat'].astype(float)
df['long'] = df['long'].astype(float)
df.info()
# Clean hikes which faultily have the 'hpoint' assigned to the "length" to NaN
df.loc[df['length'] == df['hpoint'], 'length'] = np.nan
# Good! All length types exist with a length preceding it.
df[df['length'].notna() & df['length'].isna()]
# +
# Accurately update length type of "of" to "miles_of_trails" like on website
df[df.lengthtype == "of"]
df.loc[df['lengthtype'] == "of", 'lengthtype'] = "miles_of_trails"
# Clean hikes which faultily have the 'gain assigned to the "length" to NaN
df.loc[df['length'] == df['gain'], 'length'] = np.nan
df[df.length == df.gain].shape
# rename author1 and author2
df.rename(columns={'author1': 'org_author', "author2":"author"},inplace=True)
# do some renaming of dfs and create new one dropping a few columns and all NaNs
old_df = df.copy()
df = df.drop(columns=['trailhead1', 'trailhead2','org_author', 'author', 'subregion','lat','long'])
df.dropna(inplace=True)
# drop hikes without any votes and ratings remaining
df = df.loc[df['votes'] != 0, :]
# -
# There are 7 passes, and also none and n/a
df.fee.unique()
# +
##### consider limitations on extreme values in length and gain at this point
# +
##### should I require votes to be above a certain threshold? 1-5?
# -
# # Assumptions and changes to data on first pass:
# * Must have 3 votes are greater
# * Remove "one-way" hikes
# * Remove extreme continuous variables (length, high-point, elevation gain)
Run below (commented out) tweaks to make tweaks
# +
# df = df[df.votes > 3] # Remove hikes with 3 or less votes contributing to rating
# df = df[df.length > 30] # Remove hikes greater than 30 miles
# df = df[df.gain > 6000] # Remove hikes greater than 6000 feet in vertical gain
# df = df[df.lengthtype != "one-way"] # Remove one way hikes
# -
# # To-Do
# * Feature engineer lat/long
# * Change and/or remove one-way and miles of trails locations.
# +
# import pickle
# with open('data/cleaned_wta_df.pkl', 'wb') as picklefile:
# pickle.dump(df, picklefile)
# -
| wta_data_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ensemble Methods for Machine Learning
# In this notebook, we'll take a look on how to implement three different ensemble learning algorithms, using some functionatilies from numpy and sklearn. The algorithms are: AdaBoost, Gradient Boosting and Random Forests.
# ## AdaBoost
# The AdaBoost is an ensemble model for supervised learning first described by <NAME> and <NAME>. The algorithm consists of a boosting technique, that takes a weak classifier,e.g., decision bump, and improves its performance by applying weights to each sucessive prediction on each iteration. At the end of the iteration loop, the algorithm makes a weighted "vote" on the final prediction. This implementation of the AdaBoost only works for classification problems with $y \in \{-1,1\} $.
#
# Dependencies
import numpy as np
from sklearn.tree import DecisionTreeClassifier
# ### Implementation
# +
def misclassification(y,prediction):
"""
Calculates the number of misclassifications between the prediction and the output.
Inputs:
y: int
Input a single value of the output variable y.
prediction: int
Input a single valut of the predicted output.
Returns:
misclassifications: array
Output 1 if the values do not match and 0 if they do.
"""
y=y.reshape((-1,1))
prediction = prediction.reshape((-1,1))
misclassifications = 1*(y != prediction)
return misclassifications #returns the number of misclassifications on the prediction
class AdaBoostClassifier:
"""
AdaBoost algorithm for weak classifiers, that can fit discrete classification problems.
Methods:
fit(x,y) -> Performs the boosting algorithm on the training set(x,y).
predict(x) -> Predict class for X.
get_tree_weights() -> Returns the weights for each of the n_iter trees generated during the boosting task.
"""
def __init__(self,n_estimators):
"""
Initialize self.
Inputs:
n_estimators: int
input the number of trees(stumps) to grow.
"""
self.n_estimators = n_estimators
def fit(self,X,y):
"""
Fits the AdaBooster on a given dataset.
Inputs:
X: array
input the array of input points
y: array
input the array of output points, with y E {-1,1}
"""
self.input_train = X
self.output_train = y
alphas = np.zeros((self.n_estimators,1))
predictions_train = np.zeros((len(self.output_train),self.n_estimators))
staged_weights = np.zeros((len(self.output_train),self.n_estimators))
weighted_error = np.zeros((self.n_estimators,1))
stumps = list()
for i in range(len(self.output_train)):
staged_weights[i,0] = 1/len(self.output_train) #initialized the weights with value 1/num_samples
for m in range(self.n_estimators):
curr_weights = staged_weights[:,m] #current staged weights
stump = DecisionTreeClassifier(max_depth=1, max_leaf_nodes=2)
stump = stump.fit(self.input_train,self.output_train,sample_weight=curr_weights) # fits a decision stump with curr_weights as the sample weight
predictions_train[:,m] = stump.predict(self.input_train) # stores the current predictions
curr_weights = curr_weights.reshape((-1,1))
weighted_misclassification = curr_weights * misclassification(self.output_train,predictions_train[:,m]) #sets the weighted misclassification as the product between the current weights and the misclassification between the current prediction and the original output
weighted_error[m] = np.sum(weighted_misclassification)/np.sum(curr_weights) #calculates the weighted error
alphas[m] = np.log((1 - weighted_error[m])/weighted_error[m]) # calculates the constants alpha
curr_weights = curr_weights * np.exp(alphas[m] * misclassification(self.output_train,predictions_train[:,m])) #sets the new weights
curr_weights = curr_weights.reshape((-1,)) # reshapes it to be 1D
if m + 1 < self.n_estimators: # if this iteration isn't the last one, sets the weights for the next iteration
staged_weights[:,m+1] = curr_weights
stumps.append(stump)
self.tree_weights = alphas
self.stumps_list = stumps
self.weighted_error = weighted_error
def predict(self,x):
"""
Makes a prediction based on the weights and classifiers calculated by the Adabooster
Inputs:
x: array_like
input the array of input points
Returns:
self.predict: array_like
outputs the array of predictions
"""
indiv_predictions = np.array([self.classifier.predict(x) for self.classifier in self.stumps_list])
prediction = indiv_predictions.T @ self.tree_weights #multiplies each tree prediction by the trees calculated weight
prediction = np.sign(prediction) #the adaboost final prediction will be the sign of the previous operation.
return prediction
def get_tree_weights(self):
"""
Gives the weights calculated by the AdaBooster
"""
return self.tree_weights
# -
# ## Gradient Boosting
# The Gradient Boosting is an ensemble model for supervised learning first described by <NAME>. It consists of a boosting model, that iteratively calculates the generalized(or partial) residuals at each iteration M, and then fits a new decision tree(regression or classification) targeting these residuals. The algorithm also finds the constant values(gamma) for each split on the decision tree that minimizes the inserted loss function. This model differs from AdaBoost in the sense that it allows for any type of loss function to be used, so long as the function is differentiable. This means that the Gradient Boosting algorithm can fit any type of classification or regression task, which is a big improvement with respect to AdaBoost, since it's most basic implementation can only fit classification problems with target values $y \in \{-1,1\} $. There are versions of AdaBoost that allow for regression tasks, but the model itself is based on the aforementioned assumption about the nature of the output set, so it is far more limited than the Gradient Boosting technique. In this example, we'll be implementing a Gradient Boosted Regressor.
# Dependencies
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.tree._tree import TREE_LEAF
# ### Implementation
# +
def residuals_func(y,prediction,loss_func):
"""
Calcultes the generalized(or partial) residuals between the prediction and the output. The residuals are defined to be the negative value of the gradient of the loss function w.r.t. prediction.
Inputs:
y: array
Input the array of outputs.
prediction: array
Input the array of predictions.
loss_func: string
Input the string that identifies the loss function to use when calculating the residuals; loss_func can be 'multinomial deviance'(multinomial deviance loss), 'entropy'(Cross-entropy loss), 'mse'(Mean Squared Error), 'mae'(Mean Absolute error).
"""
possible_loss_funcs = ['mse','mae']
assert loss_func in possible_loss_funcs
if loss_func == possible_loss_funcs[0]:
return (y - prediction)
else:
return (2*(y - prediction) - 1).reshape((-1,1))
def optimal_gamma(y,prediction,loss_func):
"""
Calculates the optimal value for the gamma constant based on a certain loss_func.
Inputs:
y: array
input the array of outputs/targets.
prediction: array
input the array of predictions.
loss_func: string
input the string that identifies the loss function to be used; loss_func can be: 'mse'(Mean Squared Error) or 'mae'(Mean Absolute Error)
"""
assert loss_func in ["mse","mae"]
if loss_func == 'mae':
res = y - prediction
return np.median(res)
elif loss_func == 'mse':
res = y - prediction
return np.mean(res)
class GBRegressor:
"""
GradientBoost algorithm for supervised learning, that can fit regression problems.
Methods:
fit(X,y) -> Performs the gradient boosting algorithm on the training set(x,y).
predict(x) -> Predict value for X.
"""
def __init__(self,n_estimators,loss_func,max_depth=None,random_state=None):
"""
Inilialize self.
Inputs:
n_estimators: int
input the number of trees to grow.
loss_func: string
input the string that identifies the loss function to use when calculating the residuals; loss_func can be 'mse'(Mean Squared Error), 'mae'(Mean Absolute error).
max_depth: int
input the maximum depth of the tree; default is set to None.
random_state: int
input the random_state to be used on the sklearn DecisionTreeClassifier; default is set to None.
"""
possible_params = ['mse', 'mae']
assert n_estimators > 0
assert loss_func in possible_params
if max_depth != None:
assert max_depth >= 1
self.n_estimators = n_estimators
self.loss_func = loss_func
self.random_state = random_state
self.max_depth = max_depth
def fit(self,X,y):
"""
Fits the GradientBooster model.
Inputs:
X: array
input array of input points.
y: array
input array of output points.
"""
self.input_train = X
self.output_train = y
self.trained_trees_list = list()
self.gammas = list()
self.gamma_0 = optimal_gamma(self.output_train,np.zeros(self.output_train.shape[0]),self.loss_func) #initializes gamma as the optimal value between the output and a vector of zeroes
raw_pred = np.ones((self.output_train.shape[0])) * self.gamma_0 #makes the initial prediction
for m in range(self.n_estimators):
residuals = residuals_func(self.output_train,raw_pred,self.loss_func) # calculates the residuals between the initial prediction and the output
model = DecisionTreeRegressor(criterion = self.loss_func, random_state = self.random_state,max_depth=self.max_depth)
tree = model.fit(self.input_train,residuals) #fits a tree targeting those residuals
terminal_regions = tree.apply(self.input_train) # gets terminal nodes for the tree
gamma = np.zeros((len(tree.tree_.children_left))) # generates a gamma vector, with size = number of terminal nodes
for leaf in np.where(tree.tree_.children_left == TREE_LEAF)[0]: # searches through the tree for terminal nodes(leafs)
mask = np.where(terminal_regions == leaf) # stores the position of each leaf
gamma[leaf] = optimal_gamma(self.output_train[mask],raw_pred[mask],self.loss_func) #finds the best gamma for each leaf
raw_pred += gamma[terminal_regions] # those gammas are then summed to the initial prediction
self.trained_trees_list.append(tree)
self.gammas.append(gamma)
def predict(self,x):
"""
Predicts the value or class of a given group of inputs points based on the trained trees.
Inputs:
x: array_like
input the input point/array to be predicted.
Returns:
final_prediction: array_like
outputs the class/value prediction of the input made by the gradient booster model.
"""
raw_pred = np.ones(x.shape[0])* self.gamma_0
for tree,gamma in zip(self.trained_trees_list,self.gammas):
terminal_regions = tree.apply(x)
raw_pred += gamma[terminal_regions] #the final prediction of the gradient boosting regressor will be the initial predictions summed with all the optimal gammas found for all leafs
return raw_pred
# -
# ## Random Forests
# The Random Forests is an ensemble model for supervised learning first described by Breiman (2004). It uses bagging to build a group of subsets of the initial dataset, and then builds a tree(decision or regression) for each subset, by randomly choosing a group of features in each subset, so that n_features_used < n_features_total. Here I implement two different Random Forest models, for classification and regression. These implementations are built on top of the sklearn and numpy libraries.
# Dependencies
import numpy as np
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import DecisionTreeRegressor
# ### Implementation
# #### Random Forests for Classification
# +
class RandomForestClassifier:
"""
Class of the Random Forest Classifier Model.
Methods:
fit(X,y) -> Performs the random forests algorithm on the training set(x,y).
predict(x) -> Predict class for X.
"""
def __init__(self,n_estimators,n_classes,max_depth=None,criterion='gini',random_state=None,max_features='sqrt'):
"""
Initialize self.
Inputs:
n_estimators: int
input the number of trees to grow.
n_classes: int
input the number of classes of the classification task.
max_depth: int
input the maximum depth of the tree; default is set to None.
criterion: string
input string that identifies the criterion to be used when deciding how to split each node. criterion can be: 'gini' or 'entropy' .default is set to 'gini'.
max_features = string or int/float:
input string or int/float that identifies the maximum number of features to be used when splitting each decision tree; if string can be: 'sqrt' or 'log2'; if int max_fetures will be the maximum number of features; if float the maximum number of features will be int(max_features * n_features).
random_state: int
input the random_state to be used on the sklearn DecisionTreeClassifier. default is set to None.
"""
self.n_estimators = n_estimators
self.criterion = criterion
self.random_state = random_state
self.n_classes = n_classes
self.max_features = max_features
self.max_depth = max_depth
possible_criterion = ['gini','entropy']
assert self.criterion in possible_criterion
def fit(self,X,y):
"""
Fits the RandomForestClassifier model.
Inputs:
X: array
input array of input points.
y: array
input array of output points.
"""
self.input_train = X
self.output_train = y.reshape((-1,1))
self.trained_trees_list = list()
for i in range(self.n_estimators):
train_inds = np.random.choice(int(self.input_train.shape[0]/2),int(self.input_train.shape[0]/2),False) #generates the indices for a bootstrap sample with size N/2 and with values that don't repeat(doing this improves the efficiency of the algorithm and does not deprecate performance)
model_tree = DecisionTreeClassifier(criterion = self.criterion, random_state = self.random_state,max_features = self.max_features,max_depth=self.max_depth)
model_tree = model_tree.fit(self.input_train[train_inds,:],self.output_train[train_inds,:]) #fits a decision tree with the bootstrap sample
self.trained_trees_list.append(model_tree)
def predict(self,x):
"""
Predicts the class of a given group of inputs points based on the trained trees.
Inputs:
x: array_like
input the input point/array to be predicted.
Returns:
final_prediction: array_like
outputs the class prediction of the input made by the random forest model.
