code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Problem 3
# (10 points) The princess wedding ring can be made from four types of gold $1,2,3$ or $4$ with the following amounts of milligrams of impurity per gram.
#
# | Type | 1 | 2 | 3 | 4 |
# |-------------:|---|---|---|---|
# | mg of lead | 1 | 2 | 2 | 1 |
# | mg of cobalt | 0 | 1 | 1 | 2 |
# | value | 1 | 2 | 3 | 2 |
# **(a)** Set up a linear program that finds the most valuable ring that can be made containing at most $6mg$ of lead and $10mg$ of cobalt.
# Maximize $\sum_{i=1}^{4} V_i*X_i$
#
# such that
#
# $\sum_{i=1}^{4} L_i*X_i \leq 6$
#
# $\sum_{i=1}^{4} C_i*X_i \leq 10$
#
# $ X_i \geq 0 \forall i \in \{1,...,4\}$, where $X_i$ is the amount of type $i$ gold being used to make the ring.
#
#
#
# **(b)** Put the linear program in standard form.
# Maximize $\sum_{i=1}^{4} V_i*X_i$
#
# such that
#
# $\sum_{i=1}^{4} L_i*X_i + s_1 = 6$
#
# $\sum_{i=1}^{4} C_i*X_i + s_2 = 10$
#
# $ X_i \geq 0 \forall i \in \{1,...,4\}$, where $X_i$ is the amount of type $i$ gold being used to make the ring.
#
# $s_1, s_2 \geq 0 $
#
# and where
#
# $V_i \in \{1,2,3,4\}$\
# $L_i \in \{1,2,2,1\}$\
# $C_i \in \{0,1,1,2\}$
# **(c)** Solve the linear program using the simplex method by hand and by Gurobi.
# By hand:
#
# $$\begin{bmatrix}
# 1 & 2 & 1 \\
# 3 & 0 & 1 \\
# 0 & 2 & 4 \end{bmatrix}$$
| HW3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
from dotenv import load_dotenv
import requests
import time
from datetime import date
import json
import csv
import pandas as pd
import warnings
import matplotlib.pyplot as plt
#Load TOKEN
load_dotenv()
cryptocompare_api_key=os.getenv('API-COMPARE-KEY')
print(cryptocompare_api_key)
TODAY = str(date.today())
MY_CRYPTO_CURRENCIES = ["BTC","MANA","ETH","LTC","ALGO","SHIB","MANA"]
def get_prices(token):
url = "https://min-api.cryptocompare.com/data/v2/histoday?fsym="+token+"&tsym=USD&limit=1000&api_key="
response = requests.get(url+cryptocompare_api_key)
my_dict = response.json()
dictionary = my_dict['Data']
columns = ['Date','Value']
rows = []
for entries in dictionary['Data']:
epoch = entries['time']
date = time.strftime('%m/%d/%Y',time.localtime(epoch))
closing_price = entries['close']
rows.append((date,closing_price))
filename = '/Users/rebecaangulorojas/Dropbox/Mi Mac (Rebeca’s MacBook Air)/Documents/Trabajo/DFiBot/Crypto-data-mining/data/'+token+'.csv'
with open(filename,'w') as csvfile:
csvwriter = csv.writer(csvfile)
csvwriter.writerow(columns)
csvwriter.writerows(rows)
get_prices("BTC") # guardamos BTC y MANA
get_prices("MANA")
# # Lectura de Dataframes
btc_df = pd.read_csv('data/BTC.csv')
mana_df = pd.read_csv('data/MANA.csv')
# +
btc_df['Date'] = pd.to_datetime(btc_df['Date'])
btc_df = btc_df.sort_values('Date',ascending=True)
mana_df['Date'] = pd.to_datetime(mana_df['Date'])
mana_df = mana_df.sort_values('Date',ascending=True)
# +
fig, ax = plt.subplots(figsize=(14,6))
plt.title('Bitcoin'+' & '+ 'Mana')
ax.plot(btc_df['Date'],btc_df['Value'],color='red',label='BTC')
ax.legend(loc = 'lower right')
ax2 = ax.twinx()
plt.plot(mana_df['Date'],mana_df['Value'],color='blue',label='MANA')
ax2.legend(loc='upper left')
plt.grid(True)
plt.show()
# +
# btc
DSR_btc =btc_df['Value'].pct_change(1).to_frame()
DSR_btc.rename(columns = {'Value':'BTC-percChange'}, inplace = True)
# mana
DSR_mana =mana_df['Value'].pct_change(1).to_frame()
DSR_mana.rename(columns = {'Value':'MANA-percChange'}, inplace = True)
# +
ETL_table = pd.concat([btc_df, DSR_btc, DSR_mana], axis=1)
DSR = ETL_table[['BTC-percChange','MANA-percChange']]
# -
print("maximo de Mana", ETL_table['MANA-percChange'].max(), " \n ",
ETL_table.iloc[ETL_table['MANA-percChange'].idxmax()])
# +
# Visualize the daily simple returns
plt.figure(figsize=(20,5))
for c in DSR.columns.values:
plt.plot(DSR.index,DSR[c], label = c,lw=2, alpha=0.7)
plt.grid(True)
plt.title("Daily Simple Returns")
plt.ylabel("percentage (in decimal form)")
plt.xlabel("Days")
plt.legend(DSR.columns.values, loc = "upper right")
plt.show()
# -
# Get Volatility
print("Cryptocurrency Volatility")
DSR.std()
# Show the mean daily simple return
DSR.mean()
# Correlation
DSR.corr()
# +
# Visualize the correlation
import seaborn as sns
plt.subplots(figsize=(11,11))
sns.heatmap(DSR.corr(),annot=True,fmt='.2%')
# +
# Get the daily cumulative simple returns
DCSR = (DSR + 1).cumprod()
DCSR
# +
# Visualize the daily cumulative simple returns
plt.figure(figsize=(12.2,4.5))
for c in DCSR.columns.values:
plt.plot(DCSR.index,DCSR[c],lw=2,label=c)
plt.title("Daily Cumulative Simple Return")
plt.xlabel("Days")
plt.ylabel("Growth of $1 investment")
plt.legend(DCSR.columns.values, loc = 'upper left', fontsize=10)
plt.grid(True)
plt.show()
# -
| API-CryptoCompare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Estimating the carbon content of marine bacteria and archaea
# In order to estimate the characteristic carbon content of marine bacteria and archaea, we collected data on the carbon content of marine prokaryotes from 5 studies. Here is a summary of the data collected
import pandas as pd
import numpy as np
import sys
sys.path.insert(0, '../../../statistics_helper')
from CI_helper import *
pd.options.display.float_format = '{:,.1f}'.format
summary = pd.read_excel('marine_prok_carbon_content_data.xlsx','Summary')
summary.head()
# We use the geometric mean of the carbon content from each study as our best estimate for the carbon content of marine bacteria and archaea.
best_estimate = 10**(np.log10(summary['fg C cell-1']).mean())
print('Our best estimate for the carbon content of marine bacteria and arcaea is %0.1f fg C cell^-1' % best_estimate)
# # Uncertainty analysis
#
# In order to assess the uncertainty associated with our estimate of the carbon content of marine bacteria and archaea, we survey all availabe measures of uncertainty.
#
# ## Intra-study uncertainty
# We collected the uncertainties reported in each of the studies. Below is a list of the uncertainties reported in each of the studies. The highest uncertainty reported is lower than 1.5-fold.
print(summary[['Reference','Intra-study uncertainty']])
# ## Interstudy uncertainty
# We estimate the 95% multiplicative confidence interval around the geometric mean of the values from the different studies.
mul_CI = geo_CI_calc(summary['fg C cell-1'])
print('The interstudy uncertainty is ≈%.1f-fold' % mul_CI)
# We thus take the highest uncertainty from our collection of intra-study and interstudy uncertainties which is ≈1.4-fold.
# Our final parameters are:
# +
print('Carbon content of marine bacteria and archaea: %.1f fg C cell^-1' % best_estimate)
print('Uncertainty associated with the carbon content of marine bacteria and archaea: %.1f-fold' % mul_CI)
old_results = pd.read_excel('../marine_prok_biomass_estimate.xlsx')
result = old_results.copy()
result.loc[1] = pd.Series({
'Parameter': 'Carbon content',
'Value': "{0:.1f}".format(best_estimate),
'Units': 'fg C cell^-1',
'Uncertainty': "{0:.1f}".format(mul_CI)
})
result.to_excel('../marine_prok_biomass_estimate.xlsx',index=False)
| bacteria_archaea/marine/carbon_content/marine_prok_carbon_content.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Question 1
#
# >Response: looking at the data plotted, the `myPower` looks fairly definitively linear, whereas the `myPowerSmart` shows the logarithmic "flattening" that we hope for. It "might" be faster at first, but `mySmartPower` has a better asymptotic behaviour.
#
# # Question 2
#
# - (a) seems like a trick question. often the CPU architecture determins, but a 'byte' is 8 bits. in a 32bit CPU, an int can occupry 32 bits. I started on 8 bit CPU's --- 2^8, then 2^16, then 2^32, now we have mostly 2^64.
#
# - (b) $ \large log_2(n) $ -- in python:`log(n, 2)`
#
# - (c) $ \large log_2(n) $ -- in python:`log(n, 2)`
#
# - (d) $ \large log_c(n) $ -- in python:`log(n, c)`
#
#
# # Question 3
#
# - (a) given an underlying array of fixed size, that we keep track of the total # elements in that array with some value `n` -- we can
#
# ```
# while i < n:
# L[i] = L[i+1]
# if i == n:
# break
# i = i + 1
# ```
#
# - (b) move the last element into the slot that we want to delete... Google asked me this question in an interview once.
#
# ```
# L[i] = L[n]
# n = n - 1
# ```
#
#
# # Question 4
#
# - (a) since both $ log_2(n) $ and $ log_c(n) $ represent algorithms that "flatten" as `n` increases, the relative importance is that they are more efficient than `O(n)` or even an exponential growth. Given `c` is constant, with the log identity we have
#
# $ \large log_c * n = log_2 * c * log_2 * n $ -- so, essentially we have $ log_2 * n $ as THE varying part of the running time.
#
#
# - (b)
# Ranking:
#
# - $ n^n $
# - $ n! $
# - $ n^3 $
# - $ n^2 $
# - $ n $
# - $ log_2 n $
#
#
# # Question 6
# Nested looping via brute force is the simple answer.
#
# - $ running time =O(n^3) $
#
# >Note: this looked promising: [Efficient Ramanujan](https://www.ocf.berkeley.edu/~wwu/cgi-bin/yabb/YaBB.cgi?board=riddles_cs;action=display;num=1288192473)
# +
## Question 6 Code:
from math import pow, log
from collections import namedtuple
ramanujan = namedtuple('ramanujan', 'a b c d')
# brute force...
def pr(n):
rv = []
for a in range(n+1): # O(n)
for b in range(n+1): # O(n)
for c in range(n+1): # O(n)
d = n - (a+b+c)
if (pow(a,3)+pow(b,3) == pow(c,3)+ pow(d,3)):
rv.append(ramanujan(a,b,c,d))
return rv
result = pr(20)
result
# -
import math
math.pow(3,2)
# # Addendum
#
# ## Question 1 Work.
# +
def myPower(x,k,p):
y = 1
for i in range(k+1):
y = y*x % p
return y # running time O(k)
# x**k mod p: Simple recursive computation
# x^k = x* x^(k-1)
def myPowerRec(x,k,p):
if k==1:
return x % p
z = myPowerRec(x,k-1,p)
y = x*z % p
return y # running time O(k)
# x**k mod p: Smart recursive computation
# x^k = x^(k/2)*x^(k/2) if k is even
# x^k = x^(k/2)*x^(k/2)*x if k is odd
def myPowerSmart(x,k,p):
if k==1:
return x % p
r = k % 2
k = k // 2
z = myPowerSmart(x,k,p)
if r == 0:
y = z*z % p
else:
y = z*z*x % p
return y # running time O(log k)
# +
import datetime
from time import time as tm
import timeit
def getMinsSeconds(s):
return str(datetime.timedelta(seconds=s))
def timeStatement(rawCode):
rv = []
repeat = 1
kRange = [10, 100, 1000, 10000, 100000, 1000000]#, 10000000]
for k in kRange:
statement = rawCode.format(k)
start_time = tm()
#s = (timeit.timeit(statement, number = repeat)) / repeat
s = (timeit.timeit(statement, number=repeat))
print("--- {} seconds ---".format(tm() - start_time))
rv.append([k,s])
print("for k = {} --- hh:mm::ss.ms: {}".format(k, getMinsSeconds(s)))
return rv
statement_Smart='''
from __main__ import myPowerSmart
from __main__ import myPower
x = 2
p = 10
myPowerSmart(x, {}, p)
'''
statement_Regular='''
from __main__ import myPowerSmart
from __main__ import myPower
x = 2
p = 10
myPowerSmart(x, {}, p)
'''
t_one = timeStatement(statement_Regular)
t_two = timeStatement(statement_Smart)
import matplotlib.pyplot as plt
figsize = (40,10)
fix, ax = plt.subplots(1,1, figsize=figsize)
y1 = [row[1] for row in t_one]
x1 = [row[0] for row in t_one]
y2 = [row[1] for row in t_two]
x2 = [row[0] for row in t_two]
plt.plot(x1, y1, 'tab:red', label='myPower')
plt.plot(x2, y2, 'tab:blue', label='mySmartPower')
#ax.ticklabel_format(useOffset=False, style='plain')
ax.get_xaxis().get_major_formatter().set_useOffset(False)
ax.get_xaxis().get_major_formatter().set_scientific(False)
ax.legend()
plt.show()
# -
# +
def timeStatementEasy(f):
rv = []
x = 2
p = 10
kRange = [10, 100, 1000, 10000, 100000, 1000000]#, 10000000]
for k in kRange:
start_time = tm()
f(x,k,p)
s = tm() - start_time
rv.append([k,s])
print("for k = {} --- hh:mm::ss.ms: {}".format(k, getMinsSeconds(s)))
return rv
t_one_b = timeStatementEasy(myPower)
t_two_b = timeStatementEasy(myPowerSmart)
import matplotlib.pyplot as plt
figsize = (40,10)
fix, ax = plt.subplots(1,1, figsize=figsize)
y1 = [row[1] for row in t_one_b]
x1 = [row[0] for row in t_one_b]
y2 = [row[1] for row in t_two_b]
x2 = [row[0] for row in t_two_b]
plt.plot(x1, y1, 'tab:red', label='myPower')
plt.plot(x2, y2, 'tab:blue', label='mySmartPower')
#ax.ticklabel_format(useOffset=False, style='plain')
ax.get_xaxis().get_major_formatter().set_useOffset(False)
ax.get_xaxis().get_major_formatter().set_scientific(False)
ax.legend()
plt.show()
# +
## Question 2 work...
from math import log
print(log(16,2))
print(log(64,2))
print(log(16,1/2))
print(log(64,1/2))
print(log(16,4))
print(log(64,4))
# -
math.factorial(4)
math.pow(4, 4)
log(2,2)
log(3,2)
| notebooks/work/prior/shawn.cicoria-CS610.j31-assignment 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting started with DoWhy: A simple example
# This is quick introduction to DoWhy causal inference library.
# We will load in a sample dataset and estimate causal effect from a (pre-specified)treatment variable to a (pre-specified) outcome variable.
#
# First, let us add required path for python to find DoWhy code and load required packages.
import os, sys
sys.path.append(os.path.abspath("../../"))
# +
import numpy as np
import pandas as pd
import dowhy
from dowhy.do_why import CausalModel
import dowhy.datasets
# -
# Let us first load a dataset. For simplicity, we simulate a dataset with linear relationships between common causes and treatment, and common causes and outcome.
#
# Beta is the true causal effect.
data = dowhy.datasets.linear_dataset(beta=10,
num_common_causes=5,
num_instruments = 2,
num_samples=10000,
treatment_is_binary=True)
df = data["df"]
print(df.head())
print(data["dot_graph"])
# Note that we are using a pandas dataframe to load the data. At present, DoWhy only supports pandas dataframe as input.
# ## Interface 1 (recommended): Input causal graph
# We now input a causal graph in the DOT graph format.
# With graph
model=CausalModel(
data = df,
treatment=data["treatment_name"],
outcome=data["outcome_name"],
graph=data["dot_graph"],
)
model.view_model()
# <img src="causal_model.png">
# We get the same causal graph. Now identification and estimation is done as before.
identified_estimand = model.identify_effect()
print(identified_estimand)
causal_estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression")
print(causal_estimate)
print("Causal Estimate is " + str(causal_estimate.value))
# ## Interface 2: Specify common causes and instruments
# Without graph
model= CausalModel(
data=df,
treatment=data["treatment_name"],
outcome=data["outcome_name"],
common_causes=data["common_causes_names"])
model.view_model()
# <img src="causal_model.png" />
# The above causal graph shows the assumptions encoded in the cauasl model. We can now use this graph to first identify
# the causal effect (go from a causal estimand to a probability expression), and then estimate the causal effect.
# **DoWhy philosophy: Keep identification and estimation separate**
#
# Identification can be achieved without access to data, only the graph. This results in an expression to computed. This expression can then be computed using the available data in the estimation step.
# Important to understand that these are orthogonal steps.
#
# * Identification
identified_estimand = model.identify_effect()
# * Estimation
estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression",
test_significance=True
)
print(estimate)
print("Causal Estimate is " + str(estimate.value))
# ## Refuting the estimate
#
# Now refuting the obtained estimate.
# ### Adding a random common cause variable
res_random=model.refute_estimate(identified_estimand, estimate, method_name="random_common_cause")
print(res_random)
# ### Replacing treatment with a random (placebo) variable
res_placebo=model.refute_estimate(identified_estimand, estimate,
method_name="placebo_treatment_refuter", placebo_type="permute")
print(res_placebo)
# ### Removing a random subset of the data
res_subset=model.refute_estimate(identified_estimand, estimate,
method_name="data_subset_refuter", subset_fraction=0.9)
print(res_subset)
# As you can see, the linear regression estimator is very sensitive to simple refutations.
| docs/source/do_why_simple_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="sK-RhlsGxwpd" colab_type="text"
# ##### Copyright 2020 Google LLC.
#
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="fuN9k6Gux-yD" colab_type="text"
# This colab contains TensorFlow code for implementing the constrained optimization methods presented in the paper:
# > <NAME>, <NAME>, <NAME>, <NAME>, "Pairwise Fairness for Ranking and Regression", AAAI 2020. [<a href="https://arxiv.org/pdf/1906.05330.pdf">link</a>]
#
# First, let's install and import the relevant libraries.
# + id="JXgLyAJm0UyB" colab_type="code" colab={}
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
# + [markdown] id="iTUyZk_A0XnF" colab_type="text"
# We will need the TensorFlow Constrained Optimization (TFCO) library.
# + id="DvhGP5TW0V_J" colab_type="code" colab={}
# !pip install git+https://github.com/google-research/tensorflow_constrained_optimization
# + id="XGFoSFuX0XJc" colab_type="code" colab={}
import tensorflow_constrained_optimization as tfco
# + [markdown] id="gBUr48pLzsqK" colab_type="text"
# ## Constrained Optimization Problem
# + [markdown] id="_Wgq7N73rPHV" colab_type="text"
# We will be training a linear ranking model $f(x) = w^\top x$ where $x \in \mathbb{R}^d$ is a set of features for a query-document pair (the protected attribute is not a feature). Our goal is to train the model such that it accurately ranks the positive documents in a query above the negative ones.
#
# + [markdown] id="6tZrx9BfOB_Q" colab_type="text"
# Specifically, for the ranking model $f$, we denote:
# - $err(f)$ as the pairwise ranking error for model $f$ over all pairs of positive and negative documents
# $$
# err(f) = \mathbf{E}\big[\mathbb{I}\big(f(x) < f(x')\big) \,\big|\, y = 1,~ y' = 0\big]
# $$
#
#
# - $err_{i,j}(f)$ as the pairwise ranking error over positive-negative document pairs where the pos. document is from group $i$, and the neg. document is from group $j$.
#
# $$
# err_{i, j}(f) = \mathbf{E}\big[\mathbb{I}\big(f(x) < f(x')\big) \,\big|\, y = 1,~ y' = 0,~ grp(x) = i, ~grp(x') = j\big]
# $$
# <br>
#
# We then wish to solve the following constrained problem:
# $$min_f\; err(f)$$
# $$\text{ s.t. } |err_{i,j}(f) - err_{k,\ell}(f)| \leq \epsilon \;\;\; \forall ((i,j), (k,\ell)) \in \mathcal{G},$$
#
# where $\mathcal{G}$ contains the pairs we are interested in constraining.
# + [markdown] id="qM-PzAuykOmN" colab_type="text"
# ## Generate 2D Synthetic Data
#
# We begin by generating a synthetic ranking dataset.
#
# **Setting**:
# - Set of $m$ queries, a set of $n$ documents associated with each query.
# - One document per query has a positive label, while the remaining have negative labels.
# - Each document is associated with one of two protected groups.
#
#
# **Generative process**:
#
# For convenience, the first document for each query is labeled positive, and remaining $n-1$ documents are labeled negative.
#
# For each query-document pair:
# - Draw the protected group for the document from a $Bernoulli(\pi)$ distribution
# - Draw the document features from the following 2D Gaussian distributions:
# - $\mathcal{N}(\mu_{0,0}, \Sigma_{0,0})$ if label is -ve and group is 0
# - $\mathcal{N}(\mu_{1,0}, \Sigma_{1,0})$ if label is +ve and group is 0
# - $\mathcal{N}(\mu_{0,1}, \Sigma_{0,1})$ if label is -ve and group is 1
# - $\mathcal{N}(\mu_{1,1}, \Sigma_{1,1})$ if label is +ve and group is 1
#
# + id="6WvqQsmokWJM" colab_type="code" colab={}
def create_dataset(num_queries, num_docs):
# Create a synthetic 2-dimensional training dataset with 1000 queries,
# with 1 positive document and 10 negative document each
# and with two protected groups for each document.
num_posdocs = 1
num_negdocs = num_docs - 1
dimension = 2
num_groups = 2
tot_pairs = num_queries * num_posdocs * num_negdocs
# Conditional distributions are Gaussian: Conditioned on the label and the
# protected group, the feature distribution is a Gaussian.
# Positive documents, Group 0
mu_10 = [1,0]
sigma_10 = np.array([[1, 0], [0, 1]])
# Positive documents, Group 1
mu_11 = [-1.5, 0.75]
sigma_11 = np.array([[1, 0], [0, 1]]) * 0.5
# Negative documents, Group 0
mu_00 = [-1,-1]
sigma_00 = np.array([[1, 0], [0, 1]])
# Negative documents, Group 1
mu_01 = [-2,-1]
sigma_01 = np.array([[1, 0], [0, 1]])
# Generate positive documents
posdocs_groups = (np.random.rand(num_queries, num_posdocs) <= 0.1) * 1
posdocs_mask = np.dstack([posdocs_groups] * dimension)
posdocs0 = np.random.multivariate_normal(mu_10, sigma_10,
size=(num_queries, num_posdocs))
posdocs1 = np.random.multivariate_normal(mu_11, sigma_11,
size=(num_queries, num_posdocs))
posdocs = (1 - posdocs_mask) * posdocs0 + posdocs_mask * posdocs1
# Generate negative documents
negdocs_groups = (np.random.rand(num_queries, num_negdocs) <= 0.1) * 1
negdocs_mask = np.dstack([negdocs_groups] * dimension)
negdocs0 = np.random.multivariate_normal(mu_00, sigma_00,
size=(num_queries, num_negdocs))
negdocs1 = np.random.multivariate_normal(mu_01, sigma_01,
size=(num_queries, num_negdocs))
negdocs = (1 - negdocs_mask) * negdocs0 + negdocs_mask * negdocs1
# Concatenate positive and negative documents for each query
# (along axis 1, where documents are arranged)
features = np.concatenate((posdocs, negdocs), axis=1)
# Concatenate the associated labels:
# (for each query, first num_posdocs documents are positive, remaining negative)
poslabels = np.tile([1], reps=(num_queries, num_posdocs))
neglabels = np.tile([-1], reps=(num_queries, num_negdocs))
labels = np.concatenate((poslabels, neglabels), axis=1)
# Concatenate the protected groups
groups = np.concatenate((posdocs_groups, negdocs_groups), axis=1)
dataset = {
'features': features,
'labels': labels,
'groups': groups,
'num_queries': num_queries,
'num_posdocs': num_posdocs,
'num_negdocs': num_negdocs,
'dimension': dimension,
'num_groups': num_groups,
'tot_pairs': tot_pairs
}
return dataset
# + [markdown] id="m3YfQJ5NzRSE" colab_type="text"
# ## Plotting Functions
# + [markdown] id="lVxfbRjJy41l" colab_type="text"
# Next, let's write functions to plot the data and the linear decision boundary.
# + id="sRxmpzKV1sPV" colab_type="code" outputId="aa727ba2-f37e-491d-b0b8-83c5e35c498e" colab={"base_uri": "https://localhost:8080/", "height": 283}
def plot_data(dataset, ax=None):
# Plot data set.
features = dataset["features"]
labels = dataset["labels"]
groups = dataset["groups"]
# Create axes if not specified
if not ax:
_, ax = plt.subplots(1, 1, figsize=(4.0, 4.0))
ax.set_xlabel("Feature 0")
ax.set_ylabel("Feature 1")
# Plot positive points in group 0
data = features[(labels==1) & (groups==0), :]
ax.plot(data[:,0], data[:,1], 'bx', label="Pos, Group 0")
# Plot positive points in group 1
data = features[(labels==1) & (groups==1), :]
ax.plot(data[:,0], data[:,1], 'bo', label="Pos, Group 1")
# Plot negative points in group 0
data = features[(labels==-1) & (groups==0), :]
ax.plot(data[:,0], data[:,1], 'rx', label="Neg, Group 0")
# Plot negative points in group 1
data = features[(labels==-1) & (groups==1), :]
ax.plot(data[:,0], data[:,1], 'ro', label="Neg, Group 1")
ax.legend(loc = "upper right")
def plot_model(weights, ax, x_range, y_range, fmt):
# Plot model decision boundary.
ax.plot([x_range[0], x_range[1]],
[-x_range[0] * weights[0] / weights[1],
-x_range[1] * weights[0] / weights[1]],
fmt)
ax.set_ylim(y_range)
# Sample data set.
dataset = create_dataset(num_queries=500, num_docs=10)
plot_data(dataset)
# + [markdown] id="JxBMPRJA2wvW" colab_type="text"
# ## Evaluation Metrics
# + [markdown] id="H4HV7a7wq7rm" colab_type="text"
# We will also need functions to evaluate the pairwise error rates for a linear model.
# + id="K8OQ4ado20p-" colab_type="code" colab={}
def get_mask(dataset, pos_group, neg_group=None):
# Returns a boolean mask selecting positive-negative document pairs where
# the protected group for the positive document is pos_group and
# the protected group for the negative document (if specified) is neg_group.
groups = dataset['groups']
num_negdocs = dataset['num_negdocs']
# Repeat group membership positive docs as many times as negative docs.
mask_pos = groups[:,0] == pos_group
mask_pos_rep = np.repeat(mask_pos.reshape(-1,1), num_negdocs, axis=1)
if neg_group is None:
return mask_pos_rep
else:
mask_neg = groups[:,1:] == neg_group
return mask_pos_rep & mask_neg
def error_rate(model, dataset):
# Returns error rate of model on dataset.
features = dataset["features"]
num_negdocs = dataset['num_negdocs']
scores = np.matmul(features, model)
pos_scores_rep = np.repeat(scores[:, 0].reshape(-1,1), num_negdocs, axis=1)
neg_scores = scores[:, 1:]
diff = pos_scores_rep - neg_scores
return np.mean(diff.reshape((-1)) <= 0)
def group_error_rate(model, dataset, pos_group, neg_group=None):
# Returns error rate of model on data set, considering only document pairs,
# where the protected group for the positive document is pos_group, and the
# protected group for the negative document (if specified) is neg_group.
features = dataset['features']
num_negdocs = dataset['num_negdocs']
scores = np.matmul(features, model)
pos_scores_rep = np.repeat(scores[:, 0].reshape(-1,1), num_negdocs, axis=1)
neg_scores = scores[:, 1:]
mask = get_mask(dataset, pos_group, neg_group)
diff = pos_scores_rep - neg_scores
diff = diff * mask
return np.sum(diff.reshape((-1)) < 0) * 1.0 / np.sum(mask)
# + [markdown] id="kI8xNJDcpQYP" colab_type="text"
# ## Create Linear Model
#
#
#
# + [markdown] id="lY4hvJAOra6s" colab_type="text"
# We then write a function to create the linear ranking model in Tensorflow.
# + id="eTQOebAepXSu" colab_type="code" colab={}
def group_tensors(predictions, dataset, pos_group, neg_group=None):
# Returns predictions and labels for document-pairs where the protected group
# for the positive document is pos_group, and the protected group for the
# negative document (if specified) is neg_group.
mask = get_mask(dataset, pos_group, neg_group)
mask = np.reshape(mask, (-1))
group_labels = lambda: tf.constant(np.ones(np.sum(mask)), dtype=tf.float32)
group_predictions = lambda: tf.boolean_mask(predictions(), mask)
return group_predictions, group_labels
def linear_model(dataset):
# Creates a linear ranking model, and returns a nullary function returning
# predictions on the dataset, and the model weights.
# Create variables containing the model parameters.
weights = tf.Variable(tf.ones(dataset["dimension"], dtype=tf.float32),
name="weights")
# Create a constant tensor containing the features.
features_tensor = tf.constant(dataset["features"], dtype=tf.float32)
# Create a nullary function that returns applies the linear model to the
# features and returns the tensor with the predictions.
def predictions():
predicted_scores = tf.tensordot(features_tensor, weights, axes=(2, 0))
# Compute ranking errors and flatten tensor.
pos_scores = tf.slice(predicted_scores, begin=[0,0],
size=[-1, dataset["num_posdocs"]])
neg_scores = tf.slice(predicted_scores, begin=[0, dataset["num_posdocs"]],
size=[-1,-1])
pos_scores_rep = tf.tile(pos_scores, multiples=(1, dataset["num_negdocs"]))
predictions_tensor = tf.reshape(pos_scores_rep - neg_scores, shape=[-1])
return predictions_tensor
return predictions, weights
# + [markdown] id="WIBvG3Arv7zR" colab_type="text"
# ## Formulate Optimization Problem
# + [markdown] id="SfZd-XPt0A8E" colab_type="text"
# We are ready to formulate the constrained optimization problem using the TFCO library.
# + id="0AfVknixv9So" colab_type="code" colab={}
def formulate_problem(dataset, constraint_groups=[], constraint_slack=None):
# Returns a RateMinimizationProblem object and the linear model weights.
#
# Formulates a constrained problem that optimizes the error rate for a linear
# model on the specified dataset, subject to pairwise fairness constraints
# specified by the constraint_groups and the constraint_slack.
#
# Args:
# dataset: Dataset dictionary returned by create_dataset()
# constraint_groups: List containing tuples of the form
# ((pos_group0, neg_group0), (pos_group1, neg_group1)), specifying the
# group memberships for the document pairs to compare in the constraints.
# constraint_slack: slackness "\epsilon" allowed in the constraints.
# Create linear model: we get back a nullary function returning the
# predictions on the dataset, and the model weights.
predictions, weights = linear_model(dataset)
# Create a nullary function returning a constant tensor with the labels.
labels = lambda: tf.constant(np.ones(dataset["tot_pairs"]), dtype=tf.float32)
# Context for the optimization objective.
context = tfco.rate_context(predictions, labels)
# Constraint set.
constraint_set = []
# Context for the constraints.
for ((pos_group0, neg_group0), (pos_group1, neg_group1)) in constraint_groups:
# Context for group 0.
group0_predictions, group0_labels = group_tensors(
predictions, dataset, pos_group0, neg_group0)
context_group0 = tfco.rate_context(group0_predictions, group0_labels)
# Context for group 1.
group1_predictions, group1_labels = group_tensors(
predictions, dataset, pos_group1, neg_group1)
context_group1 = tfco.rate_context(group1_predictions, group1_labels)
# Add constraints to constraint set.
constraint_set.append(
tfco.false_negative_rate(context_group0) <= (
tfco.false_negative_rate(context_group1) + constraint_slack)
)
constraint_set.append(
tfco.false_negative_rate(context_group1) <= (
tfco.false_negative_rate(context_group0) + constraint_slack)
)
# Formulate constrained minimization problem.
problem = tfco.RateMinimizationProblem(
tfco.error_rate(context), constraint_set)
return problem, weights
# + [markdown] id="P1x4yEllRKjH" colab_type="text"
# ## Train Model
# + [markdown] id="16nddoPIrmuj" colab_type="text"
# The following function then trains the linear model by solving the above constrained optimization problem. We will handle three type of pairwise fairness constraints.
# + id="TWKxylqXPbR4" colab_type="code" colab={}
# Pairwise fairness constraint types.
MARGINAL_EQUAL_OPPORTUNITY = 0
CROSS_GROUP_EQUAL_OPPORTUNITY = 1
CROSS_AND_WITHIN_GROUP_EQUAL_OPPORTUNITY = 2
# + id="Md5pDHyBRN83" colab_type="code" colab={}
def train_model(train_set, params):
# Trains the model and returns the trained model weights, objective and
# maximum constraint violation values.
# Set up problem and model.
if params['flag_constrained']:
# Constrained optimization.
if params['constraint_type'] == MARGINAL_EQUAL_OPPORTUNITY:
constraint_groups = [((0, None), (1, None))]
elif params['constraint_type'] == CROSS_GROUP_EQUAL_OPPORTUNITY:
constraint_groups = [((0, 1), (1, 0))]
else:
constraint_groups = [((0, 1), (1, 0)), ((0, 0), (1, 1))]
else:
# Unconstrained optimization.
constraint_groups = []
problem, weights = formulate_problem(
train_set, constraint_groups, params["constraint_bound"])
# Set up optimization problem.
optimizer = tfco.ProxyLagrangianOptimizerV2(
tf.keras.optimizers.Adagrad(learning_rate=params["learning_rate"]),
num_constraints=problem.num_constraints)
# List of trainable variables.
var_list = (
[weights] + problem.trainable_variables + optimizer.trainable_variables())
# List of objectives and constraint violations during course of training.
objectives = []
constraints = []
# Run loops * iterations_per_loop full batch iterations.
for ii in xrange(params['loops']):
for ii in xrange(params['iterations_per_loop']):
optimizer.minimize(problem, var_list=var_list)
objectives.append(problem.objective())
if params['flag_constrained']:
constraints.append(max(problem.constraints()))
return weights.numpy(), objectives, constraints
# + [markdown] id="WxFJV0tvKvyR" colab_type="text"
# ## Summarize and Plot Results
# + [markdown] id="i7In7Ra7M_S7" colab_type="text"
# Having trained a model, we will need functions to summarize the various evaluation metrics and plot the trained decision boundary
# + id="CBl5KfEOPApl" colab_type="code" colab={}
def evaluate_results(model, test_set, params):
# Returns overall and group error rates for model on test set.
if params['constraint_type'] == MARGINAL_EQUAL_OPPORTUNITY:
return (error_rate(model, test_set),
[group_error_rate(model, test_set, 0),
group_error_rate(model, test_set, 1)])
else:
return (error_rate(model, test_set),
[[group_error_rate(model, test_set, 0, 0),
group_error_rate(model, test_set, 0, 1)],
[group_error_rate(model, test_set, 1, 0),
group_error_rate(model, test_set, 1, 1)]])
def display_results(model, objectives, constraints, test_set, params, ax):
# Prints evaluation results for model on test data
# and plots its decision boundary
# Evaluate model on test set and print results.
error, group_error = evaluate_results(model, test_set, params)
if params['constraint_type'] == MARGINAL_EQUAL_OPPORTUNITY:
if params['flag_constrained']:
print 'Constrained', '\t \t',
else:
print 'Test Error\t \t', 'Overall', '\t', 'Group 0', '\t', 'Group 1',
print '\t', 'Diff'
print 'Unconstrained', '\t \t',
print("%.3f" % (error_rate(model, test_set)) + '\t\t' +
"%.3f" % group_error[0] + '\t\t' + "%.3f" % group_error[1] + '\t\t'
"%.3f" % abs(group_error[0] - group_error[1]))
elif params['constraint_type'] == CROSS_GROUP_EQUAL_OPPORTUNITY:
if params['flag_constrained']:
print 'Constrained', '\t \t',
else:
print 'Test Error\t \t', 'Overall', '\t', 'Group 0/1', '\t', 'Group 1/0',
print '\t', 'Diff'
print 'Unconstrained', '\t \t',
print("%.3f" % error + '\t\t' +
"%.3f" % group_error[0][1] + '\t\t' +
"%.3f" % group_error[1][0] + '\t\t' +
"%.3f" % abs(group_error[0][1] - group_error[1][0]))
else:
if params['flag_constrained']:
print 'Constrained', '\t \t',
else:
print 'Test Error\t \t', 'Overall', '\t', 'Group 0/1', '\t', 'Group 1/0',
print '\t', 'Diff', '\t',
print '\t', 'Group 0/0', '\t', 'Group 1/1', '\t', 'Diff'
print 'Unconstrained', '\t \t',
print("%.3f" % error + '\t\t' +
"%.3f" % group_error[0][1] + '\t\t' +
"%.3f" % group_error[1][0] + '\t\t' +
"%.3f" % abs(group_error[0][1] - group_error[1][0]) + '\t\t' +
"%.3f" % group_error[0][0] + '\t\t' +
"%.3f" % group_error[1][1] + '\t\t' +
"%.3f" % abs(group_error[0][0] - group_error[1][1]))
# Plot decision boundary and progress of training objective/constraint viol.
if params['flag_constrained']:
ax[0].set_title("Model: Constrained")
else:
ax[0].set_title("Model: Unconstrained")
features = train_set['features']
plot_data(train_set, ax[0])
plot_model(model, ax[0],
[features[:, :, 0].min(), features[:, :, 0].max()],
[features[:, :, 1].min(), features[:, :, 1].max()],
"k--")
if params['flag_constrained']:
ax[1].set_title("Objective (Hinge)")
ax[1].set_xlabel("Number of epochs")
objective_curve, = ax[1].plot(range(params['loops']), objectives)
ax[2].set_title("Constraint Violation")
ax[2].set_xlabel("Number of epochs")
constraint_curve, = ax[2].plot(range(params['loops']), constraints)
# + [markdown] id="PQR0nnORRedG" colab_type="text"
# # Experimental Results
# + [markdown] id="jTYOW_EOsrWV" colab_type="text"
# We now run experiments with two types of pairwise fairness criteria: (1) marginal equal opportunity and (2) pairwise equal opportunity. In each case, we compare an unconstrained model trained to optimize the error rate and a constrained model trained with pairwise fairness constraints.
# + [markdown] id="jqxzaPTEwEIn" colab_type="text"
# ## (1) Marginal Equal Opportunity
#
#
# For a ranking model $f: \mathbb{R}^d \rightarrow \mathbb{R}$, recall:
# - $err(f)$ as the pairwise ranking error for model $f$ over all pairs of positive and negative documents
# $$
# err(f) ~=~ \mathbf{E}\big[\mathbb{I}\big(f(x) < f(x')\big) \,\big|\, y = 1,~ y' = 0\big]
# $$
#
# and we additionally define:
#
# - $err_i(f)$ as the row-marginal pairwise error over positive-negative document pairs where the pos. document is from group $i$, and the neg. document is from either groups
#
# $$
# err_i(f) = \mathbf{E}\big[\mathbb{I}\big(f(x) < f(x')\big) \,\big|\, y = 1,~ y' = 0,~ grp(x) = i\big]
# $$
#
# The constrained optimization problem we solve constraints the row-marginal pairwise errors to be similar:
#
# $$min_f\;err(f)$$
#
# $$\text{s.t. }\;|err_0(f) - err_1(f)| \leq 0.01$$
#
# + id="2iboCFdiHV4r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 260} outputId="9577981e-2c99-4b08-d351-ed3223456c89"
np.random.seed(121212)
train_set = create_dataset(num_queries=500, num_docs=10)
test_set = create_dataset(num_queries=500, num_docs=10)
model_params = {
'loops': 25,
'iterations_per_loop': 10,
'learning_rate': 1.0,
'constraint_type': MARGINAL_EQUAL_OPPORTUNITY,
'constraint_bound': 0.01}
# Plot training stats, data and model.
ff, ax = plt.subplots(1, 4, figsize=(16.0, 3.5))
# Unconstrained optimization.
model_params['flag_constrained'] = False
model, objectives, constraints = train_model(train_set, model_params)
display_results(model, objectives, constraints, test_set, model_params, [ax[0]])
# Constrained optimization.
model_params['flag_constrained'] = True
model, objectives, constraints = train_model(train_set, model_params)
display_results(model, objectives, constraints, test_set, model_params, ax[1:])
ff.tight_layout()
# + [markdown] colab_type="text" id="CorY2URkop1Y"
# ## (2) Pairwise Equal Opportunity
#
# Recall that we denote
# $err_{i,j}(f)$ as the ranking error over positive-negative document pairs where the pos. document is from group $i$, and the neg. document is from group $j$.
# $$
# err_{i, j}(f) ~=~ \mathbf{E}\big[\mathbb{I}\big(f(x) < f(x')\big) \,\big|\, y = 1,~ y' = 0,~ grp(x) = i, ~grp(x') = j\big]
# $$
#
#
# We first constrain only the cross-group errors, highlighted below.
#
# <br>
# <table border="1" bordercolor="black">
# <tr >
# <td bgcolor="white"> </td>
# <td bgcolor="white"> </td>
# <td bgcolor="white" colspan=2 align=center><b>Negative</b></td>
# </tr>
# <tr>
# <td bgcolor="white"></td>
# <td bgcolor="white"></td>
# <td>Group 0</td>
# <td>Group 1</td>
# </tr>
# <tr>
# <td bgcolor="white" rowspan=2><b>Positive</b></td>
# <td bgcolor="white">Group 0</td>
# <td bgcolor="white">$err_{0,0}$</td>
# <td bgcolor="white">$\mathbf{err_{0,1}}$</td>
# </tr>
# <tr>
# <td>Group 1</td>
# <td bgcolor="white">$\mathbf{err_{1,0}}$</td>
# <td bgcolor="white">$err_{1,1}$</td>
# </tr>
# </table>
# <br>
#
# The optimization problem we solve constraints the cross-group pairwise errors to be similar:
#
# $$min_f\; err(f)$$
# $$\text{s.t. }\;\; |err_{0,1}(f) - err_{1,0}(f)| \leq 0.01$$
#
# + id="ejzFg8McNswG" colab_type="code" outputId="0f62f733-7c86-445d-c1ec-3199df446cd4" colab={"base_uri": "https://localhost:8080/", "height": 260}
model_params = {
'loops': 50,
'iterations_per_loop': 10,
'learning_rate': 1.0,
'constraint_type': CROSS_GROUP_EQUAL_OPPORTUNITY,
'constraint_bound': 0.01}
# Plot training stats, data and model
ff, ax = plt.subplots(1, 4, figsize=(16.0, 3.5))
model_params['flag_constrained'] = False
model, objectives, constraints = train_model(train_set, model_params)
display_results(model, objectives, constraints, test_set, model_params, [ax[0]])
model_params['flag_constrained'] = True
model, objectives, constraints = train_model(train_set, model_params)
display_results(model, objectives, constraints, test_set, model_params, ax[1:])
ff.tight_layout()
# + [markdown] id="q23bS58DxPsm" colab_type="text"
# ### Cross-group + Within-group Comparisons
# We next constrain both the cross-group errors to be similar, and the within-group errors to be similar:
# <br>
# <table border="1" bordercolor="black">
# <tr >
# <td bgcolor="white"> </td>
# <td bgcolor="white"> </td>
# <td bgcolor="white" colspan=2 align=center><b>Negative</b></td>
# </tr>
# <tr>
# <td bgcolor="white"></td>
# <td bgcolor="white"></td>
# <td>Group 0</td>
# <td>Group 1</td>
# </tr>
# <tr>
# <td bgcolor="white" rowspan=2><b>Positive</b></td>
# <td bgcolor="white">Group 0</td>
# <td bgcolor="white">$\mathbf{err_{0,0}}$</td>
# <td bgcolor="white">$\mathbf{err_{0,1}}$</td>
# </tr>
# <tr>
# <td>Group 1</td>
# <td bgcolor="white">$\mathbf{err_{1,0}}$</td>
# <td bgcolor="white">$\mathbf{err_{1,1}}$</td>
# </tr>
# </table>
# <br>
#
# The constrained optimization problem we wish to solve is given by:
# $$min_f \;err(f)$$
# $$\text{s.t. }\;\;|err_{0,1}(f) - err_{1,0}(f)| \leq 0.01
# \;\;\; |err_{0,0}(f) - err_{1,1}(f)| \leq 0.01
# $$
# + colab_type="code" outputId="fd2ca829-9c3f-4672-9417-372e48c2f6b7" id="OMXzmOLpqexb" colab={"base_uri": "https://localhost:8080/", "height": 280}
model_params = {
'loops': 250,
'iterations_per_loop': 10,
'learning_rate': 0.2,
'constraint_type': CROSS_AND_WITHIN_GROUP_EQUAL_OPPORTUNITY,
'constraint_bound': 0.01
}
# Plot training stats, data and model
ff, ax = plt.subplots(1, 4, figsize=(16.0, 3.5))
model_params['flag_constrained'] = False
model, objectives, constraints = train_model(train_set, model_params)
display_results(model, objectives, constraints, test_set, model_params, [ax[0]])
model_params['flag_constrained'] = True
model, objectives, constraints = train_model(train_set, model_params)
display_results(model, objectives, constraints, test_set, model_params, ax[1:])
ff.tight_layout()
# + [markdown] id="Tg9uNCwiFP3T" colab_type="text"
# ### Conclusion
# The experimental results show that unconstrained training performs poorly on the pairwise fairness metrics, while the constrained optimization methods perform yield significantly lower fairness violations at the cost of a higher overall error rate. The constrained model ends up substatially lowering the error rates associated with the minorty group (group 1), at the cost of a slightly higher error rate for the majority group (group 0).
#
# Also, note that the quality of the learned ranking function depends on the slope of the hyperplane and is unaffected by its intercept. The hyperplane learned by the unconstrained approach ranks the majority examples (the group marked with x) well, but is not accurate at ranking the minority examples (the group marked with o).
| pairwise_fairness/synthetic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Linear classifiers demo: `predict`
#
# CPSC 340: Machine Learning and Data Mining
#
# The University of British Columbia
#
# 2017 Winter Term 2
#
# <NAME>
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['font.size'] = 16
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from plot_classifier import plot_classifier
# %autosave 0
# -
# ### Linear decision boundaries and the coefficients
#
# - Key idea: compute $w^Tx$ (or $w^Tx+\beta$) just like in linear regression, but then look at the **sign** of this function to make a prediction.
#
# $$\hat{y}_i = \textrm{sign}(w^Tx_i)$$
#
# with the convention that $y_i \in \{-1,+1\}$.
#
# We're actually fitting a hyperplane through the space, that divides it in half. The $w$ represents the tilt of the hyperplane and $\beta$ represents the intercept (offset from the origin).
N = 20
X = np.random.randn(N,2)
y = np.random.choice((-1,+1),size=N)
X[y>0,0] -= 1
X[y>0,1] += 1
model = LogisticRegression() # can change to LinearSVC and get the same plot
model.fit(X,y);
# +
# Let's play with the coefficients and see what they do.
model.coef_ = np.array([[4,2]]) # this is w
model.intercept_ = np.array([2]) # this is β
# Plot the data and decision boundary
plot_classifier(X,y,model);
# -
# Demo: change the coefficients and see what happens.
#
# Question: why do we need 2 coefficients to encode a slope? Indeed, what if we double the coefficients? (Answer coming soon!)
# #### Actual learned boundaries
lr = LogisticRegression(C=1e6) # very little regularization
lr.fit(X,y)
plot_classifier(X,y,lr);
# support vector machine
svm = LinearSVC(C=1e6) # very little regularization
svm.fit(X,y)
plot_classifier(X,y,svm);
# They look the same! Let's try a non-separable data set.
# ### Probabilities and `predict_proba`
lr.predict(X)
lr.predict_proba(X)
# Logistic regression can output probabilities of each class. Note that the probabilities add up to 1 for each row. We can plot these probabilities (say the second column).
lr = LogisticRegression(C=1e6) # very little regularization
lr.fit(X,y)
plot_classifier(X,y,lr, proba=True);
# Going back to the issue of the scale of the coefficents, let's try changing them now.
# +
lr.fit(X,y)
lr.coef_ *= 10
lr.intercept_ *= 10
plot_classifier(X,y,lr, proba=True);
# +
lr.fit(X,y)
lr.coef_ /= 5
lr.intercept_ /= 5
plot_classifier(X,y,lr, proba=True);
# -
# Thus, we have an interpretation for regularization here: smaller coefficients means less confident predictions.
#
# This makes sense -- there's a relationship between overconfidence and overfitting.
# ### Where do these probabilities come from?
#
# - The probabilities come from a softer version of taking $\textrm{sign}(w^Tx)$.
# - Instead we take $h(w^Tx)$ where $$h(z)\equiv \frac{1}{1+\exp(-z)}$$
# - This is "squishing" the output from any number to $[0,1]$.
z = np.linspace(-7,7,1000)
plt.plot(z, 1/(1+np.exp(-z)));
plt.plot((0,0),(0,0.5),'--',color='black');
plt.plot((-7,0),(0.5,0.5),'--',color='black');
plt.yticks((0,0.25,0.5,0.75,1));
plt.xlabel("$w^Tx$ (raw model output)");
plt.ylabel("predicted probability");
plt.title("logistic (sigmoid) function, $1/(1+\exp(-w^Tx))$");
# - Thus, checking whether the raw model output is positive or negative corresponds to checking whether the probability is greater or less than 0.5. Makes sense!
# - Furthermore, making the coefficients larger (e.g. doubling) pushes $w^Tx$ away from zero and towards the edges of the sigmoid. Hence the behaviour we observed.
# - We can sort of see the sigmoid in the probability contours above: for small coefficients it appears linear; for large coefficients we see the saturation.
# ### Interpretation of the coefficients
#
# - Above we've seen a geometric interpretation
# - The interpretation from linear regression also holds:
# - the $j$th coefficient tells us how feature $j$ affects the prediction
# - if $w_j>0$ then increasing $x_{ij}$ moves us toward predicting $+1$
# - if $w_j<0$ then increasing $x_{ij}$ moves us toward prediction $-1$
# - if $w_j=0$ then the feature is not used in making a prediction
# ### Preview of next lecture
#
# - We need a loss function for our linear classifiers.
# - Number of errors is not smooth/convex.
# - But, maybe, higher probabilities on the correct class?
| lectures/L18demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 한국어 적용 코드
import matplotlib.font_manager as fm
font_location = "C:\\Windows\Fonts\malgunbd.ttf"
font_name = fm.FontProperties(fname=font_location).get_name()
mpl.rc("font", family=font_name)
mu = 0 # 평균 : 0
std = 1 # 표준편차 : 1
rv = sp.stats.norm(mu, std)
xx = np.linspace(-5, 5, 100)
plt.plot(xx, rv.pdf(xx))
plt.ylabel("p(x)")
plt.title("정규 분포의 확률 밀도 함수(pdf)")
plt.show()
x = rv.rvs(100, random_state=0)
x
# kde : 본 함수를 유연하게 그리기, fit : 비교할 함수
sns.distplot(x, kde=True, fit=sp.stats.norm)
plt.show()
# ### Q-Q Plot
# - 가지고 있는 분포를 분위로 쪼개서 정규분포의 분위와 일대일 대응시켜봐서
# - 정규분포인지 아닌지 시각적으로 판단하도록 도와줌
np.random.seed(0)
x = np.random.randn(100) # randn 자체가 정규분포를 따라 값이 뽑힘
plt.figure(figsize=(7, 7))
sp.stats.probplot(x, plot=plt)
# plt.axis("equal")
plt.show()
np.random.seed(0)
x = np.random.rand(100) # uniform 분포이기 때문에 short tail형태로 그래프가 나옴.
plt.figure(figsize=(7, 7))
sp.stats.probplot(x, plot=plt)
plt.ylim(-0.5, 1.5)
plt.show()
# +
X = np.random.rand(2,10) # 2 X 10 matrix
print(X,"\n\n",X.mean(axis=1),"\n\n", X.mean(axis=0))
# axis = 1 : 각 행의 연산(평균, 합계....)
# axis = 0 : 각 열의 연산
# +
# 여기에서는 0부터 1까지의 균일 분포의 기댓값이 0.5 ,
# 분산이 1/12 라는 사실을 이용
np.random.seed(0)
xx = np.linspace(-2, 2, 100)
plt.figure(figsize=(6,9))
for i, N in enumerate([1, 2, 20]):
X = np.random.rand(5000, N) # uniform 분포
# 세트당(각 행벡터) 평균을 내고 정규화까지 하는 과정...총 5000세트
Xbar = (X.mean(axis=1) - 0.5) * np.sqrt(12 * N)
ax = plt.subplot(3, 2, 2 * i + 1)
sns.distplot(Xbar, bins=10, kde=False, norm_hist=True)
plt.xlim(-5, 5)
plt.yticks([])
ax.set_title("N = {0}".format(N))
plt.subplot(3, 2, 2 * i + 2)
sp.stats.probplot(Xbar, plot=plt)
plt.tight_layout()
plt.show()
# +
# N = 1 인 경우를 좀 더 자세하게 보자.
X = np.random.rand(5000, 1) # uniform 분포
# 세트당(각 행벡터) 평균을 내고 정규화까지 하는 과정...총 5000세트
Xbar = (X.mean(axis=1) - 0.5) * np.sqrt(12 * 1)
print(type(Xbar))
sns.distplot(Xbar, bins=10, kde=False, norm_hist=True)
| probability/gaussian_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0. Python Basics
#
# ### Finanzas Cuantitativas y Ciencia de Datos
# #### <NAME> y <NAME>
# #### ITAM Primavera 2019
#
#
# Jugando con este notebook pueden familiarizarse un poco con python y su uso para Ciencia de Datos.
#
# ---
#
# _INSTRUCCIONES:_
# * Todas las celdas se corren haciendo __Shift + Enter__ o __Ctrl + Enter__
#
# _NOTAS:_
# * _Notebook adaptado de distintas fuentes (Coursera y Github) y proyectos personales_
x = 1
y = 2
x + y
x
# ##### Crear funciones:
# `add_numbers` is a function that takes two numbers and adds them together.
# +
def add_numbers(x, y):
return x + y
add_numbers(1, 2)
# -
# `add_numbers` updated to take an optional 3rd parameter.
#
# Using `print` allows printing of multiple expressions within a single cell.
# +
def add_numbers(x,y,z=None):
if (z==None):
return x+y
else:
return x+y+z
print(add_numbers(1, 2))
print(add_numbers(1, 2, 3))
# -
# `add_numbers` updated to take an optional flag parameter.
# +
def add_numbers(x, y, z=None, flag=False):
if (flag):
print('Flag is true!')
if (z==None):
return x + y
else:
return x + y + z
print(add_numbers(1, 2, flag=True))
# -
# Assign function `add_numbers` to variable `a`.
# +
def add_numbers(x,y):
return x+y
a = add_numbers
a(1,2)
# -
# ## Types and Sequences
# <br>
# Use `type` to return the object's type.
type('This is a string')
type(None)
type(1)
type(1.0)
type(add_numbers)
# Tuples are an immutable data structure (cannot be altered).
x = (1, 'a', 2, 'b')
type(x)
# Lists are a mutable data structure.
x = [1, 'a', 2, 'b']
type(x)
# Use `append` to append an object to a list.
x.append('t')
print(x)
# This is an example of how to loop through each item in the list.
for item in x:
print(item)
# Or using the indexing operator:
i=0
while( i != len(x) ):
print(x[i])
i = i + 1
# Use `+` to concatenate lists.
[1,2] + [3,4]
# Use `*` to repeat lists.
[1]*3
# Use the `in` operator to check if something is inside a list.
't' in x
# Use bracket notation to slice a string.
x = 'This is a string'
print(x[0]) #first character
print(x[0:1]) #first character, but we have explicitly set the end character
print(x[0:2]) #first two characters
# This will return the last element of the string.
x[-1]
# Return the slice starting from the 4th element from the end and stopping before the 2nd element from the end.
x[-4:-2]
# This is a slice from the beginning of the string and stopping before the 3rd element.
x[:3]
# <br>
# And this is a slice starting from the 3rd element of the string and going all the way to the end.
x[3:]
# +
firstname = 'Rodrigo'
lastname = 'Lugo-Frias'
print(firstname + ' ' + lastname)
print(firstname*3)
print('Rodrigo' in firstname)
# -
# `split` returns a list of all the words in a string, or a list split on a specific character.
firstname = '<NAME>'.split(' ')[0] # [0] selects the first element of the list
lastname = '<NAME>'.split(' ')[-1] # [-1] selects the last element of the list
print(firstname)
print(lastname)
# Make sure you convert objects to strings before concatenating.
'RLF' + 2 # This instruction returns an error
'RLF' + str(2) # This instruction does not return an error
# <br>
# Dictionaries associate keys with values.
x = {'<NAME>': '<EMAIL>', '<NAME>': '<EMAIL>'}
x['<NAME>'] # Retrieve a value by using the indexing operator
x['<NAME>'] = None
x['<NAME>'] = '<EMAIL>'
x
# Iterate over all of the keys:
for name in x:
print(x[name])
# Iterate over all of the values:
for email in x.values():
print(email)
# Iterate over all of the items in the list:
for name, email in x.items():
print(name)
print(email)
# You can unpack a sequence into different variables:
x = ('Rodrigo', 'Lugo', '<EMAIL>')
fname, lname, email = x # Variables assigned to the tuple
fname
lname
# <br>
# Make sure the number of values you are unpacking matches the number of variables being assigned.
x = ('Rodrigo', 'Lugo', '<EMAIL>', 'Mexico City')
fname, lname, email = x # Too many values to unpack
# Built in method for convenient string formatting.
# +
sales_record = {
'price': 3.24,
'num_items': 4,
'person': 'Chris'}
sales_statement = '{} bought {} item(s) at a price of {} each for a total of {}'
# Each {} means python is waiting for a value from a particular source
# See example below
# Call the format method for this particular string
print(sales_statement.format(sales_record['person'],
sales_record['num_items'],
sales_record['price'],
sales_record['num_items']*sales_record['price']))
# -
# <br>
# # Reading and Writing CSV files
# <br>
# Let's import our datafile mpg.csv, which contains fuel economy data for 234 cars.
#
# * mpg : miles per gallon
# * class : car classification
# * cty : city mpg
# * cyl : # of cylinders
# * displ : engine displacement in liters
# * drv : f = front-wheel drive, r = rear wheel drive, 4 = 4wd
# * fl : fuel (e = ethanol E85, d = diesel, r = regular, p = premium, c = CNG)
# * hwy : highway mpg
# * manufacturer : automobile manufacturer
# * model : model of car
# * trans : type of transmission
# * year : model year
# +
import csv
# %precision 2
with open('Data/mpg.csv') as csvfile:
mpg = list(csv.DictReader(csvfile))
mpg[:3] # The first three dictionaries in our list.
# -
# `csv.Dictreader` has read in each row of our csv file as a dictionary. `len` shows that our list is comprised of 234 dictionaries.
len(mpg)
# `keys` gives us the column names of our csv.
mpg[0].keys()
# This is how to find the average cty fuel economy across all cars. All values in the dictionaries are strings, so we need to convert to float.
sum(float(d['cty']) for d in mpg) / len(mpg)
# Similarly this is how to find the average hwy fuel economy across all cars.
sum(float(d['hwy']) for d in mpg) / len(mpg)
# Use `set` to return the unique values for the number of cylinders the cars in our dataset have.
cylinders = set(d['cyl'] for d in mpg)
cylinders
# Here's a more complex example where we are grouping the cars by number of cylinder, and finding the average cty mpg for each group.
# +
CtyMpgByCyl = []
for c in cylinders: # iterate over all the cylinder levels
summpg = 0
cyltypecount = 0
for d in mpg: # iterate over all dictionaries
if d['cyl'] == c: # if the cylinder level type matches,
summpg += float(d['cty']) # add the cty mpg
cyltypecount += 1 # increment the count
CtyMpgByCyl.append((c, summpg / cyltypecount)) # append the tuple ('cylinder', 'avg mpg')
CtyMpgByCyl.sort(key=lambda x: x[0])
CtyMpgByCyl
# -
# <br>
# Use `set` to return the unique values for the class types in our dataset.
vehicleclass = set(d['class'] for d in mpg) # what are the class types
vehicleclass
# And here's an example of how to find the average hwy mpg for each class of vehicle in our dataset.
# +
HwyMpgByClass = []
for t in vehicleclass: # iterate over all the vehicle classes
summpg = 0
vclasscount = 0
for d in mpg: # iterate over all dictionaries
if d['class'] == t: # if the cylinder amount type matches,
summpg += float(d['hwy']) # add the hwy mpg
vclasscount += 1 # increment the count
HwyMpgByClass.append((t, summpg / vclasscount)) # append the tuple ('class', 'avg mpg')
HwyMpgByClass.sort(key=lambda x: x[1])
HwyMpgByClass
# -
# ## Dates and Times
import datetime as dt
import time as tm
# `time` returns the current time in seconds since the Epoch. (January 1st, 1970)
tm.time()
# Convert the timestamp to datetime.
dtnow = dt.datetime.fromtimestamp(tm.time())
dtnow
# Datetime attributes:
dtnow.year, dtnow.month, dtnow.day, dtnow.hour, dtnow.minute, dtnow.second # get year, month, day, etc.from a datetime
# `timedelta` is a duration expressing the difference between two dates.
delta = dt.timedelta(days = 100) # create a timedelta of 100 days
delta
# `date.today` returns the current local date.
today = dt.date.today()
today - delta # the date 100 days ago
today > today-delta # compare dates
# ## Lambda and List Comprehensions
# Example of lambda that takes in three parameters and adds the first two.
my_function = lambda a, b, c : a + b
my_function(1, 2, 3)
# +
# Exercise
people = ['Dr. <NAME>', 'Dr. <NAME>', 'Dr. VG <NAME>', 'Dr. <NAME>']
def split_title_and_name(person):
return person.split()[0] + ' ' + person.split()[-1]
split_title_and_name(people[0])
sp_ti_na = lambda person: person.split()[0] + ' ' + person.split()[-1]
sp_ti_na(people[0])
# Convert this function into a lambda
# +
# Solution
#option 1
for person in people:
print(split_title_and_name(person) == (lambda x: x.split()[0] + ' ' + x.split()[-1])(person))
#option 2
list(map(split_title_and_name, people)) == list(map(lambda person: person.split()[0] + ' ' + person.split()[-1], people))
# -
# Let's iterate from 0 to 999 and return the even numbers.
my_list = []
for number in range(0, 1000):
if number % 2 == 0:
my_list.append(number)
my_list
# <br>
# Now the same thing but with list comprehension.
my_list = [number for number in range(0,1000) if number % 2 == 0]
my_list
# ## Numerical Python (NumPy)
import numpy as np
# ### Creating Arrays
# Create a list and convert it to a numpy array
mylist = [1, 2, 3]
x = np.array(mylist)
x
# Or just pass in a list directly
y = np.array([4, 5, 6])
y
# <br>
# Pass in a list of lists to create a multidimensional array.
m = np.array([[7, 8, 9], [10, 11, 12]])
m
# <br>
# Use the shape method to find the dimensions of the array. (rows, columns)
m.shape
# <br>
# `arange` returns evenly spaced values within a given interval.
n = np.arange(0, 30, 2) # start at 0 count up by 2, stop before 30
n
# <br>
# `reshape` returns an array with the same data with a new shape.
n = n.reshape(3, 5) # reshape array to be 3x5
n
# <br>
# `linspace` returns evenly spaced numbers over a specified interval.
o = np.linspace(0, 4, 9) # return 9 evenly spaced values from 0 to 4
o
# <br>
# `resize` changes the shape and size of array in-place.
o.resize(3, 3)
o
# <br>
# `ones` returns a new array of given shape and type, filled with ones.
np.ones((3, 2))
# <br>
# `zeros` returns a new array of given shape and type, filled with zeros.
np.zeros((2, 3))
# <br>
# `eye` returns a 2-D array with ones on the diagonal and zeros elsewhere.
np.eye(3)
# <br>
# `diag` extracts a diagonal or constructs a diagonal array.
np.diag(y)
# <br>
# Create an array using repeating list (or see `np.tile`)
np.array([1, 2, 3] * 3)
# <br>
# Repeat elements of an array using `repeat`.
np.repeat([1, 2, 3], 3)
# #### Combining Arrays
p = np.ones([2, 3], int)
p
# <br>
# Use `vstack` to stack arrays in sequence vertically (row wise).
np.vstack([p, 2*p])
# <br>
# Use `hstack` to stack arrays in sequence horizontally (column wise).
np.hstack([p, 2*p])
# <br>
# ## Operations
# Use `+`, `-`, `*`, `/` and `**` to perform element wise addition, subtraction, multiplication, division and power.
print(x + y) # elementwise addition [1 2 3] + [4 5 6] = [5 7 9]
print(x - y) # elementwise subtraction [1 2 3] - [4 5 6] = [-3 -3 -3]
print(x * y) # elementwise multiplication [1 2 3] * [4 5 6] = [4 10 18]
print(x / y) # elementwise divison [1 2 3] / [4 5 6] = [0.25 0.4 0.5]
print(x**2) # elementwise power [1 2 3] ^2 = [1 4 9]
# <br>
# **Dot Product:**
#
# $ \begin{bmatrix}x_1 \ x_2 \ x_3\end{bmatrix}
# \cdot
# \begin{bmatrix}y_1 \\ y_2 \\ y_3\end{bmatrix}
# = x_1 y_1 + x_2 y_2 + x_3 y_3$
x.dot(y) # dot product 1*4 + 2*5 + 3*6
z = np.array([y, y**2])
print(len(z)) # number of rows of array
# <br>
# Let's look at transposing arrays. Transposing permutes the dimensions of the array.
z = np.array([y, y**2])
z
# <br>
# The shape of array `z` is `(2,3)` before transposing.
z.shape
# <br>
# Use `.T` to get the transpose.
z.T
# <br>
# The number of rows has swapped with the number of columns.
z.T.shape
# <br>
# Use `.dtype` to see the data type of the elements in the array.
z.dtype
# <br>
# Use `.astype` to cast to a specific type.
z = z.astype('f')
z.dtype
# <br>
# ## Math Functions
# Numpy has many built in math functions that can be performed on arrays.
a = np.array([-4, -2, 1, 3, 5])
a.sum()
a.max()
a.min()
a.mean()
a.std()
# <br>
# `argmax` and `argmin` return the index of the maximum and minimum values in the array.
a.argmax()
a.argmin()
# <br>
# ## Indexing / Slicing
s = np.arange(13)**2
s
# <br>
# Use bracket notation to get the value at a specific index. Remember that indexing starts at 0.
s[0], s[4], s[-1]
# <br>
# Use `:` to indicate a range. `array[start:stop]`
#
#
# Leaving `start` or `stop` empty will default to the beginning/end of the array.
s[1:5]
# <br>
# Use negatives to count from the back.
s[-4:]
# <br>
# A second `:` can be used to indicate step-size. `array[start:stop:stepsize]`
#
# Here we are starting 5th element from the end, and counting backwards by 2 until the beginning of the array is reached.
s[-5::-2]
# <br>
# Let's look at a multidimensional array.
r = np.arange(36)
r.resize((6, 6))
r
# <br>
# Use bracket notation to slice: `array[row, column]`
r[2, 2]
# <br>
# And use : to select a range of rows or columns
r[3, 3:6]
# <br>
# Here we are selecting all the rows up to (and not including) row 2, and all the columns up to (and not including) the last column.
r[:2, :-1]
# <br>
# This is a slice of the last row, and only every other element.
r[-1, ::2]
# <br>
# We can also perform conditional indexing. Here we are selecting values from the array that are greater than 30. (Also see `np.where`)
r[r > 30]
# <br>
# Here we are assigning all values in the array that are greater than 30 to the value of 30.
r[r > 30] = 30
r
# <br>
# ## Copying Data
# Be careful with copying and modifying arrays in NumPy!
#
#
# `r2` is a slice of `r`
r2 = r[:3,:3]
r2
# <br>
# Set this slice's values to zero ([:] selects the entire array)
r2[:] = 0
r2
# <br>
# `r` has also been changed!
r
# <br>
# To avoid this, use `r.copy` to create a copy that will not affect the original array
r_copy = r.copy()
r_copy
# <br>
# Now when r_copy is modified, r will not be changed.
r_copy[:] = 10
print(r_copy, '\n')
print(r)
# <br>
# ### Iterating Over Arrays
# Let's create a new 4 by 3 array of random numbers 0-9.
test = np.random.randint(0, 10, (4,3))
test
# <br>
# Iterate by row:
for row in test:
print(row)
# <br>
# Iterate by index:
for i in range(len(test)):
print(test[i])
# <br>
# Iterate by row and index:
for i, row in enumerate(test):
print('row', i, 'is', row)
# <br>
# Use `zip` to iterate over multiple iterables.
test2 = test**2
test2
for i, j in zip(test, test2):
print(i,'+',j,'=',i+j)
| 0. Python Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # SageMaker PySpark Custom Estimator MNIST Example
#
# 1. [Introduction](#Introduction)
# 2. [Setup](#Setup)
# 3. [Loading the Data](#Loading-the-Data)
# 4. [Create a custom SageMakerEstimator](#Create-a-custom-SageMakerEstimator)
# 5. [Inference](#Inference)
# 6. [Clean-up](#Clean-up)
# 7. [More on SageMaker Spark](#More-on-SageMaker-Spark)
#
# ## Introduction
# This notebook will show how to cluster handwritten digits through the SageMaker PySpark library.
#
# We will manipulate data through Spark using a SparkSession, and then use the SageMaker Spark library to interact with SageMaker for training and inference.
# We will use a custom estimator to perform the classification task, and train and infer using that custom estimator.
#
# You can visit SageMaker Spark's GitHub repository at https://github.com/aws/sagemaker-spark to learn more about SageMaker Spark.
#
# This notebook was created and tested on an ml.m4.xlarge notebook instance.
# ## Setup
#
# First, we import the necessary modules and create the `SparkSession` with the SageMaker-Spark dependencies attached.
# +
import os
import boto3
from pyspark import SparkContext, SparkConf
from pyspark.sql import SparkSession
import sagemaker
from sagemaker import get_execution_role
import sagemaker_pyspark
role = get_execution_role()
# Configure Spark to use the SageMaker Spark dependency jars
jars = sagemaker_pyspark.classpath_jars()
classpath = ":".join(sagemaker_pyspark.classpath_jars())
# See the SageMaker Spark Github to learn how to connect to EMR from a notebook instance
spark = SparkSession.builder.config("spark.driver.extraClassPath", classpath)\
.master("local[*]").getOrCreate()
spark
# -
# ## Loading the Data
#
# Now, we load the MNIST dataset into a Spark Dataframe, which dataset is available in LibSVM format at
#
# `s3://sagemaker-sample-data-[region]/spark/mnist/`
#
# where `[region]` is replaced with a supported AWS region, such as us-east-1.
#
# In order to train and make inferences our input DataFrame must have a column of Doubles (named "label" by default) and a column of Vectors of Doubles (named "features" by default).
#
# Spark's LibSVM DataFrameReader loads a DataFrame already suitable for training and inference.
#
# Here, we load into a DataFrame in the SparkSession running on the local Notebook Instance, but you can connect your Notebook Instance to a remote Spark cluster for heavier workloads. Starting from EMR 5.11.0, SageMaker Spark is pre-installed on EMR Spark clusters. For more on connecting your SageMaker Notebook Instance to a remote EMR cluster, please see [this blog post](https://aws.amazon.com/blogs/machine-learning/build-amazon-sagemaker-notebooks-backed-by-spark-in-amazon-emr/).
# +
import boto3
region = boto3.Session().region_name
trainingData = spark.read.format('libsvm')\
.option('numFeatures', '784')\
.load('s3a://sagemaker-sample-data-{}/spark/mnist/train/'.format(region))
testData = spark.read.format('libsvm')\
.option('numFeatures', '784')\
.load('s3a://sagemaker-sample-data-{}/spark/mnist/test/'.format(region))
trainingData.show()
# -
# MNIST images are 28x28, resulting in 784 pixels. The dataset consists of images of digits going from 0 to 9, representing 10 classes.
#
# In each row:
# * The `label` column identifies the image's label. For example, if the image of the handwritten number is the digit 5, the label value is 5.
# * The `features` column stores a vector (`org.apache.spark.ml.linalg.Vector`) of `Double` values. The length of the vector is 784, as each image consists of 784 pixels. Those pixels are the features we will use.
#
#
#
# As we are interested in clustering the images of digits, the number of pixels represents the feature vector, while the number of classes represents the number of clusters we want to find.
# ## Create a custom SageMakerEstimator
#
# SageMaker-Spark provides several classes that extend SageMakerEstimator to use SageMaker-provided algorithms, like `KMeansSageMakerEstimator` to run the SageMaker-provided K-Means algorithm. These classes are `SageMakerEstimator` with certain default values passed in. You can use SageMaker-Spark with any algorithm (provided by Amazon or your own model) that runs on Amazon SageMaker by creating a `SageMakerEstimator`.
#
# In this example, we'll re-create the `KMeansSageMakerEstimator` into an equivalent SageMakerEstimator.
# +
from sagemaker.amazon.amazon_estimator import get_image_uri
from sagemaker_pyspark import SageMakerEstimator
from sagemaker_pyspark.transformation.deserializers import KMeansProtobufResponseRowDeserializer
from sagemaker_pyspark.transformation.serializers import ProtobufRequestRowSerializer
from sagemaker_pyspark import IAMRole
from sagemaker_pyspark import RandomNamePolicyFactory
from sagemaker_pyspark import EndpointCreationPolicy
# Create an Estimator from scratch
estimator = SageMakerEstimator(
trainingImage = get_image_uri(region, 'kmeans'), # Training image
modelImage = get_image_uri(region, 'kmeans'), # Model image
requestRowSerializer = ProtobufRequestRowSerializer(),
responseRowDeserializer = KMeansProtobufResponseRowDeserializer(),
hyperParameters = {"k": "10", "feature_dim": "784"}, # Set parameters for K-Means
sagemakerRole = IAMRole(role),
trainingInstanceType = "ml.m4.xlarge",
trainingInstanceCount = 1,
endpointInstanceType = "ml.t2.medium",
endpointInitialInstanceCount = 1,
trainingSparkDataFormat = "sagemaker",
namePolicyFactory = RandomNamePolicyFactory("sparksm-4-"),
endpointCreationPolicy = EndpointCreationPolicy.CREATE_ON_TRANSFORM
)
# -
# The main parts of a `SageMakerEstimator` are:
# * `trainingImage`: the Docker Registry path where the training image is hosted - can be a custom Docker image hosting your own model, or one of the [Amazon provided images](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html)
# * `modelImage`: the Docker Registry path where the inference image is used - can be a custom Docker image hosting your own model, or one of the [Amazon provided images](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html)
# * `hyperparameters`: the hyper-parameters of the algorithm being created - the values passed in need to be of type string
#
# To put this `SageMakerEstimator` back into context, let's look at the below architecture that shows what actually runs on the notebook instance and on SageMaker.
# 
# Let's train this estimator by calling fit on it with the training data. Please note the below code will take several minutes to run and create all the resources needed for this model.
customModel = estimator.fit(trainingData)
# ## Inference
#
# Now we transform our DataFrame.
# To do this, we serialize each row's "features" Vector of Doubles into a Protobuf format for inference against the Amazon SageMaker Endpoint. We deserialize the Protobuf responses back into our DataFrame. This serialization and deserialization is handled automatically by the `transform()` method:
transformedData = customModel.transform(testData)
transformedData.show()
# How well did the custom algorithm perform? Let us display the digits from each of the clusters and manually inspect the results:
# +
from pyspark.sql.types import DoubleType
import matplotlib.pyplot as plt
import numpy as np
import string
# Helper function to display a digit
def showDigit(img, caption='', xlabel='', subplot=None):
if subplot==None:
_,(subplot)=plt.subplots(1,1)
imgr=img.reshape((28,28))
subplot.axes.get_xaxis().set_ticks([])
subplot.axes.get_yaxis().set_ticks([])
plt.title(caption)
plt.xlabel(xlabel)
subplot.imshow(imgr, cmap='gray')
def displayClusters(data):
images = np.array(data.select("features").cache().take(250))
clusters = data.select("closest_cluster").cache().take(250)
for cluster in range(10):
print('\n\n\nCluster {}:'.format(string.ascii_uppercase[cluster]))
digits = [ img for l, img in zip(clusters, images) if int(l.closest_cluster) == cluster ]
height=((len(digits)-1)//5)+1
width=5
plt.rcParams["figure.figsize"] = (width,height)
_, subplots = plt.subplots(height, width)
subplots=np.ndarray.flatten(subplots)
for subplot, image in zip(subplots, digits):
showDigit(image, subplot=subplot)
for subplot in subplots[len(digits):]:
subplot.axis('off')
plt.show()
displayClusters(transformedData)
# -
# ## Clean-up
# Since we don't need to make any more inferences, now we delete the resources (endpoints, models, configurations, etc):
# +
# Delete the resources
from sagemaker_pyspark import SageMakerResourceCleanup
def cleanUp(model):
resource_cleanup = SageMakerResourceCleanup(model.sagemakerClient)
resource_cleanup.deleteResources(model.getCreatedResources())
cleanUp(customModel)
# -
# ## More on SageMaker Spark
#
# The SageMaker Spark Github repository has more about SageMaker Spark, including how to use SageMaker Spark using the Scala SDK: https://github.com/aws/sagemaker-spark
#
| sagemaker-spark/pyspark_mnist/pyspark_mnist_custom_estimator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''vk-igibson'': conda)'
# name: python3
# ---
import pickle
# import torch
import numpy
import matplotlib.pyplot as plt
with open("../pkl_files/sample_states.pkl", "rb") as f:
save_dict = pickle.load(f)
save_dict["state_hist"][0].keys()
save_dict["state_hist"][0]["sensor"].shape
import numpy as np
np.sort(save_dict["state"][0]["task_obs"])
plt.imshow(save_dict["state"][850]["rgb"])
plt.imshow(save_dict["state"][865]["depth"])
save_dict["state"][850]["task_obs"]
save_dict.keys()
plt.plot(save_dict["reward"])
| igibson_usage/notebooks/tiago_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python Multi
# language: python
# name: multi
# ---
# !cd
import matplotlib.pyplot as plt
import numpy as np
# +
# nrows, ncols int, default: 1
# -
fig, ax = plt.subplots(nrows=2, ncols=2)
ax[1,1].plot([1, 2, 3, 4], [1, 4, 2, 3])
ax[0,0].plot([1, 2, 3, 4], [3, 2, 1, 2])
ax[1,1].set_title('Simple plot')
fig.suptitle('Vertically stacked subplots')
fig.supxlabel('Vertically stacked subplots')
fig.supylabel('Vertically stacked subplots')
# +
#fig.suptitle('Vertically stacked subplots')
# -
fig
fig
ax[1,0].plot([1, 2, 3, 4], [1, 4, 2, 3])
#plt.show()
plt.plot([1, 2, 3, 4], [1, 4, 2, 3])
fig = plt.figure() # an empty figure with no Axes
fig, ax = plt.subplots(2,2) # a figure with a single Axes
# +
x = np.linspace(0, 2, 100)
# Note that even in the OO-style, we use `.pyplot.figure` to create the figure.
fig, ax = plt.subplots() # Create a figure and an axes.
ax.plot(x, x, label='linear') # Plot some data on the axes.
ax.plot(x, x**2, label='quadratic') # Plot more data on the axes...
ax.plot(x, x**3, label='cubic') # ... and some more.
ax.set_xlabel('x label') # Add an x-label to the axes.
ax.set_ylabel('y label') # Add a y-label to the axes.
ax.set_title("Simple Plot") # Add a title to the axes.
ax.legend(loc=10) # Add a legend.
# -
ax[1,0]
fig, axs = plt.subplots(2, 2) # a figure with a 2x2 grid of Axes
import pandas
a = pandas.DataFrame(np.random.rand(4, 5), columns = list('abcde'))
import numpy as np
np_array
np_array = a.values # DataFrame -> Array
np_array.mean()
np_array.var()
np_array.std()
a_asarray = a.values
b = np.matrix([[1, 2], [3, 4]])
b
b_asarray = np.asarray(b)
x=a
x.std()
type(x)
x.div(10)
# Note that even in the OO-style, we use `.pyplot.figure` to create the figure.
fig, ax = plt.subplots() # Create a figure and an axes.
ax.plot(x, x, label='linear') # Plot some data on the axes.
ax.plot(x, x**2, label='quadratic') # Plot more data on the axes...
ax.plot(x, x**3, label='cubic') # ... and some more.
ax.set_xlabel('x label') # Add an x-label to the axes.
ax.set_ylabel('y label') # Add a y-label to the axes.
ax.set_title("Simple Plot") # Add a title to the axes.
ax.legend() # Add a legend.
# +
x = np.linspace(0, 2, 100)
# Note that even in the OO-style, we use `.pyplot.figure` to create the figure.
fig, ax = plt.subplots() # Create a figure and an axes.
ax.plot(x, x, label='linear') # Plot some data on the axes.
ax.plot(x, x**2, label='quadratic') # Plot more data on the axes...
ax.plot(x, x**3, label='cubic') # ... and some more.
ax.set_xlabel('x label') # Add an x-label to the axes.
ax.set_ylabel('y label') # Add a y-label to the axes.
ax.set_title("Simple Plot") # Add a title to the axes.
ax.legend() # Add a legend.
# +
x = np.linspace(0, 2, 100)
plt.plot(x, x, label='linear') # Plot some data on the (implicit) axes.
plt.plot(x, x**2, label='quadratic') # etc.
plt.plot(x, x**3, label='cubic')
# Note that even in the OO-style, we use `.pyplot.figure` to create the figure.
fig, ax = plt.subplots() # Create a figure and an axes.
ax.plot(x, x, label='linear') # Plot some data on the axes.
ax.plot(x, x**2, label='quadratic') # Plot more data on the axes...
ax.plot(x, x**3, label='cubic') # ... and some more.
ax.legend(loc=0, fontsize='xx-large') # Add a legend.
# -
plt.xlabel('x label')
plt.ylabel('y label')
plt.title("Simple Plot")
plt.legend()
| plot1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring ways of finding hair in images
# This is an exploratory notebook - it doesn't really go in order, but I wanted to save some of the things I tried, for posterity. I did not use anything in this notebook for the final product.
import cv2
import numpy as np
import matplotlib.pyplot as plt
import pymeanshift as pms
import os
from glob import glob
from PIL import Image
# ## Resize images
# This function takes the images and resizes them to 300x400 pixels.
# +
imgs = glob('./hair_images/*')
def resize(imgs):
for img in imgs:
filename = img.split('/')[2]
filename = filename.split('.')[0]
if filename == 'image_urls':
continue
if os.path.exists('./resized_hair_images/'+filename+'_resized.jpg') == True:
continue
im = Image.open(img)
imResize = im.resize((300,400), Image.ANTIALIAS)
imResize.save('./resized_hair_images/'+filename+'_resized.jpg', 'JPEG', quality=90)
resize(imgs)
# -
# ## Find hair via contours
# This code explores using skin tones to find adjacent hair.
min_YCrCb = np.array([0,133,77],np.uint8)
max_YCrCb = np.array([255,173,127],np.uint8)
# +
img = cv2.imread('wavy.jpg')
imageYCrCb = cv2.cvtColor(img,cv2.COLOR_BGR2YCR_CB)
skinRegion = cv2.inRange(imageYCrCb,min_YCrCb,max_YCrCb)
contours, hierarchy = cv2.findContours(skinRegion, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
# Draw the contour on the source image
for i, c in enumerate(contours):
area = cv2.contourArea(c)
if area > 9000:
cv2.drawContours(img, contours, i, (0, 255, 0), 3)
# Display the source image
cv2.imwrite('output1.jpg', img)
plt.figure(figsize=(12,12))
plt.imshow(img[:,:,::-1])
plt.show()
# -
# ## Find faces
# Using the the built-in Haar Cascades in OpenCV to find faces.
face_cascade = cv2.CascadeClassifier('/Users/joanna/anaconda2/envs/py37/lib/python3.7/site-packages/cv2/data/haarcascade_frontalface_default.xml')
eye_cascade = cv2.CascadeClassifier('/Users/joanna/anaconda2/envs/py37/lib/python3.7/site-packages/cv2/data/haarcascade_eye.xml')
# +
img = cv2.imread('hair.jpg')
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
faces = face_cascade.detectMultiScale(gray, scaleFactor=1.5, minNeighbors=3)
for (x,y,w,h) in faces:
cv2.rectangle(img,(x,y),(x+w,y+h),(255,0,0),2)
roi_gray = gray[y:y+h, x:x+w]
roi_color = img[y:y+h, x:x+w]
eyes = eye_cascade.detectMultiScale(roi_gray,minSize=(100, 100), maxSize=(150,150))
for (ex,ey,ew,eh) in eyes:
cv2.rectangle(roi_color,(ex,ey),(ex+ew,ey+eh),(0,255,0),2)
cv2.imwrite('output_hair.jpg', img)
plt.figure(figsize=(12,12))
plt.imshow(img[:,:,::-1])
plt.show()
# +
img = cv2.imread('hair.jpg')
img = cv2.cvtColor(img ,cv2.COLOR_BGR2GRAY)
cv2.imwrite('blackwhite.jpg', img)
(segmented_image, labels_image, number_regions) = pms.segment(img, spatial_radius=6,
range_radius=4.5, min_density=1500)
# +
plt.figure(figsize=(12,12))
plt.imshow(segmented_image)
plt.show()
cv2.imwrite('blackwhite_seg.jpg', segmented_image)
# -
# ## Playing with edge detection
# This didn't work very well.
# +
img = cv2.imread('hair_images/903ff49749.jpg',0)
edges = cv2.Canny(img,100,250)
plt.figure(figsize=(20,20))
plt.subplot(121),plt.imshow(img,cmap = 'gray')
plt.title('Original Image'), plt.xticks([]), plt.yticks([])
plt.subplot(122),plt.imshow(edges,cmap = 'gray')
plt.title('Edge Image'), plt.xticks([]), plt.yticks([])
plt.show()
# +
def auto_canny(image, sigma=0.33):
# compute the median of the single channel pixel intensities
v = np.median(image)
# apply automatic Canny edge detection using the computed median
lower = int(max(0, (1.0 - sigma) * v))
upper = int(min(255, (1.0 + sigma) * v))
edged = cv2.Canny(image, lower, upper)
# return the edged image
return edged
imagePath = 'hair_images/903ff49749.jpg'
image = cv2.imread(imagePath)
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
blurred = cv2.GaussianBlur(gray, (3, 3), 0)
# +
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
blurred = cv2.GaussianBlur(gray, (3, 3), 0)
# apply Canny edge detection using a wide threshold, tight
# threshold, and automatically determined threshold
wide = cv2.Canny(blurred, 10, 200)
tight = cv2.Canny(blurred, 225, 250)
auto = auto_canny(blurred)
# +
plt.figure(figsize=(20,20))
plt.subplot(121),plt.imshow(img,cmap = 'gray')
plt.title('Original Image'), plt.xticks([]), plt.yticks([])
plt.subplot(122),plt.imshow(wide,cmap = 'gray')
plt.title('Edge Image'), plt.xticks([]), plt.yticks([])
plt.show()
| notebooks/exploratory_image_masking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
Use movie_data_general_cleaned to build a basic Linear Regression model for general movies.
functions used:
get_Xy(df) from model.py
get_score(X_train,X_val,y_train,y_val) from model.py
"""
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler, PolynomialFeatures, OneHotEncoder
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression, Lasso, LassoCV, Ridge, RidgeCV
from sklearn.metrics import r2_score, mean_squared_error
import patsy
import scipy.stats as stats
import sys
sys.path.append('/Users/katiehuang/Documents/metis/projects/onl_ds5_project_2/py')
import importlib
from model import *
# -
# ### Separate columns into target candidates, continuous features, categorical features
# Load the cleaned complete data
all_df = pd.read_pickle('../data/movie_data_general_cleaned')
con_feature = ['budget','runtime','release_year']
cat_feature = ['MPAA','genre','distributor','language','country','keywords']
target_candidate = ['opening_weekend_usa','gross_usa','gross_world','rating','vote']
# Choose only the continuous variables for now
c_movie_df = all_df[['opening_weekend_usa'] + con_feature].copy()
# ### Baseline Model
# Only consider the continuous predictors and use 'opening_weekend_usa' as target.
# +
# Input: whole df
# Output: features_df(X) and target_Series(y)
X, y = get_Xy(c_movie_df)
# -
# Separate X, y into train/validation/test sets (60/20/20)
X_, X_test, y_, y_test = train_test_split(X, y, test_size=.2, random_state=20)
X_train, X_val, y_train, y_val = train_test_split(X_, y_, test_size=.25, random_state=21)
# %run -i "../py/model.py"
# Input: X_train,X_val,y_train,y_val
# lr_model = LinearRegression()
# Output: Train R^2, Validation R^2, RMSE, coefficcients, actual vs prediction plot
# Return: plot, lr_model,y_pred
fig,lr,y_pred = get_score(X_train,X_val,y_train,y_val)
fig.savefig('../plot/lr_prediction.pdf', bbox_inches='tight', transparent = True)
# ### Residuals
res = y_val - y_pred
plt.scatter(y_pred, res)
plt.title("Residual plot")
plt.xlabel("prediction")
plt.ylabel("residuals");
# diagnose/inspect residual normality using qqplot:
stats.probplot(res, dist="norm", plot=plt)
plt.title("Normal Q-Q plot")
plt.show()
# +
# right skew/ heavily tailed
# -
# ### Cross-validation
# +
# Will not use X_test, y_test
lr = LinearRegression()
X, y = get_Xy(c_movie_df)
X_, X_test, y_, y_test = train_test_split(X, y, test_size=.2, random_state=20)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=.25, random_state=21)
result = cross_val_score(lr, X_train, y_train, # estimator, features, target
cv=5, # number of folds
scoring='r2') # scoring metric
# mean of val_score
print(f"Mean of validation score is: {np.mean(result):.2f} +- {np.std(result):.2f}")
print("All validation scores are", result)
# -
| notebooks/12_movie_general_lr_model_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import cv2
import json
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sn
import pandas as pd
from tensorflow.keras.preprocessing.image import ImageDataGenerator, img_to_array, load_img
from tensorflow.keras.utils import to_categorical
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from imutils import paths
input_shape = (300, 300, 3)
BATCH_SIZE = 32
trainPath = os.path.sep.join(['dataset/dfdc' ,"training"])
valPath = os.path.sep.join(['dataset/dfdc' ,"validation"])
testPath = os.path.sep.join(['dataset/dfdc', "evaluation"])
# determine the total number of image paths in training, validation,
# and testing directories
totalTrain = len(list(paths.list_images(trainPath)))
totalVal = len(list(paths.list_images(valPath)))
totalTest = len(list(paths.list_images(testPath)))
# initialize the training data augmentation object
trainAug = ImageDataGenerator(
rotation_range=3,
zoom_range=0.05,
width_shift_range=0.1,
height_shift_range=0.1,
shear_range=0.05,
horizontal_flip=False,
fill_mode="nearest")
# trainAug = ImageDataGenerator()
# initialize the validation/testing data augmentation object (which
# we'll be adding mean subtraction to)
valAug = ImageDataGenerator()
# define the ImageNet mean subtraction (in RGB order) and set the
# the mean subtraction value for each of the data augmentation
# objects
# mean = np.array([123.68, 116.779, 103.939], dtype="float32")
# trainAug.mean = mean
# valAug.mean = mean
# initialize the training generator
trainGen = trainAug.flow_from_directory(
trainPath,
class_mode="binary",
target_size=input_shape,
color_mode="rgb",
shuffle=True,
batch_size=BATCH_SIZE)
# initialize the validation generator
valGen = valAug.flow_from_directory(
valPath,
class_mode="binary",
target_size=input_shape,
color_mode="rgb",
shuffle=False,
batch_size=BATCH_SIZE)
# initialize the testing generator
testGen = valAug.flow_from_directory(
testPath,
class_mode="binary",
target_size=input_shape,
color_mode="rgb",
shuffle=False,
batch_size=BATCH_SIZE)
labels = (trainGen.class_indices)
print(labels)
# +
# models can be build with Keras or Tensorflow frameworks
# use keras and tfkeras modules respectively
# efficientnet.keras / efficientnet.tfkeras
import os
import sys
import numpy as np
from skimage.io import imread
import matplotlib.pyplot as plt
sys.path.append('..')
from keras.applications.imagenet_utils import decode_predictions
from efficientnet.efficientnet.tfkeras import EfficientNetB6
from efficientnet.efficientnet.tfkeras import center_crop_and_resize, preprocess_input
model = EfficientNetB6(weights='noisy-student', include_top=False) # or weights='noisy-student'
### applying finetuning
for layer in model.layers:
layer.trainable = False
for layer in model.layers[50:]:
layer.trainable = True
model.summary()
model.compile(loss='binary_crossentropy', optimizer=optimizers.Adam(lr=1e-4,
beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.001, amsgrad=False),metrics=['accuracy'])
checkpoint = ModelCheckpoint('./weights.{epoch:02d}-{val_loss:.2f}.model',
save_weights_only=False, monitor='loss', verbose=0, save_best_only=True, mode='min')
callbacks_list = [checkpoint]
H = efficinet_model.fit_generator(
trainGen,
steps_per_epoch=totalTrain // BATCH_SIZE,
validation_data=valGen,
validation_steps=totalVal // BATCH_SIZE,
epochs=20, verbose = 1, callbacks=callbacks_list)
# -
| src/CNNs/EfficientNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kernel SHAP explanation for multinomial logistic regression models
# <div class="alert alert-info">
# Note
#
# To enable SHAP support, you may need to run
#
# ```bash
# pip install alibi[shap]
# ```
#
# </div>
# ## Introduction
# In a previous [example](kernel_shap_wine_intro.ipynb), we showed how the KernelSHAP algorithm can be aplied to explain the output of an arbitrary classification model so long the model outputs probabilities or operates in margin space. We also showcased the powerful visualisations in the `shap` library that can be used for model investigation. In this example we focus on understanding, in a simple setting, how conclusions drawn from the analysis of the KernelShap output relate to conclusions drawn from interpreting the model directly. To make this possible, we fit a logistic regression model on the Wine dataset.
# +
import shap
shap.initjs()
import matplotlib.pyplot as plt
import numpy as np
from alibi.explainers import KernelShap
from scipy.special import logit
from sklearn.datasets import load_wine
from sklearn.metrics import confusion_matrix, plot_confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
# -
# ## Data preparation: load and split Wine dataset
wine = load_wine()
wine.keys()
data = wine.data
target = wine.target
target_names = wine.target_names
feature_names = wine.feature_names
# Split data into testing and training sets and normalize it.
X_train, X_test, y_train, y_test = train_test_split(data,
target,
test_size=0.2,
random_state=0,
)
print("Training records: {}".format(X_train.shape[0]))
print("Testing records: {}".format(X_test.shape[0]))
scaler = StandardScaler().fit(X_train)
X_train_norm = scaler.transform(X_train)
X_test_norm = scaler.transform(X_test)
# ## Fitting a multinomial logistic regression classifier to the Wine dataset
# ### Training
classifier = LogisticRegression(multi_class='multinomial',
random_state=0,
)
classifier.fit(X_train_norm, y_train)
# ### Model assessment
y_pred = classifier.predict(X_test_norm)
cm = confusion_matrix(y_test, y_pred)
title = 'Confusion matrix for the logistic regression classifier'
disp = plot_confusion_matrix(classifier,
X_test_norm,
y_test,
display_labels=target_names,
cmap=plt.cm.Blues,
normalize=None,
)
disp.ax_.set_title(title)
# ## Interpreting the logistic regression model
# <a id='src_2'></a>
# One way to arrive at the multinomial logistic regression model is to consider modelling a categorical response
# variable $y \sim \text{Cat} (y| \beta x)$ where $\beta$ is $K \times D$ matrix of distribution parameters with $K$ being the number of classes and $D$ the feature dimensionality. Because the probability of outcome $k$ being observed given $x$, $p_{k} = p(y=k|x, \mathbf{\beta})$, is bounded by $[0, 1]$, the logistic regression assumes that a linear relationship exists between the *logit* transformation of the output and the input. This can be formalised as follows:
#
# $$
# \log \left( \frac{p_{k}}{1 - p_{k}} \right) = \beta_{0,k} + \beta_{1,k} x_{1} + \beta_{2,k} x_{2} + \cdots + \beta_{D,k} x_{D} = \mathbf{\beta}_k \cdot x
# $$
#
#
# The RHS is a function of the expected value of the categorical distribution (sometimes referred to as a _link function_ in the literature). The coefficients $\beta$ of the linear relations used to fit the logit transformation are estimated jointly given a set of training examples $\mathcal{D}= \{(x_i, y_i)\}_{i=1}^N$.
#
# For each class, the vector of coefficients $\mathbb{\beta}_k$ can be used to interpret the model *globally*; in the absence of interaction terms, the coefficient of a predictor (i.e., independent variable) represents the *change in log odds* when the predictor changes by one unit while all other variables are kept at fixed values. Equivalently, the exponentiated coefficient is equivalent to a change in odds. Since the transformation from odds to outcome probabilities is monotonic, a change in odds also implies a change in the outcome probability in the same direction. Thus, the magnitudes of the feature coefficients measure the effect of a predictor on the output and thus one can globally interpret the logistic regression model.
#
# However, the log odds ratios and odds ratios are known to be sensitive to *unobserved heterogenity*, that is, omission of a variable with good explanatory power from a logistic regression model *assumed true*. While we will not be concerned directly with this issue and refer the interested reader to [[2]](#References), we will be using the *estimated percentage unit effect* (or the *marginal effect*)
#
# $$
# \beta_{j,k} \times p_{i,k}(1 - p_{i, k})
# $$
#
# as a means of estimating the effect of a predictor $j$ on individual $i$ ($x_{i, j})$ with respect to predicting the $k^{th}$ class and thus *locally* interpret the model. The average marginal effect is more robust measure of effects in situations where effects are compared across different groups or models. Consider a logistic model where an independent variable $x_1$ is used to predict an outcome and a logistic model where $x_2$, known to be uncorrelated with $x_1$, is also included. Since the two models assign different probabilities to the different outcomes and since the distribution of the outcome across values of $x_1$ should be the same across the two models (due to the independence assumption), we expected the second model will scale the coeffcient of $\beta_1$. Hence, the log-odds and odds ratios are not robust to unobserved heterogeneity so directly comparing the two across models or groups can be misleading. As discussed in [[2]](#References), the marginal effect is generally robust to the effect.
#
#
# The average marginal effect (AME) of a predictor
#
# $$
# \frac{1}{n} \sum_{i=1}^{n}\beta_{j,k} \times p_{i,k} (1 - p_{i,k})
# $$
#
# is equivalent to simply using $\beta_{j,k}$ to *globally* explain the model.
# +
def issorted(arr, reverse=False):
"""
Checks if a numpy array is sorted.
"""
if reverse:
return np.all(arr[::-1][:-1] <=arr[::-1][1:])
return np.all(arr[:-1] <= arr[1:])
def get_importance(class_idx, beta, feature_names, intercepts=None):
"""
Retrive and sort abs magnitude of coefficients from model.
