code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import excursion
import excursion.optimize as optimize
import excursion.diagnosis as diagnosis
from excursion.testcases.fast import masked_single as scandetails
import numpy as np
import datetime
import excursion.plotting.twodim as plots
import excursion.utils as utils
np.warnings.filterwarnings('ignore')
# %pylab inline
# +
def plot_and_save(workdir, index, *args,**kwargs):
fig, axarr = plt.subplots(1, 2, sharey=True)
fig.set_size_inches(6.5, 3.5)
plt.title('Iteration {}'.format(index))
plots.plot(axarr,gps,X,y_list,scandetails,**kwargs)
plt.tight_layout()
plt.savefig(os.path.join(workdir,'update_{}.png'.format(str(index).zfill(3))), bbox_inches = 'tight')
plt.show()
N_UPDATES = 20
N_BATCH = 6
workdir = datetime.datetime.now().strftime('%Y-%m-%d-%H-%M-%S-2dmulti')
os.mkdir(workdir)
X,y_list,gps = optimize.init(scandetails)
testX = scandetails.plotX[~scandetails.invalid_region(scandetails.plotX)]
testy_list = [func(testX) for func in scandetails.functions]
testdata = testX, testy_list
for index in range(N_UPDATES):
plot_and_save(workdir, index, gps,X,y_list,scandetails, batchsize = (N_BATCH if index > 0 else len(X)), evaluate_truth = True)
t = diagnosis.diagnose(testdata,gps,scandetails)
print(t)
newX = optimize.suggest(gps, X, scandetails, batchsize=N_BATCH)
X,y_list,gps = optimize.evaluate_and_refine(X,y_list,newX,scandetails)
plot_and_save(workdir, index+1, gps,X,y_list,scandetails, batchsize = (N_BATCH if index > 0 else len(X)), evaluate_truth = True)
# -
| examples/MaskedExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8vs4O0IglwpY"
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
# + id="AvETfaFFnK8D" outputId="e84785e1-87fd-488f-b44b-8515d591bcaa" colab={"base_uri": "https://localhost:8080/", "height": 402}
from sklearn.datasets import load_boston
data=load_boston()
dt = pd.DataFrame(data.data, columns = data.feature_names)
dt
# + id="x84ihudTnvhv" outputId="a681ac87-c661-4192-aee2-b0a715a4d79b" colab={"base_uri": "https://localhost:8080/", "height": 369}
dt.info()
# + id="W2EH7TEQou2o" outputId="2b1ebc45-f9af-449e-bf30-d43c6e8d503e" colab={"base_uri": "https://localhost:8080/", "height": 34}
data.keys()
# + id="YCq0IO96o6sv" outputId="6d6fd66a-a815-43e1-e041-f8eae7168e18" colab={"base_uri": "https://localhost:8080/", "height": 934}
print(data.DESCR)
# + id="KpQoXz0y2FwE"
dt['MEDV'] = data.target
# + id="S_TIL_DEqxxe"
X = dt[:]
Y = dt['MEDV']
# + id="EeVm_z8Jp-me"
X_train, X_test, Y_train, Y_test = train_test_split(x, y, test_size = 0.25, random_state=0)
X_train = X
Y_train = Y
# + id="btRwZPZTsoR_"
from sklearn.preprocessing import StandardScaler
FS = StandardScaler()
X_train = FS.fit_transform(X_train)
X_test = FS.fit_transform(X_test)
# + id="sGlbUUigtX6s" outputId="f21bd674-53bd-4cf1-e229-77d305b57e59" colab={"base_uri": "https://localhost:8080/", "height": 34}
model = LinearRegression()
model.fit(X_train,Y_train)
# + id="fOQr3ZGZ13zW"
Y_pred = model.predict(X_test)
# + id="1ahGQCIzepV2" outputId="335677d8-f9f0-4a7b-8119-05f5139c98bc" colab={"base_uri": "https://localhost:8080/", "height": 34}
model.score(X_train, Y_train)*100
# + id="InVHz170u2IT" outputId="9b652555-5371-4a87-f9e0-2539c8528643" colab={"base_uri": "https://localhost:8080/", "height": 34}
model.score(X_test, Y_test)*100
# + id="wpABKmFRvQKa" outputId="1e035309-bbc4-4a20-cf15-036d4214bdaa" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.metrics import mean_squared_error
y_tr_pred = model.predict(X_train)
rmse = (np.sqrt(mean_squared_error(Y_train, y_tr_pred)))
print("Root Mean Square Error for Training set {}".format(rmse))
y_test_pred = model.predict(X_test)
rmse = (np.sqrt(mean_squared_error(Y_test, y_test_pred)))
print("Root Mean Square Error for Testing set {}".format(rmse))
| MachineLearning/28_Housing_Price/linearRegression/Boston_Housing_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PromoterArchitecturePipeline] *
# language: python
# name: conda-env-PromoterArchitecturePipeline-py
# ---
# this script extracts transcription start sites of promoters from Molly's dataset
# +
##needs mysql installed
#Need to be aware that fasta file sequence can be multiline or single line, databases provide it multilined
# -
from pybedtools import BedTool
from BCBio.GFF import GFFExaminer
from BCBio import GFF
import pprint
import pymysql.cursors
from pyfaidx import Fasta
import pandas as pd
import os
import numpy as np
import io
#examine the gff3 file
in_file = "/home/witham/Documents/Pipeline/data/TSS_data/AnnotatedPEATPeaks.txt"
examiner = GFFExaminer()
in_handle = open(in_file)
pprint.pprint(examiner.available_limits(in_handle))
in_handle.close()
#examine the gff3 file
in_file = "/home/witham/Documents/Pipeline/data/genomes/Araport11_GFF3_genes_transposons.201606.gff"
examiner = GFFExaminer()
in_handle = open(in_file)
pprint.pprint(examiner.available_limits(in_handle))
in_handle.close()
#examine the gff3 file
in_file = "../../data/genomes/Arabidopsis_thaliana/annotation/Arabidopsis_thaliana.TAIR10.47.gff3"
examiner = GFFExaminer()
in_handle = open(in_file)
pprint.pprint(examiner.available_limits(in_handle))
in_handle.close()
# +
# def download_chromosome_sizes(genome_assembly_id,output_file):
# """download chromosome sizes from the UCSC Genome browser's mysql database, using the UCSC Genome Browser assembly ID"""
# #connect to the database
# connection = pymysql.connect(host='genome-euro-mysql.soe.ucsc.edu',
# user='genome',
# port=3306)
# output = open(output_file, 'w') #make output file with write capability
# try:
# with connection.cursor() as cursor:
# sql = f"SELECT 'chrom', 'size' FROM {genome_assembly_id}.chromInfo" #specify the columns and genome assembly
# cursor.execute(sql) #grab the columns and genome assembly
# result = cursor.fetchone()
# output.write(result)
# finally:
# connection.close()
# output.close()
# -
def fasta_chromsizes(genome, output_file):
"""extracts chromosome sizes in a form compatible with BedTools.flank"""
the_genome = Fasta(genome) #using pyfaidx Fasta, parse the genome file as a fasta
chromsizes = {} #make dictionary called chromsizes
for key in the_genome.keys():
chromsizes[f'{key}'] = f'({len(the_genome[key])})' #add the chromosome name and length to dictionary
#create empty string
chromsizes_string = ''
#iterate over chromsizes dictionary key/values. Add key, tab, value, newline to the string iteratively
for k,v in chromsizes.items():
chromsizes_string = chromsizes_string + f'{k}\t{v}\n'
#write output file, deleting the parentheses created from the dictionary. This file is suitable for use in BedTools.flank
with open(output_file, 'w') as output:
output.write(chromsizes_string.replace('(','').replace(')',''))
# +
# def remove_overlapping_proms(promoter_bed,output_bed):
# #read in promoters
# proms_df = pd.read_table(promoter_bed, sep='\t', header=0)
# cols = ['chr','start','stop','AGI','dot1','strand','source','type','dot2','attributes']
# proms_df.columns = cols
# #create bedtools object of promoters
# proms_bed = BedTool(promoter_bed)
# #c = columns to apply function to
# #o = count number of merged promoters, name the first and last promoter that were merged
# merged = proms_bed.merge(c=4, o=['count_distinct','first', 'last'])
# #write to buffer
# merged_buffer = io.StringIO()
# merged_buffer.write(str(merged))
# merged_buffer.seek(0)
# #read as dataframe
# overlapping = pd.read_table(merged_buffer, sep='\t', header=0)
# cols2 = ['chr','start','stop', 'number_of_overlaps', 'first_overlap','second_overlap']
# overlapping.columns = cols2
# #select only features made of more than one promoter that were merged as overlapping
# overlapping_only = overlapping[overlapping.number_of_overlaps >= 2]
# #write to file before removing them
# # overlapping_only.to_csv(output_bed,index=False,sep='\t',header=0)
# # #remove overlapping promoters
# # for i,data in proms_df.iterrows():
# return overlapping_only
# +
# def extract_genes(gene_gff,output_file):
# """This function extracts all whole genes from a gff3 file, ignoring gene features, and adds them to an output file"""
# stringToMatch = 'ID=gene'
# matchedLine = ''
# with open(gene_gff, 'r') as file:
# for line in file:
# if stringToMatch in line:
# matchedLine = matchedLine + line + '\n'
# with open(output_file, 'w') as output:
# output.write(matchedLine)
# -
def extract_genes(gene_gff,output_overlapping,output_file):
"""This function extracts all whole genes from a gff3 file, ignoring gene features, and adds them to an output file. It also exports a file containing overlapping genes, and removes any overlapping genes from the final output"""
#limit dictionary to genes
limit_info = dict(gff_type = ['gene'])
#open temporary buffer
#output = open(temp, 'w')
tempbuffer = io.StringIO()
#open gff file, parse it, limiting to genes only. Save the file.
with open(gene_gff, 'r') as in_handle:
GFF.write(GFF.parse(in_handle, limit_info=limit_info),tempbuffer)
#go back to beginning of the buffer
tempbuffer.seek(0)
#now remove the unwanted annotations that were added by GFF.write eg. Chr1 annotation remark 1 30425192 . . . gff-version=3
#remove lines beginning with ##
with open(output_file, 'w') as newfile:
for line in tempbuffer:
line = line.strip() # removes hidden characters/spaces
if line[0] == "#":
pass
else:
#don't include lines beginning with ##
newfile.write(line + '\n') #output to new file
#remove temporary buffer
tempbuffer.close()
#read in gff file
genes = pd.read_table(output_file, sep='\t', header=None)
cols2 = ['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']
genes.columns = cols2
#remove all lines where source is annotation
no_annotation = genes[~genes.source.str.contains("annotation")]
#remove all lines that are not protein coding
protein_coding = no_annotation[no_annotation.attributes.str.contains('biotype=protein_coding')]
#add AGI column
gene_agi = protein_coding.assign(AGI=protein_coding.attributes.str.extract(r'ID=gene:(.*?)\;'))
#remove all features which overlap:
#make buffer of genes
genes_buffer = io.StringIO()
gene_agi.to_csv(genes_buffer,index=False,sep='\t',header=None)
genes_buffer.seek(0)
#create bedtools object of genes
genes_bed = BedTool(genes_buffer)
#c = columns to apply function to
#o = count number of merged promoters, name the first and last promoter that were merged
merged = genes_bed.merge(c=10, o=['count_distinct','first', 'last'])
#write to bufer
merged_buffer = io.StringIO()
merged_buffer.write(str(merged))
merged_buffer.seek(0)
#read as dataframe
overlapping = pd.read_table(merged_buffer, sep='\t', header=None)
cols3 = ['chr','start','stop', 'number_of_overlaps', 'first_overlap','second_overlap']
overlapping.columns = cols3
#select only features made of more than one promoter that were merged as overlapping
overlapping_only = overlapping[overlapping.number_of_overlaps >= 2]
#save these overlapping genes to file
overlapping_only.to_csv(output_overlapping,index=False,sep='\t',header=None)
#keep only non-overlapping genes
final_genes_list = overlapping[overlapping.number_of_overlaps == 1]
final_genes = gene_agi[gene_agi.AGI.isin(final_genes_list.first_overlap)]
final_genes_noAGI = final_genes[cols2]
#create gff file containing no lines with annotation and no lines starting with ##
final_genes_noAGI.to_csv(output_file,index=False,sep='\t',header=None)
return no_annotation
# +
# def add_promoter(genes_gff,chromsize,promoter_length,output_file):
# """This function adds a promoter of a certain length to each gene in the input file and exports to an output file"""
# #output = open(output_location, 'w') #make output file with write capability
# genes = BedTool(genes_gff)
# promoters = genes.flank(g=chromsize, l=promoter_length, r=0, s=True)
# #promoters = a.flank(genome=chromsize, l=1000)
# #open gff file as dataframe and separate by strand, and add column names
# # df = pd.read_csv(genes_gff, sep='\t', encoding='utf-8',
# #names=["chromosome", "source", "type", "start", "end", "score", "strand", "phase", "attributes"])
# #positive = BedTool.from_dataframe(df[df.strand.isin(['+'])]) #add only positive strand genes to bedtools
# #pos_promoters = positive.flank(g=chromsize, l=promoter_length, r=0, s=True) #add promoter upstream of gene
# #make negative signs positive
# # negative = df[df.strand.isin(['-'])].copy()
# #negpos = negative.strand.replace('-','+')
# # negative2 = BedTool.from_dataframe(negpos) #add only negative strand genes to bedtools
# # neg_promoters = negative2.flank(g=chromsize, l=0, r=promoter_length) #add promoter after gene as it is on -ve strand
# #merge the positive and negative promoters, with postmerge option so that promoters themselves aren't merged
# #promoters = pos_promoters.cat(neg_promoters, postmerge=False)
# with open(output_file,'w') as f:
# f.write(str(promoters))
# # count = 0
# # while count < 10:
# # for line in promoters:
# # print(line)
# # count = count + 1
# # return negpos
# -
def add_promoter(genes_gff,chromsize,promoter_length):
"""This function adds a promoter of a certain length to each gene in the input file and exports an output pybedtools object"""
#output = open(output_location, 'w') #make output file with write capability
#parse gff file containing only genes.
genes = BedTool(genes_gff)
#extract promoters upsteam using chromsize file and specified promoter length. r, no. of bp to add to end coordinate. s, based on strand.
#-l =The number of base pairs to subtract from the start coordinate.
promoters = genes.flank(g=chromsize, l=promoter_length, r=0, s=True)
return promoters
# +
# def promoter_overlap(promoter_gff,allfeatures_gff,output_file):
# """function to create file containing promoters which overlap other genome features"""
# promoters = BedTool(promoter_gff) #read in files using BedTools
# features = BedTool(allfeatures_gff)
# #report chromosome position of overlapping feature, along with the promoter which overlaps it (only reports the overlapping nucleotides, not the whole promoter length. Can use u=True to get whole promoter length)
# #f, the minimum overlap as fraction of A. F, nucleotide fraction of B (genes) that need to be overlapping with A (promoters)
# #wa, Write the original entry in A for each overlap.
# #u, write original A entry only once even if more than one overlap
# intersect = promoters.intersect(features,f=0.001, F=0.001, u=True, wa=True) #could add u=True which indicates we want to see the promoters that overlap features in the genome
# with open(output_file, 'w') as output:
# output.write(str(intersect))
# -
def remove_promoter_overlap(promoter_gff, all_genes_df, output_file):
"""function to create file containing promoters which overlap other genome features.
Then create a df with only promoters which overlap other genes. Then shorten them so they are no longer overlapping.
Then merge back with all the extracted promoters, keeping only the shortened ones"""
#create buffer to save the protein_coding genes
all_genes_gff = io.StringIO()
all_genes_df.to_csv(all_genes_gff,sep='\t', header=None, index = False)
all_genes_gff.seek(0)
all_proms = BedTool(promoter_gff) #read in files using BedTools
features = BedTool(all_genes_gff)
#report chromosome position of overlapping feature, along with the promoter which overlaps it (only reports the overlapping nucleotides, not the whole promoter length. Can use u=True to get whole promoter length)
#f, the minimum overlap as fraction of A. F, nucleotide fraction of B (genes) that need to be overlapping with A (promoters)
#wa, Write the original entry in A for each overlap.
#wo, Write the original A and B entries plus the number of base pairs of overlap between the two features. Only A features with overlap are reported.
#u, write original A entry only once even if more than one overlap
intersect = all_proms.intersect(features, wo=True) #could add u=True which indicates we want to see the promoters that overlap features in the genome
#Write to output_file
with open(output_file, 'w') as output:
#Each line in the file contains gff entry a and gff entry b that it overlaps plus the number of bp in the overlap so 19 columns
output.write(str(intersect))
#read in gff file
overlapping_proms = pd.read_table(output_file, sep='\t', header=None)
cols = ['chrA', 'sourceA', 'typeA', 'startA','stopA','dot1A','strandA','dot2A','attributesA','chrB', 'sourceB', 'typeB', 'startB','stopB','dot1B','strandB','dot2B','attributesB','bp_overlap']
overlapping_proms.columns = cols
#make new columns for the new start and stop for feature A
#overlapping_proms['new_startA'] = overlapping_proms.startA
#overlapping_proms['new_stopA'] = overlapping_proms.stopA
#create dictionaary for AGI as key and starts and stops as values
temp_dict = {}
#iterate over rows
for i,data in overlapping_proms.iterrows():
#create AGI key
key = overlapping_proms.loc[i,'attributesA'].split('ID=gene:')[1].split(';')[0]
#if positive strand feature A, reduce length to the stop position of feature B + 1
if overlapping_proms.loc[i,'strandA'] == '+':
if key not in temp_dict:
if overlapping_proms.loc[i,'stopA'] <= overlapping_proms.loc[i, 'stopB']:
start = overlapping_proms.loc[i, 'stopB'] + 1
stop = overlapping_proms.loc[i, 'stopB'] + 1
temp_dict[key] = [start,stop]
else:
start = overlapping_proms.loc[i, 'stopB'] + 1
#update dictionary
temp_dict[key] = [start,overlapping_proms.loc[i,'stopA']]
#if key is in the dictionary
else:
#if overlapping feature is downstream of the promoter or equal to promoter end, reduce both start and stop to the stop of the feature + 1
if temp_dict[key][1] <= overlapping_proms.loc[i, 'stopB']:
start = overlapping_proms.loc[i, 'stopB'] + 1
stop = overlapping_proms.loc[i, 'stopB'] + 1
#overlapping_proms.drop(i, inplace=True)
temp_dict[key] = [start,stop]
#else reduce length of promoter to stop position of feature +1
else:
#if promoter has already been shortened by another overlapping feature, pass
if temp_dict[key][0] > overlapping_proms.loc[i, 'stopB']:
pass
#else shorten the promoter
else:
start = overlapping_proms.loc[i, 'stopB'] + 1
#update dictionary
temp_dict[key][0] = start
#if negative strand feature A, reduce length of promoter to start position of feature -1
elif overlapping_proms.loc[i,'strandA'] == '-':
if key not in temp_dict:
#if overlapping feature is downstream of the promoter, change promoter start and stop to the start position of the feature -1
if overlapping_proms.loc[i,'startA'] >= overlapping_proms.loc[i, 'startB']:
stop = overlapping_proms.loc[i, 'startB'] - 1
start = overlapping_proms.loc[i, 'startB'] - 1
temp_dict[key] = [start,stop]
else:
#if overlapping feature is downstream of the promoter, change promtoer start and stop to the start position of the feature -1
stop = overlapping_proms.loc[i, 'startB'] - 1
temp_dict[key] = [overlapping_proms.loc[i,'startA'],stop]
#if key is in dictionary already
else:
#if overlapping feature is downstream of the promoter, change promoter start and stop to the start position of the feature -1
if temp_dict[key][0] >= overlapping_proms.loc[i, 'startB']:
stop = overlapping_proms.loc[i, 'startB'] - 1
start = overlapping_proms.loc[i, 'startB'] - 1
temp_dict[key] = [start,stop]
#overlapping_proms.drop(i, inplace=True)
else:
#if promoter has already been shortened by another overlapping feature, pass
if temp_dict[key][1] < overlapping_proms.loc[i, 'startB']:
pass
else:
stop = overlapping_proms.loc[i, 'startB'] - 1
temp_dict[key][1] = stop
#create df with just feature A containing new start and stop locations
new_feature_A = overlapping_proms[['chrA', 'sourceA', 'typeA', 'startA', 'stopA', 'dot1A', 'strandA', 'dot2A', 'attributesA']]
#rename columns
cols2 = ['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']
new_feature_A.columns = cols2
#remove duplicates based on attributes
no_dups = new_feature_A.drop_duplicates('attributes')
#replace start and stop with those from dictionary
no_dups_replaced = no_dups.copy()
for i,data in no_dups.iterrows():
key = no_dups_replaced.loc[i,'attributes'].split('ID=gene:')[1].split(';')[0]
no_dups_replaced.loc[i,'start'] = temp_dict[key][0]
no_dups_replaced.loc[i,'stop'] = temp_dict[key][1]
#create a buffer of promoter_gff
promoter_gff_buffer = io.StringIO()
promoter_gff_buffer.write(str(promoter_gff))
#go back to beginning of the buffer
promoter_gff_buffer.seek(0)
#read in all_proms as df
all_proms_df = pd.read_table(promoter_gff_buffer, sep='\t', header=None)
all_proms_df.columns = cols2
#remove overlapping promoters from all_proms_df
#first add AGI columns
#add AGI column to promoters
all_proms_df_agi = all_proms_df.assign(AGI=all_proms_df.attributes.str.extract(r'ID=gene:(.*?)\;'))
new_feature_A_agi = no_dups_replaced.assign(AGI=no_dups_replaced.attributes.str.extract(r'ID=gene:(.*?)\;'))
overlapping_proms_agi = overlapping_proms.assign(AGI=overlapping_proms.attributesA.str.extract(r'ID=gene:(.*?)\;'))
#remove any promoters in all_prom_df that overlap other genes
all_prom_removed_overlaps = all_proms_df_agi[~all_proms_df_agi.AGI.isin(overlapping_proms_agi.AGI)]
#merge all_prom_removed_overlaps with new_feature_A_agi
#outer merge adding suffix to new_feature_A
merged = pd.merge(all_prom_removed_overlaps, new_feature_A_agi, how='outer')
#remove unwanted columns
merged = merged[cols2]
all_genes_gff.close()
promoter_gff_buffer.close()
#sort on chr and start
merged.sort_values(['chr','start'], inplace=True, ignore_index=True)
return merged
def add_5UTR(promoter_gff, all_features_gff):
"""Function to extend the promoters to include the 5'UTR region until the start codon of the first CDS feature of the same gene.
Also remove mitochondira and chloroplast features."""
promoters = pd.read_table(promoter_gff, sep='\t', header=None, low_memory=False)
cols = ['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']
promoters.columns = cols
#make chr dtype string
#add AGI column to promoters
promoters = promoters.assign(AGI=promoters.attributes.str.extract(r'ID=gene:(.*?)\;'))
#remove lines beginning with ##
#create a buffer for feature output
feature_buffer = io.StringIO()
with open(all_features_gff, 'r') as f:
for line in f:
line = line.strip() # removes hidden characters/spaces
if line[0] == "#":
pass
else:
#don't include lines beginning with ##
feature_buffer.write(line + '\n') #output to buffer
#go back to beginning of the buffer
feature_buffer.seek(0)
#read in feature buffer to df
features = pd.read_table(feature_buffer, sep='\t', header=None, low_memory=False)
features.columns = cols
#filter features to contain only cds
cds = features[features.type == 'CDS']
#filter to only contain cds 1 in parent .1
#remove all features whose parent isnt Parent=AGI.1
# cds_1 = cds_1[cds_1.attributes.str.contains('Parent=transcript[A-Z]{2}\d{1}[A-Z]{1}\d{5}.1')]
# keep only CDS:1;
#cds_1 = cds_1[cds_1.attributes.str.contains(':CDS:1;')]
#make chromosome column string
cds = cds.astype({'chr': 'str'})
cds_1 = cds.copy()
#remove mitochondria and chloroplast features
cds_1 = cds[cds['chr'].isin(['1', '2', '3', '4', '5'])]
#add AGI column to CDSs
cds2 = cds_1.assign(AGI=cds_1.attributes.str.extract(r'ID=CDS:(.*?)\.'))
#Sort based on chromosome then start
cds2 = cds2.sort_values(['chr','start']).reset_index(drop=True)
#remove duplicates keeping the CDS closest to the promoter
#if positive strand, remove the first AGI duplicate
no_dups_pos = cds2[cds2.strand == '+'].drop_duplicates('AGI',keep='first')
#if negative strand, remove the last AGI duplicate
no_dups_neg = cds2[cds2.strand == '-'].drop_duplicates('AGI',keep='last')
no_dups = pd.merge(no_dups_pos, no_dups_neg, how='outer')
#sort by chr and start again
no_dups = no_dups.sort_values(['chr','start']).reset_index(drop=True)
#remove mitochondria and chloroplast features from promoters
promoters = promoters[promoters['chr'].isin(['1', '2', '3', '4', '5'])]
#merge on AGI
merged = pd.merge(promoters, no_dups, on='AGI', how='left',suffixes=('','_cds'))
for i, v in merged.iterrows():
#if positive strand feature A, reduce length to the stop position of feature B + 1
if merged.loc[i,'strand'] == '+':
#if overlapping feature is upstream or equal to start of the promoter, drop row
if merged.loc[i,'start'] >= merged.loc[i, 'start_cds']:
merged.drop(i, inplace=True)
else:
#extend promoter to include 5'UTR region up until the start of the cds
merged.loc[i, 'stop'] = merged.loc[i, 'start_cds'] - 1
elif merged.loc[i,'strand'] == '-':
#if overlapping feature is upstream or equal to start of the promoter, drop row
if merged.loc[i,'stop'] <= merged.loc[i, 'stop_cds']:
merged.drop(i, inplace=True)
else:
#extend promoter to include 5'UTR region up until the start of the cds
merged.loc[i, 'start'] = merged.loc[i, 'stop_cds'] + 1
#remove unwanted columns
merged_new = merged[['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']]
return merged_new
def filter_bad_proms(proms):
"""function to remove promoters whose start is higher or equal to the stop.
Then calculate lengths and sort by length. Drop duplicates keeping the shortest promoter."""
# #Remove promoters with start higher than stop
# for i, v in proms.iterrows():
# if proms.loc[i,'start'] >= proms.loc[i, 'stop']:
# proms.drop(i, inplace=True)
##sort based on length then keep the shortest, and remove lengths that are 0
#Make promoter length column
proms_length = proms.assign(length=(proms.stop - proms.start))
#remove promoters with length <100
removed= proms_length.loc[proms_length.length >= 100]
#remove duplicates keeping the shortest promoter
no_dups = removed.sort_values('length',ascending=True).drop_duplicates('attributes',keep='first')
proms_dropped = no_dups[['chr', 'source', 'type', 'start', 'stop', 'dot1', 'strand', 'dot2', 'attributes']]
#sort by chr then start
proms_dropped = proms_dropped.sort_values(['chr','start']).reset_index(drop=True)
return proms_dropped
# +
#will come back to this later once pipline is finished
# def rename_promoter_feature(promoter_gff, output_file):
# """This function renames the 3rd column of the promoter gff file to promoter"""
# output = open(output_file, 'w')
# with open(promoter_gff, 'r') as in_handle:
# GFF.write(GFF.parse(in_handle, ??????????????????),output)
# output.close()
# +
# def find_closest_TSS(gene_gff,TSS_gff,output_location):
# """this reads in the genes gff file and TSS gff file, finds the closest gene each TSS belongs to"""
# output = open(output_location, 'w') #make output file with write capability
# genes = BedTool(gene_gff)
# TSS = BedTool(TSS_gff)
# non_overlapping_TSS = TSS.subtract(genes)
# #intersect = TSS.intersect(genes)
# # for gene in intersect:
# # print(gene.name)
# nearby = genes.closest(TSS, d=True, stream=True)
# for gene in nearby:
# if int(gene[-1]) < 1000:
# output.write(gene.name + '\n')
# output.close()
# -
def find_closest_TSS(gene_gff,TSS_gff,output_location):
"""this reads in the genes gff file and TSS gff file, finds the closest gene each TSS belongs to"""
#gff columns:
# 1:seqname 2:source 3:feature 4:start 5:end 6:score 7:filter 8:strand 9:group 10:attribute
def remove_characters_linestart(input_location,output_location,oldcharacters,newcharacters,linestart):
"""this function removes characters from the start of each line in the input file and sends modified lines to output"""
output = open(output_location, 'w') #make output file with write capability
#open input file
with open(input_location, 'r') as infile:
#iterate over lines in fuile
for line in infile:
line = line.strip() # removes hidden characters/spaces
if line[0] == linestart:
line = line.replace(oldcharacters, newcharacters) #remove characters from start of line, replace with new characters
output.write(line + '\n') #output to new file
output.close()
def remove_empty_lines(input_file):
"""function to strip empty line from end of file"""
#make temporary buffer file
temp_buffer = io.StringIO()
with open(input_file) as f_input:
data = f_input.read().rstrip('\n')
temp_buffer.write(data)
temp_buffer.seek(0)
data2 = temp_buffer.read().rstrip('\n')
with open(input_file, 'w') as f_output:
f_output.write(data2)
# +
def bidirectional_proms(in_file, out_file):
"""this function create a file containing all promoters with an upstream gene going in the other direction ie. potential bidirectional promoters"""
#read in gff file
promoters = pd.read_table(in_file, sep='\t', header=None)
cols2 = ['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']
promoters.columns = cols2
#make sure lines are sorted
promoters = promoters.sort_values(['chr','start']).reset_index(drop=True)
#if bidirectional
promoters['bidirectional'] = 'no'
# promoters['shift_f_start']
# promoters['shift_f_stop']
#print(promoters)
for i,data in promoters.iterrows():
if i-1 >= 0:
#print(i,data)
if promoters.loc[i, 'strand'] == '+' and promoters.loc[i-1, 'strand'] == '-' and promoters.loc[i, 'start'] - promoters.loc[i-1, 'stop'] < 2000:
promoters.loc[i, 'bidirectional'] = 'yes'
promoters.loc[i-1, 'bidirectional'] = 'yes'
#write file
with open(out_file, 'w') as output:
promoters[promoters.bidirectional == 'no'][['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']].to_csv(out_file,index=False,sep='\t',header=0)
# for i,data in promoters.iterrows():
# print(i, data)
# if i-1 > 0:
# if promoters.loc[i, 'strand'] == '+' and promoters.loc[i-1, 'strand'] == '-' and promoters.loc[i, 'start'] - promoters.loc[i-1, 'stop'] < 2000:
# promoters.loc[i, 'bidirectional'] = 'yes'
# promoters.loc[i-1, 'bidirectional'] = 'yes'
#print(len(promoters.bidirectional[promoters.bidirectional == 'yes']))
#print(promoters)
# -
directory_path = '../..'
file_names='non-overlapping_includingbidirectional_all_genes_newannotation'
# +
genome = f'{directory_path}/data/genomes/TAIR10_chr_all.fas'
#genes = f"{args.directory_path}/data/genomes/Araport11_GFF3_genes_transposons.201606.gff"
genes = f"{directory_path}/data/genomes/Arabidopsis_thaliana/annotation/Arabidopsis_thaliana.TAIR10.47.gff3"
#genes_renamedChr = "{args.directory_path}//data/genomes/Araport11_GFF3_genes_transposons.201606_renamedChr.gff"
#test_genes = f"{args.directory_path}/data/genomes/test_genes.gff3"
#testgenesonly_gff = f"{args.directory_path}/data/genomes/testgenes_only.gff3"
#temp = f"{args.directory_path}/data/TSS_data/temp.txt"
#TSS = "{args.directory_path}//data/TSS_data/AnnotatedPEATPeaks_renamedcol.gff"
#TSS = "{args.directory_path}//data/TSS_data/TSStest.txt"
#find_closest_TSS(genes,output,temp)
genesoverlapping_gff = f"{directory_path}/data/output/{file_names}/genesoverlapping.gff3"
genesonly_gff = f"{directory_path}/data/output/{file_names}/genesonly.gff3"
promoters = f"{directory_path}/data/output/{file_names}/promoters.gff3"
promoters_5UTR = f"{directory_path}/data/output/{file_names}/promoters_5UTR.gff3"
#genes_bed = "{args.directory_path}//data/genomes/genes.bed"
overlapping_promoters = f'{directory_path}/data/output/{file_names}/promoters_overlapping.gff3'
promoterandgenes_only_overlap = f'{directory_path}/data/output/{file_names}/promoterandgenes_only_overlap.gff3'
chromsizes_file = f'{directory_path}/data/output/{file_names}/chromsizes.chr'
#need to change temporary files to scratch directory
chromsizes_file_renamedChr_temp = f'{directory_path}/data/output/{file_names}/chromsizes_renamedChr_temp.chr'
chromsizes_file_renamedChr = f'{directory_path}/data/output/{file_names}/chromsizes_renamedChr.chr'
#chromsizes_file2 = '{args.directory_path}//data/genomes/chromsizes2.chr'
promoters_renamedChr_temp = f'{directory_path}/data/output/{file_names}/promoters_renamedChr_temp.gff3'
#promoters_renamedChr_temp2 = f'{args.directory_path}/data/genomes/{args.file_names}/promoters_renamedChr_temp2.gff3'
promoters_renamedChr = f'{directory_path}/data/output/{file_names}/promoters_renamedChr.gff3'
promoters_5UTR_renamedChr = f'{directory_path}/data/output/{file_names}/promoters_5UTR_renamedChr.gff3'
nonbidirectional_promoters = f'{directory_path}/data/output/{file_names}/nonbidirectional_proms.gff3'
# +
# TSS_raw = '/home/witham/Documents/Pipeline/data/TSS_data/AnnotatedPEATPeaks.txt'
# #TSS_renamedChr = '/home/witham/Documents/Pipeline/data/TSS_data/AnnotatedPEATPeaks_renamedChr.gff'
# #genome2 = "/home/witham/Documents/Pipeline/data/genomes/Arabidopsis_thaliana.TAIR10.dna.toplevel_sl_simpleChr.fasta"
# genome = '../../data/genomes/TAIR10_chr_all.fas'
# nonbidirectional_promoters = '../../data/genomes/nonbidirectional_proms.gff3'
# +
# genes_old = "../../data/genomes/Araport11_GFF3_genes_transposons.201606.gff"
# genes = "../../data/genomes/Arabidopsis_thaliana/annotation/Arabidopsis_thaliana.TAIR10.47.gff3"
# #genes_renamedChr = "/home/witham/Documents/Pipeline/data/genomes/Araport11_GFF3_genes_transposons.201606_renamedChr.gff"
# #test_genes = "/home/witham/Documents/Pipeline/data/genomes/test_genes.gff3"
# #testgenesonly_gff = "/home/witham/Documents/Pipeline/data/genomes/testgenes_only.gff3"
# #TSS = "/home/witham/Documents/Pipeline/data/TSS_data/AnnotatedPEATPeaks_renamedcol.gff"
# #TSS = "/home/witham/Documents/Pipeline/data/TSS_data/TSStest.txt"
# #find_closest_TSS(genes,output,temp)
# genesonly_gff = "../../data/genomes/genesonly.gff3"
# promoters = "../../data/genomes/promoters.gff3"
# promoters_5UTR = '../../data/genomes/promoters_5UTR.gff3'
# #genes_bed = "/home/witham/Documents/Pipeline/data/genomes/genes.bed"
# overlapping_promoters = "../../data/genomes/promoters_overlapping.gff3"
# promoterandgenes_only_overlap = "../../data/genomes/promoterandgenes_only_overlap.gff3"
# +
#remove 'Chr' from seqname column in TSS gff file so that it matches the naming in the gene gff file
#also remove Chr from araport gff file
#remove_characters_linestart(TSS_raw, TSS_renamedChr, 'Chr')
#remove_characters_linestart(genes, genes_renamedChr, 'Chr')
# +
#num = len([1 for line in open("/home/witham/Documents/Pipeline/data/genomes/Arabidopsis_thaliana.TAIR10.dna.toplevel_sl_simpleChr.fasta") if line.startswith(">")])
#print(num)
# +
# chromsizes_file = '../../data/genomes/chromsizes.chr'
# chromsizes_file_renamedChr_temp = '../../data/genomes/chromsizes_renamedChr_temp.chr'
# chromsizes_file_renamedChr = '../../data/genomes/chromsizes_renamedChr.chr'
# #chromsizes_file2 = '/home/witham/Documents/Pipeline/data/genomes/chromsizes2.chr'
# -
fasta_chromsizes(genome, chromsizes_file)
#rename mitochondria and chloroplast to Mt and Pt
#input_location,output_location,oldcharacters,newcharacters,linestart
remove_characters_linestart(chromsizes_file, chromsizes_file_renamedChr_temp, 'mitochondria','Mt', 'm')
remove_characters_linestart(chromsizes_file_renamedChr_temp, chromsizes_file_renamedChr, 'chloroplast','Pt','c')
os.remove(chromsizes_file_renamedChr_temp)
# +
#fasta_chromsizes(genome2, chromsizes_file2)
# +
#extract_genes(genes,genesonly_gff)
#extract_genes(genes,genesonly_gff)
#this changes chromosome no. to 1 rather than Chr1
# +
#remove genes which are bidirectional (ie. upsteam promoter of another gene)
#bidirectional_proms(genesonly_gff, nonbidirectional_promoters)
# +
#add_promoter(nonbidirectional_promoters,chromsizes_file_renamedChr,1000,promoters)
# +
# df = pd.read_csv(genesonly_gff, sep='\t', encoding='utf-8',
# names=["chromosome", "source", "type", "start", "end", "score", "strand", "phase", "attributes"])
# positive = BedTool.from_dataframe(df[df.strand.isin(['+'])]) #add only positive strand genes to bedtools
# pos_promoters = positive.flank(g=chromosome_sizes_dict, l=1000, r=0) #add promoter upstream of gene
# with open("/home/witham/Documents/Pipeline/data/genomes/promoters.gff3",'w') as f:
# f.write(str(pos_promoters))
# +
#find_closest_TSS(genesonly_gff,TSS_raw,'/home/witham/Documents/Pipeline/data/genomes/closest.test')
# +
#create file containing only promoters which overlap other genome features
#promoter_overlap(promoters,genes,overlapping_promoters)
# -
#examine this gff3 promoter file and compare to all promoters
in_file = overlapping_promoters
examiner = GFFExaminer()
in_handle = open(in_file)
pprint.pprint(examiner.available_limits(in_handle))
in_handle.close()
# all promoters
in_file = promoters
examiner = GFFExaminer()
in_handle = open(in_file)
pprint.pprint(examiner.available_limits(in_handle))
in_handle.close()
# +
# #promoters overlapping only genes
# promoter_overlap(promoters,genesonly_gff,promoterandgenes_only_overlap)
# -
# promoters overlapping genes only
in_file = promoterandgenes_only_overlap
examiner = GFFExaminer()
in_handle = open(in_file)
pprint.pprint(examiner.available_limits(in_handle))
in_handle.close()
protein_coding = extract_genes(genes,genesoverlapping_gff,genesonly_gff)
protein_coding
# +
selected_genes = genesonly_gff
promoters_incl_overlap = add_promoter(selected_genes,chromsizes_file_renamedChr,1000)
# -
#note, this only removes overlap for protein coding genes
subtracted = remove_promoter_overlap(promoters_incl_overlap,protein_coding,promoterandgenes_only_overlap)
subtracted
df = pd.read_table(subtracted,sep='\t',header=None)
df
subtracted = subtracted.sort_values(['chr','start']).reset_index(drop=True)
cleaned_proms = filter_bad_proms(subtracted)
with open(promoters,'w') as f:
cleaned_proms.to_csv(f,index=False,sep='\t',header=0)
cds = add_5UTR(promoters, genes)
cleaned_proms_fiveUTR = filter_bad_proms(cds)
cleaned_proms_fiveUTR = cleaned_proms_fiveUTR[['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']]
with open(promoters_5UTR,'w') as f:
cleaned_proms_fiveUTR.to_csv(f,index=False,sep='\t',header=0)
cleaned_proms = cleaned_proms[['chr', 'source', 'type', 'start','stop','dot1','strand','dot2','attributes']]
with open(promoters,'w') as f:
cleaned_proms.to_csv(f,index=False,sep='\t',header=0)
# +
import seaborn as sns
import matplotlib.pyplot as plt
graph = cleaned_proms_fiveUTR.copy()
graph['length'] = (graph.start - graph.stop).abs()
#graph.sort_values('length',ascending=True)
sns.set(color_codes=True)
sns.set_style("whitegrid")
dist_plot = graph['length']
#create figure with no transparency
dist_plot_fig = sns.distplot(dist_plot).get_figure()
# -
| src/data_sorting/extract_promoter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PROMISE12 prostate segmentation demo
# ## Preparation:
# 1) Make sure you have set up the PROMISE12 data set. If not, download it from https://promise12.grand-challenge.org/ (registration required) and run data/PROMISE12/setup.py
#
# 2) Make sure you are in NiftyNet root, setting niftynet_path correctly to the path with the niftynet folder in it
import os,sys
niftynet_path=r'path/to/NiftyNet'
os.chdir(niftynet_path)
# 3) Make sure you have all the dependencies installed (replacing gpu with cpu for cpu-only mode):
import pip
#pip.main(['install','-r','requirements-gpu.txt'])
pip.main(['install','-r','requirements-cpu.txt'])
pip.main(['install', 'SimpleITK>=1.0.0'])
# ## Training a network from the command line
# The simplest way to use NiftyNet is via the commandline net_segment.py script. Normally, this is done on the command line with a command like this from the NiftyNet root directory:
#
# ```python net_segment.py train --conf demo/PROMISE12/promise12_demo_train_config.ini --max_iter 10```
#
# Notice that we use configuration file that is specific to this experiment. This file contains default settings. Also note that we can override these settings on the command line.
#
# To execute NiftyNet from within the notebook, you can run the following python code:
import os
import sys
import niftynet
sys.argv=['','train','-a','net_segment','--conf',os.path.join('demos','PROMISE12','promise12_demo_train_config.ini'),'--max_iter','10']
niftynet.main()
# Now you have trained (a few iterations of) a deep learning network for medical image segmentation. If you have some time on your hands, you can finish training the network (by leaving off the max_iter argument) and try it out, by running the following command
#
# ```python net_segment.py inference --conf demos/PROMISE12/promise12_demo_inference_config.ini```
#
# or the following python code in the Notebook
import os
import sys
import niftynet
sys.argv=['', 'inference','-a','net_segment','--conf',os.path.join('demos','PROMISE12','promise12_demo_inference_config.ini')]
niftynet.main()
# Otherwise, you can load up some pre-trained weights for the network:
#
# ```python net_segment.py inference --conf demo/PROMISE12/promise12_demo_config.ini --model_dir demo/PROMISE12/pretrained```
# or the following python code in the Notebook
import os
import sys
import niftynet
sys.argv=['', 'inference','-a','net_segment','--conf',os.path.join('demos','PROMISE12','promise12_demo_inference_config.ini'), '--model_dir', os.path.join('demos','PROMISE12','pretrained')]
niftynet.main()
# You can find your segmented images in output/promise12_demo
#
# NiftyNet has taken care of a lot of details behind the scenes:
# 1. Organizing data into a dataset of images and segmentation labels
# 2. Building a deep leaning network (in this case, it is based on VNet by Milletari et al.)
# 3. Added deep learning infrastruture, such as a loss function for segmentation, the ADAM optimizer.
# 4. Added augmentation, where the images are zoomed and rotated a little bit for every training step so that you do not over-fit the data
# 5. Run the training algorithm
#
# All of this was controlled by the configuration file.
#
# ## The configuration file
#
# Let's take a closer look at the configuration file. Further details about the configuration settings are available in ```config/readme.md```
# + active=""
# [promise12]
# path_to_search = data/PROMISE12/TrainingData_Part1,data/PROMISE12/TrainingData_Part2,data/PROMISE12/TrainingData_Part3
# filename_contains = Case,mhd
# filename_not_contains = Case2,segmentation
# spatial_window_size = (64, 64, 64)
# interp_order = 3
# axcodes=(A, R, S)
#
# [label]
# path_to_search = data/PROMISE12/TrainingData_Part1,data/PROMISE12/TrainingData_Part2,data/PROMISE12/TrainingData_Part3
# filename_contains = Case,_segmentation,mhd
# filename_not_contains = Case2
# spatial_window_size = (64, 64, 64)
# interp_order = 3
# axcodes=(A, R, S)
# -
# These lines define how NiftyNet organizes your data. In this case, in the ./data/PROMISE12 folder there is one T2-weighted MR image named 'Case??_T2.nii.gz' and one reference segmentation named 'Case??_segmentation.nii.gz' per patient. The images for each patient are automatically grouped because they share the same prefix 'Case??'. For training, we exclude patients Case20-Case26, and for inference, we only include patients Case20-Case26, so that our training and inference data are mutually exclusive.
# + active=""
# [SYSTEM]
# cuda_devices = ""
# num_threads = 2
# num_gpus = 1
# model_dir = ./promise12_model
# -
# These lines are setting up some system parameters: which GPUs to use (in this case whatever is available), where to save the trained network parameters, and how many threads to use for queuing them up.
#
# The following lines specify network properties.
# + active=""
# [NETWORK]
# name = dense_vnet
# activation_function = prelu
# batch_size = 1
#
# # volume level preprocessing
# volume_padding_size = 0
# # histogram normalisation
# histogram_ref_file = standardisation_models.txt
# norm_type = percentile
# cutoff = (0.01, 0.99)
# normalisation = True
# whitening = True
# normalise_foreground_only=True
# foreground_type = otsu_plus
# multimod_foreground_type = and
# window_sampling = resize
#
# #how many images to queue up in advance so that the GPU isn't waiting for data
# queue_length = 8
#
# -
# ## Summary
#
# In this demo
# 1. you learned to run training and testing for a deep-learning-based segmentation pipeline from the command-line and from python code directly;
# 2. you also learned about the NiftyNet configuration files, and how they control the learning and inference process; and
# 3. you learned multiple ways to tell NiftyNet which data to use.
| demos/PROMISE12/PROMISE12_Demo_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Introduction : Email discovery
# + active=""
# # Date of creation : 17 july 2021
# # Version : 1.0
# # Description : This notebook helps in discovery of email address
# # Tools used here are:
# 1- hunter
#
# # Other tools which you can use are:
# -------------------------------------------------------------------------------------------
# Skrapp: Browser addon to find emails on Linkedin
#
# Email Extractor: Chrome extension to extract emails from web pages
#
# Convertcsv: Online tool to extract email addresses in text, web pages, data files etc.
#
# linkedin2username: OSINT Tool: Generate username lists for companies on LinkedIn
#
# Office365UserEnum: Enumerate valid usernames from Office 365 using ActiveSync.
# -
# # Importing package
# Importing packages
import os
# # Hunter.io API KEY
hunter_api=''
# # Getting the location of current directory
# +
# Getting the location of current directory
def current_directory():
cwd=os.getcwd()
print("Location of current directory :", cwd)
current_directory()
# -
# # Making the directory name results to save the result
# +
# Making directory name results
def directory_results():
try:
os.mkdir("Results")
except Exception as e:
print(e)
directory_results()
# -
# # Performing email discovery
# # Find email address using domain name :
# +
# Find email address using domain name :
domain=input("Enter the domain name : ")
def email_address():
try:
os.system(f"python3 /root/Desktop/offensivenotebook/Email_Discovery/hunter/emailfinder.py -d {domain} -a {hunter_api} -o Results/email_addr.txt ")
except Exception as e:
print(e)
email_address()
# -
# # Finding Online presence of email address :
# +
# Finding Online presence of email address :
domain = input("Enter the domain name : ")
firstname = input("Enter the firstname : ")
lastname = input("Enter the lastname : ")
def email_online_presence():
try:
os.system(f"python3 /root/Desktop/offensivenotebook/Email_Discovery/hunter/emailpresence.py -d {domain} -f {firstname} -l {lastname} -a {hunter_api} -o Results/email_presence.txt")
except Exception as e:
print(e)
email_online_presence()
# -
# # Verify the email address (FAKE, OR TRASH ONE ):
# +
# Verify the email address (FAKE, OR TRASH ONE ):
email_address=input("Enter the email address : ")
def verify_email_address():
try:
os.system(f"python3 /root/Desktop/offensivenotebook/Email_Discovery/hunter/emailverify.py -e {email_address} -a {hunter_api} -o Results/email_address_verify")
except Exception as e:
print(e)
verify_email_address()
# -
| Email_Discovery/Email_discovery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 7 - Strings
#
# Until now, we have mainly been dealing with numeric data (numbers). However, it is often necessary to process text sequences, referred to as 'character strings'. In this exercise, we will see some of the mechanisms Python provides for text processing.
#
# As we have already seen, any literal text sequence can be entered between a pair of inverted commas (" " or ' '), and stored to a variable. Python allows both single and double quotes to be used (not all languages do, so be careful if translating this sort of thinking to another language in the future).
# ```python
# name = "Hamlet"
# quote = 'To be or not to be: that is the question.'
# ```
# If we look at `type(name)`, we see that Python has a `str` type to represent strings.
#
# To join two strings together, we can simply combine them using the `+` operator:
# ```python
# x = name+" says "+quote
# print(x)
# ```
# We can enter quotation marks in a string literal by using the *other* kind of quotation mark as the string delimiter: either
# ```python
# x = name+" says '"+quote+"'"
# y = name+' says "'+quote+'"'
# print(x)
# print(y)
# ```
# We can also 'escape' the quotation mark, by placing a `\` in front of it. This forces Python to treat it as a literal character, without any special meaning:
# ```python
# x = name+" says \""+quote+"\""
# print(x)
# ```
# Finally, we can use triple-quotes (""" """) to mark the beginning and end of the string, which allows ' and " to appear within the string without difficulties:
# ```python
# difficult = """ Apparently "we can't just use single quotes" around this string """
# ```
# We already encountered triple-quoted strings, in the context of docstrings at the start of a function.
#
# We can also use the multiplication operator to repeat a string multiple times:
# ```python
# y = 3*name
# print(y)
# ```
# However, most other 'mathematical' operators (such as subtraction and division) have no meaning with strings.
#
# **➤ Try it out!**
# Try it here!
# Note that string variables containing numbers are not the same as integer or float variables containing the same numbers:
# ```python
# x = '6'
# y = '3'
# print(x + y) # prints '63'
# ```
# However, we can obtain an integer or float variable by using the `int` and `float` functions:
# ```python
# print(int(x) + int(y)) # prints '9'
# print(float(x) + float(y)) # prints '9.0'
# ```
# This does not work for strings containing two or more numbers separated by whitespace: `int('3 6')` is assumed to correspond to two distinct numbers, and so they cannot be converted to a single integer. Similarly, while numbers containing a decimal point can be converted to floating-point form, they cannot be interpreted as an integer.
#
# **➤ Try it out!**
# Try it here!
# If we want to extract a 'substring' - a letter, or sequence of letters, from the middle of the string - we can use syntax similar to that for extracting a subset of a list:
# ```python
# quote = 'To be or not to be: that is the question.'
# print(quote[2:35:3])
# ```
# This will print every 3rd character, starting from the character at position 2. Remember, Python counts from 0:
# ```text
# 0123456789...
# To be or n...
# ```
# We therefore see that the character at position 2 is a space, ' '. In general, a substring specification takes the form
# ```python
# variable[istart:istop:istep]
# ```
# Omitting `istart` means the substring should begin from the start of `variable`; omitting `istep` means it should go to the end of `variable`; and omitting `istep` implies that all intervening characters should be printed:
# ```python
# print(quote[:35:3])
# print(quote[2::3])
# print(quote[2:35])
# ```
# If we wish to extract only a single character, we simply provide its index:
# ```python
# print(quote[4])
# ```
# We can also iterate over the letters in a string:
# ```python
# for letter in quote:
# print(letter)
# ```
# **➤ Try it out!**
# Try it here!
# A new-line (carriage return) can be represented within a string by entering `'\n'`, for example:
# ```python
# multiline = 'This\nstring\noccupies\nfive\nlines.'
# print(multiline)
# ```
# Similarly, `\t` can be used to enter a Tab character. Spaces and tab characters are collectively known as 'whitespace'.
#
# **➤ Try it out!**
# Try it here!
# As with other data types, Python provides a number of functions to work with a string, `s`. Some of the more important ones are:
# - `len(s)` - Return the number of characters in `s`.
# - `s.count(x)` - Count the number of occurrences of string `x` within string `s`. Again, this is case-sensitive.
#
# - `s.join(x)` - Here, `x` is assumed to be an iterable (typically a list or tuple) of strings. This function returns a single string, containing all the strings from `x` with a copy of `s` between each. For example, `':'.join(['a','b','c'])` will return `'a:b:c'`.
# - `s.split()` - Return a list of all of the 'words' (substrings separated by whitespace) within `s`. Optionally, provide a character to be regarded as the word separator. For example, `'a,b,c'.split(',')` will return `['a','b','c']`. It is also possible to specify the maximum number of words to be returned; once this limit is reached, no further splitting is performed. A variant `s.rsplit()` works backwards from the end of the string.
# - `s.replace(x,y)` - Return a version of `s` where every occurrence of string `x` is replaced by string `y`.
#
# - `s.find(x)` - Return the index of the start of the *first* occurrence of string `x` in string `s`. Note that this is case-sensitive: compare `quote.find('to')` with `quote.find('To')`. A variant `s.rfind(x)` finds the *last* occurrence of `x`. Variants `s.index()` and `s.rindex()` are almost identical, except that they have different behaviour if `x` cannot be found within `s`: whereas `s.find()` raises an error, `s.index()` returns `-1`.
#
# - `s.upper()`, `s.lower()` and `s.title()` - Return a copy of the string `s` converted to be entirely in UPPER CASE/lower case/Title Case respectively.
# - `s.isupper()`, `s.islower()`, `s.istitle()` - Return `True` if `s` is entirely in UPPER CASE/lower case/Title Case respectively, otherwise `False`.
# - `s.capitalize()` - Return a version of `s` where the first character is in UPPER CASE and the remainder in lower case.
# - `s.swapcase()` - Return a version of `s` where all UPPER CASE characters are converted to lower case and vice versa.
#
# - `s.center(n)` - Create a string of length `n` containing a copy of `s` centered within this. By default, this is achived by padding with spaces (' ') before and after `s`; optionally, you can specify a different charater. For example, `'hello'.center(11, '_')` returns `'___hello___'`.
# - `s.ljust(n)` - Create a string of length `n` containing a copy of `s` at its left. Optionally, specify a character to use for padding. Similarly, `s.rjust(n)` places `s` at the right of the `n`-character string.
# - `s.strip()` - Return a copy of `s` with all whitespace removed. `s.lstrip()` and `s.rstrip()` are variants removing whitespace only at the start or end of the string, respectively. For example, `'elephant'.strip('e')` returns `'lephant'`.
#
# **➤ Try out all of these functions, and satisfy yourself that you understand how they work.**
#
# Try it here!
# **➤Write a function which takes in strings like `'apples:3; pears:17; bananas:21'` and produces a nicely-formatted table:**
# ```text
# +-----------+-----+
# | Apples | 3 |
# | Pears | 17 |
# | Bananas | 21 |
# +-----------+-----+
# | TOTAL | 41 |
# +-----------+-----+
# ```
# Here are some additional input strings it should be able to cope with:
# ```text
# cheese:0; BREAD:5; milk:1; Bacon:2
# Ham:15000; salami:36030; corned beef:1836;
# triangles:3;squares:4;pentagons:5;hexagons:6;heptagons:7;octagons:8;nonagons:9
# ```
# Try it here!
# As you may have noticed, Python's `print` function often displays information to a large number of decimal places, and it does not generally produce nicely-formatted output. To achieve this, we must make use of Python's string-formatting facilities. These provide a mechanism for converting numbers into strings, and controlling the exact form this takes.
#
# Python 3 provides two different frameworks for string formatting. In each, you create a string containing placeholders for the contents of each variable you want to output, then insert the data into these.
#
# This is best illustrated by an example, using the first (older) formatting framework:
# ```python
# x = 1/11
# print(x)
# s = "One eleventh is approximately %.3f"
# print(s)
# print(s%x)
# ```
# Here, the string `s` contains the text we wish to produce, and the entry `%.3f` is a placeholder representing a floating point number with three decimal places. `x` is a floating point variable, calculated to many decimal places. The code `s%x` combines the two, resulting in the contents of `x` being inserted into the string `s`, formatted as required.
#
# All placeholders begin with the '%' symbol. Integer placeholders end with the letter 'i', floating-point placeholders end with the letter 'f', and string placeholders end with the letter 's'. Between the '%' and the letter, one can specify various options controlling the exact form of output:
#
# | Placeholder | Description | Example | Output |
# |---|---|---|---|
# | `%i` | General integer (no further formatting specified) | `'%i'%3` | '3' |
# | `%3i` | Integer, at least 3 characters wide | `'%3i'%3` | `' 3'` |
# | `%03i` | Integer, at least 3 characters wide, zero-padded | `'%03i'%3` | `'003'` |
# | `%f` | General floating-point number (no format specified) | `'%f'%2.9` | '2.900000' |
# | `%12f` | Floating-point number, occupying at least 12 characters | `'%12f'%2.9` | `' 2.900000'`|
# | `%012f` | Floating-point number, occupying at least 12 characters, zero-padded | `'%012f'%2.9` | `'00002.900000'`|
# | `%8.2f` | Floating-point number, occupying at least 8 characters, rounded to two decimal places | `'%8.2f'%2.9` | `' 2.90'` |
# | `%s` | General string (no format specified) | `'%s'%'test'` | `'test'` |
# | `%10s` | String, occupying at least 10 characters | `'%10s'%'test'` | `' test'`|
# | `%%` | Literal '%' character | `'%6.2f%%'%2.9` | `' 2.90%'` |
#
# Where a string contains more than one placeholder, we can pass the required information as a tuple:
# ```python
# phrase = '%i litres of %s at $%.2f/L costs a total of $%.2f'
# print(phrase%(2, 'milk', 1.29, 2*1.29))
# print(phrase%(40, 'petrol', 1.53, 40*1.53))
# print(phrase%(7.5, 'water', 0.17, 7.5*0.17))
# ```
# **➤ Try it out!**
#
# Try it here!
# Sometimes, it may be necessary to use string formatting to write the placeholders, allowing the style of output to be set at runtime:
# ```python
# def print_result(result,number_of_decimal_places):
# fmt = "The result is %%.%if"
# print((fmt%number_of_decimal_places)%result)
# ```
# However, this is best avoided if possible.
#
# The second, newer approach to formatting uses braces `{}` instead of `%...` to represent a placeholder, and a `.format()` function that can act on any string. The syntax of the format specifiers is also different. Our example would become:
# ```python
# x = 1/11
# print(x)
# s = "One eleventh is approximately {:.2f}"
# print(s)
# print(s.format(x))
# ```
# Similarly,
# ```python
# phrase = '{} litres of {} at ${:.2f}/L costs a total of ${:.2f}'
# print(phrase.format(2, 'milk', 1.29, 2*1.29))
# print(phrase,format(40, 'petrol', 1.53, 40*1.53))
# print(phrase.format(7.5, 'water', 0.17, 7.5*0.17))
# ```
# The new approach provides a much richer set of formatting options, described in full [in the online documentation](https://docs.python.org/3.4/library/string.html#format-string-syntax). One benefit of the new style is that it is no longer necessary to pass information to format in the same order as it is used: we can number the placeholders. For example,
# ```python
# phrase = '{3} litres of {0} at ${1:.2f}/L costs a total of ${2:.2f}'
# print(phrase.format('milk', 1.29, 2*1.29, 2))
# print(phrase,format('petrol', 1.53, 40*1.53, 40))
# print(phrase.format('water', 0.17, 7.5*0.17, 7.5))
# ```
# This is particularly useful if you need to repeat the same information several times in a string:
# ```python
# phrase = "This sentence has the word {0} {0} {0} repeated three times and the word {1} {1} repeated twice."
# print(phrase.format('cat', 'dog'))
# ```
# **➤ Try it out!**
# Try it here!
# **➤ Repeat the table-formatting example, making use of the string formatting capabilities**
# Try it here!
# We will encounter many more text-formatting examples in later exercises.
# ## The 'Caesar' cipher
#
# As we have already discussed, every piece of information within a computer must be organised and represented in binary form. This implies that the sequence of letters in the alphabet can be mapped onto the set of integers, and this is usually done via the 'ASCII' code sequence.
#
# Python provides the function `chr(integer)` to convert integers into their ASCII alphanumeric equivalent.
#
# **➤ Write a loop to print the integers from 33 up to 128, and their ASCII equivalents.** (The first 32 ASCII codes are special control characters, and do not have alphanumeric equivalents).
# Try it here!
# A 'Caesar cipher' is a very simple way to hide a message making it difficult for someone to read. To encode a piece of text with a Caesar cipher, we simply shift each letter $N$ places up (or down) the alphabet. For example, choosing $N=1$, the message
# ```text
# I like Python
# ```
# would become
# ```text
# J mjlf Qzuipm
# ```
# because 'J' is one letter after 'I', 'm' is one after 'l', and so on.
#
# **➤ Write a function to encode messages using Caesar ciphers (for any choice of $N$).** Note that the 'decoder' is simply the 'encoder', but instead using $-N$.
# Try it here!
# Here is a message encoded using a Caesar cipher:
# ```text
# Pfl yrmv wzezjyvu kyzj vovitzjv
# ```
# **➤ By looping through all possible values of $N$, find the $N$ used to encode this message and decode it.**
# Try it here!
| jupyterbook/content-de/python/lab/ex07-strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # House Prices : Advanced Regression Techniques(submission2)
# ## Before get started, I referred to another kernal from Kaggle.
# 1. <NAME> : https://www.kaggle.com/surya635/house-price-prediction
# 2. https://www.kaggle.com/surya635/house-price-prediction
# 3. https://www.kaggle.com/pmarcelino/comprehensive-data-exploration-with-python - He is so good!
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
# %matplotlib inline
# -
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
train.head()
test.head()
train.info()
# ## EDA
# ### 1. Target Variable
fig, axes = plt.subplots(2,1,figsize = (10,5))
sns.distplot(train['SalePrice'], ax = axes[0], kde_kws = {'label' : 'kde'}) # 히스토그램 및 kde(커널 밀도 함수, Kernel density function)
sns.distplot(train['SalePrice'], fit = stats.norm,
fit_kws = {'label' : 'norm'},
kde_kws = {'label' : 'kde'}, ax = axes[1]) # fit = stats.norm 정규분포(mean, sigma)
plt.tight_layout()
(mu, sigma) = stats.norm.fit(train['SalePrice'])
mu, sigma
stats.probplot(train['SalePrice'], plot = plt)
# Transform this variable to normal distribution.<br>
# In this case, we use log.**(but i don't know why use log for normalization yet.)**
# +
# Apply log to variables
train['SalePrice'] = np.log1p(train['SalePrice'])
# Check if it is applied
plt.figure(figsize = (10,5))
sns.distplot(train['SalePrice'], fit = stats.norm)
# -
(mu, sigma) = stats.norm.fit(train['SalePrice'])
mu, sigma
fig = plt.figure(figsize = (10,5))
stats.probplot(train['SalePrice'], plot = plt)
plt.show()
# ### 2. Relation with catergorical features
df = train[['OverallQual', 'SalePrice']]
plt.figure(figsize = (15, 8))
sns.boxplot(x = 'OverallQual', y = 'SalePrice', data = df)
train_columns_10 = train.columns[1:11]
sns.set()
cols = ['SalePrice', 'OverallQual', 'GrLivArea', 'GarageCars', 'TotalBsmtSF', 'FullBath', 'YearBuilt']
sns.pairplot(train[cols])
# ## 배운 개념들
# ### 1. stats.probplot(data, plot = plt)
# - Calculate quantiles for a probability plot, and optionally show the plot.
# - 확률밀도함수를 plot할 때 위치척도를 계산해주며, plot도 해준다.(다만 plot은 template를 줘야함. ex) plt)
# - probplot optionally calculates a best-fit line for the data
# - 데이터에 알맞는 최적의 line을 계산해준다. <<< ? 무슨 뜻인지 모르겠음
#
# ### 2. boxplot이 무엇을 의미하는지
# - 전형적은 boxplot을 생각해보면 맨 위와 아래는 각각 max, min
# - 그리고 중앙 상자 안에는 밑과 아래에 각각 first quartile, thrid quartile이 존재한다.
# - 마지막으로 상자 중앙 선은 median값을 의미한다.
# - **sns.boxplot일 경우 outliers까지 표시해 줌(점들로 표시)**
#
# ### 3. sns.pairplot()의 기능(미친 기능이다 이건... seaborn 공부해야할듯)
# - 일단 여러 그래프를 grid형식으로 만들어 줌
# - 두 쌍의 값(x,y - corr 모양처럼)을 이용해서 각각의 correlation을 한 번에 확인할 수 있다.
#
| House-Prices/House-Prices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# ## Import the relevant libraries
#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from sklearn.cluster import KMeans
import sklearn
# ## Load the data
# +
data = pd.read_csv('Notebook_1.csv')
data.head()
# -
# ## Plot the data
plt.scatter(data['Longitude'],data['Latitude'])
plt.xlim(-180,180)
plt.ylim(-90,90)
plt.show()
# ## Select the features
# +
x = data.iloc[:, 1:3]
data['Language'] = data['Language'].map({'English': 0, 'French': 1 , 'German': 2})
data
# -
data.head(5)
# ## Clustering
# +
km = KMeans(5)
km.fit(x)
# -
# ## Clustering results
# +
id_cluster = km.fit_predict(x)
id_cluster
# +
newdf = data.copy()
newdf['pred'] = id_cluster
newdf
# -
plt.scatter(newdf['Longitude'],newdf['Latitude'], c = newdf['pred'], cmap = 'rainbow')
plt.xlim(-180,180)
plt.ylim(-90,90)
plt.show()
| Notebook_1_Basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Currently stacks using weighted average on predicted submissions files.
#
# To do:
# - Use folds in the initial layers and train a stacking a layer (boosted tree)
# ## Import data
import numpy as np
import pandas as pd
# +
path = 'data/raw/train.csv'
full_path = os.path.join(dir_path, path)
df_train = pd.read_csv(full_path, header=0, index_col=0)
print("Dataset has {} rows, {} columns.".format(*df_train.shape))
# +
path = 'data/raw/test.csv'
full_path = os.path.join(dir_path, path)
df_test = pd.read_csv(full_path, header=0, index_col=0)
print("Dataset has {} rows, {} columns.".format(*df_test.shape))
# -
# fill NaN with string "unknown"
df_train.fillna('unknown',inplace=True)
df_test.fillna('unknown',inplace=True)
# ## Pre-processing
from sklearn.model_selection import train_test_split
# +
seed = 42
np.random.seed(seed)
test_size = 0.2
target = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
corpus = 'comment_text'
X = df_train[corpus]
y = df_train[target]
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=test_size, random_state=seed)
# -
# ## Load and predict using trained models
from sklearn.model_selection import ParameterGrid
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.pipeline import Pipeline
from keras.callbacks import CSVLogger
from sklearn.metrics import log_loss
# +
from keras.models import model_from_json
def load_model(model_name):
# load json and create model
json_file = open(model_name+'.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights(model_name+".h5")
print("Loaded model from disk")
return loaded_model
# +
# LSTM
model_name = ''
loaded_model = load_model(model_name)
y_pred_LSTM = loaded_model.predict(padded_test, verbose=1)
# LR
model_name = ''
loaded_model = load_model(model_name)
y_pred_LR = loaded_model.predict(padded_test, verbose=1)
# -
# ## Stacking layer
# +
import os
def load_data(path):
full_path = os.path.join(os.path.realpath('..'), path)
df = pd.read_csv(full_path, header=0, index_col=0)
print("Dataset has {} rows, {} columns.".format(*df.shape))
return df
# -
LR = load_data('data/submissions/LR.csv')
LSTM = load_data('data/submissions/LSTM1.csv')
# Weighted averaging
score_lr = 0.188
score_lstm = 0.065
total = score_lr + score_lstm
df_stack = (1-score_lr/total)*LR + (1-score_lstm/total)*LSTM
df_stack.head()
# ## Submissions
model_name = 'LR_LSTM'
path = 'data/submissions/' + model_name + '.csv'
full_path = os.path.join(os.path.realpath('..'), path)
df_stack.to_csv(full_path, header=True, index=True)
# ## Misc
# +
# Create stacking layer training set - LSTM predictions with true labels
stacking = pd.concat([predictions[0], predictions[1]])
for col in stacking.columns:
stacking.rename(columns={col: col+'_LSTM'}, inplace=True)
stacking = stacking.join(X)
stacking = stacking.join(y)
# Evaluate 1st layer
for label in target:
loss = log_loss(stacking[label], stacking[label+'_LSTM'])
losses.append(loss)
print("{} log loss is {} .".format(label, loss))
print("Combined log loss: {} .".format(np.mean(losses)))
# Save file
path = 'data/processed/stacking.csv'
full_path = os.path.join(dir_path, path)
ytest.to_csv(full_path, header=True, index=True)
# +
# Use HOO to evaluate first layer combined log loss
| notebooks/18-jc-stacking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <figure>
# <IMG SRC="https://mamba-python.nl/images/logo_basis.png" WIDTH=125 ALIGN="right">
#
# </figure>
#
# # Common pitfalls
#
# <br>
#
#
#
#
# This notebook is created for the MAMBA Python course to explain the common pitfalls using Python.
# <br><br>
# <div style="text-align: right"> developed by MAMBA </div>
# Table of content:<a class="anchor" id="0"></a>
# 1. [variable names](#1)
# 1. [reusing variable names](#11)
# 2. [built-in functions](#12)
# 2. [nested loops and statements](#2)
# 3. [copy-paste](#4)
# 4. [answers to exercises](#answers)
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
# [back to TOC](#0)
# ## 1. variable names<a class="anchor" id="1"></a>
# Naming your variables may sound trivial but it is actually quite important for the readability of your code. Some naming conventions are written down in PEP-8. Keeping to these allows for the next reader to easily understand which are function names, class names, packages etc.
#
# An example of a widely adopted naming convention is shown below. The convention is that <b>variables and function names</b> should be <b>lowercase</b>, with words separated by <b>underscores</b> as necessary to improve readability.
# +
# according to conventions (PEP-8)
list_of_animals = ['dog', 'cat', 'mouse', 'duck']
# not according to conventions (PEP-8)
ListOfAnimals = ['dog','cat','mouse','duck']
# -
# ### A. reusing variable names<a class="anchor" id="11"></a>
# #### Exercise 1
#
# #### answer
# ### B. built-in functions<a class="anchor" id="12"></a>
#
# There are a number of built-in functions that are available without importing them via a package. These functions can be shown with the command below.
dir(__builtin__)
# If you use a built-in function as a variable name, you overwrite the built-in function and cannot use it anymore. See the two examples below. You have to restart the kernel <button class='btn btn-default btn-xs'><i class='fa fa-repeat icon-repeat'></i></button> in order to retrieve the built-in function.
# Note: after you understood the examples and did the exercise it may be helpful to comment out the cells below. Otherwise you constantly have to restart the kernel if you accidentally run these cells. You can comment out a cell by selecting its code and press `[ctrl][/]`.
# +
# example 1
range(0,3) #The built-in function range generates the integer numbers between the given start integer to the stop integer
print([x for x in range(0,3)])
range = (3,6) #this overwrites the built-in function 'range'
print([x for x in range])
range(0,3) #now you cannot call this function anymore!
# -
# example 2
list = [1,2,3] #this overwrites the built-in function 'list'
a = (5,4,7) #if you want to convert this tuple to a list you get an error
list(a)
# #### Exercise 2
# You have an integer, a float and a list. You want to divide the integer by the float and use the remainder as an index for the list. The code below gives a TypeError. Try to find out what causes the error and change the code to avoid it.
# +
int = 2
float = 2.0
a_list = [1, 2, 3, 4]
index_a = int/float
print(a[int(index_a)])
# -
# #### answer
# [back to TOC](#0)
#
# ## 2. loops<a class="anchor" id="2"></a>
#
# loops are a very powerful tool but can make code slow and hard to read. The common approach is to avoid loops, especially nested loops, if possible.