"""
indiv_predictions = np.array([self.classifier.predict(x) for self.classifier in self.trained_trees_list]).T
final_prediction = np.zeros((indiv_predictions.shape[0],))
counter_vec = np.zeros((self.n_classes,1))
for i in range(indiv_predictions.shape[0]):
for j in range(indiv_predictions.shape[1]):
counter_vec[indiv_predictions[i][j]] += 1
final_prediction[i] = np.argmax(counter_vec) #the final prediction of the random forest classifier will be a majority vote made w.r.t. to all trees,i.e., for each output point, predict the class that appears the most on the group of all prediction made by the forest
counter_vec = np.zeros((counter_vec.shape))
return final_prediction
# -
# #### Random Forests for Regression
class RandomForestRegressor:
"""
Class of the Random Forest Classifier Model.
Methods:
fit(X,y) -> Performs the random forests algorithm on the training set(x,y).
predict(x) -> Predict regression value for X.
"""
def __init__(self,n_estimators,max_features =1/3,max_depth=None,criterion='mse',random_state=None):
"""
Initialize self.
Inputs:
n_estimators: int
input the number of trees to grow.
max_depth: int
input the maximum depth of the tree; default is set to None.
criterion: string
input string that identifies the criterion to be used when deciding how to split each node. criterion can be: 'mse', 'friedman_mse' and 'mae' .default is set to 'mse'.
max_features = string or int/float:
input string or int/float that identifies the maximum number of features to be used when splitting each decision tree; if string can be: 'sqrt' or 'log2'; if int max_fetures will be the maximum number of features; if float the maximum number of features will be int(max_features * n_features); default is set to 1/3.
random_state: int
input the random_state to be used on the sklearn DecisionTreeClassifier. default is set to None.
"""
self.n_estimators = n_estimators
self.criterion = criterion
self.random_state = random_state
self.max_features = max_features
self.max_depth = max_depth
possible_criterion = ['mse', 'friedman_mse', 'mae']
assert self.criterion in possible_criterion
def fit(self,X,y):
"""
Fits the RandomForestRegressor model.
Inputs:
X: array
input array of input points.
y: array
input array of output points.
"""
self.input_train = X
self.output_train = y.reshape((-1,1))
self.trained_trees_list = list()
for i in range(self.n_estimators):
train_inds = np.random.choice(self.input_train.shape[0],self.input_train.shape[0],True) #generates the indices for a bootstrap sample with size N aand with repeating values(in the case of regression, there is a slight increase in error when using the technique described in the Classifier code)
model_tree = DecisionTreeRegressor(criterion = self.criterion, random_state = self.random_state,max_features = self.max_features,max_depth=self.max_depth)
model_tree = model_tree.fit(self.input_train[train_inds,:],self.output_train[train_inds,:]) #fits a decision tree with the bootstrap sample
self.trained_trees_list.append(model_tree)
def predict(self,x):
"""
Predicts the value of a given group of inputs points based on the trained trees.
Inputs:
x: array_like
input the input point/array to be predicted.
Returns:
final_prediction: array_like
outputs the value prediction of the input made by the random forest model.
"""
indiv_predictions = np.array([self.classifier.predict(x) for self.classifier in self.trained_trees_list]).T
final_prediction = np.zeros((indiv_predictions.shape[0],))
for i in range(indiv_predictions.shape[0]):
final_prediction[i] = np.sum(indiv_predictions[i][:])/indiv_predictions.shape[1] #the final prediction of the Random Forest Regressor will be the average of all predictions made by the group of trees
return final_prediction
# ## Testing and comparing the models
# Scikitlearn models for comparison
import sklearn.ensemble as ske
from sklearn.datasets import make_classification
from basicMLpy.classification import acc_and_loss #calculates the accuracy of classfication tasks. basicMLpy is available at: https://github.com/HenrySilvaCS/basicMLpy
from basicMLpy.regression import mse_and_huber
# #### Random Forests Classifier on the Wisconsin Breast Cancer Dataset
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X,y = load_breast_cancer(return_X_y=True)
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X) # scales the data for easier computation
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size = 0.3, random_state=5)
our_rfclassifier = RandomForestClassifier(n_estimators = 100, n_classes = 2)
sk_rfclassifier = ske.RandomForestClassifier(n_estimators = 100)
our_rfclassifier.fit(X_train,Y_train)
sk_rfclassifier.fit(X_train,Y_train)
our_prediction = our_rfclassifier.predict(X_test)
sk_prediction = sk_rfclassifier.predict(X_test)
print(f"The accuracy of our model is: {np.round(acc_and_loss(our_prediction,Y_test)[0],3)}%")
print(f"The accuracy of the sklearn model is: {np.round(acc_and_loss(sk_prediction,Y_test)[0],3)}%")
# #### Random Forests Regressor on the Boston Housing Dataset
from sklearn.datasets import load_boston
X,y = load_boston(return_X_y=True)
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X) # scales the data for easier computation
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size = 0.3, random_state=5)
our_rfregressor = RandomForestRegressor(n_estimators = 100, criterion = 'mse', max_features = 1/3)
sk_rfregressor = ske.RandomForestRegressor(n_estimators = 100, criterion = 'mse', max_features = 1/3)
our_rfregressor.fit(X_train,Y_train)
sk_rfregressor.fit(X_train,Y_train)
our_prediction = our_rfregressor.predict(X_test)
sk_prediction = sk_rfregressor.predict(X_test)
print(f"The Mean Squared Error of our model is: {np.round(mse_and_huber(our_prediction,Y_test)[0],3)}")
print(f"The Mean Squared Error of the sklearn model is: {np.round(mse_and_huber(sk_prediction,Y_test)[0],3)}")
# #### Gradient Boosting Regressor on the Boston Housing Dataset
from sklearn.datasets import load_boston
X,y = load_boston(return_X_y=True)
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X) # scales the data for easier computation
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size = 0.3, random_state=5)
our_gbregressor = GBRegressor(n_estimators = 100, loss_func = 'mse', max_depth = 3)
sk_gbregressor = ske.GradientBoostingRegressor(n_estimators = 100, criterion = 'mse', loss='ls', max_depth = 3)
our_gbregressor.fit(X_train,Y_train)
sk_gbregressor.fit(X_train,Y_train)
our_prediction = our_gbregressor.predict(X_test)
sk_prediction = sk_gbregressor.predict(X_test)
print(f"The Mean Squared Error of our model is: {np.round(mse_and_huber(our_prediction,Y_test)[0],3)}")
print(f"The Mean Squared Error of the sklearn model is: {np.round(mse_and_huber(sk_prediction,Y_test)[0],3)}")
# #### AdaBoost Classifier on a toy dataset
# +
from sklearn.datasets import make_gaussian_quantiles
def make_toy_dataset(n: int = 100, random_seed: int = None):
""" Generate a toy dataset for evaluating AdaBoost classifiers """
n_per_class = int(n/2)
if random_seed:
np.random.seed(random_seed)
X, y = make_gaussian_quantiles(n_samples=n, n_features=4, n_classes=2)
return X, y*2-1
X, y = make_toy_dataset(n=100, random_seed=10)
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size = 0.2, random_state=5)
# -
our_abclassifier = AdaBoostClassifier(n_estimators = 100)
sk_abclassifier = ske.AdaBoostClassifier(n_estimators = 100)
our_abclassifier.fit(X_train,Y_train)
sk_abclassifier.fit(X_train,Y_train)
our_prediction = our_abclassifier.predict(X_test)
sk_prediction = sk_abclassifier.predict(X_test)
print(f"The accuracy of our model is: {np.round(acc_and_loss(our_prediction,Y_test)[0],3)}%")
print(f"The accuracy of the sklearn model is: {np.round(acc_and_loss(sk_prediction,Y_test)[0],3)}%")
# ## Conclusions
# As we can see, our models have a similar performance to the scikitlearn pre-built ones, which is a good indication that our implementations are efficient and competitive when compared to the ensemble algorithms available at other libraries.
| notebooks/Ensemble methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# Datashader provides a flexible series of processing stages that map from raw data into viewable images. As shown in the [Introduction](1-Introduction.ipynb), using datashader can be as simple as calling ``datashade()``, but understanding each of these stages will help you get the most out of the library.
#
# The stages in a datashader pipeline are similar to those in a [3D graphics shading pipeline](https://en.wikipedia.org/wiki/Graphics_pipeline):
#
# 
#
# Here the computational steps are listed across the top of the diagram, while the data structures or objects are listed along the bottom. Breaking up the computations in this way is what makes Datashader able to handle arbitrarily large datasets, because only one stage (Aggregation) requires access to the entire dataset. The remaining stages use a fixed-sized data structure regardless of the input dataset, allowing you to use any visualization or embedding methods you prefer without running into performance limitations.
#
# In this notebook, we'll first put together a simple, artificial example to get some data, and then show how to configure and customize each of the data-processing stages involved:
#
# 1. [Projection](#Projection)
# 2. [Aggregation](#Aggregation)
# 3. [Transformation](#Transformation)
# 4. [Colormapping](#Colormapping)
# 5. [Embedding](#Embedding)
#
# ## Data
#
# For an example, we'll construct a dataset made of five overlapping 2D Gaussian distributions with different σs (spatial scales). By default we'll have 10,000 datapoints from each category, but you should see sub-second response times even for 1 million datapoints per category if you increase `num`.
# +
import pandas as pd
import numpy as np
from collections import OrderedDict as odict
num=10000
np.random.seed(1)
dists = {cat: pd.DataFrame(odict([('x',np.random.normal(x,s,num)),
('y',np.random.normal(y,s,num)),
('val',val),
('cat',cat)]))
for x, y, s, val, cat in
[( 2, 2, 0.03, 10, "d1"),
( 2, -2, 0.10, 20, "d2"),
( -2, -2, 0.50, 30, "d3"),
( -2, 2, 1.00, 40, "d4"),
( 0, 0, 3.00, 50, "d5")] }
df = pd.concat(dists,ignore_index=True)
df["cat"]=df["cat"].astype("category")
# -
# Datashader can work with columnar data in [Pandas](http://pandas.pydata.org) or [Dask](http://dask.pydata.org) dataframes, or with gridded data using [xarray](http://xarray.pydata.org). Here, we're using a Pandas dataframe, with 50,000 rows by default:
df.tail()
# To illustrate this dataset, we'll make a quick-and-dirty Datashader plot that dumps these x,y coordinates into an image:
# +
import datashader as ds
import datashader.transfer_functions as tf
# %time tf.shade(ds.Canvas().points(df,'x','y'))
# -
# Without any special tweaking, datashader is able to reveal the overall shape of this distribution faithfully: four summed 2D normal distributions of different variances, arranged at the corners of a square, overlapping another very high-variance 2D normal distribution centered in the square. This immediately obvious structure makes a great starting point for exploring the data, and you can then customize each of the various stages involved as described below.
#
# Of course, this is just a static plot, and you can't see what the axes are, so we can instead embed this data into an interactive plot if we prefer:
# Here, if you are running a live Python process, you can enable the "wheel zoom" tool on the right, zoom in anywhere in the distribution, and datashader will render a new image that shows the full distribution at that new location. If you are viewing this on a static web site, zooming will simply make the existing set of pixels larger, because this dynamic updating requires Python.
#
# Now that you can see the overall result, we'll unpack each of the steps in the Datashader pipeline and show how this image is constructed from the data.
#
#
# ## Projection
#
# Datashader is designed to render datasets projected on to a 2D rectangular grid, eventually generating an image where each pixel corresponds to one cell in that grid. The ***Projection*** stage is primarily conceptual, as it consists of you deciding what you want to plot and how you want to plot it:
#
# - **Variables**: Select which variable you want to have on the *x* axis, and which one for the *y* axis. If those variables are not already columns in your dataframe (e.g. if you want to do a coordinate transformation), you'll need to create suitable columns mapping directly to *x* and *y* for use in the next step. For this example, the "x" and "y" columns are conveniently named `x` and `y` already, but any column name can be used for these axes.
# - **Ranges**: Decide what ranges of those values you want to map onto the scene. If you omit the ranges, datashader will calculate the ranges from the data values, but you will often wish to supply explicit ranges for three reasons:
# 1. Calculating the ranges requires a complete pass over the data, which takes nearly as much time as actually aggregating the data, so your plots will be about twice as fast if you specify the ranges.
# 2. Real-world datasets often have some outliers with invalid values, which can make it difficult to see the real data, so after your first plot you will often want to specify only the range that appears to have valid data.
# 3. Over the valid range of data, you will often be mainly interested in a specific region, allowing you to zoom in to that area (though with an interactive plot you can always do that as needed).
# - **Axis types**: Decide whether you want `'linear'` or `'log'` axes.
# - **Resolution**: Decide what size of aggregate array you are going to want.
#
# Here's an example of specifying a ``Canvas`` (a.k.a. "Scene") object for a 200x200-pixel image covering the range ±8.0 on both axes:
canvas = ds.Canvas(plot_width=300, plot_height=300,
x_range=(-8,8), y_range=(-8,8),
x_axis_type='linear', y_axis_type='linear')
# At this stage, no computation has actually been done -- the `canvas` object is a purely declarative, recording your preferences to be applied in the next stage.
#
# <!-- Need to move the Points/Lines/Rasters discussion into the section above once the API is rationalized, and rename Canvas to Scene. -->
#
#
# ## Aggregation
#
# <!-- This section really belongs under Scene, above-->
# Once a `Canvas` object has been specified, it can then be used to guide aggregating the data into a fixed-sized grid. You'll first need to know what your data points represent, i.e., what form each datapoint should take as it maps onto the rectangular grid. The library currently supports:
# - **Canvas.points**: mapping each datapoint into the single closest grid cell to that datapoint's location
# - **Canvas.lines**: mapping each datapoint into every grid cell falling between this point's location and the next.
# - **Canvas.raster**: mapping each datapoint into an axis-aligned rectangle forming a regular grid with adjacent points.
#
# Datashader can be extended to add additional types here and in each section below; see [Extending Datashader](../user_guide/7-Extending.ipynb) for more details. Other plots like time series and network graphs are constructed out of these basic primitives.
# <!-- (to here) -->
#
# ### Reductions
#
# One you have determined your mapping, you'll next need to choose a reduction operator to use when aggregating multiple datapoints into a given pixel. All of the currently supported reduction operators are incremental, which means that we can efficiently process datasets in a single pass. Given an aggregate bin to update (typically corresponding to one eventual pixel) and a new datapoint, the reduction operator updates the state of the bin in some way. (Actually, datapoints are normally processed in batches for efficiency, but it's simplest to think about the operator as being applied per data point, and the mathematical result should be the same.) A large number of useful [reduction operators]((http://datashader.readthedocs.org/en/latest/api.html#reductions) are supplied in `ds.reductions`, including:
#
# **`count(column=None)`**:
# increment an integer count each time a datapoint maps to this bin.
#
# **`any(column=None)`**:
# the bin is set to 1 if any datapoint maps to it, and 0 otherwise.
#
# **`sum(column)`**:
# add the value of the given column for this datapoint to a running total for this bin.
#
# **`count_cat(column)`**:
# given a bin with categorical data (i.e., [Pandas' `categorical` datatype](https://pandas-docs.github.io/pandas-docs-travis/categorical.html)), count each category separately, adding the given datapoint to an appropriate category within this bin. These categories can later be collapsed into a single count if needed; see example below.