"""
# sort the absolute value of model coef from largest to smallest
srt_beta_k = np.argsort(np.abs(beta[class_idx, :]))[::-1]
feat_names = [feature_names[idx] for idx in srt_beta_k]
feat_imp = beta[class_idx, srt_beta_k]
# include bias among feat importances
if intercepts is not None:
intercept = intercepts[class_idx]
bias_idx = len(feat_imp) - (np.searchsorted(np.abs(feat_imp)[::-1], np.abs(intercept)))
# bias_idx = np.searchsorted(np.abs(feat_imp)[::-1], np.abs(intercept)) + 1
feat_imp = np.insert(feat_imp, bias_idx, intercept.item(), )
intercept_idx = np.where(feat_imp == intercept)[0][0]
feat_names.insert(intercept_idx, 'bias')
return feat_imp, feat_names
def plot_importance(feat_imp, feat_names, **kwargs):
"""
Create a horizontal barchart of feature effects, sorted by their magnitude.
"""
left_x, right_x = kwargs.get("left_x"), kwargs.get("right_x")
eps_factor = kwargs.get("eps_factor", 4.5)
xlabel = kwargs.get("xlabel", None)
ylabel = kwargs.get("ylabel", None)
labels_fontsize = kwargs.get("labels_fontsize", 15)
tick_labels_fontsize = kwargs.get("tick_labels_fontsize", 15)
# plot
fig, ax = plt.subplots(figsize=(10, 5))
y_pos = np.arange(len(feat_imp))
ax.barh(y_pos, feat_imp)
# set lables
ax.set_yticks(y_pos)
ax.set_yticklabels(feat_names, fontsize=tick_labels_fontsize)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel(xlabel, fontsize=labels_fontsize)
ax.set_ylabel(ylabel, fontsize=labels_fontsize)
ax.set_xlim(left=left_x, right=right_x)
# add text
for i, v in enumerate(feat_imp):
eps = 0.03
if v < 0:
eps = -eps_factor*eps
ax.text(v + eps, i + .25, str(round(v, 3)))
return ax, fig
# -
# We now retrieve the estimated coefficients, and plot them sorted by their maginitude.
beta = classifier.coef_
intercepts = classifier.intercept_
all_coefs = np.concatenate((beta, intercepts[:, None]), axis=1)
class_idx = 0
feat_imp, feat_names = get_importance(class_idx,
beta,
feature_names,
)
_, class_0_fig = plot_importance(feat_imp,
feat_names,
left_x=-1.,
right_x=1.25,
xlabel = f"Feature effects (class {class_idx})",
ylabel = "Features"
)
# Note that these effects are with respect to the model bias (displayed below).
classifier.intercept_
# This plot shows that features such as `proline`, `flavanoids`, `od280/od315_of_diluted_wines`, `alcohol` increase the odds of *any* sample being classified as `class_0` whereas the `alcalinity_of_ash` decreases them.
feat_imp, feat_names = get_importance(1, # class_idx
beta,
feature_names,
)
# The plot below shows that, however, `alcalinity_of_ash` increases the odds of a wine being in `class_1`. Predictors such as `proline`, `alcohol` or `ash`, which increase the odds of predicting a wine as a member of `class_0`, decrease the odds of predicting it as a member of `class_1`.
_, class_1_fig = plot_importance(feat_imp,
feat_names,
left_x=-1.5,
right_x=1,
eps_factor = 5, # controls text distance from end of bar for negative examples
xlabel = "Feature effects (class {})".format(1),
ylabel = "Features"
)
feat_imp, feat_names = get_importance(2, # class_idx
beta,
feature_names,
)
# Finally, for `class_2`, the `color_intensity`, `ash` are the features that increase the `class_2` odds.
_, class_2_fig = plot_importance(feat_imp,
feat_names,
left_x=-1.25,
right_x=1,
xlabel = "Feature effects (class {})".format(2),
ylabel = "Features"
# eps_factor = 5.
)
# ## Apply KernelSHAP to explain the model
# <a id='src_1'></a>
# Note that the *local accuracy* property of SHAP (eq. (5) in [[1]](#References)) requires
#
# $$
# f(x) = g(x') = \phi_0 + \sum_{j=1}^D \phi_j x_j'.
# $$
#
# Hence, sum of the feature importances, $\phi_j$, should be equal to the model output, $f(x)$. By passing `link='logit'` to the explainer, we ensure that $\phi_0$, the *base value* (see _**Local explanation**_ section [here](kernel_shap_wine_intro.ipynb)) will be calculated in the correct units. Note that here $x' \in \mathbb{R}^D$ represents a *simplified input* for which the shap value is computed. A simple example of a simplified input in the image domain, justified by the dimensionality of the input space, is a *superpixel mask*: we formulate the task of explaining the outcome of an image prediction task as determining the effects of each superpixel in a segmenented image upon the outcome. The interested reader is referred to [[1]](#References) for more details about simplified inputs.
pred_fcn = classifier.predict_proba
lr_explainer = KernelShap(pred_fcn, link='logit')
lr_explainer.fit(X_train_norm)
# passing the logit link function to the explainer ensures the units are consistent ...
mean_scores_train = logit(pred_fcn(X_train_norm).mean(axis=0))
print(mean_scores_train - lr_explainer.expected_value)
lr_explanation = lr_explainer.explain(X_test_norm, l1_reg=False)
# Because the dimensionality of the feature space is relatively small, we opted not to regularise the regression that computes the Shapley values. For more information about the regularisation options available for higher dimensional data see the introductory example [here](kernel_shap_wine_intro.ipynb).
# ### Locally explaining multi-output models with KernelShap
# ### Explaining the logitstic regression model globally with KernelSHAP
# #### Summary plots
# To visualise the impact of the features on the decision scores associated with class `class_idx`, we can use a summary plot. In this plot, the features are sorted by the sum of their SHAP values magnitudes across all instances in `X_test_norm`. Therefore, the features with the highest impact on the decision score for class `class_idx` are displayed at the top of the plot.
shap.summary_plot(lr_explanation.shap_values[class_idx], X_test_norm, feature_names)
# Because the logistic regression model uses a linear predictor function, the exact shap values for each class $k$ can be computed exactly according to ([[1]](#References))
# $$
# \phi_{i,j}(f, x_i) = \beta_{j,k}(x_{i,j} - \mathbb{E}_{\mathcal{D}}[x_{j}]).
# $$
#
# Here we introduced an additional index $i$ to emphasize that we compute a shap value for *each predictor* and *each instance* in a set to be explained.This allows us to check the accuracy of the SHAP estimate. Note that we have already applied the normalisation so the expectation is not subtracted below.
exact_shap = beta[:, None, :]*X_test_norm
feat_name = 'alcohol'
feat_idx = feature_names.index(feat_name)
x = np.linspace(-3, 4, 1000)
plt.scatter(exact_shap[class_idx,...][:, feat_idx], lr_explanation.shap_values[class_idx][:, feat_idx])
plt.plot(x, x, linestyle='dashed', color='red')
plt.xlabel(r'Exact $\phi_j$', fontsize=18)
plt.ylabel(r'Estimated $\phi_j$', fontsize=18)
plt.title(fr"Comparison of estimated and exact shap values for feature '{feat_name}'")
plt.grid(True)
# The plot below shows that the exact shap values and the estimate values give rise to similar ranking of the features, and only the order of the `flavanoids` and `alcohol`features is swapped.
shap.summary_plot(exact_shap[class_idx, ...], X_test_norm, feature_names)
# An simlar plot can be create for the logistic regression model by plotting the marginal effects. Note that the plot labelling cannot be changed, so the x axis is incorrectly labeled as `SHAP value` below.
p = classifier.predict_proba(X_test_norm)
prb = p * (1. - p)
marg_effects = all_coefs[:, None, :] * prb.T[..., None]
assert (all_coefs[0, 0] * prb[:, 0] - marg_effects[0, :, 0]).sum() == 0.0
avg_marg_effects = np.mean(marg_effects, axis=1) # nb: ranking of the feature coefs should be preserved
mask = np.ones_like(X_test_norm) # the effect (postive vs negative) on the output depend on the sign of the input
mask[X_test_norm < 0] = -1
shap.summary_plot(marg_effects[class_idx, :, :-1]*mask, X_test_norm, feature_names) # exclude bias
# As expected, the ranking of the marginal effects is the same as that provided the ranking the raw coefficients (see below). However, this effect measure allows us to assess the effects at instance level. Note that both the approximate computation and the exact method yield the same group of features as the most important, although their rankings are not identical. It is important to note that the exact effects ranking and absolute values is a function of the entire data (due to the dependence of the model coefficients) whereas the approximate computation is *local*: the explanation model is fitted locally around each instance. We also notice that the approximate and exact shap value computation both identify the same relationship between the feature value and the effect on the evidence of a sample belonging to `class_idx`.
class_0_fig
# Looking at the 6 most important features for this classification in `class_0`, we see that both the `KernelSHAP` method and the logistic regression rank the `proline` feature as the one with the most significant effect. While the order of the subsequent 5 features is permuted, the effects of these features are also very similar so, in effect, similar conclusions would be drawn from analysing either output.
# ### References
# <a id='References'></a>
#
# [[1]](#src_1) <NAME>. and <NAME>., 2017. A unified approach to interpreting model predictions. In Advances in neural information processing systems (pp. 4765-4774).
#
# [[2]](#src_2) <NAME>., 2017. "Logistic regression: Uncovering unobserved heterogeneity."
| doc/source/examples/kernel_shap_wine_lr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dataset loading and handling utilities in sklearn
from sklearn.datasets import load_iris
iris = load_iris()
iris.keys()
print(iris['data'])
print(iris['feature_names'])
print(iris['target'])
print(iris['target_names'])
# # Classification & Regression
# * ## Classification - output set is discrete - categorizing the input in one of a finite set of labels or classes
# * ## Regression - output set is continuous - the output consists of one or more continuous variables
# # Classification with Sklearn
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
X = iris['data']
y = iris['target']
knn.fit(X, y)
prediction = knn.predict(X)
correct = (prediction == y).sum()
accuracy = correct / len(y) * 100
print("Accuracy: {} %".format(accuracy))
# ## Explanation of Nearest Neighbour Classification
# <ul>
# <item><h3>1. Memorize the dataset</h3></item>
# <item><h3>2. For each test sample:</h3></item>
# <ul>
# <item><h4>1. Find the most similar <i>training</i> sample</h4></item>
# <item><h4>2. Assign the label of the most similar training sample to the test sample.</h4></item>
# <item><h4>3. Generalize for $k$ nearest samples, have them vote on the label.</h4></item>
# </ul>
# <item><h3>3. What is _similarity_?</h3></item>
# <ul>
# <item><h4>1. Depends on _distance_. Generally, $similarity \propto \frac{1}{distance}$</h4></item>
# <item><h4>2. Distances can be of multiple types.</h4></item>
# <item><h4>3. Choosing a distance metrics depends on the distribution of data</h4></item>
# </ul>
# </ul>
# # Regression with sklearn
from sklearn.datasets import load_boston
boston = load_boston()
print(boston.keys())
print(boston['data'])
print(boston['feature_names'])
print(boston['DESCR'])
print(boston['target'])
from sklearn.linear_model import LinearRegression
X = boston['data']
y = boston['target']
lr = LinearRegression()
lr.fit(X, y)
prediction = lr.predict(X)
from sklearn.metrics import r2_score
r2_score(y, prediction)
# ## Exercise: Use DecisionTreeClassifier and LogisticRegression on the MNIST dataset (provided below) and compare their performance.
from sklearn.datasets import load_digits
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
# +
# enter code here
# -
# ## Exercise: Use LinearRegressor and Support Vector regressor on the diabetes dataset (provided below) and compare their performance
from sklearn.svm import SVR
from sklearn.datasets import load_diabetes
# +
# enter code here
| 01_intro_to_ml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 38
# language: python
# name: python38
# ---
# + language="html"
# <!--Script block to left align Markdown Tables-->
# <style>
# table {margin-left: 0 !important;}
# </style>
# -
# Preamble script block to identify host, user, and kernel
import sys
# ! hostname
# ! whoami
print(sys.executable)
print(sys.version)
print(sys.version_info)
# # Lesson 23 - From Regression To Classification
#
# ## Prediction Machine
#
# Imagine a basic machine that takes a question, does some “thinking” and pushes out an answer. Just like the example above with ourselves taking input through our eyes, using our brains to analyse the scene, and coming to the conclusion about what objects are in that scene. Here’s what this looks like:
#
# 
#
# Computers don’t really think, they’re just glorified calculators remember, so let’s use more appropriate words to describe what’s going on:
#
# 
#
# A computer takes some input, does some calculation and poops out an output. The following illustrates this. An input of “3 x 4” is processed, perhaps by turning multiplication into an easier set of additions, and the output answer “12” poops out.
#
# 
#
# Not particularly impressive - we could even write a function!
# +
def threeByfour(a,b):
value = a * b
return(value)
a = 3; b=4
print('a times b =',threeByfour(a,b))
# -
# Next, Imagine a machine that converts kilometres to miles, like the following:
#
# 
#
# But imagine we don’t know the formula for converting between kilometres and miles. All we know is the the relationship between the two is **linear**. That means if we double the number in miles, the same distance in kilometres is also doubled.
#
# This linear relationship between kilometres and miles gives us a clue about that mysterious calculation it needs to be of the form “miles = kilometres x **c**”, where **c** is a constant. We don’t know what this constant **c** is yet. The only other clues we have are some examples pairing kilometres with the correct value for miles. These are like real world observations used to test scientific theories - they’re examples of real world truth.
#
# |Truth Example|Kilometres|Miles|
# |:---|---:|---:|
# |1| 0| 0|
# |2| 100| 62.137|
#
# To work out that missing constant **c** just pluck a value at random and give it a try! Let’s try **c** = 0.5 and see what happens.
#
# 
#
# Here we have miles = kilometres x **c**, where kilometres is 100 and **c** is our current guess at 0.5. That gives 50 miles. Okay. That’s not bad at all given we chose **c** = 0.5 at random! But we know it’s not exactly right because our truth example number 2 tells us the answer should be 62.137. We’re wrong by 12.137. That’s the **error**, the difference between our calculated answer and the actual truth from our list of examples. That is,
#
# error = truth - calculated = 62.137 - 50 = 12.137
#
#
# +
def km2miles(km,c):
value = km*c
return(value)
x=100
c=0.5
y=km2miles(x,c)
t=62.137
print(x, 'kilometers is estimated to be ',y,' miles')
print('Estimation error is ', t-y , 'miles')
# -
# 
#
# So what next? We know we’re wrong, and by how much. Instead of being a reason to despair, we use this error to guide a second, better, guess at **c**. Look at that error again. We were short by 12.137. Because the formula for converting kilometres to miles is linear, miles = kilometres x **c**, we know that increasing **c** will increase the output. Let’s nudge **c** up from 0.5 to 0.6 and see what happens.
#
# 
#
# With **c** now set to 0.6, we get miles = kilometres x **c** = 100 x 0.6 = 60. That’s better than the previous answer of 50. We’re clearly making progress! Now the error is a much smaller 2.137. It might even be an error we’re happy to live with.
# +
def km2miles(km,c):
value = km*c
return(value)
x=100
c=0.6
y=km2miles(x,c)
t=62.137
print(x, 'kilometers is estimated to be ',y,' miles')
print('Estimation error is ', t-y , 'miles')
# -
# The important point here is that we used the error to guide how we nudged the value of c. We wanted to increase the output from 50 so we increased **c** a little bit. Rather than try to use algebra to work out the exact amount **c** needs to change, let’s continue with this approach of refining **c**. If you’re not convinced, and think it’s easy enough to work out the exact answer, remember that many more interesting problems won’t have simple mathematical formulae relating the output and input. That’s why we use more sophisticated "machine learning" methods. Let’s do this again. The output of 60 is still too small. Let’s nudge the value of **c** up again from 0.6 to 0.7.
#
# <NAME>. Make Your Own Neural Network (Page 16). . Kindle Edition.
# +
def km2miles(km,c):
value = km*c
return(value)
x=100
c=0.7
y=km2miles(x,c)
t=62.137
print(x, 'kilometers is estimated to be ',y,' miles')
print('Estimation error is ', t-y , 'miles')
# -
# 
#
# Oh no! We’ve gone too far and **overshot** the known correct answer. Our previous error was 2.137 but now it’s -7.863. The minus sign simply says we overshot rather than undershot, remember the error is (correct value - calculated value). Ok so **c** = 0.6 was way better than c = 0.7. We could be happy with the small error from **c** = 0.6 and end this exercise now. But let’s go on for just a bit longer.
#
# Let's split the difference from our last guess - we still have overshot, but not as much (-2.8629).
# Split again to **c**=0.625, and overshoot is only (-0.3629) (we could sucessively split the **c** values until we are close enough. The method just illustrated is called bisection, and the important point is that we avoided any mathematics other than bigger/smaller and multiplication and subtraction; hence just arithmetic.)
#
# That’s much much better than before. We have an output value of 62.5 which is only wrong by 0.3629 from the correct 62.137. So that last effort taught us that we should moderate how much we nudge the value of **c**. If the outputs are getting close to the correct answer - that is, the error is getting smaller - then don’t nudge the constant so much. That way we avoid overshooting the right value, like we did earlier. Again without getting too distracted by exact ways of working out **c**, and to remain focussed on this idea of successively refining it, we could suggest that the correction is a fraction of the error. That’s intuitively right - a big error means a bigger correction is needed, and a tiny error means we need the teeniest of nudges to **c**. What we’ve just done, believe it or not, is walked through the very core process of learning in a neural network - we’ve trained the machine to get better and better at giving the right answer. It is worth pausing to reflect on that - we’ve not solved a problem exactly in one step. Instead, we’ve taken a very different approach by trying an answer and improving it repeatedly. Some use the term **iterative** and it means repeatedly improving an answer bit by bit.
#
# +
def km2miles(km,c):
value = km*c
return(value)
x=100
c=0.65
y=km2miles(x,c)
t=62.137
print(x, 'kilometers is estimated to be ',y,' miles')
print('Estimation error is ', t-y , 'miles')
# +
def km2miles(km,c):
value = km*c
return(value)
x=100
c=0.625
y=km2miles(x,c)
t=62.137
print(x, 'kilometers is estimated to be ',y,' miles')
print('Estimation error is ', t-y , 'miles')
# -
# ## Classification
#
# We called the above simple machine a **predictor**, because it takes an input and makes a prediction of what the output should be. We refined that prediction by adjusting an internal parameter, informed by the error we saw when comparing with a known-true example.
#
# Now look at the following graph showing the measured widths and lengths of garden bugs.
#
# 
#
# You can clearly see two groups. The caterpillars are thin and long, and the ladybirds are wide and short. Remember the predictor that tried to work out the correct number of miles given kilometres? That predictor had an adjustable linear function at it’s heart. Remember, linear functions give straight lines when you plot their output against input. The adjustable parameter **c** changed the slope of that straight line.
#
# <NAME>. Make Your Own Neural Network (Page 19). . Kindle Edition.
# +
import numpy as np
import pandas as pd
import statistics
import scipy.stats
from matplotlib import pyplot as plt
import statsmodels.formula.api as smf
import sklearn.metrics as metrics
# plot the predictor machine here
kilometers = [0, 100]
miles = [0,62.137]
x = np.array(kilometers)
Y = np.array(miles)
#We already know these parameters from last week but let's assume that we don't!
# alpha = -16.78636363636364
# beta = 11.977272727272727
#Our linear model: ypred = alpha + beta * x
import statsmodels.api as sm #needed for linear regression
from statsmodels.sandbox.regression.predstd import wls_prediction_std #needed to get prediction interval
X = sm.add_constant(x)
re = sm.OLS(Y, X).fit()
#print(re.summary())
#print(re.params)
prstd, iv_l, iv_u = wls_prediction_std(re) #iv_l and iv_u give you the limits of the prediction interval for each point.
#print(iv_l)
#print(iv_u)
from statsmodels.stats.outliers_influence import summary_table
st, data, ss2 = summary_table(re, alpha=0.05)
fittedvalues = data[:, 2]
predict_mean_se = data[:, 3]
predict_mean_ci_low, predict_mean_ci_upp = data[:, 4:6].T
predict_ci_low, predict_ci_upp = data[:, 6:8].T
c = 0.6125
yyyy = km2miles(x,c)
plt.plot(x, Y, 'o')
plt.plot(x, yyyy , '-',color='red', lw=2)
#plt.plot(x, predict_ci_low, '--', color='green',lw=2) #Lower prediction band
#plt.plot(x, predict_ci_upp, '--', color='green',lw=2) #Upper prediction band
#plt.plot(x, predict_mean_ci_low,'--', color='orange', lw=2) #Lower confidence band
#plt.plot(x, predict_mean_ci_upp,'--', color='orange', lw=2) #Upper confidence band
plt.show()
# -
# What happens if we place a straight line over that plot?
#
# 
#
# We can’t use the line in the same way we did before - to convert one number (kilometres) into another (miles), but perhaps we can use the line to separate different kinds of things. In the plot above, if the line was dividing the caterpillars from the ladybirds, then it could be used to **classify** an unknown bug based on its measurements. The line above doesn’t do this yet because half the caterpillars are on the same side of the dividing line as the ladybirds. Let’s try a different line, by adjusting the slope again, and see what happens.
#
# 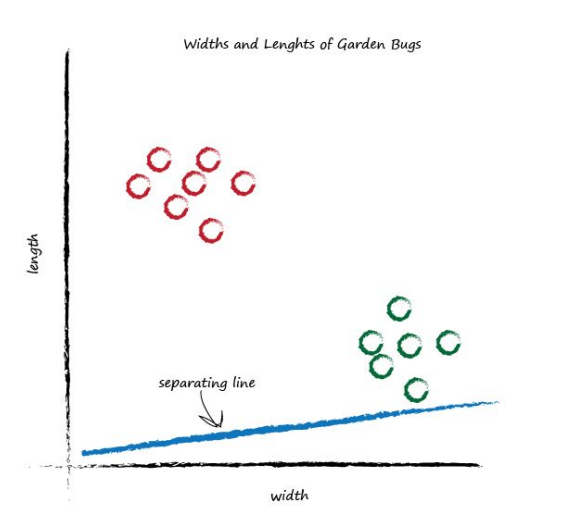
#
# This time the line is even less useful! It doesn’t separate the two kinds of bugs at all. Let’s have another go:
#
# 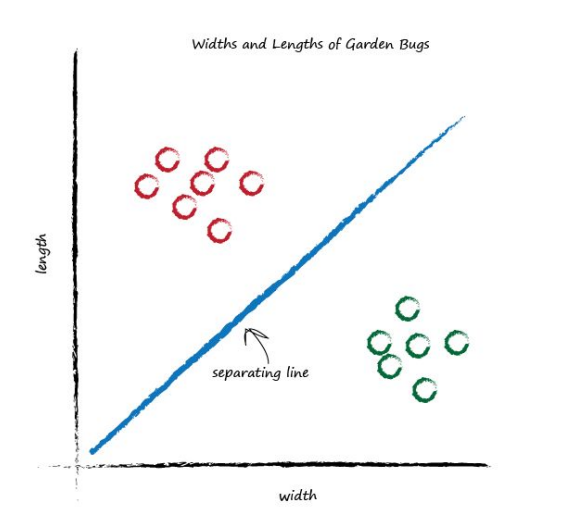
#
# That’s much better! This line neatly separates caterpillars from ladybirds. We can now use this line as a **classifier** of bugs. We are assuming that there are no other kinds of bugs that we haven’t seen - but that’s ok for now, we’re simply trying to illustrate the idea of a simple classifier. Imagine next time our computer used a robot arm to pick up a new bug and measured its width and height, it could then use the above line to classify it correctly as a caterpillar or a ladybird. Look at the following plot, you can see the unknown bug is a caterpillar because it lies above the line. This classification is simple but pretty powerful already!
#
# 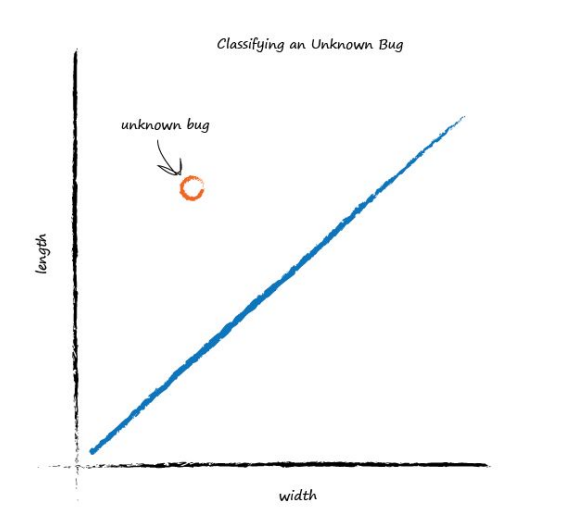
#
# We’ve seen how a linear function inside our simple predictors can be used to classify previously unseen data. But we’ve skipped over a crucial element. How do we get the right slope? How do we improve a line we know isn’t a good divider between the two kinds of bugs? The answer to that is again at the very heart of how machines learn, and we’ll look at this next.
# ### Training A Simple Classifier
#
# We want to **train** our linear classifier to correctly classify bugs as ladybirds or caterpillars. We saw above this is simply about refining the slope of the dividing line that separates the two groups of points on a plot of big width and height.
#
# How do we do this? We need some examples to learn from. The following table shows two examples, just to keep this exercise simple.
#
# |Example| Width| Length| Bug |
# |:---|:---|:---|:---|
# |1 | 3.0 | 1.0 | ladybird |
# |2 | 1.0 | 3.0 | caterpillar |
#
# We have an example of a bug which has width 3.0 and length 1.0, which we know is a ladybird. We also have an example of a bug which is longer at 3.0 and thinner at 1.0, which is a caterpillar. This is a set of examples which we declare to be the truth.
#
# It is these examples which will help refine the slope of the classifier function. Examples of truth used to teach a predictor or a classifier are called the **training data.**
# Let’s plot these two training data examples. Visualising data is often very helpful to get a better understand of it, a feel for it, which isn’t easy to get just by looking at a list or table of numbers.
#
# 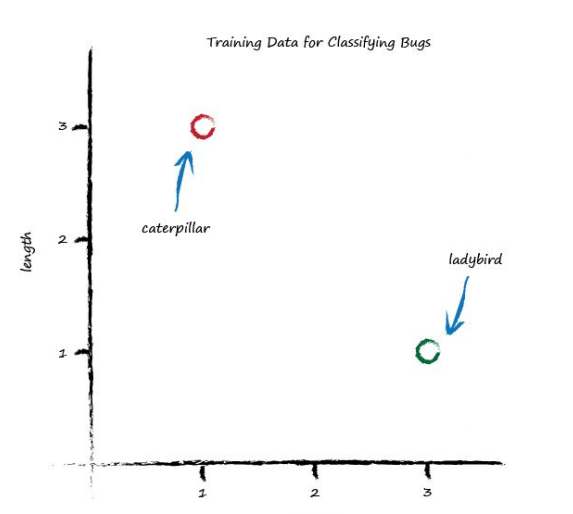
#
# Let’s start with a random dividing line, just to get started somewhere. Looking back at our miles to kilometre predictor, we had a linear function whose parameter we adjusted. We can do the same here, because the dividing line is a straight line: $y = Ax+b$
#
# We’ve deliberately used the names $y$ and $x$ instead of length and width, because strictly speaking, the line is not a predictor here. It doesn’t convert width to length, like we previously converted miles to kilometres. Instead, it is a dividing line, a classifier. To keep the garden bug scenario as simple as possible we will choose a zero intercept $b=0$.
#
# We saw before that the parameter $A$ controls the slope of the line. The larger $A$ is the larger the slope. Let’s go for $A$ is 0.25 to get started. The dividing line is $y = 0.25x$. Let’s plot this line on the same plot of training data to see what it looks like:
#
# 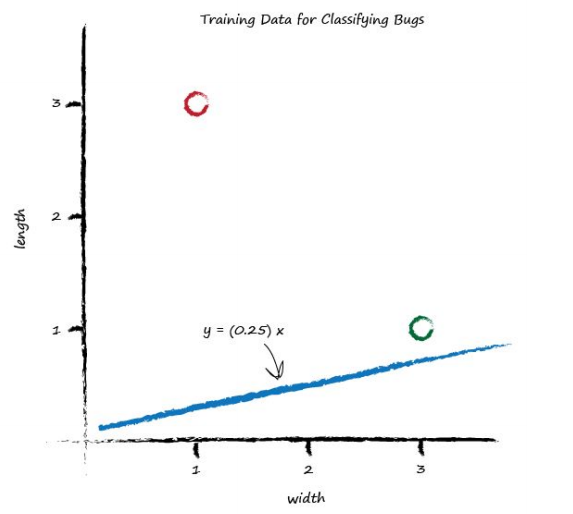
#
# Well, we can see that the line $y = 0.25x$ isn’t a good classifier already without the need to do any calculations. The line doesn’t divide the two types of bug - We can’t say “if the bug is above the line then it is a caterpillar” because the ladybird is above the line too.
#
# So intuitively we need to move the line up a bit. We’ll resist the temptation to do this by looking at the plot and drawing a suitable line. We want to see if we can find a repeatable recipe to do this, a series of computer instructions, which computer scientists call an **algorithm**.
#
# Let’s look at the first training example: the width is 3.0 and length is 1.0 for a ladybird.
# If we tested the $y = Ax$ function with this example where $x$ is 3.0, we’d get $y = (0.25) * (3.0) = 0.75$
# The function, with the parameter $A$ set to the initial arbitrary chosen value of 0.25, is suggesting that for a bug of width 3.0, the length should be 0.75.
# We know that’s too small because the training data example tells us it must be a length of 1.0. So we have a difference, an **error**.
# Just as before, with the miles to kilometres predictor, we can use this error to inform how we adjust the parameter $A$. But let’s think about what $y$ should be again.
# If $y$ was 1.0 then the line goes right through the point where the ladybird sits at $(x,y) = (3.0, 1.0)$.
# It’s a subtle point but we don’t actually want that.
# We want the line to go above that point.
# Why? Because we want all the ladybird points to be below the line, not on it.
# The line needs to be a *dividing line* between ladybirds and caterpillars, not a predictor of a bug’s length given its width.
# So let’s try to aim for $y = 1.1$ when $x = 3.0$.
# It’s just a small number above 1.0, We could have chosen 1.2, or even 1.3, but we don’t want a larger number like 10 or 100 because that would make it more likely that the line goes above both ladybirds and caterpillars, resulting in a separator that wasn’t useful at all. So the desired target is 1.1, and the error **E** is
#
# error = (desired target actual output)
#
# Which is, $E = 1.1 \times 0.75 = 0.35$
#
# Let’s examine the error, the desired target and the calculated value visually.
#
# 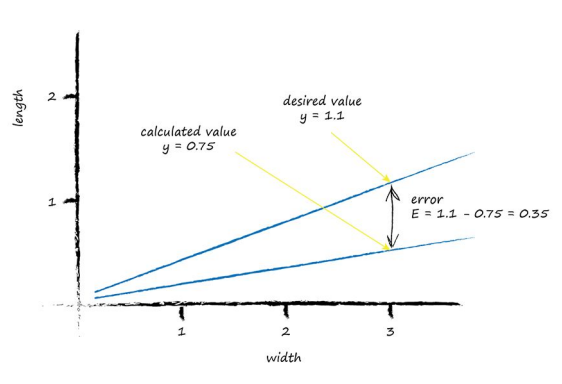
#
# Now, what do we do with this **E** to guide us to a better refined parameter $A$?
#
# We want to use the error in $y$, which we call **E**, to inform the required change in parameter $A$.
# To do this we need to know how the two are related. How is $A$ related to **E**?
#
# If we know this, then we can understand how changing one affects the other (correlation anyone?).
#
# Let’s start with the linear function for the classifier: $y = Ax$
# We know that for initial guesses of $A$ this gives the wrong answer for $y$, which should be the value given by the training data.
# Let’s call the correct desired value, $t$ for target value. To get that value $t$, we need to adjust $A$ by a small amount; $ t = (A + \Delta A)x$ Let’s picture this to make it easier to understand. You can see the new slope $(A + \Delta A)$.
#
# 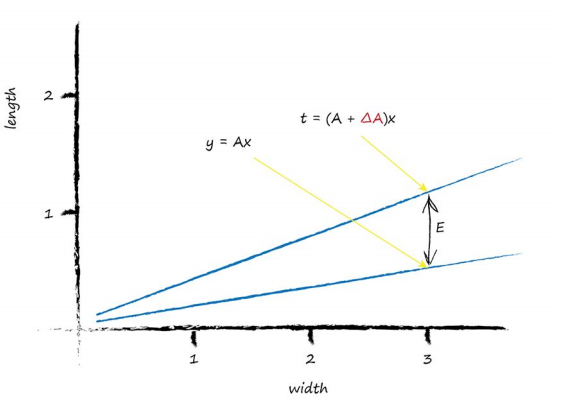
#
# Remember the error **E** was the difference between the desired correct value and the one we calculate based on our current guess for $A$. That is, **E** was $t - y$ (Kind of smells like a residual!);
#
# $$ t - y = (A + \Delta A)x - Ax$$
#
# Expanding out the terms and simplifying:
#
# $$ \textbf{E} = t - y = Ax + (\Delta A)x Ax E = (\Delta A)x $$
#
# That’s remarkable! The error **E** is related to $\Delta A$ in a very simple way.
#
# We wanted to know how much to adjust $A$ by to improve the slope of the line so it is a better classifier, being informed by the error **E**.
# To do this we simply re-arrange that last equation: $\Delta A = \textbf{E}/ x$
# That’s the magic expression we’ve been looking for. We can use the error **E** to refine the slope $A$ of the classifying line by an amount $\Delta A$.
#
# Let’s update that initial slope. The error was 0.35 and the $x$ was 3.0.
# That gives $\Delta A = \textbf{E}/ x$ as 0.35/ 3.0 = 0.1167.
# That means we need to change the current $A = 0.25$ by $0.1167$.
# That means the new improved value for $A$ is (A + ΔA) which is 0.25 + 0.1167 = 0.3667. As it happens, the calculated value of $y$ with this new $A$ is 1.1 as you’d expect - it’s the desired target value.
#
# Now we have a method for refining that parameter $A$, informed by the current error. Now we’re done with one training example, let’s learn from the next one. Here we have a known true pairing of $x$ = 1.0 and $y$ = 3.0. Let’s see what happens when we put $x$ = 1.0 into the linear function which is now using the updated $A$ = 0.3667. We get $y$ = 0.3667 * 1.0 = 0.3667.
# That’s not very close to the training example with $y$ = 3.0 at all.
#
# Using the same reasoning as before that we want the line to not cross the training data but instead be just above or below it, we can set the desired target value at 2.9. This way the training example of a caterpillar is just above the line, not on it. The error E is (2.9 0.3667) = 2.5333. That’s a bigger error than before, but if you think about it, all we’ve had so far for the linear function to learn from is a single training example, which clearly biases the line towards that single example.
#
# Let’s update the $A$ again, just like we did before. The $\Delta A$ is $\textbf{E}/x$ which is 2.5333/ 1.0 = 2.5333. That means the even newer $A$ is 0.3667 + 2.5333 = 2.9. That means for $x = 1.0$ the function gives 2.9 as the answer, which is what the desired value was.
#
# 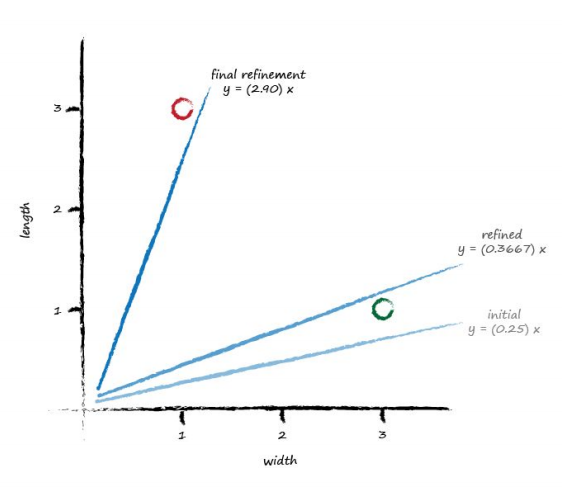
# The plot shows the initial line, the line updated after learning from the first training example, and the final line after learning from the second training example.
#
# Looking at that plot, we don’t seem to have improved the slope in the way we had hoped. It hasn’t divided neatly the region between ladybirds and caterpillars.
# The line updates to give each desired value for y.
# If we keep doing this, updating for each training data example, all we get is that the final update simply matches the last training example closely. We might as well have not bothered with all previous training examples. In effect we are throwing away any learning that previous training examples might gives us and just learning from the last one. How do we fix this?
#
# Easy! And this is an important idea in machine learning.**We moderate** the updates. That is, we calm them down a bit. Instead of jumping enthusiastically to each new $A$, we take a fraction of the change $\Delta A$, not all of it. This way we move in the direction that the training example suggests, but do so slightly cautiously, keeping some of the previous value which was arrived at through potentially many previous training iterations. We saw this idea of moderating our refinements before - with the simpler miles to kilometres predictor, where we nudged the parameter **c** as a fraction of the actual error.
#
# This moderation, has another very powerful and useful side effect. When the training data itself can’t be trusted to be perfectly true, and contains errors or noise, both of which are normal in real world measurements, the moderation can dampen the impact of those errors or noise. It smooths them out. Ok let’s rerun that again, but this time we’ll add a moderation into the update formula: $\Delta A = L (E/ x)$
#
# The moderating factor is often called a **learning rate**, and we’ve called it $L$. Let’s pick $L$ = 0.5 as a reasonable fraction just to get started. It simply means we only update half as much as would have done without moderation.
#
# Running through that all again, we have an initial $A$ = 0.25. The first training example gives us y = 0.25 * 3.0 = 0.75. A desired value of 1.1 gives us an error of 0.35. The $\Delta A = L (E/ x)$ = 0.5 * 0.35/ 3.0 = 0.0583. The updated $A$ is 0.25 + 0.0583 = 0.3083.
#
# Trying out this new A on the training example at $x$ = 3.0 gives y = 0.3083 * 3.0 = 0.9250. The line now falls on the wrong side of the training example because it is below 1.1 but it’s not a bad result if you consider it a first refinement step of many to come. It did move in the right direction away from the initial line.
#
# Let’s press on to the second training data example at $x$ = 1.0. Using $A$ = 0.3083 we have y = 0.3083 * 1.0 = 0.3083. The desired value was 2.9 so the error is (2.9 * 0.3083) = 2.5917. The $\Delta A = L (E/ x)$ = 0.5 * 2.5917/ 1.0 = 1.2958. The even newer $A$ is now 0.3083 + 1.2958 = 1.6042. Let’s visualise again the initial, improved and final line to see if moderating updates leads to a better dividing line between ladybird and caterpillar regions.
#
# 
#
# This is really good! Even with these two simple training examples, and a relatively simple update method using a moderating **learning rate**, we have very rapidly arrived at a good dividing line $y = Ax$ where $A$ is 1.6042. Let’s not diminish what we’ve achieved. We’ve achieved an automated method of learning to classify from examples that is remarkably effective given the simplicity of the approach.
# ### Multiple Classifiers (future revisions)
#
# ### Neuron Analog (future revisions)
# - threshold
# - step-function
# - logistic function
# - computational linear algebra
#
# ## Classifiers in Python (future revisions)
# - KNN Nearest Neighbor (use concrete database as example, solids as homework)
# - ANN Artifical Neural Network (use minst database as example, something from tensorflow as homework)
# - Clustering(K means, heriarchial (random forests))
# - SVM
# - PCA (? how is this machine learning we did this in the 1970s?)
#
# ## References
#
# <NAME>. Make Your Own Neural Network. . Kindle Edition.
| 1-Lessons/Lesson22/.ipynb_checkpoints/Lesson23-DEV-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <div class="alert block alert-info alert">
#
# # <center> Scientific Programming in Python
# ## <center><NAME><br>Bonn-Rhein-Sieg University of Applied Sciences<br>Sankt Augustin, Germany
#
# # <center> Significant Figures
#
# <hr style="border:2px solid gray"></hr>
# <p><img alt="Accuracy vs Precision" width="800" src="00_images/alexander-muzenhardt-753371-unsplash.jpg" align="center" hspace="10px" vspace="0px"></p>
#
# ---
# ## Accuracy versus Precision
# #### What is the difference between the two?
#
# (A visual attempt to place the idea of significant figures into context)
# <font color='red'>Red</font> = target value
#
# <font color='dodgerblue'>Blue</font> and <font color='orange'>Orange</font> = two different data sets, measure by two different people
#
# All of the points look close together - the data all appears to be equally good.
#
# <p><img alt="Accuracy vs Precision" width="400" src="00_images/accuracy_vs_precision_1.png" align="left" hspace="10px" vspace="0px"></p>
# <br><br><br>
# <br><br><br>
# <br><br><br>
# Create a more appropriate scale - things look a bit different now.
#
# <p><img alt="Accuracy vs Precision" width="400" src="00_images/accuracy_vs_precision_2.png" align="left" hspace="10px" vspace="0px"></p>
#
# <br>
# <br>
# <br>
# <br>
#
# The most <font color='dodgerblue'>**accurate**</font> value is **one** (i.e. a singluar data point) of the <font color='dodgerblue'>**blue**</font> dots, while the higher <font color='orange'>**precision**</font> values are represented by the <font color='orange'>**orange**</font> dots.
# <br><br><br>
# <br><br><br>
# <br><br><br>
# However, note that even if you look at the highly precise values (i.e. orange dots), you notice a distribution in the data.
#
# <p><img alt="Accuracy vs Precision" width="400" src="00_images/accuracy_vs_precision_3.png" align="left" hspace="10px" vspace="0px"></p>
# **Important**: Notice how **using additional digits** to report the tick lables is **provides** us with **more knowledge** about the data that we obtained.
#
# ### Take Home Message: It seems that the <font color='dodgerblue'>number of digits</font> for a data value is <font color='dodgerblue'>related</font> to its <font color='dodgerblue'>information content</font>.
# <br><br><br>
# <br><br><br>
# <br><br><br>
# <hr style="border:2px solid gray"></hr>
#
# ## Question
#
# Three researchers are measuring the size of a device that will be fitted onto a satellite. This is done independently from one another.
#
# Researcher 1: 10.39 cm
#
# Researcher 2: 10.3 cm
#
# Researcher 3: 10.37 cm
#
# <br><br>
# To give their boss their best measurement value, they **average their values**.
#
# **What is the average value that they should report?**
# <!-- <br><br>
# What about the a different data set?
#
# Researcher 1: 10.36
#
# Researcher 2: 10.3
#
# Researcher 3: 10.35
#
# What value is reported to the boss?
#
# <br><br><br>
# <br><br><br>
# <br><br><br> -->
# **Possible** ways to answer this:
## First data set
(10.39 + 10.3 + 10.37)/3
# If we were to write/print this out, we could then give the following:
## round the number to two decimal places, which we arbitrarily decided:
round((10.39 + 10.3 + 10.37)/3, 2)
# Or if we do it more properly:
#
# $\frac{10.39 + 10.3 + 10.37}{3} = 10.35\bar{3}$
# <br><br><br>
# <br><br><br>
# <br><br><br>
# <!-- # ## Second data set
# # (10.36 + 10.3 + 10.35)/3 -->
# <!-- $\frac{10.36 + 10.3 + 10.35}{3} = 10.33\bar{6}$
# <br><br><br>
# <br><br><br>
# <br><br><br> -->
# or you do some **pre-rounding** of each individual data point **before** you do the mathematics:
#
# $\frac{10.4 + 10.3 + 10.4}{3} = 10.3\bar{6}$
#
# All of this seems correct, right?
# <br><br><br>
# <br><br><br>
# <br><br><br>
# However, the **correct answers** is:
#
# Data set 1: <font color='dodgerblue'>**10.4**</font>
#
# <!-- Data set 2 (i.e.10.36, 10.3, 10.35): 10.3 -->
#
# **Why report the number as XX.Y?**
# <br><br><br>
# <br><br><br>
# <br><br><br>
# ## Significant figures / significant digits / sigfigs
#
# **What are sigfigs?**
#
# Significant figures corresponds to how **precise** a measurement is.
#
# ## Firm Rules for Sigfigs
#
# 1. <font color='dodgerblue'>All non-zero digits are significant</font>: **1, 2, 3, 4, 5, 6, 7, 8, 9**
# <br>
# <br>
# 2. <font color='dodgerblue'>Zeros between non-zero digits are significant</font>: 1**0**2.0, 2**00**5.0, 5**000**9.0
#
# (However the total number of sigfigs in the entire number <font color='red'>101</font> -- i.e. without the decimal point -- is actually **unclear**. We will come to this soon.)
# <br>
# <br>
# 3. <font color='dodgerblue'>Leading zeros are <font color='red'>never</font> significant</font>: <font color='red'>00</font>1, <font color='red'>0.0</font>2, <font color='red'>00</font>1.887, <font color='red'>0.000</font>515
#
# They are **placeholders**, putting the decimal point in the right place.
#
# (In **scientific notation**, they disappear $\rightarrow$ 0.000515 becomes 5.15 x 10$^{-4}$)
# <br>
# <br>
# 4. <font color='dodgerblue'>For number within the decimals - trailing (i.e. after a significant number) zeros are significant</font>: 0.02**0**
#
#
# (Note: alternative scientific notation: 5.15E-4 and 5.15e-4 - https://en.wikipedia.org/wiki/Scientific_notation)
# ## Issues surrounding zeros
#
# Most **number with zero** are complicated.
#
# Ask: are they or are they not place holders?
#
# 1. Zeros after a decimal (e.g. 120.00) - always significant if there is a significant number before the decimal (i.e. 0.01 has one sigfig since the leading zero is not significant).
# <br>
# <br>
# 2. **With a decimal** (e.g. **120.**) - then the zero is significant (i.e. 120. has 3 sigfigs.) This states that all of the values to the left of the decimal are exact (i.e. no rounding has occurred).
#
#
#
# 3. **Without a decimal** the numbers are <font color='red'>*inexact*</font> (e.g. **120**) - the zero is formally not significant, which results in 2 sigfigs being indicated. This leads to uncertainty, because a person questions if there should or should not be a decimal place.
#
# **Example**: a measured weight of 100 g. How precise is this number? Where did the number come from? Have two numbers been averaged to give this number? This leads to assumptions having to be made.
#
# **However**, it is often **assumed** to be at the one's place, but it could be 100.0 or 99.8 g.
#
# Possibilities:
#
# $\frac{120 + 80}{2} = 100$ (imprecise, consider the following examples)
#
# $\frac{119.7 + 80.0}{2} \approx 100$ (rounding the final value of 99.85 up to 100)
#
# $\frac{120. + 80.}{2} = 100.$ (i.e. precise to the one's place)
#
# $\frac{120.0 + 80.0}{2} = 100.0$ (i.e. precise to the tenth's place)
# **Examples**
#
# 1. 1.23 x 10$^2$ and 110. $\leftarrow$ both have 3 sigfigs
# <br><br>
# 2. 123.00 and 0.0012300 $\leftarrow$ both have 5 sigfigs
# <br><br>
# 3. 100 versus 100. or 1.00 x 10$^2$ $\leftarrow$ unclear, but safe to *assume* at least 1 versus 3 sigfigs
# <br><br>
# 4. 120 versus 1.2 x 10$^2$ $\leftarrow$ unclear (safe to *assume* at least 2) versus 2 (clear) sigfigs
# <hr style="border:2px solid gray"></hr>
#
# ## Sigfigs and Math
#
# **Addition and Subtraction** - requires that the answer's number has places <font color='dodgerblue'>after the **decimal** that is equal to the least number of **decimals** present in the sum</font>
#
# Examples:
# <ol type = "1">
# <li> 7.1 + 2.3 = 9.4
# <li> 5,500.000 + 111.111 = 5,611.111
# <li> 1.6 + 12.4 = 14.0
# <li> 10.4 + 10. = 2.0 x 10$^1$ (Convert 10. to 1.0 x 10$^1$)
# <li> 0.003 + 0.02 = 0.02 (There is less precision in 0.02 in comparison to 0.020. Thus the 0.003 is lost within the impreciseness of 0.02.)
# <li> 0.003 + 0.020 = 0.023
# <li> 2,333.3333 + 22.22 = 2,355.55
# </ol>
#
# **Multiplication and Division** - requires that the answer's number has significantn figure places <font color='dodgerblue'>equal to the number of the least amount of significant figures used **in the calculation**</font>
#
# Examples:
# <ol type = "1">
# <li> 1. 8.21 x 2.32 = 19.0 (Unrounded value = 19.0472)
# <li> 8.210 x 2.32 = 19.0
# <li> 8.210 x 2.320 = 19.05
# <li> 0.21 x 10.0 = 0.021
# <li> 1230 x 2.000 = 2.46 x 10$^3$ (Uncertain value of 1230, but it is "fairly" safe to assume it has at least three sig figs)
# <li> 1230\. x 2.000 = 2.460 x 10$^3$
# <li> 1.00 x 20.07 = 20.1
# <li> 8.2 x 12.3 = 1.0 x 10$^2$ (Unrounded value = 100.86)
# </ol>
# <hr style="border:2px solid gray"></hr>
#
# ## Numbers with Infinite Sigfigs
#
# **Exact Number**: have an infinite number of significant figures - must have some context given
# <ol type = "1">
# <li> 100 cm in 1 m - both numbers are exact
# <li> 2 molecules (e.g. a quantity without factions)
# <li> 2 times c (i.e. the speed of light)
# </ol>
#
# <ol type = "1">
# **Mathematical Constants**: are infinite or approximate provided:
# <li> c (speed of light in a vacuum): 299 792 458 m s$^{-1}$ (exact)
# <li> $\pi$ (Pi): ca. 3.14159 26535 89793 23846
# <li> $e$ (Euler's constant): ca. 2.71828 18284 59045 23536
# <li> $\sqrt{2}$ (Pythagoras's constant): ca. 1.41421 35623 73095 04880
# </ol>
#
# https://physics.nist.gov/cuu/Constants
# **Additional Resources**
#
# https://www.khanacademy.org/math/arithmetic-home/arith-review-decimals/arithmetic-significant-figures-tutorial/v/significant-figures
| significant_figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DeepF-g
# language: python
# name: deepf-g
# ---
# ---
#
# # Resample data to achieve more consistent training series
# - assuming that data has been organized into folders as ./base/xxxxx_class/imgxxx.jpg
# ---
import os
import sys
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.utils import resample
import matplotlib.pyplot as plt
import shutil as sh
def mean(vec):
return sum(vec)/len(vec) if len(vec) != 0 else None
"""def resample(X, y, sample_type=None, sample_size=None, class_weights=None, seed=None):
# Nothing to do if sample_type is 'abs' or not set. sample_size should then be int
# If sample type is 'min' or 'max' then sample_size should be float
if sample_type == 'min':
sample_size_ = np.round(sample_size * y.value_counts().min()).astype(int)
elif sample_type == 'max':
sample_size_ = np.round(sample_size * y.value_counts().max()).astype(int)
else:
sample_size_ = max(int(sample_size), 1)
if seed is not None:
np.random.seed(seed)
if class_weights is None:
class_weights = dict()
X_resampled = pd.DataFrame()
for yi in y.unique():
size = np.round(sample_size_ * class_weights.get(yi, 1.)).astype(int)
X_yi = X[y == yi]
sample_index = np.random.choice(X_yi.index, size=size)
X_resampled = X_resampled.append(X_yi.reindex(sample_index))
return X_resampled
"""
# ---
sourceDir = '../dataCropSQ/Img/'
trainTargetDir = '../fdata_CropSQRS/train'
testTargetDir = '../fdata_CropSQRS/test'
#sourceDir = '/Volumes/SanDiskSSD/BFH_backup/CAS_PML/Project/DeepFashion/dataCropSQ/Img/'
#trainTargetDir = '/Volumes/SanDiskSSD/BFH_backup/CAS_PML/Project/DeepFashion/fdata_CropSQRS/train'
#testTargetDir = '/Volumes/SanDiskSSD/BFH_backup/CAS_PML/Project/DeepFashion/fdata_CropSQRS/test'
with open('../data/Anno/list_category_cloth.txt','r') as fp:
lines = [ l.rstrip().replace('1','upper') \
.replace('2','lower') \
.replace('3','full').split() for l in fp][2:]
mainCatagories = { l[0]:l[1] for l in lines}
# +
classDict = {}
for d,_,files in os.walk(sourceDir,topdown=False):
cl_name = d.split('_')[-1]
if mainCatagories.get(cl_name) is None: continue
if classDict.get(cl_name) is None: classDict[cl_name] = []
classDict[cl_name].extend([ d + '/' + f for f in files \
if not f.startswith('.') or not f.endswith('.jpg') ])
# clean out some old classes which were ignored
bunkKeys = []
for cl_name in mainCatagories.keys():
if classDict.get(cl_name) is None: bunkKeys.append(cl_name)
# can't do it in one step, because you're modifing the object you're working on
for b in bunkKeys:
mainCatagories.pop(b,None)
# -
classCount = { k:len(v) for k,v in classDict.items()}
print('Total image files',sum([v for v in classCount.values()]))
print('UpperClass:',sum([classCount[cl] for cl,ca in mainCatagories.items() if ca == 'upper' ]))
print('LowerClass:',sum([classCount[cl] for cl,ca in mainCatagories.items() if ca == 'lower' ]))
print('FullClass:',sum([classCount[cl] for cl,ca in mainCatagories.items() if ca == 'full' ]))
# +
names = sorted(classCount.items(),key=lambda kv: kv[1])
namesSorted = [ v[0] for v in names]
valuesSorted = [ v[1] for v in names]
labelsSorted = [ mainCatagories[n] for n in namesSorted]
colorSorted = []
for n in namesSorted:
color = 'red' if mainCatagories[n] == 'upper' \
else 'blue' if mainCatagories[n] == 'lower' \
else 'green'
colorSorted.append(color)
plt.figure(figsize=(15,5))
plt.bar(range(len(valuesSorted)),valuesSorted,color=colorSorted)
plt.xticks(range(len(valuesSorted)),namesSorted,rotation='vertical')
plt.title('Class distribution of dataset',fontsize=16)
plt.show()
# +
def plotClassCount(clCountDict,title='',color='blue'):
names = sorted(clCountDict.items(),key=lambda kv: kv[1])
namesSorted = [ v[0] for v in names]
valuesSorted = [ v[1] for v in names]
plt.bar(range(len(valuesSorted)),valuesSorted,color=color)
plt.xticks(range(len(valuesSorted)),namesSorted,rotation='vertical')
plt.title(title,fontsize=16)
plt.show()
def summarizeClassCount(clCountDict):
print(' min=',min(clCountDict.values()))
print(' avg=',mean(clCountDict.values()))
print(' max=',max(clCountDict.values()))
# -
catagories = ['upper','lower','full']
ctClDict = { ct:{ k:v for k,v in classDict.items() if mainCatagories[k] == ct} for ct in catagories }
ctClCount = { ct:{ k:len(v) for k,v in ctClDict[ct].items()} for ct in catagories }
print('Summary: Upper Class')
summarizeClassCount(ctClCount['upper'])
plotClassCount(ctClCount['upper'],title='Upper Class distribution',color='red')
print('Summary: Lower Class')
summarizeClassCount(ctClCount['lower'])
plotClassCount(ctClCount['lower'],title='Lower class distribution',color='blue')
ctClCount['lower']['Sweatpants']
print('Summary: Full Class')
summarizeClassCount(ctClCount['full'])
plotClassCount(ctClCount['full'],title='Full class distribution',color='green')
print('Robe=',ctClCount['full']['Robe'])
print('Coat=',ctClCount['full']['Coat'])
threshhold = 2000
selectedClasses = {ct:[k for k,v in ctClCount[ct].items() if v >= threshhold] for ct in catagories}
selectedClasses
testSplit = 0.15
trainClDict = {}
testClDict = {}
for ct,cls in selectedClasses.items():
trainClDict[ct] = {}
testClDict[ct] = {}
for cl in cls:
trainClDict[ct][cl], testClDict[ct][cl] = train_test_split(ctClDict[ct][cl],test_size = testSplit)
print('train[Hoodie]=',len(trainClDict['upper']['Hoodie']))
print(' test[Hoodie]=',len(testClDict['upper']['Hoodie']))
print(' percentage Check=',len(testClDict['upper']['Hoodie'])/len(trainClDict['upper']['Hoodie']))
# +
trainClDictRS = {}
testClDictRS = {}
nsamples = 5000
nsamples_t = int(nsamples*testSplit)
for ct in trainClDict.keys():
trainClDictRS[ct] = {}
testClDictRS[ct] = {}
for cl in trainClDict[ct].keys():
useReplace_train = not len(trainClDict[ct][cl]) > nsamples
useReplace_test = not len(testClDict[ct][cl]) > nsamples_t
trainClDictRS[ct][cl] = resample(trainClDict[ct][cl],n_samples=nsamples,replace=useReplace_train)
testClDictRS[ct][cl] = resample(testClDict[ct][cl],n_samples=nsamples_t,replace=useReplace_test)
# -
len(trainClDictRS['upper']['Hoodie'])
len(testClDictRS['upper']['Hoodie'])
plt.bar(range(len(list(trainClDictRS['upper'].keys()))),[len(v) for k,v in trainClDictRS['upper'].items()])
plt.xticks(range(len(list(trainClDictRS['upper'].keys()))), trainClDictRS['upper'].keys(),rotation='vertical')
plt.show()
# ---
# # Copy files to new training directory
# - resampling of redundant files will be handled by the augmenter
# ---
def copyAndRenameFile(sourceFile,classTargetDir):
# if file exists, renames the file to unique name
labelDir,imgFileOrig = sourceFile.split('/')[-2:]
fullTargetDir = os.path.join(classTargetDir,labelDir)
if not os.path.exists(fullTargetDir): os.mkdir(fullTargetDir)
ind = 0
imgFile = imgFileOrig
#print(imgFile)
while os.path.exists(os.path.join(fullTargetDir,imgFile)):
name,ext = imgFileOrig.split('.')
imgFile = name + f"R{ind:03d}." + ext
ind += 1
sh.copy(sourceFile,fullTargetDir + '/' + imgFile)
print('sourceDir=',sourceDir)
assert os.path.exists(sourceDir),'Failed to find sourceDir'
print('trainTargetDir=',trainTargetDir)
assert os.path.exists(trainTargetDir),'Failed to find trainTargetDir'
print('testTargetDir=',testTargetDir)
assert os.path.exists(testTargetDir),'Failed to find testTargetDir'
# +
for ct,cls in trainClDictRS.items():
ctDir = os.path.join(trainTargetDir,ct)
if not os.path.exists(ctDir): os.mkdir(ctDir)
for cl,files in cls.items():
clDir = os.path.join(ctDir,cl)
if not os.path.exists(clDir): os.mkdir(clDir)
for f in files:
copyAndRenameFile(f,clDir)
for ct,cls in testClDictRS.items():
ctDir = os.path.join(testTargetDir,ct)
if not os.path.exists(ctDir): os.mkdir(ctDir)
for cl,files in cls.items():
clDir = os.path.join(ctDir,cl)
if not os.path.exists(clDir): os.mkdir(clDir)
for f in files:
copyAndRenameFile(f,clDir)
# -
| DeepFashionModel/classDetect/ResampleDataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aharrisonau/gym-ReverseTruck/blob/main/ReverseTruck.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VzVejHlnd_sM"
# ReverseTruck uses the ReverseTruck_env to simulate the reversing of a truck and trailer combination into a loading bay
#
# The environment and the simulation have been developed by <NAME> as a submission for SIT796 - Reinforcement Learning at Deakin University
# + [markdown] id="Y4cnu7OKe846"
# First step is to install the environment for Github
# + id="1KAfz1h3e75r"
# helper code from https://stackoverflow.com/questions/55835318/how-to-create-and-use-a-custom-openai-gym-environment-on-google-colab
# %%capture
# !rm -r ReverseTruck_env
# !git clone https://github.com/aharrisonau/gym-ReverseTruck.git
# !pip install -e gym-ReverseTruck
# + id="Lz4Uh0VSIgoV"
import gym_ReverseTruck
# + id="6bM2MG1UelJ0"
# preliminaries
import pandas as pd
import numpy as np
import math as math
import gym
from gym import error, spaces, utils
from gym.utils import seeding
# + [markdown] id="ItJ4eMfefI6R"
# Then we can create the environment
#
# Note that the environment requires a truck starting position, a truck setup file and a dictionary of obstacles passed to it
# + id="VO_96lW8fcja"
# the initial truck model has a prime mover length of 5m and a trailer of 10m
# width of both is set as 2.5m, but not used yet
TruckDefinition = np.array( [5.0, 2.5, 10.0, 2.5])
# The truck starts with the back of the truck 20m out and 20m forward of the
# end point (0,0) and perpendicular to the finish position
# Note that the position is consistent with the truck definition
StartPosition = np.array( [[20.0,35.0],
[20.0,30],
[20.0,20.0]])
# No obstacles are used yet
Obstacles = {}
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="asY3fHLVllT1" outputId="381ecd76-a330-4018-9f41-ef089ba81078"
env = gym.make('ReverseTruck-v0',TruckDefinition=TruckDefinition, StartPosition=StartPosition, Obstacles=Obstacles)
# + [markdown] id="d61IsZAcK-Wf"
# Follow the method of Practical 1 to run the model
# + [markdown] id="fZBhE9dCLEaO"
# We need to define a policy to take actions.
#
# The simplest policy is to make the back of the truck follow a 20m radius curve till it gets to the loading dock (x=0)
#
# The geometry of this means that the trailer angle should be held at 0.253 rad to the direction of the prime mover.
#
# The policy is then if the trailer angle is less than this, apply "alpha" right steering to increase the angle
# If angle greater than this, apply "alpha" left steering to decrease the angle.
#
# "alpha" is an adjustable parameter. 1 means full 45 deg steerign lock. Lower it to prevent overcorrection if needed
# + id="hh2jU1tmO7lt"
def policy(obs, t):
# Write the code for your policy here. You can use the observation
# (a tuple of position and velocity), the current time step, or both,
# if you want.
pmX, pmY, pivX, pivY, trlX, trlY = obs
alpha=0.2
pmVect=np.array([pmX-pivX,pmY-pivY]) # prime mover vector
trlVect=np.array([pivX-trlX,pivY-trlY]) # trailer vector
pm_trlSin = np.cross(trlVect,pmVect,)/np.linalg.norm(pmVect)/np.linalg.norm(trlVect) #sine of the trailer angle relative to the PM
pm_trlAngle=math.asin(np.clip(pm_trlSin,-1,1)) #relative trailer angle. Clip eliminate rounding errors giving arguments >1
if pm_trlAngle<0.253:
actions=[-1.0,-alpha]
else:
actions=[-1.0,alpha]
return actions
# + colab={"base_uri": "https://localhost:8080/"} id="AfyClfcrKVz3" outputId="daefa39a-2c24-4096-da38-c06cca478224"
o = env.reset()
for t in range(350):
action = policy(o,t) # Call your policy
o, r, d, _ = env.step(action) # Pass the action chosen by the policy to the environment
if t%20==0:
print(t,o,action)
if d and t<TIME_LIMIT-1:
print("Task completed in", t, "time steps")
break
else:
print("Time limit exceeded. Try again.")
env.reset()
| ReverseTruck.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Processing with Python and Pandas Part One
#
#
# ## Today's Topics
#
# * What/Why Pandas
# * Data Structures
# * Loading Data
# * Basic Data Manipulation
# ## What is Pandas
#
# * Pandas is a 3rd-party library for doing data analysis
# * It is a foundational component of Python data science
# * Developed by [<NAME>](http://wesmckinney.com/pages/about.html) while working in the finance industry, so it has some...warts
# * Vanilla Python (what we did previously) can do many of the same things, but Pandas does them *faster* and usually in fewer lines of code
# * To do this, is built on top of another 3rd party library called [numpy](http://www.numpy.org/)
# * If you have TONS of numerical data you can use Numpy directly
# * Pandas gives Python some R like functionality (Dataframes)
# ## Why Pandas?
#
# * Pandas provides a powerful set of data structure and functions for working with data.
# * Once you learn these structures and functions (which takes time) you can begin to quickly ask questions and get answers from data.
# * Pandas integrates nicely with other libraries in the Python data science ecosysem like:
# * [Jupyter Notebooks](http://jupyter.org/) - pretty display of Dataframes as HTML tables
# * [Matplotlib](https://matplotlib.org/) - Easy plotting from Dataframes
# * [Scikit Learn](http://scikit-learn.org/stable/) - Integrates with the machine learning api
#
#
import pandas as pd
# %matplotlib inline
# +
# load the CSV file
data = pd.read_csv("community-center-attendance.csv", index_col="date", parse_dates=True)
# drop the id column because we don't need it
data = data.drop(columns="_id")
# look at the first ten rows of the data
data.head(10)
# -
# What does the data look like?
data.plot();
# We can pivot the data so the center names are columns and each row is the number of people attending that community center per day. This is basically rotating the data.
# Use the pivot function to make column values into columns
data.pivot(columns="center_name", values="attendance_count").head()
# That is a lot of NaN, and not the tasty garlicy kind either.
#
# We might want to break this apart for each Community Center. We can start by inspecting the number rows per center.
# count the number of rows per center and sort the list
data.groupby("center_name").count().sort_values(by=["attendance_count"],
ascending=False)
# We can look at this visually too!
# plot the total attendance
data.groupby("center_name").count().sort_values(by=["attendance_count"],
ascending=False).plot(kind="bar");
# There are a lot of community centers that don't have a lot of numbers because either 1) they are not very popular or 2) they don't report their daily attendance (more likely given how man NaNs we saw above).
#
# What we will do is create a custom filter function that we will apply to ever row in the dataframe using the [groupby filter function](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.core.groupby.DataFrameGroupBy.filter.html). This is some knarly stuff we are doing here. This isn't the plain old filter function, this is a special filter fuction (part of the groupby functionality) that requires you to create a special function to apply to each row. In our case we will make a little function that takes a value and tests to see if it is create than a threshold value (in our case 1000).
# +
# create a function we will use to perform a filtering operation on the data
# filter out centers that have less then 1000 total entries
def filter_less_than(x, threshold):
if len(x) > threshold:
return True
else:
return False
# use the custom function to filter out rows
popular_centers = data.groupby("center_name").filter(filter_less_than,
threshold=1000)
# look at what centers are in the data now
popular_centers.groupby("center_name").count()
# -
# Now we have a more meaty subset of the data to examine.
# look at the first 5 rows
popular_centers.head()
# plot the popular community centers
popular_centers.plot();
# This isn't the most informative representation of the data. Perhaps we can reshape it to make it more useful.
# Use the pivot function to make rows into columns with only the popular community centers
pivoted_data = popular_centers.pivot_table(columns="center_name",
values="attendance_count",
index="date")
pivoted_data.head()
# Still NaN-y, but not as bad. Now we can look at the attendance at the more popular community centers over time.
# plot the data
pivoted_data.plot(figsize=(10,10));
# Still pretty messy. Let's look at the cumulative sum.
# compute the cumulative sum for every column and make a chart
pivoted_data.cumsum().plot(figsize=(10,10));
# Looks like Brookline is the winner here, but attendance has tapered off in the past couple years.
# Resample and compute the monthly totals for the popular community centers
pivoted_data.resample("M").sum().plot(figsize=(10,10));
# Looks like monthly is too messy, maybe by year?
# yearly resample to monthly, compute the totals, and plot
pivoted_data.resample("Y").sum().plot(figsize=(10,10));
| day-two/1-why-pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="QOpPUH3FR179"
# # Laboratory 2: Computer Vision
#
# # Part 2: Debiasing Facial Detection Systems
#
# In the second portion of the lab, we'll explore two prominent aspects of applied deep learning: facial detection and algorithmic bias.
#
# Deploying fair, unbiased AI systems is critical to their long-term acceptance. Consider the task of facial detection: given an image, is it an image of a face? This seemingly simple, but extremely important, task is subject to significant amounts of algorithmic bias among select demographics.
#
# In this lab, we'll investigate [one recently published approach](https://github.com/Jagadambass/Intro-to-TensorFlow-Music-Generation/blob/main/AI_MitigatingAlgorithmicBias.pdf) to addressing algorithmic bias. We'll build a facial detection model that learns the *latent variables* underlying face image datasets and uses this to adaptively re-sample the training data, thus mitigating any biases that may be present in order to train a *debiased* model.
#
#
# Run the next code block for a short video from Google that explores how and why it's important to consider bias when thinking about machine learning:
# + id="XQh5HZfbupFF" colab={"base_uri": "https://localhost:8080/", "height": 321} outputId="3e3c7968-64b6-417e-9bf0-e867c647adb5"
import IPython
IPython.display.YouTubeVideo('59bMh59JQDo')
# + [markdown] id="3Ezfc6Yv6IhI"
# Let's get started by installing the relevant dependencies:
# + id="E46sWVKK6LP9" colab={"base_uri": "https://localhost:8080/"} outputId="a7052c6e-07ae-4282-e3e2-9a9c265dbffc"
# Import Tensorflow 2.0
# %tensorflow_version 2.x
import tensorflow as tf
import IPython
import functools
import matplotlib.pyplot as plt
import numpy as np
from tqdm import tqdm
# Download and import the MIT 6.S191 package
# !pip install mitdeeplearning
import mitdeeplearning as mdl
# + [markdown] id="V0e77oOM3udR"
# ## 2.1 Datasets
#
# We'll be using three datasets in this lab. In order to train our facial detection models, we'll need a dataset of positive examples (i.e., of faces) and a dataset of negative examples (i.e., of things that are not faces). We'll use these data to train our models to classify images as either faces or not faces. Finally, we'll need a test dataset of face images. Since we're concerned about the potential *bias* of our learned models against certain demographics, it's important that the test dataset we use has equal representation across the demographics or features of interest. In this lab, we'll consider skin tone and gender.
#
# 1. **Positive training data**: [CelebA Dataset](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html). A large-scale (over 200K images) of celebrity faces.
# 2. **Negative training data**: [ImageNet](http://www.image-net.org/). Many images across many different categories. We'll take negative examples from a variety of non-human categories.
# [Fitzpatrick Scale](https://en.wikipedia.org/wiki/Fitzpatrick_scale) skin type classification system, with each image labeled as "Lighter'' or "Darker''.
#
# Let's begin by importing these datasets. We've written a class that does a bit of data pre-processing to import the training data in a usable format.
# + id="RWXaaIWy6jVw" colab={"base_uri": "https://localhost:8080/"} outputId="535ce1c9-b0c0-4eff-b17e-d48fedb186d8"
# Get the training data: both images from CelebA and ImageNet
path_to_training_data = tf.keras.utils.get_file('train_face.h5', 'https://www.dropbox.com/s/hlz8atheyozp1yx/train_face.h5?dl=1')
# Instantiate a TrainingDatasetLoader using the downloaded dataset
loader = mdl.lab2.TrainingDatasetLoader(path_to_training_data)
# + [markdown] id="yIE321rxa_b3"
# We can look at the size of the training dataset and grab a batch of size 100:
# + id="DjPSjZZ_bGqe"
number_of_training_examples = loader.get_train_size()
(images, labels) = loader.get_batch(100)
# + [markdown] id="sxtkJoqF6oH1"
# Play around with displaying images to get a sense of what the training data actually looks like!
# + id="Jg17jzwtbxDA" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="42791624-98ec-4f0e-e308-c147cb114320"
### Examining the CelebA training dataset ###
#@title Change the sliders to look at positive and negative training examples! { run: "auto" }
face_images = images[np.where(labels==1)[0]]
not_face_images = images[np.where(labels==0)[0]]
idx_face = 17 #@param {type:"slider", min:0, max:50, step:1}
idx_not_face = 14 #@param {type:"slider", min:0, max:50, step:1}
plt.figure(figsize=(5,5))
plt.subplot(1, 2, 1)
plt.imshow(face_images[idx_face])
plt.title("Face"); plt.grid(False)
plt.subplot(1, 2, 2)
plt.imshow(not_face_images[idx_not_face])
plt.title("Not Face"); plt.grid(False)
# + [markdown] id="NDj7KBaW8Asz"
# ### Thinking about bias
#
# Remember we'll be training our facial detection classifiers on the large, well-curated CelebA dataset (and ImageNet), and then evaluating their accuracy by testing them on an independent test dataset. Our goal is to build a model that trains on CelebA *and* achieves high classification accuracy on the the test dataset across all demographics, and to thus show that this model does not suffer from any hidden bias.
#
# What exactly do we mean when we say a classifier is biased? In order to formalize this, we'll need to think about [*latent variables*](https://en.wikipedia.org/wiki/Latent_variable), variables that define a dataset but are not strictly observed. As defined in the generative modeling lecture, we'll use the term *latent space* to refer to the probability distributions of the aforementioned latent variables. Putting these ideas together, we consider a classifier *biased* if its classification decision changes after it sees some additional latent features. This notion of bias may be helpful to keep in mind throughout the rest of the lab.
# + [markdown] id="AIFDvU4w8OIH"
# ## 2.2 CNN for facial detection
#
# First, we'll define and train a CNN on the facial classification task, and evaluate its accuracy. Later, we'll evaluate the performance of our debiased models against this baseline CNN. The CNN model has a relatively standard architecture consisting of a series of convolutional layers with batch normalization followed by two fully connected layers to flatten the convolution output and generate a class prediction.
#
# ### Define and train the CNN model
#
# Like we did in the first part of the lab, we'll define our CNN model, and then train on the CelebA and ImageNet datasets using the `tf.GradientTape` class and the `tf.GradientTape.gradient` method.
# + id="82EVTAAW7B_X"
### Define the CNN model ###
n_filters = 12 # base number of convolutional filters
'''Function to define a standard CNN model'''
def make_standard_classifier(n_outputs=1):
Conv2D = functools.partial(tf.keras.layers.Conv2D, padding='same', activation='relu')
BatchNormalization = tf.keras.layers.BatchNormalization
Flatten = tf.keras.layers.Flatten
Dense = functools.partial(tf.keras.layers.Dense, activation='relu')
model = tf.keras.Sequential([
Conv2D(filters=1*n_filters, kernel_size=5, strides=2),
BatchNormalization(),
Conv2D(filters=2*n_filters, kernel_size=5, strides=2),
BatchNormalization(),
Conv2D(filters=4*n_filters, kernel_size=3, strides=2),
BatchNormalization(),
Conv2D(filters=6*n_filters, kernel_size=3, strides=2),
BatchNormalization(),
Flatten(),
Dense(512),
Dense(n_outputs, activation=None),
])
return model
standard_classifier = make_standard_classifier()
# + [markdown] id="c-eWf3l_lCri"
# Now let's train the standard CNN!
# + id="eJlDGh1o31G1" colab={"base_uri": "https://localhost:8080/", "height": 530} outputId="c57530c5-d667-47fd-c632-a5ad10a5ac4c"
### Train the standard CNN ###
# Training hyperparameters
batch_size = 32
num_epochs = 2 # keep small to run faster
learning_rate = 5e-4
optimizer = tf.keras.optimizers.Adam(learning_rate) # define our optimizer
loss_history = mdl.util.LossHistory(smoothing_factor=0.99) # to record loss evolution
plotter = mdl.util.PeriodicPlotter(sec=2, scale='semilogy')
if hasattr(tqdm, '_instances'): tqdm._instances.clear() # clear if it exists
@tf.function
def standard_train_step(x, y):
with tf.GradientTape() as tape:
# feed the images into the model
logits = standard_classifier(x)
# Compute the loss
loss = tf.nn.sigmoid_cross_entropy_with_logits(labels=y, logits=logits)
# Backpropagation
grads = tape.gradient(loss, standard_classifier.trainable_variables)
optimizer.apply_gradients(zip(grads, standard_classifier.trainable_variables))
return loss
# The training loop!
for epoch in range(num_epochs):
for idx in tqdm(range(loader.get_train_size()//batch_size)):
# Grab a batch of training data and propagate through the network
x, y = loader.get_batch(batch_size)
loss = standard_train_step(x, y)
# Record the loss and plot the evolution of the loss as a function of training
loss_history.append(loss.numpy().mean())
plotter.plot(loss_history.get())
# + [markdown] id="AKMdWVHeCxj8"
# ### Evaluate performance of the standard CNN
#
# Next, let's evaluate the classification performance of our CelebA-trained standard CNN on the training dataset.
#
# + id="35-PDgjdWk6_" colab={"base_uri": "https://localhost:8080/"} outputId="494e37bc-bca3-429b-97ee-7ff389ace543"
### Evaluation of standard CNN ###
# TRAINING DATA
# Evaluate on a subset of CelebA+Imagenet
(batch_x, batch_y) = loader.get_batch(5000)
y_pred_standard = tf.round(tf.nn.sigmoid(standard_classifier.predict(batch_x)))
acc_standard = tf.reduce_mean(tf.cast(tf.equal(batch_y, y_pred_standard), tf.float32))
print("Standard CNN accuracy on (potentially biased) training set: {:.4f}".format(acc_standard.numpy()))
# + [markdown] id="Qu7R14KaEEvU"
# We will also evaluate our networks on an independent test dataset containing faces that were not seen during training. For the test data, we'll look at the classification accuracy across four different demographics, based on the Fitzpatrick skin scale and sex-based labels: dark-skinned male, dark-skinned female, light-skinned male, and light-skinned female.
#
# Let's take a look at some sample faces in the test set.
# + id="vfDD8ztGWk6x" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="077d7560-22cf-4d6d-e983-b0e5f43f7120"
### Load test dataset and plot examples ###
test_faces = mdl.lab2.get_test_faces()
keys = ["Light Female", "Light Male", "Dark Female", "Dark Male"]
for group, key in zip(test_faces,keys):
plt.figure(figsize=(5,5))
plt.imshow(np.hstack(group))
plt.title(key, fontsize=15)
# + [markdown] id="uo1z3cdbEUMM"
# Now, let's evaluate the probability of each of these face demographics being classified as a face using the standard CNN classifier we've just trained.
# + id="GI4O0Y1GAot9" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="13bbecea-30ad-4463-fe6d-78dc06710b3e"
### Evaluate the standard CNN on the test data ###
standard_classifier_logits = [standard_classifier(np.array(x, dtype=np.float32)) for x in test_faces]
standard_classifier_probs = tf.squeeze(tf.sigmoid(standard_classifier_logits))
# Plot the prediction accuracies per demographic
xx = range(len(keys))
yy = standard_classifier_probs.numpy().mean(1)
plt.bar(xx, yy)
plt.xticks(xx, keys)
plt.ylim(max(0,yy.min()-yy.ptp()/2.), yy.max()+yy.ptp()/2.)
plt.title("Standard classifier predictions");
# + [markdown] id="j0Cvvt90DoAm"
# Take a look at the accuracies for this first model across these four groups. What do you observe? Would you consider this model biased or unbiased? What are some reasons why a trained model may have biased accuracies?
# + [markdown] id="0AKcHnXVtgqJ"
# ## 2.3 Mitigating algorithmic bias
#
# Imbalances in the training data can result in unwanted algorithmic bias. For example, the majority of faces in CelebA (our training set) are those of light-skinned females. As a result, a classifier trained on CelebA will be better suited at recognizing and classifying faces with features similar to these, and will thus be biased.
#
# How could we overcome this? A naive solution -- and one that is being adopted by many companies and organizations -- would be to annotate different subclasses (i.e., light-skinned females, males with hats, etc.) within the training data, and then manually even out the data with respect to these groups.
#
# But this approach has two major disadvantages. First, it requires annotating massive amounts of data, which is not scalable. Second, it requires that we know what potential biases (e.g., race, gender, pose, occlusion, hats, glasses, etc.) to look for in the data. As a result, manual annotation may not capture all the different features that are imbalanced within the training data.
#
# Instead, let's actually **learn** these features in an unbiased, unsupervised manner, without the need for any annotation, and then train a classifier fairly with respect to these features. In the rest of this lab, we'll do exactly that.
# + [markdown] id="nLemS7dqECsI"
# ## 2.4 Variational autoencoder (VAE) for learning latent structure
#
# As you saw, the accuracy of the CNN varies across the four demographics we looked at. To think about why this may be, consider the dataset the model was trained on, CelebA. If certain features, such as dark skin or hats, are *rare* in CelebA, the model may end up biased against these as a result of training with a biased dataset. That is to say, its classification accuracy will be worse on faces that have under-represented features, such as dark-skinned faces or faces with hats, relevative to faces with features well-represented in the training data! This is a problem.
#
# Our goal is to train a *debiased* version of this classifier -- one that accounts for potential disparities in feature representation within the training data. Specifically, to build a debiased facial classifier, we'll train a model that **learns a representation of the underlying latent space** to the face training data. The model then uses this information to mitigate unwanted biases by sampling faces with rare features, like dark skin or hats, *more frequently* during training. The key design requirement for our model is that it can learn an *encoding* of the latent features in the face data in an entirely *unsupervised* way. To achieve this, we'll turn to variational autoencoders (VAEs).
#
# 
#
# As shown in the schematic above and in Lecture 4, VAEs rely on an encoder-decoder structure to learn a latent representation of the input data. In the context of computer vision, the encoder network takes in input images, encodes them into a series of variables defined by a mean and standard deviation, and then draws from the distributions defined by these parameters to generate a set of sampled latent variables. The decoder network then "decodes" these variables to generate a reconstruction of the original image, which is used during training to help the model identify which latent variables are important to learn.
#
# Let's formalize two key aspects of the VAE model and define relevant functions for each.
#
# + [markdown] id="KmbXKtcPkTXA"
# ### Understanding VAEs: loss function
#
# In practice, how can we train a VAE? In learning the latent space, we constrain the means and standard deviations to approximately follow a unit Gaussian. Recall that these are learned parameters, and therefore must factor into the loss computation, and that the decoder portion of the VAE is using these parameters to output a reconstruction that should closely match the input image, which also must factor into the loss. What this means is that we'll have two terms in our VAE loss function:
#
# 1. **Latent loss ($L_{KL}$)**: measures how closely the learned latent variables match a unit Gaussian and is defined by the Kullback-Leibler (KL) divergence.
# 2. **Reconstruction loss ($L_{x}{(x,\hat{x})}$)**: measures how accurately the reconstructed outputs match the input and is given by the $L^1$ norm of the input image and its reconstructed output.
# + [markdown] id="Ux3jK2wc153s"
# The equation for the latent loss is provided by:
#
# $$L_{KL}(\mu, \sigma) = \frac{1}{2}\sum_{j=0}^{k-1} (\sigma_j + \mu_j^2 - 1 - \log{\sigma_j})$$
#
# The equation for the reconstruction loss is provided by:
#
# $$L_{x}{(x,\hat{x})} = ||x-\hat{x}||_1$$
#
# Thus for the VAE loss we have:
#
# $$L_{VAE} = c\cdot L_{KL} + L_{x}{(x,\hat{x})}$$
#
# where $c$ is a weighting coefficient used for regularization. Now we're ready to define our VAE loss function:
# + id="S00ASo1ImSuh"
### Defining the VAE loss function ###
def vae_loss_function(x, x_recon, mu, logsigma, kl_weight=0.0005):
# TODO: Define the latent loss. Note this is given in the equation for L_{KL}
latent_loss = 0.5 * tf.reduce_sum(tf.exp(logsigma) + tf.square(mu) - 1.0 - logsigma, axis=1)
# https://www.tensorflow.org/api_docs/python/tf/math/reduce_mean
reconstruction_loss = tf.reduce_mean(tf.abs(x-x_recon), axis=(1,2,3))
vae_loss = kl_weight * latent_loss + reconstruction_loss
return vae_loss
# + [markdown] id="E8mpb3pJorpu"
# Great! Now that we have a more concrete sense of how VAEs work, let's explore how we can leverage this network structure to train a *debiased* facial classifier.
# + [markdown] id="DqtQH4S5fO8F"
# ### Understanding VAEs: reparameterization
#
# As you may recall from lecture, VAEs use a "reparameterization trick" for sampling learned latent variables. Instead of the VAE encoder generating a single vector of real numbers for each latent variable, it generates a vector of means and a vector of standard deviations that are constrained to roughly follow Gaussian distributions. We then sample from the standard deviations and add back the mean to output this as our sampled latent vector. Formalizing this for a latent variable $z$ where we sample $\epsilon \sim \mathcal{N}(0,(I))$ we have:
#
# $$z = \mu + e^{\left(\frac{1}{2} \cdot \log{\Sigma}\right)}\circ \epsilon$$
#
# where $\mu$ is the mean and $\Sigma$ is the covariance matrix. This is useful because it will let us neatly define the loss function for the VAE, generate randomly sampled latent variables, achieve improved network generalization, **and** make our complete VAE network differentiable so that it can be trained via backpropagation. Quite powerful!
#
# Let's define a function to implement the VAE sampling operation:
# + id="cT6PGdNajl3K"
### VAE Reparameterization ###
"""Reparameterization trick by sampling from an isotropic unit Gaussian.
# Arguments
z_mean, z_logsigma (tensor): mean and log of standard deviation of latent distribution (Q(z|X))
# Returns
z (tensor): sampled latent vector
"""
def sampling(z_mean, z_logsigma):
# By default, random.normal is "standard" (ie. mean=0 and std=1.0)
batch, latent_dim = z_mean.shape
epsilon = tf.random.normal(shape=(batch, latent_dim))
# TODO: Define the reparameterization computation!
# Note the equation is given in the text block immediately above.
z = z_mean + tf.math.exp(0.5 * z_logsigma) * epsilon
# z = # TODO
return z
# + [markdown] id="qtHEYI9KNn0A"
# ## 2.5 Debiasing variational autoencoder (DB-VAE)
#
# Now, we'll use the general idea behind the VAE architecture to build a model, termed a [*debiasing variational autoencoder*](https://github.com/Jagadambass/Intro-to-TensorFlow-Music-Generation/blob/main/AIES-19_paper_220.pdf) or DB-VAE, to mitigate (potentially) unknown biases present within the training idea. We'll train our DB-VAE model on the facial detection task, run the debiasing operation during training, evaluate on the PPB dataset, and compare its accuracy to our original, biased CNN model.
#
# ### The DB-VAE model
#
# The key idea behind this debiasing approach is to use the latent variables learned via a VAE to adaptively re-sample the CelebA data during training. Specifically, we will alter the probability that a given image is used during training based on how often its latent features appear in the dataset. So, faces with rarer features (like dark skin, sunglasses, or hats) should become more likely to be sampled during training, while the sampling probability for faces with features that are over-represented in the training dataset should decrease (relative to uniform random sampling across the training data).
#
# A general schematic of the DB-VAE approach is shown here:
#
# [DB-VAE](https://github.com/Jagadambass/Intro-to-TensorFlow-Music-Generation/blob/main/lab2/img/DB-VAE.png)
# + [markdown] id="ziA75SN-UxxO"
# Recall that we want to apply our DB-VAE to a *supervised classification* problem -- the facial detection task. Importantly, note how the encoder portion in the DB-VAE architecture also outputs a single supervised variable, $z_o$, corresponding to the class prediction -- face or not face. Usually, VAEs are not trained to output any supervised variables (such as a class prediction)! This is another key distinction between the DB-VAE and a traditional VAE.
#
# Keep in mind that we only want to learn the latent representation of *faces*, as that's what we're ultimately debiasing against, even though we are training a model on a binary classification problem. We'll need to ensure that, **for faces**, our DB-VAE model both learns a representation of the unsupervised latent variables, captured by the distribution $q_\phi(z|x)$, **and** outputs a supervised class prediction $z_o$, but that, **for negative examples**, it only outputs a class prediction $z_o$.
# + [markdown] id="XggIKYPRtOZR"
# ### Defining the DB-VAE loss function
#
# This means we'll need to be a bit clever about the loss function for the DB-VAE. The form of the loss will depend on whether it's a face image or a non-face image that's being considered.
#
# For **face images**, our loss function will have two components:
#
#
# 1. **VAE loss ($L_{VAE}$)**: consists of the latent loss and the reconstruction loss.
# 2. **Classification loss ($L_y(y,\hat{y})$)**: standard cross-entropy loss for a binary classification problem.
#
# In contrast, for images of **non-faces**, our loss function is solely the classification loss.
#
# We can write a single expression for the loss by defining an indicator variable $\mathcal{I}_f$which reflects which training data are images of faces ($\mathcal{I}_f(y) = 1$ ) and which are images of non-faces ($\mathcal{I}_f(y) = 0$). Using this, we obtain:
#
# $$L_{total} = L_y(y,\hat{y}) + \mathcal{I}_f(y)\Big[L_{VAE}\Big]$$
#
# Let's write a function to define the DB-VAE loss function:
#
# + id="VjieDs8Ovcqs"
### Loss function for DB-VAE ###
"""Loss function for DB-VAE.
# Arguments
x: true input x
x_pred: reconstructed x
y: true label (face or not face)
y_logit: predicted labels
mu: mean of latent distribution (Q(z|X))
logsigma: log of standard deviation of latent distribution (Q(z|X))
# Returns
total_loss: DB-VAE total loss
classification_loss = DB-VAE classification loss
"""
def debiasing_loss_function(x, x_pred, y, y_logit, mu, logsigma):
vae_loss = vae_loss_function(x, x_pred, mu, logsigma)
classification_loss = tf.nn.sigmoid_cross_entropy_with_logits(labels=y, logits=y_logit)
face_indicator = tf.cast(tf.equal(y, 1), tf.float32)
total_loss = tf.reduce_mean(
classification_loss +
face_indicator * vae_loss
)
# total_loss = # TODO
return total_loss, classification_loss
# + [markdown] id="YIu_2LzNWwWY"
# ### DB-VAE architecture
#
# Now we're ready to define the DB-VAE architecture. To build the DB-VAE, we will use the standard CNN classifier from above as our encoder, and then define a decoder network. We will create and initialize the two models, and then construct the end-to-end VAE. We will use a latent space with 100 latent variables.
#
# The decoder network will take as input the sampled latent variables, run them through a series of deconvolutional layers, and output a reconstruction of the original input image.
# + id="JfWPHGrmyE7R"
### Define the decoder portion of the DB-VAE ###
n_filters = 12 # base number of convolutional filters, same as standard CNN
latent_dim = 100 # number of latent variables
def make_face_decoder_network():
# Functionally define the different layer types we will use
Conv2DTranspose = functools.partial(tf.keras.layers.Conv2DTranspose, padding='same', activation='relu')
BatchNormalization = tf.keras.layers.BatchNormalization
Flatten = tf.keras.layers.Flatten
Dense = functools.partial(tf.keras.layers.Dense, activation='relu')
Reshape = tf.keras.layers.Reshape
# Build the decoder network using the Sequential API
decoder = tf.keras.Sequential([
# Transform to pre-convolutional generation
Dense(units=4*4*6*n_filters), # 4x4 feature maps (with 6N occurances)
Reshape(target_shape=(4, 4, 6*n_filters)),
# Upscaling convolutions (inverse of encoder)
Conv2DTranspose(filters=4*n_filters, kernel_size=3, strides=2),
Conv2DTranspose(filters=2*n_filters, kernel_size=3, strides=2),
Conv2DTranspose(filters=1*n_filters, kernel_size=5, strides=2),
Conv2DTranspose(filters=3, kernel_size=5, strides=2),
])
return decoder
# + [markdown] id="yWCMu12w1BuD"
# Now, we will put this decoder together with the standard CNN classifier as our encoder to define the DB-VAE. Note that at this point, there is nothing special about how we put the model together that makes it a "debiasing" model -- that will come when we define the training operation. Here, we will define the core VAE architecture by sublassing the `Model` class; defining encoding, reparameterization, and decoding operations; and calling the network end-to-end.
# + id="dSFDcFBL13c3"
### Defining and creating the DB-VAE ###
class DB_VAE(tf.keras.Model):
def __init__(self, latent_dim):
super(DB_VAE, self).__init__()
self.latent_dim = latent_dim
# Define the number of outputs for the encoder. Recall that we have
# `latent_dim` latent variables, as well as a supervised output for the
# classification.
num_encoder_dims = 2*self.latent_dim + 1
self.encoder = make_standard_classifier(num_encoder_dims)
self.decoder = make_face_decoder_network()
# function to feed images into encoder, encode the latent space, and output
# classification probability
def encode(self, x):
# encoder output
encoder_output = self.encoder(x)
# classification prediction
y_logit = tf.expand_dims(encoder_output[:, 0], -1)
# latent variable distribution parameters
z_mean = encoder_output[:, 1:self.latent_dim+1]
z_logsigma = encoder_output[:, self.latent_dim+1:]
return y_logit, z_mean, z_logsigma
# VAE reparameterization: given a mean and logsigma, sample latent variables
def reparameterize(self, z_mean, z_logsigma):
# TODO: call the sampling function defined above
z = sampling(z_mean, z_logsigma)
# z = # TODO
return z
# Decode the latent space and output reconstruction
def decode(self, z):
# TODO: use the decoder to output the reconstruction
reconstruction = self.decoder(z)
# reconstruction = # TODO
return reconstruction
# The call function will be used to pass inputs x through the core VAE
def call(self, x):
# Encode input to a prediction and latent space
y_logit, z_mean, z_logsigma = self.encode(x)
# TODO: reparameterization
z = self.reparameterize(z_mean, z_logsigma)
# z = # TODO
# TODO: reconstruction
recon = self.decode(z)
# recon = # TODO
return y_logit, z_mean, z_logsigma, recon
# Predict face or not face logit for given input x
def predict(self, x):
y_logit, z_mean, z_logsigma = self.encode(x)
return y_logit
dbvae = DB_VAE(latent_dim)
# + [markdown] id="M-clbYAj2waY"
# As stated, the encoder architecture is identical to the CNN from earlier in this lab. Note the outputs of our constructed DB_VAE model in the `call` function: `y_logit, z_mean, z_logsigma, z`. Think carefully about why each of these are outputted and their significance to the problem at hand.
#
#
# + [markdown] id="nbDNlslgQc5A"
# ### Adaptive resampling for automated debiasing with DB-VAE
#
# So, how can we actually use DB-VAE to train a debiased facial detection classifier?
#
# Recall the DB-VAE architecture: as input images are fed through the network, the encoder learns an estimate $\mathcal{Q}(z|X)$ of the latent space. We want to increase the relative frequency of rare data by increased sampling of under-represented regions of the latent space. We can approximate $\mathcal{Q}(z|X)$ using the frequency distributions of each of the learned latent variables, and then define the probability distribution of selecting a given datapoint $x$ based on this approximation. These probability distributions will be used during training to re-sample the data.
#
# You'll write a function to execute this update of the sampling probabilities, and then call this function within the DB-VAE training loop to actually debias the model.
# + [markdown] id="Fej5FDu37cf7"
# First, we've defined a short helper function `get_latent_mu` that returns the latent variable means returned by the encoder after a batch of images is inputted to the network:
# + id="ewWbf7TE7wVc"
# Function to return the means for an input image batch
def get_latent_mu(images, dbvae, batch_size=1024):
N = images.shape[0]
mu = np.zeros((N, latent_dim))
for start_ind in range(0, N, batch_size):
end_ind = min(start_ind+batch_size, N+1)
batch = (images[start_ind:end_ind]).astype(np.float32)/255.
_, batch_mu, _ = dbvae.encode(batch)
mu[start_ind:end_ind] = batch_mu
return mu
# + [markdown] id="wn4yK3SC72bo"
# Now, let's define the actual resampling algorithm `get_training_sample_probabilities`. Importantly note the argument `smoothing_fac`. This parameter tunes the degree of debiasing: for `smoothing_fac=0`, the re-sampled training set will tend towards falling uniformly over the latent space, i.e., the most extreme debiasing.
# + id="HiX9pmmC7_wn"
### Resampling algorithm for DB-VAE ###
'''Function that recomputes the sampling probabilities for images within a batch
based on how they distribute across the training data'''
def get_training_sample_probabilities(images, dbvae, bins=10, smoothing_fac=0.001):
print("Recomputing the sampling probabilities")
# TODO: run the input batch and get the latent variable means
mu = get_latent_mu(images, dbvae)
# mu = get_latent_mu('''TODO''') # TODO
# sampling probabilities for the images
training_sample_p = np.zeros(mu.shape[0])
# consider the distribution for each latent variable
for i in range(latent_dim):
latent_distribution = mu[:,i]
# generate a histogram of the latent distribution
hist_density, bin_edges = np.histogram(latent_distribution, density=True, bins=bins)
# find which latent bin every data sample falls in
bin_edges[0] = -float('inf')
bin_edges[-1] = float('inf')
# TODO: call the digitize function to find which bins in the latent distribution
# every data sample falls in to
# https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.digitize.html
bin_idx = np.digitize(latent_distribution, bin_edges)
# bin_idx = np.digitize('''TODO''', '''TODO''') # TODO
# smooth the density function
hist_smoothed_density = hist_density + smoothing_fac
hist_smoothed_density = hist_smoothed_density / np.sum(hist_smoothed_density)
# invert the density function
p = 1.0/(hist_smoothed_density[bin_idx-1])
# TODO: normalize all probabilities
p = p / np.sum(p)
# p = # TODO
# TODO: update sampling probabilities by considering whether the newly
# computed p is greater than the existing sampling probabilities.
training_sample_p = np.maximum(p, training_sample_p)
# training_sample_p = # TODO
# final normalization
training_sample_p /= np.sum(training_sample_p)
return training_sample_p
# + [markdown] id="pF14fQkVUs-a"
# Now that we've defined the resampling update, we can train our DB-VAE model on the CelebA/ImageNet training data, and run the above operation to re-weight the importance of particular data points as we train the model. Remember again that we only want to debias for features relevant to *faces*, not the set of negative examples. Complete the code block below to execute the training loop!
# + id="xwQs-Gu5bKEK" colab={"base_uri": "https://localhost:8080/", "height": 780} outputId="ea52f4d7-bcb4-4147-893b-4a366d9c41e2"
### Training the DB-VAE ###
# Hyperparameters
batch_size = 32
learning_rate = 5e-4
latent_dim = 100
# DB-VAE needs slightly more epochs to train since its more complex than
# the standard classifier so we use 6 instead of 2
num_epochs = 6
# instantiate a new DB-VAE model and optimizer
dbvae = DB_VAE(100)
optimizer = tf.keras.optimizers.Adam(learning_rate)
# To define the training operation, we will use tf.function which is a powerful tool
# that lets us turn a Python function into a TensorFlow computation graph.
@tf.function
def debiasing_train_step(x, y):
with tf.GradientTape() as tape:
# Feed input x into dbvae. Note that this is using the DB_VAE call function!
y_logit, z_mean, z_logsigma, x_recon = dbvae(x)
'''TODO: call the DB_VAE loss function to compute the loss'''
loss, class_loss = debiasing_loss_function(x, x_recon, y, y_logit, z_mean, z_logsigma)
# loss, class_loss = debiasing_loss_function('''TODO arguments''') # TODO
'''TODO: use the GradientTape.gradient method to compute the gradients.
Hint: this is with respect to the trainable_variables of the dbvae.'''
grads = tape.gradient(loss, dbvae.trainable_variables)
# grads = tape.gradient('''TODO''', '''TODO''') # TODO
# apply gradients to variables
optimizer.apply_gradients(zip(grads, dbvae.trainable_variables))
return loss
# get training faces from data loader
all_faces = loader.get_all_train_faces()
if hasattr(tqdm, '_instances'): tqdm._instances.clear() # clear if it exists
# The training loop -- outer loop iterates over the number of epochs
for i in range(num_epochs):
IPython.display.clear_output(wait=True)
print("Starting epoch {}/{}".format(i+1, num_epochs))
# Recompute data sampling proabilities
'''TODO: recompute the sampling probabilities for debiasing'''
p_faces = get_training_sample_probabilities(all_faces, dbvae)
# p_faces = get_training_sample_probabilities('''TODO''', '''TODO''') # TODO
# get a batch of training data and compute the training step
for j in tqdm(range(loader.get_train_size() // batch_size)):
# load a batch of data
(x, y) = loader.get_batch(batch_size, p_pos=p_faces)
# loss optimization
loss = debiasing_train_step(x, y)
# plot the progress every 200 steps
if j % 500 == 0:
mdl.util.plot_sample(x, y, dbvae)
# + [markdown] id="uZBlWDPOVcHg"
# Wonderful! Now we should have a trained and (hopefully!) debiased facial classification model, ready for evaluation!
# + [markdown] id="Eo34xC7MbaiQ"
# ## 2.6 Evaluation of DB-VAE on Test Dataset
#
# Finally let's test our DB-VAE model on the test dataset, looking specifically at its accuracy on each the "Dark Male", "Dark Female", "Light Male", and "Light Female" demographics. We will compare the performance of this debiased model against the (potentially biased) standard CNN from earlier in the lab.