# ### A. avoid loops with pandas<a class="anchor" id="21"></a>
#
# The example below is about a pandas DataFrame with data from a whatsapp conversation. Every row has a datetime, user and message (the text is ommited because of privacy issues).
df = pd.read_csv(r'..\..\..\..\Practical_examples\whatsapp data\data\_chat_df.csv')
df['datetime'] = pd.to_datetime(df['Unnamed: 0'])
del df['Unnamed: 0']
df.head()
# Let's say you want to get a new column with the hour of the day of each row. You could use a for-loop to loop over every row and get the hour of the day from the datetime column. See the example below. This will take some time. Using the `%%time` command you can see how much time.
# %%time
df['hour'] = 0
for row_no, row in df.iterrows():
df.loc[row_no]['hour'] = row['datetime'].hour
print(df.head())
# There is an alternative for the for-loop, this is in general referred to as a vectorized solution. The operation is not executed row-by-row but on the whole vector (column) at once. The example below shows a vectorized solution.
df['hour'] = df['datetime'].dt.hour
df.head()
# #### exercise 3
#
# Create a new colum 'user_int' that only contains the number of the user as an integer. You can check the dtype of the column with `df['user_int'].dtype`.
# #### answer
# ### B. avoid nested for loops<a class="anchor" id="22"></a>
#
# Consider the code below in which an image of a ship is blurred by taking the average gray value of a 10x10 pixel patch.
#
# source: https://realpython.com/numpy-array-programming/
from skimage import io
url = ('https://www.history.navy.mil/bin/imageDownload?image=/'
'content/dam/nhhc/our-collections/photography/images/'
'80-G-410000/80-G-416362&rendition=cq5dam.thumbnail.319.319.png')
img = io.imread(url, as_grey=True)
fig, ax = plt.subplots()
ax.imshow(img, cmap='gray')
# +
# using a nested for loop
m, n = img.shape
mm, nn = m - size + 1, n - size + 1
patch_means = np.empty((mm, nn))
for i in range(mm):
for j in range(nn):
patch_means[i, j] = img[i: i+size, j: j+size].mean()
fig, ax = plt.subplots()
ax.imshow(patch_means, cmap='gray')
ax.grid(False)
# +
shape = (img.shape[0] - size + 1, img.shape[1] - size + 1, size, size)
patches = stride_tricks.as_strided(img, shape=shape, strides=img.strides*2)
veclen = 10**2
patches.reshape(*patches.shape[:2], veclen).mean(axis=-1).shape
strided_means = patches.mean(axis=(-1, -2))
fig, ax = plt.subplots()
ax.imshow(strided_means, cmap='gray')
ax.grid(False)
# -
# #### Exercise 4
#
# Below you see some code to calculate the sum of the rows and the columns of a numpy array. The code contains a nested for loop. Find a way to calculate the sum of the rows and columns without using a nested for-loop.
#
# Bonus: find a way to calculate the sum of the rows and columns without using a for-loop at all.
rand_arr = np.random.random(size=(3,4))
rand_arr
# +
row_sum_list = []
col_sum_list = []
for i in range(len(rand_arr)):
row_sum = 0
for j in range(len(rand_arr[i])):
row_sum += rand_arr[i][j]
if i==0:
col_sum_list.append(np.sum(rand_arr[:,j]))
row_sum_list.append(row_sum)
print('sum of rows',row_sum_list)
print('sum of columns',col_sum_list)
# -
# #### answer
# [back to TOC](#0)
#
# ## 3. copy-paste <a class="anchor" id="3"></a>
#
# # copy and paste is both a blessing and a curse for programming. It may look obvious to copy-paste a piece of code if you need it again. However this may turn out to be very time consuming in the end.
df2 = pd.DataFrame({ 'A' : 1.,
'B' : pd.Timestamp('20130102'),
'C' : pd.Series(1,index=list(range(4)),dtype='float32'),
'D' : np.array([3] * 4,dtype='int32'),
'E' : pd.Categorical(["test","train","test","train"]),
'F' : 'foo' })
# #### Exercise 5
#
# #### answer
# [back to TOC](#0)
#
# ## 4. answers to exercises <a class="anchor" id="answers"></a>
# #### answer to exercise 1
# #### answer to exercise 2
# You need to rename the variables int and float to avoid overriding the built-in functions `int()` and `float()`.
#
# Note: if you used `int` as a variable name you need to restart the kernel <button class='btn btn-default btn-xs'><i class='fa fa-repeat icon-repeat'></i></button> to be able to use the `int()` built-in function.
# +
no_int = 2
no_float = 2.0
a_list = ['item 0','item 1', 'item 2', 'item 3']
index_a = no_int/no_float
print(a_list[int(index_a)])
# -
# #### answer to exercise 3
df['user_int'] = df.user.str[-1].astype('int')
df.head()
# #### answer to exercise 4
# +
# answer 1 without nested for-loops
row_sum_list = []
for i in range(len(rand_arr)):
row_sum_list.append(np.sum(rand_arr[i,:]))
col_sum_list = []
for j in range(len(rand_arr[0])):
col_sum_list.append(np.sum(rand_arr[:,j]))
print('sum of rows',row_sum_list)
print('sum of columns',col_sum_list)
# -
# answer 2 without for-loops
print('sum of rows', rand_arr.sum(axis=1))
print('sum of columns', rand_arr.sum(axis=0))
# answer 3 without for-loops
print('sum of rows', np.sum(rand_arr, axis=1))
print('sum of columns', np.sum(rand_arr, axis=0))
# #### answer to exercise 5
| Exercise_notebooks/On_topic/09_Code_Quality/02_common_pitfalls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Practice-Problems" data-toc-modified-id="Practice-Problems-1"><span class="toc-item-num">1 </span>Practice Problems</a></div><div class="lev2 toc-item"><a href="#2E1." data-toc-modified-id="2E1.-11"><span class="toc-item-num">1.1 </span>2E1.</a></div><div class="lev2 toc-item"><a href="#2E2." data-toc-modified-id="2E2.-12"><span class="toc-item-num">1.2 </span>2E2.</a></div><div class="lev2 toc-item"><a href="#2E3." data-toc-modified-id="2E3.-13"><span class="toc-item-num">1.3 </span>2E3.</a></div><div class="lev2 toc-item"><a href="#2E4." data-toc-modified-id="2E4.-14"><span class="toc-item-num">1.4 </span>2E4.</a></div><div class="lev2 toc-item"><a href="#2M1." data-toc-modified-id="2M1.-15"><span class="toc-item-num">1.5 </span>2M1.</a></div><div class="lev2 toc-item"><a href="#2M2." data-toc-modified-id="2M2.-16"><span class="toc-item-num">1.6 </span>2M2.</a></div><div class="lev2 toc-item"><a href="#2M3." data-toc-modified-id="2M3.-17"><span class="toc-item-num">1.7 </span>2M3.</a></div><div class="lev2 toc-item"><a href="#2M4." data-toc-modified-id="2M4.-18"><span class="toc-item-num">1.8 </span>2M4.</a></div><div class="lev2 toc-item"><a href="#2M5." data-toc-modified-id="2M5.-19"><span class="toc-item-num">1.9 </span>2M5.</a></div><div class="lev2 toc-item"><a href="#2M6." data-toc-modified-id="2M6.-110"><span class="toc-item-num">1.10 </span>2M6.</a></div><div class="lev2 toc-item"><a href="#2M7." data-toc-modified-id="2M7.-111"><span class="toc-item-num">1.11 </span>2M7.</a></div><div class="lev2 toc-item"><a href="#2H1." data-toc-modified-id="2H1.-112"><span class="toc-item-num">1.12 </span>2H1.</a></div><div class="lev2 toc-item"><a href="#2H2." data-toc-modified-id="2H2.-113"><span class="toc-item-num">1.13 </span>2H2.</a></div><div class="lev2 toc-item"><a href="#2H3." data-toc-modified-id="2H3.-114"><span class="toc-item-num">1.14 </span>2H3.</a></div><div class="lev2 toc-item"><a href="#2H4." data-toc-modified-id="2H4.-115"><span class="toc-item-num">1.15 </span>2H4.</a></div>
# -
# ## Import Statements
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pymc3 as pm
import scipy.stats as stats
# %config InlineBackend.figure_format = 'retina'
plt.style.use(['seaborn-colorblind', 'seaborn-darkgrid'])
# -
# ## Helper Functions
# +
def posterior_grid_approx(prior, success=6, tosses=9):
"""
This function helps calculate a grid approximation of the posterior distribution.
Parameters:
prior: np.array
A distribution representing our state of knowledge before seeing the data.
Number of items should be the same as number of grid points.
success: integer
Number of successes.
tosses: integer
Number of tosses (i.e. successes + failures).
Returns:
p_grid: np.array
Evenly-spaced out grid between 0 and 1.
posterior: np.array
The posterior distribution.
"""
# define grid
p_grid = np.linspace(0, 1, prior.shape[0])
# compute likelihood at each point in the grid
likelihood = stats.binom.pmf(success, tosses, p_grid)
# compute product of likelihood and prior
unstd_posterior = likelihood * prior
# standardize the posterior, so it sums to 1
posterior = unstd_posterior / unstd_posterior.sum()
return p_grid, posterior
def grid_approx_plot(
w=6,
n=9,
prior=np.repeat(5,100),
x_label='probability of water'
):
"""
This function helps calculate a grid approximation of the posterior distribution,
and then plots it.
Parameters:
prior: np.array
A distribution representing our state of knowledge before seeing the data.
Number of items should be the same as number of grid points.
w: integer
Number of successes.
n: integer
Number of trials.
"""
p_grid, posterior = posterior_grid_approx(
success=w,
tosses=n,
prior=prior
)
plt.plot(p_grid, posterior, 'o-', label=f'success = {w}\ntosses = {n}')
plt.xlabel(x_label)
plt.ylabel('posterior probability')
# plt.title('{} points'.format(points))
plt.legend(loc=0)
return p_grid, posterior
# -
# # Practice Problems
# ## 2E1.
#
# Which of the expressions below correspond to the statement: _the probability of rain on Monday_?
#
# (1) $Pr(rain)$
# * Nope. This is the averaged probability of rain.
#
# (2) $Pr(rain\mid Monday)$
# * Yes. Given that it is Monday, what is the probability of rain?
#
# (3) $Pr(Monday \mid rain)$
# * No. This is the probability of being Monday given that it is raining.
#
# (4) $Pr(rain, Monday) / Pr(Monday)$
# * Yes. This is equivalent to the second option.
# ## 2E2.
#
# Which of the expressions below correspond to the expression: $Pr(Monday \mid rain)$?
#
# (1) The probability of rain on Monday.
# * No. This is $Pr(rain \mid Monday)$.
#
# (2) The probability of rain, given that it is Monday.
# * No. This is $Pr(rain \mid Monday)$.
#
# (3) The probability that it is Monday, given that it is raining.
# * Yes. This is $Pr(Monday \mid rain)$.
#
# (4) The probability that it is Monday and that it is raining.
# * No. This is $Pr(Monday, rain)$.
# ## 2E3.
#
# Which of the expressions below correspond to the statement: _the probability that it is Monday, given that it is raining_?
#
# (1) $Pr(Monday \mid rain)$
# * Yes. By definition.
#
# (2) $Pr(rain \mid Monday)$
# * No. This is probability of rain given that it is a Monday.
#
# (3) $Pr(rain\mid Monday)Pr(Monday)$
# * No. This is equivalent to $Pr(rain, Monday)$, or the probability that it is raining and it is Monday.
#
# (4) $Pr(rain\mid Monday)Pr(Monday) / Pr(rain)$
# * This is equivalent to $Pr(rain, Monday) / Pr(rain)$, which then simplifies to $Pr(Monday \mid rain)$. So yes.
#
# (5) $Pr(Monday \mid rain)Pr(rain) / Pr(Monday)$
# * This simplifies to $Pr(Monday, rain) / Pr(Monday) = Pr(rain \mid Monday)$, so nope.
# ## 2E4.
#
# The Bayesian statistician <NAME> (1906-1985) began his book on probability theory with the declaration: "PROBABILITY DOES NOT EXIST." The capitals appeared in the original, so I imagine <NAME> wanted us to shout this statement. What he meant is that probabilty is a device for describing uncertainty from the perspective of an observer with limited knowledge; it has no objective reality. Discuss the globe tossing example from the chapter, in light of this statement. What does it mean to say "the probability of water is 0.7?"
#
# Ans:
#
# I think he meant that probability only exists in the "small world." It represents a person's state of uncertainty on some aspect of the real world. For example, in the real world, water is objectively some proportion of the whole Earth. The true proportion of water in Earth is probably some precise number, for example: 0.712345678... However, when a Bayesian says that "the probability of water is 0.7," this implies that this person _believes_ the ratio is about 0.7. The person is betting that the ratio is close to 0.7.
# ## 2M1.
#
# Recall the globe tossing model from the chapter. Compute and plot the grid approximate posterior distribution for each of the following sets of observations. In each case, assume a uniform prior _p_.
#
# (1) W,W,W
_, _ = grid_approx_plot(n=3,w=3, prior=np.repeat(5,int(1e3)) )
# (2) W,W,W,L
_,_ = grid_approx_plot(n=4, w=3, prior=np.repeat(5,int(1e3)) )
# (3) L,W,W,L,W,W,W
_, _ = grid_approx_plot(n=7, w=5, prior=np.repeat(5,int(1e3)))
# ## 2M2.
#
# Now assume a prior for _p_ that is equal to zero when $p < 0.5$ and is a positive constant when $p >= 0.5$. Again, compute and plot the grid approximate posterior distribution for each of the sets of observations in the problem just above.
_, _ = grid_approx_plot(
n=7,
w=5,
prior=( np.linspace(0, 1, int(1e3)) >= 0.5).astype(int))
# ## 2M3.
#
# Suppose there are two globes, one for Earth and one for Mars. The Earth globe is 70% covered in water. The Mars globe is 100% land. Further suppose that one of these globes--you don't know which--was tossed in the air and produced a "land" observation. Assume that each globe was equally likely to be tossed. Show that the posterior probability that the globe was the Earth, conditional on seeing "land" $Pr(\text{Earth}\mid\text{land})$ is 0.23.
# $Pr(\text{water} \mid \text{Earth}) = 0.7$
#
# $Pr(\text{land} \mid \text{Earth}) = 0.3$
#
# $Pr(\text{Earth}) = 0.5$
#
# $Pr(\text{land} \mid \text{Mars}) = 1.0$
#
# $Pr(\text{Mars}) = 0.5$
#
# \begin{equation}
# \begin{aligned}
# Pr(\text{land}) &= Pr(\text{land} \mid \text{Earth})Pr(\text{Earth}) + Pr(\text{land} \mid \text{Mars})Pr(\text{Mars}) \\
# &= 0.3 * 0.5 + 1.0 * 0.5 \\
# &= 0.65
# \end{aligned}
# \end{equation}
#
#
# \begin{equation}
# \begin{aligned}
# Pr(\text{Earth} \mid \text{land}) &= \frac{0.3 * 0.5}{0.65} \\
# &= 0.23 \\
# \end{aligned}
# \end{equation}
#
#
#
#
#
# ## 2M4.
#
# Suppose you have a deck with only three cards. Each card has two sides, and each side is either black or white. One card has two black sides. The second card has one black and one white side. The third card has two white sides. Now suppose all three cards are placed in a bag and shuffled. Someone reaches into the bag and pulls out a card and places it flat on a table. A black side is shown facing up, but you don't know the color of the side facing down. Show that the probability that the other side is also black is 2/3. Use the counting method (Section 2 of the chapter) to approach this problem. This means counting up the ways that each card could produce the observed data (a black side facing up on the table).
# Note: $B$ stands for black and $W$ stands for white. $B_1$ means that side 1 of a card is black. The left side is the side facing up (e.g. $B_2 \mid B_1$ means that the second side is black and it is the one facing up, while the other side is also black).
#
# Here are the combinations:
#
# $B_1 \mid B_2$ Black side is facing up and other side is black
#
# $B_2 \mid B_1$ Black side is facing up and other side is black
#
# $B_1 \mid W_2$ Black side is facing up, but other side is white.
#
# $W_2 \mid B_1$ Not relevant, since white side up.
#
# $W_1 \mid W_2$ Not relevant, since white side up.
#
# $W_2 \mid W_1$ Not relevant, since white side up.
#
# There are three events consistent with "Black side facing up," and only two of them are consistent with the the event "other side is also black." Thus the probability that the other side is black, given that the known side is black is $2/3$.
# ## 2M5.
#
# Now suppose there are four cards: B/B, B/W, W/W, and another B/B. Again suppose a card is drawn from the bag and a black side appears face up. Again calculate the probability that the other side is black.
# Previously:
#
# $B_1 \mid B_2$ Black side is facing up and other side is black
#
# $B_2 \mid B_1$ Black side is facing up and other side is black
#
# $B_1 \mid W_2$ Black side is facing up, but other side is white.
#
# $W_2 \mid B_1$ Not relevant, since white side up.
#
# $W_1 \mid W_2$ Not relevant, since white side up.
#
# $W_2 \mid W_1$ Not relevant, since white side up.
#
# New Data:
#
# $B_1 \mid B_2$ Black side is facing up and other side is black
#
# $B_2 \mid B_1$ Black side is facing up and other side is black
#
# There are now five events consistent with "Black side facing up," and four of them are consistent with the the event "other side is also black," so the probability of the other side being black, given that the side being shown is black is $4/5$.
# ## 2M6.
#
# Imagine that the black ink is heavy, and so cards with black sides are heavier than cards with white sides. As a result, it's less likely that a card with black sides is pulled from the bag. So again assume there are three cards: B/B, B/W, and W/W. After experimenting a number of times, you conclude that for every way to pull the B/B card from the bag there are 2 ways to pull the B/W card and 3 ways to pull the W/W card. Again suppose that a card is pulled and a black side appears face up. Show that the probability the other side is black is now 0.5. Use the counting method, as before.
# A probabilistically equal statement to this is if we pretend to have the bag have one $B/B$ card, two $B/W$ and three $W/W$ card in the bag.
#
# $B/B$ gives us two ways of picking a black side up: $B_1 / B_2$ and $B_2 / B_1$.
#
# If we pick one of the $B/W$ cards, there is one way that a card has black side up: $B_1$. However, because picking up a $B/W$ is twice as likely as picking up a $B/B$, then there are $2\times 1 = 2$ ways to get a card where we know that one side is black.
#
# There are $4$ total ways of picking a black card, but only half of them has black on the other side. Therefore the probability that the other side is black, given that a black side is face up, is 0.5.
# ## 2M7.
#
# Assume again the original card problem, with a single card showing a black side face up. Before looking at the other side, we draw another card from the bag and lay it face up on the table. The face that is shown on the new card is white. Show that the probability that the first card, the one showing a black side, has black on its other side is now 0.75. Use the counting method, if you can. Hint: Treat this like the sequence of globe tosses, counting all the ways to see each observation, for each possible first card.
# $B_1 \mid B_2$ -> $W_1 \mid B_2$ Black on the other side.
#
# $B_1 \mid B_2$ -> $W_1 \mid W_2$ Black on the other side.
#
# $B_1 \mid B_2$ -> $W_2 \mid W_1$ Black on the other side.
#
# $B_2 \mid B_1$ -> $W_1 \mid B_2$ Black on the other side.
#
# $B_2 \mid B_1$ -> $W_1 \mid W_2$ Black on the other side.
#
# $B_2 \mid B_1$ -> $W_2 \mid W_1$ Black on the other side.
#
# $B_1 \mid W_2$ -> $W_1 \mid W_2$ Not black on the other side.
#
# $B_1 \mid W_2$ -> $W_2 \mid W_1$ Not black on the other side.
#
# Given that the second card has white face-up and the first card has black face-up, the probability that the first card's other side is black is 0.75.
#
# ## 2H1.
#
# Suppose there are two species of panda bear. Both are equally common in the wild and live in the same places. They look exactly alike and eat the same food, and there is yet no genetic assay capable of telling them apart. They differ however in their family sizes. Species A gives birth to twins 10% of the time, otherwise birthing a single infant. Species B births twins 20% of the time, otherwise birthing singleton infants. Assume these numbers are known with certainty, from many years of field research.
#
# Now suppose you are managing a captive panda breeding program. You have a new female panda of unknown species, and she has just given birth to twins. What is the probability that her next birth will also be twins?
# Note: $C_1=2$ stands for twins during the first birthing event, $C_2=1$ stands for a singleton infant during the second birthing event. $S=a$ means that the species which gave birth was from Species A.
#
# $P(C_1=2 \mid S=a) = 0.1$
#
# $P(C_1=1 \mid S=a) = 0.9$
#
# $P(C_1=2 \mid S=b) = 0.2$
#
# $P(C_1=1 \mid S=b) = 0.8$
#
# $P(S=a) = 0.5$
#
# $P(S=b) = 0.5$
#
# $$
# \begin{equation}
# \begin{aligned}
# P(S=a \mid C_1=2) &= \frac{P(C_1=2 \mid S=a)P(S=a)}{P(C_1=2)} \\
# &= \frac{0.05}{0.05+0.1} \\
# &= \frac{1}{3} \\
# \end{aligned}
# \end{equation}
# $$
#
# $$
# \begin{equation}
# \begin{aligned}
# P(S=b \mid C_1=2) &= 1 - P(S=a \mid C_1=2) \\
# &= \frac{2}{3}
# \end{aligned}
# \end{equation}
# $$
#
# $$
# \begin{equation}
# \begin{aligned}
# P(C_2=2 \mid C_1=2) &= \sum_S P(C_2=2, S, \mid C_1=2) \\
# &= P(C_2=2, S=a \mid C_1=2) + P(C_2=2, S=b \mid C_1=2) \\
# &= P(C_2=2 \mid S=a, C_1=2) \cdot P(S=a \mid C_1=2) \\
# &\quad+ P(C_2=2 \mid S=b, C_1=2) \cdot P(S=b \mid C_1=2) \\
# &= P(C_2=2 \mid S=a) \cdot P(S=a \mid C_1=2) \\
# &\quad+ P(C_2=2 \mid S=b) \cdot P(S=b \mid C_1=2) & \text{Conditional independence. See Note below}\\
# &= \frac{1}{10} \times \frac{1}{3} + \frac{2}{10} \times \frac{2}{3} \\
# &= \frac{1}{30} + \frac{4}{30} \\
# &= \frac{1}{6}
# \end{aligned}
# \end{equation}
# $$
#
# _Note_: The causal diagram implied here is the following:
#
# 
#
# The causal diagram implies that if we already know the Species ($S$), then Birth at Time 1 ($C_1$) and Birth at Time 2 ($C_2$) are independent from each other. In mathematical notation:
#
# $$ C_2 \mathrel{\unicode{x2AEB} C_1 \mid S}$$
#
#
# In other words, if we know the species, knowing about the birth at time 1 does not give us any more information about birth at time 2. Therefore, we have the license to remove $C_1$ from the conditioning section:
#
# $$
# \begin{equation}
# \begin{aligned}
# P(C_2=2 \mid S=a, C_1=2) &= P(C_2=2 \mid S=a) \\
# P(C_2=2 \mid S=b, C_1=2) &= P(C_2=2 \mid S=b) \\
# \end{aligned}
# \end{equation}
# $$
# ## 2H2.
#
# Recall all the facts from the problem above. Now compute the probability that the panda we have is from species A, assuming we have observed only the first birth and that it was twins.
# $$
# \begin{equation}
# \begin{aligned}
# P(S=a \mid C_1=2) &= \frac{P(C_1=2 \mid S=a)P(S=a)}{P(C_1=2)} \\
# &= \frac{0.05}{0.05+0.1} \\
# &= \frac{1}{3} \\
# \end{aligned}
# \end{equation}
# $$
# ## 2H3.
#
# Continuing on from the previous problem, suppose the same panda mother has a second birth and that it is not twins, but a singleton infant. Compute the posterior probability that this panda is species A.
# $$
# \begin{equation}
# \begin{aligned}
# P(S=a \mid C_2=1, C_1=2) &= \frac{P(C_2=1, C_1=2 \mid S=a)P(S=a)}{P(C_2=1, C_1=2)} \\
# &= \frac{P(C_2=1 \mid S=a)P(C_1=2 \mid S=a)P(S=a)}{P(C_2=1, C_1=2)} \\
# &= \frac{P(C_2=1 \mid S=a)P(C_1=2 \mid S=a)P(S=a)}{P(C_2=1 \mid S=a)P(C_1=2 \mid S=a)P(S=a) + P(C_2=1 \mid S=b)P(C_1=2 \mid S=b)P(S=b)} \\
# &= \frac{0.9 \times 0.1 \times 0.5}{0.9 \times 0.1 \times 0.5 + 0.8 \times 0.2 \times 0.5} \\
# &= 0.36
# \end{aligned}
# \end{equation}
# $$
#
#
# ## 2H4.
#
# A common boast of Bayesian statisticians is that Bayesian inference makes it easy to use all the data, even if the data are of different types.
#
# So suppose now that a veterinarian comes along who has a new genetic test that she claims can identify the species of our mother panda. But the test, like all tests, is imperfect. This in the information you have about the test:
#
# * The probability it correctly identifies a species A panda is 0.8.
# * The probability it correctly identifies a species B panda is 0.65.
#
# The vet administers the test to your panda and tells you that the test is positive for species A. First ignore your previous information from the births and compute the posterior probability that your panda is species A. Then redo your calculation now using the birth data as well.
# Let $T$ stand for the test result.
#
# $P(T=a \mid S=a) = 0.8$
#
# $P(T=b \mid S=b) = 0.65$, therefore: $P(T=a \mid S=b) = 0.35$.
#
# Without birth data:
#
# $$
# \begin{equation}
# \begin{aligned}
# P(S=a \mid T=a) &= \frac{P(T=a \mid S=a)P(S=a)}{P(T=a)} \\
# &= \frac{P(T=a \mid S=a)P(S=a)}{P(T=a \mid S=a)P(S=a) + P(T=a \mid S=b)P(S=b)} \\
# &= \frac{0.8 \times 0.5}{0.8 \times 0.5 + 0.35 \times 0.5} \\
# &= \frac{0.8}{1.15} \\
# &\approx 0.70
# \end{aligned}
# \end{equation}
# $$
#
# With birth data:
#
# We use the posterior of the previous answer $P(S=a \mid C_2=1, C_1=2) = 0.36$ as the new prior $P(S=a)$:
#
# $$
# \begin{equation}
# \begin{aligned}
# P(S=a \mid T=a) &= \frac{P(T=a \mid S=a)P(S=a)}{P(T=a)} \\
# &= \frac{P(T=a \mid S=a)P(S=a)}{P(T=a \mid S=a)P(S=a) + P(T=a \mid S=b)P(S=b)} \\
# &= \frac{0.8 \times 0.36}{0.8 \times 0.36 + 0.35 \times 0.64} \\
# &\approx 0.56
# \end{aligned}
# \end{equation}
# $$
#
#
| Rethinking/end-of-chapter-practice-problems/ch-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GUARDADO DE DATOS EN BRUTO DE CADA CICLÓN
import xarray as xr
import numpy as np
import sys
import os
import pandas as pd
from geopy import distance
from mpl_toolkits.axes_grid1 import make_axes_locatable
from geographiclib.geodesic import Geodesic
import time
def get_bearing(lat1, lat2, long1, long2):
brng = Geodesic.WGS84.Inverse(lat1, long1, lat2, long2)['azi1']
return brng
# # Gráfico de celdas
from matplotlib.pylab import *
# +
start_time = time.time()
total_obs = []
#creación del mesh discretizado que quiero
th = np.array(range(0,370,10)) #discretización de ángulos
radius = np.array(range(0,525,25)) #discretización de radios
r, theta = np.meshgrid(radius,th) #mesh de r y theta
data = xr.open_dataset('WNP_cyclones/ciclon1.nc4')
data_id1 = np.array(data.id1)
data_id2 = np.array(data.id2)
dist = np.array(xr.open_dataarray('WNP_cyclones/dist_ciclon1.nc4'))
rad_ang = np.array(xr.open_dataarray('WNP_cyclones/angs_ciclon1.nc4'))
prec = np.array(xr.open_dataarray('WNP_cyclones/precip_ciclon1.nc4'))
#para seleccionar y cargar automaticamente el dato
for k in range(0,len(data.time)):
obs = np.zeros((r.shape))
for i in range(obs.shape[0]):
for j in range(obs.shape[1]):
if j == 20 and i == 36 :
index = np.intersect1d(np.where((np.ravel(dist[k,:data_id1[k],:data_id2[k]]) <= 500) & (np.ravel(dist[k,:data_id1[k],:data_id2[k]]) >= radius[j])),
np.where(((np.ravel(rad_ang[k,:data_id1[k],:data_id2[k]]) >= np.radians(th[i])))))
elif j == 20:
index = np.intersect1d(np.where((np.ravel(dist[k,:data_id1[k],:data_id2[k]]) <= 500) & (np.ravel(dist[k,:data_id1[k],:data_id2[k]]) >= radius[j])),
np.where((np.ravel(rad_ang[k,:data_id1[k],:data_id2[k]]) <= np.radians(th[i+1])) & (np.ravel(rad_ang[k,:data_id1[k],:data_id2[k]]) >= np.radians(th[i]))))
elif i == 36:
index = np.intersect1d(np.where((np.ravel(dist[k,:data_id1[k],:data_id2[k]]) <= radius[j+1]) & (np.ravel(dist[k,:data_id1[k],:data_id2[k]]) >= radius[j])),
np.where(((np.ravel(rad_ang[k,:data_id1[k],:data_id2[k]]) >= np.radians(th[i])))))
else:
index = np.intersect1d(np.where((np.ravel(dist[k,:data_id1[k],:data_id2[k]]) <= radius[j+1]) & (np.ravel(dist[k,:data_id1[k],:data_id2[k]]) >= radius[j])),
np.where((np.ravel(rad_ang[k,:data_id1[k],:data_id2[k]]) <= np.radians(th[i+1])) & (np.ravel(rad_ang[k,:data_id1[k],:data_id2[k]]) >= np.radians(th[i]))))
precip = prec[k,:data_id1[k],:data_id2[k]]
obs[i,j] = np.nanmean(np.ravel(precip)[index]) #cuando el index es un array vacío retorna nan, es decir no hay datos
total_obs.append(obs)
#plot
obs = np.nanmean(np.array(total_obs), axis = 0)
fig = plt.figure(figsize=(16, 10))
ax0 = fig.add_axes([0,0,0.4,0.4], projection='polar')
im = ax0.pcolormesh(np.radians(theta),r, obs, cmap='inferno_r', vmin = 0, vmax = 12)
ax0.set_theta_zero_location('N', offset=0)
ax0.set_theta_direction(-1)
ax0.set_title('Storm mean P(mm) inside 500 km radius\n', fontweight='bold')
ax0.set_thetagrids(range(0, 360, 10), labels = '')
ax0.set_rgrids(np.arange(0, 500, 25), labels='')
fig.colorbar(im)
ax0.grid('on')
plt.show()
print("--- %s seconds ---" % (time.time() - start_time))
# -
# # WNP
# ### EJEMPLO PARA TODOS LOS CICLONES
#
# Una vez verificado que funciona, lo hacemos para todos los ciclones
xds_tracks = xr.open_dataset('Allstorms.ibtracs_wmo.v03r09.nc')
#ciclones de la cuenca WNP, hay 418
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
wnp = xr.open_dataset('basinWNP.nc4')
wnp
# +
start_time = time.time()
for t in range(0,418):
storm = wnp.sel(storm = wnp.storm[t]) #seleccionamos la storm que corresponde en cada bucle
storm_pres = storm.presion[~np.isnan(storm.presion)]
storm_lat = storm.latitud[~np.isnan(storm.latitud)]
storm_lon = storm.longitud[~np.isnan(storm.longitud)]
storm_time = storm.fechas[~np.isnat(storm.fechas)]
all_dist = []
all_ang = []
all_prec = []
all_direction = []
all_sst = []
all_id1 = []
all_id2 = []
os.chdir('D:\oscar\dataset_modificado')
for k in range(0,len(storm_time)-1):
#parseamos a string la fecha correspondiente para leer el fichero nc4 correspondiente al instante de tiempo
ts = pd.to_datetime(str(storm_time.values[k]))
#necesario porque al ser una base de datos 3 horarias se han "colado" observaciones externas en ese rango
hora = ts.strftime('%H')
if hora != '00' and hora != '03' and hora != '06' and hora != '09' and hora != '12' and hora != '15' and hora != '18' and hora != '21':
continue
day = ts.strftime('%Y%m%d-%H.nc4')
precip = xr.open_dataset(day)
precip_lat = np.array(precip.nlat[~np.isnan(precip.nlat)])
precip_lon = np.array(precip.nlon[~np.isnan(precip.nlon)])
#seleccionamos el centro y su sucesivo
x = np.array(storm_lon[k])
y = np.array(storm_lat[k])
xnext = np.array(storm_lon[k+1])
ynext = np.array(storm_lat[k+1])
#seleccionamos sst del ciclón en ese instante
sst = xr.open_dataset('SST\sst.day.mean.'+str(ts.strftime('%Y'))+'.nc')
idx_time = np.where(pd.to_datetime(sst.time.values) == ts.strftime('%Y-%m-%d'))
idx_lat = np.where(np.round(sst.lat.values, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
#si idx_lat es vacío, redondeamos al decimal por abajo o por arriba
idx_lat2 = np.where(np.round(np.round(sst.lat.values, decimals = 1)-0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
idx_lat3 = np.where(np.round(np.round(sst.lat.values, decimals = 1)+0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
if len(idx_lat[0]) == 0:
idx_lat = idx_lat2
if len(idx_lat2[0]) == 0:
idx_lat = idx_lat3
#como la longitud en sst va de 0-360 y en storm_lon de -180-180, hacemos una pequeña conversión para buscar el índice
storm_lon_converted = storm_lon[k].values
if storm_lon_converted < 0:
storm_lon_converted = storm_lon_converted + 360
idx_lon = np.where(np.round(sst.lon.values, decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon2 = np.where(np.round(np.round(sst.lon.values, decimals = 1)-0.1,decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon3 = np.where(np.round(np.round(sst.lon.values, decimals = 1)+0.1, decimals = 1)== np.round(storm_lon_converted,decimals=1))
if len(idx_lon[0]) == 0:
idx_lon = idx_lon2
if len(idx_lon2[0]) == 0:
idx_lon = idx_lon3
#ahora que tenemos los 3 indices tiempo,longitud y latitud seleccionamos el valor de sst correspondiente
sst_value = sst.sst[idx_time[0],idx_lat[0],idx_lon[0]].values
#restringiremos los datos para optimizar el tiempo de computo a una resolucion 5x5
precip_lon = precip_lon[np.where((precip_lon <= x + 5) & (precip_lon >= x - 5))]
precip_lat = precip_lat[np.where((precip_lat <= y + 5) & (precip_lat >= y - 5))]
direction = get_bearing(y,ynext,x,xnext)
lons, lats = np.meshgrid(precip_lon,precip_lat)
dist = np.empty((40,40))
ang = np.empty((40,40))
dist[:] = np.nan
ang[:] = np.nan
#cálculo de mallas de ángulos y radios
for j in range(lons.shape[1]):
for i in range(lats.shape[0]):
coords = (lats[i,0], lons[0,j])
dist[j,i] = distance.geodesic(coords, (y,x)).km
ang[j,i] = np.rad2deg(np.arctan2((lats[i,0]-y), (lons[0,j]-x)))
if precip.precipitation.nlat.max() < y:
continue
x_min = np.min(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
x_max = np.max(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
y_min = np.min(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
y_max = np.max(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
precs = np.empty((40,40))
precs[:] = np.nan
prec = precip['precipitation'][0,x_min:x_max+1,y_min:y_max+1].values
for i in range(prec.shape[0]):
for j in range(prec.shape[1]):
if prec[i,j] > 500:
precs[i,j] = np.nan
else:
precs[i,j] = prec[i,j]
rad_ang = np.radians(ang)
change = np.radians(np.full((40,40), -direction))
rad_ang = rad_ang + change
rad_ang[rad_ang < 0] = np.radians(360+ np.degrees(rad_ang[rad_ang < 0]))
rad_ang = np.array(rad_ang)
dist = np.array(dist)
prec = np.array(prec)
all_direction.append(direction)
all_prec.append(precs)
all_ang.append(rad_ang)
all_dist.append(dist)
all_sst.append(sst_value[0][0][0])
all_id1.append(lons.shape[1])
all_id2.append(lats.shape[0])
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
#guardo archivos
multidata=xr.Dataset({'minPressure': ('time',storm_pres[:len(storm_time)-1]),
'latitude': ('time',storm_lat[:len(storm_time)-1]),
'longitude': ('time',storm_lon[:len(storm_time)-1]),
'azimuth': all_direction,
'sst': all_sst,
'id1': all_id1,
'id2': all_id2
},
{'time': storm_time[:len(storm_time)-1]})
multidata.to_netcdf('WNP_cyclones/ciclon'+str(t)+'.nc4')
dist = xr.DataArray(all_dist)
dist.to_netcdf('WNP_cyclones/dist_ciclon'+str(t)+'.nc4')
angs = xr.DataArray(all_ang)
angs.to_netcdf('WNP_cyclones/ang_ciclon'+str(t)+'.nc4')
precs = xr.DataArray(all_prec)
precs.to_netcdf('WNP_cyclones/precip_ciclon'+str(t)+'.nc4')
print('cyclone'+str(t)+'done!')
# -
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
import xarray as xr
xr.open_dataset('WNP_cyclones/ciclon417.nc4').azimuth
# # SP
sp = xr.open_dataset('basinSP.nc4')
sp
# +
start_time = time.time()
for t in range(0,178):
storm = sp.sel(storm = sp.storm[t]) #seleccionamos la storm que corresponde en cada bucle
storm_pres = storm.presion[~np.isnan(storm.presion)]
storm_lat = storm.latitud[~np.isnan(storm.latitud)]
storm_lon = storm.longitud[~np.isnan(storm.longitud)]
storm_time = storm.fechas[~np.isnat(storm.fechas)]
all_dist = []
all_ang = []
all_prec = []
all_direction = []
all_sst = []
all_id1 = []
all_id2 = []
os.chdir('D:\oscar\dataset_modificado')
for k in range(0,len(storm_time)-1):
#parseamos a string la fecha correspondiente para leer el fichero nc4 correspondiente al instante de tiempo
ts = pd.to_datetime(str(storm_time.values[k]))
#necesario porque al ser una base de datos 3 horarias se han "colado" observaciones externas en ese rango
hora = ts.strftime('%H')
if hora != '00' and hora != '03' and hora != '06' and hora != '09' and hora != '12' and hora != '15' and hora != '18' and hora != '21':
continue
day = ts.strftime('%Y%m%d-%H.nc4')
precip = xr.open_dataset(day)
precip_lat = np.array(precip.nlat[~np.isnan(precip.nlat)])
precip_lon = np.array(precip.nlon[~np.isnan(precip.nlon)])
#seleccionamos el centro y su sucesivo
x = np.array(storm_lon[k])
y = np.array(storm_lat[k])
xnext = np.array(storm_lon[k+1])
ynext = np.array(storm_lat[k+1])
#seleccionamos sst del ciclón en ese instante
sst = xr.open_dataset('SST\sst.day.mean.'+str(ts.strftime('%Y'))+'.nc')
idx_time = np.where(pd.to_datetime(sst.time.values) == ts.strftime('%Y-%m-%d'))
idx_lat = np.where(np.round(sst.lat.values, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
#si idx_lat es vacío, redondeamos al decimal por abajo o por arriba
idx_lat2 = np.where(np.round(np.round(sst.lat.values, decimals = 1)-0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
idx_lat3 = np.where(np.round(np.round(sst.lat.values, decimals = 1)+0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
if len(idx_lat[0]) == 0:
idx_lat = idx_lat2
if len(idx_lat2[0]) == 0:
idx_lat = idx_lat3
#como la longitud en sst va de 0-360 y en storm_lon de -180-180, hacemos una pequeña conversión para buscar el índice
storm_lon_converted = storm_lon[k].values
if storm_lon_converted < 0:
storm_lon_converted = storm_lon_converted + 360
idx_lon = np.where(np.round(sst.lon.values, decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon2 = np.where(np.round(np.round(sst.lon.values, decimals = 1)-0.1,decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon3 = np.where(np.round(np.round(sst.lon.values, decimals = 1)+0.1, decimals = 1)== np.round(storm_lon_converted,decimals=1))
if len(idx_lon[0]) == 0:
idx_lon = idx_lon2
if len(idx_lon2[0]) == 0:
idx_lon = idx_lon3
#ahora que tenemos los 3 indices tiempo,longitud y latitud seleccionamos el valor de sst correspondiente
sst_value = sst.sst[idx_time[0],idx_lat[0],idx_lon[0]].values
#restringiremos los datos para optimizar el tiempo de computo a una resolucion 5x5
precip_lon = precip_lon[np.where((precip_lon <= x + 5) & (precip_lon >= x - 5))]
precip_lat = precip_lat[np.where((precip_lat <= y + 5) & (precip_lat >= y - 5))]
direction = get_bearing(y,ynext,x,xnext)
lons, lats = np.meshgrid(precip_lon,precip_lat)
dist = np.empty((40,40))
ang = np.empty((40,40))
dist[:] = np.nan
ang[:] = np.nan
#cálculo de mallas de ángulos y radios
for j in range(lons.shape[1]):
for i in range(lats.shape[0]):
coords = (lats[i,0], lons[0,j])
dist[j,i] = distance.geodesic(coords, (y,x)).km
ang[j,i] = np.rad2deg(np.arctan2((lats[i,0]-y), (lons[0,j]-x)))
if precip.precipitation.nlat.min() > y:
continue
x_min = np.min(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
x_max = np.max(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
y_min = np.min(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
y_max = np.max(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
precs = np.empty((40,40))
precs[:] = np.nan
prec = precip['precipitation'][0,x_min:x_max+1,y_min:y_max+1].values
for i in range(prec.shape[0]):
for j in range(prec.shape[1]):
if prec[i,j] > 500:
precs[i,j] = np.nan
else:
precs[i,j] = prec[i,j]
rad_ang = np.radians(ang)
change = np.radians(np.full((40,40), -direction))
rad_ang = rad_ang + change
rad_ang[rad_ang < 0] = np.radians(360+ np.degrees(rad_ang[rad_ang < 0]))
rad_ang = np.array(rad_ang)
dist = np.array(dist)
prec = np.array(prec)
all_direction.append(direction)
all_prec.append(precs)
all_ang.append(rad_ang)
all_dist.append(dist)
all_sst.append(sst_value[0][0][0])
all_id1.append(lons.shape[1])
all_id2.append(lats.shape[0])
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
#guardo archivos
multidata=xr.Dataset({'minPressure': ('time',storm_pres[:len(storm_time)-1]),
'latitude': ('time',storm_lat[:len(storm_time)-1]),
'longitude': ('time',storm_lon[:len(storm_time)-1]),
'azimuth': all_direction,
'sst': all_sst,
'id1': all_id1,
'id2': all_id2
},
{'time': storm_time[:len(storm_time)-1]})
multidata.to_netcdf('SP_cyclones/ciclonsp'+str(t)+'.nc4')
dist = xr.DataArray(all_dist)
dist.to_netcdf('SP_cyclones/dist_ciclonsp'+str(t)+'.nc4')
angs = xr.DataArray(all_ang)
angs.to_netcdf('SP_cyclones/ang_ciclonsp'+str(t)+'.nc4')
precs = xr.DataArray(all_prec)
precs.to_netcdf('SP_cyclones/precip_ciclonsp'+str(t)+'.nc4')
print('cyclone'+str(t)+'done!')
# -
xr.open_dataset('SP_cyclones/ciclonsp177.nc4')
# ### Modificando el ángulo en SP
sp = xr.open_dataset('basinSP.nc4')
sp
# +
start_time = time.time()
for t in range(0,178):
storm = sp.sel(storm = sp.storm[t]) #seleccionamos la storm que corresponde en cada bucle
storm_pres = storm.presion[~np.isnan(storm.presion)]
storm_lat = storm.latitud[~np.isnan(storm.latitud)]
storm_lon = storm.longitud[~np.isnan(storm.longitud)]
storm_time = storm.fechas[~np.isnat(storm.fechas)]
all_dist = []
all_ang = []
all_prec = []
all_direction = []
all_sst = []
all_id1 = []
all_id2 = []
os.chdir('D:\oscar\dataset_modificado')
for k in range(0,len(storm_time)-1):
#parseamos a string la fecha correspondiente para leer el fichero nc4 correspondiente al instante de tiempo
ts = pd.to_datetime(str(storm_time.values[k]))
#necesario porque al ser una base de datos 3 horarias se han "colado" observaciones externas en ese rango
hora = ts.strftime('%H')
if hora != '00' and hora != '03' and hora != '06' and hora != '09' and hora != '12' and hora != '15' and hora != '18' and hora != '21':
continue
day = ts.strftime('%Y%m%d-%H.nc4')
precip = xr.open_dataset(day)
precip_lat = np.array(precip.nlat[~np.isnan(precip.nlat)])
precip_lon = np.array(precip.nlon[~np.isnan(precip.nlon)])
#seleccionamos el centro y su sucesivo
x = np.array(storm_lon[k])
y = np.array(storm_lat[k])
xnext = np.array(storm_lon[k+1])
ynext = np.array(storm_lat[k+1])
#seleccionamos sst del ciclón en ese instante
sst = xr.open_dataset('SST\sst.day.mean.'+str(ts.strftime('%Y'))+'.nc')
idx_time = np.where(pd.to_datetime(sst.time.values) == ts.strftime('%Y-%m-%d'))
idx_lat = np.where(np.round(sst.lat.values, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
#si idx_lat es vacío, redondeamos al decimal por abajo o por arriba
idx_lat2 = np.where(np.round(np.round(sst.lat.values, decimals = 1)-0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
idx_lat3 = np.where(np.round(np.round(sst.lat.values, decimals = 1)+0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
if len(idx_lat[0]) == 0:
idx_lat = idx_lat2
if len(idx_lat2[0]) == 0:
idx_lat = idx_lat3
#como la longitud en sst va de 0-360 y en storm_lon de -180-180, hacemos una pequeña conversión para buscar el índice
storm_lon_converted = storm_lon[k].values
if storm_lon_converted < 0:
storm_lon_converted = storm_lon_converted + 360
idx_lon = np.where(np.round(sst.lon.values, decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon2 = np.where(np.round(np.round(sst.lon.values, decimals = 1)-0.1,decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon3 = np.where(np.round(np.round(sst.lon.values, decimals = 1)+0.1, decimals = 1)== np.round(storm_lon_converted,decimals=1))
if len(idx_lon[0]) == 0:
idx_lon = idx_lon2
if len(idx_lon2[0]) == 0:
idx_lon = idx_lon3
#ahora que tenemos los 3 indices tiempo,longitud y latitud seleccionamos el valor de sst correspondiente
sst_value = sst.sst[idx_time[0],idx_lat[0],idx_lon[0]].values
#restringiremos los datos para optimizar el tiempo de computo a una resolucion 5x5
precip_lon = precip_lon[np.where((precip_lon <= x + 5) & (precip_lon >= x - 5))]
precip_lat = precip_lat[np.where((precip_lat <= y + 5) & (precip_lat >= y - 5))]
direction = get_bearing(y,ynext,x,xnext)
lons, lats = np.meshgrid(precip_lon,precip_lat)
dist = np.empty((40,40))
ang = np.empty((40,40))
dist[:] = np.nan
ang[:] = np.nan
#cálculo de mallas de ángulos y radios
for j in range(lons.shape[1]):
for i in range(lats.shape[0]):
coords = (lats[i,0], lons[0,j])
dist[j,i] = distance.geodesic(coords, (y,x)).km
ang[j,i] = np.rad2deg(np.arctan2((lats[i,0]-y), (lons[0,j]-x)))
if precip.precipitation.nlat.min() > y:
continue
x_min = np.min(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
x_max = np.max(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
y_min = np.min(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
y_max = np.max(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
precs = np.empty((40,40))
precs[:] = np.nan
prec = precip['precipitation'][0,x_min:x_max+1,y_min:y_max+1].values
for i in range(prec.shape[0]):
for j in range(prec.shape[1]):
if prec[i,j] > 500:
precs[i,j] = np.nan
else:
precs[i,j] = prec[i,j]
ang = 360 - ang
rad_ang = np.radians(ang)
change = np.radians(np.full((40,40), -direction))
rad_ang = rad_ang + change
rad_ang[rad_ang < 0] = np.radians(360+ np.degrees(rad_ang[rad_ang < 0]))
rad_ang = np.array(rad_ang)
dist = np.array(dist)
prec = np.array(prec)
all_direction.append(direction)
all_prec.append(precs)
all_ang.append(rad_ang)
all_dist.append(dist)
all_sst.append(sst_value[0][0][0])
all_id1.append(lons.shape[1])
all_id2.append(lats.shape[0])
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
#guardo archivos
multidata=xr.Dataset({'minPressure': ('time',storm_pres[:len(storm_time)-1]),
'latitude': ('time',storm_lat[:len(storm_time)-1]),
'longitude': ('time',storm_lon[:len(storm_time)-1]),
'azimuth': all_direction,
'sst': all_sst,
'id1': all_id1,
'id2': all_id2
},
{'time': storm_time[:len(storm_time)-1]})
multidata.to_netcdf('SP_cyclones_mod/ciclonsp_mod'+str(t)+'.nc4')
dist = xr.DataArray(all_dist)
dist.to_netcdf('SP_cyclones_mod/dist_ciclonsp_mod'+str(t)+'.nc4')
angs = xr.DataArray(all_ang)
angs.to_netcdf('SP_cyclones_mod/ang_ciclonsp_mod'+str(t)+'.nc4')
precs = xr.DataArray(all_prec)
precs.to_netcdf('SP_cyclones_mod/precip_ciclonsp_mod'+str(t)+'.nc4')
print('cyclone'+str(t)+'done!')
# -
# # NA + Caribbean
na = xr.open_dataset('basinNA.nc4')
na
# +
start_time = time.time()
for t in range(0,300):
storm = na.sel(storm = na.storm[t]) #seleccionamos la storm que corresponde en cada bucle
storm_pres = storm.presion[~np.isnan(storm.presion)]
storm_lat = storm.latitud[~np.isnan(storm.latitud)]
storm_lon = storm.longitud[~np.isnan(storm.longitud)]
storm_time = storm.fechas[~np.isnat(storm.fechas)]
all_dist = []
all_ang = []
all_prec = []
all_direction = []
all_sst = []
all_id1 = []
all_id2 = []
os.chdir('D:\oscar\dataset_modificado')
for k in range(0,len(storm_time)-1):
#parseamos a string la fecha correspondiente para leer el fichero nc4 correspondiente al instante de tiempo
ts = pd.to_datetime(str(storm_time.values[k]))
#necesario porque al ser una base de datos 3 horarias se han "colado" observaciones externas en ese rango
hora = ts.strftime('%H')
if hora != '00' and hora != '03' and hora != '06' and hora != '09' and hora != '12' and hora != '15' and hora != '18' and hora != '21':
continue
day = ts.strftime('%Y%m%d-%H.nc4')
precip = xr.open_dataset(day)
precip_lat = np.array(precip.nlat[~np.isnan(precip.nlat)])
precip_lon = np.array(precip.nlon[~np.isnan(precip.nlon)])
#seleccionamos el centro y su sucesivo
x = np.array(storm_lon[k])
y = np.array(storm_lat[k])
xnext = np.array(storm_lon[k+1])
ynext = np.array(storm_lat[k+1])
#seleccionamos sst del ciclón en ese instante
sst = xr.open_dataset('SST\sst.day.mean.'+str(ts.strftime('%Y'))+'.nc')
idx_time = np.where(pd.to_datetime(sst.time.values) == ts.strftime('%Y-%m-%d'))
idx_lat = np.where(np.round(sst.lat.values, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
#si idx_lat es vacío, redondeamos al decimal por abajo o por arriba
idx_lat2 = np.where(np.round(np.round(sst.lat.values, decimals = 1)-0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
idx_lat3 = np.where(np.round(np.round(sst.lat.values, decimals = 1)+0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
if len(idx_lat[0]) == 0:
idx_lat = idx_lat2
if len(idx_lat2[0]) == 0:
idx_lat = idx_lat3
#como la longitud en sst va de 0-360 y en storm_lon de -180-180, hacemos una pequeña conversión para buscar el índice
storm_lon_converted = storm_lon[k].values
if storm_lon_converted < 0:
storm_lon_converted = storm_lon_converted + 360
idx_lon = np.where(np.round(sst.lon.values, decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon2 = np.where(np.round(np.round(sst.lon.values, decimals = 1)-0.1,decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon3 = np.where(np.round(np.round(sst.lon.values, decimals = 1)+0.1, decimals = 1)== np.round(storm_lon_converted,decimals=1))
if len(idx_lon[0]) == 0:
idx_lon = idx_lon2
if len(idx_lon2[0]) == 0:
idx_lon = idx_lon3
#ahora que tenemos los 3 indices tiempo,longitud y latitud seleccionamos el valor de sst correspondiente
sst_value = sst.sst[idx_time[0],idx_lat[0],idx_lon[0]].values
#restringiremos los datos para optimizar el tiempo de computo a una resolucion 5x5
precip_lon = precip_lon[np.where((precip_lon <= x + 5) & (precip_lon >= x - 5))]
precip_lat = precip_lat[np.where((precip_lat <= y + 5) & (precip_lat >= y - 5))]
direction = get_bearing(y,ynext,x,xnext)
lons, lats = np.meshgrid(precip_lon,precip_lat)
dist = np.empty((40,40))
ang = np.empty((40,40))
dist[:] = np.nan
ang[:] = np.nan
#cálculo de mallas de ángulos y radios
for j in range(lons.shape[1]):
for i in range(lats.shape[0]):
coords = (lats[i,0], lons[0,j])
dist[j,i] = distance.geodesic(coords, (y,x)).km
ang[j,i] = np.rad2deg(np.arctan2((lats[i,0]-y), (lons[0,j]-x)))
if precip.precipitation.nlat.max() < y:
continue
x_min = np.min(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
x_max = np.max(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
y_min = np.min(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
y_max = np.max(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
precs = np.empty((40,40))
precs[:] = np.nan
prec = precip['precipitation'][0,x_min:x_max+1,y_min:y_max+1].values
for i in range(prec.shape[0]):
for j in range(prec.shape[1]):
if prec[i,j] > 500:
precs[i,j] = np.nan
else:
precs[i,j] = prec[i,j]
rad_ang = np.radians(ang)
change = np.radians(np.full((40,40), -direction))
rad_ang = rad_ang + change
rad_ang[rad_ang < 0] = np.radians(360+ np.degrees(rad_ang[rad_ang < 0]))
rad_ang = np.array(rad_ang)
dist = np.array(dist)
prec = np.array(prec)
all_direction.append(direction)
all_prec.append(precs)
all_ang.append(rad_ang)
all_dist.append(dist)
all_sst.append(sst_value[0][0][0])
all_id1.append(lons.shape[1])
all_id2.append(lats.shape[0])
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
#guardo archivos
multidata=xr.Dataset({'minPressure': ('time',storm_pres[:len(storm_time)-1]),
'latitude': ('time',storm_lat[:len(storm_time)-1]),
'longitude': ('time',storm_lon[:len(storm_time)-1]),
'azimuth': all_direction,
'sst': all_sst,
'id1': all_id1,
'id2': all_id2
},
{'time': storm_time[:len(storm_time)-1]})
multidata.to_netcdf('NA_cyclones/ciclonna'+str(t)+'.nc4')
dist = xr.DataArray(all_dist)
dist.to_netcdf('NA_cyclones/dist_ciclonna'+str(t)+'.nc4')
angs = xr.DataArray(all_ang)
angs.to_netcdf('NA_cyclones/ang_ciclonna'+str(t)+'.nc4')
precs = xr.DataArray(all_prec)
precs.to_netcdf('NA_cyclones/precip_ciclonna'+str(t)+'.nc4')
print('cyclone'+str(t)+'done!')
# -
# # EP
ep = xr.open_dataset('basinEP.nc4')
ep
# +
start_time = time.time()
for t in range(0,335):
storm = ep.sel(storm = ep.storm[t]) #seleccionamos la storm que corresponde en cada bucle
storm_pres = storm.presion[~np.isnan(storm.presion)]
storm_lat = storm.latitud[~np.isnan(storm.latitud)]
storm_lon = storm.longitud[~np.isnan(storm.longitud)]
storm_time = storm.fechas[~np.isnat(storm.fechas)]
all_dist = []
all_ang = []
all_prec = []
all_direction = []
all_sst = []
all_id1 = []
all_id2 = []
os.chdir('D:\oscar\dataset_modificado')
for k in range(0,len(storm_time)-1):
#parseamos a string la fecha correspondiente para leer el fichero nc4 correspondiente al instante de tiempo
ts = pd.to_datetime(str(storm_time.values[k]))
#necesario porque al ser una base de datos 3 horarias se han "colado" observaciones externas en ese rango
hora = ts.strftime('%H')
if hora != '00' and hora != '03' and hora != '06' and hora != '09' and hora != '12' and hora != '15' and hora != '18' and hora != '21':
continue
day = ts.strftime('%Y%m%d-%H.nc4')
precip = xr.open_dataset(day)
precip_lat = np.array(precip.nlat[~np.isnan(precip.nlat)])
precip_lon = np.array(precip.nlon[~np.isnan(precip.nlon)])
#seleccionamos el centro y su sucesivo
x = np.array(storm_lon[k])
y = np.array(storm_lat[k])
xnext = np.array(storm_lon[k+1])
ynext = np.array(storm_lat[k+1])
#seleccionamos sst del ciclón en ese instante
sst = xr.open_dataset('SST\sst.day.mean.'+str(ts.strftime('%Y'))+'.nc')
idx_time = np.where(pd.to_datetime(sst.time.values) == ts.strftime('%Y-%m-%d'))
idx_lat = np.where(np.round(sst.lat.values, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
#si idx_lat es vacío, redondeamos al decimal por abajo o por arriba
idx_lat2 = np.where(np.round(np.round(sst.lat.values, decimals = 1)-0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
idx_lat3 = np.where(np.round(np.round(sst.lat.values, decimals = 1)+0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
if len(idx_lat[0]) == 0:
idx_lat = idx_lat2
if len(idx_lat2[0]) == 0:
idx_lat = idx_lat3
#como la longitud en sst va de 0-360 y en storm_lon de -180-180, hacemos una pequeña conversión para buscar el índice
storm_lon_converted = storm_lon[k].values
if storm_lon_converted < 0:
storm_lon_converted = storm_lon_converted + 360
idx_lon = np.where(np.round(sst.lon.values, decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon2 = np.where(np.round(np.round(sst.lon.values, decimals = 1)-0.1,decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon3 = np.where(np.round(np.round(sst.lon.values, decimals = 1)+0.1, decimals = 1)== np.round(storm_lon_converted,decimals=1))
if len(idx_lon[0]) == 0:
idx_lon = idx_lon2
if len(idx_lon2[0]) == 0:
idx_lon = idx_lon3
#ahora que tenemos los 3 indices tiempo,longitud y latitud seleccionamos el valor de sst correspondiente
sst_value = sst.sst[idx_time[0],idx_lat[0],idx_lon[0]].values
#restringiremos los datos para optimizar el tiempo de computo a una resolucion 5x5
precip_lon = precip_lon[np.where((precip_lon <= x + 5) & (precip_lon >= x - 5))]
precip_lat = precip_lat[np.where((precip_lat <= y + 5) & (precip_lat >= y - 5))]
direction = get_bearing(y,ynext,x,xnext)
lons, lats = np.meshgrid(precip_lon,precip_lat)
dist = np.empty((40,40))
ang = np.empty((40,40))
dist[:] = np.nan
ang[:] = np.nan
#cálculo de mallas de ángulos y radios
for j in range(lons.shape[1]):
for i in range(lats.shape[0]):
coords = (lats[i,0], lons[0,j])
dist[j,i] = distance.geodesic(coords, (y,x)).km
ang[j,i] = np.rad2deg(np.arctan2((lats[i,0]-y), (lons[0,j]-x)))
if precip.precipitation.nlat.max() < y:
continue
x_min = np.min(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
x_max = np.max(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
y_min = np.min(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
y_max = np.max(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
precs = np.empty((40,40))
precs[:] = np.nan
prec = precip['precipitation'][0,x_min:x_max+1,y_min:y_max+1].values
for i in range(prec.shape[0]):
for j in range(prec.shape[1]):
if prec[i,j] > 500:
precs[i,j] = np.nan
else:
precs[i,j] = prec[i,j]
rad_ang = np.radians(ang)
change = np.radians(np.full((40,40), -direction))
rad_ang = rad_ang + change
rad_ang[rad_ang < 0] = np.radians(360+ np.degrees(rad_ang[rad_ang < 0]))
rad_ang = np.array(rad_ang)
dist = np.array(dist)
prec = np.array(prec)
all_direction.append(direction)
all_prec.append(precs)
all_ang.append(rad_ang)
all_dist.append(dist)
all_sst.append(sst_value[0][0][0])
all_id1.append(lons.shape[1])
all_id2.append(lats.shape[0])
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
#guardo archivos
multidata=xr.Dataset({'minPressure': ('time',storm_pres[:len(storm_time)-1]),
'latitude': ('time',storm_lat[:len(storm_time)-1]),
'longitude': ('time',storm_lon[:len(storm_time)-1]),
'azimuth': all_direction,
'sst': all_sst,
'id1': all_id1,
'id2': all_id2
},
{'time': storm_time[:len(storm_time)-1]})
multidata.to_netcdf('EP_cyclones/ciclonep'+str(t)+'.nc4')
dist = xr.DataArray(all_dist)
dist.to_netcdf('EP_cyclones/dist_ciclonep'+str(t)+'.nc4')
angs = xr.DataArray(all_ang)
angs.to_netcdf('EP_cyclones/ang_ciclonep'+str(t)+'.nc4')
precs = xr.DataArray(all_prec)
precs.to_netcdf('EP_cyclones/precip_ciclonep'+str(t)+'.nc4')
print('cyclone'+str(t)+'done!')
# -
# # Todos los ciclones
all_basin = xr.open_dataset('All_basin.nc4')
all_basin
# +
start_time = time.time()
for t in range(0,1636):
storm = all_basin.sel(storm = all_basin.storm[t]) #seleccionamos la storm que corresponde en cada bucle
storm_pres = storm.presion[~np.isnan(storm.presion)]
storm_lat = storm.latitud[~np.isnan(storm.latitud)]
storm_lon = storm.longitud[~np.isnan(storm.longitud)]
storm_time = storm.fechas[~np.isnat(storm.fechas)]
storm_basin = storm.cuenca[~np.isnan(storm.cuenca)]
storm_distland = storm.distancia_tierra[~np.isnan(storm.distancia_tierra)]
all_dist = []
all_ang = []
all_prec = []
all_direction = []
all_sst = []
all_id1 = []
all_id2 = []
os.chdir('D:\oscar\dataset_modificado')
for k in range(0,len(storm_time)-1):
#parseamos a string la fecha correspondiente para leer el fichero nc4 correspondiente al instante de tiempo
ts = pd.to_datetime(str(storm_time.values[k]))
#necesario porque al ser una base de datos 3 horarias se han "colado" observaciones externas en ese rango
hora = ts.strftime('%H')
if hora != '00' and hora != '03' and hora != '06' and hora != '09' and hora != '12' and hora != '15' and hora != '18' and hora != '21':
continue
day = ts.strftime('%Y%m%d-%H.nc4')
precip = xr.open_dataset(day)
precip_lat = np.array(precip.nlat[~np.isnan(precip.nlat)])
precip_lon = np.array(precip.nlon[~np.isnan(precip.nlon)])
#seleccionamos el centro y su sucesivo
x = np.array(storm_lon[k])
y = np.array(storm_lat[k])
xnext = np.array(storm_lon[k+1])
ynext = np.array(storm_lat[k+1])
#seleccionamos sst del ciclón en ese instante
sst = xr.open_dataset('SST\sst.day.mean.'+str(ts.strftime('%Y'))+'.nc')
idx_time = np.where(pd.to_datetime(sst.time.values) == ts.strftime('%Y-%m-%d'))
idx_lat = np.where(np.round(sst.lat.values, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
#si idx_lat es vacío, redondeamos al decimal por abajo o por arriba
idx_lat2 = np.where(np.round(np.round(sst.lat.values, decimals = 1)-0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
idx_lat3 = np.where(np.round(np.round(sst.lat.values, decimals = 1)+0.1, decimals = 1) == np.round(storm_lat[k].values,decimals=1))
if len(idx_lat[0]) == 0:
idx_lat = idx_lat2
if len(idx_lat2[0]) == 0:
idx_lat = idx_lat3
#como la longitud en sst va de 0-360 y en storm_lon de -180-180, hacemos una pequeña conversión para buscar el índice
storm_lon_converted = storm_lon[k].values
if storm_lon_converted < 0:
storm_lon_converted = storm_lon_converted + 360
idx_lon = np.where(np.round(sst.lon.values, decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon2 = np.where(np.round(np.round(sst.lon.values, decimals = 1)-0.1,decimals = 1) == np.round(storm_lon_converted,decimals=1))
idx_lon3 = np.where(np.round(np.round(sst.lon.values, decimals = 1)+0.1, decimals = 1)== np.round(storm_lon_converted,decimals=1))
if len(idx_lon[0]) == 0:
idx_lon = idx_lon2
if len(idx_lon2[0]) == 0:
idx_lon = idx_lon3
#ahora que tenemos los 3 indices tiempo,longitud y latitud seleccionamos el valor de sst correspondiente
sst_value = sst.sst[idx_time[0],idx_lat[0],idx_lon[0]].values
#restringiremos los datos para optimizar el tiempo de computo a una resolucion 5x5
precip_lon = precip_lon[np.where((precip_lon <= x + 5) & (precip_lon >= x - 5))]
precip_lat = precip_lat[np.where((precip_lat <= y + 5) & (precip_lat >= y - 5))]
direction = get_bearing(y,ynext,x,xnext)
lons, lats = np.meshgrid(precip_lon,precip_lat)
dist = np.empty((40,40))
ang = np.empty((40,40))
dist[:] = np.nan
ang[:] = np.nan
#cálculo de mallas de ángulos y radios
for j in range(lons.shape[1]):
for i in range(lats.shape[0]):
coords = (lats[i,0], lons[0,j])
dist[j,i] = distance.geodesic(coords, (y,x)).km
ang[j,i] = np.rad2deg(np.arctan2((lats[i,0]-y), (lons[0,j]-x)))
if precip.precipitation.nlat.max() < np.abs(y):
continue
x_min = np.min(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
x_max = np.max(np.where((precip.precipitation.nlon <= x + 5) & (precip.precipitation.nlon >= x - 5)))
y_min = np.min(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
y_max = np.max(np.where((precip.precipitation.nlat <= y + 5) & (precip.precipitation.nlat >= y - 5)))
precs = np.empty((40,40))
precs[:] = np.nan
prec = precip['precipitation'][0,x_min:x_max+1,y_min:y_max+1].values
for i in range(prec.shape[0]):
for j in range(prec.shape[1]):
if prec[i,j] > 500:
precs[i,j] = np.nan
else:
precs[i,j] = prec[i,j]
rad_ang = np.radians(ang)
change = np.radians(np.full((40,40), -direction))
rad_ang = rad_ang + change
rad_ang[rad_ang < 0] = np.radians(360+ np.degrees(rad_ang[rad_ang < 0]))
rad_ang = np.array(rad_ang)
dist = np.array(dist)
prec = np.array(prec)
all_direction.append(direction)
all_prec.append(precs)
all_ang.append(rad_ang)
all_dist.append(dist)
all_sst.append(sst_value[0][0][0])
all_id1.append(lons.shape[1])
all_id2.append(lats.shape[0])
os.chdir('C:\\Users\\usuario\\Desktop\\trazasIB')
#guardo archivos
multidata=xr.Dataset({'minPressure': ('time',storm_pres[:len(storm_time)-1]),
'latitude': ('time',storm_lat[:len(storm_time)-1]),
'longitude': ('time',storm_lon[:len(storm_time)-1]),
'azimuth': all_direction,
'sst': all_sst,
'id1': all_id1,
'id2': all_id2,
'distland': ('time',storm_distland[:len(storm_time)-1]),
'basin':('time', storm_basin[:len(storm_time)-1])
},
{'time': storm_time[:len(storm_time)-1]})
multidata.to_netcdf('All_cyclones/ciclon_all'+str(t)+'.nc4')
dist = xr.DataArray(all_dist)
dist.to_netcdf('All_cyclones/dist_ciclon_all'+str(t)+'.nc4')
angs = xr.DataArray(all_ang)
angs.to_netcdf('All_cyclones/ang_ciclon_all'+str(t)+'.nc4')
precs = xr.DataArray(all_prec)
precs.to_netcdf('All_cyclones/precip_ciclon_all'+str(t)+'.nc4')
print('cyclone'+str(t)+'done!')
| script_generacion_dataset/cyclone_files_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## How to complete a Kaggle Competition with Machine Learning
# In this code along session, you'll build several algorithms of increasing complexity that predict whether any given passenger on the Titanic survived or not, given data on them such as the fare they paid, where they embarked and their age.
#
# <img src="img/nytimes.jpg" width="500">
# In particular, you'll build _supervised learning_ models. _Supervised learning_ is the branch of machine learning (ML) that involves predicting labels, such as 'Survived' or 'Not'. Such models:
#
# * it learns from labelled data, e.g. data that includes whether a passenger survived (called model training).
# * and then predicts on unlabelled data.
#
# On Kaggle, a platform for predictive modelling and analytics competitions, these are called train and test sets because
#
# * You want to build a model that learns patterns in the training set
# * You _then_ use the model to make predictions on the test set!
#
# Kaggle then tells you the **percentage that you got correct**: this is known as the _accuracy_ of your model.
# ## Approach
#
# A good way to approach supervised learning:
#
# * Exploratory Data Analysis (EDA);
# * Build a quick and dirty model (baseline);
# * Iterate;
# * Engineer features;
# * Get model that performs better.
#
# In this code along session, we'll do all of these! We also have free courses that get you up and running with machine learning for the Titanic dataset in [Python](https://campus.datacamp.com/courses/kaggle-python-tutorial-on-machine-learning) and [R](https://campus.datacamp.com/courses/kaggle-r-tutorial-on-machine-learning).
# **Note:** We may move quickly at some points in order to get a bit further along. I'll answer questions in the live event but also feel free to chime in and help each other in the comments.
# ## Import you data and check it out
# +
# Import modules
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import tree
from sklearn.metrics import accuracy_score
# Figures inline and set visualization style
# %matplotlib inline
sns.set()
# +
# Import test and train datasets
df_train = ____
df_test = ____
# View first lines of training data
____
# -
# * What are all these features? Check out the Kaggle data documentation [here](https://www.kaggle.com/c/titanic/data).
#
# **Important note on terminology:**
# * The _target variable_ is the one you are trying to predict;
# * Other variables are known as _features_ (or _predictor variables_).
# View first lines of test data
____
# * Use the DataFrame `.info()` method to check out datatypes, missing values and more (of `df_train`).
____
# * Use the DataFrame `.describe()` method to check out summary statistics of numeric columns (of `df_train`).
____
# **Recap:**
# * you've loaded your data and had a look at it.
#
# **Up next:** Explore your data visually and build a first model!
#
# For more on `pandas`, check out our [Data Manipulation with Python track](https://www.datacamp.com/tracks/data-manipulation-with-python).
#
# If you're enoying this session, retweet or share on FB now and follow us on Twitter: [@hugobowne](https://twitter.com/hugobowne) & [@DataCamp](https://twitter.com/datacamp).
# ## Visual exploratory data analysis and your first model
# * Use `seaborn` to build a bar plot of Titanic survival (your _target variable_).
____
# **Take-away:** In the training set, less people survived than didn't. Let's then build a first model that **predict that nobody survived**.
#
# This is a bad model as we know that people survived. But it gives us a _baseline_: any model that we build later needs to do better than this one.
# * Create a column 'Survived' for `df_test` that encodes 'did not survive' for all rows;
# * Save 'PassengerId' and 'Survived' columns of `df_test` to a .csv and submit to Kaggle.
df_test['Survived'] = 0
df_test[['PassengerId', 'Survived']].____
# * What accuracy did this give you?
#
# Accuracy on Kaggle = ??
#
# **Essential note!** There are metrics other than accuracy that you may want to use.
# **Recap:**
# * you've loaded your data and had a look at it.
# * you've explored your target variable visually and made your first predictions.
#
# **Up next:** More EDA and you'll build another model.
# ## EDA on feature variables
# * Use `seaborn` to build a bar plot of the Titanic dataset feature 'Sex' (of `df_train`).
____
# * Use `seaborn` to build bar plots of the Titanic dataset feature 'Survived' split (faceted) over the feature 'Sex'.
____
# **Take-away:** Women were more likely to survive than men.
# * Use `pandas` to figure out how many women and how many men survived.
____
# * Use `pandas` to figure out the proportion of women that survived, along with the proportion of men:
print(df_train[df_train.Sex == 'female'].Survived.sum()/df_train[df_train.Sex == 'female'].Survived.count())
print(df_train[df_train.Sex == 'male'].Survived.sum()/df_train[df_train.Sex == 'male'].Survived.count())
# 74% of women survived, while 18% of men survived.
#
# Let's now build a second model and predict that all women survived and all men didn't. Once again, this is an unrealistic model, but it will provide a baseline against which to compare future models.