#
# **`summary(name1=op1,name2=op2,...)`**:
# allows multiple reduction operators to be computed in a single pass over the data; just provide a name for each resulting aggregate and the corresponding reduction operator to use when creating that aggregate.
#
# The API documentation contains the complete list of [reduction operators]((http://datashader.readthedocs.org/en/latest/api.html#reductions) provided, including `mean`, `min`, `max`, `var` (variance), `std` (standard deviation). The reductions are also imported into the ``datashader`` namespace for convenience, so that they can be accessed like ``ds.mean()`` here.
#
# For the operators above, those accepting a `column` argument will only do the operation if the value of that column for this datapoint is not `NaN`. E.g. `count` with a column specified will count the datapoints having non-`NaN` values for that column.
#
# Once you have selected your reduction operator, you can compute the aggregation for each pixel-sized aggregate bin:
canvas.points(df, 'x', 'y', agg=ds.count())
# The result of will be an [xarray](http://xarray.pydata.org) `DataArray` data structure containing the bin values (typically one value per bin, but more for multiple category or multiple-aggregate operators) along with axis range and type information.
#
# We can visualize this array in many different ways by customizing the pipeline stages described in the following sections, but for now we'll use HoloViews to render images using the default parameters to show the effects of a few different aggregate operators:
tf.Images(tf.shade( canvas.points(df,'x','y', ds.count()), name="count()"),
tf.shade( canvas.points(df,'x','y', ds.any()), name="any()"),
tf.shade( canvas.points(df,'x','y', ds.mean('y')), name="mean('y')"),
tf.shade(50-canvas.points(df,'x','y', ds.mean('val')), name="50- mean('val')"))
# Here ``count()`` renders each bin's count in a different color, to show the true distribution, while ``any()`` turns on a pixel if any point lands in that bin, and ``mean('y')`` averages the `y` column for every datapoint that falls in that bin. Of course, since ever datapoint falling into a bin happens to have the same `y` value, the mean reduction with `y` simply scales each pixel by its `y` location.
#
# For the last image above, we specified that the `val` column should be used for the `mean` reduction, which in this case results in each category being assigned a different color, because in our dataset all items in the same category happen to have the same `val`. Here we also manipulated the result of the aggregation before displaying it by subtracting it from 50, as detailed in the next section.
#
#
#
# ## Transformation
#
# Now that the data has been projected and aggregated into a gridded data structure, it can be processed in any way you like, before converting it to an image as will be described in the following section. At this stage, the data is still stored as bin data, not pixels, which makes a very wide variety of operations and transformations simple to express.
#
# For instance, instead of plotting all the data, we can easily plot only those bins in the 99th percentile by count (left), or apply any [NumPy ufunc](http://docs.scipy.org/doc/numpy/reference/ufuncs.html) to the bin values (whether or not it makes any sense!:
# +
agg = canvas.points(df, 'x', 'y')
tf.Images(tf.shade(agg.where(agg>=np.percentile(agg,99)), name="99th Percentile"),
tf.shade(np.power(agg,2), name="Numpy square ufunc"),
tf.shade(np.sin(agg), name="Numpy sin ufunc"))
# -
# The [xarray documentation](http://xarray.pydata.org/en/stable/computation.html) describes all the various transformations you can apply from within xarray, and of course you can always extract the data values and operate on them outside of xarray for any transformation not directly supported by xarray, then construct a suitable xarray object for use in the following stage. Once the data is in the aggregate array, you generally don't have to worry much about optimization, because it's a fixed-sized grid regardless of your data size, and so it is very straightforward to apply arbitrary transformations to the aggregates.
#
# The above examples focus on a single aggregate, but there are many ways that you can use multiple data values per bin as well. For instance, if you collect categorical data, you will have an aggregate value for each category for each bin:
aggc = canvas.points(df, 'x', 'y', ds.count_cat('cat'))
aggc
# Currently only counts are supported for categories, but other reduction operators can be implemented as well (a [to-do item](https://github.com/bokeh/datashader/issues/140)).
#
# You can then select a specific category or subset of them for further processing, where `.sum(dim='cat')` will collapse across such a subset to give a single aggregate array:
# +
agg_d3_d5=aggc.sel(cat=['d3', 'd5']).sum(dim='cat')
tf.Images(tf.shade(aggc.sel(cat='d3'), name="Category d3"),
tf.shade(agg_d3_d5, name="Categories d3 and d5"))
# -
# You can also combine multiple aggregates however you like, as long as they were all constructed using the same Canvas object (which ensures that their aggregate arrays are the same size) and cover the same axis ranges:
tf.Images(tf.shade(agg_d3_d5.where(aggc.sel(cat='d3') == aggc.sel(cat='d5')), name="d3+d5 where d3==d5"),
tf.shade( agg.where(aggc.sel(cat='d3') == aggc.sel(cat='d5')), name="d1+d2+d3+d4+d5 where d3==d5"))
# The above two results are using the same mask (only those bins `where` the counts for 'd3' and 'd5' are equal), but applied to different aggregates (either just the `d3` and `d5` categories, or the entire set of counts).
#
# ## Colormapping
#
# As you can see above, the usual way to visualize an aggregate array is to map from each array bin into a color for a corresponding pixel in an image. The above examples use the `tf.shade()` method, which maps a scalar aggregate bin value into an RGB (color) triple and an alpha (opacity) value. By default, the colors are chosen from the colormap ['lightblue','darkblue'] (i.e., `#ADD8E6` to `#00008B`), with intermediate colors chosen as a linear interpolation independently for the red, green, and blue color channels (e.g. `AD` to `00` for the red channel, in this case). The alpha (opacity) value is set to 0 for empty bins and 1 for non-empty bins, allowing the page background to show through wherever there is no data. You can supply any colormap you like, including Bokeh palettes, Matplotlib colormaps, or a list of colors (using the color names from `ds.colors`, integer triples, or hexadecimal strings):
from bokeh.palettes import RdBu9
tf.Images(tf.shade(agg, cmap=["darkred", "yellow"], name="darkred, yellow"),
tf.shade(agg,cmap=[(230,230,0), "orangered", "#300030"], name="yellow, orange red, dark purple"),
tf.shade(agg,cmap=RdBu9,name="Bokeh RdBu9"))
# #### Colormapping categorical data
#
# If you want to use `tf.shade` with a categorical aggregate, you can use a colormap just as for a non-categorical aggregate if you first select a single category using something like `aggc.sel(cat='d3')` or else collapse all categories into a single aggregate using something like
# `aggc.sum(dim='cat')`. Or you can instead use `tf.shade` with the categorical aggregate directly, which will assign a color to each category and then mix the colors according to the values of each category:
# +
color_key = dict(d1='blue', d2='green', d3='red', d4='orange', d5='purple')
tf.Images(tf.shade(aggc, name="Default color key"),
tf.shade(aggc, color_key=color_key, name="Custom color key"))
# -
# Here the different colors mix not just on the page, but per pixel, with pixels having non-zero counts from multiple categories taking intermediate color values. The actual data values are used to calculate the alpha channel, with this computed color being revealed to a greater or lesser extent depending on the value of the aggregate for that bin. The default color key for categorical data provides distinguishable colors for a couple of dozen categories, but you can provide an explicit color_key if you prefer.
# Choosing colors for different categories is more of an art than a science, because the colors not only need to be distinguishable, their combinations also need to be distinguishable if those categories ever overlap in nearby pixels, or else the results will be ambiguous. In practice, only a few categories can be reliably distinguished in this way, but zooming in (as shown below) can be used to help disambiguate overlapping colors, as long as the basic set of colors is itself distinguishable.
#
#
# #### Transforming data values for colormapping
#
# In each of the above examples, you may have noticed that we never needed to specify any parameters about the data values; the plots just appear like magic. That magic is implemented in `tf.shade`. What `tf.shade` does for a 2D aggregate (non-categorical) is:
#
# 1. **Mask** out all bins with a `NaN` value (for floating-point arrays) or a zero value (for integer arrays); these bins will not have any effect on subsequent computations. Unfortunately, integer arrays do not support `NaN`; using zero as a pseudo-`NaN` works well for counts but not for all integer data, which is something that may need to be generalized in a future version of the library (a [to-do item](https://github.com/bokeh/datashader/issues/142)).
#
# 2. **Transform** the bin values using a specified scalar function `how`. Calculates the value of that function for the difference between each bin value and the minimum non-masked bin value. E.g. for `how="linear"`, simply returns the difference unchanged. Other `how` functions are discussed below.
#
# 3. **Map** the resulting transformed data array into the provided colormap. First finds the value span (*l*,*h*) for the resulting transformed data array -- what are the lowest and highest non-masked values? -- and then maps the range (*l*,*h*) into the full range of the colormap provided. Masked values are given a fully transparent alpha value, and non-masked ones are given a fully opaque alpha value.
#
# The result is thus auto-ranged to show whatever data values are found in the aggregate bins (though the `span` argument can be used to set the range explicitly where appropriate).
#
# As described in plotting_pitfalls.ipynb, auto-ranging is only part of what is required to reveal the structure of the dataset; it's also crucial to automatically and potentially nonlinearly map from the aggregate values (e.g. bin counts) into the colormap. If we used a linear mapping, we'd see very little of the structure of the data:
tf.shade(agg,how='linear')
# In the linear version, you can see that the bins that have zero count show the background color, since they have been masked out using the alpha channel of the image, and that the rest of the pixels have been mapped to colors near the bottom of the colormap. If you peer closely at it, you may even be able to see that one pixel (from the smallest Gaussian) has been mapped to the highest color in the colormap (here dark blue). But no other structure is visible, because the highest-count bin is so much higher than all of the other bins:
top15=agg.values.flat[np.argpartition(agg.values.flat, -15)[-15:]]
print(sorted(top15))
print(sorted(np.round(top15*255.0/agg.values.max()).astype(int)))
# I.e., if using a colormap with 255 colors, the largest bin (`agg.values.max()`) is mapped to the highest color, but with a linear scale all of the other bins map to only the first 24 colors, leaving all intermediate colors unused. If we want to see any structure for these intermediate ranges, we need to transform these numerical values somehow before displaying them. For instance, if we take the logarithm of these large values, they will be mapped into a more tractable range:
print(np.log1p(sorted(top15)))
# So we can plot the logarithms of the values (``how='log'``, below), which is an arbitrary transform but is appropriate for many types of data. Alternatively, we can make a histogram of the numeric values, then assign a pixel color to each equal-sized histogram bin to ensure even usage of every displayable color (``how='eq_hist'``; see [plotting pitfalls](../user_guide/1-Plotting_pitfalls.ipynb). We can even supply any arbitrary transformation to the colormapper as a callable, such as a twenty-third root:
tf.Images(tf.shade(agg,how='log', name="log"),
tf.shade(agg,how='eq_hist', name="eq_hist"),
tf.shade(agg,how=lambda d, m: np.where(m, np.nan, d)**(1/23.), name="23rd root"))
# Usually, however, such custom operations are done directly on the aggregate during the ***Transformation*** stage; the `how` operations are meant for simple, well-defined transformations solely for the final steps of visualization, which allows the main aggregate array to stay in the original units and scale in which it was measured. Using `how` also helps simplify the subsequent ***Embedding*** stage, letting it provide one of a fixed set of legend types, either linear (for `how=linear`), logarithmic (for `how=log`) or percentile (for `how=eq_hist`). See the [shade docs](http://datashader.readthedocs.org/en/latest/api.html#datashader.transfer_functions.shade) for more details on the `how` functions. The `shade` function applies the `how` method similarly for categorical aggregates, based on the total across all categories, but then uses it for the alpha (opacity) channel of the image, rather than to index into a separate colormap.
#
#
# #### Spreading
#
# Once an image has been created, it can be further transformed with a set of functions from `ds.transfer_functions`.
#
# For instance, because it can be difficult to see individual dots, particularly for zoomed-in plots, you can transform the image to replace each non-transparent pixel with a shape, such as a circle (default) or square. This process is called spreading:
# +
img = tf.shade(aggc, name="Original image")
tf.Images(img,
tf.spread(img, name="spread 1px"),
tf.spread(img, px=2, name="spread 2px"),
tf.spread(img, px=3, shape='square', name="spread square"))
# -
# As you can see, spreading is very effective for isolated datapoints, which is what it's normally used for, but it has overplotting-like effects for closely spaced points like in the green and purple regions above, and so it would not normally be used when the datapoints are dense.
#
# Spreading can be used with a custom mask, as long as it is square and an odd width and height (so that it will be centered over the original pixel):
# +
mask = np.array([[1, 1, 1, 1, 1],
[1, 0, 0, 0, 1],
[1, 0, 0, 0, 1],
[1, 0, 0, 0, 1],
[1, 1, 1, 1, 1]])
tf.spread(img, mask=mask)
# -
# To support interactive zooming, where spreading would be needed only in sparse regions of the dataset, we provide the dynspread function. `dynspread` will dynamically calculate the spreading size to use by counting the fraction of non-masked bins that have non-masked neighbors; see the
# [dynspread docs](http://datashader.readthedocs.org/en/latest/api.html#datashader.transfer_functions.dynspread) for more details.
#
#
# #### Other image transfer_functions
#
# Other useful image operations are also provided, such as setting the background color or combining images:
tf.Images(tf.set_background(img,"black", name="Black bg"),
tf.stack(img,tf.shade(aggc.sel(cat=['d2', 'd3']).sum(dim='cat')), name="Sum d2 and d3 colors"),
tf.stack(img,tf.shade(aggc.sel(cat=['d2', 'd3']).sum(dim='cat')), how='saturate', name="d2+d3 saturated"))
# See [the API docs](http://datashader.readthedocs.org/en/latest/api.html#transfer-functions) for more details. Image composition operators to provide for the `how` argument of `tf.stack` (e.g. `over` (default), `source`, `add`, and `saturate`) are listed in [composite.py](https://raw.githubusercontent.com/bokeh/datashader/master/datashader/composite.py) and illustrated [here](http://cairographics.org/operators).