# + id="bgK77aB9oDtX" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="cd577c00-01c0-42d7-e2a3-b55719565138"
dbvae_logits = [dbvae.predict(np.array(x, dtype=np.float32)) for x in test_faces]
dbvae_probs = tf.squeeze(tf.sigmoid(dbvae_logits))
xx = np.arange(len(keys))
plt.bar(xx, standard_classifier_probs.numpy().mean(1), width=0.2, label="Standard CNN")
plt.bar(xx+0.2, dbvae_probs.numpy().mean(1), width=0.2, label="DB-VAE")
plt.xticks(xx, keys);
plt.title("Network predictions on test dataset")
plt.ylabel("Probability"); plt.legend(bbox_to_anchor=(1.04,1), loc="upper left");
# + [markdown] id="rESoXRPQo_mq"
# ## 2.7 Conclusion and submission information
#
# We encourage you to think about and maybe even address some questions raised by the approach and results outlined here:
#
# * How does the accuracy of the DB-VAE across the four demographics compare to that of the standard CNN? Do you find this result surprising in any way?
# * How can the performance of the DB-VAE classifier be improved even further? We purposely did not optimize hyperparameters to leave this up to you!
# * In which applications (either related to facial detection or not!) would debiasing in this way be desired? Are there applications where you may not want to debias your model?
# * Do you think it should be necessary for companies to demonstrate that their models, particularly in the context of tasks like facial detection, are not biased? If so, do you have thoughts on how this could be standardized and implemented?
# * Do you have ideas for other ways to address issues of bias, particularly in terms of the training data?
#
# Try to optimize your model to achieve improved performance. **MIT students and affiliates will be eligible for prizes during the IAP offering.** To enter the competition, MIT students and affiliates should upload the following to the course Canvas:
#
# * Jupyter notebook with the code you used to generate your results;
# * copy of the bar plot from section 2.6 showing the performance of your model;
# * a description and/or diagram of the architecture and hyperparameters you used -- if there are any additional or interesting modifications you made to the template code, please include these in your description;
# * discussion of why these modifications helped improve performance.
#
# Hopefully this lab has shed some light on a few concepts, from vision based tasks, to VAEs, to algorithmic bias. We like to think it has, but we're biased ;).
#
# <img src="https://i.ibb.co/BjLSRMM/ezgif-2-253dfd3f9097.gif" />
# + id="tbWEwAzR6SIP"
| lab2/solutions/Part2_Debiasing_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import astropy.units as u
import astropy.coordinates as coord
from astropy.table import Table
from astropy.io import fits
import scipy.stats
# -
plt.style.use('notebook')
# +
t = Table.read('../data/rcat.fits')
ind = np.isfinite(t['GAIADR2_PMRA']) & (t['FLAG']==0) & (t['SNR']>10) & (t['E_tot_pot1']<0)
t = t[ind]
print(len(t))
age = 10**t['logAge']*1e-9
age_lerr = age - 10**(t['logAge']-t['logAge_lerr'])*1e-9
age_uerr = 10**(t['logAge']+t['logAge_uerr'])*1e-9 - age
age_err = 0.5 * (age_lerr + age_uerr)
age_err_cons = np.maximum(age_lerr, age_uerr)
age_err_min = 1
age_err_cons[age_err_cons<age_err_min] = age_err_min
age_invar = (age_err_cons)**-2
# -
blue = '#0039d3'
purple = '#4700d3'
orange = '#ff6200'
pink = '#ee0449'
ind_all = np.ones(len(t), dtype='bool')
ind_z = (np.abs(t['Z_gal'])<1)
loggmin, loggmax = 3.8, 4.3
ind_safeage = (t['logg']>loggmin) & (t['logg']<loggmax) #& (t['logg_err']<0.1)
# ind_safeage = (t['logg']>3.8) & (t['logg']<4.3) & (age_err/age<0.15)
print(np.sum(ind_safeage))
# +
ind_highe = (t['eccen_pot1']>0.75)
ind_lowe = (t['eccen_pot1']<0.25)
p_ge = [-0.32,-0.02]
poly_ge = np.poly1d(p_ge)
p_splash = [-0.1,0.18]
p_splash = [-0.14,0.18]
poly_splash = np.poly1d(p_splash)
p_lowa = [-0.16,0.1]
p_lowa = [-0.14,0.15]
# p_lowa = [-0.2,0.1]
poly_lowa = np.poly1d(p_lowa)
ind_trap = ((t['init_FeH']>-0.5) & (t['init_FeH']<-0.1) & (t['init_aFe']>0.0)
& (t['init_aFe']<poly_lowa(t['init_FeH'])))
ind_lowa = ind_trap & ind_lowe
ind_lowa = ind_lowe & (t['init_FeH']>-0.45) & (t['init_aFe']<poly_lowa(t['init_FeH'])) & (t['init_FeH']<-0.1)
ind_mpoor = (t['init_FeH']<-0.6) & (t['init_aFe']<poly_ge(t['init_FeH']))
ind_ge = ind_mpoor & ind_highe
ind_tdchem = (t['init_FeH']>-0.75) & (t['init_aFe']>poly_splash(t['init_FeH']))
ind_splash = ind_tdchem & ind_highe
ind_thick = ind_tdchem & ind_lowe
ind_overlap = (t['init_FeH']>-0.75) & (t['init_FeH']<-0.7)
# -
labels = ['Distance [kpc]', 'Age [Gyr]', 'Metallicity [Fe/H]']
for e, x in enumerate([t['dist_adpt'], age, t['init_FeH']]):
print(labels[e])
print('Accreted: {:.2f}\tIn-situ: {:.2f}\n'.format(np.median(x[ind_safeage & ind_ge]),
np.median(x[ind_safeage & ind_splash])))
labels = ['Distance [kpc]', 'Age [Gyr]', 'Metallicity [Fe/H]']
for e, x in enumerate([t['dist_adpt'], age, t['init_FeH']]):
print(labels[e])
print('Accreted: {:.2f}\tIn-situ: {:.2f}\n'.format(np.median(x[ind_safeage & ind_ge & ind_overlap]),
np.median(x[ind_safeage & ind_splash & ind_overlap])))
# +
isochrones = []
for age_ in [11.5, 10]:
for feh_ in [-1,-0.5]:
iso = Table.read('../data/mist_{:.1f}_{:.2f}.cmd'.format(age_, feh_),
format='ascii.commented_header', header_start=12)
phasecut = (iso['phase']>=0) & (iso['phase']<3)
iso = iso[phasecut]
isochrones += [iso]
# -
print(t.colnames)
# +
dm = 5*np.log10(t['dist_adpt']*1e3) - 5
N = min(np.sum(ind_splash), np.sum(ind_ge))
N = -1
plt.plot(t['PS_G'][ind_splash][:N] - t['PS_I'][ind_splash][:N], t['PS_G'][ind_splash][:N]-dm[ind_splash][:N],
'ro', alpha=0.8, ms=2, mew=0, label='In-situ')
plt.plot(t['PS_G'][ind_ge][:N] - t['PS_I'][ind_ge][:N], t['PS_G'][ind_ge][:N]-dm[ind_ge][:N],
'bo', alpha=0.8, ms=2, mew=0, label='Accreted')
label_age = [11.5, 11.5, 10, 10]
label_feh = [-1, -0.5, -1, -0.5]
for e, iso in enumerate(isochrones):
# dm = 5 * np.log10(2800) - 5
i_gi = iso['PS_g']-iso['PS_i'] + 0.05
i_g = iso['PS_g'] #+dm
plt.plot(i_gi, i_g, '-', label='{:.1f} Gyr [Fe/H]={:.1f}'.format(label_age[e], label_feh[e]))
plt.legend(markerscale=2, handlelength=1, fontsize='small')
plt.xlim(-0.2,1.7)
plt.ylim(7,-3)
plt.xlabel('g - i [mag]')
plt.ylabel('$M_g$ [mag]')
plt.tight_layout()
# plt.savefig('../plots/cmd.png')
# +
dm = 5*np.log10(t['dist_adpt']*1e3) - 5
# N = min(np.sum(ind_splash & ind_overlap), np.sum(ind_ge & ind_overlap))
N = -1
plt.plot(t['PS_G'][ind_splash & ind_overlap][:N] - t['PS_I'][ind_splash & ind_overlap][:N],
t['PS_G'][ind_splash & ind_overlap & ind_overlap][:N]-dm[ind_splash & ind_overlap][:N],
'ro', alpha=0.8, ms=2, mew=0, label='In-situ')
plt.plot(t['PS_G'][ind_ge & ind_overlap][:N] - t['PS_I'][ind_ge & ind_overlap][:N],
t['PS_G'][ind_ge & ind_overlap][:N]-dm[ind_ge & ind_overlap][:N],
'bo', alpha=0.8, ms=2, mew=0, label='Accreted')
label_age = [11.5, 11.5, 10, 10]
label_feh = [-1, -0.5, -1, -0.5]
for e, iso in enumerate(isochrones):
# dm = 5 * np.log10(2800) - 5
i_gi = iso['PS_g']-iso['PS_i'] #+ 0.05
i_g = iso['PS_g'] #+dm
# plt.plot(i_gi, i_g, '-', label='{:.1f} Gyr [Fe/H]={:.1f}'.format(label_age[e], label_feh[e]))
plt.legend(markerscale=2, handlelength=1, fontsize='small')
plt.xlim(-0.2,1.7)
plt.ylim(7,-3)
plt.xlabel('g - i [mag]')
plt.ylabel('$M_g$ [mag]')
plt.tight_layout()
# plt.savefig('../plots/cmd.png')
# +
ind_mede = (t['eccen_pot1']>0.)
plt.figure(figsize=(10,6))
plt.plot(age[ind_mede & ind_safeage], t['init_FeH'][ind_mede & ind_safeage], 'k.', ms=0.5
)
plt.errorbar(age[ind_mede & ind_safeage], t['init_FeH'][ind_mede & ind_safeage],
xerr=(age_lerr[ind_mede & ind_safeage], age_uerr[ind_mede & ind_safeage]),
yerr=(t['init_FeH_lerr'][ind_mede & ind_safeage], t['init_FeH_uerr'][ind_mede & ind_safeage]),
fmt='none', color='k', alpha=0.1, lw=0.1)
plt.xlabel('Age [Gyr]')
plt.ylabel('[Fe/H]$_{init}$')
plt.tight_layout()
# -
| notebooks/cmd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import packages
import os
from math import log
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
# Import AuTuMN modules
from autumn.settings import Models, Region
from autumn.settings.folders import OUTPUT_DATA_PATH
from autumn.tools.project import get_project
from autumn.tools import db
from autumn.tools.plots.calibration.plots import calculate_r_hats, get_output_from_run_id, get_posterior
from autumn.tools.plots.uncertainty.plots import _plot_uncertainty, _get_target_values
from autumn.tools.plots.plotter.base_plotter import COLOR_THEME
from autumn.tools.plots.utils import get_plot_text_dict, change_xaxis_to_date, REF_DATE, ALPHAS, COLORS, _apply_transparency, _plot_targets_to_axis, split_mcmc_outputs_by_chain
from autumn.dashboards.calibration_results.plots import get_uncertainty_df
import yaml
# +
# Specify model details
model = Models.COVID_19
region = Region.VICTORIA_2020
dirname_lhs = "2021-09-05"
dirname_main = "2021-09-04"
main_burn_in = 8000
# +
# get the relevant project and output data
project = get_project(model, region)
project_calib_dir = os.path.join(
OUTPUT_DATA_PATH, "calibrate", project.model_name, project.region_name
)
# load data from LHS analysis
calib_path = os.path.join(project_calib_dir, dirname_lhs)
mcmc_tables = db.load.load_mcmc_tables(calib_path)
mcmc_params = db.load.load_mcmc_params_tables(calib_path)
# Load data from main analysis
main_calib_path = os.path.join(project_calib_dir, dirname_main)
main_mcmc_tables = db.load.load_mcmc_tables(main_calib_path)
main_mcmc_params = db.load.load_mcmc_params_tables(main_calib_path)
# param_names = list(mcmc_params[0]["name"].unique())
param_names = ['victorian_clusters.metro.mobility.microdistancing.face_coverings_adjuster.parameters.effect', 'sojourn.compartment_periods_calculated.active.total_period', 'contact_rate', 'victorian_clusters.intercluster_mixing', 'infectious_seed', 'infection_fatality.top_bracket_overwrite', 'clinical_stratification.props.hospital.multiplier', 'testing_to_detection.assumed_cdr_parameter', 'sojourn.compartment_periods.icu_early', 'victorian_clusters.metro.mobility.microdistancing.behaviour_adjuster.parameters.effect']
# -
# # Param traces
# +
# Get median estimates
medians, lower, upper = {}, {}, {}
for param_name in param_names:
param_values = get_posterior(main_mcmc_params, main_mcmc_tables, param_name, burn_in=main_burn_in)
medians[param_name] = np.quantile(param_values, 0.5)
lower[param_name] = np.quantile(param_values, 0.025)
upper[param_name] = np.quantile(param_values, 0.975)
# +
n_rows, n_cols = 5, 2
fig, axes = plt.subplots(n_rows, n_cols, sharex=True, sharey=False, figsize=(15, 18))
plt.style.use("ggplot")
chain_ids = list(mcmc_params[0]["chain"].unique())
mcmc_params_list, mcmc_tables_list = split_mcmc_outputs_by_chain(mcmc_params, mcmc_tables)
i_row, i_col = 0, 0
for param_name in param_names:
axis = axes[i_row, i_col]
for i_chain in range(len(mcmc_params_list)):
param_values = get_posterior([mcmc_params_list[i_chain]], [mcmc_tables_list[i_chain]], param_name, burn_in=0)
axis.plot(param_values, alpha=0.8, linewidth=0.5) #, color=COLOR_THEME[i_chain])
h_color = "black"
axis.hlines(y=medians[param_name], xmin = 0, xmax=len(param_values), zorder=100, color=h_color, linestyle="solid")
axis.hlines(y=lower[param_name], xmin = 0, xmax=len(param_values), zorder=100, color=h_color, linestyle="dotted")
axis.hlines(y=upper[param_name], xmin = 0, xmax=len(param_values), zorder=100, color=h_color, linestyle="dotted")
i_col += 1
if i_col == n_cols:
i_row += 1
i_col = 0
axis.set_ylabel(get_plot_text_dict(param_name), fontsize=15)
# axis.set_ylim((min_ll - 2, 3))
plt.tight_layout()
plt.savefig("lhs_start_traces_median.png", dpi=150)
plt.savefig("lhs_start_traces_median.pdf")
# -
# # Posterior vs params
def plot_param_vs_loglike(mcmc_tables, mcmc_params, param_name, burn_in, axis, posterior=False):
var_key = "ap_loglikelihood" if posterior else "loglikelihood"
for mcmc_df, param_df in zip(mcmc_tables, mcmc_params):
df = param_df.merge(mcmc_df, on=["run", "chain"])
mask = (df["accept"] == 1) & (df["name"] == param_name) & (df["run"] > burn_in)
df = df[mask]
max_loglike = max(df[var_key]) + 1
min_loglike = min(df[var_key])
chain_ids = list(df["chain"].unique())
# chain_ids.reverse()
for chain_id in chain_ids:
chain_df = df[df["chain"] == chain_id]
param_values = chain_df["value"]
# apply transformation to improve readability
# trans_loglikelihood_values = [-log(-v + max_loglike) for v in chain_df[var_key]]
trans_loglikelihood_values = [-log(-v + max_loglike) for v in chain_df[var_key]]
zorders = list(np.random.randint(0, 10, size=len(param_values)))
axis.plot(list(param_values)[0], list(trans_loglikelihood_values)[0], "*", color="violet", markersize=15)
axis.plot(param_values, trans_loglikelihood_values, ".", color=COLOR_THEME[chain_id], markersize=5)
return -log(-min_loglike + max_loglike)
# +
n_rows, n_cols = 4, 3
fig, axes = plt.subplots(n_rows, n_cols, sharex=False, sharey=True, figsize=(15, 18))
# fig = plt.figure(figsize=(12, 8))
plt.style.use("ggplot")
i_row, i_col = 0, 0
for param_name in param_names:
axis = axes[i_row, i_col]
min_ll = plot_param_vs_loglike(mcmc_tables, mcmc_params, param_name, 0, axis, posterior=False)
i_col += 1
if i_col == n_cols:
i_row += 1
i_col = 0
axis.set_title(get_plot_text_dict(param_name))
if i_col == 1:
axis.set_ylabel("likelihood (transformed)")
axis.set_ylim((min_ll - 2, 0.5))
axis_to_shut = [ [3, 1], [3, 2] ]
for ax_ids in axis_to_shut:
axis = axes[ax_ids[0], ax_ids[1]]
axis.set_axis_off()
plt.tight_layout()
plt.savefig("likelihood_against_params.png", dpi=150)
plt.savefig("likelihood_against_params.pdf")
# -
| notebooks/user/rragonnet/vic_lhs_start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from particletracking import dataframes, statistics
import trackpy as tp
from tqdm import tqdm as tqdm
data = dataframes.DataStore("/media/data/Data/FirstOrder/Histograms/2000_flat.hdf5")
calculator = statistics.PropertyCalculator(data)
calculator.order_long()
particles = data.df['particle'].unique()
| first_order/histograms_linked.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Polygon
from matplotlib.collections import PatchCollection
from matplotlib import rcParams
import seaborn as sns
import pandas as pd
sns.set_context('paper')
rcParams['font.size'] = 10
# %matplotlib inline
# +
group = 100
delta = '0-05'
sample = 10
meas1 = 'alpha_score'
meas2 = 'chord_arc_infinity'
measure_one = '$\\alpha$ fatness'
measure_two = 'Arc $L_\infty$'
input_dir = f'../data/input/mitchell_polygons/{group}/'
output_dir =f'../data/output/mitchell_polygons/{group}/'
u_meas_1_scores = pd.read_csv(output_dir + f'u_{meas1}.csv')
u_meas_2_scores = pd.read_csv(output_dir + f'u_{meas2}.csv')
u_polygons = []
for i in range(1,11):
u_polygons.append(Polygon(np.loadtxt(input_dir + '{}.poly'.format(i), skiprows=1)))
u_meas_1_scores['Polygon'] = u_polygons
unsmoothed = u_meas_2_scores.set_index('n').join(u_meas_1_scores.set_index('n'), lsuffix=f' {measure_two}', rsuffix=f' {measure_one}').iloc[:sample]
u_fine_asorted = unsmoothed.sort_values(f'0.05 {measure_one}',ascending=True)[[f'0.05 {measure_two}', f'0.05 {measure_one}', 'Polygon']]
u_fine_csorted = unsmoothed.sort_values(f'0.05 {measure_two}',ascending=False)[[f'0.05 {measure_two}', 'Polygon']]
# +
fig_u, axes_u = plt.subplots(5,2,figsize=(10,30))
for (i, ax) in enumerate(axes_u.reshape(sample,)):
m1_rank = i
polygon_name = u_fine_asorted.iloc[i].name
m2_rank = u_fine_csorted.index.get_loc(u_fine_asorted.iloc[i].name)
p = PatchCollection([u_fine_asorted['Polygon'].iloc[i]])
ax.add_collection(p)
ax.get_yaxis().set_visible(False)
ax.set_xticks([])
ax.set_title(f'{measure_one} rank: {sample - m1_rank}\n{measure_two} rank: {sample - m2_rank}', fontsize=30)
ax.set_xlabel(f'{measure_one}: {(u_fine_asorted[f"0.05 {measure_one}"].iloc[i]):.4f}\n{measure_two}: {(u_fine_asorted[f"0.05 {measure_two}"].iloc[i]):.4f}', fontsize=20)
sns.despine(ax=ax)
plt.subplots_adjust(hspace=0.5)
fig_u.savefig(f'../plots/u_{group}_{meas1}_{meas2}_vertices_{delta}_delta_ranking.jpg', bbox_inches='tight')
# +
group = 10
#delta = '0-05'
sample = 10
meas1 = 'alpha_score'
meas2 = 'chord_arc_infinity'
measure_one = '$\\alpha$ fatness'
measure_two = 'Arc $L_\infty$'
input_dir = f'../data/input/mitchell_polygons/{group}/'
output_dir =f'../data/output/mitchell_polygons/{group}/'
u_meas_1_scores = pd.read_csv(output_dir + f'u_{meas1}.csv')
u_meas_2_scores = pd.read_csv(output_dir + f'u_{meas2}.csv')
u_polygons = []
for i in range(1,11):
u_polygons.append(Polygon(np.loadtxt(input_dir + '{}.poly'.format(i), skiprows=1)))
#u_meas_1_scores['Polygon'] = u_polygons
unsmoothed = u_meas_2_scores.set_index('n').join(u_meas_1_scores.set_index('n'), lsuffix=f' {measure_two}', rsuffix=f' {measure_one}').iloc[:sample]
u_fine_asorted = unsmoothed.sort_values(f'0.05 {measure_one}',ascending=True)
u_fine_csorted = unsmoothed.sort_values(f'0.05 {measure_two}',ascending=False)
# -
data = u_fine_asorted.loc[1]
print(u_meas_1_scores.transpose().iloc[1:].to_latex())
print(u_meas_1_scores.transpose().iloc[1:].to_latex())
# +
# pd.DataFrame.to_latex?
# -
data
| notebooks/Analysis of Mitchell Polygons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import re
from pprint import pprint as pp
from typing import Set, List, Dict, NamedTuple
class Page(NamedTuple):
pid: int
title: str
class Entry(NamedTuple):
page: Page
attr: str
text: str
fn = "/home/ebal/Data/wikipedia/shinra2019/JP-5/annotation/Company_dist.json"
entries = []
with open(fn) as f:
for line in f:
data = json.loads(line)
entries.append(Entry(Page(int(data["page_id"]), data["title"]), data["attribute"], data["text_offset"]["text"]))
allpages = {entry.page for entry in entries}
fn = "/home/ebal/Data/wikipedia/shinra2019/JP-5/EXTRACTED/Company_train.json"
myentries = []
with open(fn) as f:
reg = re.compile(r"<[^>]*?>")
for line in f:
data = json.loads(line)
myentries.append(Entry(Page(int(data["page_id"]), data["title"]), data["attribute"], reg.sub("", data["html_offset"]["text"])))
def searchr(reg: str, attr: str, pages: Set[Page], entries: Set[Entry]) -> Set[Page]:
reg = re.compile(reg)
return {et.page for et in entries if et.page in pages and et.attr == attr and reg.match(et.text)}
def searchf(fun, attr: str, pages: Set[Page], entries: Set[Entry]) -> Set[Page]:
return {et.page for et in entries if et.page in pages and et.attr == attr and fun(et.text)}
# +
tokyo = searchr(r"東京都?", "本拠地", allpages, entries)
ltd = searchr(r"株式(会社)?", "種類", tokyo, entries)
from show_a_table.model.refiner import date
f1996t2020 = date.DateRangeRefiner("設立年")
f1996t2020._start = "1996-*-*"
f1996t2020._end = "2030-*-*"
exam = f1996t2020._mk_exam()
ans = searchf(exam, "設立年", ltd, entries)
# -
tokyo = searchr(r"東京都?", "本拠地", allpages, myentries)
ltd = searchr(r"株式(会社)?", "種類", tokyo, myentries)
myans = searchf(exam, "設立年", ltd, entries)
myans
ans
| scripts/jupyter/get answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Single-Machine Model Parallel Best Practices
# ================================
# **Author**: `<NAME> <https://mrshenli.github.io/>`_
#
# Model parallel is widely-used in distributed training
# techniques. Previous posts have explained how to use
# `DataParallel <https://pytorch.org/tutorials/beginner/blitz/data_parallel_tutorial.html>`_
# to train a neural network on multiple GPUs; this feature replicates the
# same model to all GPUs, where each GPU consumes a different partition of the
# input data. Although it can significantly accelerate the training process, it
# does not work for some use cases where the model is too large to fit into a
# single GPU. This post shows how to solve that problem by using **model parallel**,
# which, in contrast to ``DataParallel``, splits a single model onto different GPUs,
# rather than replicating the entire model on each GPU (to be concrete, say a model
# ``m`` contains 10 layers: when using ``DataParallel``, each GPU will have a
# replica of each of these 10 layers, whereas when using model parallel on two GPUs,
# each GPU could host 5 layers).
#
# The high-level idea of model parallel is to place different sub-networks of a
# model onto different devices, and implement the ``forward`` method accordingly
# to move intermediate outputs across devices. As only part of a model operates
# on any individual device, a set of devices can collectively serve a larger
# model. In this post, we will not try to construct huge models and squeeze them
# into a limited number of GPUs. Instead, this post focuses on showing the idea
# of model parallel. It is up to the readers to apply the ideas to real-world
# applications.
#
# <div class="alert alert-info"><h4>Note</h4><p>For distributed model parallel training where a model spans multiple
# servers, please refer to
# `Getting Started With Distributed RPC Framework <rpc_tutorial.html>`__
# for examples and details.</p></div>
#
# Basic Usage
# -----------
#
#
# Let us start with a toy model that contains two linear layers. To run this
# model on two GPUs, simply put each linear layer on a different GPU, and move
# inputs and intermediate outputs to match the layer devices accordingly.
#
#
#
# +
import torch
import torch.nn as nn
import torch.optim as optim
class ToyModel(nn.Module):
def __init__(self):
super(ToyModel, self).__init__()
self.net1 = torch.nn.Linear(10, 10).to('cuda:0')
self.relu = torch.nn.ReLU()
self.net2 = torch.nn.Linear(10, 5).to('cuda:1')
def forward(self, x):
x = self.relu(self.net1(x.to('cuda:0')))
return self.net2(x.to('cuda:1'))
# -
# Note that, the above ``ToyModel`` looks very similar to how one would
# implement it on a single GPU, except the five ``to(device)`` calls which
# place linear layers and tensors on proper devices. That is the only place in
# the model that requires changes. The ``backward()`` and ``torch.optim`` will
# automatically take care of gradients as if the model is on one GPU. You only
# need to make sure that the labels are on the same device as the outputs when
# calling the loss function.
#
#
# +
model = ToyModel()
loss_fn = nn.MSELoss()
optimizer = optim.SGD(model.parameters(), lr=0.001)
optimizer.zero_grad()
outputs = model(torch.randn(20, 10))
labels = torch.randn(20, 5).to('cuda:1')
loss_fn(outputs, labels).backward()
optimizer.step()
# -
# Apply Model Parallel to Existing Modules
# ----------------------------------------
#
# It is also possible to run an existing single-GPU module on multiple GPUs
# with just a few lines of changes. The code below shows how to decompose
# ``torchvision.models.reset50()`` to two GPUs. The idea is to inherit from
# the existing ``ResNet`` module, and split the layers to two GPUs during
# construction. Then, override the ``forward`` method to stitch two
# sub-networks by moving the intermediate outputs accordingly.
#
#
# +
from torchvision.models.resnet import ResNet, Bottleneck
num_classes = 1000
class ModelParallelResNet50(ResNet):
def __init__(self, *args, **kwargs):
super(ModelParallelResNet50, self).__init__(
Bottleneck, [3, 4, 6, 3], num_classes=num_classes, *args, **kwargs)
self.seq1 = nn.Sequential(
self.conv1,
self.bn1,
self.relu,
self.maxpool,
self.layer1,
self.layer2
).to('cuda:0')
self.seq2 = nn.Sequential(
self.layer3,
self.layer4,
self.avgpool,
).to('cuda:1')
self.fc.to('cuda:1')
def forward(self, x):
x = self.seq2(self.seq1(x).to('cuda:1'))
return self.fc(x.view(x.size(0), -1))
# -
# The above implementation solves the problem for cases where the model is too
# large to fit into a single GPU. However, you might have already noticed that
# it will be slower than running it on a single GPU if your model fits. It is
# because, at any point in time, only one of the two GPUs are working, while
# the other one is sitting there doing nothing. The performance further
# deteriorates as the intermediate outputs need to be copied from ``cuda:0`` to
# ``cuda:1`` between ``layer2`` and ``layer3``.
#
# Let us run an experiment to get a more quantitative view of the execution
# time. In this experiment, we train ``ModelParallelResNet50`` and the existing
# ``torchvision.models.reset50()`` by running random inputs and labels through
# them. After the training, the models will not produce any useful predictions,
# but we can get a reasonable understanding of the execution times.
#
#
# +
import torchvision.models as models
num_batches = 3
batch_size = 120
image_w = 128
image_h = 128
def train(model):
model.train(True)
loss_fn = nn.MSELoss()
optimizer = optim.SGD(model.parameters(), lr=0.001)
one_hot_indices = torch.LongTensor(batch_size) \
.random_(0, num_classes) \
.view(batch_size, 1)
for _ in range(num_batches):
# generate random inputs and labels
inputs = torch.randn(batch_size, 3, image_w, image_h)
labels = torch.zeros(batch_size, num_classes) \
.scatter_(1, one_hot_indices, 1)
# run forward pass
optimizer.zero_grad()
outputs = model(inputs.to('cuda:0'))
# run backward pass
labels = labels.to(outputs.device)
loss_fn(outputs, labels).backward()
optimizer.step()
# -
# The ``train(model)`` method above uses ``nn.MSELoss`` as the loss function,
# and ``optim.SGD`` as the optimizer. It mimics training on ``128 X 128``
# images which are organized into 3 batches where each batch contains 120
# images. Then, we use ``timeit`` to run the ``train(model)`` method 10 times
# and plot the execution times with standard deviations.
#
#
# +
import matplotlib.pyplot as plt
plt.switch_backend('Agg')
import numpy as np
import timeit
num_repeat = 10
stmt = "train(model)"
setup = "model = ModelParallelResNet50()"
# globals arg is only available in Python 3. In Python 2, use the following
# import __builtin__
# __builtin__.__dict__.update(locals())
mp_run_times = timeit.repeat(
stmt, setup, number=1, repeat=num_repeat, globals=globals())
mp_mean, mp_std = np.mean(mp_run_times), np.std(mp_run_times)
setup = "import torchvision.models as models;" + \
"model = models.resnet50(num_classes=num_classes).to('cuda:0')"
rn_run_times = timeit.repeat(
stmt, setup, number=1, repeat=num_repeat, globals=globals())
rn_mean, rn_std = np.mean(rn_run_times), np.std(rn_run_times)
def plot(means, stds, labels, fig_name):
fig, ax = plt.subplots()
ax.bar(np.arange(len(means)), means, yerr=stds,
align='center', alpha=0.5, ecolor='red', capsize=10, width=0.6)
ax.set_ylabel('ResNet50 Execution Time (Second)')
ax.set_xticks(np.arange(len(means)))
ax.set_xticklabels(labels)
ax.yaxis.grid(True)
plt.tight_layout()
plt.savefig(fig_name)
plt.close(fig)
plot([mp_mean, rn_mean],
[mp_std, rn_std],
['Model Parallel', 'Single GPU'],
'mp_vs_rn.png')
# -
# .. figure:: /_static/img/model-parallel-images/mp_vs_rn.png
# :alt:
#
# The result shows that the execution time of model parallel implementation is
# ``4.02/3.75-1=7%`` longer than the existing single-GPU implementation. So we
# can conclude there is roughly 7% overhead in copying tensors back and forth
# across the GPUs. There are rooms for improvements, as we know one of the two
# GPUs is sitting idle throughout the execution. One option is to further
# divide each batch into a pipeline of splits, such that when one split reaches
# the second sub-network, the following split can be fed into the first
# sub-network. In this way, two consecutive splits can run concurrently on two
# GPUs.
#
#
# Speed Up by Pipelining Inputs
# -----------------------------
#
# In the following experiments, we further divide each 120-image batch into
# 20-image splits. As PyTorch launches CUDA operations asynchronizely, the
# implementation does not need to spawn multiple threads to achieve
# concurrency.
#
#
# +
class PipelineParallelResNet50(ModelParallelResNet50):
def __init__(self, split_size=20, *args, **kwargs):
super(PipelineParallelResNet50, self).__init__(*args, **kwargs)
self.split_size = split_size
def forward(self, x):
splits = iter(x.split(self.split_size, dim=0))
s_next = next(splits)
s_prev = self.seq1(s_next).to('cuda:1')
ret = []
for s_next in splits:
# A. s_prev runs on cuda:1
s_prev = self.seq2(s_prev)
ret.append(self.fc(s_prev.view(s_prev.size(0), -1)))
# B. s_next runs on cuda:0, which can run concurrently with A
s_prev = self.seq1(s_next).to('cuda:1')
s_prev = self.seq2(s_prev)
ret.append(self.fc(s_prev.view(s_prev.size(0), -1)))
return torch.cat(ret)
setup = "model = PipelineParallelResNet50()"
pp_run_times = timeit.repeat(
stmt, setup, number=1, repeat=num_repeat, globals=globals())
pp_mean, pp_std = np.mean(pp_run_times), np.std(pp_run_times)
plot([mp_mean, rn_mean, pp_mean],
[mp_std, rn_std, pp_std],
['Model Parallel', 'Single GPU', 'Pipelining Model Parallel'],
'mp_vs_rn_vs_pp.png')
# -
# Please note, device-to-device tensor copy operations are synchronized on
# current streams on the source and the destination devices. If you create
# multiple streams, you have to make sure that copy operations are properly
# synchronized. Writing the source tensor or reading/writing the destination
# tensor before finishing the copy operation can lead to undefined behavior.
# The above implementation only uses default streams on both source and
# destination devices, hence it is not necessary to enforce additional
# synchronizations.
#
# .. figure:: /_static/img/model-parallel-images/mp_vs_rn_vs_pp.png
# :alt:
#
# The experiment result shows that, pipelining inputs to model parallel
# ResNet50 speeds up the training process by roughly ``3.75/2.51-1=49%``. It is
# still quite far away from the ideal 100% speedup. As we have introduced a new
# parameter ``split_sizes`` in our pipeline parallel implementation, it is
# unclear how the new parameter affects the overall training time. Intuitively
# speaking, using small ``split_size`` leads to many tiny CUDA kernel launch,
# while using large ``split_size`` results to relatively long idle times during
# the first and last splits. Neither are optimal. There might be an optimal
# ``split_size`` configuration for this specific experiment. Let us try to find
# it by running experiments using several different ``split_size`` values.
#
#
# +
means = []
stds = []
split_sizes = [1, 3, 5, 8, 10, 12, 20, 40, 60]
for split_size in split_sizes:
setup = "model = PipelineParallelResNet50(split_size=%d)" % split_size
pp_run_times = timeit.repeat(
stmt, setup, number=1, repeat=num_repeat, globals=globals())
means.append(np.mean(pp_run_times))
stds.append(np.std(pp_run_times))
fig, ax = plt.subplots()
ax.plot(split_sizes, means)
ax.errorbar(split_sizes, means, yerr=stds, ecolor='red', fmt='ro')
ax.set_ylabel('ResNet50 Execution Time (Second)')
ax.set_xlabel('Pipeline Split Size')
ax.set_xticks(split_sizes)
ax.yaxis.grid(True)
plt.tight_layout()
plt.savefig("split_size_tradeoff.png")
plt.close(fig)
# -
# .. figure:: /_static/img/model-parallel-images/split_size_tradeoff.png
# :alt:
#
# The result shows that setting ``split_size`` to 12 achieves the fastest
# training speed, which leads to ``3.75/2.43-1=54%`` speedup. There are
# still opportunities to further accelerate the training process. For example,
# all operations on ``cuda:0`` is placed on its default stream. It means that
# computations on the next split cannot overlap with the copy operation of the
# prev split. However, as prev and next splits are different tensors, there is
# no problem to overlap one's computation with the other one's copy. The
# implementation need to use multiple streams on both GPUs, and different
# sub-network structures require different stream management strategies. As no
# general multi-stream solution works for all model parallel use cases, we will
# not discuss it in this tutorial.
#
# **Note:**
#
# This post shows several performance measurements. You might see different
# numbers when running the same code on your own machine, because the result
# depends on the underlying hardware and software. To get the best performance
# for your environment, a proper approach is to first generate the curve to
# figure out the best split size, and then use that split size to pipeline
# inputs.
#
#
#
| docs/_downloads/03a48646520c277662581e858e680809/model_parallel_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding=utf-8
import requests
from bs4 import BeautifulSoup
import re
# Encoding problems
# import urllib.request
# import urllib3
headers = {
# 'accept':'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8',
# 'accept-encoding':'gzip, deflate, br',
# 'accept-language':'en-US,en;q=0.9,ru;q=0.8',
# 'upgrade-insecure-requests':'1',
'user-agent':'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_0) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/62.0.3202.75 Safari/537.36'
}
def get_frequency(phrase="abracadabrr"):
url='https://www.google.ru/search?&q='+'"'+phrase+'"'
try:
r = requests.get(url, headers=headers)
print(r)
r.raise_for_status()
except requests.HTTPError:
print('Sorry!!!', r)
return -1;
html=r.text
# print(html)
# Encoding problems
# http = urllib3.PoolManager()
# r = http.request('GET', url)
# print(r.status)
# html=r.data
# Encoding problems
# with urllib.request.urlopen(url) as f:
# html = f.read()
soup = BeautifulSoup(html, 'html.parser')
#span that says there is no such phrase
span = soup.find('span', class_="spell_orig")
if span:
print (span.get_text())
# div with stats
div = soup.find(id="resultStats")
# remove all exept digits from the inner text
t=div.get_text()
if div:
t = re.sub( r'\(.*\)', '', t)
num = re.sub( r'[^\d]', '', t)
return (int(num) if num else 0)
else:
return -1
def get_frequency_y(phrase="abracadabrr"):
url='http://yandex.ru/search/?text='+'"'+phrase+'"'
r = requests.get(url, headers=headers)
r.raise_for_status()
html = r.text
# Encoding problems
# http = urllib3.PoolManager()
# r = http.request('GET', url)
# print(r.status)
# html=r.data
# Encoding problems
# with urllib.request.urlopen(url) as f:
# html = f.read()
soup = BeautifulSoup(html, 'html.parser')
#span that says there is no such phrase
span = soup.find('div', class_="misspell__message")
if span:
print (span.get_text())
# div with stats
div = soup.find('div', class_="serp-adv__found")
# remove all exept digits from the inner text
if div:
t=div.get_text()
print(t)
num = re.sub( r'[^\d]', '', t)
return (int(num) if num else 0)
else:
return -1
print(get_frequency_y("hello"))
print(get_frequency("hello"))
# +
? %pdb
| APIS/bigrams/.ipynb_checkpoints/bigrams-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.8 64-bit
# name: python36864bit277cdecdadcc4196a67241552660cd03
# ---
import numpy as np
from itertools import permutations
from collections import defaultdict
import random
# # load and parse dataset
# + tags=[]
# !file umls -I
# +
raw_data = []
entities = set()
with open('umls', 'r') as to_read:
for i, line in enumerate(to_read.readlines()):
s, p, o = line.strip().split(' ')
entities.add(s)
entities.add(o)
raw_data += [(s,p,o)]
# +
A_implies_B_rules = [
# body: head
('Process_of', 'Affects')
]
transitive_rules = ['Isa', 'Interacts_with']
#java -jar /Users/simon/Office/Dokumente/Uni/Data\ Science\ and\ Machine\ Learning\ Master/Masters\ Project/Libraries/amie-dev.jar -d " " -minc 1 -minpca 1 -mins 300 -maxad 3 umls
#java -jar /Users/simon/Office/Dokumente/Uni/Data\ Science\ and\ Machine\ Learning\ Master/Masters\ Project/Libraries/amie-dev.jar -d " " -minc 1 -minpca 1 -mins 100 -maxad 2 umls
# + tags=[]
train = set()
valid = set()
test = set()
entities = set()
A_implies_B_rule_examples = defaultdict(lambda: [])
transitive_rule_examples = defaultdict(lambda: [])
counter_A_implies_B_rules = defaultdict(lambda: 0)
counter_transitive_rules = defaultdict(lambda: 0)
for s,p,o in raw_data:
entities.add(s)
entities.add(o)
for x1, x2 in permutations(entities, 2):
for (A, B) in A_implies_B_rules:
if (x1, A, x2) in raw_data and (x1, B, x2) in raw_data:
valid.add((x1, B, x2))
A_implies_B_rule_examples[(A, B)] += [(x1, x2)]
counter_A_implies_B_rules[(A, B)] += 1
triples = [triple for triple in raw_data if triple[1] in transitive_rules]
for (x1, t, x2) in triples:
for x3 in entities:
if (x1, t, x2) in raw_data and (x2, t, x3) in raw_data and (x1, t, x3) in raw_data:
valid.add((x1, C, x3))
A_B_implies_C_rule_examples[(A, B, C)] += [(x1, x2, x3)]
counter_A_B_implies_C_rules[(A, B, C)] += 1
for s,p,o in raw_data:
if (s,p,o) not in valid:
train.add((s,p,o))
# -
train = list(train)
valid = list(valid)
random.Random(42).shuffle(valid)
valid, test = valid[:len(valid) // 2], valid[len(valid) // 2:]
# + tags=[]
print(len(train))
print(len(valid))
print(len(test))
# -
# # check that splits are mutually exclusive
for triple in train:
if triple in valid:
print("valid", triple)
if triple in test:
print("valid", triple)
for triple in valid:
if triple in train:
print("train", triple)
if triple in test:
print("test", triple)
for triple in test:
if triple in train:
print("train", triple)
if triple in valid:
print("valid", triple)
# # save splits as .tsv
with open("train.tsv", "w", encoding='utf-8') as f:
for triple in train:
f.write("{}\t{}\t{}\n".format(*triple))
with open("valid.tsv", "w", encoding='utf-8') as f:
for triple in valid:
f.write("{}\t{}\t{}\n".format(*triple))
with open("test.tsv", "w", encoding='utf-8') as f:
for triple in test:
f.write("{}\t{}\t{}\n".format(*triple))
# +
for (A, B), examples in A_implies_B_rule_examples.items():
with open("{}=>{}.tsv".format(A,B), "w", encoding='utf-8') as f:
for (x1, x2) in examples:
f.write("{}\t{}\t{}\n".format(x1, B, x2))
for (A, B, C), examples in A_B_implies_C_rule_examples.items():
with open("{},{}=>{}.tsv".format(A,B,C), "w", encoding='utf-8') as f:
for (x1, x2, x3) in examples:
f.write("{}\t{}\t{}\n".format(x1, C, x3))
| metakbc/data/umls/generate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Method to determine outliers
# ===============================
#
# The data we use in this example are publically available microarray data.
#
# * [NCBI GEO - GSE18965](http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE18965)
#
# The basic idea here is to summarize a data set with a large number of features in lower dimensional space. Then using that lower dimensional projection determine in a systematic way if there are outlier samples.
#
# The data are atopic-asthmatic (AA) subjects and healthy-nonasthmatic (HN) controls.
#
# Load data and add some outliers
# ----------------------------------
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
from sklearn import preprocessing
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.covariance import EllipticEnvelope
mat = np.loadtxt("gse18965.txt")
covs = pd.read_csv("gse18965-targets.csv",usecols=[0,1,2,4,5])
print(covs)
covsAdded = pd.DataFrame([["X","OT",999,11.1,'M'],["Y","OT",998,12.5,"F"]],
columns=['sample','phenotype','subject','age','gender'])
covs = covs.append(covsAdded,ignore_index=True)
print(covs)
print mat.shape
print mat.mean(axis=0).shape, mat.mean(axis=0).mean()
mat = np.vstack([mat,mat.mean(axis=0)*1.5])
mat = np.vstack([mat,mat.mean(axis=0)*1.6])
print mat.shape
# Run both PCA and tSNE
# ------------------------
# +
from IPython.display import Image
matScaled = preprocessing.scale(mat)
fit1 = PCA(n_components=2).fit_transform(mat)
fit2 = TSNE(learning_rate=100,perplexity=10,n_iter=2000).fit_transform(mat)
fit3 = PCA(n_components=2).fit_transform(matScaled)
fit4 = TSNE(learning_rate=100,perplexity=10,n_iter=2000).fit_transform(matScaled)
def make_subplot(fit,covs,covariate,ax,pcX=0,pcY=1,fontSize=10,fontName='sans serif',ms=20,leg=True,title=None):
colors = ['k','cyan','r','orange','g','b','magenta']
cvNames = np.sort(np.unique(covs[covariate]))
lines = []
for _i,i in enumerate(cvNames):
indices = np.where(covs[covariate]==i)[0]
s = ax.scatter(fit[indices,pcX],fit[indices,pcY],c=colors[_i],s=ms,label=covariate,alpha=0.9)
lines.append(s)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontSize-2)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontSize-2)
buff = 0.02
bufferX = buff * (fit[:,pcX].max() - fit[:,pcX].min())
bufferY = buff * (fit[:,pcY].max() - fit[:,pcY].min())
ax.set_xlim([fit[:,pcX].min()-bufferX,fit[:,pcX].max()+bufferX])
ax.set_ylim([fit[:,pcY].min()-bufferY,fit[:,pcY].max()+bufferY])
ax.set_xlabel("D-%s"%str(pcX+1),fontsize=fontSize,fontname=fontName)
ax.set_ylabel("D-%s"%str(pcY+1),fontsize=fontSize,fontname=fontName)
plt.locator_params(axis='x',nbins=5)
ax.set_aspect(1./ax.get_data_ratio())
if title:
ax.set_title(title,fontsize=fontSize+2,fontname=fontName)
if leg:
legend = ax.legend(lines,cvNames,loc='upper right',scatterpoints=1,
handletextpad=0.01,labelspacing=0.01,borderpad=0.1,handlelength=1.0)
for label in legend.get_texts():
label.set_fontsize(fontSize-2)
label.set_fontname(fontName)
plt.clf()
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax2 = fig.add_subplot(222)
ax3 = fig.add_subplot(223)
ax4 = fig.add_subplot(224)
make_subplot(fit1,covs,'phenotype',ax1,pcX=0,pcY=1,leg=True,title='PCA-raw')
make_subplot(fit2,covs,'phenotype',ax2,pcX=0,pcY=1,leg=False,title='tSNE-raw')
make_subplot(fit3,covs,'phenotype',ax3,pcX=0,pcY=1,leg=False,title='PCA-scaled')
make_subplot(fit4,covs,'phenotype',ax4,pcX=0,pcY=1,leg=False,title='tSNE-scaled')
ax2.set_xlabel("")
ax4.set_xlabel("")
plt.subplots_adjust(hspace=0.3,wspace=0.05)
plt.savefig("outliers-projection.png",dpi=600)
# -
Image(filename='outliers-projection.png')
# Because we are not that interested in relative fold change differences, i.e. we do not want the scale of expression differences to dominate the projection we use a standarization. Based on these plot it appears that PCA does a better job than tSNE of separating outliers. Depending on how far we put the outliers it is often the case that PCA does better when scaled, but tSNE performs better when using the original data.
# Run the outlier algorithm
# ---------------------------
#
# This comes from the [scikit-learn example](http://scikit-learn.org/stable/auto_examples/covariance/plot_outlier_detection.html#example-covariance-plot-outlier-detection-py)
#
# +
from IPython.display import Image
outliers_fraction = 0.15
classifiers = {"robust covariance estimator": EllipticEnvelope(contamination=outliers_fraction)}
def make_subplot_again(X,covs,covariate,ax,pcX=0,pcY=1,fontSize=10,fontName='sans serif',ms=20,leg=True,title=None):
## variables
colors = ['k','cyan','r','orange','g','b','magenta']
clf_name = "robust covariance estimator"
clf = EllipticEnvelope(contamination=.1)
X =preprocessing.scale(X.copy())
## figure out scale
buff = 0.02
bufferX = buff * (X[:,pcX].max() - X[:,pcX].min())
bufferY = buff * (X[:,pcY].max() - X[:,pcY].min())
mm = [(X[:,pcX].min()-bufferX,X[:,pcX].max()+bufferX),(X[:,pcY].min()-bufferY,X[:,pcY].max()+bufferY)]
xx, yy = np.meshgrid(np.linspace(mm[0][0],mm[0][1], 500), np.linspace(mm[1][0],mm[1][1],500))
# fit the data and tag outliers
clf.fit(X)
y_pred = clf.decision_function(X).ravel()
threshold = stats.scoreatpercentile(y_pred,100 * outliers_fraction)
y_pred = y_pred > threshold
print y_pred
# plot the levels lines and the points
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
ax.contourf(xx, yy, Z, levels=np.linspace(Z.min(), threshold, 7),cmap=plt.cm.Blues_r)
a = ax.contour(xx, yy, Z, levels=[threshold],linewidths=2, colors='red')
ax.contourf(xx, yy, Z, levels=[threshold, Z.max()],colors='orange')
ax.axis('tight')
cvNames = np.sort(np.unique(covs[covariate]))
lines = []
for _i,i in enumerate(cvNames):
indices = np.where(covs[covariate]==i)[0]
s = ax.scatter(X[indices,pcX],X[indices,pcY],c=colors[_i],s=ms,label=covariate,alpha=0.9)
lines.append(s)
## axes
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontSize-2)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontSize-2)
ax.set_xlabel("D-%s"%str(pcX+1),fontsize=fontSize,fontname=fontName)
ax.set_ylabel("D-%s"%str(pcY+1),fontsize=fontSize,fontname=fontName)
plt.locator_params(axis='x',nbins=5)
ax.set_aspect(1./ax.get_data_ratio())
ax.set_xlim(mm[0])
ax.set_ylim(mm[1])
if title:
ax.set_title(title,fontsize=fontSize+2,fontname=fontName)
if leg:
legend = ax.legend(lines,cvNames,loc='upper right',scatterpoints=1,
handletextpad=0.01,labelspacing=0.01,borderpad=0.1,handlelength=1.0)
for label in legend.get_texts():
label.set_fontsize(fontSize-2)
label.set_fontname(fontName)
## make the figure again
plt.clf()
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax2 = fig.add_subplot(222)
ax3 = fig.add_subplot(223)
ax4 = fig.add_subplot(224)
make_subplot_again(fit1,covs,'phenotype',ax1,pcX=0,pcY=1,leg=True,title='PCA-raw')
make_subplot_again(fit2,covs,'phenotype',ax2,pcX=0,pcY=1,leg=False,title='tSNE-raw')
make_subplot_again(fit3,covs,'phenotype',ax3,pcX=0,pcY=1,leg=False,title='PCA-scaled')
make_subplot_again(fit4,covs,'phenotype',ax4,pcX=0,pcY=1,leg=False,title='tSNE-scaled')
ax1.set_xlabel("")
ax2.set_xlabel("")
ax2.set_ylabel("")
ax4.set_ylabel("")
plt.subplots_adjust(hspace=0.3,wspace=0.05)
plt.savefig("outliers-detection.png",dpi=600)
# -
Image(filename='outliers-detection.png')
# Discussion
# ------------------------
# The method appears to work fairly well. Care needs to be taken on which project method and whether or not the results need to be scaled, but for the most part the results are the same.
| archive/munging/outlier-detection/projection_outliers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Core Idea
#
# Despite a lot of creeping Physics and Chemistry knowledge introduced in the description, this competition is more about Geometry and pattern matching.
#
# The hypothesis of this kernel is next:
# 1. If we have two similar sets of atoms with the same distances between them and the same types - the scalar coupling constant should be very close.
# 2. More closest atoms to the pair of atoms under prediction have higher influence on scalar coupling constant then those with higher distance
#
# So, basically, this problem could be dealt with some kind of K-Nearest Neighbor algorithm or any tree-based - e.g. LightGBM, in case we can find some representation which would describe similar configurations with similar feature sets.
#
# Each atom is described with 3 cartesian coordinates. This representation is not stable. Each coupling pair is located in a different point in space and two similar coupling sets would have very different X,Y,Z.
#
# So, instead of using coordinates let's consider next system:
# 1. Take each pair of atoms as two first core atoms
# 2. Calculate the center between the pair
# 3. Find all n-nearest atoms to the center (excluding first two atoms)
# 4. Take two closest atoms from step 3 - they will be 3rd and 4th core atoms
# 5. Calculate the distances from 4 core atoms to the rest of the atoms and to the core atoms as well
#
# Using this representation each atom position can be described by 4 distances from the core atoms. This representation is stable to rotation and translation. And it's suitable for pattern-matching. So, we can take a sequence of atoms, describe each by 4 distances + atom type(H,O,etc) and looking up for the same pattern we can find similar configurations and detect scalar coupling constant.
#
# Here I used LightGBM, because sklearn KNN can't deal with the amount of data. My blind guess is that hand-crafted KNN can outperform LightGBM.
#
# Let's code the solution!
# ## Load Everything
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import math
import gc
import copy
from sklearn.model_selection import KFold, train_test_split
from sklearn.metrics import mean_absolute_error
import matplotlib.pyplot as plt
import seaborn as sns
from lightgbm import LGBMRegressor
# -
DATA_PATH = '../input'
SUBMISSIONS_PATH = './'
# use atomic numbers to recode atomic names
ATOMIC_NUMBERS = {
'H': 1,
'C': 6,
'N': 7,
'O': 8,
'F': 9
}
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.max_rows', 120)
pd.set_option('display.max_columns', 120)
# ## Load Dataset
# By default all data is read as `float64` and `int64`. We can trade this uneeded precision for memory and higher prediction speed. So, let's read with Pandas all the data in the minimal representation:
train_dtypes = {
'molecule_name': 'category',
'atom_index_0': 'int8',
'atom_index_1': 'int8',
'type': 'category',
'scalar_coupling_constant': 'float32'
}
train_csv = pd.read_csv(f'{DATA_PATH}/train.csv', index_col='id', dtype=train_dtypes)
train_csv['molecule_index'] = train_csv.molecule_name.str.replace('dsgdb9nsd_', '').astype('int32')
train_csv = train_csv[['molecule_index', 'atom_index_0', 'atom_index_1', 'type', 'scalar_coupling_constant']]
train_csv.head(10)
print('Shape: ', train_csv.shape)
print('Total: ', train_csv.memory_usage().sum())
train_csv.memory_usage()
submission_csv = pd.read_csv(f'{DATA_PATH}/sample_submission.csv', index_col='id')
test_csv = pd.read_csv(f'{DATA_PATH}/test.csv', index_col='id', dtype=train_dtypes)
test_csv['molecule_index'] = test_csv['molecule_name'].str.replace('dsgdb9nsd_', '').astype('int32')
test_csv = test_csv[['molecule_index', 'atom_index_0', 'atom_index_1', 'type']]
test_csv.head(10)
structures_dtypes = {
'molecule_name': 'category',
'atom_index': 'int8',
'atom': 'category',
'x': 'float32',
'y': 'float32',
'z': 'float32'
}
structures_csv = pd.read_csv(f'{DATA_PATH}/structures.csv', dtype=structures_dtypes)
structures_csv['molecule_index'] = structures_csv.molecule_name.str.replace('dsgdb9nsd_', '').astype('int32')
structures_csv = structures_csv[['molecule_index', 'atom_index', 'atom', 'x', 'y', 'z']]
structures_csv['atom'] = structures_csv['atom'].replace(ATOMIC_NUMBERS).astype('int8')
structures_csv.head(10)
print('Shape: ', structures_csv.shape)
print('Total: ', structures_csv.memory_usage().sum())
structures_csv.memory_usage()
# ## Build Distance Dataset
def build_type_dataframes(base, structures, coupling_type):
base = base[base['type'] == coupling_type].drop('type', axis=1).copy()
base = base.reset_index()
base['id'] = base['id'].astype('int32')
structures = structures[structures['molecule_index'].isin(base['molecule_index'])]
return base, structures
def add_coordinates(base, structures, index):
df = pd.merge(base, structures, how='inner',
left_on=['molecule_index', f'atom_index_{index}'],
right_on=['molecule_index', 'atom_index']).drop(['atom_index'], axis=1)
df = df.rename(columns={
'atom': f'atom_{index}',
'x': f'x_{index}',
'y': f'y_{index}',
'z': f'z_{index}'
})
return df
def add_atoms(base, atoms):
df = pd.merge(base, atoms, how='inner',
on=['molecule_index', 'atom_index_0', 'atom_index_1'])
return df
def merge_all_atoms(base, structures):
df = pd.merge(base, structures, how='left',
left_on=['molecule_index'],
right_on=['molecule_index'])
df = df[(df.atom_index_0 != df.atom_index) & (df.atom_index_1 != df.atom_index)]
return df
# +
def add_center(df):
df['x_c'] = ((df['x_1'] + df['x_0']) * np.float32(0.5))
df['y_c'] = ((df['y_1'] + df['y_0']) * np.float32(0.5))
df['z_c'] = ((df['z_1'] + df['z_0']) * np.float32(0.5))
def add_distance_to_center(df):
df['d_c'] = ((
(df['x_c'] - df['x'])**np.float32(2) +
(df['y_c'] - df['y'])**np.float32(2) +
(df['z_c'] - df['z'])**np.float32(2)
)**np.float32(0.5))
def add_distance_between(df, suffix1, suffix2):
df[f'd_{suffix1}_{suffix2}'] = ((
(df[f'x_{suffix1}'] - df[f'x_{suffix2}'])**np.float32(2) +
(df[f'y_{suffix1}'] - df[f'y_{suffix2}'])**np.float32(2) +
(df[f'z_{suffix1}'] - df[f'z_{suffix2}'])**np.float32(2)
)**np.float32(0.5))
# -
def add_distances(df):
n_atoms = 1 + max([int(c.split('_')[1]) for c in df.columns if c.startswith('x_')])
for i in range(1, n_atoms):
for vi in range(min(4, i)):
add_distance_between(df, i, vi)
def add_n_atoms(base, structures):
dfs = structures['molecule_index'].value_counts().rename('n_atoms').to_frame()
return pd.merge(base, dfs, left_on='molecule_index', right_index=True)
def build_couple_dataframe(some_csv, structures_csv, coupling_type, n_atoms=10):
base, structures = build_type_dataframes(some_csv, structures_csv, coupling_type)
base = add_coordinates(base, structures, 0)
base = add_coordinates(base, structures, 1)
base = base.drop(['atom_0', 'atom_1'], axis=1)
atoms = base.drop('id', axis=1).copy()
if 'scalar_coupling_constant' in some_csv:
atoms = atoms.drop(['scalar_coupling_constant'], axis=1)
add_center(atoms)
atoms = atoms.drop(['x_0', 'y_0', 'z_0', 'x_1', 'y_1', 'z_1'], axis=1)
atoms = merge_all_atoms(atoms, structures)
add_distance_to_center(atoms)
atoms = atoms.drop(['x_c', 'y_c', 'z_c', 'atom_index'], axis=1)
atoms.sort_values(['molecule_index', 'atom_index_0', 'atom_index_1', 'd_c'], inplace=True)
atom_groups = atoms.groupby(['molecule_index', 'atom_index_0', 'atom_index_1'])
atoms['num'] = atom_groups.cumcount() + 2
atoms = atoms.drop(['d_c'], axis=1)
atoms = atoms[atoms['num'] < n_atoms]
atoms = atoms.set_index(['molecule_index', 'atom_index_0', 'atom_index_1', 'num']).unstack()
atoms.columns = [f'{col[0]}_{col[1]}' for col in atoms.columns]
atoms = atoms.reset_index()
# downcast back to int8
for col in atoms.columns:
if col.startswith('atom_'):
atoms[col] = atoms[col].fillna(0).astype('int8')
atoms['molecule_index'] = atoms['molecule_index'].astype('int32')
full = add_atoms(base, atoms)
add_distances(full)
full.sort_values('id', inplace=True)
return full
def take_n_atoms(df, n_atoms, four_start=4):
labels = []
for i in range(2, n_atoms):
label = f'atom_{i}'
labels.append(label)
for i in range(n_atoms):
num = min(i, 4) if i < four_start else 4
for j in range(num):
labels.append(f'd_{i}_{j}')
if 'scalar_coupling_constant' in df:
labels.append('scalar_coupling_constant')
return df[labels]
# ## Check LightGBM with the smallest type
# %%time
full = build_couple_dataframe(train_csv, structures_csv, '1JHN', n_atoms=10)
print(full.shape)
# We don't calculate distances for `d_0_x`, `d_1_1`, `d_2_2`, `d_2_3`, `d_3_3` because we already have them in later atoms(`d_0_1` == `d_1_0`) or they are equal to zeros(e.g. `d_1_1`, `d_2_2`).
full.columns
# For experiments, full dataset can be built with higher number of atoms, and for building a training/validation sets we can trim them:
df = take_n_atoms(full, 7)
# LightGBM performs better with 0-s then with NaN-s
df = df.fillna(0)
df.columns
# +
X_data = df.drop(['scalar_coupling_constant'], axis=1).values.astype('float32')
y_data = df['scalar_coupling_constant'].values.astype('float32')
X_train, X_val, y_train, y_val = train_test_split(X_data, y_data, test_size=0.2, random_state=128)
X_train.shape, X_val.shape, y_train.shape, y_val.shape
# -
# configuration params are copied from @artgor kernel:
# https://www.kaggle.com/artgor/brute-force-feature-engineering
LGB_PARAMS = {
'objective': 'regression',
'metric': 'mae',
'verbosity': -1,
'boosting_type': 'gbdt',
'learning_rate': 0.2,
'num_leaves': 128,
'min_child_samples': 79,
'max_depth': 9,
'subsample_freq': 1,
'subsample': 0.9,
'bagging_seed': 11,
'reg_alpha': 0.1,
'reg_lambda': 0.3,
'colsample_bytree': 1.0
}
# +
model = LGBMRegressor(**LGB_PARAMS, n_estimators=1500, n_jobs = -1)
model.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_val, y_val)], eval_metric='mae',
verbose=100, early_stopping_rounds=200)
y_pred = model.predict(X_val)
np.log(mean_absolute_error(y_val, y_pred))
# -
# Not a bad score for such a simple set of features.
cols = list(df.columns)
cols.remove('scalar_coupling_constant')
cols
df_importance = pd.DataFrame({'feature': cols, 'importance': model.feature_importances_})
sns.barplot(x="importance", y="feature", data=df_importance.sort_values('importance', ascending=False));
# It's funny, but looks like atom types aren't used a lot in the final decision. Quite a contrary to what a man would do.
# ## Submission Model
def build_x_y_data(some_csv, coupling_type, n_atoms):
full = build_couple_dataframe(some_csv, structures_csv, coupling_type, n_atoms=n_atoms)
df = take_n_atoms(full, n_atoms)
df = df.fillna(0)
print(df.columns)
if 'scalar_coupling_constant' in df:
X_data = df.drop(['scalar_coupling_constant'], axis=1).values.astype('float32')
y_data = df['scalar_coupling_constant'].values.astype('float32')
else:
X_data = df.values.astype('float32')
y_data = None
return X_data, y_data
def train_and_predict_for_one_coupling_type(coupling_type, submission, n_atoms, n_folds=5, n_splits=5, random_state=128):
print(f'*** Training Model for {coupling_type} ***')
X_data, y_data = build_x_y_data(train_csv, coupling_type, n_atoms)
X_test, _ = build_x_y_data(test_csv, coupling_type, n_atoms)
y_pred = np.zeros(X_test.shape[0], dtype='float32')
cv_score = 0
if n_folds > n_splits:
n_splits = n_folds
kfold = KFold(n_splits=n_splits, shuffle=True, random_state=random_state)
for fold, (train_index, val_index) in enumerate(kfold.split(X_data, y_data)):
if fold >= n_folds:
break
X_train, X_val = X_data[train_index], X_data[val_index]
y_train, y_val = y_data[train_index], y_data[val_index]
model = LGBMRegressor(**LGB_PARAMS, n_estimators=1500, n_jobs = -1)
model.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_val, y_val)], eval_metric='mae',
verbose=100, early_stopping_rounds=200)
y_val_pred = model.predict(X_val)
val_score = np.log(mean_absolute_error(y_val, y_val_pred))
print(f'{coupling_type} Fold {fold}, logMAE: {val_score}')
cv_score += val_score / n_folds
y_pred += model.predict(X_test) / n_folds
submission.loc[test_csv['type'] == coupling_type, 'scalar_coupling_constant'] = y_pred
return cv_score
# Let's build a separate model for each type of coupling. Dataset is split into 5 pieces and in this kernel we will use only 3 folds for speed up.
#
# Main tuning parameter is the number of atoms. I took good numbers, but accuracy can be improved a bit by tuning them for each type.
# +
model_params = {
'1JHN': 7,
'1JHC': 10,
'2JHH': 9,
'2JHN': 9,
'2JHC': 9,
'3JHH': 9,
'3JHC': 10,
'3JHN': 10
}
N_FOLDS = 3
submission = submission_csv.copy()
cv_scores = {}
for coupling_type in model_params.keys():
cv_score = train_and_predict_for_one_coupling_type(
coupling_type, submission, n_atoms=model_params[coupling_type], n_folds=N_FOLDS)
cv_scores[coupling_type] = cv_score
# -
# Checking cross-validation scores for each type:
pd.DataFrame({'type': list(cv_scores.keys()), 'cv_score': list(cv_scores.values())})
# And cv mean score:
np.mean(list(cv_scores.values()))
# Sanity check for all cells to be filled with predictions:
submission[submission['scalar_coupling_constant'] == 0].shape
submission.head(10)
submission.to_csv(f'{SUBMISSIONS_PATH}/submission.csv')
# ## Room for improvement
# There are many steps, how to improve the score for this kernel:
# * Tune LGB hyperparameters - I did nothing for this
# * Tune number of atoms for each type
# * Try to add other features
# * Play with categorical features for atom types (one-hot-encoding, CatBoost?)
# * Try other tree libraries
#
# Also, this representation fails badly on `*JHC` coupling types. The main reason for this is that 3rd and 4th atoms are usually located on the same distance and representation starts "jittering" randomly picking one of them. So, two similar configurations will have different representation due to usage of 3/4 of 4/3 distances.
#
# The biggest challenge would be to implement handcrafted KNN with some compiled language(Rust, C++, C).
#
# Would be cool to see this kernel forked and addressed some of the issues with higher LB score.
| download_code/distance-is-all-you-need-lb-1-481.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 2: Sorting
# %matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
from random import shuffle
# ## Insertion Sort
# This implements the most naive version of insertion sort, following the logic: "insert the thing where it needs to go."
#
# It makes a blank list, and inserts elements one-at-a-time.
#
# This is not a very good way to implement insertion sort.
def naiveInsertionSort(A):
B = [None for i in range(len(A))] # B is a blank list of the same length as A
for x in A:
for i in range(len(B)):
if B[i] == None or B[i] > x:
# then x goes in spot i, and we should move everything over.
j = len(B)-1
while j > i:
B[j] = B[j-1]
j -= 1
B[i] = x
break # okay we are done placing x
return B
# Let's sanity-check:
A = [6,4,3,8,5]
B = naiveInsertionSort(A)
print(B)
# The code above faithfully captures the notion of "insertion sort" that we first described on the slides: insert the elements into a new list, one-at-a-time, where they should go. But we can be a bit more slick about it, and do it in-place. The following implements the description of insertion sort in-place, which we gave a demonstration of in the slides.
def InsertionSort(A):
for i in range(1,len(A)):
current = A[i]
j = i-1
while j >= 0 and A[j] > current:
A[j+1] = A[j]
j -= 1
A[j+1] = current
# Let's sanity-check:
A = [6,4,3,8,5]
InsertionSort(A)
print(A)
# Let's do it a bunch more times to make sure! (Note: this is *not* a formal proof of correctness!)
# +
def is_sorted(A):
for i in range(len(A) - 1):
if A[i] > A[i+1]:
return False
return True
A = [1,2,3,4,5,6,7,8,9,10]
for trial in range(100):
shuffle(A)
InsertionSort(A)
if is_sorted(A):
print('YES IT IS SORTED!')
# -
# Okay, now we have two working algorithms. Which one is faster?
from tryItABunch import tryItABunch
nValuesNaive, tValuesNaive = tryItABunch( naiveInsertionSort, startN = 50, endN = 1050, stepSize=50, numTrials=10, listMax = 10 )
nValues, tValues = tryItABunch( InsertionSort, startN = 50, endN = 1050, stepSize=50, numTrials=5, listMax = 10)
plt.plot(nValuesNaive, tValuesNaive, color="red", label="Naive version")
plt.plot(nValues, tValues, color="blue", label="Less naive version")
plt.xlabel("n")
plt.ylabel("Time(ms)")
plt.legend()
plt.title("Naive vs. non-naive insertion sort")
# Well, that does agree with our intuition that the naive version should take longer. But it looks like the performance is getting worse and worse! Instead of growing roughly linearly with the size of the input, it's growing quadratically!
# ## MergeSort
#
# Okay, so InsertionSort was slow. Can we do better?
# +
# merge two sorted arrays to return a sorted array
def merge(L, R):
i = 0 # current index in the L array
j = 0 # current index in the R array
ret = []
while i < len(L) and j < len(R):
if L[i] < R[j]:
ret.append(L[i])
i += 1
else: # ties go to R. Doesn't really matter.
ret.append(R[j])
j += 1
while i < len(L):
ret.append(L[i])
i += 1
while j < len(R):
ret.append(R[j])
j+= 1
return ret
def mergeSort(A):
n = len(A)
if n <= 1:
return A
L = mergeSort(A[:round(n/2)])
R = mergeSort(A[round(n/2):n])
return merge(L,R)
# -
# Let's sanity-check:
A = [27,17,2,4,52,3,4,3]
B = mergeSort(A)
print(B)
nValuesMerge, tValuesMerge = tryItABunch( mergeSort, startN = 50, endN = 1050, stepSize=50, numTrials=10, listMax = 10)
# First we can take a look at how that fits in with what we saw before
plt.plot(nValuesNaive, tValuesNaive, color="red", label="Naive version of insertion sort")
plt.plot(nValues, tValues, color="blue", label="Less naive version of insertion sort")
plt.plot(nValuesMerge, tValuesMerge, color="orange", label="Not very slick implementation of mergeSort")
plt.xlabel("n")
plt.ylabel("Time(ms)")
plt.legend()
plt.title("All sorts of sorts")
# + active=""
# For reference, this is how the theoretical running time looks:
# -
plt.plot(nValuesMerge, [x**2 for x in nValuesMerge], color="green", label="n^2")
plt.plot(nValuesMerge, [x*np.log(x) for x in nValuesMerge], "--", color="orange", label="nlog(n)")
plt.plot(nValuesMerge, [11*x*np.log(x) for x in nValuesMerge], "--", color="red", label="11*nlog(n)")
plt.plot(nValuesMerge, [100*x*np.log(x) for x in nValuesMerge], "--", color="purple", label="100*nlog(n)")
plt.plot(nValuesMerge, [200*x*np.log(x) for x in nValuesMerge], "--", color="blue", label="200*nlog(n)")
plt.xlabel("n")
plt.ylim(0,800000)
plt.legend()
plt.title("n^2 vs nlog(n)")
| notebooks/lecture2_sorting/lecture2_sorting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Correlating neutrino data from different experiments
# +
import pyvo as vo
import matplotlib.pyplot as plt
import numpy as np
#retrieve the full Antares test dataset
service_km3net = vo.dal.TAPService("http://vo.km3net.de/__system__/tap/run/tap")
dataset_antares = service_km3net.search("SELECT * FROM ant20_01.main")
# retrieve the full icecube dataset
service_icecube = vo.dal.TAPService("https://dc.zah.uni-heidelberg.de/__system__/tap/run/tap")
dataset_icecube = service_icecube.search("SELECT * FROM icecube.nucand")
# +
# superimpose the RA vs Declination scatter plots for both Antares & IC data
ra_antares = []
decl_antares = []
i=0
while i < len(dataset_antares):
for row in dataset_antares:
ra_antares.append(dataset_antares['ra', i ])
decl_antares.append(dataset_antares['decl', i ])
i=i+1
ra_icecube = []
decl_icecube = []
i=0
while i < len(dataset_icecube):
for row in dataset_icecube:
ra_icecube.append(dataset_icecube['nualpha', i ])
decl_icecube.append(dataset_icecube['nudelta', i ])
i=i+1
plt.scatter( ra_antares , decl_antares , s=3 , label="Antares" , c="red" )
plt.scatter( ra_icecube , decl_icecube , s=3 , label="IC-40" , c="cyan" )
plt.xlabel("Right Ascension")
plt.ylabel("Declination")
plt.legend()
plt.show()
# +
# superimpose the distributions of the number of hits for both datasets
nhits_antares = []
i=0
while i < len(dataset_antares):
for row in dataset_antares:
nhits = dataset_antares['nhit', i ]
i=i+1
nhits_antares.append(nhits)
nhits_icecube = []
i=0
while i < len(dataset_icecube):
for row in dataset_icecube:
nhits = dataset_icecube['nch', i ]
i=i+1
nhits_icecube.append(nhits)
plt.hist( nhits_antares , bins=50 , range=[0,200] , label = "Antares" , fill=False , color = "red" , histtype = 'step' )
plt.hist( nhits_icecube , bins=50 , range=[0,200] , label = "IC-40" , fill=False , color = "cyan" , histtype = 'step' )
plt.legend()
plt.show()
| neutrinos/pyvo_multiple_experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Check CCM against HGHI
#
# This notebook is for comparing the numbers of CCM and HGHI.
#
# **Note:** You must run the `processing/Merge_HGHI_and_CCM_data.ipynb` notebook to produce
# the merged datasets to compare.
# +
import pandas as pd
import geopandas as gpd
from covidcaremap.data import published_data_path, processed_data_path
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', 100)
# -
state_path = processed_data_path('hghi_ccm_data_by_state.geojson')
state_gdf = gpd.read_file(state_path, encoding='utf-8')
state_gdf[state_gdf['State'] == 'PA'].to_dict(orient='record')
compare_cols =[
'State',
'Licensed All Beds',
'HGHI - Total Hospital Beds',
'Staffed All Beds',
'Staffed ICU Beds',
'HGHI - Total ICU Beds',
'All Bed Occupancy Rate',
'HGHI - Hospital Bed Occupancy Rate',
'ICU Bed Occupancy Rate',
'HGHI - ICU Bed Occupancy Rate'
]
state_gdf[compare_cols].to_csv('hghi-ccm-compare.csv')
hrr_path = processed_data_path('hghi_ccm_data_by_hrr.geojson')
hrr_gdf = gpd.read_file(hrr_path, encoding='utf-8')
hrr_gdf
compare_cols_HRR =[
'HRRCITY',
'Licensed All Beds',
'Staffed All Beds',
'HGHI - Total Hospital Beds',
'All Bed Occupancy Rate',
'HGHI - Available Hospital Beds',
'Staffed ICU Beds',
'ICU Bed Occupancy Rate',
'HGHI - Available ICU Beds',
'Staffed ICU Beds',
'HGHI - Total ICU Beds',
'Population',
'Population (20+)',
'Population (65+)',
'HGHI - Adult Population',
'HGHI - Population 65+',
]
hrr_gdf[compare_cols_HRR].to_csv(processed_data_path('hghi-ccm-compare-hrr.csv'))
# +
# TODO - Quantify variance between HGHI and CCM data.
| notebooks/validation/Check_CCM_against_HGHI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # WikiNetworking Stallion Demo
#
# ### Introduction
#
# This notebook creates both interactive and high resolution graphs of social networks from Wikipedia articles. Several demonstration data sets are included.
#
# ### Getting started
#
# Run the cell below first. It will install the necessary packages and define a helper function and some variables for sample data URLs.
# +
# !pip install git+https://github.com/jchuahtacc/WikiNetworking.git
# Just in case we don't want to re-run the crawl, we will load the data directly
import wikinetworking as wn
import networkx as nx
import matplotlib.pyplot as plt
import urllib2
import json
# %matplotlib inline
bet_hiphop_directed = "https://raw.githubusercontent.com/jchuahtacc/WikiNetworking/master/lessons/bet_directed.json"
bet_hiphop_undirected = "https://raw.githubusercontent.com/jchuahtacc/WikiNetworking/master/lessons/bet_undirected.json"
forbes_400 = "https://raw.githubusercontent.com/jchuahtacc/WikiNetworking/master/lessons/forbes400.json"
nba_allstars = "https://raw.githubusercontent.com/jchuahtacc/WikiNetworking/master/lessons/nba_allstars.json"
nfl_most_games = "https://raw.githubusercontent.com/jchuahtacc/WikiNetworking/master/lessons/nfl_players.json"
marvel_cinematic_universe = "https://raw.githubusercontent.com/jchuahtacc/WikiNetworking/master/lessons/mcu_network.json"
def make_graph(url, minimum_weight=2):
graph_data = json.loads(urllib2.urlopen(url).read())
return wn.create_graph(graph_data, minimum_weight=minimum_weight)
# -
# ### Creating a graph and a layout
#
# The `make_graph` function loads a URL that contains our graph data and creates a `networkx` graph. You may optionally specify a `minimum_weight` for links between nodes to be registered on our graph. Once we have the graph, we also need to use a layout algorithm to generate the position of the nodes. Possible layouts include:
#
# - `circular_layout`
# - `random_layout`
# - `shell_layout`
# - `spring_layout`
# - `spectral_layout`
# - `fruchterman_reingold_layout`
# +
# Make a graph object (optionally, specify minimum_weight)
graph = make_graph(marvel_cinematic_universe, minimum_weight=3)
# Generate a layout object
layout = nx.spring_layout(graph)
# -
# ### Create a small, interactive graph
#
# Now we can create a small graph using embedded HTML. You may optionally specify a [`matplotlib` color map](https://matplotlib.org/examples/color/colormaps_reference.html) and a `node_size_factor`.
graph_html = wn.make_interactive_graph(graph, pos=layout, cmap=plt.cm.viridis, edge_cmap=plt.cm.Blues, node_size_factor=5)
# ### Save an extremely high resolution graph for a Massive Pixel Environment
#
# This will take some time to run. You may specify your color maps, font sizes and node sizes here as well. Remember - what looks good on a small interactive screen may not work well on a display like [TACC's Stallion](https://www.tacc.utexas.edu/vislab/stallion)
# +
wn.save_big_graph(graph,
pos=layout,
cmap=plt.cm.viridis,
edge_cmap=plt.cm.Blues,
width=3,
height=15,
dpi=1600,
font_size=1,
node_size_factor=5,
output_file="mcu_network.png")
print("OK")
| examples/Stallion Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 02 : Pooling layer - demo
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
from google.colab import drive
drive.mount('/content/gdrive')
file_name = 'pool_layer_demo.ipynb'
import subprocess
path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode("utf-8")
print(path_to_file)
path_to_file = path_to_file.replace(file_name,"").replace('\n',"")
os.chdir(path_to_file)
# !pwd
import torch
import torch.nn as nn
# ### Make a pooling module
# * inputs: activation maps of size n x n
# * output: activation maps of size n/p x n/p
# * p: pooling size
mod = nn.MaxPool2d(2,2)
# ### Make an input 2 x 6 x 6 (two channels, each one has 6 x 6 pixels )
# +
bs=1
x=torch.rand(bs,2,6,6)
print(x)
print(x.size())
# -
# ### Feed it to the pooling layer: the output size should be divided by 2
# +
y=mod(x)
print(y)
print(y.size())
# -
| codes/labs_lecture08/lab02_pool_layer/pool_layer_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="69-03V1a4jbc" colab_type="code" outputId="4cf8163d-eb79-41cf-dccd-a08e54d22812" executionInfo={"status": "ok", "timestamp": 1577545162823, "user_tz": -120, "elapsed": 97918, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 235}
# !apt-get install -y -qq software-properties-common python-software-properties module-init-tools
# !add-apt-repository -y ppa:alessandro-strada/ppa 2>&1 > /dev/null
# !apt-get update -qq 2>&1 > /dev/null
# !apt-get -y install -qq google-drive-ocamlfuse fuse
from google.colab import auth
auth.authenticate_user()
from oauth2client.client import GoogleCredentials
creds = GoogleCredentials.get_application_default()
import getpass
# !google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret} < /dev/null 2>&1 | grep URL
vcode = getpass.getpass()
# !echo {vcode} | google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret}
# + id="jf_C1xvb7E8Z" colab_type="code" outputId="b69c8084-0ebe-4ab4-c76c-badab4c790a4" executionInfo={"status": "ok", "timestamp": 1577545172650, "user_tz": -120, "elapsed": 6781, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 64}
# !mkdir -p drive
# !google-drive-ocamlfuse drive
import tensorflow as tf
# + id="pbtCd2147Jj7" colab_type="code" colab={}
tf.compat.v1.disable_eager_execution()
# + id="I5Zv8j3W7M5m" colab_type="code" colab={}
path = '/content/drive/ColabNotebooks/cities/Jaipur2016-2019.csv'
# + id="c9eGxKYo7Toy" colab_type="code" colab={}
import pandas as pd
df = pd.read_csv(path)
# + id="XcO3DEjX7xPP" colab_type="code" outputId="28612597-6ce4-48d0-8b16-b90743edae31" executionInfo={"status": "ok", "timestamp": 1577545189654, "user_tz": -120, "elapsed": 1178, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + [markdown] id="CxS3tFKz8y--" colab_type="text"
#
# + id="QKv9LPIM7sF7" colab_type="code" colab={}
df['utc'] =pd.to_datetime(df.utc)
# + id="qNZ6gz7p71uL" colab_type="code" outputId="86b105dd-f69d-478c-bd43-bfd6219f4ee6" executionInfo={"status": "ok", "timestamp": 1577545205516, "user_tz": -120, "elapsed": 1176, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 235}
df.index = df['utc']
df.head()
# + id="tgY-DFCuD8fo" colab_type="code" outputId="6a8969eb-de3f-42e9-d78d-57800c428edb" executionInfo={"status": "ok", "timestamp": 1577545212068, "user_tz": -120, "elapsed": 1126, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
len(df[df.value > 500])
# + id="-nhA5QNM9LTl" colab_type="code" colab={}
df = df[df.value != 0.00]
# + id="c3B6i3SI9ReT" colab_type="code" colab={}
df = df[df.value > 0]
# + id="XopGw1MV9VOn" colab_type="code" colab={}
df = df[df.value < 500]
# + id="mQ87Wct9EY1O" colab_type="code" outputId="71b5e5fb-e302-47b3-9aa1-28da5b5915e4" executionInfo={"status": "ok", "timestamp": 1577545224459, "user_tz": -120, "elapsed": 1081, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
df.isnull().values.any()
# + id="zQwI8ktC-Rio" colab_type="code" outputId="c1a75aea-5814-4649-bbb1-fbb3fc1b5dc6" executionInfo={"status": "ok", "timestamp": 1577545228455, "user_tz": -120, "elapsed": 1068, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 235}
df.head()
# + id="fXo3KHSc-Y5G" colab_type="code" outputId="a9798bcf-f4af-428d-f855-1cb8113929fe" executionInfo={"status": "ok", "timestamp": 1577545233086, "user_tz": -120, "elapsed": 1026, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
len(df)
# + id="WzjBa04b8Bea" colab_type="code" colab={}
daily_groups = df.resample('60T')
# + id="yG2WjYgN8Div" colab_type="code" colab={}
hourly_data = daily_groups.mean()
# + id="XcYxMMAzF5rh" colab_type="code" colab={}
hourly_data = hourly_data.dropna()
# + id="6RQKGOe5FK80" colab_type="code" outputId="6b092e07-188e-4fdd-e31a-8ae8890dd34e" executionInfo={"status": "ok", "timestamp": 1577545247189, "user_tz": -120, "elapsed": 1096, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
len(hourly_data[hourly_data.value == 0])
# + id="zJP5_7jZ8IiB" colab_type="code" outputId="09c4af7a-fc62-40b3-f9fc-9dd46765c3b0" executionInfo={"status": "ok", "timestamp": 1577545265961, "user_tz": -120, "elapsed": 1016, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 305}
print(hourly_data.shape)
print(hourly_data)
# + id="ImOwgdfz-iGy" colab_type="code" outputId="66dc6d98-dad0-4123-e9de-b59e3ba27589" executionInfo={"status": "ok", "timestamp": 1577445412898, "user_tz": -120, "elapsed": 1502, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 450}
hourly_data
# + id="EdqcrLtS8TH5" colab_type="code" colab={}
hourly_data['value'] = hourly_data['value'].astype(float)
# + id="Ey-Z3y6xDcRt" colab_type="code" outputId="af9242ac-6354-48f2-a322-68499ecf4c09" executionInfo={"status": "ok", "timestamp": 1577445447324, "user_tz": -120, "elapsed": 1462, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 53}
features.dtypes
# + id="XKY2qMlS8UzE" colab_type="code" outputId="99a39e96-f238-4716-b46f-8bfccc748ed1" executionInfo={"status": "ok", "timestamp": 1577545279871, "user_tz": -120, "elapsed": 1104, "user": {"displayName": "<NAME>en", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 235}
features_considered = ['value']
features = hourly_data[features_considered]
features.head()
# + id="h5lrWj6-8fYN" colab_type="code" colab={}
def load_timeseries(data, params):
"""Load time series dataset"""
data = data.values
adjusted_window = params['window_size']+ 1
# Split data into windows
raw = []
for index in range(len(data) - adjusted_window):
raw.append(data[index: index + adjusted_window])
# Normalize data
result = normalize_windows(raw)
raw = np.array(raw)
result = np.array(result)
# Split the input dataset into train and test
split_ratio = round(params['train_test_split'] * result.shape[0])
train = result[:int(split_ratio), :]
np.random.shuffle(train)
# x_train and y_train, for training
x_train = train[:, :-1]
y_train = train[:, -1]
# x_test and y_test, for testing
x_test = result[int(split_ratio):, :-1]
y_test = result[int(split_ratio):, -1]
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1], 1))
x_test_raw = raw[int(split_ratio):, :-1]
y_test_raw = raw[int(split_ratio):, -1]
# Last window, for next time stamp prediction
last_raw = [data[-params['window_size']:]]
last = normalize_windows(last_raw)
last = np.array(last)
last = np.reshape(last, (last.shape[0], last.shape[1], 1))
return [x_train, y_train, x_test, y_test, x_test_raw, y_test_raw, last_raw, last]
def normalize_windows(window_data):
"""Normalize data"""
normalized_data = []
for window in window_data:
normalized_window = [((float(p) / float(window[0])) - 1) for p in window]
normalized_data.append(normalized_window)
return normalized_data
# + id="mIK4V0FY8k-c" colab_type="code" outputId="b19154e5-d9d8-45a8-f622-da6fa0f6c10a" executionInfo={"status": "ok", "timestamp": 1577545293597, "user_tz": -120, "elapsed": 1184, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
from keras.layers.core import Dense, Activation, Dropout
from keras.layers.recurrent import LSTM
from keras.models import Sequential
def rnn_lstm(layers, params):
"""Build RNN (LSTM) model on top of Keras and Tensorflow"""
model = Sequential()
model.add(LSTM(input_shape=(layers[1], layers[0]), output_dim=layers[1], return_sequences=True))
model.add(Dropout(params['dropout_keep_prob']))
model.add(LSTM(layers[2], return_sequences=False))
model.add(Dropout(params['dropout_keep_prob']))
model.add(Dense(output_dim=layers[3]))
model.add(Activation("tanh"))
model.compile(loss="mean_squared_error", optimizer="rmsprop")
return model
def predict_next_timestamp(model, history):
"""Predict the next time stamp given a sequence of history data"""
prediction = model.predict(history)
prediction = np.reshape(prediction, (prediction.size,))
return prediction
# + id="T6Xad2pd8rXq" colab_type="code" colab={}
import sys
import json
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
sys.path.append('/content/drive/ColabNotebooks/')
def train_predict(path, config_data):
train_file = path
parameter = config_data
params = json.loads(parameter)
x_train, y_train, x_test, y_test, x_test_raw, y_test_raw, last_window_raw, last_window = load_timeseries(train_file, params)
lstm_layer = [1, params['window_size'], params['hidden_unit'], 1]
model = rnn_lstm(lstm_layer, params)
model.fit(x_train, y_train, batch_size=params['batch_size'], epochs=params['epochs'], validation_split=params['validation_split'])
model.save('/content/drive/ColabNotebooks/models/Jaipur-Dec-28-19.h5')
predicted = predict_next_timestamp(model, x_test)
predicted_raw = []
for i in range(len(x_test_raw)):
predicted_raw.append((predicted[i] + 1) * x_test_raw[i][0])
plt.subplot(111)
plt.plot(predicted_raw, label='Actual')
plt.plot(y_test_raw, label='Predicted')
plt.legend()
plt.show()
next_timestamp = predict_next_timestamp(model, last_window)
next_timestamp_raw = (next_timestamp[0] + 1) * last_window_raw[0][0]
print('The next time stamp forecasting is: {}'.format(next_timestamp_raw))
# + id="DUL1hDeZ81R7" colab_type="code" colab={}
js = json.dumps({
"epochs": 20,
"batch_size": 256,
"window_size": 480,
"train_test_split": 0.8,
"validation_split": 0.2,
"dropout_keep_prob": 0.2,
"hidden_unit": 50
})
# + id="eesozYFY8-3w" colab_type="code" outputId="e340d16d-7ceb-4251-c73b-64fd402b3184" executionInfo={"status": "ok", "timestamp": 1577547249124, "user_tz": -120, "elapsed": 1913783, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCaY8RlKpYLlScTslI1yipGXu-VSsx47HgOYEvWGA=s64", "userId": "10525774958010629458"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
train_predict(features, js)
| DelhiOriginal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bagasbudhi/Data-Smoking_Machine-Learning-Algorithm/blob/master/_HME_DS_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="oGmSTqw1lSuW"
import pandas as pd #Package untuk manipulasi tabel/ dataframe
import numpy as np # Package untuk perhitungan pada pandas
import matplotlib.pyplot as plt # Package untuk membuat plot/ grafik
import seaborn as sns # Package untuk mempercantik grafik
# + [markdown] id="mVKj9IVulkk7"
# ## Import Data
# + id="50brBGtVgIWF"
# + [markdown] id="jRYzJYG7gJXO"
# ## Ini Judul
# + id="knmUuloilmVq" outputId="d2b79acc-ddfd-402c-b5ef-63d4c207af2d" colab={"base_uri": "https://localhost:8080/", "height": 347}
# Import Test Data dan Raw Data
raw_data = pd.read_csv("train.csv") # data train yang akan dianalisis
test_data = pd.read_csv("test.csv") # data test yang akan diprediksi
raw_data.head(10)
# + id="yqJsPZAFl6JD" outputId="5925b171-3a60-4efd-c7c2-845d45adba1f" colab={"base_uri": "https://localhost:8080/", "height": 70}
# Cek panjang dataset
print(len(raw_data))
print(len(test_data))
len(raw_data) /(len(test_data) + len(raw_data))
# + [markdown] id="-a8F8sPhlnAL"
# ## Preprocessing
# + id="GZbz670wlolJ" outputId="2c8c9928-58f5-4274-c54d-9879783aa4a7" colab={"base_uri": "https://localhost:8080/", "height": 307}
raw_data.describe()
# + id="ZozOaK2TlpDw" outputId="c4863fca-9bef-45ed-8353-ff89f423f06d" colab={"base_uri": "https://localhost:8080/", "height": 87}
raw_data.columns
# + [markdown] id="o60Mbgfxj3IW"
#
# + id="hPYnXtWNmAeM" outputId="f162180d-52d8-4cb5-adab-04e63382564f" colab={"base_uri": "https://localhost:8080/", "height": 336}
raw_data.isnull().sum()
# + [markdown] id="Id3Fggpzkm0A"
# aku bukan heading
# + id="BtohRpznmAjo" outputId="8a5a1a06-0c88-4baf-a47c-30204a21286c" colab={"base_uri": "https://localhost:8080/", "height": 336}
raw_data.dtypes #ini komen
# + id="qVB1TGDSmAmM" outputId="b0281c5b-3b43-418e-9eae-44d35d8c8787" colab={"base_uri": "https://localhost:8080/", "height": 70}
raw_data["sex"].value_counts()
# + id="S8Yd_8iAlpdR" outputId="4768d5c3-820b-4407-821a-fbd01fadac0f" colab={"base_uri": "https://localhost:8080/", "height": 197}
raw_data.head()
# + id="jCKXihcDmAqI" outputId="ab733041-7b89-493a-c398-641eea5adbc3" colab={"base_uri": "https://localhost:8080/", "height": 197}
raw_data["sex"] = raw_data["sex"].replace({"F":1, "M" :0})
raw_data.head()
# + id="QrYP8FcWBPL6" outputId="3d8542ce-1a7b-454e-96f4-f2a2ea351d75" colab={"base_uri": "https://localhost:8080/", "height": 197}
raw_data["is_smoking"] = raw_data["is_smoking"].replace({"YES":1, "NO":0})
raw_data.head()
# + id="huajW0n3BX18" outputId="a4424878-4721-4004-d964-d5a4a9e04727" colab={"base_uri": "https://localhost:8080/", "height": 336}
raw_data.isnull().sum()
# + id="7vSLGwIZqpFg"
# Imputasi Nilai dengan Nilai Modus
raw_data_imputed["education"] = raw_data_imputed["education"].fillna(raw_data_imputed["education"].mode().iloc[0])
# Imputasi Nilai dengan Nilai Rata- Rata
raw_data_imputed["cigsPerDay"] = raw_data_imputed["cigsPerDay"].fillna(np.mean(raw_data_imputed["cigsPerDay"]))
# + [markdown] id="En4v3mj7Cr2k"
# ### Imputasi
# + id="O0HswCOwB-Mg" outputId="bb449344-243e-484e-f710-25a5dbec6e48" colab={"base_uri": "https://localhost:8080/", "height": 336}
raw_data_imputed = raw_data
raw_data_imputed["education"] = raw_data_imputed["education"].fillna(raw_data_imputed["education"].mode().iloc[0])
raw_data_imputed["cigsPerDay"] = raw_data_imputed["cigsPerDay"].fillna(np.mean(raw_data_imputed["cigsPerDay"]))
raw_data_imputed["BPMeds"] = raw_data_imputed["BPMeds"].fillna(raw_data_imputed["BPMeds"].mode().iloc[0])
raw_data_imputed["totChol"] = raw_data_imputed["totChol"].fillna(np.mean(raw_data_imputed["totChol"]))
raw_data_imputed["BMI"] = raw_data_imputed["BMI"].fillna(np.mean(raw_data_imputed["BMI"]))
raw_data_imputed["heartRate"] = raw_data_imputed["heartRate"].fillna(np.mean(raw_data_imputed["heartRate"]))
raw_data_imputed["glucose"] = raw_data_imputed["glucose"].fillna(np.mean(raw_data_imputed["glucose"]))
raw_data_imputed.isnull().sum()
# + [markdown] id="MIQwHLZ4lpeA"
# ## EDA
# + id="3LmAgUlGBOBl" outputId="c007fe4a-6e21-4c5d-d3ce-4fb9304d539b" colab={"base_uri": "https://localhost:8080/", "height": 296}
sns.distplot(raw_data["cigsPerDay"])
# + id="_wSMwpMRrXq-" outputId="bd46e98e-1af3-4373-862b-a4f1ebd279c7" colab={"base_uri": "https://localhost:8080/", "height": 352}
sns.distplot(raw_data["cigsPerDay"])
# + id="I_xe6iPXlrnc" outputId="7ca58949-532b-4430-80d6-27d73d872ce0" colab={"base_uri": "https://localhost:8080/", "height": 296}
sns.distplot(raw_data["age"])
# + id="z9EaU2Z7lr__"
# + [markdown] id="RFasYmDKUCC0"
# ### a. Correlation Heatmap
# + id="8WEqg-4PUE6S" outputId="81e47d50-2f18-4f3d-d63f-b761c1688226" colab={"base_uri": "https://localhost:8080/", "height": 974}
corr= raw_data_imputed.corr() # Ganti raw_data_imputed dengan nama dataframe yang ingin dicek
sns.set(font_scale=1.15) # Set besar tulisan
plt.figure(figsize=(20, 15)) # Set besar heatmap
sns.heatmap(corr, vmax=.8, linewidths=0.01,
square=True,annot=True,linecolor="black") # Annpt= False jika tidak ingin terlihat nilainya
plt.title('Ini judul'); # Buat judul heatmap
# + id="t5qjaFV8UFAb"
# + id="SjAscTRfUE-A"
# + [markdown] id="7mJihd-OlsL2"
# ## Feature Engineering
# + id="Zsa9frMGluDW"
# Nambah Kolom
# Standardisasi
# Kurangin Kolom
# + [markdown] id="kswWZB6fGYRb"
# ### a. Standardization
# + id="6bt_mWrKludK" outputId="5022b424-b0ab-4b60-fbc2-556d8605d4e6" colab={"base_uri": "https://localhost:8080/", "height": 247}
data_standarded = raw_data_imputed
from sklearn.preprocessing import StandardScaler # Import fungsi yang dibutuhkan
sc=StandardScaler() # Panggil Fungsi untuk standardization
data_standarded = sc.fit_transform(data_standarded) # Fit dan Transform suatu dataframe
# data_standarded = sc.transform(data_standarded) # Jika hanya ingin transform tanpa fit
data_standarded
# + id="MtudaBqoHscE" outputId="0fc52808-353a-423f-f34c-28453d400ba4" colab={"base_uri": "https://localhost:8080/", "height": 217}
data_standarded= pd.DataFrame(data_standarded, columns = raw_data_imputed.columns)
data_standarded["TenYearCHD"] = raw_data_imputed["TenYearCHD"]
data_standarded.head()
# + [markdown] id="bU4Dxe3WMcxv"
# ### b. Pemilihan Fitur
# + id="nGVD7NhrIMHx" outputId="f4a75d97-e623-4855-e516-f80e2ea83a4c" colab={"base_uri": "https://localhost:8080/", "height": 217}
# Buang id
data_standarded = data_standarded.drop(["id"], axis = 1)
data_standarded.head()
# + [markdown] id="KGonHbQMluqQ"
# ## Model and Evaluation
# + id="61_C-bG0Jsg5" outputId="1c69eb96-7772-46f5-b19a-fd2529213379" colab={"base_uri": "https://localhost:8080/", "height": 217}
# Buat data train dan data validasi
from sklearn.model_selection import train_test_split
X = data_standarded.drop(["TenYearCHD"], axis = 1)
Y = data_standarded["TenYearCHD"]
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2) # Data train dan validasi
X_train.head()
# + id="093TCU3wlvZE"
# Import needed library
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
# + [markdown] id="N8A_Ej9dIfMe"
# ### 1. Logistic Regression
# + id="Tw_fSrHdLTGP"
# + id="0MWMd9RUGdoH" outputId="e857f313-d3c5-496c-ed1b-6699e25036de" colab={"base_uri": "https://localhost:8080/", "height": 105}
# Train
lr = LogisticRegression()
lr.fit(X_train,Y_train)
# + id="u-QZgPfnGdwK" outputId="787315b0-a413-4584-8f46-56981aebe270" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Test
acc = lr.score(X_test,Y_test)*100
#accuracies['Logistic Regression'] = acc
print("Test Accuracy {:.2f}%".format(acc))
# + id="sqW3lywIMIlc"
# + id="vsS_8xiUMIpf"
# + [markdown] id="3Lnv-I1wIlRk"
# ### 2. SVM
# + id="tVp_0fsgGd4Y" outputId="44ca5664-add4-45d4-fc09-25acc09f88a8" colab={"base_uri": "https://localhost:8080/", "height": 87}
# Train SVM
svm = SVC()
svm.fit(X_train,Y_train)
# + [markdown] id="tosaeeaNwZMj"
#
# + id="age2zSFlIojY" outputId="16248f61-db82-4a87-9c98-fcd99d5f525c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Test
acc = svm.score(X_test,Y_test)*100
#accuracies['Logistic Regression'] = acc
print("Test Accuracy {:.2f}%".format(acc))
# + id="Zf5FsLIDIooX"
# + [markdown] id="qsRvb6UKIpht"
# ### 3. KNN
# + id="BB35cSQqIomA" outputId="0066a6f9-3564-4f90-ddcf-36676ad6eb4e" colab={"base_uri": "https://localhost:8080/", "height": 70}
# Train
KNN = KNeighborsClassifier()
KNN.fit(X_train,Y_train)
# + id="WRp8_3RqI9rC" outputId="7298fafb-ec37-4541-d3f0-ca072c82317d" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Test
acc = KNN.score(X_test,Y_test)*100
#accuracies['Logistic Regression'] = acc
print("Test Accuracy {:.2f}%".format(acc))
# + id="RbDZQGV0L4iu"
# + id="nLptBVQsMJVs"
# + [markdown] id="0M01bwNbMJ6Z"
# ### 4. Decision Tree
# + id="jgDUWE9NMJfh" outputId="a324b72f-03ee-476a-dbc8-bb6fbe6a321a" colab={"base_uri": "https://localhost:8080/", "height": 123}
# Train
tree = DecisionTreeClassifier()
tree.fit(X_train,Y_train)
# + id="T9xEPX2EMWPk" outputId="63880067-fa5a-4431-9750-5a38a0f8ef7d" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Test
acc = tree.score(X_test,Y_test)*100
#accuracies['Logistic Regression'] = acc
print("Test Accuracy {:.2f}%".format(acc))
# + id="ZNRbDJkmMX6R"
# + [markdown] id="ChMEcAQIM1Po"
# ## Predict
# + id="NvRdqKtVM2Qi" outputId="96942833-29c9-45e3-8ed9-b211a8f157c9" colab={"base_uri": "https://localhost:8080/", "height": 217}
test_data.head()
# + id="XKFyO0i4M8oj" outputId="5beebf6e-cde3-4b3d-9e11-c867b19063e4" colab={"base_uri": "https://localhost:8080/", "height": 247}
# Preprocessing dan FE data test
test_data["sex"] = test_data["sex"].replace({"M":0, "F":1})
test_data["is_smoking"] = test_data["is_smoking"].replace({"YES":1, "NO":0})
test_data_imputed = test_data
test_data_imputed["education"] = test_data_imputed["education"].fillna(raw_data_imputed["education"].mode().iloc[0])
test_data_imputed["cigsPerDay"] = test_data_imputed["cigsPerDay"].fillna(np.mean(raw_data_imputed["cigsPerDay"]))
test_data_imputed["BPMeds"] = test_data_imputed["BPMeds"].fillna(raw_data_imputed["BPMeds"].mode().iloc[0])
test_data_imputed["totChol"] = test_data_imputed["totChol"].fillna(np.mean(raw_data_imputed["totChol"]))
test_data_imputed["BMI"] = test_data_imputed["BMI"].fillna(np.mean(raw_data_imputed["BMI"]))
test_data_imputed["heartRate"] = test_data_imputed["heartRate"].fillna(np.mean(raw_data_imputed["heartRate"]))
test_data_imputed["glucose"] = test_data_imputed["glucose"].fillna(np.mean(raw_data_imputed["glucose"]))
data_test_standarded = test_data_imputed
data_test_standarded = sc.fit_transform(data_test_standarded)
data_test_standarded
# + id="D5uxD4RaQH15" outputId="e84ce018-d3dd-41d5-cd12-f3a79265675d" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(test_data)
# + id="hR5V_1J9N0HW" outputId="7da42f44-ebc6-41e7-8627-7c0ba368ea9a" colab={"base_uri": "https://localhost:8080/", "height": 217}
# Ubah hasil standardisasi jadi dataframe
data_test_standarded= pd.DataFrame(data_test_standarded, columns = test_data_imputed.columns)
data_test_standarded.head()
# + id="eP19Je9wPdt1"
# Buang fitur "id"
data_test_standarded = data_test_standarded.drop(["id"], axis = 1)
# + id="Hnh4VcSwQCfv"
# + id="RAHRMRHcN0Qt" outputId="f28d7895-d0cb-4552-87c1-efc0782df172" colab={"base_uri": "https://localhost:8080/", "height": 105}
# Bikin prediksi
# Train dulu
lr = LogisticRegression()
lr.fit(X,Y)
# + id="nY67Rj36N0Zs" outputId="b75a2a0f-ce17-4b3c-c0b2-b83f56941968" colab={"base_uri": "https://localhost:8080/", "height": 710}
# Lalu Prediksi
y_pred = lr.predict(data_test_standarded)
y_pred
# + id="0URSB1xHP63I" outputId="de9d340e-497c-4bd2-c5d3-9f9c014b335e" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(y_pred)
# + id="YG9QRNADRpHq" outputId="8e216c58-59a1-45b9-89be-fa57c137bc34" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.sum(y_pred)
# + id="FNNu3u2SPX9T" outputId="48c7ed49-0230-45c7-bb7d-5600c9d7ffc1" colab={"base_uri": "https://localhost:8080/", "height": 304}
# Buat dataframe untuk submission
sub = test_data[["id","age"]]
sub["TenYearCHD"] = y_pred
sub = sub.drop(["age"], axis = 1)
sub.head()
# + id="sMaz51gLP1ts"
# Export to csv
export_csv = sub.to_csv("hasil_1.csv", index= False)
# + id="jmk7Jd_MRgwz"
# + id="aCQK-ZAtTAvu"
| _HME_DS_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem set 2: Counting words
#
# ## Description
#
# The goal of this problem set is to create the world's least visually-sophisticated word count graphic.
#
# Along the way, you'll experiment with stopword removal, case folding, and other processing steps.
#
# ## Count words, naïvely
#
# We'll work with *Moby-Dick*, as we did in class.
#
# **Read the text of *Moby-Dick* from a file (it's on the class GitHub site, in the `data/texts` directory), tokenize it with NLTK's `word_tokenize` function, and count the resulting tokens in a `Counter` object.**
#
# You can refer to the lecture notebook from Monday, September 7, to borrow code to do all of this. But you must get that code working in the cell below. This cell should produce a `Counter` object that holds the token counts from the novel.