# * Create a column 'Survived' for `df_test` that encodes the above prediction.
# * Save 'PassengerId' and 'Survived' columns of `df_test` to a .csv and submit to Kaggle.
df_test['Survived'] = ____
df_test[['PassengerId', 'Survived']].to_csv('data/predictions/women_survive.csv', index=False)
# * What accuracy did this give you?
#
# Accuracy on Kaggle = ??
# **Recap:**
# * you've loaded your data and had a look at it.
# * you've explored your target variable visually and made your first predictions.
# * you've explored some of your feature variables visually and made more predictions that did better based on your EDA.
#
# **Up next:** EDA of other feature variables, categorical and numeric.
#
# For more on `pandas`, check out our [Data Manipulation with Python track](https://www.datacamp.com/tracks/data-manipulation-with-python).
#
# For more on `seaborn`, check out Chapter 3 of our [Intro. to Datavis with Python course](https://www.datacamp.com/courses/introduction-to-data-visualization-with-python).
#
# If you're enoying this session, retweet or share on FB now and follow us on Twitter: [@hugobowne](https://twitter.com/hugobowne) & [@DataCamp](https://twitter.com/datacamp).
# ## Explore your data more!
# * Use `seaborn` to build bar plots of the Titanic dataset feature 'Survived' split (faceted) over the feature 'Pclass'.
____
# **Take-away:** [Include take-away from figure here]
# * Use `seaborn` to build bar plots of the Titanic dataset feature 'Survived' split (faceted) over the feature 'Embarked'.
____
# **Take-away:** [Include take-away from figure here]
# ## EDA with numeric variables
# * Use `seaborn` to plot a histogram of the 'Fare' column of `df_train`.
____
# **Take-away:** [Include take-away from figure here]
# * Use a `pandas` plotting method to plot the column 'Fare' for each value of 'Survived' on the same plot.
____
# **Take-away:** [Include take-away from figure here]
# * Use `seaborn` to plot a histogram of the 'Age' column of `df_train`. _Hint_: you may need to drop null values before doing so.
df_train_drop = ____
____
# **Take-away:** [Include take-away from figure here]
# * Plot a strip plot & a swarm plot of 'Fare' with 'Survived' on the x-axis.
____
____
# **Take-away:** [Include take-away from figure here]
# * Use the DataFrame method `.describe()` to check out summary statistics of 'Fare' as a function of survival.
____
# * Use `seaborn` to plot a scatter plot of 'Age' against 'Fare', colored by 'Survived'.
____
# **Take-away:** [Include take-away from figure here]
# * Use `seaborn` to create a pairplot of `df_train`, colored by 'Survived'.
____
# **Take-away:** [Include take-away from figure here]
# **Recap:**
# * you've loaded your data and had a look at it.
# * you've explored your target variable visually and made your first predictions.
# * you've explored some of your feature variables visually and made more predictions that did better based on your EDA.
# * you've done some serious EDA of feature variables, categorical and numeric.
#
# **Up next:** Time to build some Machine Learning models, based on what you've learnt from your EDA here. Open the notebook `2-titanic_first_ML-model.ipynb`.
#
# For more on `pandas`, check out our [Data Manipulation with Python track](https://www.datacamp.com/tracks/data-manipulation-with-python).
#
# For more on `seaborn`, check out Chapter 3 of our [Intro. to Datavis with Python course](https://www.datacamp.com/courses/introduction-to-data-visualization-with-python).
#
# If you're enoying this session, retweet or share on FB now and follow us on Twitter: [@hugobowne](https://twitter.com/hugobowne) & [@DataCamp](https://twitter.com/datacamp).
| 1-titanic_EDA_first_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# MODIFY!
notebook_name = 'score'
# # Import Libaries & Define Functions
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import glob
sns.set(style='whitegrid')
def frame_it(path):
csv_files = glob.glob(path + '/*.csv')
df_list = []
for filename in csv_files:
df = pd.read_csv(filename, index_col='Unnamed: 0', header=0)
df_list.append(df)
return pd.concat(df_list, axis=1)
def show_values_on_bars(axs, h_v="v", space=0.4,pct=False,neg=False):
def _show_on_single_plot(ax):
if h_v == "v":
for p in ax.patches:
_x = p.get_x() + p.get_width() / 2
_y = p.get_y() + p.get_height()
if pct == True:
value = '{:.2%}'.format(p.get_height())
else:
value = '{:.2f}'.format(p.get_height())
ax.text(_x, _y, value, ha="center")
elif h_v == "h":
for p in ax.patches:
_x = p.get_x() + p.get_width() + float(space)
_y = p.get_y() + p.get_height()
if pct == True:
value = '{:.2%}'.format(p.get_width())
else:
value = '{:.2f}'.format(p.get_width())
if neg == True:
ax.text(_x, _y, value, ha="right")
else:
ax.text(_x, _y, value, ha="left")
if isinstance(axs, np.ndarray):
for idx, ax in np.ndenumerate(axs):
_show_on_single_plot(ax)
else:
_show_on_single_plot(axs)
# # Analysis
# MODIFY!
df = frame_it('./scores')
# we tranpose the data for the analysis
df = df.reindex(['MAE','MSE','RMSE','Score Time','Fit Time'])
# we don't consider the standard deviation of the metrics
df = df.T
# we transpose the data frame due to way we exported the data
df_rmse = df.sort_values('RMSE')
df_rmse
df_rmse.to_csv(f'./analysis/{notebook_name}.csv')
# ## ERR Values [MBit/s] and [(MBit/s)^2]
df_rmse.style.highlight_min(color = 'lightgrey', axis = 0).set_table_styles([{'selector': 'tr:hover','props': [('background-color', '')]}])
# ## RMSE Performance Decline based on Best Performance [%]
df_rmse_min = df_rmse.apply(lambda value : -((value/df.min())-1),axis=1)
df_rmse_min = df_rmse_min.sort_values('RMSE',ascending=False)
df_rmse_min.to_csv(f'./analysis/{notebook_name}-min.csv')
df_rmse_min.style.highlight_max(color = 'lightgrey', axis = 0).set_table_styles([{'selector': 'tr:hover','props': [('background-color', '')]}]).format('{:.2%}')
# ## RMSE Performance Increment based on Worst Performance [%]
df_rmse_max = df.apply(lambda value : abs((value/df.max())-1),axis=1)
df_rmse_max = df_rmse_max.sort_values('RMSE',ascending=False)
df_rmse_max.to_csv(f'./analysis/{notebook_name}-max.csv')
df_rmse_max.style.highlight_max(color = 'lightgrey', axis = 0).set_table_styles([{'selector': 'tr:hover','props': [('background-color', '')]}]).format('{:.2%}')
# the information in this table is not that meaningful / useful
# # Visualization
# +
ax = sns.barplot(data=df_rmse, x='RMSE',y=df_rmse.index, palette='mako')
show_values_on_bars(ax, "h", 0.1)
ax.set(ylabel='Model',xlabel='RMSE [MBit/s]')
ax.tick_params(axis=u'both', which=u'both',length=0)
ax.set_title('Opt Model RMSE (Cross Validated Scores)');
# +
fig,axes = plt.subplots(nrows=3, ncols=1,figsize=(16,20))
# scores = ['RMSE', 'Score Time', 'Fit Time']
for i, column in enumerate(df_rmse.drop(['MAE', 'MSE'],axis=1)):
ax = sns.barplot(data=df_rmse, x=column,y=df_rmse.index, palette='mako',ax=axes[i])
show_values_on_bars(ax, "h", 0)
ax.set(ylabel='Model',xlabel=f'{column}')
# ax.set_title(f'CV {column}')
axes[0].set_title('Cross Validated Model Scores',fontsize=14,fontweight='bold');
# plt.savefig('dl-abs-result.png')
# +
ax = sns.barplot(data=df_rmse_min,x='RMSE',y=df_rmse_min.index,palette='mako')
ax.set(ylabel='Model',xlabel='RMSE Performance Decline [%]')
ax.yaxis.set_label_position("right")
ax.yaxis.set_ticks_position("right")
ax.tick_params(axis=u'both', which=u'both',length=0)
show_values_on_bars(ax,"h",0.001,True,True)
ax.set_title('Opt Model RMSE Perfomance Decline based on Best Performance (Cross Validated Scores)');
# +
fig,axes = plt.subplots(nrows=3, ncols=1,figsize=(16,20))
# scores = ['RMSE', 'Score Time', 'Fit Time']
for i, column in enumerate(df_rmse_min.drop(['MAE', 'MSE'],axis=1)):
ax = sns.barplot(data=df_rmse_min, x=column,y=df_rmse_min.index, palette='mako',ax=axes[i])
ax.set(ylabel='Model',xlabel=f'{column} Performance Decline [%]')
ax.yaxis.set_label_position("right")
ax.yaxis.set_ticks_position("right")
ax.tick_params(axis=u'both', which=u'both',length=0)
show_values_on_bars(ax,"h",0,True,True)
# ax.set_title(f'CV {column}')
axes[0].set_title('Cross Validated Model Perfomance Decline based on Best Performance',fontsize=14,fontweight='bold');
# plt.savefig('dl-result.png')
# +
ax = sns.barplot(data=df_rmse_max,x='RMSE',y=df_rmse_max.index,palette='mako')
show_values_on_bars(ax,"h",0.001,True)
ax.tick_params(axis=u'both', which=u'both',length=0)
ax.set(ylabel='Model',xlabel='RMSE Performance Increment [%]')
ax.set_title('Opt Model RMSE Perfomance Increment based on Worst Performance (Cross Validated Scores)');
# +
fig,axes = plt.subplots(nrows=3, ncols=1,figsize=(16,20))
# scores = ['RMSE', 'Score Time', 'Fit Time']
for i, column in enumerate(df_rmse_max.drop(['MAE', 'MSE'],axis=1)):
ax = sns.barplot(data=df_rmse_max, x=column,y=df_rmse_max.index, palette='mako',ax=axes[i])
ax.set(ylabel='Model',xlabel=f'{column} Performance Increment [%]')
ax.tick_params(axis=u'both', which=u'both',length=0)
show_values_on_bars(ax,"h",0,True)
# ax.set_title(f'CV {column}')
axes[0].set_title('Cross Validated Model Performance Increment based on Worst Performance',fontsize=14,fontweight='bold');
# -
# DONE!
| 1-dl-project/dl-9-score-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 模块使用
# # 1 scipy.io
# ## 读取矩阵数据
import numpy as np
from scipy import io as spio
a = np.ones((3,3))
spio.savemat('file.mat',{'a':a})
data = spio.loadmat('file.mat',struct_as_record=True)
data['a']
# ## 读取图像
from scipy import misc
misc.imread('fname.png')
import matplotlib.pyplot as plt
plt.imread('fname.png')
# + **文本文件**
# numpy.loadtxt() / numpy.savetxt()
# + **txt/csv文件**
# numpy.genfromtxt()/numpy.recfromcsv()
# + **二进制文件**
# numpy.load() / numpy.save()
# # 2 scipy.linalg
# ## 计算行列式
from scipy import linalg
arr= np.array([[1,2],
[3,4]])
linalg.det(arr)
arr = np.array([[3,2],
[6,4]])
linalg.det(arr)
linalg.det(np.ones(3,4))
# ## 计算逆矩阵
arr = np.array([[1,2],[3,4]])
iarr = linalg.inv(arr)
iarr
# 验证
np.allclose(np.dot(arr,iarr),np.eye(2))
# 奇异矩阵求逆抛出异常
arr = np.array([[3,2],[6,4]])
linalg.inv(arr)
# ## 奇异值分解
arr = np.arange(9).reshape((3,3)) + np.diag([1,0,1])
uarr,spec,vharr = linalg.svd(arr)
spec
sarr = np.diag(spec)
svd_mat = uarr.dot(sarr).dot(vharr)
np.allclose(svd_mat,arr)
# SVD常用于统计和信号处理领域。其他的一些标准分解方法(QR, LU, Cholesky, Schur) 在 scipy.linalg 中也能够找到。
# # 3 优化
from scipy import optimize
def f(x):
return x**2 + 10*np.sin(x)
x = np.arange(-10,10,0.1)
plt.plot(x,f(x))
plt.show()
# 此函数有一个全局最小值,约为-1.3,含有一个局部最小值,约为3.8.
# 在寻找最小值的过程中,确定初始值,用梯度下降的方法,bfgs是一个很好的方法。
optimize.fmin_bfgs(f,0)
# 但是方法的缺陷是陷入局部最优解
optimize.fmin_bfgs(f,5)
# 可以在一个区间中找到一个最小值
xmin_local = optimize.fminbound(f,0,10)
xmin_local
# 寻找函数的零点
# guess 1
root = optimize.fsolve(f,1)
root
# guess -2.5
root = optimize.fsolve(f,-2.5)
root
# ## 曲线拟合
# 从函数f中采样得到一些含有噪声的数据
xdata = np.linspace(-10,10,num=20)
ydata = f(xdata)+np.random.randn(xdata.size)
# 我们已经知道函数的形式$x^2+\sin(x)$,但是每一项的系数不清楚,因此进行拟合处理
def f2(x,a,b):
return a*x**2 + b*np.sin(x)
guess=[3,2]
params,params_covariance = optimize.curve_fit(f2, xdata, ydata, guess)
params
# ## 绘制结果
x = np.arange(-10,10,0.1)
def f(x):
return x**2 + 10 * np.sin(x)
grid = (-10,10,0.1)
xmin_global = optimize.brute(f,(grid,))
xmin_local = optimize.fminbound(f,0,10)
root = optimize.fsolve(f,1)
root2 = optimize.fsolve(f,-2.5)
xdata = np.linspace(-10,10,num=20)
np.random.seed(1234)
ydata = f(xdata)+np.random.randn(xdata.size)
def f2(x,a,b):
return a*x**2 + b * np.sin(x)
guess=[2,2]
params,_ =optimize.curve_fit(f2,xdata,ydata,guess)
fig = plt.figure()
ax = fig.add_subplot(111)
ax.plot(x,f(x),'b-',label='f(x)')
ax.plot(x,f2(x,*params),'r--',label='Curve fit result')
xmins = np.array([xmin_global[0],xmin_local])
ax.plot(xmins,f(xmins),'go',label='minize')
roots = np.array([root,root2])
ax.plot(roots,f(roots),'kv',label='Roots')
ax.legend()
ax.set_xlabel('x')
ax.set_ylabel('f(x)')
plt.show()
# # 4 统计
# ## 直方图和概率密度统计
a = np.random.normal(size=1000)
bins = np.arange(-4,5)
bins
histogram = np.histogram(a,bins=bins,normed=True)[0]
bins = 0.5*(bins[1:]+bins[:-1])
bins
from scipy import stats
b =stats.norm.pdf(bins)
plt.plot(bins,histogram)
plt.plot(bins,b)
plt.show()
# ## 百分位数
# 百分位是累计概率分布函数的一个估计
np.median(a)
stats.scoreatpercentile(a,50)
stats.scoreatpercentile(a,90)
# ## 统计检验
# 统计检验的结果常用作一个决策指标。例如,如果我们有两组观察点,它们都来自高斯过程,我们可以使用 T-检验 来判断两组观察点是都显著不同:
a = np.random.normal(0,1,size=100)
b = np.random.normal(0,1,size=10)
stats.ttest_ind(a,b)
# 返回结果分成连个部分
# + T检验统计量
# 使用检验的统计量的值
# + P值
# 如果结果接近1,表明符合预期,接近0表明不符合预期。
# # 5 插值
measured_time = np.linspace(0,1,10)
noise = (np.random.random(10)*2-1) *1e-1
measure = np.sin(2*np.pi*measured_time) + noise
from scipy.interpolate import interp1d
linear_interp = interp1d(measured_time, measure)
computed_time = np.linspace(0, 1, 50)
linear_results = linear_interp(computed_time)
cublic_interp = interp1d(measured_time, measure, kind='cubic')
cublic_results = cublic_interp(computed_time)
plt.plot(measured_time,measure,'o',label='points')
plt.plot(computed_time,linear_results,'r-',label='linear interp')
plt.plot(computed_time,cublic_results,'y-',label='cublic interp')
plt.legend()
3 plt.show()
# # 练习
# ## 温度曲线拟合
# 阿拉斯加每个月温度的最大值和最小值数据见下表:
#
# 最小值 | 最大值 | 最小值 | 最大值
# --- | --- | --- | ---
# -62 | 17 | -9 | 37
# -59 | 19 | -13 | 37
# -56 | 21 | -25 | 31
# -46 | 28 | -46 | 23
# -32 | 33 | -52 | 19
# -18 | 38 | -48 | 18
#
# **要求**
# + 绘制温度图像
# + 拟合出一条函数曲线
# + 使用scipy.optimize.curvie_fit()来拟合函数
# + 画出函数图像。
# + 判断最大值和最小值的偏置是否合理
import numpy as np
import matplotlib.pyplot as plt
months = np.arange(1,13)
mins = [-62,-59,-56,-46,-32,-18,-9,-13,-25,-46,-52,-48]
maxes = [17,19,21,28,33,38,37,37,31,23,19,18]
fig,ax = plt.subplots()
plt.plot(months,mins,'b-',label='min')
plt.plot(months,maxes,'r-',label='max')
plt.ylim(-80,80)
plt.xlim(0.5,12.5)
plt.xlabel('month')
plt.ylabel('temperature')
plt.xticks([1,2,3,4,5,6,7,8,9,10,11,12],
['Jan.','Feb.','Mar.','Apr.','May.','Jun.','Jul.','Aug.','Sep,','Oct.','Nov.','Dec.'])
plt.legend()
plt.title('Alaska temperature')
plt.show()
# 从图像上来看,温度的最高值和最低值都符合二次函数的特点,$y = at^2+bt+c$,其中$c$为时间$t$的偏置。
from scipy import optimize
def f(t,a,b,c):
return a * t**2+b*t+c
guess = [-1,8,50]
params_min,_ = optimize.curve_fit(f,months,mins,guess)
params_max,_ = optimize.curve_fit(f,months,maxes,guess)
times = np.linspace(1,12,30)
plt.plot(times,f(times,*params_min),'b--',label='min_fit')
plt.plot(times,f(times,*params_max),'r--',label='max_fit')
plt.plot(months,mins,'bo',label='min')
plt.plot(months,maxes,'ro',label='max')
plt.ylim(-80,80)
plt.xlim(0.5,12.5)
plt.xlabel('month')
plt.ylabel('temperature')
plt.xticks([1,2,3,4,5,6,7,8,9,10,11,12],
['Jan.','Feb.','Mar.','Apr.','May.','Jun.','Jul.','Aug.','Sep,','Oct.','Nov.','Dec.'])
plt.title('Alaska temperature')
plt.show()
# 温度最高值拟合效果较好,但温度最低值拟合效果不太好
# ## 求解最小值
# 驼峰函数 $$f(x,y)=(4-2.1x^2+\frac{x^4}{3})x^2+xy+(4y^2-4)y^2$$
# + 限制变量范围: $-2<x<2,-1<y<1$
# + 使用 numpy.meshgrid() 和 pylab.imshow() 目测最小值所在区域
# + 使用 scipy.optimize.fmin_bfgs() 或者其他的用于可以求解多维函数最小值的算法
# +
import numpy as np
from scipy import optimize
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
def sixhump(x):
return (4 - 2.1*x[0]**2 + x[0]**4 / 3.) * x[0]**2 + x[0] * x[1] + (-4 + 4*x[1]**2) * x[1] **2
x = np.linspace(-2, 2)
y = np.linspace(-1, 1)
xg, yg = np.meshgrid(x, y)
#plt.figure() # simple visualization for use in tutorial
#plt.imshow(sixhump([xg, yg]))
#plt.colorbar()
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
surf = ax.plot_surface(xg, yg, sixhump([xg, yg]), rstride=1, cstride=1,
cmap=plt.cm.jet, linewidth=0, antialiased=False)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('f(x, y)')
ax.set_title('Six-hump Camelback function')
plt.show()
# -
min1 = optimize.fmin_bfgs(sixhump,[0,-0.5])
min2 = optimize.fmin_bfgs(sixhump,[0,0.5])
min3 = optimize.fmin_bfgs(sixhump,[-1.4,1.0])
local1 = sixhump(min1)
local2 = sixhump(min2)
local3 = sixhump(min3)
print local1,local2,local3
| python-statatics-tutorial/basic-theme/scipy_basic/details.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas
from datetime import datetime as dt
from pytz import utc
#Use pandas to read data from csv file into dataframe
#parse_dates tells df to read particular column as a datetime object column
df = pandas.read_csv("reviews.csv", parse_dates=["Timestamp"])
df
# -
# ## Rating average and count by day
# +
#Create a new column for the date which is taken from the timestamp datetime object
#This is done so we can then group data by the day value
df["Day"] = df["Timestamp"].dt.date
#We then group the data by the "Day" column and find the mean rating of each group
#day_avg = df.groupby(["Day"])["Rating"].mean()
day_avg = df.groupby(["Day"]).mean() #.mean() only works with columns that have numbers thus only rating appears
day_avg.head()
# +
import matplotlib.pyplot as plt
#Gives the plotted graph a size -> 25 length and 7 height
plt.figure(figsize=(25,7))
#plot function takes a list of x and y coordinates
#x coordinates are the day values -> index of the dataframe
#y coordinates are the ratings for each day value
plt.plot(day_avg.index, day_avg["Rating"])
#It reads data and finds min and max values and plots only between them
#day_avg["Rating"].min()
#day_avg["Rating"].max()
# +
#We also group the data by the "Day" column and count how many of each data occured each day
day_avg = df.groupby(["Day"]).count() #.count() gives a count of all columns that can be counted
#Gives the plotted graph a size -> 25 length and 7 height
plt.figure(figsize=(25,7))
#If we plot the count method dataframe we will get a graph of number of ratings per day
#plot function takes a list of x and y coordinates
plt.plot(day_avg.index, day_avg["Rating"])
# -
# ## Rating average by week
# +
#Create a new column for the week which is taken from the timestamp datetime object
#This is done so we can then group data by the week value
#We split the timestamp using strftime to get year and week (Otherwise same weeks in different years count as one)
# #%Y gets time and %U gets the week value
df["Week"] = df["Timestamp"].dt.strftime("%Y-%U")
#Group the data by the week column and find its mean
week_avg = df.groupby(["Week"]).mean()
#Thus we get a dataframe with the week column as the index and the mean ratings per week
week_avg
# +
#Gives the plotted graph a size -> 27 length and 7 height
plt.figure(figsize=(25,7))
#plot function takes a list of x and y coordinates
#We plot the weeks w.r.t the mean ratings per week
plt.plot(week_avg.index, week_avg["Rating"])
# -
# ## Rating average by month
# +
#Create a new column for the month which is taken from the timestamp datetime object
#This is done so we can then group data by the month value
#We split the timestamp using strftime to get year and month (Otherwise same months in different years count as one)
# #%Y gets time and %m gets the month value
df["Month"] = df["Timestamp"].dt.strftime("%Y-%m")
#Group the data by the Month column and find its mean
mon_avg = df.groupby(["Month"]).mean()
#Thus we get a dataframe with the month column as the index and the mean ratings per month
mon_avg
# +
#Gives the plotted graph a size -> 25 length and 7 height
plt.figure(figsize=(25,7))
#plot function takes a list of x and y coordinates
#We plot the months w.r.t the mean ratings per month
plt.plot(mon_avg.index, mon_avg["Rating"])
# -
# ## Average rating and count per month by course
# +
#Create a new column for the month which is taken from the timestamp datetime object
#This is done so we can then group data by the month value
#We split the timestamp using strftime to get year and month (Otherwise same months in different years count as one)
# #%Y gets time and %m gets the month value
df["Month"] = df["Timestamp"].dt.strftime("%Y-%m")
#Group the data by the Month column and the Course Name column and find its mean
#We then unstack the data to make it have better structure because we have two index columns
mon_avg_course = df.groupby(["Month", "Course Name"]).mean().unstack()
#Thus we get a dataframe with the month column and the course name column as the index
#The mean ratings are per month per course
mon_avg_course
# -
#Shows that we have more than 1 Index -> Indexes are Month column and the course name column
#And mon_avg_course.columns returns only "Rating"
mon_avg_course.index
#plot function directly on dataframe
#We plot the months w.r.t the mean ratings per month for each course and give the graph a particular size
mon_avg_course.plot(figsize=(25,8))
# +
#Create a new column for the month which is taken from the timestamp datetime object
#This is done so we can then group data by the month value
#We split the timestamp using strftime to get year and month (Otherwise same months in different years count as one)
# #%Y gets time and %m gets the month value
df["Month"] = df["Timestamp"].dt.strftime("%Y-%m")
#Group the data by the Month column and the Course Name column and count its data for all ratings
#We then unstack the data to make it have better structure because we have two index columns
mon_avg_course = df.groupby(["Month", "Course Name"])["Rating"].count().unstack()
#Thus we get a dataframe with the month column and the course name column as the index
#The count of ratings are per month per course
mon_avg_course
# -
#plot function directly on dataframe
#We plot the months w.r.t the count of ratings per month for each course and give the graph a particular size
mon_avg_course.plot(figsize=(25,8))
# ## What day of the week are people most positive
# +
#Create a new column for the days of the Week which are taken from the timestamp datetime object
#This is done so we can then group data by the weekday value
#We split the timestamp using strftime to get the Weekday using %A
df["Weekday"] = df["Timestamp"].dt.strftime("%A")
#Create a new column for the day numbers for each day which are taken from the timestamp datetime object
#This is done so we can then group data by the day number values
#We split the timestamp using strftime to get the day number using %w
df["DayNumber"] = df["Timestamp"].dt.strftime("%w")
#We then group all the data by the weekday and daynumber columns and find the mean
wkdy_avg = df.groupby(["Weekday", "DayNumber"]).mean()
#We also sort the data by DayNumber
wkdy_avg = wkdy_avg.sort_values("DayNumber")
#We get a dataframe with the weekdays and daynumbers as indexes and the mean ratings per day
wkdy_avg
# +
#Gives the plotted graph a size -> 15 length and 3 height
plt.figure(figsize=(15,3))
#plot function takes a list of x and y coordinates
#We plot the Weekdays w.r.t the mean ratings per weekday
plt.plot(wkdy_avg.index.get_level_values(0), wkdy_avg["Rating"])
# -
| Practice/Data Analysis & Visualisation/Plotting Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2値分類の例題
# 映画のレビュー文章を肯定的なレビューと否定的なレビューに分類する
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras import layers, models, initializers
from tensorflow.keras.datasets import imdb
np.random.seed(20201231)
tf.random.set_seed(20201231)
print("tensorflow version : " + tf.__version__)
# -
# # Memory setting if you use GPU
# +
#config = tf.compat.v1.ConfigProto()
#config.gpu_options.per_process_gpu_memory_fraction = 0.8
# -
# # Load dataset
# +
#データセットのロード(単語数は10000を上限にする)
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=10000)
#評価データの作成
x_val = x_train[:10000]
x_train = x_train[10000:]
y_val = y_train[:10000]
y_train = y_train[10000:]
print('x_train.shape: ', x_train.shape)
print('y_train.shape: ', y_train.shape)
print('x_val.shape: ', x_val.shape)
print('y_val.shape: ', y_val.shape)
print('x_test.shape: ', x_test.shape)
print('y_test.shape: ', y_test.shape)
print('\nx_train')
print(x_train)
print('\ny_train')
print(y_train)
print(y_train.shape)
# -
#データセット0番目の内容を文章に戻してみる
print('\nデータセット0番目の内容を文章に戻してみる')
word_index = imdb.get_word_index()
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
#0はパディング、1はシーケンスの開始、2が不明として予約されているため除外する
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in x_train[0]])
print(decoded_review)
# # Modify input data
# +
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1.
return results
#ベクトル化
x_train = vectorize_sequences(x_train)
x_val = vectorize_sequences(x_val)
x_test = vectorize_sequences(x_test)
print('\nx_train')
print(x_train)
#既にOneHot化しているため方をfloatに変換する
y_train = np.asarray(y_train).astype('float32')
y_val = np.asarray(y_val).astype('float32')
y_test = np.asarray(y_test).astype('float32')
print('\ny_train')
print(y_train)
# -
# # Create deep learning layer
# +
model = models.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(16, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.summary()
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
# -
# # Trainning
# +
training = model.fit(x_train, y_train,
validation_data=(x_val, y_val),
batch_size=512,
epochs=5)
plt.plot(training.history['accuracy'])
plt.plot(training.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
#loss
plt.plot(training.history['loss'])
plt.plot(training.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
# # Evaluate
results = model.evaluate(x_test, y_test)
# # Save the models
# +
model.save('models\\bin_class.h5')
model.save('models\\bin_class_pb', save_format="tf")
print(model.input)
print(model.output)
| binary_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Training a Neural Network using Augmentor and Keras
#
# In this notebook, we will train a simple convolutional neural network on the MNIST dataset using Augmentor to augment images on the fly using a generator.
#
# ## Import Required Libraries
#
# We start by making a number of imports:
# +
import Augmentor
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
import numpy as np
# -
# ## Define a Convolutional Neural Network
#
# Once the libraries have been imported, we define a small convolutional neural network. See the Keras documentation for details of this network: <https://github.com/fchollet/keras/blob/master/examples/mnist_cnn.py>
#
# It is a three layer deep neural network, consisting of 2 convolutional layers and a fully connected layer:
# +
num_classes = 10
input_shape = (28, 28, 1)
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
# -
# Once a network has been defined, you can compile it so that the model is ready to be trained with data:
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
# You can view a summary of the network using the `summary()` function:
model.summary()
# ## Use Augmentor to Scan Directory for Data
#
# Now we will use Augmentor to scan a directory containing our data that we will eventually feed into the neural network in order to train it.
#
# When you point a pipeline to a directory, it will scan each subdirectory and treat each subdirectory as a class for your machine learning problem.
#
# For example, within the directory `mnist`, there are subdirectories for each digit:
#
# ```
# mnist/
# ├── 0/
# │ ├── 0001.png
# │ ├── 0002.png
# │ ├── ...
# │ └── 5985.png
# ├── 1/
# │ ├── 0001.png
# │ ├── 0002.png
# │ ├── ...
# │ └── 6101.png
# ├── 2/
# │ ├── 0000.png
# │ ├── 0001.png
# │ ├── ...
# │ └── 5801.png
# │ ...
# ├── 9/
# │ ├── 0001.png
# │ ├── 0002.png
# │ ├── ...
# │ └── 6001.png
# └
# ```
#
# The directory `0` contains all the images corresponding to the 0 class.
#
# To do this, we instantiate a pipeline object in the `mnist` parent directory:
p = Augmentor.Pipeline("/home/marcus/Documents/mnist/train/")
# ## Add Operations to the Pipeline
#
# Now that a pipeline object `p` has been created, we can add operations to the pipeline. Below we add several simple operations:
p.flip_top_bottom(probability=0.1)
p.rotate(probability=0.3, max_left_rotation=5, max_right_rotation=5)
# You can view the status of pipeline using the `status()` function, which shows information regarding the number of classes in the pipeline, the number of images, and what operations have been added to the pipeline:
p.status()
# ## Creating a Generator
#
# A generator will create images indefinitely, and we can use this generator as input into the model created above. The generator is created with a user-defined batch size, which we define here in a variable named `batch_size`. This is used later to define number of steps per epoch, so it is best to keep it stored as a variable.
batch_size = 128
g = p.keras_generator(batch_size=batch_size)
# The generator can now be used to created augmented data. In Python, generators are invoked using the `next()` function - the Augmentor generators will return images indefinitely, and so `next()` can be called as often as required.
#
# You can view the output of generator manually:
images, labels = next(g)
# Images, and their labels, are returned in batches of the size defined above by `batch_size`. The `image_batch` variable is a tuple, containing the augmentented images and their corresponding labels.
#
# To see the label of the first image returned by the generator you can use the array's index:
print(labels[0])
# ## Train the Network
#
# We train the network by passing the generator, `g`, to the model's fit function. In Keras, if a generator is used we used the `fit_generator()` function as opposed to the standard `fit()` function. Also, the steps per epoch should roughly equal the total number of images in your dataset divided by the `batch_size`.
#
# Training the network over 5 epochs, we get the following output:
h = model.fit_generator(g, steps_per_epoch=len(p.augmentor_images)/batch_size, epochs=5, verbose=1)
# ## Summary
#
# Using Augmentor with Keras means only that you need to create a generator when you are finished creating your pipeline. This has the advantage that no images need to be saved to disk and are augmented on the fly.
| notebooks/Augmentor_Keras.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Q# notebooks #
#
# This notebook will show you how to use IQ# to write your own quantum application notebooks.
#
# ## Compiling Q# operations
# An operation is the basic unit of quantum execution in Q#. It is roughly equivalent to a function in C or C++ or Python, or a static method in C# or Java.
#
# IQ#, the Q# Jupyter kernel, allows you to write Q# operations directly on a code cell and compile them by running the cell (pressing Ctrl/⌘+Enter). For example:
# +
operation SayHello(name: String) : Unit {
// The following line will simply write a message to the console:
Message($"Hello {name}!");
}
# -
# When you **Run** the cell, Q# compiles the code and returns the name of the operations it found.
# In this case, it found only one operation (`HelloQ`).
#
# If the compiler detects any errors, it will instead show the list of errors in the output. For example:
operation InvalidQ() : Unit {
// The `FooBar` operation doesn't exist, so the following line
// will generate a `No variable with that name exists.` error:
FooBar("Hello again!");
// `Message` takes only one string argument, so the following line
// will generate a `Unexpected argument tuple.` error:
Message(1, 2);
}
# Q# operations can call other operations previously defined; they can also call all the operations defined in the
# [Microsoft.Quantum.Intrinsic](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic) and
# [Microsoft.Quantum.Canon](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.canon) namespaces (like [Message](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.message)).
#
# For example, you can create a new operation that calls the previously compiled `HelloQ`:
#
operation HelloAndres() : Unit {
SayHello("Andres");
}
# They can also use all [Q# standard library](https://docs.microsoft.com/qsharp/api/) operations defined in other namespaces by importing the namespace using the `open` statement. For example, to use [`PI`](https://docs.microsoft.com/qsharp/api/prelude/microsoft.quantum.math.pi) you would need to open the ` Microsoft.Quantum.Math` namespace; optionally you can call the operation providing its fully qualified name, for example:
#
# +
open Microsoft.Quantum.Math;
operation HelloPi() : Unit {
let pi = Microsoft.Quantum.Convert.DoubleAsString(PI());
SayHello(pi);
}
# -
# You can define multiple operations in a single cell and use any valid Q# code, for example:
# +
open Microsoft.Quantum.Math;
/// # Summary
/// Sets the qubit's state to |+⟩
operation SetPlus(q: Qubit) : Unit {
Reset(q);
H(q);
}
/// # Summary
/// Sets the qubit's state to |-⟩
operation SetMinus(q: Qubit) : Unit {
Reset(q);
X(q);
H(q);
}
/// # Summary
/// Randomly prepares the qubit into |+⟩ or |-⟩
operation PrepareRandomMessage(q: Qubit) : Unit {
let choice = RandomInt(2);
if (choice == 0) {
Message("Prepared |-⟩");
SetMinus(q);
} else {
Message("Prepared |+⟩");
SetPlus(q);
}
}
# -
# and create other operations that uses them:
# +
open Microsoft.Quantum.Diagnostics;
open Microsoft.Quantum.Measurement;
operation NextRandomBit() : Result {
using (q = Qubit()) {
SetPlus(q);
return MResetZ(q);
}
}
operation TestPrepareQubits() : Result {
mutable r = Zero;
using (qubits = Qubit[5]) {
ApplyToEach(PrepareRandomMessage, qubits);
DumpMachine();
set r = Measure([PauliX, PauliX, PauliX, PauliX, PauliX], qubits);
ResetAll(qubits);
}
return r;
}
# -
# ## Simulating Q# operations
#
#
# Once a Q# operation has been successfully compiled, you can use the `%simulate` command to simulate it. For example:
%simulate HelloPi
%simulate NextRandomBit
# `%simulate` will print any console output on the notebook, and it will return the operation's return value. If the operation returns `Unit` it prints `()`, otherwise it prints the actual value.
#
# `%simulate` only accepts operations that take no arguments. If you want to call an operation that accepts parameters, like `HelloQ`, create a wrapper operation that calls it with the corresponding values, like `HelloPi`.
#
# As mentioned, all messages are printed on the notebook; this includes calls to [`DumpMachine`](https://docs.microsoft.com/en-us/qsharp/api/prelude/microsoft.quantum.extensions.diagnostics.dumpmachine):
%config dump.basisStateLabelingConvention = "bitstring"
%simulate TestPrepareQubits
# ## Estimating resources
#
# The `%estimate` command lets you estimate the resources a given quantum operation will need to execute, without actually executing the operation. Similar to `%simulate` it takes the name of a no-arguments operation. However, `%estimate` does not keep track of the qubit's state and will not return the output of the operation, instead it returns the estimated values of how many resources, like Qubits and CNOT gates, the corresponding operation will use:
%estimate TestPrepareQubits
# To learn more about resources estimation, take a look at [The ResourcesEstimator Target Machine](https://docs.microsoft.com/en-us/quantum/machines/resources-estimator?) documentation.
# ## The Workspace
#
# The notebook uses the folder it lives on disk to define a workspace. It will try to compile all the Q# files (i.e. all files with a `.qs` extension) it finds under the current folder and will make the operations it finds available to operations in the notebook. For example, the [Operations.qs](/Operations.qs) file in this folder defines two operations:
# * Microsoft.Quantum.Samples.IsMinus
# * Microsoft.Quantum.Samples.IsPlus
#
# To get the list of operations defined in the workspace, you can use the `%workspace` command:
%workspace
# These operations can be used in this notebook, for example:
# +
open Microsoft.Quantum.Samples;
operation CheckPlus() : Bool {
mutable result = false;
using (q = Qubit()) {
SetPlus(q);
set result = IsPlus(q);
Reset(q);
}
return result;
}
# -
%simulate CheckPlus
# To pick up any changes you make to a Q# file in the workspace, use `%workspace reload`.
%workspace reload
# ## Getting Help ##
#
# Q# supports adding documentation to operations via comments in the code. When such documentation exists, you can access it from the notebook by adding a question mark before or after the operation name on a code cell, for example:
Microsoft.Quantum.Intrinsic.X?
# This documentation is available for any operations in the Prelude, Canon and Workspace, or even those defined locally in the notebook:
PrepareRandomMessage?
# ## Other commands ##
# ### `%who`
#
# `%who` returns the list of all local and workspace operations available.
%who
# ### `%package`
#
# `%package` allows you to load nuget packages and makes available any Q# operations defined on them. For example, to use the operations from [Q#'s Quantum Chemistry Library](https://docs.microsoft.com/en-us/quantum/libraries/chemistry/?view=qsharp-preview), you must load the [Microsoft.Quantum.Chemistry](https://www.nuget.org/packages/Microsoft.Quantum.Chemistry/) nuget package:
%package Microsoft.Quantum.Chemistry
# `%package` returns the list of nuget packages currently loaded and their version.
# ### `%version`
#
# `%version` simply returns the current versions of IQ# and of Jupyter Core (a library used by IQ#):
%version
| samples/getting-started/intro-to-iqsharp/Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_boston
boston = load_boston()
boston
boston.feature_names
X = boston.data
Y = boston.target
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=0.3,
random_state=87)
regr = LinearRegression()
regr.fit(x_train, y_train)
y_predict = regr.predict(x_test)
plt.scatter(y_test, y_predict)
plt.plot([0,50],[0,50],'r')
boston.feature_names
x = np.linspace(-10,10,200)
plt.subplot(2,2,1)
plt.plot(x,np.sin(x))
plt.subplot(2,2,2)
plt.plot(x,np.cos(x))
plt.subplot(2,2,3)
plt.plot(x,x)
plt.subplot(2,2,4)
plt.plot(x,x**2)
X = boston.data
X
plt.figure(figsize=(8,10))
for i, feature in enumerate(boston.feature_names):
plt.subplot(5, 3, i+1)
plt.scatter(X[:,i], Y, s=1)
plt.ylabel("price")
plt.xlabel(feature)
plt.tight_layout()
| Python/boston_price.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/ryanleeallred/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module2-sampling-confidence-intervals-and-hypothesis-testing/LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="838Dmw1kM2LK"
# # Lambda School Data Science Module 132
# ## Sampling, Confidence Intervals, and Hypothesis Testing
# -
# ---
# Imports
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from scipy import stats
# + [markdown] colab_type="text" id="dbcPKIo5M6Ny"
# ---
#
# ## Prepare - examine other available hypothesis tests
# -
# If you had to pick a single hypothesis test in your toolbox, t-test would probably be the best choice - but the good news is you don't have to pick just one!
#
# Here are some of the others to be aware of:
#
# - $\chi^2$ (chi-squared) test
# - Distribution tests (e.g. normal test)
# - Kruskal-Wallis H-Test
# ---
#
# ### One-way chi square test
# [Wikipedia Article](https://en.wikipedia.org/wiki/Chi-squared_test)
#
# - Chi square can take any crosstab/table and test the independence of rows/cols
# - The null hypothesis is that the rows/cols are independent -> low chi square
# - The alternative is that there is a dependence -> high chi square
# - Be aware! Chi square does *not* tell you direction/causation
# + colab={"base_uri": "https://localhost:8080/", "height": 215} colab_type="code" id="tlBel8j9M6tB" outputId="811623c9-885a-42e3-c3f7-159ced2ce330"
# One-way chi squared test using Python
ind_obs = np.array([[1, 1], [2, 2]]).T
print(ind_obs)
print(stats.chisquare(ind_obs, axis=None))
dep_obs = np.array([[16, 18, 16, 14, 12, 12], [32, 24, 16, 28, 20, 24]]).T
print(dep_obs)
print(stats.chisquare(dep_obs, axis=None))
# -
# ---
#
# ### Distribution Tests
# We often assume that something is normal, but it can be important to *check*.
#
# For example, later on with predictive modeling, a typical assumption is that residuals (prediction errors) are normal - checking is a good diagnostic
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="nN0BdNiDPxbk" outputId="36426de9-d1b9-4790-ae20-9d5eb578a77a"
# The Poisson distribution models arrival times and is related to the binomial (coinflip)
sample = np.random.poisson(5, 1000)
print(stats.normaltest(sample)) # Pretty clearly not normal
# -
# ---
#
# ### Kruskal-Wallis H-Test
# - Compare the median rank between 2+ groups
# - Can be applied to ranking decisions/outcomes/recommendations
# - The underlying math comes from chi-square distribution, and is best for n>5
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="P5t0WhkDReFO" outputId="7d6438bf-8042-4297-a8f7-cef083d22444"
x1 = [1, 3, 5, 7, 9]
y1 = [2, 4, 6, 8, 10]
print(stats.kruskal(x1, y1)) # x1 is a little better, but not "significantly" so
x2 = [1, 1, 1]
y2 = [2, 2, 2]
z = [2, 2] # Hey, a third group, and of different size!
print(stats.kruskal(x2, y2, z)) # x clearly dominates
# + [markdown] colab_type="text" id="7pT3IP36Rh0b"
# And there's many more! `scipy.stats` is fairly comprehensive, though there are even more available if you delve into the extended world of statistics packages. As tests get increasingly obscure and specialized, the importance of knowing them by heart becomes small - but being able to look them up and figure them out when they *are* relevant is still important.
# -
# ---
# ### Degrees of Freedom
# [What Are Degrees of Freedom in Statistics?](https://blog.minitab.com/blog/statistics-and-quality-data-analysis/what-are-degrees-of-freedom-in-statistics)
#
# - Technically defined as:
#
# > The dimension of the domain of a vector.
#
# - Less technically / more broadly defined as:
#
# > The number of "observations" in the data that are free to vary when estimating statistical parameters.
#
# - Or...
#
# > The freedom to vary.
#
# - 7 hats; one for each day of the week. As the week progresses, one's freedom of choice between the remaining hats decreases.
# - On the last day, one doesn't have a choice—no freedom to vary.
mean = 20
n = 7
s = [5, 9, 10, 20, 15, 12, 69]
# - The first 6 days add up to 71
# - The mean has to be 20
# - I need the sum of all the values in the list to be 140
#
# $7 \cdot 20 = 140$
#
# - The last value in the list ***has*** to be:
#
# $140 - 71 = 69$
# ---
# + [markdown] colab_type="text" id="3JqroCQYQqhy"
# ## T-test Assumptions
#
# <https://statistics.laerd.com/statistical-guides/independent-t-test-statistical-guide.php>
#
# 1. Independence of means
# 2. "Dependent Variable" (sample means) are Distributed Normally
# 3. "Homogeneity" of Variance?
# -
# ##### ---- Ø ----
# #### 1. Independence of Means
#
# *aka: Unrelated groups*
#
# > Are the means of our voting data independent (do not affect the outcome of one another)?
#
# The best way to increase thel likelihood of our means being independent is to randomly sample (which we did not do).
# *Note about variance calculation:
#
# > *By default, pandas uses the sample variance of $n-1$,
# > while numpy uses the population variance by default.
# + colab={} colab_type="code" id="sqy2hEFRZnvI"
# Take a look at the 2-sample t-test method documentation
# ?stats.ttest_ind
# + [markdown] colab_type="text" id="xI-PcK5sZ1A9"
# #### 2. "Homogeneity" of Variance?
#
# > Is the magnitude of the variance between the two roughly the same?
#
# I think we're OK on this one for the voting data, although it probably could be better, one party was larger than the other.
#
# If we suspect this to be a problem then we can use Welch's T-test.
# + colab={} colab_type="code" id="P02dL0waauN5"
# ?ttest_ind
# + [markdown] colab_type="text" id="tjgoHHwGayoC"
# #### 3. "Dependent Variable" (sample means) are Distributed Normally
#
# <https://stats.stackexchange.com/questions/9573/t-test-for-non-normal-when-n50>
#
# Lots of statistical tests depend on normal distributions. We can test for normality using Scipy as was shown above.
#
# This assumption is often assumed even if the assumption is a weak one. If you strongly suspect that things are not normally distributed, you can transform your data to get it looking more normal and then run your test.
#
# > This problem is reduced with larger sample sizes (yay Central Limit Theorem) and is often why you don't hear it brought up.
#
# People declare the assumption to be satisfied either way.
# -
# ---
# + [markdown] colab_type="text" id="bvvPV-RJN2vA"
# ## Central Limit Theorem
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 72} colab_type="code" id="FBLoOF8qOJeJ" outputId="0a4e7e48-ef94-497b-b119-cd27a6660167"
# Create a ton of simulated observations in a similar way to a Monte Carlo
sample_means = []
for _ in range(0,3000):
coinflips = np.random.binomial(n=1, p=.5, size=12)
one_sample = coinflips
sample_means.append(coinflips.mean())
print(len(sample_means))
print(sample_means[:50])
# + colab={"base_uri": "https://localhost:8080/", "height": 198} colab_type="code" id="rfeA06evOT2K" outputId="62a06b54-3ecd-496c-bac5-b1874f405cd5"
df = pd.DataFrame({'single_sample': one_sample})
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="GlMSNFX6OmBV" outputId="9122b3d0-a59f-496d-9c7b-bc2eacc489f8"
df.single_sample.hist();
# + colab={"base_uri": "https://localhost:8080/", "height": 296} colab_type="code" id="Jie4ypgLOs5M" outputId="13e41ba1-8118-4a7b-99e2-9966565e65ee"
ax = plt.hist(sample_means, bins=13)
plt.title('Distribution of 3000 sample means \n (of 8 coinflips each)');
# + [markdown] colab_type="text" id="LsEAjc4rOylm"
# What does the Central Limit Theorem State?
#
# > That no matter the initial distribution of the population, the distribution of sample means taken will approximate a normal distribution as $n \rightarrow \infty$.
#
# This has very important implications for hypothesis testing and is precisely the reason why the t-distribution begins to approximate the normal distribution as our sample size increases.
# + colab={"base_uri": "https://localhost:8080/", "height": 72} colab_type="code" id="F15l-J788ApQ" outputId="afb637fe-330d-4292-a718-70587f0a1d97"
sample_means_small = []
sample_means_large = []
for _ in range(0,3000):
coinflips_small = np.random.binomial(n=1, p=.5, size=20)
coinflips_large = np.random.binomial(n=1, p=.5, size=100)
one_small_sample = coinflips_small
one_small_large = coinflips_large
sample_means_small.append(coinflips_small.mean())
sample_means_large.append(coinflips_large.mean())
print(len(sample_means_small))
print(sample_means_small[:100])
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="JdES8jXz8kAG" outputId="7f1ca5c1-62cc-4bea-c808-12f943166c2a"
# Stack the distributions of the sample means on one plot
fig, ax = plt.subplots()
for sample in [sample_means_small, sample_means_large]:
sns.distplot(sample)
# -
# - The yellow / orange distribution
# - more accurately represents the sample mean
# - less standard error
# - The blue distribution
# - confidence interval is wider / not as confident
#
# > Visualizes how the increase in $n$ tightens the confidence interval
# ---
# + [markdown] colab_type="text" id="EYqo5vZZSFUr"
# ## Standard Error of the Mean
#
# What does it mean to "estimate"? the Population mean?
# + colab={} colab_type="code" id="puGXH6vbSIE4"
# Sample mean for a single sample
df.single_sample.mean()
# -
# ---
# + [markdown] colab_type="text" id="nfdQf8QYUUmw"
# ## Build and Interpret a Confidence Interval
#
# <img src="https://github.com/ryanallredblog/ryanallredblog.github.io/blob/master/img/Confidence_Interval.png?raw=true" width=400>
# -
# ## $\bar{X} ± t \frac{s}{\sqrt{n}}$
# + colab={} colab_type="code" id="tBx71Kf0UjT3"
def confidence_interval(data, confidence=0.95):
"""
Calculates a confidence interval around a sample mean for given data,
using t-distribution and two-tailed test, default 95% confidence.
Arguments:
data - iterable (list or numpy array) of sample observations
confidence - level of confidence for the interval
Returns:
tuple of (mean, lower bound, upper bound)
"""
data = np.array(data) # Standardize to numpy array
mean = np.mean(data)
n = len(data)
# Stdev divided by sqare root of n (degrees of freedom / # observations)
stderr = np.std(data, ddof=1) / np.sqrt(n)
# stderr = stats.sem(data)
# Std error multiplied by t-statistic
margin_of_error = stderr * stats.t.ppf((1 + confidence) / 2.0, n - 1)
print(margin_of_error)
return (mean, mean - margin_of_error, mean + margin_of_error)
# -
# Create some more coinflips
coinflips_42 = np.random.binomial(n=1, p=0.5, size=42)
print(np.std(coinflips_42, ddof=1)) # ddof=1 tells numpy to use sample var/stdev
print(coinflips_42)
# Calculate the (mean, lower_bound, upper_bound)
confidence_interval(coinflips_42)
# More coinflips
coinflips_500 = np.random.binomial(n=1, p=0.5, size=500)
print(np.std(coinflips_500, ddof=1))
print(coinflips_500[:50])
# Calculate the (mean, lower_bound, upper_bound)
confidence_interval(coinflips_500)
# #### Looking at stats.t.ppf
# `stats.t.ppf(probability cutoff, degrees of freedom)`
#
# 95% confidence_level -> 0.25
#
# (1 - confidence_level) == 0.05 / 2 -> 0.25
# +
# Break up the one-liner in the function into its component atoms
n = 42
confidence_level = 0.95
dof = n - 1
stats.t.ppf((1 + confidence_level) / 2, dof) # The plus gives the positive t-statistic
# -
# ---
# + [markdown] colab_type="text" id="C4rtc8luVUAK"
# ## Graphically Represent a Confidence Interval
# + colab={} colab_type="code" id="pz6F9_3_VmKr"
coinflips_69 = np.random.binomial(n=1, p=0.5, size=69)
sns.kdeplot(coinflips_69)
ci = confidence_interval(coinflips_42)
plt.axvline(x=ci[1], color="r")
plt.axvline(x=ci[2], color="r")
plt.axvline(x=ci[0], color="k")
# -
# ---
# + [markdown] colab_type="text" id="_oy0uoBGeoEb"
# ## Relationship between Confidence Intervals and T-tests
#
# Confidence Interval == Bounds of statistical significance for our t-test
#
# A sample mean that falls inside of our confidence interval will "FAIL TO REJECT" our null hypothesis
#
# A sample mean that falls outside of our confidence interval will "REJECT" our null hypothesis
# + colab={"base_uri": "https://localhost:8080/", "height": 55} colab_type="code" id="Y7HwdMwDfL1N" outputId="43309626-838a-4d17-8507-038e7a0f6b74"
# More coinflips
coinflip_means = []
for x in range(0,100):
coinflips = np.random.binomial(n=1, p=.5, size=30)
coinflip_means.append(coinflips.mean())
print(coinflip_means[:16])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="nQDo-ZXlfOvR" outputId="c2e8386e-a0c4-49a4-f2ca-447fa92463f9"
# Sample Size
n = len(coinflip_means)
# Degrees of Freedom
dof = n-1
# The Mean of Means
mean = np.mean(coinflip_means)
# Sample Standard Deviation
sample_std = np.std(coinflip_means, ddof=1)
# Standard Error
std_err = sample_std/n**.5
CI = stats.t.interval(.95, dof, loc=mean, scale=std_err)
print("95% Confidence Interval: ", CI)
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="PiaALHSNfWou" outputId="612dbe6c-be4e-4cbb-9aeb-81309fb54529"
'''You can roll your own CI calculation pretty easily.
The only thing that's a little bit challenging
is understanding the t stat lookup'''
# 95% confidence interval
t_stat = stats.t.ppf(.975, dof)
print("t Statistic:", t_stat)
CI = (mean - (t_stat * std_err), mean + (t_stat * std_err))
print("Confidence Interval", CI)
# + [markdown] colab_type="text" id="EamZNJhAf-fY"
# A null hypothesis that's just inside of our confidence interval == fail to reject
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="cNpzYbjpfirR" outputId="ec1072cb-5130-43c8-c026-df6728f521e8"
stats.ttest_1samp(coinflip_means, .4901)
# + [markdown] colab_type="text" id="hO34mbL9gHn1"
# A null hypothesis that's just outside of our confidence interval == reject
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="N4SUjj82gKlv" outputId="8acaad7f-e23a-4c7a-a6aa-dc5c6a19b337"
stats.ttest_1samp(coinflip_means, .4899)
# -
# ---
# + [markdown] colab_type="text" id="pTIzrkKdUaLl"
# ## Run a $\chi^{2}$ Test "by hand" (Using Numpy)
# -
# \begin{align}
# \chi^2 = \sum \frac{(observed_{ij}-expected_{ij})^2}{(expected_{ij})}
# \end{align}
# + [markdown] colab={} colab_type="code" id="DDsovHUyUj3v"
# Chi-squared test is testing for relationships between two categorical variables.
#
# Inner cells of the table is called a "contingency table".
# + colab={} colab_type="code" id="X52Nwt7AVlvk"
# Load the "adults" dataset from github
df = pd.read_csv('https://raw.githubusercontent.com/ryanleeallred/datasets/master/adult.csv', na_values=" ?")
print(df.shape)
df.head()
# -
df.describe()
df.describe(exclude="number")
# +
cut_points = [0, 9, 19, 29, 39, 49, 1000]
label_names = ['0-9', '10-19', '20-29', '30-39', '40-49', '50+']
df['hours_per_week_categories'] = pd.cut(df['hours-per-week'], cut_points, labels=label_names)
df.head()
# -
df['sex'].value_counts()
df['hours_per_week_categories'].value_counts()
# +
# Sort first to workaround a Pandas crosstab bug
df = df.sort_values(by='hours_per_week_categories', ascending=True)
df.head()
# +
# Create contingency table
contingency_table = pd.crosstab(df['sex'], df['hours_per_week_categories'], margins=True)
contingency_table
# -
# Female count / row
femalecount = contingency_table.iloc[0][0:6].values
femalecount
# Male couint / row
malecount = contingency_table.iloc[1][0:6].values
malecount
# Plot male / female with bar chart
fig = plt.figure(figsize=(10, 5))
sns.set(font_scale=1.8)
categories = ["0-9","10-19","20-29","30-39","40-49","50+"]
p1 = plt.bar(categories, malecount, 0.55, color='#d62728')
p2 = plt.bar(categories, femalecount, 0.55, bottom=malecount)
plt.legend((p2[0], p1[0]), ('Female', 'Male'))
plt.xlabel('Hours per Week Worked')
plt.ylabel('Count')
plt.show()
# ### Expected Value Calculation
# \begin{align}
# expected_{i,j} =\frac{(row_{i} \text{total})(column_{j} \text{total}) }{(\text{total observations})}
# \end{align}
# +
row_sums = contingency_table.iloc[0:2, 6].values
col_sums = contingency_table.iloc[2, 0:6].values
print(row_sums)
print(col_sums)
# -
total = contingency_table.loc['All','All']
total
df.shape[0]
# +
expected = []
for i in range(len(row_sums)):
expected_row = []
for column in col_sums:
expected_val = column*row_sums[i]/total
expected_row.append(expected_val)
expected.append(expected_row)
expected = np.array(expected)
print(expected.shape)
print(expected)
# -
observed = pd.crosstab(df['sex'], df['hours_per_week_categories']).values
print(observed.shape)
observed
# ### Chi-Squared Statistic with Numpy
# \begin{align}
# \chi^2 = \sum \frac{(observed_{i}-expected_{i})^2}{(expected_{i})}
# \end{align}
# For the $observed$ values we will just use a version of our contingency table without the margins as a numpy array. In this way, if our observed values array and our expected values array are the same shape, then we can subtract them and divide them directly which makes the calculations a lot cleaner. No for loops!
# Array broadcasting will work with numpy arrays but not python lists
chi_squared = ((observed - expected)**2/(expected)).sum()
print(f"Chi-Squared: {chi_squared}") # Chi-Squared numbers are generally much larger
# #### Degrees of Freedom of a Chi-squared test
#
# degrees_of_freedom = (num_rows - 1)(num_columns - 1)
# Calculate Degrees of Freedom
dof = (len(row_sums) - 1) * (len(col_sums) - 1)
print(f"Degrees of Freedom: {dof}")
# > *Note:* Chi-Square will be on sprint challenge, but can use the NumPy one-liner method
# ---
# + [markdown] colab_type="text" id="7Igz-XHcVbW3"
# ## Run a $\chi^{2}$ Test using Scipy
# +
chi_squared, p_value, dof, expected = stats.chi2_contingency(observed)
print(f"Chi-Squared: {chi_squared}")
print(f"P-value: {p_value}")
print(f"Degrees of Freedom: {dof}")
print("Expected: \n", np.array(expected))
# -
# Null Hypothesis: Hours worked per week bins is **independent** of sex.
#
# Due to a p-value of 0, we REJECT the null hypothesis that hours worked per week and sex are independent, and conclude that there is an association between hours worked per week and sex.
| module2-sampling-confidence-intervals-and-hypothesis-testing/LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Needed in general
from __future__ import division
import numpy as np
import matplotlib.pyplot as plt
import sys
import h5py as h5
import gc
#Quick fudge to make import from ../Scripts work
sys.path.append('../../Scripts')
#Custom imports
import ClassCOMPAS as CC ###
# import ClassFormationChannels as FC
# import script that has formation channel classification functions:
from PostProcessingScripts import *
from ClassFormationChannels_5mainchannels import *
from astropy import units as u
from astropy import constants as const
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
print(fs)
import seaborn as sns
# +
fs=20
def plot2DdistributionScatter(axe, var, BPSmodelName='A'):
DCOtype='BHNS'
# path for files
path_ = '/Volumes/Andromeda/DATA/AllDCO_bugfix/' + alphabetDirDict[BPSmodelName] +'/'
path = path_ + 'COMPASCompactOutput_'+ DCOtype +'_' + BPSmodelName + '.h5'
# read in data
fdata = h5.File(path)
# M1 will be the most massive, M2 the least massive compact object.
M1, M2 = obtainM1BHandM2BHassymetric(m1=fdata['doubleCompactObjects']['M1'][...].squeeze(), m2=fdata['doubleCompactObjects']['M2'][...].squeeze())
Chirpmass = chirpmass(M1, M2)
MassRatio = M2/M1
### read in MSSFR weights: ###
mssfr = '112' # mssfr that matches LIGO observed rates.
# get detected weights
fparam_key = 'weights_detected'
weightheader = 'w_' + mssfr
w = fdata[fparam_key][weightheader][...].squeeze()
labellist=[ r'$\rm{M}_{\rm{chirp}}$', r'$\rm{M}_{\rm{BH}} \ [\rm{M}_{\odot}]$', r'$\rm{M}_{\rm{NS}} \ [\rm{M}_{\odot}]$',\
r'$q $'] #, r'$\log_{10}(t) $']
varlist = [Chirpmass, M1, M2, MassRatio]
maskGW200105 = ((M1 <= (8.9+1.2)) & (M1>=(8.9-1.5))) & ((M2 <= (1.9+0.3)) & (M2>=(1.9-0.2))) & ((chirpmass(M1,M2)>=(3.41-0.07)) & (chirpmass(M1,M2)<=(3.41+0.08))) & ((M2/M1>=(0.26-0.1)) & (M2/M1<=(0.26+0.35)))
maskGW200115 = ((M1 <= (5.7+1.8)) & (M1>=(5.7-2.1))) & ((M2 <= (1.5+0.7)) & (M2>=(1.5-0.3))) & ((chirpmass(M1,M2)>=(2.42-0.07)) & (chirpmass(M1,M2)<=(2.42+0.05))) & ((M2/M1>=(0.26-0.1)) & (M2/M1<=(0.26+0.35)))
fs_l = 20 # label fontsize
cGW1 = 'lightskyblue'
cGW0 = 'orange'
LEGEND = True
for jj, param_x in enumerate(var):
jj_minn,jj_maxx = min(varlist[jj]), max(varlist[jj])
for ii, param_y in enumerate(var):
ii_minn,ii_maxx = min(varlist[ii]), max(varlist[ii])
if ii==jj:
nrbins=np.linspace(jj_minn, jj_maxx, 100)
hist, bin_edges = np.histogram(varlist[jj], bins=nrbins, weights=w)
yy = hist / np.max(hist) # normalize by max to set highest bin to fixed height 1
xx = (bin_edges[:-1]+ bin_edges[1:])/2 # center of bins
axe[ii,jj].plot(xx, yy, color='lightgray', lw=2, zorder=2)
axe[ii,jj].fill_between(xx, y1=np.zeros_like(xx), y2=yy, color='lightgray', zorder=2, alpha=0.3)
hist, bin_edges = np.histogram(varlist[jj][maskGW200105], bins=nrbins, weights=w[maskGW200105])
yy = hist / np.max(hist) # normalize by max to set highest bin to fixed height 1
xx = (bin_edges[:-1]+ bin_edges[1:])/2 # center of bins
axe[ii,jj].plot(xx, yy, color=cGW0, lw=2, zorder=3)
axe[ii,jj].fill_between(xx, y1=np.zeros_like(xx), y2=yy, color=cGW0, zorder=2, alpha=0.3)
hist, bin_edges = np.histogram(varlist[jj][maskGW200115], bins=nrbins, weights=w[maskGW200115])
yy = hist / np.max(hist) # normalize by max to set highest bin to fixed height 1
xx = (bin_edges[:-1]+ bin_edges[1:])/2 # center of bins
axe[ii,jj].plot(xx, yy, color=cGW1, lw=2, zorder=3)
axe[ii,jj].fill_between(xx, y1=np.zeros_like(xx), y2=yy, color=cGW1, zorder=2, alpha=0.3)
ylabel='PDF'
print('ii,jj=', ii, jj)
axe[ii,jj].set_xlim(jj_minn, jj_maxx)
axe[ii,jj].set_ylim(0,1)
if param_x==r'$q $':
xx = [-100, -50]
axe[ii,jj].fill_between(xx, y1=np.zeros_like(xx), y2=np.ones_like(xx), color='lightgray', zorder=2, alpha=0.3, label='All BHNS')
axe[ii,jj].fill_between(xx, y1=np.zeros_like(xx), y2=np.ones_like(xx), color='orange', zorder=2, alpha=0.3, label = 'GW200105')
axe[ii,jj].fill_between(xx, y1=np.zeros_like(xx), y2=np.ones_like(xx), color='lightskyblue', zorder=2, alpha=0.3, label = 'GW200115')
# ax[ii,jj].set_legend(fontsize=24, loc='top right')
ax[ii,jj].legend(fontsize=26, loc='upper center', bbox_to_anchor=(0.65, 0.9), frameon=False)
if jj==(len(var)-1):
ax[ii,jj] == layoutAxesNoYlabel(ax[ii,jj], nameX=labellist[jj], nameY='NA', setMinor=False, labelpad=10, fontsize=fs_l)
if ii!=0:
axe[ii,jj].set_yticks([])
axe[ii,jj].set_yticklabels( () )
if ii==0:
ax[ii,jj] == layoutAxesNoXlabel(ax[ii,jj], nameX='NA', nameY=r'$\textbf{PDF}$', setMinor=False, labelpad=10, fontsize=fs_l)
elif jj>ii:
ax[ii,jj].set_axis_off()
else:
print('ii,jj=', ii, jj)
axe[ii,jj].set_xlim(jj_minn, jj_maxx)
axe[ii,jj].set_ylim(ii_minn, ii_maxx)
x = varlist[jj] #np.linspace(0,1,100)
y = np.ones_like(varlist[jj])
print(np.shape(varlist[jj]))
print(np.shape(x))
N = 10000
sns.kdeplot(varlist[jj][0:N], varlist[ii][0:N], weights=w[0:N], ax=ax[ii, jj], fill=True, color='gray')# , levels=[0.5, 0.68, 0.98], bw_method=10)
# ax[ii,jj] = sns.kdeplot(x=varlist[jj], y=varlist[ii], fill=True, color='r', shade=True, Label='Iris_Setosa',
# cmap="Reds", shade_lowest=False)
# ax[ii,jj].scatter(varlist[jj], varlist[ii], s=140*w, color='lightgray', zorder=1)
# ax[ii,jj].scatter(varlist[jj][maskGW200105], varlist[ii][maskGW200105], s=140*w[maskGW200105], color='orange', zorder=10)
# ax[ii,jj].scatter(varlist[jj][maskGW200115], varlist[ii][maskGW200115], s=140*w[maskGW200115], color='lightskyblue', zorder=10)
if (ii==(len(var)-1)):
print(ii)
# ax[ii,jj].set_xlabel(labellist[jj], fontsize=fs)
ax[ii,jj] == layoutAxesNoYlabel(ax[ii,jj], nameX=labellist[jj], nameY='NA', setMinor=False, labelpad=10, fontsize=fs_l)
if jj==0:
# ax[ii,jj].set_ylabel(labellist[ii], fontsize=fs)
ax[ii,jj] == layoutAxesNoXlabel(ax[ii,jj], nameX='NA', nameY=labellist[ii], setMinor=False, labelpad=10, fontsize=fs_l)
else:
axe[ii,jj].set_yticks([])
axe[ii,jj].set_yticklabels( () )
labellist=[ r'$M_{\rm{chirp}}$', r'$M_{\rm{BH}} \ [M_{\odot}]$', r'$M_{\rm{NS}} \ [M_{\odot}]$',\
r'$q $'] #, r'$\log_{10}(t) $']
f, ax= plt.subplots(ncols=len(labellist),nrows=len(labellist),figsize=(20,20),
gridspec_kw={"width_ratios":1.5*np.ones(len(labellist)), "height_ratios":2*np.ones(len(labellist))})
mssfr = '112' # mssfr that matches LIGO observed rates.
plt.tight_layout()
plt.subplots_adjust(wspace=0, hspace=0)#2)
model='P'
plot2DdistributionScatter(axe=ax, var=labellist, BPSmodelName=model)
plt.savefig('./Scatter_Final_kde' + mssfr +'_' + model+ '.png', transparent=False, bbox_inches="tight")
plt.show()
| plottingCode/.ipynb_checkpoints/Spin -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="oBlTecNJP2WJ"
# # Reference
#
# This example is taken from the book [DL with Python](https://www.manning.com/books/deep-learning-with-python) by <NAME>. It explains how to retrain a pre-trained CNN classifier
#
# All the notebooks from the book are available for free on [Github](https://github.com/fchollet/deep-learning-with-python-notebooks)
#
# If you like to run the example locally follow the instructions provided on [Keras website](https://keras.io/#installation)
#
# [](https://colab.research.google.com/github/digitalideation/comppx_h2101/blob/master/samples/week07/04-using-a-pretrained-convnet.ipynb)
#
# ---
# + [markdown] id="efiveAHoP2WN"
# # Using a pre-trained convnet
#
# This notebook contains the code sample found in Chapter 5, Section 3 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
#
# ----
#
# A common and highly effective approach to deep learning on small image datasets is to leverage a pre-trained network. A pre-trained network
# is simply a saved network previously trained on a large dataset, typically on a large-scale image classification task. If this original
# dataset is large enough and general enough, then the spatial feature hierarchy learned by the pre-trained network can effectively act as a
# generic model of our visual world, and hence its features can prove useful for many different computer vision problems, even though these
# new problems might involve completely different classes from those of the original task. For instance, one might train a network on
# ImageNet (where classes are mostly animals and everyday objects) and then re-purpose this trained network for something as remote as
# identifying furniture items in images. Such portability of learned features across different problems is a key advantage of deep learning
# compared to many older shallow learning approaches, and it makes deep learning very effective for small-data problems.
#
# In our case, we will consider a large convnet trained on the ImageNet dataset (1.4 million labeled images and 1000 different classes).
# ImageNet contains many animal classes, including different species of cats and dogs, and we can thus expect to perform very well on our cat
# vs. dog classification problem.
#
# We will use the VGG16 architecture, developed by <NAME> and <NAME> in 2014, a simple and widely used convnet architecture
# for ImageNet. Although it is a bit of an older model, far from the current state of the art and somewhat heavier than many other recent
# models, we chose it because its architecture is similar to what you are already familiar with, and easy to understand without introducing
# any new concepts. This may be your first encounter with one of these cutesie model names -- VGG, ResNet, Inception, Inception-ResNet,
# Xception... you will get used to them, as they will come up frequently if you keep doing deep learning for computer vision.
#
# There are two ways to leverage a pre-trained network: *feature extraction* and *fine-tuning*. We will cover both of them. Let's start with
# feature extraction.
# + [markdown] id="8K6kizPZP2WN"
# ## Feature extraction
#
# Feature extraction consists of using the representations learned by a previous network to extract interesting features from new samples.