#
# ## Moving on
#
# The steps outlined above represent a complete pipeline from data to images, which is one way to use Datashader. However, in practice one will usually want to add one last additional step, which is to embed these images into a plotting program to be able to get axes, legends, interactive zooming and panning, etc. The [next notebook](3_Interactivity.ipynb) shows how to do such embedding.
| datashader-work/datashader-examples/getting_started/2_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division, absolute_import, print_function
import sys
import os
import pickle
import numpy as np
import random
import h5py
from sklearn import metrics
from collections import Counter
from glob import glob
import gc
import tensorflow as tf
import matplotlib.pyplot as plt
import pandas as pd
from tables import *
from keras import backend as K
from keras.preprocessing import sequence
from keras.utils import np_utils
from keras.backend import manual_variable_initialization
from keras.models import load_model, Model
from keras.layers import Dense, concatenate, Flatten, Conv1D, BatchNormalization, Input, Dropout
from keras.optimizers import Adam
from keras.callbacks import CSVLogger, ModelCheckpoint, TerminateOnNaN
#import keras
#root
absPath = '/home/angela3/imbalance_pcm_benchmark/'
sys.path.insert(0, absPath)
from src.model_functions import *
from src.Target import Target
from src.postproc_auxiliar_functions import *
os.environ['PYTHONHASHSEED'] = '0'
np.random.seed(8)
random.seed(8)
tf.random.set_seed(8)
# -
absPath
nfolds = 10
batch_size = 128
epochss = 100
type_padding_prot = "pre_padding"
protein_type= "GPCRs" #"kinases"
# +
#Opening HDF5 with data
file_h5 = "".join((absPath, "data/", protein_type,"/resampling_before_clustering/compounds_activity.h5"))
f = h5py.File(file_h5, 'r')
group = '/activity'
table = "prot_comp"
#Loading maximum lengths of proteins and compounds
with open("".join((absPath, 'data/prot_max_len.pickle')), "rb") as input_file:
max_len_prot = pickle.load(input_file)
#Defining protein dictionary
instarget = Target("AAA")
prot_dict = instarget.predefining_dict()
# -
learning_rate = 5e-4
decay_rate = learning_rate/epochss
adamm = Adam(lr=learning_rate, beta_1=0.1, beta_2=0.001, epsilon=1e-08, decay=decay_rate)
# +
# LEFT BLOCK (to analyse amino acid sequences)
input_seq = Input(shape=(max_len_prot, len(prot_dict)), dtype='float32')
conv_seq = Conv1D(filters=64, padding='same', strides=1, kernel_size=3, activation='relu')(input_seq)
dropout_1 = Dropout(0.4)(conv_seq)
flatten_seq = Flatten()(dropout_1)#(dense_seq)
dense_seq_2 = Dense(50)(flatten_seq)
dropout_2 = Dropout(0.4)(dense_seq_2)
#RIGHT BRANCH (to analyse fingerprints)
input_fps = Input(shape=(881,), dtype='float32')
dense_fps = Dense(50)(input_fps)
dropout_3 = Dropout(0.4)(dense_fps)
#bn_3 = BatchNormalization()(dense_fps)#(dense_seq_2)#(conv_seq)
#MERGE BOTH BRANCHES
main_merged = concatenate([dropout_2, dropout_3],axis=1)#([dense_seq_2, dense_fps], axis=1)
main_dense = Dense(2, activation='softmax')(main_merged)
#build and compile model
model = Model(inputs=[input_seq, input_fps], outputs=[main_dense])
model.compile(loss='categorical_crossentropy', optimizer = adamm, metrics=['accuracy'])
model.summary()
# -
for fold in range(nfolds):
print("Fold:", str(fold))
file_list = "".join((absPath, "data/", protein_type, "/resampling_before_clustering/splitting_lists/splitting_",
str(fold), "_list.pickle"))
with open(file_list, "rb") as input_file:
splitting_list = pickle.load(input_file)
splitting_list[0].sort()
splitting_list[1].sort()
#Defining generators
train_generator = batch_generator_DL(batch_size, f, group, table, splitting_list[0],
max_len_prot, type_padding_prot=type_padding_prot)
val_generator = batch_generator_DL(batch_size, f, group, table, splitting_list[1],
max_len_prot, type_padding_prot=type_padding_prot)
#defining callbacks
if not os.path.exists("".join((absPath, "data/", protein_type,
"/resampling_before_clustering/logs/", str(fold), "/"))):
os.makedirs("".join((absPath, "data/", protein_type,
"/resampling_before_clustering/logs/", str(fold), "/")))
log_path = "".join((absPath, "data/", protein_type,
"/resampling_before_clustering/logs/", str(fold), "/training_log.csv"))
csv_logger = CSVLogger(log_path)
if not os.path.exists("".join((absPath, "data/", protein_type,
"/resampling_before_clustering/checkpoint/", str(fold), "/"))):
os.makedirs("".join((absPath, "data/", protein_type,
"/resampling_before_clustering/checkpoint/", str(fold), "/")))
#if there are already files in the folder, it removes them
r = glob("".join((absPath, "data/", protein_type,
"/resampling_before_clustering/checkpoint/", str(fold), "/*")))
for i in r:
os.remove(i)
terminan = TerminateOnNaN()
checkpoint_path = "".join((absPath, "data/", protein_type,
"/resampling_before_clustering/checkpoint/", str(fold),
"/weights-improvement-{epoch:03d}-{val_accuracy:.4f}.hdf5"))
mcheckpoint = ModelCheckpoint(checkpoint_path, monitor='val_accuracy', verbose=0,
save_best_only=True, save_weights_only=False)
callbacks_list = [csv_logger, terminan, mcheckpoint ]
print("Training")
# fitting the model
history = model.fit_generator(generator=train_generator,
validation_data=val_generator,
steps_per_epoch= int(len(splitting_list[0])/batch_size),
validation_steps=int(len(splitting_list[1])/batch_size),
epochs=epochss,
callbacks=callbacks_list,
verbose=1)
#saving history
if not os.path.exists("".join((absPath, "data/", protein_type, "/resampling_before_clustering/results/",
str(fold), "/"))):
os.makedirs("".join((absPath, "data/", protein_type, "/resampling_before_clustering/results/", str(fold), "/")))
with open("".join((absPath, "data/", protein_type, "/resampling_before_clustering/results/", str(fold), "/history.pickle")), 'wb') as handle:
pickle.dump(history, handle)
print("Prediction on test data")
splitting_list[2].sort()
#PROTEINS
batch_sequences = list(f[group][table][splitting_list[2]]["sequence"])
#COMPOUNDS
batch_compounds = list(f[group][table][splitting_list[2]]["fingerprint"])
#LABELS
batch_y = list(f[group][table][splitting_list[2]]["label"])
#processing sequences and compounds
seqs_onehot = np.asarray(processing_sequences(batch_sequences, max_len_prot, type_padding_prot))
comps_batch = np.asarray(processing_fingerprints(batch_compounds))
batch_labels = np.asarray(bin_to_onehot(batch_y))
history_path = "".join((absPath, "data/", protein_type, "/resampling_before_clustering/results/",
str(fold), "/history.pickle"))
path_to_confusion = "".join((absPath, "data/", protein_type, "/resampling_before_clustering/results/",
str(fold), "/"))
path_to_auc = "".join((absPath, "data/", protein_type, "/resampling_before_clustering/results/",
str(fold), "/"))
history = plot_history(history_path, "".join((absPath, "data/", protein_type,
"/resampling_before_clustering/results/",
str(fold), "/")))
path_to_cp = ''.join((absPath, "data/", protein_type, "/resampling_before_clustering/checkpoint/",
str(fold), "/"))
model, best_path = load_best_model(history, path_to_cp)
cps_loc = ''.join((absPath, "data/", protein_type, "/resampling_before_clustering/checkpoint/",
str(fold), "/*.hdf5"))
#removing the rest of weights
fileList = glob(cps_loc, recursive=True)
fileList.remove(best_path)
if len(fileList) >1:
for filePath in fileList:
try:
os.remove(filePath)
except OSError:
print("Error while deleting file")
y_predprob = model.predict([seqs_onehot, comps_batch])
y_prob = y_predprob[:,1]
y_pred = y_predprob.argmax(-1)
y_test = batch_labels.argmax(-1)
print("Counting predicted: ", Counter(y_pred))
batch_compID_test = list(f[group][table][splitting_list[2]]["da_comp_id"])
batch_protID_test = list(f[group][table][splitting_list[2]]["da_prot_id"])
#confusion matrix
confusion_matrix(y_test, y_pred, path_to_confusion)
#AUC
file_auc = ''.join((absPath, "data/", protein_type, "/resampling_before_clustering/results/",
str(fold), "/AUC.pickle"))
compute_roc(y_test, y_prob, path_to_auc)
# saving predictions on test set
predictions_test = pd.DataFrame({"y_test":y_test, "y_prob":y_prob, "y_pred":y_pred, "comp_ID": batch_compID_test,
"DeepAffinity Protein ID": batch_protID_test})
if not os.path.exists("".join((absPath, "data/", protein_type, "/resampling_before_clustering/predictions/", str(fold), "/"))):
os.makedirs("".join((absPath, "data/", protein_type, "/resampling_before_clustering/predictions/", str(fold), "/")))
predictions_test.to_csv("".join((absPath, "data/", protein_type, "/resampling_before_clustering/predictions/", str(fold), "/test.csv")))
| scripts/resampling_before_clustering/01_training_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
import numpy as np
plt.style.use("fivethirtyeight")
width=0.25
ages_x = [25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35]
index=np.arange(len(ages_x))
dev_y = [38496, 42000, 46752, 49320, 53200,
56000, 62316, 64928, 67317, 68748, 73752]
plt.bar(index-width, dev_y, color="#444444", width=width,label="All Devs")
py_dev_y = [45372, 48876, 53850, 57287, 63016,
65998, 70003, 70000, 71496, 75370, 83640]
plt.bar(index, py_dev_y, color="#008fd5",width=width, label="Python")
js_dev_y = [37810, 43515, 46823, 49293, 53437,
56373, 62375, 66674, 68745, 68746, 74583]
plt.bar(index+width, js_dev_y, color="#e5ae38",width=width,label="JavaScript")
plt.legend()
plt.xticks(ticks=index,labels=ages_x)
plt.title("Median Salary (USD) by Age")
plt.xlabel("Ages")
plt.ylabel("Median Salary (USD)")
plt.tight_layout()
plt.show()
#here we separate the each bar by just 0.25 width first and last
# -
#we read the file as dict mean by keys and we only take the first rows of the file by using next fun
#the csvreader is a iterator it will return the all rows but we need one rows so for that we use next
import csv
with open('F://jupyter//new.csv','r') as csv_file:
csv_reader=csv.DictReader(csv_file)
row=next(csv_reader)
print(row['events'].split(';'))
| mat2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# + [markdown] id="3a84dd0791f54f2eba25d609d4f4f322"
# # Install OpenVINO in IBM Cloud Pak for Data v3.5.0 - Watson Studio Jupyter Env
# + [markdown] id="7450cea6960b437f8309beea421ddf4b"
# ### Use Python 3.6 in IBM CP4D v3.5.0 Watson Studio Jupyter Env
#
# ### OpenVINO currently supports Python 3.6, 3.8 on Red Hat Enterprise Linux 8, 64-bit.
#
# + [markdown] id="26cce525dbbb4c05883ec830764ee424"
# ### Sections in the notebook:
# 1. Install OpenVINO
# 2. Test OpenVINO python imports
# 3. Test OpenVINO tools: Model Optimizer, Benchmark App
# 4. Sanity check OpenVINO by donwloading, converting and benchmarking googlenet-v1-tf model
# + [markdown] id="f654368fde164285a10ef2c79811d8cb"
# ### Resources:
# 1. OpenVINO PyPi: https://pypi.org/project/openvino-dev/
# 2. IBM CP4D: [Customizing environment definitions (Watson Studio)](https://www.ibm.com/support/producthub/icpdata/docs/content/SSQNUZ_latest/wsj/analyze-data/cust-env-parent.html)
# + [markdown] id="0e9acd13bd3d48cd9d6064a4abd30bb8"
# ## 1. Install OpenVINO
# + id="7aff8282-127b-43aa-8738-c0096a11e805"
# Install this specific version of OpenCV to prevent libGl errors
# !pip uninstall -y opencv-python
# !pip install -U opencv-python-headless==4.2.0.32 --user
# + id="ef47c5b9189a4e9a9207d4d88861ce89"
# Install OpenVINO
# !pip install --ignore-installed PyYAML openvino-dev
# + id="8f19bcb3f1954e1c80746c937f1329fc"
# !pip show openvino
# + [markdown] id="05d697550317482189bbcdd29477ecb2"
# ### After installing, Restart Kernel just to be sure...
# + [markdown] id="0bdb111a86d24c9885985ba7d91268d6"
# ## 2. Test OpenVINO python imports
# + id="f5ebc3e9eadc40ca8716d1dacc6489c3"
from openvino.inference_engine import IENetwork, IECore
from openvino.tools.benchmark.main import main
# + [markdown] id="aa8d6023fabf490683bf4d75e0c9e74e"
# ## 3. Test OpenVINO tools: Model Optimizer, Benchmark App, ...
# + [markdown] id="cf04e2e81b1049e383b0bb62db50fb79"
# ### Test Model Optimizer
# + id="615ca08063454ee59d5d3a0afacf6ac7"
# !mo --version
# + [markdown] id="71fd9d4272f840d3b41c5c90d20ad4b1"
# ### Test Benchmark App
# + id="681bb6c8b36e40b68b05311697cb3d1f"
# !benchmark_app
# + [markdown] id="aea7148993b54701844377a75ef2af66"
# ### See other tools:
# + id="4c62e4c8a0d84964ba3aaae8ea9fccc3"
# !ls /opt/conda/envs/Python-3.6-WMLCE/bin/omz*
# + [markdown] id="911bad7f35bb40978017463432b088f8"
# ## 4. Sanity check OpenVINO by donwloading, converting and benchmarking googlenet-v1-tf model
#
# + [markdown] id="3015d3c1b6da4e4ba6e1084701a2108b"
# ### Resources:
#
# 1. OpenVINO Model Zoo (OMZ): https://github.com/openvinotoolkit/open_model_zoo
# 1. OMZ Intel Pre-Trained Models : https://github.com/openvinotoolkit/open_model_zoo/blob/master/models/intel/index.md
# 1. OMZ Public Pre-Trained Models: See Column 3 for OMZ model name: https://github.com/openvinotoolkit/open_model_zoo/blob/master/models/public/index.md