# Use standard Python file commands to open Moby-Dick,
# then count the words in that file.
import os
from collections import Counter
from nltk import word_tokenize
moby_name= 'A-Melville-Moby_Dick-1851-M.txt'
moby_path= os.path.join('..','..','data','texts', moby_name)
print(moby_path)
# +
import nltk
nltk.download('punkt')
moby_simple_c= Counter()
with open(moby_path, 'r') as f:
for line in f:
tokens = word_tokenize(line.strip())
for token in tokens:
moby_simple_c[token] += 1
# -
moby_simple_c.most_common(5)
# **Print the total number of words (hint: use `Counter`'s `.values()` method, along with the `sum` function) in your text, as well as the 20 most frequently occurring terms and their counts.**
#
# We'll do this a lot, so wrap it up as a function that takes as input a `Counter` object and an optional number of top terms to print:
#
# ```
# def word_stats(data, n=20):
# ```
#
# The output of your fuction should look like this:
#
# ```
# Total words in the text: 255380
#
# Top 20 words by frequency:
# , 19204
# the 13715
# . 7432
#
# [and so on ...]
# ```
# Your word_stats function here
def word_stats(data, n=20):
'''
Print total wordcount and n top terms.
Takes a Counter object and a number of terms to print.
Returns None.
'''
total = sum(data.values())
print('Total Tokens:', total)
print('\nMost common tokens:')
for word in data.most_common(n):
print(word[0], '\t', word[1])
return None
# Call word_stats on your data
word_stats(moby_simple_c)
# ## Case folding and stopwords
#
# As you can see, the top words that we counted aren't super informative. That said, list two things that you **can** say about the text with reasonable confidence on the basis of our results above:
# **Two things you *can* tell about *Moby-Dick* from the naïve word counts:**
#
# 1. Moby Dick's most used *words* are stop words
# 2. Punctuation is counted to be the most used token
# If we want our word list to be informative, we need to find a way to ignore high-frequency, low-information words. We can do this either by not counting them in the first place, or by excluding them from our reporting after we've collected them. Both methods have advantages and drawbacks. The one you pursue is up to you.
#
# **Modify the original code to ignore token case (e.g., 'The' and 'the' are both counted as occurrences of the same token; note the `.lower()` method for strings) and to remove the English-language stopwords defined by NLTK (`from nltk.corpus import stopwords`). Then display the total token count and top-20 tokens.**
# +
# Count tokens with case folding and NLTK English stopwords removed
nltk.download('stopwords')
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english')) #set stopwords code off stack overflow
moby_simple_c= Counter()
with open(moby_path, 'r') as f:
for line in f:
tokens = word_tokenize(line.strip())
for token in tokens:
token = token.lower()
if not token in stop_words:
moby_simple_c[token] += 1
word_stats(moby_simple_c)
# -
# Is this better? Maybe! **Note one advantage of this stopword-removed count, as well as one disadvantage:**
# **Advantage:**
#
# * We are getting more words that aren't stop words and we can see, possibly, the type of book this is without even having to read it. We also get to understand what is really the important tokens to look at.
#
# **Disadvantage:**
#
# * A disadvantange might be that we are unable to see all of the tokens that should be included and might be leaving out important tokens
# Let's see if we can further improve/refine our approach to continue narrowing our word list. Our goal is to produce a list that contains *only* interesting words and ranks them by frequency.
#
# **List at least two ideas for modifying the stopword list to better approach our goal:**
#
# 1. Personally, I do not think commas or periods or semicolons tell much of the story, unlike other punctuation, so I would get rid of those. I believe that these punctuations do not give any additional information or context for the story.
# 2. I would also like to see elimination of shortened words, such as 's. I think n't gives more context than 's so, other than n't I would like to see stopwords try to modify its list by better identifying stop words that have been shortened.
# 3. Also just as a side note, I got rid of the '--' because I do not think it really adds anything, but I did keep in the quotation marks and stylized quotation marks since they give context to the reading.
# **Implement one or more of your ideas to improve the stopword list, then display the output of your new version using `word_stats()`.**
# +
# Better stopwords in action!
lst = [',','.',';','\'s', '--']
for i in lst:
stop_words.add(i)
moby_simple_c= Counter()
with open(moby_path, 'r') as f:
for line in f:
tokens = word_tokenize(line.strip())
for token in tokens:
token = token.lower()
if not token in stop_words:
moby_simple_c[token] += 1
word_stats(moby_simple_c)
# -
# Refine your stoplist until you're satisfied with it. Make sure your code above displays the final output of your `word_stats` function. Then move on.
# # Visualization
#
# Now, make the world's least visually-impressive word count graphic. Your task is to produce a visual representation of your top 10 words that shows the relative frquency of those terms.
#
# The simplest acceptable version of this visualization is a bar chart. **Complete the starter code below to produce a bar chart of the top ten words in the text.**
#
# Your output might look like this:
#
# 
def words(d, n=10):
total = sum(d.values())
print('Total Tokens:', total)
print('\nMost common tokens:')
for word in d.most_common(n):
print(word[0], '\t', word[1])
return None
# +
lst = [',','.',';','\'s', '--', '``', '\'\'', '!', '?']
for i in lst:
stop_words.add(i)
moby_simple= Counter()
with open(moby_path, 'r') as f:
for line in f:
tokens = word_tokenize(line.strip())
for token in tokens:
token = token.lower()
if not token in stop_words:
moby_simple[token] += 1
words(moby_simple)
# +
# Make a bar chart of the top 10 words
# %matplotlib inline
import matplotlib.pyplot as plt
moby_chart = moby_simple.most_common(10)
x = []
for i in moby_chart:
x.append(i[0])
y = []
for i in moby_chart:
y.append(i[1])
# Get labels and counts
labels = x
counts = y
# Create the figure
fig, ax = plt.subplots()
ax.barh(x, y, color='royalblue') #like the ocean
ax.invert_yaxis()
ax.set_xlabel('Count')
ax.set_ylabel('Words')
ax.set_title('Words v Count')
plt.show()
# -
# ## Optional: word clouds
#
# **This is optional.** Make a word cloud. You can do this the ugly way in pure `matplotlib` or the easy-and-pretty way by using the [`wordcloud`](https://github.com/amueller/word_cloud) library:
#
# ```
# conda install -c conda-forge wordcloud
# ```
# If you use `wordcloud`, you'll be interested in the [`.generate_from_frequencies()` method](http://amueller.github.io/word_cloud/auto_examples/frequency.html).
#
# Here are examples of the ugly and the pretty outputs. Your specific results might vary.
#
# 
# 
# +
# The ugly way (matplotlib)
# Hint: you'll want to use the .text() plotting method
# Strictly optional
# +
# The pretty way
# Strictly optional
#code from link posted
import numpy as np
import os
import re
from PIL import Image
from os import path
from wordcloud import WordCloud
import matplotlib.pyplot as plt
whale = np.array(Image.open(path.join("moby-dick-stencil.png"))) #image from online, https://www.pinterest.com/pin/2111131054358686/
moby_dick = open(moby_path).read()
wc = WordCloud(background_color="white", max_words=100, mask=whale,
stopwords=stop_words)
wc.generate(moby_dick)
plt.imshow(wc, interpolation='bilinear')
plt.axis("off")
plt.show()
# +
wc = WordCloud(background_color="white", max_words=100,
stopwords=stop_words)
wc.generate(moby_dick)
plt.imshow(wc, interpolation='nearest')
plt.axis("off")
plt.show()
# -
| problem_sets/ps_02/ps_02_word_counts_jh976.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + cell_id="e2a2070f-2473-4736-b71d-c0fc8facca0a" deepnote_cell_height=117 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1429 execution_start=1647027891407 source_hash="35374e32" tags=[]
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
/ + [markdown] cell_id="a2d58bd5-7a1a-44c2-9dbf-b4d10d418ba3" deepnote_cell_height=52.390625 deepnote_cell_type="markdown" tags=[]
/ Importar datos
/ + cell_id="3dbdf21f-89a6-4e15-8cbc-4e0947ac68df" deepnote_cell_height=445.796875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=23 execution_start=1647027892875 source_hash="56a46059" tags=[]
data = pd.read_csv('/work/Bot-de-Trafing//GOOG.csv')
data.head()
/ + cell_id="eb308bec-3501-44ef-a184-1c866df656d5" deepnote_cell_height=516 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=71 execution_start=1647027892910 source_hash="d786443f" tags=[]
data.describe()
/ + [markdown] cell_id="42550a17-42c9-439b-8b36-4989513fb160" deepnote_cell_height=52.390625 deepnote_cell_type="markdown" tags=[]
/ Graficar los datos
/ + cell_id="7fc34702-03d1-4a48-9e11-5ab9f049c5c0" deepnote_cell_height=540 deepnote_cell_type="code" deepnote_output_heights=[335] deepnote_to_be_reexecuted=false execution_millis=464 execution_start=1647027892991 source_hash="bb6fbfa4" tags=[]
plt.figure(figsize=(10, 5))
plt.plot(data['Close'], label =('Google stock'))
plt.title('Precio de las acciones de google desde el 2016-2022')
plt.xlabel('3 de junio 2016 hasta 1 de marzo 2022')
plt.ylabel('precio de cierre en ($)')
plt.legend(loc ='upper left')
plt.show()
/ + cell_id="45633028-2f92-433d-9d11-5dd7f09e87e5" deepnote_cell_height=634 deepnote_cell_type="code" deepnote_table_loading=false deepnote_table_state={"filters": [], "pageIndex": 0, "pageSize": 10, "sortBy": []} deepnote_to_be_reexecuted=false execution_millis=88 execution_start=1647027893368 source_hash="216ac24b" tags=[]
mvs30 = pd.DataFrame()
mvs30['Close'] = data['Close'].rolling(window=30).mean()
mvs30
/ + cell_id="8ca009a7-ae59-455d-9a5d-e44e9688e091" deepnote_cell_height=634 deepnote_cell_type="code" deepnote_table_loading=false deepnote_table_state={"filters": [], "pageIndex": 0, "pageSize": 10, "sortBy": []} deepnote_to_be_reexecuted=false execution_millis=87 execution_start=1647027893370 source_hash="1a235bf8" tags=[]
mvs100 = pd.DataFrame()
mvs100['Close'] = data['Close'].rolling(window=100).mean()
mvs100
/ + cell_id="815eed6f-1069-41f3-8f1d-611a1f7336a7" deepnote_cell_height=576 deepnote_cell_type="code" deepnote_output_heights=[335] deepnote_to_be_reexecuted=false execution_millis=500 execution_start=1647027893423 source_hash="d4fbe277" tags=[]
plt.figure(figsize=(10, 5))
plt.plot(data['Close'], label =('Google stock'))
plt.plot(mvs30['Close'], label =('Media movil de 30 periodos'))
plt.plot(mvs100['Close'], label =('Media movil de 100 periodos'))
plt.title('Precio de las acciones de google desde el 2016-2022')
plt.xlabel('3 de junio 2016 hasta 1 de marzo 2022')
plt.ylabel('precio de cierre en ($)')
plt.legend(loc ='upper left')
plt.show()
/ + cell_id="b04d90d8-03f4-4260-8ad3-47a251cfc964" deepnote_cell_height=670 deepnote_cell_type="code" deepnote_table_loading=false deepnote_table_state={"filters": [], "pageIndex": 0, "pageSize": 10, "sortBy": []} deepnote_to_be_reexecuted=false execution_millis=220 execution_start=1647027893706 source_hash="ed28337e" tags=[]
datos = pd.DataFrame()
datos['Google'] = data['Close']
datos['mvs30'] = mvs30['Close']
datos['mvs100'] = mvs100['Close']
datos
/ + cell_id="56925112-5643-47c6-ad08-004e29c63955" deepnote_cell_height=621 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1647027893749 source_hash="d28d7332" tags=[]
def senal(datos):
compra = []
venta = []
condicion = 0
for dia in range(len(datos)):
if datos['mvs30'][dia] > datos['mvs100'][dia]:
if condicion != 1:
compra.append(datos['Google'][dia])
venta.append(np.nan)
condicion = 1
else:
compra.append(np.nan)
venta.append(np.nan)
elif datos['mvs30'][dia] < datos['mvs100'][dia]:
if condicion != -1:
venta.append(datos['Google'][dia])
compra.append(np.nan)
condicion = -1
else:
compra.append(np.nan)
venta.append(np.nan)
else:
compra.append(np.nan)
venta.append(np.nan)
return(compra,venta)
/ + cell_id="b9887021-7ea0-473c-87e0-b8b5e97c7b27" deepnote_cell_height=652 deepnote_cell_type="code" deepnote_output_heights=[611] deepnote_table_invalid=false deepnote_table_loading=false deepnote_table_state={"filters": [], "pageIndex": 9, "pageSize": 10, "sortBy": []} deepnote_to_be_reexecuted=false execution_millis=131 execution_start=1647027893801 source_hash="59baffaf" tags=[]
senales = senal(datos)
datos['Compra'] = senales[0]
datos['Venta'] = senales[1]
datos
/ + cell_id="39955eb2-ade3-439f-a2af-6bab47f0af6e" deepnote_cell_height=612 deepnote_cell_type="code" deepnote_output_heights=[335, 305] deepnote_to_be_reexecuted=false execution_millis=330 execution_start=1647027893865 source_hash="536758c2" tags=[]
plt.figure(figsize=(10,5))
plt.plot(datos['Google'], label = 'Google', alpha = 0.3)
plt.plot(datos['mvs30'], label = 'mvs30', alpha = 0.3)
plt.plot(datos['mvs100'], label = 'mvs100', alpha = 0.3)
plt.scatter(datos.index, datos['Compra'], label = 'Precio de Compra', marker = '^', color = 'green')
plt.scatter(datos.index, datos['Venta'], label = 'Precio de Venta', marker = 'v', color = 'red')
plt.title('Acciones de Google - Precio de sus acciones desde el 2016 - 2022')
plt.xlabel('3 de junio 2016 - 1 marzo. 2022')
plt.ylabel('Precio de cierre ($)')
plt.legend(loc = 'upper left')
plt.show()
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=add430d6-c4b8-458c-a6b4-6a764b1d4b2b' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ<KEY> > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CNNs
# In this notebook you will learn how to build Convolutional Neural Networks (CNNs) for image processing.
# ## Imports
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import sklearn
import sys
import tensorflow as tf
from tensorflow import keras
import time
print("python", sys.version)
for module in mpl, np, pd, sklearn, tf, keras:
print(module.__name__, module.__version__)
assert sys.version_info >= (3, 5) # Python ≥3.5 required
assert tf.__version__ >= "2.0" # TensorFlow ≥2.0 required
# 
# ## Exercise 1 – Simple CNN
# ### 1.1)
# Load CIFAR10 using `keras.datasets.cifar10.load_data()`, and split it into a training set (45,000 images), a validation set (5,000 images) and a test set (10,000 images). Make sure the pixel values range from 0 to 1. Visualize a few images using `plt.imshow()`.
classes = [
"airplane",
"automobile",
"bird",
"cat",
"deer",
"dog",
"frog",
"horse",
"ship",
"truck",
]
(x_train, y_train), (x_test, y_test) = keras.datasets.cifar10.load_data()
x_train, x_valid = x_train[:-5000], x_train[-5000:]
y_train, y_valid = y_train[:-5000], y_train[-5000:]
y_valid.shape
x_test.shape
# ### 1.2)
# Build and train a baseline model with a few dense layers, and plot the learning curves. Use the model's `summary()` method to count the number of parameters in this model.
#
# **Tip**:
#
# * Recall that to plot the learning curves, you can simply create a Pandas `DataFrame` with the `history.history` dict, then call its `plot()` method.
# ### 1.3)
# Build and train a Convolutional Neural Network using a "classical" architecture: N * (Conv2D → Conv2D → Pool2D) → Flatten → Dense → Dense. Before you print the `summary()`, try to manually calculate the number of parameters in your model's architecture, as well as the shape of the inputs and outputs of each layer. Next, plot the learning curves and compare the performance with the previous model.
# ### 1.4)
# Looking at the learning curves, you can see that the model is overfitting. Add a Batch Normalization layer after each convolutional layer. Compare the model's performance and learning curves with the previous model.
#
# **Tip**: there is no need for an activation function just before the pooling layers.
# 
# ## Exercise 1 – Solution
# ### 1.1)
# Load CIFAR10 using `keras.datasets.cifar10.load_data()`, and split it into a training set (45,000 images), a validation set (5,000 images) and a test set (10,000 images). Make sure the pixel values range from 0 to 1. Visualize a few images using `plt.imshow()`.
classes = [
"airplane",
"automobile",
"bird",
"cat",
"deer",
"dog",
"frog",
"horse",
"ship",
"truck",
]
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.cifar10.load_data()
X_train = X_train_full[:-5000] / 255
y_train = y_train_full[:-5000]
X_valid = X_train_full[-5000:] / 255
y_valid = y_train_full[-5000:]
X_test = X_test / 255
plt.figure(figsize=(10, 7))
n_rows, n_cols = 10, 15
for row in range(n_rows):
for col in range(n_cols):
i = row * n_cols + col
plt.subplot(n_rows, n_cols, i + 1)
plt.axis("off")
plt.imshow(X_train[i])
# Let's print the classes of the images in the first row:
for i in range(n_cols):
print(classes[y_train[i][0]], end=" ")
# ### 1.2)
# Build and train a baseline model with a few dense layers, and plot the learning curves. Use the model's `summary()` method to count the number of parameters in this model.
#
# **Tip**:
#
# * Recall that to plot the learning curves, you can simply create a Pandas `DataFrame` with the `history.history` dict, then call its `plot()` method.
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[32, 32, 3]),
keras.layers.Dense(64, activation="selu"),
keras.layers.Dense(64, activation="selu"),
keras.layers.Dense(64, activation="selu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=0.01), metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
pd.DataFrame(history.history).plot()
plt.axis([0, 19, 0, 1])
plt.show()
model.summary()
# ### 1.3)
# Build and train a Convolutional Neural Network using a "classical" architecture: N * (Conv2D → Conv2D → Pool2D) → Flatten → Dense → Dense. Before you print the `summary()`, try to manually calculate the number of parameters in your model's architecture, as well as the shape of the inputs and outputs of each layer. Next, plot the learning curves and compare the performance with the previous model.
model = keras.models.Sequential([
keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu", input_shape=[32, 32, 3]),
keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu"),
keras.layers.MaxPool2D(pool_size=2),
keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
keras.layers.MaxPool2D(pool_size=2),
keras.layers.Flatten(),
keras.layers.Dense(128, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=0.01), metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
pd.DataFrame(history.history).plot()
plt.axis([0, 19, 0, 1])
plt.show()
# Number of params in a convolutional layer =
# (kernel_width * kernel_height * channels_in + 1 for bias) * channels_out
(
(3 * 3 * 3 + 1) * 32 # in: 32x32x3 out: 32x32x32 Conv2D
+ (3 * 3 * 32 + 1) * 32 # in: 32x32x32 out: 32x32x32 Conv2D
+ 0 # in: 32x32x32 out: 16x16x32 MaxPool2D
+ (3 * 3 * 32 + 1) * 64 # in: 16x16x32 out: 16x16x64 Conv2D
+ (3 * 3 * 64 + 1) * 64 # in: 16x16x64 out: 16x16x64 Conv2D
+ 0 # in: 16x16x64 out: 8x8x64 MaxPool2D
+ 0 # in: 8x8x64 out: 4096 Flatten
+ (4096 + 1) * 128 # in: 4096 out: 128 Dense
+ (128 + 1) * 10 # in: 128 out: 10 Dense
)
# Let's check:
model.summary()
# ### 1.4)
# Looking at the learning curves, you can see that the model is overfitting. Add a Batch Normalization layer after each convolutional layer. Compare the model's performance and learning curves with the previous model.
#
# **Tip**: there is no need for an activation function just before the pooling layers.
model = keras.models.Sequential([
keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu", input_shape=[32, 32, 3]),
keras.layers.BatchNormalization(),
keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.MaxPool2D(pool_size=2),
keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.MaxPool2D(pool_size=2),
keras.layers.Flatten(),
keras.layers.Dense(128, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=0.01), metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
pd.DataFrame(history.history).plot()
plt.axis([0, 19, 0, 1])
plt.show()
# 
# ## Exercise 2 – Separable Convolutions
# ### 2.1)
# Replace the `Conv2D` layers with `SeparableConv2D` layers (except the first one), fit your model and compare its performance and learning curves with the previous model.
# ### 2.2)
# Try to estimate the number of parameters in your network, then check your result with `model.summary()`.
#
# **Tip**: the batch normalization layer adds two parameters for each feature map (the scale and bias).
# 
# ## Exercise 2 – Solution
# ### 2.1)
# Replace the `Conv2D` layers with `SeparableConv2D` layers (except the first one), fit your model and compare its performance and learning curves with the previous model.
model = keras.models.Sequential([
keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu", input_shape=[32, 32, 3]),
keras.layers.BatchNormalization(),
keras.layers.SeparableConv2D(filters=32, kernel_size=3, padding="same", activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.MaxPool2D(pool_size=2),
keras.layers.SeparableConv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.SeparableConv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.MaxPool2D(pool_size=2),
keras.layers.Flatten(),
keras.layers.Dense(128, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=0.01), metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
pd.DataFrame(history.history).plot()
plt.axis([0, 19, 0, 1])
plt.show()
# ### 2.2)
# Try to estimate the number of parameters in your network, then check your result with `model.summary()`.
#
# **Tip**: the batch normalization layer adds two parameters for each feature map (the scale and bias).
# Number of params in a depthwise separable 2D convolution layer =
# kernel_width * kernel_height * channels_in + (channels_in + 1 for bias) * channels_out
(
(3 * 3 * 3 + 1) * 32 # in: 32x32x3 out: 32x32x32 Conv2D
+ 32 * 2 # in: 32x32x32 out: 32x32x32 BN
+ 3 * 3 * 32 + (32 + 1) * 32 # in: 32x32x32 out: 32x32x32 SeparableConv2D
+ 32 * 2 # in: 32x32x32 out: 32x32x32 BN
+ 0 # in: 32x32x32 out: 16x16x32 MaxPool2D
+ 3 * 3 * 32 + (32 + 1) * 64 # in: 16x16x32 out: 16x16x64 SeparableConv2D
+ 64 * 2 # in: 16x16x64 out: 16x16x64 BN
+ 3 * 3 * 64 + (64 + 1) * 64 # in: 16x16x64 out: 16x16x64 SeparableConv2D
+ 64 * 2 # in: 16x16x64 out: 16x16x64 BN
+ 0 # in: 16x16x64 out: 8x8x64 MaxPool2D
+ 0 # in: 8x8x64 out: 4096 Flatten
+ (4096 + 1) * 128 # in: 4096 out: 128 Dense
+ (128 + 1) * 10 # in: 128 out: 10 Dense
)
# Let's check:
model.summary()
# 
# ## Exercise 3 – Pretrained CNNs
# ### 3.1)
# Using `keras.preprocessing.image.load_img()` followed by `keras.preprocessing.image.img_to_array()`, load one or more images (e.g., `fig.jpg` or `ostrich.jpg` in the `images` folder). You should set `target_size=(299, 299)` when calling `load_img()`, as this is the shape that the Xception network expects.
# ### 3.2)
# Create a batch containing the image(s) you just loaded, and preprocess this batch using `keras.applications.xception.preprocess_input()`. Verify that the features now vary from -1 to 1: this is what the Xception network expects.
# ### 3.3)
# Create an instance of the Xception model (`keras.applications.xception.Xception`) and use its `predict()` method to classify the images in the batch. You can use `keras.applications.resnet50.decode_predictions()` to convert the output matrix into a list of top-N predictions (with their corresponding class labels).
# 
# ## Exercise 3 – Solution
# ### 3.1)
# Using `keras.preprocessing.image.load_img()` followed by `keras.preprocessing.image.img_to_array()`, load one or more images (e.g., `fig.jpg` or `ostrich.jpg` in the `images` folder). You should set `target_size=(299, 299)` when calling `load_img()`, as this is the shape that the Xception network expects.
img_fig_path = os.path.join("images", "fig.jpg")
img_fig = keras.preprocessing.image.load_img(img_fig_path, target_size=(299, 299))
img_fig = keras.preprocessing.image.img_to_array(img_fig)
plt.imshow(img_fig / 255)
plt.axis("off")
plt.show()
img_fig.shape
img_ostrich_path = os.path.join("images", "ostrich.jpg")
img_ostrich = keras.preprocessing.image.load_img(img_ostrich_path, target_size=(299, 299))
img_ostrich = keras.preprocessing.image.img_to_array(img_ostrich)
plt.imshow(img_ostrich / 255)
plt.axis("off")
plt.show()
img_ostrich.shape
# ### 3.2)
# Create a batch containing the image(s) you just loaded, and preprocess this batch using `keras.applications.xception.preprocess_input()`. Verify that the features now vary from -1 to 1: this is what the Xception network expects.
X_batch = np.array([img_fig, img_ostrich])
X_preproc = keras.applications.xception.preprocess_input(X_batch)
X_preproc.min(), X_preproc.max()
# ### 3.3)
# Create an instance of the Xception model (`keras.applications.xception.Xception`) and use its `predict()` method to classify the images in the batch. You can use `keras.applications.resnet50.decode_predictions()` to convert the output matrix into a list of top-N predictions (with their corresponding class labels).
model = keras.applications.xception.Xception()
Y_proba = model.predict(X_preproc)
Y_proba.shape
np.argmax(Y_proba, axis=1)
decoded_predictions = keras.applications.resnet50.decode_predictions(Y_proba)
for preds in decoded_predictions:
for wordnet_id, name, proba in preds:
print("{} ({}): {:.1f}%".format(name, wordnet_id, 100 * proba))
print()
# 
# ## Exercise 4 – Data Augmentation and Transfer Learning
# In this exercise you will reuse a pretrained Xception model to build a flower classifier.
# First, let's download the dataset:
# +
import tensorflow as tf
from tensorflow import keras
import os
flowers_url = "http://download.tensorflow.org/example_images/flower_photos.tgz"
flowers_path = keras.utils.get_file("flowers.tgz", flowers_url, extract=True)
flowers_dir = os.path.join(os.path.dirname(flowers_path), "flower_photos")
# -
for root, subdirs, files in os.walk(flowers_dir):
print(root)
for filename in files[:3]:
print(" ", filename)
if len(files) > 3:
print(" ...")
# ### 4.1)
# Build a `keras.preprocessing.image.ImageDataGenerator` that will preprocess the images and do some data augmentation (the [documentation](https://keras.io/preprocessing/image/) contains useful examples):
#
# * It should at least perform horizontal flips and keep 10% of the data for validation, but you may also make it perform a bit of rotation, rescaling, etc.
# * Also make sure to apply the Xception preprocessing function (using the `preprocessing_function` argument).
# * Call this generator's `flow_from_directory()` method to get an iterator that will load and preprocess the flower photos from the `flower_photos` directory, setting the target size to (299, 299) and `subset` to `"training"`.
# * Call this method again with the same parameters except `subset="validation"` to get a second iterator for validation.
# * Get the next batch from the validation iterator and display the first image from the batch.
# ### 4.2)
# Now let's build the model:
# * Create a new `Xception` model, but this time set `include_top=False` to get the model without the top layer. **Tip**: you will need to access its `input` and `output` properties.
# * Make all its layers non-trainable.
# * Using the functional API, add a `GlobalAveragePooling2D` layer (feeding it the Xception model's output), and add a `Dense` layer with 5 neurons and the Softmax activation function.
# * Compile the model. **Tip**: don't forget to add the `"accuracy"` metric.
# * Fit your model using `fit_generator()`, passing it the training and validation iterators (and setting `steps_per_epoch` and `validation_steps` appropriately).
# 
# ## Exercise 4 – Solution
# ### 4.1)
# Build a `keras.preprocessing.image.ImageDataGenerator` that will preprocess the images and do some data augmentation (the [documentation](https://keras.io/preprocessing/image/) contains useful examples):
#
# * It should at least perform horizontal flips and keep 10% of the data for validation, but you may also make it perform a bit of rotation, rescaling, etc.
# * Also make sure to apply the Xception preprocessing function (using the `preprocessing_function` argument).
# * Call this generator's `flow_from_directory()` method to get an iterator that will load and preprocess the flower photos from the `flower_photos` directory, setting the target size to (299, 299) and `subset` to `"training"`.
# * Call this method again with the same parameters except `subset="validation"` to get a second iterator for validation.
# * Get the next batch from the validation iterator and display the first image from the batch.
# +
datagen = keras.preprocessing.image.ImageDataGenerator(
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
validation_split=0.1,
preprocessing_function=keras.applications.xception.preprocess_input)
train_generator = datagen.flow_from_directory(
flowers_dir,
target_size=(299, 299),
batch_size=32,
subset="training")
valid_generator = datagen.flow_from_directory(
flowers_dir,
target_size=(299, 299),
batch_size=32,
subset="validation")
# -
X_batch, y_batch = next(valid_generator)
plt.imshow((X_batch[0] + 1)/2)
plt.axis("off")
plt.show()
# ### 4.2)
# Now let's build the model:
# * Create a new `Xception` model, but this time set `include_top=False` to get the model without the top layer. **Tip**: you will need to access its `input` and `output` properties.
# * Make all its layers non-trainable.
# * Using the functional API, add a `GlobalAveragePooling2D` layer (feeding it the Xception model's output), and add a `Dense` layer with 5 neurons and the Softmax activation function.
# * Compile the model. **Tip**: don't forget to add the `"accuracy"` metric.
# * Fit your model using `fit_generator()`, passing it the training and validation iterators (and setting `steps_per_epoch` and `validation_steps` appropriately).
# +
n_classes = 5
base_model = keras.applications.xception.Xception(include_top=False)
for layer in base_model.layers:
layer.trainable = False
global_pool = keras.layers.GlobalAveragePooling2D()(base_model.output)
predictions = keras.layers.Dense(n_classes, activation='softmax')(global_pool)
model = keras.models.Model(base_model.input, predictions)
model.compile(loss="categorical_crossentropy",
optimizer="sgd", metrics=["accuracy"])
# -
history = model.fit_generator(
train_generator,
steps_per_epoch=3306 // 32,
epochs=50,
validation_data=valid_generator,
validation_steps=364 // 32)
pd.DataFrame(history.history).plot()
plt.axis([0, 19, 0, 1])
plt.show()
# 
# ## Object Detection Project
# The Google [Street View House Numbers](http://ufldl.stanford.edu/housenumbers/) (SVHN) dataset contains pictures of digits in all shapes and colors, taken by the Google Street View cars. The goal is to classify and locate all the digits in large images.
# * Train a Fully Convolutional Network on the 32x32 images.
# * Use this FCN to build a digit detector in the large images.
| 05_cnns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import json
import csv
import pandas as pd
import re
from sklearn.preprocessing import MultiLabelBinarizer
from collections import Counter
# +
year_list = [str(year) for year in range(1978,2019 + 1)]
data = []
with open('../nominate_movie_meta_data.json', 'r', encoding = 'utf-8') as f:
json_dict = json.load(f)
for year in year_list:
data += json_dict[year]
data = pd.DataFrame(data).set_index('id')
def dict2list_other_nominate(series):
l = []
other_nominates = series['other_nominates']
for each in other_nominates:
if each['prized'] == 1:
l.append(each['award'])
return pd.Series([l])
data['other_nominates'] = data.apply(dict2list_other_nominate, axis = 'columns')
data['production_studio'] = data['production_studio'].map(lambda each: re.split(r'[、==]',each))
mlb = MultiLabelBinarizer()
onehot = mlb.fit_transform(data['other_nominates'])
columns = mlb.classes_
nomination_onehot = pd.DataFrame(onehot, columns = columns, index = data.index) #####################
performers = []
data['performers'].map(lambda each: performers.extend(each))
count = Counter(performers)
countSeries = pd.Series(count)
selected_performers = countSeries[countSeries > 9].index ######10回以上受賞作品に出演
data['selected_performers'] = data['performers'].map(lambda each: list( set(each) & set(selected_performers) ))
onehot = mlb.fit_transform(data['selected_performers'])
columns = mlb.classes_
selected_performers_onehot = pd.DataFrame(onehot, columns = columns, index = data.index) #####################10回以上出た出演者のonehot
directors = []
data['director'].map(lambda each: directors.extend(each))
count = Counter(directors)
countSeries = pd.Series(count)
selected_directors = countSeries[countSeries > 3].index
data['selected_directors'] = data['director'].map(lambda each: list( set(each) & set(selected_directors) ))
onehot = mlb.fit_transform(data['selected_directors'])
columns = mlb.classes_
selected_directors_onehot = pd.DataFrame(onehot, columns = columns, index = data.index) #####################4回以上出た監督のonehot
studio = []
data['production_studio'].map(lambda each: studio.extend(each))
count = Counter(studio)
countSeries = pd.Series(count)
selected_studios = countSeries[countSeries > 20].index ########################### 20回以上受賞
data['selected_studio'] = data['production_studio'].map(lambda each: list( set(each) & set(selected_studios) ))
onehot = mlb.fit_transform(data['selected_studio'])
columns = mlb.classes_
selected_studio_onehot = pd.DataFrame(onehot, columns = columns, index = data.index) #####################4回以上出た監督のonehot
scriptwriters = []
data['scriptwriter'].map(lambda each: scriptwriters.extend(each))
count = Counter(scriptwriters)
countSeries = pd.Series(count)
selected_scriptwriters = countSeries[countSeries > 2].index
data['selected_scriptwriter'] = data['scriptwriter'].map(lambda each: list( set(each) & set(selected_scriptwriters) ))
onehot = mlb.fit_transform(data['selected_scriptwriter'])
columns = mlb.classes_
selected_scriptwriter_onehot = pd.DataFrame(onehot, columns = columns, index = data.index) #############
# +
reviews_json = {}
filepaths = ['../coco_reviews.json', '../eigacom_review.json']
for each in data.index:
reviews_json[str(each)] = []
temp_json = {}
for each in filepaths:
with open(each, 'r', encoding= 'utf-8') as f:
temp_json[each] = json.load(f)
for each in data.index:
for file in filepaths:
reviews_json[str(each)].extend(temp_json[file][str(each)]['reviews'])
for each in data.index:
[temp_json[file][str(each)] for file in filepaths]
reviews = [
pd.DataFrame(
reviews_json[str(i)],
columns = ['date', 'review','rating','star'] if len(reviews_json[str(i)]) == 0 else None
) for i in data.index
];
for each in reviews:
each['date'] = pd.to_datetime(each['date'])
review_dataframe = pd.DataFrame(reviews, columns = ['reviews'], index = data.index)
# +
# tfidf
l = []
with open('../nlp/tfidf.csv', 'r', encoding= 'utf-8') as f:
reader = csv.reader(f, delimiter=',')
for nomination_id, row in enumerate(reader, 1):
data_dict = {}
for i, each in enumerate(row):
data_dict[each] = len(row) - i
l.append(pd.DataFrame(data_dict, index = [nomination_id]))
tfidf = pd.concat(l, sort=False)
tfidf.index.name = 'id'
# -
# 保存
"""
data.to_pickle('data.pkl')
nomination_onehot.to_pickle('nomination_onehot.pkl')
selected_performers_onehot.to_pickle('selected_performers_onehot.pkl')
selected_directors_onehot.to_pickle('selected_directors_onehot.pkl')
selected_studio_onehot.to_pickle('selected_studio_onehot.pkl')
selected_scriptwriter_onehot.to_pickle('selected_scriptwriter_onehot.pkl')
review_dataframe.to_pickle('review_dataframe.pkl')
tfidf.to_pickle('tfidf.pkl')
""";
# 読み出し
data = pd.read_pickle('data.pkl')
nomination_onehot = pd.read_pickle('nomination_onehot.pkl')
selected_performers_onehot = pd.read_pickle('selected_performers_onehot.pkl')
selected_directors_onehot = pd.read_pickle('selected_directors_onehot.pkl')
selected_studio_onehot = pd.read_pickle('selected_studio_onehot.pkl')
selected_scriptwriter_onehot = pd.read_pickle('selected_scriptwriter_onehot.pkl')
review_dataframe = pd.read_pickle('review_dataframe.pkl')
tfidf = pd.read_pickle('tfidf.pkl')
| neo_review/scripts/xgb/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="45ab671ce78132c4e89737347df00708faa6ecd4"
# # Microsoft Malware Detection XGBoost Blending CV predictions
#
# 
#
# 
#
# 
#
#
# The Goal For this Kernel is to blend / mix predictions from different folds when training the XGBoost classifier. The idea stems from different Kernels which I want to credit for such as:
#
# - https://www.kaggle.com/artgor/is-this-malware-eda-fe-and-lgb-updated
#
#
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
print(os.listdir("../input"))
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import time
# due to Kaggle memory limitations and the enormous dataset size, a sample from the whole
# trainset will be used for ML modeling
train_sample_fraction = 0.2
# another global variable that must be defined is the NA values rate / theshold to ommit columns with
# NA values that pass this rate
na_rate_threshold = 0.9
# theshold to remove columns with unbalanced features to their values
unbalanced_feature_rate_threshold = 0.9
# Any results you write to the current directory are saved as output.
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# I am grateful for the help of author of this kernel for the main idea to load the dataset and save memory space!!
# https://www.kaggle.com/theoviel/load-the-totality-of-the-data
dtypes = {
'MachineIdentifier': 'category',
'ProductName': 'category',
'EngineVersion': 'category',
'AppVersion': 'category',
'AvSigVersion': 'category',
'IsBeta': 'int8',
'RtpStateBitfield': 'float16',
'IsSxsPassiveMode': 'int8',
'DefaultBrowsersIdentifier': 'float16',
'AVProductStatesIdentifier': 'float32',
'AVProductsInstalled': 'float16',
'AVProductsEnabled': 'float16',
'HasTpm': 'int8',
'CountryIdentifier': 'int16',
'CityIdentifier': 'float32',
'OrganizationIdentifier': 'float16',
'GeoNameIdentifier': 'float16',
'LocaleEnglishNameIdentifier': 'int8',
'Platform': 'category',
'Processor': 'category',
'OsVer': 'category',
'OsBuild': 'int16',
'OsSuite': 'int16',
'OsPlatformSubRelease': 'category',
'OsBuildLab': 'category',
'SkuEdition': 'category',
'IsProtected': 'float16',
'AutoSampleOptIn': 'int8',
'PuaMode': 'category',
'SMode': 'float16',
'IeVerIdentifier': 'float16',
'SmartScreen': 'category',
'Firewall': 'float16',
'UacLuaenable': 'float32',
'Census_MDC2FormFactor': 'category',
'Census_DeviceFamily': 'category',
'Census_OEMNameIdentifier': 'float16',
'Census_OEMModelIdentifier': 'float32',
'Census_ProcessorCoreCount': 'float16',
'Census_ProcessorManufacturerIdentifier': 'float16',
'Census_ProcessorModelIdentifier': 'float16',
'Census_ProcessorClass': 'category',
'Census_PrimaryDiskTotalCapacity': 'float32',
'Census_PrimaryDiskTypeName': 'category',
'Census_SystemVolumeTotalCapacity': 'float32',
'Census_HasOpticalDiskDrive': 'int8',
'Census_TotalPhysicalRAM': 'float32',
'Census_ChassisTypeName': 'category',
'Census_InternalPrimaryDiagonalDisplaySizeInInches': 'float16',
'Census_InternalPrimaryDisplayResolutionHorizontal': 'float16',
'Census_InternalPrimaryDisplayResolutionVertical': 'float16',
'Census_PowerPlatformRoleName': 'category',
'Census_InternalBatteryType': 'category',
'Census_InternalBatteryNumberOfCharges': 'float32',
'Census_OSVersion': 'category',
'Census_OSArchitecture': 'category',
'Census_OSBranch': 'category',
'Census_OSBuildNumber': 'int16',
'Census_OSBuildRevision': 'int32',
'Census_OSEdition': 'category',
'Census_OSSkuName': 'category',
'Census_OSInstallTypeName': 'category',
'Census_OSInstallLanguageIdentifier': 'float16',
'Census_OSUILocaleIdentifier': 'int16',
'Census_OSWUAutoUpdateOptionsName': 'category',
'Census_IsPortableOperatingSystem': 'int8',
'Census_GenuineStateName': 'category',
'Census_ActivationChannel': 'category',
'Census_IsFlightingInternal': 'float16',
'Census_IsFlightsDisabled': 'float16',
'Census_FlightRing': 'category',
'Census_ThresholdOptIn': 'float16',
'Census_FirmwareManufacturerIdentifier': 'float16',
'Census_FirmwareVersionIdentifier': 'float32',
'Census_IsSecureBootEnabled': 'int8',
'Census_IsWIMBootEnabled': 'float16',
'Census_IsVirtualDevice': 'float16',
'Census_IsTouchEnabled': 'int8',
'Census_IsPenCapable': 'int8',
'Census_IsAlwaysOnAlwaysConnectedCapable': 'float16',
'Wdft_IsGamer': 'float16',
'Wdft_RegionIdentifier': 'float16',
'HasDetections': 'int8'
}
def reduce_mem_usage(df, verbose=True):
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
start_mem = df.memory_usage(deep=True).sum() / 1024**2
for col in df.columns:
col_type = df[col].dtypes
if col_type in numerics:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == 'int':
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage(deep=True).sum() / 1024**2
if verbose: print('Mem. usage decreased to {:5.2f} Mb ({:.1f}% reduction)'.format(end_mem, 100 * (start_mem - end_mem) / start_mem))
return df
# + _uuid="a9fa66bdccee4111ac7ed2acde1e2748edb13c46"
# %%time
train = pd.read_csv('../input/train.csv', dtype=dtypes)
# + _uuid="e20433faf47d6bd83456a35f78a3314adbdc7986"
good_cols = list(train.columns)
for col in train.columns:
# remove columns with high NA rate
na_rate = train[col].isnull().sum() / train.shape[0]
# remove columns with high Unbalanced values rate
unbalanced_rate = train[col].value_counts(normalize=True, dropna=False).values[0]
if na_rate > na_rate_threshold:
good_cols.remove(col)
elif unbalanced_rate > unbalanced_feature_rate_threshold:
good_cols.remove(col)
# + _uuid="f60e134c2fcf4e65a0551d6a0b925c1d33f97265"
good_cols
# + _uuid="cd629eab88cebfdb7a354103dc9c6767c6d775ea"
train = train[good_cols]
# + _uuid="7178d430eaccbdf1e099647434e7e44272bba7c9"
import gc
gc.collect()
# + _uuid="2afcda836a7188b58e82ba9a145d689bab1b1c57"
categorical_columns = list(train.loc[:, train.dtypes =="category"].columns)
numerical_and_binary_columns = list(train.loc[:, train.dtypes !="category"].columns)
numerical_columns = numerical_and_binary_columns
categorical_columns.remove("MachineIdentifier")
binary_columns = []
for col in (numerical_and_binary_columns):
if train[col].nunique() == 2:
binary_columns.append(col)
numerical_columns.remove(col)
# + [markdown] _uuid="b4a4eafe356cf04082ab3ee284f0b352048ce50a"
# ## Machine Learning Modeling and Tuning
# + [markdown] _uuid="0a7bcdda6c53f3cea7059178104fdaed58be7c9f"
# 
#
# 
# + _uuid="2fb705fbc15a9009968d7d5a229a5f27ead6e704"
train_sample = train.sample(frac=train_sample_fraction, random_state=42)
del train
gc.collect()
# + _uuid="41fbee6299db8ee267c963d65b7a59336d485098"
train_sample.shape
# + _uuid="0cbf8c7f54df79442941de2afd6cedebed96b1d2"
test_dtypes = {k: v for k, v in dtypes.items() if k in good_cols}
# get all columns except
test = pd.read_csv('../input/test.csv', dtype=test_dtypes, usecols=good_cols[:-1])
#test = reduce_mem_usage(test)
# + _uuid="a3f304766c39f7e2c6c8fda0056f3e1bdbca3eff"
test.head()
# + _uuid="513121ff0b8730ddca7cb469f0219d268b4d4605"
test.shape
# + _uuid="ac15de9d17b067234afdae7e020a6b5de02a7a70"
train_sample = train_sample.drop(['MachineIdentifier'], axis=1)
test = test.drop(['MachineIdentifier'], axis=1)
# + _uuid="a83ccfddbb26500e88c312778908a35753d0cdcf"
train_sample = train_sample.reset_index(drop=True)
# + [markdown] _uuid="5a140c3ce337be0bfe665626d6b3ae2c270056c0"
# ### Filling NA values with the statistical Mode
# + _uuid="5baca4dd7ec3d6a07a53274671334a2f786f28fc"
modes = train_sample.mode()
for col in train_sample.columns:
train_sample[col] = np.where(train_sample[col].isnull(), modes[col], train_sample[col])
del modes
# + _uuid="6ac26a55808e7cc4731241e4eeb625724dd36a24"
modes_test = test.mode()
for col in test.columns:
test[col] = np.where(test[col].isnull(), modes_test[col], test[col])
#train_sample.shape
del modes_test
# + [markdown] _uuid="5cc83fc5edbee697391c02c63f36478584444ec6"
# ### Concatenate both train_sample and test sets before label encoding
# + _uuid="<KEY>"
train_shape = train_sample.shape
test_shape = test.shape
train_and_test = pd.concat([train_sample,test], axis="rows", sort=False)
del train_sample
del test
gc.collect()
# + _uuid="4f51e3d3dafc1bd44f24993bdbe838f16eb9e05c"
train_and_test.head()
# + _uuid="18ce9f155121b34b9d0f01952f912bf6edbd5b80"
train_and_test.tail()
# + [markdown] _uuid="3b53abfda614ae7c587c7701d6ec1ff602660322"
# ### Encode the Categorical features before machine learning modeling
# + _uuid="6ca351ce6699aa9c16555e0d4fc88bcc8a03d515"
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
def MultiLabelEncoder(columnlist,dataframe):
for i in columnlist:
#print(i)
labelencoder_X=LabelEncoder()
dataframe[i]=labelencoder_X.fit_transform(dataframe[i])
MultiLabelEncoder(categorical_columns, train_and_test)
# + _uuid="8ebf152f7d3859c64b8f5a59de631ef2283ec51d"
gc.collect()
# + [markdown] _uuid="cbc688ee912088949b5ad0457d8fd65e0bfb5ac0"
# ### Back to train and test set after Label Encoding
# + _uuid="f32ea815d2d1f28940d9beaf7c2b851fc65837a6"
train_sample = train_and_test[0:train_shape[0]]
test = train_and_test[(train_shape[0]):(train_and_test.shape[0]+1)]
# + _uuid="18c499bed0d4f25fb6895332ce7f375b134c1201"
del train_and_test
# + [markdown] _uuid="b53deef880d7a28701805c1a626e5976be578b2a"
# ### Remove the HasDetections columns from test set, it has been added during dataframe concatenation.
# + _uuid="364fe1f26469600443c904641ba009a3518f282d"
test = test.drop(["HasDetections"], axis = 1)
# + _uuid="3a1c7d626d0aa12f9809e2f077a1856ebccffd2c"
y = train_sample['HasDetections']
X = train_sample.drop(['HasDetections'], axis=1)
# + _uuid="bf909b41fa2e2a66dee6c95f483c67a78a5e648a"
del train_sample
gc.collect()
# + [markdown] _uuid="70772cfcf66c4def9848474a92299cc78a64a2d1"
# ### XGBoost Baseline model and blending Folds
# + _uuid="8fab97ec9c5a8c38155e5e1c77480c28f204e9f6"
# main idea:
# https://www.kaggle.com/infinitewing/k-fold-cv-xgboost-example-0-28?scriptVersionId=1553202
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
import xgboost as xgb
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import f1_score
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score
from sklearn import metrics
import time
import random
K = 5
index = 0
predictions_proba_test_list = np.zeros(len(test))
fold_auc_list = []
fold_accuracy_list = []
kf = KFold(n_splits = K, random_state = 42, shuffle = True)
for train_index, test_index in kf.split(X):
print("Fold:", index)
index = index + 1
train_X, valid_X = X.iloc[train_index, :], X.iloc[test_index, :]
train_y, valid_y = y[train_index], y[test_index]
new_seed = random.randint(1, 2000)
clf_xgb = xgb.XGBClassifier(learning_rate=0.03,
n_estimators=1300,
max_depth=8,
min_child_weight=4,
gamma=0,
subsample=0.8,
colsample_bytree=0.7,
objective= 'binary:logistic',
nthread=-1,
scale_pos_weight=1,
reg_alpha = 0.1,
reg_lambda = 1,
seed=new_seed)
clf_xgb.fit(train_X, train_y, eval_set=[(train_X, train_y), (valid_X, valid_y)],
early_stopping_rounds=100, eval_metric='auc', verbose=100)
temp_predictions_proba_test_list = []
# read test set in chunks
chunck = 400000
test_times = test.shape[0] // chunck
test_rest = test.shape[0] % chunck
for i in np.arange(0,(chunck * (test_times+1)), chunck):
# create predictions in chunks due ot memory limitations
predictions_proba_test = list(clf_xgb.predict_proba(test[i:(i+chunck)])[:,1])
temp_predictions_proba_test_list.append(predictions_proba_test)
#print("times:", i)
# flatten the list of lists
temp_predictions_proba_test_list = [y for x in temp_predictions_proba_test_list for y in x]
#print(np.shape(predictions_proba_test_list))
predictions_proba_test_list = [sum(x) for x in zip(predictions_proba_test_list, temp_predictions_proba_test_list)]
#print(test.shape)
#print(np.shape(predictions_proba_test_list))
predictions = clf_xgb.predict(valid_X, ntree_limit=clf_xgb.n_estimators)
print()
print(classification_report(valid_y, predictions))
print()
print("accuracy_score", accuracy_score(valid_y, predictions))
predictions_probas = clf_xgb.predict_proba(valid_X)[:,1]
print("auc score", roc_auc_score(valid_y, predictions_probas))
print()
fold_accuracy_list.append(accuracy_score(valid_y, predictions))
fold_auc_list.append(roc_auc_score(valid_y, predictions_probas))
print()
print("Mean auc:", np.mean(fold_auc_list))
print("Std auc:", np.std(fold_auc_list))
print("Mean accuracy:", np.mean(fold_accuracy_list))
print("Std accuracy:", np.std(fold_accuracy_list))
gc.collect()
# + _uuid="48105472ae1b6f3ced834d90fe46098368dc96ca"
predictions_proba_test_list = [x / kf.n_splits for x in predictions_proba_test_list]
# + _uuid="dc687a13542e759b06b39e0561156c74ce2cd364"
from sklearn.metrics import confusion_matrix
import scikitplot as skplt
sns.set(rc={'figure.figsize':(8,8)})
skplt.metrics.plot_confusion_matrix(valid_y, predictions, cmap="BrBG")
# + _uuid="02492c7c98e2d3032e002f340ab66ced0b7207d9"
sns.set(rc={'figure.figsize':(8,8)})
predictions_probas = clf_xgb.predict_proba(valid_X)
skplt.metrics.plot_roc(valid_y, predictions_probas)
# + _uuid="984bbd49fd0884e96fb6df070b02488e2f32e9e9"
sns.set(rc={'figure.figsize':(8,8)})
skplt.metrics.plot_ks_statistic(valid_y, predictions_probas)
# + _uuid="acc1ec59d4522c5317849a06c93ddd1a4401b4fa"
sns.set(rc={'figure.figsize':(8,8)})
skplt.metrics.plot_precision_recall(valid_y, predictions_probas)
# + [markdown] _uuid="502f36a6106fb160d505c1f4a644a33b6f4baf74"
# ### Tuning
# #### The following part is commented, I have run tuning in previous versions of this kernel and figured out the optimal (so far) values of the xgboost parameters.
#
# ### XGBoost Grid Search Part 1
# #### Tuning parameters:
# - 'max_depth'
# - 'min_child_weight'
# + _uuid="3ad65deff0365c802e4972b82b52856ae77aa33e"
# I may implement tuning in the future but I am afraid of variance - bias tradeoff
'''
#idea and a big thank you to https://www.analyticsvidhya.com/blog/2016/03/complete-guide-parameter-tuning-xgboost-with-codes-python/
from sklearn.model_selection import GridSearchCV #Perforing grid search
gc.collect()
param_test1 = {
# based on previous personal kernels both parameters show better result having high numbers
'max_depth':[3, 5, 7, 9, 11],
'min_child_weight':[1, 3, 5, 7, 9]
}
gsearch1 = GridSearchCV(estimator = xgb.XGBClassifier( learning_rate=0.05, n_estimators=70, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=-1, scale_pos_weight=1, reg_alpha = 0,
reg_lambda =1, seed=42),
param_grid = param_test1, scoring='roc_auc', n_jobs=1, iid=False, cv=3, verbose = 1)
gsearch1.fit(xtrain, ytrain)
gsearch1.best_params_, gsearch1.best_score_
'''
# + [markdown] _uuid="ad276e45246c6eb85273133b1c1c25054a353e47"
# ### Prepare for submission
# + _uuid="a4e3263d4bf6a80b92f0c14e5ab358c45ff26864"
del X
del y
del train_X
del train_y
del valid_X
del valid_y
del predictions
del predictions_probas
del temp_predictions_proba_test_list
del clf_xgb
gc.collect()
# + _uuid="3a984b61c5bd6e9881f76b9a281aae6bb4e22930"
submission = pd.read_csv('../input/sample_submission.csv')
submission['HasDetections'] = predictions_proba_test_list
submission.to_csv('xgboost.csv', index=False)
# + _uuid="e163232fc5e4ee1cd248ccf8d319a399912b8239"
# + _uuid="155ba1e850542bb709a19e03f1c177a4b9c1e02e"
| microsoft-malware-detection-xgboost-blends.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''base'': conda)'
# name: python3
# ---
# Description: implementation required by Question 2 of Coursework 2
#
# Version: 1.0.3.20210709
#
# Author: <NAME>
# **NOTE:** All locations are given as `[longitude, latitude]` in this work.
# ## Importing modules
# +
import json
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# -
# ## Reading data from the given dataset
#
# The given dataset contains JSON objects. Each line represents a tweet object. We can read them as raw data, load the data as a DataFrame object, and display each tweet object's coordinates.
#
# For this part only, `data = pd.read_json('Data/geoLondonAug2020-1', lines = True)` to read the file as a JSON object per line can be sufficient. The `coordinates` values containing multiple levels (e.g., `coordinates.coordinates`) are flattened here to make it convenient for further processing.
# + tags=[]
data = []
id_col = 'ID'
coordinate_col = 'Coordinates'
for tweet in open('Data/geoLondonAug2020-1', encoding = 'utf-8'):
data.append(json.loads(tweet))
data = pd.json_normalize(data)
data.rename(columns = {'_id': id_col, 'coordinates.coordinates': coordinate_col}, inplace = True)
data[[id_col, coordinate_col]]
# -
# ## Implementing the algorithm of the Haversine distance
def compute_hav_distance(loc_1: list, loc_2: list) -> float:
'''
Calculate the Haversine distance between two location points.
Parameters
----------
loc_1 : a location point given as a pair of longitude and latitude in a list
loc_2 : another location point given as a pair of longitude and latitude in a list
Returns
-------
hav_distance : the haversine distance in kilometres between two location points
'''
long_1, lat_1, long_2, lat_2 = map(np.radians, [loc_1[0], loc_1[1], loc_2[0], loc_2[1]]) # Covert angles from degrees to radians.
hav = np.sin((lat_2 - lat_1) / 2) ** 2 + np.cos(lat_1) * np.cos(lat_2) * np.sin((long_2 - long_1) / 2) ** 2 # Compute the haversine of the central angle.
return 6371 * 2 * np.arcsin(np.sqrt(hav))
# ## Dividing a specified London area into grids of 1 km * 1 km
#
# The specified rectangle area of London is determined by two coordinates - `[-0.563, 51.261318]` and `[0.28036, 51.686031]`.
long_1, lat_1 = [-0.563, 51.261318]
long_2, lat_2 = [0.28036, 51.686031]
n_row = np.ceil(compute_hav_distance([long_1, lat_1], [long_1, lat_2])).astype(int)
n_col = np.ceil(compute_hav_distance([long_1, lat_1], [long_2, lat_1])).astype(int)
n_grid = n_row * n_col
print('The number of rows:', n_row)
print('The number of columns:', n_col)
print('The number of grids:', n_grid)
# ## Assigning tweets to grids
# + tags=[]
grids_map = np.zeros((n_row, n_col), dtype = int)
for coordinate in data[coordinate_col]:
row_index = np.ceil(compute_hav_distance([long_1, lat_1], [long_1, coordinate[1]])).astype(int)
col_index = np.ceil(compute_hav_distance([long_1, lat_1], [coordinate[0], lat_1])).astype(int)
grids_map[row_index, col_index] += 1
# -
# ## Showing the number of tweets per grid
#
# The grids are numbered from 1. All grids containing at least 1 tweet are listed in a table.
n_tweets_list = np.ravel(grids_map)
grid_col = 'Grid'
n_tweets_col = 'Tweets count'
n_tweets_df = pd.DataFrame({grid_col: np.arange(1, n_grid + 1), n_tweets_col: n_tweets_list}) # Create a DataFrame containing the number of tweets per grid.
n_tweets_df[n_tweets_df[n_tweets_col] != 0].style.hide_index()
# ## Drawing a heatmap
#
# The location `[-0.563, 51.261318]` is used as the coordinate system's origin.
#
# **NOTE:**
#
# 1. The axes are displayed in a way reflecting the actual situation (0 on the x-axis would on the top by default).
# 2. The logarithm is applied to the number of tweets per grid to make the heatmap more readable.
heatmap_fig, heatmap_ax = plt.subplots(figsize = (15, 18))
heatmap_ax.set_title('Heatmap of the logarithm of the number of tweets per grid')
heatmap_ax.set_xticks(np.arange(n_col))
heatmap_ax.set_yticks(np.arange(n_row))
grids_map_log = np.log(grids_map, out = np.zeros_like(grids_map, dtype = float), where = (grids_map != 0)) # Apply logarithm to the number of tweets per grid.
heatmap_img = heatmap_ax.imshow(grids_map_log, cmap = 'GnBu', origin = 'lower')
heatmap_fig.colorbar(heatmap_img, cax = make_axes_locatable(heatmap_ax).append_axes('right', size='5%', pad=0.05)) # Format the colour bar.
plt.show()
# ## Drawing a histogram
#
# It can be observed from the heatmap that multiple grids contain no tweet at all, causing that the first bar in the histogram could be incredibly tall.
#
# **NOTE:** The logarithm is applied to the number of grids to increase readability.
hist_fig, hist_ax = plt.subplots(figsize = (16, 8))
hist_ax.set_title('Tweet distribution (the logarithm applied to the y-axis)')
hist_ax.set_xlabel('Number of tweets')
hist_ax.set_ylabel('Logarithm of the number of grids')
hist_ax.hist(n_tweets_list, bins = np.max(n_tweets_list), log = True)
plt.show()
| Coursework2/Q2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Exercises
#
# #### 1. Load Credit Risk dataset
credit_data <- read.csv("credit.data.manage.csv", sep=",", stringsAsFactors=TRUE)
summary(credit_data)
head(credit_data)
# <p>
# <b>2. Replace negative values in Age column with median age.</b>
# </p>
# calculate how many values column Age has
dim(credit_data["Age"])
# calculate the number of negative values in Age column
sum(credit_data["Age"] < 0)
# So, there are 116 negatiove values in the `Age` column
# calculate median age
median_age = median(credit_data[,2])
median_age
# import dplyr package to use %>%
library(dplyr)
# replace the negative values in the Age column with it's median
credit_data <- credit_data %>%
mutate(Age = ifelse(Age < 0, median_age, Age))
# count the negative values again
sum(credit_data["Age"] < 0)
# <p>
# <b>3. Using IQR rule and empirical rule with −2.5𝜎 and 2.5𝜎, determine the valid range of Credit.amount column. Use only positive values when determining the valid range</b>
# </p>
#
#
# ## determine outlier using IQR
# +
# defining and calculating the 1st quartile q1
q1 <- quantile(credit_data$Credit.amount, 0.25)
# defining and calculating the 3rd quartile q3
q3 <- quantile(credit_data$Credit.amount, 0.75)
q3
# -
iqr_rule <- function(q1, q3){
#calculate iqr and bound limits
iqr <<- (q3 - q1)
low_bound <<- (q1 - (1.5*iqr))
up_bound <<- (q3 + (1.5*iqr))
#printing
cat("iqr:", iqr,
"\nlower bound:", low_bound,
"\nupper bound:", up_bound)
}
iqr_rule(q1, q3)
cat("iqr:", iqr,
"\nlower bound:", low_bound,
"\nupper bound:", up_bound)
# For IQR rule, the valid range for `Credit.amount` is between 0 and the upper bound (7762.625)
# +
# calculate how many data points are below the lower bound
# and above the upper bound
sum(credit_data["Credit.amount"] < low_bound)
sum(credit_data["Credit.amount"] > up_bound)
# -
# <mark>Now we know that there are `24` and `72` data points which are respectively below and above the bound limit set by the IQR rule. So, that makes it `96` data points that are considered as outliers</mark>
library(ggplot2)
library(MASS)
install.packages("MASS")
# plot the distribution of Credit.amount
ggplot(credit_data, aes(x = Credit.amount)) +
geom_histogram(alpha = 1, bins = 50, fill = "#d6a51e") +
geom_vline(aes(xintercept = mean(Credit.amount, na.rm = FALSE)),
colour = "red", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = low_bound),
colour = "#2c4fdb", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = up_bound),
colour = "#2c4fdb", linetype = "longdash", size = .8)
# ## determine outlier by using empirical rule (Chebyshev's Theorem)
#
# Chebychshev's Theorem: For any number $k$ greater than 1, at least $1-(\frac{1}{k^2})$ of the data values lie within $k$ standard deviation of the mean
#
# In this case, the $k$ is set as ($k=2.5$). Thus, the range of value becomes:
#
# $$(\mu-2.5\sigma)<\mu < (\mu+2.5\sigma)$$
#
# In addition, for the context of credit amount, the valid and meaningful value should be positive integer. The negative value could indicates debts thus omitted for the range of valid credit amount values.
#
# Therefore, the range of our valid credit amount adhering to the empirical rule becomes:
#
# $$0 < \ Credit.amount_{valid} \ < (\mu+2.5\sigma)$$
#
# define a function to calculate limits set by the emprical rule
empirical_rule <- function(k, mean, stdev){
#calculate lower and upper limit
empirical_up_limit <<- (mean+(k*stdev))
empirical_low_limit <<- (mean-(k*stdev))
#print
cat("Lower limit:", empirical_low_limit,
"\nUpper limit:", empirical_up_limit)
}
# calculate range limits using k=2.5
empirical_rule(2.5, mean(credit_data[,6]), sd(credit_data[,6]))
# Thus, the valid range for the `Credit.amount` column is:
#
# $$ $0 \ < \ Credit.amount_{valid} \ < \ $10957$$
# plot the distribution of Credit.amount
ggplot(credit_data, aes(x = Credit.amount)) +
geom_histogram(alpha = 1, bins = 50, fill = "#d6a51e") +
geom_vline(aes(xintercept = mean(Credit.amount, na.rm = FALSE)),
colour = "red", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = low_bound),
colour = "#2c4fdb", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = up_bound),
colour = "#2c4fdb", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = empirical_up_limit),
colour = "#a949cc", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = empirical_low_limit),
colour = "#a949cc", linetype = "longdash", size = .8)
# #### interesting finding:
# - the empirical rule seems a better measure to use for this particular dataset. When assessing the lower limit set by both empirical and IQR rule, the values between the purple and blue line doesn't really looks like an outlier visually. However, the IQR rule here omits those values.
#
# #### 4. Explain what to be done with the outliers in Credit.amount column
#
# - The outliers should be dropped before this dataset being used for modeling
# #### 5. Replace negative values in Credit.amount column with median value
#
# calculate the number of negative values in Credit.amount column
sum(credit_data["Credit.amount"] < 0)
cat("Percentage of negative values in Credit.amount:",
(sum(credit_data["Credit.amount"]<0))/
(nrow(credit_data["Credit.amount"]))*100, "%")
# calculate the median valeue for Credit.amount
median_cred_amount = median(credit_data[,6])
median_cred_amount
# replace the negative values in the Credit.amount column with it's median
credit_data <- credit_data %>%
mutate(Credit.amount = ifelse(Credit.amount < 0, median_cred_amount, Credit.amount))
# calculate the number of negative values in Credit.amount column again
sum(credit_data["Credit.amount"] < 0)
# +
# recalculate some parameters after replacing negative values
# defining and calculating the updated 1st quartile
q1_new <- quantile(credit_data$Credit.amount, 0.25)
# defining and calculating the updated 3rd quartile
q3_new <- quantile(credit_data$Credit.amount, 0.75)
# -
# recalculate mean and limits after replacing negative values
empirical_rule(2.5, mean(credit_data[,6]), sd(credit_data[,6]))
iqr_rule(q1_new, q3_new)
# plot the distribution of Credit.amount after updating
ggplot(credit_data, aes(x = Credit.amount)) +
geom_histogram(alpha = 1, bins = 50, fill = "#d6a51e") +
geom_vline(aes(xintercept = mean(Credit.amount, na.rm = FALSE)),
colour = "red", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = low_bound),
colour = "#2c4fdb", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = up_bound),
colour = "#2c4fdb", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = empirical_up_limit),
colour = "#a949cc", linetype = "longdash", size = .8) +
geom_vline(aes(xintercept = empirical_low_limit),
colour = "#a949cc", linetype = "longdash", size = .8)
# **The bound limits also changed!**
# #### 5. Derive a new attribute called Credit amount per duration attribute.
head(credit_data[,6])
unique(credit_data[,7])
# +
# defining credit amount per duration attribute
credit_data$credit_per_duration <- credit_data[,6]/credit_data[,7]
# -
head(credit_data)
| Courses (USM)/CDS501 Principles & Practices of Data Science/Lab 4/Lab 4 exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
m,n =200,3
adv = (100+100*np.random.rand(m,n)).round(2)
adv_df = pd.DataFrame(adv,columns=['TV','Radio','Newspaper'])
adv_df.head()
sales = (1 + 2*adv[:,0:1] + 3*adv[:,1:2] + 4*adv[:,2:3] + np.random.randn(m,1)).round(2)
# np.concatenate((adv,sales))
sales[:5]
plt.plot(adv[:, 2:3], sales, 'b.')
plt.xlabel('Newspaper')
plt.ylabel('Sales')
plt.show()
adv
X = np.c_[np.ones((m,1)),adv]
X[:5]
sales.shape
y = sales
theta_hat = (np.linalg.inv(X.T.dot(X))).dot(X.T.dot(y))
theta_hat
# ## Predicted value
example = X[100].dot(theta_hat)
example
# ## Actual value
sales[100]
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(adv,y)
lin_reg.coef_
lin_reg.intercept_
lin_reg.predict([adv[100]])
| Understanding Python/Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kishkath/Data_Structures-Hashing-/blob/main/iNeuron_Ann.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="2QyXyARnt-2Q"
# + colab={"base_uri": "https://localhost:8080/"} id="BNbvdmEBudcm" outputId="605e0e49-3028-4d59-f575-6e66671f96ff"
from google.colab import drive
drive.mount('/content/drive')
# + id="Su3QcLfXuekX"
import tensorflow as tf
from tensorflow import keras
# + id="zljBxw4vv15B"
import os
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="El989L-5v7cc" outputId="9e574774-b388-4f19-ab46-532cc7a370fd"
tf.keras.__version__
# + colab={"base_uri": "https://localhost:8080/"} id="VocuL9kFwW1_" outputId="377b31bf-cf02-47b4-bab2-8dc779d3c624"
tf.config.list_physical_devices('GPU')
# + id="6CiR56W2weN2"
| iNeuron_Ann.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# ## Obtain the train and test data
# +
train = pd.read_csv('UCI_HAR_dataset/csv_files/train.csv')
test = pd.read_csv('UCI_HAR_dataset/csv_files/test.csv')
print(train.shape, test.shape)
# -
train.head(3)
# get X_train and y_train from csv files
X_train = train.drop(['subject', 'Activity', 'ActivityName'], axis=1)
y_train = train.ActivityName
# get X_test and y_test from test csv file
X_test = test.drop(['subject', 'Activity', 'ActivityName'], axis=1)
y_test = test.ActivityName
print('X_train and y_train : ({},{})'.format(X_train.shape, y_train.shape))
print('X_test and y_test : ({},{})'.format(X_test.shape, y_test.shape))
#
#
# # Let's model with our data
# ### Labels that are useful in plotting confusion matrix
labels=['LAYING', 'SITTING','STANDING','WALKING','WALKING_DOWNSTAIRS','WALKING_UPSTAIRS']
# ### Function to plot the confusion matrix
# +
import itertools
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
plt.rcParams["font.family"] = 'DejaVu Sans'
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=90)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# -
# ### Generic function to run any model specified
# +
from datetime import datetime
def perform_model(model, X_train, y_train, X_test, y_test, class_labels, cm_normalize=True, \
print_cm=True, cm_cmap=plt.cm.Greens):
# to store results at various phases
results = dict()
# time at which model starts training
train_start_time = datetime.now()
print('training the model..')
model.fit(X_train, y_train)
print('Done \n \n')
train_end_time = datetime.now()
results['training_time'] = train_end_time - train_start_time
print('training_time(HH:MM:SS.ms) - {}\n\n'.format(results['training_time']))
# predict test data
print('Predicting test data')
test_start_time = datetime.now()
y_pred = model.predict(X_test)
test_end_time = datetime.now()
print('Done \n \n')
results['testing_time'] = test_end_time - test_start_time
print('testing time(HH:MM:SS:ms) - {}\n\n'.format(results['testing_time']))
results['predicted'] = y_pred
# calculate overall accuracty of the model
accuracy = metrics.accuracy_score(y_true=y_test, y_pred=y_pred)
# store accuracy in results
results['accuracy'] = accuracy
print('---------------------')
print('| Accuracy |')
print('---------------------')
print('\n {}\n\n'.format(accuracy))
# confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred)
results['confusion_matrix'] = cm
if print_cm:
print('--------------------')
print('| Confusion Matrix |')
print('--------------------')
print('\n {}'.format(cm))
# plot confusin matrix
plt.figure(figsize=(8,8))
plt.grid(b=False)
plot_confusion_matrix(cm, classes=class_labels, normalize=True, title='Normalized confusion matrix', cmap = cm_cmap)
plt.show()
# get classification report
print('-------------------------')
print('| Classifiction Report |')
print('-------------------------')
classification_report = metrics.classification_report(y_test, y_pred)
# store report in results
results['classification_report'] = classification_report
print(classification_report)
# add the trained model to the results
results['model'] = model
return results
# -
# ### Method to print the gridsearch Attributes
# +
def print_grid_search_attributes(model):
# Estimator that gave highest score among all the estimators formed in GridSearch
print('--------------------------')
print('| Best Estimator |')
print('--------------------------')
print('\n\t{}\n'.format(model.best_estimator_))
# parameters that gave best results while performing grid search
print('--------------------------')
print('| Best parameters |')
print('--------------------------')
print('\tParameters of best estimator : \n\n\t{}\n'.format(model.best_params_))
# number of cross validation splits
print('---------------------------------')
print('| No of CrossValidation sets |')
print('--------------------------------')
print('\n\tTotal numbre of cross validation sets: {}\n'.format(model.n_splits_))
# Average cross validated score of the best estimator, from the Grid Search
print('--------------------------')
print('| Best Score |')
print('--------------------------')
print('\n\tAverage Cross Validate scores of best estimator : \n\n\t{}\n'.format(model.best_score_))
# -
#
#
#
# # 1. Logistic Regression with Grid Search
# +
from sklearn import linear_model
from sklearn import metrics
from sklearn.model_selection import GridSearchCV
# +
# start Grid search
parameters = {'C':[0.01, 0.1, 1, 10, 20, 30], 'penalty':['l2','l1']}
log_reg = linear_model.LogisticRegression()
log_reg_grid = GridSearchCV(log_reg, param_grid=parameters, cv=3, verbose=1, n_jobs=-1)
log_reg_grid_results = perform_model(log_reg_grid, X_train, y_train, X_test, y_test, class_labels=labels)
# -
plt.figure(figsize=(8,8))
plt.grid(b=False)
plot_confusion_matrix(log_reg_grid_results['confusion_matrix'], classes=labels, cmap=plt.cm.Greens, )
plt.show()
# observe the attributes of the model
print_grid_search_attributes(log_reg_grid_results['model'])
#
#
#
# # 2. Linear SVC with GridSearch
from sklearn.svm import LinearSVC
parameters = {'C':[0.125, 0.5, 1, 2, 8, 16]}
lr_svc = LinearSVC(tol=0.00005)
lr_svc_grid = GridSearchCV(lr_svc, param_grid=parameters, n_jobs=-1, verbose=1)
lr_svc_grid_results = perform_model(lr_svc_grid, X_train, y_train, X_test, y_test, class_labels=labels)
print_grid_search_attributes(lr_svc_grid_results['model'])
# # 3. Kernel SVM with GridSearch
from sklearn.svm import SVC
parameters = {'C':[2,8,16],\
'gamma': [ 0.0078125, 0.125, 2]}
rbf_svm = SVC(kernel='rbf')
rbf_svm_grid = GridSearchCV(rbf_svm,param_grid=parameters, n_jobs=-1)
rbf_svm_grid_results = perform_model(rbf_svm_grid, X_train, y_train, X_test, y_test, class_labels=labels)
print_grid_search_attributes(rbf_svm_grid_results['model'])
# # 4. Decision Trees with GridSearchCV
from sklearn.tree import DecisionTreeClassifier
parameters = {'max_depth':np.arange(3,10,2)}
dt = DecisionTreeClassifier()
dt_grid = GridSearchCV(dt,param_grid=parameters, n_jobs=-1)
dt_grid_results = perform_model(dt_grid, X_train, y_train, X_test, y_test, class_labels=labels)
print_grid_search_attributes(dt_grid_results['model'])
# # 5. Random Forest Classifier with GridSearch
from sklearn.ensemble import RandomForestClassifier
params = {'n_estimators': np.arange(10,201,20), 'max_depth':np.arange(3,15,2)}
rfc = RandomForestClassifier()
rfc_grid = GridSearchCV(rfc, param_grid=params, n_jobs=-1)
rfc_grid_results = perform_model(rfc_grid, X_train, y_train, X_test, y_test, class_labels=labels)
print_grid_search_attributes(rfc_grid_results['model'])
#
# # 7. Comparing all models
# +
print('\n Accuracy Error')
print(' ---------- --------')
print('Logistic Regression : {:.04}% {:.04}%'.format(log_reg_grid_results['accuracy'] * 100,\
100-(log_reg_grid_results['accuracy'] * 100)))
print('Linear SVC : {:.04}% {:.04}% '.format(lr_svc_grid_results['accuracy'] * 100,\
100-(lr_svc_grid_results['accuracy'] * 100)))
print('rbf SVM classifier : {:.04}% {:.04}% '.format(rbf_svm_grid_results['accuracy'] * 100,\
100-(rbf_svm_grid_results['accuracy'] * 100)))
print('DecisionTree : {:.04}% {:.04}% '.format(dt_grid_results['accuracy'] * 100,\
100-(dt_grid_results['accuracy'] * 100)))
print('Random Forest : {:.04}% {:.04}% '.format(rfc_grid_results['accuracy'] * 100,\
100-(rfc_grid_results['accuracy'] * 100)))
# -
# > We can choose ___Logistic regression___ or ___Linear SVC___ or ___rbf SVM___.
# # Conclusion :
# In the real world, domain-knowledge, EDA and feature-engineering matter most.
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
| HAR_PREDICTION_MODELS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Kaprekar's constant
#
# The number 6174 is known as Kaprekar's contant, after the mathematician who discovered an associated property: for all four-digit numbers with at least two distinct digits, repeatedly applying a simple procedure eventually results in this value.
#
# The procedure is as follows:
#
# - For a given input x, create two new numbers that consist of the digits in x in ascending and descending order.
# - Subtract the smaller number from the larger number.
#
# For example, this algorithm terminates in three steps when starting from 1234:
#
# ```js
# 4321 - 1234 = 3087
# 8730 - 0378 = 8352
# 8532 - 2358 = 6174
# ```
# Write a function that returns how many steps this will take for a given input N.
# ## Solution
# To solve this imperatively, we can implement a while loop that continually runs the procedure described above until obtaining the number 6174.
#
# For each iteration of the loop we will increment a counter for the number of steps, and return this value at the end.
#
# We also use a helper function that prepends zeros if necessary so that the number always remains four digits long, before creating the ascending and descending integers.
# +
def get_digits(n):
digits = str(n)
if len(digits) == 4:
return digits
else:
return '0' * (4 - len(digits)) + digits
def count_steps(n):
count = 0
while n != 6174:
n = int(''.join(sorted(get_digits(n), reverse=True))) - int(''.join(sorted(get_digits(n))))
count += 1
return count
# -
count_steps(12)
### Recursive solution
def count_steps(n, steps=0):
if n == 6174:
return steps
num = int(''.join(sorted(get_digits(n), reverse=True))) - int(''.join(sorted(get_digits(n))))
return count_steps(num, steps + 1)
count_steps(1234)
| others/kaprekar's_constant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#based on pierian_data - Machine Learning course on Udemy
# -
# ## K Nearest Neighbors
#
# Based on given features of data preparing a KNN model to predict the target class
#import necessary libraries
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# Loading data
df = pd.read_csv("Classified Data",index_col=0)
#Check head of the df
df.head()
#check the details of the data
df.info()
# ### Variable Standardisation
#
# KNN classifier gives output predictions based on the observations that are near. If variables are on a large scale they will have large effect on the distance between observations than the variables in a small scale, which will reflect on the classifier performance. Hence it is really important to standardise variables in a KNN data.
from sklearn.preprocessing import StandardScaler
#creating an instance of StandardScaler
scaler = StandardScaler()
#compute the mean and standad deviation of the data
scaler.fit(df.drop('TARGET CLASS',axis=1))
#standardising the data based on the mean computed
scaled_features = scaler.transform(df.drop('TARGET CLASS',axis=1))
scaled_features
#output of scaling is an array which needs to be converted back to a dataframe
df_feat = pd.DataFrame(scaled_features,columns=df.columns[:-1])
df_feat.head()
# ### Perform Train Test Split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(scaled_features,df['TARGET CLASS'],
test_size=0.30, random_state = 101)
# ### Using KNNClassifier
from sklearn.neighbors import KNeighborsClassifier
#n_neighbors indicate value of k
knn = KNeighborsClassifier(n_neighbors=1)
#fit the data
knn.fit(X_train,y_train)
#prediction
pred = knn.predict(X_test)
# ### Evaluating the model
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,pred))
print(classification_report(y_test,pred))
# ## Choosing a K value
#
# Using elbow method to come up with a suitable K value.
# +
error_rate = []
#Trying differet values and saving the error
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train,y_train)
pred_i = knn.predict(X_test)
error_rate.append(np.mean(pred_i != y_test))
# -
#Plotting the error rate to do visual analysis(EDA)
plt.figure(figsize=(10,6))
plt.plot(range(1,40),error_rate,color='blue', linestyle='dashed',
marker='o', markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
# Here we can see that that after arouns K>23 the error rate just tends to hover around 0.06-0.05 Let's retrain the model with that and check the classification report!
# +
# NOW WITH K=23
knn = KNeighborsClassifier(n_neighbors=23)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
print('WITH K=23')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
# -
| MachineLearning/04-KNN/KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Projeto: Teste Técnico
# ## Desafio
#
# Faça uma análise exploratória para avaliar a consistência dos dados e identifcar possíveis variáveis que impactam sua variável resposta.
#
# Para a realização deste teste você pode utilizar o software de sua preferência (Python ou R).
#
# Sua solução deverá ser entregue no formato Jupyter, por meio de um repositório Git. Inclua também um arquivo README.md no qual você deve cobrir as respostas para os 5 pontos abaixo.
#
# **a. Como foi a definição da sua estratégia de modelagem?**<br>
# **b. Como foi definida a função de custo utilizada?**<br>
# **c. Qual foi o critério utilizado na seleção do modelo final?**<br>
# **d. Qual foi o critério utilizado para validação do modelo? Por que escolheu utilizar esse método?**
# **e. Quais evidências você possui de que seu modelo é suficientemente bom?**
#
#
# ## Sobre o dataset
#
# O dataset utilizado é uma extração de dados do `Airbnb Rio de Janeiro`, conforme
# [fonte](http://insideairbnb.com/get-the-data.html).
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import LabelEncoder
from sklearn.impute import SimpleImputer
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
import xgboost as xgb
from sklearn.impute import KNNImputer
from sklearn.preprocessing import MinMaxScaler
from sklearn.ensemble import RandomForestRegressor
from sklearn.svm import SVR
from lightgbm import LGBMRegressor
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.model_selection import RandomizedSearchCV
from pprint import pprint
pd.set_option('display.max_columns', 100)
# +
# cols = ['id', 'name', 'host_id', 'host_name', 'neighbourhood_group', 'neighbourhood', 'latitude', 'longitude', 'room_type',
# 'price', 'minimum_nights', 'number_of_reviews', 'last_review', 'reviews_per_month', 'calculated_host_listings_count',
# 'availability_365']
# -
# df = pd.read_csv("../dados/listings.csv.gz", compression='gzip', names=cols)
df = pd.read_csv("../dados/listings_new.csv", index_col="id")
# df_neighb = pd.read_csv("../dados/neighbourhoods.csv")
# df_reviews = pd.read_csv("../dados/reviews.csv")
df.head()
df.shape
# ## Machine Learning
# ### Preparação dos dados
# +
seed = 1
df_test = df.sample(frac=0.1, random_state=seed)
df_train_val = df.drop(df_test.index.tolist())
# -
# +
# feature_list = ['host_name', 'neighbourhood', 'room_type', 'minimum_nights', 'number_of_reviews',
# 'reviews_per_month', 'calculated_host_listings_count', 'availability_365']
# feature_list = ['room_type', 'minimum_nights', 'number_of_reviews',
# 'reviews_per_month', 'calculated_host_listings_count', 'availability_365']
feature_list = ['host_id', 'latitude', 'longitude', 'room_type', 'minimum_nights', 'number_of_reviews',
'reviews_per_month', 'calculated_host_listings_count', 'availability_365']
# -
X = df_train_val[feature_list]
y = df_train_val.price
X
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.3, random_state=seed)
print(X_train.shape, X_val.shape, y_train.shape, y_val.shape)
# ### Tranformação dos dados
X_train_enc = X_train.copy()
X_val_enc = X_val.copy()
X_train_enc
# +
# X_train['neighbourhood'] = X_train.neighbourhood.astype('category')
# +
def LabelEncoderFunc(train, val, col):
# train[col] = train[col].astype(str)
# val[col] = val[col].astype(str)
# le = LabelEncoder()
# le.fit(train[col])
# le_dict = dict(zip(le.classes_, le.transform(le.classes_)))
# le_dict.get(-1, '<Unknown>')
# # train[col] = le.transform(train[col])
# # val[col] = le.transform(val[col])
# train[col] = train[col].apply(lambda x: le_dict.get(x, <unknown_value>))
# val[col] = val[col].apply(lambda x: le_dict.get(x, <unknown_value>))
# return train[col], val[col]
le = LabelEncoder()
le.fit(train[col])
train[col] = le.transform(train[col])
val[col] = le.transform(val[col])
return train[col], val[col]
# +
# X_train_enc['host_name'] = X_train_enc['host_name'].str.replace('&','')
# X_train_enc['host_name'] = X_train_enc['host_name'].str.replace('/','')
# X_train_enc['host_name'] = X_train_enc['host_name'].str.replace(' ',' ')
# X_train_enc['host_name'] = X_train_enc['host_name'].str.replace(' ','_')
# -
# X_train_enc['host_name'], X_val_enc['host_name'] = LabelEncoderFunc(X_train_enc, X_val_enc, 'host_name')
# X_train_enc['neighbourhood'], X_val_enc['neighbourhood'] = LabelEncoderFunc(X_train_enc, X_val_enc, 'neighbourhood')
X_train_enc['room_type'], X_val_enc['room_type'] = LabelEncoderFunc(X_train_enc, X_val_enc, 'room_type')
# +
# # X_train.room_type
# le = LabelEncoder()
# le.fit(X_train['room_type'])
# X_train_enc['room_type'] = le.transform(X_train['room_type'])
# X_val_enc['room_type'] = le.transform(X_val['room_type'])
# -
X_train_enc['reviews_per_month'] = X_train_enc['reviews_per_month'].fillna(X_train_enc.reviews_per_month.mean())
X_val_enc['reviews_per_month'] = X_val_enc['reviews_per_month'].fillna(X_train_enc.reviews_per_month.mean())
# +
scaler = MinMaxScaler()
def MinMaxFunc(train, val, col):
scaler = MinMaxScaler()
scaler.fit(train[[col]])
train[col] = scaler.transform(train[[col]])
val[col] = scaler.transform(val[[col]])
return train[col], val[col]
# +
# X_train_enc['minimum_nights', 'number_of_reviews', 'reviews_per_month',
# 'calculated_host_listings_count', 'availability_365']
X_train_enc['minimum_nights'], X_val_enc['minimum_nights'] = MinMaxFunc(X_train_enc, X_val_enc, 'minimum_nights')
X_train_enc['number_of_reviews'], X_val_enc['number_of_reviews'] = MinMaxFunc(X_train_enc, X_val_enc, 'number_of_reviews')
X_train_enc['reviews_per_month'], X_val_enc['reviews_per_month'] = MinMaxFunc(X_train_enc, X_val_enc, 'reviews_per_month')
X_train_enc['calculated_host_listings_count'], X_val_enc['calculated_host_listings_count'] = MinMaxFunc(X_train_enc, X_val_enc, 'calculated_host_listings_count')
X_train_enc['availability_365'], X_val_enc['availability_365'] = MinMaxFunc(X_train_enc, X_val_enc, 'availability_365')
# -
# ### 1º Tunning do modelo selecionado
# +
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 2000, num = 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 4]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the random grid
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
pprint(random_grid)
# +
# Use the random grid to search for best hyperparameters
# First create the base model to tune
rf = RandomForestRegressor()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random = RandomizedSearchCV(estimator = rf, param_distributions = random_grid, n_iter = 100, cv = 3, verbose=2, random_state=seed, n_jobs = -1)
# Fit the random search model
rf_random.fit(X_train_enc, y_train)
# -
rf_random.best_params_
best_random = rf_random.best_estimator_
best_random.fit(X_train_enc, y_train)
# +
y_pred_train = best_random.predict(X_train_enc)
y_pred_val = best_random.predict(X_val_enc)
print(f'Previsão nos dados de treino:')
print('-----------------------------------------------------')
print(f'Mean Squared Error: {mean_squared_error(y_train, y_pred_train)}')
print(f'R2 score: {r2_score(y_train, y_pred_train)}\n')
print(f'Previsão nos dados de Validação:')
print('-----------------------------------------------------')
print(f'Mean Squared Error: {mean_squared_error(y_val, y_pred_val)}')
print(f'R2 score: {r2_score(y_val, y_pred_val)}')
# -
sns.regplot(x=y_val, y=y_pred_val)
# ### 2º Tunning do modelo selecionado
# +
room_type_dummies_train = X_train.room_type.str.get_dummies()
room_type_dummies_valid = X_val.room_type.str.get_dummies()
X_train_enc2 = pd.concat([X_train, room_type_dummies_train], axis=1)
X_val_enc2 = pd.concat([X_val, room_type_dummies_valid], axis=1)
# -
X_train_enc2.drop(['room_type'], axis=1, inplace=True)
X_val_enc2.drop(['room_type'], axis=1, inplace=True)
# +
inputer = KNNImputer(n_neighbors=5)
# inputer.fit_transform(X_train_enc2.reviews_per_month.values.reshape(1,-1))
# X_val_enc2['reviews_per_month'] = inputer.fit_transform(X_val_enc2.reviews_per_month.values.reshape(1,-1))
X_train_enc2 = pd.DataFrame(inputer.fit_transform(X_train_enc2), columns=X_train_enc2.columns)
X_val_enc2 = pd.DataFrame(inputer.fit_transform(X_val_enc2), columns=X_val_enc2.columns)
# -
X_train_enc2
X_train_enc2['minimum_nights'], X_val_enc2['minimum_nights'] = MinMaxFunc(X_train_enc2, X_val_enc2, 'minimum_nights')
X_train_enc2['number_of_reviews'], X_val_enc2['number_of_reviews'] = MinMaxFunc(X_train_enc2, X_val_enc2, 'number_of_reviews')
X_train_enc2['reviews_per_month'], X_val_enc2['reviews_per_month'] = MinMaxFunc(X_train_enc2, X_val_enc2, 'reviews_per_month')
X_train_enc2['calculated_host_listings_count'], X_val_enc2['calculated_host_listings_count'] = MinMaxFunc(X_train_enc2, X_val_enc2, 'calculated_host_listings_count')
X_train_enc2['availability_365'], X_val_enc2['availability_365'] = MinMaxFunc(X_train_enc2, X_val_enc2, 'availability_365')
X_train_enc2
rfr2 = RandomForestRegressor(n_estimators=1000)
rfr2.fit(X_train_enc2, y_train)
# +
y_pred_train2 = rfr2.predict(X_train_enc2)
y_pred_val2 = rfr2.predict(X_val_enc2)
print(f'Previsão nos dados de treino:')
print('-----------------------------------------------------')
print(f'Mean Squared Error: {mean_squared_error(y_train, y_pred_train2)}')
print(f'R2 score: {r2_score(y_train, y_pred_train2)}\n')
print(f'Previsão nos dados de Validação:')
print('-----------------------------------------------------')
print(f'Mean Squared Error: {mean_squared_error(y_val, y_pred_val2)}')
print(f'R2 score: {r2_score(y_val, y_pred_val2)}')
# +
# Use the random grid to search for best hyperparameters
# First create the base model to tune
rf = RandomForestRegressor()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random2 = RandomizedSearchCV(estimator = rf, param_distributions = random_grid, n_iter = 100, cv = 3, verbose=2, random_state=seed, n_jobs = -1)
# Fit the random search model
rf_random2.fit(X_train_enc2, y_train)
# -
rf_random2.best_params_
best_random2 = rf_random2.best_estimator_
best_random2.fit(X_train_enc2, y_train)
# +
y_pred_train2 = best_random2.predict(X_train_enc2)
y_pred_val2 = best_random2.predict(X_val_enc2)
print(f'Previsão nos dados de treino:')
print('-----------------------------------------------------')
print(f'Mean Squared Error: {mean_squared_error(y_train, y_pred_train2)}')
print(f'R2 score: {r2_score(y_train, y_pred_train2)}\n')
print(f'Previsão nos dados de Validação:')
print('-----------------------------------------------------')
print(f'Mean Squared Error: {mean_squared_error(y_val, y_pred_val2)}')
print(f'R2 score: {r2_score(y_val, y_pred_val2)}')
# -
sns.regplot(x=y_val, y=y_pred_val2)
# ### 3º Tunning do modelo selecionado
| Notebook/.ipynb_checkpoints/Machine Learning 1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''SingleC'': conda)'
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import scanpy as sc
sc.settings.verbosity = 3 # verbosity: errors (0), warnings (1), info (2), hints (3)
sc.logging.print_header()
sc.settings.set_figure_params(dpi=80, facecolor='white')
DataPath = "../../../Dataset/GSE94820_cleared/"
# +
genes = pd.read_csv('genes.csv',header=0)
umapValue = pd.read_csv('umap_RISC.csv',header=0)
anno = pd.read_csv('anno_RISC.csv',header=0,index_col=0)
anno
# -
anno.rename(columns={'Set':'batch'},inplace=True)
anno
from anndata import AnnData
adata = AnnData(np.ones((576, 26593)), var=pd.DataFrame(index=genes['Unnamed: 0']),obs=anno)
adata.obsm['X_umap'] = umapValue.values[:,1:3]
# +
def output(Scadata,method):
sc.pl.umap(Scadata, color=['batch','celltype'],s=10,save='UMAP_' + method + '.svg') #如何用矢量格式存?
pd.DataFrame(Scadata.obsm['X_umap'],index = Scadata.obs.index,columns=['umap1','umap2']).to_csv(DataPath + method +'_lisi.csv')
pd.DataFrame(Scadata.obsm['X_umap'].T, index=['umap1','umap2'], columns=Scadata.obs.index).to_csv(DataPath + method+'_kBET.csv')
Scadata.obs.to_csv(DataPath + method+'_anno.csv')
output(adata,'RISC')
| BatchCorrection/Script/GSE94820_cleared/forR/visual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# As you've already seen in the [introduction to variables](variables_intro),
# Python gives names to values using an *assignment* statement. In an
# assignment, a *name* is followed by `=`, which is followed by any
# [expression](Expressions). The value of the expression to the right of `=` is
# *assigned* to the name. Once a name has a value assigned to it, the value will
# be substituted for that name in future expressions.
#
# To repeat, when Python gives a name to a value, that is an *assignment
# statement*.
#
# A *statement* is a piece of code that performs an action.
# Here are two assignment statements, giving values to the names `a` and `b`.
# Then we use these variables in an expression. The Notebook shows the human-readable display of the expression return value.
a = 10
b = 20
a + b
# In the expression `a + b` above, Python evaluates the variable `a` to get
# a computer representation (CR) of the number 10, it evaluates the variable `b`
# to get the CR of 20, and then adds them, to give the final result of the
# expression; a CR of 30.
#
# Finally, the notebook creates a human-readable (HR) version of the result, and
# shows that to us.
#
# A variable can be used in the expression to the right of `=`.
quarter = 1/4
half = 2 * quarter
half
# We can change the value for variables. Here we change the value of variable
# `quarter` from 0.25 to 4.
quarter = 4
# Now we have changed the value of `quarter` What do you think will happen to
# the value of `half` above? Will it change, or will it stay the same?
#
# Try working out the answer before reading further.
#
# Remember that an expression *evaluates* its variables before returning the
# result. In the cell above, the expression `half = 2 * quarter` evaluated
# `quarter`, to get the CR of 0.25, and then evaluated 2 * 0.25, to give the CR
# of 0.5. `half` has this value - the CR of 0.5. When we changed `quarter` in
# the next cell, it did not affect the value that we have already given to
# `half`:
half
# ## Reassigning variables
#
# You can change (reassign) the value of a variable. Let's say we start with:
a = 5
a
# At this stage the variable `a` has the value 5.
#
# If you want, you can then *change* the value of `a`, like this:
a = 7
a
# Now the variable `a` has the value 7. You can call this *reassigning* a variable.
#
# Now consider this piece of code:
a = 10
a = a + 1
# What value will `a` have?
#
# Remember that Python will evaluate the right hand side (RHS) of `a = a + 1`.
# The RHS is `a + 1`. This has two sub-expressions, `a` (which evaluates to the
# CR of 10) and `1` (which evaluates to a CR of 1), so the value returned from `a
# \+ 1` is the CR of 11. After this Python sets the left-hand-side name `a` to
# have this value. So:
a
# See [this chapter in Think Like a Computer
# Scientist](https://runestone.academy/runestone/books/published/thinkcspy/SimplePythonData/Reassignment.html)
# for more explanation of variables and reassignment.
#
# ## Rules for variable names
#
# Variable names must start with a letter, but can contain both letters and
# numbers. A name cannot contain a space; instead, it is common to use an
# underscore character `_` to replace each space. Names are only as useful as
# you make them; it's up to the programmer to choose names that are easy to
# interpret. Typically, more meaningful names can be invented than `a` and `b`.
# For example, let's say you were calculating the 20% Value Added Tax for a
# restaurant bill, as well as 15% tip, on top of that. The following names
# clarify the meaning of the various quantities involved.
meal_price = 25
vat_rate = 0.2
vat = meal_price * vat_rate
meal_price_with_vat = meal_price + vat
meal_price_with_vat
tip_rate = 0.15
tip = meal_price_with_vat * tip_rate
meal_price_total = meal_price_with_vat + tip
meal_price_total
# See [the Names exercises](../../exercises/names) to practice the material in this section.
#
# {% data8page Names %}
| notebooks/02/Names.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment-1
# # Question-1
# Given the following jumbled word, OBANWRI guess the correct English word.
#
# <NAME>
# <NAME>
# <NAME>
# <NAME>
Answer: B,RAINBOW
# # Question-2
print("LETS UPGRADE")
# # Question-3
cp=int(input())
sp=int(input())
if cp<sp:
print("Profit")
elif cp>sp:
print("Loss")
else:
print("Neither")
# # Question-4
Rupees=int(input())
n=80
Euro=Rupees*n
print(Euro)
| Data Science Assignment-1 Day-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Construir o gráfico e encontrar o foco e uma equação da diretriz.</b>
# <b>1)</b> $x^2 = -4y$
# <b> $2p = -4$, logo </b><br><br>
# $P = -2$<br><br>
# <b>Calculando o foco</b><br><br>
# $F = \frac{P}{2}$<br><br>
# $F = -\frac{2}{2}$<br><br>
# $F = 1$, $F(0,-1)$<br><br>
# <b>Calculando a diretriz</b><br><br>
# $d = -\frac{p}{2}$<br><br>
# $d = -\frac{2}{2}$<br><br>
# $d : y = - 1$<br><br>
# $V(0,0)$<br><br>
# $F(0,-1)$
# <b>Gráfico da parábola</b>
from sympy import *
from sympy.plotting import plot_implicit
x = symbols("x")
plot(x**2,(x,-3,3),
title=u'Gráfico da parábola', xlabel='x', ylabel='y');
| Problemas Propostos. Pag. 172 - 175/01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import keras, grid
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.optimizers import SGD
from grid.clients.keras import KerasClient
import grid.lib.coinbase_helper as helper
g = KerasClient()
# -
g.set_coinbase_api("DO NOT COMMIT", "DO NOT COMMIT")
# +
input = np.array([[0,0],[0,1],[1,0],[1,1]])
target = np.array([[0],[1],[1],[0]])
model = Sequential()
model.add(Dense(8, input_dim=2))
model.add(Activation('tanh'))
model.add(Dense(1))
model.add(Activation('sigmoid'))
sgd = SGD(lr=0.1)
model.compile(loss='binary_crossentropy', optimizer=sgd)
model, train_spec = g.fit(model,input,target,epochs=20,log_interval=100,preferred_node='QmSCXtNJGfQD6dkYWinDu6jjAcetqg8Je4uAbtShPHfPQS')
# -
| notebooks/experimental/CoinbasePayment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# THIS CELL SETS STUFF UP FOR DEMO / COLLAB. THIS CELL CAN BE IGNORED.