# These features are then run through a new classifier, which is trained from scratch.
#
# As we saw previously, convnets used for image classification comprise two parts: they start with a series of pooling and convolution
# layers, and they end with a densely-connected classifier. The first part is called the "convolutional base" of the model. In the case of
# convnets, "feature extraction" will simply consist of taking the convolutional base of a previously-trained network, running the new data
# through it, and training a new classifier on top of the output.
#
# 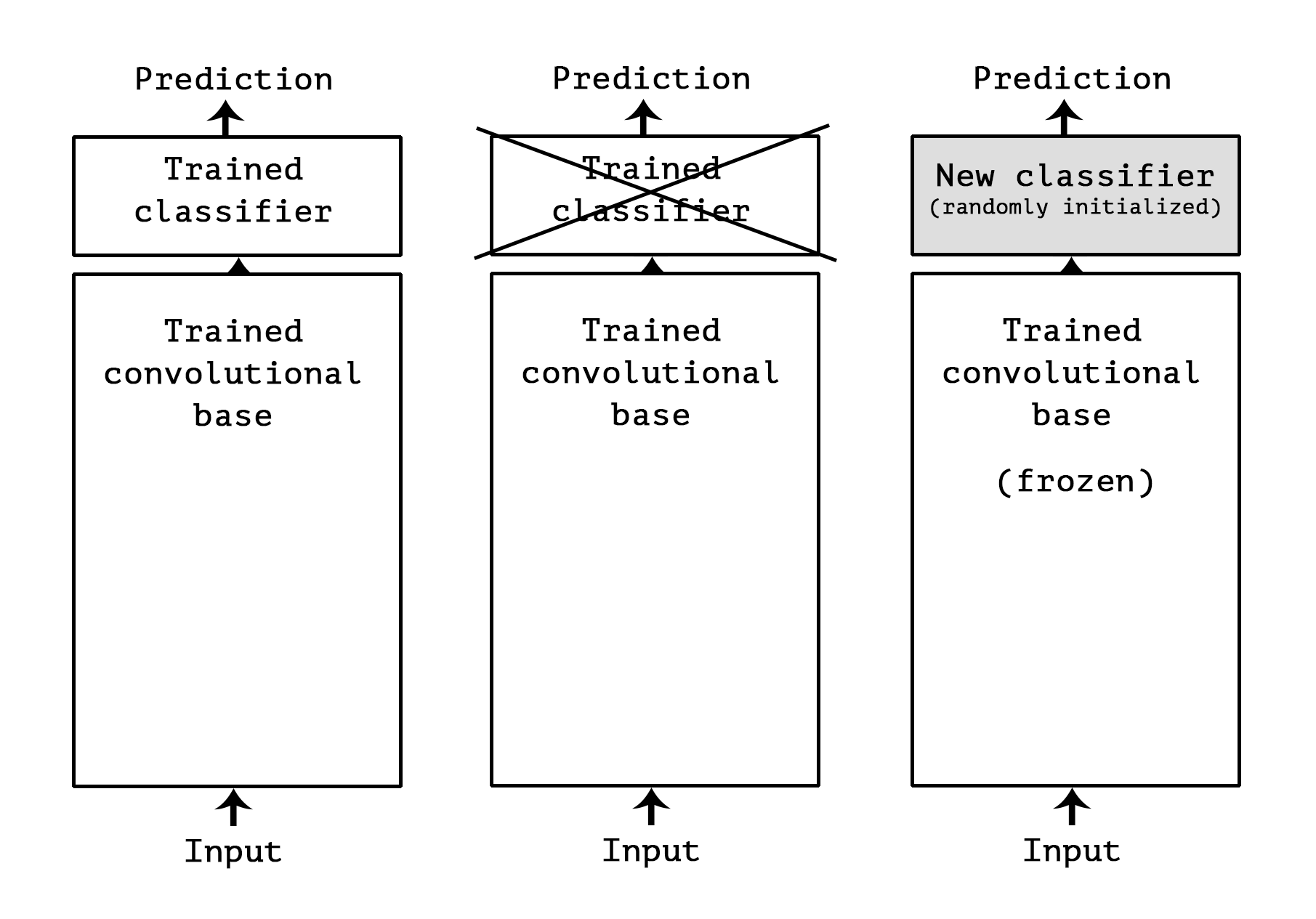
#
# Why only reuse the convolutional base? Could we reuse the densely-connected classifier as well? In general, it should be avoided. The
# reason is simply that the representations learned by the convolutional base are likely to be more generic and therefore more reusable: the
# feature maps of a convnet are presence maps of generic concepts over a picture, which is likely to be useful regardless of the computer
# vision problem at hand. On the other end, the representations learned by the classifier will necessarily be very specific to the set of
# classes that the model was trained on -- they will only contain information about the presence probability of this or that class in the
# entire picture. Additionally, representations found in densely-connected layers no longer contain any information about _where_ objects are
# located in the input image: these layers get rid of the notion of space, whereas the object location is still described by convolutional
# feature maps. For problems where object location matters, densely-connected features would be largely useless.
#
# Note that the level of generality (and therefore reusability) of the representations extracted by specific convolution layers depends on
# the depth of the layer in the model. Layers that come earlier in the model extract local, highly generic feature maps (such as visual
# edges, colors, and textures), while layers higher-up extract more abstract concepts (such as "cat ear" or "dog eye"). So if your new
# dataset differs a lot from the dataset that the original model was trained on, you may be better off using only the first few layers of the
# model to do feature extraction, rather than using the entire convolutional base.
#
# In our case, since the ImageNet class set did contain multiple dog and cat classes, it is likely that it would be beneficial to reuse the
# information contained in the densely-connected layers of the original model. However, we will chose not to, in order to cover the more
# general case where the class set of the new problem does not overlap with the class set of the original model.
# + [markdown] id="s1Se6YjyP2WO"
# Let's put this in practice by using the convolutional base of the VGG16 network, trained on ImageNet, to extract interesting features from
# our cat and dog images, and then training a cat vs. dog classifier on top of these features.
#
# The VGG16 model, among others, comes pre-packaged with Keras. You can import it from the `keras.applications` module. Here's the list of
# image classification models (all pre-trained on the ImageNet dataset) that are available as part of `keras.applications`:
#
# * Xception
# * InceptionV3
# * ResNet50
# * VGG16
# * VGG19
# * MobileNet
#
# Let's instantiate the VGG16 model:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} id="kK5ru0ijP2WO" outputId="851c33f3-f4ec-4868-9f96-ca1dbe46d137"
from tensorflow.keras.applications import VGG16
conv_base = VGG16(weights='imagenet',
include_top=False,
input_shape=(150, 150, 3))
# + [markdown] id="xd-5yw_QP2WR"
# We passed three arguments to the constructor:
#
# * `weights`, to specify which weight checkpoint to initialize the model from
# * `include_top`, which refers to including or not the densely-connected classifier on top of the network. By default, this
# densely-connected classifier would correspond to the 1000 classes from ImageNet. Since we intend to use our own densely-connected
# classifier (with only two classes, cat and dog), we don't need to include it.
# * `input_shape`, the shape of the image tensors that we will feed to the network. This argument is purely optional: if we don't pass it,
# then the network will be able to process inputs of any size.
#
# Here's the detail of the architecture of the VGG16 convolutional base: it's very similar to the simple convnets that you are already
# familiar with.
# + colab={"base_uri": "https://localhost:8080/", "height": 799} id="pBKP9CPTP2WS" outputId="139b3f15-55b6-405b-e903-6534353a4f5f"
conv_base.summary()
# + [markdown] id="rSVkEfPvP2WU"
# The final feature map has shape `(4, 4, 512)`. That's the feature on top of which we will stick a densely-connected classifier.
#
# At this point, there are two ways we could proceed:
#
# * Running the convolutional base over our dataset, recording its output to a Numpy array on disk, then using this data as input to a
# standalone densely-connected classifier similar to those you have seen in the first chapters of this book. This solution is very fast and
# cheap to run, because it only requires running the convolutional base once for every input image, and the convolutional base is by far the
# most expensive part of the pipeline. However, for the exact same reason, this technique would not allow us to leverage data augmentation at
# all.
# * Extending the model we have (`conv_base`) by adding `Dense` layers on top, and running the whole thing end-to-end on the input data. This
# allows us to use data augmentation, because every input image is going through the convolutional base every time it is seen by the model.
# However, for this same reason, this technique is far more expensive than the first one.
#
# We will cover both techniques. Let's walk through the code required to set-up the first one: recording the output of `conv_base` on our
# data and using these outputs as inputs to a new model.
#
# We will start by simply running instances of the previously-introduced `ImageDataGenerator` to extract images as Numpy arrays as well as
# their labels. We will extract features from these images simply by calling the `predict` method of the `conv_base` model.
# + [markdown] id="qFbYacarP2WX"
# ## Run those cells If you're running the code on Colab
#
# We will be using the data that has been preprocessed in `week04-03-using-convnets-with-small-datasets.ipynb`
# + colab={"base_uri": "https://localhost:8080/", "height": 85} id="q4ulF5ZWP2WX" outputId="e9edfa14-15c5-428d-feab-5ea020f2e857"
# !gdown https://drive.google.com/uc?id=1-Evs12qbw6DV13JYJJjz-0zyY_qsZtdL
# + id="lGk5D4BDQdzu"
# !unzip dogs-vs-cats-small.zip -d .
# + colab={"base_uri": "https://localhost:8080/", "height": 68} id="2nlCft2EP2Wg" outputId="4e01f58b-cc10-4c3b-8dc6-681421a5e4f3"
import os
import numpy as np
from tensorflow.keras.preprocessing.image import ImageDataGenerator
base_dir = 'dogs-vs-cats-small'
models_dir = 'models'
os.makedirs(models_dir, exist_ok=True)
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
datagen = ImageDataGenerator(rescale=1./255)
batch_size = 20
def extract_features(directory, sample_count):
features = np.zeros(shape=(sample_count, 4, 4, 512))
labels = np.zeros(shape=(sample_count))
generator = datagen.flow_from_directory(
directory,
target_size=(150, 150),
batch_size=batch_size,
class_mode='binary')
i = 0
for inputs_batch, labels_batch in generator:
features_batch = conv_base.predict(inputs_batch)
features[i * batch_size : (i + 1) * batch_size] = features_batch
labels[i * batch_size : (i + 1) * batch_size] = labels_batch
i += 1
if i * batch_size >= sample_count:
# Note that since generators yield data indefinitely in a loop,
# we must `break` after every image has been seen once.
break
return features, labels
train_features, train_labels = extract_features(train_dir, 2000)
validation_features, validation_labels = extract_features(validation_dir, 1000)
test_features, test_labels = extract_features(test_dir, 1000)
# + [markdown] id="Ke1avKojP2Wi"
# The extracted features are currently of shape `(samples, 4, 4, 512)`. We will feed them to a densely-connected classifier, so first we must
# flatten them to `(samples, 8192)`:
# + id="ksUV4TyFP2Wj"
train_features = np.reshape(train_features, (2000, 4 * 4 * 512))
validation_features = np.reshape(validation_features, (1000, 4 * 4 * 512))
test_features = np.reshape(test_features, (1000, 4 * 4 * 512))
# + [markdown] id="fbXaJBXuP2Wl"
# At this point, we can define our densely-connected classifier (note the use of dropout for regularization), and train it on the data and
# labels that we just recorded:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="a1d0eSCoP2Wl" outputId="cf65a8c8-0d36-4ae3-9cd0-38295be7a24d"
from tensorflow.keras import models
from tensorflow.keras import layers
from tensorflow.keras import optimizers
model = models.Sequential([
layers.Dense(256, activation='relu', input_dim=4 * 4 * 512),
layers.Dropout(0.5),
layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer=optimizers.RMSprop(learning_rate=2e-5),
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(train_features, train_labels,
epochs=30,
batch_size=20,
validation_data=(validation_features, validation_labels))
# + [markdown] id="7AtqgmuTP2Wn"
# Training is very fast, since we only have to deal with two `Dense` layers -- an epoch takes less than one second even on CPU.
#
# Let's take a look at the loss and accuracy curves during training:
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="ArLxUx7AP2Wn" outputId="c8f8e80a-ced8-4435-8ef5-ca696800c565"
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="aVP_JpM6P2Wp"
#
# We reach a validation accuracy of about 90%, much better than what we could achieve in the previous section with our small model trained from
# scratch. However, our plots also indicate that we are overfitting almost from the start -- despite using dropout with a fairly large rate.
# This is because this technique does not leverage data augmentation, which is essential to preventing overfitting with small image datasets.
#
# Now, let's review the second technique we mentioned for doing feature extraction, which is much slower and more expensive, but which allows
# us to leverage data augmentation during training: extending the `conv_base` model and running it end-to-end on the inputs. Note that this
# technique is in fact so expensive that you should only attempt it if you have access to a GPU: it is absolutely intractable on CPU. If you
# cannot run your code on GPU, then the previous technique is the way to go.
#
# Because models behave just like layers, you can add a model (like our `conv_base`) to a `Sequential` model just like you would add a layer.
# So you can do the following:
# + id="e8LIIYjgP2Wq"
model = models.Sequential([
conv_base,
layers.Flatten(),
layers.Dense(256, activation='relu'),
layers.Dense(1, activation='sigmoid')
])
# + [markdown] id="dgmFwoDKP2Ws"
# This is what our model looks like now:
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="ygFEDhLsP2Ws" outputId="12ae2fb5-8ea3-4a0e-83ac-412ed1a9460d"
model.summary()
# + [markdown] id="qRvaKzC1P2Wu"
# As you can see, the convolutional base of VGG16 has 14,714,688 parameters, which is very large. The classifier we are adding on top has 2
# million parameters.
#
# Before we compile and train our model, a very important thing to do is to freeze the convolutional base. "Freezing" a layer or set of
# layers means preventing their weights from getting updated during training. If we don't do this, then the representations that were
# previously learned by the convolutional base would get modified during training. Since the `Dense` layers on top are randomly initialized,
# very large weight updates would be propagated through the network, effectively destroying the representations previously learned.
#
# In Keras, freezing a network is done by setting its `trainable` attribute to `False`:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="ZVXICheNP2Wu" outputId="fa02fbbb-f91f-4084-f456-19d42ebf6c5c"
print('This is the number of trainable weights '
'before freezing the conv base:', len(model.trainable_weights))
# + id="OVwbf6J9P2Ww"
conv_base.trainable = False
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="jI-HMwxMP2Wy" outputId="9f078d75-2ae1-407a-f1fa-7b1e6b21efcc"
print('This is the number of trainable weights '
'after freezing the conv base:', len(model.trainable_weights))
# + [markdown] id="vKVSSQbIP2W0"
# With this setup, only the weights from the two `Dense` layers that we added will be trained. That's a total of four weight tensors: two per
# layer (the main weight matrix and the bias vector). Note that in order for these changes to take effect, we must first compile the model.
# If you ever modify weight trainability after compilation, you should then re-compile the model, or these changes would be ignored.
#
# Now we can start training our model, with the same data augmentation configuration that we used in our previous example:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="gr3PCjZVP2W0" outputId="a76b89c5-d475-45d2-b09e-0cfd530a9059"
from tensorflow.keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
# Note that the validation data should not be augmented!
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
# This is the target directory
train_dir,
# All images will be resized to 150x150
target_size=(150, 150),
batch_size=20,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(learning_rate=2e-5),
metrics=['acc'])
history = model.fit(
train_generator,
steps_per_epoch=2000//32-1,
epochs=30,
validation_data=validation_generator,
validation_steps=50,
verbose=2)
# + id="EQHNLwzFP2W2"
model.save(os.path.join(models_dir, 'cats_and_dogs_small_3.h5'))
# + [markdown] id="RBd9ZmtcP2W4"
# Let's plot our results again:
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="TbS52YMqP2W4" outputId="992c1173-dbf2-40cf-aba8-96cf9c2150d2"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="PAPRUQNsP2W6"
# As you can see, we reach a validation accuracy of about 96%. This is much better than our small convnet trained from scratch.
# + [markdown] id="D90E7y2jP2W7"
# ## Fine-tuning
#
# Another widely used technique for model reuse, complementary to feature extraction, is _fine-tuning_.
# Fine-tuning consists in unfreezing a few of the top layers
# of a frozen model base used for feature extraction, and jointly training both the newly added part of the model (in our case, the
# fully-connected classifier) and these top layers. This is called "fine-tuning" because it slightly adjusts the more abstract
# representations of the model being reused, in order to make them more relevant for the problem at hand.
#
# 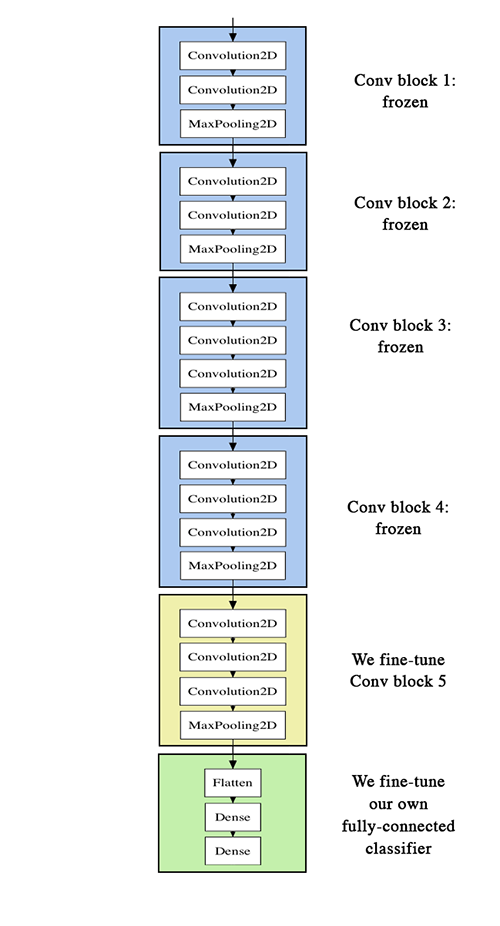
# + [markdown] id="3miSdJm6P2W7"
# We have stated before that it was necessary to freeze the convolution base of VGG16 in order to be able to train a randomly initialized
# classifier on top. For the same reason, it is only possible to fine-tune the top layers of the convolutional base once the classifier on
# top has already been trained. If the classified wasn't already trained, then the error signal propagating through the network during
# training would be too large, and the representations previously learned by the layers being fine-tuned would be destroyed. Thus the steps
# for fine-tuning a network are as follow:
#
# * 1) Add your custom network on top of an already trained base network.
# * 2) Freeze the base network.
# * 3) Train the part you added.
# * 4) Unfreeze some layers in the base network.
# * 5) Jointly train both these layers and the part you added.
#
# We have already completed the first 3 steps when doing feature extraction. Let's proceed with the 4th step: we will unfreeze our `conv_base`,
# and then freeze individual layers inside of it.
#
# As a reminder, this is what our convolutional base looks like:
# + colab={"base_uri": "https://localhost:8080/", "height": 799} id="yRuKlKjAP2W7" outputId="13791ae5-64e0-4734-d374-cec446a7840f"
conv_base.summary()
# + [markdown] id="7-8JoEbaP2W9"
#
# We will fine-tune the last 3 convolutional layers, which means that all layers up until `block4_pool` should be frozen, and the layers
# `block5_conv1`, `block5_conv2` and `block5_conv3` should be trainable.
#
# Why not fine-tune more layers? Why not fine-tune the entire convolutional base? We could. However, we need to consider that:
#
# * Earlier layers in the convolutional base encode more generic, reusable features, while layers higher up encode more specialized features. It is
# more useful to fine-tune the more specialized features, as these are the ones that need to be repurposed on our new problem. There would
# be fast-decreasing returns in fine-tuning lower layers.
# * The more parameters we are training, the more we are at risk of overfitting. The convolutional base has 15M parameters, so it would be
# risky to attempt to train it on our small dataset.
#
# Thus, in our situation, it is a good strategy to only fine-tune the top 2 to 3 layers in the convolutional base.
#
# Let's set this up, starting from where we left off in the previous example:
# + id="V7K9mHifP2W9"
conv_base.trainable = True
set_trainable = False
for layer in conv_base.layers:
if layer.name == 'block5_conv1':
set_trainable = True
if set_trainable:
layer.trainable = True
else:
layer.trainable = False
# + [markdown] id="a1UMhyFBP2XA"
# Now we can start fine-tuning our network. We will do this with the RMSprop optimizer, using a very low learning rate. The reason for using
# a low learning rate is that we want to limit the magnitude of the modifications we make to the representations of the 3 layers that we are
# fine-tuning. Updates that are too large may harm these representations.
#
# Now let's proceed with fine-tuning:
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="PBEavRY6P2XA" outputId="6e724ebf-3fb3-4eac-8aa5-84385d5c49ca"
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(learning_rate=1e-5),
metrics=['acc'])
history = model.fit(
train_generator,
steps_per_epoch=2000//32-1,
epochs=100,
validation_data=validation_generator,
validation_steps=50)
# + id="Mnl7oxUOP2XC"
model.save(os.path.join(models_dir, 'cats_and_dogs_small_4.h5'))
# + [markdown] id="C9Pir04wP2XF"
# Let's plot our results using the same plotting code as before:
# + id="4S-ySR-2P2XF"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="RoPB198CP2XH"
#
# These curves look very noisy. To make them more readable, we can smooth them by replacing every loss and accuracy with exponential moving
# averages of these quantities. Here's a trivial utility function to do this:
# + id="j1zXhxiRP2XH"
def smooth_curve(points, factor=0.8):
smoothed_points = []
for point in points:
if smoothed_points:
previous = smoothed_points[-1]
smoothed_points.append(previous * factor + point * (1 - factor))
else:
smoothed_points.append(point)
return smoothed_points
plt.plot(epochs, smooth_curve(acc), 'bo', label='Smoothed training acc')
plt.plot(epochs, smooth_curve(val_acc), 'b', label='Smoothed validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, smooth_curve(loss), 'bo', label='Smoothed training loss')
plt.plot(epochs, smooth_curve(val_loss), 'b', label='Smoothed validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="GLTuIvSgP2XJ"
#
# These curves look much cleaner and more stable. We are seeing a nice 1% absolute improvement.
#
# Note that the loss curve does not show any real improvement (in fact, it is deteriorating). You may wonder, how could accuracy improve if the
# loss isn't decreasing? The answer is simple: what we display is an average of pointwise loss values, but what actually matters for accuracy
# is the distribution of the loss values, not their average, since accuracy is the result of a binary thresholding of the class probability
# predicted by the model. The model may still be improving even if this isn't reflected in the average loss.
#
# We can now finally evaluate this model on the test data:
# + id="-XNDXRn4P2XJ"
test_generator = test_datagen.flow_from_directory(
test_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
test_loss, test_acc = model.evaluate_generator(test_generator, steps=50)
print('test acc:', test_acc)
# + [markdown] id="UPS6xz7rP2XL"
#
# Here we get a test accuracy of 97%. In the original Kaggle competition around this dataset, this would have been one of the top results.
# However, using modern deep learning techniques, we managed to reach this result using only a very small fraction of the training data
# available (about 10%). There is a huge difference between being able to train on 20,000 samples compared to 2,000 samples!
# + [markdown] id="RgXnKbsLP2XM"
# ## Take-aways: using convnets with small datasets
#
# Here's what you should take away from the exercises of these past two sections:
#
# * Convnets are the best type of machine learning models for computer vision tasks. It is possible to train one from scratch even on a very
# small dataset, with decent results.
# * On a small dataset, overfitting will be the main issue. Data augmentation is a powerful way to fight overfitting when working with image
# data.
# * It is easy to reuse an existing convnet on a new dataset, via feature extraction. This is a very valuable technique for working with
# small image datasets.
# * As a complement to feature extraction, one may use fine-tuning, which adapts to a new problem some of the representations previously
# learned by an existing model. This pushes performance a bit further.
#
# Now you have a solid set of tools for dealing with image classification problems, in particular with small datasets.
| samples/week07/04-using-a-pretrained-convnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Mask-RCNN evaluate model. Balloon dataset
# +
import os
import tqdm
import matplotlib.pyplot as plt
os.chdir('..')
from samples.balloon import balloon
from preprocess import preprocess
from preprocess import augmentation as aug
from model import mask_rcnn_functional
import evaluating
from common import utils
from common import inference_utils
from common.inference_utils import process_input
from common.config import CONFIG
from common.inference_optimize import maskrcnn_to_onnx, modify_onnx_model
import numpy as np
import tensorflow as tf
utils.tf_limit_gpu_memory(tf, 2000)
# %matplotlib inline
# -
# %load_ext watermark
# %watermark
# %watermark --iversions
base_dir = os.getcwd().replace('src', 'balloon')
eval_dir = os.path.join(base_dir, 'val')
# +
from common.config import CONFIG
CONFIG.update(balloon.BALLON_CONFIG)
# -
eval_dataset = balloon.BalloonDataset(images_dir=eval_dir,
class_key='object',
classes_dict=CONFIG['class_dict'],
augmentation=aug.get_validation_augmentation(
image_size=CONFIG['img_size'],
normalize=CONFIG['normalization']
),
json_annotation_key=None,
**CONFIG
)
eval_dataloader = preprocess.DataLoader(eval_dataset,
shuffle=True,
cast_output=False,
return_original=True,
**CONFIG
)
weights_path = os.path.join('..', 'tests', 'samples', 'balloon',
'maskrcnn_mobilenet_ed3e7dd4c2e064d9dd92df2088834243_cp-0029.ckpt'
)
weights_path
# Loading inference graph and import weights
inference_config = CONFIG
inference_config.update({'training': False})
inference_model = mask_rcnn_functional(config=inference_config)
inference_model = inference_utils.load_mrcnn_weights(model=inference_model,
weights_path=weights_path,
verbose=True
)
# #### Evaluate data on a single batch with tensorflow
def tf_mrcnn_inference(model, infer_batch, eval_batch):
"""
Args:
model: tensorflow tf.keras.Model
infer_batch: prepared data for inference
eval_batch: ground truth data for evaluation
Returns: boxes,
class_ids,
scores,
ull_masks,
eval_gt_boxes,
eval_gt_class_ids,
eval_gt_masks
"""
# Extract inference inputs from dataloader
batch_images, batch_image_meta, batch_rpn_match, batch_rpn_bbox, \
batch_gt_class_ids, batch_gt_boxes, batch_gt_masks = infer_batch
# Extract original inputs from dataloader
eval_gt_image = eval_batch[0][0]
eval_gt_boxes = eval_batch[3][0]
eval_gt_class_ids = eval_batch[2][0]
eval_gt_masks = eval_batch[1][0]
# Make inference
output = model([batch_images, batch_image_meta])
detections, mrcnn_probs, mrcnn_bbox, mrcnn_mask, rpn_rois, rpn_class, rpn_bbox = output
# Extract bboxes, class_ids, scores and full-size masks
boxes, class_ids, scores, full_masks = \
utils.reformat_detections(detections=detections[0].numpy(),
mrcnn_mask=mrcnn_mask[0].numpy(),
original_image_shape=eval_gt_image.shape,
image_shape=batch_images[0].shape,
window=batch_image_meta[0][7:11]
)
return boxes, class_ids, scores, full_masks, eval_gt_boxes, eval_gt_class_ids, eval_gt_masks
def evaluate_mrcnn(model, inference_function, eval_dataloader, iou_limits=(0.5, 1), iou_step=0.05):
"""
Evaluate Mask-RCNN model
Args:
model: tensorflow tf.keras.Model
inference_function:
eval_dataloader:
iou_limits: start and end for IoU in mAP
iou_step: step for IoU in mAP
Returns:
"""
# Evaluate mAP
for eval_iou_threshold in np.arange(iou_limits[0], iou_limits[1], iou_step):
# Metrics lists
ap_list = []
precisions_list = []
recalls_list = []
eval_iterated = iter(eval_dataloader)
pbar = tqdm.tqdm(eval_iterated, total=eval_dataloader.__len__())
for eval_input, _ in pbar:
# Split batch into prepared data for inference and original data for evaluation
infer_batch = eval_input[:-4]
eval_batch = eval_input[-4:]
try:
boxes, class_ids, scores, full_masks, eval_gt_boxes, eval_gt_class_ids, eval_gt_masks = \
inference_function(model=model, infer_batch=infer_batch, eval_batch=eval_batch)
# Get AP, precisions, recalls, overlaps
ap, precisions, recalls, overlaps = \
evaluating.compute_ap(gt_boxes=eval_gt_boxes,
gt_class_ids=eval_gt_class_ids,
gt_masks=eval_gt_masks,
pred_boxes=boxes,
pred_class_ids=class_ids,
pred_scores=scores,
pred_masks=full_masks,
iou_threshold=eval_iou_threshold
)
postfix = ''
except:
postfix = 'Passed an image. AP added as zero.'
ap = 0.0
precisions = 0.0
recalls = 0.0
ap_list.append(ap)
precisions_list.append(precisions)
recalls_list.append(recalls)
# Update tqdm mAP
pbar.set_description(f"IoU: {eval_iou_threshold:.2f}. mAP: {np.mean(ap_list):.4f} ")# {postfix}
print(f'mAP={np.mean(ap_list):.4f}, IoU: {eval_iou_threshold:.2f}')
evaluate_mrcnn(model=inference_model,
inference_function=tf_mrcnn_inference,
eval_dataloader=eval_dataloader
)
# #### Evaluate data on a single batch with TensorRT
import tensorrt as trt
import pycuda.autoinit
import pycuda.driver as cuda
def trt_mrcnn_inference(model, infer_batch, eval_batch):
"""
Args:
model: tensorflow tf.keras.Model
infer_batch: prepared data for inference
eval_batch: ground truth data for evaluation
Returns: boxes,
class_ids,
scores, f
ull_masks,
eval_gt_boxes,
eval_gt_class_ids,
eval_gt_masks
"""
# Extract inference inputs from dataloader
batch_images, batch_image_meta, batch_rpn_match, batch_rpn_bbox, \
batch_gt_class_ids, batch_gt_boxes, batch_gt_masks = infer_batch
# Extract original inputs from dataloader
eval_gt_image = eval_batch[0][0]
eval_gt_boxes = eval_batch[3][0]
eval_gt_class_ids = eval_batch[2][0]
eval_gt_masks = eval_batch[1][0]
# Extract trt-variables from a dict for transparency
engine = model['engine']
stream = model['stream']
context = model['context']
device_input = model['device_input']
device_output1 = model['device_output1']
device_output2 = model['device_output2']
host_output1 = model['host_output1']
host_output2 = model['host_output2']
output_nodes = model['output_nodes']
graph_type = model['graph_type']
if graph_type == 'uff':
# Prepare image for uff original graph
input_image, window, scale, padding, crop = utils.resize_image(
eval_gt_image,
min_dim=800,
min_scale=0,
max_dim=1024,
mode='square')
# Substract channel-mean
input_image = input_image.astype(np.float32) - np.array([123.7, 116.8, 103.9])
image_shape_reformat = input_image.shape
# Add batch dimension
batch_images = np.expand_dims(input_image, 0)
# (batch, w, h, 3) -> (batch, 3, w, h)
batch_images = np.moveaxis(batch_images, -1, 1)
else:
window = batch_image_meta[0][7:11]
image_shape_reformat = batch_images[0].shape
# Make inference
host_input = batch_images.astype(dtype=np.float32, order='C')
cuda.memcpy_htod_async(device_input, host_input, stream)
context.execute_async(bindings=[int(device_input),
int(device_output1),
int(device_output2),
],
stream_handle=stream.handle)
cuda.memcpy_dtoh_async(host_output1, device_output1, stream)
cuda.memcpy_dtoh_async(host_output2, device_output2, stream)
stream.synchronize()
output_shape1 = engine.get_binding_shape(output_nodes[0])
output_shape2 = engine.get_binding_shape(output_nodes[1])
if graph_type == 'onnx':
trt_mrcnn_detection = host_output1.reshape(output_shape1).astype(dtype=np.float32)
trt_mrcnn_mask = host_output2.reshape(output_shape2).astype(dtype=np.float32)
elif graph_type == 'uff':
# (batch, 100, 6)
trt_mrcnn_detection = host_output1.reshape(
(engine.max_batch_size, *output_shape1)).astype(dtype=np.float32)
# (batch, 100, 2, 28, 28)
trt_mrcnn_mask = host_output2.reshape(
(engine.max_batch_size, *output_shape2)).astype(dtype=np.float32)
# (batch, 100, 2, 28, 28) -> (batch, 100, 28, 28, 2)
trt_mrcnn_mask = np.moveaxis(trt_mrcnn_mask, 2, -1)
else:
raise ValueError(f'Only onnx and uff graph types. Passed: {graph_type}')
# Extract bboxes, class_ids, scores and full-size masks
trt_boxes, trt_class_ids, trt_scores, trt_full_masks = \
utils.reformat_detections(detections=trt_mrcnn_detection[0],
mrcnn_mask=trt_mrcnn_mask[0],
original_image_shape=eval_gt_image.shape,
image_shape=image_shape_reformat,
window=window
)
return trt_boxes, trt_class_ids, trt_scores, trt_full_masks, eval_gt_boxes, eval_gt_class_ids, eval_gt_masks
def set_mrcnn_trt_engine(model_path, output_nodes=['mrcnn_detection', 'mrcnn_mask'], graph_type='onnx'):
"""
Load TensorRT engine via pycuda
Args:
model_path: model path to TensorRT-engine
output_nodes: output nodes names
graph_type: onnx or uff
Returns: python dict of attributes for pycuda model inference
"""
trt_logger = trt.Logger(trt.Logger.VERBOSE)
trt.init_libnvinfer_plugins(trt_logger, "")
with open(model_path, "rb") as f, trt.Runtime(trt_logger) as runtime:
engine = runtime.deserialize_cuda_engine(f.read())
context = engine.create_execution_context()
# Inputs
input_shape = engine.get_binding_shape('input_image')
input_size = trt.volume(input_shape) *\
engine.max_batch_size * np.dtype(np.float32).itemsize
device_input = cuda.mem_alloc(input_size)
# Outputs
output_names = list(engine)[1:]
# mrcnn_detection output
output_shape1 = engine.get_binding_shape(output_nodes[0])
host_output1 = cuda.pagelocked_empty(trt.volume(output_shape1) *
engine.max_batch_size,
dtype=np.float32)
device_output1 = cuda.mem_alloc(host_output1.nbytes)
# mrcnn_mask output
output_shape2 = engine.get_binding_shape(output_nodes[1])
host_output2 = cuda.pagelocked_empty(trt.volume(output_shape2) * engine.max_batch_size,
dtype=np.float32)
device_output2 = cuda.mem_alloc(host_output2.nbytes)
# Setting a cuda stream
stream = cuda.Stream()
return {'engine': engine,
'stream': stream,
'context': context,
'device_input': device_input,
'device_output1': device_output1,
'device_output2':device_output2,
'host_output1': host_output1,
'host_output2': host_output2,
'output_nodes': output_nodes,
'graph_type': graph_type
}
evaluate_mrcnn(model=set_mrcnn_trt_engine(
model_path=f"""../weights/maskrcnn_{CONFIG['backbone']}_512_512_3_trt_mod_fp32.engine"""),
inference_function=trt_mrcnn_inference,
eval_dataloader=eval_dataloader
)
evaluate_mrcnn(model=set_mrcnn_trt_engine(
model_path=f"""../weights/maskrcnn_{CONFIG['backbone']}_512_512_3_trt_mod_fp16.engine"""),
inference_function=trt_mrcnn_inference,
eval_dataloader=eval_dataloader
)
| src/notebooks/example_evaluate_model_balloon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import re
# +
POLITICA = []
CIENCIA = []
DEPORTE = []
with open("./Ciencia.txt","r") as RF:
CIENCIA.append([])
for l in RF:
if l == "###\n": #Es un nuevo documento, abrimos una nueva lista
CIENCIA.append([])
continue
if l != "\n":
CIENCIA[-1].append(l[:-1])
#CIENCIA_WORDS = [w.lower() for w in np.concatenate([np.concatenate([l.split(" ") for l in] CIENCIA[0]]))]
print "Cargados ",len(CIENCIA)," documentos de ciencia"
with open("./Deportes.txt","r") as RF:
DEPORTE.append([])
for l in RF:
if l == "###\n": #Es un nuevo documento, abrimos una nueva lista
DEPORTE.append([])
continue
if l != "\n":
DEPORTE[-1].append(l[:-1])
print "Cargados ",len(DEPORTE)," documentos de deporte"
with open("./Politica.txt","r") as RF:
POLITICA.append([])
for l in RF:
if l == "###\n": #Es un nuevo documento, abrimos una nueva lista
POLITICA.append([])
continue
if l != "\n":
POLITICA[-1].append(l[:-1])
print "Cargados ",len(POLITICA)," documentos de politica"
# -
def preprocess(Docs):
WOR = [np.concatenate([[w.lower() for w in l.split(" ")] for l in d]) for d in Docs]
for w in xrange(len(WOR)):
W = WOR[w]
W = filter(lambda w : len(w) != 0, W)
W = map(lambda w : re.sub(r'[\.\,-]',"",w) , W)
W = filter(lambda w : len(w) != 0, W)
W = map(lambda w : re.sub(r'[\"\'><\\\/\)\(]',"",w) , W)
W = filter(lambda w : len(w) != 0, W)
W = map(lambda w : re.sub(r'\d',"",w) , W)
W = filter(lambda w : len(w) != 0, W)
WOR[w] = W
return WOR
POLITICA_W = preprocess(POLITICA)
CIENCIA_W = preprocess(DEPORTE)
DEPORTE_W = preprocess(CIENCIA)
TOT_D = POLITICA_W + CIENCIA_W + DEPORTE_W
TOT_W = list(set(np.concatenate(TOT_D)))
ITF_W = [np.log(len(TOT_D)*1.0/len(filter(lambda d :w in d , TOT_D))) for w in TOT_W]
TOT_WU=np.array(TOT_W)[np.where(np.array(ITF_W)> 2.0)[0]]
POLW = []
count = 0
for d in TOT_D:
print count
count+=1
#POLW.append([])
POLW.append([(len(np.where(np.asanyarray(d) == w)[0]) *1.0/ len(d)) * np.log(len(TOT_D)*1.0/len(filter(lambda d :w in d,TOT_D))) for w in TOT_WU])
#for w in TOT_W:
# ft = len(np.where(np.asanyarray(d) == w)[0]) *1.0/ len(d)
# itf = len(filter(lambda d :w in d,TOT_D))
# #for d in TOT_D:
# # if w in d:
# # itf+=1
# itf = np.log(len(TOT_D)*1.0/itf)
# POLW[-1].append(ft*itf)
A = TOT_D[0]
len(filter(lambda d :"rajoy" in d,TOT_D))
np.log(1)
[len(filter(lambda w : w != 0.0, l)) for l in POLW]
np.exp(3.5)
| DocumentRetrival.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Tensorflow Sessions statically run Tensorflow Graphs.
#
# If you have some Tensorflow Graph, whether you downloaded or created it from scratch using the tensorflow’s **Graph API**. You will need to use the **Session API** to run it.
#
# We will take the following graph as a simple example. When we call `sess.run()` on `tf.Tensor` types.
#
# `g1.get_operation_by_name` will simply return the `tf.Operation` object. We can get an array of the `tf.Tensor` objects that this operation produces with the `.outputs` attribute. **Remember, it is an important distinction to make that `tf.Tensor` and `tf.Operation` are different**! `tf.Operation` is a node `tf.Tensor` is an edge.
#
#
# +
import tensorflow as tf
g1 = tf.Graph()
with g1.as_default():
my_input = tf.constant([-1,0,1], dtype=tf.float16, name="input")
a = tf.square(my_input, name="A")
b = tf.cos(a, name="B")
c = tf.sin(a, name="C")
d = tf.add(b, c, name="D")
e = tf.floor(b, name="E")
f = tf.sqrt(d, name="F")
sess = tf.Session(graph=g1);
print("A:{}".format(sess.run(g1.get_operation_by_name("A").outputs)))
print("B:{}".format(sess.run(g1.get_operation_by_name("B").outputs)))
print("C:{}".format(sess.run(g1.get_operation_by_name("C").outputs)))
print("D:{}".format(sess.run(g1.get_operation_by_name("D").outputs)))
print("E:{}".format(sess.run(g1.get_operation_by_name("E").outputs)))
print("F:{}".format(sess.run(g1.get_operation_by_name("F").outputs)))
# -
# The way we run the graph above is actually very inefficient. It is running 6 times. You can see this by [running a quick `tf.Print`](https://gist.github.com/Ouwen/66326b796311f0cca3c60067b9237124). `tf.Print` is a graph node that outputs values and messages to `stderr` from the Tensorflow C++ Runtime.
#
# <img src="../figures/yikes.png" width="400"/>
# *Yikes*
#
# We only want to run the graph once and retrieve all outputs. We can easily tweak our code so that we only make one `run` call. We will also notice that the graph is run just once instead of 6 times.
# +
import tensorflow as tf
g1 = tf.Graph()
with g1.as_default():
my_input = tf.constant([-1,0,1], dtype=tf.float16, name="input")
my_printed_input = tf.Print(my_input, [my_input], message="Running the graph", name="print")
a = tf.square(my_printed_input, name="A")
b = tf.cos(a, name="B")
c = tf.sin(a, name="C")
d = tf.add(b, c, name="D")
e = tf.floor(b, name="E")
f = tf.sqrt(d, name="F")
sess = tf.Session(graph=g1);
sess.run({
"A": g1.get_operation_by_name("A").outputs,
"B": g1.get_operation_by_name("B").outputs,
"C": g1.get_operation_by_name("C").outputs,
"D": g1.get_operation_by_name("D").outputs,
"E": g1.get_operation_by_name("E").outputs,
"F": g1.get_operation_by_name("F").outputs
})
# -
# We may want to add some more dynamic elements to our graph. This can be done with `tf.Variable` operations and `tf.Placeholder` operations.
#
# `tf.Placeholder` is a simple operation that takes in a value during the session runtime. Rather than having `my_input` be a constant we can instead use a placeholder.
#
# `tf.Variable` is a bit more interesting… We will just create a graph with one variable to our graph.
# +
import tensorflow as tf
from src import cloud_visualizer
g1 = tf.Graph()
with g1.as_default():
# Add a single variable to our graph
v = tf.get_variable(name="v", shape=(), initializer=tf.glorot_uniform_initializer())
sess = tf.Session(graph=g1)
sess.run(v.initializer) # Run just the initializer on our variable
# sess.run(tf.global_variables_initializer()) # This will initialize all variables
print(sess.run(v))
cloud_visualizer.show_graph(g1)
# -
# When we inspect what is added to our graph, we notice that `tf.Variable` is actually a group of many different operations: `tf.Identity`, `tf.Assign`, `tf.VariableV2` and more operations within the the `Initializer`. **These exist to help `tf.Variable` store state.**
#
# When the variable is first willed into existence, it has no value. This is why when you begin a session you must first initialize your variables, and why there is an `Initializer` is attached to the `tf.Variable` class (in our example we use a random distribution. The snippet above can be run and `sess.run(v)` will print out a random number. However, try placing `sess.run(v)` before the variable is init, and you will receive an error.
#
# When using the python API to work with variables, there is under the hood syntax sugar that exists so that you are able to use the variables as though they are normal tensor outputs from a normal operation. A big difference to note is that when we save the graph as a protobuf, the `tf.Variable` group is saved; however the value stored in the `tf.Variable` is lost. In order to save this we must utilize `tf.train.Saver`.
# +
import tensorflow as tf
g1 = tf.Graph()
with g1.as_default():
v = tf.get_variable(name="v", shape=(), initializer=tf.glorot_uniform_initializer())
saver = tf.train.Saver()
sess = tf.Session(graph=g1);
sess.run(v.initializer)
sess.run(v.assign_add(1))
sess.run(v.assign_add(1))
sess.run(v.assign_add(1))
sess.run(v.assign_add(1))
saver.save(sess, "/tmp/model.ckpt")
# -
# *We perform some adds to our variable, then we run the saver. **Note** the files produced are actually checkpoint, model.ckpt.data-…, model.ckpt.index, and model.ckpt.meta. When we call /tmp/model.ckpt later, these files will need to exist so don’t move them around.*
# +
import tensorflow as tf
g1 = tf.Graph()
with g1.as_default():
v = tf.get_variable(name="v", shape=(), initializer=tf.glorot_uniform_initializer())
saver = tf.train.Saver()
sess = tf.Session(graph=g1);
saver.restore(sess, "/tmp/model.ckpt")
sess.run(v)
# -
# *We can run the saver to restore our session variable. Since the value is coming from our checkpoint file, no init is needed.*
# To put most simply, the `.ckpt` file is just a map of our variable names to the value that was stored during the session. If I created an entirely new graph, as long as I had the variable name `v` in my graph I would be able to restore the value in the checkpoint.
#
# In production these `.ckpt` files can be run multiple times to snapshot the progress of a Tensorflow graph that is running in a session.
#
# I hope this was a helpful short overview of the low level tensorflow API.
| notebooks/.ipynb_checkpoints/Tensorflow Sessions statically run Tensorflow Graphs.-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 12. Pandas
# pandas is great way to manipilate data
import pandas as pd
import numpy as np
# pandas has many diffrent read function
iris = pd.read_csv("Dataset/iris.data")
iris
print(type(iris))
df = iris.copy() # creating copy (non-linked) set
df.head()
print(df.shape)
print(df.dtypes)
# +
# change column headers
# -
df.columns # change this to new entry's
df.columns = ['sl','sw','pl','pw','flower_type'] # changed to new entries
print(df.dtypes)
df.columns
# describe -> it describe the data for us (shows valid value, it will not count invalid data i.e empty/NaN etc)
df.describe()
# +
# in above 25%,or 50% or 75% are percentile not percentage (we have 25 percentage heigh compare to other)
# -
# checking a particular column
df.sl
# or
df["sl"]
# some narrator
df.isnull()
df.isnull().sum() # how many null entries are present in each column
# +
# taking slice from set, access data from particulr set
# -
df.iloc[1:4,2:4]
# ### Manupliate Data In DataFrame
df.head()
# remove some row
df.drop(0)
df.head() # 0th row not removed
# drop by lables
df.drop(0,inplace = True) # by default inplace id false so turn it into True
df.head() # so it remove 0 ffrom all dataset do we need to rearrange the data
df.index
# drop by index
df.index[0],df.index[3]
df.drop(df.index[0],inplace = True) # remove 0th index that is lable 1
df.head()
df.sl > 5 # condtion checking
df[df.sl > 5] # codition crospondance value
df[df.flower_type == 'Iris-virginica'].describe() # v.imp data eg.
# add addition row in dataset
print(df.head())
print(df.iloc[0]) # doing this by index
print(df.loc[8]) # doing this by lable
# add addition row in dataset
df.loc[0] = [2.4,3.2,5.2,4.2,'Iris-setosa']
df.tail() # added at last
# or
df.loc[149] = [2.4,3.2,5.2,4.2,'Iris-setosa']
df.tail()
# reset index of dataset [v.imp****]
df.reset_index() # creating addition column of preivious index do we need to remove that
df.reset_index(drop = True,inplace = True)
df.index
# remove whole column at once
df.drop('sl',axis = 1,inplace = True)
# or -> del df['sl']
df.describe()
df.index
# how to add a column in datasets
# re-gaining prev datasets
df = iris.copy() # creating copy (non-linked) set
df.columns = ['sl','sw','pl','pw','flower_type']
df.describe()
# add col 'pl-pw', which is difference btw 'pl-pw'
df['pl-pw'] = df['pl'] - df['pw']
df.tail()
# ### Handling N/A entries
# making some N/A entries
df.iloc[2:4,1:3] = 0
df.head()
# exploration
df.iloc[2:4,1:3] = np.nan # amking N/A/N
df.head()
# two way to handle Nan
df.dropna(inplace = True)
df.head()
df.reset_index(drop = True, inplace = True)
df.head()
# 2nd way of handling
df.iloc[2:4,1:3] = np.nan # amking N/A/N
df.head()
# replace Nan with valid entries (filling with mean of sl)
df.sw.fillna(df.sw.mean(),inplace = True)
df.head()
# replace Nan with valid entries (filling with mean of pl)
df.pl.fillna(df.pl.mean(),inplace = True)
df.head()
# ### handling string data
# string based data
df['Gender'] = 'Female'
df.iloc[0:10,6] = 'Male'
df.head()
# +
def f(str):
if str == "Male":
return 0
else:
return 1
df["sex"] = df.Gender.apply(f)
df.head()
del df["Gender"]
df.sex
# -
# ## Assignment Solution
| 12. Pandas/.ipynb_checkpoints/main-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from os import path
from backbone.dihedral import DihedralMaker
import numpy as np
import MDAnalysis as mda
big_traj_folder = '/home/yizaochen/codes/dna_rna/all_systems'
backbone_data_folder = '/home/yizaochen/codes/dna_rna/backbone_data'
# ### Part 1: Initialize
host = 'a_tract_21mer'
strand_id = 'STRAND2'
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
# ### Part 2: Set Dihedral Name
dihedral_name = 'C4prime-P' # 'C2prime-P', 'C4prime-P'
# ### Part 3: Calculate Dihedral by PLUMED
# +
#d_agent.make_all_out(dihedral_name)
# -
# ### Batch Run
"""
hosts = ['a_tract_21mer', 'g_tract_21mer', 'atat_21mer', 'gcgc_21mer']
strand_lst = ['STRAND1', 'STRAND2']
for host in hosts:
for strand_id in strand_lst:
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
d_agent.make_all_out(dihedral_name)
"""
"""
hosts = ['a_tract_21mer', 'g_tract_21mer', 'atat_21mer', 'gcgc_21mer']
strand_lst = ['STRAND1', 'STRAND2']
dihedral_name_lst = ['C1prime-N3orO2', 'C1prime-N7orC5']
for host in hosts:
for strand_id in strand_lst:
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
for dihedral_name in dihedral_name_lst:
d_agent.make_all_out_with_resname(dihedral_name)
"""
"""
hosts = ['a_tract_21mer', 'g_tract_21mer', 'atat_21mer', 'gcgc_21mer']
strand_lst = ['STRAND1', 'STRAND2']
dihedral_name_lst = ['O4prime-C4orC2', 'C2prime-C4orC2']
for host in hosts:
for strand_id in strand_lst:
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
for dihedral_name in dihedral_name_lst:
d_agent.make_all_out_with_resname(dihedral_name)
"""
"""
hosts = ['a_tract_21mer', 'g_tract_21mer', 'atat_21mer', 'gcgc_21mer']
strand_lst = ['STRAND1', 'STRAND2']
dihedral_name_lst = ['O4prime-C8orC6', 'C2prime-C8orC6']
for host in hosts:
for strand_id in strand_lst:
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
for dihedral_name in dihedral_name_lst:
d_agent.make_all_out_with_resname(dihedral_name)
"""
"""
hosts = ['a_tract_21mer', 'g_tract_21mer', 'atat_21mer', 'gcgc_21mer']
strand_lst = ['STRAND1', 'STRAND2']
dihedral_name_lst = ['C3prime-O5prime']
for host in hosts:
for strand_id in strand_lst:
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
for dihedral_name in dihedral_name_lst:
d_agent.make_all_out(dihedral_name)
"""
hosts = ['a_tract_21mer', 'g_tract_21mer', 'atat_21mer', 'gcgc_21mer']
strand_lst = ['STRAND1', 'STRAND2']
dihedral_name_lst = ['O4prime-O5prime']
for host in hosts:
for strand_id in strand_lst:
d_agent = DihedralMaker(host, strand_id, big_traj_folder, backbone_data_folder)
for dihedral_name in dihedral_name_lst:
d_agent.make_all_out(dihedral_name)
| notebooks/calc_dihedral.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false}
# %matplotlib inline
# -
#
# The plasma dispersion function
# ==============================
#
# Let's import some basics (and `PlasmaPy`!)
#
# + jupyter={"outputs_hidden": false}
import matplotlib.pyplot as plt
import numpy as np
# + jupyter={"outputs_hidden": false}
import plasmapy.dispersion.dispersionfunction
help(plasmapy.dispersion.dispersionfunction.plasma_dispersion_func)
# + raw_mimetype="text/restructuredtext" active=""
# Take a look at the docs to :func:`~plasmapy.dispersion.dispersionfunction.plasma_dispersion_func` for more information on this.
# -
# We'll now make some sample data to visualize the dispersion function:
#
#
# + jupyter={"outputs_hidden": false}
x = np.linspace(-1, 1, 1000)
X, Y = np.meshgrid(x, x)
Z = X + 1j * Y
print(Z.shape)
# -
# Before we start plotting, let's make a visualization function first:
#
#
# + jupyter={"outputs_hidden": false}
def plot_complex(X, Y, Z, N=50):
fig, (real_axis, imag_axis) = plt.subplots(1, 2)
real_axis.contourf(X, Y, Z.real, N)
imag_axis.contourf(X, Y, Z.imag, N)
real_axis.set_title("Real values")
imag_axis.set_title("Imaginary values")
for ax in [real_axis, imag_axis]:
ax.set_xlabel("Real values")
ax.set_ylabel("Imaginary values")
fig.tight_layout()
plot_complex(X, Y, Z)
# -
# We can now apply our visualization function to our simple dispersion relation
#
#
# + jupyter={"outputs_hidden": false} tags=["nbsphinx-thumbnail"]
# sphinx_gallery_thumbnail_number = 2
F = plasmapy.dispersion.dispersionfunction.plasma_dispersion_func(Z)
plot_complex(X, Y, F)
# -
# So this is going to be a hack and I'm not 100% sure the dispersion function
# is quite what I think it is, but let's find the area where the dispersion
# function has a lesser than zero real part because I think it may be important
# (brb reading Fried and Conte):
#
#
# + jupyter={"outputs_hidden": false}
plot_complex(X, Y, F.real < 0)
# -
# We can also visualize the derivative:
#
#
# + jupyter={"outputs_hidden": false}
F = plasmapy.dispersion.dispersionfunction.plasma_dispersion_func_deriv(Z)
plot_complex(X, Y, F)
# -
# Plotting the same function on a larger area:
#
#
# + jupyter={"outputs_hidden": false}
x = np.linspace(-2, 2, 2000)
X, Y = np.meshgrid(x, x)
Z = X + 1j * Y
print(Z.shape)
# + jupyter={"outputs_hidden": false}
F = plasmapy.dispersion.dispersionfunction.plasma_dispersion_func(Z)
plot_complex(X, Y, F, 100)
# -
# Now we examine the derivative of the dispersion function as a function
# of the phase velocity of an electromagnetic wave propagating through
# the plasma. This is recreating figure 5.1 in:
# <NAME>, <NAME>, <NAME>, and <NAME>,
# Plasma scattering of electromagnetic radiation: theory and measurement
# techniques. Chapter 5 Pg 106 (Academic press, 2010).
#
#
# + jupyter={"outputs_hidden": false}
xs = np.linspace(0, 4, 100)
ws = (-1 / 2) * plasmapy.dispersion.dispersionfunction.plasma_dispersion_func_deriv(xs)
wRe = np.real(ws)
wIm = np.imag(ws)
plt.plot(xs, wRe, label="Re")
plt.plot(xs, wIm, label="Im")
plt.axis([0, 4, -0.3, 1])
plt.legend(
loc="upper right", frameon=False, labelspacing=0.001, fontsize=14, borderaxespad=0.1
)
plt.show()
| docs/notebooks/dispersion/dispersion_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow
import pandas as pd
import numpy as np
from sklearn.preprocessing import OneHotEncoder
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
# +
health_data = pd.read_csv('../merged_final_data_clean.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="ZlGgwLy3Ue53" outputId="c8445da5-b78c-41cb-a139-55849a01bba9"
print(health_data.columns)
# +
columns_to_encode = ['Tobacco smoking status NHIS',
"Stress is when someone feels tense nervous anxious or can't sleep at night because their mind is troubled. How stressed are you?",
"Are you a refugee?",
"In the past year have you or any family members you live with been unable to get any of the following when it was really needed?",
"Are you worried about losing your housing?",
"What is your main insurance?",
"In the past year have you spent more than 2 nights in a row in a jail prison detention center or juvenile correctional facility?",
"What is your current work situation?",
"Are you Hispanic or Latino?",
"Do you feel physically and emotionally safe where you currently live?",
"What is the highest level of school that you have finished?",
"Which race(s) are you?",
"In the past year have you been afraid of your partner or ex-partner?",
"Has lack of transportation kept you from medical appointments meetings work or from getting things needed for daily living?",
"What is your housing situation today?",
"Tobacco smoking status NHIS",
"At any point in the past 2 years has season or migrant farm work been your or your family's main source of income?",
"Have you been discharged from the armed forces of the United States?",
"Gender",
"Race",
"How often do you see or talk to people that you care about and feel close to (For example: talking to friends on the phone visiting friends or family going to church or club meetings)?"]
OHE = OneHotEncoder(sparse=False)
encoded_columns = OHE.fit_transform(health_data[columns_to_encode])
# -
print(encoded_columns.shape)
only_scalar = health_data.drop(columns_to_encode, axis=1).values
print(only_scalar)
# +
only_scalar_filtered =only_scalar[:,2:-3] #filter time and patient AND DIASTALIC AND SYSTOLIC
labels =only_scalar[:,-1]
labels = np.expand_dims(labels, axis=-1)
only_scalar_filtered = np.concatenate([only_scalar_filtered, labels], axis=1)
print(only_scalar_filtered)
# -
processed_data = np.concatenate([only_scalar_filtered, encoded_columns], axis=1)
print(processed_data.shape)
# + [markdown] id="kDSzK674s-4k"
# Shuffle the dataset.
# + id="dZiwd2-oVirj"
from sklearn.utils import shuffle
processed_data = shuffle(processed_data)
# -
X = processed_data[:,:-1].astype(float)
y = processed_data[:,-1].astype(int)
# + [markdown] id="QsoPOklKlxMl"
# Split the data into train and test set. 18% of it preserved for test set, and remaining 72% for train set.
# + id="UloEuNuXXe8-"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1)
# -
X_train[0].shape[0]
print(X_train.dtype)
print(y_train)
# + [markdown] id="wDi9jfnRqUgJ"
# Fully connected neural network model with 3 hidden layers:
# + id="bVz5-8xuqEsy"
import tensorflow as tf
model = tf.keras.models.Sequential([
tf.keras.Input(shape=(X_train[0].shape[0],)), #Xtrain[0].shape[0] = 104
tf.keras.layers.Dense(15, activation='relu'), #represents 1st hidden layer
tf.keras.layers.Dropout(0.2), # dropout regularization with dropout probability 20% percent
tf.keras.layers.Dense(7, activation='relu'), #represents the 2nd hidden layer
tf.keras.layers.Dropout(0.2), # dropout regularization with dropout probability 20% percent
tf.keras.layers.Dense(4, activation='relu'), #represents the 3rd hidden layer
tf.keras.layers.Dropout(0.2), # dropout regularization with dropout probability 20% percent
tf.keras.layers.Dense(1, activation='sigmoid') # sigmoid activation at output since it is binary classification
])
# + id="5FNnHp_QrGCv"
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
# + [markdown] id="aS2gk-MarTGl"
# Train the model with 10 epochs:
# + colab={"base_uri": "https://localhost:8080/"} id="HxyIThDGP_gj" outputId="360b23ea-a569-412d-a2da-90a2a7eec26a"
r2 = model.fit(X_train, y_train, epochs=150)
# + [markdown] id="cdbGFMKmrPsU"
# Plot the losses:
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="eFCAh_9Srkc9" outputId="2b1d48ad-5d05-445f-be6e-53c6c69e5da3"
plt.plot(r2.history['loss'], label='loss')
#plt.plot(r2.history['val_loss'], label='val_loss')
plt.legend()
# + [markdown] id="--olrolUrRdx"
# Plot the accuracies:
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="LL2qwSSvrmWy" outputId="c2d1311e-7c43-4832-de14-f4b2e280520d"
plt.plot(r2.history['accuracy'], label='acc')
#plt.plot(r2.history['val_accuracy'], label='val_acc')
plt.legend()
# + colab={"base_uri": "https://localhost:8080/"} id="uzNf6IUxro9G" outputId="68d8f4a1-10dd-4ab8-e5cc-23761d184b0d"
print(model.evaluate(X_test, y_test))
# -
model.save_weights('./weights/model_full/')
# + [markdown] id="oEjqnSkkKwkP"
# **Applying this dataset on different models to create benchmarks to compare with Fully Connected model:**
# + [markdown] id="Up6ggCZ6K7_h"
# *Logistic Regression* **(linear classifier):**
# + colab={"base_uri": "https://localhost:8080/"} id="LbK0ykIJLls0" outputId="7bbf0310-2368-45e4-d576-e39c9f440b6d"
from sklearn.linear_model import LogisticRegression
model =LogisticRegression()
model.fit(X_train, y_train)
print(model.score(X_train, y_train))
print(model.score(X_test,y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="cLvXlm3Eis1R" outputId="b1cbb427-a4bd-4d82-abb2-dc382eb439d8"
for i in range(X_train[0].shape[0]):
print("weight for the feature",i,":",model.coef_[0][i])
model.coef_[0].sort()
print("sorted list, positive weights contribute MORE TO DETERMINE potential hypertension case and negative features conribute more to make prediction as NOT hypertension. So we can say 0 valued weights do not contribute to anything:",model.coef_[0])
# + [markdown] id="F2cS87lMLDDT"
# *Decision Tree*
# + colab={"base_uri": "https://localhost:8080/"} id="BLXjjRAFLmGd" outputId="383a288e-c694-4d7b-c7fc-dbfb311ce006"
from sklearn.tree import DecisionTreeClassifier
model =DecisionTreeClassifier()
model.fit(X_train, y_train)
print(model.score(X_train, y_train))
print(model.score(X_test,y_test))
# + [markdown] id="KUe9EcWyLFhl"
# *AdaBoost*
# + colab={"base_uri": "https://localhost:8080/"} id="PEk2Fd8eLmhY" outputId="6e6071e3-e2ec-4315-ffa5-1f741b628138"
from sklearn.ensemble import AdaBoostClassifier
model =AdaBoostClassifier()
model.fit(X_train, y_train)
print(model.score(X_train, y_train))
print(model.score(X_test,y_test))
# +
# Visualize the data (tsne is great but slow.)
from sklearn.manifold import TSNE
tsne = TSNE(n_components=2)
transformed = tsne.fit_transform(X_train)
#visualize in the 2d
plt.scatter(transformed[:,0], transformed[:,1], c=y_train) #take first two columns in order to get 2d plot.
plt.show()
# +
transformed = tsne.fit_transform(X_test)
# visualize the clouds in 2-D
plt.scatter(transformed[:,0], transformed[:,1], c=y_test) #take first two columns in order to get 2d plot.
plt.show()
| utils/notebooks/model_systolic_diastolic_excluded.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Standard problem 3
#
# ## Problem specification
#
# This problem is to calculate a single domain limit of a cubic magnetic particle. This is the size $L$ of equal energy for the so-called flower state (which one may also call a splayed state or a modified single-domain state) on the one hand, and the vortex or curling state on the other hand.
#
# Geometry:
#
# A cube with edge length, $L$, expressed in units of the intrinsic length scale, $l_\text{ex} = \sqrt{A/K_\text{m}}$, where $K_\text{m}$ is a magnetostatic energy density, $K_\text{m} = \frac{1}{2}\mu_{0}M_\text{s}^{2}$.
#
# Material parameters:
#
# - uniaxial anisotropy $K_\text{u}$ with $K_\text{u} = 0.1 K_\text{m}$, and with the easy axis directed parallel to a principal axis of the cube (0, 0, 1),
# - exchange energy constant is $A = \frac{1}{2}\mu_{0}M_\text{s}^{2}l_\text{ex}^{2}$.
#
# More details about the standard problem 3 can be found in Ref. 1.
#
# ## Simulation
#
# Firstly, we import all necessary modules.
import discretisedfield as df
import micromagneticmodel as mm
import oommfc as oc
# The following two functions are used for initialising the system's magnetisation [1].
# +
import numpy as np
# Function for initiaising the flower state.
def m_init_flower(pos):
x, y, z = pos[0]/1e-9, pos[1]/1e-9, pos[2]/1e-9
mx = 0
my = 2*z - 1
mz = -2*y + 1
norm_squared = mx**2 + my**2 + mz**2
if norm_squared <= 0.05:
return (1, 0, 0)
else:
return (mx, my, mz)
# Function for initialising the vortex state.
def m_init_vortex(pos):
x, y, z = pos[0]/1e-9, pos[1]/1e-9, pos[2]/1e-9
mx = 0
my = np.sin(np.pi/2 * (x-0.5))
mz = np.cos(np.pi/2 * (x-0.5))
return (mx, my, mz)
# -
# The following function is used for convenience. It takes two arguments:
#
# - $L$ - the cube edge length in units of $l_\text{ex}$, and
# - the function for initialising the system's magnetisation.
#
# It returns the relaxed system object.
#
# Please refer to other tutorials for more details on how to create system objects and drive them using specific drivers.
def minimise_system_energy(L, m_init):
print("L={:7}, {} ".format(L, m_init.__name__), end="")
N = 16 # discretisation in one dimension
cubesize = 100e-9 # cube edge length (m)
cellsize = cubesize/N # discretisation in all three dimensions.
lex = cubesize/L # exchange length.
Km = 1e6 # magnetostatic energy density (J/m**3)
Ms = np.sqrt(2*Km/mm.consts.mu0) # magnetisation saturation (A/m)
A = 0.5 * mm.consts.mu0 * Ms**2 * lex**2 # exchange energy constant
K = 0.1*Km # Uniaxial anisotropy constant
u = (0, 0, 1) # Uniaxial anisotropy easy-axis
p1 = (0, 0, 0) # Minimum sample coordinate.
p2 = (cubesize, cubesize, cubesize) # Maximum sample coordinate.
cell = (cellsize, cellsize, cellsize) # Discretisation.
mesh = df.Mesh(p1=(0, 0, 0), p2=(cubesize, cubesize, cubesize),
cell=(cellsize, cellsize, cellsize)) # Create a mesh object.
system = mm.System(name='stdprob3')
system.energy = mm.Exchange(A=A) + mm.UniaxialAnisotropy(K=K, u=u) + mm.Demag()
system.m = df.Field(mesh, dim=3, value=m_init, norm=Ms)
md = oc.MinDriver()
md.drive(system, overwrite=True)
return system
# ### Relaxed magnetisation states
#
# Now, we show the magnetisation configurations of two relaxed states.
#
# **Vortex** state:
# NBVAL_IGNORE_OUTPUT
system = minimise_system_energy(8, m_init_vortex)
system.m.plane('y').mpl()
# **Flower** state:
# NBVAL_IGNORE_OUTPUT
system = minimise_system_energy(8, m_init_flower)
system.m.plane('y').mpl()
# ### Energy crossing
#
# We can plot the energies of both vortex and flower states as a function of cube edge length $L$. This will give us an idea where the state transition occurrs. We can achieve that by simply looping over the edge lengths $L$ of interest, computing the energy of both vortex and flower states, and finally, plotting the energy dependence.
# +
# NBVAL_IGNORE_OUTPUT
L_array = np.linspace(8, 9, 5)
vortex_energies, flower_energies = [], []
for L in L_array:
vortex = minimise_system_energy(L, m_init_vortex)
flower = minimise_system_energy(L, m_init_flower)
vortex_energies.append(vortex.table.data.tail(1)['E'][0])
flower_energies.append(flower.table.data.tail(1)['E'][0])
import matplotlib.pyplot as plt
plt.figure(figsize=(8, 4))
plt.plot(L_array, vortex_energies, 'o-', label='vortex')
plt.plot(L_array, flower_energies, 'o-', label='flower')
plt.xlabel('L (lex)')
plt.ylabel('E (J)')
plt.grid()
plt.legend();
# -
# From the plot, we can see that the energy crossing occurrs between $8.4l_\text{ex}$ and $8.6l_\text{ex}$, so we can employ a root-finding (e.g. bisection) algorithm to find the exact crossing.
# +
# NBVAL_IGNORE_OUTPUT
from scipy.optimize import bisect
def energy_difference(L):
vortex = minimise_system_energy(L, m_init_vortex)
flower = minimise_system_energy(L, m_init_flower)
return (vortex.table.data.tail(1)['E'][0] -
flower.table.data.tail(1)['E'][0])
cross_section = bisect(energy_difference, 8.4, 8.6, xtol=0.02)
print(f'\nThe energy crossing occurs at {cross_section}*lex')
# -
# ## References
#
# [1] µMAG Site Directory http://www.ctcms.nist.gov/~rdm/mumag.org.html
| docs/07-tutorial-standard-problem3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
def fit(x_train,y_train):
res = {}
classes = set(y_train)
res["total_data"] = len(y_train)
for curr_class in classes:
res[curr_class]= {}
num_features = x_train.shape[1]
curr_class_rows = (y_train==curr_class)
x_train_curr = x_train[curr_class_rows]
y_train_curr = y_train[curr_class_rows]
res[curr_class]["total_count"] =len(y_train_curr)
# res['total_data']+=len(y_train_curr)
for j in range(1,num_features+1):
res[curr_class][j] = {}
values = set(x_train[:,j-1])
for curr_val in values:
res[curr_class][j][curr_val] = (x_train_curr[:,j-1]==curr_val).sum()
return res
def probability(curr_class,dictionary,x):
num_features = len(dictionary[curr_class].keys()) -1
prob = np.log(dictionary[curr_class]['total_count'])- np.log(dictionary['total_data'])
for j in range(1,num_features +1):
prob += np.log(dictionary[curr_class][j][x[j-1]] + 1) - np.log(dictionary[curr_class]["total_count"]+ len(dictionary[curr_class][j].keys()))
return prob
def predictSinglePoint(x,dictionary):
max_prob = 0
max_class = 0
first_run= True
total_classes = dictionary.keys()
for curr_class in total_classes:
if curr_class== 'total_data':
continue
prob = probability(curr_class,dictionary,x)
if(first_run or prob>max_prob):
max_prob = prob
max_class = curr_class
first_run = False
return max_class
def predict(x_test,dictionary):
y_test = np.zeros(len(x_test),int)
for i in range(len(x_test)):
y_test[i] = predictSinglePoint(x_test[i],dictionary)
return y_test
def make_labelled(column):
second_limit = column.mean()
first_limit = (1/2) * second_limit
third_limit = 1.5 * second_limit
for i in range(len(column)):
if column[i]<first_limit:
column[i] = 0
elif column[i]<second_limit:
column[i] = 1
elif column[i]<third_limit:
column[i] = 2
else:
column[i] = 3
return column
# +
from sklearn import datasets
iris = datasets.load_iris()
x = iris.data
y = iris.target
for i in range(x.shape[-1]):
x[:,i] = make_labelled(x[:,i])
from sklearn import model_selection
x_train,x_test,y_train,y_test = model_selection.train_test_split(x,y,random_state = 0,test_size = 0.32)
dictionary = fit(x_train,y_train)
y_pred = predict(x_test,dictionary)
dictionary['total_data']
# print(dictionary)
print(y_pred)
# -
from sklearn.metrics import classification_report , confusion_matrix
print(classification_report(y_test,y_pred))
print(confusion_matrix(y_test,y_pred))
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
clf.fit(x_train,y_train)
y_pred = clf.predict(x_test)
print(classification_report(y_test,y_pred))
print(confusion_matrix(y_test,y_pred))
| naive bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [nlu]
# language: python
# name: Python [nlu]
# ---
# # CS 224U Jupyter Notebook Tutorial
#
# Lucy will demo this notebook during the Week 1 Python special session, but this notebook should also be workable on your own if you decide to do so.
#
# The contents of this notebook were tested on a MacBook computer. Feel free to stop by office hours w/ questions or ask on Piazza if you run into problems.
# ## Table of Contents
#
# 1. [Starting up](#start)
# 2. [Cells](#cells)
# 1. [Code](#code)
# 2. [Markdown](#markdown)
# 3. [Kernels](#kernels)
# 4. [Shortcuts](#shortcuts)
# 5. [Shutdown](#shutdown)
# 6. [Extra](#noness)
# 7. [More resources](#res)
#
# ## Starting up <a id="start"></a>
#
# This tutorial assumes that you have followed the [course setup](https://nbviewer.jupyter.org/github/cgpotts/cs224u/blob/master/setup.ipynb) instructions. This means Jupyter is installed using Conda.
#
# 1. Open up Terminal (Mac/Linux) or Command Prompt (Windows).
# 2. Enter a directory that you'd like to have as your `Home`, e.g. where your cloned `cs224u` Github repo resides.
# 3. Type `jupyter notebook` and enter. After a few moments, a new browser window should open, listing the contents of your `Home` directory.
# - Note that on your screen, you'll see something like `[I 17:23:47.479 NotebookApp] The Jupyter Notebook is running at: http://localhost:8888/`. This tells you where your notebook is located. So if you were to accidentally close the window, you can open it again while your server is running. For this example, navigating to `http://localhost:8888/` on your favorite web browser should open it up again.
# - You may also specify a port number, e.g. `jupyter notebook --port 5656`. In this case, `http://localhost:5656/` is where your directory resides.
# 4. Click on a notebook with `.ipynb` extension to open it. If you want to create a new notebook, in the top right corner, click on `New` and under `Notebooks`, click on `Python`. If you have multiple environments, you should choose the one you want, e.g. `Python [nlu]`.
# - You can rename your notebook by clicking on its name (originally "Untitled") at the top of the notebook and modifying it.
# - Files with `.ipynb` are formatted as a JSON and so if you open them in vim, emacs, or a code editor, it's much harder to read and edit.
#
# Jupyter Notebooks allow for **interactive computing**.
# ## Cells <a id="cells"></a>
#
# Cells help you organize your work into manageable chunks.
#
# The top of your notebook contains a row of buttons. If you hover over them, the tooltips explain what each one is for: saving, inserting a new cell, cut/copy/paste cells, moving cells up/down, running/stopping a cell, choosing cell types, etc. Under Edit, Insert, and Cell in the toolbar, there are more cell-related options.