# + [markdown] id="f8c88985ca16414082ec60115a4f754a"
# ### Download `googlenet-v1-tf` model from OMZ
# + id="bb14aaa7793b413383142b1bc383e91c"
# !omz_downloader --name googlenet-v1-tf
# + [markdown] id="0f9c1cb31a874f8db5d37b550d5b45d3"
# # # !ls public/googlenet-v1-tf/
# + [markdown] id="97f8ea53e843446c83f45c797a04c50f"
# ### Convert `googlenet-v1-tf` model to OpenVINO IR
# + id="fe5f01a0bdd147e087f26ed5f9858755"
# !omz_converter --name googlenet-v1-tf
# + [markdown] id="6477f366a16643c2855766615bb88f9b"
# ### Benchmark `googlenet-v1-tf` model with OpenVINO Benchmark App...
# + id="96711ed8d70540618b0ac8af8999b8e3"
# !benchmark_app -m public/googlenet-v1-tf/FP32/googlenet-v1-tf.xml
# + id="123e7b28deba4d8babd79b8766a38091"
| ibm-cp4d/ov-install-ibm-cp4d-jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="dzhATTc3qK69" outputId="d45be3f2-f7b7-44f0-a2b8-071826c5a06a"
import os
import numpy as np
import pandas as pd
import tensorflow as tf
import FinanceDataReader as fdr
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM, Conv1D, Lambda
from tensorflow.keras.losses import Huber
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
# + [markdown] id="ZnJBSxJXT6sT"
# # 수집한 데이터
# + colab={"base_uri": "https://localhost:8080/", "height": 626} id="d-ww6k5PBiVB" outputId="22eb0595-cf98-46e5-ac3d-a737b9b5b8d9"
data = pd.read_excel('./data/samsung.xlsx')
data
# -
data.info()
# +
# 실수형으로 변환
data['CBOE'] = data['CBOE'].astype('float')
data['futures2y'] = data['futures2y'].astype('float')
data['futures10y'] = data['futures10y'].astype('float')
# +
# 실수형으로 변환(값에 ','가 포함된 경우)
data['NASDAQ'] = data['NASDAQ'].str.replace(',', '').astype(float)
data['S&P'] = data['S&P'].str.replace(',', '').astype(float)
data['Exchange rate'] = data['Exchange rate'].str.replace(',', '').astype(float)
# + id="7INm1u1sTtQH"
scaler = MinMaxScaler()
# + id="LlyJbMMvT5Wv"
scale_cols = ['거래량', 'PER', 'PBR', '기관 합계', '기타법인', '개인', '외국인 합계', 'ATR', 'NASDAQ', 'S&P', 'CBOE', 'Exchange rate', 'futures2y', 'futures10y', 'y']
# + colab={"base_uri": "https://localhost:8080/"} id="QufpuJmUUfW2" outputId="b8b563fe-135d-4beb-9a20-cccc14dd880a"
scaled = scaler.fit_transform(data[scale_cols])
scaled
# + id="4sdnuwYIUh8u"
df = pd.DataFrame(scaled, columns=scale_cols)
# + id="Qed_t_AeV9ZH"
x_train, x_test, y_train, y_test = train_test_split(df.drop('y', 1), df['y'], test_size=0.2, random_state=0, shuffle=False)
# + colab={"base_uri": "https://localhost:8080/"} id="oNv0hRddWAP_" outputId="ac42ef5a-fcdb-483c-c680-13ccfe32bff4"
x_train.shape, y_train.shape, x_test.shape, y_test.shape
# + id="lpJ1UjDqWFbn"
def windowed_dataset(series, window_size, batch_size, shuffle):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
if shuffle:
ds = ds.shuffle(1000)
ds = ds.map(lambda w: (w[:-1], w[-1]))
return ds.batch(batch_size).prefetch(1)
# + id="U55izni6WI9_"
WINDOW_SIZE=120
BATCH_SIZE=32
# + id="tPb1RIjeWLLH"
train_data = windowed_dataset(y_train, WINDOW_SIZE, BATCH_SIZE, True)
test_data = windowed_dataset(y_test, WINDOW_SIZE, BATCH_SIZE, False)
# + colab={"base_uri": "https://localhost:8080/"} id="wgQW-BaiWNMP" outputId="b55f0185-9eaf-449e-f410-ec9757f8158a"
for data in train_data.take(1):
print(f'데이터셋(X) 구성(batch_size, window_size, feature갯수): {data[0].shape}')
print(f'데이터셋(Y) 구성(batch_size, window_size, feature갯수): {data[1].shape}')
# + id="N0KzBMj5WPin"
model = Sequential([
# 1차원 feature map 생성
Conv1D(filters=32, kernel_size=5,
padding="causal",
activation="relu",
input_shape=[WINDOW_SIZE, 1]),
# LSTM
LSTM(16, activation='tanh'),
Dense(16, activation="relu"),
Dense(1),
])
# + id="Za90FsTRWSP_"
loss = Huber()
optimizer = Adam(0.0005)
model.compile(loss=Huber(), optimizer=optimizer, metrics=['mse'])
# + id="T5Sru53qWU5v"
earlystopping = EarlyStopping(monitor='val_loss', patience=100, mode='min')
filename = os.path.join('tmp', 'ckeckpointer.ckpt')
checkpoint = ModelCheckpoint(filename,
save_weights_only=True,
save_best_only=True,
monitor='val_loss',
verbose=1)
# + colab={"base_uri": "https://localhost:8080/"} id="2srj41YlWXBf" outputId="f32823f9-7864-4b68-a488-02487a3b851e"
history = model.fit(train_data,
validation_data=(test_data),
epochs=1000,
callbacks=[checkpoint, earlystopping])
# + colab={"base_uri": "https://localhost:8080/"} id="rF7O8hNLWZx3" outputId="01cb10cf-ac64-49dc-cbf1-5af695c7d2c7"
model.load_weights(filename)
# + id="Bo6efgfSXHtI"
pred = model.predict(test_data)
# + colab={"base_uri": "https://localhost:8080/"} id="dzVhSz7ZXJpI" outputId="572236b9-15e6-4baf-8470-66b0a20e4ef1"
pred.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="CsZEi_1GXL9t" outputId="f8e91b31-3b8c-472c-fc24-93c0d489a1f4"
plt.figure(figsize=(12, 9))
plt.plot(np.asarray(y_test)[120:], label='actual')
plt.plot(pred, label='prediction')
plt.legend()
plt.show()
# +
def RMSE(y_test, pred):
return np.sqrt(mean_squared_error(y_test[120:], pred))
print('RMSE : ', RMSE(y_test, pred))
# +
r2_y_predict = r2_score(y_test[120:], pred)
print('R2 : ', r2_y_predict)
# -
val_lose 0.00013
RMSE : 0.01586393814653742
R2 : 0.9922155763036076
| modeling/notebook/lstm_samsung.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Support Vector Machines (SVMs) in Python #
# ##### <NAME> #####
# ### Overview ###
#
# 1. [What is a Support Vector Machine](#section1)
# 2. [Pros and Cons of SVMs](#section2)
# 3. [When to use SVMs](#section3)
# 4. [Key Parameters](#section4)
# 5. [SVM Classifier Walkthrough](#section5)
# 6. [Conclusion](#section6)
# 7. [Additional Reading](#section7)
# 8. [Sources](#section8)
# <a id='section1'></a>
# ### What is a Support Vector Machine? ###
#
# The support vector machine (svm) is a supervised learning algorithm that is primarily used for classification (although it can be used for regression). Svms are currently one of the most popular machine learning algorithms because they are capable of performing non linear classifications on both standard and high dimensional data. The objective of support vector machines is to find the optimal hyperplane in an N dimensional space (where N is the number of features). The optimal hyperplane has the largest margin between different classes of data.
#
# <img src='svm_example.png'>
# <a id='section2'></a>
# ### Pros and Cons of SVMs ###
#
# #### Pros ####
# 1. **Gives optimal solution**: Unlike some algorithms that can get caught in locally optimum solutions, svms always give the global optimum.
# 2. **Overfitting Resistance**: svm's regularization parameters helps prevent overfitting.
# 3. **Accurate**: svms are accurate and tunable thanks to their regularization and kernel parameters.
# 4. **Resistant to Class Imbalances**: Svms continue to perform well on classification tasks where there are many more of a certain classs than the others.
#
# #### Cons ####
# 1. **Slow Training Time**: Svm's can take a while to train on larger datasets.
# 2. **Noise Sensitive**: Svm's can be adversely affected by noise in datasets, especially with overlapping classes.
# <a id='section3'></a>
# ### When to use SVMs? ###
#
# Svms are a great option for most classification problems and should also be considered in regression. Because of their popularity, it is easier to talk about some of the instances when svms are not appropriate.
#
# 1. They can be somewhat cumbersome for large multiclass classification problems since each class needs a new model
# 2. On perceptual tasks (Speech, vision, etc.) svm's are usually worse the deep neural networks
# 3. Gradient boosted trees tend to perform better on structured data than svms
# 4. It can be difficult to interpret the output of svms
# 5. They can take a long time to train on larger datasets
# 6. Choosing a good [kernel function](https://towardsdatascience.com/understanding-the-kernel-trick-e0bc6112ef78) can be difficult
# <a id='section4'></a>
# ### Key SVM Parameters ###
#
# There are three key parameters for svms: kernel, cost (lambda), and gamma
#
# 1. Kernel: kernel is the type of svm we want to create (which could be linear, polynomial, sigmoid, or radial). We choose this based on the underlying shape of our data (can be tough to know without testing). For example, if we want to classify/separate nonlinear data, we wouldn't use a linear kernel.
# 2. Lambda: serves as a degree of importance given to miss classifications of the svm. Higher lambda values necessitate more accurate models at the cost of generalizing on new data.
# 3. Gamma: gamma is a parameter for gaussian kernels i.e. high dimensional data spaces (see below). Gamma controls the shapes of peaks in a high dimensional setting. A small gamma provides low bias with high variance and vice versa. Grid search can be used to find ideal lambda and gamma values.
#
# <img src='hdd_example.jpg'>
#
# <a id='section5'></a>
# ### SVM Classifier Walkthrough: Simple and Kernel ###
#
# Let's walk through how to build a svm classifier on some sample data. More specifically, we will walk through building a kernel svm. We will be using the iris dataset to classify species of iris based on their characteristics.
# +
# building a simple svm
# -
import numpy as mp
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.metrics import classification_report, confusion_matrix
# +
# importing the data
#url for the dataset
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data"
# Assign colum names to the dataset
colnames = ['sepal-length', 'sepal-width', 'petal-length', 'petal-width', 'Class']
# Read dataset to pandas dataframe
irisdata = pd.read_csv(url, names=colnames)
# +
# preprocessing the data
#drop class
X = irisdata.drop('Class', axis=1)
y = irisdata['Class']
# +
#create a train test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20)
# +
# here we implement a polynomial kernel svm using scikit learn
#the degree parameter is the degree of the polynomial
svclassifier = SVC(kernel='poly', degree=8)
svclassifier.fit(X_train, y_train)
# +
# making predictions on new test data
y_pred = svclassifier.predict(X_test)
# +
# evaluating our algorithm
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# +
# now lets build a svm with a gaussian kernel
svclassifier = SVC(kernel='rbf')
svclassifier.fit(X_train, y_train)
# +
# getting predictions with our svm model
y_pred = svclassifier.predict(X_test)
# +
# evaluating the accuracy of the model
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# +
#lets fit one last svm model with a sigmoid kernel
svclassifier = SVC(kernel='sigmoid')
svclassifier.fit(X_train, y_train)
# +
# generating new predictions
y_pred = svclassifier.predict(X_test)
# +
# model accuracy evaluation
# here we can see the model is not very accurate when we make the assumption the data can be fit with a sigmoid svm
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
# <a id='section6'></a>
# ### Conclusion ###
#
# In this tutorial we stepped through how support vector machines can be used effectively in both classification and regression settings. Next, we examined the key parameters (kernel, lambda, and gamma) of svms and then fit three different svms to the iris dataset to classify different flower species.
# <a id='section7'></a>
# ### Additional Reading ###
# 1. [The math behind svms](https://www.svm-tutorial.com/2014/11/svm-understanding-math-part-1/)
# 2. [svms in depth](https://med.nyu.edu/chibi/sites/default/files/chibi/Final.pdf)
# <a id='section8'></a>
# ### Sources ###
#
# 1. https://www.quora.com/What-are-C-and-gamma-with-regards-to-a-support-vector-machine
# 2. https://stackabuse.com/implementing-svm-and-kernel-svm-with-pythons-scikit-learn/
# 3. https://www.google.com/search?rlz=1C1CHBD_enUS811US811&biw=1396&bih=641&tbm=isch&sa=1&ei=2tlIXIroIJKJjwTE4IGwCg&q=3d+data&oq=3d+data&gs_l=img.3..0i67l2j0l8.20519.21706..21839...1.0..0.74.332.5......1....1..gws-wiz-img.......0i7i30j0i8i7i30.6AmvZzYaikE#imgdii=E5kmTvFhCiHPlM:&imgrc=DhVTosm32bZMtM:
# 4. https://www.google.com/search?rlz=1C1CHBD_enUS811US811&biw=1396&bih=641&tbm=isch&sa=1&ei=sLlIXL-ENei-jwSwib_IBg&q=support+vector+machine+optimal+hyperplane&oq=support+vector+machine+optimal+hyperplane&gs_l=img.3...13058.16730..16892...0.0..0.180.1768.15j4......1....1..gws-wiz-img.......0j0i8i30j0i24.-zmN7H2K3RU#imgrc=QhS3ivfEb21sNM:
#
| support_vector_machines/support_vector_machines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Journey 1: 1st year PhD student
# ====
#
# ## 1 Introduction
#
# This notebook is a one of the possible journeys into HARK - the Python package designed to solve economic models with the heterogeneous agents. As it is a "journey", it is not one big tutorial, but a set of links to notebooks and other resources which will help you understand the different HARK objects and functionalities.
#
# This journey does not require a special skill in programing. However, we recommend you take a few introductury tutorials in Python and object-oriented programing (OOP) to make you familiar with the basic concepts. Moreover, we assume some knowledge in the economic theory.