#-------------------------------------GET RID OF TF DEPRECATION WARNINGS--------------------------------------#
import warnings
warnings.filterwarnings('ignore', category=FutureWarning)
import tensorflow as tf
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR)
#----------------------------------INSTALL PSYCHRNN IF IN A COLAB NOTEBOOK-------------------------------------#
# Installs the correct branch / release version based on the URL. If no branch is provided, loads from master.
# Loads saved weights from correct branch and saves a local copy for later use.
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
import json
import re
import ipykernel
import requests
from requests.compat import urljoin
from io import BytesIO
import numpy as np
import os
from notebook.notebookapp import list_running_servers
kernel_id = re.search('kernel-(.*).json',
ipykernel.connect.get_connection_file()).group(1)
servers = list_running_servers()
for ss in servers:
response = requests.get(urljoin(ss['url'], 'api/sessions'),
params={'token': ss.get('token', '')})
for nn in json.loads(response.text):
if nn['kernel']['id'] == kernel_id:
relative_path = nn['notebook']['path'].split('%2F')
if 'blob' in relative_path:
blob = relative_path[relative_path.index('blob') + 1]
# !pip install git+https://github.com/murraylab/PsychRNN@$blob
file_location = "https://github.com/murraylab/PsychRNN/blob/" + blob + "/docs/notebooks/weights/saved_weights.npz?raw=true"
else:
# !pip install git+https://github.com/murraylab/PsychRNN
file_location = "https://github.com/murraylab/PsychRNN/docs/notebooks/weights/saved_weights.npz?raw=true"
r = requests.get(file_location, stream = True)
data = dict(np.load(BytesIO(r.raw.read()), allow_pickle = True))
if not os.path.exists("./weights"):
os.makedirs("./weights")
np.savez("./weights/saved_weights.npz", **data)
# -
# # Accessing and Modifying Weights
# In [Simple Example](PerceptualDiscrimination.ipynb#Get-&-Save-Model-Weights), we saved weights to ``./weights/saved_weights``. Here we will load those weights, and modify them by silencing a few recurrent units.
import numpy as np
weights = dict(np.load('./weights/saved_weights.npz', allow_pickle = True))
weights['W_rec'][:10, :10] = 0
# Here are all the different weights you have access to for modifying. The ones that don't end in ``Adam`` or ``Adam_1`` will be read in when loading a model from weights.
print(weights.keys())
# Save the modified weights at ``'./weights/modified_saved_weights.npz'``.
np.savez('./weights/modified_saved_weights.npz', **weights)
# # Loading Model with Weights
from psychrnn.backend.models.basic import Basic
network_params = {'N_batch': 50,
'N_in': 2,
'N_out': 2,
'dt': 10,
'tau': 100,
'T': 2000,
'N_steps': 200,
'N_rec': 50
}
# ### Load from File
# Set network parameters.
file_network_params = network_params.copy()
file_network_params['name'] = 'file'
file_network_params['load_weights_path'] = './weights/modified_saved_weights.npz'
# Instantiate model.
fileModel = Basic(file_network_params)
# Verify that the W_rec weights are modified as expected.
print(fileModel.get_weights()['W_rec'][:10,:10])
fileModel.destruct()
# ### Load from Weights Dictionary
# Set network parameters.
dict_network_params = network_params.copy()
dict_network_params['name'] = 'dict'
dict_network_params.update(weights)
type(dict_network_params['dale_ratio']) == np.ndarray and dict_network_params['dale_ratio'].item() is None
# Instantiate model.
dictModel = Basic(dict_network_params)
# Verify that the W_rec weights are modified as expected.
print(dictModel.get_weights()['W_rec'][:10,:10])
dictModel.destruct()
| docs/notebooks/SavingLoadingWeights.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Word Embeddings First Steps: Data Preparation
#
# In this series of ungraded notebooks, you'll try out all the individual techniques that you learned about in the lectures. Practicing on small examples will prepare you for the graded assignment, where you will combine the techniques in more advanced ways to create word embeddings from a real-life corpus.
#
# This notebook focuses on data preparation, which is the first step of any machine learning algorithm. It is a very important step because models are only as good as the data they are trained on and the models used require the data to have a particular structure to process it properly.
#
# To get started, import and initialize all the libraries you will need.
# +
import re
import nltk
nltk.download('punkt')
import emoji
import numpy as np
from nltk.tokenize import word_tokenize
from utils2 import get_dict
# -
# # Data preparation
# In the data preparation phase, starting with a corpus of text, you will:
#
# - Clean and tokenize the corpus.
#
# - Extract the pairs of context words and center word that will make up the training data set for the CBOW model. The context words are the features that will be fed into the model, and the center words are the target values that the model will learn to predict.
#
# - Create simple vector representations of the context words (features) and center words (targets) that can be used by the neural network of the CBOW model.
# ## Cleaning and tokenization
#
# To demonstrate the cleaning and tokenization process, consider a corpus that contains emojis and various punctuation signs.
# Define a corpus
corpus = 'Who ❤️ "word embeddings" in 2020? I do!!!'
# First, replace all interrupting punctuation signs — such as commas and exclamation marks — with periods.
# +
# Print original corpus
print(f'Corpus: {corpus}')
# Do the substitution
data = re.sub(r'[,!?;-]+', '.', corpus)
# Print cleaned corpus
print(f'After cleaning punctuation: {data}')
# -
# Next, use NLTK's tokenization engine to split the corpus into individual tokens.
# +
# Print cleaned corpus
print(f'Initial string: {data}')
# Tokenize the cleaned corpus
data = nltk.word_tokenize(data)
# Print the tokenized version of the corpus
print(f'After tokenization: {data}')
# -
# Finally, as you saw in the lecture, get rid of numbers and punctuation other than periods, and convert all the remaining tokens to lowercase.
# +
# Print the tokenized version of the corpus
print(f'Initial list of tokens: {data}')
# Filter tokenized corpus using list comprehension
data = [ ch.lower() for ch in data
if ch.isalpha()
or ch == '.'
or emoji.get_emoji_regexp().search(ch)
]
# Print the tokenized and filtered version of the corpus
print(f'After cleaning: {data}')
# -
# Note that the heart emoji is considered as a token just like any normal word.
#
# Now let's streamline the cleaning and tokenization process by wrapping the previous steps in a function.
# Define the 'tokenize' function that will include the steps previously seen
def tokenize(corpus):
data = re.sub(r'[,!?;-]+', '.', corpus)
data = nltk.word_tokenize(data) # tokenize string to words
data = [ ch.lower() for ch in data
if ch.isalpha()
or ch == '.'
or emoji.get_emoji_regexp().search(ch)
]
return data
# Apply this function to the corpus that you'll be working on in the rest of this notebook: "I am happy because I am learning"
# +
# Define new corpus
corpus = 'I am happy because I am learning'
# Print new corpus
print(f'Corpus: {corpus}')
# Save tokenized version of corpus into 'words' variable
words = tokenize(corpus)
# Print the tokenized version of the corpus
print(f'Words (tokens): {words}')
# -
# **Now try it out yourself with your own sentence.**
# Run this with any sentence
tokenize("Now it's your turn: try with your own sentence!")
# ## Sliding window of words
# Now that you have transformed the corpus into a list of clean tokens, you can slide a window of words across this list. For each window you can extract a center word and the context words.
#
# The `get_windows` function in the next cell was introduced in the lecture.
# Define the 'get_windows' function
def get_windows(words, C):
i = C
while i < len(words) - C:
center_word = words[i]
context_words = words[(i - C):i] + words[(i+1):(i+C+1)]
yield context_words, center_word
i += 1
# The first argument of this function is a list of words (or tokens). The second argument, `C`, is the context half-size. Recall that for a given center word, the context words are made of `C` words to the left and `C` words to the right of the center word.
#
# Here is how you can use this function to extract context words and center words from a list of tokens. These context and center words will make up the training set that you will use to train the CBOW model.
# Print 'context_words' and 'center_word' for the new corpus with a 'context half-size' of 2
for x, y in get_windows(['i', 'am', 'happy', 'because', 'i', 'am', 'learning'], 2):
print(f'{x}\t{y}')
# The first example of the training set is made of:
#
# - the context words "i", "am", "because", "i",
#
# - and the center word to be predicted: "happy".
#
# **Now try it out yourself. In the next cell, you can change both the sentence and the context half-size.**
# Print 'context_words' and 'center_word' for any sentence with a 'context half-size' of 1
for x, y in get_windows(tokenize("Now it's your turn: try with your own sentence!"), 1):
print(f'{x}\t{y}')
# ## Transforming words into vectors for the training set
# To finish preparing the training set, you need to transform the context words and center words into vectors.
#
# ### Mapping words to indices and indices to words
#
# The center words will be represented as one-hot vectors, and the vectors that represent context words are also based on one-hot vectors.
#
# To create one-hot word vectors, you can start by mapping each unique word to a unique integer (or index). We have provided a helper function, `get_dict`, that creates a Python dictionary that maps words to integers and back.
# Get 'word2Ind' and 'Ind2word' dictionaries for the tokenized corpus
word2Ind, Ind2word = get_dict(words)
# Here's the dictionary that maps words to numeric indices.
# Print 'word2Ind' dictionary
word2Ind
# You can use this dictionary to get the index of a word.
# Print value for the key 'i' within word2Ind dictionary
print("Index of the word 'i': ",word2Ind['i'])
# And conversely, here's the dictionary that maps indices to words.
# Print 'Ind2word' dictionary
Ind2word
# Print value for the key '2' within Ind2word dictionary
print("Word which has index 2: ",Ind2word[2] )
# Finally, get the length of either of these dictionaries to get the size of the vocabulary of your corpus, in other words the number of different words making up the corpus.
# +
# Save length of word2Ind dictionary into the 'V' variable
V = len(word2Ind)
# Print length of word2Ind dictionary
print("Size of vocabulary: ", V)
# -
# ### Getting one-hot word vectors
#
# Recall from the lecture that you can easily convert an integer, $n$, into a one-hot vector.
#
# Consider the word "happy". First, retrieve its numeric index.
# +
# Save index of word 'happy' into the 'n' variable
n = word2Ind['happy']
# Print index of word 'happy'
n
# -
# Now create a vector with the size of the vocabulary, and fill it with zeros.
# +
# Create vector with the same length as the vocabulary, filled with zeros
center_word_vector = np.zeros(V)
# Print vector
center_word_vector
# -
# You can confirm that the vector has the right size.
# Assert that the length of the vector is the same as the size of the vocabulary
len(center_word_vector) == V
# Next, replace the 0 of the $n$-th element with a 1.
# Replace element number 'n' with a 1
center_word_vector[n] = 1
# And you have your one-hot word vector.
# Print vector
center_word_vector
# **You can now group all of these steps in a convenient function, which takes as parameters: a word to be encoded, a dictionary that maps words to indices, and the size of the vocabulary.**
# Define the 'word_to_one_hot_vector' function that will include the steps previously seen
def word_to_one_hot_vector(word, word2Ind, V):
one_hot_vector = np.zeros(V)
one_hot_vector[word2Ind[word]] = 1
return one_hot_vector
# Check that it works as intended.
# Print output of 'word_to_one_hot_vector' function for word 'happy'
word_to_one_hot_vector('happy', word2Ind, V)
# **What is the word vector for "learning"?**
# Print output of 'word_to_one_hot_vector' function for word 'learning'
word_to_one_hot_vector('learning', word2Ind, V)
# Expected output:
#
# array([0., 0., 0., 0., 1.])
# ### Getting context word vectors
# To create the vectors that represent context words, you will calculate the average of the one-hot vectors representing the individual words.
#
# Let's start with a list of context words.
# Define list containing context words
context_words = ['i', 'am', 'because', 'i']
# Using Python's list comprehension construct and the `word_to_one_hot_vector` function that you created in the previous section, you can create a list of one-hot vectors representing each of the context words.
# +
# Create one-hot vectors for each context word using list comprehension
context_words_vectors = [word_to_one_hot_vector(w, word2Ind, V) for w in context_words]
# Print one-hot vectors for each context word
context_words_vectors
# -
# And you can now simply get the average of these vectors using numpy's `mean` function, to get the vector representation of the context words.
# Compute mean of the vectors using numpy
np.mean(context_words_vectors, axis=0)
# Note the `axis=0` parameter that tells `mean` to calculate the average of the rows (if you had wanted the average of the columns, you would have used `axis=1`).
#
# **Now create the `context_words_to_vector` function that takes in a list of context words, a word-to-index dictionary, and a vocabulary size, and outputs the vector representation of the context words.**
# Define the 'context_words_to_vector' function that will include the steps previously seen
def context_words_to_vector(context_words, word2Ind, V):
context_words_vectors = [word_to_one_hot_vector(w, word2Ind, V) for w in context_words]
context_words_vectors = np.mean(context_words_vectors, axis=0)
return context_words_vectors
# And check that you obtain the same output as the manual approach above.
# Print output of 'context_words_to_vector' function for context words: 'i', 'am', 'because', 'i'
context_words_to_vector(['i', 'am', 'because', 'i'], word2Ind, V)
# **What is the vector representation of the context words "am happy i am"?**
# Print output of 'context_words_to_vector' function for context words: 'am', 'happy', 'i', 'am'
context_words_to_vector(['am', 'happy', 'i', 'am'], word2Ind, V)
# Expected output:
#
# array([0.5 , 0. , 0.25, 0.25, 0. ])
#
# ## Building the training set
# You can now combine the functions that you created in the previous sections, to build a training set for the CBOW model, starting from the following tokenized corpus.
# Print corpus
words
# To do this you need to use the sliding window function (`get_windows`) to extract the context words and center words, and you then convert these sets of words into a basic vector representation using `word_to_one_hot_vector` and `context_words_to_vector`.
# Print vectors associated to center and context words for corpus
for context_words, center_word in get_windows(words, 2): # reminder: 2 is the context half-size
print(f'Context words: {context_words} -> {context_words_to_vector(context_words, word2Ind, V)}')
print(f'Center word: {center_word} -> {word_to_one_hot_vector(center_word, word2Ind, V)}')
print()
# In this practice notebook you'll be performing a single iteration of training using a single example, but in this week's assignment you'll train the CBOW model using several iterations and batches of example.
# Here is how you would use a Python generator function (remember the `yield` keyword from the lecture?) to make it easier to iterate over a set of examples.
# Define the generator function 'get_training_example'
def get_training_example(words, C, word2Ind, V):
for context_words, center_word in get_windows(words, C):
yield context_words_to_vector(context_words, word2Ind, V), word_to_one_hot_vector(center_word, word2Ind, V)
# The output of this function can be iterated on to get successive context word vectors and center word vectors, as demonstrated in the next cell.
# Print vectors associated to center and context words for corpus using the generator function
for context_words_vector, center_word_vector in get_training_example(words, 2, word2Ind, V):
print(f'Context words vector: {context_words_vector}')
print(f'Center word vector: {center_word_vector}')
print()
# Your training set is ready, you can now move on to the CBOW model itself which will be covered in the next lecture notebook.
#
# **Congratulations on finishing this lecture notebook!** Hopefully you now have a better understanding of how to prepare your data before feeding it to a continuous bag-of-words model.
#
# **Keep it up!**
| Part2_Probabilistic_Models/C2_W4_lecture_nb_1_data_prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# PREDICTING ANALYTE(S) FROM ANALYTE(S)
#
# ---
# ### 1. Google Colab runtime setup [Optional]
# Clone and install spectrai package
# !git clone https://github.com/franckalbinet/spectrai.git
# !pip install /content/spectrai
from google.colab import drive
drive.mount('/content/drive')
# Prepare /root folder content
# !cp -r /content/drive/My\ Drive/Colab\ Notebooks/data/data_spectrai /root
# Create configuration file
# !mkdir /root/.spectrai_config & cp /content/spectrai/config.toml /root/.spectrai_config
# ### 2. Import packages
# +
from spectrai.datasets.kssl import (get_tax_orders_lookup_tbl, load_data,
load_analytes, load_data_analytes, load_fact_tbl, load_spectra)
from spectrai.vis.spectra import (plot_spectra)
import pandas as pd
import numpy as np
from tqdm import tqdm
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LinearRegression
import xgboost as xgb
# %load_ext autoreload
# %autoreload 2
# -
# ### 3. Load KSSL dataset for analyte(s) -> analyte(s) modeling
# +
# Choosing analytes to be used as features and target(s)
targets = [725]
# Selected analytes where nb_samples > 30000 (i.e most frequently measured analytes)
features = [622, 623, 624, 420, 723, 724, 722,726, 750, 481, 268, 339, 334, 342, 338, 337, 343, 340, 341, 417, 65, 66, 67]
# Tetsuya features
tetsuya_features = [383, 368]
# Selected features
features_selected = [383, 723, 481, 726, 268, 417]
# -
X, X_names, y, y_names, instances_id = load_data_analytes(features=features, targets=targets)
print('X shape: ', X.shape)
print('X approx. memory size: {:.2f} MB'.format(X.nbytes / 10**6))
print('y approx. memory size: {:.2f} MB'.format(y.nbytes / 10**6))
print('Features: ', X_names)
print('Target variable: ', y_names)
# ### 4. Modeling
# * **Utilities functions**
def prettify_features_importance(features_importance, ascending_sort=False, label='feature_importance'):
df_analytes = load_analytes()
df = df_analytes.set_index('analyte_id').loc[X_names,:]
df[label] = features_importance
return df \
.sort_values(by=[label], ascending=ascending_sort) \
.reset_index()[['analyte_id', 'analyte_name', label]]
# * **Assess Correlation between features and target**
prettify_features_importance(np.corrcoef(np.c_[X, y].T)[:, -1][:-1], label='corr. with potassium (725)')
# * **Data preparation**
X_subset = X
y_subset = y
X_subset = StandardScaler().fit_transform(X_subset)
# +
# Classical train, test split
X_train, X_test, y_train, y_test = train_test_split(X_subset, y_subset, test_size=0.20, random_state=42)
print('X train shape: ', X_train.shape)
print('X test shape: ', X_test.shape)
print('y train shape: ', y_train.shape)
# -
# * **Linear Regression**
lr = LinearRegression()
lr.fit(X_train, y_train)
print('# of training sample: {}'.format(X_train.shape[0]))
print('Train R2 score: {:.3f}'.format(lr.score(X_train, y_train)))
print('Test R2 score: {:.3f}'.format(lr.score(X_test, y_test)))
prettify_features_importance(lr.coef_.tolist()[0])
# * **XGBoost**
xgb_reg = xgb.XGBRegressor(n_estimators=500)
xgb_reg.fit(X_train, y_train)
print('# of training sample: {}'.format(X_train.shape[0]))
print('Train R2 score: {:.3f}'.format(xgb_reg.score(X_train, y_train)))
print('Test R2 score: {:.3f}'.format(xgb_reg.score(X_test, y_test)))
prettify_features_importance(xgb_reg.feature_importances_)
# ### 5. In-depth analysis of analytes correlation
corrs = []
for analyte in tqdm(df_fact['analyte_id'].unique().tolist()):
X, _, y, _, _ = load_data_analytes(features=[analyte], targets=[725])
corrs.append({
'analyte_id': analyte,
'correlation': np.corrcoef(np.c_[X, y].T)[:, -1][:-1].item(),
'nb_samples': X.shape[0]
})
# +
df_corrs = pd.DataFrame(corrs).dropna()
mask_not_target = df_corrs['analyte_id'] != 725
mask_min_nb = df_corrs['nb_samples'] > 30
df_corrs = df_corrs.loc[mask_not_target & mask_min_nb, :]
df_corrs['abs_correlation'] = np.abs(df_corrs['correlation'])
df_corrs = df_corrs.sort_values(by=['abs_correlation', 'nb_samples'], ascending=False)
# -
df_corrs.head()
df_corrs = df_corrs.merge(df_analytes[['analyte_id', 'analyte_name', 'uom_abbrev']],
on='analyte_id')[['analyte_id', 'analyte_name', 'uom_abbrev', 'correlation', 'nb_samples', 'abs_correlation']]
df_corrs.head(25)
df_corrs.to_excel('data/correlations_with_potassium.xlsx', float_format="%.2f", index=False)
| examples/predicting-analytes-from-analytes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:genpen]
# language: python
# name: conda-env-genpen-py
# ---
# + heading_collapsed="false" tags=[]
import itertools
import numpy as np
import os
import seaborn as sns
from tqdm import tqdm
from dataclasses import asdict, dataclass, field
import vsketch
import shapely.geometry as sg
from shapely.geometry import box, MultiLineString, Point, MultiPoint, Polygon, MultiPolygon, LineString
import shapely.affinity as sa
import shapely.ops as so
import matplotlib.pyplot as plt
import pandas as pd
import vpype_cli
from typing import List, Generic
from genpen import genpen as gp, utils as utils
from scipy import stats as ss
import geopandas
from shapely.errors import TopologicalError
import functools
# %load_ext autoreload
# %autoreload 2
import vpype
from skimage import io
from pathlib import Path
import fn
import bezier
from sklearn.preprocessing import minmax_scale
from skimage import feature
from genpen.utils import Paper
from genpen.genpen import *
from genpen import subdivide as sd
from functools import partial
from genpen.grower import Grower, GrowerParams
# + heading_collapsed="false"
# make page
paper_size = 'A2'
border:float=45
paper = Paper(paper_size)
drawbox = paper.get_drawbox(border)
split_func = functools.partial(sd.split_random_bezier, x0=0.2, x1=0.75, n_eval_points=50)
xgen = ss.uniform(loc=0.4, scale=0.01).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
# x0gen = ss.uniform(loc=0.15, scale=0.01).rvs
# x1gen = ss.uniform(loc=0.65, scale=0.01).rvs
# split_func = functools.partial(sd.split_random_line_gen, x0gen=x0gen, x1gen=x1gen)
target = Point(140, 325)
target = drawbox.centroid
dist_from_center = partial(sd.distance_from_pt, target=target, p_range=(0.99, 0.3,), d_range=(0, 200))
cp = sd.ContinuePolicy(dist_from_center)
polys = sd.very_flex_rule_recursive_split(poly=drawbox, split_func=split_func, continue_func=cp, depth_limit=14, buffer_kwargs={'distance':1e-6})
bps = gp.merge_Polygons(polys)
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.5mm')
sk.geometry(bps.boundary)
# tolerance=0.5
sk.display()
# + heading_collapsed="false"
n_layers = 1
layers = []
for ii in range(n_layers):
fills = []
for p in bps:
xjitter_func = 0
yjitter_func = ss.norm(loc=0, scale=np.random.uniform(0.005, 0.01)).rvs
bhf = BezierHatchFill(
spacing=np.random.uniform(0.2, 0.22),
degrees=np.random.uniform(40,60),
poly_to_fill=p,
xjitter_func=xjitter_func,
yjitter_func=yjitter_func,
fill_inscribe_buffer=1.4,
n_nodes_per_line=5,
n_eval_points=6,
)
fills.append(bhf.p)
fills = [f for f in fills if f.length > 0]
layer = gp.merge_LineStrings(fills)
layers.append(layer)
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
for i, layer in enumerate(layers):
sk.stroke(i+1)
sk.geometry(layer)
for tolerance in [0.1, 0.3, 0.5, 0.7]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='layer')
# -
plot_id = fn.new_plot_id()
savedir='/home/naka/art/plotter_svgs'
savepath = Path(savedir).joinpath(f'{plot_id}.svg').as_posix()
sk.save(savepath)
# # more interesting initialization
# + heading_collapsed="false"
# make page
paper_size = 'A2'
border:float=45
paper = Paper(paper_size)
drawbox = paper.get_drawbox(border)
# -
params = GrowerParams(
rad_func='loss_scaled_rad',
rad_range=(70, 60),
loss_range=(40, 100),
n_pts_eval_per_iter=55,
n_pts_add_per_iter=1,
pt_to_poly_func='buffer_pt'
)
g = Grower(poly=drawbox.centroid.buffer(0.1), params=params)
# + tags=[]
g.grow(6)
# +
# polys = g.mpoly.buffer(-5)
# polys = g.mpoly
polys = gp.make_like(g.mpoly, drawbox)
# -
bg = drawbox.difference(polys)
polys = list(polys)
# polys.append(bg)
# +
# split_func = functools.partial(sd.split_random_bezier, x0=0.2, x1=0.75, n_eval_points=50)
xgen = ss.uniform(loc=0.5, scale=0.01).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
# x0gen = ss.uniform(loc=0.15, scale=0.01).rvs
# x1gen = ss.uniform(loc=0.65, scale=0.01).rvs
# split_func = functools.partial(sd.split_random_line_gen, x0gen=x0gen, x1gen=x1gen)
# -
target = Point(140, 325)
# target = drawbox.centroid
# +
xgen = ss.uniform(loc=0.4, scale=0.01).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
split_polys = []
for poly in polys:
dist_from_center = partial(sd.distance_from_pt, target=target, p_range=(0.99, 0.3,), d_range=(0, 200))
cp = sd.ContinuePolicy(dist_from_center)
sps = sd.very_flex_rule_recursive_split(poly=poly, split_func=split_func, continue_func=cp, depth_limit=2, buffer_kwargs={'distance':1e-6})
split_polys.append(sps)
xgen = ss.uniform(loc=0.5, scale=0.01).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
dist_from_center = partial(sd.distance_from_pt, target=target, p_range=(0.99, 0.3,), d_range=(0, 200))
cp = sd.ContinuePolicy(dist_from_center)
sps = sd.very_flex_rule_recursive_split(poly=bg, split_func=split_func, continue_func=cp, depth_limit=5, buffer_kwargs={'distance':1e-6})
split_polys.append(sps)
bps = gp.merge_Polygons(split_polys)
# + heading_collapsed="false"
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.5mm')
sk.geometry(bps.boundary)
# tolerance=0.5
sk.display()
# + heading_collapsed="false"
n_layers = 1
# -
layers = []
# + heading_collapsed="false"
for ii in range(n_layers):
fills = []
for p in bps:
xjitter_func = 0
yjitter_func = ss.norm(loc=0, scale=np.random.uniform(0.001, 0.005)).rvs
bhf = BezierHatchFill(
spacing=np.random.uniform(0.2, 0.21),
degrees=np.random.uniform(40,60),
poly_to_fill=p,
xjitter_func=xjitter_func,
yjitter_func=yjitter_func,
fill_inscribe_buffer=1.4,
n_nodes_per_line=5,
n_eval_points=6,
)
fills.append(bhf.p)
fills = [f for f in fills if f.length > 0]
layer = gp.merge_LineStrings(fills)
layers.append(layer)
# + heading_collapsed="false"
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
for i, layer in enumerate(layers):
sk.stroke(i+1)
sk.geometry(layer)
for tolerance in [0.1, 0.3, 0.5, 0.7]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='layer')
# -
plot_id = fn.new_plot_id()
savedir='/home/naka/art/plotter_svgs'
savepath = Path(savedir).joinpath(f'{plot_id}.svg').as_posix()
sk.save(savepath)
# # more interesting initialization
# + heading_collapsed="false"
# make page
paper_size = '6x6 inches'
border:float=15
paper = Paper(paper_size)
drawbox = paper.get_drawbox(border)
# -
params = GrowerParams(
rad_func='loss_scaled_rad',
rad_range=(20, 10),
loss_range=(20, 100),
n_pts_eval_per_iter=55,
n_pts_add_per_iter=1,
pt_to_poly_func='reg_poly'
)
g = Grower(poly=drawbox.centroid.buffer(40), params=params)
# + tags=[]
g.grow(2)
# +
# polys = g.mpoly.buffer(-1)
polys = g.mpoly
# polys = gp.make_like(g.mpoly, drawbox)
# -
bg = drawbox.difference(g.agg_poly)
polys = gp.merge_Polygons(polys)
# polys.append(bg)
# +
# split_func = functools.partial(sd.split_random_bezier, x0=0.2, x1=0.75, n_eval_points=50)
xgen = ss.uniform(loc=0.5, scale=0.01).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
# x0gen = ss.uniform(loc=0.15, scale=0.01).rvs
# x1gen = ss.uniform(loc=0.65, scale=0.01).rvs
# split_func = functools.partial(sd.split_random_line_gen, x0gen=x0gen, x1gen=x1gen)
# -
# target = Point(140, 325)
target = drawbox.centroid
# +
split_polys =[]
x0gen = ss.uniform(loc=0.15, scale=0.01).rvs
x1gen = ss.uniform(loc=0.65, scale=0.1).rvs
split_func = functools.partial(sd.split_random_line_gen, x0gen=x0gen, x1gen=x1gen)
dist_from_center = partial(sd.distance_from_pt, target=target, p_range=(1, 0.3,), d_range=(0, 20))
cp = sd.ContinuePolicy(dist_from_center)
sps = sd.very_flex_rule_recursive_split(poly=drawbox, split_func=split_func, continue_func=cp, depth_limit=2, buffer_kwargs={'distance':1e-6})
sps = gp.merge_Polygons(sps)
split_polys.append(sps)
bps = gp.merge_Polygons(split_polys)
# + heading_collapsed="false"
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.5mm')
sk.geometry(bps.boundary)
# tolerance=0.5
sk.display()
# + heading_collapsed="false"
n_layers = 1
# -
layers = []
# + heading_collapsed="false"
for ii in range(n_layers):
fills = []
for p in bps:
xjitter_func = 0
yjitter_func = ss.norm(loc=0, scale=np.random.uniform(0.001, 0.005)).rvs
bhf = BezierHatchFill(
spacing=np.random.uniform(0.2, 0.21),
degrees=np.random.uniform(40,60),
poly_to_fill=p,
xjitter_func=xjitter_func,
yjitter_func=yjitter_func,
fill_inscribe_buffer=1.4,
n_nodes_per_line=5,
n_eval_points=6,
)
fills.append(bhf.p)
fills = [f for f in fills if f.length > 0]
layer = gp.merge_LineStrings(fills)
layers.append(layer)
# + heading_collapsed="false"
for ii in range(n_layers):
fills = []
for p in bps:
xjitter_func = 0
yjitter_func = ss.norm(loc=0, scale=np.random.uniform(0.001, 0.005)).rvs
bhf = BezierHatchFill(
spacing=np.random.uniform(0.2, 0.21),
degrees=np.random.uniform(20,40),
poly_to_fill=p,
xjitter_func=xjitter_func,
yjitter_func=yjitter_func,
fill_inscribe_buffer=1.4,
n_nodes_per_line=5,
n_eval_points=6,
)
fills.append(bhf.p)
fills = [f for f in fills if f.length > 0]
layer = gp.merge_LineStrings(fills)
layers.append(layer)
# + heading_collapsed="false"
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
for i, layer in enumerate(layers):
sk.stroke(i+1)
sk.geometry(layer)
for tolerance in [0.1, 0.3, 0.5, 0.7]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='layer')
# -
plot_id = fn.new_plot_id()
savedir='/home/naka/art/plotter_svgs'
savepath = Path(savedir).joinpath(f'{plot_id}.svg').as_posix()
sk.save(savepath)
# # remove
# + heading_collapsed="false"
# make page
paper_size = '5x7 inches'
border:float=10
paper = Paper(paper_size)
drawbox = paper.get_drawbox(border)
# -
poly = drawbox
# +
# split_func = functools.partial(sd.split_random_bezier, x0=0.2, x1=0.75, n_eval_points=50)
xgen = ss.uniform(loc=0.5, scale=0.01).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
# x0gen = ss.uniform(loc=0.15, scale=0.01).rvs
# x1gen = ss.uniform(loc=0.65, scale=0.01).rvs
# split_func = functools.partial(sd.split_random_line_gen, x0gen=x0gen, x1gen=x1gen)
# -
# target = Point(140, 325)
target = drawbox.centroid
drawbox.bounds
# +
split_polys =[]
x0gen = ss.uniform(loc=0.25, scale=0.01).rvs
x1gen = ss.uniform(loc=0.75, scale=0.1).rvs
# split_func = functools.partial(sd.split_random_line_gen, x0gen=x0gen, x1gen=x1gen)
xgen = ss.uniform(loc=0.6, scale=0.001).rvs
split_func = functools.partial(sd.split_along_longest_side_of_min_rectangle, xgen=xgen)
dist_from_center = partial(sd.distance_from_pt, target=target, p_range=(1, 0.1,), d_range=(0, 60))
cp = sd.ContinuePolicy(dist_from_center)
sps = sd.very_flex_rule_recursive_split(poly=poly, split_func=split_func, continue_func=cp, depth_limit=5, buffer_kwargs={'distance':1e-6})
sps = gp.merge_Polygons(sps)
split_polys.append(sps)
bps = gp.merge_Polygons(split_polys)
# -
bps = gp.make_like(bps, drawbox)
# + heading_collapsed="false"
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.5mm')
sk.geometry(bps.boundary)
# tolerance=0.5
sk.display()
# + heading_collapsed="false"
n_layers = 1
# -
layers = []
# + heading_collapsed="false"
for ii in range(n_layers):
fills = []
for p in bps:
xjitter_func = 0
yjitter_func = ss.norm(loc=0, scale=np.random.uniform(0.001, 0.005)).rvs
d = p.distance(drawbox.centroid)
angle= (np.interp(d, (0, 160), (0, 360)) // 30) * 30
bhf = BezierHatchFill(
spacing=np.random.uniform(0.2, 0.21),
degrees=angle,
poly_to_fill=p,
xjitter_func=xjitter_func,
yjitter_func=yjitter_func,
fill_inscribe_buffer=1.4,
n_nodes_per_line=5,
n_eval_points=6,
)
fills.append(bhf.p)
fills = [f for f in fills if f.length > 0]
layer = gp.merge_LineStrings(fills)
layers.append(layer)
# + heading_collapsed="false"
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.08mm')
for i, layer in enumerate(layers):
sk.stroke(i+1)
sk.geometry(layer)
for tolerance in [0.05, 0.1, 0.2, 0.3, 0.5, 0.7]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
sk.vpype('linesimplify --tolerance 0.1 linesort')
# sk.display(color_mode='layer')
# -
def vsketch_to_shapely(sketch):
return [[LineString([Point(pt.real, pt.imag) for pt in lc]) for lc in layer] for layer in sketch.document.layers.values()]
layer = sk.document.layers[1]
mls = gp.make_like(MultiLineString([LineString([Point(pt.real, pt.imag) for pt in lc]) for lc in layer]), drawbox)
# + tags=[]
ds = [ls.distance(drawbox.centroid) for ls in mls]
# -
ds = np.array(ds) ** 0.5
ds = ds/ ds.sum()
frac_keep = 0.85
n_keep = int(frac_keep * len(mls))
pmls = MultiLineString(list(np.random.choice(mls, size=n_keep, replace=False, p=ds)))
rlayers = [pmls]
# +
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.08mm')
for i, layer in enumerate(rlayers):
sk.stroke(i+1)
sk.geometry(layer)
for tolerance in [0.05, 0.1, 0.2, 0.3, 0.5, 0.7]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='layer')
# -
plot_id = fn.new_plot_id()
savedir='/home/naka/art/plotter_svgs'
savepath = Path(savedir).joinpath(f'{plot_id}.svg').as_posix()
sk.save(savepath)
sk.vpype('stat')
pmls.length
| scratch/053_bez_shading_again_again.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''venv'': venv)'
# name: python37364bitvenvvenve6e237c6e45e46d28de089fb5b2763f0
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from IPython.display import display
from sklearn.metrics import plot_confusion_matrix
names = [
"Konst, kultur och nöje",
"Brott, lag och rätt",
"Katastrofer och olyckor",
"Ekonomi, affärer och finans",
"Utbildning",
"Miljö och natur",
"Medicin och hälsa",
"Mänskligt",
"Arbete",
"Fritid och livsstil",
"Politik",
"Etik och religion",
"Teknik och vetenskap",
"Samhälle",
"Sport",
"Krig, konflikter och oroligheter",
"Väder",
]
# -
# <h1>Evaluation Results 2020-07-30</h1>
# Training on the 17 IPTC top categories, using 8804 articles from TT and MittMedia split 85/15 into train/test.
#
# * Evaluation accuracy = 0.9162381089376118
# * Evaluation loss = 0.22331761486000484
# + tags=[]
epoch = [1,2,3,4]
loss = [0.3895855526129405, 0.2734635619890122, 0.2296229791073572, 0.2083784256662641]
plt.title("Epoch / Training Loss")
plt.plot(epoch, loss)
plt.show()
# -
#
#
# + tags=[]
# Individual ROC-AUC's
auc = [0.9523777748120469, 0.9421631576327709,0.946391594447427, 0.8660729383912971,0.9639164598842018, 0.9275587197273946, 0.8838577132726942, 0.8411159633799854, 0.9482721880882197, 0.8245653032997431, 0.8829590315658428, 0.9764546566486109, 0.7954117122982559, 0.748578789362744, 0.9579801324503311, 0.8984403805029127, 0.9725423246550007]
data ={"Category":names, "AUC":auc}
df = pd.DataFrame(data)
display(df)
## Number of articles where the category is used
num_articles = [681, 576, 415, 716, 383, 429, 827, 500, 364, 524, 913, 282, 407, 708, 442, 326, 311]
plt.title("AUC / Number of Articles")
plt.scatter(num_articles, auc)
plt.show()
# Number of articles where the category is the only one
num_articles_single = [85, 131, 154, 107, 70, 125, 68, 61, 53, 135, 41, 57, 88, 44, 206, 34, 129]
plt.title("AUC / Number of Articles with Single Category")
plt.scatter(num_articles_single, auc)
plt.show()
# The average number of categories appearing in conjuntion with each category
num_other_cats = [2.3098384728340675, 1.9114583333333333, 1.4409638554216868, 1.899441340782123, 1.6005221932114881, 1.3356643356643356, 1.841596130592503, 2.222, 1.6043956043956045, 1.383587786259542, 2.2694414019715223, 1.1702127659574468, 1.7616707616707616, 2.281073446327684, 1.6515837104072397, 2.144171779141104, 1.234726688102894]
plt.title("AUC / Average Number of Other Categories")
plt.scatter(num_other_cats, auc)
plt.show()
# -
# <h1>Evaluation Results 2020-08-03</h1>
# Training on the 17 IPTC top categories, using 14664 articles from TT and MittMedia, split 85/15 into train/test.
#
# * Evaluation accuracy = 0.9492188182821706
# * Evaluation loss = 0.13287404469092903
# +
import matplotlib.pyplot as plt
epoch = [1,2,3,4]
loss = [0.23655671807011078, 0.13767188245415363, 0.11091446202522402, 0.09424879136990137]
plt.title("Epoch / Training Loss")
plt.plot(epoch, loss)
plt.show()
# + tags=[]
auc = [0.9744482095429985, 0.9746570643990476, 0.9746545457194263, 0.9450286922903556, 0.9777901355518438, 0.9542101673574314, 0.9546278524205588, 0.9348027861984398, 0.9680333186740457, 0.9578485506760959, 0.9303148409918123, 0.9925167815882003, 0.9145604576779767, 0.872212791156731, 0.9903408381302724, 0.9498844992542471, 0.9808168316831682]
data ={"Category":names, "AUC":auc}
df = pd.DataFrame(data)
display(df)
num_articles = [2221, 1825, 1093, 2372, 869, 1057, 2522, 1388, 823, 2070, 2291, 348, 908, 1885, 1381, 456, 509]
plt.title("AUC / Number of Articles")
plt.scatter(num_articles, auc)
plt.show()
num_articles_single = [952, 933, 632, 854, 235, 376, 582, 395, 200, 1000, 355, 77, 326, 383, 921, 71, 256]
plt.title("AUC / Number of Articles with Single Category")
plt.scatter(num_articles_single, auc)
plt.show()
num_other_cats = [1.1656911301215669, 0.9895890410958904, 0.8225068618481244, 1.1538785834738616, 1.2508630609896432, 1.0586565752128667, 1.3057097541633624, 1.3638328530259367, 1.3183475091130012, 0.8309178743961353, 1.6154517677869926, 1.132183908045977, 1.2147577092511013, 1.543236074270557, 0.8117306299782766, 1.888157894736842, 0.9607072691552063]
plt.title("AUC / Average Number of Other Categories")
plt.scatter(num_other_cats, auc)
plt.show()
# -
# <h1>Evaluation Results 2020-08-10</h1>
# Training on the sub-categories to "Kultur och Nöje", using 9627 articles from TT and MittMedia, split 85/15 into train/test. Two model variants were evaluated:
#
# * One where KB-BERT was trained as the top-level classfier, finetuning all model parameters but using the sub-categories instead.
# * Evaluation accuracy = 0.9611858489440749
# * Evalaution loss = 0.10951708459659763
# * Training time = 39:59:18
# * Precision = 0.7685688436031342
# * Recall = 0.699886789140494
# * F1 score = 0.7094580330278563
# * One where the model from 2020-08-03 was loaded, freezing the parameters of the embedding layers and transformer blocks. Then, an additional transformer block was injected and trained together with the classification head and the freezed "base" model.
# * Evaluation accuracy = 0.9297256357451885
# * Evalaution loss = 0.1965097065853036
# * Training time = 10:31:09
#
# A third model variant has been designed but is not yet trained/evaluated. There, KB-BERT would be trained on all sub-categories in a similar fashion as the top-level classifier but with the results from the top-level classifier concatenated with the intermediate output from the last transformer block, right before the classification layer.
# +
mm_codes = [
"RYF-XKI-JIJ",
"RYF-XKI-YFJ",
"RYF-XKI-FEY",
"RYF-XKI-GJH",
"RYF-XKI-BUS",
"RYF-XKI-DEG",
"RYF-XKI-YBJ",
"RYF-XKI-LNE",
"RYF-XKI-HLO",
"RYF-XKI-FXL",
"RYF-XKI-WEG",
"RYF-XKI-ISL",
"RYF-XKI-KKX",
"RYF-XKI-IUV",
"RYF-XKI-SFU",
"RYF-XKI-IHA",
"RYF-XKI-TMS",
"RYF-XKI-JUJ",
"RYF-XKI-PGP",
"RYF-XKI-VME",
"RYF-XKI-CDA",
]
num_articles = [1297, 1351, 2307, 1198, 1200, 595, 534, 1030, 520, 768, 694, 660, 272, 271, 779, 580, 60, 149, 263, 48, 10]
auc_all_params = [0.9605303653112419, 0.9717462566491757, 0.9396335458456435, 0.9425151597429633, 0.9265674920127795,0.9514572197729313, 0.9422706582633054, 0.9690571870170015, 0.9512155730151342, 0.91587584528761, 0.9656285878300802, 0.9397872340425532, 0.8916607270135424, 0.9874176252842655, 0.9516981400476546, 0.9657394582948599, 0.8414641882890314, 0.9978498091705639, 0.976411379396454, 0.7219512195121952, 0.3224687933425797]
auc_one_transformer = [0.8007082779991147, 0.8146876238277639, 0.8100036282589541, 0.7736944519866051, 0.7637571552183174, 0.8475986110569234, 0.8608368347338936, 0.7781504379185986, 0.8752125708965504, 0.7734035028152676, 0.8457161308840413, 0.8126319936958233, 0.7354101837525859, 0.8959879156084525, 0.8092321917564637, 0.8242536164973837, 0.4047452896022331, 0.5901736279094769, 0.8328050865364298, 0.3713511420828494, 0.3467406380027739]
plt.figure(figsize=(10,7))
plt.scatter(num_articles, auc_all_params, label="All Parameters")
plt.scatter(num_articles, auc_one_transformer, label="One Transformer Block")
plt.title("AUC / Number of Articles")
plt.legend()
plt.show()
plt.figure(figsize=(26,12))
bar_width = 0.4
r1 = np.arange(len(auc_all_params))
r2 = [x + bar_width for x in r1]
plt.bar(r1, auc_all_params, width=bar_width, edgecolor="white", label="All Parameters")
plt.bar(r2, auc_one_transformer, width=bar_width, edgecolor="white", label="One Transformer Block")
plt.xticks([r + bar_width/2 for r in range(len(auc_all_params))], mm_codes)
plt.title("Individual AUC's for Sub-Categories to \"Kultur och nöje\"")
plt.legend()
plt.show()
# -
# <h1>Junk</h1>
[[270 5 1 11 3 8 1 11 0 8 4 1 3 1 1 0 0]
[ 12 184 9 7 4 2 2 0 3 4 4 0 0 2 2 3 0]
[ 3 2 87 2 0 1 6 0 0 1 1 0 0 4 0 2 7]
[ 9 1 0 220 2 15 6 1 8 13 17 1 4 8 2 0 1]
[ 4 2 0 1 76 1 5 1 3 4 5 0 2 1 0 0 1]
[ 6 0 0 7 2 79 2 0 0 12 1 0 3 1 0 1 0]
[ 25 0 0 3 4 2 158 7 12 6 4 6 3 6 6 0 0]
[ 15 0 2 1 3 4 3 46 0 8 0 5 0 2 1 2 2]
[ 5 0 0 4 0 0 2 0 33 1 5 1 0 2 0 0 0]
[ 10 1 0 2 1 11 1 2 0 123 0 0 9 2 3 0 3]
[ 13 2 1 8 2 3 10 1 3 2 53 2 0 6 2 3 1]
[ 1 0 0 0 0 0 0 0 0 0 0 9 0 0 1 0 0]
[ 8 0 0 1 1 2 1 1 1 3 0 0 31 1 0 0 0] # Teknik och vetenskap
[ 4 3 2 4 2 6 7 0 1 2 2 0 0 32 1 1 4] # Samhälle
[ 0 0 0 0 0 0 0 0 0 0 0 0 0 0 136 0 1] # Sport
[ 1 1 0 0 0 0 0 0 0 0 1 0 0 0 0 8 0]
[ 2 0 2 0 0 1 0 0 0 0 0 0 0 0 0 0 36]]
# +
pred
actual [[TP FN]
[FP TN]]
Samhälle & Konst, kultur och nöje
[[ 89 178]
[ 0 0]]
Samhälle & Brott, lag och rätt
[[ 84 191]
[ 0 0]]
Samhälle & Katastrofer och olyckor
[[ 91 201]
[ 0 0]]
Samhälle & Ekonomi, affärer och finans
[[ 75 193]
[ 0 0]]
Samhälle & Utbildning
[[ 85 200]
[ 0 0]]
Samhälle & Miljö och natur
[[ 90 197]
[ 0 0]]
Samhälle & Medicin och hälsa
[[ 72 155]
[ 0 0]]
Samhälle & Mänskligt
[[ 89 180]
[ 0 0]]
Samhälle & Arbete
[[ 88 196]
[ 0 0]]
Samhälle & Fritid och livsstil
[[ 88 196]
[ 0 0]]
Samhälle & Politik
[[ 68 152]
[ 0 0]]
Samhälle & Etik och religion
[[ 93 208]
[ 0 0]]
Samhälle & Teknik och vetenskap
[[ 93 204]
[ 0 0]]
Samhälle & Samhälle
[[0 0]
[0 0]]
Samhälle & Sport
[[ 91 199]
[ 0 0]]
Samhälle & Krig, konflikter och oroligheter
[[ 93 206]
[ 0 0]]
Samhälle & Väder
[[ 91 208]
[ 0 0]]
# +
# Samhälle vs all
# Jämföra accuracy e.d. för samtliga kategoripar
13 0
[[ 89 178]
[ 1 287]]
13 1
[[ 84 191]
[ 1 237]]
13 2
[[ 91 201]
[ 5 125]]
13 3
[[ 73 195]
[ 8 307]]
13 4
[[ 85 200]
[ 2 106]]
13 5
[[ 90 197]
[ 1 136]]
13 6
[[ 66 161]
[ 3 307]]
13 7
[[ 86 183]
[ 2 163]]
13 8
[[ 85 199]
[ 4 107]]
13 9
[[ 88 196]
[ 3 274]]
13 10
[[ 62 158]
[ 11 270]]
13 11
[[ 93 208]
[ 0 34]]
13 12
[[ 93 204]
[ 3 114]]
13 13
[[0 0]
[0 0]]
13 14
[[ 91 199]
[ 1 177]]
13 15
[[ 93 206]
[ 1 49]]
13 16
[[ 89 210]
[ 0 71]]
| mltc/evaluation_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ipywidgets
import pandas as pd
import pymongo
from bson.objectid import ObjectId
# +
client = pymongo.MongoClient()
database = 'uwnet'
run_collection_name = 'runs'
metric_collection_name = 'metrics'
# -
client[database].list_collection_names()
# +
collection = client[database][run_collection_name]
metrics = client[database][metric_collection_name]d
# -
# this is what one record looks like
# +
runs_with_artifacts = collection\
.find({"$where": "this.artifacts.length > 0"})\
.sort([("start_time", pymongo.DESCENDING)])
run = runs_with_artifacts.next()
# Drop all entries without artifacts
collection.find({"$where": "this.artifacts.length == 0", 'status': {"$ne": "RUNNING"}})
collection.delete_many({"$where": "this.artifacts.length == 0", 'status': {"$ne": "RUNNING"}})
metrics_dict = {m['name']: ObjectId(m['id']) for m in run['info']['metrics']}
loss = metrics.find_one({'_id': metrics_dict['loss']})
loss_series = pd.Series(loss['values'], index=loss['steps'])
loss_series.plot()
| notebooks/sacred-runs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py38pytorch
# language: python
# name: py38pytorch
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('ggplot')
plt.rcParams['figure.figsize'] = (12, 6)
# +
"""
This file contains code that will kick off training and testing processes
"""
import os
import json
import numpy as np
from experiments.UNetExperiment import UNetExperiment
from data_prep.HippocampusDatasetLoader import LoadHippocampusData
class Config:
"""
Holds configuration parameters
"""
def __init__(self):
"""
Defines important parameters:
name: name of algorithm (neural network)
root_dir:
n_epochs: number of epochs to train
learning_rate: initial learning rate
batch_size: size of image/label batches to feed the neural network at training at once
patch_size:
test_results_dir:
"""
self.name = 'Basic_unet'
# self.root_dir = r'../data/TrainingSet/'
self.root_dir = r'../../section1/data/TrainingSet/'
self.n_epochs = 1
self.learning_rate = 0.0002
self.batch_size = 8
self.patch_size = 64
self.test_results_dir = '../results/'
# +
# set random seed generator:
np.random.seed(seed=5)
# define relative training, validation and testing size:
train_size = 0.8
valid_size = 0.1
test_size = 0.1
# Get configuration
# TASK: Fill in parameters of the Config class and specify directory where the data is stored and
# directory where results will go
c = Config()
# +
# Load data
print("Loading data...")
# TASK: LoadHippocampusData is not complete. Go to the implementation and complete it.
data = LoadHippocampusData(root_dir=c.root_dir, y_shape=c.patch_size,
z_shape=c.patch_size)
# -
data[0]['filename']
data[0]['image'].shape
# +
# Create test-train-val split
# In a real world scenario you would probably do multiple splits for
# multi-fold training to improve your model quality
keys = range(len(data))
# Here, random permutation of keys array would be useful in case if we do something like
# a k-fold training and combining the results.
keys = np.array(keys)
np.random.shuffle(keys)
split = dict()
# TASK: create three keys in the dictionary: "train", "val" and "test". In each key, store
# the array with indices of training volumes to be used for training, validation
# and testing respectively.
# <YOUR CODE GOES HERE>
split['train'] = keys[:int(train_size*len(keys))]
split['val'] = keys[int(train_size*len(keys)):int((train_size+valid_size)*len(keys))]
split['test'] = keys[int((train_size+valid_size)*len(keys)):]
# -
split['test']
# +
# Set up and run experiment
# TASK: Class UNetExperiment has missing pieces. Go to the file and fill them in
exp = UNetExperiment(config=c, split=split, dataset=data)
# +
# You could free up memory by deleting the dataset
# as it has been copied into loaders
# del dataset
# run training
exp.run()
# +
# prep and run testing
# TASK: Test method is not complete. Go to the method and complete it
results_json = exp.run_test()
# +
results_json["config"] = vars(c)
with open(os.path.join(exp.out_dir, 'results.json'), 'w') as out_file:
json.dump(results_json, out_file, indent=2, separators=(',', ': '))
# -
j
| section2/src/run_ml_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Script to build Markdown pages that provide term metadata for simple vocabularies
# <NAME> 2020-06-28 CC0
# This script merges static Markdown header and footer documents with term information tables (in Markdown) generated from data in the rs.tdwg.org repo from the TDWG Github site
# Note: this script calls a function from http_library.py, which requires importing the requests, csv, and json modules
import re
import requests # best library to manage HTTP transactions
import csv # library to read/write/parse CSV files
import json # library to convert JSON to Python data structures
import pandas as pd
# -----------------
# Configuration section
# -----------------
# !!!! Note !!!!
# This is an example of a simple vocabulary without categories. For a complex example
# with multiple namespaces and several categories, see build-page-categories.ipynb
# This is the base URL for raw files from the branch of the repo that has been pushed to GitHub. In this example,
# the branch is named "pathway"
githubBaseUri = 'https://raw.githubusercontent.com/tdwg/rs.tdwg.org/master/'
headerFileName = 'termlist-header.md'
footerFileName = 'termlist-footer.md'
outFileName = '../../docs/subtype.md'
# This is a Python list of the database names of the term lists to be included in the document.
termLists = ['acsubtype']
# NOTE! There may be problems unless every term list is of the same vocabulary type since the number of columns will differ
# However, there probably aren't any circumstances where mixed types will be used to generate the same page.
vocab_type = 2 # 1 is simple vocabulary, 2 is simple controlled vocabulary, 3 is c.v. with broader hierarchy
# Terms in large vocabularies like Darwin and Audubon Cores may be organized into categories using tdwgutility_organizedInClass
# If so, those categories can be used to group terms in the generated term list document.
organized_in_categories = False
# If organized in categories, the display_order list must contain the IRIs that are values of tdwgutility_organizedInClass
# If not organized into categories, the value is irrelevant. There just needs to be one item in the list.
display_order = ['']
display_label = ['Vocabulary'] # these are the section labels for the categories in the page
display_comments = [''] # these are the comments about the category to be appended following the section labels
display_id = ['Vocabulary'] # these are the fragment identifiers for the associated sections for the categories
# ---------------
# Function definitions
# ---------------
# replace URL with link
#
def createLinks(text):
def repl(match):
if match.group(1)[-1] == '.':
return '<a href="' + match.group(1)[:-1] + '">' + match.group(1)[:-1] + '</a>.'
return '<a href="' + match.group(1) + '">' + match.group(1) + '</a>'
pattern = '(https?://[^\s,;\)"<]*)'
result = re.sub(pattern, repl, text)
return result
# 2021-08-05 Add code to convert backticks copied from the DwC QRG build script written by <NAME>
def convert_code(text_with_backticks):
"""Takes all back-quoted sections in a text field and converts it to
the html tagged version of code blocks <code>...</code>
"""
return re.sub(r'`([^`]*)`', r'<code>\1</code>', text_with_backticks)
def convert_link(text_with_urls):
"""Takes all links in a text field and converts it to the html tagged
version of the link
"""
def _handle_matched(inputstring):
"""quick hack version of url handling on the current prime versions data"""
url = inputstring.group()
return "<a href=\"{}\">{}</a>".format(url, url)
regx = "(http[s]?://[\w\d:#@%/;$()~_?\+-;=\\\.&]*)(?<![\)\.,])"
return re.sub(regx, _handle_matched, text_with_urls)
# +
term_lists_info = []
frame = pd.read_csv(githubBaseUri + 'term-lists/term-lists.csv', na_filter=False)
for termList in termLists:
term_list_dict = {'list_iri': termList}
term_list_dict = {'database': termList}
for index,row in frame.iterrows():
if row['database'] == termList:
term_list_dict['pref_ns_prefix'] = row['vann_preferredNamespacePrefix']
term_list_dict['pref_ns_uri'] = row['vann_preferredNamespaceUri']
term_list_dict['list_iri'] = row['list']
term_lists_info.append(term_list_dict)
print(term_lists_info)
# +
# Create column list
column_list = ['pref_ns_prefix', 'pref_ns_uri', 'term_localName', 'label', 'definition', 'usage', 'notes', 'examples', 'term_modified', 'term_deprecated', 'type']
if vocab_type == 2:
column_list += ['controlled_value_string']
elif vocab_type == 3:
column_list += ['controlled_value_string', 'skos_broader']
if organized_in_categories:
column_list.append('tdwgutility_organizedInClass')
column_list.append('version_iri')
# Create list of lists metadata table
table_list = []
for term_list in term_lists_info:
# retrieve versions metadata for term list
versions_url = githubBaseUri + term_list['database'] + '-versions/' + term_list['database'] + '-versions.csv'
versions_df = pd.read_csv(versions_url, na_filter=False)
# retrieve current term metadata for term list
data_url = githubBaseUri + term_list['database'] + '/' + term_list['database'] + '.csv'
frame = pd.read_csv(data_url, na_filter=False)
for index,row in frame.iterrows():
row_list = [term_list['pref_ns_prefix'], term_list['pref_ns_uri'], row['term_localName'], row['label'], row['definition'], row['usage'], row['notes'], row['examples'], row['term_modified'], row['term_deprecated'], row['type']]
if vocab_type == 2:
row_list += [row['controlled_value_string']]
elif vocab_type == 3:
if row['skos_broader'] =='':
row_list += [row['controlled_value_string'], '']
else:
row_list += [row['controlled_value_string'], term_list['pref_ns_prefix'] + ':' + row['skos_broader']]
if organized_in_categories:
row_list.append(row['tdwgutility_organizedInClass'])
# Borrowed terms really don't have implemented versions. They may be lacking values for version_status.
# In their case, their version IRI will be omitted.
found = False
for vindex, vrow in versions_df.iterrows():
if vrow['term_localName']==row['term_localName'] and vrow['version_status']=='recommended':
found = True
version_iri = vrow['version']
# NOTE: the current hack for non-TDWG terms without a version is to append # to the end of the term IRI
if version_iri[len(version_iri)-1] == '#':
version_iri = ''
if not found:
version_iri = ''
row_list.append(version_iri)
table_list.append(row_list)
# Turn list of lists into dataframe
terms_df = pd.DataFrame(table_list, columns = column_list)
terms_sorted_by_label = terms_df.sort_values(by='label')
terms_sorted_by_localname = terms_df.sort_values(by='term_localName')
terms_sorted_by_label
# -
# Run the following cell to generate an index sorted alphabetically by lowercase term local name. Omit this index if the terms have opaque local names.
# +
# generate the index of terms grouped by category and sorted alphabetically by lowercase term local name
text = '### 3.1 Index By Term Name\n\n'
text += '(See also [3.2 Index By Label](#32-index-by-label))\n\n'
for category in range(0,len(display_order)):
text += '**' + display_label[category] + '**\n'
text += '\n'
if organized_in_categories:
filtered_table = terms_sorted_by_localname[terms_sorted_by_localname['tdwgutility_organizedInClass']==display_order[category]]
filtered_table.reset_index(drop=True, inplace=True)
else:
filtered_table = terms_sorted_by_localname
filtered_table.reset_index(drop=True, inplace=True)
for row_index,row in filtered_table.iterrows():
curie = row['pref_ns_prefix'] + ":" + row['term_localName']
curie_anchor = curie.replace(':','_')
text += '[' + curie + '](#' + curie_anchor + ') |\n'
text = text[:len(text)-2] # remove final trailing vertical bar and newline
text += '\n\n' # put back removed newline
index_by_name = text
print(index_by_name)
# -
# Run the following cell to generate an index by term label
# +
text = '\n\n'
# Comment out the following two lines if there is no index by local names
#text = '### 3.2 Index By Label\n\n'
#text += '(See also [3.1 Index By Term Name](#31-index-by-term-name))\n\n'
for category in range(0,len(display_order)):
if organized_in_categories:
text += '**' + display_label[category] + '**\n'
text += '\n'
filtered_table = terms_sorted_by_label[terms_sorted_by_label['tdwgutility_organizedInClass']==display_order[category]]
filtered_table.reset_index(drop=True, inplace=True)
else:
filtered_table = terms_sorted_by_label
filtered_table.reset_index(drop=True, inplace=True)
for row_index,row in filtered_table.iterrows():
if row_index == 0 or (row_index != 0 and row['label'] != filtered_table.iloc[row_index - 1].loc['label']): # this is a hack to prevent duplicate labels
curie_anchor = row['pref_ns_prefix'] + "_" + row['term_localName']
text += '[' + row['label'] + '](#' + curie_anchor + ') |\n'
text = text[:len(text)-2] # remove final trailing vertical bar and newline
text += '\n\n' # put back removed newline
index_by_label = text
print(index_by_label)
# +
decisions_df = pd.read_csv('https://raw.githubusercontent.com/tdwg/rs.tdwg.org/master/decisions/decisions-links.csv', na_filter=False)
# generate a table for each term, with terms grouped by category
# generate the Markdown for the terms table
text = '## 4 Vocabulary\n'
for category in range(0,len(display_order)):
if organized_in_categories:
text += '### 4.' + str(category + 1) + ' ' + display_label[category] + '\n'
text += '\n'
text += display_comments[category] # insert the comments for the category, if any.
filtered_table = terms_sorted_by_localname[terms_sorted_by_localname['tdwgutility_organizedInClass']==display_order[category]]
filtered_table.reset_index(drop=True, inplace=True)
else:
filtered_table = terms_sorted_by_localname
filtered_table.reset_index(drop=True, inplace=True)
for row_index,row in filtered_table.iterrows():
text += '<table>\n'
curie = row['pref_ns_prefix'] + ":" + row['term_localName']
curieAnchor = curie.replace(':','_')
text += '\t<thead>\n'
text += '\t\t<tr>\n'
text += '\t\t\t<th colspan="2"><a id="' + curieAnchor + '"></a>Term Name ' + curie + '</th>\n'
text += '\t\t</tr>\n'
text += '\t</thead>\n'
text += '\t<tbody>\n'
text += '\t\t<tr>\n'
text += '\t\t\t<td>Term IRI</td>\n'
uri = row['pref_ns_uri'] + row['term_localName']
text += '\t\t\t<td><a href="' + uri + '">' + uri + '</a></td>\n'
text += '\t\t</tr>\n'
text += '\t\t<tr>\n'
text += '\t\t\t<td>Modified</td>\n'
text += '\t\t\t<td>' + row['term_modified'] + '</td>\n'
text += '\t\t</tr>\n'
if row['version_iri'] != '':
text += '\t\t<tr>\n'
text += '\t\t\t<td>Term version IRI</td>\n'
text += '\t\t\t<td><a href="' + row['version_iri'] + '">' + row['version_iri'] + '</a></td>\n'
text += '\t\t</tr>\n'
text += '\t\t<tr>\n'
text += '\t\t\t<td>Label</td>\n'
text += '\t\t\t<td>' + row['label'] + '</td>\n'
text += '\t\t</tr>\n'
if row['term_deprecated'] != '':
text += '\t\t<tr>\n'
text += '\t\t\t<td></td>\n'
text += '\t\t\t<td><strong>This term is deprecated and should no longer be used.</strong></td>\n'
text += '\t\t</tr>\n'
text += '\t\t<tr>\n'
text += '\t\t\t<td>Definition</td>\n'
text += '\t\t\t<td>' + row['definition'] + '</td>\n'
text += '\t\t</tr>\n'
if row['usage'] != '':
text += '\t\t<tr>\n'
text += '\t\t\t<td>Usage</td>\n'
text += '\t\t\t<td>' + convert_link(convert_code(row['usage'])) + '</td>\n'
text += '\t\t</tr>\n'
if row['notes'] != '':
text += '\t\t<tr>\n'
text += '\t\t\t<td>Notes</td>\n'
text += '\t\t\t<td>' + convert_link(rconvert_code(ow['notes'])) + '</td>\n'
text += '\t\t</tr>\n'
if row['examples'] != '':
text += '\t\t<tr>\n'
text += '\t\t\t<td>Examples</td>\n'
text += '\t\t\t<td>' + convert_link(convert_code(row['examples'])) + '</td>\n'
text += '\t\t</tr>\n'
if (vocab_type == 2 or vocab_type == 3) and row['controlled_value_string'] != '': # controlled vocabulary
text += '\t\t<tr>\n'
text += '\t\t\t<td>Controlled value</td>\n'
text += '\t\t\t<td>' + row['controlled_value_string'] + '</td>\n'
text += '\t\t</tr>\n'
if vocab_type == 3 and row['skos_broader'] != '': # controlled vocabulary with skos:broader relationships
text += '\t\t<tr>\n'
text += '\t\t\t<td>Has broader concept</td>\n'
curieAnchor = row['skos_broader'].replace(':','_')
text += '\t\t\t<td><a href="#' + curieAnchor + '">' + row['skos_broader'] + '</a></td>\n'
text += '\t\t</tr>\n'
text += '\t\t<tr>\n'
text += '\t\t\t<td>Type</td>\n'
if row['type'] == 'http://www.w3.org/1999/02/22-rdf-syntax-ns#Property':
text += '\t\t\t<td>Property</td>\n'
elif row['type'] == 'http://www.w3.org/2000/01/rdf-schema#Class':
text += '\t\t\t<td>Class</td>\n'
elif row['type'] == 'http://www.w3.org/2004/02/skos/core#Concept':
text += '\t\t\t<td>Concept</td>\n'
else:
text += '\t\t\t<td>' + row['type'] + '</td>\n' # this should rarely happen
text += '\t\t</tr>\n'
# Look up decisions related to this term
for drow_index,drow in decisions_df.iterrows():
if drow['linked_affected_resource'] == uri:
text += '\t\t<tr>\n'
text += '\t\t\t<td>Executive Committee decision</td>\n'
text += '\t\t\t<td><a href="http://rs.tdwg.org/decisions/' + drow['decision_localName'] + '">http://rs.tdwg.org/decisions/' + drow['decision_localName'] + '</a></td>\n'
text += '\t\t</tr>\n'
text += '\t</tbody>\n'
text += '</table>\n'
text += '\n'
text += '\n'
term_table = text
print(term_table)
# -
# Modify to display the indices that you want
text = index_by_label + term_table
#text = index_by_name + index_by_label + term_table
# +
# read in header and footer, merge with terms table, and output
headerObject = open(headerFileName, 'rt', encoding='utf-8')
header = headerObject.read()
headerObject.close()
footerObject = open(footerFileName, 'rt', encoding='utf-8')
footer = footerObject.read()
footerObject.close()
output = header + text + footer
outputObject = open(outFileName, 'wt', encoding='utf-8')
outputObject.write(output)
outputObject.close()
print('done')
# -
| code/build_subtype_cv/build-page-simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Tensorflow DALI plugin: DALI and tf.data
#
# ### Overview
#
# DALI offers integration with [tf.data API](https://www.tensorflow.org/guide/data). Using this approach you can easily connect DALI pipeline with various TensorFlow APIs and use it as a data source for your model. This tutorial shows how to do it using well known [MNIST](http://yann.lecun.com/exdb/mnist/) converted to LMDB format. You can find it in [DALI_extra](https://github.com/NVIDIA/DALI_extra) - DALI test data repository.
#
# We start with creating a DALI pipeline to read, decode and normalize MNIST images and read corresponding labels.
#
# `DALI_EXTRA_PATH` environment variable should point to the place where data from [DALI extra repository](https://github.com/NVIDIA/DALI_extra) is downloaded. Please make sure that the proper release tag is checked out.
# +
import nvidia.dali as dali
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
import os
# Path to MNIST dataset
data_path = os.path.join(os.environ['DALI_EXTRA_PATH'], 'db/MNIST/training/')
class MnistPipeline(Pipeline):
def __init__(self, batch_size, device, device_id=0, num_threads=4, seed=0):
super(MnistPipeline, self).__init__(
batch_size, num_threads, device_id, seed)
self.device = device
self.reader = ops.Caffe2Reader(path=data_path, random_shuffle=True)
self.decode = ops.ImageDecoder(
device='mixed' if device is 'gpu' else 'cpu',
output_type=types.GRAY)
self.cmn = ops.CropMirrorNormalize(
device=device,
output_dtype=types.FLOAT,
image_type=types.GRAY,
std=[255.],
output_layout="CHW")
def define_graph(self):
inputs, labels = self.reader(name="Reader")
images = self.decode(inputs)
if self.device is 'gpu':
labels = labels.gpu()
images = self.cmn(images)
return (images, labels)
# -
# Now we define some parameters of the training:
BATCH_SIZE = 64
DROPOUT = 0.2
IMAGE_SIZE = 28
NUM_CLASSES = 10
HIDDEN_SIZE = 128
EPOCHS = 5
ITERATIONS_PER_EPOCH = 100
# Next step is to wrap an instance of `MnistPipeline` with a `DALIDataset` object from DALI TensorFlow plugin. This class is compatible with `tf.data.Dataset`. Other parameters are shapes and types of the outputs of the pipeline. Here we return images and labels. It means we have two outputs one of type `tf.float32` for images and on of type `tf.int32` for labels.
# +
import nvidia.dali.plugin.tf as dali_tf
import tensorflow.compat.v1 as tf
tf.logging.set_verbosity(tf.logging.ERROR)
tf.disable_eager_execution()
# Create pipeline
mnist_pipeline = MnistPipeline(BATCH_SIZE, device='cpu', device_id=0)
# Define shapes and types of the outputs
shapes = [
(BATCH_SIZE, IMAGE_SIZE, IMAGE_SIZE),
(BATCH_SIZE)]
dtypes = [
tf.float32,
tf.int32]
# Create dataset
mnist_set = dali_tf.DALIDataset(
pipeline=mnist_pipeline,
batch_size=BATCH_SIZE,
shapes=shapes,
dtypes=dtypes,
device_id=0)
# -
# We are ready to start the training. Following sections show how to do it with different APIs availible in TensorFlow.
#
# ### Keras
#
# First, we pass `mnist_set` to model created with `tf.keras` and use `model.fit` method to train it.
# +
# Create the model
model = tf.keras.models.Sequential([
tf.keras.layers.Input(shape=(IMAGE_SIZE, IMAGE_SIZE), name='images'),
tf.keras.layers.Flatten(input_shape=(IMAGE_SIZE, IMAGE_SIZE)),
tf.keras.layers.Dense(HIDDEN_SIZE, activation='relu'),
tf.keras.layers.Dropout(DROPOUT),
tf.keras.layers.Dense(NUM_CLASSES, activation='softmax')])
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train using DALI dataset
model.fit(
mnist_set,
epochs=EPOCHS,
steps_per_epoch=ITERATIONS_PER_EPOCH)
# -
# As you can see, it was very easy to integrate DALI pipeline with `tf.keras` API.
#
# The code above performed the training using the CPU. Both the DALI pipeline and the model were using the CPU.
#
# We can easily move the whole processing to the GPU. First, we create a pipeline that uses the GPU with ID = 0. Next we place both the DALI dataset and the model on the same GPU.
# +
# Create pipeline
mnist_pipeline = MnistPipeline(BATCH_SIZE, device='gpu', device_id=0)
# Define the model and place it on the GPU
with tf.device('/gpu:0'):
mnist_set = dali_tf.DALIDataset(
pipeline=mnist_pipeline,
batch_size=BATCH_SIZE,
shapes=shapes,
dtypes=dtypes,
device_id=0)
model = tf.keras.models.Sequential([
tf.keras.layers.Input(shape=(IMAGE_SIZE, IMAGE_SIZE), name='images'),
tf.keras.layers.Flatten(input_shape=(IMAGE_SIZE, IMAGE_SIZE)),
tf.keras.layers.Dense(HIDDEN_SIZE, activation='relu'),
tf.keras.layers.Dropout(DROPOUT),
tf.keras.layers.Dense(NUM_CLASSES, activation='softmax')])
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# -
# We move the training to the GPU as well. This allows TensorFlow to pick up GPU instance of DALI dataset.
# Train on the GPU
with tf.device('/gpu:0'):
model.fit(
mnist_set,
epochs=EPOCHS,
steps_per_epoch=ITERATIONS_PER_EPOCH)
# It is important to note here, that there is no intermediate CPU buffer between DALI and TensorFlow in the execution above. DALI GPU outputs are copied straight to TF GPU Tensors used by the model.
#
# In this particular toy example performance of the GPU variant is lower than the CPU one. The MNIST images are small and nvJPEG decoder used in the GPU DALI pipeline to decode them is not well suited for such circumstance. We use it here to show how to integrate it properly in the real life case.
#
#
# ### Estimators
#
# Another popular TensorFlow API is `tf.estimator` API. This section shows how to use DALI dataset as a data source for model based on this API.
#
# First we create the model.
# +
# Define the feature columns for Estimator
feature_columns = [tf.feature_column.numeric_column(
"images", shape=[IMAGE_SIZE, IMAGE_SIZE])]
# And the run config
run_config = tf.estimator.RunConfig(
model_dir='/tmp/tensorflow-checkpoints',
device_fn=lambda op: '/gpu:0')
# Finally create the model based on `DNNClassifier`
model = tf.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[HIDDEN_SIZE],
n_classes=NUM_CLASSES,
dropout=DROPOUT,
config=run_config,
optimizer='Adam')
# -
# In `tf.estimator` API data is passed to the model with the function returning the dataset. We define this function to return DALI dataset placed on the GPU.
def train_data_fn():
with tf.device('/gpu:0'):
mnist_pipeline = MnistPipeline(BATCH_SIZE, device='gpu', device_id=0)
mnist_set = dali_tf.DALIDataset(
pipeline=mnist_pipeline,
batch_size=BATCH_SIZE,
shapes=shapes,
dtypes=dtypes,
device_id=0)
mnist_set = mnist_set.map(
lambda features, labels: ({'images': features}, labels))
return mnist_set
# With everything set up we are ready to run the training.
# Running the training on the GPU
model.train(input_fn=train_data_fn, steps=EPOCHS * ITERATIONS_PER_EPOCH)
model.evaluate(input_fn=train_data_fn, steps=ITERATIONS_PER_EPOCH)
# ### Custom models and training loops
#
# Finally, the last part of this tutorial focuses on integrating DALI dataset with custom models and training loops. A complete example below shows from start to finish how to use DALI dataset with native TensorFlow model and run training using `tf.Session`.
#
# First step is to define the model and the dataset and place both on the GPU.
# +
tf.reset_default_graph()
options = tf.data.Options()
options.experimental_optimization.apply_default_optimizations = False
options.experimental_optimization.autotune = False
with tf.device('/gpu:0'):
mnist_set = dali_tf.DALIDataset(
pipeline=MnistPipeline(BATCH_SIZE, device='gpu', device_id=0),
batch_size=BATCH_SIZE,
shapes=shapes,
dtypes=dtypes,
device_id=0).with_options(options)
iterator = tf.data.make_initializable_iterator(mnist_set)
images, labels = iterator.get_next()
labels = tf.reshape(
tf.one_hot(labels, NUM_CLASSES),
[BATCH_SIZE, NUM_CLASSES])
with tf.variable_scope('mnist_net', reuse=False):
images = tf.layers.flatten(images)
images = tf.layers.dense(images, HIDDEN_SIZE, activation=tf.nn.relu)
images = tf.layers.dropout(images, rate=DROPOUT, training=True)
images = tf.layers.dense(images, NUM_CLASSES, activation=tf.nn.softmax)
logits_train = images
loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
logits=logits_train, labels=labels))
train_step = tf.train.AdamOptimizer().minimize(loss_op)
correct_pred = tf.equal(
tf.argmax(logits_train, 1), tf.argmax(labels, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# -
# With `tf.Session` we can run this model and train it on the GPU.
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
sess.run(iterator.initializer)
for i in range(EPOCHS * ITERATIONS_PER_EPOCH):
sess.run(train_step)
if i % ITERATIONS_PER_EPOCH == 0:
train_accuracy = sess.run(accuracy)
print("Step %d, accuracy: %g" % (i, train_accuracy))
final_accuracy = 0
for _ in range(ITERATIONS_PER_EPOCH):
final_accuracy = final_accuracy + sess.run(accuracy)
final_accuracy = final_accuracy / ITERATIONS_PER_EPOCH
print('Final accuracy: ', final_accuracy)
| docs/examples/frameworks/tensorflow/tensorflow-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
log1=pd.read_csv('../vol1_ext_alcore/log_real.csv')
log2=pd.read_csv('../vol2_ext_alcore/log_real.csv')
log3=pd.read_csv('../vol12/log_real.csv')
def gen_grad_acc(df):
df3=df
import transforms3d as tf3d
acc_ned_grad=np.zeros((len(df3),3))
acc_ned_grad[:,0]=np.diff(df3['speed[0]'],append=0)/np.diff(df3["t"],append=4e-3)
acc_ned_grad[:,0]=np.array([i if abs(i)<30 else 0 for i in acc_ned_grad[:,0]])
acc_ned_grad[:,1]=np.diff(df3['speed[1]'],append=0)/np.diff(df3["t"],append=4e-3)
acc_ned_grad[:,1]=np.array([i if abs(i)<30 else 0 for i in acc_ned_grad[:,1]])
acc_ned_grad[:,2]=np.diff(df3['speed[2]'],append=0)/np.diff(df3["t"],append=4e-3)
acc_ned_grad[:,2]=np.array([i if abs(i)<30 else 0 for i in acc_ned_grad[:,2]])
acc_body_grad=np.zeros((len(df3),3))
speed_body=np.zeros((len(df3),3))
gamma=np.zeros((len(df3),3))
for i in df3.index:
q0,q1,q2,q3=df3["q[0]"][i],df3["q[1]"][i],df3["q[2]"][i],df3["q[3]"][i]
# print(i,q0,q1,q2,q3)
R=tf3d.quaternions.quat2mat(np.array([q0,q1,q2,q3]))
acc_body_grad[i]=(R.T@(acc_ned_grad[i].reshape((3,1)))).flatten()
speed_body[i]=(R.T@(np.array([df3['speed[0]'][i],df3['speed[1]'][i],df3['speed[2]'][i]]).reshape((3,1)))).flatten()
gamma[i]=(R.T@np.array([[0],[0],[9.81]])).flatten()
df3['acc_ned_grad[0]'],df3['acc_ned_grad[1]'],df3['acc_ned_grad[2]']=acc_ned_grad.T
df3['acc_body_grad[0]'],df3['acc_body_grad[1]'],df3['acc_body_grad[2]']=acc_body_grad.T
df3['speed_body[0]'],df3['speed_body[1]'],df3['speed_body[2]']=speed_body.T
df3['gamma[0]'],df3['gamma[1]'],df3['gamma[2]']=gamma.T
return df3
log1_new=gen_grad_acc(log1)
log2_new=gen_grad_acc(log2)
log3_new=gen_grad_acc(log3)
log1_new.to_csv('../vol1_ext_alcore/log_real_processed.csv')
log2_new.to_csv('../vol2_ext_alcore/log_real_processed.csv')
log3_new.to_csv('../vol12/log_real_processed.csv')
# -
log1_new
log2_new
plt.plot(log1.t,log1['speed[2]'])
plt.plot(log2.t,log2['speed[2]'])
# +
log1_r=log1[log1["t"]>40]
log1_r=log1_r[log1_r["t"]<255]
log2_r=log2[log2["t"]>10]
log2_r=log2_r[log2_r['t']<140]
# -
plt.plot(log1_r.t,log1_r['speed[2]'])
plt.plot(log2_r.t,log2_r['speed[2]'])
log1_r["t"]=log1_r["t"]-log1_r['t'].min()
log2_r["t"]=log2_r["t"]-log2_r['t'].min()+5e-3+log1_r["t"].max()
plt.plot(log1_r.t,log1_r['speed[2]'])
plt.plot(log2_r.t,log2_r['speed[2]'])
df3=pd.concat([log1_r,log2_r])
plt.plot(df3.t,df3['speed[2]'])
df3=df3.drop(columns=["Unnamed: 0", "level_0","Unnamed: 0.1"])
df3
df3=df3.reset_index()
import transforms3d as tf3d
# +
acc_ned_grad=np.zeros((len(df3),3))
acc_ned_grad[:,0]=np.diff(df3['speed[0]'],append=0)/np.diff(df3["t"],append=4e-3)
acc_ned_grad[:,0]=np.array([i if abs(i)<30 else 0 for i in acc_ned_grad[:,0]])
acc_ned_grad[:,1]=np.diff(df3['speed[1]'],append=0)/np.diff(df3["t"],append=4e-3)
acc_ned_grad[:,1]=np.array([i if abs(i)<30 else 0 for i in acc_ned_grad[:,1]])
acc_ned_grad[:,2]=np.diff(df3['speed[2]'],append=0)/np.diff(df3["t"],append=4e-3)
acc_ned_grad[:,2]=np.array([i if abs(i)<30 else 0 for i in acc_ned_grad[:,2]])
# +
acc_body_grad=np.zeros((len(df3),3))
for i in df3.index:
q0,q1,q2,q3=df3["q[0]"][i],df3["q[1]"][i],df3["q[2]"][i],df3["q[3]"][i]
# print(i,q0,q1,q2,q3)
R=tf3d.quaternions.quat2mat(np.array([q0,q1,q2,q3]))
acc_body_grad[i]=R.T@(acc_ned_grad[i].reshape((3,1))).flatten()
# +
df3['acc_ned_grad[0]'],df3['acc_ned_grad[1]'],df3['acc_ned_grad[2]']=acc_ned_grad.T
df3['acc_body_grad[0]'],df3['acc_body_grad[1]'],df3['acc_body_grad[2]']=acc_body_grad.T
# -
df3.to_csv('./log_real.csv')
[(plt.figure(),plt.plot(df3["t"],df3['acc[%i]'%(i)]),plt.plot(df3["t"],df3['acc_ned_grad[%i]'%(i)])) for i in range(3)]
plt.gca().set_ylim(-15,15)
df3['q[0]'].plot()
df3['q[1]'].plot()
df3['q[2]'].plot()
df3['q[3]'].plot()
| logs/copter/vol2/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # tree.py
# ```python
# # Copyright 2013, <NAME>
# #
# # Developed for use with the book:
# #
# # Data Structures and Algorithms in Python
# # <NAME>, <NAME>, and <NAME>
# # <NAME>, 2013
# #
# # This program is free software: you can redistribute it and/or modify
# # it under the terms of the GNU General Public License as published by
# # the Free Software Foundation, either version 3 of the License, or
# # (at your option) any later version.
# #
# # This program is distributed in the hope that it will be useful,
# # but WITHOUT ANY WARRANTY; without even the implied warranty of
# # MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# # GNU General Public License for more details.
# #
# # You should have received a copy of the GNU General Public License
# # along with this program. If not, see <http://www.gnu.org/licenses/>.
#
# import collections
#
# class Tree:
# """Abstract base class representing a tree structure."""
#
# #------------------------------- nested Position class -------------------------------
# class Position:
# """An abstraction representing the location of a single element within a tree.
#
# Note that two position instaces may represent the same inherent location in a tree.
# Therefore, users should always rely on syntax 'p == q' rather than 'p is q' when testing
# equivalence of positions.
# """
#
# def element(self):
# """Return the element stored at this Position."""
# raise NotImplementedError('must be implemented by subclass')
#
# def __eq__(self, other):
# """Return True if other Position represents the same location."""
# raise NotImplementedError('must be implemented by subclass')
#
# def __ne__(self, other):
# """Return True if other does not represent the same location."""
# return not (self == other) # opposite of __eq__
#
# # ---------- abstract methods that concrete subclass must support ----------
# def root(self):
# """Return Position representing the tree's root (or None if empty)."""
# raise NotImplementedError('must be implemented by subclass')
#
# def parent(self, p):
# """Return Position representing p's parent (or None if p is root)."""
# raise NotImplementedError('must be implemented by subclass')
#
# def num_children(self, p):
# """Return the number of children that Position p has."""
# raise NotImplementedError('must be implemented by subclass')
#
# def children(self, p):
# """Generate an iteration of Positions representing p's children."""
# raise NotImplementedError('must be implemented by subclass')
#
# def __len__(self):
# """Return the total number of elements in the tree."""
# raise NotImplementedError('must be implemented by subclass')
#
# # ---------- concrete methods implemented in this class ----------
# def is_root(self, p):
# """Return True if Position p represents the root of the tree."""
# return self.root() == p
#
# def is_leaf(self, p):
# """Return True if Position p does not have any children."""
# return self.num_children(p) == 0
#
# def is_empty(self):
# """Return True if the tree is empty."""
# return len(self) == 0
#
# def depth(self, p):
# """Return the number of levels separating Position p from the root."""
# if self.is_root(p):
# return 0
# else:
# return 1 + self.depth(self.parent(p))
#
# def _height1(self): # works, but O(n^2) worst-case time
# """Return the height of the tree."""
# return max(self.depth(p) for p in self.positions() if self.is_leaf(p))
#
# def _height2(self, p): # time is linear in size of subtree
# """Return the height of the subtree rooted at Position p."""
# if self.is_leaf(p):
# return 0
# else:
# return 1 + max(self._height2(c) for c in self.children(p))
#
# def height(self, p=None):
# """Return the height of the subtree rooted at Position p.
#
# If p is None, return the height of the entire tree.
# """
# if p is None:
# p = self.root()
# return self._height2(p) # start _height2 recursion
#
# def __iter__(self):
# """Generate an iteration of the tree's elements."""
# for p in self.positions(): # use same order as positions()
# yield p.element() # but yield each element
#
# def positions(self):
# """Generate an iteration of the tree's positions."""
# return self.preorder() # return entire preorder iteration
#
# def preorder(self):
# """Generate a preorder iteration of positions in the tree."""
# if not self.is_empty():
# for p in self._subtree_preorder(self.root()): # start recursion
# yield p
#
# def _subtree_preorder(self, p):
# """Generate a preorder iteration of positions in subtree rooted at p."""
# yield p # visit p before its subtrees
# for c in self.children(p): # for each child c
# for other in self._subtree_preorder(c): # do preorder of c's subtree
# yield other # yielding each to our caller
#
# def postorder(self):
# """Generate a postorder iteration of positions in the tree."""
# if not self.is_empty():
# for p in self._subtree_postorder(self.root()): # start recursion
# yield p
#
# def _subtree_postorder(self, p):
# """Generate a postorder iteration of positions in subtree rooted at p."""
# for c in self.children(p): # for each child c
# for other in self._subtree_postorder(c): # do postorder of c's subtree
# yield other # yielding each to our caller
# yield p # visit p after its subtrees
# ```
# # binary_tree.py
#
# ```python
# # Copyright 2013, <NAME>
# #
# # Developed for use with the book:
# #
# # Data Structures and Algorithms in Python
# # <NAME>, <NAME>, and <NAME>
# # <NAME>, 2013
# #
# # This program is free software: you can redistribute it and/or modify
# # it under the terms of the GNU General Public License as published by
# # the Free Software Foundation, either version 3 of the License, or
# # (at your option) any later version.
# #
# # This program is distributed in the hope that it will be useful,
# # but WITHOUT ANY WARRANTY; without even the implied warranty of
# # MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# # GNU General Public License for more details.
# #
# # You should have received a copy of the GNU General Public License
# # along with this program. If not, see <http://www.gnu.org/licenses/>.
#
# from tree import Tree
#
# class BinaryTree(Tree):
# """Abstract base class representing a binary tree structure."""
#
# # --------------------- additional abstract methods ---------------------
# def left(self, p):
# """Return a Position representing p's left child.
#
# Return None if p does not have a left child.
# """
# raise NotImplementedError('must be implemented by subclass')
#
# def right(self, p):
# """Return a Position representing p's right child.
#
# Return None if p does not have a right child.
# """
# raise NotImplementedError('must be implemented by subclass')
#
# # ---------- concrete methods implemented in this class ----------
# def sibling(self, p):
# """Return a Position representing p's sibling (or None if no sibling)."""
# parent = self.parent(p)
# if parent is None: # p must be the root
# return None # root has no sibling
# else:
# if p == self.left(parent):
# return self.right(parent) # possibly None
# else:
# return self.left(parent) # possibly None
#
# def children(self, p):
# """Generate an iteration of Positions representing p's children."""
# if self.left(p) is not None:
# yield self.left(p)
# if self.right(p) is not None:
# yield self.right(p)
#
# def inorder(self):
# """Generate an inorder iteration of positions in the tree."""
# if not self.is_empty():
# for p in self._subtree_inorder(self.root()):
# yield p
#
# def _subtree_inorder(self, p):
# """Generate an inorder iteration of positions in subtree rooted at p."""
# if self.left(p) is not None: # if left child exists, traverse its subtree
# for other in self._subtree_inorder(self.left(p)):
# yield other
# yield p # visit p between its subtrees
# if self.right(p) is not None: # if right child exists, traverse its subtree
# for other in self._subtree_inorder(self.right(p)):
# yield other
#
# # override inherited version to make inorder the default
# def positions(self):
# """Generate an iteration of the tree's positions."""
# return self.inorder() # make inorder the default
# ```
# # linked_binary_tree.py
#
# ```python
# # Copyright 2013, <NAME>
# #
# # Developed for use with the book:
# #
# # Data Structures and Algorithms in Python
# # <NAME>, <NAME>, and <NAME>
# # <NAME>, 2013
# #
# # This program is free software: you can redistribute it and/or modify
# # it under the terms of the GNU General Public License as published by
# # the Free Software Foundation, either version 3 of the License, or
# # (at your option) any later version.
# #
# # This program is distributed in the hope that it will be useful,
# # but WITHOUT ANY WARRANTY; without even the implied warranty of
# # MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# # GNU General Public License for more details.
# #
# # You should have received a copy of the GNU General Public License
# # along with this program. If not, see <http://www.gnu.org/licenses/>.
#
# from binary_tree import BinaryTree
#
# class LinkedBinaryTree(BinaryTree):
# """Linked representation of a binary tree structure."""
#
# #-------------------------- nested _Node class --------------------------
# class _Node:
# """Lightweight, nonpublic class for storing a node."""
# __slots__ = '_element', '_parent', '_left', '_right' # streamline memory usage
#
# def __init__(self, element, parent=None, left=None, right=None):
# self._element = element
# self._parent = parent
# self._left = left
# self._right = right
#
# #-------------------------- nested Position class --------------------------
# class Position(BinaryTree.Position):
# """An abstraction representing the location of a single element."""
#
# def __init__(self, container, node):
# """Constructor should not be invoked by user."""
# self._container = container
# self._node = node
#
# def element(self):
# """Return the element stored at this Position."""
# return self._node._element
#
# def __eq__(self, other):
# """Return True if other is a Position representing the same location."""
# return type(other) is type(self) and other._node is self._node
#
# #------------------------------- utility methods -------------------------------
# def _validate(self, p):
# """Return associated node, if position is valid."""
# if not isinstance(p, self.Position):
# raise TypeError('p must be proper Position type')
# if p._container is not self:
# raise ValueError('p does not belong to this container')
# if p._node._parent is p._node: # convention for deprecated nodes
# raise ValueError('p is no longer valid')
# return p._node
#
# def _make_position(self, node):
# """Return Position instance for given node (or None if no node)."""
# return self.Position(self, node) if node is not None else None
#
# #-------------------------- binary tree constructor --------------------------
# def __init__(self):
# """Create an initially empty binary tree."""
# self._root = None
# self._size = 0
#
# #-------------------------- public accessors --------------------------
# def __len__(self):
# """Return the total number of elements in the tree."""
# return self._size
#
# def root(self):
# """Return the root Position of the tree (or None if tree is empty)."""
# return self._make_position(self._root)
#
# def parent(self, p):
# """Return the Position of p's parent (or None if p is root)."""
# node = self._validate(p)
# return self._make_position(node._parent)
#
# def left(self, p):
# """Return the Position of p's left child (or None if no left child)."""
# node = self._validate(p)
# return self._make_position(node._left)
#
# def right(self, p):
# """Return the Position of p's right child (or None if no right child)."""
# node = self._validate(p)
# return self._make_position(node._right)
#
# def num_children(self, p):
# """Return the number of children of Position p."""
# node = self._validate(p)
# count = 0
# if node._left is not None: # left child exists
# count += 1
# if node._right is not None: # right child exists
# count += 1
# return count
#
# #-------------------------- nonpublic mutators --------------------------
# def _add_root(self, e):
# """Place element e at the root of an empty tree and return new Position.
#
# Raise ValueError if tree nonempty.
# """
# if self._root is not None:
# raise ValueError('Root exists')
# self._size = 1
# self._root = self._Node(e)
# return self._make_position(self._root)
#
# def _add_left(self, p, e):
# """Create a new left child for Position p, storing element e.
#
# Return the Position of new node.
# Raise ValueError if Position p is invalid or p already has a left child.
# """
# node = self._validate(p)
# if node._left is not None:
# raise ValueError('Left child exists')
# self._size += 1
# node._left = self._Node(e, node) # node is its parent
# return self._make_position(node._left)
#
# def _add_right(self, p, e):
# """Create a new right child for Position p, storing element e.
#
# Return the Position of new node.
# Raise ValueError if Position p is invalid or p already has a right child.
# """
# node = self._validate(p)
# if node._right is not None:
# raise ValueError('Right child exists')
# self._size += 1
# node._right = self._Node(e, node) # node is its parent
# return self._make_position(node._right)
#
# def _replace(self, p, e):
# """Replace the element at position p with e, and return old element."""
# node = self._validate(p)
# old = node._element
# node._element = e
# return old
#
# def _delete(self, p):
# """Delete the node at Position p, and replace it with its child, if any.
#
# Return the element that had been stored at Position p.
# Raise ValueError if Position p is invalid or p has two children.
# """
# node = self._validate(p)
# if self.num_children(p) == 2:
# raise ValueError('Position has two children')
# child = node._left if node._left else node._right # might be None
# if child is not None:
# child._parent = node._parent # child's grandparent becomes parent
# if node is self._root:
# self._root = child # child becomes root
# else:
# parent = node._parent
# if node is parent._left:
# parent._left = child
# else:
# parent._right = child
# self._size -= 1
# node._parent = node # convention for deprecated node
# return node._element
#
# def _attach(self, p, t1, t2):
# """Attach trees t1 and t2, respectively, as the left and right subtrees of the external Position p.
#
# As a side effect, set t1 and t2 to empty.
# Raise TypeError if trees t1 and t2 do not match type of this tree.
# Raise ValueError if Position p is invalid or not external.
# """
# node = self._validate(p)
# if not self.is_leaf(p):
# raise ValueError('position must be leaf')
# if not type(self) is type(t1) is type(t2): # all 3 trees must be same type
# raise TypeError('Tree types must match')
# self._size += len(t1) + len(t2)
# if not t1.is_empty(): # attached t1 as left subtree of node
# t1._root._parent = node
# node._left = t1._root
# t1._root = None # set t1 instance to empty
# t1._size = 0
# if not t2.is_empty(): # attached t2 as right subtree of node
# t2._root._parent = node
# node._right = t2._root
# t2._root = None # set t2 instance to empty
# t2._size = 0
# ```
# +
# Copyright 2013, <NAME>
#
# Developed for use with the book:
#
# Data Structures and Algorithms in Python
# <NAME>, <NAME>, and <NAME>
# <NAME>, 2013
#
# This program is free software: you can redistribute it and/or modify
# it under the terms of the GNU General Public License as published by
# the Free Software Foundation, either version 3 of the License, or
# (at your option) any later version.
#
# This program is distributed in the hope that it will be useful,
# but WITHOUT ANY WARRANTY; without even the implied warranty of
# MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# GNU General Public License for more details.
#
# You should have received a copy of the GNU General Public License
# along with this program. If not, see <http://www.gnu.org/licenses/>.
from linked_binary_tree import LinkedBinaryTree
class ExpressionTree(LinkedBinaryTree):
"""An arithmetic expression tree."""
def __init__(self, token, left=None, right=None):
"""Create an expression tree.