#
# Notice how the bar on the left of the cell changes color depending on whether you're in edit mode or command mode. This is useful for knowing when certain keyboard shortcuts apply (discussed later).
#
# There are three main types of cells: **code**, **markdown**, and raw.
#
# Raw cells are less common than the other two, and you don't need to understand it to get going for cs224u. If you put anything in this type of cell, you can't run it. They are used for situations where you might want to convert your notebook to HTML or LaTeX using the `nbconvert` tool or File -> Download as a format that isn't `.ipynb`. Read more about raw cells [here](https://nbsphinx.readthedocs.io/en/0.4.2/raw-cells.html) if you're curious.
#
# ### Code <a id="code"></a>
#
# Use the following code cells to explore various operations.
#
# Typically it's good practice to put import statements in the first cell or at least in their own cell.
#
# The square brackets next to the cell indicate the order in which you run cells. If there is an asterisk, it means the cell is currently running.
#
# The output of a cell is usually any print statements in the cell and the value of the last line in the cell.
import time
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
print("cats")
# run this cell and notice how both strings appear as outputs
"cheese"
# cut/copy and paste this cell
# move this cell up and down
# run this cell
# toggle the output
# toggle scrolling to make long output smaller
# clear the output
for i in range(100):
print("cats")
# run this cell and stop before it finishes
# stop acts like a KeyboardInterrupt
for i in range(100):
time.sleep(1) # make loop run slowly
print("cats")
# +
# running this cell leads to no output
def function1():
print("dogs")
# put cursor in front of this comment and split and merge this cell.
def function2():
print("cheese")
# -
function1()
function2()
# One difference between coding a Python script and a notebook is how you can run code "out of order" for the latter. This means you should be careful about variable reuse. It is good practice to order cells in the order which you expect someone to use the notebook, and organize code in ways that prevent problems from happening.
#
# Clearing the output doesn't remove the old variable value. In the example below, we need to rerun cell A to start with a new `a`. If we don't keep track of how many times we've run cell B or cell C, we might encounter unexpected bugs.
# Cell A
a = []
# Cell B
# try running this cell multiple times to add more pineapple
a.append('pineapple')
# Cell C
# try running this cell multiple times to add more cake
a.append('cake')
# depending on the number of times you ran
# cells B and C, the output of this cell will
# be different.
a
# Even deleting cell D's code after running it doesn't remove list `b` from this notebook. This means if you are modifying code, whatever outputs you had from old code may still remain in the background of your notebook.
# Cell D
# run this cell, delete/erase it, and run the empty cell
b = ['apple pie']
# b still exists after cell C is gone
b
# Restart the kernel (Kernel -> Restart & Clear Output) to start anew. To check that things run okay in its intended order, restart and run everything (Kernel -> Restart & Run All). This is especially good to do before sharing your notebook with someone else.
#
# Jupyter notebooks are handy for telling stories using your code. You can view dataframes and plots directly under each code cell.
# dataframe example
d = {'ingredient': ['flour', 'sugar'], '# of cups': [3, 4], 'purchase date': ['April 1', 'April 4']}
df = pd.DataFrame(data=d)
df
# plot example
plt.title("pineapple locations")
plt.ylabel('latitude')
plt.xlabel('longitude')
_ = plt.scatter(np.random.randn(5), np.random.randn(5))
# ### Markdown <a id="markdown"></a>
#
# The other type of cell is Markdown, which allows you to write blocks of text in your notebook. Double click on any Markdown cell to view/edit it. Don't worry if you don't remember all of these things right away. You'll write more code than Markdown essays for cs224u, but the following are handy things to be aware of.
#
# You may notice that this cell's header is prefixed with `###`. The fewer hashtags, the larger the header. You can go up to five hashtags for the smallest level header.
#
# Here is a table. You can emphasize text using underscores or asterisks. You can also include links.
#
# | Markdown | Outcome |
# | ----------------------------- | ---------------------------- |
# | `_italics_ or *italics*` | _italics_ or *italics* |
# | `__bold__ or **bold**` | __bold__ or **bold** |
# | `[link](http://web.stanford.edu/class/cs224u/)` | [link](http://web.stanford.edu/class/cs224u/) |
# | `[jump to Cells section](#cells)` | [jump to Cells section](#cells) |
#
# Try removing/adding the `python` in the code formatting below to toggle code coloring.
#
# ```python
# if text == code:
# print("You can write code between a pair of triple backquotes, e.g. ```long text``` or `short text`")
# ```
#
# Latex also works:
# $y = \int_0^1 2x dx$
# $$y = x^2 + x^3$$
# > You can also format quotes by putting a ">" in front of each line.
# >
# > You can space your lines apart with ">" followed by no text.
# There are three different ways to write a bullet list (asterisk, dash, plus):
# * sugar
# * tea
# * earl gray
# * english breakfast
# - cats
# - persian
# - dogs
# + pineapple
# + apple
# + granny smith
#
# Example of a numbered list:
# 1. tokens
# 2. vectors
# 3. relations
#
# You can also insert images:
#
# ``
#
# (Try removing the backquotes and look at what happens.)
#
# A line of dashes, e.g. `----------------`, becomes a divider.
#
# ------------------
# ## Kernels <a id="kernels"></a>
#
# A kernel executes code in a notebook.
#
# You may have multiple conda environments on your computer. You can change which environment your notebook is using by going to Kernel -> Change kernel.
#
# When you open a notebook, you may get a message that looks something like "Kernel not found. I couldn't find a kernel matching ____. Please select a kernel." This just means you need to choose the version of Python or environment that you want to have for your notebook.
#
# If you have difficulty getting your conda environment to show up as a kernel, [this](https://stackoverflow.com/questions/39604271/conda-environments-not-showing-up-in-jupyter-notebook) may help.
#
# In our class we will be using IPython notebooks, which means the code cells run Python.
#
# Fun fact: there are also kernels for other languages, e.g. Julia. This means you can create notebooks in these other languages as well, if you have them on your computer.
# ## Shortcuts <a id="shortcuts"></a>
#
# Go to Help -> Keyboard Shortcuts to view the shortcuts you may use in Jupyter Notebook.
#
# Here are a few that I find useful on a regular basis:
# - **run** a cell, select below: shift + enter
# - **save** and checkpoint: command + S (just like other file types)
# - enter **edit** mode from command mode: press enter
# - enter **command** mode from edit mode: esc
# - **delete** a cell (command mode): select a cell and press D
# - **dedent** while editing: command + [
# - **indent** while editing: command + ]
# play around with this cell with shortcuts
# delete this cell
# Edit -> Undo Delete Cells
for i in range(10):
print("jelly beans")
# ## Shutdown <a id="shutdown"></a>
#
# Notice that when you are done working and exit out of this notebook's window, the notebook icon in the home directory listing next to this notebook is green. This means your kernel is still running. If you want to shut it down, check the box next to your notebook in the directory and click "Shutdown."
#
# To shutdown the jupyter notebook app as a whole, use Control-C in Terminal to stop the server and shut down all kernels.
# ## Extra <a id="noness"></a>
# + [markdown] slideshow={"slide_type": "slide"}
# These are some extra things that aren't top priority to know but may be interesting.
#
# **Checkpoints**
#
# When you create a notebook, a checkpoint file is also saved in a hidden directory called `.ipynb_checkpoints`. Every time you manually save the notebook, the checkpoint file updates. Jupyter autosaves your work on occasion, which only updates the `.ipynb` file but not the checkpoint. You can revert back to the latest checkpoint using File -> Revert to Checkpoint.
#
# **NbViewer**
#
# We use this in our class for viewing jupyter notebooks from our course website. It allows you to render notebooks on the Internet. Check it out [here](https://nbviewer.jupyter.org/).
#
# View -> **Cell toolbar**
# - **Edit Metadata**: Modify the metadata of a cell by editing its json representation. Example of metadata: whether cell output should be collapsed, whether it should be scrolled, deletability of cell, name, and tags.
# - **Slideshow**: For turning your notebook into a presentation. This means different cells fall under slide types, e.g. Notes, Skip, Slide.
#
# -
# ## More resources <a id="res"></a>
#
# If you click on "Help" in the toolbar, there is a list of references for common Python tools, e.g. numpy, pandas.
#
# [IPython website](https://ipython.org/)
#
# [Markdown basics](https://daringfireball.net/projects/markdown/)
#
# [Jupyter Notebook Documentation](https://jupyter-notebook.readthedocs.io/en/stable/index.html)
#
# [Real Python Jupyter Tutorial](https://realpython.com/jupyter-notebook-introduction/)
#
# [Dataquest Jupyter Notebook Tutorial](https://www.dataquest.io/blog/jupyter-notebook-tutorial/)
#
# [Stack Overflow](https://stackoverflow.com/)
| jupyter_notebook_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pydicom as dicom
import numpy as np
import matplotlib.pyplot as plt
import os
import glob
path = 'D:\\ILBS_14_8_21_Sanjeev\\Ilbs_14-08-2021\\'
pid = []
fpath = []
for fname in os.listdir(path):
if fname.startswith('S'):
id_ = int(fname.split('_')[1])
pid.append(id_)
for fname_ins in os.listdir(path+fname):
fpath.append(path+fname+'\\'+fname_ins+'\\DICOMOBJ')
slice_list = []
for files_ in fpath:
ct_images = os.listdir(files_)
print(files_)
slices = [dicom.read_file(files_+'/'+s, force=True) for s in ct_images]
pixel_spacing = slices[0].PixelSpacing
slice_thickness = slices[0].SliceThickness
axial_aspect_ratio = pixel_spacing[1]/pixel_spacing[0]
image_shape = list(slices[0].pixel_array.shape)
image_shape.append(len(slices))
volume3d = np.zeros(image_shape)
for i,s in enumerate(slices):
array2d = s.pixel_array
if i==498 or i==499:
pass
else:
#print("Index",i,array2d.shape)
volume3d[:,:,i] = array2d
axial = plt.subplot(2,2,1)
plt.title('Axial')
plt.imshow(volume3d[:,:,image_shape[2]//2])
axial.set_aspect(axial_aspect_ratio)
break
# +
plt.plot(np.array(volume3d.reshape(1,-1)))
# -
| All img test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''pandas-tutorial'': conda)'
# name: python3
# ---
# # 07 - Data Visualization
# ## Import Package
# +
# Import package as pandas does not ship along as native package
# use pip install pandas if not installed yet, or follow setup guide
import pandas as pd
import matplotlib.pyplot as plt
# -
# Use Pokemon dataset
dataframe = pd.read_csv("https://raw.githubusercontent.com/KianYang-Lee/pandas-tutorial/main/datasets/Pokemon.csv")
# ## Practice Section
# Create the following visualizations, complete with styling that provides context to readers:
# - Bar chart of Type 2 Pokemon count
# - Line chart of Pokemon count for each generation
# - Boxplot for Pokemon Speed distribution
# - Bar chart of Type 2 Pokemon count
dataframe["Type 2"].value_counts().plot(
kind="bar",
title="Type 2 Pokemon Count",
xlabel="Secondary type (Type 2) for Pokemon",
ylabel="Count",
)
plt.show()
# - Line chart of Pokemon count for each generation
pokemon_count_by_gen = dataframe["Generation"].value_counts()
pokemon_count_by_gen.index = dataframe["Generation"].value_counts().index.map(lambda x: str(x)+'_Gen')
pokemon_count_by_gen.sort_values(ascending=False).plot(
kind="line",
title="Number of Pokemon by Generation",
xlabel="Pokemon Generation",
ylabel="Number of Pokemon",
)
plt.show()
# - Boxplot for Pokemon Speed distribution
dataframe.Speed.plot(
kind="box",
title="Pokemon Speed Distribution",
xlabel="Speed",
ylabel="Value",
)
plt.show()
# **Copyright (C) 2021 <NAME>**
#
# This program is licensed under MIT license.
| solutions/07_data_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 前言
# 这几年机器学习这种从经验学习的软件技术重现光明。在计算机诞生的早期,机器学习的概念已经出现,各种理论天马行空,限于计算成本而未能普及。随着计算设备的普及,日常生活中越来越多的机器学习应用,可以说它的成功开始变得习以为常。新应用如雨后春笋一般出现,很多都从机器学习中获得动力。
#
# <!-- TEASER_END-->
# 在这本书里,我们将看到一些机器学习的模型和算法。我们会介绍一些常用的机器学习任务和模型的效果评估方法。而这些模型和算法都是通过十分流行的Python机器学习库scikit-learn来完成,里面有许多机器学习的模型和算法,每个API都简单易用。
#
# 本书特点主要有:
#
# - 内容通俗易懂。本书只需要基本的编程和数学知识
# - 案例实用。本书的案例都很容易上手,读者可以调整后解决自己的问题。
# ## 本书内容简介
# [*第1章,机器学习基础*](http://muxuezi.github.io/posts/1-the-fundamentals-of-machine-learning.html),将机器学习定义成一种通过学习经验改善工作效果的程序研究与设计过程。其他章节都以这个定义为基础,后面每一章里介绍的机器学习模型都是按照这个思路解决任务,评估效果。
#
# [*第2章,线性回归*](http://muxuezi.github.io/posts/2-linear-regression.html),介绍线性回归模型,一种解释变量和模型参数与连续的响应变量相关的模型。本章介绍成本函数的定义,通过最小二乘法求解模型参数获得最优模型。
#
# [*第3章,特征提取与处理*](http://muxuezi.github.io/posts/3-feature-extraction-and-preprocessing.html),介绍了常见的机器学习对象如文本,图像与分类变量的特征提取与处理方法。
#
# [*第4章,从线性回归到逻辑回归*](http://muxuezi.github.io/posts/4-from-linear-regression-to-logistic-regression.html),介绍广义线性回归模型如何解决分类任务。将逻辑回归模型与特征提取技术结合起来实现一个垃圾短信分类器。
#
# [*第5章,决策树——非线性回归与分类*](http://muxuezi.github.io/posts/5-nonlinear-classification-and-regression-with-decision-trees.html),介绍了一种回归和分类的非线性模型——决策树。用决策树集成方法实现了一个网页广告图片屏蔽器。
#
# [*第6章,K-Means聚类*](http://muxuezi.github.io/posts/6-clustering-with-k-means.html),介绍非监督学习的K-Means聚类算法,并与逻辑回归组合起来实现一个照片分类器。
#
# [*第7章,用PCA降维*](http://muxuezi.github.io/posts/7-dimensionality-reduction-with-pca.html),介绍另一种非监督学习任务——降维。我们用主成分分析实现高维数据的可视化,建立一个脸部识别器。
#
# [*第8章,感知器*](http://muxuezi.github.io/posts/8-the-perceptron.html),介绍一种实时的,二元分类器——感知器。后面两章都是针对感知器的缺点发展起来的。
#
# [*第9章,从感知器到支持向量机*](http://muxuezi.github.io/posts/9-from-the-perceptron-to-support-vector-machines.html),介绍支持向量机,是一种有效的非线性回归与分类模型。我们用支持向量机识别街景照片中的字母。
#
# [*第10章,从感知器到人工神经网络*](http://muxuezi.github.io/posts/10-from-the-perceptron-to-artificial-neural-networks.html),介绍了人工神经网络,是一种强大的有效的非线性回归与分类模型。我们用人工神经网络识别手写数字。
| posts/0-perface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_amazonei_tensorflow_p36
# language: python
# name: conda_amazonei_tensorflow_p36
# ---
# # Distributed DeepRacer RL training with SageMaker and RoboMaker
#
# ---
# ## Introduction
#
#
# In this notebook, we will train a fully autonomous 1/18th scale race car using reinforcement learning using Amazon SageMaker RL and AWS RoboMaker's 3D driving simulator. [AWS RoboMaker](https://console.aws.amazon.com/robomaker/home#welcome) is a service that makes it easy for developers to develop, test, and deploy robotics applications.
#
# This notebook provides a jailbreak experience of [AWS DeepRacer](https://console.aws.amazon.com/deepracer/home#welcome), giving us more control over the training/simulation process and RL algorithm tuning.
#
# 
#
#
# ---
# ## How it works?
#
# 
#
# The reinforcement learning agent (i.e. our autonomous car) learns to drive by interacting with its environment, e.g., the track, by taking an action in a given state to maximize the expected reward. The agent learns the optimal plan of actions in training by trial-and-error through repeated episodes.
#
# The figure above shows an example of distributed RL training across SageMaker and two RoboMaker simulation envrionments that perform the **rollouts** - execute a fixed number of episodes using the current model or policy. The rollouts collect agent experiences (state-transition tuples) and share this data with SageMaker for training. SageMaker updates the model policy which is then used to execute the next sequence of rollouts. This training loop continues until the model converges, i.e. the car learns to drive and stops going off-track. More formally, we can define the problem in terms of the following:
#
# 1. **Objective**: Learn to drive autonomously by staying close to the center of the track.
# 2. **Environment**: A 3D driving simulator hosted on AWS RoboMaker.
# 3. **State**: The driving POV image captured by the car's head camera, as shown in the illustration above.
# 4. **Action**: Six discrete steering wheel positions at different angles (configurable)
# 5. **Reward**: Positive reward for staying close to the center line; High penalty for going off-track. This is configurable and can be made more complex (for e.g. steering penalty can be added).
# ## Prequisites
# ### Imports
# To get started, we'll import the Python libraries we need, set up the environment with a few prerequisites for permissions and configurations.
#
# You can run this notebook from your local machine or from a SageMaker notebook instance. In both of these scenarios, you can run the following to launch a training job on SageMaker and a simulation job on RoboMaker.
import boto3
import sagemaker
import sys
import os
import re
import numpy as np
import subprocess
sys.path.append("common")
from misc import get_execution_role, wait_for_s3_object
from docker_utils import build_and_push_docker_image
from sagemaker.rl import RLEstimator, RLToolkit, RLFramework
from time import gmtime, strftime
import time
from IPython.display import Markdown
from markdown_helper import *
# ### Initializing basic parameters
# +
# Select the instance type
#instance_type = "ml.c4.2xlarge"
instance_type = "ml.p2.xlarge"
#instance_type = "ml.c5.4xlarge"
# Starting SageMaker session
sage_session = sagemaker.session.Session()
# Create unique job name.
job_name_prefix = 'sahika-neuripschallenge-2019'
# Duration of job in seconds (1 hours)
job_duration_in_seconds = 60 * 20
# AWS Region
aws_region = sage_session.boto_region_name
if aws_region not in ["us-west-2", "us-east-1", "eu-west-1"]:
raise Exception("This notebook uses RoboMaker which is available only in US East (N. Virginia),"
"US West (Oregon) and EU (Ireland). Please switch to one of these regions.")
# -
# ### Setup S3 bucket
# Set up the linkage and authentication to the S3 bucket that we want to use for checkpoint and metadata.
# +
# S3 bucket
s3_bucket = sage_session.default_bucket()
# SDK appends the job name and output folder
s3_output_path = 's3://{}/'.format(s3_bucket)
#Ensure that the S3 prefix contains the keyword 'sagemaker'
s3_prefix = job_name_prefix + "-sagemaker-" + strftime("%y%m%d-%H%M%S", gmtime())
# Get the AWS account id of this account
sts = boto3.client("sts")
account_id = sts.get_caller_identity()['Account']
print("Using s3 bucket {}".format(s3_bucket))
print("Model checkpoints and other metadata will be stored at: \ns3://{}/{}".format(s3_bucket, s3_prefix))
# -
# ### Create an IAM role
# Either get the execution role when running from a SageMaker notebook `role = sagemaker.get_execution_role()` or, when running from local machine, use utils method `role = get_execution_role('role_name')` to create an execution role.
# +
try:
sagemaker_role = sagemaker.get_execution_role()
except:
sagemaker_role = get_execution_role('sagemaker')
print("Using Sagemaker IAM role arn: \n{}".format(sagemaker_role))
# -
# > Please note that this notebook cannot be run in `SageMaker local mode` as the simulator is based on AWS RoboMaker service.
# ### Permission setup for invoking AWS RoboMaker from this notebook
# In order to enable this notebook to be able to execute AWS RoboMaker jobs, we need to add one trust relationship to the default execution role of this notebook.
display(Markdown(generate_help_for_robomaker_trust_relationship(sagemaker_role)))
# ### Permission setup for Sagemaker to S3 bucket
#
# The sagemaker writes the Redis IP address, models to the S3 bucket. This requires PutObject permission on the bucket. Make sure the sagemaker role you are using as this permissions.
display(Markdown(generate_s3_write_permission_for_sagemaker_role(sagemaker_role)))
# ### Permission setup for Sagemaker to create KinesisVideoStreams
#
# The sagemaker notebook has to create a kinesis video streamer. You can observer the car making epsiodes in the kinesis video streamer.
display(Markdown(generate_kinesis_create_permission_for_sagemaker_role(sagemaker_role)))
# ### Build and push docker image
#
# The file ./Dockerfile contains all the packages that are installed into the docker. Instead of using the default sagemaker container. We will be using this docker container.
# %%time
cpu_or_gpu = 'gpu' if instance_type.startswith('ml.p') else 'cpu'
repository_short_name = "sagemaker-docker-%s" % cpu_or_gpu
docker_build_args = {
'CPU_OR_GPU': cpu_or_gpu,
'AWS_REGION': boto3.Session().region_name,
}
custom_image_name = build_and_push_docker_image(repository_short_name, build_args=docker_build_args)
print("Using ECR image %s" % custom_image_name)
# ### Configure VPC
#
# Since SageMaker and RoboMaker have to communicate with each other over the network, both of these services need to run in VPC mode. This can be done by supplying subnets and security groups to the job launching scripts.
# We will check if the deepracer-vpc stack is created and use it if present (This is present if the AWS Deepracer console is used atleast once to create a model). Else we will use the default VPC stack.
# +
ec2 = boto3.client('ec2')
#
# Check if the user has Deepracer-VPC and use that if its present. This will have all permission.
# This VPC will be created when you have used the Deepracer console and created one model atleast
# If this is not present. Use the default VPC connnection
#
deepracer_security_groups = [group["GroupId"] for group in ec2.describe_security_groups()['SecurityGroups']\
if group['GroupName'].startswith("deepracer-vpc")]
if(deepracer_security_groups):
print("Using the DeepRacer VPC stacks")
deepracer_vpc = [vpc['VpcId'] for vpc in ec2.describe_vpcs()['Vpcs'] \
if "Tags" in vpc for val in vpc['Tags'] \
if val['Value'] == 'deepracer-vpc'][0]
deepracer_subnets = [subnet["SubnetId"] for subnet in ec2.describe_subnets()["Subnets"] \
if subnet["VpcId"] == deepracer_vpc]
else:
print("Using the default VPC stacks")
deepracer_vpc = [vpc['VpcId'] for vpc in ec2.describe_vpcs()['Vpcs'] if vpc["IsDefault"] == True][0]
deepracer_security_groups = [group["GroupId"] for group in ec2.describe_security_groups()['SecurityGroups'] \
if 'VpcId' in group and group["GroupName"] == "default" and group["VpcId"] == deepracer_vpc]
deepracer_subnets = [subnet["SubnetId"] for subnet in ec2.describe_subnets()["Subnets"] \
if subnet["VpcId"] == deepracer_vpc and subnet['DefaultForAz']==True]
print("Using VPC:", deepracer_vpc)
print("Using security group:", deepracer_security_groups)
print("Using subnets:", deepracer_subnets)
# -
# ### Create Route Table
# A SageMaker job running in VPC mode cannot access S3 resourcs. So, we need to create a VPC S3 endpoint to allow S3 access from SageMaker container. To learn more about the VPC mode, please visit [this link.](https://docs.aws.amazon.com/sagemaker/latest/dg/train-vpc.html)
# +
#TODO: Explain to customer what CREATE_ROUTE_TABLE is doing
CREATE_ROUTE_TABLE = True
def create_vpc_endpoint_table():
print("Creating ")
try:
route_tables = [route_table["RouteTableId"] for route_table in ec2.describe_route_tables()['RouteTables']\
if route_table['VpcId'] == deepracer_vpc]
except Exception as e:
if "UnauthorizedOperation" in str(e):
display(Markdown(generate_help_for_s3_endpoint_permissions(sagemaker_role)))
else:
display(Markdown(create_s3_endpoint_manually(aws_region, deepracer_vpc)))
raise e
print("Trying to attach S3 endpoints to the following route tables:", route_tables)
if not route_tables:
raise Exception(("No route tables were found. Please follow the VPC S3 endpoint creation "
"guide by clicking the above link."))
try:
ec2.create_vpc_endpoint(DryRun=False,
VpcEndpointType="Gateway",
VpcId=deepracer_vpc,
ServiceName="com.amazonaws.{}.s3".format(aws_region),
RouteTableIds=route_tables)
print("S3 endpoint created successfully!")
except Exception as e:
if "RouteAlreadyExists" in str(e):
print("S3 endpoint already exists.")
elif "UnauthorizedOperation" in str(e):
display(Markdown(generate_help_for_s3_endpoint_permissions(role)))
raise e
else:
display(Markdown(create_s3_endpoint_manually(aws_region, default_vpc)))
raise e
if CREATE_ROUTE_TABLE:
create_vpc_endpoint_table()
# -
# ## Setup the environment
#
# The environment is defined in a Python file called “deepracer_racetrack_env.py” and the file can be found at `src/markov/environments/`. This file implements the gym interface for our Gazebo based RoboMakersimulator. This is a common environment file used by both SageMaker and RoboMaker. The environment variable - `NODE_TYPE` defines which node the code is running on. So, the expressions that have `rospy` dependencies are executed on RoboMaker only.
#
# We can experiment with different reward functions by modifying `reward_function` in `src/markov/rewards/`. Action space and steering angles can be changed by modifying `src/markov/actions/`.json file
#
# ### Configure the preset for RL algorithm
#
# The parameters that configure the RL training job are defined in `src/markov/presets/`. Using the preset file, you can define agent parameters to select the specific agent algorithm. We suggest using Clipped PPO for this example.
# You can edit this file to modify algorithm parameters like learning_rate, neural network structure, batch_size, discount factor etc.
# +
# Uncomment the pygmentize code lines to see the code
# Environmental File
# #!pygmentize src/markov/environments/deepracer_racetrack_env.py
# Reward function
# #!pygmentize src/markov/rewards/default.py
# Action space
# #!pygmentize src/markov/actions/model_metadata_10_state.json
# Preset File
# #!pygmentize src/markov/presets/default.py
# #!pygmentize src/markov/presets/preset_attention_layer.py
# -
# ### Copy custom files to S3 bucket so that sagemaker & robomaker can pick it up
# +
s3_location = "s3://%s/%s" % (s3_bucket, s3_prefix)
print(s3_location)
# Clean up the previously uploaded files
# !aws s3 rm --recursive {s3_location}
# Make any changes to the environment and preset files below and upload these files
# !aws s3 cp src/markov/environments/deepracer_racetrack_env.py {s3_location}/environments/deepracer_racetrack_env.py
# !aws s3 cp src/markov/rewards/default.py {s3_location}/rewards/reward_function.py
# !aws s3 cp src/markov/actions/model_metadata_10_state.json {s3_location}/model_metadata.json
# !aws s3 cp src/markov/presets/default.py {s3_location}/presets/preset.py
# #!aws s3 cp src/markov/presets/preset_attention_layer.py {s3_location}/presets/preset.py
# -
# ### Train the RL model using the Python SDK Script mode
#
# Next, we define the following algorithm metrics that we want to capture from cloudwatch logs to monitor the training progress. These are algorithm specific parameters and might change for different algorithm. We use [Clipped PPO](https://coach.nervanasys.com/algorithms/policy_optimization/cppo/index.html) for this example.
metric_definitions = [
# Training> Name=main_level/agent, Worker=0, Episode=19, Total reward=-102.88, Steps=19019, Training iteration=1
{'Name': 'reward-training',
'Regex': '^Training>.*Total reward=(.*?),'},
# Policy training> Surrogate loss=-0.32664725184440613, KL divergence=7.255815035023261e-06, Entropy=2.83156156539917, training epoch=0, learning_rate=0.00025
{'Name': 'ppo-surrogate-loss',
'Regex': '^Policy training>.*Surrogate loss=(.*?),'},
{'Name': 'ppo-entropy',
'Regex': '^Policy training>.*Entropy=(.*?),'},
# Testing> Name=main_level/agent, Worker=0, Episode=19, Total reward=1359.12, Steps=20015, Training iteration=2
{'Name': 'reward-testing',
'Regex': '^Testing>.*Total reward=(.*?),'},
]
# We use the RLEstimator for training RL jobs.
#
# 1. Specify the source directory which has the environment file, preset and training code.
# 2. Specify the entry point as the training code
# 3. Specify the choice of RL toolkit and framework. This automatically resolves to the ECR path for the RL Container.
# 4. Define the training parameters such as the instance count, instance type, job name, s3_bucket and s3_prefix for storing model checkpoints and metadata. **Only 1 training instance is supported for now.**
# 4. Set the RLCOACH_PRESET as "deepracer" for this example.
# 5. Define the metrics definitions that you are interested in capturing in your logs. These can also be visualized in CloudWatch and SageMaker Notebooks.
# +
estimator = RLEstimator(entry_point="training_worker.py",
source_dir='src',
image_name=custom_image_name,
dependencies=["common/"],
role=sagemaker_role,
train_instance_type=instance_type,
train_instance_count=1,
output_path=s3_output_path,
base_job_name=job_name_prefix,
metric_definitions=metric_definitions,
train_max_run=job_duration_in_seconds,
hyperparameters={
"s3_bucket": s3_bucket,
"s3_prefix": s3_prefix,
"aws_region": aws_region,
"preset_s3_key": "%s/presets/preset.py"% s3_prefix,
"model_metadata_s3_key": "%s/model_metadata.json" % s3_prefix,
"environment_s3_key": "%s/environments/deepracer_racetrack_env.py" % s3_prefix,
},
subnets=deepracer_subnets,
security_group_ids=deepracer_security_groups,
)
estimator.fit(wait=False)
job_name = estimator.latest_training_job.job_name
print("Training job: %s" % job_name)
# -
# ### Create the Kinesis video stream
# +
kvs_stream_name = "dr-kvs-{}".format(job_name)
# !aws --region {aws_region} kinesisvideo create-stream --stream-name {kvs_stream_name} --media-type video/h264 --data-retention-in-hours 24
print ("Created kinesis video stream {}".format(kvs_stream_name))
# -
# ### Start the Robomaker job
robomaker = boto3.client("robomaker")
# ### Create Simulation Application
robomaker_s3_key = 'robomaker/simulation_ws.tar.gz'
robomaker_source = {'s3Bucket': s3_bucket,
's3Key': robomaker_s3_key,
'architecture': "X86_64"}
simulation_software_suite={'name': 'Gazebo',
'version': '7'}
robot_software_suite={'name': 'ROS',
'version': 'Kinetic'}
rendering_engine={'name': 'OGRE',
'version': '1.x'}
# Download the DeepRacer bundle provided by RoboMaker service and upload it in our S3 bucket to create a RoboMaker Simulation Application
# +
# Download Robomaker simApp for the deepracer public s3 bucket
simulation_application_bundle_location = "s3://deepracer-managed-resources-us-east-1/deepracer-simapp.tar.gz"
#simulation_application_bundle_location = "s3://sahika-neuripschallenge-2019/deepracer-simapp.tar.gz"
# !aws s3 cp {simulation_application_bundle_location} ./
# Remove if the Robomaker sim-app is present in s3 bucket
# !aws s3 rm s3://{s3_bucket}/{robomaker_s3_key}
# Uploading the Robomaker SimApp to your S3 bucket
# !aws s3 cp ./deepracer-simapp.tar.gz s3://{s3_bucket}/{robomaker_s3_key}
# Cleanup the locally downloaded version of SimApp
# #!rm deepracer-simapp.tar.gz
# +
app_name = "sahika-neuripschallenge-2019" + strftime("%y%m%d-%H%M%S", gmtime())
print(app_name)
try:
response = robomaker.create_simulation_application(name=app_name,
sources=[robomaker_source],
simulationSoftwareSuite=simulation_software_suite,
robotSoftwareSuite=robot_software_suite,
renderingEngine=rendering_engine)
simulation_app_arn = response["arn"]
print("Created a new simulation app with ARN:", simulation_app_arn)
except Exception as e:
if "AccessDeniedException" in str(e):
display(Markdown(generate_help_for_robomaker_all_permissions(role)))
raise e
else:
raise e
# -
# ### Launch the Simulation job on RoboMaker
#
# We create [AWS RoboMaker](https://console.aws.amazon.com/robomaker/home#welcome) Simulation Jobs that simulates the environment and shares this data with SageMaker for training.
# +
num_simulation_workers = 1
envriron_vars = {
"WORLD_NAME": "reinvent_base",
"KINESIS_VIDEO_STREAM_NAME": kvs_stream_name,
"SAGEMAKER_SHARED_S3_BUCKET": s3_bucket,
"SAGEMAKER_SHARED_S3_PREFIX": s3_prefix,
"TRAINING_JOB_ARN": job_name,
"APP_REGION": aws_region,
"METRIC_NAME": "TrainingRewardScore",
"METRIC_NAMESPACE": "AWSDeepRacer",
"REWARD_FILE_S3_KEY": "%s/rewards/reward_function.py" % s3_prefix,
"MODEL_METADATA_FILE_S3_KEY": "%s/model_metadata.json" % s3_prefix,
"METRICS_S3_BUCKET": s3_bucket,
"METRICS_S3_OBJECT_KEY": s3_bucket + "/training_metrics.json",
"TARGET_REWARD_SCORE": "None",
"NUMBER_OF_EPISODES": "0",
"ROBOMAKER_SIMULATION_JOB_ACCOUNT_ID": account_id
}
simulation_application = {"application":simulation_app_arn,
"launchConfig": {"packageName": "deepracer_simulation_environment",
"launchFile": "distributed_training.launch",
"environmentVariables": envriron_vars}
}
vpcConfig = {"subnets": deepracer_subnets,
"securityGroups": deepracer_security_groups,
"assignPublicIp": True}
client_request_token = strftime("%<KEY>", gmtime())
responses = []
for job_no in range(num_simulation_workers):
response = robomaker.create_simulation_job(iamRole=sagemaker_role,
clientRequestToken=client_request_token,
maxJobDurationInSeconds=job_duration_in_seconds,
failureBehavior="Continue",
simulationApplications=[simulation_application],
vpcConfig=vpcConfig
)
responses.append(response)
print("Created the following jobs:")
job_arns = [response["arn"] for response in responses]
for response in responses:
print("Job ARN", response["arn"])
# -
# ### Visualizing the simulations in RoboMaker
# You can visit the RoboMaker console to visualize the simulations or run the following cell to generate the hyperlinks.
display(Markdown(generate_robomaker_links(job_arns, aws_region)))
# ### Creating temporary folder top plot metrics
tmp_dir = "/tmp/{}".format(job_name)
os.system("mkdir {}".format(tmp_dir))
print("Create local folder {}".format(tmp_dir))
# ### Plot metrics for training job
# +
# %matplotlib inline
import pandas as pd
import json
training_metrics_file = "training_metrics.json"
training_metrics_path = "{}/{}".format(s3_bucket, training_metrics_file)
wait_for_s3_object(s3_bucket, training_metrics_path, tmp_dir)
json_file = "{}/{}".format(tmp_dir, training_metrics_file)
with open(json_file) as fp:
data = json.load(fp)
df = pd.DataFrame(data['metrics'])
x_axis = 'episode'
y_axis = 'reward_score'
plt = df.plot(x=x_axis,y=y_axis, figsize=(12,5), legend=True, style='b-')
plt.set_ylabel(y_axis);
plt.set_xlabel(x_axis);
# -
# ### Clean up RoboMaker and SageMaker training job
#
# Execute the cells below if you want to kill RoboMaker and SageMaker job.
# +
# # Cancelling robomaker job
# for job_arn in job_arns:
# robomaker.cancel_simulation_job(job=job_arn)
# # Stopping sagemaker training job
# sage_session.sagemaker_client.stop_training_job(TrainingJobName=estimator._current_job_name)
# -
# ### Evaluation - ReInvent Track
# +
sys.path.append("./src")
num_simulation_workers = 1
envriron_vars = {
"WORLD_NAME": "reinvent_base",
"KINESIS_VIDEO_STREAM_NAME": "SilverstoneStream",
"MODEL_S3_BUCKET": s3_bucket,
"MODEL_S3_PREFIX": s3_prefix,
"APP_REGION": aws_region,
"MODEL_METADATA_FILE_S3_KEY": "%s/model_metadata.json" % s3_prefix,
"METRICS_S3_BUCKET": s3_bucket,
"METRICS_S3_OBJECT_KEY": s3_bucket + "/evaluation_metrics.json",
"NUMBER_OF_TRIALS": "5",
"ROBOMAKER_SIMULATION_JOB_ACCOUNT_ID": account_id
}
simulation_application = {
"application":simulation_app_arn,
"launchConfig": {
"packageName": "deepracer_simulation_environment",
"launchFile": "evaluation.launch",
"environmentVariables": envriron_vars
}
}
vpcConfig = {"subnets": deepracer_subnets,
"securityGroups": deepracer_security_groups,
"assignPublicIp": True}
responses = []
for job_no in range(num_simulation_workers):
response = robomaker.create_simulation_job(clientRequestToken=strftime("%<KEY>", gmtime()),
outputLocation={
"s3Bucket": s3_bucket,
"s3Prefix": s3_prefix
},
maxJobDurationInSeconds=job_duration_in_seconds,
iamRole=sagemaker_role,
failureBehavior="Continue",
simulationApplications=[simulation_application],
vpcConfig=vpcConfig)
responses.append(response)
# print("Created the following jobs:")
for response in responses:
print("Job ARN", response["arn"])
# -
# ### Creating temporary folder top plot metrics
# +
evaluation_metrics_file = "evaluation_metrics.json"
evaluation_metrics_path = "{}/{}".format(s3_bucket, evaluation_metrics_file)
wait_for_s3_object(s3_bucket, evaluation_metrics_path, tmp_dir)
json_file = "{}/{}".format(tmp_dir, evaluation_metrics_file)
with open(json_file) as fp:
data = json.load(fp)
df = pd.DataFrame(data['metrics'])
# Converting milliseconds to seconds
df['elapsed_time'] = df['elapsed_time_in_milliseconds']/1000
df = df[['trial', 'completion_percentage', 'elapsed_time']]
display(df)
# -
# ### Clean Up Simulation Application Resource
# +
# robomaker.delete_simulation_application(application=simulation_app_arn)
# -
# ### Clean your S3 bucket (Uncomment the awscli commands if you want to do it)
# +
## Uncomment if you only want to clean the s3 bucket
# sagemaker_s3_folder = "s3://{}/{}".format(s3_bucket, s3_prefix)
# # !aws s3 rm --recursive {sagemaker_s3_folder}
# robomaker_s3_folder = "s3://{}/{}".format(s3_bucket, job_name)
# # !aws s3 rm --recursive {robomaker_s3_folder}
# robomaker_sim_app = "s3://{}/{}".format(s3_bucket, 'robomaker')
# # !aws s3 rm --recursive {robomaker_sim_app}
# model_output = "s3://{}/{}".format(s3_bucket, s3_bucket)
# # !aws s3 rm --recursive {model_output}
# -
# ### Clean the docker images
# Remove this only when you want to completely remove the docker or clean up the space of the sagemaker instance
# +
# # !docker rmi -f $(docker images -q)
| Advanced workshops/AI Driving Olympics 2019/challenge_train_DQN/deepracer_rl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Project: Linear Regression
#
# Reggie is a mad scientist who has been hired by the local fast food joint to build their newest ball pit in the play area. As such, he is working on researching the bounciness of different balls so as to optimize the pit. He is running an experiment to bounce different sizes of bouncy balls, and then fitting lines to the data points he records. He has heard of linear regression, but needs your help to implement a version of linear regression in Python.
#
# _Linear Regression_ is when you have a group of points on a graph, and you find a line that approximately resembles that group of points. A good Linear Regression algorithm minimizes the _error_, or the distance from each point to the line. A line with the least error is the line that fits the data the best. We call this a line of _best fit_.
#
# We will use loops, lists, and arithmetic to create a function that will find a line of best fit when given a set of data.
#
# ## Part 1: Calculating Error
#
# The line we will end up with will have a formula that looks like:
# ```
# y = m*x + b
# ```
# `m` is the slope of the line and `b` is the intercept, where the line crosses the y-axis.
#
# Fill in the function called `get_y()` that takes in `m`, `b`, and `x`. It should return what the `y` value would be for that `x` on that line!
#
# +
def get_y(m, b, x):
y = m*x + b
return y
print(get_y(1, 0, 7) == 7)
print(get_y(5, 10, 3) == 25)
# -
#
# Reggie wants to try a bunch of different `m` values and `b` values and see which line produces the least error. To calculate error between a point and a line, he wants a function called `calculate_error()`, which will take in `m`, `b`, and an [x, y] point called `point` and return the distance between the line and the point.
#
# To find the distance:
# 1. Get the x-value from the point and store it in a variable called `x_point`
# 2. Get the y-value from the point and store it in a variable called `y_point`
# 3. Use `get_y()` to get the y-value that `x_point` would be on the line
# 4. Find the difference between the y from `get_y` and `y_point`
# 5. Return the absolute value of the distance (you can use the built-in function `abs()` to do this)
#
# The distance represents the error between the line `y = m*x + b` and the `point` given.
#
#Write your calculate_error() function here
def calculate_error(m, b, point):
x_point, y_point = point
y = m*x + b
distance = abs(y - y_point)
return distance
# Let's test this function!
#this is a line that looks like y = x, so (3, 3) should lie on it. thus, error should be 0:
print(calculate_error(1, 0, (3, 3)))
#the point (3, 4) should be 1 unit away from the line y = x:
print(calculate_error(1, 0, (3, 4)))
#the point (3, 3) should be 1 unit away from the line y = x - 1:
print(calculate_error(1, -1, (3, 3)))
#the point (3, 3) should be 5 units away from the line y = -x + 1:
print(calculate_error(-1, 1, (3, 3)))
# Great! Reggie's datasets will be sets of points. For example, he ran an experiment comparing the width of bouncy balls to how high they bounce:
#
datapoints = [(1, 2), (2, 0), (3, 4), (4, 4), (5, 3)]
# The first datapoint, `(1, 2)`, means that his 1cm bouncy ball bounced 2 meters. The 4cm bouncy ball bounced 4 meters.
#
# As we try to fit a line to this data, we will need a function called `calculate_all_error`, which takes `m` and `b` that describe a line, and `points`, a set of data like the example above.
#
# `calculate_all_error` should iterate through each `point` in `points` and calculate the error from that point to the line (using `calculate_error`). It should keep a running total of the error, and then return that total after the loop.
#
# +
#Write your calculate_all_error function here
# -
# Let's test this function!
# +
#every point in this dataset lies upon y=x, so the total error should be zero:
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(1, 0, datapoints))
#every point in this dataset is 1 unit away from y = x + 1, so the total error should be 4:
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(1, 1, datapoints))
#every point in this dataset is 1 unit away from y = x - 1, so the total error should be 4:
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(1, -1, datapoints))
#the points in this dataset are 1, 5, 9, and 3 units away from y = -x + 1, respectively, so total error should be
# 1 + 5 + 9 + 3 = 18
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(-1, 1, datapoints))
# -
# Great! It looks like we now have a function that can take in a line and Reggie's data and return how much error that line produces when we try to fit it to the data.
#
# Our next step is to find the `m` and `b` that minimizes this error, and thus fits the data best!
#
# ## Part 2: Try a bunch of slopes and intercepts!
#
# The way Reggie wants to find a line of best fit is by trial and error. He wants to try a bunch of different slopes (`m` values) and a bunch of different intercepts (`b` values) and see which one produces the smallest error value for his dataset.
#
# Using a list comprehension, let's create a list of possible `m` values to try. Make the list `possible_ms` that goes from -10 to 10 inclusive, in increments of 0.1.
#
# Hint (to view this hint, either double-click this cell or highlight the following white space): <font color="white">you can go through the values in range(-100, 100) and multiply each one by 0.1</font>
#
#
possible_ms = #your list comprehension here
# Now, let's make a list of `possible_bs` to check that would be the values from -20 to 20 inclusive, in steps of 0.1:
possible_bs = #your list comprehension here
# We are going to find the smallest error. First, we will make every possible `y = m*x + b` line by pairing all of the possible `m`s with all of the possible `b`s. Then, we will see which `y = m*x + b` line produces the smallest total error with the set of data stored in `datapoint`.
#
# First, create the variables that we will be optimizing:
# * `smallest_error` — this should start at infinity (`float("inf")`) so that any error we get at first will be smaller than our value of `smallest_error`
# * `best_m` — we can start this at `0`
# * `best_b` — we can start this at `0`
#
# We want to:
# * Iterate through each element `m` in `possible_ms`
# * For every `m` value, take every `b` value in `possible_bs`
# * If the value returned from `calculate_all_error` on this `m` value, this `b` value, and `datapoints` is less than our current `smallest_error`,
# * Set `best_m` and `best_b` to be these values, and set `smallest_error` to this error.
#
# By the end of these nested loops, the `smallest_error` should hold the smallest error we have found, and `best_m` and `best_b` should be the values that produced that smallest error value.
#
# Print out `best_m`, `best_b` and `smallest_error` after the loops.
#
#
# ## Part 3: What does our model predict?
#
# Now we have seen that for this set of observations on the bouncy balls, the line that fits the data best has an `m` of 0.3 and a `b` of 1.7:
#
# ```
# y = 0.3x + 1.7
# ```
#
# This line produced a total error of 5.
#
# Using this `m` and this `b`, what does your line predict the bounce height of a ball with a width of 6 to be?
# In other words, what is the output of `get_y()` when we call it with:
# * m = 0.3
# * b = 1.7
# * x = 6
# Our model predicts that the 6cm ball will bounce 3.5m.
#
# Now, Reggie can use this model to predict the bounce of all kinds of sizes of balls he may choose to include in the ball pit!
| Reggie's Linear Regression (2)/Reggie_Linear_Regression_Skeleton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Forecasting the future incidence of highly pathogenic contagious diseases
# +
import os
import time
from termcolor import colored
import pandas as pd
import numpy as np
import random
import tensorflow as tf
import tensorflow.keras.backend as K
import matplotlib.pyplot as plt
from python.models import model_TCN
from python.transform import log_df, inv_log_df
# %matplotlib inline
# -
# ## Loading data
# +
# n_his is the number of days used for model input
n_his = 30
n_inf_days = 30
n_pred_days = 1
data_path = './dataset/train_data_all'
output_path = './dataset/data_processed'
city_list = ["A", "B", "C", "D", "E", "F", "G", "H", "I", "J", "K"]
region_nums = []
for city in city_list:
filename = os.path.join(data_path, f'city_{city}', 'grid_attr.csv')
df = pd.read_csv(filename, header=None, names=['x', 'y', 'region'])
region_nums.append(len(set(df['region'])))
tot_regions = sum(region_nums)
population_dict = (pd
.read_csv(os.path.join(output_path, 'density.csv'))[:6]
.drop(columns=['date'])
.mean(axis=0)
.to_dict())
df_inf = pd.read_csv(os.path.join(output_path, 'infection.csv'))
df_dens_int = pd.read_csv(os.path.join(output_path, 'density_int.csv'))
df_mig_in = pd.read_csv(os.path.join(output_path, 'migration_in.csv'))
df_mig_out = pd.read_csv(os.path.join(output_path, 'migration_out.csv'))
del df_inf['date']
del df_dens_int['date']
del df_mig_in['date']
del df_mig_out['date']
df_inf = df_inf.sort_index(axis=1)
cols = df_inf.columns.to_list()
df_dens_int = df_dens_int.sort_index(axis=1)
df_mig_in = df_mig_in.sort_index(axis=1)
df_mig_out = df_mig_out.sort_index(axis=1)
# -
# ## Processing data
# Calculate some new features
# +
# dinsity (flux)
df_popu = df_dens_int.copy()
for col in df_popu.columns:
df_popu[col] = population_dict[col]
# average migration
df_mig_avg = (df_mig_in + df_mig_out) / 2.
# cumulitive infection
df_cumu = df_inf.cumsum()
# inverse increasing rate = 1 / (cumu_inf_today / cumu_inf_yesterday)
# call it "decrease", but it's not the real decreasing rate
df_dcrs = (df_cumu.shift() / df_cumu).fillna(0)
# -
# Most data are transformed by taking log plus 1.
# +
# transforming datasets and taking lags
df_inf, sc_inf, max_inf = log_df(df_inf)
df_inf_lag = df_inf.shift(periods=1)
df_inf_lag[:1] = 0
df_cumu, sc_cumu, max_cumu = log_df(df_cumu)
df_cumu_lag = df_cumu.shift(periods=1)
df_cumu_lag[:1] = 0
df_dcrs = df_dcrs.shift(periods=1)
df_dcrs[:1] = 0
df_popu, sc_popu, max_popu = log_df(df_popu)
df_mig_avg, sc_mig_avg, max_mig_avg = log_df(df_mig_avg)
df_dens_int, sc_dens_int, max_dens_int = log_df(df_dens_int)
# +
df_feat_dict = {'inf_lag' : df_inf_lag,
'cumu_lag' : df_cumu_lag,
'dcrs' : df_dcrs,
'popu' : df_popu,
'mig_avg' : df_mig_avg,
'dens' : df_dens_int,
}
df_feat_list = list(df_feat_dict.values())
# append y to the list
list_df = df_feat_list + [df_inf]
# -
# ## Data Formatting
# +
# split a multivariate sequence into samples
def split_sequences(sequences, n_his, n_pred=1):
X, y = list(), list()
for i in range(len(sequences)):
# find the end of this pattern
end_ix = i + n_his
out_end_ix = end_ix + n_pred - 1
# check if we are beyond the dataset
if out_end_ix > len(sequences):
break
# gather input and output parts of the pattern
seq_x, seq_y = sequences[i:end_ix, :-1], sequences[end_ix-1:out_end_ix, -1]
X.append(seq_x)
y.append(seq_y)
return np.array(X), np.array(y)
def get_xy(list_df, col='A_10', n_his=10, n_pred=1):
list_seq = []
# loop over features
for df in list_df:
seq = df[col].values.reshape((len(df), 1))
list_seq.append(seq)
dataset = np.hstack(list_seq)
X, y = split_sequences(dataset, n_his, n_pred)
# city_region_day
label = np.char.add(col+'_', np.arange(y.shape[0]).astype(str))
return X, y, label
def get_xy_all(list_df, n_his=10, n_pred=1):
X_list = []
y_list = []
label_list = []
for col in cols:
X, y, label = get_xy(list_df, col=col, n_his=n_his, n_pred=n_pred)
X_list.append(X)
y_list.append(y)
label_list.append(label)
X = np.vstack(X_list)
y = np.vstack(y_list)
label = np.hstack(label_list)
return X, y, label
def get_xy_inf(list_df, n_his=10, n_pred=1):
X_list = []
y_list = []
label_list = []
for col in cols:
X, y, label = get_xy(list_df, col=col, n_his=n_his, n_pred=1)
X_list.append(X[-1:])
y_list.append(y[-1:])
label_list.append(label[-1:])
X = np.vstack(X_list)
y = np.vstack(y_list)
label = np.hstack(label_list)
return X, y, label
# -
# ## Inference
# +
# take the contral value over last 10 values for inference
mig_avg_last = df_mig_avg[-10:].median(axis=0).values
dens_int_last = df_dens_int[-10:].median(axis=0).values
# linear fit
class Fitter():
def __init__(self):
self.n_data_fit = 20 # number of data used for fitting
self.active = False # if fitter is triggered
self.n_preds = 0 # prediciont counter
self.coef = np.array([99,99]) # (slope, intercept)
self.poly1d_fn = None # fitting function
def reset(self):
self.active = False
self.n_preds = 0
self.coef = np.array([99,99])
self.poly1d_fn = None
def passed_peak(self, a_in):
# fitting might be unstable if there's a lot of missing data
has_data = sum(a_in>0) > self.n_data_fit
max_idx = a_in.argmax()
# the peak should be in front of fitting data
return max_idx < (len(a_in) - self.n_data_fit)
def predict(self, a_in):
# a_in non-transformed
if a_in[-1] < 1:
return 0 # infection number should be an integer
if not self.active:
if not self.passed_peak(a_in):
return -99
x = np.arange(self.n_data_fit)
y = a_in[-self.n_data_fit:]
mask = y>0
x = x[mask]
y = np.log(y[mask]) # assume non-transformed
coef = np.polyfit(x, y, 1)
if (coef[0]>=0):
return -99
elif (coef[0]<=self.coef[0]):
self.coef = coef
return -99
else:
# trigger of the linear fit
self.active = True
self.poly1d_fn = np.poly1d(self.coef)
pred = self.poly1d_fn(self.n_data_fit+self.n_preds)
self.n_preds += 1 # increase by 1 after each prediction
# transform back to original infection
pred = np.exp(pred)
return pred
# +
pred_cols = list(df_feat_dict.keys())
id_IL = pred_cols.index('inf_lag') if 'inf_lag' in pred_cols else -99
id_CML = pred_cols.index('cumu_lag') if 'cumu_lag' in pred_cols else -99
id_DC = pred_cols.index('dcrs') if 'dcrs' in pred_cols else -99
id_PP = pred_cols.index('popu') if 'popu' in pred_cols else -99
id_MA = pred_cols.index('mig_avg') if 'mig_avg' in pred_cols else -99
id_DENS = pred_cols.index('dens') if 'dens' in pred_cols else -99
def inference(X_in, y_in, n_pred, model_list, int_inf=True, linear_fit=False):
single_model = len(model_list)==1
# X = (n_region, n_his, n_feat)
X = X_in.copy()
pred_size = len(X)
batch_size = 1
while batch_size < pred_size:
batch_size = batch_size * 2
pred_list = []
fitters = [Fitter() for i in range(tot_regions)]
for i in range(n_pred):
# pred = (n_region,)
if i==0:
pred = y_in.reshape((-1,1))
else:
# prediction from all models
preds = []
for model in model_list:
pred = model.predict(X, verbose=0, batch_size=1000)[:,0,:].reshape((-1,1))
pred[pred < 0] = 0
preds.append(pred)
# use the central model
pred = preds[0] if single_model else np.median(preds, axis=0)
# feat_new = (n_region, n_feat)
feat_new = np.zeros(shape=X[:,-1,:].shape)
##### new infection #####
if id_IL >= 0:
feat_new[:, id_IL] = pred[:, -1].copy()
inf_today = inv_log_df(feat_new[:, id_IL], sc_inf, max_inf)
if linear_fit and i>0:
inf_his = inv_log_df(X[:,:,id_IL], sc_inf, max_inf)
inf_linear = np.array([fitters[i].predict(inf_his[i,:]) for i in range(tot_regions)])
# linear fit is done for log scale, so inf_linear must be >= 0
_mask = inf_linear >= 0
inf_today[_mask] = inf_linear[_mask]
if int_inf and i>0:
inf_today = inf_today.astype(int)
feat_new[:, id_IL], _, _ = log_df(inf_today, sc_inf, max_inf)
if i>0:
inf_last = inv_log_df(X[:, -1, id_IL], sc_inf, max_inf)
feat_new[:, id_IL][inf_last<1] = 0
##### cumulative #####
if id_CML >= 0:
cumu_last = inv_log_df(X[:, -1, id_CML], sc_cumu, max_cumu)
inf_today = inv_log_df(feat_new[:, id_IL], sc_inf, max_inf)
cumu_today = cumu_last + inf_today
feat_new[:, id_CML], _, _ = log_df(cumu_today, sc_cumu, max_cumu)
##### inversed increasing rate #####
if id_DC >= 0:
dcrs_rate = cumu_last / np.where(cumu_today==0, np.inf, cumu_today)
feat_new[:, id_DC] = dcrs_rate
##### migration #####
if id_MA >= 0:
feat_new[:, id_MA] = mig_avg_last
##### density #####
if id_DENS >= 0:
feat_new[:, id_DENS] = dens_int_last
##### population #####
if id_PP >= 0:
popu_last = X[:, -1, id_PP]
feat_new[:, id_PP] = popu_last
# add prediction to input #####
X[:, 0:-1, :] = X[:, 1:, :]
X[:, -1, :] = feat_new
pred_list.append(feat_new)
# return np.append(pred_list, axis=0)
return np.concatenate(pred_list)
# -
# ## Training and taking average
# +
from tensorflow.keras.optimizers import Adam, RMSprop
from sklearn.model_selection import train_test_split
n_models = 50
max_models = 100
n_epochs = 50
n_batch = 64
learning_rate = 0.001 # default:0.001
test_size = 0.2
n_pred_dim = 1
n_features = len(list_df) - n_pred_dim
def mean_squared_error(y_true, y_pred):
return K.mean(K.square(y_pred - y_true))
def root_mean_squared_error(y_true, y_pred):
return K.sqrt(K.mean(K.square(y_pred - y_true)))
# +
current_time = time.strftime("%H:%M:%S", time.localtime())
print('starting local time:', current_time)
model_list = []
his_fit_list = []
for ith_model in range(max_models):
# timer
start_time = time.time()
# load and split training and tesing data
X, y, label = get_xy_all(list_df, n_his=n_his, n_pred=n_pred_days)
city_label = np.array([l.split('_', 1)[0] for l in label])
region_label = np.array([l.rsplit('_', 1)[0] for l in label])
(X_train, X_val,
y_train, y_val,
l_train, l_val) = train_test_split(X, y, label,
test_size=test_size,
shuffle=True,
# random_state=666,
stratify=region_label)
# calculate weights based on cities and regions
city_label_train = np.array([l.split('_', maxsplit=1)[0] for l in l_train])
city_label_test = np.array([l.split('_', maxsplit=1)[0] for l in l_val])
unique, counts = np.unique(city_label, return_counts=True)
weight_map = dict(zip(unique, sum(counts)/counts))
w_train = np.array([weight_map[_city] for _city in city_label_train])
w_test = np.array([weight_map[_city] for _city in city_label_test])
# load model
model = model_TCN(n_his=n_his, n_feat=n_features, n_out=n_pred_days, dropout_rate=0.2)
opt = Adam(learning_rate=learning_rate)
model.compile(optimizer=opt,
loss='mean_squared_error',
metrics=[root_mean_squared_error, 'mean_squared_error'])
# calculate batch size
train_size = len(X_train)
batch_size = 1
while batch_size < train_size / n_batch:
batch_size = batch_size * 2
# start training
his_fit = model.fit(x=X_train,
y=y_train,
batch_size = batch_size,
epochs=n_epochs,
verbose=0,
validation_data=(X_val, y_val))
last_loss = his_fit.history["loss"][-1]
last_val_loss = his_fit.history["val_loss"][-1]
# save if model is okay
model_ok = last_val_loss < 0.001
if model_ok:
his_fit_list.append(his_fit)
model_list.append(model)
model_path = os.path.join('models/running', f'model_{len(model_list)-1}.h5')
model.save(model_path)
model_info = ''.join([f'model number: {ith_model:02}',
f' time: {int(time.time() - start_time) // 60} min',
f' X shape: {X_train.shape}',
f' batch_size: {batch_size}',
f' loss: {last_loss:.5f}',
f' val_loss: {last_val_loss:.5f}'])
print(colored(model_info, 'green' if model_ok else 'red'))
if len(model_list)==n_models:
break
# +
def plot_loss(his_fit_list):
loss = np.median([fit_his.history['loss'] for fit_his in his_fit_list], axis=0)
val_loss = np.median([fit_his.history['val_loss'] for fit_his in his_fit_list], axis=0)
mse = np.median([fit_his.history['mean_squared_error'] for fit_his in his_fit_list], axis=0)
val_mse = np.median([fit_his.history['val_mean_squared_error'] for fit_his in his_fit_list], axis=0)
fig, axes = plt.subplots(nrows=1, ncols=4, figsize=(16,4))
axes[0].plot(loss)
axes[0].plot(val_loss)
axes[0].set_title('loss')
axes[1].plot(loss)
axes[1].plot(val_loss)
axes[1].set_title('log_loss')
axes[1].set_yscale('log')
axes[2].plot(mse)
axes[2].plot(val_mse)
axes[2].set_title('MSE')
axes[3].plot(mse)
axes[3].plot(val_mse)
axes[3].set_title('log_MSE')
axes[3].set_yscale('log')
plot_loss(his_fit_list)
# -
# ## Save result
# +
def save_to_submit(int_inf=False, linear_fit=False):
X_inf, y_inf, _ = get_xy_inf(list_df, n_his=n_his, n_pred=n_pred_days)
inf_all = inference(X_inf, y_inf, n_inf_days, model_list, int_inf=int_inf, linear_fit=linear_fit)[:,id_IL]
# (n_region, n_pred)
inf_all = np.reshape(inf_all, (n_inf_days,-1)).T
dates = pd.date_range('2120-06-30', periods=n_inf_days).strftime('%Y%m%d')
pred_list = []
for col, inf in zip(cols, inf_all):
city = [col.split('_')[0]]*n_inf_days
region = [col.split('_')[1]]*n_inf_days
inf = inv_log_df(inf, sc_inf, max_inf)
pred_list += [list(_z) for _z in zip(city, region, dates, inf)]
df_pred = pd.DataFrame(np.array(pred_list))
df_pred.to_csv('outputs/submission.csv', header=False, index=False)
save_to_submit(int_inf=False, linear_fit=True)
| training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import Libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import statsmodels.api as sm
from scipy.stats import norm
# Read in excess returns from excel data
#avaiable sheets: descriptions, hedge_fund_series, merrill_factors, other_data
funds = pd.read_excel('proshares_analysis_data.xlsx',sheet_name ='hedge_fund_series',
converters= {'date': pd.to_datetime}, index_col=0)
funds.head()
#fn to quickly calc mean, vol, sharpe
#returns = pandas df annualization = int
def performanceMetrics(returns,annualization=1):
metrics = pd.DataFrame(index=returns.columns)
metrics['Mean'] = returns.mean() * annualization
metrics['Vol'] = returns.std() * np.sqrt(annualization)
metrics['Sharpe'] = (returns.mean() / returns.std()) * np.sqrt(annualization)
return metrics
# ### Question 1
# **For the series in the “hedge fund series” tab, report the following summary statistics (annualize): <br />
#  (a) mean <br />
#  (b) volatility <br />
#  (c) Sharpe ratio**
stats = performanceMetrics(funds, 12)
stats
# ### Question 2
# **For the series in the “hedge fund series” tab, , calculate the following statistics related to tailrisk (do not annualize): <br />
#  (a) Skewness   $\zeta = \frac{\mathbb{E}[(x-\mu)^{3}]}{\sigma^{3}}$<br />
#  (b) Excess Kurtosis (in excess of 3)   $\kappa = \frac{\mathbb{E}[(x-\mu)^{4}]}{\sigma^{4}}$<br />
#  (c) VaR (.05) - the fifth quantile of historic returns   $r^{Var}_{\pi,\tau} = F_{\tau}^{r(-1)}(\pi)$<br />
#  (d) CVaR (.05) - the mean of the returns at or below the fifth quantile<br />
#  (e) Maximum drawdown - include the dates of the max/min/recovery within the max drawdown
# period.**
#measure of data symmetry, is it skewed pos or neg
stats["Skew"] = funds.skew()
#mesaure of tail heaviness of the data, pandas default is k-3 which is what we need
stats["Kurtosis"] = funds.kurtosis()
stats
# VaR = there's a pi % chance that over the next t days, the portfolio will lose more than VaR
#VaR in term of returns is simply the Pithe quantile of the observed returns
#aggregate runs the percentile function on each column
#https://s3.amazonaws.com/assets.datacamp.com/production/course_5612/slides/chapter4.pdf
stats["VaR (0.05)"] = funds.aggregate(lambda x: np.percentile(x, 5))
stats
#CVaR = if the portfolio does lose at least amount VaR, what is the expected loss
#we are looking for the average of the worst 5% of outcomes for our portfolio
#aggregate runs our lambda fn over each col
#find the subset of returns that are less than or equal to the VaR @ the 5th percentile and avg them
#https://s3.amazonaws.com/assets.datacamp.com/production/course_5612/slides/chapter4.pdf
stats["CVaR (0.05)"] = funds.aggregate(lambda x: x[x <= np.percentile(x, 5)].mean())
stats
# +
def drawdownMetrics(returns):
#some code from in class demo
cum_returns = (1 + returns).cumprod()
rolling_max = cum_returns.cummax()
drawdown = (cum_returns - rolling_max) / rolling_max
# Make the index datetime
drawdown.index = pd.to_datetime(drawdown.index)
metrics = pd.DataFrame(index=returns.columns)
metrics['Max Drawdown'] = drawdown.min()
metrics['Peak'] = drawdown.agg(lambda x: x[(x == 0) & (x.index < x.idxmin())].last_valid_index())
bottom = drawdown.idxmin()
metrics['Bottom'] = bottom
recover = drawdown.agg(lambda x: x[(x == 0) & (x.index > x.idxmin())].first_valid_index())
metrics['Recover'] = recover
metrics['Time to Recover'] = recover - bottom
return metrics
#calculate drawdown metrics
drawdownMetrics(funds)
# -
# ### Question 3
# **For the series in the “hedge fund series” tab, run a regression of each against SPY (found in the
# “merrill factors” tab.) Include an intercept. Report the following regression-based statistics (annualize as appropriate): <br />
#  (a) Market Beta<br />
#  (b) Treynor Ratio<br />
#  (c) Information ratio<br />**
#already read in hedge fund series data, sotred in "fund" variable
#read in merrill factors data to get SPY returns
factors = pd.read_excel('proshares_analysis_data.xlsx',sheet_name ='merrill_factors',
converters= {'date': pd.to_datetime}, index_col=0)
factors.head()
# +
#create fresh df
regMetrics = pd.DataFrame(index=funds.columns)
#regress each fund return on SPY, code from in class demo used as a reference
for fund in funds:
y= funds[fund]
#add const to regression
x = sm.add_constant(factors["SPY US Equity"])
#regress fund on SPY
reg = sm.OLS(y, x, missing='drop').fit()
#add metrics to a df
const, beta = reg.params
#add beta
regMetrics.loc[fund, "mkt beta"] = beta
#add treynor ratio
#treynor = expected return/Beta times 12 to annualize
regMetrics.loc[fund, "treynor ratio"] = (y.mean() / beta) *12
#add information ratio
#info ratio = sharpe ratio of the non-factor component (alpha+error) of the return sqrt(12) to annualize
regMetrics.loc[fund, "info ratio"] = (const*12) / (reg.resid.std() * np.sqrt(12))
regMetrics
# -
# ### Question 4
# **Relative Performance - Discuss the previous statistics, and what they tell us about: <br />
#  (a) the differences between SPY and the hedge-fund series? <br />
#  (b) which performs better between HDG and QAI.<br />
#  (c) whether HDG and the ML series capture the most notable properties of HFRI.**
# <span style="color:#00008B"> **Solution**:<br>4a. When we regress each fund to SPY, we can see that the hedge fund series all have market betas of between 0.2 and 0.4, which suggests little to no correlation to SPY. This is expected, as these hedge funds are acting as investment alternatives to the S&P 500, so we would want them to have little correlation to the S&P 500 as it moves up or down.
# 4b. After analyzing the treynor ratio and information ratio, we can see that QAI performs better than HDG on both metrics. The treynor ratio determines how much excess return is generated per each unit of risk taken on in a portfolio. Since QAI has a higher treynor ratio than HDG, it means that it generated more return compared to the risk it took on. The information ratio measures the returns beyond a benchmark compared to the volatility of those returns. Once again, QAI has a higher information ratio than HDG, meaning that it generated more return beyond the benchmark when taking into account those returns' volatility. <br>
# 4c. HFRI was meant to be an investment alternative to the S&P 500 for investors that wanted an option that had little correlation to the S&P. When analyzing the market beta, we can see that HDG and ML had betas of between 0.35 and 0.4, meaning little correlation to the S&P, so they effectively capture HFRI's most notable property of low correlation to the S&P 500.