#
# As you have found this journey, you probably have a concept of what a heterogeneous agent model is, but here is a short recap. Think about a basic infinitely lived consumer problem as you know from first-year graduate courses (letting alone the companies and general equilibrium). Using the Bellman equation, we can write it as:
#
# \begin{eqnarray*}
# V(M_t) &=& \max_{C_t} U(C_t) + \beta V(M_{t+1}), \\
# & s.t. & \\
# A_t &=& M_t - C_t, \\
# M_{t+1} &=& R (M_{t}-C_{t}) + Y_t, \\
# \end{eqnarray*}
#
#
# Where $\beta <1$ is a discount factor, $C_t$ is a consumption, $A_t$ - assets, $Y_t$ - income and $U(C)$ is a standard CRRA utility function:
#
# $$
# U(C)=\frac{C^{1-\rho}}{1-\rho}
# $$
#
# Now assume that every consumer faces some uncertainty on her income (e.g. it follows AR (1) process), which is idiosyncratic - the realizations of each shock is (potentially) different for each agent. In this setting the bellman equation looks like:
#
# \begin{eqnarray*}
# V(M_t, Y_t) &=& \max_{C_t} U(C_t) + E[\beta V(M_{t+1}, Y_{t+1})], \\
# & s.t. & \\
# A_t &=& M_t - C_t, \\
# M_{t+1} &=& R (M_{t}-C_{t}) + Y_t, \\
# \end{eqnarray*}
#
# Therefore, finding a distribution of agent assets (consumption, savings) involves many much more advanced numerical tools than in the case of a representative agent. Obviously, this is more demanding to master. Moreover, the knowledge about involved numerical methods is less systematic, and often hard to find. It was mentioned in the HARK manual:
#
# *"After months of effort, you may have had the character-improving experience of
# proudly explaining to your adviser that not only had you grafted two ideas
# together, you also found a trick that speeded the solution by an order of
# magnitude, only to be told that your breathtaking insight had been understood
# for many years, as reflected in an appendix to a 2008 paper; or, worse, your
# discovery was something that “everybody knows” but did not exist at all in
# published form!"*
#
#
# HARK was designed to help you avoid similar experiences. We see two main ways how you can use this package:
#
# - To simulate the standard heterogeneous agent models without learning all the numerical methods
# - To solve your own models building-on the already implemented algorithms
#
# This journey will help you mostly with using HARK in the first way. We do not elaborate here the numerical methods, however in the last sections you can find some guidance which were used and how the source code is structured.
#
# Although using the prepared package is easier than writing your own solution (what sooner or later you will need to do if you create an original heterogeneous agent model), you still need some effort to comprehend the main classes and functionalities of HARK. We hope that this journey will make this easier! We believe that it also will be your first step into the world of the heterogeneous agents modeling.
#
# ---
# NOTE
# ***
# We will be very happy to see your feedback. If you have any questions regarding this tutorial or HARK as a whole please see our [Github page](https://github.com/econ-ark/HARK).
#
# ---
# ## 2 Before you start
#
# As we have mentioned before, this journey does not require a special skill in programing. However, some knowledge about Python and object-oriented programing (OOP) is needed. We propose two possible ways to gather the basic concepts, however, plenty of others are available:
#
# - Quick introduction to Python and OOP: the first three chapters from [Quantecon](https://lectures.quantecon.org/py/index_postgrad.html) should familiarize you with everything what you need for the first tutorials.
# - A little longer introduction (if you want to learn something about used numerical methods):
# - Start with the basic Python [tutorial](https://docs.python.org/3/tutorial)
# - Get some knowledge about [Numpy](https://docs.scipy.org/doc/numpy/user/quickstart.html)
# - You can also learn Python by learning Machine learning, as there are many tutorials constructed in that way. For example [scikit-learn tutorials](https://scikit-learn.org/stable/tutorial/index.html).
# ## 3 Few words about HARK structure
#
# HARK was written using OOP (we hope that you skimmed the tutorials and have some understanding of this). This means that different parts of the model, like different type of consumers', firms, general equilibrium conditions (if you have these components in the model) are implemented as different objects.
#
# Such structure enables you to build your own models with different consumer type distributions / company structure (if you want some). Importantly, learning the package with a such structure implies learning the different types of objects (classes). In HARK there are two main classes: $\texttt{Agent-type}$ (think consumers, macroeconomic models) and $\texttt{Market}$ (think general equilibrium, macro models). As Agent-type objects are the attributes of the Market, we first present you this type (additionally, if you are interested only in microeconomic research, you may not want to study the Market class).
#
# However, only two classes cannot accommodate the huge variety of the currently used models. Thus, each of the classes have subclasses and they have their own subclasses... In general more sophisticated class is a subclass. This journey will reflect this structure, by showing you first the most primitive models, then go ahead to the more fancy ones.
#
# ---
# NOTE
# ***
# In OOP objects are organized in **classes** (the general structure of the objects) and more specific **subclasses**. The subclass inherits the methods and attributes from the its parent class. Thus everything which you can do with the object from a general class can be done with the object from its subclass. Therefore, in case of the economic models the basic one are always the parent classes of the more sophisticated ones.
#
# ---
#
# ## 4 Agent-type class
# Agent-type class enables you to build the macroeconomic models, such as presented in the introduction. It is also the essential part of the macroeconomic model in HARK. Therefore, to use HARK, you always need to use agent-type classes!
#
# ### 4.1 Introductory example
# As an example, let's solve the stochastic model from the introduction. Assume the income process of the agent i in the period t: $Y_{i,t}$, is given by:
#
# \begin{eqnarray*}
# Y_{i,t} &=& \varepsilon_t(\theta_{i,t} p_{i,t}) \\
# p_{i,t+1} &=& p_{i,t}\psi_{i,t+1}\\
# \psi_{i,t} & \sim & N(1,\sigma_{\varrho})\\
# \theta_{i,t} & \sim & N(1,\sigma_{\theta})\\
# \end{eqnarray*}
#
# To get a universal solution of this problem we need to find a policy function (in this case consumption function), we can easily use the HARK solve function. Before we need to declare our model (we assume standard parametrization: R= 1.03, $\rho = 2$, $\beta = 0.96$, $P(\varepsilon=0)= 0.005$, $P(\varepsilon=1)= 0.995$, $\sigma_{\psi}= \sigma_{\theta}=0.1)$:
#
# +
import sys #set path of the notebook
import os
sys.path.insert(0, os.path.abspath('../../.'))
from HARK.ConsumptionSaving.ConsIndShockModel import * #import the module for the idiosyncratic shocks
#we previously defined the paramters to not bother you about it now
import Journey_1_param as Params #imported paramters
from HARK.utilities import plotFuncs #useful function
Example = IndShockConsumerType()
# -
# Next we can solve the model and plot the consumption function:
Example.solve()
min_v = Example.solution[0].mNrmMin #minimal value for which the consumption function is defined
max_v = 20
print("Consumption function")
plotFuncs([Example.solution[0].cFunc],min_v,max_v)
# ### 4.2 The Agent-Type structure
# To understand the microeconomic models in HARK, you need to have some concept of the Agent-type class structure. As it was mentioned, in HARK more advanced models are subclasses of the more primitive ones. The diagram, illustrates this structure: the deterministic class $\texttt{PerfForesightConsumerType}$, is then a parent for the class of the consumers with idiosyncratic income shocks $\texttt{IndShockConsumerType}$. Next there is a class with the idiosyncratic and aggregate income shocks $\texttt{𝙼𝚊𝚛𝚔𝚘𝚟ConsumerType}$. However, it is not the end! There are subclass of the $\texttt{AggShockConsumerType}$ which are designed to be integrated with the macroeconomic models (we will discuss them in the section devoted to the Market class), as well as there are many other subclasses (which we will mention in the supplementary section).
#
# 
#
# ### 4.3 Main tutorials
#
# To reflect the agent-type structure, we propose Quickstart to be your first HARK notebook as it is devoted to the deterministic case. Then proceed to the idiosyncratic consumers and then consumers with aggregate and idiosyncratic shocks. The exact order of the suggested tutorials is given in the table.
#
#
# |Number | Tutorial | Description|
# | :---- | :---- | :---- |
# |1 |[Quickstart](../notebooks/Quickstart_tutorial/Quick_start_with_solution.ipynb) |This tutorial familiarize you with the basic HARK objects and functionalities.<br /> You will learn how to create, solve, plot and simulate the deterministic<br /> microeconomic models ($\texttt{PerfForesightConsumerType}$ class).|
# |2 |[Idiosyncratic consumers](../notebooks/IndShockConsumerType.ipynb) |In this tutorial you will learn how to deal<br /> with the microeconomic models with agents with idiosyncratic shocks:<br /> individual productivity shocks ($\texttt{IndShockConsumerType}$ class). It builds on the Quickstart. |
# |3|[Nondurables during great recession](../notebooks/Nondurables-During-Great-Recession.ipynb)| Use you knowledge about HARK to conduct a few economic experiments!<br /> You will examine the effects of the uncertinity increase on the heterogenous<br /> agents with idiosyncratic income risk.|
# |4|[Chinese-Growth](../notebooks/Chinese-Growth.ipynb.ipynb)|Learn how to dealt with models with idiosyncratic <br /> and aggregate risk ($\texttt{𝙼𝚊𝚛𝚔𝚘𝚟ConsumerType}$ class). <br />Next build advanced simulation with many agent types.|
#
# ### 4.4 Supplementary tutorials
#
# The aforementioned four tutorials are the most essential ones. However, in HARK there are a few other classes, with a similar but, not-the same structure as three basic ones. Here is a list of the notebooks which familiarize you with them (if you so wish, as it is not required to understand the next topics).
#
# |Number | Tutorial | Description|
# | :---- | :---- | :---- |
# |1* |[Kinked consumer](../notebooks/KinkedRconsumerType.ipynb) | $\texttt{KinkedConsumerType}$ is a subclass of $\texttt{IndShockConsumerType}$. <br /> In enables to set different borrowing and lending interest rate. |
# |2* |[Buffer-stock consumer](../notebooks/TractableBufferStockQuickDemo.ipynb) | In the Buffer Stock model, the unemployment state (zero income stat) is irreversible.<br /> This framework is implemented by $\texttt{TractableConsumerType}$ class.<br /> For the analytical properties of buffer stock model check this [lecture notes](http://www.econ2.jhu.edu/people/ccarroll/public/LectureNotes/Consumption/TractableBufferStock/).|
# |3*|[Generalized income process](../notebooks/IndShockConsumerType.ipynb)| In $\texttt{IndShockConsumerType}$ class, the idiosyncratic income shocks<br /> were assumed to be or purely permanent or purely transitory. In the similar class <br /> $\texttt{PersistentShockConsumerType}$ the income shocks follows AR(1) process with parameter <1,<br /> thus there are not full permanent nor transitory <br />(it was called generalized income process).|
#
#
# ## 5 Market class
#
# In macroeconomic models, the consumers are only one type of agents. In such models, the economy contains also firms and a government (or other types of agents). In HARK, several standard macro models were implemented using the **Market** class and its subclasses.
#
#
# ### 5.1 Introductory example
#
# Let's extend our model from the previous section. Assume the prefect competition and Cobb-Douglas production function:
#
# \begin{eqnarray*}
# y_t = k_t^{\alpha} n_t^{1-\alpha}
# \end{eqnarray*}
# Thus producers' problem is:
# \begin{eqnarray*}
# \max_{k_t, n_t} &\: k_t^{\alpha} n_t^{1-\alpha} - (R_t +\delta)k_t-w_t n_t
# \end{eqnarray*}
#
# Where $k_t$ is a capital, $n_t$ labour, $\delta$ is a depreciation rate.
#
# In this case, consumers' income is determined by the wage:
#
# \begin{eqnarray*}
# V(M_{i,t}, Y_{i,t}) &=& \max_{C_{i,t}, M_{i,t+1}} U(C_{i,t}) + E[\beta V(M_{i,t+1}, Y_{i,t+1})], \\
# & s.t. & \\
# A_{i,t} &=& M_{i,t} - C_{i,t}, \\
# M_{i,t+1} &=& R_{t+1} (M_{i,t}-C_{i,t}) + w_{t+1} Y_{i,t+1}, \\
# \end{eqnarray*}
#
# Additionally, assume that the distribution of the consumers over capital is given by the measure $\Gamma_t$. To close the economy, there are the market clearing conditions:
# \begin{eqnarray*}
# n_t &= \int Y{_i,t} d \Gamma_t \\
# k_{t+1} &= \int A_{i,t}^i d \Gamma_t \\
# k_{t+1}+ \int C_{i,t} d\Gamma_t &= y_t+(1-\delta)k_t
# \end{eqnarray*}
#
# In HARK, you can solve this basic case by using **CobbDouglasEconomy** class. However, to add the consumers to the economy you need **AggShockConsumerType** class, which is a subclass of **IndShockConsumerType** Let's declare the economy (assuming depreciation rate $delta = 0.025$):
#
# +
from HARK.ConsumptionSaving.ConsAggShockModel import * #module with the economy classes
AggShockExample = AggShockConsumerType(**Params.init_agg_shocks) #declare the consumer, using the previously prepared parameters
# Make a Cobb-Douglas economy for the agents
EconomyExample = CobbDouglasEconomy(agents=[AggShockExample], **Params.init_cobb_douglas)
# -
# Now, you can solve the economy and plot the aggregate savings function:
# +
EconomyExample.makeAggShkHist() # Simulate a history of aggregate shocks
# Have the consumers inherit relevant objects from the economy
AggShockExample.getEconomyData(EconomyExample)
AggShockExample.solve() #solve the model
print("capital-level steady state: ", EconomyExample.kSS) #print the capital-level steady stae
plotFuncs(AggShockExample.AFunc,0.1,2*EconomyExample.kSS) # plot the aggregate savings function
# -
# ### 5.2 Market class structure
#
# As in case of the agent-type the more complicated macroeconomic models are the subclasses of the more primitive ones. The subclasses of Market include $\texttt{CobbDouglasEconomy}$ and $\texttt{SmallOpenEconomy}$. The main difference between them is that for $\texttt{CobbDouglasEconomy}$, the capital and labour prices are endogenous, while in the (small) open economy class there are set exogenously. Nevertheless, both basic classes enable the aggregate fluctuation in the economy, that is:
#
# \begin{eqnarray*}
# Y_{i,t} &=& \varepsilon_t(\epsilon_{i,t}p_{i,t}\Theta_t P_t )\\
# P_{t+1} &=& P_{t}\Psi_{t+1}\\
# \Psi_{t} &\sim & {N}(1,\sigma_{\Psi})\\
# \Theta_t &\sim &{N}(1,\sigma_{\Theta})\\
# \end{eqnarray*}
#
# Therefore, the consumers, which are attributes of such market classes, need to include the aggregate fluctuations of the whole economy in their optimization problem. This is the reason why the $\texttt{AggShockConsumerType}$ consumer type class (and their subclasses) must be used to construct the macro-model.
#
# The subclass of $\texttt{CobbDouglasEconomy}$ is $\texttt{CobbDouglasMarkovEconomy}$. In this setting, in the economy there exist an additional aggregate fluctuation, which distribution is given by the finite Markov matrix.
#
#
# 
#
#
#
# ### 5.3 Tutorial
#
# To learn the functionalities of the market type classes in HARK we suggest to study a notebook devoted to [Krussel-Smith economy](../notebooks/KrusellSmith.ipynb). In this notebook classical [Krussell-Smith model](https://www.journals.uchicago.edu/doi/abs/10.1086/250034?journalCode=jpe) is implemented (with some extensions) using $\texttt{CobbDouglasMarkovEconomy}$ class.
#
# Before, you can also check the main function from [ConsAggShockModel module](https://github.com/econ-ark/HARK/blob/master/HARK/ConsumptionSaving/ConsAggShockModel.py) to see the basic steps to create the market type objects.