In a single parameter form, token should be a leaf value (e.g., '42'),
and the expression tree will have that value at an isolated node.
In a three-parameter version, token should be an operator,
and left and right should be existing ExpressionTree instances
that become the operands for the binary operator.
"""
super().__init__() # LinkedBinaryTree initialization
if not isinstance(token, str):
raise TypeError('Token must be a string')
self._add_root(token) # use inherited, nonpublic method
if left is not None: # presumably three-parameter form
if token not in '+-*x/':
raise ValueError('token must be valid operator')
self._attach(self.root(), left, right) # use inherited, nonpublic method
def __str__(self):
"""Return string representation of the expression."""
pieces = [] # sequence of piecewise strings to compose
self._parenthesize_recur(self.root(), pieces)
return ''.join(pieces)
def _parenthesize_recur(self, p, result):
"""Append piecewise representation of p's subtree to resulting list."""
if self.is_leaf(p):
result.append(str(p.element())) # leaf value as a string
else:
result.append('(') # opening parenthesis
self._parenthesize_recur(self.left(p), result) # left subtree
result.append(p.element()) # operator
self._parenthesize_recur(self.right(p), result) # right subtree
result.append(')') # closing parenthesis
def evaluate(self):
"""Return the numeric result of the expression."""
return self._evaluate_recur(self.root())
def _evaluate_recur(self, p):
"""Return the numeric result of subtree rooted at p."""
if self.is_leaf(p):
return float(p.element()) # we assume element is numeric
else:
op = p.element()
left_val = self._evaluate_recur(self.left(p))
right_val = self._evaluate_recur(self.right(p))
if op == '+':
return left_val + right_val
elif op == '-':
return left_val - right_val
elif op == '/':
return left_val / right_val
else: # treat 'x' or '*' as multiplication
return left_val * right_val
def tokenize(raw):
"""Produces list of tokens indicated by a raw expression string.
For example the string '(43-(3*10))' results in the list
['(', '43', '-', '(', '3', '*', '10', ')', ')']
"""
SYMBOLS = set('+-x*/() ') # allow for '*' or 'x' for multiplication
mark = 0
tokens = []
n = len(raw)
for j in range(n):
if raw[j] in SYMBOLS:
if mark != j:
tokens.append(raw[mark:j]) # complete preceding token
if raw[j] != ' ':
tokens.append(raw[j]) # include this token
mark = j+1 # update mark to being at next index
if mark != n:
tokens.append(raw[mark:n]) # complete preceding token
return tokens
def build_expression_tree(tokens):
"""Returns an ExpressionTree based upon by a tokenized expression.
tokens must be an iterable of strings representing a fully parenthesized
binary expression, such as ['(', '43', '-', '(', '3', '*', '10', ')', ')']
"""
S = [] # we use Python list as stack
for t in tokens:
if t in '+-x*/': # t is an operator symbol
S.append(t) # push the operator symbol
elif t not in '()': # consider t to be a literal
S.append(ExpressionTree(t)) # push trivial tree storing value
elif t == ')': # compose a new tree from three constituent parts
right = S.pop() # right subtree as per LIFO
op = S.pop() # operator symbol
left = S.pop() # left subtree
S.append(ExpressionTree(op, left, right)) # repush tree
# we ignore a left parenthesis
return S.pop()
# -
unit = build_expression_tree(tokenize('(1+2)'))
print(unit, '=', unit.evaluate())
small = build_expression_tree(tokenize('((2x(5-1))+(3x10))'))
print(small, '=', small.evaluate())
big = build_expression_tree(tokenize('((((3+1)x3)/((9-5)+2))-((3x(7-4))+6))'))
print(big, '=', big.evaluate())
for item in small.preorder():
print(item.element())
for item in small.postorder():
print(item.element())
for item in small.inorder():
print(item.element())
| Lecture Codes/Lecture 4 notebooks/BinaryTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Did above-average rains contribute to the disaster in Brumadinho?</h1>
# <h3>Analysis of the possible impact of rainfalls on the rupture of Brumadinho's dam</h3>
#
# <br>
# <br>
#
# On January 25, 2019, <NAME> Vale’s Córrego do Feijão mine in Brumadinho collapsed. At the time it burst, it was holding 11.7m cubic metres of iron ore tailings, or almost 5,000 Olympic-sized swimming pools, and stood 86m high<sup><a href="#link1">[1]</a></sup>. The rupture gave rise to one of the greatest human and environmental disasters in the history of Brazil. One year later, 259 people had been found dead and 11 were still missing<sup><a href="#link2">[2]</a></sup>.
#
# In its 2017 _"Mine Tailing Storage: Safety is no Accident"_ report<sup><a href="#link3">[3]</a></sup>, the United Nations states that heavy and prolonged rains, hurricanes and earthquakes can even be triggers for disruptions and overflows, but even in these cases the UN considers that there was an error since the risk planning for the construction and maintenance of the dam must take into account the climatic conditions of the site. About the report, <NAME>, professor of geology at the Federal University of Espírito Santo and member of the UN committee on ore dams, summarizes: "The conclusion of the study is that there are two reasons that cause disruptions: error in the risk analysis and negligence in the maintenance of the dam. That is, if there was a torrential rain that caused the dam to overflow, there was an error in the risk analysis. If the region is subject to rain like this, the structure of the dam should be different. It has to be accurate."<sup><a href="#link4">[4]</a></sup>
#
# According to an independent report<sup><a href="#link5">[5]</a></sup> commissioned by the company, released in December 2019, one of the factors identified as a cause of the tragedy was an increase in the amount of rain in the region. “The Panel concluded that the sudden strength loss and resulting failure of the marginally stable dam were due to a critical combination of ongoing internal strains due to creep, and a strength reduction due to loss of suction in the unsaturated zone caused by the intense rainfall towards the end of 2018. This followed a number of years of increasing rainfall after tailings deposition ceased in July 2016.”, the report said.
#
# Anyway, before analyzing the participation of the precipitation factor in the disaster, we need to ask ourselves: were there, in fact, above average rains that could have impacted the rupture of the Brumadinho dam? That is the question that we will try to answer in this article in Jupyter Notebook format.
#
# ## Analyzed Data
#
# To conduct our study, publicly available data sets in the National Water Resources Information System (SNIRH) of the National Water Agency (ANA) and in CEMADEN (National Center for Natural Disaster Monitoring and Alerts), agencies of the Brazilian government, were used to collect information on precipitation in the vicinity of the dam and to analyze if there was any abnormal pattern of rain that could have contributed to the disaster.
#
# * <a href="http://www.snirh.gov.br/hidroweb/rest/api/documento/convencionais?tipo=3&documentos=2044008" target="_blank">Melo Franco station data from ANA's SNIRH<sup>[6]</sup></a>
# * <a href="http://www.cemaden.gov.br/mapainterativo/download/downpluv.php" target="_blank">CEMADEN stations data<sup>[7]</sup></a>
#
# The surrounding meteorological stations were reduced to the 6 closest to the dam, varying from 5.47 to 1.34 miles in distance, so that the data of the nearest station that were available in each period were always used (in the case of CEMADEN stations). In the following map it is possible to see the geographical position of ANA's Melo Franco station (blue dot), CEMADEN's Aranha, Alberto Flores, Centro, Córrego do Feijão and Casa Branca District stations (green dots) and the dam site (x in red).
# +
import matplotlib.pyplot as plt
import mplleaflet
plt.figure(figsize=(6, 6))
# ANA's station
plt.plot(-44.120881, -20.197752, 'bo', ms=10) # Melo Franco (02044008) [8,81 km (5.47 mi)]
# CEMADEN's stations
# plt.plot(-44.2, -20.143, 'go', ms=10) # Centro (310900601A) [8,98 km (5.58 mi)]
# plt.plot(-44.216, -20.146, 'go', ms=10) # Progresso (310900602A) [10,67 km (6.63 mi)]
plt.plot(-44.047, -20.094, 'go', ms=10) # Casa Branca district (310900603A) [7,91 km (4.92 mi)]
# plt.plot(-44.023, -20.165, 'go', ms=10) # Prefeito Maciel street (310900604A) [11,18 km (6.95 mi)]
plt.plot(-44.107, -20.135, 'go', ms=10) # Córrego do Feijão (310900605A) [2,16 km (1.34 mi)]
# plt.plot(-44.2009, -20.1409, 'go', ms=10) # Rio Paraopeba (310900605H) [9,00 km (5.59 mi)]
plt.plot(-44.147, -20.156, 'go', ms=10) # <NAME> (310900606A) [5,14 km (3.19 mi)]
plt.plot(-44.198, -20.142, 'go', ms=10) # Centro (310900607A) [8,74 km (5.43 mi)]
plt.plot(-44.105, -20.196, 'go', ms=10) # Aranha (310900608A) [8,72 km (5.42 mi)]
# plt.plot(-44.227, -20.12, 'go', ms=10) # Inhotim (310900609A) [11,38 km (7.07 mi)]
# Dam I
plt.plot(-44.118047, -20.118579, 'rX', ms=10)
mplleaflet.display()
# -
# Since data come from different sources, the formats and dispositions of the files are quite different and will need specific handling. The data for each ANA station is consolidated into a single CSV file, which has some headers and captions that will need to be ignored. CEMADEN data, on the other hand, are divided by city and month, that is, although a file contains data from all stations in the city, several files are needed to analyze a period of several years.
#
# Below is a preview of two of these files:
# +
from IPython.display import display, Pretty
# ANA's Melo Franco station data (first 50 file lines)
display(Pretty(filename ='data/chuvas_C_02044008[sample].csv'))
# -
# CEMADEN's stations data for January 2014 in Brumadinho (first 50 file lines)
display(Pretty(filename ='data/3277_MG_2014_1[sample].csv'))
# ## Data Wrangling
#
# Our final objective is to consolidate, in a single DataFrame by dataset, the accumulated rainfall per month, so that we can create visualizations that allow us to make our analysis. ANA's data already have a monthly consolidation and are all arranged in a single file, so let's start with it.
#
# A brief inspection shows us some details that we will need to take into account when telling Pandas how to load this file:
#
# * the separator is ';' and not ',' as the default for CSV files
# * the first 12 lines are a kind of legend that we will need to ignore
# * the numbers are using Brazilian standard, with a comma separating the decimals, which is also not Pandas standard
# * lastly, we could already indicate which column to use as an index for the rows, but as we will need to do some treatments on it too, for now we will load the file with the standard index
#
# That said, we can load our data like this:
# +
import pandas as pd
# For the sake of performance, Pandas by default truncates the total rows and columns to be displayed.
# The following lines modify this setting and instruct the library to display more or less data.
pd.set_option('display.max_rows', 20)
pd.set_option('display.max_columns', None)
"""
Here we use _ as the name of our DataFrame being processed. Although _ is a valid Python variable name,
it is generally not good practice in software engineering to name a variable with such a generic name. However,
as we will see later, many operations require frequent repetition of the DataFrame name, which can make the lines
very long, so we’ll use this trick to keep the lines of code as short as possible until we’ve finished cleaning
and processing our data.
"""
_ = pd.read_csv('data/chuvas_C_02044008.csv', decimal=',', index_col=False, sep=';', skiprows=12)
_
# -
# What really interests us is in the 'Data' and 'Total' columns. They tell us how much it rained each month in the region of Melo Franco station. So let's cut out this data to get a more detailed view. First, however, let's transform the data in the 'Data' column so that Pandas can work with this information in a time format.
_['Data'] = pd.to_datetime(_['Data'], dayfirst=True)
_[['Data','Total']]
# In a perfect world the data would almost ready, but for a better quality in our results, we have to pay attention to two other points:
#
# 1. From the beginning of 1941 to the end of 2019 there is a total of 948 months, but we have more than 1700 lines in our DataFrame, which means we have repeated dates.
# 2. In the file legend, in addition to 'NivelConsistencia' and 'TipoMedicaoChuvas', there is also the definition of 'Status' and not all values are useful to us.
#
# We will start with the treatment of repeated dates. In fact, if we analyze it carefully, the dates are repeated but with different levels of consistency (indicated in the column 'NivelConsistencia'). A quick look at ANA website reveals what this means:
# > "The analysis of consistency of rainfall data should aim to identify and correct errors, as well as to fill in faults in the rainfall series."<sup><a href="#link8">[8]</a></sup>
#
# Therefore, "consisted data" are those that have already undergone this analysis and correction of possible failures. So, whenever possible, we will give priority to data that is already consisted ('NivelConsistencia' = 2), we will do this by creating a Boolean mask that when applied will filter the lines and return only the version with the highest level of consistency for each date. We will also create another mask to remove data after January 2019.
#
# Finally, let's take the opportunity to sort the data in ascending order by date and transform our 'Data' column into the index of our DataFrame.
# +
mask1 = _.groupby(['Data'])['NivelConsistencia'].transform(max) == _['NivelConsistencia']
mask2 = (_['Data'].dt.year < 2019) | ((_['Data'].dt.year == 2019) & (_['Data'].dt.month == 1))
_ = _[mask1 & mask2].sort_values('Data')
_.set_index('Data', inplace=True)
_
# -
# We solved the first point, the problem of status remains. First, let's see how much of each status we have for the data that is meaningful to us:
_['TotalStatus'].value_counts()
# The first good news is that we do not have status 4 ('Accumulated') for 'Total', which makes sense since this is already a cumulative measure. Another is the fact that only 42 of the 930 entries are not in the status we want, which represents less than 5% of the sample. Since we want to generate a series of averages, it makes sense to give up the data with great uncertainty (status 2 and 3) to prevent them from contaminating our sample and consequently distorting our results.
#
# However, three entries are with status 0, which means an inconsistency. Let's see:
_[_['TotalStatus'] == 0]
# For some reason, the 'Total' column was not correctly measured but all daily measurements are present and, better, they all have status 1, indicating that they are real measurements. These inconsistencies would probably be corrected when the data were consisted (note that 'NivelConsistencia' is 1 for these lines), but luckily we have enough data to do this ourselves.
# +
# Updating 'Total' e 'TotalStatus'
# Note that if any of the days had a status other than 1, our 'TotalStatus' would also be different
_.at['2011-10-01', 'Total'] = _.loc['2011-10-01'].filter(regex=("Chuva[0-9]{2}$")).sum().round(1)
_.at['2011-10-01', 'TotalStatus'] = _.loc['2011-10-01'].filter(regex=("Chuva[0-9]{2}Status$")).max()
_.at['2011-12-01', 'Total'] = _.loc['2011-12-01'].filter(regex=("Chuva[0-9]{2}$")).sum().round(1)
_.at['2011-12-01', 'TotalStatus'] = _.loc['2011-12-01'].filter(regex=("Chuva[0-9]{2}Status$")).max()
_.at['2014-10-01', 'Total'] = _.loc['2014-10-01'].filter(regex=("Chuva[0-9]{2}$")).sum().round(1)
_.at['2014-10-01', 'TotalStatus'] = _.loc['2014-10-01'].filter(regex=("Chuva[0-9]{2}Status$")).max()
_.loc[[pd.to_datetime('2011-10-01'), pd.to_datetime('2011-12-01'), pd.to_datetime('2014-10-01')]]
# -
# Great! Now we can get rid of the inaccurate measures and reduce the DataFrame to a Series that contains only the column that is really meaningful to us, finishing the treatment for this dataset.
df_ana = _[_['TotalStatus'] == 1]['Total'].rename('ANA')
df_ana.index.name = None
df_ana
# Now we are going to clean and process data from CEMADEN stations. As previously stated, this data is arranged in several files, so we will have to use a new library to go through all these files and consolidate the data in a single DataFrame. Right after loading, in addition to the treatment for the separator, decimal and index, we will already define which columns we want to load. The ideal is to restrict the data to just what we need whenever possible, as this decreases the amount of memory and cpu required to handle the data.
#
# In the previous case, as we only had one file, it was easier to check the format in which the information was recorded, but now we are dealing with a dataset divided into more than 60 files and by inspecting only one of them is not possible to have this guarantee. Therefore, by concatenating the data from each file, we will preserve the original index and take advantage of that to print the first line from each one, checking if everything is as expected.
# +
from glob import glob
pd.set_option('display.max_rows', 70)
df1 = pd.concat([pd.read_csv(f,
sep=';',
index_col=False,
decimal=',',
usecols=['codEstacao','datahora','valorMedida']) for f in glob('data/3277_MG_*.csv')])
df1.loc[0]
# -
# It was good that we were careful because, from what we see, the way of recording the data has changed over time. In fact, if we look line by line, we can identify three distinct patterns:
#
# 1. Until March 2015, both dates and values were using the Brazilian standard (day/month/year and decimals separated by commas)
# 2. In April, May, June and July 2015, the dates are in international format and the values are with decimals separated by periods
# 3. As of August 2015, the dates remain using the international standard but the values are recorded again using the Brazilian standard
#
# Knowing this, we can rename the files so that the names distinguish the pattern used in each one, this way we'll be able to load them independently to make the necessary treatment.
# +
df_br_date = pd.concat([pd.read_csv(f,
sep=';',
index_col=False,
decimal=',',
usecols=['codEstacao','datahora','valorMedida']) for f in glob('data/renamed/_3277_MG_*.csv')],
ignore_index=True)
df_dot_decimal = pd.concat([pd.read_csv(f,
sep=';',
index_col=False,
usecols=['codEstacao','datahora','valorMedida']) for f in glob('data/renamed/dot_3277_MG_*.csv')],
ignore_index=True)
df_us_date = pd.concat([pd.read_csv(f,
sep=';',
index_col=False,
decimal=',',
usecols=['codEstacao','datahora','valorMedida']) for f in glob('data/renamed/3277_MG_*.csv')],
ignore_index=True)
df_br_date['datahora'] = pd.to_datetime(df_br_date['datahora'], dayfirst=True)
df_dot_decimal['datahora'] = pd.to_datetime(df_dot_decimal['datahora'])
df_us_date['datahora'] = pd.to_datetime(df_us_date['datahora'])
print(df_br_date.head())
print(df_dot_decimal.head())
print(df_us_date.head())
# -
# Great, now that we have all of our data in the same format we can combine them again into a single DataFrame.
df2 = pd.concat([df_br_date, df_dot_decimal, df_us_date], ignore_index=True)
df2.shape
# The amount of data we have here is still huge, it's almost one and a half million cells, which still slows down our operations. We can restrict the data a little more by filtering the distant stations. As we said, in order to facilitate the analysis and not need to resort to more complex methods like Thiessen Polygons to make a weighted average between all stations, we decided to always use the data from the geographically closest station to the dam. CEMADEN stations started to operate in 2014, providing 10 new data sources, and following our initial idea we could simply use the data from the nearest station (Córrego do Feijão). Then, let's see if we can follow this approach:
df3 = df2[df2['codEstacao'] == '310900605A'].copy()
df3.shape
# Okay, analyzing only the Córrego do Feijão station we reduced the amount of data by more than 90%, but they are still arranged by time and we need them consolidated by month. The next step is to standardize the times so that we can make an aggregation. At the moment this would not be possible because, as we can see in the previous blocks, the minutes and seconds are not always the same for all entries. CEMADEN website explains the different times recorded on our DataFrame:
# > "... automatic rain gauges connect to CEMADEN's servers and transmit data of accumulated rainfall, in millimeters, every 10 minutes, and are then processed and made available to the CEMADEN Situation Room on a specialized platform developed by the Center. On the other hand, if it is not raining, the rain gauge will connect only once every hour, sending the accumulated 0 millimeter of the last 60 minutes."<sup><a href="#link9">[9]</a></sup>
#
# So we must assume that we need to have at least one measurement per hour. In a month of 28 days we have 672 hours, in a month of 29 we have 696, 30 days have 720 hours and, finally, a month of 31 days has 744 hours. Knowing this, let's check the consistency of the data we have:
def hours_in_month(month, year):
if month in (1,3,5,7,8,10,12):
return 744
elif month in (4,6,9,11):
return 720
elif year == 2016:
return 696
else:
return 672
_ = df3[['valorMedida']].groupby(df3['datahora'].dt.floor('H')).sum().groupby(pd.Grouper(freq='MS')).count()
_['%'] = _.index.map(lambda dt: (_.loc[dt]['valorMedida']/hours_in_month(dt.month, dt.year)*100).round(2))
_ = _.rename(columns={'valorMedida': 'Hours with measurements'})
_
# From what we see, Córrego do Feijão station stopped recording or transmitting information in several moments. This demonstrates that only this station's data is not sufficient to get the whole sample we need. We will then use an approach similar to the one we did previously with 'NivelConsistencia' column in ANA dataset and fill in the gaps with data from the other CEMADEN stations in order of proximity, until we have enough data.
# +
df4 = df2[df2['codEstacao'].isin(['310900603A', '310900605A', '310900606A', '310900607A', '310900608A'])].copy()
df4['datahora'] = df4['datahora'].dt.floor('H')
df_corrego_feijao = df4[df4['codEstacao'] == '310900605A'][['datahora','valorMedida']]
df_alberto_flores = df4[df4['codEstacao'] == '310900606A'][['datahora','valorMedida']]
df_casa_branca = df4[df4['codEstacao'] == '310900603A'][['datahora','valorMedida']]
df_aranha = df4[df4['codEstacao'] == '310900608A'][['datahora','valorMedida']]
df_centro = df4[df4['codEstacao'] == '310900607A'][['datahora','valorMedida']]
df_corrego_feijao = df_corrego_feijao.groupby('datahora').sum()
df_alberto_flores = df_alberto_flores.groupby('datahora').sum()
df_casa_branca = df_casa_branca.groupby('datahora').sum()
df_aranha = df_aranha.groupby('datahora').sum()
df_centro = df_centro.groupby('datahora').sum()
df_cemaden = (df_corrego_feijao.combine_first(df_alberto_flores)
.combine_first(df_casa_branca)
.combine_first(df_aranha)
.combine_first(df_centro))
# -
_ = pd.DataFrame(df_cemaden.groupby(pd.Grouper(freq='MS')).count()).rename(columns={'valorMedida': 'Hours with measurements'})
_['%'] = _.index.map(lambda dt: (_.loc[dt]['Hours with measurements']/hours_in_month(dt.month, dt.year)*100).round(2))
_
# Now we have a relevant sample. We still have some months with missing data, but these gaps do not even represent 2% of the total hours in the month.
#
# So we just need to consolidate this data in a Series format, just as we did with the ANA dataset, and we will be ready to generate the visualizations that will help us to answer our initial question.
df_cemaden = df_cemaden.groupby(pd.Grouper(freq='MS')).sum().round(1)['valorMedida'].rename('CEMADEN')
df_cemaden.index.name = None
df_cemaden
# ## Visualizations
#
# After wrangling the data, we can now create graphical visualizations that not only help us to understand the data but also allow us to identify patterns, anomalies and also answer several questions immediately.
# +
import locale
locale.setlocale(locale.LC_TIME, 'pt_BR')
import calendar
import matplotlib.ticker as ticker
import matplotlib.lines as mlines
import numpy as np
# %matplotlib notebook
# -
# The first visualization that we are going to create is one that shows the amount of rain accumulated in the fourth semester of the year in Brumadinho in three sections: historical maximum, historical average and the measurements of 2018, for each of the datasets.
def precipitation_fourth_quarter(ana, cemaden):
_ = ana[:-1].groupby(pd.Grouper(freq='QS')).sum()
__ = cemaden[:-1].groupby(pd.Grouper(freq='QS')).sum()
_ = _.groupby(_.index.month)
__ = __.groupby(__.index.month)
maxs_ana = _.max()
maxs_cemaden = __.max()
maxs_ana = maxs_ana.loc[10].sum()
maxs_cemaden = maxs_cemaden.loc[10].sum()
means_ana = _.mean()
means_cemaden = __.mean()
means_ana = means_ana.loc[10].sum()
means_cemaden = means_cemaden.loc[10].sum()
g_labels = ['ANA (1941 to 2018)', 'CEMADEN (2014 to 2018)']
g_maxs = [maxs_ana, maxs_cemaden]
g_means = [means_ana, means_cemaden]
g_72p = [means_ana*1.72, means_cemaden*1.72]
g_2018 = [ana.iloc[-4:-1].sum(), cemaden.iloc[-4:-1].sum()]
x = np.arange(2)
width = 0.20
fig, ax = plt.subplots()
fig.set_size_inches(6.4, 4.8)
rects1 = ax.bar(x - width, g_maxs, width, label='All-time high', color='tab:blue')
rects2 = ax.bar(x, g_means, width, label='All-time average', color='tab:cyan')
rects3 = ax.bar(x + width, g_2018, width, label='2018 Q4', color='tab:orange')
for i, rects in enumerate([rects1, rects2, rects3]):
for rect in rects:
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width()/2., height + 15,
'%d mm' % int(height), ha='center', va='bottom')
ax.set_title('Accumulated precipitation for the 4th quarter in Brumadinho', pad=17)
ax.set_xticks(x)
ax.set_xticklabels(g_labels)
ax.set_yticklabels([])
plt.tick_params(axis='both', length=0)
ax.legend(loc='upper center', bbox_to_anchor=(0.5, -0.07), frameon=False, ncol=3)
fig.tight_layout()
plt.ylim(0, 1300)
for s in ax.spines:
ax.spines[s].set_visible(False)
plt.show()
# Next, we are going to plot the accumulated annual precipitation in Brumadinho since the beginning of ANA’s station measurements.
def precipitation_by_year(df):
plt.figure(figsize=(6.4, 4.8))
df_by_year = df.groupby(pd.Grouper(freq='Y')).sum()[1:-1]
precip_plot = df_by_year.plot()
blue_line = mlines.Line2D([], [], color='blue')
means_plot, = plt.plot([-29, 48], [df_by_year.mean(), df_by_year.mean()], color='tab:orange')
plt.tick_params(axis='x', length=0)
plt.title('Annual precipitation in Brumadinho [{}]'.format(df.name), pad=17)
plt.ylim(0, 2000)
ax = plt.gca()
span_area = ax.axvspan(46, 48, alpha=0.3, color='red')
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('%d mm'))
plt.legend(handles=[blue_line, means_plot, span_area],
labels=['Annual Precipitation', 'Average', '2016 to 2018 span'],
loc='upper center',
fontsize='small',
ncol=3,
frameon=False,
bbox_to_anchor=(0.5, -0.07))
for s in ['bottom','right','top']:
ax.spines[s].set_visible(False)
plt.show()
# The last visualization will show details of Brumadinho's rainfall profile. We will display in a single graph the historical rainfall averages per month (and the standard error), the historical maximums per month and the actual measurements from July 2018 to January 2019. The idea is to compare the amount of rain of the months prior to the tragedy with the standard behavior of rainfall and also the most extreme behavior ever recorded in the region.
def precipitation_profile(df):
plt.figure(figsize=(6.4, 4.8))
_ = df.iloc[:-7].groupby(df.iloc[:-7].index.month)
maxs = _.max().rename(index=lambda x: calendar.month_abbr[x])
means = _.mean().rename(index=lambda x: calendar.month_abbr[x])
std_errors = (_.std()/np.sqrt(_.count())).rename(index=lambda x: calendar.month_abbr[x])
shift_mask = calendar.month_abbr[7:14] + calendar.month_abbr[1:7]
maxs = maxs.loc[shift_mask]
means = means.loc[shift_mask]
std_errors = std_errors.loc[shift_mask]
x_axis = np.arange(12)
means_plot = plt.bar(x_axis, means, yerr=std_errors*2, ecolor='tab:gray', color='tab:cyan')
maxs_plot, = plt.plot(x_axis, maxs, '.:', color='tab:blue')
measured_plot, = plt.plot(np.arange(7), df.iloc[-7:], 'o-', color='tab:orange')
plt.xticks(x_axis, shift_mask)
plt.tick_params(axis='x', length=0)
plt.title('Brumadinho\'s precipitation profile [{}]'.format(df.name), pad=17)
plt.ylim(0, 700)
ax = plt.gca()
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('%d mm'))
plt.legend(handles=[maxs_plot, means_plot, measured_plot],
labels=['All-time highs', 'All-time averages', 'Jul/18 to Jan/19'],
loc='upper center',
fontsize='small',
ncol=3,
frameon=False,
bbox_to_anchor=(0.5, -0.07))
for s in ['bottom','right','top']:
ax.spines[s].set_visible(False)
plt.show()
# ## Analysis
#
# With the help of visualizations, we can return to the questions we set out to answer:
#
# **- Was there more rain than usual in late 2018?**
#
# The first thing I’ll try to answer is if rainfall at the end of 2018 was, indeed, intense as Vale's report said. Although the definition of “intense” is somewhat subjective in this case, I set out to analyze the issue from the perspective of the entire historical series that we had available, and the chart below has the results.
precipitation_fourth_quarter(df_ana, df_cemaden)
# CEMADEN data, only available after 2014, form a small sample to generate a relevant statistical value, but because it is an agency created specifically for monitoring natural disasters, we decided to check if the independent panel had used these numbers for make its statement. But the fact is the rains of the last quarter of 2018, recorded by CEMADEN, were only 7.8% above the average. For ANA data, 2018 records were even lower than average. Therefore, we have sufficient evidence to affirm that the information conveyed in the report is hardly true.
#
# **- Has rainfall increased in the years that followed 2016?**
#
# Next, I want to validate the statement that the collapse “followed a number of years of increasing rainfall after tailings deposition ceased in July 2016” and for that I’m going to use the chart with the accumulated annual precipitation in Brumadinho since the beginning of ANA’s station measurements.
precipitation_by_year(df_ana)
# Again, it is not possible to notice any significant increase in the volume of annual precipitation in the region. Being quite strict, the volume was actually lower in the year following the end of tailings deposition.
#
# **- Were there unexpected rain patterns that could have impacted the rupture of Brumadinho’s dam?**
#
# Finally, in order to understand if any meteorological anomaly hit the region in the months that preceded the disaster, I’m going to use the charts with precipitation profile of the region of the dam.
precipitation_profile(df_ana)
precipitation_profile(df_cemaden)
# _*It is worth noting that the standard error for CEMADEN data is much larger than that of ANA data, which was to be expected, since the sample is much smaller._
#
# The first analysis we should do is to compare the records from July 2018 to January 2019 and the historical averages. The charts show that, of the 7 months prior to the disaster, only August and September showed rains consistently above average, but even so at levels that do not even reach half the historical average of the wettest month, for example. October and November were within the average, fluctuating to more or less depending on the dataset but still within the margins of error. However, December and mainly January, typically the wettest months, had considerably below average rainfall at the turn of 2019.
#
# ## Conclusion
#
# Throughout this article, we used rainfall data to analyze the possible impact of the amount of rain in one of the greatest natural disasters in world’s history, and we could see that:
#
# - The measured rainfall for the end of 2018 was close to the average
# - There’s no evidence of rainfall increasing after 2016
# - The wettest months for the region even had bellow average precipitation levels just before the disaster
#
# That said, if we are going to make an analysis purely from the perspective of the search for the cause of the tragedy, we must emphasize that **the amount of rain should not be considered a causative factor for the disaster** since there were no anomalies and, since the beginning, the most conservative security parameters should have been adopted when dealing with a structure with such potential for destruction.
# ## External links
#
# 1. <a name="link1"></a>"Heavy rain, design and poor drainage factors in Vale dam breach", Financial Times. https://www.ft.com/content/e805b142-1cde-11ea-97df-cc63de1d73f4
# 2. <a name="link2"></a>"‘Vale ended our lives’: Broken Brumadinho a year after dam collapse", BBC. https://www.bbc.com/news/world-latin-america-51220373
# 3. <a name="link3"></a>"Mine Tailings Storage: Safety Is No Accident", United Nations. http://www.grida.no/publications/383
# 4. <a name="link4"></a>"Tragédia com barragem da Vale em Brumadinho pode ser a pior no mundo em 3 décadas", BBC Brasil. https://www.bbc.com/portuguese/brasil-47034499
# 5. <a name="link5"></a>"Report of the Expert Panel on the Technical Causes of the Failure of Feijão Dam I". https://bdrb1investigationstacc.z15.web.core.windows.net/assets/Feijao-Dam-I-Expert-Panel-Report-ENG.pdf
# 6. <a name="link6"></a>Melo Franco station data from ANA's SNIRH. http://www.snirh.gov.br/hidroweb/rest/api/documento/convencionais?tipo=3&documentos=2044008
# 7. <a name="link7"></a>CEMADEN stations data. http://www.cemaden.gov.br/mapainterativo/download/downpluv.php
# 8. <a name="link8"></a>"DIRETRIZES E ANÁLISES RECOMENDADAS PARA A CONSISTÊNCIA DE DADOS PLUVIOMÉTRICOS". http://arquivos.ana.gov.br/infohidrologicas/cadastro/DiretrizesEAnalisesRecomendadasParaConsistenciaDeDadosPluviometricos-VersaoJan12.pdf
# 9. <a name="link9"></a>"Automatic Pluviometers". https://www.cemaden.gov.br/pluviometros-automatico/
| Brumadinho Dam Collapse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sanity Check for Bandit Sequence Design
#
# In this notebook, we show the sanity check for bandit sequence design.
# The RBS sequence is 20-base, and we divide it into three parts: 7-base (Pre) + 6-base (Center) + 7-base (Post). For the central part, we use spectrum kernel (l = 3), and for the other parts, we use dot product with one-hot encoding. Then the kernel matrix of 20-based RBS is the sum of the three kernels, i.e. K(rbs_i, rbs_j) = k_dotpro (pre_i, pre_j) + k_spectrum (center_i, center_j) + k_dotpro (post_i, post_j). The reason for summing up the kernels is that the central part is the design part and the other parts are similar in terms of different data points.
# +
# direct to proper path
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from collections import defaultdict
from itertools import product
import math
from codes.embedding import Embedding
from codes.environment import Rewards_env
from codes.ucb import GPUCB, Random
from codes.evaluations import evaluate, plot_eva
from codes.regression import Regression
from codes.kernels import spectrum_kernel
from codes.kernels_pairwise import spectrum_kernel_pw, sum_onehot_spectrum_kernel_pw, mixed_spectrum_kernel_pw, WD_kernel_pw, WD_shift_kernel_pw
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import PairwiseKernel, DotProduct, RBF
from sklearn.kernel_ridge import KernelRidge
from ipywidgets import IntProgress
from IPython.display import display
import warnings
# %matplotlib inline
# -
# ## Reading Dataset
# +
# Data downloaded from https://github.com/synbiochem/opt-mva
# Paper https://pubs.acs.org/doi/abs/10.1021/acssynbio.8b00398
# A: whole RBS sequence (len: 29)
# B: extracted 20-base RBS seqeunce (A[7:27], len: 20), training features
# C: TIR labels
# D: the design part RBS (B[7:13], len: 6)
Path = '../data/RBS_seqs.csv'
df = pd.read_csv(Path)
df.columns = ['A', 'B', 'C']
df['D'] = df['B'].str[7:13]
df.head()
# +
# drop the exact same record (same B and C)
df = df.drop_duplicates(subset = ['B', 'C'])
df.shape
# +
Log_flag = False # indicates whether take log label
Norm_method = 'minmax' # indicates how to normalize label (one of 'mean', 'minmax', None)
def normalize(df):
# take log FC -- possiblely provide Gaussain distribution?
if Log_flag:
df['C'] = np.log(df['C'])
if Norm_method == 'mean':
# mean normalization
df['C'] = (df['C'] - df['C'].mean())/df['C'].std()
elif Norm_method == 'minmax':
# min-max normalization [-1,1]
df['C'] = (df['C'] - df['C'].min())/(df['C'].max() - df['C'].min())
else:
assert Norm_method == None
return df
# -
# data: num_data * 2, columns: [20-base RBS seq (B), TIR labels (C)]
data = np.asarray(normalize(df)[['B', 'C']])
data.shape
# data6: num_data * 2, columns: [6-base RBS seq (D), TIR labels (C)]
data6 = np.asarray(normalize(df)[['D', 'C']])
data6.shape
# ## Construct feature spaces
#
# We fix the part other the designing 6-base RBS, the 20-based RBS would be: 'TTTAAGA' + 6-base design + 'TATACAT'. The 6-base design has totally 4^6 = 4096 combinations.
# +
# create all combos
combos = [] # 20-base
combos_6 = [] # 6-base
labels = []
char_sets = ['A', 'G', 'C', 'T']
design_len = 6
# to be changed
pre_design = 'TTTAAGA'
pos_design = 'TATACAT'
for combo in product(char_sets, repeat= design_len):
combos_6.append(''.join(combo))
combo = pre_design + ''.join(combo) + pos_design
combos.append(combo)
labels.append(math.inf)
assert len(combos) == len(char_sets) ** design_len
# -
todesign_data = np.asarray(list(zip(combos, labels))) # 20-base
todesign6_data = np.asarray(list(zip(combos_6, labels))) # 6-base
# ## Setting
embedding = 'label'
# sum_spectrum_kernel_pw: spectrum kernels for [:7], [7:13], [13:] respectively
kernel = [spectrum_kernel, sum_onehot_spectrum_kernel_pw]
string_kernel_flag = True
# ### 1. Check influence of the parameters of kernels.
#
# Check whether changing the parameters of kernels by a small amount changes the recommendations dramatically.
# But since we use spectrum kernel for the main part, the only parameter is the l (the length of features) and we fixed it to 3. There is not much to test.
#
# ### 2. Check whether permutating the data change the recommendations.
# +
num_rounds = 1
num_exper = 1
num_rec = 60
my_env = Rewards_env(data, embedding)
new_env = Rewards_env(todesign_data, embedding)
if string_kernel_flag:
agent = GPUCB(new_env, num_rounds, init_list = my_env.rewards_dict, num_rec = num_rec,
model = GaussianProcessRegressor(kernel = PairwiseKernel(metric = kernel[-1])))
else:
agent = GPUCB(new_env, num_rounds, init_list = my_env.rewards_dict, num_rec = num_rec,
model = GaussianProcessRegressor(kernel = kernel[-1]))
rec_arms = agent.play(label_avaiable = False)
# -
from sklearn.utils import shuffle
shuffled_df = shuffle(df)
shuffled_df.reset_index(inplace=True, drop=True)
# +
# data: num_data * 2, columns: [20-base RBS seq (B), TIR labels (C)]
shuffled_data = np.asarray(normalize(shuffled_df)[['B', 'C']])
# data6: num_data * 2, columns: [6-base RBS seq (D), TIR labels (C)]
shuffled_data6 = np.asarray(normalize(shuffled_df)[['D', 'C']])
# +
num_rounds = 1
num_exper = 1
num_rec = 60
my_env = Rewards_env(shuffled_data, embedding)
new_env = Rewards_env(todesign_data, embedding)
if string_kernel_flag:
agent_with_shuffle_data = GPUCB(new_env, num_rounds, init_list = my_env.rewards_dict, num_rec = num_rec,
model = GaussianProcessRegressor(kernel = PairwiseKernel(metric = kernel[-1])))
else:
agent_with_shuffle_data = GPUCB(new_env, num_rounds, init_list = my_env.rewards_dict, num_rec = num_rec,
model = GaussianProcessRegressor(kernel = kernel[-1]))
rec_arms_with_shuffled_data = agent_with_shuffle_data.play(label_avaiable = False)
# -
set(rec_arms) == set(rec_arms_with_shuffled_data)
# #### Conclusion 2.
# From the above experiment, shuffling data does not change the recommendations.
# ### 3. Check the kernel matrix of the Bandit Top60 vs 1by1 sequences.
#
# Since the designed bandit sequences use about 1/3 same pre- and post- sequences as the 1by1 sequences, it's kind of cheating to test the similarities for the whole sequence. We provide the plot for both: 1) kernel matrix for 20-base bandit vs. 1by1; 2) kernel matrix for only 6-base central part for bandit vs. 1by1.
# #### 1) kernel matrix for 20-base bandit vs. 1by1
rbs1by1seq = pd.read_csv('../data/1by1_recs.csv')
rbs1by1 = np.asarray(rbs1by1seq['1by1'].str.upper())
rbs1by1_bandit_kernel = spectrum_kernel(todesign_data[np.asarray(rec_arms)][:,0], np.asarray(rbs1by1))
plt.imshow(rbs1by1_bandit_kernel, cmap = 'hot', interpolation='nearest')
#plt.imshow(sorted_new_kernel_matrix, cmap = 'hot')
plt.colorbar()
plt.title('kernel matrix for 20-base bandit vs. 1by1')
plt.xlabel('1by1')
plt.ylabel('Bandit')
# #### 2) kernel matrix for only 6-base central part for bandit vs. 1by1.
rbs1by1_6 = np.asarray(rbs1by1seq['1by1'].str[7:13].str.upper())
rbs1by1_6
rbs1by1_bandit_kernel = spectrum_kernel(todesign6_data[np.asarray(rec_arms)][:,0], np.asarray(rbs1by1_6))
plt.imshow(rbs1by1_bandit_kernel, cmap = 'hot', interpolation='nearest')
#plt.imshow(sorted_new_kernel_matrix, cmap = 'hot')
plt.colorbar()
plt.title('kernel matrix for 6-base bandit vs. 1by1')
plt.xlabel('1by1')
plt.ylabel('Bandit')
# #### Ps. Random recs vs. 1by1 seqs
# +
random_recs = pd.read_csv('../data/random_recs.csv')
random = np.asarray(random_recs['uniform'].append(random_recs['ppm']))
random_rbs1by1_kernel = spectrum_kernel(random, np.asarray(rbs1by1_6))
plt.imshow(random_rbs1by1_kernel, cmap = 'hot', interpolation='nearest')
#plt.imshow(sorted_new_kernel_matrix, cmap = 'hot')
plt.colorbar()
plt.title('kernel matrix for 6-base Random vs. 1by1')
plt.xlabel('1by1')
plt.ylabel('Random')
# -
# #### Conclusion 3.
#
# As expected, similarities between 20-base sequences are high. For the 6-base comparison, we show two plots: bandit (y-axis) vs. 1by1 (x-axis); random (y-axis) vs. 1by1 (x-axis). The 1-30 random sequences are uniformly random, and 31-60 are random based on PPM. We can see the similarity of PPM random vs. 1by1 is generally higher than uniform random vs. 1by1. And the similarity of bandit vs. 1by1 is bewteen the above two random designs, which makes sense (bandit is desinged to recommend seqeucnes tends to have higher predictions or with high uncertainty).
# ### 4. Clustering the Top60 recommendations (we expect there are a few clusters)
bandit_kernel = spectrum_kernel(todesign6_data[np.asarray(rec_arms)][:,0])
plt.imshow(bandit_kernel, cmap = 'hot', interpolation='nearest')
#plt.imshow(sorted_new_kernel_matrix, cmap = 'hot')
plt.colorbar()
plt.title('kernel matrix for 6-base bandit (Top 60 UCB)')
# #### Conclusion 4.
#
# Actually, before any clustering, the recommendation sequences are already shown in several blocks in the kernel heatmap plot. The recommendations are actually sorted in terms of the UCB (mu + sigma) in decreasing order. The first 10 arms are similar to each other within the small group, so are 10-20, 20-40, 40-60 groups.
# One thing to notice is that the 20-40 arms are also similar to 1-10.
# So there are roughly 4-5 clusters among the Top 60 arms. This is good. On the one hand, arms are not all similar to each other; on the other hand, arms are all not relevant to each other.
# ### 5. Check the intersection of recommendations based on mu, mu+sigma, sigma
#
# Ideally, there should be some interactions but not a lot. We use the [jaccard similarity](https://en.wikipedia.org/wiki/Jaccard_index) score and kernel matrix to show the interactions.
def jaccard_similarity(list1, list2):
intersection = len(list(set(list1).intersection(list2)))
union = (len(list1) + len(list2)) - intersection
return float(intersection) / union
recs_mu = np.asarray(sorted(range(len(agent.mu)), key=lambda k: (agent.mu)[k]))[-60:]
recs_sigma = np.asarray(sorted(range(len(agent.sigma)), key=lambda k: (agent.sigma)[k]))[-60:]
jaccard_similarity(recs_mu, rec_arms)
jaccard_similarity(recs_sigma, rec_arms)
bandit_kernel = spectrum_kernel(todesign6_data[np.asarray(rec_arms)][:,0], todesign6_data[np.asarray(recs_mu)][:,0])
plt.imshow(bandit_kernel, cmap = 'hot', interpolation='nearest')
#plt.imshow(sorted_new_kernel_matrix, cmap = 'hot')
plt.colorbar()
plt.title('kernel matrix for 6-base Top UCB vs. Top mu')
plt.ylabel('Top UCB')
plt.xlabel('Top mu')
bandit_kernel = spectrum_kernel(todesign6_data[np.asarray(rec_arms)][:,0], todesign6_data[np.asarray(recs_sigma)][:,0])
plt.imshow(bandit_kernel, cmap = 'hot', interpolation='nearest')
#plt.imshow(sorted_new_kernel_matrix, cmap = 'hot')
plt.colorbar()
plt.title('kernel matrix for 6-base Top UCB vs. Top sigma')
plt.ylabel('Top UCB')
plt.xlabel('Top sigma')
# #### Conclusion 5.
#
# Measuring the intersections might not be a good idea, since sorting the total more than 4,000 sequences by different metrics will highly possibly lead to very different sorting. Then the intersections will be small. But the intersection cannot take similar arms into consideration. That is, if different metrics return similar but not the same arms, then the effect of recommendation is similar as well, but the Jaccard score will be small.
#
# So we plot the kernel matrix for "Top UCB vs. Top mu", and "Top UCB vs. Top sigma". The plot is pretty interesting. The similarity trend for these two groups is on the contrary. 1-10, 20-40 arms for UCB are similar to top mu arms; 10-20, 40-60 arms for UCB are similar to top sigma arms. This trend can be explained by that the UCB recommendations combine mu and sigma, try to balance the exploration and exploitation.
#
# Note that most of the highly ranked UCB arms are similar to top mu groups. This shows the front part of Top 60 UCB recommendations prefers the high predictions mean (mu), which controls the exploitation; and the back part of Top 60 UCB arms are similar to the top sigma group, which leads the exploration.
| notebook/rec_design/Sanity_check/Sanity_Check_[0,1]label.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Multilabel classification on PASCAL using python data-layers
# In this tutorial we will do multilabel classification on PASCAL VOC 2012.
#
# Multilabel classification is a generalization of multiclass classification, where each instance (image) can belong to many classes. For example, an image may both belong to a "beach" category and a "vacation pictures" category. In multiclass classification, on the other hand, each image belongs to a single class.
#
# Caffe supports multilabel classification through the SigmoidCrossEntropyLoss layer, and we will load data using a Python data layer. Data could also be provided through HDF5 or LMDB data layers, but the python data layer provides endless flexibility, so that's what we will use.
# ### 1. Preliminaries
#
# * First, make sure you compile caffe using
# WITH_PYTHON_LAYER := 1
#
# * Second, download PASCAL VOC 2012. It's available here: http://host.robots.ox.ac.uk/pascal/VOC/voc2012/index.html
#
# * Third, import modules:
# +
import sys
import os
import numpy as np
import os.path as osp
import matplotlib.pyplot as plt
from copy import copy
% matplotlib inline
plt.rcParams['figure.figsize'] = (6, 6)
caffe_root = '../' # this file is expected to be in {caffe_root}/examples
sys.path.append(caffe_root + 'python')
import caffe # If you get "No module named _caffe", either you have not built pycaffe or you have the wrong path.
from caffe import layers as L, params as P # Shortcuts to define the net prototxt.
sys.path.append("pycaffe/layers") # the datalayers we will use are in this directory.
sys.path.append("pycaffe") # the tools file is in this folder
import tools #this contains some tools that we need
# -
# * Fourth, set data directories and initialize caffe
# +
# set data root directory, e.g:
pascal_root = osp.join(caffe_root, 'data/pascal/VOC2012')
# these are the PASCAL classes, we'll need them later.
classes = np.asarray(['aeroplane', 'bicycle', 'bird', 'boat', 'bottle', 'bus', 'car', 'cat', 'chair', 'cow', 'diningtable', 'dog', 'horse', 'motorbike', 'person', 'pottedplant', 'sheep', 'sofa', 'train', 'tvmonitor'])
# make sure we have the caffenet weight downloaded.
if not os.path.isfile(caffe_root + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'):
print("Downloading pre-trained CaffeNet model...")
# !../scripts/download_model_binary.py ../models/bvlc_reference_caffenet
# initialize caffe for gpu mode
caffe.set_mode_gpu()
caffe.set_device(0)
# -
# ### 2. Define network prototxts
#
# * Let's start by defining the nets using caffe.NetSpec. Note how we used the SigmoidCrossEntropyLoss layer. This is the right loss for multilabel classification. Also note how the data layer is defined.
# +
# helper function for common structures
def conv_relu(bottom, ks, nout, stride=1, pad=0, group=1):
conv = L.Convolution(bottom, kernel_size=ks, stride=stride,
num_output=nout, pad=pad, group=group)
return conv, L.ReLU(conv, in_place=True)
# another helper function
def fc_relu(bottom, nout):
fc = L.InnerProduct(bottom, num_output=nout)
return fc, L.ReLU(fc, in_place=True)
# yet another helper function
def max_pool(bottom, ks, stride=1):
return L.Pooling(bottom, pool=P.Pooling.MAX, kernel_size=ks, stride=stride)
# main netspec wrapper
def caffenet_multilabel(data_layer_params, datalayer):
# setup the python data layer
n = caffe.NetSpec()
n.data, n.label = L.Python(module = 'pascal_multilabel_datalayers', layer = datalayer,
ntop = 2, param_str=str(data_layer_params))
# the net itself
n.conv1, n.relu1 = conv_relu(n.data, 11, 96, stride=4)
n.pool1 = max_pool(n.relu1, 3, stride=2)
n.norm1 = L.LRN(n.pool1, local_size=5, alpha=1e-4, beta=0.75)
n.conv2, n.relu2 = conv_relu(n.norm1, 5, 256, pad=2, group=2)
n.pool2 = max_pool(n.relu2, 3, stride=2)
n.norm2 = L.LRN(n.pool2, local_size=5, alpha=1e-4, beta=0.75)
n.conv3, n.relu3 = conv_relu(n.norm2, 3, 384, pad=1)
n.conv4, n.relu4 = conv_relu(n.relu3, 3, 384, pad=1, group=2)
n.conv5, n.relu5 = conv_relu(n.relu4, 3, 256, pad=1, group=2)
n.pool5 = max_pool(n.relu5, 3, stride=2)
n.fc6, n.relu6 = fc_relu(n.pool5, 4096)
n.drop6 = L.Dropout(n.relu6, in_place=True)
n.fc7, n.relu7 = fc_relu(n.drop6, 4096)
n.drop7 = L.Dropout(n.relu7, in_place=True)
n.score = L.InnerProduct(n.drop7, num_output=20)
n.loss = L.SigmoidCrossEntropyLoss(n.score, n.label)
return str(n.to_proto())
# -
# ### 3. Write nets and solver files
#
# * Now we can crete net and solver prototxts. For the solver, we use the CaffeSolver class from the "tools" module
# +
workdir = './pascal_multilabel_with_datalayer'
if not os.path.isdir(workdir):
os.makedirs(workdir)
solverprototxt = tools.CaffeSolver(trainnet_prototxt_path = osp.join(workdir, "trainnet.prototxt"), testnet_prototxt_path = osp.join(workdir, "valnet.prototxt"))
solverprototxt.sp['display'] = "1"
solverprototxt.sp['base_lr'] = "0.0001"
solverprototxt.write(osp.join(workdir, 'solver.prototxt'))
# write train net.
with open(osp.join(workdir, 'trainnet.prototxt'), 'w') as f:
# provide parameters to the data layer as a python dictionary. Easy as pie!
data_layer_params = dict(batch_size = 128, im_shape = [227, 227], split = 'train', pascal_root = pascal_root)
f.write(caffenet_multilabel(data_layer_params, 'PascalMultilabelDataLayerSync'))
# write validation net.
with open(osp.join(workdir, 'valnet.prototxt'), 'w') as f:
data_layer_params = dict(batch_size = 128, im_shape = [227, 227], split = 'val', pascal_root = pascal_root)
f.write(caffenet_multilabel(data_layer_params, 'PascalMultilabelDataLayerSync'))
# -
# * This net uses a python datalayer: 'PascalMultilabelDataLayerSync', which is defined in './pycaffe/layers/pascal_multilabel_datalayers.py'.
#
# * Take a look at the code. It's quite straight-forward, and gives you full control over data and labels.
#
# * Now we can load the caffe solver as usual.
solver = caffe.SGDSolver(osp.join(workdir, 'solver.prototxt'))
solver.net.copy_from(caffe_root + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel')
solver.test_nets[0].share_with(solver.net)
solver.step(1)
# * Let's check the data we have loaded.
transformer = tools.SimpleTransformer() # This is simply to add back the bias, re-shuffle the color channels to RGB, and so on...
image_index = 0 # First image in the batch.
plt.figure()
plt.imshow(transformer.deprocess(copy(solver.net.blobs['data'].data[image_index, ...])))
gtlist = solver.net.blobs['label'].data[image_index, ...].astype(np.int)
plt.title('GT: {}'.format(classes[np.where(gtlist)]))
plt.axis('off');
# * NOTE: we are readin the image from the data layer, so the resolution is lower than the original PASCAL image.
# ### 4. Train a net.
#
# * Let's train the net. First, though, we need some way to measure the accuracy. Hamming distance is commonly used in multilabel problems. We also need a simple test loop. Let's write that down.
# +
def hamming_distance(gt, est):
return sum([1 for (g, e) in zip(gt, est) if g == e]) / float(len(gt))
def check_accuracy(net, num_batches, batch_size = 128):
acc = 0.0
for t in range(num_batches):
net.forward()
gts = net.blobs['label'].data
ests = net.blobs['score'].data > 0
for gt, est in zip(gts, ests): #for each ground truth and estimated label vector
acc += hamming_distance(gt, est)
return acc / (num_batches * batch_size)
# -
# * Alright, now let's train for a while
for itt in range(6):
solver.step(100)
print 'itt:{:3d}'.format((itt + 1) * 100), 'accuracy:{0:.4f}'.format(check_accuracy(solver.test_nets[0], 50))
# * Great, the accuracy is increasing, and it seems to converge rather quickly. It may seem strange that it starts off so high but it is because the ground truth is sparse. There are 20 classes in PASCAL, and usually only one or two is present. So predicting all zeros yields rather high accuracy. Let's check to make sure.
# +
def check_baseline_accuracy(net, num_batches, batch_size = 128):
acc = 0.0
for t in range(num_batches):
net.forward()
gts = net.blobs['label'].data
ests = np.zeros((batch_size, len(gts)))
for gt, est in zip(gts, ests): #for each ground truth and estimated label vector
acc += hamming_distance(gt, est)
return acc / (num_batches * batch_size)
print 'Baseline accuracy:{0:.4f}'.format(check_baseline_accuracy(solver.test_nets[0], 5823/128))
# -
# ### 6. Look at some prediction results
test_net = solver.test_nets[0]
for image_index in range(5):
plt.figure()
plt.imshow(transformer.deprocess(copy(test_net.blobs['data'].data[image_index, ...])))
gtlist = test_net.blobs['label'].data[image_index, ...].astype(np.int)
estlist = test_net.blobs['score'].data[image_index, ...] > 0
plt.title('GT: {} \n EST: {}'.format(classes[np.where(gtlist)], classes[np.where(estlist)]))
plt.axis('off')
| examples/pascal-multilabel-with-datalayer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="sm0KVbgx7hmv"
# Inheritance
#
# When you use inheritance to create your new class, you can use all the code from the old class (copying none of it) and you can also add the additional functionality you need.
#
# In the new class, we only need to define what we add or change from the old class. We call the original class either the parent, superclass, or base class. We call the new class the child, subclass, or derived class.
# + id="QE4zcQ6GgrmZ"
# parent class
class Bicycle():
pass
# child class is created by passing Bicycle class into
# the Specialzed class
# Specialized class will inherit whatever Bicycle class can do
class Specialized(Bicycle):
pass
# driver code
a_bike = Bicycle()
a_specialized = Specialized()
# + [markdown] id="jZrP_FmM7h2P"
# In this example, both these classes are useless unless we add to our code:
#
# + colab={"base_uri": "https://localhost:8080/"} id="NIHtbsNF7iUp" outputId="a760b280-1f59-4598-e052-ae2d0a730835"
class Bicycle():
def exclaim(self):
print("I'm a Bicycle!")
class Specialized(Bicycle):
pass
# driver code
a_bike = Bicycle()
a_specialized = Specialized()
a_bike.exclaim()
# I'm a Bicycle!
# + [markdown] id="q1-1a7Sa7h5W"
# Now, Any instance of the Specialized class has access to the methods on the parent class Bicycle.
#
#
# + [markdown] id="AvTagdtT7h9Z"
# **Overriding**
#
# If we override the exclaim method, an instance of Specialized will tell us what it is...
# + colab={"base_uri": "https://localhost:8080/"} id="AN5825mJ7iX_" outputId="75d2b963-6ada-4b76-fee4-8b9420166513"
class Bicycle():
def exclaim(self):
print("I'm a Bicycle!")
class Specialized(Bicycle):
def exclaim(self):
print("I'm Specialized! I'm a more specialized version of a Bicycle!")
a_bike = Bicycle()
a_specialized = Specialized()
a_bike.exclaim()
a_specialized.exclaim()
# I'm a Bicycle!
# I'm Specialized! I'm a more specialized version of a Bicycle!
# + [markdown] id="k2kYZDz5knqQ"
# **Using *Super***
#
# Calling the Parent Method
#
# 1. create a Student parent class
# 2. create a Graduate child class that inherits everthing in the parent class but also needs to hold a graduation_date
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="lSEjr1gs7ico" outputId="5a867af9-80c9-464e-bb42-0c404ca728e0"
class Student():
def __init__(self, name):
self.name = name
class Graduate(Student):
def __init__(self, name, graduation_date):
"""
overides __init__ method in Student
while using Student portion of child class to run initialiation
when instantiating a Graduate instance
"""
super().__init__(name)
# note: argument self is passed to parent class automatically
self.graduation_date = graduation_date
john_lennon = Graduate('<NAME>', 'December 8, 1980')
print(john_lennon.name)
print(john_lennon.graduation_date)
# <NAME>
# December 8, 1980
# + [markdown] id="9VAggIsqkoI6"
# **Class Association**
#
# What types of relationship exhist in inheritance?
#
#
# 1. **is_a**
#
#
# * parent/child
# * more general parent class is created to define shared attributes & behaviors
# * more specific child class is used for unique details
# * type/subtype
#
# Ex. Restraurant Mgmt Syst
#
# A chef is an employee
# A waiter is an emplyee
#
#
# 2. **has_a**
#
# * there is a link between two classes
# * Composition
# * class A CANNOT exist without class B
# * Aggregation
# * class A CAN exist independently of class B
#
# Ex. Restraurant Mgmt Syst
#
# If the resaurant disappeared there would be no employees, no menues, etc. (composition)
#
# There are several different drinks that are available with or without a menu (off menu items). (aggregation)
#
#
# >*We can define association reltionships between classes by creating attributes within class A that are instances of class B.*
#
#
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 144} id="AcVrQGms7ih9" outputId="bd034987-ed3d-49e9-ea5c-bc191666bf75"
class Restuarant:
def __init__(self, name, staff, dinner_menu):
self.name = name
self.staff = Employees[]
# has_an
self.dinner_menu = Menu()
# has_a
# class functions would be entered here...
class Employee:
def __init__(self, name, wage):
self.name = name
self.wage = wage
def cal_pay(self, hours):
return self.wage * hours
class Menu:
def __init__(self, name):
self.name = name
self.dishes = Food[]
self.drinks = Drink[]
def add_item(self, item):
pass
class Food:
def __init__(self, is_vegetarian=False):
self.name = name
self.is_vegetarian = is_vegetarian
class Drink:
def __init__(self, name):
pass
# + [markdown] id="dsG8haFmGHpx"
# **Composition**
#
# when we collect several objects together to create a new one
#
# has_a relationship
#
# Ex. A Duck is composed of a Tail, Bill, Eyes, Feet, etc.
#
# + colab={"base_uri": "https://localhost:8080/"} id="qBEtfZNaAmCf" outputId="55d8a99a-a9a1-4696-f1e4-58d7d9912826"
class Bill():
def __init__(self, description):
self.description = description
class Tail():
def __init__(self, length):
self.length = length
class Duck():
def __init__(self, bill, tail):
self.bill = bill
self.tail = tail
def about(self):
print(f'This duck has a {self.bill.description} bill and a {self.tail.length} tail. ')
duck = Duck(Bill('wide, orange'), Tail('long'))
duck.about()
# This duck has a wide orange bill and a long tail. ')
# + [markdown] id="8vFYL4feIPvi"
# **Aggregation**
#
# Aggregation is almost exactly like composition. The difference is that aggregate objects can exist independently. It would be impossible for a bill to be associated with a different duck. But what if there was a Tag or Feathers class? A Duck could have a tag or feathers, and, unlike a Bill or Tail, the tag or feathers could exist independently from the Duck.
#
# Another way to differentiate between aggregation and composition is to consider the object's lifespan. If the outside object (Duck in this example) controls when the inside objects are created and destroyed (like Bill or Tail), composition is most suitable. If the related object (like tag or feathers) can outlast the composition object (Duck in this example), then an aggregate relationship makes more sense.
#
#
#
| CS41long_U1S1M2_Warmup_RJProctor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="poK9I4m9wSTW"
# # Catalyst - customizing what happens in `train()`
# based on `Keras customizing what happens in fit`
# + [markdown] colab_type="text" id="NJb_eBKEwXS6"
# ## Introduction
#
# When you're doing supervised learning, you can use `train()` and everything works smoothly.
#
# A core principle of Catalyst is **progressive disclosure of complexity**. You should always be able to get into lower-level workflows in a gradual way. You shouldn't fall off a cliff if the high-level functionality doesn't exactly match your use case. You should be able to gain more control over the small details while retaing a commensurate amount of high-level convenience.
#
# When you need to customize what `train()` does, you should **override the `handle_batch` function of the `Runner` class**. This is the function that is called by `train()` for every batch of data. You will then be able to call `train()` as usual -- and it will be running your own learning algorithm.
#
# Note that this pattern does not prevent you from building models with the Functional API. You can do this with **any** PyTorch model.
#
# Let's see how that works.
# + [markdown] colab_type="text" id="E6R34jh5xKkW"
# ## Setup
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 9418, "status": "ok", "timestamp": 1587014070467, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjYXcxiGiIDQYhW2wTkdrNLwx68llP5BzH91oGlAQ=s64", "userId": "07081474162282073276"}, "user_tz": -180} id="S1rkIIaKaG2O" outputId="5c4d2dc1-74a0-4b04-f3f5-9a08e8efc28c"
# !pip install catalyst[ml]==21.4.2
# don't forget to restart runtime for correct `PIL` work with Colab
# + colab={"base_uri": "https://localhost:8080/", "height": 255} colab_type="code" executionInfo={"elapsed": 1572, "status": "ok", "timestamp": 1587014153359, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjYXcxiGiIDQYhW2wTkdrNLwx68llP5BzH91oGlAQ=s64", "userId": "07081474162282073276"}, "user_tz": -180} id="3eLP4fR6wCYc" outputId="1e36766e-6d62-46da-894f-8d2e4967544c"
import catalyst
from catalyst import dl, metrics, utils
catalyst.__version__
# + [markdown] colab_type="text" id="5F8q4oByxt2T"
# ## A first simple example
#
# Let's start from a simple example:
#
# - We create a new runner that subclasses `dl.Runner`.
# - We just override the `handle_batch(self, batch)` method for custom train step logic
# - And update `on_loader_start`/`on_loader_start` handlers for correct custom metrics aggregation.
#
# The input argument `batch` is what gets passed to fit as training data. If you pass a `torch.utils.data.DataLoader`, by calling `train(loaders={"train": loader, "valid": loader}, ...)`, then `batch` will be what gets yielded by `loader` at each batch.
#
# In the body of the `handle_batch` method, we implement a regular training update, similar to what you are already familiar with. Importantly, **we log batch-based metrics via `self.batch_metrics`**, which passes them to the loggers.
#
# Addiionally, we have to use [`AdditiveValueMetric`](https://catalyst-team.github.io/catalyst/api/metrics.html#additivevaluemetric) during `on_loader_start` and `on_loader_start` for correct metrics aggregation for the whole loader. Importantly, **we log loader-based metrics via `self.loader_metrics`**, which passes them to the loggers.
# + colab={} colab_type="code" id="MbTkRLQUxQmC"
import torch
from torch.nn import functional as F
class CustomRunner(dl.Runner):
def on_loader_start(self, runner):
super().on_loader_start(runner)
self.meters = {
key: metrics.AdditiveValueMetric(compute_on_call=False)
for key in ["loss", "mae"]
}
def handle_batch(self, batch):
# Unpack the data. Its structure depends on your model and
# on what you pass to `train()`.
x, y = batch
y_pred = self.model(x) # Forward pass
# Compute the loss value
loss = F.mse_loss(y_pred, y)
# Update metrics (includes the metric that tracks the loss)
self.batch_metrics.update({"loss": loss, "mae": F.l1_loss(y_pred, y)})
for key in ["loss", "mae"]:
self.meters[key].update(self.batch_metrics[key].item(), self.batch_size)
if self.is_train_loader:
# Compute gradients
loss.backward()
# Update weights
# (the optimizer is stored in `self.state`)
self.optimizer.step()
self.optimizer.zero_grad()
def on_loader_end(self, runner):
for key in ["loss", "mae"]:
self.loader_metrics[key] = self.meters[key].compute()[0]
super().on_loader_end(runner)
# + [markdown] colab_type="text" id="nAEiVP4IzNj-"
# Let's try this out:
# + colab={"base_uri": "https://localhost:8080/", "height": 562} colab_type="code" executionInfo={"elapsed": 17386, "status": "error", "timestamp": 1587015544733, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjYXcxiGiIDQYhW2wTkdrNLwx68llP5BzH91oGlAQ=s64", "userId": "07081474162282073276"}, "user_tz": -180} id="AlUHnIG6zPV9" outputId="bcc53cac-174d-4a1e-c3e8-441c102609cb"
import numpy as np
import torch
from torch.utils.data import DataLoader, TensorDataset
# Construct custom data
num_samples, num_features = int(1e4), int(1e1)
X, y = torch.rand(num_samples, num_features), torch.rand(num_samples, 1)
dataset = TensorDataset(X, y)
loader = DataLoader(dataset, batch_size=32, num_workers=1)
loaders = {"train": loader, "valid": loader}
# and model
model = torch.nn.Linear(num_features, 1)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters())
# and use `train`
runner = CustomRunner()
runner.train(
model=model,
optimizer=optimizer,
loaders=loaders,
num_epochs=3,
verbose=True, # you can pass True for more precise training process logging
timeit=False, # you can pass True to measure execution time of different parts of train process
)
# + [markdown] colab_type="text" id="NGJgVd9lzkQc"
# ## Going high-level
#
# Naturally, you could skip a loss function backward in `handle_batch()`, and instead do everything with `Callbacks` in `train` params. Likewise for metrics. Here's a high-level example, that only uses `handle_batch()` for model forward pass and metrics computation:
# +
import numpy as np
import torch
from torch.nn import functional as F
from torch.utils.data import DataLoader, TensorDataset
class CustomRunner(dl.Runner):
def on_loader_start(self, runner):
super().on_loader_start(runner)
self.meters = {
key: metrics.AdditiveValueMetric(compute_on_call=False)
for key in ["loss", "mae"]
}
def handle_batch(self, batch):
# Unpack the data. Its structure depends on your model and
# on what you pass to `train()`.
x, y = batch
y_pred = self.model(x) # Forward pass
# Compute the loss value
# (the criterion is stored in `self.state` also)
loss = self.criterion(y_pred, y)
# Update metrics (includes the metric that tracks the loss)
self.batch_metrics.update({"loss": loss, "mae": F.l1_loss(y_pred, y)})
for key in ["loss", "mae"]:
self.meters[key].update(self.batch_metrics[key].item(), self.batch_size)
def on_loader_end(self, runner):
for key in ["loss", "mae"]:
self.loader_metrics[key] = self.meters[key].compute()[0]
super().on_loader_end(runner)
# Construct custom data
num_samples, num_features = int(1e4), int(1e1)
X, y = torch.rand(num_samples, num_features), torch.rand(num_samples, 1)
dataset = TensorDataset(X, y)
loader = DataLoader(dataset, batch_size=32, num_workers=1)
loaders = {"train": loader, "valid": loader}
# and model
model = torch.nn.Linear(num_features, 1)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters())
# Just use `train` as usual
runner = CustomRunner()
runner.train(
model=model,
optimizer=optimizer,
criterion=criterion, # you could also pass any PyTorch criterion for loss computation
scheduler=None, # or scheduler, but let's simplify the train loop for now :)
loaders=loaders,
num_epochs=3,
verbose=True,
timeit=False,
callbacks={
"optimizer": dl.OptimizerCallback(
metric_key="loss", # you can also pass 'mae' to optimize it instead
# generaly, you can optimize any differentiable metric from `runner.batch_metrics`
accumulation_steps=1, # also you can pass any number of steps for gradient accumulation
grad_clip_fn=None, # or you can use `grad_clip_fn=nn.utils.clip_grad_norm_`
grad_clip_params=None, # with `grad_clip_params={max_norm=1, norm_type=2}`
# or `grad_clip_fn=nn.utils.clip_grad_value_`
# with `grad_clip_params={clip_value=1}`
# for gradient clipping during training!
# for more information about gradient clipping please follow pytorch docs
# https://pytorch.org/docs/stable/nn.html#clip-grad-norm
)
}
)
# + [markdown] colab_type="text" id="vVZtc6P61icn"
# ## Metrics support through Callbacks
#
# Let's go even deeper! Could we transfer different metrics/criterions computation to `Callbacks` too? Of course! If you want to support different losses, you'd simply do the following:
#
# - Do your model forward pass as usual.
# - Save all batch-based artefacts to `self.batch`, so Callbacks can find it.
# - Add extra callbacks, that will use data from `runner.batch` during training.
#
# That's it. That's the list. Let's see the example:
# +
import numpy as np
import torch
from torch.nn import functional as F
from torch.utils.data import DataLoader, TensorDataset
class CustomRunner(dl.Runner):
def handle_batch(self, batch):
# Unpack the data. Its structure depends on your model and
# on what you pass to `train()`.
x, y = batch
y_pred = self.model(x) # Forward pass
# pass all batch-based artefacts to `self.batch`
# we recommend to use key-value storage to make it Callbacks-friendly
self.batch = {"features": x, "targets": y, "logits": y_pred}
# Construct custom data
num_samples, num_features = int(1e4), int(1e1)
X, y = torch.rand(num_samples, num_features), torch.rand(num_samples, 1)
dataset = TensorDataset(X, y)
loader = DataLoader(dataset, batch_size=32, num_workers=1)
loaders = {"train": loader, "valid": loader}
# and model
model = torch.nn.Linear(num_features, 1)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters())
# Just use `train` as usual
runner = CustomRunner()
runner.train(
model=model,
optimizer=optimizer,
criterion=criterion,
scheduler=None,
loaders=loaders,
num_epochs=3,
verbose=True,
timeit=False,
callbacks={
# alias for
# `runner.batch_metrics[metric_key] = \
# runner.criterion[criterion_key](runner.batch[input_key], runner.batch[target_key])`
"criterion": dl.CriterionCallback( # special Callback for criterion computation
input_key="logits", # `input_key` specifies model predictions (`y_pred`) from `runner.batch`
target_key="targets", # `target_key` specifies correct labels (or `y_true`) from `runner.batch`
metric_key="loss", # `metric_key` - key to use with `runner.batch_metrics`
criterion_key=None, # `criterion_key` specifies criterion in case of key-value runner.criterion
# if `criterion_key=None`, runner.criterion used for computation
),
# alias for
# `runner.batch_metrics[metric_key] = \
# metric_fn(runner.batch[input_key], runner.batch[target_key])`
"metric": dl.FunctionalMetricCallback( # special Callback for metrics computation
input_key="logits", # the same logic as with `CriterionCallback`
target_key="targets", # the same logic as with `CriterionCallback`
metric_key="loss_mae", # the same logic as with `CriterionCallback`
metric_fn=F.l1_loss, # metric function to use
),
"optimizer": dl.OptimizerCallback(
metric_key="loss",
accumulation_steps=1,
grad_clip_fn=None,
grad_clip_params=None,
)
}
)
# -
# ## Simplify it a bit - SupervisedRunner
#
# But can we simplify last example a bit? <br/>
# What if we know, that we are going to train `supervised` model, that will take some `features` in and output some `logits` back? <br/>
# Looks like commom case... could we automate it? Let's check it out!
# +
import numpy as np
import torch
from torch.nn import functional as F
from torch.utils.data import DataLoader, TensorDataset
# Construct custom data
num_samples, num_features = int(1e4), int(1e1)
X, y = torch.rand(num_samples, num_features), torch.rand(num_samples, 1)
dataset = TensorDataset(X, y)
loader = DataLoader(dataset, batch_size=32, num_workers=1)
loaders = {"train": loader, "valid": loader}
# and model
model = torch.nn.Linear(num_features, 1)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters())
# Just use `train` as usual
runner = dl.SupervisedRunner( # `SupervisedRunner` works with any model like `some_output = model(some_input)`
input_key="features", # if your dataloader yields (x, y) tuple, it will be transformed to
output_key="logits", # {input_key: x, target_key: y} and stored to runner.batch
target_key="targets", # then the model will be used like
loss_key="loss", # runner.batch[runner.output_key] = model(runner.batch[input_key])
) # loss computation suppose to looks like
# loss = criterion(runner.batch[runner.output_key], runner.batch[runner.target_key])
# and stored to `runner.batch_metrics[runner.loss_key]`
# thanks to prespecified `input_key`, `output_key`, `target_key` and `loss_key`
# `SupervisedRunner` automatically adds required `CriterionCallback` and `OptimizerCallback`
# moreover, with specified `logdir`, `valid_loader` and `valid_metric`
# `SupervisedRunner` automatically adds `CheckpointCallback` and tracks best performing based on selected metric
runner.train(
model=model,
optimizer=optimizer,
criterion=criterion,
scheduler=None,
loaders=loaders,
num_epochs=3,
verbose=True,
timeit=False,
valid_loader="valid", # `loader_key` from loaders to use for model selection
valid_metric="loss", # `metric_key` to use for model selection
logdir="./logs_supervised", # logdir to store models checkpoints
callbacks={
# "criterion_mse": dl.CriterionCallback(
# input_key="logits",
# target_key="targets",
# metric_key="loss",
# ),
"criterion_mae": dl.FunctionalMetricCallback(
input_key="logits",
target_key="targets",
metric_key="mae",
metric_fn=F.l1_loss,
),
# "optimizer": dl.OptimizerCallback(
# metric_key="loss",
# accumulation_steps=1,
# grad_clip_fn=None,
# grad_clip_params=None,
# )
}
)
# + [markdown] colab_type="text" id="rn1q6NCP2dtR"
# ## Providing your own inference step
#
# But let's return to the basics.
#
# What if you want to do the same customization for calls to `runner.predict_*()`? Then you would override `predict_batch` in exactly the same way. Here's what it looks like:
# +
import torch
from torch.nn import functional as F
class CustomRunner(dl.Runner):
def predict_batch(self, batch): # here is the trick
return self.model(batch[0].to(self.device)) # you can write any prediciton logic here
# our first time example
def on_loader_start(self, runner):
super().on_loader_start(runner)
self.meters = {
key: metrics.AdditiveValueMetric(compute_on_call=False)
for key in ["loss", "mae"]
}
def handle_batch(self, batch):
# Unpack the data. Its structure depends on your model and
# on what you pass to `train()`.
x, y = batch
y_pred = self.model(x) # Forward pass
# Compute the loss value
loss = F.mse_loss(y_pred, y)
# Update metrics (includes the metric that tracks the loss)
self.batch_metrics.update({"loss": loss, "mae": F.l1_loss(y_pred, y)})
for key in ["loss", "mae"]:
self.meters[key].update(self.batch_metrics[key].item(), self.batch_size)
if self.is_train_loader:
# Compute gradients
loss.backward()
# Update weights
# (the optimizer is stored in `self.state`)
self.optimizer.step()
self.optimizer.zero_grad()
def on_loader_end(self, runner):
for key in ["loss", "mae"]:
self.loader_metrics[key] = self.meters[key].compute()[0]
super().on_loader_end(runner)
# +
import numpy as np
import torch
from torch.utils.data import DataLoader, TensorDataset
# Construct custom data
num_samples, num_features = int(1e4), int(1e1)
X, y = torch.rand(num_samples, num_features), torch.rand(num_samples, 1)
dataset = TensorDataset(X, y)
loader = DataLoader(dataset, batch_size=32, num_workers=1)
loaders = {"train": loader, "valid": loader}
# and model
model = torch.nn.Linear(num_features, 1)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters())
# Just use `train` as usual
runner = CustomRunner()
runner.train(
model=model,
optimizer=optimizer,
loaders=loaders,
num_epochs=3,
verbose=True,
timeit=False,
valid_loader="valid", # `loader_key` from loaders to use for model selection
valid_metric="loss", # `metric_key` to use for model selection
load_best_on_end=True, # flag to load best model at the end of the training process
logdir="./logs", # logdir to store models checkpoints (required for `load_best_on_end`)
)
# and use `batch` prediciton
prediction = runner.predict_batch(next(iter(loader))) # let's sample first batch from loader
# or `loader` prediction
for prediction in runner.predict_loader(loader=loader):
assert prediction.detach().cpu().numpy().shape[-1] == 1 # as we have 1-class regression
# -
# Finally, after model training and evaluation, it's time to prepare it for deployment. PyTorch upport model tracing for production-friendly Deep Leanring models deployment.
#
# Could we make it quick with Catalyst? Sure!
features_batch = next(iter(loaders["valid"]))[0].to(runner.device)
# model stochastic weight averaging
model.load_state_dict(utils.get_averaged_weights_by_path_mask(logdir="./logs", path_mask="*.pth"))
# model tracing
utils.trace_model(model=runner.model, batch=features_batch)
# model quantization
utils.quantize_model(model=runner.model)
# model pruning
utils.prune_model(model=runner.model, pruning_fn="l1_unstructured", amount=0.8)
# onnx export, catalyst[onnx] or catalyst[onnx-gpu] required
# utils.onnx_export(model=runner.model, batch=features_batch, file="./logs/mnist.onnx", verbose=True)
# + [markdown] colab_type="text" id="9-CMpP5a3Wcp"
# ## Wrapping up: an end-to-end GAN example
#
# Let's walk through an end-to-end example that leverages everything you just learned.
#
# Let's consider:
#
# - A generator network meant to generate 28x28x1 images.
# - A discriminator network meant to classify 28x28x1 images into two classes ("fake" - 1 and "real" - 0).
#
#
# +
import torch
from torch import nn
from torch.nn import functional as F
from catalyst.contrib.nn.modules import Flatten, GlobalMaxPool2d, Lambda
# Create the discriminator
discriminator = nn.Sequential(
nn.Conv2d(1, 64, (3, 3), stride=(2, 2), padding=1),
nn.LeakyReLU(0.2, inplace=True),
nn.Conv2d(64, 128, (3, 3), stride=(2, 2), padding=1),
nn.LeakyReLU(0.2, inplace=True),
GlobalMaxPool2d(),
Flatten(),
nn.Linear(128, 1),
)
# Create the generator
latent_dim = 128
generator = nn.Sequential(
# We want to generate 128 coefficients to reshape into a 7x7x128 map
nn.Linear(128, 128 * 7 * 7),
nn.LeakyReLU(0.2, inplace=True),
Lambda(lambda x: x.view(x.size(0), 128, 7, 7)),
nn.ConvTranspose2d(128, 128, (4, 4), stride=(2, 2), padding=1),
nn.LeakyReLU(0.2, inplace=True),
nn.ConvTranspose2d(128, 128, (4, 4), stride=(2, 2), padding=1),
nn.LeakyReLU(0.2, inplace=True),
nn.Conv2d(128, 1, (7, 7), padding=3),
nn.Sigmoid(),
)
# Final model
model = {"generator": generator, "discriminator": discriminator}
criterion = {"generator": nn.BCEWithLogitsLoss(), "discriminator": nn.BCEWithLogitsLoss()}
optimizer = {
"generator": torch.optim.Adam(generator.parameters(), lr=0.0003, betas=(0.5, 0.999)),
"discriminator": torch.optim.Adam(discriminator.parameters(), lr=0.0003, betas=(0.5, 0.999)),
}
# + [markdown] colab_type="text" id="POY42XRf5Jbd"
# Here's a feature-complete `GANRunner`, overriding `predict_batch()` to use its own signature, and implementing the entire GAN algorithm in 16 lines in `handle_batch`:
# + colab={} colab_type="code" id="iyKOtjfn5RL3"
class GANRunner(dl.Runner):
def __init__(self, latent_dim: int):
super().__init__()
self.latent_dim = latent_dim
def predict_batch(self, batch):
batch_size = 1
# Sample random points in the latent space
random_latent_vectors = torch.randn(batch_size, self.latent_dim).to(self.device)
# Decode them to fake images
generated_images = self.model["generator"](random_latent_vectors).detach()
return generated_images
def handle_batch(self, batch):
real_images, _ = batch
batch_size = real_images.shape[0]
# Sample random points in the latent space
random_latent_vectors = torch.randn(batch_size, self.latent_dim).to(self.device)
# Decode them to fake images
generated_images = self.model["generator"](random_latent_vectors).detach()
# Combine them with real images
combined_images = torch.cat([generated_images, real_images])
# Assemble labels discriminating real from fake images
labels = \
torch.cat([torch.ones((batch_size, 1)), torch.zeros((batch_size, 1))]).to(self.device)
# Add random noise to the labels - important trick!
labels += 0.05 * torch.rand(labels.shape).to(self.device)
# Discriminator forward
combined_predictions = self.model["discriminator"](combined_images)
# Sample random points in the latent space
random_latent_vectors = torch.randn(batch_size, self.latent_dim).to(self.device)
# Assemble labels that say "all real images"
misleading_labels = torch.zeros((batch_size, 1)).to(self.device)
# Generator forward
generated_images = self.model["generator"](random_latent_vectors)
generated_predictions = self.model["discriminator"](generated_images)
self.batch = {
"combined_predictions": combined_predictions,
"labels": labels,
"generated_predictions": generated_predictions,
"misleading_labels": misleading_labels,
}
# + [markdown] colab_type="text" id="zYGZRIJh6ZYu"
# Let's test-drive it:
# +
import os
from torch.utils.data import DataLoader
from catalyst.contrib.datasets import MNIST
from catalyst.data import ToTensor
loaders = {
"train": DataLoader(
MNIST(os.getcwd(), train=True, download=True, transform=ToTensor()),
batch_size=64),
}
runner = GANRunner(latent_dim=latent_dim)
runner.train(
model=model,
criterion=criterion,
optimizer=optimizer,
loaders=loaders,
callbacks=[
dl.CriterionCallback(
input_key="combined_predictions",
target_key="labels",
metric_key="loss_discriminator",
criterion_key="discriminator",
),
dl.CriterionCallback(
input_key="generated_predictions",
target_key="misleading_labels",
metric_key="loss_generator",
criterion_key="generator",
),
dl.OptimizerCallback(
model_key="generator",
optimizer_key="generator",
metric_key="loss_generator"
),
dl.OptimizerCallback(
model_key="discriminator",
optimizer_key="discriminator",
metric_key="loss_discriminator"
),
],
valid_loader="train",
valid_metric="loss_generator",
minimize_valid_metric=True,
num_epochs=20,
verbose=True,
logdir="./logs_gan",
)
# + [markdown] colab_type="text" id="M9Fz5_u68FqW"
# The idea behind deep learning are simple, so why should their implementation be painful?
# +
import matplotlib.pyplot as plt
# %matplotlib inline
utils.set_global_seed(42)
generated_image = runner.predict_batch(None)
plt.imshow(generated_image[0, 0].detach().cpu().numpy())
# -
# %load_ext tensorboard
# %tensorboard --logdir ./logs_gan
| examples/notebooks/customizing_what_happens_in_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="hWkYCPEDXobK"
# # COVID-19
# #### Finding the countries with similar spread
# + [markdown] colab_type="text" id="uT5KobbVXobN"
# ***
# + [markdown] colab_type="text" id="mUfunpOsXobR"
# This notebook aims to clarify how the nearest neighbours for each country are found. This is useful because there is a high chance that alike countries will keep evolving in the same way.
# + colab={} colab_type="code" id="XL1IKK0IXobU"
import os
os.chdir('..')
# + colab={} colab_type="code" id="NH0vUJe9Xobk"
import warnings
import statistics
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
# sns.set()
from core.data import utils as dataUtils
from pandas_profiling import ProfileReport
from sklearn.metrics import mean_squared_error, mean_absolute_error
# + [markdown] colab_type="text" id="9hWbKMxHXobu"
# ## 1. Loading data
# + [markdown] colab_type="text" id="G0LCathCXobv"
# The latest data can be fetched by running ```python scripts/fetch_latest_data.py```. This will download the new global data from Johns Hopkins University GitHub [repo](https://github.com/CSSEGISandData/COVID-19) (they update the data on a daily basis.) Also, it will convert it to a more convenient format, namely the one used in [Kaggles COVID-19 spread](https://www.kaggle.com/c/covid19-global-forecasting-week-4) competition. The main reason for converting the data was that when I first started working on this project only the Kaggle data was available, and rather that rewriting a good portion of the code, I found this solution to be easier.
#
# The dataset contains the spread evolution for each country (except USA). Large countries are divided into zones for better granularity.
# + colab={} colab_type="code" id="yfecPBI0Xoby" outputId="31088945-5822-4a00-fd5d-fe84ce7c755b"
allData = pd.read_csv('assets/covid_spread.csv', parse_dates=['Date'])
allData.head()
# + [markdown] colab_type="text" id="88T_DjJPXob8"
# Preprocess the input dataframe. Fill empty states with their country names (this will make working with regions much easier).
# + colab={} colab_type="code" id="XwA-5HxbXob-"
def preprocess_data(df):
renameState = df['Province_State'].fillna(0).values
renameCountries = df['Country_Region'].values
renameState[renameState == 0] = renameCountries[renameState == 0]
df['Province_State'] = renameState
return df
# + colab={} colab_type="code" id="WHU8NcrFXocH" outputId="36dec400-8dc4-4bcf-f9b3-8fa990a87c4a"
allData = preprocess_data(allData)
allData.head()
# + [markdown] colab_type="text" id="vEACFMyWXocS"
# Because exploratory data analysis is beyond the purpose of this notebook, I leave you the following reccomandation on Kaggle:
# - [COVID19-Explained through Visualizations](https://www.kaggle.com/anshuls235/covid19-explained-through-visualizations)
# + [markdown] colab_type="text" id="rEt6VxayXocT"
# ## Approach
# + [markdown] colab_type="text" id="csetdTj4XocU"
# ### Comparing 2 countries
# + [markdown] colab_type="text" id="AFkgavG6XocV"
# Two countries are compared by incrementally sliding the source country **S** over the cadidate country **C** and computing the error (difference) between them. If **C** is not as evolved as **S**, we do not make the comparison.
# + colab={} colab_type="code" id="YNNDkiMxXocX"
def compare_sequence(source, candidate, errorFunc):
minError = np.inf
minIdx = -1
# only check the countries that can influence
if len(candidate) > len(source):
noWindows = len(candidate) - len(source)
windowSize = len(source)
# sliding window over candidate country
for i in range(0, noWindows):
# compute loss
error = errorFunc(source, candidate[i:i + windowSize])
# save the min error and its location
if error <= minError:
minError = error
minIdx = i
return minError, minIdx
# return none if invalid
return None, None
# + [markdown] colab_type="text" id="sNr9IN9jXoce"
# ### Getting all neighbours
# + [markdown] colab_type="text" id="QPUttT2QXocf"
# We discard the entries (days) which are below a specified alignment threshold for each feature **$TA_f$** (have less than a specified number of cases), for every country (**S** included). Then, we take a candidate country, **$C_n$**. **$C_n$** must be more evolved than **S** (this means it reached **$T_a$** earlier). We start sliding **S** over **$C_n$**, beginning with the first day it reached the treshold, until **$C_n$** ends. For each such step, an error is computed. The smallest error will be the error associated with **$C_n$**. We do this for all countries that are available in the dataset, taking one feature f, f in {confirmedCases, fatalities}, at a time. During training, the neighbours will be filtered by applying an error threshold **$T_{error}$**.
#
# **Note:** in Pandas, sort_values is a very time expensive operation, this can be avoided if we are sure that the input data is sorted. I opted for doing this for safety reasons.
# + colab={} colab_type="code" id="pndeknu1Xocg"
def get_nearest_sequence(df, state, alignThreshConf = 50, alignThreshDead = 10, errorFunc = mean_absolute_error):
resDf = pd.DataFrame(columns=['Province_State', 'deathError', 'confirmedError', 'deathIdx', 'confirmedIdx'])
confDf = df[df['ConfirmedCases'] > alignThreshConf]
deadDf = df[df['Fatalities'] > alignThreshDead]
# get source region data
regionDfConf = confDf[confDf['Province_State'] == state].sort_values(by='Date', ascending=True)
regionDfDead = deadDf[deadDf['Province_State'] == state].sort_values(by='Date', ascending=True)
regionConf = regionDfConf['ConfirmedCases'].values
regionDead = regionDfDead['Fatalities'].values
# check all possible candidates
for neighbour in df['Province_State'].unique():
# skip comparing with the same country
if neighbour == state:
continue
# get country candidate
confNeighDf = confDf[confDf['Province_State'] == neighbour].sort_values(by='Date', ascending = True)
deadNeighDf = deadDf[deadDf['Province_State'] == neighbour].sort_values(by='Date', ascending = True)
neighConf = confNeighDf['ConfirmedCases'].values
neighDead = deadNeighDf['Fatalities'].values
# get error for confirmed and neighbour
confErr, confIdx = compare_sequence(regionConf, neighConf, errorFunc)
deadErr, deadIdx = compare_sequence(regionDead, neighDead, errorFunc)
# the candidate will be ignored if it does not have enough data
if confErr is None or deadErr is None:
continue
# append result
res = {'Province_State':neighbour, 'deathError':deadErr, 'confirmedError':confErr,
'deathIdx':deadIdx, 'confirmedIdx':confIdx}
resDf = resDf.append(res, ignore_index=True)
return resDf
# + [markdown] colab_type="text" id="76tLJ9tnXoco"
# Now, let's display the results:
# + colab={} colab_type="code" id="YWhWy_4tXocq" outputId="acc037f9-0a25-4e66-8f95-984539d3176c"
r = get_nearest_sequence(allData, 'Germany', 40, 10)
r.head()
# + [markdown] colab_type="text" id="WTn9szRJXocy"
# ## Choosing the right metric
# + [markdown] colab_type="text" id="3to3gkzPXoc0"
# It is essential to choose a right metric such that the results are relevant. In essence, there should not be a huge difference between the losses, but it is safer to investigate at least some of the major error functions and see which one yields the best result.
# + [markdown] colab_type="text" id="lgp333e0Xoc2"
# ### Metrics
# + [markdown] colab_type="text" id="DkBvmY1YXoc3"
# #### 3.1.3 Mean absolute error
# + [markdown] colab_type="text" id="l7VA8fZbXoc4"
# Arithmetic average of absolute errors.
#
# $$MAE = \frac{1}{n}\sum_{t=1}^n\left| S_t-C_t\right|$$
# + [markdown] colab_type="text" id="F60avdAlXoc6"
# #### 3.1.1 Mean average percentage error
# + [markdown] colab_type="text" id="Fdc0RzLtXoc7"
# This yields the arithmetic average of the errors in percentage, relative to the source. This can be very easily thresholded and is easy to understand.
#
# $$MAPE = \frac{1}{n}\sum_{t=1}^n \left|\frac{S_t-C_t}{S_t}\right|$$
#
# Below is a naive implementation.
# + colab={} colab_type="code" id="AzBx3jfUXoc8"
def l1_norm_error(source, candidate):
error = (abs(source - candidate))
source[source == 0] = 1e-30 # add for numerical stability
error = error/source # normalize the error
error = error.mean()
return error
# + [markdown] colab_type="text" id="qo1TA0mVXodG"
# #### 3.1.2 Root mean squared log error
# + [markdown] colab_type="text" id="8_zT6b-nXodK"
# It implies a larger penalty for the underestimation of the result rather than the overestimating of it. This is the metric used in Kaggle competition for evaluation.
# $$ RMSLE = \sqrt{ \frac{1}{n} \sum_{t=1}^{n} (\log{S_t} - \log{C_t})^2} $$
#
# Below is a naive implementation.
# + colab={} colab_type="code" id="pasKkdRYXodS"
def rmsle_error(source, candidate):
candidate += 1e-30
error = np.log10((source+1)/(candidate+1)) # 1 is added for numerical stability
error = error * error
error = error.mean()
error = np.sqrt(error)
return error
# + [markdown] colab_type="text" id="ulL_394YXodY"
# ### Comparing metrics
# + [markdown] colab_type="text" id="oxsh3by6XodZ"
# Now lets compare the afore mentioned losses. For this, is important to consider 2 types of countries:
# - **top countries** - where the disease spreaded first and have the highest number of Confirmed Cases and Fatalities; this type of countries won't have many possible neighbours.
# - **average countries** - where the disease has spread in a normal way, this countries will have lots of possible neighbours
# + colab={} colab_type="code" id="SBaxYjXwXoda"
def show_country_nn(data, sourceState, alignThreshConf, alignThreshDead, listErrorDf, errorNames):
SHOW_FIRST = 3 # only show the first top neighbours
# setup plot figures
fig, axes = plt.subplots(len(listErrorDf), 2,
figsize=(15, len(listErrorDf) * 3),
gridspec_kw={'hspace': 0.3})
# get rid of the annoying
axes = axes.flatten()
fig.suptitle(sourceState.title(), fontsize=20)
colors = sns.color_palette()[:SHOW_FIRST + 1]
# only keep aligned data
showDataConf = data[data['ConfirmedCases'] > alignThreshConf].copy()
showDataDead = data[data['Fatalities'] > alignThreshDead].copy()
showData = [showDataConf, showDataDead]
for i, (attr, err) in enumerate(zip(['ConfirmedCases', 'Fatalities'],
['confirmedError', 'deathError'])):
for j, (error, name) in enumerate(zip(listErrorDf, errorNames)):
legend = []
axIdx = j * 2 + i
tempError = error.sort_values(by=err, ascending=True)
# only show available neighbours (if they are less than SHOW_FIRST)
show = min(SHOW_FIRST, tempError.shape[0])
for k in range(1, show + 1):
# plot neighbours
neighbour = tempError['Province_State'].iloc[k - 1]
tempShow = showData[i][showData[i]['Province_State'] == neighbour][attr]
xAxisValues = [z for z in range(tempShow.shape[0])]
if len(xAxisValues) > 0:
legend.append(neighbour)
sns.lineplot(x=xAxisValues, y=tempShow, color=colors[k],
ax=axes[axIdx], linewidth=4.5)
# plot source country
tempShow = showData[i][showData[i]['Province_State'] == sourceState][attr]
xAxisValues = [z for z in range(tempShow.shape[0])]
sns.lineplot(x=xAxisValues, y=tempShow, color=colors[0],
ax=axes[axIdx], linewidth=4.5)
# final touches to figure
axes[axIdx].legend(legend + [sourceState])
axes[axIdx].set_title(name.title() + ' error')
axes[axIdx].grid(True)
axes[axIdx].box = True
return axes
# + [markdown] colab_type="text" id="Kie38UF3Xodj"
# This will show individual results for a country.
# + colab={} colab_type="code" id="4Cd6m75BXodk"
def test_metrics(trainData, sourceCountry, alignThreshConf, alignThreshDead):
results = []
errorNames = ['MAPE', 'MSE', 'RMSLE']
errors = [l1_norm_error, mean_absolute_error, rmsle_error]
# compute error df for each metric
for error in errors:
r = get_nearest_sequence(trainData, sourceCountry, alignThreshConf, alignThreshDead , error)
results.append(r)
# diplay for both fatalities and confirmed cases
show_country_nn(trainData, sourceCountry, alignThreshConf, alignThreshDead, results, errorNames)
# + [markdown] colab_type="text" id="ZlbQpDmMXod3"
# #### Evolved country
# + [markdown] colab_type="text" id="t8nDJjI_Xod3"
# An evolved country is a country having a high number of confirmed cases and fatalities.For this reason, such a country will not have many candidate countries with which they can be compared. Examples of evolved countries could be considered Italy or Spain.
# + colab={} colab_type="code" id="51xtSrKjXod4" outputId="e9050bc6-e6e5-413a-deaf-98adc24d997a"
test_metrics(allData, 'Italy', 500, 100)
# + [markdown] colab_type="text" id="kiyeD9-2Xod-"
# In this edge cases, the algorithm does not produce relevant results, hence there are not many countries that resemble the same growth. The algorithm was set to display the first 3 neighbours, but hence there aren't that many, only the possible ones are shown.
# + [markdown] colab_type="text" id="0-lwAimLXods"
# #### Normal spread country
# + [markdown] colab_type="text" id="EFYa1v5-Xodt"
# To precisly choose a strong representant for this category, it would be ideal to iterate over all countries, compute the average number of possible neighbours and then choose a country which comes close to that number. To avoid doing all of this, an educated guess would be a country from Eastern Europe, for example Romania or Czechia.
#
# The threshold for each category is chosen such that the beggining plateou of the spreading is ignored (that period that has 1 confirmed case or so). A large threshold means using more recent data, and this is relevant for the prediction on short term, but it could also lead to irellevant results.
# + colab={} colab_type="code" id="4Li19MrvXodu" outputId="26d498fe-2173-4426-cbfb-b052759e29ce"
test_metrics(allData, 'Romania', 500, 100)
# + [markdown] colab_type="text" id="dhj0ER21Xod_"
# ## 4. Results
# -
# Below are the graphs for the results obtained for confirmed cases (left) and
# fatalities (right). The thresholds applied are 500 and 40 respectively.
# The data that was used was last updated on ```26.04.2020``` .
# + colab={} colab_type="code" id="bpJwhNsaXoeB" outputId="8a3bd6cd-7f0f-46aa-d443-910bd851a4e7"
for country in ['Romania', 'Germany', 'Czechia']:
r = get_nearest_sequence(allData, country, 500, 40, l1_norm_error)
show_country_nn(allData, country, 500, 40, [r], ['MAPE'])
| notebooks/Covid_19_Country_growth_similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Emerging technology and trends
# ## OGC API
# The OGC is undergoing (2021) an evolution of their API standards. The [OGC API](https://ogcapi.org) effort is focused on lowering the barrier and being more developer friendly. Key OGC API principles include (but are not limited to):
#
# - [W3C Spatial Data on the Web Best Practices](https://www.w3.org/TR/sdw-bp)
# - make APIs more 'of the web'/webby
# - use of JSON and HTML
# - [OpenAPI](https://github.com/OAI/OpenAPI-Specification)
# - **Resource** oriented
# - REST patterns:
# - `GET /foo/bar, not GET /foo request=bar`
# - ease of implementation for a wide audience of web developers
#
# OGC API efforts focus on key resource types:
#
# |OGC first generation | OGC API |
# |------------|---------|
# |OGC OWS Common | OGC API - Common|
# |OGC:WFS | OGC API - Features|
# |OGC:WCS | OGC API - Coverages|
# |OGC:CSW | OGC API - Records|
# |OGC:WPS | OGC API - Processes|
# |OGC:WMS | OGC API - Maps|
# |OGC:WMTS | OGC API - Tiles|
# |OGC:SLD | OGC API - Styles|
# While development of these standards is ongoing, OGC API - Features formally adopted. The [OGC API Roadmap](https://ogcapi.ogc.org/apiroadmap.html) provides a timeline and schedule for the development of the various APIs.