# Functionalize the regression procedures
def regression(Y,X):
# output dataframe with stats
df_out = pd.DataFrame(index = Y.columns)
# Add constant for regression
X = sm.add_constant(X)
# Run regression
results = sm.OLS(Y, X, missing='drop').fit()
# Market beta
df_out['Beta'] = results.params[1]
# Treynor Ratio - annualized portfolio mean / market beta
df_out['Treynor Ratio'] = (Y.mean() * 12) / df_out['Beta']
# Information Ratio - excess annualized return over market (alpha) / non-factor vol (residuals)
df_out['Info Ratio'] = (results.params[0] * 12) / (results.resid.std()*np.sqrt(12))
print(df_out)
return df_out
regression(funds[['HDG US Equity']],funds[['HFRIFWI Index']])
regression(funds[['MLEIFCTR Index']],funds[['HFRIFWI Index']])
regression(funds[['MLEIFCTX Index']],funds[['HFRIFWI Index']])
# ### Question 5
# **Report the correlation matrix for these assets: <br />
#  (a) Show the correlations as a heat map.<br />
#  (b) Which series have the highest and lowest correlations?**
# +
# Create correlation matrix from data frame with hudge funds and spy returns
corrmat = pd.merge(funds,factors.loc[:,["SPY US Equity"]], left_index=True, right_index=True).corr()
#dont record correlation=1 values (aka the asset vs itself)
#this comes in handy below when we need to find the max correleation pair
corrmat[corrmat==1] = None
#adjust heatmap size
plt.figure(figsize=(7, 5))
#set bounds for corrleation: -1=perfectly uncorr 1=perfectly corr
sns.heatmap(corrmat, vmin=-1, vmax=1, annot=True)
corr_rank = corrmat.unstack().sort_values().dropna()
# Get pairs with min and max correlations
pair_max = corr_rank.index[-1]
pair_min = corr_rank.index[0]
print(f'MIN Correlation pair is:{pair_min} {corr_rank[0]}')
print(f'MAX Correlation pair is:{pair_max} {corr_rank[-1]}')
# -
# ### Question 6
# **Replicate HFRI with the six factors listed on the “merrill factors” tab. Include a constant, and
# run the unrestricted regression:** <br />
#
# $$r^{hfri}_{t} = \alpha^{merr} + x^{merr}_{t} \beta^{merr} + \epsilon^{merr}_{t}$$
# $$\hat{r}^{hfri}_{t} = \hat{\alpha}^{merr} + x^{merr}_{t} \hat{\beta}^{merr}$$
#
# **Note that the second equation is just our notation for the fitted replication <br />
#  (a) Report the intercept and betas.<br />
#  (b) Are the betas realistic position sizes, or do they require huge long-short positions?<br />
#  (c) Report the R-squared.<br />
#  (d) Report the volatility of $\epsilon^{merr}$, (the tracking error.)**
#
#regress HFRI on all factors in merrill factors data set
y= funds["HFRIFWI Index"]
#add const to regression
x = sm.add_constant(factors)
#regress fund on SPY
reg = sm.OLS(y, x, missing='drop').fit()
#find predicted values for later
alpha = reg.params["const"]
beta = reg.params.drop(index='const')
HFRI_pred_Int = alpha + factors @ beta
#show results
reg.params
# <span style="color:#00008B"> (6b) We can see a range of betas that should all be at an appropriate level for replication loadings.
print(f"R Squared: {reg.rsquared}")
# sqrt(12) to annualize
print(f"Tracking Error: {reg.resid.std() * np.sqrt(12)}")
# ### Question 7
# **Let’s examine the replication out-of-sample. Starting with t = 61 month of the sample, do the following: <br />**
# - **Use the previous 60 months of data to estimate the regression equation, (1). This gives time-t estimates of the regression parameters, $\~{\alpha}^{merr}_{t}$ and $\~{\beta}^{merr}_{t}$**
# - **Use the estimated regression parameters, along with the time-t regressor values, $x^{merr}_{t}$, to calculate the time-t replication value that is, with respect to the regression estimate, built “out-of-sample” (OOS)**
# $$\~{r}^{hfri}_{t} = \~{\alpha}^{merr}_{t} + (x^{merr}_{t})^{'} \~{\beta}^{merr}_{t}$$
# - **Step forward to t = 62, and now use t = 2 through t = 61 for the estimation. Re-run the steps above, and continue this process throughout the data series. Thus, we are running a rolling, 60-month regression for each point-in-time.**
#
# **How well does the out-of-sample replication perform with respect to the target?**
date_range = factors['08-2016':'09-2021'].index
oos_fitted = pd.Series(index=date_range, name='OOS_fit')
for date in date_range:
date_prior = pd.DatetimeIndex([date]).shift(periods=-60, freq='M')[0]
rhs = sm.add_constant(factors[date_prior:date])
lhs = funds["HFRIFWI Index"][date_prior:date]
res = sm.OLS(lhs, rhs, drop="missing").fit()
alpha = res.params['const']
beta = res.params.drop(index='const')
x_t = factors.loc[date]
predicted_next_value = alpha + x_t @ beta
oos_fitted[date] = predicted_next_value
#showing returns
oos_fitted['08-2016':].plot(figsize=(14,3))
funds["HFRIFWI Index"].loc['08-2016':].plot()
plt.legend()
plt.show()
#showing total return
(oos_fitted['08-2016':] + 1).cumprod().plot(figsize=(14,3))
(funds["HFRIFWI Index"].loc['08-2016':] + 1).cumprod().plot()
plt.legend()
plt.show()
display((pd.DataFrame([oos_fitted, funds["HFRIFWI Index"].loc['08-2016':]])).T.corr())
# <span style="color:#00008B"> **Solution**: The out of sample result is good and we achieved $96.6\%$ correlation level
# between the replication portfolio and the HFRI.
# ### Question 8
# **We estimated the replications using an intercept. Try the full-sample estimation, but this time
# without an intercept.**<br />
#
# $$r^{hfri}_{t} = \alpha^{merr} + x^{merr}_{t} \beta^{merr} + \epsilon^{merr}_{t}$$
# $$\v{r}^{hfri}_{t} = \v{\alpha}^{merr} + x^{merr}_{t} \v{\beta}^{merr}$$
#
#  **(a) the regression beta. How does it compare to the estimated beta with an intercept, $\hat{\beta}^{merr}$ ?<br />
#  (b) the mean of the fitted value, $\v{r}^{hfri}_{t}$. How does it compare to the mean of the HFRI? <br />
#  (c) the correlations of the fitted values, $\v{r}^{hfri}_{t}$ to the HFRI. How does the correlation compare to that of the fitted values with an intercept, $\hat{r}^{hfri}_{t}$**<br />
#
# **Do you think Merrill and ProShares fit their replicators with an intercept or not?**
#regress HFRI on all factors in merrill factors data set
y= funds["HFRIFWI Index"]
#add const to regression
x = factors
#regress fund on SPY
reg = sm.OLS(y, x, missing='drop').fit()
#show results
reg.params
# +
#HFRI mean
HFRI_mean = funds["HFRIFWI Index"].mean() * 12
#Predicted HFRI mean
beta = reg.params
HFRI_pred_noInt = factors @ beta
HFRI_pred_mean = HFRI_pred_noInt.mean() * 12
print("Actual HFRI mean: ", HFRI_mean)
print("Predicted HFRI mean: ", HFRI_pred_mean)
# -
#compare correlations of fitted HFRI (with no intercept) to actaual HFRI
display((pd.DataFrame([HFRI_pred_noInt, funds["HFRIFWI Index"]])).T.corr())
#compare correlations of fitted HFRI (with intercept) to actaual HFRI
display((pd.DataFrame([HFRI_pred_Int, funds["HFRIFWI Index"]])).T.corr())
#
# <span style="color:#00008B"> **Solution**:
# (a) For all the estimated beta except the negative one(USGG3M Index), they are slightly diffrent. But for the negative estimated beta calculated in the regression with intercept, it changed a lot.<br>
# (b) The predicted HFRI is roughly $16\%$ lower than the actual HFRI mean.<br>
# (c) According to our calculation, they are almost the same. Both are around $92.5\%$.<br>
# We think that Merrill and Proshares should not include the intercept in order to achieve a higher mean returns.
| solutions/hw2/FINM36700_HW2-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Yt3mUS3Jx5qE"
# # Large-scale multi-label text classification
#
# **Author:** [<NAME>](https://twitter.com/RisingSayak), [<NAME>](https://github.com/soumik12345)<br>
# **Date created:** 2020/09/25<br>
# **Last modified:** 2020/12/23<br>
# **Description:** Implementing a large-scale multi-label text classification model.
# + [markdown] id="Qo5Cxt9Ix5qF"
# ## Introduction
#
# In this example, we will build a multi-label text classifier to predict the subject areas
# of arXiv papers from their abstract bodies. This type of classifier can be useful for
# conference submission portals like [OpenReview](https://openreview.net/). Given a paper
# abstract, the portal could provide suggestions for which areas the paper would
# best belong to.
#
# The dataset was collected using the
# [`arXiv` Python library](https://github.com/lukasschwab/arxiv.py)
# that provides a wrapper around the
# [original arXiv API](http://arxiv.org/help/api/index).
# To learn more about the data collection process, please refer to
# [this notebook](https://github.com/soumik12345/multi-label-text-classification/blob/master/arxiv_scrape.ipynb).
# Additionally, you can also find the dataset on
# [Kaggle](https://www.kaggle.com/spsayakpaul/arxiv-paper-abstracts).
# + [markdown] id="bZjJVQCox5qG"
# ## Imports
# + id="mjgi0EpNx5qG"
from tensorflow.keras import layers
from tensorflow import keras
import tensorflow as tf
from sklearn.model_selection import train_test_split
from ast import literal_eval
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# + [markdown] id="m9QFdzHUx5qG"
# ## Perform exploratory data analysis
#
# In this section, we first load the dataset into a `pandas` dataframe and then perform
# some basic exploratory data analysis (EDA).
# + id="Xfhv6xkMx5qH"
arxiv_data = pd.read_csv(
"https://github.com/soumik12345/multi-label-text-classification/releases/download/v0.2/arxiv_data.csv"
)
arxiv_data.head()
# + [markdown] id="ZzD-jc99x5qH"
# Our text features are present in the `summaries` column and their corresponding labels
# are in `terms`. As you can notice, there are multiple categories associated with a
# particular entry.
# + id="XFcyPElIx5qH"
print(f"There are {len(arxiv_data)} rows in the dataset.")
# + [markdown] id="2xythF--x5qI"
# Real-world data is noisy. One of the most commonly observed source of noise is data
# duplication. Here we notice that our initial dataset has got about 13k duplicate entries.
# + id="dVBYRBNtx5qI"
total_duplicate_titles = sum(arxiv_data["titles"].duplicated())
print(f"There are {total_duplicate_titles} duplicate titles.")
# + [markdown] id="dybEIZdyx5qI"
# Before proceeding further, we drop these entries.
# + id="QjoTehzSx5qI"
arxiv_data = arxiv_data[~arxiv_data["titles"].duplicated()]
print(f"There are {len(arxiv_data)} rows in the deduplicated dataset.")
# There are some terms with occurrence as low as 1.
print(sum(arxiv_data["terms"].value_counts() == 1))
# How many unique terms?
print(arxiv_data["terms"].nunique())
# + [markdown] id="gCVZ6veIx5qJ"
# As observed above, out of 3,157 unique combinations of `terms`, 2,321 entries have the
# lowest occurrence. To prepare our train, validation, and test sets with
# [stratification](https://en.wikipedia.org/wiki/Stratified_sampling), we need to drop
# these terms.
# + id="Fex7gjXJx5qJ"
# Filtering the rare terms.
arxiv_data_filtered = arxiv_data.groupby("terms").filter(lambda x: len(x) > 1)
arxiv_data_filtered.shape
# + [markdown] id="ZZUmPLo6x5qJ"
# ## Convert the string labels to lists of strings
#
# The initial labels are represented as raw strings. Here we make them `List[str]` for a
# more compact representation.
# + id="C03tnaGex5qJ"
arxiv_data_filtered["terms"] = arxiv_data_filtered["terms"].apply(
lambda x: literal_eval(x)
)
arxiv_data_filtered["terms"].values[:5]
# + [markdown] id="lwQ2FYMKx5qJ"
# ## Use stratified splits because of class imbalance
#
# The dataset has a
# [class imbalance problem](https://developers.google.com/machine-learning/glossary/#class-imbalanced-dataset).
# So, to have a fair evaluation result, we need to ensure the datasets are sampled with
# stratification. To know more about different strategies to deal with the class imbalance
# problem, you can follow
# [this tutorial](https://www.tensorflow.org/tutorials/structured_data/imbalanced_data).
# For an end-to-end demonstration of classification with imbablanced data, refer to
# [Imbalanced classification: credit card fraud detection](https://keras.io/examples/structured_data/imbalanced_classification/).
# + id="8vJaopjhx5qJ"
test_split = 0.1
# Initial train and test split.
train_df, test_df = train_test_split(
arxiv_data_filtered,
test_size=test_split,
stratify=arxiv_data_filtered["terms"].values,
)
# Splitting the test set further into validation
# and new test sets.
val_df = test_df.sample(frac=0.5)
test_df.drop(val_df.index, inplace=True)
print(f"Number of rows in training set: {len(train_df)}")
print(f"Number of rows in validation set: {len(val_df)}")
print(f"Number of rows in test set: {len(test_df)}")
# + [markdown] id="WN0lc9uJx5qK"
# ## Multi-label binarization
#
# Now we preprocess our labels using the
# [`StringLookup`](https://keras.io/api/layers/preprocessing_layers/categorical/string_lookup)
# layer.
# + id="1_xdRY81x5qK"
terms = tf.ragged.constant(train_df["terms"].values)
lookup = tf.keras.layers.StringLookup(output_mode="multi_hot")
lookup.adapt(terms)
vocab = lookup.get_vocabulary()
def invert_multi_hot(encoded_labels):
"""Reverse a single multi-hot encoded label to a tuple of vocab terms."""
hot_indices = np.argwhere(encoded_labels == 1.0)[..., 0]
return np.take(vocab, hot_indices)
print("Vocabulary:\n")
print(vocab)
# + [markdown] id="9UbiyJ7Ex5qK"
# Here we are separating the individual unique classes available from the label
# pool and then using this information to represent a given label set with 0's and 1's.
# Below is an example.
# + id="LXr5Qrvix5qK"
sample_label = train_df["terms"].iloc[0]
print(f"Original label: {sample_label}")
label_binarized = lookup([sample_label])
print(f"Label-binarized representation: {label_binarized}")
# + [markdown] id="8iYWy9ZPx5qK"
# ## Data preprocessing and `tf.data.Dataset` objects
#
# We first get percentile estimates of the sequence lengths. The purpose will be clear in a
# moment.
# + id="pgwaWvBLx5qL"
train_df["summaries"].apply(lambda x: len(x.split(" "))).describe()
# + [markdown] id="vlIsAmw_x5qL"
# Notice that 50% of the abstracts have a length of 154 (you may get a different number
# based on the split). So, any number close to that value is a good enough approximate for the
# maximum sequence length.
#
# Now, we implement utilities to prepare our datasets.
# + id="2KbaUIcix5qL"
max_seqlen = 150
batch_size = 128
padding_token = "<pad>"
auto = tf.data.AUTOTUNE
def make_dataset(dataframe, is_train=True):
labels = tf.ragged.constant(dataframe["terms"].values)
label_binarized = lookup(labels).numpy()
dataset = tf.data.Dataset.from_tensor_slices(
(dataframe["summaries"].values, label_binarized)
)
dataset = dataset.shuffle(batch_size * 10) if is_train else dataset
return dataset.batch(batch_size)
# + [markdown] id="E8xLzKwnx5qL"
# Now we can prepare the `tf.data.Dataset` objects.
# + id="PkqvcBwgx5qL"
train_dataset = make_dataset(train_df, is_train=True)
validation_dataset = make_dataset(val_df, is_train=False)
test_dataset = make_dataset(test_df, is_train=False)
# + [markdown] id="k4arOINSx5qL"
# ## Dataset preview
# + id="J1cLBfQjx5qL"
text_batch, label_batch = next(iter(train_dataset))
for i, text in enumerate(text_batch[:5]):
label = label_batch[i].numpy()[None, ...]
print(f"Abstract: {text}")
print(f"Label(s): {invert_multi_hot(label[0])}")
print(" ")
# + [markdown] id="gIECoX2Sx5qL"
# ## Vectorization
#
# Before we feed the data to our model, we need to vectorize it (represent it in a numerical form).
# For that purpose, we will use the
# [`TextVectorization` layer](https://keras.io/api/layers/preprocessing_layers/text/text_vectorization).
# It can operate as a part of your main model so that the model is excluded from the core
# preprocessing logic. This greatly reduces the chances of training / serving skew during inference.
#
# We first calculate the number of unique words present in the abstracts.
# + id="qVvcDyNOx5qL"
# Source: https://stackoverflow.com/a/18937309/7636462
vocabulary = set()
train_df["summaries"].str.lower().str.split().apply(vocabulary.update)
vocabulary_size = len(vocabulary)
print(vocabulary_size)
# + [markdown] id="DCCrDEAqx5qL"
# We now create our vectorization layer and `map()` to the `tf.data.Dataset`s created
# earlier.
# + id="XPTNnJLZx5qL"
text_vectorizer = layers.TextVectorization(
max_tokens=vocabulary_size, ngrams=2, output_mode="tf_idf"
)
# `TextVectorization` layer needs to be adapted as per the vocabulary from our
# training set.
with tf.device("/CPU:0"):
text_vectorizer.adapt(train_dataset.map(lambda text, label: text))
train_dataset = train_dataset.map(
lambda text, label: (text_vectorizer(text), label), num_parallel_calls=auto
).prefetch(auto)
validation_dataset = validation_dataset.map(
lambda text, label: (text_vectorizer(text), label), num_parallel_calls=auto
).prefetch(auto)
test_dataset = test_dataset.map(
lambda text, label: (text_vectorizer(text), label), num_parallel_calls=auto
).prefetch(auto)
# + [markdown] id="uzuzjmp9x5qL"
# A batch of raw text will first go through the `TextVectorization` layer and it will
# generate their integer representations. Internally, the `TextVectorization` layer will
# first create bi-grams out of the sequences and then represent them using
# [TF-IDF](https://wikipedia.org/wiki/Tf%E2%80%93idf). The output representations will then
# be passed to the shallow model responsible for text classification.
#
# To learn more about other possible configurations with `TextVectorizer`, please consult
# the
# [official documentation](https://keras.io/api/layers/preprocessing_layers/text/text_vectorization).
#
# **Note**: Setting the `max_tokens` argument to a pre-calculated vocabulary size is
# not a requirement.
# + [markdown] id="EBwEUhYdx5qL"
# ## Create a text classification model
#
# We will keep our model simple -- it will be a small stack of fully-connected layers with
# ReLU as the non-linearity.
# + id="TtrXSndRx5qL"
def make_model():
shallow_mlp_model = keras.Sequential(
[
layers.Dense(512, activation="relu"),
layers.Dense(256, activation="relu"),
layers.Dense(lookup.vocabulary_size(), activation="sigmoid"),
] # More on why "sigmoid" has been used here in a moment.
)
return shallow_mlp_model
# + [markdown] id="02hLU29Gx5qL"
# ## Train the model
#
# We will train our model using the binary crossentropy loss. This is because the labels
# are not disjoint. For a given abstract, we may have multiple categories. So, we will
# divide the prediction task into a series of multiple binary classification problems. This
# is also why we kept the activation function of the classification layer in our model to
# sigmoid. Researchers have used other combinations of loss function and activation
# function as well. For example, in
# [Exploring the Limits of Weakly Supervised Pretraining](https://arxiv.org/abs/1805.00932),
# Mahajan et al. used the softmax activation function and cross-entropy loss to train
# their models.
# + id="E2GhnV1_x5qM"
epochs = 20
shallow_mlp_model = make_model()
shallow_mlp_model.compile(
loss="binary_crossentropy", optimizer="adam", metrics=["categorical_accuracy"]
)
history = shallow_mlp_model.fit(
train_dataset, validation_data=validation_dataset, epochs=epochs
)
def plot_result(item):
plt.plot(history.history[item], label=item)
plt.plot(history.history["val_" + item], label="val_" + item)
plt.xlabel("Epochs")
plt.ylabel(item)
plt.title("Train and Validation {} Over Epochs".format(item), fontsize=14)
plt.legend()
plt.grid()
plt.show()
plot_result("loss")
plot_result("categorical_accuracy")
# + [markdown] id="A-M-d9Eox5qM"
# While training, we notice an initial sharp fall in the loss followed by a gradual decay.
# + [markdown] id="meNmaWDfx5qM"
# ### Evaluate the model
# + id="HlSn4xSpx5qM"
_, categorical_acc = shallow_mlp_model.evaluate(test_dataset)
print(f"Categorical accuracy on the test set: {round(categorical_acc * 100, 2)}%.")
# + [markdown] id="plIfr6kmx5qM"
# The trained model gives us an evaluation accuracy of ~87%.
# + [markdown] id="ZKhcXVOAx5qM"
# ## Inference
#
# An important feature of the
# [preprocessing layers provided by Keras](https://keras.io/guides/preprocessing_layers/)
# is that they can be included inside a `tf.keras.Model`. We will export an inference model
# by including the `text_vectorization` layer on top of `shallow_mlp_model`. This will
# allow our inference model to directly operate on raw strings.
#
# **Note** that during training it is always preferable to use these preprocessing
# layers as a part of the data input pipeline rather than the model to avoid
# surfacing bottlenecks for the hardware accelerators. This also allows for
# asynchronous data processing.
# + id="mKNRsFF7x5qM"
# Create a model for inference.
model_for_inference = keras.Sequential([text_vectorizer, shallow_mlp_model])
# Create a small dataset just for demoing inference.
inference_dataset = make_dataset(test_df.sample(100), is_train=False)
text_batch, label_batch = next(iter(inference_dataset))
predicted_probabilities = model_for_inference.predict(text_batch)
# Perform inference.
for i, text in enumerate(text_batch[:5]):
label = label_batch[i].numpy()[None, ...]
print(f"Abstract: {text}")
print(f"Label(s): {invert_multi_hot(label[0])}")
predicted_proba = [proba for proba in predicted_probabilities[i]]
top_3_labels = [
x
for _, x in sorted(
zip(predicted_probabilities[i], lookup.get_vocabulary()),
key=lambda pair: pair[0],
reverse=True,
)
][:3]
print(f"Predicted Label(s): ({', '.join([label for label in top_3_labels])})")
print(" ")
# + [markdown] id="DrUCFUoPx5qM"
# The prediction results are not that great but not below the par for a simple model like
# ours. We can improve this performance with models that consider word order like LSTM or
# even those that use Transformers ([Vaswani et al.](https://arxiv.org/abs/1706.03762)).
# + [markdown] id="JYjeYts5x5qM"
# ## Acknowledgements
#
# We would like to thank [<NAME>](https://github.com/mattdangerw) for helping us
# tackle the multi-label binarization part and inverse-transforming the processed labels
# to the original form.
| examples/nlp/ipynb/multi_label_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Regression plots
# +
# %matplotlib inline
from __future__ import print_function
from statsmodels.compat import lzip
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.formula.api import ols
# -
# ### Duncan's Prestige Dataset
prestige = sm.datasets.get_rdataset("Duncan", "carData", cache=True).data
prestige.head()
prestige_model = ols('prestige ~ income + education', data = prestige).fit()
print(prestige_model.summary())
# ### Influence plots
fig, ax = plt.subplots(figsize=(12,8))
fig = sm.graphics.influence_plot(prestige_model, ax=ax, criterion='cooks')
# Both contractor and reporter have low leverage but a large residual. PR.engineer has large leverage and small residual. Conductor and minister have high leverage and large residual, and therefore, large influence.
# ### Partial Regression Plots
fig, ax = plt.subplots(figsize=(12,8))
fig = sm.graphics.plot_partregress('prestige', 'income', ['income', 'education'], data = prestige, ax=ax)
fig, ax = plt.subplots(figsize=(12,14))
fig = sm.graphics.plot_partregress('prestige', 'income', ['education'], data = prestige, ax = ax)
# As you can see the partial regression plot confirms the influence of conductor, minister, and RR.engineer on the partial relationship between income and prestige. The cases greatly decrease the effect of income on prestige. Dropping these cases confirms this.
subset = ~ prestige.index.isin(['conductor', 'PR.engineer', 'minister'])
prestige_model2 = ols('prestige ~ income + education', data = prestige, subset = subset).fit()
print(prestige_model2.summary())
# For a quick check of all the regressors, you can use plot_partregress_grid. These plots will not label the
# points, but you can use them to identify problems and then use plot_partregress to get more information.
fig = plt.figure(figsize=(12,8))
fig = sm.graphics.plot_partregress_grid(prestige_model, fig=fig)
# ### Component-Component plus Residual (CCPR) Plots
fig, ax = plt.subplots(figsize=(12,8))
fig = sm.graphics.plot_ccpr(prestige_model, 'education', ax = ax)
# As you can see the relationship between the variation in prestige explained by education conditional on income seems to be linear, though you can see there are some observations that are exerting considerable influence on the relationship. We can quickly look at more than one variable.
fig = plt.figure(figsize=(12,8))
fig = sm.graphics.plot_ccpr_grid(prestige_model, fig=fig)
# ### Regression plot
#
# The plot_regress_exog function is a convenience function that gives a 2x2 plot containing the dependent variable and fitted values with confidence intervals vs. the independent variable chosen, the residuals of the model vs. the chosen independent variable, a partial regression plot, and a CCPR plot. This function can be used for quickly checking modeling assumptions with respect to a single regressor.
fig = plt.figure(figsize=(12, 8))
fig = sm.graphics.plot_regress_exog(prestige_model, 'education', fig=fig)
# ### Fit plot
#
# The plot_fit function plots the fitted values versus a chosen independent variable. It includes prediction confidence intervals and optionally plots the true dependent variable.
fig, ax = plt.subplots(figsize=(12,8))
fig = sm.graphics.plot_fit(prestige_model, 'education', ax=ax)
# ## Statewide Crime 2009 Dataset
dta = sm.datasets.statecrime.load_pandas().data
dta.head(10)
crime_model = ols('murder ~ hs_grad + poverty + single + urban', data = dta).fit()
print(crime_model.summary())
# ### Partial Regression Plots
fig = plt.figure(figsize=(12,8))
fig = sm.graphics.plot_partregress_grid(crime_model, fig=fig)
fig, ax = plt.subplots(figsize = (12,8))
fig = sm.graphics.plot_partregress('murder', 'hs_grad', ['urban', 'poverty', 'single'], ax = ax, data = dta)
# ### Leverage-Resid2 Plot
#
# Closely related to the influence_plot is the leverage-resid2 plot.
fig, ax = plt.subplots(figsize=(8,6))
fig = sm.graphics.plot_leverage_resid2(crime_model, ax=ax)
# ### Influence plot
fig, ax = plt.subplots(figsize=(8,6))
fig = sm.graphics.influence_plot(crime_model, ax=ax)
# ### Using robust regression to correct for outliers.
#
# Part of the problem here in recreating the Stata results is that M-estimators are not robust to leverage points. MM-estimators should do better with this examples.
from statsmodels.formula.api import rlm
rob_crime_model = rlm('murder ~ urban + poverty + single + hs_grad', data = dta, M=sm.robust.norms.TukeyBiweight(3)).fit(conv='weights')
print(rob_crime_model.summary())
# There isn't yet an influence diagnostics method as part of RLM, but we can recreate them.
weights = rob_crime_model.weights
idx = weights > 0
X = rob_crime_model.model.exog[idx.values]
ww = weights[idx] / weights[idx].mean()
hat_matrix_diag = ww*(X*np.linalg.pinv(X).T).sum(1)
resid = rob_crime_model.resid
resid2 = resid**2
resid2 /= resid2.sum()
nobs = int(idx.sum())
hm = hat_matrix_diag.mean()
rm = resid2.mean()
# +
from statsmodels.graphics import utils
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(resid2[idx], hat_matrix_diag, 'o')
ax = utils.annotate_axes(range(nobs), labels = rob_crime_model.model.data.row_labels[idx],
points = lzip(resid2[idx], hat_matrix_diag,), offset_points = [(-5,5)]*nobs,
size='large', ax=ax)
ax.set_xlabel('resid2')
ax.set_ylabel('leverage')
ylim = ax.get_ylim()
ax.vlines(rm, *ylim)
xlim = ax.get_xlim()
ax.hlines(hm, *xlim)
ax.margins(0,0)
# -
| Regression Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **DSFM Exercise**: Natural language processing
# Creator: [Data Science for Managers - EPFL Program](https://www.dsfm.ch)
# Source: [https://github.com/dsfm-org/code-bank.git](https://github.com/dsfm-org/code-bank.git)
# License: [MIT License](https://opensource.org/licenses/MIT). See open source [license](LICENSE) in the Code Bank repository.
# -------------
# ## Overview
# In this exercise, we are revisiting the EPFL course book data. We would like to understand how similar courses are based on their textual description. Think about this case: maybe you liked a course very much and would now like to take the most similar one to that.
#
# ### Business Objective:
#
# * To discover similarity relationship between EPFL courses based on their textual description
#
# ### Learning Objectives:
#
# * Getting familiar with text preprocessing facilities in the `nltk` library
# * Understanding intuition behind different vector space models to work with text data, e.g. TFIDF
# * Learning how to transform a raw corpus into the vector space model of choice
# * Learning how to query similar documents to a focal document in a given space
# * Learning how to visualize text data from high-dimensional space into low dimansions for visualization
# -------
# # Part 0: Setup
# +
# Standard imports
import pandas as pd
# Natural Language Toolkit (NLTK) and spaCy
import nltk
nltk.download('wordnet')
import spacy
# Sklearn TFIDF function and PCA
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.decomposition import PCA
# Plotting packages
import matplotlib.pyplot as plt
# Python math package
import math
# Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# +
# Define constant(s)
SEED = 42
# -
# # Part 1: Load .csv data
#
# In this part, simply load the EPFL course file from `data/epfl_description.csv`.
# **Q 1**: Load EPFL course data. Look at the shape and the first 5 rows. What shape does the data have?
# **Q 2**: Concatenate the course title and the course description column. Why is this useful?
# # Part 2: Clean data
# **Q 1**: Draw a random sample of 5 course descriptions and look at the entire description. Are there any issues with the text data? If so, what are they?
#
# Hint: Look at the element in row 8.
# Some of the issues that require cleaning include:
#
# - `\r` character is the carriage return
# - punctuation marks like `.`, `,`, `?`, `!`, etc.
# - quotation marks and other symbols like `$`, `(`, `)`, etc.
# - etc.
# **Q 2**: Remove the parts of the text identified above. Also remove multiple white spaces. How did the element in row 8 change?
#
# Hint: use a "regular expressions" (regex), which defines a search pattern for strings - a very handy tool for pre-processing text. You can visit https://regex101.com/ to test your regex expressions.
# **Q 3**: Clean data by removing rows with missing data in any column. How many clean rows are left?
# # Part 3: Tokenize and lemmatize course descriptions
#
# For grammatical reasons, documents are going to use different forms of a word, such as organize, organizes, and organizing. The goal of both stemming and lemmatization is to reduce inflectional forms and sometimes derivationally related forms of a word to a common base form. For example:
#
# - am, are, is $\Rightarrow$ be
# - car, cars, car's, cars' $\Rightarrow$ car
#
# For details about lemmatization and stemming visit: https://nlp.stanford.edu/IR-book/html/htmledition/stemming-and-lemmatization-1.html
# **Q 1**: Define a simple function that takes a course description as input and outputs the tokenized and lemmatized text as a list.
# **Q 2**: Apply the function to the course description in your Pandas dataframe.
#
# Hint: Use the Pandas `apply` function. Look at the second row - what changed? Did lemmatization work?
# # Part 4: Create a term frequency inverse document frequency (TFIDF) matrix
#
# We now have to ensure that the text description is stored as a string in our dataframe, not as a list. In the code below, replace the variable names with the ones you are using.
# Transform list data to text data
df_clean['description_lemmatized_text'] = df_clean['description_lemmatized'].str.join(' ')
df_clean['description_lemmatized_text'].head()
# Extract all the text data
data = df_clean['description_lemmatized_text']
len(data)
# **Q 1**: Fit and transform your text data using TFIDF.
#
# Hint: use the `TfidfVectorizer()` function in sklearn with the parameter `max_features = 400`.
# **Q 2**: What shape does the TFIDF matrix have? What's the meaning of the number of columns? Use the `toarray()` function to show some of the TFIDF entries.
# # Part 5: Apply a principal component analysis (PCA)
#
# We now project the high-dimensional TFIDF matrix into its 2 principal components.
# **Q 1**: Run a PCA on the TFIDF matrix. Hint: use the `PCA.fit_transform()` function in sklearn.
# **Q 2**: What's the shape of the PCA output? Why?
# **Q 3**: This is done for you: add the PCA values to the cleaned dataframe. We want a dataframe with the course name and the PCA values.
# # Part 6: Visualize how similar EPFL courses to each other
#
# Now we returning to the initial business objective: to discover similarity relationship between EPFL courses based on their textual description.
# **Q 1**: Visualize the PCA values using a simple scatter plot. Each dot represents a course.
# **Q 2**: What's the most similar course to COM-421? Compute the Euclidean distance between COM-421 and every other course. To do so, you can use this function below:
def euclideanDistance(p1, p2):
"""
Compute euclidean distance
Parameter:
p1 (list): input point defined as a [x,y] list
p2 (list): input point defined as a [x,y] list
Returns:
float: euclidean distance between p1 and p2
"""
return math.sqrt( ((p1[0]-p2[0])**2)+((p1[1]-p2[1])**2) )
| exercises/nlp-text/nlp-text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vadManuel/Machine-Learning-UCF/blob/master/Homework/hw3/hw3_training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="S3OqUKFZL5eJ" colab_type="code" outputId="5705b08a-1432-4164-b8f8-1f25eabfba84" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="o5_-cA4JHOrY" colab_type="code" colab={}
import time
import numpy as np
import cv2
from pathlib import Path
import os
import matplotlib.pyplot as plt
# + id="1806WXhsMkmT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 522} outputId="f084d9b0-0254-4d49-bb21-655468af4b89"
from tensorflow.python.client import device_lib
device_lib.list_local_devices()
# + id="G3OvDfv-SFio" colab_type="code" outputId="9da9796f-483f-4ef6-9617-3b22e4d2b934" colab={"base_uri": "https://localhost:8080/", "height": 119}
rootdir = Path('gdrive/My Drive/RuTanks7000_v1/dataset/RuTanks7000_v1')
labels = [label.name for label in rootdir.glob('train/*')]
for label in labels:
train_len = len(list(rootdir.glob(f'train/{label}/*')))
test_len = len(list(rootdir.glob(f'test/{label}/*')))
print('%s%sTrain: %s, Test: %s' % (label, ' '*(11-len(label))+'| ', train_len, test_len))
# + id="OmZM6-0qBXxJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="ffe122b2-38a4-49ab-ae45-2ef745d93608"
for i,label in enumerate(labels):
s = 'gdrive/My Drive/RuTanks7000_v1/dataset/RuTanks7000_v1/train/%s/%s_2.jpg'%(label,label)
image = cv2.imread(s, cv2.IMREAD_COLOR)
plt.subplot(2,3,i+1)
plt.imshow(image)
plt.axis('off')
plt.title(label)
plt.show()
# + id="HfNGFVTqR60u" colab_type="code" outputId="d131c7fc-ca88-44b1-d4ce-e9d239e7928f" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# https://github.com/UgurUysal86/MLS4MIL/tree/master/Image%20Classification
# imports
import os
from keras.applications import NASNetLarge
from keras import models, layers, optimizers, backend
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint, ReduceLROnPlateau, TensorBoard
import matplotlib.pyplot as plt
import numpy as np
from timeit import default_timer as timer
def main():
start = timer()
level1() # Training the classifier only
level2() # Training the pretrained model + the trained classifier from level 1
end = timer()
print("Time elapsed in minutes: ", ((end - start)/60))
# Setting Parameters ##################################################################
# image settings
img_height,img_width = 331, 331 # For NASNetLarge
# classes
classnames = labels.copy()
classes = len(classnames)
# path settings
path = 'gdrive/My Drive/RuTanks7000_v1/'
dataset_path = path+'dataset/RuTanks7000_v1/'
weights_path = path+'trained_models/trained_weights/weights_temp.h5'
model_path = path+'trained_models/RuTanks7000_v1_'
TensorBoardLogDir = path+'logs'
nbrTrainImages = 7000 # per class
nbrTestImages = 0 # Value gets accurate after counting (Total Number of test images)
for ImagesClass in os.listdir(dataset_path+'test/'):
nbrTestImages += len(os.listdir(dataset_path+'test/'+ImagesClass))
# unfreezing the base network up to a specific layer in Level2:
freezeUptoLayer = "normal_add_1_15" # NASNetLarge
# hyperparameters
learning_rate = 0.0002 # Learning_rate in Level 2 = learning_rate/10
lr_decay = 0.0001
batch = 64
fcLayer1 = 32
dropout = 0.5
epochsL1 = 10
patiencel1 = 1
factorL1 = 0.5
epochsL2 = 10
patiencel2 = 1
factorL2 = 0.5
verbose_train = 1
# datagenerators https://keras.io/preprocessing/image/
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=30,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
fill_mode='nearest',
horizontal_flip=True)
train_generator = train_datagen.flow_from_directory(
dataset_path+'train',
target_size=(img_height, img_width),
batch_size=batch,
shuffle=True,
classes=classnames,
class_mode='categorical')
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_directory(
dataset_path+'test',
target_size=(img_height, img_width),
batch_size=1,
shuffle=True,
classes=classnames,
class_mode='categorical')
#########################################################################################
# function to plot results of model performance
def plot(h,t,e):
history_dict = h[0]
loss_values = history_dict['loss']
validation_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
validation_acc_values = history_dict['val_acc']
epochs_range = range(1, e + 1)
# Plotting Training and Validation loss of the corresponding Model
plt.plot(epochs_range, loss_values, 'bo', label='Training loss')
plt.plot(epochs_range, validation_loss_values, 'ro', label='Validation loss')
plt.title('Training and validation loss of ' + t)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.yticks(np.arange(0, 3.1, step=0.2))
plt.legend()
plt.show()
# Plotting Training and Validation accuracy of the corresponding Model
plt.plot(epochs_range, acc_values, 'bo', label='Training accuracy')
plt.plot(epochs_range, validation_acc_values, 'ro', label='Validation accuracy')
plt.title('Training and validation accuracy of ' + t)
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.yticks(np.arange(0.3, 1.1, step=0.1))
plt.legend()
plt.show()
# LEVEL1 - Training of densely connected layers
def level1():
# Building the model using the pretrained model
conv_base1 = NASNetLarge(weights='imagenet', include_top=False, input_shape=(img_height, img_width, 3))
print("\n### LEVEL1 ###\npretrained network:")
conv_base1.summary()
model = models.Sequential()
model.add(conv_base1)
model.add(layers.GlobalAveragePooling2D())
model.add(layers.Dense(fcLayer1, activation='relu'))
model.add(layers.Dropout(dropout))
model.add(layers.Dense(classes, activation='softmax'))
# freezing the base network
print("trainable layers bevor freezing:", int(len(model.trainable_weights)/2)) # weights = weights + bias = 2 pro layer
conv_base1.trainable = False
print("trainable layers after freezing:", int(len(model.trainable_weights)/2))
print("\npretrained network + densely connected classifier")
model.summary()
# training the added layers only
model.compile(loss='categorical_crossentropy', optimizer=optimizers.RMSprop(lr=learning_rate, decay=lr_decay), metrics=['acc'])
callbacks_list_L1 = [ModelCheckpoint(filepath=weights_path, save_weights_only=True, monitor='val_acc', verbose=1, save_best_only=True),
ReduceLROnPlateau(monitor='val_acc', factor=factorL1, patience=patiencel1, verbose=1),
TensorBoard(log_dir=TensorBoardLogDir+'\\level1')]
print("\n### Level1 Training ... ")
# training the model
history = model.fit_generator(
train_generator,
steps_per_epoch=(nbrTrainImages * classes) // (batch*10),
epochs=epochsL1,
callbacks=callbacks_list_L1,
validation_data=test_generator,
validation_steps=nbrTestImages,
verbose=verbose_train)
history_val1 = [history.history] # saving all results of the final test
plot(history_val1, "LEVEL1:", epochsL1)
print("\n### LEVEL1 Training finished successfully ###")
print("\nLoading trained weights from " + weights_path + " ...")
model.load_weights(weights_path)
model.compile(loss='categorical_crossentropy', optimizer=optimizers.RMSprop(lr=learning_rate), metrics=['acc'])
print("\n### Saving Level1 Model to ", model_path+'l1.h5', " ... ")
model.save(model_path+'l1.h5')
# LEVEL2 - Training pretrained network and trained densely connected layers
def level2():
# Destroying the current TF graph - https://keras.io/backend/
backend.clear_session()
print("\n### LEVEL2 ###")
conv_base2 = NASNetLarge(weights='imagenet', include_top=False, input_shape=(img_height, img_width, 3))
model2 = models.Sequential()
model2.add(conv_base2)
model2.add(layers.GlobalAveragePooling2D())
model2.add(layers.Dense(fcLayer1, activation='relu'))
model2.add(layers.Dropout(dropout))
model2.add(layers.Dense(classes, activation='softmax'))
print("\nLoading trained weights from " + weights_path + " ...")
model2.load_weights(weights_path)
# unfreezing the base network up to a specific layer:
if freezeUptoLayer == "":
conv_base2.trainable = True
print ("\ntrainable layers: ",int(len(model2.trainable_weights) / 2))
else:
print("\ntrainable layers before unfreezing the base network up to " + freezeUptoLayer + ": ",int(len(model2.trainable_weights) / 2)) # weights = weights + bias = 2 pro layer
conv_base2.trainable = True
set_trainable = False
for layer in conv_base2.layers:
if layer.name == freezeUptoLayer: set_trainable = True
if set_trainable: layer.trainable = True
else: layer.trainable = False
print("trainable layers after the base network unfreezed from layer " + freezeUptoLayer + ": ", int(len(model2.trainable_weights)/2))
print("\nLEVEL2 Model after unfreezing the base network")
model2.summary()
model2.compile(loss='categorical_crossentropy', optimizer=optimizers.RMSprop(lr=learning_rate/10, decay=lr_decay), metrics=['acc'])
print ("\n### Validating ... ")
val_loss, val_acc = model2.evaluate_generator(test_generator, steps=nbrTestImages, verbose=0)
print('Validation Results before training unfreeze layers and trained densely connected layers:\nValidation loss:',val_loss,",",'Validation accuracy:', val_acc, "\n")
# Jointly training both the unfreeze layers and the added trained densely connected layers
callbacks_list_L2 = [ModelCheckpoint(filepath=model_path+'l2.h5', save_weights_only=False, monitor='val_acc', verbose=1, save_best_only=True),
ReduceLROnPlateau(monitor='val_acc', factor=factorL2, patience=patiencel2, verbose=1),
TensorBoard(log_dir=TensorBoardLogDir+'\\level2')]
print ("\n### Level2 Training ... ")
history = model2.fit_generator(
train_generator,
steps_per_epoch=(nbrTrainImages * classes) // (batch*10),
epochs=epochsL2,
callbacks=callbacks_list_L2,
validation_data=test_generator,
validation_steps=nbrTestImages,
verbose=verbose_train)
history_val2 = [history.history] # saving all results of the final test
plot(history_val2, "LEVEL2:", epochsL2)
print("\n###LEVEL2 Training finished successfully ###")
main()
# References
# <NAME>. (2018). Deep learning with Python. Section 5.3 - Using a pretrained convnet.
# + id="Xv8L5KOAExQU" colab_type="code" colab={}
| Homework/hw3/hw3_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example jupyternotebook workflow for pyjugex
#
# This notebook demonstrates how a gene differential analysis can be carried out with pyjugex. Also demonstrated is the use of atlas-core to retrieve probability maps.
# ## Download probability maps via atlas core (optional)
#
# You may choose to use `ebrains_atlascore` to download probabilistic maps. Alternatively, you can use a variety of tools, such as `wget` on command line or `requests` in python to download the necessary probabilistic maps.
#
# It should be noted that Allen Brain provides data in MNI152 reference space. In normal circumstances, you should probably also use probabilistic maps in that space.
# !pip install ebrains_atlascore
from ebrains_atlascore import regions
from ebrains_atlascore.util.hemisphere import Hemisphere
from ebrains_atlascore.region import Region
pmap_hoc1 = regions.get_probability_map_for_region(Region('Area-hOc1', parcellation='JuBrain Cytoarchitectonic Atlas', referencespace='MNI152'), Hemisphere.LEFT.value, 0.2)
pmap_hoc2 = regions.get_probability_map_for_region(Region('Area-hOc2', parcellation='JuBrain Cytoarchitectonic Atlas', referencespace='MNI152'), Hemisphere.LEFT.value, 0.2)
# As `nibabel` does not load file in memory, write them to disk.
with open('hoc1.nii', 'wb') as fp:
fp.write(pmap_hoc1.data)
with open('hoc2.nii', 'wb') as fp:
fp.write(pmap_hoc2.data)
# ## pyjugex analysis
#
# This section details how one may set up parameters for gene differential analysis.
# +
# install pyjugex and import dependencies
# !pip install pyjugex
import pyjugex
import nibabel as nib
# +
# setup parameters
gene_list=['MAOA','TAC1']
nii1 = nib.load('hoc1.nii')
nii2 = nib.load('hoc2.nii')
# +
# load parameters and setup analysis
analysis = pyjugex.analysis(
n_rep=1000,
gene_list=gene_list,
roi1 = nii1,
roi2 = nii2
)
# +
# prior to analysis, one can retrieve the coordinates of the probes in MNI152 space
filtered_coord = analysis.get_filtered_coord()
assert(len(filtered_coord['roi1']) == 12)
assert(len(filtered_coord['roi2']) == 11)
# -
analysis.run() # Go grab a coffee
# +
# results of the differential analysis is saved in the result object
maoa = analysis.anova.result.get('MAOA')
tac1 = analysis.anova.result.get('TAC1')
assert(0.95 <= maoa <= 1.0)
assert(0.35 <= tac1 <= 0.55)
# +
# alter the parameter and start another run
analysis.n_rep = 10000
analysis.run() # Really go grab a coffee
# +
maoa = analysis.anova.result.get('MAOA')
tac1 = analysis.anova.result.get('TAC1')
assert(0.95 <= maoa <= 1.0)
assert(0.35 <= tac1 <= 0.52)
| nb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cflows
# language: python
# name: cflows
# ---
# ## Config
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '3'
# +
# %load_ext autoreload
# %autoreload 2
from pathlib import Path
from experiment import data_path, device
model_name = 'mnist-cef-joint'
checkpoint_path = data_path / 'cef_models' / model_name
gen_path = data_path / 'generated' / model_name
# -
# ## Data
# +
from torch.utils.data import DataLoader, random_split
from torchvision import transforms, datasets
# Pad images from 28x28 to 32x32 to make it a power of 2
transform = transforms.Compose([
transforms.Pad(2),
transforms.ToTensor(),
])
train_data = datasets.MNIST(
root=data_path, train=True, download=True, transform=transform)
train_data, val_data = random_split(train_data, [50000, 10000])
test_data = datasets.MNIST(
root=data_path, train=False, download=True, transform=transform)
# -
# ## Model
# +
from nflows import cef_models
flow = cef_models.MNISTCEFlow(128).to(device)
# -
# ## Train
# +
import torch.optim as opt
from experiment import train_injective_flow
optim = opt.Adam(flow.parameters(), lr=0.001)
scheduler = opt.lr_scheduler.CosineAnnealingLR(optim, 1000)
def weight_schedule():
'''Yield epoch weights for likelihood and recon loss, respectively'''
for _ in range(1000):
yield 0.01, 100000
scheduler.step()
train_loader = DataLoader(train_data, batch_size=512, shuffle=True, num_workers=30)
val_loader = DataLoader(val_data, batch_size=512, shuffle=True, num_workers=30)
train_injective_flow(flow, optim, scheduler, weight_schedule, train_loader, val_loader,
model_name, checkpoint_path=checkpoint_path, checkpoint_frequency=10)
# -
# ## Generate Images
# +
from experiment import save_samples
save_samples(flow, num_samples=10000, gen_path=gen_path, checkpoint_epoch=-1, batch_size=512)
| experiments/mnist-cef-joint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
# Cute Gists
# +
% matplotlib inline
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
# +
# Create grid coordinates for plotting
Z = np.zeros((3, 5))
# Alias/abstract lambda function
npenum = lambda x: np.ndenumerate(x)
# Calculate Z-values (RSS) based on grid of coefficients
for (i, j), v in npenum(Z):
print(i, j, v)
# +
def min_max(data):
from operator import itemgetter
from collections import namedtuple
min_index, min_value = min(enumerate(data), key=itemgetter(1))
max_index, max_value = max(enumerate(data), key=itemgetter(1))
min_max_tup = namedtuple('min_max', ['min_index', 'min_value', 'max_index', 'max_value'])
return min_max_tup(min_index, min_value, max_index, max_value)
tup = min_max([1, 2, 3, 4, 5])
print('min[{}]: {}, max[{}]: {}'.format(tup.min_index, tup.min_value, tup.max_index, tup.max_value))
# -
def xx_yy_grid(X1, X2):
x_min, x_max = np.min(X1), np.max(X1)
y_min, y_max = np.min(X2), np.max(X2)
# Create a coordinate grid - take max. feature value for range
xr = np.arange(int(x_min*(1-margin)), int(x_max*(1+margin)))
yr = np.arange(int(y_min*(1-margin)), int(y_max*(1+margin)))
xx, yy = np.meshgrid(xr, yr, indexing='xy')
Z = np.zeros((xr.size, yr.size))
return xx, yy, Z
# +
for (i,j), v in np.ndenumerate(Z):
Z[i,j] =(regr.intercept_ + xx[i,j]*regr.coef_[0] + yy[i,j]*regr.coef_[1])
# -
# Plot the decision boundary by assigning a color in the color map
# to each mesh point.
mesh_step_size = .01 # step size in the mesh
plot_symbol_size = 50
x_min, x_max = X_mat[:, 0].min() - 1, X_mat[:, 0].max() + 1
y_min, y_max = X_mat[:, 1].min() - 1, X_mat[:, 1].max() + 1
xx, yy = numpy.meshgrid(numpy.arange(x_min, x_max, mesh_step_size),
numpy.arange(y_min, y_max, mesh_step_size))
Z = clf.predict(numpy.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light)
# Plot training points
plt.scatter(X_mat[:, 0], X_mat[:, 1], s=plot_symbol_size, c=y, cmap=cmap_bold, edgecolor = 'black')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
# +
from collections import namedtuple
my_namedtuple = namedtuple('field_name', ['x', 'y', 'z', 'bla', 'blub'])
p = my_namedtuple(1, 2, 3, 4, 5)
print(p.x, p.y, p.z)
# -
# ###
list(map(np.shape, (X_train, X_test, y_train, y_test)))
# ###
# +
# %%file cmd_line_args_1_sysarg.py
import sys
def error(msg):
"""Prints error message, sends it to stderr, and quites the program."""
sys.exit(msg)
args = sys.argv[1:] # sys.argv[0] is the name of the python script itself
try:
arg1 = int(args[0])
arg2 = args[1]
arg3 = args[2]
print("Everything okay!")
except ValueError:
error("First argument must be integer type!")
except IndexError:
error("Requires 3 arguments!")
# -
# %run cmd_line_args_1_sysarg.py
# ###
# +
import time
print('current time: {} - {}'.format(time.strftime("%d/%m/%Y"), time.strftime("%H:%M:%S")))
# -
# ###
# +
def subtract(a, b):
"""
Subtracts second from first number and returns result.
>>> subtract(10, 5)
5
>>> subtract(11, 0.7)
10.3
"""
return a-b
if __name__ == "__main__": # is 'false' if imported
import doctest
doctest.testmod()
print('ok')
# -
# ###
# +
import os
import shutil
import glob
# working directory
c_dir = os.getcwd() # show current working directory
os.listdir(c_dir) # shows all files in the working directory
os.chdir('~/Data') # change working directory
# get all files in a directory
glob.glob('/Users/sebastian/Desktop/*')
# e.g., ['/Users/sebastian/Desktop/untitled folder', '/Users/sebastian/Desktop/Untitled.txt']
# walk
tree = os.walk(c_dir)
# moves through sub directories and creates a 'generator' object of tuples
# ('dir', [file1, file2, ...] [subdirectory1, subdirectory2, ...]),
# (...), ...
#check files: returns either True or False
os.exists('../rel_path')
os.exists('/home/abs_path')
os.isfile('./file.txt')
os.isdir('./subdir')
# file permission (True or False
os.access('./some_file', os.F_OK) # File exists? Python 2.7
os.access('./some_file', os.R_OK) # Ok to read? Python 2.7
os.access('./some_file', os.W_OK) # Ok to write? Python 2.7
os.access('./some_file', os.X_OK) # Ok to execute? Python 2.7
os.access('./some_file', os.X_OK | os.W_OK) # Ok to execute or write? Python 2.7
# join (creates operating system dependent paths)
os.path.join('a', 'b', 'c')
# 'a/b/c' on Unix/Linux
# 'a\\b\\c' on Windows
os.path.normpath('a/b/c') # converts file separators
# os.path: direcory and file names
os.path.samefile('./some_file', '/home/some_file') # True if those are the same
os.path.dirname('./some_file') # returns '.' (everythin but last component)
os.path.basename('./some_file') # returns 'some_file' (only last component
os.path.split('./some_file') # returns (dirname, basename) or ('.', 'some_file)
os.path.splitext('./some_file.txt') # returns ('./some_file', '.txt')
os.path.splitdrive('./some_file.txt') # returns ('', './some_file.txt')
os.path.isabs('./some_file.txt') # returns False (not an absolute path)
os.path.abspath('./some_file.txt')
# create and delete files and directories
os.mkdir('./test') # create a new direcotory
os.rmdir('./test') # removes an empty direcotory
os.removedirs('./test') # removes nested empty directories
os.remove('file.txt') # removes an individual file
shutil.rmtree('./test') # removes directory (empty or not empty)
os.rename('./dir_before', './renamed') # renames directory if destination doesn't exist
shutil.move('./dir_before', './renamed') # renames directory always
shutil.copytree('./orig', './copy') # copies a directory recursively
shutil.copyfile('file', 'copy') # copies a file
# Getting files of particular type from directory
files = [f for f in os.listdir(s_pdb_dir) if f.endswith(".txt")]
# Copy and move
shutil.copyfile("/path/to/file", "/path/to/new/file")
shutil.copy("/path/to/file", "/path/to/directory")
shutil.move("/path/to/file","/path/to/directory")
# Check if file or directory exists
os.path.exists("file or directory")
os.path.isfile("file")
os.path.isdir("directory")
# Working directory and absolute path to files
os.getcwd()
os.path.abspath("file")
# -
# ### Closure
# +
def create_message(msg_txt):
def _priv_msg(message): # private, no access from outside
print("{}: {}".format(msg_txt, message))
return _priv_msg # returns a function
new_msg = create_message("My message")
# note, new_msg is a function
new_msg("Hello, World")
# -
# ### Snippets
http://nbviewer.jupyter.org/github/rasbt/python_reference/blob/master/python_patterns/patterns.ipynb#Data-and-time-basics
# ### Regex
http://nbviewer.jupyter.org/github/rasbt/python_reference/blob/master/tutorials/useful_regex.ipynb
# ### Markdown
https://medium.com/ibm-data-science-experience/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed
# ###
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
x = np.linspace(0, 20, 1000) # 100 evenly-spaced values from 0 to 50
y = np.sin(x)
fig = plt.figure()
ax1 = fig.add_subplot(1,2,1) # 1 row, 2 cols, graphic 1
ax2 = fig.add_subplot(1,2,2) # graphic 2
ax1.plot(x, y)
ax2.hist(np.random.randn(1000), alpha=0.5, histtype='stepfilled')
ax2.hist(0.75*np.random.randn(1000)+1, alpha=0.5, histtype='stepfilled')
plt.show();
# -
# ###
# foo <font color='red'>bar</font> foo
# ### Matplotlib
http://www.labri.fr/perso/nrougier/teaching/matplotlib/
# +
def f(x,y): return (1-x/2+x**5+y**3)*np.exp(-x**2-y**2)
n = 256
x = np.linspace(-3,3,n)
y = np.linspace(-3,3,n)
X,Y = np.meshgrid(x,y)
plt.contourf(X, Y, f(X,Y), 8, alpha=.75, cmap='jet')
C = plt.contour(X, Y, f(X,Y), 8, colors='black')
plt.show();
# -
n = 20
Z = np.random.uniform(0, 1, n)
plt.pie(Z)
plt.show();
# +
from matplotlib import cm
plt.axes([0,0,1,1])
N = 20
theta = np.arange(0.0, 2*np.pi, 2*np.pi/N)
radii = 10*np.random.rand(N)
width = np.pi/4*np.random.rand(N)
bars = plt.bar(theta, radii, width=width, bottom=0.0)
for r, bar in zip(radii, bars):
bar.set_facecolor(cm.jet(r/10.))
bar.set_alpha(0.5)
plt.show()
# +
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = Axes3D(fig)
X = np.arange(-4, 4, 0.25)
Y = np.arange(-4, 4, 0.25)
X, Y = np.meshgrid(X, Y)
R = np.sqrt(X**2 + Y**2)
Z = np.sin(R)
ax.plot_surface(X, Y, Z, rstride=1, cstride=1, cmap='hot')
plt.show();
# +
eqs = []
eqs.append((r"$W^{3\beta}_{\delta_1 \rho_1 \sigma_2} = U^{3\beta}_{\delta_1 \rho_1} + \frac{1}{8 \pi 2} \int^{\alpha_2}_{\alpha_2} d \alpha^\prime_2 \left[\frac{ U^{2\beta}_{\delta_1 \rho_1} - \alpha^\prime_2U^{1\beta}_{\rho_1 \sigma_2} }{U^{0\beta}_{\rho_1 \sigma_2}}\right]$"))
eqs.append((r"$\frac{d\rho}{d t} + \rho \vec{v}\cdot\nabla\vec{v} = -\nabla p + \mu\nabla^2 \vec{v} + \rho \vec{g}$"))
eqs.append((r"$\int_{-\infty}^\infty e^{-x^2}dx=\sqrt{\pi}$"))
eqs.append((r"$E = mc^2 = \sqrt{{m_0}^2c^4 + p^2c^2}$"))
eqs.append((r"$F_G = G\frac{m_1m_2}{r^2}$"))
plt.axes([0.025,0.025,0.95,0.95])
for i in range(24):
index = np.random.randint(0,len(eqs))
eq = eqs[index]
size = np.random.uniform(12,32)
x,y = np.random.uniform(0,1,2)
alpha = np.random.uniform(0.25,.75)
plt.text(x, y, eq, ha='center', va='center', color="#11557c", alpha=alpha,
transform=plt.gca().transAxes, fontsize=size, clip_on=True)
plt.xticks([]), plt.yticks([])
# savefig('../figures/text_ex.png',dpi=48)
plt.show();
# -
# ### Import warnings
import warnings
warnings.filterwarnings(action="ignore", module="scipy", message="^internal gelsd")
plt.gca().set_aspect('equal')
# +
### Double loop: np.ndenumerate(Z)
# -
# !find /Users/frank/Documents/Python_workspace/ISLR_python/Notebooks/data/*.csv
# +
import seaborn as sns
# View color palette
colors = ['#FFFF00', '#00AAFF', '#000000', '#FF00AA']
cmap_bold = sns.color_palette(colors)
sns.palplot(cmap_bold)
# Set color palette
sns.set_palette("husl")
sns.set_palette(cmap_bold)
# Plot current color palatte
sns.palplot(sns.color_palette())
cmap_mpl_discrete = mpl.colors.ListedColormap(sns.color_palette().as_hex()) # discrete
cmap_mpl_continuous = mpl.colors.LinearSegmentedColormap.from_list(sns.color_palette().as_hex()) # discrete continuousified
# -
import matplotlib.cm as cm
colmap = cm.jet #['#FFFF00', '#00AAFF', '#000000', '#FF00AA']
gradient = np.linspace(0, 1, 256)
plt.imshow(np.linspace(0, 1, 256), aspect='auto', cmap=plt.get_cmap(colmap))
# plt.get_cmap(colmap)
plt.show()
# +
# Create gradient
import matplotlib as mpl
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
cmap_continuous = LinearSegmentedColormap.from_list("", ["red","white","blue"])
cmap_discrete = ListedColormap(["red","gray","blue"], name='from_list', N=None)
data = np.arange(100, 0, -1).reshape(10, 10)
fig, ax = plt.subplots()
cax = fig.add_axes([1, 0.2, 0.5, 0.1]) # figure in figure
im = ax.imshow(data, cmap=cmap_discrete)
# fig.colorbar(im, cax=cax, orientation='horizontal')
plt.show()
# +
# Create gradient
import matplotlib as mpl
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
cmap_continuous = LinearSegmentedColormap.from_list("", ["red","white","blue"])
cmap_discrete = ListedColormap(["red","gray","blue"], name='from_list', N=None)
fig = plt.figure()
ax1 = fig.add_subplot(1,2,1) # two rows, one column, first plot
ax1.scatter(np.arange(15), np.arange(15)+1, c=np.arange(15), cmap=cmap_discrete)
fig.colorbar(ax=ax1)
ax2 = fig.add_subplot(1,2,2) # two rows, one column, first plot
ax2.scatter(np.arange(15), np.arange(15)+1, c=np.arange(15), cmap=cmap_continuous)
fig.colorbar()
plt.show();
# +
def color_in_tokens(tokens, color_token_contains="_"):
"""
Highlights the tokens which contain 'color_token_contains'
:param tokens: list of strings
:param color_token_contains: str (the string for marking a token red)
:return: str
"""
return " ".join(["\x1b[31m%s\x1b[0m" % i if color_token_contains in i else i for i in tokens])
print(color_in_tokens(['I', 'am' 'in_blue']))
# +
from IPython.display import HTML as html_print
def cstr(s, color='black'):
return "<text style=color:{}>{}</text>".format(color, s)
left, word, right = 'foo' , 'abc' , 'bar'
html_print(cstr(' '.join([left, cstr(word, color='red'), right]), color='black') )
# +
import seaborn as sns
# https://seaborn.pydata.org/tutorial/color_palettes.html
colors = ["#67E568","#257F27","#08420D","#FFF000","#FFB62B","#E56124","#E53E30","#7F2353","#F911FF","#9F8CA6"]
current_palette = sns.color_palette(colors)
sns.palplot(current_palette, 1)
current_palette = sns.color_palette(colors, 4)
sns.palplot(current_palette, 2)
# +
flatui = ["#9b59b6", "#3498db", "#95a5a6", "#e74c3c", "#34495e", "#2ecc71"]
sns.palplot(sns.color_palette(flatui))
vibe = ['#98dd1e', '#f27c1a', '#e10976', '#088edf', '#8140ff']
sns.palplot(sns.color_palette(vibe))
# +
import seaborn as sns
# http://www.color-hex.com/color-palettes/?page=2
# View color palette
colors = ['#FFFF00', '#00AAFF', '#000000', '#FF00AA']
cmap_bold = sns.color_palette(colors)
sns.palplot(cmap_bold)
# Set color palette
sns.set_palette("husl")
sns.set_palette(cmap_bold)
# Plot current color palatte
sns.palplot(sns.color_palette())
# -
def plot_labelled_scatter(X, y, class_labels, ax_pad=1):
import matplotlib.cm as cm
from matplotlib.colors import ListedColormap, BoundaryNorm
import matplotlib.patches as mpatches
import seaborn as sns
# Colors
colors = ['#FFFF00', '#00AAFF', '#000000', '#FF00AA']
col_pal = sns.color_palette(colors).as_hex()
cmap = ListedColormap(col_pal)
# BoundaryNorm maps from data points(labels) to colors based on discrete intervals.
# Boundaries defines the edges of bins, and data falling within a bin is mapped to the color with the same index.
# If the number of bins doesn’t equal ncolors, the color is chosen by linear interpolation of the bin number onto color numbers.
# c=y, cmap=cmap, norm=bnorm => map y to color from pallete cut by bounderies
num_labels = len(class_labels)
bounderies = np.arange(num_labels+1)
bnorm = BoundaryNorm(boundaries=bounderies, ncolors=num_labels)
plt.figure()
plt.scatter(X[:, 0], X[:, 1], s=65, c=y, cmap=cmap, norm=bnorm,
alpha=0.40, edgecolor='black', lw=1) #
plt.xlim(X[:, 0].min()-ax_pad, X[:, 0].max()+ax_pad)
plt.ylim(X[:, 1].min()-ax_pad, X[:, 1].max()+ax_pad)
# Legend
legend_handle = [mpatches.Patch(color=color_array[c], label=class_labels[c])
for c in range(num_labels)]
plt.legend(handles=legend_handle)
# +
import seaborn as sns
# View color palette
colors = ['#FFFF00', '#00AAFF', '#000000', '#FF00AA']
cmap_bold = sns.color_palette(colors)
sns.palplot(cmap_bold)
# Set color palette
sns.set_palette("husl")
sns.set_palette(cmap_bold)
# Plot current color palatte
sns.palplot(sns.color_palette())
cmap_mpl = mpl.colors.ListedColormap(sns.color_palette().as_hex()) # discrete
cmap_mpl = mpl.colors.LinearSegmentedColormap.from_list(sns.color_palette().as_hex()) # continuousified
# -
#### Sanity check dataset format
def format_check(X, y):
import numpy as np
assert type(X) == type(np.zeros(2))
assert X.shape[1] > 0
assert type(y) == type(np.zeros(y.shape))
try:
y.shape[1]
print('{} must be of shape: (n,)'.format(y.shape))
except:
pass
print('X:\t {} {}\ny:\t {} {}\nclasses: {}\n'.format(X.shape, type(X), y.shape, type(y), set(y)))
# +
# http://walkerke.github.io/geog30323/slides/data-visualization/#/20
# -
df = pd.read_csv('../_data/col_hex_palettes.csv')
df.info()
df['name'] = [x.lower() for x in df['name'].astype('str')]
df = df.set_index('name')
# Remove duplicates
df.drop_duplicates(subset=None, keep='first', inplace=True)
df.info()
df.to_csv('../_data/col_hex_palettes.csv')
df.tail()
# +
import re
find = input('Search: ')
cols = df.filter(like=find.lower(), axis=0)['hexs'].unique()
names = df.filter(like=find.lower(), axis=0).index.unique()
for name, col in zip(names, cols):
col = re.sub('[\[\]\' \']', '', col).split(',')
cmap = sns.color_palette(col)
_ = sns.palplot(cmap)
_ = plt.gca().set_title(name);
# +
import ipywidgets as widgets
from IPython.display import display
from ipywidgets import interact, interactive, fixed, interact_manual, IntSlider
@interact(search = (df.index.sort_values()))
def get_palette(search):
import re
try:
cols = df.filter(like=search.lower(), axis=0)['hexs'].unique()
names = df.filter(like=search.lower(), axis=0).index.unique()
for name, col in zip(names, cols):
col = re.sub('[\[\]\' \']', '', col).split(',')
cmap = sns.color_palette(col)
_ = sns.palplot(cmap)
_ = plt.gca().set_title(name);
except:
pass
return col
# -
import requests, urllib
urllib.request.urlopen('http://http://www.color-hex.com/color-palettes/')
import os
os.system("start \"\" http://google.com")
# !jupyter lab --version
# !pip list | grep ipywidgets
# !pip list | grep qgrid
# !jupyter labextension list
# !jupyter labextension uninstall qgrid
| _productivity/genious_gists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# ## GRCh37 New chainSelf and Segdups
# This notebook shows the process to generate a new chainSelf file for GRCh37 using `chainSelf.txt.gz` and `chainSelfLink.txt.gz` from the UCSC table browser. This process also uses Aaron Wegners (PacBio) script `hg19.README_annotation.md` (5/22/19)
#
# UCSC files previously downloaded can be obtained using the following commands:
#
# `rsync -avzP rsync://hgdownload.cse.ucsc.edu/goldenPath/hg19/database/chainSelf.txt.gz`
# `rsync -avzP rsync://hgdownload.cse.ucsc.edu/goldenPath/hg19/database/chainSelfLink.txt.gz`
# Note: this process was first run by JM with shell script `Aaron_hg19_step_by_step.sh`. All output files from what was called the "DM" run for datamash, are found in `/Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_beds/selfchain/new_selfchains/Files_from_DM_selfchain_prep`
# **Working Directory**
# `/Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_beds/selfchain/new_selfchains`
# #### Summary of Process
#
# **1. Pre-processing**
#
# UCSC chainSelf/Link --> remove trivial (chainSelf only) --> remove alts (chainSelf and Link)
#
# **2. Run Aaron Script to generate selfchain**
#
# **3. Post-Process selfchain**
#
# selfchain --> sort/remove non 1-22,XY --> merge -d100 --> filter merged >10kb --> sum regions
display < /Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_beds/selfchain/new_selfchains/img/GRCh37_chainSelf_process.png
# ### 1) Remove Trivial Regions
# This step will remove "trivial" regions, aka regions that map to themselves.
gzcat GRCh37_chainSelf.txt.gz | awk '$3!=$7 || $5!=$10' | bgzip > GRCh37_chainSelf_notrivial.txt.gz
gzcat GRCh37_chainSelf_notrivial.txt.gz | wc -l
gzcat GRCh37_chainSelf.txt.gz | wc -l
# chainSelf did not contain any regions that mapped to itself.
# ### 2) Remove Alternate Loci
# Remove any alternate loci regions. These are regions where "hap" is included in the chromosome name. Alternates will be removed from both the chainSelf and Link files and outputs will be used as input into Aaron script.
# **chainSelf**
gzcat GRCh37_chainSelf_notrivial.txt.gz | grep -v hap | bgzip > GRCh37_chainSelf_notrivial_noalts.txt.gz
gzcat GRCh37_chainSelf_notrivial.txt.gz | wc -l
gzcat GRCh37_chainSelf_notrivial_noalts.txt.gz | wc -l
echo $((1058543-988267))
# approximately ~70K alternate sites removed from chainSelf.
# **chainSelfLink**
gzcat GRCh37_chainSelfLink.txt.gz | grep -v hap | bgzip > GRCh37_chainSelfLink_noalts.txt.gz
gzcat GRCh37_chainSelfLink.txt.gz | wc -l
gzcat GRCh37_chainSelfLink.txt.gz | wc -l
gzcat GRCh37_chainSelfLink_noalts.txt.gz | wc -l
echo $((6882618-6183406))
# approximately ~700K alternate sites removed from chainSelfLink.
# ### 3) Run Aaron Script
# code below is modified version of Aaron's selfchain preparation in `hg.19.README_annotation.md`. His script was broken in to smaller chunks to monitor running and eliminate some issues with running from terminal on the Mac.
echo "began script at" `date`
gzcat GRCh37_chainSelfLink_noalts.txt.gz \
| awk '{ print $6 "\t" $3 "\t" $4 "\t" $5 "\t" $5+$4-$3; }' \
| sort -k1,1 -k2,2g > ./script_intermediates/37_chainSelfLink_intermediate.bed
echo "intermediate finished" `date`
echo "began script at" `date`
gzcat GRCh37_chainSelfLink_noalts.txt.gz | awk '{ print $6 "\t" $3 "\t" $4; }' | bedtools merge -d 100 -i - | bedtools intersect -wa -wb -a stdin -b ./script_intermediates/37_chainSelfLink_intermediate.bed | sort -k1,1 -k2,2 -k3,3 | datamash -g1,2,3 min 7 max 8 > ./script_intermediates/37_datamash_intermediate.bed
echo "finished through datamash" `date`
echo "began script at" `date`
gzcat GRCh37_chainSelf_notrivial_noalts.txt.gz | awk '{ print $12 "\t" $3 "\t" $7 "\t" $8 "\t" $9 "\t" $13; }' | sort -k1,1 > ./script_intermediates/37_chainSelf_intermediate.bed
echo "finished 37_chainSelf_intermediate.bed" `date`
echo "began script at" `date`
join -t $'\t' ./script_intermediates/37_datamash_intermediate.bed ./script_intermediates/37_chainSelf_intermediate.bed > ./script_intermediates/37_joined_intermediate.bed
echo "finished join" `date`
echo "began script at" `date`
awk '{ qs=($9=="+")?$4:$8-$5; qe=($9=="+")?$5:$8-$4; printf "%s\t%d\t%d\t%s:%'\''d-%'\''d\t%s\t%d\n", $6, $2, $3, $7, qs+1, qe, $9, int(10*$10); }' ./script_intermediates/37_joined_intermediate.bed| sort -k1,1 -k2,2g | bgzip -c > ./annotation/GRCh37_Aaron_code_chainSelf.bed.gz
tabix -f ./annotation/GRCh37_Aaron_code_chainSelf.bed.gz
echo "finished selfchain GRCh37_Aaron_code_chainSelf.bed.gz" `date`
md5 ./annotation/GRCh37_Aaron_code_chainSelf.bed.gz
# ### 4) Selfchain post processing
# post processing done to remove undesired chromosomes/contigs, merge and filter regions to >10kb
# **check to see if sorted and what needs to be removed... "M" and other "un_" extra chroms and needs sorting.**
gzcat ./annotation/GRCh37_Aaron_code_chainSelf.bed.gz | cut -f 1 | uniq
# **Remove non 1-22, XY chromosomes (Remove chrM, chr#_, chr##_ and chrUn) and sort.**
gzcat ./annotation/GRCh37_Aaron_code_chainSelf.bed.gz | sed 's/^chr//' | grep -Ev '^M|^[0-9][0-9]_|[0-9]_|^Un' | sed 's/^X/23/;s/^Y/24/' | sort -k1,1n -k2,2n | sed 's/^23/X/;s/^24/Y/'| bgzip > ./annotation/GRCh37_Aaron_code_chainSelf_sorted.bed.gz
# **Merge Regions**
# This processs will merge regions with maximum distance of 100 bps
# Note: filenames changed as this will be a distributed outputfile, "Aaron_code" removed from filename
gzcat ./annotation/GRCh37_Aaron_code_chainSelf_sorted.bed.gz | bedtools merge -i stdin -d 100 | bgzip > ./annotation/GRCh37_chainSelf_sorted_merged.bed.gz
# **merged line count and regions sum for merged output file**
gzcat ./annotation/GRCh37_chainSelf_sorted_merged.bed.gz | wc -l
gzcat ./annotation/GRCh37_chainSelf_sorted_merged.bed.gz | awk '{ sum+=$3;sum-=$2 } END { print sum }'
# **Filter to regions >10kb**
gzcat ./annotation/GRCh37_chainSelf_sorted_merged.bed.gz | awk '($3-$2 > 10000)' | bgzip > ./annotation/GRCh37_chainSelf_sorted_merged_gt10kb.bed.gz
# **line count and regions sum for filtered output file**
gzcat ./annotation/GRCh37_chainSelf_sorted_merged_gt10kb.bed.gz | wc -l
gzcat ./annotation/GRCh37_chainSelf_sorted_merged_gt10kb.bed.gz | awk '{ sum+=$3;sum-=$2 } END { print sum }'
echo $((190079658-131331831))
# ## Segdups
# JZ would like to use segmental duplications file in conjunction with self chains. Like self chain, this file will be merged and filtered to regions >10kb. The starting file `segdups.bed.gz` was generated by <NAME> (PacBio) using his script `hg19.README_annotation.md`
#
# UCSC Segdups file was downloaded from:
# rsync -avzP rsync://hgdownload.cse.ucsc.edu/goldenPath/hg19/database/genomicSuperDups.txt.gz
#
# **Summary of Processing**
# segdups.bed.gz --> sort/remove non 1-22,XY --> merge -d100 --> filter merged >10kb
gzcat hg19.segdups.bed.gz | cut -f 1 | grep -E "_|chr[a-zA-Z]"| uniq
# #### Remove non 1-22, XY chromosomes (Remove chrM, chr#, chr## and chrUn) and sort.
gzcat hg19.segdups.bed.gz | sed 's/^chr//' | grep -Ev '^M|^[0-9][0-9]_|[0-9]_|^Un' | sed 's/^X/23/;s/^Y/24/' | sort -k1,1n -k2,2n | sed 's/^23/X/;s/^24/Y/'| bgzip > ./new_segdups/hg19.segdups_sorted.bed.gz
gzcat hg19.segdups.bed.gz | wc -l
gzcat ./new_segdups/hg19.segdups_sorted.bed.gz | wc -l
echo $((51599-49948))
# removal of extra chrom/contigs removed 1651 lines (locations)
# #### **Merge Regions**
# This processs will merge regions with maximum distance of 100 bps
gzcat ./new_segdups/hg19.segdups_sorted.bed.gz | bedtools merge -i stdin -d 100 | bgzip > ./new_segdups/hg19.segdups_sorted_merged.bed.gz
# Stats before merging (line count, regions sum)
gzcat ./new_segdups/hg19.segdups_sorted.bed.gz | wc -l
gzcat ./new_segdups/hg19.segdups_sorted.bed.gz | awk '{ sum+=$3;sum-=$2 } END { print sum }'
# Stats after merging (line count, regions sum)
gzcat ./new_segdups/hg19.segdups_sorted_merged.bed.gz | wc -l
gzcat ./new_segdups/hg19.segdups_sorted_merged.bed.gz | awk '{ sum+=$3;sum-=$2 } END { print sum }'
# #### Filter to regions >10kb
gzcat ./new_segdups/hg19.segdups_sorted_merged.bed.gz| awk '($3-$2 > 10000)' | bgzip > ./new_segdups/hg19.segdups_sorted_merged_gt10kb.bed.gz
# Stats after filtering (line count, regions sum)
gzcat ./new_segdups/hg19.segdups_sorted_merged_gt10kb.bed.gz | wc -l
gzcat ./new_segdups/hg19.segdups_sorted_merged_gt10kb.bed.gz | awk '{ sum+=$3;sum-=$2 } END { print sum }'
# ## Create "notin" files
subtractBed -a /Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_stratification_validation/data/stratifications/reference_beds/intermediates/human.b37.1_22XY.genome.sorted.bed -b ./annotation/GRCh37_chainSelf_sorted_merged_gt10kb.bed.gz | bgzip > ./annotation/GRCh37_notinchainSelf_sorted_merged_gt10kb.bed.gz
subtractBed -a /Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_stratification_validation/data/stratifications/reference_beds/intermediates/human.b37.1_22XY.genome.sorted.bed -b ./annotation/GRCh37_chainSelf_sorted_merged.bed.gz | bgzip > ./annotation/GRCh37_notinchainSelf_sorted_merged.bed.gz
subtractBed -a /Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_stratification_validation/data/stratifications/reference_beds/intermediates/human.b37.1_22XY.genome.sorted.bed -b ./new_segdups/hg19.segdups_sorted_merged_gt10kb.bed.gz | bgzip > ./new_segdups/hg19.notinsegdups_sorted_merged_gt10kb.bed.gz
subtractBed -a /Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_stratification_validation/data/stratifications/reference_beds/intermediates/human.b37.1_22XY.genome.sorted.bed -b ./new_segdups/hg19.segdups_sorted_merged.bed.gz | bgzip > ./new_segdups/hg19.notinsegdups_sorted_merged.bed.gz
# ## md5 checksums
md5 ./new_segdups/hg19.segdups_sorted_merged.bed.gz
md5 ./new_segdups/hg19.segdups_sorted_merged_gt10kb.bed.gz
md5 ./annotation/GRCh37_chainSelf_sorted_merged_gt10kb.bed.gz
md5 ./annotation/GRCh37_chainSelf_sorted_merged.bed.gz
md5 ./new_segdups/hg19.notinsegdups_sorted_merged.bed.gz
md5 ./new_segdups/hg19.notinsegdups_sorted_merged_gt10kb.bed.gz
md5 ./annotation/GRCh37_notinchainSelf_sorted_merged_gt10kb.bed.gz
md5 ./annotation/GRCh37_notinchainSelf_sorted_merged.bed.gz
# ## New GRCh37 SelfChain and Segdup Output files
# chainself files on JM computer in `/Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_beds/selfchain/new_selfchains/annotation`
#
# sedup files on JM computer in `/Users/jmcdani/Documents/GiaB/Benchmarking/GRCh38_beds/selfchain/new_selfchains/new_segdups`
# |File|md5|File Size|Date Generated|
# |-|-|-|-|
# |GRCh37_chainSelf_sorted_merged_gt10kb.bed.gz|54026470ca8fc602648356db063e34ba|22 KB |12/11/19|
# |GRCh37_chainSelf_sorted_merged.bed.gz |a997899308d2343150ae7cfc1cf6d4af|470 KB |12/11/19|
# |hg19.segdups_sorted_merged_gt10kb.bed.gz|ff88d362fb0ba763b52defdb300a4849|17 KB| 12/11/19|
# |hg19.segdups_sorted_merged.bed.gz|14e02e2092d24886c297d71170bfefb0|65 KB|12/11/19|
# |hg19.notinsegdups_sorted_merged.bed.gz|f4c8d96e208ae4056fa4d8bec3768e99|67 KB|12/11/19|
# |hg19.notinsegdups_sorted_merged_gt10kb.bed.gz|2c4a6a79352366716eda70696c09f82a|17 KB|12/11/19|
# |GRCh37_notinchainSelf_sorted_merged_gt10kb.bed.gz|e03447454eaa5b64b225547c7d819662|22 KB|12/11/19|
# |GRCh37_notinchainSelf_sorted_merged.bed.gz|78724aeeb4977a080695010062d2a29a|481 KB|12/11/19|
| GRCh37/SegmentalDuplications/GRCh37_new_chainSelf_and_Segdups.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Flattening a nested linked list
#
# Suppose you have a linked list where the value of each node is a sorted linked list (i.e., it is a _nested_ list). Your task is to _flatten_ this nested list—that is, to combine all nested lists into a single (sorted) linked list.