#
#
# #### 5.3.1 If you want to learn (a little) how the Market class works
#
# The Market class was designed to be a general framework for many different macro models. It involves a procedure of aggregating the agents' choices: eg. aggregating consumption and savings ($\texttt{reap_vars}$ in the code) and then transforming the aggregated variables ($\texttt{mill_rule}$ n the code).
#
# If you would like to get better knowledge about this structure firstly look at the [Hark manual](../HARKmanual/index.html). Next, to understand how the HARK Market class works in less standard setting look at the [Fashion victim model](../notebooks/Fashion-Victim-Model.ipynb).
#
# ## 6 If you need to study a source code
#
# In the previous sections we showed how to solve different models using HARK. However, we know that you may also need to work with the source code for a few reasons (e.g. to learn used numerical methods, write your own code).
#
# Obviously, working with the code, even well-written, is much more complicated tasks than just working with finished functions, and no tutorial will let you go through this painlessly. However, we hope that this part which elaborate a little the HARK structure and numerical methods, will help you with this task.
#
# ### 6.1 A few more words on HARK structure
#
# When you look at the [HARK](https://github.com/econ-ark/HARK) sources, you find the subdirectory called HARK. Next there is a script called "core. py". Surprisingly, you will not find this code in many of the subclasses which you learned during this journey!
#
# The reason for this is that HARK.core is a core of the package, kind of a framework for all models which can be coded in HARK. It contains the general framework of the Agent-type classes (AgentType class) and for the market. The exact structure of modules in the HARK core you can find in the [manual](../HARKmanual/index.html) in section 0.2 General Purpose Tools. For the general structure of the AgentType and Market classes also look at the manual, for the sections 0.3 and 0.4 (you can skip the examples, as you already made a lot of them in the tutorials).
#
# Where are the subclasses which you learned during the journey? In HARK, the subclasses are in the separate directories. For the AgentType subclasses, you need to look at HARK.ConsumptionSaving directory. For example, $\texttt{PerfForesightConsumerType}$ and $\texttt{IndShockConsumerType}$ can be found in ConsIndShockModel.py. Nevertheless, if you want to understand any of the HARK modules, you firstly need to understand HARK.core.
#
#
# ### 6.2 HARK solution
#
# For the consumer problems, solutions of the one-period consumer's problem are found using the attribute function $\texttt{solveOnePeriod}$. The inputs passed to this function includes also data from the subsequent periods. Before solveOnePeriod is called, the function presolve() is applied, which prepare the solution (eg. transmit the solution of the sub-sequent period as an input).
#
# The structure of the functions which are used as solveOnePeriod reflects the agent-type class structures. Thus when you will study the source code, you firstly will read the solve classes.
#
# 
#
#
# #### 6.2.1 Solution method for agent problem
# However, knowing the structure of the code does not be very beneficial if you do not know the solution method! While for the perfect foresight consumer it is analytic, for the stochastic consumer (thus with the idiosyncratic or the aggregate shocks) the policy functions are solved by the **endogenous grid method**.
#
# The endogenous grid method is now widely used in the macroeconomic simulations. There are a few resources to learn it, we suggest professor Carroll [lecture notes](http://www.econ2.jhu.edu/people/ccarroll/SolvingMicroDSOPs/). If you prefer a very quick version, we suggest appendix to the Kruger and Kindermann [paper](https://www.nber.org/papers/w20601.pdf) (they develop a little bigger model with a different notation, but the idea is the same).
#
# #### 6.2.2 Finding general equilibrium
# In the most basic case the rational expectations general equilibrium is found by updating the agents' expectations and the aggregate choices to the point when actual aggregated variables (like intrest rate or capital) are equal to the expected ones. However, refer to the papers cited in the notebooks, to understand the exact used mathods.
#
#
# ### 6.3 How to study HARK codes
#
# We hope that this section gave you some idea how the HARK codes work. However, HARK contains a pretty high number of separate modules and directories. Here we give you some guidance how to start:
#
# - Before you start make sure that you understand the endogenous grid method, and general framework structure for AgentType and Market from manual.
# - Start with the HARK.core, make sure that you see the connection between the structure in the manual and the code, check autodoc from the [HARK documentation](https://hark.readthedocs.io/en/latest/generated/HARK.core.html) webpage.
# - Proceed to the ConsumptionSaving\ConsIndShockModel.py and compare the tutorials with the source code.
# - Proceed to the ConsumptionSaving\ConsAggShockModel.py and compare the tutorial on the Market class with the source code, check [autodoc](https://hark.readthedocs.io/en/latest/generated/HARK.ConsumptionSaving.ConsAggShockModel.html).
# - When you want to learn any of the modules, always firstly check autodoc from the [HARK documentation](https://hark.readthedocs.io/en/latest/generated/HARK.core.html) webpage.
#
| examples/Journeys/Journey_1_PhD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
import numpy as np
import pandas as pd
final=pd.read_csv('new_all.csv')
final
final['e_type'].describe()
final=final[final['e_type'] =='드라이기']
final.shape
final.describe()
import seaborn as sns
import matplotlib.pyplot as plt
# +
fig = plt.figure(figsize =(10, 7))
# Creating plot
plt.boxplot(final['H_ele'])
# show plot
plt.show()
# -
final.loc[(final.H_ele >1000),'H_ele'] = final.H_ele.median()
# +
fig = plt.figure(figsize =(10, 7))
# Creating plot
plt.boxplot(final['H_ele'])
# show plot
plt.show()
# -
final.loc[(final.H_ele >390),'H_ele'] = final.H_ele.median()
# +
fig = plt.figure(figsize =(10, 7))
# Creating plot
plt.boxplot(final['H_ele'])
# show plot
plt.show()
# -
final.describe()
# # 드라이기는 50보다 작으면 꺼진걸로 보고, 그 이상이면 켜져 있는 것으로 간주합니다.
# y_list =[]
# for i in range(0,21961):
# if final['H_ele'][i] > 50:
# a = ['H_ele'][i]
# a = '1'
# a.append.y_list
#
# else:
# b = ['H_ele'][i]
# b = '0'
# b.append.y_list
final= final.drop(['ID','date_time', 'e_type'], axis=1)
final['onoff'] = final['H_ele']
final['onoff'].round(2)
final['onoff_2'] = pd.cut(final['onoff'], bins=[0,50, 999], labels=[0,1])
final['onoff_2'].unique()
final= final.drop(['onoff'], axis=1)
feature= final.drop(['onoff_2'],axis=1)
label = final['onoff_2']
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
encoder.fit(feature['size'])
labels = encoder.transform(feature['size'])
# +
from sklearn.preprocessing import OneHotEncoder
import numpy as np
labels = labels.reshape(-1,1)
# 원-핫 인코딩을 적용합니다.
oh_encoder = OneHotEncoder()
oh_encoder.fit(labels)
oh_labels = oh_encoder.transform(labels)
# -
labels = np.asarray(labels ).astype(np.float32)
labels= feature
# +
# 모델 돌리기
# +
# 4. Train set, Test set 나누기
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(feature,label, test_size=0.2,random_state=100)
X_val, X_test, y_val, y_test = train_test_split(feature,label, test_size=0.5, random_state=100)
print(X_train.shape)
print(X_val.shape)
print('======'*2)
print(y_train.shape)
print(y_val.shape)
# -
# from tensorflow import keras
# from tensorflow.keras import layers
# from tensorflow.keras.layers import Dense, Dropout
# model = keras.Sequential()
#
# model.add(layers.Dense(input_dim =17568*8, units=64, kernel_regularizer = keras.regularizers.L2(0.1)))
# model.add(layers.Activation('relu'))
# # model.add(kernel_regularizer = keras.regularizers.L2(0.1))
#
# model.add(layers.Dense(units=128))
# model.add(layers.Activation('relu'))
# #model.add(layers.Dropout(0.2)) # 앞에있는 layer에 영향이 간다.
#
# model.add(layers.Dense(units=64))
# model.add(layers.Activation('relu'))
#
# model.add(layers.Dense(units=1))
# model.add(layers.Activation('sigmoid'))
# model.summary() # summary로 layer의 형태 파악 해주기
#
# sgd= keras.optimizers.SGD(lr=0.1)
# model.compile(loss='binary_crossentropy', optimizer='adam',
# metrics=['accuracy'])
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV, cross_val_score, StratifiedKFold
RFC = RandomForestClassifier()
RFC.fit(X_train, y_train)
model =RandomForestClassifier()
model.fit(X_train, y_train)
y_pred = RFC.predict(X_test)
print(y_pred)
print(np.array(y_test))
# +
from sklearn.metrics import accuracy_score, precision_score, recall_score
acc = accuracy_score(y_test, y_pred)
pres = precision_score(y_test, y_pred)
recall= recall_score(y_test, y_pred)
print(acc)
print(pres)
print(recall)
# +
from sklearn.model_selection import KFold, cross_val_score
data_kf = KFold(n_splits=5, shuffle=True, random_state=9)
score = cross_val_score(model, feature, label, cv=data_kf, scoring= 'accuracy')
print("5_fold_acc:", score)
print("Mean_Acc:", score.mean())
| Mini_Data_Anal/Mini_Project_Hairdryer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="kN_I0YGEupvp"
from sklearn import preprocessing
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.impute import SimpleImputer
from sklearn.model_selection import RandomizedSearchCV
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.tree import DecisionTreeRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn import linear_model
from sklearn import svm
from sklearn import tree
import xgboost as xgb
from sklearn.ensemble import BaggingRegressor
import numpy as np
import pandas as pd
from sklearn.preprocessing import scale
from sklearn.metrics import roc_auc_score
from sklearn.naive_bayes import GaussianNB
from sklearn.model_selection import cross_val_predict
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from xgboost import XGBClassifier
from sklearn.ensemble import VotingClassifier
import seaborn as sns
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt
import calendar
from sklearn.cluster import KMeans
# + colab={} colab_type="code" id="-DO_Z-bRvLDT"
df = pd.read_csv('US_WeatherEvents_2016-2019.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 190} colab_type="code" id="B157WDDYvLrp" outputId="eb51ae0e-4e8f-4670-c2b0-9ce4c91a866b"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 277} colab_type="code" id="ljubl8OMvNsz" outputId="9e9c4333-85dc-47c1-a44f-8c66e991a076"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 319} colab_type="code" id="TNlJkDBfvh-R" outputId="d7ff7e15-b32d-497b-9ae0-64c74f13e970"
df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 252} colab_type="code" id="cqdaU4Yhvkc-" outputId="9778c4e0-d7b0-4440-dc78-198791b721a0"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 117} colab_type="code" id="ZT_HK3wovopP" outputId="47004a6e-ed45-4523-e9ee-420307c4c5a5"
df = df.fillna(df.median())
table_1 = df.groupby(['City','Type']).count()
table_1 = table_1.reset_index()
list_of_dic = []
label = table_1.City[0]
list_val = {}
for index,row in table_1.iterrows():
if row['City'] != label:
label = row['City']
list_of_dic.append(list_val)
list_val = {}
list_val['City'] = row['City']
list_val[row['Type']] = row['EventId']
else:
list_val['City'] = row['City']
list_val[row['Type']] = row['EventId']
data = pd.DataFrame()
for dic in list_of_dic:
data = data.append(dic, ignore_index=True)
data = data.fillna(0)
df = df.merge(data,on = 'City')
# +
df['StartTime(UTC)'] = pd.to_datetime(df['StartTime(UTC)'])
df['EndTime(UTC)'] = pd.to_datetime(df['EndTime(UTC)'])
df['Start_year'] = df['StartTime(UTC)'].dt.year
df['Start_month'] = df['StartTime(UTC)'].dt.month
df['Start_week'] = df['StartTime(UTC)'].dt.week
df['Start_weekday'] = df['StartTime(UTC)'].dt.weekday
df['Start_day'] = df['StartTime(UTC)'].dt.day
df['end_year'] = df['EndTime(UTC)'].dt.year
df['end_month'] = df['EndTime(UTC)'].dt.month
df['end_week'] = df['EndTime(UTC)'].dt.week
df['end_weekday'] = df['EndTime(UTC)'].dt.weekday
df['end_day'] = df['EndTime(UTC)'].dt.day
# + colab={"base_uri": "https://localhost:8080/", "height": 948} colab_type="code" id="B1f10cnWcyQ7" outputId="3e250d79-1bea-48c2-8e8b-e15d144402d4"
weather_type_df = df['Type'].value_counts(ascending=True)
fig=plt.figure(figsize=(18, 16))
plt.title("Frequency of Weathers")
plt.xlabel("Frequency of Weather")
plt.ylabel("Type of Weather")
ax = weather_type_df.plot(kind='barh')
ax.get_xaxis().set_major_formatter(plt.FuncFormatter(lambda x, loc: "{:,}".format(int(x))))
# + colab={} colab_type="code" id="zxFJ8NeAeRl7"
df.head()
# + colab={} colab_type="code" id="vUvMQA_T96dT"
weather_categories = df['Type'].value_counts()
weather_category_names = weather_categories.index
# + colab={} colab_type="code" id="wB_JUDQee7Vh"
def plot_temporal_feature(df, time_feature, weather_category_names, feature, xaxis_formatter=None, xtick_inc=None):
fig = plt.figure(figsize=(50, 100))
for i in range(len(weather_category_names)):
p = plt.subplot(10, 4, i+1)
weather = weather_category_names[i]
cur_weather_data = df[df[feature] == weather]
temporal_data = cur_weather_data[time_feature].value_counts().sort_index()
sns.lineplot(data=temporal_data)
if xtick_inc:
plt.xticks(np.arange(df[time_feature].unique().min(),df[time_feature].unique().max()+1, xtick_inc))
plt.tick_params(axis = 'both', which = 'major', labelsize = 13)
if xaxis_formatter:
p.get_xaxis().set_major_formatter(plt.FuncFormatter(lambda x, loc: xaxis_formatter(x)))
plt.ylabel("Frequency of Weather", fontsize=25)
time_feature_str = str(time_feature).capitalize()
plt.xlabel(time_feature_str, fontsize=25)
plt.title("%s (%s)" % (weather, 'per '+ time_feature_str), fontsize=30)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="aYeLiMyaahQp" outputId="c4186b71-1cf1-4287-8fc9-f366b93e0638"