#
# While OGC API is not limited to Python and is programming language agnostic, Python's natural support for JSON makes things even easier to work with the new OGC standards.
#
# Let's interact with an OGC API - Features server:
# # Using Python requests
# +
import json
import requests
url = 'https://demo.pygeoapi.io/master'
# get the root service page
print(json.dumps(requests.get(url).json(), indent=4))
# -
# see all collections
print(json.dumps(requests.get(f'{url}/collections').json(), indent=4))
# grab features from a specific collections
print(json.dumps(requests.get(f'{url}/collections/obs/items').json(), indent=4))
# # Using OWSLib
# Let's try OWSLib's OGC API - Features support
# +
from owslib.ogcapi.features import Features
w = Features(url)
w.links
# -
w.url
w.collections()
features = w.collection_items('obs')
features['numberMatched']
features['numberReturned']
len(features['features'])
features
# # Using a web browser
# Check out the OGC API - Features server in your [browser](https://demo.pygeoapi.io/master). Also take note of the [Swagger API page](https://demo.pygeoapi.io/master/openapi?f=html) which allows developers easy access and perusal of various API functionality, including testing the API as part of the interface. Wow!
# # Using QGIS
#
# QGIS currently (2021) supports the following OGC API standards:
#
# - OGC API - Features
# - OGC API - Records
# [<- Remote data](10-remote-data.ipynb) | [Conclusion ->](12-conclusion.ipynb)
| workshop/jupyter/content/notebooks/11-emerging-technology-trends.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
# %matplotlib inline
import joblib as jb
import torch
from torch.nn import functional as F
from torch import nn
import json
import dask
import itertools
import joblib
import time
import tqdm
from dask.diagnostics import ProgressBar
ProgressBar().register()
# -
train = pd.read_parquet("./data/train.parquet")
test = pd.read_parquet("./data/test.parquet")
train.head()
train.shape, test.shape
# +
item_data = pd.read_parquet("./data/item_data.parquet")
item_title_map = item_data[['title', 'item_id']].set_index("item_id").squeeze().to_dict()
item_data.sample(10)
# -
# # data
class Dataset(torch.utils.data.Dataset):
def __init__(self, item_data):
self.item_data = item_data
def __len__(self):
return self.item_data.shape[0]
def __getitem__(self, index):
title = self.item_data.iloc[index]['title']
return title
# +
from sentence_transformers import SentenceTransformer, SentencesDataset, InputExample
from torch.utils.data import DataLoader
pretrained = 'neuralmind/bert-large-portuguese-cased'
model = SentenceTransformer(pretrained, device='cuda')
train_data = Dataset(item_data)
train_loader = DataLoader(train_data, batch_size=2048)
# -
# %%time
embs_list = list()
for data in tqdm.tqdm(train_loader):
embs = model.encode(data)
embs_list.append(embs)
embs_np = np.vstack(embs_list)
joblib.dump(embs_np, "22a_embs_np.pkl.z")
import nmslib
# %%time
index = nmslib.init(method='hnsw', space='cosinesimil')
index.addDataPointBatch(data=embs_np, ids=item_data['item_id'].values)
index.createIndex(print_progress=True)
item_emb_map = {t: embs_np[i] for i, t in enumerate(item_data['item_id'].values)}
recall = 0
hs = list()
for elist, t in tqdm.tqdm(train[['user_history', 'item_bought']].values):
elist = json.loads(elist)
rep = list()
for e in elist:
if isinstance(e['event_info'], int):
rep.append(item_emb_map[e['event_info']])
#print(item_title_map[e['event_info']])
h = np.mean(rep, axis=0)
#hs.append(h)
#h = rep[0]
#t = item_emb_map[t]
#print()
try:
k = index.knnQuery(h, k=50)
recall += int(t in set(k[0]))
except:
continue
#for i,d in zip(k[0], k[1]):
# print(d, item_title_map[i])
#print(recall)
#print(int(t in k[0]))
#print()
#print(item_title_map[t])
#print("-"*10+"\n"*5)
print(recall/train.shape[0])
# recall@10 - 0.13778097264275843
# recall@20 - 0.15457821731374785
# recall@100 - 0.18157240604797623
# recall@1000 - 0.18950632074992194
# recall cs = viewed - 0.29388401187908886
index.saveIndex("22a_sbert_neuralmind.nms")
# # search
class Dataset(torch.utils.data.Dataset):
def __init__(self, search_data):
self.search_data = search_data
def __len__(self):
return len(self.search_data)
def __getitem__(self, index):
seq_index = self.search_data[index][0]
search = self.search_data[index][1]
#print(search)
return seq_index, search
# #%%time
search_data = set()
seq_index = 0
for hist, bought in tqdm.tqdm(train[['user_history', 'item_bought']].values):
for item in json.loads(hist):
i = item['event_info']
if item['event_type'] == 'search':
search_data.add((seq_index, i.lower()))
seq_index += 1
search_data = list(search_data)
# +
from sentence_transformers import SentenceTransformer, SentencesDataset, InputExample
from torch.utils.data import DataLoader
pretrained = 'neuralmind/bert-large-portuguese-cased'
model = SentenceTransformer(pretrained, device='cuda')
train_data = Dataset(search_data)
train_loader = DataLoader(train_data, batch_size=2048)
# +
#seq_index_embs_map = np.zeros((train.shape[0], 1024))
res = list()
for seq_ix, search in tqdm.tqdm(train_loader):
#print(seq_i
#print(search_list)
emb = model.encode(search)
seq_ix = seq_ix.numpy()
for i in range(emb.shape[0]):
res.append((seq_ix[i], emb[i, :]))
# +
from collections import Counter
ctr = Counter([e[0] for e in res])
seq_index_embs_map = np.zeros((train.shape[0], 1024))
for seqix, emb in tqdm.tqdm(res):
seq_index_embs_map[seqix, :] += emb
for i in tqdm.tqdm(range(train.shape[0])):
seq_index_embs_map[i, :] /= ctr.get(i, 1)
# -
joblib.dump(seq_index_embs_map, "22a_embs_search_np.pkl.z")
# # teste
# #%%time
search_data = set()
seq_index = 0
for hist in tqdm.tqdm(test['user_history'].values):
for item in json.loads(hist):
i = item['event_info']
if item['event_type'] == 'search':
search_data.add((seq_index, i.lower()))
seq_index += 1
search_data = list(search_data)
# +
from sentence_transformers import SentenceTransformer, SentencesDataset, InputExample
from torch.utils.data import DataLoader
pretrained = 'neuralmind/bert-large-portuguese-cased'
model = SentenceTransformer(pretrained, device='cuda')
test_data = Dataset(search_data)
test_loader = DataLoader(test_data, batch_size=2048)
# +
#seq_index_embs_map = np.zeros((train.shape[0], 1024))
res = list()
for seq_ix, search in tqdm.tqdm(test_loader):
#print(seq_i
#print(search_list)
emb = model.encode(search)
seq_ix = seq_ix.numpy()
for i in range(emb.shape[0]):
res.append((seq_ix[i], emb[i, :]))
# +
from collections import Counter
ctr = Counter([e[0] for e in res])
seq_index_embs_map = np.zeros((test.shape[0], 1024))
for seqix, emb in tqdm.tqdm(res):
seq_index_embs_map[seqix, :] += emb
for i in tqdm.tqdm(range(test.shape[0])):
seq_index_embs_map[i, :] /= ctr.get(i, 1)
# -
joblib.dump(seq_index_embs_map, "22a_embs_search_test_np.pkl.z")
| 1a_prep_sbert_neuralmind.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# Select events from a catalogue based on the fault they're on
# + pycharm={"name": "#%%\n"}
#import relevant modules
from rsqsim_api.catalogue.catalogue import RsqSimCatalogue
from rsqsim_api.fault.multifault import RsqSimMultiFault
import fnmatch
import os
# + pycharm={"name": "#%%\n"}
# Tell python where field paths etc are relative to
script_dir = os.path.abspath('')
fault_dir = "../../../data/shaw2021/rundir5091"
catalogue_dir = fault_dir
outdir=os.path.join(catalogue_dir,"by_fault")
# + [markdown] pycharm={"name": "#%% md\n"}
# Read in faults and earthquake catalogue. The files are quite large and can be downloaded from https://doi.org/10.5281/zenodo.5534462
# The models have a spin up period so it's best to remove the first ~2e12s (60kyr) - see below.
# + pycharm={"name": "#%%\n"}
fault_model = RsqSimMultiFault.read_fault_file_bruce(os.path.join(script_dir, fault_dir, "zfault_Deepen.in"),
os.path.join(script_dir, fault_dir, "znames_Deepen.in"),
transform_from_utm=True)
whole_catalogue = RsqSimCatalogue.from_catalogue_file_and_lists(os.path.join(catalogue_dir, "eqs..out"),
list_file_directory=catalogue_dir, list_file_prefix="catalog")
# + [markdown] pycharm={"name": "#%% md\n"}
# Remove first 2e12s
# + pycharm={"name": "#%%\n"}
no_spin_up_cat=whole_catalogue.filter_whole_catalogue(min_t0=2.0e12)
# + [markdown] pycharm={"name": "#%% md\n"}
# Select faults of interest (the possible fault names can be listed using fault_model.names)
# + pycharm={"name": "#%%\n"}
#e.g. single fault
faultName="alpinef2k"
fault_selection1=RsqSimMultiFault([fault_model.name_dic[faultName]])
#e.g. all faults with a particular string in the name
baseFault="hope"
faults2select = [name for name in fault_model.names if fnmatch.fnmatch(name, baseFault+"*")]
fault_selection2=RsqSimMultiFault([fault_model.name_dic[name] for name in faults2select])
#e.g. a list of faults
faultNames=["fidget","jorkekneed","clarencene"]
fault_selection3=RsqSimMultiFault([fault_model.name_dic[name] for name in faultNames])
#check these have worked correctly
#fault_selection1.names
# + [markdown] pycharm={"name": "#%% md\n"}
# Extract events on these faults.
# + pycharm={"name": "#%%\n"}
event_selection1=no_spin_up_cat.filter_by_fault(fault_selection1, minimum_patches_per_fault=1)
event_selection2=no_spin_up_cat.filter_by_fault(fault_selection2, minimum_patches_per_fault=1)
event_selection3=no_spin_up_cat.filter_by_fault(fault_selection3, minimum_patches_per_fault=1)
# + [markdown] pycharm={"name": "#%% md\n"}
# Write out to new catalogues.
# + pycharm={"name": "#%%\n"}
#first make sure output directory exists
if not os.path.exists(outdir):
os.mkdir(outdir)
#write out
event_selection1.write_csv_and_arrays(prefix=faultName,directory=outdir)
event_selection2.write_csv_and_arrays(prefix=baseFault+"_all",directory=outdir)
#might want to change naming convention here especially for a long list
#currently just concatenates all list elements to get prefix
event_selection3.write_csv_and_arrays(prefix=''.join(faultNames[:]),directory=outdir)
# + [markdown] pycharm={"name": "#%% md\n"}
# Example of how to read one of these catalogues back in:
# + pycharm={"name": "#%%\n"}
alp_cat=RsqSimCatalogue.from_csv_and_arrays(prefix=os.path.join(outdir,faultName))
| examples/rsqsim_api/read_and_filter_catalogue/find_events_by_fault.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Z-UQUOMnhf2n" executionInfo={"status": "ok", "timestamp": 1616314892923, "user_tz": -480, "elapsed": 2293, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="f4a80421-f859-49df-f70c-57e810a5bddd"
from google.colab import drive
drive.mount('/content/drive')
# %cd /content/drive/My Drive/SQuAD/squad
# + [markdown] id="p6k2u9pqAlFs"
# # Data loading and package installation
# + colab={"base_uri": "https://localhost:8080/"} id="tJdT0Vglg5bs" executionInfo={"status": "ok", "timestamp": 1615956828473, "user_tz": -480, "elapsed": 4901, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="a69a0bb0-9fc0-4f75-ef7b-8073753ef24c"
# # !git clone https://github.com/minggg/squad.git
# + colab={"base_uri": "https://localhost:8080/"} id="Vln1hl-4iJl3" executionInfo={"status": "ok", "timestamp": 1615957118798, "user_tz": -480, "elapsed": 6906, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="eebe565d-d22a-4025-e4f5-092fefaf584d"
# # !pip install spacy
# # !pip install ujson
# + colab={"base_uri": "https://localhost:8080/"} id="tIj7-F1cjP3X" executionInfo={"status": "ok", "timestamp": 1615958291179, "user_tz": -480, "elapsed": 994957, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="28c975e6-6bc8-44d5-c98e-fd42c83a7f46"
# # !python setup.py
# + [markdown] id="yNvEmchoL_Pk"
# # read CS224 squad data json files
# + [markdown] id="ojXByuKFrr_d"
# The dev + test set here = full development set in official SQuAD data
# + colab={"base_uri": "https://localhost:8080/"} id="NSIxBnA-Y1yd" executionInfo={"status": "ok", "timestamp": 1616308442201, "user_tz": -480, "elapsed": 2490, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="d0d54486-8736-4f37-9893-52c190cdb76f"
import os, json
import pandas as pd
path_to_json = '/content/drive/My Drive/SQuAD/squad/data/'
json_files = [pos_json for pos_json in os.listdir(path_to_json) if pos_json.endswith('.json')]
print(json_files)
# + id="zuIgdnvIaW93"
# # we need both the json and an index number so use enumerate()
# for index, js in enumerate(json_files):
# with open(os.path.join(path_to_json, js)) as json_file:
# json_text = json.load(json_file)
# + id="IZFlVIUUY9ok"
# dev = pd.DataFrame.from_dict(many_jsons[0])
# test = pd.DataFrame.from_dict(many_jsons[1])
# train = pd.DataFrame.from_dict(many_jsons[2])
# test_eval = pd.DataFrame.from_dict(many_jsons[3])
# test_meta = pd.DataFrame.from_dict(many_jsons[4])
# word_emb = pd.DataFrame.from_dict(many_jsons[5])
# char_emb= pd.DataFrame.from_dict(many_jsons[6])
# train_eval= pd.DataFrame.from_dict(many_jsons[7])
# dev_eval= pd.DataFrame.from_dict(many_jsons[8])
# word2idx= pd.DataFrame.from_dict(many_jsons[9])
# char2idx= pd.DataFrame.from_dict(many_jsons[10])
# dev_meta= pd.DataFrame.from_dict(many_jsons[11])
# print montreal_json['features'][0]['geometry']
# + id="kdqkB_ojtD4X" executionInfo={"status": "ok", "timestamp": 1616308449422, "user_tz": -480, "elapsed": 3917, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# load train
import json
# Opening JSON file
f = open('/content/drive/My Drive/SQuAD/squad/data/train-v2.0.json',)
# returns JSON object as
# a dictionary
train = json.load(f)
# Closing file
f.close()
# + id="BYD7suE_Ebtv" executionInfo={"status": "ok", "timestamp": 1616308451802, "user_tz": -480, "elapsed": 2589, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# load dev
import json
# Opening JSON file
f = open('/content/drive/My Drive/SQuAD/squad/data/dev-v2.0.json',)
# returns JSON object as
# a dictionary
dev = json.load(f)
# Closing file
f.close()
# + id="SHJ6WbjVEvIg" executionInfo={"status": "ok", "timestamp": 1616308460155, "user_tz": -480, "elapsed": 1608, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# load dev_meta
import json
# Opening JSON file
f = open('/content/drive/My Drive/SQuAD/squad/data/dev_meta.json',)
# returns JSON object as
# a dictionary
dev_meta = json.load(f)
# Closing file
f.close()
# + id="55NbQDNCFBMu" executionInfo={"status": "ok", "timestamp": 1616308465953, "user_tz": -480, "elapsed": 5748, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# load dev_eval
import json
# Opening JSON file
f = open('/content/drive/My Drive/SQuAD/squad/data/dev_eval.json',)
# returns JSON object as
# a dictionary
dev_eval = json.load(f)
# Closing file
f.close()
# + id="6cXyQOEZCOgE"
# import pandas as pd
# train_df = pd.read_json (r'/content/drive/My Drive/SQuAD/squad/data/train-v2.0.json')
# + id="N9xNBwYJCWDA"
pd.options.display.max_rows = 10
# train_df
# + colab={"base_uri": "https://localhost:8080/"} id="gRzF2vcWjmhV" executionInfo={"status": "ok", "timestamp": 1616308465954, "user_tz": -480, "elapsed": 2038, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="66142993-e12e-4b0c-d0df-452a26cbda71"
print(train.keys(),dev.keys(),dev_meta.keys(),dev_eval.keys())
# + [markdown] id="-zCmhmK5MQk8"
# ## examine json dictionaries
# + colab={"base_uri": "https://localhost:8080/"} id="zkl92xwufjTL" executionInfo={"status": "ok", "timestamp": 1616308489206, "user_tz": -480, "elapsed": 1127, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="7645e432-67a1-4297-fb79-ee096bd96353"
type(train)
# + colab={"base_uri": "https://localhost:8080/"} id="XwV-c9YBfnqD" executionInfo={"status": "ok", "timestamp": 1616308510219, "user_tz": -480, "elapsed": 770, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="d97835ea-5e02-414e-d910-22364f0e74a7"
type(train['data'])
# + colab={"base_uri": "https://localhost:8080/"} id="d7JWZyjkgTgC" executionInfo={"status": "ok", "timestamp": 1616308686064, "user_tz": -480, "elapsed": 835, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="a01f9e1c-3795-4613-c325-b27fe8f8555c"
type(train['data'][0])
# + colab={"base_uri": "https://localhost:8080/"} id="KIFNfnTNgsfj" executionInfo={"status": "ok", "timestamp": 1616308816194, "user_tz": -480, "elapsed": 849, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="344f92b9-794d-4b6b-8f5d-1b1366007dc9"
len(train['data'])*0.05
# + colab={"base_uri": "https://localhost:8080/"} id="dIZ-Sflj4XCH" executionInfo={"status": "ok", "timestamp": 1616315014141, "user_tz": -480, "elapsed": 783, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="bcef49ce-9f06-494c-df5f-6aaca1297ad9"
type(train['data'][0]['paragraphs'])
# + colab={"base_uri": "https://localhost:8080/"} id="Iq6jUnsXfzA1" executionInfo={"status": "ok", "timestamp": 1616308602009, "user_tz": -480, "elapsed": 1092, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="7fcec119-84df-47ea-a4a9-f928fc322885"
train['data'][1]
# + colab={"base_uri": "https://localhost:8080/"} id="hYaOH9ukFJFr" executionInfo={"status": "ok", "timestamp": 1616050004288, "user_tz": -480, "elapsed": 726, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="6430721f-09f0-4dd6-abf4-810b4341d1c2"
dev_meta
# + colab={"base_uri": "https://localhost:8080/"} id="hadv6F8nFnhi" executionInfo={"status": "ok", "timestamp": 1616050057140, "user_tz": -480, "elapsed": 729, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="50ac4514-3a40-4327-dbba-17a07bd39f78"
# eval_examples[str(total)] = {"context": context,
# "question": ques,
# "spans": spans,
# "answers": answer_texts,
# "uuid": qa["id"]}
dev_eval['1']
# + colab={"base_uri": "https://localhost:8080/"} id="uL7VsgsqFwMq" executionInfo={"status": "ok", "timestamp": 1616050077220, "user_tz": -480, "elapsed": 733, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="b639ac53-7174-4df2-fc5c-5cbb1598d964"
dev['data'][0]['paragraphs'][0]
# + colab={"base_uri": "https://localhost:8080/"} id="E6JlUYe7lms0" executionInfo={"status": "ok", "timestamp": 1616052399730, "user_tz": -480, "elapsed": 740, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="68f8d708-d360-4bd5-89dd-7255096b6f9d"
train['data'][0]['paragraphs'][0]['qas']
# + colab={"base_uri": "https://localhost:8080/"} id="qeKSbxweuCHZ" executionInfo={"status": "ok", "timestamp": 1616051851838, "user_tz": -480, "elapsed": 2371, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="c090e1b8-a547-494c-a72e-1f7b6940e436"
import pprint
i=0
j = 0
total_qa = 0
total_context = 0
for article in train['data']:
if i < 1:
pprint.pprint(article) # pretty print 1 article
i+=1
for paragraph in article['paragraphs']:
if j < 1:
pprint.pprint(paragraph) # pretty print 1 paragraph/context in that article with many questions
j+=1
print(len(paragraph['context'])) # inside 'context', there is only one word string, so length of 'context' returns the character length of the string
print(len(paragraph['qas'])) # inside 'qas', there are multiple questions, so length returns the number of questions
total_qa += len(paragraph['qas'])
total_context += len(paragraph['context'])
# + [markdown] id="GZstmkeHMY0G"
# ## basic stats abt train data
# + id="pU1vTQbenYeD" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1616051855862, "user_tz": -480, "elapsed": 897, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="55f7170f-b5d9-4bbe-b26e-393cccda37de"
print("total number of questions =",total_qa)
print('total number of characters of contexts =',total_context)
print('total number of articles',len(train['data']))
# + colab={"base_uri": "https://localhost:8080/"} id="iBJPqnJJK3Fj" executionInfo={"status": "ok", "timestamp": 1616051857803, "user_tz": -480, "elapsed": 828, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="585fc6f0-2371-43a5-8bff-14b88c768da9"
import pprint
i = 0
j = 0
for article in train['data']:
i+=1
for paragraph in article['paragraphs']:
j+=1
print("total number of articles =",i)
print("total number of contexts / paragraphs =",j)
# + [markdown] id="MJPPgTllMpBU"
# # data preprocessing
# + [markdown] id="W0Np36gO79qF"
# # # ## !Create a subset of data 5%
# + colab={"base_uri": "https://localhost:8080/"} id="CwxxVOLH9bp_" executionInfo={"status": "ok", "timestamp": 1616316320819, "user_tz": -480, "elapsed": 843, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="ce0df7cf-2d13-4636-9e8d-b526cb23dc9a"
type(train['data'][0]['paragraphs'])
# + colab={"base_uri": "https://localhost:8080/"} id="JpVomXn_4nUp" executionInfo={"status": "ok", "timestamp": 1616316392533, "user_tz": -480, "elapsed": 1067, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="4ece50d7-6710-4b4e-cfc1-f5b4b6510ed1"
int(round(len(train['data'][0]['paragraphs'])*0.05,0))
# + id="RH3ysU_56oLn" executionInfo={"status": "ok", "timestamp": 1616316601794, "user_tz": -480, "elapsed": 787, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
import random
random.seed(1)
context_sample = random.sample(train['data'][0]['paragraphs'],int(round(len(train['data'][0]['paragraphs'])*0.05,0)))
# context_sample
# + colab={"base_uri": "https://localhost:8080/"} id="Iz-WGMahgLNR" executionInfo={"status": "ok", "timestamp": 1616317157427, "user_tz": -480, "elapsed": 863, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="46c78ba7-f13c-4d62-8417-3b0de0686b56"
count_para = 0
para_context = []
train_05 = {'version':train['version'],'data':[]}
for paragraph in train['data']:
count_para +=1
para_context.append(len(paragraph['paragraphs']))
context_sample = random.sample(paragraph['paragraphs'],int(round(len(paragraph['paragraphs'])*0.05,0)))
new_paragraph = {'paragraphs':context_sample}
train_05['data'].append(new_paragraph)
print(sum(para_context))
# + id="lu5LhnEeAp5h" executionInfo={"status": "ok", "timestamp": 1616317397677, "user_tz": -480, "elapsed": 780, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# train_05
# + colab={"base_uri": "https://localhost:8080/"} id="AnD-xyIRBlHx" executionInfo={"status": "ok", "timestamp": 1616318280798, "user_tz": -480, "elapsed": 843, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="434f21ec-728e-49ad-eaa6-2c7c01e0054e"
count_para_05 = 0
para_context_05 = []
for paragraph in train_05['data']:
count_para_05 +=1
para_context_05.append(len(paragraph['paragraphs']))
print(sum(para_context_05),"contexts, which is",sum(para_context_05)/sum(para_context),'of original data')
# + id="a6c2pT9PCSEB" executionInfo={"status": "ok", "timestamp": 1616317670475, "user_tz": -480, "elapsed": 837, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
import json
with open("/content/drive/My Drive/SQuAD/train_05.json", "w") as write_file:
json.dump(train_05, write_file)
# + id="yFa_stLFCeYb" executionInfo={"status": "ok", "timestamp": 1616317994413, "user_tz": -480, "elapsed": 1206, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# load train
import json
# Opening JSON file
f = open('/content/drive/My Drive/SQuAD/squad/train_05.json',)
# returns JSON object as
# a dictionary
train_05_load = json.load(f)
# Closing file
f.close()
# + id="aKE-w4iBD17n" executionInfo={"status": "ok", "timestamp": 1616318061865, "user_tz": -480, "elapsed": 765, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
# train_05_load['data'][0]['paragraphs'][2]
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="_KcE5qMoBkps" executionInfo={"status": "ok", "timestamp": 1616318114629, "user_tz": -480, "elapsed": 1866, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="2355cd6a-4453-4610-fa11-55c2da600c36"
import matplotlib.pyplot as plt
import seaborn as sns
# Show 4 different binwidths
for i, binwidth in enumerate([1, 5, 10, 15]):
# Set up the plot
ax = plt.subplot(2, 2, i + 1)
# Draw the plot
ax.hist(para_context_05, bins = int(180/binwidth),
color = 'blue', edgecolor = 'black')
# Title and labels
ax.set_title('Histogram with Binwidth = %d' % binwidth)
ax.set_xlabel('Number of Context per Article')
ax.set_ylabel('Count of Articles')
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="s52OJZHlCOhY" executionInfo={"status": "ok", "timestamp": 1616317576619, "user_tz": -480, "elapsed": 1883, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="0a2b6845-7e1c-401f-e758-09b909b6d697"
import matplotlib.pyplot as plt
import seaborn as sns
# Show 4 different binwidths
for i, binwidth in enumerate([1, 5, 10, 15]):
# Set up the plot
ax = plt.subplot(2, 2, i + 1)
# Draw the plot
ax.hist(para_context, bins = int(180/binwidth),
color = 'blue', edgecolor = 'black')
# Title and labels
ax.set_title('Histogram with Binwidth = %d' % binwidth)
ax.set_xlabel('Number of Context per Article')
ax.set_ylabel('Count of Articles')
plt.tight_layout()
plt.show()
# + [markdown] id="upuwshyxEmfj"
# # # ## !Create a subset of data 10%
# + colab={"base_uri": "https://localhost:8080/"} id="LIh8fQNHEmp4" executionInfo={"status": "ok", "timestamp": 1616318257864, "user_tz": -480, "elapsed": 1076, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="1011ec51-8ddd-422f-b5b7-badafe94b5de"
count_para = 0
para_context = []
train_10 = {'version':train['version'],'data':[]}
for paragraph in train['data']:
count_para +=1
para_context.append(len(paragraph['paragraphs']))
context_sample = random.sample(paragraph['paragraphs'],int(round(len(paragraph['paragraphs'])*0.1,0)))
new_paragraph = {'paragraphs':context_sample}
train_10['data'].append(new_paragraph)
print(sum(para_context))
# + colab={"base_uri": "https://localhost:8080/"} id="QvTVaUfnEmp5" executionInfo={"status": "ok", "timestamp": 1616318272355, "user_tz": -480, "elapsed": 927, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="d7d5e91c-a417-412d-95cc-e09012d2416f"
count_para_10 = 0
para_context_10 = []
for paragraph in train_10['data']:
count_para_10 +=1
para_context_10.append(len(paragraph['paragraphs']))
print(sum(para_context_10),"contexts, which is",sum(para_context_10)/sum(para_context),'of original data')
# + id="eTpuxaCuEmp5" executionInfo={"status": "ok", "timestamp": 1616318292952, "user_tz": -480, "elapsed": 1126, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}}
import json
with open("/content/drive/My Drive/SQuAD/train_10.json", "w") as write_file:
json.dump(train_10, write_file)
# + [markdown] id="Ydlu7gTqM18p"
# ## convert json to dataframe
# + id="syrbm6V8vZcf" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1616053547596, "user_tz": -480, "elapsed": 768, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="c3465acb-83a4-4296-fd0f-e3eae1c31144"
# https://www.kaggle.com/sanjay11100/squad-stanford-q-a-json-to-pandas-dataframe
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import json # to read json
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
print(check_output(["ls", "/content/drive/My Drive/SQuAD/official_data"]).decode("utf8"))
# + id="ib2j2S8hM8_7"
# defined for train-v2.0.json
def squad_json_to_dataframe_train(input_file_path, record_path = ['data','paragraphs','qas','answers'],verbose = 1):
"""
input_file_path: path to the squad json file.
record_path: path to deepest level in json file default value is
['data','paragraphs','qas','answers']
verbose: 0 to suppress it default is 1
"""
if verbose:
print("Reading the json file")
file = json.loads(open(input_file_path).read())
if verbose:
print("processing...")
# parsing different level's in the json file
js = pd.io.json.json_normalize(file , record_path )
m = pd.io.json.json_normalize(file, record_path[:-1])
r = pd.io.json.json_normalize(file,record_path[:-2])
# l = pd.io.json.json_normalize(file,record_path[:-3])
#combining it into single dataframe
ndx = np.repeat(m['id'].values,m['answers'].str.len())
idx = np.repeat(r['context'].values, r.qas.str.len())
# rdx = np.repeat(l['paragraphs'].values, l.paragraphs.str.len())
m['context'] = idx
js['q_idx'] = ndx
# r['paragraphs'] = rdx
main = pd.concat([ m[['id','question','context','is_impossible']].set_index('id'),js.set_index('q_idx') ],1,sort=False).reset_index()
main['context_id'] = main['context'].factorize()[0]
# main['paragraph_id'] = main['paragraphs'].factorize()[0]
if verbose:
print("shape of the dataframe is {}".format(main.shape))
print("Done")
return main
# + colab={"base_uri": "https://localhost:8080/"} id="VIw1-63zP6O3" executionInfo={"status": "ok", "timestamp": 1616065081766, "user_tz": -480, "elapsed": 9800, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="5d9b6949-b12b-4124-b90e-c5d113731e88"
# training data
input_file_path = '/content/drive/My Drive/SQuAD/official_data/train-v2.0.json'
record_path = ['data','paragraphs','qas','answers']
train_df = squad_json_to_dataframe_train(input_file_path=input_file_path,record_path=record_path)
# + colab={"base_uri": "https://localhost:8080/", "height": 580} id="j4aLqwUg2HZW" executionInfo={"status": "ok", "timestamp": 1616065081768, "user_tz": -480, "elapsed": 8558, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="a181326e-b7d1-47b2-88f9-2d42717c9370"
train_df
# + colab={"base_uri": "https://localhost:8080/", "height": 407} id="Er-uWMvVQjSW" executionInfo={"status": "ok", "timestamp": 1616065476896, "user_tz": -480, "elapsed": 784, "user": {"displayName": "Akshay Arun", "photoUrl": "", "userId": "07298175230978186476"}} outputId="abcf549b-c1f3-47d9-ee21-b505c4f83fb9"
train_df[train_df.is_impossible == True]
# + id="H2tiIM83Pwjn"
def squad_json_to_dataframe_dev(input_file_path, record_path = ['data','paragraphs','qas','answers'],
verbose = 1):
"""
input_file_path: path to the squad json file.
record_path: path to deepest level in json file default value is
['data','paragraphs','qas','answers']
verbose: 0 to suppress it default is 1
"""
if verbose:
print("Reading the json file")
file = json.loads(open(input_file_path).read())
if verbose:
print("processing...")
# parsing different level's in the json file
js = pd.io.json.json_normalize(file , record_path )
m = pd.io.json.json_normalize(file, record_path[:-1] )
r = pd.io.json.json_normalize(file,record_path[:-2])
#combining it into single dataframe
idx = np.repeat(r['context'].values, r.qas.str.len())
# ndx = np.repeat(m['id'].values,m['answers'].str.len())
m['context'] = idx
# js['q_idx'] = ndx
main = m[['id','question','context','answers','is_impossible']].set_index('id').reset_index()
main['context_id'] = main['context'].factorize()[0]
if verbose:
print("shape of the dataframe is {}".format(main.shape))
print("Done")
return main
# + colab={"base_uri": "https://localhost:8080/"} id="5wre8R8FM887" executionInfo={"status": "ok", "timestamp": 1616065485106, "user_tz": -480, "elapsed": 1474, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="f5e0227e-9099-4a3a-cfd3-cfa1ec6751b2"
# dev data
input_file_path = '/content/drive/My Drive/SQuAD/official_data/dev-v2.0.json'
record_path = ['data','paragraphs','qas','answers']
verbose = 0
dev_df = squad_json_to_dataframe_dev(input_file_path=input_file_path,record_path=record_path)
# + colab={"base_uri": "https://localhost:8080/", "height": 407} id="M3wiM87XM86Q" executionInfo={"status": "ok", "timestamp": 1616065485960, "user_tz": -480, "elapsed": 567, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="fab2369e-ed45-45e3-ee0b-94e326635cea"
dev_df[dev_df.is_impossible==True]
# + [markdown] id="1WobzZ2eX9Un"
# # EDA
# + [markdown] id="IeSk5mn0TTn9"
# ## Unanswerable questions count
# **In Training set, out of 130,319 questions, 86,821 questions are answerable.**
#
# **In Development set, out of 11,873 questions, 5,928 questions are answerable**
# + colab={"base_uri": "https://localhost:8080/"} id="K9K51Tt-TFGA" executionInfo={"status": "ok", "timestamp": 1616053571710, "user_tz": -480, "elapsed": 761, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="1f068718-91cb-4fab-ccd4-4b8ae939f444"
train_df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 138} id="CftNaslEUOiR" executionInfo={"status": "ok", "timestamp": 1616053865702, "user_tz": -480, "elapsed": 749, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="f135823d-73f5-45b2-db3c-79adf65a7ccc"
train_df.groupby('is_impossible').count()
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="dsDnbZwfcyfS" executionInfo={"status": "ok", "timestamp": 1616056203532, "user_tz": -480, "elapsed": 916, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="cc944dc7-22fd-4569-bb4f-bbe87ac372fe"
train_df['is_impossible'].value_counts().plot(kind='barh',title='Count of Inanswerable Questions in Training Data')
# + colab={"base_uri": "https://localhost:8080/"} id="IUfSJ-YDTdmb" executionInfo={"status": "ok", "timestamp": 1616053660961, "user_tz": -480, "elapsed": 747, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="b50bc154-a5cb-4ca4-ea80-960603c6ad4c"
dev_df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 138} id="qfAmSS5CSy5y" executionInfo={"status": "ok", "timestamp": 1616053844684, "user_tz": -480, "elapsed": 828, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="a7d6822a-f319-413d-c8c0-dba162a7f682"
dev_df.groupby('is_impossible').count()
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="g_Kv8616dFh3" executionInfo={"status": "ok", "timestamp": 1616056207845, "user_tz": -480, "elapsed": 895, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="3ff520a8-ea82-4042-e1b1-70beca053ef5"
dev_df['is_impossible'].value_counts().plot(kind='barh',title='Count of Inanswerable Questions in Dev Data')
# + [markdown] id="LZudWek3Y_-M"
# ## missing value
# **No missing values other than the unanswerable ones in training dataset**
# + colab={"base_uri": "https://localhost:8080/"} id="wCng8KjBYDZ4" executionInfo={"status": "ok", "timestamp": 1616055098378, "user_tz": -480, "elapsed": 1115, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="e729ab8a-8305-4954-d189-cb0da9c47600"
train_df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="kOcMrgXXY-Ai" executionInfo={"status": "ok", "timestamp": 1616055102667, "user_tz": -480, "elapsed": 821, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="7917d915-e790-4f35-ce9a-71f7649f0007"
dev_df.isnull().sum()
# + [markdown] id="19frL78P8nhQ"
# # # ## ! Paragraph-Context Count
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="4Fiw4xjG69JK" executionInfo={"status": "ok", "timestamp": 1616315844469, "user_tz": -480, "elapsed": 1728, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="4fb07031-5ed3-41e2-b18c-b7f7a146a476"
import matplotlib.pyplot as plt
import seaborn as sns
# Show 4 different binwidths
for i, binwidth in enumerate([1, 5, 10, 15]):
# Set up the plot
ax = plt.subplot(2, 2, i + 1)
# Draw the plot
ax.hist(para_context, bins = int(180/binwidth),
color = 'blue', edgecolor = 'black')
# Title and labels
ax.set_title('Histogram with Binwidth = %d' % binwidth)
ax.set_xlabel('Number of Context per Article')
ax.set_ylabel('Count of Articles')
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="8B7CH4XK5Gab" executionInfo={"status": "ok", "timestamp": 1616315190488, "user_tz": -480, "elapsed": 772, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="9e350a3c-333e-48d0-eee6-8c50915bd4f0"
para_context
# + [markdown] id="yprt6o9wbYzi"
# ## Context-Question-Answer Count
# **Dev set has multiple answers for each question, while training set only has one answer for each.**
#
# **For development set, min answer count = 1, max answer count = 6**
#
# **Average number of question-answer pairs per context in training set = 7**
#
# **Average number of questions per context in developmemt set = 10**
#
# **Average number of answers per question in developmemt set = 3**
#
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="vvCYVYrfY_ZZ" executionInfo={"status": "ok", "timestamp": 1616055688110, "user_tz": -480, "elapsed": 795, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="0dfa6beb-79f6-45bc-e984-13ee63992f68"
dev_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="GV4Yd44lbO_w" executionInfo={"status": "ok", "timestamp": 1616055727638, "user_tz": -480, "elapsed": 511, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="3e1f7153-0483-44dc-9e2c-24a030ccee7a"
dev_df.loc[0,'answers']
# + colab={"base_uri": "https://localhost:8080/", "height": 431} id="AS6S46lmeSfe" executionInfo={"status": "ok", "timestamp": 1616065797133, "user_tz": -480, "elapsed": 2510, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="b77f6c48-0b7a-4fe8-ad24-d482c021a640"
dev_df.groupby('context_id').question.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="9cUv_w8gDL_l" executionInfo={"status": "ok", "timestamp": 1616067128893, "user_tz": -480, "elapsed": 885, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="8d86a9ee-00c7-4d5b-a762-178418d5f246"
answer_count = [len(a) for a in dev_df[dev_df.is_impossible==False].answers]
# y = [v * 5 for v in x]
# len(dev_df[dev_df.is_impossible==True].answers)
print("For development set, min answer count =", min(answer_count),"For development set, max answer count =", max(answer_count))
# + colab={"base_uri": "https://localhost:8080/"} id="JdMRth73B5nn" executionInfo={"status": "ok", "timestamp": 1616067560496, "user_tz": -480, "elapsed": 813, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="446d2feb-0ee6-4b45-86d3-7c26bc08d41d"
print("Average number of question-answer pairs per context in training set = {:.0f}".format(train_df.groupby('context_id').question.count().mean()))
print("Average number of questions per context in developmemt set = {:.0f}".format(dev_df.groupby('context_id').question.count().mean()))
print("Average number of answers per question in developmemt set = {:.0f}".format(sum(answer_count)/len(answer_count)))
# + [markdown] id="87u4DRjmdiFm"
# ## Question Type
# + colab={"base_uri": "https://localhost:8080/"} id="zPrbeo5scGK4" executionInfo={"status": "ok", "timestamp": 1616076605362, "user_tz": -480, "elapsed": 17972, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="d94544c2-bd96-4a50-f831-5ed4a24b1513"
q_type = ['where','when','what','which','who','why','how','does','did','has','was','is','are','were']
import time
start_time = time.time()
for i in q_type:
print(i)
# for a in range(len(train_df.question)):
# train_df[i] = train_df.question[a].lower().find(i)+1
train_df[i] = [train_df.question[a].lower().find(i) for a in range(len(train_df.question))]
q_type_count = []
q_type_count.append([len(train_df[train_df[a]!=-1]) for a in q_type])
print("--- %s seconds ---" % (time.time() - start_time))
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="M3HFVCFJV32B" executionInfo={"status": "ok", "timestamp": 1616076632172, "user_tz": -480, "elapsed": 924, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="43da2e06-b674-43ce-8d79-254d41081618"
train_df[q_type]
# + [markdown] id="ja5dIKcDWcG0"
# **399 questions do not include any of above question type key word.**
# **Typos are observed like 'which' and 'wat'**
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="mxpiuCMhPMEO" executionInfo={"status": "ok", "timestamp": 1616076703086, "user_tz": -480, "elapsed": 926, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="e1c8adb5-0031-45c8-f56b-025c0af08fb5"
train_df[train_df[q_type].sum(axis=1)==-len(q_type)]
# + [markdown] id="fu2qPWawVALm"
# **Some questions belong to one type but may contain 2 question type key words**
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="2ic7iPTCU38H" executionInfo={"status": "ok", "timestamp": 1616070824135, "user_tz": -480, "elapsed": 739, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="0ae903e9-fbf5-419e-d68e-7e5717fe71f9"
train_df.question[1]
# + [markdown] id="LJUPSmK5WvsZ"
# **Plot count of each question type, as different question types have different level of difficulties.**
# + colab={"base_uri": "https://localhost:8080/"} id="iYtrSxHPXF01" executionInfo={"status": "ok", "timestamp": 1616076726913, "user_tz": -480, "elapsed": 904, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="a20974f4-0678-408f-bb31-83cbc6ed1ca6"
print(q_type)
# + colab={"base_uri": "https://localhost:8080/"} id="BUIELoXxc3WR" executionInfo={"status": "ok", "timestamp": 1616076727846, "user_tz": -480, "elapsed": 571, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="9255b316-a018-4d59-cffb-6d5a2ab91f7b"
q_type_count[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="zhhJDffyYR-x" executionInfo={"status": "ok", "timestamp": 1616077340310, "user_tz": -480, "elapsed": 1375, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="4a01e480-dd04-43fe-e204-8c9b2b05d473"
import numpy as np
import matplotlib.pyplot as plt
xs = np.arange(len(q_type))
ys = np.arange(0,80000,10000)
width = .5
fig = plt.figure(figsize=[8,8])
ax = fig.gca() #get current axes
ax.bar(xs, q_type_count[0], width, align='center', alpha = 0.5, \
color=['lightblue', 'limegreen', 'orange', 'cornflowerblue',\
'midnightblue','aqua','darkorchid','gold','gold','gold',\
'darkred','darkred','darkred','darkred'])
#Remove the default x-axis tick numbers and
#use tick numbers of your own choosing:
ax.set_xticks(xs)
#Replace the tick numbers with strings:
ax.set_xticklabels(q_type)
#Remove the default y-axis tick numbers and
#use tick numbers of your own choosing:
ax.set_yticks(ys)
labels = q_type_count[0]
def add_value_labels(ax, spacing=5):
"""Add labels to the end of each bar in a bar chart.
Arguments:
ax (matplotlib.axes.Axes): The matplotlib object containing the axes
of the plot to annotate.
spacing (int): The distance between the labels and the bars.
"""
# For each bar: Place a label
for rect in ax.patches:
# Get X and Y placement of label from rect.
y_value = rect.get_height()
x_value = rect.get_x() + rect.get_width() / 2
# Number of points between bar and label. Change to your liking.
space = spacing
# Vertical alignment for positive values
va = 'bottom'
# If value of bar is negative: Place label below bar
if y_value < 0:
# Invert space to place label below
space *= -1
# Vertically align label at top
va = 'top'
# Use Y value as label and format number with one decimal place
label = "{:.1f}".format(y_value)
# Create annotation
ax.annotate(
label, # Use `label` as label
(x_value, y_value), # Place label at end of the bar
xytext=(0, space), # Vertically shift label by `space`
textcoords="offset points", # Interpret `xytext` as offset in points
ha='center', # Horizontally center label
va=va) # Vertically align label differently for
# positive and negative values.
# Call the function above. All the magic happens there.
add_value_labels(ax)
ax.set_xlabel('Key Words for Question Type')
ax.set_ylabel('Number of Questions')
plt.title('Count of question type key words')
plt.savefig('count of question type key words.png')
# + [markdown] id="iKIK2RfHkCrT"
# ## Answer Length
# + colab={"base_uri": "https://localhost:8080/"} id="cco9znUWknxN" executionInfo={"status": "ok", "timestamp": 1616078593341, "user_tz": -480, "elapsed": 864, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="5579cce3-3b18-43c8-88b4-2d28f8c54f69"
from tokenize import tokenize, untokenize, NUMBER, STRING, NAME, OP
split_text= [sub.split() for sub in train_df[train_df.is_impossible==False].text]
split_text
# + id="d-w8bB7VzbIQ"
train_df['split_text'] = ""
# + id="1YlViNAj0rp5"
mask = train_df.is_impossible==False
train_df.loc[mask, 'split_text'] = train_df.loc[mask, 'text'].apply(lambda x: x.split())
# + id="5mh8rfAm5g4q"
train_df['ans_len'] = ""
# + colab={"base_uri": "https://localhost:8080/", "height": 736} id="ICXD55Mc5g41" executionInfo={"status": "ok", "timestamp": 1616081291983, "user_tz": -480, "elapsed": 768, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="37ffd1bd-1d2c-477e-b57e-79c336d2ec81"
mask = train_df.is_impossible==False
train_df.loc[mask, 'ans_len'] = train_df.loc[mask, 'split_text'].apply(lambda x: len(x))
train_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="VVbH_mR557HH" executionInfo={"status": "ok", "timestamp": 1616081373512, "user_tz": -480, "elapsed": 738, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="c9679107-152e-4999-f418-0c9378f8f01d"
# train_df['ans_len'].sum(axis = 0, skipna = True)
print("Average Answer Length = {:.0f}".format(train_df[train_df.is_impossible==False]['ans_len'].mean()))
# + id="jpwOtBl6F7N4"
mask = train_df.is_impossible==False
train_df.loc[mask, 'isnumeric'] = train_df.loc[mask, 'text'].apply(lambda x: x.isnumeric())
# + colab={"base_uri": "https://localhost:8080/"} id="1cUoeLxeG8Tq" executionInfo={"status": "ok", "timestamp": 1616084066948, "user_tz": -480, "elapsed": 838, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="ef298724-1c84-4f06-df5d-f8ef19aad7ee"
print("Number of questions with numeric answers are :",train_df[train_df.isnumeric==True].text.count())
# + [markdown] id="rro5jpLkHe6S"
# **Some answers have numerical meanings, but are written as numbers in english.**
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="FYf9lTICHeiN" executionInfo={"status": "ok", "timestamp": 1616084148125, "user_tz": -480, "elapsed": 765, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="2f0ad2ac-1255-47a5-bde6-f54efcbb11b3"
train_df[train_df.text=='two']
# + [markdown] id="SvpcnoV_IgPE"
# ## POS Tagging - In Progress
# + colab={"base_uri": "https://localhost:8080/"} id="ws4A0lPpIh2Q" executionInfo={"status": "ok", "timestamp": 1616084745289, "user_tz": -480, "elapsed": 3813, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="94868dfa-d3f7-4e67-ff2b-b68e4b2298fd"
# pip install stanfordnlp
# !pip install stanza
# + colab={"base_uri": "https://localhost:8080/"} id="jUdDKpAqIiht" executionInfo={"status": "ok", "timestamp": 1616084859519, "user_tz": -480, "elapsed": 84478, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="b7b05c91-f419-4192-a2f6-4ab02f4bc09b"
# import stanfordnlp
# stanfordnlp.download('en') # This downloads the English models for the neural pipeline
# nlp = stanfordnlp.Pipeline() # This sets up a default neural pipeline in English
import stanza
stanza.download('en') # download English model
nlp = stanza.Pipeline('en') # initialize English neural pipeline
# + colab={"base_uri": "https://localhost:8080/"} id="-TJGGJBTLdEP" executionInfo={"status": "ok", "timestamp": 1616085123512, "user_tz": -480, "elapsed": 1041, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="7edaa3f5-ace5-4124-ac66-7b83524e79bf"
train_df.loc[0,:]
# + colab={"base_uri": "https://localhost:8080/"} id="5s1urQITJAdd" executionInfo={"status": "ok", "timestamp": 1616085152120, "user_tz": -480, "elapsed": 725, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="85a07dec-e03a-4f3a-d296-10112c543760"
doc = nlp(train_df.text[0])
print(doc)
# + colab={"base_uri": "https://localhost:8080/"} id="e4gBeK9sKtYh" executionInfo={"status": "ok", "timestamp": 1616085190798, "user_tz": -480, "elapsed": 834, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="b7423c8f-913a-4284-de3a-44c0e8788bf5"
print(doc.entities)
# + [markdown] id="U7WTnx_iMWZB"
# universal POS (UPOS) tags, treebank-specific POS (XPOS) tags, and universal morphological features (UFeats).
# + colab={"base_uri": "https://localhost:8080/"} id="ZtlxrsHAK9EL" executionInfo={"status": "ok", "timestamp": 1616085227763, "user_tz": -480, "elapsed": 1015, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07298175230978186476"}} outputId="90b9cec9-348b-40ea-cbcd-496fe4965b4e"
nlp = stanza.Pipeline(lang='en', processors='tokenize,mwt,pos')
doc = nlp(train_df.text[0])
print(*[f'word: {word.text}\tupos: {word.upos}\txpos: {word.xpos}\tfeats: {word.feats if word.feats else "_"}' for sent in doc.sentences for word in sent.words], sep='\n')
# + [markdown] id="CXYljdFcMrG6"
# # try out bidaf
# - may use cs224 bidaf baseline py file directly instead
# + id="cMMYqzp5MADc"
import nltk
# + id="llwmiXGjP_Fl"
#following replacement is suggested in the BidAF (Seo et al., 2016)
def tokenize(sequence):
tokens = [token.replace("``", '"').replace("''", '"').lower() for token in nltk.word_tokenize(sequence)]
return tokens
# + id="c3iL53wYP9vr"
def get_char_word_loc_mapping(context, context_tokens):
"""
Return a mapping that maps from character locations to the corresponding token locations.
If we're unable to complete the mapping e.g. because of special characters, we return None.
Inputs:
context: string (unicode)
context_tokens: list of strings (unicode)
Returns:
mapping: dictionary from ints (character locations) to (token, token_idx) pairs
Only ints corresponding to non-space character locations are in the keys
e.g. if context = "hello world" and context_tokens = ["hello", "world"] then
0,1,2,3,4 are mapped to ("hello", 0) and 6,7,8,9,10 are mapped to ("world", 1)
"""
acc = '' # accumulator
current_token_idx = 0 # current word loc
mapping = dict()
for char_idx, char in enumerate(context): # step through original characters
if char != u' ' and char != u'\n': # if it's not a space:
acc += char # add to accumulator
context_token = str(context_tokens[current_token_idx]) # current word token
if acc == context_token: # if the accumulator now matches the current word token
syn_start = char_idx - len(acc) + 1 # char loc of the start of this word
for char_loc in range(syn_start, char_idx+1):
mapping[char_loc] = (acc, current_token_idx) # add to mapping
acc = '' # reset accumulator
current_token_idx += 1
if current_token_idx != len(context_tokens):
return None
else:
return mapping
| Final_Code/Preprocessing/Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''cpuml-venv'': venv)'
# language: python
# name: python37664bitcpumlvenvvenvd94f9e51502149bb8cea6e47812f52f1
# ---
# # Salary Prediction Project EDA
#
# pass
__author__ = '<NAME>'
__email__ = '<EMAIL>'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Read in files and print the information
train_feature_df = pd.read_csv('../data/raw/train_features.csv')
train_target_df = pd.read_csv('../data/raw/train_salaries.csv')
test_feature_df = pd.read_csv('../data/raw/test_features.csv')
train_feature_df.head()
train_target_df.head()
test_feature_df.head()
train_feature_df.info()
train_target_df.info()
test_feature_df.info()
train_feature_df.duplicated().sum()
train_target_df.duplicated().sum()
test_feature_df.duplicated().sum()
train_feature_df.isnull().sum()
test_feature_df.isnull().sum()
train_target_df.isnull().sum()
numeric_cols = [col for col in train_feature_df.columns if train_feature_df[col].dtype == 'int64']
numeric_cols
categorical_cols = [col for col in train_feature_df.columns if train_feature_df[col].dtype == 'object']
categorical_cols
train_feature_df.describe().transpose()
train_feature_df.describe(include='O').transpose()
# For the `categorical` features, we can see that there's `63` Company the most frequent Company is `COMP39`, `8` types of job, most frequent is `SENIRO` level, `5` typs of education degree, the most frequent is `HIGH_SCHOOL`, and `9` types of degree major, the most frequent is `NONE` which mean maybe the individual forgot what major they are studied, but we assume is the individual not willing to provide. What's more, there are `7` types of Industry, the most frequent is `WEB` industry.
# Merge the features and salaries on jobId, delete original file to save memory
train_df = pd.merge(train_feature_df, train_target_df, on='jobId')
del train_feature_df
del train_target_df
train_df.head(10)
train_df.info()
plt.figure(figsize = (14, 6))
plt.subplot(1,2,1)
sns.boxplot(train_df.salary)
plt.subplot(1,2,2)
sns.distplot(train_df.salary, bins=20)
plt.show()
stat = train_df.salary.describe()
print(stat, '\n')
IQR = stat['75%'] - stat['25%']
upper = stat['75%'] + 1.5 * IQR
lower = stat['25%'] - 1.5 * IQR
print(f'The upper and lower bounds for suspected outliers are {upper} and {lower}.')
#check potential outlier below lower bound
train_df[train_df.salary < 8.5]
#check potential outlier above upper bound
train_df.loc[train_df.salary > 220.5, 'jobType'].value_counts()
# Check most suspicious potential outliers above upper bound
train_df[(train_df.salary > 220.5) & (train_df.jobType == 'JUNIOR')]
# These entries with zero salary do not appear to be volunteer positions. We are confident that they are instances of missing/corrupt data and should be removed from the training set.
# The high-salary potential outliers all appear to be legitimate data. Most roles are C-level executive roles and the junior positions are in industries that are well known for high salaries (oil, finance). We determine these entries to be legitimate and will not remove them.
# Remove data with zero salaries
train_df = train_df[train_df.salary > 8.5]
def plot_feature(df, col):
'''
Make plot for each features
left, the distribution of samples on the feature
right, the dependance of salary on the feature
'''
plt.figure(figsize = (14, 6))
plt.subplot(1, 2, 1)
if df[col].dtype == 'int64':
df[col].value_counts().sort_index().plot()
else:
#change the categorical variable to category type and order their level by the mean salary
#in each category
mean = df.groupby(col)['salary'].mean()
df[col] = df[col].astype('category')
levels = mean.sort_values().index.tolist()
df[col].cat.reorder_categories(levels, inplace=True)
df[col].value_counts().plot()
plt.xticks(rotation=45)
plt.xlabel(col)
plt.ylabel('Counts')
plt.subplot(1, 2, 2)
if df[col].dtype == 'int64' or col == 'companyId':
#plot the mean salary for each category and fill between the (mean - std, mean + std)
mean = df.groupby(col)['salary'].mean()
std = df.groupby(col)['salary'].std()
mean.plot()
plt.fill_between(range(len(std.index)), mean.values-std.values, mean.values + std.values, \
alpha = 0.1)
else:
sns.boxplot(x = col, y = 'salary', data=df)
plt.xticks(rotation=45)
plt.ylabel('Salaries')
plt.show()
plot_feature(train_df, 'companyId')
plot_feature(train_df, 'jobType')
plot_feature(train_df, 'degree')
plot_feature(train_df, 'major')
plot_feature(train_df, 'industry')
plot_feature(train_df, 'yearsExperience')
plot_feature(train_df, 'milesFromMetropolis')
def encode_label(df, col):
#encode the categories using average salary for each category to replace label
cat_dict ={}
cats = df[col].cat.categories.tolist()
for cat in cats:
cat_dict[cat] = train_df[train_df[col] == cat]['salary'].mean()
df[col] = df[col].map(cat_dict)
for col in train_df.columns:
if train_df[col].dtype.name == "category":
encode_label(train_df, col)
# Correlations between selected features and response
# jobId is discarded because it is unique for individual
fig = plt.figure(figsize=(12, 10))
features = ['companyId', 'jobType', 'degree', 'major', 'industry', 'yearsExperience', 'milesFromMetropolis']
sns.heatmap(train_df[features + ['salary']].corr(), cmap='Blues', annot=True)
plt.xticks(rotation=45)
plt.show()
| notebooks/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit
# metadata:
# interpreter:
# hash: 3e4985cd88b3a56a52b003f0882d181d8cd7ec7ac144250e5d6b471c317680e4
# name: python3
# ---
# +
# Run this!
# Here we set a randomisation seed for replicatability.
import os
os.environ['PYTHONHASHSEED'] = '0'
seed = 6
import random as rn
rn.seed(seed)
import numpy as np
np.random.seed(seed)
import warnings
warnings.filterwarnings("ignore")
from keras import backend as K
import keras
print('keras using %s backend'%keras.backend.backend())
import pandas as pd
from sklearn.preprocessing import OneHotEncoder
# Sets up the graphing configuration
import matplotlib.pyplot as graph
# %matplotlib inline
graph.rcParams['figure.figsize'] = (15,5)
graph.rcParams["font.family"] = 'DejaVu Sans'
graph.rcParams["font.size"] = '12'
graph.rcParams['image.cmap'] = 'rainbow'
# +
# Run this too!
# This gets our data ready
# Load the data
dataset = pd.read_csv('Data/dog_data.csv')
# Separate out the features
features = dataset.drop(['breed'], axis = 1)
# Sets the target one-hot vectors
target = OneHotEncoder(sparse = False).fit_transform(np.transpose([dataset['breed']]))
# Take the first 4/5 of the data and assign it to training
train_X = features.values[:160]
train_Y = target[:160]
# Take the last 1/5 of the data and assign it to testing
test_X = features.values[160:]
test_Y = target[160:]
# +
# Run this!
# Below are a few helper methods. Do not edit these.
def train_network(structure, activation, optimizer, epochs):
os.environ['PYTHONHASHSEED'] = '0'
rn.seed(seed)
np.random.seed(seed)
# This initialises the model
model = keras.models.Sequential()
# This is our input + the first hidden layer 1
model.add(keras.layers.Dense(units = structure[1], input_dim = structure[0], activation = activation))
# Hidden layer 2, if not ignored (of size 0)
if structure[2] > 0:
model.add(keras.layers.Dense(units = structure[2], activation = activation))
# Output layer
model.add(keras.layers.Dense(units=structure[-1], activation = "softmax"))
# Compiles the model with parameters
model.compile(loss = 'categorical_crossentropy', optimizer = optimizer, metrics = ['accuracy'])
# This tells the us training has started, so we know that it's actually running
print('training... ', end = '')
# This trains the network
training_stats = model.fit(train_X, train_Y, batch_size = 1, epochs = epochs, verbose = 0, shuffle = False)
# Results!
print('train_acc: %0.3f, test_acc: %0.3f' %(training_stats.history['accuracy'][-1],
model.evaluate(test_X, test_Y, verbose = 0)[1]))
# This returns the results and the model for use outside the function
return training_stats, model
# Plots our evaluations in a line graph to see how they compare
def plot_acc(train_acc, test_acc, title):
# Plots the training and testing accuracy lines
training_accuracy, = graph.plot(train_acc, label = 'Training Accuracy')
testing_accuracy, = graph.plot(test_acc, label = 'Testing Accuracy')
graph.legend(handles = [training_accuracy, testing_accuracy])
# Plots guide lines along y = 0 and y = 1 to help visualise
xp = np.linspace(0, train_acc.shape[0] - 1, 10 * train_acc.shape[0])
graph.plot(xp, np.full(xp.shape, 1), c = 'k', linestyle = ':', alpha = 0.5)
graph.plot(xp, np.full(xp.shape, 0), c = 'k', linestyle = ':', alpha = 0.5)
graph.xticks(range(0, train_acc.shape[0]), range(1, train_acc.shape[0] + 1))
graph.ylim(0,1)
graph.title(title)
graph.show()
# Plots our evaluations in a bar chart to see how they compare
def bar_acc(train_acc, test_acc, title, xticks):
index = range(1, train_acc.shape[0] + 1)
# Plots the training and testing accuracy bars
training_accuracy = graph.bar(index, train_acc, 0.4, align = 'center')
testing_accuracy = graph.bar(index, test_acc, 0.4, align = 'edge')
graph.legend((training_accuracy[0], testing_accuracy[0]), ('Training Accuracy', 'Testing Accuracy'))
graph.xticks(index, xticks)
graph.title(title)
graph.show()
# +
# Initialises empty arrays into which to append new values.
train_acc = np.empty((0))
test_acc = np.empty((0))
for hidden1 in range (1,11):
print('Evaluating model with %i hidden neurons... ' %hidden1, end = '')
training_stats, model = train_network(structure = [3, hidden1, hidden1, 3],
activation = 'relu', optimizer = 'RMSprop', epochs = 12)
train_acc = np.append(train_acc, training_stats.history['accuracy'][-1])
test_acc = np.append(test_acc, model.evaluate(test_X, test_Y, verbose = 0)[1])
plot_acc(train_acc, test_acc, 'hidden layer size performance comparison')
# +
train_acc = np.empty((0))
test_acc = np.empty((0))
# Makes a list of the activation functions we wish to compare
activation_functions = ['elu', 'selu', 'relu', 'tanh', 'sigmoid',
'hard_sigmoid', 'softplus', 'softsign', 'linear']
for activation in activation_functions:
print('Evaluating model with %s hidden layer activation function... ' %activation, end = '')
training_stats, model = train_network(structure = [3, 4, 2, 3],
activation = activation, optimizer = 'RMSprop', epochs = 12)
train_acc = np.append(train_acc, training_stats.history['accuracy'][-1])
test_acc = np.append(test_acc, model.evaluate(test_X, test_Y, verbose=0)[1])
bar_acc(train_acc, test_acc, 'activation function performance comparison using (4,2) hidden layer', activation_functions)
# +
train_acc = np.empty((0))
test_acc = np.empty((0))
activation_functions = ['elu', 'selu', 'relu', 'tanh', 'sigmoid',
'hard_sigmoid', 'softplus', 'softsign', 'linear']
for activation in activation_functions:
print('Evaluating model with %s hidden layer activation function... ' %activation, end='')
# The value you choose for <updateHere> below will change the size of the hidden layers. Lets try changing them both to 3 for now
# (but you can have a play around with different numbers if you want)
training_stats, model = train_network(structure = [3, 3, 3, 3],
activation = activation, optimizer = 'RMSprop', epochs = 12)
train_acc = np.append(train_acc, training_stats.history['accuracy'][-1])
test_acc = np.append(test_acc, model.evaluate(test_X, test_Y, verbose=0)[1])
bar_acc(train_acc, test_acc, 'activation function performance comparison using (3,3) hidden layer', activation_functions)
# +
train_acc = np.empty((0))
test_acc = np.empty((0))
# This is a list of the optimisation functions for us to compare
optimization_functions = ['SGD', 'RMSprop', 'Adagrad', 'Adadelta',
'Adam', 'Adamax', 'Nadam']
for optimizer in optimization_functions:
print('Evaluating model with %s optimizer... ' %optimizer, end='')
# The <addOptimizer> below is where we specify the optimizer in the code
training_stats, model = train_network(structure = [3, 4, 2, 3],
activation = 'relu', optimizer = optimizer, epochs = 12)
# This is recording our data for the plot
train_acc = np.append(train_acc, training_stats.history['accuracy'][-1])
test_acc = np.append(test_acc, model.evaluate(test_X, test_Y, verbose=0)[1])
# And now, the plot!
bar_acc(train_acc, test_acc, 'optimizer performance comparison using (4,2) hidden layer', optimization_functions)
# +
structure = [3, 4, 2, 3]
# REPLACE <activationFunction> WITH ONE OF THE FOLLOWING: 'relu', 'softsign', 'tanh', 'elu', 'selu', 'softplus', 'linear'
###
activation = 'relu'
###
###
# REPLACE <optimiser> WITH ONE OF THE FOLLOWING: 'SGD', 'adam', 'RMSprop', 'Adagrad', 'Adadelta', 'Adamax', 'Nadam'
###
optimizer = 'SGD'
###
training_stats, model = train_network(structure, activation, optimizer, epochs = 24)
# We can plot our training statistics to see how it developed over time
accuracy, = graph.plot(training_stats.history['accuracy'], label = 'Accuracy')
training_loss, = graph.plot(training_stats.history['loss'], label = 'Training Loss')
graph.legend(handles = [accuracy, training_loss])
loss = np.array(training_stats.history['loss'])
xp = np.linspace(0, loss.shape[0], 10 * loss.shape[0])
graph.plot(xp, np.full(xp.shape, 1), c = 'k', linestyle = ':', alpha = 0.5)
graph.plot(xp, np.full(xp.shape, 0), c = 'k', linestyle = ':', alpha = 0.5)
graph.show()
| Curso-de-iniciacion-sobre-aprendizaje-automatico/Realizacion-de-predicciones-a-partir-de-datos-complejos-con-redes-neuronales/redes_neuronales_avanzadas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task 2
#
# #### Vypracovali: <NAME>, <NAME>
#
# Implement basic backward pass in MLP. Perform forward and backward propagation through your network and check your gradients.
# This time, the forward pass is implemented for you. Notice the matrix notation - the dimensions are in form $[m,nX,1]$, where $m$ is batch size (number of samples) and $nX$ is the size of sample vector.
# Import
import numpy as np
# ## Activations
#
# Implement derivations of standard activation functions (ReLU, Sigmoid), which are used in your task.
# +
#------------------------------------------------------------------------------
# ActivationFunction class
#------------------------------------------------------------------------------
class ActivationFunction:
def __init__(self):
pass
def __call__(self, z):
pass
#------------------------------------------------------------------------------
# LinearActivationFunction class
#------------------------------------------------------------------------------
class LinearActivationFunction(ActivationFunction):
def __call__(self, z):
return z
def derivation(self, z):
return 1
#------------------------------------------------------------------------------
# RELUActivationFunction class
#------------------------------------------------------------------------------
class RELUActivationFunction(ActivationFunction):
def __call__(self, z):
return np.maximum(z, 0)
def derivation(self, z):
return (z>0).astype("float")
#------------------------------------------------------------------------------
# SigmoidActivationFunction class
#------------------------------------------------------------------------------
class SigmoidActivationFunction(ActivationFunction):
def __call__(self, z):
return 1.0/(1.0+np.exp(-z))
def derivation(self, z):
a = self(z)
return np.multiply(self(z),(1-self(z)))
# Activation mapping
MAP_ACTIVATION_FUCTIONS = {
"linear": LinearActivationFunction,
"relu": RELUActivationFunction,
"sigmoid": SigmoidActivationFunction
}
def CreateActivationFunction(kind):
if (kind in MAP_ACTIVATION_FUCTIONS):
return MAP_ACTIVATION_FUCTIONS[kind]()
raise ValueError(kind, "Unknown activation function {0}".format(kind))
# -
# ## Layer
#
# This is the main class which can hold different types of layers and provides us with standard tasks like forward propagation. Implement backward functions for defined classes.
#
# nUnits - number of neuron units in your layer
#
# prevLayer - previous layer (need it to know the shape of it to create appropriate number of weights for you to use in current layer)
# +
#------------------------------------------------------------------------------
# Layer class
#------------------------------------------------------------------------------
class Layer:
def __init__(self, act="linear", name="layer"):
self.shape = (0, 0)
self.activation = CreateActivationFunction(act)
self.name = name
def initialize(self, prevLayer,debug=False):
pass
def forward(self, x):
pass
#------------------------------------------------------------------------------
# InputLayer class
#------------------------------------------------------------------------------
class InputLayer(Layer):
def __init__(self, nUnits, name="Input"):
super().__init__(act="linear", name=name)
self.nUnits = nUnits
def initialize(self, prevLayer,debug=False):
self.shape = (self.nUnits, 1)
self.debug=debug
def forward(self, x):
self.a=x
return x
def backward(self, X):
return None
#------------------------------------------------------------------------------
# Basic Dense Layer class
#------------------------------------------------------------------------------
class DenseLayer(Layer):
def __init__(self, nUnits, act="linear", name="Dense"):
super().__init__(act, name=name,)
# init each neuron into list
self.nUnits = nUnits
self.W = None
self.b = None
def initialize(self, prevLayer,debug=False):
#initialize all neurons
self.shape = (self.nUnits, prevLayer.shape[0])
self.debug=debug
# Initialize weights and bias
prev_nUnits, _ = prevLayer.shape
self.W = np.random.randn(self.nUnits, prev_nUnits)
self.b = np.zeros((self.nUnits, 1), dtype=float)
def forward(self, X):
if self.debug:
print("Forward of", self.name)
print("X:",X.shape)
print("W:",self.W.shape)
self.z = np.matmul(self.W, X) + self.b # Z = W*x + b
self.a = self.activation(self.z) # a = activation(Z)
return self.a
def backward(self, da, aPrev):
# da = dLoss -> dL/da of previous layer - with respect to backward pass
# aPrev = activation of previous layer needed for weights - with respect to forward pass
batch_size = aPrev.shape[1]
print("Backward of", self.name)
dz = np.multiply(da,self.activation.derivation(self.z))
self.dW = (1./batch_size)*np.matmul(dz, aPrev.T)
self.db = (1./batch_size)*np.sum(dz,axis=1, keepdims=True)
if self.debug:
print("dz:",dz.shape," aPrev.T:", aPrev.T.shape)
print("w:", self.W.shape," dw:", self.dW.shape)
print("db:", self.db.shape)
daPrev = np.matmul(self.W.T,dz)
return daPrev
# -
# ## Loss Functions
#
# Implement two standard loss functions (Binary Cross Entropy and Mean Squared Error), which you will/can use in your implementation of MLP backward pass.
# +
#------------------------------------------------------------------------------
# LossFunction class
#------------------------------------------------------------------------------
class LossFunction:
def __init__(self,debug=False):
self.debug=debug
pass
def __call__(self, A, Y):
pass
def derivation(self, A, Y):
pass
#------------------------------------------------------------------------------
# BinaryCrossEntropyLossFunction class
#------------------------------------------------------------------------------
class BinaryCrossEntropyLossFunction(LossFunction):
def __call__(self, A, Y):
# Warning! Use of logarithm - Take care about definition scope
return -(np.multiply(Y,np.log(A))+np.multiply((1-Y),np.log(1-A)))
def derivation(self, A, Y):
# Warning! Use of logarithm - Take care about definition scope
if self.debug:
print("A",A.shape)
print("Y",Y.shape)
return (-np.divide(Y,A)+np.divide((1-Y),(1-A)))
class MeanSquaredErrorLossFunction(LossFunction):
def __call__(self, A, Y): # loss = (Y-A)^2
return np.mean(np.square((Y-A)),axis=0,keepdims=True)
def derivation(self, A, Y): #dLoss = -1*(2(Y-A))
return np.mean(-2*(Y-A),axis=0,keepdims=True)
MAP_LOSS_FUNCTIONS = {
"bce": BinaryCrossEntropyLossFunction,
"mse": MeanSquaredErrorLossFunction
}
def CreateLossFunction(kind,debug=False):
if (kind in MAP_LOSS_FUNCTIONS):
return MAP_LOSS_FUNCTIONS[kind](debug)
raise ValueError(kind, "Unknown loss function {0}".format(kind))
# -
# ## Model class
#
# This is the basic class which holds all of your layers and encapsulate functionality to predict results from your input as a forward pass through all the layers after you create your model and initialize all the layers.
#
# Implemet backpropagation.
#------------------------------------------------------------------------------
# Model class
#------------------------------------------------------------------------------
class Model:
def __init__(self, lossName, debug = False):
self.layers = []
# Initialize loss function
self.loss_fn = CreateLossFunction(lossName,debug)
self.debug = debug
def addLayer(self, layer):
self.layers.append(layer)
def initialize(self):
# Call initialization sequentially on all layers
prevLayer = None
for l in self.layers:
l.initialize(prevLayer, self.debug)
prevLayer = l
def forward(self, X):
# Single feed forward
A = X
for l in self.layers:
A = l.forward(A)
return A
def backward(self, dLoss):
da = dLoss
if self.debug:
print("da",da.shape)
for layer, lPrev in zip(self.layers[::-1], self.layers[-2::-1]):
if self.debug:
print("L:",layer.name,"pL:",lPrev.name)
da = layer.backward(da, lPrev.a)
def compute_cost(self, A, Y):
return np.mean(self.loss_fn(A,Y), axis = -1)
def derive_loss(self, A, Y):
return self.loss_fn.derivation(A,Y)
# ### Main Processing Cell
#
# 1. Initialize dataset.
# 2. Declare a simple model (at least 4 layer) with relu on hidden layers and sigmoid on output layer.
# 3. Perform forward pass through the network.
# 4. Compute cost.
# 5. Derive loss.
# 6. Perform backward pass.
# 7. Celebrate and scroll lower.
# +
# Main processing
from dataset import dataset_Circles
# Task A:
X, Y = dataset_Circles(n=50, radius=0.7, noise=0.0)
X=X.squeeze(axis=-1).T
Y=Y.squeeze(axis=-1).T
model = Model(lossName = "bce",debug = False)
model.addLayer( InputLayer(nUnits=2, name="input_layer"))
model.addLayer( DenseLayer(nUnits=6, act="relu", name="1st_Layer"))
model.addLayer( DenseLayer(nUnits=3, act="relu", name="2nd_Layer"))
model.addLayer( DenseLayer(nUnits=2, act="relu", name="3rd_Layer"))
model.addLayer( DenseLayer(nUnits=1, act="sigmoid", name="4th_Layer"))
model.initialize()
A = model.forward(X)
loss = model.derive_loss(A,Y)
# for layer in model.layers[1:]:
# print(np.squeeze(layer.a).shape)
model.backward(loss)
print(model.compute_cost(A,Y))
# + [markdown] pycharm={"name": "#%% md\n"}
# **How does gradient checking work?**.
#
# As in 1) and 2), you want to compare "gradapprox" to the gradient computed by backpropagation. The formula is still:
#
# $$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
#
# However, $\theta$ is not a scalar anymore. It is a dictionary called "parameters". We implemented a function "`dictionary_to_vector()`" for you. It converts the "parameters" dictionary into a vector called "values", obtained by reshaping all parameters (W1, b1, W2, b2, W3, b3) into vectors and concatenating them.
#
# The inverse function is "`vector_to_dictionary`" which outputs back the "parameters" dictionary.
#
#
# We have also converted the "gradients" dictionary into a vector "grad" using gradients_to_vector(). You don't need to worry about that.
#
#
# Here is pseudo-code that will help you implement the gradient check.
#
# For each i in num_parameters:
# - To compute `J_plus[i]`:
# 1. Set $\theta^{+}$ to `np.copy(parameters_values)`
# 2. Set $\theta^{+}_i$ to $\theta^{+}_i + \varepsilon$
# 3. Calculate $J^{+}_i$ using to `forward_propagation_n(x, y, vector_to_dictionary(`$\theta^{+}$ `))`.
# - To compute `J_minus[i]`: do the same thing with $\theta^{-}$
# - Compute $gradapprox[i] = \frac{J^{+}_i - J^{-}_i}{2 \varepsilon}$
#
# Thus, you get a vector gradapprox, where gradapprox[i] is an approximation of the gradient with respect to `parameter_values[i]`. You can now compare this gradapprox vector to the gradients vector from backpropagation. Just like for the 1D case (Steps 1', 2', 3'), compute:
# $$ difference = \frac {\| grad - gradapprox \|_2}{\| grad \|_2 + \| gradapprox \|_2 } \tag{3}$$
#
#
# **The code will be added later** but soon enough ;)
# +
# GRADED FUNCTION: gradient_check_n
gradient_check_n(model,X,Y)
# -
# ## Verification cell
#
# 8. Verify your solution by gradient checking.
# 9. Start crying.
# 10. Repeat until correct ;)
# + pycharm={"name": "#%%\n"}
def gradient_check_n(network, X, Y, epsilon = 1e-7):
"""
Checks if backward_propagation_n computes correctly the gradient of the cost output by forward_propagation_n
Arguments:
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
grad -- output of backward_propagation_n, contains gradients of the cost with respect to the parameters.
x -- input datapoint, of shape (input size, 1)
y -- true "label"
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Set-up variables
gradapprox = []
grad_backward = []
for i,layer in enumerate(network.layers):
# Compute gradapprox
if i < 1:
continue
shape = layer.W.shape
# print(shape[0], ',', shape[1])
for i in range(shape[0]):
for j in range(shape[1]):
# print('i',i,'j',j)
# Compute J_plus[i]. Inputs: "parameters_values, epsilon". Output = "J_plus[i]".
# "_" is used because the function you have to outputs two parameters but we only care about the first one
origin_W = layer.W[i][j]
layer.W[i][j] = origin_W + epsilon
A_plus = network.forward(X)
J_plus = network.compute_cost(A_plus, Y)
# Compute J_minus[i]. Inputs: "parameters_values, epsilon". Output = "J_minus[i]".
layer.W[i][j] = origin_W - epsilon
A_minus = network.forward(X)
J_minus = network.compute_cost(A_minus, Y)
# Compute gradapprox[i]
gradapprox.append((J_plus - J_minus) / (2*epsilon))
# print(layer.name, layer.dW.shape)
# grad = np.mean(layer.dW, axis=0, keepdims=True)
# grad_backward.append(grad[0][i][j])
grad_backward.append(layer.dW[i][j])
layer.W[i][j] = origin_W
# Compare gradapprox to backward propagation gradients by computing difference.
gradapprox = np.reshape(gradapprox, (-1, 1))
grad_backward = np.reshape(grad_backward, (-1, 1))
numerator = np.linalg.norm(grad_backward - gradapprox)
denominator = np.linalg.norm(grad_backward) + np.linalg.norm(gradapprox)
difference = numerator / denominator
if difference > 2e-7:
print ("\033[91m" + "There is a mistake in the backward propagation! difference = " + str(difference) + "\033[0m")
else:
print ("\033[92m" + "Your backward propagation works perfectly fine! difference = " + str(difference) + "\033[0m")
| week_3/Task_2_backward_pass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring ONC Data Web Services
#
# Exploring the ONC data web services documented at https://wiki.oceannetworks.ca/display/help/API
# +
import json
import os
from urllib.parse import (
quote,
urlencode,
)
import arrow
import numpy as np
import requests
import xarray as xr
# -
# ## Contents
#
# * [Web Service Requests](#Web-Service-Requests)
# * [`stations` End-point](#stations-End-point)
# * [`scalardata` End-point](#scalardata-End-point)
#
# * [VENUS Node CTD Data](#VENUS-Node-CTD-Data)
# * [Ferry Temperature and Salinity Data](#Ferry-Temperature-and-Salinity-Data)
# * [Parsing `scalardata`](#Parsing-scalardata)
# ## Web Service Requests
#
# The web service URLS are composed of:
#
# * the base URL: http://dmas.uvic.ca/api/
# * an end-point: presently one of `archivefiles`, `dataproducts`, `rawdata`, `scalardata`, `stations`, or `status`
# * a query string that includes `method=methodName`, `token=USER_TOKEN`, and other end-point-specific key/value pairs
#
# For example:
#
# `http://dmas.uvic.ca/api/scalardata?method=getByStation&token=<yourValidToken>&station=SCHDW.O1&deviceCategory=OXYSENSOR`
#
# Access to the web services requires a user token which you can generate on the
# `Web Services API` tab of your [ONC account profile page](http://dmas.uvic.ca/Profile).
# I have stored mine in an environment variable so as not to publish it to the world
# in this notebook.
TOKEN = os.environ['ONC_USER_TOKEN']
# Here's a string template for a request URL, and a technique for composing the query string:
url_tmpl = 'http://dmas.uvic.ca/api/{endpoint}?{query}'
url_tmpl.format(
endpoint='scalardata',
query=urlencode({
'method': 'getByStation',
'token': 'USER_TOKEN',
'station': 'SCHDW.O1',
'deviceCategory': 'OXYSENSOR',
}, quote_via=quote, safe='/:'))
# Substituting my real token,
# and using the `requests` package to send the data request to the web service:
# +
data_url = url_tmpl.format(
endpoint='scalardata',
query=urlencode({
'method': 'getByStation',
'token': TOKEN,
'station': 'SCHDW.O1',
'deviceCategory': 'OXYSENSOR',
}, quote_via=quote, safe='/:'))
response = requests.get(data_url)
response.raise_for_status()
# -
# Calling the `raise_for_status()` method on the response object is a quick
# way to test for HTTP errors.
#
# The default response type from the web services is JSON
# and `requests` provides a convenience method to convert JSON
# in the response to a Python `dict` object:
response.json()
# So, let's put that all together into a function to query the ONC data web services:
def get_onc_data(endpoint, method, token, **query_params):
url_tmpl = 'http://dmas.uvic.ca/api/{endpoint}?{query}'
query = {'method': method, 'token': token}
query.update(query_params)
data_url = url_tmpl.format(
endpoint=endpoint,
query=urlencode(query, quote_via=quote, safe='/:'))
response = requests.get(data_url)
response.raise_for_status()
return response.json()
get_onc_data('scalardata', 'getByStation', TOKEN, station='SCHDW.O1', deviceCategory='OXYSENSOR')
# ## `stations` End-point
#
# The [`stations` end-point](https://wiki.oceannetworks.ca/display/help/stations)
# has a `getTree` method that returns a large, hierarchical tree data structure of
# stations, station codes, and devices.
# ```python
# get_onc_data('stations', 'getTree', TOKEN)
# ```
# Rather than reading or parsing the data structure here,
# it is perhaps easier to find station codes and device categories to use with the
# other end-points by looking at the list generated by the
# [Javascript Usage Exmaple](https://wiki.oceannetworks.ca/download/attachments/42172426/stationTreeExample.html?version=2&modificationDate=1428438144000&api=v2)
# provided by ONC.
# ## `scalardata` End-point
#
# The [`scalardata` end-point](https://wiki.oceannetworks.ca/display/help/scalardata+service)
# has a `getByStation` method that returns time series of data given a station code
# and an device category code.
# Its simplest use case is to return the most recent data for all sensors associated with
# the device category at the station.
# ### VENUS Node CTD Data
#
# Here that is for the CTD at the Salish Sea Central node VENUS Instrument Platform:
get_onc_data('scalardata', 'getByStation', TOKEN, station='USDDL', deviceCategory='CTD')
# Adding a `sensors` item to the query with a value that is a comma-separated list
# of `sensor` codes limits the response to contain only the data from the specified sensors:
get_onc_data(
'scalardata', 'getByStation', TOKEN, station='SCVIP', deviceCategory='CTD',
sensors='salinity,temperature')
# Time series of data are obtained by adding `dateFrom` and `dateTo` items to the query:
get_onc_data(
'scalardata', 'getByStation', TOKEN, station='SCVIP', deviceCategory='CTD',
sensors='salinity,temperature',
dateFrom='2016-06-28T00:26:45.000Z',
)
# With only a `dateFrom` item in the query the time series length defaults to 1 day in length.
# The `dateTo` query item controls the length of the time series by date/time stamp.
get_onc_data(
'scalardata', 'getByStation', TOKEN, station='SCVIP', deviceCategory='CTD',
sensors='salinity,temperature',
dateFrom='2016-06-21T17:58:45.000Z', dateTo='2016-06-21T17:58:50.000Z',
)
# The number of measurements returned can be specified directly with the `rowLimit` query item.
# *Note also that there is a hard limit of 100,000 measurements per sensor per request.*
get_onc_data(
'scalardata', 'getByStation', TOKEN, station='SCVIP', deviceCategory='CTD',
sensors='salinity,temperature',
dateFrom='2016-06-21T17:58:45.000Z', rowLimit=2,
)
# The values for `dateFrom` and `dateTo` are in UTC and must be strings formatted
# as `yyyy-MM-ddTHH:mm:ss.SSSZ`.
# That format is annoying enough to type,
# and timezone conversions are error-prone enough
# that it is worth writing a function to handle the details:
def onc_datetime(datetime_str, timezone='Canada/Pacific'):
d = arrow.get(datetime_str)
d_tz = arrow.get(d.datetime, timezone)
d_utc = d_tz.to('utc')
return '{}Z'.format(d_utc.format('YYYY-MM-DDTHH:mm:ss.SSS'))
# The `onc_datetime()` function has been added to the
# [`salishssea_tools.data_tools` module](http://salishsea-meopar-tools.readthedocs.io/en/latest/SalishSeaTools/api.html#module-salishsea_tools.data_tools).
onc_datetime('2016-06-21 10:58:45')
onc_datetime('2016-06-21 17:58:45', 'utc')
get_onc_data(
'scalardata', 'getByStation', TOKEN, station='SCVIP', deviceCategory='CTD',
sensors='salinity,temperature',
dateFrom=onc_datetime('2016-06-21 10:58:45'), dateTo=onc_datetime('2016-06-21 10:58:50'),
)
# ### Ferry Temperature and Salinity Data
#
# The instrumented ferries are stations.
# Here is the most recent available data from the TSG device
# aboard the Tsawwassen to Duke Point ferry:
get_onc_data('scalardata', 'getByStation', TOKEN, station='TWDP', deviceCategory='TSG')
# Note that the data may lag the present time by several hours because it is only
# transmitted from the ferry to the ONC servers when the ferry is at dock.
# Also, there appears to be a several hours long gap in the data each day,
# presumably while the ferry is docked overnight.
#
# The ferry's location is available from the NAV device.
# *Note that the times from the TSG and NAV devices do not appear to be synchronized.*
get_onc_data('scalardata', 'getByStation', TOKEN, station='TWDP', deviceCategory='NAV')
# ### Parsing `scalardata`
#
# The Python `dict` data structure that we get from the `scalarData` end-point
# has 2 top-level keys: `sensorData` and `serviceMetadata`.
data = get_onc_data(
'scalardata', 'getByStation', TOKEN, station='SCVIP', deviceCategory='CTD',
sensors='salinity,temperature',
dateFrom=onc_datetime('2016-06-21 10:58:45'), dateTo=onc_datetime('2016-06-21 10:58:50'),
)
data['serviceMetadata']
# `serviceMetadata` is a `dict` of metadata attributes of the returned data
# as a whole.
data['sensorData']
# `sensorData` is a `list` of `dict`s containing the sensor data and metadata
# for each of the sensors requested in the query
# (or all of the sensors in the `deviceCategory` if an explicit list of sensors was
# not included in the query).
#
# The metadata keys in each `list` element are:
#
# * `sensor`: the sensor id (as listed in the query)
# * `actualSamples`: the count of the data samples for the sensor
# * `sensorName`: the sensor's descriptive name
# * `unitOfMeasure`: the sensor's unit of measure
#
# The sensor data is contained in a `list` of `dict`s that are the value
# associated with the `data` key:
data['sensorData'][0]['data']
# We can parse those `dict`s into `list`s of data items with list comprehensions:
qaqcFlag = [d['qaqcFlag'] for d in data['sensorData'][0]['data']]
qaqcFlag
# The meaning of the `qaqcFlag` is
# [described in the ONC docs](http://www.oceannetworks.ca/data-tools/data-quality).
salinity = [d['value'] for d in data['sensorData'][0]['data']]
salinity
# It's convenient to use [`arrow`](http://crsmithdev.com/arrow/)
# to convert the `sampleTime` strings to Python objects:
timestamp = [arrow.get(d['sampleTime']) for d in data['sensorData'][0]['data']]
timestamp
# and from there it is easy to get timezone-aware `datetime` objects if we need them:
[t.datetime for t in timestamp]
# Rather than dealing with the layers of `dict`s and `list`s that
# we get back from the `scalardata` service it is worthwhile to
# create a function that constructs an
# [`xarray.Dataset` object](http://xarray.pydata.org/en/stable/data-structures.html#dataset)
# containing the data and metadata.
def onc_json_to_dataset(onc_json):
data_vars = {}
for sensor in data['sensorData']:
data_vars[sensor['sensor']] = xr.DataArray(
name=sensor['sensor'],
data=[d['value'] for d in sensor['data']],
coords={
'sampleTime': [arrow.get(d['sampleTime']).datetime
for d in sensor['data']],
},
attrs={
'qaqcFlag': np.array([d['qaqcFlag'] for d in sensor['data']]),
'sensorName': sensor['sensorName'],
'unitOfMeasure': sensor['unitOfMeasure'],
'actualSamples': sensor['actualSamples'],
}
)
return xr.Dataset(data_vars, attrs=onc_json['serviceMetadata'])
# The `onc_json_to_dataset()` function has been added to the
# [`salishssea_tools.data_tools` module](http://salishsea-meopar-tools.readthedocs.io/en/latest/SalishSeaTools/api.html#module-salishsea_tools.data_tools).
onc_json_to_dataset(data)
# It is noteworthy that,
# even though the `xarray.Dataset` contructor
# collapses the `sampleTime` coordinates of the 2 sensors on to a
# single dataset coordinate,
# the individual `sampleTime` arrays are preserved at the variable level:
ds = onc_json_to_dataset(data)
ds.salinity.sampleTime is ds.temperature.sampleTime
| notebooks/ONC-DataWebServices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Data Science – Practical Natural Language Processing (NLP)
# *COMP 5360 / MATH 4100, University of Utah, http://datasciencecourse.net/*
#
# In this lecture, we'll do some practical NLP following up on the theoretical lecture. We will do some basic text processing followed by analyzing sentiment for movie reviews. For this purpose, we'll introduce the [Natural Language Toolkit (NLTK)](http://www.nltk.org/), a Python library for natural language processing.
#
# We won't cover NLTK or NLP extensively here – this lecture is meant to give you a few pointers if you want to use NLP in the future, e.g., for your project.
#
# Also, there is a well-regarded alternative to NLTK: [Spacy](https://spacy.io/). If you're planning to use a lot of NLP in your project, that might be worth checking out.
#
# **Reading:**
#
# [<NAME>, <NAME>, and <NAME>, *Natural Language Processing with Python – Analyzing Text with the Natural Language Toolkit*](http://www.nltk.org/book/).
#
#
# [<NAME> and <NAME>, *Foundations of Statistical Natural Language Processing* (1999).](http://nlp.stanford.edu/fsnlp/)
#
# [<NAME> and <NAME>, *Speech and Language Processing* (2016).](https://web.stanford.edu/~jurafsky/slp3/)
#
# **In a prior lecture,** guest lecturer <NAME> gave a nice overview of Natural Language Processing (NLP). He gave several examples of NLP tasks:
# * Part of speech tagging (what are the nouns, verbs, adjectives, prepositions).
# + Information Extraction
# + Sentiment Analysis (determine the attitude of text, e.g., is it positive or negative).
# + Semantic Parsing (translate natural language into a formal meaning representation).
#
# One of the major takeaways from his talk is that the current state-of-the-art for many NLP tasks is to find a good way to represent the text ("extract features") and then to use machine learning / statistics tools, such as classification or clustering.
#
# Our goal today is to use NLTK + scikit-learn to do some basic NLP tasks.
# ### Install datasets and models
#
# To use NLTK, you must first download and install the datasets and models. Run the following:
import nltk
nltk.download('all')
# +
# imports and setup
import numpy as np
from sklearn.neighbors import NearestNeighbors
from sklearn.neighbors import KNeighborsClassifier
from sklearn import svm
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (15, 9)
plt.style.use('ggplot')
# -
# ## Basics of NLTK
#
# We have downloaded a set of text corpora above. Here is a list of these texts:
from nltk.book import *
# Let's look at the first 20 words of text1 – <NAME>:
text1[0:20]
# ### Text Statistics
#
# We can check the length of a text. The text of <NAME> is 26,0819 words, whereas Monty Python and the Holy Grail has 16,967 words.
len(text1)
len(text6)
# We can check for the frequency of a word. The word "swallow" appears 10 times in Monty Python.
text6.count("swallow")
# We might want to know the context in which "swallow" appears in the text
#
# "You shall know a word by the company it keeps." – <NAME>
#
# Use the [`concordance`](http://www.nltk.org/api/nltk.html#nltk.text.Text.concordance) function to print out the words just before and after all occurrences of the word "swallow".
text6.concordance("swallow")
# Words that occur with notable frequencey are "fly" or "flight", "unladen", "air", "African", "European". We can learn about what a swallow can do or properties of a swallow by this.
# And if we look for Ishmael in Moby Dick:
text1.concordance("Ishmael")
# Here, we see a lot of "I"s. We could probably infer that Ishmael is the narrator based on that.
# We can see what other words frequently appear in the same context using the [`similar`](http://www.nltk.org/api/nltk.html#nltk.text.Text.similar) function.
text6.similar("swallow")
text6.similar("african")
text6.similar("coconut")
# This means that 'african' and 'unladen' both appeared in the text with the same word just before and just after. To see what the phrase is, we can use the [`common_contexts`](http://www.nltk.org/api/nltk.html#nltk.text.Text.concordance) function.
text6.common_contexts(["African", "unladen"])
# We see that both "an unladen swallow" and "an african swallow" appear in the text.
text6.concordance("unladen")
print()
text6.concordance("african")
# ### Dispersion plot
#
# `text4` is the Inaugural Address Corpus which includes inaugural addresses going back to 1789.
# We can use a dispersion plot to see where in a text certain words appear, and hence how the language of the address has changed over time.
#
# +
print(" ".join(text4[:100]))
print("")
print(" ".join(text4[-100:]))
# -
text4.dispersion_plot(["citizens", "democracy", "freedom", "duty", "America", "nation", "God", "military"])
# ### Exploring texts using statistics
#
# We'll explore a text by counting the frequency of different words.
#
# The total number of words ("outcomes") in Moby Dick is 260,819 and the number of different words is 19,317.
# +
frequency_dist = FreqDist(text1)
print(frequency_dist)
# find 50 most common words
print('\n',frequency_dist.most_common(50))
# not suprisingly, whale occurs quite frequently (906 times!)
print('\n', frequency_dist['whale'])
# -
# We can find all the words in Moby Dick with more than 15 characters
unique_words = set(text1)
long_words = [w.lower() for w in unique_words if len(w) > 15]
long_words
# ### Stopword Removal
#
# Often, it is useful to ignore frequently used words, to concentrate on the meaning of the remaining words. These are referred to as *stopwords*. Examples are "the", "was", "is", etc.
#
# NLTK comes with a stopword corpus.
from nltk.corpus import stopwords
stopwords = nltk.corpus.stopwords.words('english')
print(stopwords)
# Depending on the task, these stopwords are important modifiers, or superfluous content.
# ### Exercise 1.1: Frequent Words
# Find the most frequently used words in Moby Dick that are not stopwords and not punctuation. Hint: [`str.isalpha()`](https://docs.python.org/3/library/stdtypes.html#str.isalpha) could be useful here.
# +
# your code here
frequency_dist = FreqDist(text1)
most_common = frequency_dist.most_common(500)
filtered_words = [word_tuple for word_tuple in most_common if word_tuple[0].lower not in stopwords]
filtered_words = [word_tuple for word_tuple in filtered_words if word_tuple[0].isalpha()]
filtered_words[0:50]
# -
# ### Stopwords in different corpora
# Is there a difference between the frequency in which stopwords appear in the different texts?
# +
def content_fraction(text):
stopwords = nltk.corpus.stopwords.words('english')
content = [w for w in text if w.lower() not in stopwords]
return len(content) / len(text)
for i,t in enumerate([text1,text2,text3,text4,text5,text6,text7,text8,text9]):
print(i+1,content_fraction(t))
# -
# Apparently, "text8: Personals Corpus" has the most content.
# ### Collocations
# A *collocation* is a sequence of words that occur together unusually often, we can retreive these using the [`collocations()`](http://www.nltk.org/api/nltk.html#nltk.text.Text.collocations) function.
# +
# this won't work because of a current bug in NLTK
# text2.collocations()
# -
# ## Sentiment analysis for movie reviews
# When analyzing movie reviews, we can ask the simple question: Is the attitude of a movie review positive or negative? If you're developing rotten tomatoes, that's what you want to know to certify whether a review is "fresh" or "rotten".
#
# How can we approach this question?
#
# Our data is a corpus consisting of 2000 movie reviews together with the user's sentiment polarity (positive or negative). More information about this dataset is available [from this website](https://www.cs.cornell.edu/people/pabo/movie-review-data/).
#
# Our goal is to predict the sentiment polarity from just the review.
#
# Of course, this is something that we can do very easily:
# 1. That movie was terrible. -> negative
# + That movie was great! -> positive
#
#
#
#
from nltk.corpus import movie_reviews as reviews
# The datset contains 1000 positive and 1000 negative movie reviews.
#
# The paths to / IDs for the individual reviews are accessible via the fileids() call:
reviews.fileids()[0:5]
# We can access the positives or negatives explicitly:
reviews.fileids('pos')[0:5]
# There are in fact 1000 positive and 1000 negative reviews:
num_reviews = len(reviews.fileids())
print(num_reviews)
print(len(reviews.fileids('pos')),len(reviews.fileids('neg')))
# Let's see the review for the third movie. Its a negative review for [The Mod Squad](https://www.rottentomatoes.com/m/mod_squad/) (see the [trailer](https://www.youtube.com/watch?v=67cdXuWnRKs)), which has a "rotten" rating on rotten tomatoes.
#
# 
# +
# the name of the file
fid = reviews.fileids()[2]
print(fid)
print('\n', reviews.raw(fid))
print('\n', "The Category:", reviews.categories(fid) )
print('\n', "Individual Words:",reviews.words(fid))
# -
# Let's look at some sentences that indicate that this is a negative review:
#
# * "it is movies like these that make a jaded movie viewer thankful for the invention of the timex indiglo watch"
# * "sounds like a cool movie , does it not ? after the first fifteen minutes , it quickly becomes apparent that it is not ."
# * "nothing spectacular"
# * "avoid this film at all costs"
# * "unfortunately , even he's not enough to save this convoluted mess"
# ### A Custom Algorithm
# We'll build a sentiment classifier using methods we already know to predicts the label ['neg', 'pos'] from the review text
#
# `reviews.categories(file_id)` returns the label ['neg', 'pos'] for that movie
categories = [reviews.categories(fid) for fid in reviews.fileids()]
print(categories[0:10])
labels = {'pos':1, 'neg':0}
# create the labels: 1 for positive, 0 for negative
y = [labels[x[0]] for x in categories]
y[0], y[1000]
# Here, we collect all words into a nested array datastructure:
doc_words = [list(reviews.words(fid)) for fid in reviews.fileids()]
# first 10 words of the third document - mod squad
doc_words[2][1:10]
# Here we get all of the words in the reviews and make a FreqDist, pick the most common 2000 words and remove the stopwords.
# +
# get the 2000 most common words in lowercase
most_common = nltk.FreqDist(w.lower() for w in reviews.words()).most_common(2000)
# remove stopwords
filtered_words = [word_tuple for word_tuple in most_common if word_tuple[0].lower() not in stopwords]
# remove punctuation marks
filtered_words = [word_tuple for word_tuple in filtered_words if word_tuple[0].isalpha()]
print(len(filtered_words))
filtered_words[0:50]
# -
# We extract this word list from the frequency tuple.
word_features = [word_tuple[0] for word_tuple in filtered_words]
print(word_features[:5])
len(word_features)
# We define a function that takes a document and returns a list of zeros and ones indicating which of the words in `word_features` appears in that document.
def document_features(document):
# convert each document into a set of its words
# this removes duplicates and makes "existence" tests efficient
document_words = set(document)
# a list, initalized with 0s, that we'll set to 1 for each of the words that exists in the document
features = np.zeros(len(word_features))
for i, word in enumerate(word_features):
features[i] = (word in document_words)
return features
# Let's just focus on the third document. Which words from `word_features` are in this document?
# +
words_in_doc_2 = document_features(doc_words[2])
print(words_in_doc_2)
inds = np.where(words_in_doc_2 == 1)[0]
print('\n', [word_features[i] for i in inds])
# -
# Now we build our feature set for all the reviews.
# +
X = np.zeros([num_reviews,len(word_features)])
for i in range(num_reviews):
X[i,:] = document_features(doc_words[i])
X[0:5]
# -
# The result is a feature vector for each of these reviews that we can use in classification.
# Now that we have features for each document and labels, **we have a classification problem!**
#
# NLTK has a built-in classifier, but we'll use the scikit-learn classifiers we're already familiar with.
#
# Let's try k-nearest neighbors:
k = 30
model = KNeighborsClassifier(n_neighbors=k)
scores = cross_val_score(model, X, y, cv=10)
print(scores)
# And SVM:
model = svm.SVC(kernel='rbf', C=30, gamma="auto")
scores = cross_val_score(model, X, y, cv=10)
print(scores)
# Here we can see that kNN with these parameters is less accurate than SVM, which is about 80% accurate. Of course, we could now use cross validation to find the optimal parameters, `k` and `C`
# So, let's see what our algorithm things about the Mod Squad!
# +
XTrain, XTest, yTrain, yTest = train_test_split(X, y, random_state=1, test_size=0.2)
model.fit(XTrain, yTrain)
# -
mod_squad = [X[2]]
mod_squad
model.predict(mod_squad)
# Our model says 0 - so a bad review! We have succesfully build a classifier that can detect the Mod Squad review as a bad review!