#
# First, we'll need some code for generating nodes and a linked list:
# +
# Use this class as the nodes in your linked list
class Node:
def __init__(self, value):
self.value = value
self.next = None
def __repr__(self):
return str(self.value)
class LinkedList:
def __init__(self, head):
self.head = head
def append(self, value):
if self.head is None:
self.head = Node(value)
return
node = self.head
while node.next is not None:
node = node.next
node.next = Node(value)
# -
# Now, in the cell below, see if you can solve the problem by implementing the `flatten` method.
#
# >Hint: If you first create a `merge` method that merges two linked lists into a sorted linked list, then there is an elegant recursive solution.
# +
def merge(list1, list2):
# TODO: Implement this function so that it merges the two linked lists in a single, sorted linked list.
pass
class NestedLinkedList(LinkedList):
def flatten(self):
# TODO: Implement this method to flatten the linked list in ascending sorted order.
pass
# -
# Here's some code that will generate a nested linked list that we can use to test the solution:
# +
# First Test scenario
linked_list = LinkedList(Node(1))
linked_list.append(Node(3))
linked_list.append(Node(5))
nested_linked_list = NestedLinkedList(Node(linked_list))
second_linked_list = LinkedList(Node(2))
second_linked_list.append(4)
nested_linked_list.append(Node(second_linked_list))
# -
# ### Structure
# `nested_linked_list` should now have 2 nodes. The head node is a linked list containing `1, 3, 5`. The second node is a linked list containing `2, 4`.
#
# Calling `flatten` should return a linked list containing `1, 2, 3, 4, 5`.
solution = nested_linked_list.flatten()
assert solution == [1,2,3,4,5]
# ### Solution
# First, let's implement a `merge` function that takes in two linked lists and returns one sorted linked list. Note, this implementation expects both linked lists to be sorted.
def merge(list1, list2):
merged = LinkedList(None)
if list1 is None:
return list2
if list2 is None:
return list1
list1_elt = list1.head
list2_elt = list2.head
while list1_elt is not None or list2_elt is not None:
if list1_elt is None:
merged.append(list2_elt)
list2_elt = list2_elt.next
elif list2_elt is None:
merged.append(list1_elt)
list1_elt = list1_elt.next
elif list1_elt.value <= list2_elt.value:
merged.append(list1_elt)
list1_elt = list1_elt.next
else:
merged.append(list2_elt)
list2_elt = list2_elt.next
return merged
# Let's make sure merge works how we expect:
# +
linked_list = LinkedList(Node(1))
linked_list.append(3)
linked_list.append(5)
second_linked_list = LinkedList(Node(2))
second_linked_list.append(4)
merged = merge(linked_list, second_linked_list)
node = merged.head
while node is not None:
#This will print 1 2 3 4 5
print(node.value)
node = node.next
# Lets make sure it works with a None list
merged = merge(None, linked_list)
node = merged.head
while node is not None:
#This will print 1 2 3 4 5
print(node.value)
node = node.next
# -
# Now let's implement `flatten` recursively using merge.
class NestedLinkedList(LinkedList):
def flatten(self):
return self._flatten(self.head)
def _flatten(self, node):
if node.next is None:
return merge(node.value, None)
return merge(node.value, self._flatten(node.next))
# +
nested_linked_list = NestedLinkedList(Node(linked_list))
nested_linked_list.append(second_linked_list)
flattened = nested_linked_list.flatten()
node = flattened.head
while node is not None:
#This will print 1 2 3 4 5
print(node.value)
node = node.next
# -
# ### Computational Complexity
# Lets start with the computational complexity of `merge`. Merge takes in two lists. Let's say the lengths of the lists are $N_{1}$ and $N_{2}$. Because we assume the inputs are sorted, `merge` is very efficient. It looks at the first element of each list and adds the smaller one to the returned list. Every time through the loop we are appending one element to the list, so it will take $N_{1} + N_{2}$ iterations until we have the whole list.
#
# The complexity of `flatten` is a little more complicated to calculate. Suppose our `NestedLinkedList` has $N$ linked lists and each list's length is represented by $M_{1}, M_{2}, ..., M_{N}$.
#
# We can represent this recursion as:
#
# $merge(M_{1}, merge(M_{2}, merge(..., merge(M_{N-1}, merge(M_{N}, None)))))$
#
# Let's start from the inside. The inner most merge returns the $nth$ linked list. The next merge does $M_{N-1} + M_{N}$ comparisons. The next merge does $M_{N-2} + M_{N-1} + M_{N}$ comparisons.
#
# Eventually we will do $N$ comparisons on all of the $M_{N}$ elements. We will do $N-1$ comparisons on $M_{N-1}$ elements.
#
# This can be generalized as:
#
# $$
# \sum_n^N n*M_{n}
# $$
| linked_lists/Flattening a nested linked list.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression Explained
# <table align="left"><td>
# <a target="_blank" href="https://colab.research.google.com/github/TannerGilbert/Tutorials/blob/master/Machine-Learning-Explained/Linear%20Regression%20Explained.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab
# </a>
# </td><td>
# <a target="_blank" href="https://github.com/TannerGilbert/Tutorials/blob/master/Machine-Learning-Explained/Linear%20Regression%20Explained.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td></table>
# 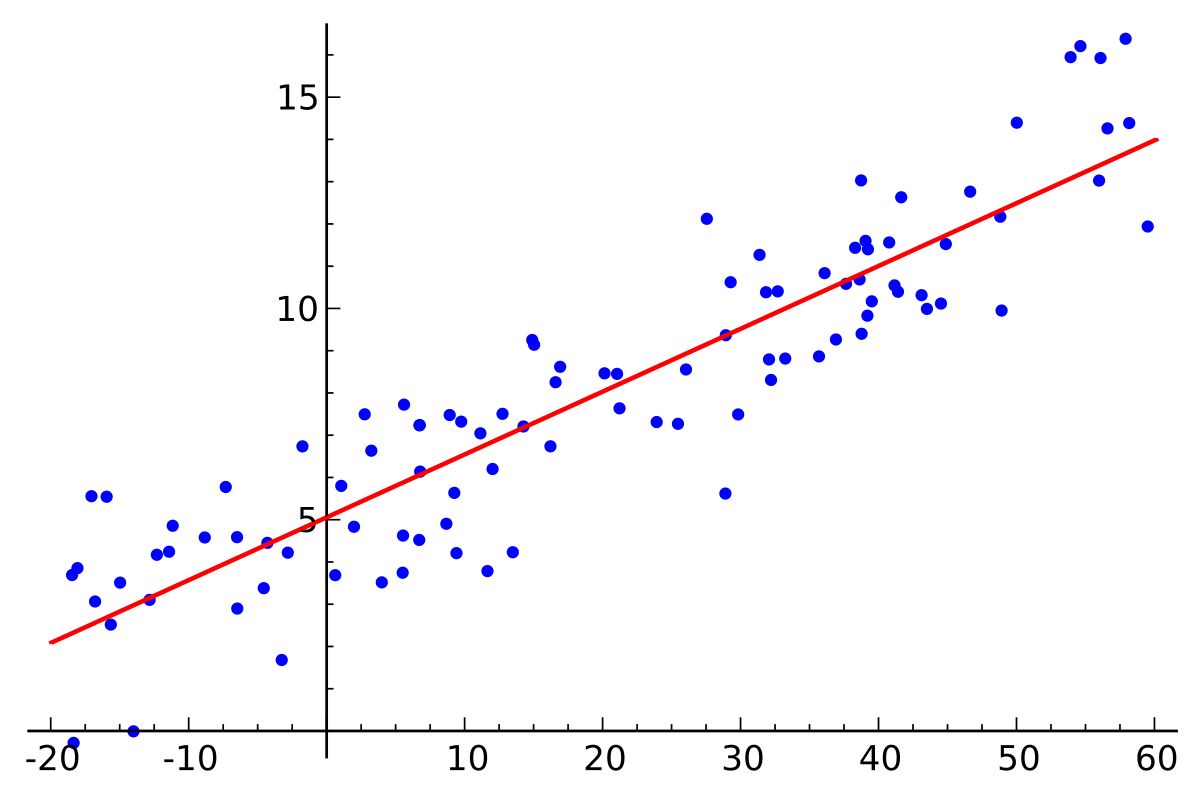
# ## What is Linear Regression?
# In statistics, linear regression is a linear approach to modelling the relationship between a dependent variable(y) and one or more independent variables(X). In linear regression, the relationships are modeled using linear predictor functions whose unknown model parameters are estimated from the data. Linear Regression is one of the most popular algorithms in Machine Learning. That's due to its relative simplicity and well known properties.
# ## Simple Linear Regression
#
# Simple linear regression is a linear regression model with only independent variable.
#
# Formula: $ f\left(x\right)=mx+b $
# ### Cost Function
# We can measure the accuracy of Linear Regression using the <b>Mean Squared Error</b> cost function.
#
# $ Error(m, b) = \frac{1}{N}\sum_{i=1}^{N}(\text{actual output}-\text{predicted output})^2$
# Code:
def cost_function(m, b, x, y):
totalError = 0
for i in range(0, len(x)):
totalError += (y[i]-(m*x[i]+b))**2
return totalError/float(len(x))
# ### Optimization
#
# To find the coefficients that minimize our error function we will use <b>gradient descent</b>. Gradient descent is a optimization algorithm which iteratively takes steps to the local minimum of the cost function. It takes the derivative of our cost function to find the direction to move towards.
# 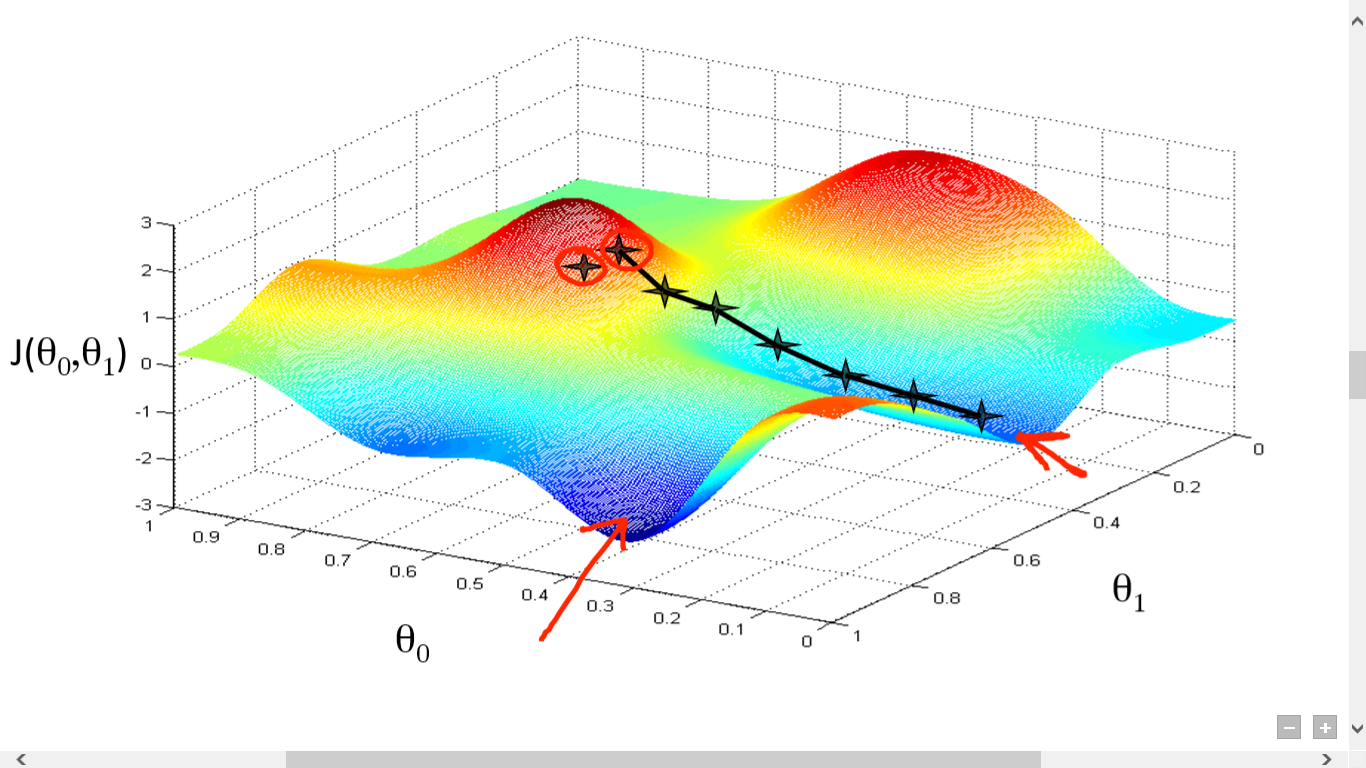
# Gradient Descent Formula:
# $$ \Theta_j:=\Theta_j-\alpha\frac{\partial}{\partial\Theta_j}J\left(\Theta_0,\Theta_1\right) $$
#
# Gradient Descent for Linear Regression:
# $$ \frac{\partial}{\partial m}=\frac{2}{N}\sum_{i=1}^{N}{-x_i(y_i-\left(mx_i+b\right))} $$
# $$ \frac{\partial}{\partial b}=\frac{2}{N}\sum_{i=1}^{N}{-(y_i-\left(mx_i+b\right))} $$
#
# Code:
def gradient_descent(b, m, x, y, learning_rate, num_iterations):
N = float(len(x))
for j in range(num_iterations):
b_gradient = 0
m_gradient = 0
for i in range(0, len(x)):
b_gradient += -(2/N) * (y[i] - ((m * x[i]) + b))
m_gradient += -(2/N) * x[i] * (y[i] - ((m * x[i]) + b))
b -= (learning_rate * b_gradient)
m -= (learning_rate * m_gradient)
if j%50==0:
print('error:', cost_function(m, b, x, y))
return [b, m]
# ### Running Linear Regression
# Creating Dataset
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 100, 50)
delta = np.random.uniform(-10, 10, x.size)
y = 0.5*x + 3 + delta
plt.scatter(x, y)
# +
learning_rate = 0.0001
initial_b = 0
initial_m = 0
num_iterations= 100
print('Initial error:', cost_function(initial_m, initial_b, x, y))
[b, m] = gradient_descent(initial_b, initial_m, x, y, learning_rate, num_iterations)
print('b:', b)
print('m:', m)
print('error:', cost_function(m, b, x, y))
# -
predictions = [(m * x[i]) + b for i in range(len(x))]
plt.scatter(x, y)
plt.plot(x, predictions, color='r')
# ## Multivariate Linear Regression
#
# Linear Regression is called mutlivariate if the data you are working with has 2 or mode independent variables.
#
# Formula: $ f\left(x\right)=b+w_1x_1+w_2x_2+\ldots+w_nx_n=b+\sum_{i=1}^{n}{w_ix}_i $
#
# It works almost the same that simple linear regression. We can use the same error function (the predicted output is different). But now we need to perform gradient descent for each feature.
# ### Cost Function
def cost_function(x, y, w):
dif = np.dot(x,w)-y
cost = np.sum(dif**2) / (2*np.shape(x)[0])
return dif, cost
# ### Optimization
def multivariate_gradient_descent(x, y, w, learning_rate, num_iterations):
for i in range(num_iterations):
dif, cost = cost_function(x, y, w)
gradient = np.dot(x.transpose(), dif) / np.shape(x)[0]
w = w - learning_rate * gradient
if i%500==0:
print('error:', cost)
return w
# ### Runing Multivariate Linear Regression
# +
import pandas as pd
from sklearn.preprocessing import LabelEncoder
iris = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data', names=['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'label'])
le = LabelEncoder()
iris['label'] = le.fit_transform(iris['label'])
X = np.array(iris.drop(['petal_width'], axis=1))
y = np.array(iris['petal_width'])
iris.head()
# -
learning_rate = 0.0001
num_iterations= 10000
_, num_features = np.shape(X)
initial_weights = np.zeros(num_features)
weights = multivariate_gradient_descent(X, y, initial_weights, learning_rate, num_iterations)
print(weights)
dif, cost = cost_function(X, y, weights)
print('error: ', cost)
# ## Resources
#
# <ul>
# <li><a href="https://en.wikipedia.org/wiki/Linear_regression">Linear Regression (Wikipedia)</a></li>
# <li><a href="https://towardsdatascience.com/simple-and-multiple-linear-regression-in-python-c928425168f9">Simple and Multiple Linear Regression in Python (Adi Bronshtein on Medium)</a></li>
# <li><a href="http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html">Linear Regression (Scikit Learn Documentation)</a></li>
# </ul>
# ## Conclusion
#
# That was my explanation of Linear Regression.
# I hope you liked this tutorial if you did consider subscribing on my <a href="https://www.youtube.com/channel/UCBOKpYBjPe2kD8FSvGRhJwA">Youtube Channel</a> or following me on Social Media. If you have any question feel free to contact me.
| Machine-Learning-Explained/Linear Regression Explained.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #%load_ext autoreload
# #%autoreload 2
from sindy_bvp import SINDyBVP
from sindy_bvp.differentiators import FiniteDifferences
# +
# %%time
outcome_var = 'd^{2}u/dx^{2}'
# Known Operator Case
sbvp_ko = SINDyBVP(file_stem = "./data/S3-P2-",
num_trials = 2,
differentiator = FiniteDifferences(diff_order = 2),
outcome_var = outcome_var,
noisemaker = None,
known_vars = ['du/dx', 'f'],
dep_var_name = 'u',
ind_var_name = 'x')
coeffs, plotter = sbvp_ko.sindy_bvp()
# Generate the analysis plots: ODE solutions (first 3), p(x) and q(x), and u_xx model coefficients (entries in Xi)
plotter.generate_analysis_plots(save_stem='3c-KO')
print("Learned function for", outcome_var ,"includes:", list(coeffs))
# Score coefficients
plotter.score_coefficients()
# +
# %%time
# Unknown Operator Case
sbvp_uo = SINDyBVP(file_stem = "./data/S3-P2-",
num_trials = 8,
differentiator = FiniteDifferences(diff_order = 2),
outcome_var = outcome_var,
noisemaker = None,
known_vars = None,
dep_var_name = 'u',
ind_var_name = 'x')
coeffs, plotter = sbvp_uo.sindy_bvp()
# Generate the analysis plots: ODE solutions (first 3), p(x) and q(x), and u_xx model coefficients (entries in Xi)
plotter.generate_analysis_plots(save_stem='3c-UO')
print("Learned function for", outcome_var ,"includes:", list(coeffs))
# Score coefficients
plotter.score_coefficients()
| Fig 3c - Second Order Poisson Operator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from Bio import SeqIO
from Bio import motifs
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
import click
import numpy as np
import os
import pandas as pd
import torch
from tqdm import tqdm
bar_format = "{percentage:3.0f}%|{bar:20}{r_bar}"
from architectures import CAM, get_metrics
from jaspar import get_figure, reformat_motif
from sequence import one_hot_encode, rc_one_hot_encoding, rc, one_hot_decode
from train import _get_data_loaders, __get_handle
from predict import _predict
device = "cpu"
if torch.cuda.is_available():
device = "cuda"
CAM_DIR = "/mnt/md1/home/oriol/CAM/results/IRF4"
ASSAY = "ChIP-seq"
LABEL = "T95R"
model_file = f"{CAM_DIR}/CAM/{ASSAY}/{LABEL}.1/best_model.pth.tar"
batch_size = 2**6
debugging = False
name = f"{ASSAY}.{LABEL}"
output_dir = f"{CAM_DIR}/CAM/{ASSAY}/{LABEL}.1"
rev_complement = True
threads = 1
FASTA_DIR = f"{CAM_DIR}/CAM/{ASSAY}/T95R-WT.1"
NAME = "test"
fasta_file = f"{FASTA_DIR}/{NAME}.fa"
# Initialize
torch.set_num_threads(threads)
# Create output dirs
if not os.path.isdir(output_dir):
os.makedirs(output_dir)
for subdir in ["predictions", "embeddings"]:
if not os.path.isdir(os.path.join(output_dir, subdir)):
os.makedirs(os.path.join(output_dir, subdir))
# Get model
selene_dict = torch.load(model_file)
model = CAM(
selene_dict["options"]["cnn_units"],
selene_dict["options"]["kernel_size"],
selene_dict["options"]["sequence_length"],
selene_dict["options"]["n_features"],
selene_dict["options"]["clamp_weights"],
selene_dict["options"]["no_padding"],
selene_dict["options"]["weights_file"],
)
model.load_state_dict(selene_dict["state_dict"])
model.to(device)
# Padding
if selene_dict["options"]["no_padding"]:
padding = 0
else:
padding = selene_dict["options"]["kernel_size"]
def _get_Xs_ys_seq_ids(fasta_file, debugging=False, reverse_complement=False):
# Initialize
Xs = []
ys = []
seq_ids = []
# Xs / ys
handle = __get_handle(fasta_file)
for record in SeqIO.parse(handle, "fasta"):
Xs.append(one_hot_encode(str(record.seq).upper()))
ys.append([1.])
seq_ids.append(record.id)
handle.close()
# Reverse complement
if reverse_complement:
n = len(Xs)
for i in range(n):
Xs.append(rc_one_hot_encoding(Xs[i]))
ys.append(ys[i])
seqs.append(rc(seqs[i]))
# Return 1,000 sequences
if debugging:
return(np.array(Xs)[:1000], np.array(ys)[:1000],
np.array(seq_ids)[:1000])
return(np.array(Xs), np.array(ys), np.array(seq_ids))
# +
##############
# Load Data #
##############
# Get data
Xs, ys, seq_ids = _get_Xs_ys_seq_ids(fasta_file, debugging, rev_complement)
# Get DataLoader
data_loader = _get_data_loaders(list(Xs), list(ys), batch_size=batch_size)
# -
for x, _ in data_loader:
for seq, seq_id in zip(x, seq_ids[:x.shape[0]]):
print(one_hot_decode(seq), seq_id)
break
# +
############
# Predict #
############
# Initialize
input_data = "binary"
if selene_dict["options"]["no_padding"]:
padding = 0
else:
padding = selene_dict["options"]["kernel_size"]
# -
predictions, _ = _predict(model, data_loader, input_data)
predictions.shape
def __get_max_predictions(predictions):
# Initialize
data = []
fwd = []
rev = []
strands = []
# DataFrame
for i, values in enumerate(predictions[:len(predictions)//2]):
fwd.append(values.tolist())
data.append([i] + fwd[-1])
for i, values in enumerate(predictions[len(predictions)//2:]):
rev.append(values.tolist())
data.append([i] + rev[-1])
df = pd.DataFrame(data)
# Get max. values
df = df.groupby(0).max()
max_predictions = df.values.tolist()
# Get strands
for i in range(len(max_predictions)):
strands.append([])
for j in range(len(max_predictions[i])):
if max_predictions[i][j] == fwd[i][j]:
strands[-1].append("+")
else:
strands[-1].append("-")
return(np.array(max_predictions), np.array(strands))
#max_predictions, strands = __get_max_predictions(predictions)
seq_ids = seq_ids.reshape(1, -1).T
#data = np.concatenate((seq_ids, max_predictions, strands), axis=1)
data = np.concatenate((seq_ids, predictions), axis=1)
df = pd.DataFrame(data)
tsv_file = f"{output_dir}/predictions/{NAME}.tsv"
df.to_csv(tsv_file, sep="\t", header=False, index=False)
predictions
seq_ids
def _get_embeddings(model, data_loader, input_data):
# Initialize
embeddings = []
labels = []
with torch.no_grad():
for x, label in tqdm(iter(data_loader), total=len(data_loader),
bar_format=bar_format):
# Get embeddings
x = x.to(device)
x = x.repeat(1, model._options["cnn_units"], 1)
out = model.linears(x)
embeddings.extend(out.detach().cpu().numpy())
# Get labels
labels.extend(label.numpy())
return(np.array(embeddings), np.array(labels))
embeddings, _ = _get_embeddings(model, data_loader, input_data)
#values = np.concatenate(np.array_split(embeddings, 2), axis=1)
#data = np.concatenate((seq_ids, values), axis=1)
data = np.concatenate((seq_ids, embeddings), axis=1)
df = pd.DataFrame(data)
tsv_file = f"{output_dir}/embeddings/{NAME}.tsv"
df.to_csv(tsv_file, sep="\t", header=False, index=False)
df
| explainn/predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!unxz mnist.out.xz
# -
from model import Model
from collections import namedtuple
import sys
import occlusion
import random
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
# +
random.seed(1)
np.random.seed(1)
Game = namedtuple('Game', ['env_name', 'input_size', 'output_size', 'wann_file', 'action_select', 'weight_bias'])
game = Game(env_name='MNISTTEST256-v0',
input_size=256,
output_size=10,
wann_file='mnist.out',
action_select='softmax', # all, soft, hard
weight_bias=0.0,
)
model = Model(game)
model.make_env()
model.env.seed(1)
model.load_model('mnist256.wann.json')
batch = model.env.reset()
output = model.get_action(batch)
hdm = occlusion.HausdorffDistanceMasks(16, 16)
hdm.generate_masks(1, 1)
# -
for index in range(10):
image = batch[index].reshape(16, 16)
label = model.env.target[model.env.currIndx][index]
print('label:', label)
baseline = model.get_action([batch[index]])
print('prediction', np.argmax(baseline[0]))
result = hdm.explain(model, image, baseline[0], output_class=label)
i = image * (255 / image.max())
i = i.astype(np.uint8)
plt.imshow(Image.fromarray(i).convert('RGB'))
plt.show()
img = plt.imshow(result.distances(occlusion.RAW))
plt.colorbar(img)
plt.show()
| wann/wann16x16_evaluate_occlusion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
# %config InlineBackend.figure_format = 'retina'
from ipywidgets import interact
import scipy.stats as stats
import pandas as pd
import pymc3 as pm
import seaborn as sns
import arviz as az ## This is new, but it is installed along with PyMC3
## This is new for working with DAGs, you will have to install it
import causalgraphicalmodels as cgm
from causalgraphicalmodels import CausalGraphicalModel
import warnings # the warning spam is pointless and annoying
warnings.simplefilter(action="ignore", category=FutureWarning)
def credible_interval_from_samples(samples, prob):
"""`samples` can be an nd array. Assume that all of the dimensions
except for the last index parameters while the last (rightmost)
dimension indexes the samples."""
samples_sorted = sort(samples, axis=-1)
N_samples = samples.shape[-1]
index = int(N_samples*(1 - prob)/2)
lower = samples_sorted[..., index]
upper = samples_sorted[..., -index]
## quantile(x, [(1 - prob)/2, (1 + prob)/2], axis=-1)
return lower, upper
# +
# # !pip install causalgraphicalmodels
# +
####### If you install causalgraphicalmodels and still get an error
############ "cannot import name 'gcd' from 'fractions'"
####### when you try to load it run this command (it takes several minutes, restart the notebook after)
# # !conda install -y networkx">=2.5"
# -
data = pd.read_csv('Data/WaffleDivorce.csv', delimiter=";")
data_standardized = pd.read_csv('Data/WaffleDivorce.csv', delimiter=";")
columns = ['MedianAgeMarriage', 'Marriage', 'Divorce']
data_standardized[columns] = (data[columns] - data[columns].mean())/data[columns].std()
data_standardized
# ## The DAG
# +
waffles_dag = CausalGraphicalModel(
nodes=['A', 'M', 'D'], edges=[('A', 'M'), ('A', 'D'), ('M', 'D')]
)
waffles_dag.draw()
# -
# ## The effect of marriage rate on divorce rate
with pm.Model() as model_D_M:
sigma = pm.Exponential('sigma', 1)
beta_M = pm.Normal('beta_M', mu=0, sd=0.5)
alpha = pm.Normal('alpha', mu=0, sd=0.2)
## Note that an underscore '_' is nothing special;
## it is just another character that can be part of a variable name.
_mu = alpha + beta_M*data_standardized['Marriage']
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
D = pm.Normal('divorce', mu=mu, sd=sigma, observed=data_standardized['Divorce'])
## MCMC
_D_M_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_D_M_posterior, var_names=['beta_M', 'alpha', 'sigma']))
D_M_posterior = pm.trace_to_dataframe(_D_M_posterior)
az.plot_forest(_D_M_posterior, var_names=["~mu"], combined=True, figsize=[5, 2]);
# ## The effect of age at marriage on divorce rate
with pm.Model() as model_D_A:
sigma = pm.Exponential('sigma', 1)
beta_A = pm.Normal('beta_A', mu=0, sd=0.5)
alpha = pm.Normal('alpha', mu=0, sd=0.2)
_mu = alpha + beta_A*data_standardized['MedianAgeMarriage']
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
D = pm.Normal('divorce', mu=mu, sd=sigma, observed=data_standardized['Divorce'])
## MCMC
_D_A_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_D_A_posterior, var_names=['beta_A', 'alpha', 'sigma']))
D_A_posterior = pm.trace_to_dataframe(_D_A_posterior)
az.plot_forest(_D_A_posterior, var_names=["~mu"], combined=True, figsize=[5, 2]);
# ## The effect of age at marriage on marriage rate
with pm.Model() as model_M_A:
## A -> M
sigma_M = pm.Exponential('sigma_M', 1)
beta_AM = pm.Normal('beta_AM', 0, 0.5)
alpha_M = pm.Normal('alpha_M', 0, 0.2)
mu_M = pm.Deterministic('mu_M', alpha_M + beta_AM*data_standardized['MedianAgeMarriage'])
age = pm.Normal('age', mu_M, sigma_M, observed=data_standardized['Marriage'])
_M_A_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_M_A_posterior, var_names=['beta_AM', 'alpha_M', 'sigma_M']))
M_A_posterior = pm.trace_to_dataframe(_M_A_posterior)
az.plot_forest(_M_A_posterior, var_names=["~mu_M"], combined=True, figsize=[5, 2]);
# ## The multiple regression model
with pm.Model() as model_D_AM:
## A -> M
sigma_M = pm.Exponential('sigma_M', 1)
beta_AM = pm.Normal('beta_AM', 0, 0.5)
alpha_M = pm.Normal('alpha_M', 0, 0.2)
mu_M = pm.Deterministic('mu_m', alpha_M + beta_AM*data_standardized["MedianAgeMarriage"])
marriage = pm.Normal('marriage', mu_M, sigma_M, observed=data_standardized["Marriage"])
## A -> D <- M
sigma = pm.Exponential('sigma', 1)
beta_A = pm.Normal('beta_A', mu=0, sd=0.5)
beta_M = pm.Normal('beta_M', mu=0, sd=0.5)
alpha = pm.Normal('alpha', mu=0, sd=0.2)
## Note that an underscore '_' is nothing special;
## it is just another character that can be part of a variable name.
_mu = alpha + beta_M*marriage + beta_A*data_standardized["MedianAgeMarriage"]
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
D = pm.Normal('divorce', mu=mu, sd=sigma, observed=data_standardized['Divorce'])
## MCMC
_D_AM_posterior = pm.sample(1000, tune=1000)
print(pm.summary(_D_AM_posterior, var_names=['beta_A', 'beta_M', 'beta_AM']))
D_AM_posterior = pm.trace_to_dataframe(_D_AM_posterior)
az.plot_forest(_D_AM_posterior, var_names=['~mu_M', '~mu'], combined=True, figsize=[5, 2]);
# ## Prediction is that $M$ and $D$ are nearly conditionally independent given $A$
CausalGraphicalModel(
nodes=['A', 'M', 'D'], edges=[('A', 'D'), ('A', 'M')]
).draw()
# ## What if we instead assume that $M$ causes $A$ in our model?
# +
waffles_alt_dag = CausalGraphicalModel(
nodes=['A', 'M', 'D'], edges=[('M', 'A'), ('A', 'D'), ('M', 'D')]
)
waffles_alt_dag.draw()
# -
with pm.Model() as model_D_MA:
## M -> A
sigma_A = pm.Exponential('sigma_A', 1)
beta_MA = pm.Normal('beta_MA', 0, 0.5)
alpha_A = pm.Normal('alpha_A', 0, 0.2)
mu_A = pm.Deterministic('mu_A', alpha_A + beta_MA*data_standardized['Marriage'])
age = pm.Normal('age', mu_A, sigma_A, observed=data_standardized['MedianAgeMarriage'])
## A -> D <- M
sigma = pm.Exponential('sigma', 1)
beta_A = pm.Normal('beta_A', mu=0, sd=0.5)
beta_M = pm.Normal('beta_M', mu=0, sd=0.5)
alpha = pm.Normal('alpha', mu=0, sd=0.2)
## Note that an underscore '_' is nothing special;
## it is just another character that can be part of a variable name.
_mu = alpha + beta_M*data_standardized['Marriage'] + beta_A*age
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
D = pm.Normal('divorce', mu=mu, sd=sigma, observed=data_standardized['Divorce'])
## MCMC
_D_MA_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_D_MA_posterior, var_names=['beta_A', 'beta_M', 'beta_MA']))
D_MA_posterior = pm.trace_to_dataframe(_D_MA_posterior)
az.plot_forest(_D_MA_posterior, var_names=['~mu_A', '~mu'], combined=True, figsize=[5, 2]);
# ### We have the same prediction that D and M are independent conditioned on A
# $$ \mu(A, M) = \alpha + \beta_A A + \beta_M M $$
# Then
# $$ \mu(0, M) = \alpha + \beta_M M \approx \alpha $$
# Note that we are using $A=0$, but the conditional independence holds for any fixed $A$.
#
# ### The reverse is not true: D and A are not independent conditioned on M
# --------------------------
# ## Is the information about the single variable relationship "in" the multiple regression model?
# We have $p(D | M, A)$ from the multiple regression.
# Is there a relationship to $p(D | M)$?
# We have
# $$ E_A[p(D | A, M)p(A | M)] = E_A[p(D, A | M)] = p(D | M)$$
#
# Suppose we have a set of iid samples $D_i, A_i \sim p(D, A | M)$ for some fixed value of $M$. Then, the set of samples $\{D_i\}$ represents (approximately if the set is finite) the marginal distribution $E_A[p(D, A | M)] = p(D | M)$. **(By the way, we will use this fact later when we study MCMC sampling methods.)**
# Is this different from a single variable regression $p(D | M)$? Notice that $E[\beta_M] \approx 0.35$ from the single variable regression, whereas the association between $D$ and $M$ from the posterior of the multiple regression model works through $A$, which can be seen by substituting $A = \alpha_A + \beta_{MA}M$ into $D = \alpha + \beta_M M + \beta_{A}A$. Notice that $\beta_A \beta_{MA} \approx 0.42$.
az.summary(_D_M_posterior, var_names=['beta_M'])
az.summary(_D_A_posterior, var_names=['beta_A'])
az.summary(_D_MA_posterior, var_names=['beta_A', 'beta_M', 'beta_MA'])
# ### However, the association between $D$ and $M$ (conditioned on $A$) is small but not exactly zero. The association between $D$ and $M$ is actually
# $$ (\beta_M + \beta_A\beta_{MA})M \approx 0.36 M$$
print(around(D_M_posterior['beta_M'].mean(), 3))
print(around(D_MA_posterior['beta_M'].mean()
+ D_MA_posterior['beta_A'].mean()*D_MA_posterior['beta_MA'].mean(), 3))
# ## Exploring the DAG with the posterior predictive model
# What do the arrows really mean? In this case we assume that
# 1. $M$ is a function of $A$
# 2. $D$ is a function of $A$ and $M$
# We can simulate how $M$ and $D$ are predicted by $A$ for a range of different values of $A$.
# +
N_cf = 30
N_posterior_samples = D_AM_posterior.shape[0]
## intervention variable is Age
A_counter_factual = linspace(-2, 2, N_cf) ## the manipulated values for age
## first generate posterior predictive samples of M, using
## parameter posterior samples and intervention variable
alpha_M_post = array(D_AM_posterior['alpha_M'])
beta_AM_post = array(D_AM_posterior['beta_AM'])
sigma_M_post = array(D_AM_posterior['sigma_M'])
mu_M = (alpha_M_post[None, :]
+ beta_AM_post[None, :]*A_counter_factual[:, None])
M = stats.norm.rvs(
mu_M,
sigma_M_post[None, :],
size=(N_cf, N_posterior_samples))
## use above generated samples of M (along with parameter
## posterior samples) to generate samples of D
alpha_post = array(D_AM_posterior['alpha'])
beta_A_post = array(D_AM_posterior['beta_A'])
beta_M_post = array(D_AM_posterior['beta_M'])
sigma_post = array(D_AM_posterior['sigma'])
mu = (alpha_post[None, :]
+ beta_A_post[None, :]*A_counter_factual[:, None]
+ beta_M_post[None, :]*M)
D = stats.norm.rvs(
mu,
sigma_post[None, :],
size=(N_cf, N_posterior_samples))
## Plot the result: credible intervals etc
prob = 0.89
M_lower, M_upper = credible_interval_from_samples(M, prob)
D_lower, D_upper = credible_interval_from_samples(D, prob)
fig = figure(1, [15, 5])
fig.add_subplot(121)
fill_between(A_counter_factual, D_upper, D_lower, color='0.5', alpha=0.3)
plot(A_counter_factual, D.mean(axis=1), 'k')
ylabel('Simulated D', fontsize=20)
xlabel('A', fontsize=20);
fig.add_subplot(122)
fill_between(A_counter_factual, M_upper, M_lower, color='0.5', alpha=0.3)
plot(A_counter_factual, M.mean(axis=1), 'k')
ylabel('Simulated M', fontsize=20)
xlabel('A', fontsize=20);
# -
# ## Suppose we hold $A$ fixed and simulate $D$ with posterior predictive model for a range of values of $M$
# In this case, $M$ is no longer a function of $A$, so we remove that arrow in our DAG
CausalGraphicalModel(
nodes=['A', 'M', 'D'], edges=[('A', 'D'), ('M', 'D')]
).draw()
# +
N_cf = 30
N_posterior_samples = D_AM_posterior.shape[0]
## intervention variable is Age
M_counter_factual = linspace(-2, 2, N_cf) ## the manipulated values for age
A = zeros(N_cf)
## use above generated samples of M (along with parameter
## posterior samples) to generate samples of D
alpha_post = array(D_AM_posterior['alpha'])
beta_A_post = array(D_AM_posterior['beta_A'])
beta_M_post = array(D_AM_posterior['beta_M'])
sigma_post = array(D_AM_posterior['sigma'])
mu = (alpha_post[None, :]
+ beta_A_post[None, :]*A[:, None]
+ beta_M_post[None, :]*M_counter_factual[:, None])
D = stats.norm.rvs(
mu,
sigma_post[None, :],
size=(N_cf, N_posterior_samples))
## Plot the result: credible intervals etc
prob = 0.89
D_lower, D_upper = credible_interval_from_samples(D, prob)
figure(1, [7, 5])
fill_between(M_counter_factual, D_upper, D_lower, color='0.5', alpha=0.3)
plot(M_counter_factual, D.mean(axis=1), 'k')
ylabel('counterfactual D', fontsize=20)
xlabel('manipulated M', fontsize=20);
# -
## M has almost no effect on D, given that we know A
CausalGraphicalModel(
nodes=['A', 'M', 'D'], edges=[('A', 'D')]
).draw()
# ## We can also break the influence of $A$ on $M$ if we hold $M$ fixed and vary $A$
# +
N_cf = 30
N_posterior_samples = D_AM_posterior.shape[0]
## intervention variable is Age
A_counter_factual = linspace(-2, 2, N_cf) ## the manipulated values for age
M = zeros(N_cf)
## use above generated samples of M (along with parameter
## posterior samples) to generate samples of D
alpha_post = array(D_AM_posterior['alpha'])
beta_A_post = array(D_AM_posterior['beta_A'])
beta_M_post = array(D_AM_posterior['beta_M'])
sigma_post = array(D_AM_posterior['sigma'])
mu = (alpha_post[None, :]
+ beta_A_post[None, :]*A_counter_factual[:, None]
+ beta_M_post[None, :]*M[:, None])
D = stats.norm.rvs(
mu,
sigma_post[None, :],
size=(N_cf, N_posterior_samples))
## Plot the result: credible intervals etc
prob = 0.89
D_lower, D_upper = credible_interval_from_samples(D, prob)
figure(1, [7, 5])
fill_between(A_counter_factual, D_upper, D_lower, color='0.5', alpha=0.3)
plot(A_counter_factual, D.mean(axis=1), 'k')
ylabel('counterfactual D', fontsize=20)
xlabel('manipulated A', fontsize=20);
# -
# # Thought experiment: predicting height from leg length
# ## ... but suppose we do multiple regression on the length of both the left and right leg.
# We will generate a dataset to explore the idea.
## U is an unknown factor that determines both leg lengths
CausalGraphicalModel(
nodes=['L', 'R', 'H', 'U'], edges=[('U', 'L'), ('U', 'R'), ('L', 'H'), ('R', 'H')]
).draw()
N = 100
height = normal(10, 2, N)
leg_prop = linspace(0.4, 0.5, N)
leg_left = leg_prop*height + normal(0, 0.02, N)
leg_right = leg_prop*height + normal(0, 0.02, N)
# ### Now we imagine that this is a real dataset and that we want to use multiple regression to study the height predicted by the height of both legs
# \begin{align*}
# H_i &\sim \text{Normal}(\mu_i, \sigma) \\
# \mu_i &= \alpha + \beta_{L}L_i + \beta_{R}R_i \\
# \alpha &\sim \text{Normal}(10, 100) \\
# \beta_L &\sim \text{Normal}(2, 10) \\
# \beta_R &\sim \text{Normal}(2, 10) \\
# \sigma &\sim \text{Exponential}(1)
# \end{align*}
# Note that just like the book, we are using "...very vague, bad priors here, just so that we can be sure that the priors aren't responsible for what is about to happen."
with pm.Model() as model_H_LR:
sigma = pm.Exponential('sigma', 1)
beta_L = pm.Normal('beta_L', mu=2, sd=10)
beta_R = pm.Normal('beta_R', mu=2, sd=10)
alpha = pm.Normal('alpha', mu=10, sd=100)
## Note that an underscore '_' is nothing special;
## it is just another character that can be part of a variable name.
_mu = alpha + beta_L*leg_left + beta_R*leg_right
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H = pm.Normal('height', mu=mu, sd=sigma, observed=height)
## MCMC
_H_LR_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H_LR_posterior, var_names=['beta_L', 'beta_R', 'alpha', 'sigma']))
H_LR_posterior = pm.trace_to_dataframe(_H_LR_posterior)
az.plot_forest(_H_LR_posterior, var_names=["~mu"], combined=True, figsize=[5, 2]);
# ### Let's compare to the single variable regression. We will see that the variability for $\beta_L$ and $\beta_R$ is unusually large.
with pm.Model() as model_H_L:
sigma = pm.Exponential('sigma', 1)
beta_L = pm.Normal('beta_L', mu=2, sd=10)
alpha = pm.Normal('alpha', mu=10, sd=100)
## Note that an underscore '_' is nothing special;
## it is just another character that can be part of a variable name.
_mu = alpha + beta_L*leg_left
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H = pm.Normal('height', mu=mu, sd=sigma, observed=height)
## MCMC
_H_L_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H_L_posterior, var_names=['beta_L', 'alpha', 'sigma']))
H_L_posterior = pm.trace_to_dataframe(_H_L_posterior)
az.plot_forest(_H_L_posterior, var_names=["~mu"], combined=True, figsize=[5, 2]);
# ## The model is telling us that we know $\beta_L + \beta_R$ very well, but not $\beta_L - \beta_R$
# It shows that $\beta_L + \beta_R \approx 2$. On the other hand, $\beta_L - \beta_R$ can be anything so long as $\beta_L + \beta_R \approx 2$.
#
# A simple scatter plot of the posterior samples shows that the two parameters are negatively correlated.
plot(H_LR_posterior['beta_R'], H_LR_posterior['beta_L'], '.')
xlabel(r'$\beta_R$', fontsize=24)
ylabel(r'$\beta_L$', fontsize=24);
# ### A plot of the parameter posterior densities for $\beta_L + \beta_R$ and $\beta_L - \beta_R$
# We also show a density for $\beta_L$ from the single variable regression we did above. It matches very well with $\beta_L + \beta_R$ from the two variable regression.
# The second density plot for $\beta_L - \beta_R$ shows **much** more variability (look at the range of credible values). In fact, I'm not sure the Gaussian-like distribution is accurate here; remember, MCMC only gives us approximations, and they can sometimes fail.
#
# ### Summary: the weird variability in the posterior for $\beta_L$ and $\beta_R$ stems from the model's inability to determine $\beta_L - \beta_R$ from the data.
# This is an example of **non-identifiability.**
#
# The height is conditionally independent both ways:
# 1. given $L$, $R$ has give no additional information about $H$: $p(H | L, R) = p(H | L)$
# 2. given $R$, $L$ has give no additional information about $H$: $p(H | L, R) = p(H | R)$
# +
fig = figure(1, [12, 4])
fig.add_subplot(121)
sns.kdeplot(H_LR_posterior['beta_L'] + H_LR_posterior['beta_R'], label=r'$\beta_L + \beta_R$')
sns.kdeplot(H_L_posterior['beta_L'], label='single variable\nregression')
xlabel(r'$\beta_L + \beta_R$', fontsize=24)
ylabel('density', fontsize=20);
fig.add_subplot(122)
sns.kdeplot(H_LR_posterior['beta_L'] - H_LR_posterior['beta_R'])
xlabel(r'$\beta_L - \beta_R$', fontsize=24)
ylabel('density', fontsize=20);
# -
# # Example: Effect of anti-fungal soil treatment on plant growth
# Fungal growth inhibits plant growth. But the anti-fungal soil treatment inhibits fungal growth. We want to show that the treatment has a positive effect on plant growth.
CausalGraphicalModel(
nodes=['H0', 'H1', 'F', 'T'],
edges=[('H0', 'H1'), ('F', 'H1'), ('T', 'F')]
).draw()
N = 100
height_initial = normal(10, 2, N)
treatment = ones(N)
treatment[:N//2] = 0 ## integer division N//2 will always return an integer (insteaed of float)
## if the treatment is present (value = 1) then the probability of fungus is 0.1
## otherwise, if treatment=0, then the fungus is present with probability 0.5
fungus = stats.bernoulli.rvs(p=0.5 - treatment*0.4, size=N)
height_t1 = normal(height_initial + 5 - 3*fungus, 1, N)
# ## We will assume the above is a real dataset and build some models
# ### First, let's see what $H_0$ by itself tells us about $H_1$
with pm.Model() as model_H1_H0:
sigma = pm.Exponential('sigma', 1)
p = pm.Lognormal('p', mu=0, sd=0.25)
_mu = p*height_initial
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H1 = pm.Normal('height1', mu=mu, sd=sigma, observed=height_t1)
## MCMC
_H1_H0_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H1_H0_posterior, var_names=['p', 'sigma']))
H1_H0_posterior = pm.trace_to_dataframe(_H1_H0_posterior)
az.plot_forest(_H1_H0_posterior, var_names=["~mu"], combined=True, figsize=[5, 2]);
with pm.Model() as model_H1_H0TF:
sigma = pm.Exponential('sigma', 1)
alpha = pm.Lognormal('alpha', mu=0, sd=0.2)
beta_T = pm.Normal('beta_T', mu=0, sd=0.5)
beta_F = pm.Normal('beta_F', mu=0, sd=0.5)
p = pm.Deterministic('p', alpha + beta_T*treatment + beta_F*fungus)
_mu = p*height_initial
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H1 = pm.Normal('height1', mu=mu, sd=sigma, observed=height_t1)
## MCMC
_H1_H0TF_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H1_H0TF_posterior, var_names=['beta_T', 'beta_F', 'alpha', 'sigma']))
H1_H0TF_posterior = pm.trace_to_dataframe(_H1_H0TF_posterior)
az.plot_forest(
_H1_H0TF_posterior,
var_names=['beta_T', 'beta_F', 'alpha', 'sigma'],
combined=True,
figsize=[5, 2]);
# ## The model is telling us that if we know $F$ then $T$ tells us nothing about growth
# If we misinterpret this, it seems to tell us that treatment has no effect on growth.
# But really it is telling us that once we know if fungus is present, knowing if the treatment was given tells us nothing about the growth.
# This is because the treatment inhibits fungal growth, so if we see fungal growth then presumably the treatment was either not given or was ineffective.
#
# ### If we condition on $F$ we make $H_1$ independent of $T$
# ### ...we block the path from $T$ to $H_1$
anti_fungal = CausalGraphicalModel(
nodes=['H0', 'H1', 'F', 'T'], edges=[('H0', 'H1'), ('F', 'H1'), ('T', 'F')]
)
anti_fungal.draw()
# ### We can show the conditional independencies implied by the graph
all_independencies = anti_fungal.get_all_independence_relationships()
for s in all_independencies:
if 'H1' in [s[0], s[1]]: ## Print only the independencies involving H1
print(s[0], 'is independent of', s[1], 'given', s[2])
# ## So we should not use $F$ in our model?
# Let's try
with pm.Model() as model_H1_H0T:
sigma = pm.Exponential('sigma', 1)
alpha = pm.Lognormal('alpha', mu=0, sd=0.2)
beta_T = pm.Normal('beta_T', mu=0, sd=0.5)
p = pm.Deterministic('p', alpha + beta_T*treatment)
_mu = p*height_initial
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H1 = pm.Normal('height1', mu=mu, sd=sigma, observed=height_t1)
## MCMC
_H1_H0T_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H1_H0T_posterior, var_names=['beta_T', 'alpha', 'sigma']))
H1_H0T_posterior = pm.trace_to_dataframe(_H1_H0T_posterior)
az.plot_forest(_H1_H0T_posterior, var_names=['beta_T', 'alpha', 'sigma'], combined=True, figsize=[5, 2]);
# ### The above shows that once we remove the conditioning on the presence of fungus, our model shows a positive effect of the treatment on growth
# ## Summary: conditioning on $F$ closes the path between $H_1$ and $T$
# # Example: Anti-fungal soil treatment: Version 2
# ## Let's change the plant growth model slightly so that fungal growth has no effect on plant growth
# Add in the unobserved effect of moisture $M$
anti_fungal2 = CausalGraphicalModel(
nodes=['H0', 'H1', 'F', 'T', '(M)'], edges=[('H0', 'H1'), ('T', 'F'), ('(M)', 'H1'), ('(M)', 'F')]
)
anti_fungal2.draw()
N = 100
height_initial = normal(10, 2, N)
treatment = ones(N)
treatment[:N//2] = 0
moisture = stats.bernoulli.rvs(p=0.5, size=N)
fungus = stats.bernoulli.rvs(p=0.5 - treatment*0.4 + moisture*0.4, size=N)
height_t1 = height_initial + normal(5 + 3*moisture, 1, N)
# ## Rerun models again
with pm.Model() as model_H1_H0TF:
sigma = pm.Exponential('sigma', 1)
alpha = pm.Lognormal('alpha', mu=0, sd=0.2)
beta_T = pm.Normal('beta_T', mu=0, sd=0.5)
beta_F = pm.Normal('beta_F', mu=0, sd=0.5)
p = pm.Deterministic('p', alpha + beta_T*treatment + beta_F*fungus)
_mu = p*height_initial
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H1 = pm.Normal('height1', mu=mu, sd=sigma, observed=height_t1)
## MCMC
_H1_H0TF_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H1_H0TF_posterior, var_names=['beta_T', 'beta_F', 'alpha', 'sigma']))
H1_H0TF_posterior = pm.trace_to_dataframe(_H1_H0TF_posterior)
# ## The above seems to tell us that fungus helps the plants grow!
# Look at $\beta_F$
# ### This time treatment should not affect growth
# Does this agree with $\beta_T$?
with pm.Model() as model_H1_H0T:
sigma = pm.Exponential('sigma', 1)
alpha = pm.Lognormal('alpha', mu=0, sd=0.2)
beta_T = pm.Normal('beta_T', mu=0, sd=0.5)
p = pm.Deterministic('p', alpha + beta_T*treatment)
_mu = p*height_initial
mu = pm.Deterministic('mu', _mu) ## we use this so that we can sample the posterior predictive later
H1 = pm.Normal('height1', mu=mu, sd=sigma, observed=height_t1)
## MCMC
_H1_H0T_posterior = pm.sample(1000, tune=1000, cores=4)
print(pm.summary(_H1_H0T_posterior, var_names=['beta_T', 'alpha', 'sigma']))
H1_H0T_posterior = pm.trace_to_dataframe(_H1_H0T_posterior)
# ## The above tells us that removing the conditioning on presence of fungus still tells us that treatment has no association with growth, as expected
## F is an example of a collider, which closes the path between H1 and T
## unless we open it by conditioning on F
anti_fungal2 = CausalGraphicalModel(
nodes=['H0', 'H1', 'F', 'T', '(M)'], edges=[('H0', 'H1'), ('T', 'F'), ('(M)', 'H1'), ('(M)', 'F')]
)
anti_fungal2.draw()
# ## Summary: conditioning on $F$ opens a path from $H_1$ to $T$
# -------------------------------
# # Example: Collider Bias:
# ## Trustworthiness, Newsworthiness, and Selection of research grants
# The path through a collider is closed unless we condition on the collider variable (in this case $S$). Conditioning on $S$ opens the path between $T$ and $N$, creating spurious correlation between them.
#
# **Note that this is the opposite of the age, marriage, divorce relationship in our first example. The arrows are pointing in the opposite direction.**
CausalGraphicalModel(
nodes=['T', 'S', 'N'], edges=[('T', 'S'), ('N', 'S')]
).draw()
N = 200 # num grant proposals
p = 0.1 # proportion to select
# uncorrelated newsworthiness and trustworthiness
nw = normal(size=N)
tw = normal(size=N)
# select top 10% of combined scores
score = nw + tw # total score
q = quantile(score, 1 - p) # top 10% threshold
selected = score >= q
cor = corrcoef(tw[selected], nw[selected])
cor
# +
figure(1, [7, 4])
plot(nw[selected == False], tw[selected == False], '.')
plot(nw[selected], tw[selected], '.', label='selected')
legend()
# correlation line
xn = array([-2, 3])
plot(xn, tw[selected].mean() + cor[0, 1] * (xn - nw[selected].mean()), 'k')
xlabel('newsworthiness', fontsize=20)
ylabel('trustworthiness', fontsize=20);
# -
# ## Summary: conditioning on a collider variable opens a path between two variables
# ---------------------
# # Example: The haunted DAG: predicting child education from parent education and grandparent education
# The DAG is "haunted" by $U$ an unobserved effect of neighborhood on parent and child education
#
# **This is an important and very counter intuitive example, and I am leaving it as an exercise.**
CausalGraphicalModel(
nodes=['G', 'P', 'C', '(U)'], edges=[('G', 'P'), ('P', 'C'), ('(U)', 'P'), ('(U)', 'C')]
).draw()
# # The four elemental confounds
# 1. The Fork
# 2. The Pipe
# 3. The Collider
# 4. The Descendant
#
# ### Think of situations where we want to predict outcome variable Y with predictor variable X
# ## 1. Fork
CausalGraphicalModel(
nodes=['X', 'Y', 'Z'], edges=[('Z', 'X'), ('Z', 'Y')]
).draw()
# ### Path from X to Y is open
# ### can be closed by conditioning on Z
# ## 2. Pipe
CausalGraphicalModel(
nodes=['X', 'Y', 'Z'], edges=[('X', 'Z'), ('Z', 'Y')]
).draw()
# ### Path from X to Y is open
# ### can be closed by conditioning on Z
# ## 3. Collider
CausalGraphicalModel(
nodes=['X', 'Y', 'Z'], edges=[('X', 'Z'), ('Y', 'Z')]
).draw()
# ### Path from X to Y is closed
# ### can be opened by conditioning on Z
# ## 4. Descendant
CausalGraphicalModel(
nodes=['X', 'Y', 'Z', 'D'], edges=[('X', 'Z'), ('Y', 'Z'), ('Z', 'D')]
).draw()
# ### Path from X to Y is closed
# ### can be opened by conditioning on Z or on D
# # Example: Removing confounds in a DAG
# ## Assume that we want to predict outcome $Y$ with predictor $X$
#
CausalGraphicalModel(
nodes=['A', 'B', 'C', 'X', 'Y', '(U)'],
edges=[('X', 'Y'), ('(U)', 'X'), ('(U)', 'B'), ('A', '(U)'), ('A', 'C'), ('C', 'B'), ('C', 'Y')]
).draw()
# ## Look for "backdoor" paths (arrows going into $X$)
# There should be two additional "backdoor" paths
#
# 1. X <- (U) -> B <- C -> Y
# 2. X <- (U) <- A -> C -> Y
#
# ## Are the paths "open" or "closed"?
# ## If any are open, how do we close it?
# # Example: Backdoor waffles
# 1. S is whether the state is in the "South"
# 2. W is the number of Waffle Houses
# 3. D is divorce rate
# 4. M is marriage rate
# 5. A is median age at marriage
#
# ## We want to understand the effect of W on D
CausalGraphicalModel(
nodes=['A', 'D', 'S', 'M', 'W'],
edges=[('A', 'M'), ('A', 'D'), ('S', 'A'), ('S', 'W'), ('S', 'M'), ('W', 'D')]
).draw()
# ## There are three backdoor paths
# 1.
# 2.
# 3.
| Week 7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
df = pd.read_csv('data/hacc.csv')
df.head()
len(df)
print(df.columns)
df.columns = ['name', 'session_type', 'start_time', 'end_time', 'duration', 'energy', 'amount', 'id', 'port_type', 'payment_mode']
print(df.columns)
df.dtypes
# +
df['start_time'] = pd.to_datetime(df['start_time'])
df['end_time'] = pd.to_datetime(df['end_time'])
# Changes the payment amount to a raw float value instead of a string of the form "$X.XX"
df['amount'] = df['amount'].replace('[\$,]', '', regex=True).astype(float)
df.head()
# -
df[df['duration'].str.contains('-')]['duration']
# ## Data Errors
# Types of errors found:
# 1. Positive energy, Zero amount -- Resolution: ?
# 2. Positive amount, Zero energy -- Resolution: ?
# Checking for any missing values
missing = df[(df['energy'] == 0) | (df['amount'] == 0)]
print(len(missing))
missing.head()
# Anyone who charges their car but doesn't pay is a "thief" -- correctable if we just use HECO formula?
# Question: Is this ALWAYS just a data issue or do some people actually get away without paying?
thieves = df[(df['energy'] > 0) & (df['amount'] == 0)]
print(len(thieves))
thieves.head()
# Anyone who used 0 energy but still paid got "jipped"
jipped = df[(df['energy'] == 0) & (df['amount'] > 0)]
print(len(jipped))
jipped.head()
dates = df['start_time'].dt.date
df['date'] = dates
df['date']
df['day_of_week'] = df['start_time'].dt.day_name()
df['day_of_week']
# ## Time of Day
# Noting from his slides:
#
# Times:
# - On Peak: 5pm - 10pm => 17:00 - 22:00, Daily
# - Mid Day: 9am - 5pm => 9:00 => 17:00, Daily
# - Off Peak: 10pm - 9am => 22:00 - 9:00, Daily
#
# Cost:
# - On Peak: \$0.57
# - Mid Day: \$0.49
# - Off Peak: \$0.54
#
import datetime as dt
start_times = df['start_time'].dt.time
df['on_peak'] = (dt.time(17, 0, 0) <= start_times) & (start_times < dt.time(22, 0, 0))
df['mid_day'] = (dt.time(9, 0, 0) <= start_times) & (start_times < dt.time(17, 0, 0))
df['off_peak'] = (dt.time(22, 0, 0) <= start_times) | (start_times < dt.time(9, 0, 0))
df.head()
# ## Error Checking
# Checking if each columns are in agreement with each other
# 1. Does cost match with the amount of energy for the given time period?
df['calculated_amount'] = df['energy'] * df['on_peak'] * 0.57 + df['energy'] * df['mid_day'] * 0.49 + df['energy'] * df['off_peak'] * 0.54
df['rounded_calculated_amount'] = np.round(df['calculated_amount'], 2)
correct = df[(df['amount'] == df['rounded_calculated_amount'])]
err = df[~(df['amount'] == df['rounded_calculated_amount'])]
correct.head()
err[np.abs(err['amount'] - err['rounded_calculated_amount']) == 0.01]#[['amount', 'rounded_calculated_amount', 'calculated_amount']]
err[np.abs(err['amount'] - err['rounded_calculated_amount']) > 1]
df = df.rename({'rounded_calculated_amount': 'correct_amount'}, axis=1)
df['error_rounding'] = np.abs(df['amount'] - df['correct_amount']) == 0.01
df['error_calculation'] = np.abs(df['amount'] - df['correct_amount']) > 0.01
df.head()
for col in ['session_type', 'port_type', 'payment_mode']:
df[col] = df[col].astype('category')
df.dtypes
# +
preproc_df = df.join(pd.get_dummies(df.select_dtypes('category')))
# preproc_df = preproc_df.join(pd.get_dummies(df['name']))
# Drop original categorical columns in favor of the "One Hot Encoding"
preproc_df = preproc_df.drop(df.select_dtypes('category'), axis=1)
# preproc_df['start_time'] = pd.to_timedelta(preproc_df['start_time'])
# preproc_df['end_time'] = pd.to_timedelta(preproc_df['end_time'])
# preproc_df['correct_duration'] = preproc_df['end_time'].dt.total_seconds() - preproc_df['start_time'].dt.total_seconds()
def get_sec(time_str):
"""Get Seconds from time."""