plot_temporal_feature(df, 'Start_year', weather_category_names, 'Type', xtick_inc=1,)
# + colab={} colab_type="code" id="2DKOLqNiCIZf"
weather_serverity = df['Severity'].value_counts()
weather_serverity_names = weather_serverity.index
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="ZoYsOLo2CRou" outputId="dd77d4e4-080f-4644-9325-67064f44adbe"
plot_temporal_feature(df, 'Start_year', weather_serverity_names, 'Severity', xtick_inc=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="BPXjFLY_FdyD" outputId="41d6c65c-bcfc-43c4-8770-be7638ba90c3"
def convert_month(num):
return calendar.month_abbr[num]
plot_temporal_feature(df, 'Start_month', weather_category_names, 'Type', convert_month, xtick_inc=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 369} colab_type="code" id="CFJBKFzCwyE4" outputId="ccced229-1712-4ac9-d9b8-1172f0b76edf"
df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 190} colab_type="code" id="9nIUVW3he3Ru" outputId="6494387f-ecbf-4911-f4cf-46bde2446421"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 277} colab_type="code" id="dpoX3E-AgQJ7" outputId="b91f0dbe-b863-446f-ff75-bf03f468e062"
df.describe()
# +
### Unsupervised Learning
#texas = df[(df['State'] == 'NV')] # & (str(df['end_year']) == '2019')]
#texas = df['City'].value_counts()
#len(texas['LocationLat'].unique())
#len(texas)
#k = texas[(texas['end_year'] == '2019')]
# +
#Group the occurences of severity based on
sevCityCom = df[['City', 'Severity']]
severity = pd.get_dummies(sevCityCom['Severity'])
severity['City'] = sevCityCom['City']
severity = severity.groupby('City').sum().reset_index()
severity = severity.drop(['UNK', 'Other'], axis=1) #dropping Unknown and Other
severityDBScan = severity
severity.head()
#len(severity)
# +
### KMEANS Clustering
#using elbow method for param tuning
kMeanDf = severity[['City']]
severity.drop('City', axis=1, inplace=True)
normalizedSeverity = preprocessing.normalize(severity)
kmeanRes = []
for i in range(1,21):
kmeans = KMeans(n_clusters=i, max_iter=500, n_init = 50, random_state = 0)
kmeans.fit(normalizedSeverity)
kmeanRes.append(kmeans.inertia_)
K = range(1,21)
plt.figure(figsize=(10,5))
plt.plot(K, kmeanRes, 'bx-')
plt.xlabel('k')
plt.ylabel('Sum Squared Distance')
plt.title('The Elbow Method')
plt.show()
#Based on the elbow method, we will be going with k=4
# +
#might wanna encode airport code rather than dropping it
#severity.drop('AirportCode', axis=1, inplace=True)
finalKMean = KMeans(n_clusters = 4, max_iter=500, n_init = 50, random_state = 0).fit(normalizedSeverity)
#kMeanDf['cluster'] = finalKMean.labels_
severity['cluster'] = finalKMean.labels_
severity['cluster'].unique()
# +
#Dimensionality Reduction using Principal Component Analysis to lower the dimensionality of severity dataframe
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
pca = PCA().fit(normalizedSeverity)
pcaSeverity = pca.transform(normalizedSeverity)
print("Explained Variance Ratio of each component: ")
for i in range(len(pca.explained_variance_ratio_)):
print("\n")
print(i+1,":", pca.explained_variance_ratio_[i]*100)
print('\n')
print("Total sum (%): ", sum(pca.explained_variance_ratio_)*100)
print("Explained variance of the first two components (%): ", sum(pca.explained_variance_ratio_[0:1])*100)
# +
arr1, arr2, arr3, arr4 = ([] for i in range(4))
for i in range(len(pcaSeverity)):
if kmeans.labels_[i] == 0:
arr1.append(pcaSeverity[i])
if kmeans.labels_[i] == 1:
arr2.append(pcaSeverity[i])
if kmeans.labels_[i] == 2:
arr3.append(pcaSeverity[i])
if kmeans.labels_[i] == 3:
arr4.append(pcaSeverity[i])
arr1 = np.array(arr1)
arr2 = np.array(arr2)
arr3 = np.array(arr3)
arr4 = np.array(arr4)
plt.figure(figsize=(9,5))
plt.scatter(arr1[:,0], arr1[:,1], c='red', label='Cluster 0')
plt.scatter(arr2[:,0], arr2[:,1], c='blue', label='Cluster 1')
plt.scatter(arr3[:,0], arr3[:,1], c='green', label='Cluster 2')
plt.scatter(arr4[:,0], arr4[:,1], c='black', label='Cluster 3')
plt.legend()
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.title('Low dimensional visualization (PCA) - City');
#for i in range(len(pca))
# +
#Param tuning for DBSCAN - Choosing the right
from sklearn.neighbors import NearestNeighbors
#from sklearn import preprocessing
import math
#dbObj = severityDBScan[['City']]
#severityDBScan.drop('City', axis=1, inplace=True)
scaler = preprocessing.StandardScaler()
scaledSeverityDB = scaler.fit_transform(severityDBScan)
neigh = NearestNeighbors(n_neighbors=2)
nbrs = neigh.fit(severityDBScan)
distances, indices = nbrs.kneighbors(severityDBScan)
distances = np.sort(distances, axis=0)
distances = distances[:,1]
plt.plot(distances)
plt.title("Finding Optimal Epsilon Value")
#minSample = math.floor(math.log2(len(normalizedSeverityDB)))
# +
#DBSCAN Visualization
#severityDBScan.drop('AirportCode', axis=1, inplace=True)
# +
#DBSCAN
from sklearn.cluster import DBSCAN
dbscan = DBSCAN(eps=240, min_samples=50).fit(severityDBScan)
dbObj['cluster'] = dbscan.labels_
severityDBScan['cluster'] = dbscan.labels_
severityDBScan['cluster'].unique()
# +
list1, list2 = ([] for i in range(2))
for i in range(len(pcaSeverity)):
if dbscan.labels_[i] == 0:
list1.append(pcaSeverity[i])
if dbscan.labels_[i] == -1:
list2.append(pcaSeverity[i])
list1 = np.array(list1)
list2 = np.array(list2)
plt.figure(figsize=(7,7))
plt.scatter(list1[:,0], list1[:,1], c='red', label='Cluster 0')
plt.scatter(list2[:,0], list2[:,1], c='blue', label='Cluster -1')
plt.legend()
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.title('DBSCAN Visualization - City');
#for i in range(len(pca))
#try calculating the entropy/trying multiple value/elbow method on eps and minpts
# -
| main_chan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def vehicle_miles_data():
"""
vehicle_miles_data()
Returns a 15 x 12 matrix with the vehicle miles traveled in the US
(in millions), per month, for the years 2000, ..., 2014 (discussed on page 252).
"""
# Vehicle Miles Traveled (VMT) (Millions)
# US Dept of Transportation, Bureau of Transportation Statistics
# www.transtats.bts.gov
# 15 x 12 matrix: monthly value for 15 years 2000-2014
import numpy as np
vmt = np.array([
[203442, 199261, 232490, 227698, 242501, 242963, 245140, 247832, 227899, 236491, 222819, 218390], # 2000
[209685, 200876, 232587, 232513, 245357, 243498, 250363, 253274, 226312, 241050, 230511, 229584], # 2001
[215215, 208237, 236070, 237226, 251746, 247868, 256392, 258666, 233625, 245556, 230648, 234260], # 2002
[218534, 203677, 236679, 239415, 253244, 252145, 262105, 260687, 237451, 254048, 233698, 238538], # 2003
[222450, 213709, 251403, 250968, 257235, 257383, 265969, 262836, 243515, 254496, 239796, 245029], # 2004
[224072, 219970, 253182, 250860, 262678, 263816, 267025, 265323, 242240, 251419, 243056, 245787], # 2005
[233302, 220730, 256645, 250665, 263393, 263805, 263442, 265229, 245624, 257961, 245367, 248208], # 2006
[233799, 219221, 259740, 252734, 267646, 265475, 267179, 271401, 246050, 261505, 245928, 240444], # 2007
[233469, 221728, 252773, 252699, 261890, 256152, 262152, 261228, 238701, 256402, 237009, 242326], # 2008
[224840, 218031, 247433, 251481, 258793, 258487, 265026, 260838, 242034, 252683, 237342, 239774], # 2009
[220177, 210968, 251858, 254014, 257401, 260159, 265861, 264358, 244712, 256867, 239656, 240932], # 2010
[222724, 213547, 250410, 249309, 254145, 258025, 260317, 260623, 241764, 252058, 238278, 244615], # 2011
[226834, 218714, 253785, 249567, 261355, 260534, 260880, 264983, 239001, 254170, 240734, 238876], # 2012
[228607, 216306, 250496, 252116, 263923, 260023, 264570, 268609, 242582, 259281, 240146, 241365], # 2013
[226444, 215166, 252089, 257947, 268075, 264868, 272335, 271018, 249125, 267185, 242816, 253618]]) # 2014
return vmt
| vehicle_miles_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.datasets import imdb
((Xtrain,Ytrain),(Xtest,Ytest))=imdb.load_data(num_words=10000)
print(Xtrain.shape,Ytrain.shape)
print(Xtrain[0])
print(len(Xtrain[0]))
word_idx=imdb.get_word_index()
print(len(word_idx))
idx_word = dict([value,key] for (key,value) in word_idx.items())
actual_review = ' '.join([idx_word.get(idx-3,'?') for idx in Xtrain[0]])
print(actual_review)
print(len(actual_review.split()))
from keras.preprocessing import sequence
X_Train=sequence.pad_sequences(Xtrain,maxlen=500)
X_Test=sequence.pad_sequences(Xtest,maxlen=500)
print(X_Train[0])
print(X_Test.shape)
# +
# creating model
from keras.layers import Embedding,SimpleRNN,Dense
from keras.models import Sequential
model=Sequential()
model.add(Embedding(10000,64))
model.add(SimpleRNN(32))
model.add(Dense(1,activation='sigmoid'))
model.summary()
# -
model.compile(optimizer='adam',loss='binary_crossentropy',metrics=['accuracy'])
model.fit(X_Train,Ytrain,validation_split=0.2,batch_size=128,epochs=10)
| DeepLearning/RNN/RNN imdb.ipynb |
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Matlab
% language: matlab
% name: matlab
% ---
% +
%test scipt to make realizations and run KPCA and SOM
% the test case is on the gradual deformation example
clear all; close all; clc
model1='gaussian_realz1';
model2='gaussian_realz2';
r1=read_model(model1);
r2=read_model(model2);
%theta vector
theta=0:0.1:1;
%realization matrix
r_ensemble=[];
r_temp=r1*sin(theta*pi)+r2*cos(theta*pi);
r_ensemble=[r_ensemble r_temp];
%run the KPCASOM_rank script
kernel_para=5;
[Y_eig,ndim,rank]=KPCASOM_rank(r_ensemble,'gaussian',kernel_para);
% -
%visualize some models
hFig =figure();
%set(hFig, 'Position', [0 0 4000 4000])
index=1:2:length(theta);
for i=1:length(index)
subplot(3,2,i)
imagesc(reshape(r_ensemble(:,index(i)), 200, 200))
%title(['\theta=' num2str(theta(index(i)))])
end
% + magic_args="second case on an emsemble of 500 realizations of 2D permeability field (68*64)"
temp=load('ModelsExample2.mat');
r_ensemble_2D=temp.perm_ensemble;
[Y_eig_2,ndim_2,rank_2]=KPCASOM_rank(r_ensemble_2D,'gaussian',kernel_para);
% -
| Code/Functions/DGSA Toolbox/KPCASOM/kpcasom_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2020년 7월 16일 목요일
# ### leetCode - Most Common Word (Python)
# ### 문제 : https://leetcode.com/problems/most-common-word/
# ### 블로그 : https://somjang.tistory.com/entry/leetCode-819-Most-Common-Word-Python
# ### 첫번째 시도
# +
from collections import Counter
import re
class Solution:
def mostCommonWord(self, paragraph: str, banned: List[str]) -> str:
paragraph = paragraph.lower()
paragraph = re.sub('[^a-z]',' ',paragraph)
paragraph_list = paragraph.split(' ')
paragraph_list = [para for para in paragraph_list if para not in banned and para != '']
cnt_dict = Counter(paragraph_list)
val_list = list(cnt_dict.items())
print(val_list)
sorted_list = sorted(val_list, key=lambda x: -x[1])
print(sorted_list)
answer = sorted_list[0][0]
return answer
| DAY 101 ~ 200/DAY161_[leetCode] Most Common Word (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualization
# +
# Import requiered packaages
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
img_path = './test/A/image_10077.png'
def visualize(path):
# load model
model = tf.keras.models.load_model('./model')
# intermediate layers in model
successive_output = [layer.output for layer in model.layers[1:]]
visualization_model = tf.keras.models.Model(inputs=model.input, outputs=successive_output)
img = tf.keras.preprocessing.image.load_img(path, color_mode="grayscale", target_size=(28, 28))
x = tf.keras.preprocessing.image.img_to_array(img)
x = x.reshape((1,) + x.shape)
# rescale by 1/255
x /= 255
# obtain all intermediate representation for this image
successive_features_maps = visualization_model.predict(x)
# these are the names of the layers, so can heve them as part of our plot
layer_names = [layer.name for layer in model.layers[1:]]
# Display representation
for layer_name, feature_map in zip(layer_names, successive_features_maps):
if len(feature_map.shape) == 4:
n_features = feature_map.shape[-1]
size = feature_map.shape[1]
display_grid = np.zeros((size, size * n_features))
for i in range(n_features):
# postprocess the features to make it visually palatable
x = feature_map[0, :, :, i]
x -= x.mean()
x /= x.std()
x *= 64
x += 128
x = np.clip(x, 0, 255).astype('uint8')
display_grid[:, i * size : (i + 1) * size] = x
# Display the grid
scale = 20./ n_features
plt.figure(figsize=(scale * n_features, scale))
plt.title(layer_name)
plt.grid(False)
plt.imshow(display_grid, aspect='auto', cmap='viridis')
plt.show()
visualize(img_path)
# +
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing import image
# load model
model = tf.keras.models.load_model('./model')
f, axarr = plt.subplots(3,4)
CONVOLUTION_NUMBER = 1
from tensorflow.keras import models
layer_outputs = [layer.output for layer in model.layers]
# print(len(layer_outputs))
activation_model = tf.keras.models.Model(inputs = model.input, outputs = layer_outputs)
# predicting images
path = './test/A/image_10077.png'
img = image.load_img(path, target_size=(28, 28), color_mode="grayscale")
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
f_image = np.vstack([x])
path = './test/G/image_10035.png'
img = image.load_img(path, target_size=(28, 28), color_mode="grayscale")
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
s_image = np.vstack([x])
path = './test/F/image_10114.png'
img = image.load_img(path, target_size=(28, 28), color_mode="grayscale")
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
t_image = np.vstack([x])
for x in range(0,4):
f1 = activation_model.predict(f_image)[x]
axarr[0,x].imshow(f1[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[0,x].grid(False)
f2 = activation_model.predict(s_image)[x]
axarr[1,x].imshow(f2[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[1,x].grid(False)
f3 = activation_model.predict(t_image)[x]
axarr[2,x].imshow(f3[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[2,x].grid(False)
# -
| project_handwritten-character-recognition-with-convolutional-neural-network-master/Codes/visualization.ipynb |