# Let's take a look at a mis-classified movie. Remember, that the first 1000 movies are negative reviews, so we can just look for the first negative one:
model.predict(X[0:10])
# Review 9, which was misclassified, is for Aberdeen, which has [generally favorable reviews](https://www.rottentomatoes.com/m/aberdeen/) with about 80% positive. Let's looks at the review:
# +
fid = reviews.fileids()[8]
print('\n', reviews.raw(fid))
print('\n', reviews.categories(fid) )
# -
# So if we read this, we can see that this is a negative review, but not a terrible review. Take this sentence for example:
#
# * "if signs & wonders sometimes feels overloaded with ideas , at least it's willing to stretch beyond what we've come to expect from traditional drama"
# * "yet this ever-reliable swedish actor adds depth and significance to the otherwise plodding and forgettable aberdeen , a sentimental and painfully mundane european drama"
# ## We could have also used the Classifier from the NLTK library
#
# Below is the sentiment analysis from [Ch. 6 of the NLTK book](http://www.nltk.org/book/ch06.html).
#
#
documents = [(list(reviews.words(fileid)), category)
for category in reviews.categories()
for fileid in reviews.fileids(category)]
# This list contains tuples where the review, stored as an array of words, is the first item in the tuple and the category is the second.
documents[1]
# Extract the features from all of the documents
# +
def document_features(document):
document_words = set(document)
features = {}
for word in word_features:
features['contains('+ word +')'] = (word in document_words)
return features
featuresets = [(document_features(d), c) for (d,c) in documents]
# -
featuresets[2]
# Split into train_set, test_set and perform classification
# +
train_set, test_set = featuresets[100:], featuresets[:100]
classifier = nltk.NaiveBayesClassifier.train(train_set)
print(nltk.classify.accuracy(classifier, test_set))
classifier.show_most_informative_features(10)
# -
# NLTK gives us 88% accuracy, which isn't bad, but our home-made naive algorithm also achieved a respectable 80%.
#
#
# What improvements could we have made? Obviously, we could have used more data, or – in our home-grown model select words that discriminate between good and bad reviews. We could have used n-grams, e.g., to catch "not bad" as a postitive sentiment.
| 20-nlp_regex/lecture-20-NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process fgmax grid results and plot
#
# To process fgmax results after doing a run.
# + tags=["hide-py"]
# %matplotlib inline
# -
from pylab import *
import os,sys
import glob
from importlib import reload
from clawpack.geoclaw import topotools, dtopotools
from clawpack.visclaw import colormaps
from scipy.interpolate import RegularGridInterpolator
import matplotlib as mpl
from matplotlib import colors
sys.path.insert(0,'../../new_python')
import region_tools, plottools
import fgmax_tools, kmltools
# ## Set some things...
#
# Specify the directory to read results from, and some other settings:
# +
save_figs = False # make png files for figures?
make_new_fgmax_nc_file = True # make new netcdf file of fgmax results (possibly overwrites)?
rundir = os.path.abspath('.')
outdir = os.path.join(rundir, '_output')
print('Will read fgmax results from outdir = \n ', outdir)
# +
use_force_dry = True
if use_force_dry:
fname_force_dry = os.path.join(rundir, 'input_files', 'force_dry_init.data')
adjust_by_dz = True
if adjust_by_dz:
dtopo_path = os.path.join(rundir, 'input_files', 'SFL.tt3')
# -
def savefigp(fname):
global save_figs
if save_figs:
savefig(fname)
print('Created ', fname)
else:
print('save_figs = False')
# ## Read in and process the fgmax results from the latest run
#
t_files = glob.glob(outdir + '/fort.t0*')
times = []
for f in t_files:
lines = open(f,'r').readlines()
for line in lines:
if 'time' in line:
t = float(line.split()[0])
times.append(t)
times.sort()
print('Output times found: ',times)
if len(times) > 0:
t_hours = times[-1] / 3600.
print('\nfgmax results are presumably from final time: %.1f seconds = %.2f hours'\
% (times[-1], t_hours))
else:
t_hours = nan
# +
# Read fgmax data:
fg = fgmax_tools.FGmaxGrid()
fgmax_input_file_name = outdir + '/fgmax_header.data'
print('fgmax input file: \n %s' % fgmax_input_file_name)
fg.read_input_data(fgmax_input_file_name)
fg.read_output(outdir=outdir)
xx = fg.X
yy = fg.Y
# +
# convert to masked array on uniform grid for .nc file and plots:
fgm = fgmax_tools.FGmaxMaskedGrid()
dx = dy = 1./(3*3600.) # For 1/3 arcsecond fgmax grid
# convert to arrays and create fgm.X etc.
fgm.convert_lists_to_arrays(fg,dx,dy)
# 1d versions of X and Y arrays:
fgm.x = fgm.X[0,:]
fgm.y = fgm.Y[:,0]
# +
# compute subsidence/uplift at each fgmax point:
if adjust_by_dz:
dtopo = dtopotools.DTopography()
dtopo.read(dtopo_path, dtopo_type=3)
x1d = dtopo.X[0,:]
y1d = dtopo.Y[:,0]
dtopo_func = RegularGridInterpolator((x1d,y1d), dtopo.dZ[-1,:,:].T,
method='linear', bounds_error=False, fill_value=0.)
dz = dtopo_func(list(zip(ravel(fgm.X), ravel(fgm.Y))))
fgm.dz = reshape(dz, fgm.X.shape)
print('Over fgmax extent, min(dz) = %.2f m, max(dz) = %.2f m' \
% (dz.min(), dz.max()))
else:
fgm.dz = zeros(fgm.X.shape)
fgm.B0 = fgm.B - fgm.dz # original topo before subsidence/uplift
# +
if use_force_dry:
print('Reading force_dry from ',fname_force_dry)
force_dry = topotools.Topography()
force_dry.read(fname_force_dry, topo_type=3)
i1 = int(round((fgm.x[0]-force_dry.x[0])/dx))
i2 = int(round((fgm.x[-1]-force_dry.x[0])/dx))
j1 = int(round((fgm.y[0]-force_dry.y[0])/dy))
j2 = int(round((fgm.y[-1]-force_dry.y[0])/dy))
if (i1<0) or (i2-i1+1 != len(fgm.x)) or \
(j1<0) or (j2-j1+1 != len(fgm.y)):
print('*** force_dry does not cover fgm extent, not using')
use_force_dry = False
fgm.force_dry_init = None
else:
fgm.force_dry_init = force_dry.Z[j1:j2+1, i1:i2+1]
else:
fgm.force_dry_init = None
print('*** use_force_dry is False')
if fgm.force_dry_init is not None:
fgm.h_onshore = ma.masked_where(fgm.force_dry_init==0, fgm.h)
else:
fgm.h_onshore = ma.masked_where(fgm.B0 < 0., fgm.h)
# -
print('Number of fgmax points: ', fgm.h.count())
# +
zmin = -60.
zmax = 40.
land_cmap = colormaps.make_colormap({ 0.0:[0.1,0.4,0.0],
0.25:[0.0,1.0,0.0],
0.5:[0.8,1.0,0.5],
1.0:[0.8,0.5,0.2]})
sea_cmap = colormaps.make_colormap({ 0.0:[0,0,1], 1.:[.8,.8,1]})
cmap, norm = colormaps.add_colormaps((land_cmap, sea_cmap),
data_limits=(zmin,zmax),
data_break=0.)
def plotZ(Z, show_cb=True):
pc = plottools.pcolorcells(fgm.X, fgm.Y, Z, cmap=cmap, norm=norm)
if show_cb:
cb = colorbar(pc,shrink=0.5)
cb.set_label('meters')
#axis([-122.76,-122.525,47.95,48.2])
gca().set_aspect(1./cos(48*pi/180.))
ticklabel_format(useOffset=False)
xticks(rotation=20);
figure(figsize=(10,6))
subplot(121)
plotZ(fgm.B, show_cb=False)
title('GeoClaw B');
if fgm.force_dry_init is not None:
print('Found force_dry_init array')
subplot(122)
mask_all_but_dryneg = logical_or(logical_or(fgm.B.mask,
logical_not(fgm.force_dry_init)),
fgm.B0>0)
B_dryneg = ma.masked_array(fgm.B.data, mask=mask_all_but_dryneg)
plotZ(fgm.B, show_cb=False)
sea_cmap_dry = colormaps.make_colormap({ 0.0:[1.0,0.6,0.6], 1.:[1.0,0.6,0.6]})
cmap_dry, norm_dry = colormaps.add_colormaps((land_cmap, sea_cmap_dry),
data_limits=(zmin,zmax),
data_break=0.)
B0_dryneg = ma.masked_array(fgm.B0.data, mask=mask_all_but_dryneg)
plottools.pcolorcells(fgm.X, fgm.Y, B0_dryneg, cmap=cmap_dry, norm=norm_dry)
title('B0, with dry regions below MHW pink')
savefigp('geoclaw_topo_and_dry.png')
else:
print('No force_dry_init array')
# -
# In the plot above, "GeoClaw B" refers to the cell-averaged topography value used by GeoClaw and stored with the fgmax output, and is generally recorded after any subsidence/uplift. The colors are blues for values of $B < 0$ and greens/brown for $B > 0$. If there's a plot on the right, it shows as pink any areas that were initialized as dry in spite of having $B_0 < 0$, where $B_0$ is the initial topography ($B$ corrected by $dz$).
# ## Plot maximum flow depth
# +
bounds_depth = array([1e-6,0.25,0.5,0.75,1,1.25,1.5])
#bounds_depth = array([1e-6,0.5,1.0,1.5,2,2.5,3.0])
cmap_depth = colors.ListedColormap([[.7,.7,1],[.5,.5,1],[0,0,1],\
[1,.7,.7], [1,.4,.4], [1,0,0]])
# Set color for value exceeding top of range to purple:
cmap_depth.set_over(color=[1,0,1])
# Set color for land points without inundation to light green:
cmap_depth.set_under(color=[.7,1,.7])
norm_depth = colors.BoundaryNorm(bounds_depth, cmap_depth.N)
figure(figsize=(8,8))
pc = plottools.pcolorcells(fgm.X, fgm.Y, fgm.h_onshore, cmap=cmap_depth, norm=norm_depth)
cb = colorbar(pc, extend='max', shrink=0.7)
cb.set_label('meters')
contour(fgm.X, fgm.Y, fgm.B0, [0], colors='g')
gca().set_aspect(1./cos(48*pi/180.))
ticklabel_format(useOffset=False)
xticks(rotation=20)
title('Maximum flow depth over %.2f hours' % t_hours)
savefigp('h_onshore.png')
# -
# In the plot above, green shows fgmax points that never got wet. The green contour shows `B0 = 0`, and note that some of the initially dry region below MHW never got wet (over the limited duration of this simulation).
#
# White areas are masked out either because they were not fgmax points or because they were initially wet.
#
# Regions colored blue or red are initially dry fgmax points that did get wet during the tsunami, with color showing the maximum depth of water recorded.
# ## Plot maximum speed
# +
bounds_speed = np.array([1e-6,0.5,1.0,1.5,2,2.5,3,4.5,6])
cmap_speed = mpl.colors.ListedColormap([[.9,.9,1],[.6,.6,1],\
[.3,.3,1],[0,0,1], [1,.8,.8],\
[1,.6,.6], [1,.3,.3], [1,0,0]])
# Set color for value exceeding top of range to purple:
cmap_speed.set_over(color=[1,0,1])
# Set color for land points without inundation to light green:
cmap_speed.set_under(color=[.7,1,.7])
norm_speed = colors.BoundaryNorm(bounds_speed, cmap_speed.N)
figure(figsize=(8,8))
pc = plottools.pcolorcells(fgm.X, fgm.Y, fgm.s, cmap=cmap_speed, norm=norm_speed)
cb = colorbar(pc, extend='max', shrink=0.7)
cb.set_label('m/s')
contour(fgm.X, fgm.Y, fgm.B0, [0], colors='g')
gca().set_aspect(1./cos(48*pi/180.))
ticklabel_format(useOffset=False)
xticks(rotation=20)
title('Maximum speed over %.2f hours' % t_hours)
savefigp('speed.png')
# -
# The plot above shows the maximum speed at each fgmax point. The points colored green remained dry over this simulation. The green contour shows `B0 = 0`.
#
# White areas are masked out because they were not fgmax points. Regions colored blue or red are either offshore (initially wet) or onshore points that got wet, colored by the maximum water speed $s = \sqrt{u^2 + v^2}$ over the simulation.
# ## Plots for Google Earth overlays
#
# Tne new version of `kmltools` includes some tools to make png files that display properly on Google Earth. The png files have no axes and have the dimension and dpi set properly so that there is an integer number of pixels in each grid cell so cell edges are sharp when zooming in.
#
# We make three png files and then make a kml file that can be used to open all three.
kml_dir = 'fgmax_results_kmlfiles'
os.system('mkdir -p %s' % kml_dir)
print('Will put png and kml files in %s' % kml_dir)
h_wet_onshore = ma.masked_where(fgm.h_onshore==0., fgm.h_onshore)
png_filename=kml_dir+'/h_onshore_max_for_kml.png'
fig,ax,png_extent,kml_dpi = kmltools.pcolorcells_for_kml(fgm.x, fgm.y, h_wet_onshore,
png_filename=png_filename,
dpc=2, cmap=cmap_depth, norm=norm_depth)
speed = ma.masked_where(fgm.h==0., fgm.s)
png_filename = '%s/speed_max_for_kml.png' % kml_dir
fig,ax,png_extent,kml_dpi = kmltools.pcolorcells_for_kml(fgm.x, fgm.y, speed,
png_filename=png_filename,
dpc=2, cmap=cmap_speed, norm=norm_speed)
stays_dry = ma.masked_where(fgm.h>0., fgm.h)
png_filename = '%s/stays_dry_for_kml.png' % kml_dir
fig,ax,png_extent,kml_dpi = kmltools.pcolorcells_for_kml(fgm.x, fgm.y, stays_dry,
png_filename=png_filename,
dpc=2, cmap=cmap_speed, norm=norm_speed)
# ### Make the kml file to display these three png files
#
# Then you can open `fgmax_results_kmlfiles/fgmax_results.kml` in Google Earth to view them.
# +
png_files=['h_onshore_max_for_kml.png', 'speed_max_for_kml.png','stays_dry_for_kml.png']
png_names=['max depth onshore','max speed','stays dry']
kmltools.png2kml(png_extent, png_files=png_files, png_names=png_names,
name='fgmax_results',
fname='%s/fgmax_results.kml' % kml_dir,
radio_style=False)
print('Contents of %s:' % kml_dir)
for f in glob.glob('%s/*' % kml_dir):
print(' ',f)
# -
# ## Make colorbars for kml files
reload(kmltools)
kmltools.kml_build_colorbar('%s/colorbar_depth.png' % kml_dir, cmap_depth,
norm=norm_depth, label='meters', title='depth')
kmltools.kml_build_colorbar('%s/colorbar_speed.png' % kml_dir, cmap_speed,
norm=norm_speed, label='meters / second', title='speed')
# +
png_files=['h_onshore_max_for_kml.png', 'speed_max_for_kml.png','stays_dry_for_kml.png']
png_names=['max depth onshore','max speed','stays dry']
cb_files = ['colorbar_depth.png', 'colorbar_speed.png']
cb_names = ['depth colorbar', 'speed colorbar']
kmltools.png2kml(png_extent, png_files=png_files, png_names=png_names,
name='fgmax_results',
fname='%s/fgmax_results.kml' % kml_dir,
radio_style=False,
cb_files=cb_files, cb_names=cb_names)
print('Contents of %s:' % kml_dir)
for f in glob.glob('%s/*' % kml_dir):
print(' ',f)
# -
# ## Write output to netCDF file
# The next cell will add fgmax results to a copy of the `input.nc` file containing input data that was created by [MakeInputFiles_Whidbey1.ipynb](MakeInputFiles_Whidbey1.ipynb). The `input.nc` file contains the original topo values as well as the `fgmax_point` array that indicates which of these are fgmax points.
#
# The new file is called `results.nc`.
#
# **WARNING:** This will overwrite any previous `results.nc` file!
fname = 'results.nc'
fname_input = 'input.nc'
print('Copying %s to %s' % (fname_input, fname))
os.system('cp %s %s' % (fname_input, fname))
fgmax_tools.write_nc_output(fname, fgm, outdir=outdir, force=False)
| examples/geoclaw_whidbey1/process_fgmax_whidbey1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Naver 쇼핑 수집한 내용 불러오기**
# ## **1 크롤링 수집함수**
# +
# Web Crawler 함수
from urllib import parse
from urllib import parse, request
def crawler_naver(url):
browser = "Mozilla/5.0 (X11; Linux i686) AppleWebKit/537.17 (KHTML, like Gecko) Chrome/24.0.1312.27 Safari/537.17"
return request.urlopen(request.Request(url, headers = {'User-Agent':browser})).read().decode('utf8')
# -
# ## **2 Loading 카테고리**
# 네이버 쇼핑 카테고리 목록 by **[Join inner list](https://stackoverflow.com/questions/716477/join-list-of-lists-in-python)**
# ```python
# import itertools
# a = [['a','b'], ['c']]
# print(list(itertools.chain.from_iterable(a)))
# ```
import pandas as pd
import itertools
category_dict = pd.read_csv('./data/categoty_naver.csv').to_dict(orient='list')
category_dict = {k: [_ for _ in items if str(_) != 'nan'] for k, items in category_dict.items()}
category_values = list(itertools.chain.from_iterable(category_dict.values()))
print(f"total_length : {len(category_values)} 개")
# ## **3 Maulo Item Result**
# 마을로 현재 Item 내용을 활용하여 수집한 내용
# ```python
# # load and uncompress.
# with gzip.open('./data/mauloItemNaver.pickle','rb') as f:
# items_maulo_category = pickle.load(f)
# ```
import pickle
with open('./data/category_Naver.pickle', 'rb') as f:
items_maulo_category = pickle.load(f)
print("loading is done.")
# %whos | grep list & str & dict
# <br/>
#
# # **Category Counting**
# ## **1 Category Matching & Counting**
items_count = {}
# Counting the matched items
for k,v in items_maulo_category.items():
mathed = False
for _ in v.split('>'):
if _ in category_values:
if _ not in list(items_count.keys()):
items_count[_] = 1; matched = True
#items_count[_] = [k]
else:
items_count[_] += 1; matched = True
#items_count[_].append(k)
if matched == False:
print(k)
len(items_count)
# ## **2 Category Matching Token List**
# Token list extracted
# +
items_token_list = {}
# Counting the matched items
for k,v in items_maulo_category.items():
mathed = False
for _ in v.split('>'):
if _ in category_values:
if _ not in list(items_token_list.keys()):
items_token_list[_] = [k]; matched = True
else:
items_token_list[_].append(k); matched = True
if matched == False:
print(k)
# "수산" Key Items List
">> ".join(items_token_list['수산'])
# -
items_token_list['모<PASSWORD>']
# ## **3 ReNewal the Category Table**
# matching the table
#
# - [dict result to Pandas Dataframe](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.from_dict.html)
# +
result = {}
for k, v_list in category_dict.items():
v_list_new = []
marker = False
# shopping item list loop
for _key in v_list:
# matching the count result roop
for _k, _v in items_count.items():
if _k == _key: _key = f"({_v}){_key}" # Adding the count number
else: pass
v_list_new.append(_key)
result[k] = v_list_new
">> ".join(result["식품"])
# -
# dict result to Pandas DataFrame Table
result_df = pd.DataFrame.from_dict(result, orient='index').T.fillna('')
result_df.to_csv('./data/categoty_naver_count.csv', index=None)
result_df.head()
| jupyters/Kj_naver2_preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
import json
import pandas as pd
import numpy as np
import time
import re
#import psycopg2
#import psycopg2-binary
from sqlalchemy import create_engine
from config import db_password
# 1. Create a function that takes in three arguments;
# Wikipedia data, Kaggle metadata, and MovieLens rating data (from Kaggle)
def extract_transform_load():
file_dir = "./Resources"
# 2. Read in the kaggle metadata and MovieLens ratings CSV files as Pandas DataFrames.
kaggle_file = pd.read_csv(f'{file_dir}/movies_metadata.csv', low_memory=False)
ratings_file = pd.read_csv(f'{file_dir}/ratings.csv')
# 3. Open the read the Wikipedia data JSON file.
with open(f'{file_dir}/wikipedia-movies.json', mode='r') as file:
wiki_movies_raw = json.load(file)
# 4. Read in the raw wiki movie data as a Pandas DataFrame.
wiki_movies_df = pd.DataFrame(wiki_movies_raw)
# 5. Return the three DataFrames
return wiki_movies_df, kaggle_file, ratings_file
# 6 Create the path to your file directory and variables for the three files.
file_dir = "./Resources"
# Wikipedia data
wiki_file = f'{file_dir}/wikipedia-movies.json'
# Kaggle metadata
kaggle_file = f'{file_dir}/movies_metadata.csv'
# MovieLens rating data.
ratings_file = f'{file_dir}/ratings.csv'
# 7. Set the three variables in Step 6 equal to the function created in Step 1.
wiki_file, kaggle_file, ratings_file = extract_transform_load()
# 8. Set the DataFrames from the return statement equal to the file names in Step 6.
#wiki_movies_df = wiki_file
#import pandas as pd
wiki_movies_df = pd.DataFrame(wiki_file)
# +
#kaggle_metadata = kaggle_file
kaggle_metadata = pd.DataFrame(kaggle_file)
#ratings = ratings_file
ratings = pd.DataFrame(ratings_file)
# -
# 9. Check the wiki_movies_df DataFrame.
wiki_movies_df.head(5)
# 10. Check the kaggle_metadata DataFrame.
kaggle_metadata.head()
# 11. Check the ratings DataFrame.
ratings.head()
| ETL_Deliverable1_function_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import sklearn
import seaborn as sn
import time
import matplotlib.pyplot as plt
from collections import defaultdict
from collections import Counter
from imblearn.under_sampling import RandomUnderSampler
from sklearn.metrics import confusion_matrix,classification_report
from sklearn.neighbors import KNeighborsClassifier
from sklearn import metrics
from sklearn.metrics import accuracy_score
from sklearn.dummy import DummyClassifier
from sklearn.linear_model import LogisticRegression
X_train = pd.read_csv("train_features.csv")
y_train = pd.read_csv("train_labels.csv")
X_valid = pd.read_csv("valid_features.csv")
y_valid = pd.read_csv("valid_labels.csv")
X_test = pd.read_csv("test_features.csv")
y_train = y_train["genre"]
y_valid = y_valid["genre"]
# +
###visualize labels
def count_inst(filestream, class_dict):
n_instances = 0
for line in filestream.readlines()[1:]:
n_instances += 1
class_dict[line.strip().split(",")[-1]] += 1
return n_instances
f = open("train_labels.csv",'r')
n_instances = 0
class_dict = defaultdict(int)
class_name = []
class_num = []
print('Our total number of instances is:',count_inst(f,class_dict))
for lbl in class_dict.keys():
class_name.append(lbl)
class_num.append(class_dict[lbl])
print('For class', lbl, 'we have', class_dict[lbl], 'instances.')
plt.figure(1, figsize=(15,5))
plt.title("labels in all training instance")
plt.bar(class_name, class_num, color = ["tomato", "cornflowerblue"])
# -
###visulize "mode" feature
X_train["mode"].value_counts().plot(kind='barh',title= "mode count", color = ["tomato", "cornflowerblue"])
###pre-processing
def undersampling (X_train, y_train): # fit with only training data
rus = RandomUnderSampler(random_state=42, replacement=True)
X_train, y_train = rus.fit_resample(X_train, y_train)
return X_train, y_train
def category_1 (X): #select both continuous and discrete data
X = X.drop(["trackID", "title", "tags"], axis = 1)
return X
def category_2 (X): #only continuous data
X =X.drop(["trackID", "title", "tags","time_signature","key","mode"], axis = 1)
return X
# +
###train function
def train_clf(clf, X_train, y_train):
start = time.time()
clf.fit(X_train, y_train)
end = time.time()
print("This model is trained in {:.4f} seconds".format(end-start))
###classifiers
Zero_R_clf = DummyClassifier(strategy='most_frequent')
lgr = LogisticRegression(solver="lbfgs", multi_class="auto",max_iter=10000)
knn = KNeighborsClassifier()
#optimal k for knn classifier
def optimal_k (X_train, y_train, X_valid, y_valid):
scores = {}
scores_list = []
start = time.time()
for k in range(1,60):
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train, y_train)
preds = knn.predict(X_valid)
scores[k] = metrics.accuracy_score(y_valid, preds)
scores_list.append(metrics.accuracy_score(y_valid, preds))
max_k = max(scores, key=scores.get)
end = time.time()
print("optimal k is found in {:.4f} seconds".format(end-start))
return max_k
# -
###evaluate function
def evaluate_clf(clf, X_train, y_train, X_valid, y_valid):
train_clf(clf, X_train, y_train)
print("Accuracy for training set: ", clf.score(X_train, y_train))
#print('training set confusion matrix:\n',confusion_matrix(y_train,clf.predict(X_train)))
print('training set report:\n',classification_report(y_train,clf.predict(X_train)))
print("Accuracy for validation set: ", clf.score(X_valid, y_valid))
print('validation set confusion matrix:')
df_valid_cm = pd.DataFrame(confusion_matrix(y_valid,clf.predict(X_valid)))
sn.set(font_scale=1.4)
valid_cm = sn.heatmap(df_valid_cm, annot=True, annot_kws={"size":16}, fmt="d",
xticklabels = ["classic pop and rock","dance and electronica", "folk",
"jazz and blues","metal","pop","punk","sould and raggae"],
yticklabels = ["classic pop and rock","dance and electronica", "folk",
"jazz and blues","metal","pop","punk","sould and raggae"])
plt.show(valid_cm)
#print('validation set confusion matrix:\n',confusion_matrix(y_valid,clf.predict(X_valid)))
print('validation set report:\n',classification_report(y_valid,clf.predict(X_valid)))
###predict function
def predict_test(clf, X_test):
start = time.time()
y_test = clf.predict(X_test)
end = time.time()
return y_test
# +
#(model 1) classify data without sampling using category 1 feature with standization
X_train_full_cate1 = category_1 (X_train)
X_valid_full_cate1 = category_1 (X_valid)
X_test_full_cate1 = category_1 (X_test)
y_train_full_cate1 = y_train
y_valid_full_cate1 = y_valid
scaler = sklearn.preprocessing.StandardScaler(copy=False)
X_train_full_cate1 = scaler.fit_transform(X_train_full_cate1)
X_valid_full_cate1 = scaler.transform(X_valid_full_cate1)
X_test_full_cate1 = scaler.transform(X_test_full_cate1)
print("#########Zero-R#########")
train_clf(Zero_R_clf, X_train_full_cate1, y_train_full_cate1)
print("Accuracy for training set: ", Zero_R_clf.score(X_train_full_cate1, y_train_full_cate1))
print("Accuracy for validation set: ", Zero_R_clf.score(X_valid_full_cate1, y_valid_full_cate1))
y_test_full_cate1_ZeroR = predict_test(Zero_R_clf, X_test_full_cate1)
print("\n#########logistic regression#########")
evaluate_clf(lgr, X_train_full_cate1, y_train_full_cate1, X_valid_full_cate1, y_valid_full_cate1)
y_test_full_cate1_lgr = predict_test(lgr, X_test_full_cate1)
print("\n#########K-Nearest Neighbour#########")
max_k = optimal_k(X_train_full_cate1, y_train_full_cate1, X_valid_full_cate1, y_valid_full_cate1)
knn = KNeighborsClassifier(n_neighbors=max_k)
print("optimal value for K:", max_k)
evaluate_clf(knn, X_train_full_cate1, y_train_full_cate1, X_valid_full_cate1, y_valid_full_cate1)
y_test_full_cate1_knn = predict_test(knn, X_test_full_cate1)
# +
#(model 2) classify data after sampling using category 1 feature with standization
X_train_sampling_cate1, y_train_sampling_cate1 = undersampling(X_train, y_train)
X_train_sampling_cate1 = category_1 (X_train_sampling_cate1)
X_valid_sampling_cate1 = category_1 (X_valid)
X_test_sampling_cate1 = category_1 (X_test)
y_valid_sampling_cate1 = y_valid
scaler = sklearn.preprocessing.StandardScaler(copy=False)
X_train_sampling_cate1 = scaler.fit_transform(X_train_sampling_cate1)
X_valid_sampling_cate1 = scaler.transform(X_valid_sampling_cate1)
X_test_sampling_cate1 = scaler.transform(X_test_sampling_cate1)
print("#########Zero-R#########")
train_clf(Zero_R_clf, X_train_sampling_cate1, y_train_sampling_cate1)
print("Accuracy for training set: ", Zero_R_clf.score(X_train_sampling_cate1, y_train_sampling_cate1))
print("Accuracy for validation set: ", Zero_R_clf.score(X_valid_sampling_cate1, y_valid_sampling_cate1))
y_test_full_cate1_ZeroR = predict_test(Zero_R_clf, X_test_sampling_cate1)
print("\n#########logistic regression#########")
evaluate_clf(lgr, X_train_sampling_cate1, y_train_sampling_cate1, X_valid_sampling_cate1, y_valid_sampling_cate1)
y_test_sampling_cate1 = predict_test(lgr, X_test_sampling_cate1)
print("\n#########K-Nearest Neighbour#########")
max_k = optimal_k(X_train_sampling_cate1, y_train_sampling_cate1, X_valid_sampling_cate1, y_valid_sampling_cate1)
print("optimal value for K:", max_k)
knn = KNeighborsClassifier(n_neighbors=max_k)
evaluate_clf(knn, X_train_sampling_cate1, y_train_sampling_cate1, X_valid_sampling_cate1, y_valid_sampling_cate1)
y_test_sampling_cate1 = predict_test(knn, X_test_sampling_cate1)
# +
#(model 3) classify data without sampling using category 2 feature with standization
X_train_full_cate2 = category_2 (X_train)
X_valid_full_cate2 = category_2 (X_valid)
X_test_full_cate2 = category_2 (X_test)
y_train_full_cate2 = y_train
y_valid_full_cate2 = y_valid
scaler = sklearn.preprocessing.StandardScaler(copy=False)
X_train_full_cate2 = scaler.fit_transform(X_train_full_cate2)
X_valid_full_cate2 = scaler.transform(X_valid_full_cate2)
X_test_full_cate2 = scaler.transform(X_test_full_cate2)
print("#########Zero-R#########")
train_clf(Zero_R_clf, X_train_full_cate2, y_train_full_cate2)
print("Accuracy for training set: ", Zero_R_clf.score(X_train_full_cate2, y_train_full_cate2))
print("Accuracy for validation set: ", Zero_R_clf.score(X_valid_full_cate2, y_valid_full_cate2))
y_test_full_cate1_ZeroR = predict_test(Zero_R_clf, X_test_full_cate2)
print("\n#########logistic regression#########")
evaluate_clf(lgr, X_train_full_cate2, y_train_full_cate2, X_valid_full_cate2, y_valid_full_cate2)
y_test_full_cate2_lgr = predict_test(lgr, X_test_full_cate2)
print("\n#########K-Nearest Neighbour#########")
max_k = optimal_k(X_train_full_cate2, y_train_full_cate2, X_valid_full_cate2, y_valid_full_cate2)
knn = KNeighborsClassifier(n_neighbors=max_k)
print("optimal value for K:", max_k)
evaluate_clf(knn, X_train_full_cate2, y_train_full_cate2, X_valid_full_cate2, y_valid_full_cate2)
y_test_full_cate2_knn = predict_test(knn, X_test_full_cate2)
# +
#(model 4) classify data after sampling using category 2 feature with standization
X_train_sampling_cate2, y_train_sampling_cate2 = undersampling(X_train, y_train)
X_train_sampling_cate2 = category_2 (X_train_sampling_cate2)
X_valid_sampling_cate2 = category_2 (X_valid)
X_test_sampling_cate2 = category_2 (X_test)
y_valid_sampling_cate2 = y_valid
scaler = sklearn.preprocessing.StandardScaler(copy=False)
X_train_sampling_cate2 = scaler.fit_transform(X_train_sampling_cate2)
X_valid_sampling_cate2 = scaler.transform(X_valid_sampling_cate2)
X_test_sampling_cate2 = scaler.transform(X_test_sampling_cate2)
print("#########Zero-R#########")
train_clf(Zero_R_clf, X_train_sampling_cate2, y_train_sampling_cate2)
print("Accuracy for training set: ", Zero_R_clf.score(X_train_sampling_cate2, y_train_sampling_cate2))
print("Accuracy for validation set: ", Zero_R_clf.score(X_valid_sampling_cate2, y_valid_sampling_cate2))
y_test_full_cate1_ZeroR = predict_test(Zero_R_clf, X_test_sampling_cate2)
print("\n#########logistic regression#########")
evaluate_clf(lgr, X_train_sampling_cate2, y_train_sampling_cate2, X_valid_sampling_cate2, y_valid_sampling_cate2)
y_test_sampling_cate2 = predict_test(lgr, X_test_sampling_cate2)
y_test_sampling_cate2_lgr = predict_test(lgr, X_test_sampling_cate2)
print("\n#########K-Nearest Neighbour#########")
max_k = optimal_k(X_train_sampling_cate2, y_train_sampling_cate2, X_valid_sampling_cate2, y_valid_sampling_cate2)
print("optimal value for K:", max_k)
knn = KNeighborsClassifier(n_neighbors=max_k)
evaluate_clf(knn, X_train_sampling_cate2, y_train_sampling_cate2, X_valid_sampling_cate2, y_valid_sampling_cate2)
y_test_sampling_cate2_knn = predict_test(knn, X_test_sampling_cate2)
| peer_review/1/music genre codes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="0giL4C7FrgfE"
# # Generate Asimov's writing
# + [markdown] id="ZXbkVMMfro4B"
# ## Parameters
# + colab={"base_uri": "https://localhost:8080/"} id="6tLJIWJGrGZ1" outputId="56dddc19-09ad-4a0f-9b6d-dacf958f0ea6"
import os
BASE_DIR = '/Users/efraflores/Desktop/EF/Diplo/Asimov' #'/content' if you're using GColab
print(os.listdir(BASE_DIR))
SPLIT_PATTERN = "[\.\?\!]" #How you want to split your text into sentences
UNITS_LIST = [100,50] #List of units for the RNN
EMBEDDING_DIM = 300 #from (50,100,200,300) possible embeddings with GloVe
EPOCHS = 100
# + [markdown] heading_collapsed=true id="vI0mi651rj7P"
# ## Functions
# + hidden=true id="x2_LSo63re9m"
import time
import numpy as np
from IPython.lib.display import Audio
start = time.time()
def time_exp(x):
#Just print how many minutes and seconds have passed
minutes, seconds = np.floor(x/60), 60*(x/60-np.floor(x/60))
print(f"{'{:.0f}'.format(minutes)} minutos con {'{:.2f}'.format(seconds)} segundos")
def tone(a=1000, b=700, play_time_seconds=1, framerate=4410):
#Make a sound! Useful while training models
t = np.linspace(0, play_time_seconds, framerate*play_time_seconds)*np.pi
return Audio(np.sin(a*t)+np.sin(b*t), rate=framerate, autoplay=True)
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="zY5SJgAbr4mT" outputId="55b2616e-e9c9-4e0e-f183-09cfc8ff9604"
#Uncomment the following lines if it's the first time you run this packages
'''import nltk
nltk.download('stopwords')
nltk.download('wordnet')
'''
import re
import unicodedata
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
lem = WordNetLemmatizer()
def clean_text(text,
language='english',
pattern="[^a-zA-Z' ]",
lemma=False,
remove_stopwords=False,
add_stopwords=[]):
#It clean and can remove stopwords or even lemmatize words if specified in params
cleaned_text = unicodedata.normalize('NFD', str(text)).encode('ascii', 'ignore')
cleaned_text = re.sub(pattern,' ',cleaned_text.decode('utf-8'),flags=re.UNICODE)
cleaned_text = ' '.join([(lem.lemmatize(word,pos='v') if lemma else word) for word in
cleaned_text.lower().split() if word not in
(stopwords.words(language)+add_stopwords if remove_stopwords else '')])
return cleaned_text
#Ex
print('\nBasic cleaning:\t\t',clean_text("I am going to run!!! I ran while I was running??? ..."))
print('Changing the pattern:\t',clean_text("I am going to run!!! I ran while I was running??? ...",pattern="[^a-zA-Z!\.]"))
print('Without stopwords:\t',clean_text("I am going to run!!! I ran while I was running??? ...",remove_stopwords=True))
print('Lemmatizing verbs:\t',clean_text("I am going to run!!! I ran while I was running??? ...",lemma=True))
print('No stopwords and lemma:\t',clean_text("I am going to run!!! I ran while I was running??? ...",remove_stopwords=True,lemma=True))
print("\nIt actually corrects the weird accents, example\n\tFROM: Èl ÉfrâïsMa's?...\n\tTO:",clean_text("Èl ÉfrâïsMa's?..."))
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="Vi0DQoSio09f" outputId="bbf2dc14-73ed-4a6e-8daa-ab3f0a0e8c7f"
#Install this package to read PDFs
'''!pip install pyPDF2'''
import re
import os
import PyPDF2
import warnings
import pandas as pd
warnings.filterwarnings("ignore",message="Xref table not zero-indexed. ID numbers for objects will be corrected.")
def pdfs_to_sentences(base_dir,split_pattern,output_name='corpus.txt',**clean_args):
total_pages = []
#Loop over all pdf docs in the path provided
for pdf_file in [pdf for pdf in os.listdir(base_dir) if pdf[-4:]=='.pdf']:
#Read every page of a PDF file
pdf_reader = PyPDF2.PdfFileReader(open(os.path.join(base_dir,pdf_file),'rb'))
pdf_text = [pdf_reader.getPage(p).extractText() for p in range(pdf_reader.numPages)]
#Concat all pages for each file
total_pages.append(' '.join(pdf_text))
#Concat all pdf files as a string object and export it as txt
pd.DataFrame(' '.join(total_pages),
index=[''],columns=['']).to_csv(os.path.join(base_dir,output_name))
with open(os.path.join(base_dir,output_name)) as txt:
#Make sentences with the split-pattern provided
corpus = re.split(split_pattern,txt.read())
#Clean text with additional parameters if specified
corpus = [clean_text(x,**clean_args).strip() for x in corpus]
#Return just the non-empty sentences
return [x for x in corpus if len(x)>0]
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="PE9ffpNH7zwT" outputId="60940396-edbe-41b9-c0bf-58c1517ef1a0"
import tensorflow.keras.utils as keras_utils
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
def seq_builder(text,fitted_tokenizer):
seq = []
for sentence in text:
#Tokenize the words in every sentence
tokens = fitted_tokenizer.texts_to_sequences([sentence])[0]
for i in range(1,len(tokens)):
#Build sequences with all possible sizes
seq.append(tokens[:i+1])
#Same lenght for every sequence, filled by 0 to the left
seq = pad_sequences(seq,maxlen=max(map(len,seq)),padding='pre')
#Split the predictors and labels
X,y = seq[:,:-1], seq[:,-1]
#Encode all posible classes (words)
y = keras_utils.to_categorical(y)
return X,y
#Ex
aux = ['The same words','will have','the same representation in X','and the same vector','in Y']
aux_tok = Tokenizer()
aux_tok.fit_on_texts(aux)
aux = seq_builder(aux,aux_tok)
print(aux[0],'\n'*2,aux[1])
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="i8xeDVobu5kP" outputId="34625a21-33a6-42a6-9527-c947b9a86e0d"
#The tutorial for Transfer Embedding is right here!
# https://keras.io/examples/nlp/pretrained_word_embeddings/
#If you're running this in GColab, uncomment the following lines to unzip the GloVe embeddings
# (more info about GloVe at https://nlp.stanford.edu/projects/glove/),
# but if you're working locally, move the files from the zip to your base_dir
'''!wget http://nlp.stanford.edu/data/glove.6B.zip
!unzip -q glove.6B.zip'''
def transfer_embedding(base_dir,fitted_tokenizer,total_words,embedding_dim_out):
#Get the pre-trained vectors as a dict
emb_dict = {}
with open(os.path.join(base_dir,f'glove.6B.{embedding_dim_out}d.txt')) as f:
for line in f:
#Each line is a word with its coef sep by \s
word, coefs = line.split(maxsplit=1)
#Like .split() transforming strings into an array
coefs = np.fromstring(coefs,'f',sep=' ')
#Update the dict with a new word and its coef
emb_dict[word] = coefs
#Keep track of the not-found words
misses = 0
#Build a matrix full of zeros
words_matrix = np.zeros((total_words+1,embedding_dim_out))
#Loop over the words-indexes...
for word, i in fitted_tokenizer.word_index.items():
#... to get its pre-trained value
try: word_vector = emb_dict[word]
#Some words won't be found, keep the zeros-vector
except: misses += 1
#Finally, transfer the pre-trained vector
else: words_matrix[i] = word_vector
print(f'Transfered {round((1-misses/total_words)*100,2)}% of {total_words} words, just {misses} were not found.')
return words_matrix
# + hidden=true id="HHG3LHSPVOTv"
from keras.models import Sequential
from tensorflow.keras import regularizers
from tensorflow.keras.initializers import Constant
from keras.layers import Embedding,Bidirectional,GRU,LSTM,Dropout,Dense,LeakyReLU
def rnn_builder(x,total_words,units,embedding_dim_out,embedded_matrix,
drop_out=0.2,reg_l2=0.01,gru=False):
n = len(units)
rnn = Sequential()
#Transfer an embedded matrix
rnn.add(Embedding(total_words+1,
embedding_dim_out,
input_length=x.shape[-1], #max seq length
embeddings_initializer=Constant(embedded_matrix),
trainable=False)) #Do not change the pre-trained weights (duh!)
#Loop over the units-per-layer provided
for i,layer in enumerate(units):
#**kwarg to return sequence unless it's the last recurrent layer
seq = {'return_sequences':True if i < n-1 else False}
#Append one GRU/LSTM layer with the units from the iteration
rnn.add(Bidirectional((GRU(units=layer,**seq))
if gru else
(LSTM(units=layer,**seq))))
#Exclude % of previous layer to avoid overtfitting
rnn.add(Dropout(drop_out))
#Fully-Connected layer with half #classes and a "Lasso" regularization
rnn.add(Dense(int(total_words/2),kernel_regularizer=regularizers.l2(reg_l2)))
#Break linearity of previous layer
rnn.add(LeakyReLU())
#Final layer with the same units as #classes
rnn.add(Dense(total_words,activation='softmax'))
#Show full structure
print(rnn.summary())
return rnn
# + hidden=true id="7OqfWnA9JjvU"
import matplotlib.pyplot as plt
def plot_metrics(training_history,metrics=['accuracy','loss']):
#Show the trend of every metrics provided after training
for metric in metrics:
trend = training_history.history[metric]
plt.figure()
plt.plot(range(len(trend)), trend, 'r', label=f'Training {metric}')
plt.title(f'Training {metric}')
plt.show()
# + [markdown] heading_collapsed=true id="I7kohmntsTQX"
# ## Preprocessing
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="0W0o9RrusHbA" outputId="83c0d2ed-37a3-4bdf-a2b0-0ca844ef8138"
corpus = pdfs_to_sentences(BASE_DIR,SPLIT_PATTERN)
print(corpus[1:3])
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="iajukYWMxS1Q" outputId="bc2ce9ca-d3c1-4b21-e369-0e5f1c2b32db"
#Tokenize the words to feed the ANN
tokenizer = Tokenizer()
tokenizer.fit_on_texts(corpus)
tot_words = len(tokenizer.word_counts)
print(f'There are {tot_words} different words, the frequency of the first 6 is:',
'\n'*2,list(tokenizer.word_counts.items())[:6])
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="qG9jsEDTytG6" outputId="4f4a4e80-ee57-42c1-fc2f-d3ac1f68d99c"
X,y = seq_builder(corpus,tokenizer)
print(X.shape,y.shape)
# + colab={"base_uri": "https://localhost:8080/"} hidden=true id="50aaSwG0mu95" outputId="1a99ed0d-306e-46d3-8392-1b5ea5b16723"
words_matrix = transfer_embedding(BASE_DIR,tokenizer,tot_words+1,EMBEDDING_DIM)
# + [markdown] id="1j8c7eIeCfyU"
# ## Arquitecture
# + colab={"base_uri": "https://localhost:8080/"} id="1aUMaoMtRi-A" outputId="31f10e8b-025b-468e-b5b4-b06f3e9f3562"
rnn = rnn_builder(X,tot_words+1,UNITS_LIST,EMBEDDING_DIM,words_matrix,gru=True)
# + [markdown] heading_collapsed=true id="EnBqx3BYJWoX"
# ## Training
# + colab={"base_uri": "https://localhost:8080/", "height": 460} hidden=true id="Nlpkrrgy5ADw" outputId="3ef5a88c-6330-4cc8-8bff-a179b52484f3"
rnn.compile(loss='categorical_crossentropy', optimizer='adam', metrics='accuracy')
training = rnn.fit(X, y, epochs=EPOCHS, verbose=1)
# + [markdown] id="_m1FPPWdQn3I"
# ## Save the model
# + id="aY7VdRJaHXko"
plot_metrics(training)
# + id="kFjbbrBrQs_k"
import pickle
#First, preserve the tokenizer
with open(os.path.join(BASE_DIR,'rnn_asimov_tokenizer.pickle'), 'wb') as f:
pickle.dump(tokenizer, f)
#Then, save the architecture
with open(os.path.join(BASE_DIR,'rnn_asimov_architecture.json'), 'w') as f:
f.write(rnn.to_json())
#Finally, the paramaters learned by itself
rnn.save_weights(os.path.join(BASE_DIR,'rnn_asimov_weights.h5'))
# + id="EHUlARTrQwJq"
time_exp(time.time()-start)
tone()
| notebooks/01_Asimov_generator_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Hessian trace estimation
#
#
# This notebook shows how to efficiently approximate the trace of a Hessian, using automatic differentiation ([PyTorch](https://pytorch.org)) and a recently published randomized algorithm called [Hutch++](https://arxiv.org/abs/2010.09649).
import torch
torch.set_printoptions(precision=3)
class LinearOperator(object):
def __init__(self, matvec):
self._matvec = matvec
def matvec(self, vecs):
return self._matvec(vecs)
def hutchpp(A, d, m):
"""https://arxiv.org/abs/2010.09649
A is the LinearOperator whose trace to estimate
d is the input dimension
m is the number of queries (larger m yields better estimates)
"""
S = torch.randn(d, m // 3)
G = torch.randn(d, m // 3)
Q, _ = torch.qr(A.matvec(S))
proj = G - Q @ (Q.T @ G)
return torch.trace(Q.T @ A.matvec(Q)) + (3./m)*torch.trace(proj.T @ A.matvec(proj))
# +
torch.manual_seed(0)
d = 1000
B = torch.randn(d, d)
A = B.T @ B
torch.trace(A)
# -
m = 100
estimate = hutchpp(LinearOperator(lambda vec: A@vec), d, m)
estimate
percent_error = 100*(estimate - torch.trace(A)).abs() / torch.trace(A)
percent_error
def make_hvp(f, x):
def hvp(vecs):
# torch.autograd.functional.vhp doesn't support batching
vecs = torch.split(vecs, 1, dim=1)
products = []
for v in vecs:
_, product = torch.autograd.functional.vhp(f, x, v.squeeze())
products.append(product)
return torch.stack(products, dim=1)
return LinearOperator(hvp)
# +
def cubic(x):
return (x**3).mean()
x = torch.arange(5, dtype=torch.float)
hvp = make_hvp(cubic, x)
hessian = hvp.matvec(torch.eye(x.nelement()))
hessian
# -
x = torch.arange(10000, dtype=torch.float)
hvp = make_hvp(cubic, x)
# %time hessian = hvp.matvec(torch.eye(x.nelement()))
torch.trace(hessian)
# %time estimate = hutchpp(hvp, d=x.nelement(), m=100)
estimate
percent_error = 100*(estimate - torch.trace(hessian)).abs() / torch.trace(hessian)
percent_error
| hessian_trace_estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="1ZZSnDJ89qoW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6cff2348-de3a-46dd-e379-5eb19ef53a42" executionInfo={"status": "ok", "timestamp": 1581540566879, "user_tz": -60, "elapsed": 8491, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
# !pip install datadotworld
# !pip install datadotworld[pandas]
# + id="Y7i8IShwIvXK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="db7ead64-386b-416c-b4fa-a1e758bb244c" executionInfo={"status": "ok", "timestamp": 1581540399326, "user_tz": -60, "elapsed": 15128, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
# !dw configure
# + id="5jcmMUSsJ6qh" colab_type="code" colab={}
from google.colab import drive
import pandas as pd
import numpy as np
import datadotworld as dw
# + id="LI1qzAokKl5G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="de0baa46-151d-483f-b06a-38ea3801745d" executionInfo={"status": "ok", "timestamp": 1581541061048, "user_tz": -60, "elapsed": 32717, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
drive.mount("/content/drive")
# + id="D3BEyX7uMYEE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b0227dce-287f-40d5-f978-c3f43e0473d9" executionInfo={"status": "ok", "timestamp": 1581541101119, "user_tz": -60, "elapsed": 6492, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
# ls
# + id="OTDzADk4MtdA" colab_type="code" colab={}
# !git add .gitignore
# + id="A70xPoG_THQI" colab_type="code" colab={}
data = dw.load_dataset("datafiniti/mens-shoe-prices")
# + id="23ZB-mvtTfur" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7eacda74-f01c-4fc9-e83b-ec75f23c22d2" executionInfo={"status": "ok", "timestamp": 1581542983873, "user_tz": -60, "elapsed": 501, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
data
# + id="74l1HA5RTuWj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1573f5e9-9b59-4de9-e3ff-926bbb2e6479" executionInfo={"status": "ok", "timestamp": 1581543012323, "user_tz": -60, "elapsed": 1431, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
data.dataframes
# + id="74pSbhwAT2GX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5eecc4de-b961-4d3d-8c12-4a34ebcbc6f2" executionInfo={"status": "ok", "timestamp": 1581543068539, "user_tz": -60, "elapsed": 1444, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
df = data.dataframes['7004_1']
df.shape
# + id="LEgV65BiUCtw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="5172de85-a867-44a7-cd7d-5ba5a0c5e36f" executionInfo={"status": "ok", "timestamp": 1581543095324, "user_tz": -60, "elapsed": 970, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
df.head()
# + id="Fj1lFHMZUV3z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="580a42c9-ee3b-4f88-99ae-6664a1e0a360" executionInfo={"status": "ok", "timestamp": 1581543187202, "user_tz": -60, "elapsed": 922, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
df.prices_currency.unique()
# + id="cQtsrbu8UpGt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="0ff1e993-de10-4886-a43b-559b9dbee0b0" executionInfo={"status": "ok", "timestamp": 1581543277271, "user_tz": -60, "elapsed": 907, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
df.prices_currency.value_counts(normalize=True)
# + id="o9-2uEF5VBC4" colab_type="code" colab={}
df_usd = df[df.prices_currency == "USD"].copy()
# + id="SMDW33YkVnO9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="4383fdc2-1db2-4b5a-f6e5-650f0899d452" executionInfo={"status": "ok", "timestamp": 1581543585879, "user_tz": -60, "elapsed": 916, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
df_usd.prices_amountmin = df_usd.prices_amountmin.astype(np.float)
df_usd.prices_amountmin.hist()
# + id="BrVepY-vWJs7" colab_type="code" colab={}
filter_max = np.percentile(df_usd.prices_amountmin, 99)
# + id="fkWLyzJ1Wj8x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="d97f9ce8-ecae-46fd-831b-3da62a755fb5" executionInfo={"status": "ok", "timestamp": 1581543870544, "user_tz": -60, "elapsed": 1071, "user": {"displayName": "<NAME>\u00f3\u017cycki", "photoUrl": "", "userId": "08610380719257478006"}}
df_usd_filter = df_usd[df_usd.prices_amountmin < filter_max]
df_usd_filter.prices_amountmin.hist(bins=100)
# + id="mHJBdBBWsbzz" colab_type="code" colab={}
# !git add matrix_one/
# + id="z0Iyxu1UtCTA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="faa76878-cd8b-4d8a-81bc-38e79870b08d" executionInfo={"status": "ok", "timestamp": 1581549637025, "user_tz": -60, "elapsed": 3074, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08610380719257478006"}}
# !git status
| matrix_one/day_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo - Freeze Bayesian Neural Network
# +
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torchbnn as bnn
from torchbnn.utils import freeze, unfreeze
# -
import matplotlib.pyplot as plt
# %matplotlib inline
# ## 2. Define Model
model = nn.Sequential(
bnn.BayesLinear(prior_mu=0, prior_sigma=0.05, in_features=2, out_features=2),
nn.ReLU(),
bnn.BayesLinear(prior_mu=0, prior_sigma=0.05, in_features=2, out_features=1),
)
# ## 3. Forward Model
model(torch.ones(1, 2))
model(torch.ones(1, 2))
# ## 3. Freeze Model
freeze(model)
model(torch.ones(1, 2))
model(torch.ones(1, 2))
freeze(model)
model(torch.ones(1, 2))
model(torch.ones(1, 2))
# ## 4. Unfreeze Model
unfreeze(model)
model(torch.ones(1, 2))
model(torch.ones(1, 2))
| freeze_bnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# URL: http://matplotlib.org/examples/pie_and_polar_charts/polar_scatter_demo.html
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# ## Define data
# +
# Compute areas and colors
N = 150
r = 2 * np.random.rand(N)
theta = 2 * np.pi * np.random.rand(N)
colors = theta
scatter = hv.Scatter((theta, r), 'theta', 'r').redim(r=dict(range=(0,2.5)))
# -
# ## Plot
scatter.options(projection='polar', size_index='r', scaling_factor=50, color_index='theta',
cmap='hsv', alpha=0.75)
| examples/gallery/demos/matplotlib/polar_scatter_demo.ipynb |
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction #
#
# In the first two lessons, we learned how to build fully-connected networks out of stacks of dense layers. When first created, all of the network's weights are set randomly -- the network doesn't "know" anything yet. In this lesson we're going to see how to train a neural network; we're going to see how neural networks *learn*.
#
# As with all machine learning tasks, we begin with a set of training data. Each example in the training data consists of some features (the inputs) together with an expected target (the output). Training the network means adjusting its weights in such a way that it can transform the features into the target. In the *80 Cereals* dataset, for instance, we want a network that can take each cereal's `'sugar'`, `'fiber'`, and `'protein'` content and produce a prediction for that cereal's `'calories'`. If we can successfully train a network to do that, its weights must represent in some way the relationship between those features and that target as expressed in the training data.
#
# In addition to the training data, we need two more things:
# - A "loss function" that measures how good the network's predictions are.
# - An "optimizer" that can tell the network how to change its weights.
#
# # The Loss Function #
#
# We've seen how to design an architecture for a network, but we haven't seen how to tell a network *what* problem to solve. This is the job of the loss function.
#
# The **loss function** measures the disparity between the the target's true value and the value the model predicts.
#
# Different problems call for different loss functions. We have been looking at **regression** problems, where the task is to predict some numerical value -- calories in *80 Cereals*, rating in *Red Wine Quality*. Other regression tasks might be predicting the price of a house or the fuel efficiency of a car.
#
# A common loss function for regression problems is the **mean absolute error** or **MAE**. For each prediction `y_pred`, MAE measures the disparity from the true target `y_true` by an absolute difference `abs(y_true - y_pred)`.
#
# The total MAE loss on a dataset is the mean of all these absolute differences.
#
# <figure style="padding: 1em;">
# <img src="https://i.imgur.com/VDcvkZN.png" width="500" alt="A graph depicting error bars from data points to the fitted line..">
# <figcaption style="textalign: center; font-style: italic"><center>The mean absolute error is the average length between the fitted curve and the data points.
# </center></figcaption>
# </figure>
#
# Besides MAE, other loss functions you might see for regression problems are the mean-squared error (MSE) or the Huber loss (both available in Keras).
#
# During training, the model will use the loss function as a guide for finding the correct values of its weights (lower loss is better). In other words, the loss function tells the network its objective.
#
# # The Optimizer - Stochastic Gradient Descent #
#
# We've described the problem we want the network to solve, but now we need to say *how* to solve it. This is the job of the **optimizer**. The optimizer is an algorithm that adjusts the weights to minimize the loss.
#
# Virtually all of the optimization algorithms used in deep learning belong to a family called **stochastic gradient descent**. They are iterative algorithms that train a network in steps. One **step** of training goes like this:
# 1. Sample some training data and run it through the network to make predictions.
# 2. Measure the loss between the predictions and the true values.
# 3. Finally, adjust the weights in a direction that makes the loss smaller.
#
# Then just do this over and over until the loss is as small as you like (or until it won't decrease any further.)
#
# <figure style="padding: 1em;">
# <img src="https://i.imgur.com/rFI1tIk.gif" width="1600" alt="Fitting a line batch by batch. The loss decreases and the weights approach their true values.">
# <figcaption style="textalign: center; font-style: italic"><center>Training a neural network with Stochastic Gradient Descent.
# </center></figcaption>
# </figure>
#
# Each iteration's sample of training data is called a **minibatch** (or often just "batch"), while a complete round of the training data is called an **epoch**. The number of epochs you train for is how many times the network will see each training example.
#
# The animation shows the linear model from Lesson 1 being trained with SGD. The pale red dots depict the entire training set, while the solid red dots are the minibatches. Every time SGD sees a new minibatch, it will shift the weights (`w` the slope and `b` the y-intercept) toward their correct values on that batch. Batch after batch, the line eventually converges to its best fit. You can see that the loss gets smaller as the weights get closer to their true values.
#
# ## Learning Rate and Batch Size ##
#
# Notice that the line only makes a small shift in the direction of each batch (instead of moving all the way). The size of these shifts is determined by the **learning rate**. A smaller learning rate means the network needs to see more minibatches before its weights converge to their best values.
#
# The learning rate and the size of the minibatches are the two parameters that have the largest effect on how the SGD training proceeds. Their interaction is often subtle and the right choice for these parameters isn't always obvious. (We'll explore these effects in the exercise.)
#
# Fortunately, for most work it won't be necessary to do an extensive hyperparameter search to get satisfactory results. **Adam** is an SGD algorithm that has an adaptive learning rate that makes it suitable for most problems without any parameter tuning (it is "self tuning", in a sense). Adam is a great general-purpose optimizer.
#
# ## Adding the Loss and Optimizer ##
#
# After defining a model, you can add a loss function and optimizer with the model's `compile` method:
#
# ```
# model.compile(
# optimizer="adam",
# loss="mae",
# )
# ```
#
# Notice that we are able to specify the loss and optimizer with just a string. You can also access these directly through the Keras API -- if you wanted to tune parameters, for instance -- but for us, the defaults will work fine.
#
# <blockquote style="margin-right:auto; margin-left:auto; background-color: #ebf9ff; padding: 1em; margin:24px;">
# <strong>What's In a Name?</strong><br>
# The <strong>gradient</strong> is a vector that tells us in what direction the weights need to go. More precisely, it tells us how to change the weights to make the loss change <em>fastest</em>. We call our process gradient <strong>descent</strong> because it uses the gradient to <em>descend</em> the loss curve towards a minimum. <strong>Stochastic</strong> means "determined by chance." Our training is <em>stochastic</em> because the minibatches are <em>random samples</em> from the dataset. And that's why it's called SGD!
# </blockquote>
# # Example - Red Wine Quality #
#
# Now we know everything we need to start training deep learning models. So let's see it in action! We'll use the *Red Wine Quality* dataset.
#
# This dataset consists of physiochemical measurements from about 1600 Portuguese red wines. Also included is a quality rating for each wine from blind taste-tests. How well can we predict a wine's perceived quality from these measurements?
#
# We've put all of the data preparation into this next hidden cell. It's not essential to what follows so feel free to skip it. One thing you might note for now though is that we've rescaled each feature to lie in the interval $[0, 1]$. As we'll discuss more in Lesson 5, neural networks tend to perform best when their inputs are on a common scale.
# +
import pandas as pd
from IPython.display import display
red_wine = pd.read_csv('../input/dl-course-data/red-wine.csv')
# Create training and validation splits
df_train = red_wine.sample(frac=0.7, random_state=0)
df_valid = red_wine.drop(df_train.index)
display(df_train.head(4))
# Scale to [0, 1]
max_ = df_train.max(axis=0)
min_ = df_train.min(axis=0)
df_train = (df_train - min_) / (max_ - min_)
df_valid = (df_valid - min_) / (max_ - min_)
# Split features and target
X_train = df_train.drop('quality', axis=1)
X_valid = df_valid.drop('quality', axis=1)
y_train = df_train['quality']
y_valid = df_valid['quality']
# -
# How many inputs should this network have? We can discover this by looking at the number of columns in the data matrix. Be sure not to include the target (`'quality'`) here -- only the input features.
print(X_train.shape)
# Eleven columns means eleven inputs.
#
# We've chosen a three-layer network with over 1500 neurons. This network should be capable of learning fairly complex relationships in the data.
# +
from tensorflow import keras
from tensorflow.keras import layers
model = keras.Sequential([
layers.Dense(512, activation='relu', input_shape=[11]),
layers.Dense(512, activation='relu'),
layers.Dense(512, activation='relu'),
layers.Dense(1),
])
# -
# Deciding the architecture of your model should be part of a process. Start simple and use the validation loss as your guide. You'll learn more about model development in the exercises.
#
# After defining the model, we compile in the optimizer and loss function.
model.compile(
optimizer='adam',
loss='mae',
)
# Now we're ready to start the training! We've told Keras to feed the optimizer 256 rows of the training data at a time (the `batch_size`) and to do that 10 times all the way through the dataset (the `epochs`).
history = model.fit(
X_train, y_train,
validation_data=(X_valid, y_valid),
batch_size=256,
epochs=10,
)
# You can see that Keras will keep you updated on the loss as the model trains.
#
# Often, a better way to view the loss though is to plot it. The `fit` method in fact keeps a record of the loss produced during training in a `History` object. We'll convert the data to a Pandas dataframe, which makes the plotting easy.
# +
import pandas as pd
# convert the training history to a dataframe
history_df = pd.DataFrame(history.history)
# use Pandas native plot method
history_df['loss'].plot();
# -
# Notice how the loss levels off as the epochs go by. When the loss curve becomes horizontal like that, it means the model has learned all it can and there would be no reason continue for additional epochs.
# # Your Turn #
#
# Now, [**use stochastic gradient descent**](https://www.kaggle.com/kernels/fork/11887330) to train your network.
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/191966) to chat with other Learners.*
| Intro to Deep Learning/3 Stochastic Gradient Descent/stochastic-gradient-descent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Aquisition
# The Wikipedia traffic data must first be retrieved from the Legacy and Pageviews API endpoints. The Legacy API provides data for desktop and mobile traffic from December 2007 to July 2016. The Pageviews API provides data for desktop, mobile web, and mobile app traffic from July 2015 and on.
#
# This section creates 5 JSON files corresponding to each API call. The calls specify user traffic when possible.
import json
import requests
# +
endpoint_legacy = 'https://wikimedia.org/api/rest_v1/metrics/legacy/pagecounts/aggregate/{project}/{access-site}/{granularity}/{start}/{end}'
endpoint_pageviews = 'https://wikimedia.org/api/rest_v1/metrics/pageviews/aggregate/{project}/{access}/{agent}/{granularity}/{start}/{end}'
headers = {
'User-Agent': 'https://github.com/rileywaters',
'From': '<EMAIL>'
}
# -
def api_call(endpoint,parameters):
call = requests.get(endpoint.format(**parameters), headers=headers)
response = call.json()
return response
# #### File 1: Desktop Legacy
# +
desktop_params_legacy = {"project" : "en.wikipedia.org",
"access-site" : "desktop-site",
"granularity" : "monthly",
"start" : "2008010100",
"end" : "2016080100"
}
desktop_monthly_legacy = api_call(endpoint_legacy, desktop_params_legacy)
with open('legacy_desktop-site_200801-201608.json', 'w', encoding='utf-8') as f:
json.dump(desktop_monthly_legacy, f, ensure_ascii=False, indent=4)
# -
# #### FIle 2: Mobile Legacy
# +
mobile_params_legacy = {"project" : "en.wikipedia.org",
"access-site" : "mobile-site",
"granularity" : "monthly",
"start" : "2008010100",
"end" : "2016080100"
}
mobile_monthly_legacy = api_call(endpoint_legacy, mobile_params_legacy)
with open('legacy_mobile-site_200801-201608.json', 'w', encoding='utf-8') as f:
json.dump(mobile_monthly_legacy, f, ensure_ascii=False, indent=4)
# -
# #### File 3: Desktop PageViews
# +
desktop_params_pageviews = {"project" : "en.wikipedia.org",
"access" : "desktop",
"agent" : "user",
"granularity" : "monthly",
"start" : "2015070100",
"end" : '2019090100'}
desktop_monthly_pageviews = api_call(endpoint_pageviews, desktop_params_pageviews)
with open('pageviews_desktop-site_201507-201909.json', 'w', encoding='utf-8') as f:
json.dump(desktop_monthly_pageviews, f, ensure_ascii=False, indent=4)
# -
# #### File 4: Mobile App PageViews
# +
mobile_app_params_pageviews = {"project" : "en.wikipedia.org",
"access" : "mobile-app",
"agent" : "user",
"granularity" : "monthly",
"start" : "2015070100",
"end" : '2019090100'
}
mobile_app_monthly_pageviews = api_call(endpoint_pageviews, mobile_app_params_pageviews)
with open('pageviews_mobile-app_201507-201909.json', 'w', encoding='utf-8') as f:
json.dump(mobile_app_monthly_pageviews, f, ensure_ascii=False, indent=4)
# -
# #### File 5: Mobile Web PageViews
# +
mobile_web_params_pageviews = {"project" : "en.wikipedia.org",
"access" : "mobile-web",
"agent" : "user",
"granularity" : "monthly",
"start" : "2014100100",
"end" : '2019090100'
}
mobile_web_monthly_pageviews = api_call(endpoint_pageviews, mobile_web_params_pageviews)
with open('pageviews_mobile-web_201507-201909.json', 'w', encoding='utf-8') as f:
json.dump(mobile_web_monthly_pageviews, f, ensure_ascii=False, indent=4)
# -
# ## Processing
# The data files must now be processed into Pandas dataframes. The pageview mobile views are combined and the timestamps are separated into year and month. NA values are filled with 0.
#
# This step produces a csv file called en-wikipedia_traffic_200712-201809.csv
import pandas as pd
# +
# Read all the json files into pandas dataframes
df_legacy_mobile = pd.DataFrame(json.load(open('legacy_mobile-site_200801-201608.json', 'r'))['items'])
df_legacy_desktop = pd.DataFrame(json.load(open('legacy_desktop-site_200801-201608.json', 'r'))['items'])
df_pageviews_mobile_web = pd.DataFrame(json.load(open('pageviews_mobile-web_201507-201909.json', 'r'))['items'])
df_pageviews_mobile_app = pd.DataFrame(json.load(open('pageviews_mobile-app_201507-201909.json', 'r'))['items'])
df_pageviews_desktop = pd.DataFrame(json.load(open('pageviews_desktop-site_201507-201909.json', 'r'))['items'])
# +
# For data collected from the Pageviews API, combine the monthly values for mobile-app and mobile-web to create a total mobile traffic count for each month.
df_pageviews_mobile_web['views'] = df_pageviews_mobile_web['views'] + df_pageviews_mobile_app['views']
# rename the columns of each view count
df_pageviews_mobile = df_pageviews_mobile_web[['timestamp', 'views']].rename(columns={'views': 'pageview_mobile_views'})
df_pageviews_desktop = df_pageviews_desktop[['timestamp', 'views']].rename(columns={'views': 'pageview_desktop_views'})
df_legacy_mobile = df_legacy_mobile[['timestamp', 'count']].rename(columns={'count': 'pagecount_mobile_views'})
df_legacy_desktop = df_legacy_desktop[['timestamp', 'count']].rename(columns={'count': 'pagecount_desktop_views'})
# merge the dataframes
df_pageview_merge = df_pageviews_mobile.merge(df_pageviews_desktop, how='outer', on='timestamp')
df_legacy_merge = df_legacy_mobile.merge(df_legacy_desktop, how='outer', on='timestamp')
df_all = df_pageview_merge.merge(df_legacy_merge, how='outer', on='timestamp')
# fill na with 0
df_all = df_all.fillna(0)
# +
# Split the timestamp into year and month
df_all['timestamp'] = pd.to_datetime(df_all['timestamp'], format='%Y%m%d%H')
# Make a total views column for each API
df_all['pagecount_all_views'] = df_all['pagecount_mobile_views'] + df_all['pagecount_desktop_views']
df_all['pageview_all_views'] = df_all['pageview_mobile_views'] + df_all['pageview_desktop_views']
df_all['month'] = df_all['timestamp'].dt.month
df_all['year'] = df_all['timestamp'].dt.year
df_all = df_all.drop(['timestamp'], axis=1)
# save to csv
df_all.to_csv('en-wikipedia_traffic_200712-201809.csv', index=False)
# -
# ## Analysis
# Using the csv file, the counts are plotted as a time series. Dotted lines are used to indicate that the data is from the legacy system. Solid lines indicate that the data is from the newer API
from datetime import datetime
# +
# Read the csv and construct the timestamp
df_all = pd.read_csv('en-wikipedia_traffic_200712-201809.csv', sep=',')
df_all['timestamp'] = pd.to_datetime(df_all['year'].astype(str) + df_all['month'].astype(str), format='%Y%m')
df_all = df_all.sort_values(by='timestamp')
df_all.head()
# +
from matplotlib import pyplot as plt
from matplotlib import dates as pltdates
# Create the plot
formatter = pltdates.DateFormatter('%Y')
fig, ax = plt.subplots(1, 1, figsize=(15, 7))
ax.plot_date(df_all['timestamp'], df_all['pageview_mobile_views'].where(df_all['pageview_mobile_views']>0)/1e6, fmt='b-', label='_nolegend_')
ax.plot_date(df_all['timestamp'], df_all['pageview_desktop_views'].where(df_all['pageview_desktop_views']>0)/1e6, fmt='g-', label='_nolegend_')
ax.plot_date(df_all['timestamp'], df_all['pagecount_mobile_views'].where(df_all['pagecount_mobile_views']>0)/1e6, fmt='b--', label = 'mobile site')
ax.plot_date(df_all['timestamp'], df_all['pagecount_desktop_views'].where(df_all['pagecount_desktop_views']>0)/1e6, fmt='g--', label='main site')
ax.plot_date(df_all['timestamp'], df_all['pagecount_all_views'].where(df_all['pagecount_all_views']>0)/1e6, fmt='k--', label='total')
ax.plot_date(df_all['timestamp'], df_all['pageview_all_views'].where(df_all['pageview_all_views']>0)/1e6, fmt='k-', label='_nolegend_')
ax.xaxis.set_major_formatter(formatter)
ax.title.set_text('Page Views on English Wikipedia (x 1,000,000)')
ax.set_ylabel('Page Views')
ax.set_xlabel('Years')
ax.grid(True)
ax.legend(loc='upper left')
plt.show()
# Save the plot as png
fig.savefig('final_graph.png')
# -
| hcds-a1-data-curation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data reclassification
#
# Reclassifying data based on specific criteria is a common task when doing GIS analysis. The purpose of this lesson is to see how we can reclassify values based on some criteria which can be whatever, such as:
#
# ```
# 1. if travel time to my work is less than 30 minutes
#
# AND
#
# 2. the rent of the apartment is less than 1000 € per month
#
# ------------------------------------------------------
#
# IF TRUE: ==> I go to view it and try to rent the apartment
# IF NOT TRUE: ==> I continue looking for something else
# ```
#
# In this tutorial, we will:
#
# 1. Use classification schemes from the PySAL [mapclassify library](https://pysal.org/mapclassify/) to classify travel times into multiple classes.
#
# 2. Create a custom classifier to classify travel times and distances in order to find out good locations to buy an apartment with these conditions:
# - good public transport accessibility to city center
# - bit further away from city center where the prices are presumably lower
#
# ## Input data
#
# We will use [Travel Time Matrix data from Helsinki](https://blogs.helsinki.fi/accessibility/helsinki-region-travel-time-matrix/) that contains travel time and distance information for
# routes between all 250 m x 250 m grid cell centroids (n = 13231) in the Capital Region of Helsinki by walking, cycling, public transportation and car.
#
# In this tutorial, we will use the geojson file generated in the previous section:
# `"data/TravelTimes_to_5975375_RailwayStation_Helsinki.geojson"`
#
# Alternatively, you can re-download [L4 data](https://github.com/AutoGIS/data/raw/master/L4_data.zip) and use `"data/Travel_times_to_5975375_RailwayStation.shp"` as input file in here.
#
#
#
# ## Common classifiers
#
# ### Classification schemes for thematic maps
#
#
# [PySAL](http://pysal.readthedocs.io/en/latest) -module is an extensive Python library for spatial analysis. It also includes all of the most common data classifiers that are used commonly e.g. when visualizing data. Available map classifiers in [pysal's mapclassify -module](https://pysal.readthedocs.io/en/v1.11.0/library/esda/mapclassify.html):
#
# - Box_Plot
# - Equal_Interval
# - Fisher_Jenks
# - Fisher_Jenks_Sampled
# - HeadTail_Breaks
# - Jenks_Caspall
# - Jenks_Caspall_Forced
# - Jenks_Caspall_Sampled
# - Max_P_Classifier
# - Maximum_Breaks
# - Natural_Breaks
# - Quantiles
# - Percentiles
# - Std_Mean
# - User_Defined
#
# - First, we need to read our Travel Time data from Helsinki:
# +
import geopandas as gpd
fp = "data/TravelTimes_to_5975375_RailwayStation_Helsinki.geojson"
# Read the GeoJSON file similarly as Shapefile
acc = gpd.read_file(fp)
# Let's see what we have
print(acc.head(2))
# -
# As we can see, there are plenty of different variables (see [from here the description](http://blogs.helsinki.fi/accessibility/helsinki-region-travel-time-matrix-2015) for all attributes) but what we are interested in are columns called `pt_r_tt` which is telling the time in minutes that it takes to reach city center from different parts of the city, and `walk_d` that tells the network distance by roads to reach city center from different parts of the city (almost equal to Euclidian distance).
#
# **The NoData values are presented with value -1**.
#
# - Thus we need to remove the No Data values first.
#
# Include only data that is above or equal to 0
acc = acc.loc[acc['pt_r_tt'] >=0]
# - Let's plot the data and see how it looks like
# - `cmap` parameter defines the color map. Read more about [choosing colormaps in matplotlib](https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html)
# - `scheme` option scales the colors according to a classification scheme (requires `mapclassify` module to be installed):
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# Plot using 9 classes and classify the values using "Natural Breaks" classification
acc.plot(column="pt_r_tt", scheme="Natural_Breaks", k=9, cmap="RdYlBu", linewidth=0, legend=True)
# Use tight layout
plt.tight_layout()
# -
# As we can see from this map, the travel times are lower in the south where the city center is located but there are some areas of "good" accessibility also in some other areas (where the color is red).
#
# - Let's also make a plot about walking distances:
# +
# Plot walking distance
acc.plot(column="walk_d", scheme="Natural_Breaks", k=9, cmap="RdYlBu", linewidth=0, legend=True)
# Use tight layour
plt.tight_layout()
# -
# Okay, from here we can see that the walking distances (along road network) reminds more or less Euclidian distances.
#
# ### Applying classifiers to data
#
# As mentioned, the `scheme` option defines the classification scheme using `pysal/mapclassify`. Let's have a closer look at how these classifiers work.
import mapclassify
# - Natural Breaks
mapclassify.NaturalBreaks(y=acc['pt_r_tt'], k=9)
# - Quantiles (default is 5 classes):
mapclassify.Quantiles(y=acc['pt_r_tt'])
# - It's possible to extract the threshold values into an array:
classifier = mapclassify.NaturalBreaks(y=acc['pt_r_tt'], k=9)
classifier.bins
# - Let's apply one of the `Pysal` classifiers into our data and classify the travel times by public transport into 9 classes
# - The classifier needs to be initialized first with `make()` function that takes the number of desired classes as input parameter
# Create a Natural Breaks classifier
classifier = mapclassify.NaturalBreaks.make(k=9)
# - Now we can apply that classifier into our data by using `apply` -function
# +
# Classify the data
classifications = acc[['pt_r_tt']].apply(classifier)
# Let's see what we have
classifications.head()
# -
type(classifications)
# Okay, so now we have a DataFrame where our input column was classified into 9 different classes (numbers 1-9) based on [Natural Breaks classification](http://wiki-1-1930356585.us-east-1.elb.amazonaws.com/wiki/index.php/Jenks_Natural_Breaks_Classification).
#
# - We can also add the classification values directly into a new column in our dataframe:
# +
# Rename the column so that we know that it was classified with natural breaks
acc['nb_pt_r_tt'] = acc[['pt_r_tt']].apply(classifier)
# Check the original values and classification
acc[['pt_r_tt', 'nb_pt_r_tt']].head()
# -
# Great, now we have those values in our accessibility GeoDataFrame. Let's visualize the results and see how they look.
# +
# Plot
acc.plot(column="nb_pt_r_tt", linewidth=0, legend=True)
# Use tight layout
plt.tight_layout()
# +
# plot them side-by-side
# %matplotlib inline
import matplotlib.pyplot as plt
#basic config
fig, (ax1,ax2) = plt.subplots(nrows=1, ncols=2, figsize=(20, 16))
#ax1, ax2 = axes
#1st plot with 9 classes
ax1 = acc.plot(ax=ax1, column="nb_pt_r_tt", linewidth=0, legend=True)
ax1.set_title('Natural Breaks')
#2nd plot with original data
ax2 = acc.plot(ax=ax2, column="pt_r_tt", linewidth=0, legend=True)
ax2.set_title('Original')
fig.tight_layout()
# -
# And here we go, now we have a map where we have used one of the common classifiers to classify our data into 9 classes.
# ## Plotting a histogram
#
# A histogram is a graphic representation of the distribution of the data. When classifying the data, it's always good to consider how the data is distributed, and how the classification shceme divides values into different ranges.
#
# - plot the histogram using [pandas.DataFrame.plot.hist](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.plot.hist.html)
# - Number of histogram bins (groups of data) can be controlled using the parameter `bins`:
# Histogram for public transport rush hour travel time
acc['pt_r_tt'].plot.hist(bins=50)
# Let's also add threshold values on thop of the histogram as vertical lines.
#
# - Natural Breaks:
# +
# Define classifier
classifier = mapclassify.NaturalBreaks(y=acc['pt_r_tt'], k=9)
# Plot histogram for public transport rush hour travel time
acc['pt_r_tt'].plot.hist(bins=50)
# Add vertical lines for class breaks
for value in classifier.bins:
plt.axvline(value, color='k', linestyle='dashed', linewidth=1)
# -
# - Quantiles:
# +
# Define classifier
classifier = mapclassify.Quantiles(y=acc['pt_r_tt'])
# Plot histogram for public transport rush hour travel time
acc['pt_r_tt'].plot.hist(bins=50)
for value in classifier.bins:
plt.axvline(value, color='k', linestyle='dashed', linewidth=1)
# -
#
# <div class="alert alert-info">
#
# **Task**
#
# Select another column from the data (for example, travel times by car: `car_r_t`). Do the following visualizations using one of the classification schemes available from [pysal/mapclassify](https://github.com/pysal/mapclassify):
#
# - histogram with vertical lines showing the classification bins, `Equal_Interval`, `Fisher_Jenks`,
# `Fisher_Jenks_Sampled`, `HeadTail_Breaks`, `Jenks_Caspall`, `Jenks_Caspall_Forced`, `Jenks_Caspall_Sampled`, `Max_P_Classifier`, `Maximum_Breaks`, `Natural_Breaks`, `Quantiles`, `Percentiles`, `Std_Mean`.
# - thematic map using the classification scheme
#
#
# </div>
# +
import random
def classify_and_plot(gdf, classification = mapclassify.NaturalBreaks):
"""
A function that accepts a geodataframe; chooses a column and generates histogram and thematic map plots based on a
classification choice
"""
list_of_column_choices = ['car_m_t', 'car_r_t', 'from_id', 'pt_m_tt', 'pt_r_tt', 'walk_t']
random_column_choice = random.choice(list_of_column_choices)
#add classification to gpd
classifier = classification(y=acc[random_column_choice], k=7)
classifier = classification.make(k=7)
# add a new 'temp' column to store classified
gdf['temp'] = gdf[[random_column_choice]].apply(classifier)
#basic plot config
fig, (ax1,ax2, ax3) = plt.subplots(nrows=3, ncols=1, figsize=(15, 11))
#ax1, ax2 = axes
#1st plot with random classsification
ax2 = gdf.plot(ax=ax2, column='temp', linewidth=0, legend=True)
#set title
printTitle = ''
if random_column_choice == 'car_m_t':
printTitle = 'Travel time in minutes from origin to destination \nby private car in midday traffic'
if random_column_choice == 'car_r_t':
printTitle = 'Travel time in minutes from origin to destination \nby private car in rush hour traffic'
if random_column_choice == 'pt_m_tt':
printTitle = 'Travel time in minutes from origin to destination \nby public transportation in midday traffic'
if random_column_choice == 'pt_r_tt':
printTitle = 'Travel time in minutes from origin to destination \nby public transportation in rush hour traffic'
if random_column_choice == 'walk_t':
printTitle = 'Travel time in minutes from origin to destination \nby walking'
ax2.set_title(printTitle, pad=20)
#2nd plot with original data
ax1 = gdf.plot(ax=ax1, column="pt_r_tt", linewidth=0, legend=True)
ax1.set_title('Original')
# Plot histogram for public transport rush hour travel time
ax3 = gdf[random_column_choice].plot.hist(ax=ax3, bins=50)
classifier2 = classification(y=acc[random_column_choice])
# Add vertical lines for class breaks
for value in classifier2.bins:
plt.axvline(value, color='k', linestyle='dashed', linewidth=1)
fig.tight_layout()
# -
classify_and_plot(acc, classification = mapclassify.EqualInterval)
# ## Creating a custom classifier
#
# **Multicriteria data classification**
#
# Let's create a function where we classify the geometries into two classes based on a given `threshold` -parameter. If the area of a polygon is lower than the threshold value (average size of the lake), the output column will get a value 0, if it is larger, it will get a value 1. This kind of classification is often called a [binary classification](https://en.wikipedia.org/wiki/Binary_classification).
#
# First we need to create a function for our classification task. This function takes a single row of the GeoDataFrame as input, plus few other parameters that we can use.
#
# It also possible to do classifiers with multiple criteria easily in Pandas/Geopandas by extending the example that we started earlier. Now we will modify our binaryClassifier function a bit so that it classifies the data based on two columns.
#
# - Let's call it `custom_classifier` that does the binary classification based on two treshold values:
#
def custom_classifier(row, src_col1, src_col2, threshold1, threshold2, output_col):
# 1. If the value in src_col1 is LOWER than the threshold1 value
# 2. AND the value in src_col2 is HIGHER than the threshold2 value, give value 1, otherwise give 0
if row[src_col1] < threshold1 and row[src_col2] > threshold2:
# Update the output column with value 0
row[output_col] = 1
# If area of input geometry is higher than the threshold value update with value 1
else:
row[output_col] = 0
# Return the updated row
return row
# Now we have defined the function, and we can start using it.
#
# - Let's do our classification based on two criteria and find out grid cells where the **travel time is lower or equal to 20 minutes** but they are further away **than 4 km (4000 meters) from city center**.
#
# - Let's create an empty column for our classification results called `"suitable_area"`.
#
# +
# Create column for the classification results
acc["suitable_area"] = None
# Use the function
acc = acc.apply(custom_classifier, src_col1='pt_r_tt',
src_col2='walk_d', threshold1=20, threshold2=4000,
output_col="suitable_area", axis=1)
# See the first rows
acc.head(2)
# -
# Okey we have new values in `suitable_area` -column.
#
# - How many Polygons are suitable for us? Let's find out by using a Pandas function called `value_counts()` that return the count of different values in our column.
#
# Get value counts
acc['suitable_area'].value_counts()
# Okay, so there seems to be nine suitable locations for us where we can try to find an appartment to buy.
#
# - Let's see where they are located:
#
# +
# Plot
acc.plot(column="suitable_area", linewidth=0);
# Use tight layour
plt.tight_layout()
# -
# A-haa, okay so we can see that suitable places for us with our criteria seem to be located in the
# eastern part from the city center. Actually, those locations are along the metro line which makes them good locations in terms of travel time to city center since metro is really fast travel mode.
#
# **Other examples**
#
# Older course materials contain an example of applying a [custom binary classifier on the Corine land cover data](https://automating-gis-processes.github.io/2017/lessons/L4/reclassify.html#classifying-data>).
| source/notebooks/L4/reclassify.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nomoto model first order PIT
# +
# # %load imports.py
"""
These is the standard setup for the notebooks.
"""
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from jupyterthemes import jtplot
jtplot.style(theme='onedork', context='notebook', ticks=True, grid=False)
import pandas as pd
pd.options.display.max_rows = 999
pd.options.display.max_columns = 999
pd.set_option("display.max_columns", None)
import numpy as np
import os
import matplotlib.pyplot as plt
#plt.style.use('paper')
import copy
import numpy as np
import os
from src.data import database
from mdldb import mdl_to_evaluation
from mdldb.tables import Run
import src.data
import os.path
from sklearn.pipeline import Pipeline
import sympy as sp
from sklearn.metrics import r2_score
import src.reporting.paper_writing as paper_writing
from src.equations import equations
from src.equations import symbols
from rolldecayestimators.substitute_dynamic_symbols import lambdify
# -
from sympy.physics.vector.printing import vpprint, vlatex
from IPython.display import display, Math, Latex
from pandas_profiling import ProfileReport
import evaluation.evaluation_helpers as evaluation_helpers
from scipy.optimize import least_squares
# ## Nomotos equation:
Math(vlatex(equations.nomoto_first_order))
db = database.get_db()
sql="""
SELECT * from run
INNER JOIN projects
ON run.project_number==projects.project_number
INNER JOIN loading_conditions
ON (run.loading_condition_id == loading_conditions.id)
INNER JOIN models
ON run.model_number == models.model_number
INNER JOIN ships
ON models.ship_name == ships.name
WHERE run.test_type=="spiral"
"""
data = pd.read_sql_query(sql=sql, con=db.engine)
data = data.loc[:,~data.columns.duplicated()]
data.describe()
# +
#profile = ProfileReport(statistics, title='Pandas Profiling Report')
#profile.to_widgets()
# -
loading_conditions = data.groupby(by=['loading_condition_id','ship_speed'])
loading_conditions.describe()
loading_condition = loading_conditions.get_group(name=(3,19))
#loading_condition = loading_conditions.get_group(name=(144,16))
loading_condition.describe()
# ### Load all data for one loading condition
df_all = pd.DataFrame()
interesting_columns = ['delta','x0','y0','z0','phi','theta','psi']
for index, run in loading_condition.iterrows():
db_run = db.session.query(Run).get(int(run.id))
df = database.load_run(db_run=db_run)
df['t'] = df.index
df_=evaluation_helpers.coord(df=df) # add psi and position etc.
df = pd.concat((df,df_), axis=1)
df['run_id'] = run.id
df_all = df_all.append(df[['t','run_id'] + interesting_columns], ignore_index=True)
df_all.describe()
# +
fig,ax=plt.subplots()
runs = df_all.groupby(by='run_id')
for run_id, df in runs:
df['x0']-=df.iloc[0]['x0']
df['y0']-=df.iloc[0]['y0']
df.plot(x='y0',y='x0', ax=ax)
ax.get_legend().remove()
ax.set_aspect('equal', 'box')
# -
def derivate(group):
df = group.set_index('t')
ddf = np.gradient(df, df.index, axis=0).mean(axis=0)
s = pd.Series(ddf, index=df.columns)
return s
# +
df = runs.mean()
ddf = runs.apply(func= derivate)
df['u']=ddf['x0']
df['v']=ddf['y0']
df['w']=ddf['z0']
df['p']=ddf['phi']
df['q']=ddf['theta']
df['r']=ddf['psi']
df.sort_values(by='r', inplace=True)
# -
fig,ax=plt.subplots()
df.plot(x='delta', y='r', ax=ax, style='o')
ax.grid(True)
ax.set_title('Reverse spiral plot')
print(loading_condition.iloc[0]['project_path'])
spiral_eq = sp.simplify(equations.nomoto_first_order.subs(symbols.r_1d,0))
Math(vlatex(spiral_eq))
r_lambda=lambdify(sp.solve(spiral_eq,symbols.r)[0])
# +
def residual(parameters, X, ys):
r = r_lambda(*parameters,delta=X['delta'])
error = r - ys
return error
initial_guess = [-1,]
kwargs={
'X':df,
'ys':df['r'],
}
result = least_squares(fun=residual, x0=initial_guess, kwargs=kwargs, method='lm')
parameters={
'K':result.x,
}
# +
r_predict = r_lambda(**parameters,delta=df['delta'])
fig,ax=plt.subplots()
df.plot(x='delta', y='r', ax=ax, style='o')
ax.plot(df['delta'],r_predict, 'b-')
ax.grid(True)
ax.set_title('Reverse spiral plot');
# +
K_3 = sp.symbols('K_3')
delta_0 = sp.symbols('delta_0')
spiral_eq_3 = sp.Eq(symbols.delta,
sp.solve(spiral_eq,symbols.delta)[0] + symbols.r**5/K_3 + delta_0)
Math(vlatex(spiral_eq_3))
# -
A, A_3 = sp.symbols('A A_3')
spiral_eq_3_A = spiral_eq_3.subs([(symbols.K,1/A),
(K_3,1/A_3),
])
delta_lambda_3=lambdify(sp.solve(spiral_eq_3_A,symbols.delta)[0])
delta_lambda_3
# +
np.random.seed()
def residual_3(parameters, X, ys):
delta = delta_lambda_3(*parameters,r=X['r'])
error = (delta - ys)**2
return error
initial_guess = [-0.1,-1000,0]
kwargs={
'X':df,
'ys':df['delta'],
}
bounds = ([-np.inf,-np.inf],
[0,0,np.inf])
result = least_squares(fun=residual_3, x0=initial_guess, kwargs=kwargs, max_nfev=1000,
loss='linear', f_scale=0.1, method='lm')
parameters_3={
'A':result.x[0],
'A_3':result.x[1],
'delta_0':result.x[2],
}
# -
result
parameters_3
# +
N=100
r=np.linspace(df['r'].min(),df['r'].max(),N)
delta_predict = delta_lambda_3(**parameters_3,r=r)
fig,ax=plt.subplots()
df.plot(x='delta', y='r', ax=ax, style='o')
ax.plot(delta_predict,r, 'b-')
ax.grid(True)
ax.set_title('Reverse spiral plot');
# +
from scipy import polyval, polyfit
df['r**5'] = df['r']**5
X = df[['r','r**5']].copy()
X['1']=1.0
x, residuals, rank, s = np.linalg.lstsq(X, df['delta'], rcond=None)
parameters_4 = {
'A':x[0],
'A_3':x[1],
'delta_0':x[2],
}
delta_predict = delta_lambda_3(**parameters_4,r=r)
fig,ax=plt.subplots()
df.plot(x='delta', y='r', ax=ax, style='o')
ax.plot(delta_predict,r, 'b-')
ax.grid(True)
ax.set_title('Reverse spiral plot');
# -
| notebooks/02.01_db_nomoto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PREMIER
# language: python
# name: premier
# ---
from pyspark.sql import functions as F
from pyspark.sql.types import StringType
from pyspark.sql.functions import isnan, when, count, col
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").appName("Osm_extraction").getOrCreate()
# %run ./00_taa_prep_funcs.ipynb
# %run ./00_taa_urbanicity_funcs.ipynb
# # 1. Load input data
# 1) Trade area analysis input data
combo_set = expand_combo_set(read_geojson(path="C:/Users/<NAME>/OneDrive - EQUATORIAL COCA-COLA BOTTLING COMPANY S.L/dynamic segmentation/urbanicty/output taa combo/", file_name="taa_combo_set_1250.json"))
# # 2. Derive density
urba_set = derive_density(combo_set, ["osm_count", "carto_pop_sum", "carto_pb_urban_poi_count", "carto_mobility_residents_total_sum", "carto_mobility_workers_total_sum", "carto_mobility_others_total_sum"])
# # combo_set["ADM_EN"]
# # 3. Derive urbanicity
#
urba_set["overall_score"] = urba_set[["carto_pop_score", "carto_pb_urban_poi_score", "carto_mobility_residents_total_score", "carto_mobility_workers_total_score", "carto_mobility_others_total_score"]].mean(axis=1)
urba_set= \
derive_urbanicity_with_filler(
urba_set,
urb_bin_num=10,
metric=["overall_score"],
pop_metric="carto_pop_sum"
, filler=0.00001
)
urba_set.loc[3117,"urbanicity_overall"]=10
hex_plot(geo_data=urba_set, metric='urbanicity_overall')
# # 3. Derive trade area distance
# +
# Form the combined GeoDataFrame with data on territory, hexagons, population, osm, urbanicity, cmd and trade area distance
# runtime: 5 min
trade_area = derive_taa_dist__(path_cmd="D:/data_quality/data", cmd_set="customer_invoice_tizi_ouzou.xlsx", # latest set: 200706_100000_cmd.csv
urba_set = urba_set,path_hex="D:/data_quality/data", taa_set="taa"
)
trade_area["urbanicity_overall"] = trade_area["urbanicity_overall"].apply(int)
# +
ta_groupby = trade_area[["CHANNEL_CUSTOM", "urbanicity_overall", "dist_km"]].groupby(["CHANNEL_CUSTOM", "urbanicity_overall"], as_index=False)
ta_mean = ta_groupby.mean().rename(columns={"dist_km": "TA_radius"})
ta_count = ta_groupby.count().rename(columns={"dist_km": "nr_outlets"})
ta_table = ta_mean.merge(ta_count).rename(columns={"urbanicity_overall": "urbanicity"})
ta_table
# -
# plot a multilayer plot of the combined metric
hex_outlet_plot(
hex_set=urba_set,
hex_metric='urbanicity_overall',
point_set=trade_area.loc[:, ['geometry', 'LONGITUDE', 'LATITUDE', 'cmd_geometry', 'dist_km']],
point_metric='cmd_geometry',
point_size='dist_km')
# Form a subset and plot a multilayer plot of the combined metric
hex_outlet_plot_(
hex_set=urba_set,
hex_metric='urbanicity_overall',
point_set=trade_area.loc[:, ['geometry', 'LONGITUDE', 'LATITUDE', 'cmd_geometry', 'dist_km']],
point_metric='cmd_geometry',
point_size='dist_km')
| 02_urbanicity_exe_trade_radii_1250.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dautil as dl
import matplotlib.pyplot as plt
from scipy.signal import savgol_filter
import pandas as pd
import numpy as np
import seaborn as sns
from IPython.display import HTML
# +
def error(data, fit):
return data - fit
def win_rng():
return range(3, 25, 2)
def calc_mape(i, j, pres):
return dl.stats.mape(pres, savgol_filter(pres, i, j))
# -
pres = dl.data.Weather.load()['PRESSURE'].dropna()
pres = pres.resample('A')
context = dl.nb.Context('eval_smooth')
lr = dl.nb.LatexRenderer(chapter=6, start=6, context=context)
lr.render(r'Y_j= \sum _{i=-(m-1)/2}^{i=(m-1)/2}C_i\, y_{j+i}\qquad \frac{m+1}{2} \le j \le n-\frac{m-1}{2}')
# %matplotlib inline
dl.nb.RcWidget(context)
dl.nb.LabelWidget(2, 2, context)
# +
sp = dl.plotting.Subplotter(2, 2, context)
cp = dl.plotting.CyclePlotter(sp.ax)
cp.plot(pres.index, pres, label='Pressure')
cp.plot(pres.index, savgol_filter(pres, 11, 2), label='Poly order 2')
cp.plot(pres.index, savgol_filter(pres, 11, 3), label='Poly order 3')
cp.plot(pres.index, savgol_filter(pres, 11, 4), label='Poly order 4')
sp.label(ylabel_params=dl.data.Weather.get_header('PRESSURE'))
cp = dl.plotting.CyclePlotter(sp.next_ax())
stds = [error(pres, savgol_filter(pres, i, 2)).std()
for i in win_rng()]
cp.plot(win_rng(), stds, label='Filtered')
stds = [error(pres, pd.rolling_mean(pres, i)).std()
for i in win_rng()]
cp.plot(win_rng(), stds, label='Rolling mean')
sp.label()
sp.label(advance=True)
sp.ax.boxplot([error(pres, savgol_filter(pres, i, 2))
for i in win_rng()])
sp.ax.set_xticklabels(win_rng())
sp.label(advance=True)
df = dl.report.map_grid(win_rng()[1:], range(1, 5),
['win_size', 'poly', 'mape'], calc_mape, pres)
sns.heatmap(df, cmap='Blues', ax=sp.ax)
HTML(sp.exit())
# -
| Module2/Python_Data_Analysis_code/Chapter 6/eval_smooth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ctgan)
# language: python
# name: ctgan
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
data=pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data",header=None)
data.columns=["age","workclass","fnlwgt","education","education-num","marital-status","occupation","relationship","race","sex","capital-gain","capital-loss",
"hours-per-week","native-country","income"]
# ### data summary
data.info()
data.dtypes.value_counts()
data.describe(include="object")
# +
data.workclass.value_counts(normalize=True)
#plot the bar graph of percentage job categories
data.workclass.value_counts(normalize=True).plot.barh()
#plt.show()
# -
fig, axs = plt.subplots(3, 3, figsize=(18, 18))
for idx,column in enumerate(data.select_dtypes("object")):
idx2=idx//3
idx3=idx%3
data[column].value_counts(normalize=True).plot(ax=axs[idx2,idx3],kind="barh")
for idx in range(0,9):
print(idx//3,idx%3)
data.describe(include="int")
# **check if the dataset has nAn or null**
data.isna().sum()
data.isnull().sum()
data['sex'].value_counts()
data['workclass'].value_counts()
# **we checked that the " ?" is the NAN or null variable in the dataset**
# we can use pandas or scikit-imputer to get around this
# pandas
data1=data.copy()
data1.replace(' ?',value=np.nan,inplace=True)
#scikit learn
from sklearn.impute import SimpleImputer
data2=data.copy()
imputer = SimpleImputer(missing_values=' ?', strategy='constant',fill_value=np.nan)
X = pd.DataFrame(imputer.fit_transform(data2))
X
# ### duplicated records
# +
data.duplicated().sum()#24 records duplicated on all columns
data.duplicated(['workclass','education','occupation']).sum()# on ony some columns (31805)
# -
#check which ones they are
data[data.duplicated()]
| data-quality/quality assessment.ipynb |