h, m, s = time_str.split(':')
return int(h) * 3600 + int(m) * 60 + int(s)
preproc_df['correct_duration'] = preproc_df['duration'].apply(lambda x: get_sec(x))
preproc_df = preproc_df.drop(['id', 'start_time', 'end_time', 'duration', 'amount', 'calculated_amount', 'day_of_week'], axis=1)
preproc_df.head()
# -
df_agg = preproc_df.groupby(['name', 'date']).agg('sum').reset_index()
# df_agg.columns = df_agg.columns.to_flat_index()
# df_agg.columns
df_agg.head()
df_agg.columns
import matplotlib.pyplot as plt
plt.scatter(df_agg['energy'].shift(-1), df_agg['energy'])
from pandas.plotting import scatter_matrix
df_temp = pd.DataFrame()
df_temp['energy'] = df_agg['energy']
for x in range(1, 7):
df_temp[f'energy_prev_{x}'] = df_temp['energy'].shift(x)
scatter_matrix(df_temp[['energy', 'energy_prev_1', 'energy_prev_2', 'energy_prev_3']])
df_temp.dropna().head(10)
# +
df_agg = preproc_df.groupby(['name', 'date']).agg('sum').reset_index()
df_agg['day_of_week'] = pd.to_datetime(df_agg['date']).dt.day_name()
df_agg['month'] = pd.to_datetime(df_agg['date']).dt.month_name()
for col in ['day_of_week', 'month']:
df_agg[col] = df_agg[col].astype('category')
df_agg = df_agg.join(pd.get_dummies(df_agg.select_dtypes('category')))
df_agg = df_agg.drop(df_agg.select_dtypes('category'), axis=1)
# df_agg.columns = df_agg.columns.to_flat_index()
# df_agg.columns
# for x in range(1, 7):
# df_agg[f'energy_prev_{x}'] = df_agg['energy'].shift(x)
# df_agg = df_agg.dropna()
# df_agg.head()
stations = [g for _, g in df_agg.groupby('name')]
def offset_col_x_days(df, col, days):
for x in range(1, days):
df[f'{col}_prev_{7+x}'] = df[col].shift(7+x)
df = df.dropna().reset_index(drop=True)
return df
for i in range(len(stations)):
for col in ['energy', 'on_peak', 'mid_day', 'off_peak', 'error_rounding', 'error_calculation', 'session_type_DEVICE', 'session_type_MOBILE', 'session_type_WEB',
'port_type_CHADEMO', 'port_type_DCCOMBOTYP1', 'payment_mode_CREDITCARD',
'payment_mode_RFID']:
stations[i] = offset_col_x_days(stations[i], col, 7)
stations[0]
# -
stations[1]['date'][0]
plt.plot(stations[0]['energy'])
plt.plot(stations[1]['energy'])
stations[1]
stations[0]
# +
X = stations[0]
y = X['energy']
train_test_split = int(len(X) * 0.8)
date_train, date_test = X[:train_test_split]['date'], X[train_test_split:]['date']
X = X.drop(['name', 'date', 'correct_amount', 'correct_duration', 'energy', 'on_peak', 'mid_day', 'off_peak', 'error_rounding', 'error_calculation', 'session_type_DEVICE', 'session_type_MOBILE', 'session_type_WEB',
'port_type_CHADEMO', 'port_type_DCCOMBOTYP1', 'payment_mode_CREDITCARD',
'payment_mode_RFID'], axis=1)
X_train, X_test = X[:train_test_split], X[train_test_split:]
y_train, y_test = y[:train_test_split], y[train_test_split:]
# -
X_train.head()
X_train.columns
plt.plot(y_train)
plt.plot(X_train['energy_prev_2'])
# +
from sklearn.tree import DecisionTreeRegressor
clf = DecisionTreeRegressor()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
score = clf.score(X_test, y_test)
plt.scatter(y_pred, y_test)
score
# +
from sklearn.linear_model import LinearRegression
clf = LinearRegression()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
score = clf.score(X_test, y_test)
plt.scatter(y_pred, y_test)
score
# +
# from sklearn.utils import check_arrays
def mape(y_true, y_pred):
# y_true, y_pred = check_arrays(y_true, y_pred)
## Note: does not handle mix 1d representation
#if _is_1d(y_true):
# y_true, y_pred = _check_1d_array(y_true, y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
mape(y_test, y_pred)
# -
sorted(y_pred - y_test)
plt.scatter(range(len(date_test)), y_test)
df_test = pd.DataFrame({'date': date_test, 'energy': y_test})
df_test.to_csv('test_run.csv',index=False)
| eda/energy_forecast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from data_loader.dataloader import *
from data_loader import dataloader
import numpy as np
import torch
import torch.nn as nn
import matplotlib.pyplot as plt
from model.model import *
from tqdm import tqdm
seed = 1
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
np.random.seed(seed)
model = torch.load('trained_models/1e-05_61.pt')
model.cuda()
metadata = './data/HAM10000_metadata.csv'
images = './data/'
batch=1024
_,_,test_data = dataloader.prepare_data(metadata,all_classes[:-1],images,create_split=True,split=(0.64,0.16,0.2),batch=batch)
outputs = []
labels = []
total = 0
correct = 0
with torch.no_grad():
for data,target in tqdm(test_data):
data,target = data.cuda(),target.cuda()
output = model(data)
_, predicted = torch.max(output.data, 1)
outputs.append(predicted.detach().cpu())
labels.append(target.detach().cpu())
# smax = F.softmax(output,1)
# print(predicted)
# total += target.size(0)
# correct += (predicted == target).sum().item()
# print(correct/total)
pred=np.hstack(outputs)
label=np.hstack(labels)
import sklearn.metrics as sk
reprt=sk.classification_report(label,pred, target_names=all_classes[:-1])
print(reprt)
model = torch.load('trained_models/1e-05_61.pt')
model.cuda()
import sklearn.metrics as sk
def classification_metrics(prediction,labels):
accuracy = sk.accuracy_score(labels,prediction)
f1_score = sk.f1_score(labels,prediction)
precision = sk.precision_score(labels,prediction)
recall = sk.recall_score(labels,prediction)
return accuracy, f1_score,precision, recall
a,f,p,r = classification_metrics(outputs,labels)
outputs
| Trial test and results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nodejs MNIST model Deployment
#
# * Wrap a nodejs tensorflow model for use as a prediction microservice in seldon-core
# * Run locally on Docker to test
#
# ## Dependencies
#
# * ```pip install seldon-core```
# * [Helm](https://github.com/kubernetes/helm)
# * [Minikube](https://github.com/kubernetes/minikube)
# * [S2I](https://github.com/openshift/source-to-image)
# * node (version>=8.11.0)
# * npm
#
# ## Train locally using npm commands
# This model takes in mnist images of size 28x28x1 as input and outputs an array of size 10 with prediction of each digits from 0-9
# !make train && make clean_build
# Training creates a model.json file and a weights.bin file which is utilized for prediction
# ## Prediction using REST API on the docker container
# !s2i build . seldonio/seldon-core-s2i-nodejs:0.2-SNAPSHOT node-s2i-mnist-model:0.1
# !docker run --name "nodejs_mnist_predictor" -d --rm -p 5000:5000 node-s2i-mnist-model:0.1
# Send some random features that conform to the contract
# !seldon-core-tester contract.json 0.0.0.0 5000 -p -t
# !docker rm nodejs_mnist_predictor --force
# ## Prediction using GRPC API on the docker container
# !s2i build -E ./.s2i/environment_grpc . seldonio/seldon-core-s2i-nodejs:0.2-SNAPSHOT node-s2i-mnist-model:0.2
# !docker run --name "nodejs_mnist_predictor" -d --rm -p 5000:5000 node-s2i-mnist-model:0.2
# Send some random features that conform to the contract
# !seldon-core-tester contract.json 0.0.0.0 5000 -p -t --grpc
# !docker rm nodejs_mnist_predictor --force
# ## Test using Minikube
#
# **Due to a [minikube/s2i issue](https://github.com/SeldonIO/seldon-core/issues/253) you will need [s2i >= 1.1.13](https://github.com/openshift/source-to-image/releases/tag/v1.1.13)**
# !minikube start --memory 4096
# !kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin --serviceaccount=kube-system:default
# !helm init
# !kubectl rollout status deploy/tiller-deploy -n kube-system
# !helm install ../../../helm-charts/seldon-core-operator --name seldon-core --set usageMetrics.enabled=true --namespace seldon-system
# !kubectl rollout status statefulset.apps/seldon-operator-controller-manager -n seldon-system
# ## Setup Ingress
# Please note: There are reported gRPC issues with ambassador (see https://github.com/SeldonIO/seldon-core/issues/473).
# !helm install stable/ambassador --name ambassador --set crds.keep=false
# !kubectl rollout status deployment.apps/ambassador
# !eval $(minikube docker-env) && s2i build . seldonio/seldon-core-s2i-nodejs:0.2-SNAPSHOT node-s2i-mnist-model:0.1
# !kubectl create -f nodejs_mnist_deployment.json
# !kubectl rollout status deploy/nodejs-mnist-deployment-nodejs-mnist-predictor-5aa9edd
# !seldon-core-api-tester contract.json `minikube ip` `kubectl get svc ambassador -o jsonpath='{.spec.ports[0].nodePort}'` \
# seldon-deployment-example --namespace seldon -p
# !minikube delete
| examples/models/nodejs_mnist/nodejs_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Reference - Encapsulation
# **Author:** <NAME>
#
# #### Definition
# one of the three pillars of object oriented programming
#
# ##### Links
# https://pythonspot.com/encapsulation/
# https://pythonprogramminglanguage.com/encapsulation/
# https://medium.com/@manjuladube/encapsulation-abstraction-35999b0a3911
# ### protected and private
# **protected** members should only be accessed in derived classes. just follow the convention by prefixing the name of the member by a single underscore **“_”**
# **private** members should only be accessed in the class itself, just follow the convention by prefixing the name of the member by double underscores **“__”**
class Car():
__maxspeed = 0
def __init__(self, maxspeed):
self.__maxspeed = maxspeed
self.__updateSoftware()
def drive(self):
print("driving with maxspeed " + str(self.__maxspeed))
def __updateSoftware(self):
print("updating software")
# ##### however...
# that is only a naming convention. **protected** members can be accessed from anywhere. **private** members can be accessed by prefixing the name of a private member with an underscore **"_"** and the name of the containing class to access it
redcar = Car(200)
redcar.drive()
# redcar.__maxspeed = 130 # will throw an error
redcar._Car__maxspeed = 130 # access the private variable like this
# redcar.__updateSoftware() # will throw an error
redcar._Car__updateSoftware() # access the private function like this
redcar.drive()
| OOP/Encapsulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lists
# +
myList=[11,42,15,64,80,32,12] #creating a list of the same data type
print(myList[0:3]) #printing items from zero to three
print(myList[:3]) #printing items from zero to three
print(myList[3:]) #printing items from three to last
print(myList[2:5]) #printing items from two to five
# -
myList=[10,12.5,'H',"Hello"] #Creating a list of different data types
print(myList) #printing the whole list
print(myList[0]) #printing the 0th indexed item of the list
print(myList[1]) #printing the 1st indexed item of the list
# +
myList = ['Maths', 'Science', 2020, 2021]
print ("Value available at index 2 : ", myList[2])
myList[2] = 2001 #Assigning new value to the index:2
print ("New value available at index 2 : ", myList[2])
# +
myList = ['Maths', 'Science', 2020, 2021]
print ("Initial list : ",myList)
del myList[2]
print ("After deleting value at index 2 : ", myList)
# -
myList=[11,42,15,64,80,32,12]
print ("Length of the list : ", len(myList))
print ("Maximum value element : ", max(myList))
print ("Minimum value element : ", min(myList))
print ("Sum of the list values : ", sum(myList))
# +
myList=[11,42,15,64,80,32,12,42]
print ("Initial list : ",myList) #Original List
myList.append(50) #Appending the value 50 to the end of the list
print ("updated list : ", myList) #Printing the list after appending the value
print ("Count for 42 : ", myList.count(42)) #Printing the no of times 42 appears
print ('Index of 15 : ', myList.index(15)) #Printing the index of the value 15
list2 = [5,10,15] #Creating a second list
myList.extend(list2) #Appending the second list to the first list
print ('Extended List : ', myList)
# -
# # Tuples
# +
tup1 = ('Maths', 'Science', 2020, 2021) #Creating a python tuple with parentheses
tup2 = "a", "b", "c", "d" #Creating a python tuple without parentheses
print ("tup1[0] : ", tup1[0]) #Printing the 0th indexed value
print ("tup2[1:5] : ", tup2[1:5]) #Printing the values from one to five
# +
myTuple = (11,42,15,64,80,32,12)
# Following action is not valid for tuples
myTuple[0] = 100
# -
myTuple = (11,42,15,64,80,32,12)
print(max(myTuple)) #printing the max value of the tuple
print(len(myTuple)) #printing the length of the tuple
# +
myTuple = (11,42,15,64,80,32,12)
myList=list(myTuple) #Converting tuple into a list
print(myList) #Print the initial list
myList.append(100) #Appending the value 100 to the list
myList[0]=20 #Assign 20 as the 0th indexed value
print(myList) #Print the changed list
# -
# # Dictionaries
# +
Student={'name':'John','maths':82,'phy':88,'chem':92,'english':80}
print(Student['name']) #Printing the value of the key:name
# +
Student={'name':'John','maths':82,'phy':88,'chem':92,'english':80}
print('Original dictionary : ',Student)
Student['name']='William' #Changing the value for the key:name
print('After changing the value : ',Student)
# -
myDict = {'Name': 'John', 'Age': 20, 'Name': 'William'}
print ("myDict['Name']: ", myDict['Name'])
myDict = {['Name']: 'John', 'Age': 20}
print ("dict['Name']: ", myDict['Name'])
# +
myDict={'name':'John','maths':82,'phy':88,'chem':92,'english':80}
print("Length of the dict : ", len (myDict))
dict2 = myDict.copy()
print("Copied Dictionary : ",dict2)
keys=myDict.keys()
print("Dictionary keys : ",keys)
vals=myDict.values()
print("Dictionary values : ",vals)
# -
| Tutorial 2 - Lists, Tuples, and Dictionaries.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
# # Plots.jl gallery
#
# Sources:
#
# - [PlotsGallery.jl](https://github.com/goropikari/PlotsGallery.jl) by goropikari
# - [Julia Plots docs](https://docs.juliaplots.org/)
# + [markdown] tags=[]
# ## Attributes
# -
using Plots, Measures
# ### Tick size and properties
plot(sin, 0, 2π,
xticks=0:0.5:2π,
xrotation=60,
xtickfontsize=25,
bottom_margin=15mm)
plot(sin, 0, 2π,
xtick=(0:0.5:2π, ["$i a" for i in 0:0.5:2π]),
ytick=-1:0.2:1,
xrotation=60, ## -> rotate xticks 60 degree
yrotation=90,
# rotation=60, # x,y ticks rotation
)
# ### No axis
#
# `plot(..., axis=false)`
plot(sin, 0, 2π, axis=false)
# ### Log scale for axes
#
# - `xscale=:log10`
# - `yscale=:log10`
# - `scale=:log10`
plot(exp, -5, 5, yscale=:log10, title="semilogy", legend=nothing)
plot(log, 1e-5, 10, xscale=:log10, title="semilogx", legend=nothing)
plot(x->x^1.7, 1e-3, 3, scale=:log10, title="log-log", legend=nothing)
# ### Axis range
#
# - `xlims=(a, b)`
# - `ylims=(c, d)`
plot(sin, 0, 2π, xlims=(-10, 10), ylims=(-2,2))
# ### Scietific notation
#
# - `xformatter=:scientific`
# - `yformatter=:scientific`
plot(exp, 0, 10, yformatter=:scientific)
# ### Flip Axis
plot(x->x, 0:0.01:2π,
proj=:polar,
xflip=true,
yflip=true,
# flip=true # x,y-flip
)
# ### Aspect ratio
using Plots
using Random
Random.seed!(2021)
s = bitrand(10, 10)
heatmap(s,
aspect_ratio=:equal,
c=:blues,
colorbar=false)
# ### Fonts
#
# LaTeX fonts are supported via `LaTeXStrings` package.
# +
using Plots, Measures, LaTeXStrings
plot(sin, 0, 2π,
title=L"y = \sin(x)",
titlefont=font(40), ## title
xlabel=L"x",
ylabel="y",
xguidefontsize=30, ## x-guide
yguidefontsize=20, ## y-guide
# guidefontsize=20, # both x,y-guide
xtick=(0:0.5:2π, ["\$ $(i) \$" for i in 0:0.5:2π]),
ytick=-1:0.5:1,
xtickfontsize=15,
ytickfontsize=20,
# tickfontsize=10, # for both x and y
label="Sin function",
legendfontsize=12,
xlims=(0,2π),
ylims=(-1,1),
bottom_margin=5mm,
left_margin=10mm,
top_margin=15mm
)
# +
fib(x) = (((1+sqrt(5))/2)^x - ((1-sqrt(5))/2)^x)/sqrt(5)
ann = L"F_n = \frac{1}{\sqrt{5}} \left[\left( \frac{1+\sqrt{5}}{2} \right)^n - \left( \frac{1-\sqrt{5}}{2} \right)^n \right]"
plot(fib, 1:12, marker=:circle, xlabel=L"n", ylabel=L"F_n", annotation=(5, 100, ann))
# -
# ## Bar plots
# +
using Plots, StatsPlots, StatsBase, Random
Plots.gr(fmt=:png)
Random.seed!(2021)
# Data
measles = [38556, 24472, 14556, 18060, 19549, 8122, 28541, 7880, 3283, 4135, 7953, 1884]
mumps = [20178, 23536, 34561, 37395, 36072, 32237, 18597, 9408, 6005, 6268, 8963, 13882]
chickenPox = [37140, 32169, 37533, 39103, 33244, 23269, 16737, 5411, 3435, 6052, 12825, 23332]
ticklabel = string.(collect('A':'L'))
# -
# ### Grouped vertical bar plots
#
# Requires `StatsPlots` package.
#
# Use `groupedbar(data, bar_position = :dodge)`
groupedbar([measles mumps chickenPox], bar_position = :dodge, bar_width=0.7, xticks=(1:12, ticklabel), label=["measles" "mumps" "chickenPox"])
# ### Stacked vertical bar plots
#
# Requires `StatsPlots` package.
#
# Use `groupedbar(data, bar_position = :stack)`
groupedbar([measles mumps chickenPox],
bar_position = :stack,
bar_width=0.7,
xticks=(1:12, ticklabel),
label=["measles" "mumps" "chickenPox"])
# ### Horizontal Bar Plot
#
# `bar(data, orientation=:h)`
bar(1:12, orientation=:h, yticks=(1:12, ticklabel), yflip=true)
# ### Categorical Histogram Plot
# +
# Data
status = ["Poor", "Fair", "Good", "Excellent"]
data = sample(status, Weights([1,1,2,2]), 100)
datamap = countmap(data)
# Plot
bar((x -> datamap[x]).(status), xticks=(1:4, status), legend=nothing)
# -
# ## Histogram
# +
using Plots
Plots.gr(fmt=:png)
using Random
Random.seed!(2021)
x = randn(1000)
y = randn(1000)
z = randn(1000)
histogram(x, bins=20, alpha=0.4, label="A")
histogram!(y, bins=20, alpha=0.4, label="B")
histogram!(z, bins=20, alpha=0.4, label="C")
# -
# ## Box plots
# +
using Plots, StatsPlots, Statistics, Random
Plots.gr(fmt=:png)
Random.seed!(2021)
n = 30
science = rand(1:10, n)
boxplot(science, label="science")
# -
english = rand(1:10, n)
boxplot([science english], label=["science" "english"])
# ## Contour Plots
# ### Over a function
#
# `contour(xs, ys, f)` where `f(x, y) = ...`
# +
using Plots
Plots.gr(fmt=:png)
xs = range(0, stop=2, length=50)
ys = range(0, stop=2, length=50)
f(x , y) = x^2 + y^2
contour(xs, ys, f)
# -
# ### Contour plot over an array
#
# `contour(x1d, y1d, xy2d)`
zz = f.(xs', ys) # Notice the transpose over xs
contour(xs, ys, zz)
# ### Filled Contour Plots
#
# - `contour(xs, ys, f, fill=true)`
# - `contourf(xs, ys, f)`
# +
#=
contourf(0:0.01:5, 0:0.01:5,
(x, y) -> sin(3x) * cos(x+y),
xlabel="x", ylabel="y")
=#
contour(0:0.01:5, 0:0.01:5, (x, y) -> sin(3x) * cos(x+y), xlabel="x", ylabel="y", fill=true)
# -
# ## Datetime plot
#
# - Use `Dates` package and `Data` data type
# - Customize ticks
# +
using Plots, Measures, Random, Dates
Plots.gr(fmt=:png)
Random.seed!(2021)
days = 31
position = cumsum(randn(days))
x = Date(2018,1,1):Day(1):Date(2018,1,31)
ticks = [x[i] for i in 1:5:length(x)]
plot(x, position,
xlabel="Date",
ylabel="Position",
title="Track of random walker",
legend=nothing,
xticks=ticks,
xrotation=45,
bottom_margin=5mm,
left_margin=5mm)
# -
# ## Error bar
#
# `plots(..., xerr=xerr, yerr=yerr)`
# +
using Plots
using Random
Random.seed!(2021)
f(x) = 2 * x + 1
x = 0:0.1:2
n = length(x)
y = f.(x) + randn(n)
plot(x, y,
xerr=0.1 * rand(n),
yerr=rand(n),
legend=nothing)
# -
# ## Heatmap
#
# `heatmap(data)`
# +
using Plots
Plots.gr(fmt=:png)
using Random
Random.seed!(2021)
a = rand(5,5)
xlabel = string.(collect('A':'E'))
ylabel = string.(collect('a':'e'))
heatmap(a, xticks=(1:5, xlabel),
yticks=(1:5, ylabel),
aspect_ratio=:equal)
fontsize = 15
nrow, ncol = size(a)
# Add number annotations to plots
ann = [(i,j, text(round(a[i,j], digits=2), fontsize, :white, :center))
for i in 1:nrow for j in 1:ncol]
annotate!(ann, linecolor=:white)
# -
# ## Line plots
#
# ```julia
# using Plots
#
# plot(x, y)
#
# plot(f, xRange)
#
# plot(f, xMin, xMax)
#
# plot(x, [y1 y2])
# ```
# +
using Plots
Plots.gr(fmt=:png)
using Random
Random.seed!(2021)
# Data
x = 0:0.1:2pi
y1 = cos.(x)
y2 = sin.(x)
# Creating a plot in steps
plot(x, y1, color="blue", linewidth=3)
plot!(x, y2, color="red", line=:dash)
title!("Trigonometric functions")
xlabel!("angle")
ylabel!("sin(x) and cos(x)")
plot!(xlims=(0,2pi), ylims=(-2, 2), size=(600, 600))
# -
plot(x, y1, line=(:blue, 3))
plot!(x, y2, line=(:dash, :red))
# One set function to rule them all
plot!(title="Trigonometric functions",
xlabel="angle",
ylabel="sin(x) and cos(x)",
xlims=(0,2pi), ylims=(-2, 2), size=(600, 600))
# ### Plotting multiple series
#
# - One row = one observation
# - One column = one species
time = 30
walker1 = cumsum(randn(time))
walker2 = cumsum(randn(time))
walker3 = cumsum(randn(time))
walker4 = cumsum(randn(time))
walker5 = cumsum(randn(time))
plot(1:time, [walker1 walker2 walker3 walker4 walker5],
xlabel="time", ylabel="position",
label=["walker1" "walker2" "walker3" "walker4" "walker5"],
legend=:bottomleft)
# ### Parameteric plots
#
# Functions can be plotted directly.
#
# - `plot(f, xmin, xmax)`
# - `plot(f, range_of_x)`
f(x) = 5exp(-x^2)
g(x) = x^2
plot([f, g], -3, 3, label=["f" "g"], legend=:top)
plot(sin, t->sin(2t), 0, 2π, leg=false, fill=(0,:orange))
# ## 3D line plot
#
# Similar to 2D line plots.
#
# `plot(fx(t), fy(t), fz(t), tmin, tmax [, kwargs...])`
# 3D parametric plot
plot(cos, sin, t->sin(5t), 0, 2pi, legend=nothing)
# ### Line colors
#
# `plot(x, y, c=color)`
# +
using Plots, SpecialFunctions
Plots.gr(fmt=:png)
x = 0:0.2:10
y0 = besselj.(0,x)
y1 = besselj.(1,x)
y2 = besselj.(2,x)
y3 = besselj.(3,x)
y4 = besselj.(4,x)
y5 = besselj.(5,x)
y6 = besselj.(6,x)
colors = [:red :green :blue :cyan :magenta :yellow :black]
plot(x, [y0 y1 y2 y3 y4 y5 y6], c=colors)
# -
# ### Line styles
# +
using Plots
Plots.gr(fmt=:png)
@show Plots.supported_styles()
style = Plots.supported_styles()[2:end]
style = reshape(style, 1, length(style))
plot(x, [y0 y1 y2 y3 y4], line=(3, style))
# -
# ## Polar Plots
#
# `plot(θ, r, proj=:polar)`
# +
using Plots
Plots.gr(fmt=:png)
using Random
Random.seed!(2021)
r(θ) = 1 + cos(θ) * sin(θ)^2
θ = range(0, stop=2π, length=50)
plot(θ, r.(θ), proj=:polar, lims=(0,1.5))
# -
# One-liner
plot(θ -> 1 + cos(θ) * sin(θ)^2, 0, 2π, proj=:polar, lims=(0,1.5))
# ### Rose Plots
#
# `plot(..., proj=:polar, line=:steppre)`
n = 24
R = rand(n+1)
plot(0:2pi/n:2pi, R, proj=:polar, line=:steppre, lims=(0, 1), legend=nothing)
# ## Quiver Plots
#
# - `quiver(x1d, y1d, quiver=(vx1d, vy1d)`
# - `quiver(x2d, y2d, quiver=(x, y)->(u, v))`
# +
using Plots
Plots.gr(fmt=:png)
n = 7
f(x,y) = hypot(x, y) |> inv
x = repeat(-3:(2*3)/n:3, 1, n) |> vec
y = repeat(-3:(2*3)/n:3, 1, n)' |> vec
vx = f.(x,y) .* cos.(atan.(y,x)) |> vec
vy = f.(x,y) .* sin.(atan.(y,x)) |> vec
quiver(x, y, quiver=(vx, vy), aspect_ratio=:equal)
# -
g(x, y) = [f(x,y) * cos(atan(y,x)), f(x,y) * sin(atan(y,x))]
xx = [x for y in -3:(2*3)/n:3, x in -3:(2*3)/n:3]
yy = [y for y in -3:(2*3)/n:3, x in -3:(2*3)/n:3]
quiver(xx, yy, quiver=g, aspect_ratio=:equal, color=:black)
# ## Scatter Plots
#
# 2D Scatter Plots: `scatter(xpos, ypos)`
# +
using Plots
Plots.gr(fmt=:png)
using Random
Random.seed!(2021)
n = 50
x = rand(n)
y = rand(n)
ms = rand(50) * 30
scatter(x, y, markersize=ms)
# -
# 3D Scatter Plots: `scatter(xpos, ypos, zpos)`
# +
z = rand(n)
scatter(x, y, z, markersize=ms)
# -
# ## Stairstep plot
#
# `plot(..., line=:steppre)`
# +
using Plots
Plots.gr(fmt=:png)
plot(sin.(0:0.3:2pi), line=:steppre, label="Steps")
# -
# ## Stem plot
#
# A.k.a lolipop plot.
#
# `plot(..., line=:stem)`
# +
using Plots
Plots.gr(fmt=:png)
plot(sin.(0:0.3:2pi), line=:stem, marker=:star, markersize=20, ylims=(-1.1, 1.1), label="Stars")
# -
# ## Subplots
#
# - `plot(p1, p2, p3, ...)`
# - `plot(..., layout=(nrow, ncol))`
# - `plot(..., layout=@layout [...])`
#
# [Source](https://docs.juliaplots.org/latest/layouts/)
# +
using Plots
Plots.gr(fmt=:png)
using Random
Random.seed!(2021)
data = rand(100, 4)
# create a 2x2 grid, and map each of the 4 series to one of the subplots
plot(data, layout = 4)
# -
# create a 4x1 grid, and map each of the 4 series to one of the subplots
plot(data, layout = (4, 1))
# More complex grid layouts can be created with the grid(...) constructor:
plot(data, layout = grid(4, 1, heights=[0.1 ,0.4, 0.4, 0.1]))
# Adding titles and labels
plot(data, layout = 4, label=["a" "b" "c" "d"], title=["1" "2" "3" "4"])
l = @layout [
a{0.3w} [grid(3,3)
b{0.2h} ]
]
plot(
rand(10, 11),
layout = l, legend = false, seriestype = [:bar :scatter :path],
title = ["($i)" for j in 1:1, i in 1:11], titleloc = :right, titlefont = font(8)
)
# Use _ to ignore a spot in the layout
plot((plot() for i in 1:7)..., layout=@layout([_ ° _; ° ° °; ° ° °]))
# ### Build subplot components indivisually
p1 = plot(sin, 0, 2pi, xlabel="x1")
p2 = plot(cos, 0, 2pi, xlabel="x2")
p3 = histogram(randn(1000), xlabel="x3")
p4 = plot(x->exp(-x^2), -3, 3, xlabel="x4")
plot(p1, p2, p3, p4)
# ## Surface plots
#
# - `surface(x, y, z)`
# - `surface(x, y, (x,y)->z)`
# - `plot(x, y, z, linetype=:surface)`
# - `plot(x, y, z, linetype=:wireframe)`
# +
using Plots
Plots.gr(fmt=:png)
f(x,y) = x^2 + y^2
x = y = -10:10
surface(x, y, f)
# -
plot(x, y, f, linetype=:surface)
plot(x, y, f, linetype=:wireframe)
# ## Two Y Axis
#
# `plot!(twinx())`
# + tags=[]
using Plots, Random, Measures
Plots.gr(fmt=:png)
Random.seed!(2021)
plot(randn(100), ylabel="y1", leg=:topright)
plot!(twinx(), randn(100)*10,
c=:red,
ylabel="y2",
leg=:bottomright,
size=(600, 400))
plot!(right_margin=15mm)
| docs/plotsjl-gallery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/britjet/Linear-Algebra_ChE_2nd-sem-2021-2022/blob/main/Assignment6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bAgYHtsaS96Q"
# #Laboratory 6: Matrix Operation
# + [markdown] id="SO0dH0reTNaf"
# Objectives
#
# At the end of this activity you will be able to:
#
# 1. Understand the fundamental matrix operations.
# 2. Solve intermediate equations using the procedures.
# 3. In engineering solutions, use matrix algebra.
# + [markdown] id="y0PmHudoViEP"
# ##Discussion
# + id="49YKsSPRS3RT"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] id="IIdckyjjVyB_"
# ##Transposition
# + [markdown] id="qRdsleXmfG8p"
# Transposition is a fundamental operation in matrix algebra and is one of its most critical applications. The transposition of a matrix is accomplished by flipping its members' values over the matrix's diagonals. The rows and columns of the original matrix will be swapped due to this. As a result, the symbol $A^T$ indicates the transpose of matrix $A$. As an illustration, consider the following:
# + [markdown] id="wgmX86vgf1HL"
# $$A = \begin{bmatrix} 2 & 5 & 8\\5 & -6 &1 \\ 10 & -8 & 3\end{bmatrix} $$
# + [markdown] id="5eme-jVWgEIJ"
# $$ A^T = \begin{bmatrix} 2 & 3 & 8\\5 & -1 &-9 \\ 7 & 10 & 5\end{bmatrix}$$
# + [markdown] id="weBLdl9HgXtg"
# This may now be accomplished programmatically through the use of the np.transpose() function or the T method.
# + id="mp3cQxQegZZq" colab={"base_uri": "https://localhost:8080/"} outputId="f3da1ba7-95bc-4643-8e97-b9a6b013a0d3"
A = np.array([
[3 ,9, 7],
[4, -5, 0],
[5, -2, 5]
])
A
# + id="oAv3cKrkg6Bn" colab={"base_uri": "https://localhost:8080/"} outputId="7cbb446f-cda8-493d-ecb6-f09fe8080dcb"
AT1 = np.transpose(A)
AT1
# + id="9XzEnuYYg8A3" colab={"base_uri": "https://localhost:8080/"} outputId="fcedc2e1-5346-4d78-92ec-cd53479b3807"
AT2 = A.T
AT2
# + id="wPbg8SKpg-Rn" colab={"base_uri": "https://localhost:8080/"} outputId="1786bb89-6a45-49f5-b1c5-e251560a9814"
np.array_equiv(AT1, AT2)
# + id="_nzdkeYYhG7P" colab={"base_uri": "https://localhost:8080/"} outputId="11986a3a-9525-4fc7-b75a-1ebe8f1d6749"
B = np.array([
[2,4,6,8],
[3,6,9,12],
])
B.shape
# + id="__fwJlh0hbPn" colab={"base_uri": "https://localhost:8080/"} outputId="7086e79e-4794-4659-9d11-3bebdc2973af"
np.transpose(B).shape
# + id="CEaLsDdfhcAX" colab={"base_uri": "https://localhost:8080/"} outputId="8486be13-981f-4b5c-c6fa-7a2aa879a314"
B.T.shape
# + [markdown] id="lo1vfuj-hg3W"
# #### Create your own matrix (you can experiment with non-squares) and use it to test transposition.
# + id="BzgtUSJ0hpCh" colab={"base_uri": "https://localhost:8080/"} outputId="ca66cbb2-7a21-483a-cfea-ccf00ea6d299"
## Try out your code here.
Z=np.array([
[5,7,9],
[4,8,10]
])
Z.shape
# + id="lAtNi3gthwqH" colab={"base_uri": "https://localhost:8080/"} outputId="5d8df8a8-a39e-472b-ee6d-03a28ae3c408"
np.transpose(Z).shape
# + id="JHaGKh3zhx6k" colab={"base_uri": "https://localhost:8080/"} outputId="4be76bd7-d7fd-45aa-9251-279d770ba33a"
Z.T.shape
# + id="agEcN_mRhz3i" colab={"base_uri": "https://localhost:8080/"} outputId="4cf1b2b0-94f6-4614-f9eb-5d2bfd4dbeb7"
ZT = Z.T
ZT
# + [markdown] id="NcQgqT4_h2Kk"
# ## Dot Product / Inner Product
# + [markdown] id="Te00XjaFh5DZ"
# Assuming you are familiar with the dot product from a previous laboratory activity, we will attempt to do the same operation using matrices. The Dot Product/Inner Product will obtain the sum of products of the vectors by row-column pairs by performing a matrix dot product operation. As an example, consider the following two matrices: $X$ and $Y$.
#
# $$X = \begin{bmatrix}x_{(1,1)}&x_{(3,3)}\\ x_{(5,8)}&x_{(5,5)}\end{bmatrix}, Y = \begin{bmatrix}y_{(1,1)}&y_{(3,3)}\\ y_{(5,8)}&y_{(5,5)}\end{bmatrix}$$
#
# The dot product will then be computed as:
# $$X \cdot Y= \begin{bmatrix} x_{(1,1)}*y_{(1,1)} + x_{(3,3)}*y_{(5,8)} & x_{(1,1)}*y_{(3,3)} + x_{(3,3)}*y_{(5,5)} \\ x_{(5,8)}*y_{(1,1)} + x_{(5,5)}*y_{(5,8)} & x_{(5,8)}*y_{(3,3)} + x_{(5,5)}*y_{(5,5)}
# \end{bmatrix}$$
#
# So if we assign values to $X$ and $Y$:
# $$X = \begin{bmatrix}2&3\\ 1&2\end{bmatrix}, Y = \begin{bmatrix}-2&0\\ 4&4\end{bmatrix}$$
# + [markdown] id="lhbL0MuqlV1d"
# $$X \cdot Y= \begin{bmatrix} 2*-2 + 3*4 & 2*0 + 4*4 \\ 0*-2 + 2*3 & 0*0 + 2*4 \end{bmatrix} = \begin{bmatrix} 8 & 16 \\ 6 & 8 \end{bmatrix}$$
# This could be achieved programmatically using `np.dot()`, `np.matmul()` or the `@` operator.
# + id="_9Fa5DZ3mbOw"
X = np.array([
[0,3],
[5,8]
])
Y = np.array([
[-5,8],
[0,7]
])
# + id="ZaGCnFhlmpyV" colab={"base_uri": "https://localhost:8080/"} outputId="7d2fe1ee-cf59-420d-d388-02a8d0787725"
np.array_equiv(X, Y)
# + id="psvECCkmmqYv" colab={"base_uri": "https://localhost:8080/"} outputId="c773a955-67fd-43ff-e6ce-944b7e6a0e35"
np.dot(X,Y)
# + id="oErrgKVRmsQu" colab={"base_uri": "https://localhost:8080/"} outputId="5a1ae1ab-44bf-406a-b4a8-6829b2827fb3"
X.dot(Y)
# + id="bdAGxGZWmtx4" colab={"base_uri": "https://localhost:8080/"} outputId="df293d89-d534-4952-b02b-9036495dadf0"
X @ Y
# + id="mSPySxNPmvIv" colab={"base_uri": "https://localhost:8080/"} outputId="a568d8a7-d188-45b5-d731-53fcdf02b18d"
np.matmul(X,Y)
# + id="QdGycMlVmwdo"
D = np.array([
[5,8,12],
[2,8,3],
[5,9,1]
])
E = np.array([
[-5,0,9],
[5,8,6],
[7,1,3]
])
# + id="t-A0PzOdnBIx" colab={"base_uri": "https://localhost:8080/"} outputId="ffb8bf0b-5312-4ab0-da08-8c140f2adb34"
D @ E
# + id="Me9khwZznBrT" colab={"base_uri": "https://localhost:8080/"} outputId="e46ce0bb-c988-47c5-ea8e-7c8845a9ca5c"
D.dot(E)
# + id="gVzrQ5qCnD1s" colab={"base_uri": "https://localhost:8080/"} outputId="0320fb74-4ffa-46fe-e130-c14e709d757b"
np.matmul(D, E)
# + id="F3qu6ArmnFIi" colab={"base_uri": "https://localhost:8080/"} outputId="9fae2529-754c-4537-de3c-3272a2da238c"
np.dot(D, E)
# + [markdown] id="3h2um01nnPdt"
# When comparing matrix dot products to vector dot products, there are certain extra criteria to follow. Due to the fact that vector dot products were just in one dimension, there are less limitations. Because we are now working with Rank 2 vectors, we must take into consideration the following rules:
#
# ###In order for the two matrices in question to be equivalent, the inner dimensions of each matrix must be the same.
#
# Consider the following scenario: you have a matrix $A$ with the structure of $(a,b)$, where $a$ and $b$ are any integers. Suppose we want to conduct a dot product between $A$ and another matrix $B$, then matrix $B$ should have the shape $(b,c)$, where $b$ and $c$ may be any integers and $A$ can be any other integer. As an example, consider the following matrices:
#
# $$A = \begin{bmatrix}3&1\\8&-5\\1&8\end{bmatrix}, B = \begin{bmatrix}1&4\\8&6\\-2&-3\end{bmatrix}, C = \begin{bmatrix}2&9&5\\3&4&5\end{bmatrix}$$
#
# $A$ has the shape $(,)$, $B$ has the shape $(,)$, and $C$ has a shape of $(,)$ in this example. As a result, the only matrix pairings that are qualified to conduct dot product are the matrices $A cdot C$ and $B cdot C$, respectively.
# + id="gMHApIMQoeu4" colab={"base_uri": "https://localhost:8080/"} outputId="964f4cf5-0cee-40ba-aeec-812cba4a4a12"
A = np.array([
[3, 1],
[8, -5],
[1, 8]
])
B = np.array([
[1,4],
[8,6],
[-2,-3]
])
C = np.array([
[2,9,5],
[3,4,5]
])
print(A.shape)
print(B.shape)
print(C.shape)
# + id="X2WHumrNptPq" colab={"base_uri": "https://localhost:8080/"} outputId="3b3b441f-89e5-440f-a878-ef601fcbdd20"
A @ C
# + id="8HtYA8gapvc8" colab={"base_uri": "https://localhost:8080/"} outputId="cc2dade6-fc42-4fa1-ba13-030df46ed2b7"
B @ C
# + [markdown] id="ebmIkQyCp7cr"
# If you look closely, you will note that the geometry of the dot product has changed and that it is no longer the same as any of the matrices that we utilized. In reality, the form of a dot product is generated by the shapes of the matrices that were utilized. As an example, consider the shapes of the matrices $A$ and $B$: $(a,b)$ and $(b,c)$, respectively. In this case, the shape of $A\cdot B$ should be $(a,c)$.
# + id="gkBEeBL3p9Vy" colab={"base_uri": "https://localhost:8080/"} outputId="2d7089cd-af2e-47be-ff3d-6d2cb25e729c"
A @ B.T
# + id="7_nCB11yqGem" colab={"base_uri": "https://localhost:8080/"} outputId="0b2e6e17-5f02-42b0-b491-4af7efa0305a"
X = np.array([
[2,4,5,10]
])
Y = np.array([
[6,1,8,-5]
])
print(X.shape)
print(Y.shape)
# + id="Y8_uYdOAqOHr" colab={"base_uri": "https://localhost:8080/"} outputId="0f0f25ea-0a5e-49a7-d283-3f914fa02b4c"
Y.T @ X
# + id="MHtVEcb6qPk_" colab={"base_uri": "https://localhost:8080/"} outputId="e4b0181b-81b9-4dd3-e082-46f1779c6a54"
X @ Y.T
# + [markdown] id="KrJqJjtuqScQ"
# Moreover, you can see that when you attempt to multiply A and B, the program produces the error `ValueError` due to a mismatch in the matrix structure.
# + [markdown] id="ibebZ6PpqkZL"
# ### Rule 2: Dot Product has special properties
#
# Due to the fact that dot products are common in matrix algebra, it follows that they have numerous distinct qualities that should be taken into consideration while formulating solutions:
#
# 1. $A \cdot B \neq B \cdot A$
# 2. $A \cdot (B \cdot C) = (A \cdot B) \cdot C$
# 3. $A\cdot(B+C) = A\cdot B + A\cdot C$
# 4. $(B+C)\cdot A = B\cdot A + C\cdot A$
# 5. $A\cdot I = A$
# 6. $A\cdot \emptyset = \emptyset$
# + id="fDUH52RVrElX"
A = np.array([
[2,5,5],
[4,7,6],
[7,4,1]
])
B = np.array([
[3,1,9],
[4,3,6],
[4,8,8]
])
C = np.array([
[9,1,4],
[0,5,8],
[7,5,1]
])
# + id="iQ-3autxrYCj" colab={"base_uri": "https://localhost:8080/"} outputId="3c94651c-b8c9-466d-d1ac-367678316df3"
np.eye(3)
# + id="LW4vlnm5raGZ" colab={"base_uri": "https://localhost:8080/"} outputId="9d5c7755-4db3-4f3f-c58a-3f7872264ed9"
A.dot(np.eye(3))
# + id="xOnbV7T_rbzi" colab={"base_uri": "https://localhost:8080/"} outputId="f83886ed-cc98-4460-a5a2-6f07031b986d"
np.array_equal(A@B, B@A)
# + id="z6CyA-qsrdf5" colab={"base_uri": "https://localhost:8080/"} outputId="d4139187-e149-4768-8daa-0816103f780c"
E = A @ (B @ C)
E
# + id="4mAlAhVFrfCT" colab={"base_uri": "https://localhost:8080/"} outputId="df400908-fd7e-4b01-f286-a66f552e64dd"
F = (A @ B) @ C
F
# + id="l8nVhLoUriEq" colab={"base_uri": "https://localhost:8080/"} outputId="140fb251-ad21-4e7e-c759-8a60f0274441"
np.array_equal(E, X)
# + id="O6u5zlpxrkY0" colab={"base_uri": "https://localhost:8080/"} outputId="a7f2b36a-7bc3-42a0-e20a-fcc5cac19c2c"
np.array_equiv(E, F)
# + id="U6LvI3zlrmkv" colab={"base_uri": "https://localhost:8080/", "height": 304} outputId="1e592494-ef4d-4755-dcc5-cdb1d913f8ac"
np.eye(A)
# + id="hzWSY6KSrrsg" colab={"base_uri": "https://localhost:8080/"} outputId="df43474b-9c0b-4b96-d9e8-7193c46143a1"
A @ E
# + id="SpAxSwHQrv_8" colab={"base_uri": "https://localhost:8080/"} outputId="471d7670-2a2a-45e9-d183-c9c0f25118ba"
z_mat = np.zeros(A.shape)
z_mat
# + id="BSdIv2EMrxc4" colab={"base_uri": "https://localhost:8080/"} outputId="8a13e9ef-4298-49a3-ec11-3a4b22c7db1a"
a_dot_z = A.dot(np.zeros(A.shape))
a_dot_z
# + id="HIjwuNfkr0EG" colab={"base_uri": "https://localhost:8080/"} outputId="5daf7f63-6974-4e68-ff84-e35ffb50f1d8"
np.array_equal(a_dot_z,z_mat)
# + id="q4liwJu_r1Ti" colab={"base_uri": "https://localhost:8080/"} outputId="22837f08-a41d-47b5-f9c0-3ba270aa6bd7"
null_mat = np.empty(A.shape, dtype=float)
null = np.array(null_mat,dtype=float)
print(null)
np.allclose(a_dot_z,null)
# + [markdown] id="5nSu-J7-r2d0"
# ##Determinant
# + [markdown] id="VxjVlIDor5zE"
# A determinant is a scalar value that may be obtained from a square matrix in two dimensions. Determinants are important values in matrix algebra because they are fundamental and important values. Although it will not be immediately apparent in this laboratory how it might be used in practice, it will be extensively utilized in subsequent lectures.
#
# The determinant of some matrix $A$ is denoted as $det(A)$ or $|A|$. So let's say $A$ is represented as:
# $$A = \begin{bmatrix}a_{(0,0)}&a_{(0,1)}\\a_{(1,0)}&a_{(1,1)}\end{bmatrix}$$
# We can compute for the determinant as:
# $$|A| = a_{(0,0)}*a_{(1,1)} - a_{(1,0)}*a_{(0,1)}$$
# So if we have $A$ as:
# $$A = \begin{bmatrix}2&5\\7&9\end{bmatrix}, |A| = $$
# However, you may question what happens to square matrices that are not of the form $(2,2)$. We may address this issue in a variety of ways, including utilizing co-factor expansion and the minors technique, among others. In the laboratory, we can learn how to do this in the lecture, but we can also programmatically compute the difficult calculation of high-dimensional matrices by using Python. This may be accomplished by the use of the function `np.linalg.det()`.
#
# + id="u8d_as6JsmiB" colab={"base_uri": "https://localhost:8080/"} outputId="c74f16a0-7973-4f98-89db-d57fd4d5465f"
A = np.array([
[2,5],
[7,9]
])
np.linalg.det(A)
# + id="9eRxJjTSstzS" colab={"base_uri": "https://localhost:8080/"} outputId="0074f9a0-eb88-40d3-b524-b2a787a92009"
B = np.array([
[2, 1, 3],
[3, -6 ,-7],
[0, -1, 4]
])
np.linalg.det(B)
# + id="PA10feTgs8ly" colab={"base_uri": "https://localhost:8080/"} outputId="4d3ae12c-9879-4a71-f1ae-0e9a7d90d64d"
## Now other mathematics classes would require you to solve this by hand,
## and that is great for practicing your memorization and coordination skills
## but in this class we aim for simplicity and speed so we'll use programming
## but it's completely fine if you want to try to solve this one by hand.
B = np.array([
[3,9,7,4],
[3,6,8,3],
[3,9,8,2],
[7,5,5,2]
])
np.linalg.det(B)
# + [markdown] id="FSY6WTfOtM6Y"
# ##Inverse
# + [markdown] id="-YVM3Z2atQcq"
# The inverse of a matrix is yet another important operation in matrix algebra that should not be overlooked. Determining the inverse of a matrix allows us to assess whether or not the matrix is solvable and has the characteristics of a system of linear equations — we'll go into more detail about this in the nect module. Another use of the inverse matrix is in the solution of the issue of divisibility between matrices between two variables. Although there is an element-by-element division method, there is no such method for splitting the whole idea of matrices. It is possible that the idea of "dividing" matrices is the same as that of "inverting" matrices in the case of inverse matrices.
#
# After that, we must go through various processes to get the inverse of the matrix we have created. So, let us suppose we have the following matrix $M$:
# $$M = \begin{bmatrix}1&7\\-3&5\end{bmatrix}$$
# First, we need to get the determinant of $M$.
# $$|M| = (1)(5)-(-3)(7) = 26$$
# Next, we need to reform the matrix into the inverse form:
# $$M^{-1} = \frac{1}{|M|} \begin{bmatrix} m_{(1,1)} & -m_{(0,1)} \\ -m_{(1,0)} & m_{(0,0)}\end{bmatrix}$$
# So that will be:
# $$M^{-1} = \frac{1}{26} \begin{bmatrix} 5 & -7 \\ 3 & 1\end{bmatrix} = \begin{bmatrix} \frac{5}{26} & \frac{-7}{26} \\ \frac{3}{26} & \frac{1}{26}\end{bmatrix}$$
# It is possible that you will need to apply co-factors, minors, adjugates, and other reduction techniques for higher-dimension matrices. We can use a computer program to address this problem.`np.linalg.inv()`.
# + id="fNPU9PEIt5ZS" colab={"base_uri": "https://localhost:8080/"} outputId="d5aea876-7adf-4579-e380-3b018d182a30"
M = np.array([
[1,7],
[-3,5 ]
])
np.array(M @ np.linalg.inv(M), dtype=int)
# + id="7dJ2cY9At_fW" colab={"base_uri": "https://localhost:8080/"} outputId="5377bf2a-34e7-483a-bb2d-c9e8015f0777"
P = np.array([
[2, 5, 8],
[3, 6, -9],
[1, 4, 7]
])
Q = np.linalg.inv(P)
Q
# + id="7yIYqTVkuHtR" colab={"base_uri": "https://localhost:8080/"} outputId="71a48ad9-f3d9-4e36-9c7c-8c00d91a0f8f"
P @ Q
# + id="623JYnhOuKGx" colab={"base_uri": "https://localhost:8080/"} outputId="86967526-4266-4a97-a3b1-c511996cd038"
## And now let's test your skills in solving a matrix with high dimensions:
N = np.array([
[18,5,23,1,0,33,5],
[0,45,0,11,2,4,2],
[5,9,20,0,0,0,3],
[1,6,4,4,8,43,1],
[8,6,8,7,1,6,1],
[-5,15,2,0,0,6,-30],
[-2,-5,1,2,1,20,12],
])
N_inv = np.linalg.inv(N)
np.array(N @ N_inv,dtype=int)
# + [markdown] id="dp97iZ4wuOQ_"
# To validate the wether if the matric that you have solved is really the inverse, we follow this dot product property for a matrix $M$:
# $$M\cdot M^{-1} = I$$
# + id="aOesMR0kuMY7" colab={"base_uri": "https://localhost:8080/"} outputId="4f22ee02-f96e-4911-8d29-78b11414ea67"
squad = np.array([
[1.5, 1.0, 2.5],
[0.8, 0.1, 1.9],
[0.1, 0.5, 3.0]
])
weights = np.array([
[1.2, 0.3, 0.9]
])
p_grade = squad @ weights.T
p_grade
# + [markdown] id="I-G0bTNNuiU8"
# ##Activity
# + [markdown] id="b1D0cC3eunvo"
# ###Task 1
# + id="TVxSI9KuunPH"
N = np.array([
[8, 6, 1, 5],
[6, 7, 1, 0],
[9, 0, 8, 8],
[-8, 9, 2, 3]
])
T = np.array([
[6, 6, 1, 2],
[6, 5, 9, 2],
[4, 9, 8, -4],
[-1, 9, 7, 2]
])
G = np.array([
[5, 1, -5, 2],
[0, 0, 9, 5],
[-4, -9, 3, 8],
[-7, 5, 1, 0]
])
# + [markdown] id="b1mYanwObhPx"
# $A \cdot B \neq B \cdot A$
# + colab={"base_uri": "https://localhost:8080/"} id="XlYcADCSbkMN" outputId="6d6f5a39-fc4c-4a0f-9f37-7c63bfe6bb9b"
T = (N@T) !=(T@N)
T
# + [markdown] id="QMnhTTRHaCw-"
# $A \cdot (B \cdot C) = (A \cdot B) \cdot C$
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="3orTXmw1aDnM" outputId="7317c726-8202-4e2a-a03f-34d3037a7c00"
A = N @ (T@G)
A
# + colab={"base_uri": "https://localhost:8080/"} id="TbqSdbGocGtA" outputId="02dd20aa-c44a-4b19-9467-a53def0708d0"
B = (N @ T) @ G
B
# + [markdown] id="JwmrqVQoabTN"
# $A\cdot(B+C) = A\cdot B + A\cdot C$:
#
# + colab={"base_uri": "https://localhost:8080/"} id="N2Ev8NeVab1z" outputId="aab3317d-7861-4e25-b24b-e42418f11172"
C = N @ (T + G)
C
# + colab={"base_uri": "https://localhost:8080/"} id="wNELDRHsdAQl" outputId="8c32bc6d-3e19-499f-a5e0-83a8692b8a2b"
D = (N@T) + (N@G)
D
# + [markdown] id="qWDxxfDqasG4"
# $(B+C)\cdot A = B\cdot A + C\cdot A$
# + colab={"base_uri": "https://localhost:8080/"} id="BMjuHWJsdEbo" outputId="f6e53b75-22f8-4762-d3ab-04a7da69f9a2"
E = (T+G) @ N
E
# + colab={"base_uri": "https://localhost:8080/"} id="DZbOJDq6eAkN" outputId="c872fd69-0555-4272-b2ca-bbc1c72c14b1"
F = (T@N) + (G@N)
F
# + [markdown] id="3EMP4G4ebGck"
# $A\cdot I = A$
# + colab={"base_uri": "https://localhost:8080/"} id="MbbKflvPd_Q3" outputId="d846b0d3-327d-40ef-9636-42783cfb0238"
np.eye(4)
# + colab={"base_uri": "https://localhost:8080/"} id="nr0X-s7YeBId" outputId="c4857fcc-c57d-4e56-ef12-3cb773655b37"
N.dot(np.eye(4))
# + [markdown] id="KUVMQv9MbJRE"
# $A\cdot \emptyset = \emptyset$
# + colab={"base_uri": "https://localhost:8080/"} id="bcEmSHqWd-Qx" outputId="43e6d86b-e43e-4670-a6f4-58666ee6dcfd"
z_mat = np.zeros(G.shape)
z_mat
# + [markdown] id="sYNIY7k5iRKg"
# ##Conclusion
# + [markdown] id="2ozDI9zRiTVC"
# The students demonstrated an understanding of and ability to use a variety of matrix operations, including transpository, dot product, determinant, and inverse. Students demonstrated their ability to put their learning into practice by successfully completing the task, which required them to create and execute the six multiplications characteristics.
| Assignment6.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: JuliaFlux 1.6.3
# language: julia
# name: juliaflux-1.6
# ---
using LinearAlgebra
using Random
using CairoMakie
using Distances
CairoMakie.activate!(type = "svg")
using StatsBase
N = 8
μ = rand(N,2)
x,y = μ[:,1], μ[:,2]
scatter(μ)
# +
function basecase(xsorted)
end
# +
def dist(p1, p2):
return math.sqrt(((p2[1]-p1[1])**2)+((p2[0]-p1[0])**2))
def closest_brute_force(points):
min_dist = float("inf")
p1 = None
p2 = None
for i in range(len(points)):
for j in range(i+1, len(points)):
d = dist(points[i], points[j])
if d < min_dist:
min_dist = d
p1 = points[i]
p2 = points[j]
return p1, p2, min_dist
def rec(xsorted, ysorted):
n = len(xsorted)
if n <= 3:
return closest_brute_force(xsorted)
else:
midpoint = xsorted[n//2]
xsorted_left = xsorted[:n//2]
xsorted_right = xsorted[n//2:]
ysorted_left = []
ysorted_right = []
for point in ysorted:
ysorted_left.append(point) if (point[0] <= midpoint[0]) else ysorted_right.append(point)
(p1_left, p2_left, delta_left) = rec(xsorted_left, ysorted_left)
(p1_right, p2_right, delta_right) = rec(xsorted_right, ysorted_right)
(p1, p2, delta) = (p1_left, p2_left, delta_left) if (delta_left < delta_right) else (p1_right, p2_right, delta_right)
in_band = [point for point in ysorted if midpoint[0]-delta < point[0] < midpoint[0]+delta]
for i in range(len(in_band)):
for j in range(i+1, min(i+7, len(in_band))):
d = dist(in_band[i], in_band[j])
if d < delta:
print(in_band[i], in_band[j])
(p1, p2, delta) = (in_band[i], in_band[j], d)
return p1, p2, delta
def closest(points):
xsorted = sorted(points, key=lambda point: point[0])
ysorted = sorted(points, key=lambda point: point[1])
return rec(xsorted, ysorted)
| Algorithms/ClosestPair.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/LeightonEstrada/2021_BD-ML_Python_Tutorials/blob/main/Tut08/08%20M3%20Tutoria18-06-2021%20Funciones%20Activacion.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="6WrnS2_uaMzy"
# # Diplomatura BD&ML UTEC 2021
#
# # Tutoría de Reforzamiento Sábado 18-06-2021
#
# ## Tutor: <NAME>
#
# ## Funciones de Activación
# + [markdown] id="dv5H_OoTC0Ht"
# ### Importar libreria
# + id="FY0tKo9rCzsf"
import math
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] id="L53lAVGy8-Dm"
# ### Función Binary
# + id="3uqWzdj89BpW"
def binary_active_function(x):
return 0 if x < 0 else 1
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="z26uX_7PF6Ag" outputId="ead960f1-27ba-4ffc-d030-21780df1215f"
x = np.linspace(-10,10,100)
y= [binary_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="ImrdrHC5RxhK"
# ### Función Linear
# + id="0Wa2k82lR2TQ"
def linear_active_function(a, x):
return a*x
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="1O61eL1tR2d3" outputId="d8b1bd44-847a-4825-8a82-d61b7454c2f1"
x = np.linspace(-10,10,100)
y= [linear_active_function(1, ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="5cu-v8LsSgFg"
# ### Funcion Sigmoid
# + id="E2v5F-EvSfcy"
def sigmoid_active_function(x):
return 1./(1+np.exp(-x))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="AX9PsVlFSl1Y" outputId="7ef4ed51-c880-498c-b129-29af7becb716"
x = np.linspace(-10,10,100)
y= [sigmoid_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="0CgzzdEVS1mQ"
# ### Función Tanh
# + id="tdIHeFKHS1Ex"
def tanh_active_function(x):
return 2*sigmoid_active_function(2*x)-1
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="JiLDTMadS7Po" outputId="cdff6244-2b06-48f0-9dc9-8667d2bd8723"
x = np.linspace(-10,10,100)
y= [tanh_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="hmp2q-8KS_FQ"
# ### Función ReLU
# + id="vREOVh82S-t4"
def relu_active_function(x):
return np.array([0, x]).max()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="DXpPj3jPTDWa" outputId="5a70d4a7-b25d-4c62-aab4-79903e7acbad"
x = np.linspace(-10,10,100)
y= [relu_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="R8wNuluyTSdx"
# ### Función Leaky ReLU
# + id="P7IIcTdHTL2Z"
def leaky_relu_active_function(x):
return 0.01*x if x < 0 else x
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="t4UKT57gTV9h" outputId="fe0af860-e90a-4f27-fffd-09b7b98f99b6"
x = np.linspace(-10,10,100)
y= [leaky_relu_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="LS5zDEQGTo1p"
# ### Función Parametric ReLU
# + id="i_TNjWQeTr4J"
def parametric_relu_active_function(a, x):
return a*x if x < 0 else x
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="4KFB4a1rTsAS" outputId="405ba5db-85e2-42f6-d004-07675cd905bf"
x = np.linspace(-10,10,100)
y= [parametric_relu_active_function(1,ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="TYXV4JWvTzQR"
# ### Función Exponential Linear Unit (eLU)
# + id="XJm9s-ZBT2Jy"
def elu_active_function(a, x):
return a*(np.exp(x)-1) if x < 0 else x
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="paSFuWeqT2Qh" outputId="9c528900-3017-48f8-83bc-b9869b82d7cf"
x = np.linspace(-10,10,100)
y= [elu_active_function(1,ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="Z7MWntsXVH8y"
# ### Función ReLU-6
# + id="OFw0rjU2VHhr"
def relu_6_active_function(x):
return np.array([0, x]).max() if x<6 else 6
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="Oe2lux7HVWER" outputId="b10ec7e1-8529-4844-c8ab-f<PASSWORD>"
x = np.linspace(-10,10,100)
y= [relu_6_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="_kv8oHKRVblR"
# ### Función Softplus
# + id="mMwnaJNBVfq6"
def softplus_active_function(x):
return math.log(1+np.exp(x))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="1dKsOdYUVfzB" outputId="59fd29ba-a8a8-4927-c74f-cc88b4567529"
x = np.linspace(-10,10,100)
y= [softplus_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="VeI_Dcf0Vvsk"
# ### Función Softsign
# + id="xEXhS2NwVzRK"
def softsign_active_function(x):
return x / (1 + abs(x) )
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="_h7vqrV_VzYx" outputId="2b6d6382-2c02-4320-df92-28ace2d9c1e8"
x = np.linspace(-10,10,100)
y= [softsign_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="KGcCp8PfV5Xi"
# ### Función Softmax
# + id="LcjDnkNMV-lq"
def softmax_active_function(x):
return np.exp(x)/np.sum(np.exp(x))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="l7EsBMa9V-pB" outputId="83840742-339a-4f69-a1e8-f49609551a40"
x = np.linspace(-10,10,100)
y= [softmax_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="ng1SJRy_WHtK"
# ### Funcion Swish
# + id="jBmS4E9FWHJq"
def swish_active_function(x):
return x/(1+np.exp(-x))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="aGobKhQXWHMy" outputId="d0b161b9-72f7-4a29-ccf0-a2453b39bee1"
x = np.linspace(-10,10,100)
y= [swish_active_function(ii) for ii in x]
plt.plot(x,y)
plt.show()
# + [markdown] id="TLdtRJMDYn-v"
# -- Fin de Tutoria --
| Tut08/08 M3 Tutoria18-06-2021 Funciones Activacion.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training a text classifier model on a standalone dataset with fastai
# - This notebook ingests the Kaggle Covid tweets dataset (https://www.kaggle.com/datatattle/covid-19-nlp-text-classification)
# - This notebook assumes you have already run text_standalone_dataset_lm.ipynb notebook to create a language model
# - The encoder from the language model is used to create the text classifier
#hide
# !pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
#hide
from fastbook import *
from fastai.text.all import *
# ensure that value of modifier matches the value of modifier in text_standalone_dataset_lm notebook
modifier = 'standalone_mar20'
# # Ingest the dataset
# - define the source of the dataset
# - create a dataframe for the training dataset
# %%time
# create dataloaders object
path = URLs.path('covid_tweets')
path.ls()
# read the training CSV into a dataframe - note that the encoding parameter is needed to avoid a decode error
df_train = pd.read_csv(path/'train/Corona_NLP_train.csv',encoding = "ISO-8859-1")
# # Define the text classifier
# %%time
# create TextDataLoaders object
dls = TextDataLoaders.from_df(df_train, path=path, text_col='OriginalTweet',label_col='Sentiment')
dls.show_batch(max_n=3)
dls.path
# save the current path
keep_path = path
print("keep_path is: ",str(keep_path))
# %%time
# define a text_classifier_learner object
learn_clas = text_classifier_learner(dls, AWD_LSTM,
metrics=accuracy).to_fp16()
# # Fine-tune the text classifier
# Use the encoder created as part of training the language model to fine tune the text classifier
# Path('/storage/data/imdb')
learn_clas.path
# %%time
# set the path to the location of the encoder
learn_clas.path = Path('/notebooks/temp')
# load the encoder that was saved when the language model was trained
learn_clas = learn_clas.load_encoder('ft_standalone'+modifier)
learn_clas.path
# set the path back to the original path
learn_clas.path = keep_path
# ch 10 style Path('/storage/data/imdb')
learn_clas.path
# %%time
# fine tune the model
learn_clas.fit_one_cycle(1, 2e-2)
x, y = first(dls.train)
x.shape, y.shape, len(dls.train)
learn_clas.summary()
# # Exercise the text classifier
# Apply the fine-tuned text classifier on some text samples.
preds = learn_clas.predict("the government's approach to the pandemic has been a complete disaster")
preds
preds = learn_clas.predict("the new vaccines hold the promise of a quick return to economic growth")
preds
preds = learn_clas.predict("this flu is about what we would expect in a normal winter")
preds
preds = learn_clas.predict("the health ministry needs to pay closer attention to the vaccine rollout")
preds
# save the classifier model
learn_clas.path = Path('/notebooks/temp')
learn_clas.save('classifier_single_epoch_'+modifier+'d')
| ch4/text_standalone_dataset_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chest X-Ray Medical Diagnosis with Deep Learning
# + [markdown] colab_type="text" id="FZYK-0rin5x7"
# <img src="xray-header-image.png" style="padding-top: 50px;width: 87%;left: 0px;margin-left: 0px;margin-right: 0px;">
#
# __Welcome to the first assignment of course 1!__
#
# In this assignment! You will explore medical image diagnosis by building a state-of-the-art chest X-ray classifier using Keras.
#
# The assignment will walk through some of the steps of building and evaluating this deep learning classifier model. In particular, you will:
# - Pre-process and prepare a real-world X-ray dataset
# - Use transfer learning to retrain a DenseNet model for X-ray image classification
# - Learn a technique to handle class imbalance
# - Measure diagnostic performance by computing the AUC (Area Under the Curve) for the ROC (Receiver Operating Characteristic) curve
# - Visualize model activity using GradCAMs
#
# In completing this assignment you will learn about the following topics:
#
# - Data preparation
# - Visualizing data
# - Preventing data leakage
# - Model Development
# - Addressing class imbalance
# - Leveraging pre-trained models using transfer learning
# - Evaluation
# - AUC and ROC curves
# -
# ## Outline
# Use these links to jump to specific sections of this assignment!
#
# - [1. Import Packages and Function](#1)
# - [2. Load the Datasets](#2)
# - [2.1 Preventing Data Leakage](#2-1)
# - [Exercise 1 - Checking Data Leakage](#Ex-1)
# - [2.2 Preparing Images](#2-2)
# - [3. Model Development](#3)
# - [3.1 Addressing Class Imbalance](#3-1)
# - [Exercise 2 - Computing Class Frequencies](#Ex-2)
# - [Exercise 3 - Weighted Loss](#Ex-3)
# - [3.3 DenseNet121](#3-3)
# - [4. Training [optional]](#4)
# - [4.1 Training on the Larger Dataset](#4-1)
# - [5. Prediction and Evaluation](#5)
# - [5.1 ROC Curve and AUROC](#5-1)
# - [5.2 Visualizing Learning with GradCAM](#5-2)
# + [markdown] colab_type="text" id="XI8PBrk_2Z4V"
# <a name='1'></a>
# ## 1. Import Packages and Functions¶
#
# We'll make use of the following packages:
# - `numpy` and `pandas` is what we'll use to manipulate our data
# - `matplotlib.pyplot` and `seaborn` will be used to produce plots for visualization
# - `util` will provide the locally defined utility functions that have been provided for this assignment
#
# We will also use several modules from the `keras` framework for building deep learning models.
#
# Run the next cell to import all the necessary packages.
# + colab={} colab_type="code" id="Je3yV0Wnn5x8"
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from keras.preprocessing.image import ImageDataGenerator
from keras.applications.densenet import DenseNet121
from keras.layers import Dense, GlobalAveragePooling2D
from keras.models import Model
from keras import backend as K
from keras.models import load_model
import util
# + [markdown] colab_type="text" id="6PMDCWQRn5yA"
# <a name='2'></a>
# ## 2 Load the Datasets
#
# For this assignment, we will be using the [ChestX-ray8 dataset](https://arxiv.org/abs/1705.02315) which contains 108,948 frontal-view X-ray images of 32,717 unique patients.
# - Each image in the data set contains multiple text-mined labels identifying 14 different pathological conditions.
# - These in turn can be used by physicians to diagnose 8 different diseases.
# - We will use this data to develop a single model that will provide binary classification predictions for each of the 14 labeled pathologies.
# - In other words it will predict 'positive' or 'negative' for each of the pathologies.
#
# You can download the entire dataset for free [here](https://nihcc.app.box.com/v/ChestXray-NIHCC).
# - We have provided a ~1000 image subset of the images for you.
# - These can be accessed in the folder path stored in the `IMAGE_DIR` variable.
#
# The dataset includes a CSV file that provides the labels for each X-ray.
#
# To make your job a bit easier, we have processed the labels for our small sample and generated three new files to get you started. These three files are:
#
# 1. `nih/train-small.csv`: 875 images from our dataset to be used for training.
# 1. `nih/valid-small.csv`: 109 images from our dataset to be used for validation.
# 1. `nih/test.csv`: 420 images from our dataset to be used for testing.
#
# This dataset has been annotated by consensus among four different radiologists for 5 of our 14 pathologies:
# - `Consolidation`
# - `Edema`
# - `Effusion`
# - `Cardiomegaly`
# - `Atelectasis`
# -
# #### Sidebar on meaning of 'class'
# It is worth noting that the word **'class'** is used in multiple ways is these discussions.
# - We sometimes refer to each of the 14 pathological conditions that are labeled in our dataset as a class.
# - But for each of those pathologies we are attempting to predict whether a certain condition is present (i.e. positive result) or absent (i.e. negative result).
# - These two possible labels of 'positive' or 'negative' (or the numerical equivalent of 1 or 0) are also typically referred to as classes.
# - Moreover, we also use the term in reference to software code 'classes' such as `ImageDataGenerator`.
#
# As long as you are aware of all this though, it should not cause you any confusion as the term 'class' is usually clear from the context in which it is used.
# #### Read in the data
# Let's open these files using the [pandas](https://pandas.pydata.org/) library
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="5JRSHB7i0t_6" outputId="69830050-af47-4ebc-946d-d411d0cbdf5b"
train_df = pd.read_csv("nih/train-small.csv")
valid_df = pd.read_csv("nih/valid-small.csv")
test_df = pd.read_csv("nih/test.csv")
train_df.head()
# + colab={} colab_type="code" id="mrDoMlsun5yE"
labels = ['Cardiomegaly',
'Emphysema',
'Effusion',
'Hernia',
'Infiltration',
'Mass',
'Nodule',
'Atelectasis',
'Pneumothorax',
'Pleural_Thickening',
'Pneumonia',
'Fibrosis',
'Edema',
'Consolidation']
# + [markdown] colab_type="text" id="iKwFwpHLn5yG"
# <a name='2-1'></a>
# ### 2.1 Preventing Data Leakage
# It is worth noting that our dataset contains multiple images for each patient. This could be the case, for example, when a patient has taken multiple X-ray images at different times during their hospital visits. In our data splitting, we have ensured that the split is done on the patient level so that there is no data "leakage" between the train, validation, and test datasets.
# -
# <a name='Ex-1'></a>
# ### Exercise 1 - Checking Data Leakage
# In the cell below, write a function to check whether there is leakage between two datasets. We'll use this to make sure there are no patients in the test set that are also present in either the train or validation sets.
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li> Make use of python's set.intersection() function. </li>
# <li> In order to match the automatic grader's expectations, please start the line of code with <code>df1_patients_unique...[continue your code here]</code> </li>
#
# </ul>
# </p>
# + colab={} colab_type="code" id="Jz6dwTSrUcKc"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def check_for_leakage(df1, df2, patient_col):
"""
Return True if there any patients are in both df1 and df2.
Args:
df1 (dataframe): dataframe describing first dataset
df2 (dataframe): dataframe describing second dataset
patient_col (str): string name of column with patient IDs
Returns:
leakage (bool): True if there is leakage, otherwise False
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
df1_patients_unique = set(df1[patient_col].values)
df2_patients_unique = set(df2[patient_col].values)
patients_in_both_groups = list(df1_patients_unique.intersection(df2_patients_unique))
# leakage contains true if there is patient overlap, otherwise false.
leakage = bool(df1_patients_unique.intersection(patients_in_both_groups)) # boolean (true if there is at least 1 patient in both groups)
### END CODE HERE ###
return leakage
# + colab={"base_uri": "https://localhost:8080/", "height": 544} colab_type="code" id="Rh2p1krrV1g5" outputId="9ee44d93-8ef1-4c98-f9fa-65b309b9b889"
# test
print("test case 1")
df1 = pd.DataFrame({'patient_id': [0, 1, 2]})
df2 = pd.DataFrame({'patient_id': [2, 3, 4]})
print("df1")
print(df1)
print("df2")
print(df2)
print(f"leakage output: {check_for_leakage(df1, df2, 'patient_id')}")
print("-------------------------------------")
print("test case 2")
df1 = pd.DataFrame({'patient_id': [0, 1, 2]})
df2 = pd.DataFrame({'patient_id': [3, 4, 5]})
print("df1:")
print(df1)
print("df2:")
print(df2)
print(f"leakage output: {check_for_leakage(df1, df2, 'patient_id')}")
# -
# ##### Expected output
#
# ```Python
# test case 1
# df1
# patient_id
# 0 0
# 1 1
# 2 2
# df2
# patient_id
# 0 2
# 1 3
# 2 4
# leakage output: True
# -------------------------------------
# test case 2
# df1:
# patient_id
# 0 0
# 1 1
# 2 2
# df2:
# patient_id
# 0 3
# 1 4
# 2 5
# leakage output: False
# ```
# + [markdown] colab_type="text" id="FCWkiLudW_Il"
# Run the next cell to check if there are patients in both train and test or in both valid and test.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="AMF3Wd3yW-RS" outputId="e417c9ea-c06b-49a7-af35-d802bc1725eb"
print("leakage between train and test: {}".format(check_for_leakage(train_df, test_df, 'PatientId')))
print("leakage between valid and test: {}".format(check_for_leakage(valid_df, test_df, 'PatientId')))
# + [markdown] colab_type="text" id="zRUvYHpYXhlQ"
# If we get `False` for both, then we're ready to start preparing the datasets for training. Remember to always check for data leakage!
# + [markdown] colab_type="text" id="JBWZ5l4ln5yH"
# <a name='2-2'></a>
# ### 2.2 Preparing Images
# + [markdown] colab_type="text" id="SPjuZHPpn5yH"
# With our dataset splits ready, we can now proceed with setting up our model to consume them.
# - For this we will use the off-the-shelf [ImageDataGenerator](https://keras.io/preprocessing/image/) class from the Keras framework, which allows us to build a "generator" for images specified in a dataframe.
# - This class also provides support for basic data augmentation such as random horizontal flipping of images.
# - We also use the generator to transform the values in each batch so that their mean is $0$ and their standard deviation is 1.
# - This will facilitate model training by standardizing the input distribution.
# - The generator also converts our single channel X-ray images (gray-scale) to a three-channel format by repeating the values in the image across all channels.
# - We will want this because the pre-trained model that we'll use requires three-channel inputs.
#
# Since it is mainly a matter of reading and understanding Keras documentation, we have implemented the generator for you. There are a few things to note:
# 1. We normalize the mean and standard deviation of the data
# 3. We shuffle the input after each epoch.
# 4. We set the image size to be 320px by 320px
# + colab={} colab_type="code" id="nAgVGOAju8pX"
def get_train_generator(df, image_dir, x_col, y_cols, shuffle=True, batch_size=8, seed=1, target_w = 320, target_h = 320):
"""
Return generator for training set, normalizing using batch
statistics.
Args:
train_df (dataframe): dataframe specifying training data.
image_dir (str): directory where image files are held.
x_col (str): name of column in df that holds filenames.
y_cols (list): list of strings that hold y labels for images.
batch_size (int): images per batch to be fed into model during training.
seed (int): random seed.
target_w (int): final width of input images.
target_h (int): final height of input images.
Returns:
train_generator (DataFrameIterator): iterator over training set
"""
print("getting train generator...")
# normalize images
image_generator = ImageDataGenerator(
samplewise_center=True,
samplewise_std_normalization= True)
# flow from directory with specified batch size
# and target image size
generator = image_generator.flow_from_dataframe(
dataframe=df,
directory=image_dir,
x_col=x_col,
y_col=y_cols,
class_mode="raw",
batch_size=batch_size,
shuffle=shuffle,
seed=seed,
target_size=(target_w,target_h))
return generator
# + [markdown] colab_type="text" id="vpRXR-3_u7cl"
# #### Build a separate generator for valid and test sets
#
# Now we need to build a new generator for validation and testing data.
#
# **Why can't we use the same generator as for the training data?**
#
# Look back at the generator we wrote for the training data.
# - It normalizes each image **per batch**, meaning that it uses batch statistics.
# - We should not do this with the test and validation data, since in a real life scenario we don't process incoming images a batch at a time (we process one image at a time).
# - Knowing the average per batch of test data would effectively give our model an advantage.
# - The model should not have any information about the test data.
#
# What we need to do is normalize incoming test data using the statistics **computed from the training set**.
# * We implement this in the function below.
# * There is one technical note. Ideally, we would want to compute our sample mean and standard deviation using the entire training set.
# * However, since this is extremely large, that would be very time consuming.
# * In the interest of time, we'll take a random sample of the dataset and calcualte the sample mean and sample standard deviation.
# + colab={} colab_type="code" id="UtWEAfAnrhMq"
def get_test_and_valid_generator(valid_df, test_df, train_df, image_dir, x_col, y_cols, sample_size=100, batch_size=8, seed=1, target_w = 320, target_h = 320):
"""
Return generator for validation set and test test set using
normalization statistics from training set.
Args:
valid_df (dataframe): dataframe specifying validation data.
test_df (dataframe): dataframe specifying test data.
train_df (dataframe): dataframe specifying training data.
image_dir (str): directory where image files are held.
x_col (str): name of column in df that holds filenames.
y_cols (list): list of strings that hold y labels for images.
sample_size (int): size of sample to use for normalization statistics.
batch_size (int): images per batch to be fed into model during training.
seed (int): random seed.
target_w (int): final width of input images.
target_h (int): final height of input images.
Returns:
test_generator (DataFrameIterator) and valid_generator: iterators over test set and validation set respectively
"""
print("getting train and valid generators...")
# get generator to sample dataset
raw_train_generator = ImageDataGenerator().flow_from_dataframe(
dataframe=train_df,
directory=IMAGE_DIR,
x_col="Image",
y_col=labels,
class_mode="raw",
batch_size=sample_size,
shuffle=True,
target_size=(target_w, target_h))
# get data sample
batch = raw_train_generator.next()
data_sample = batch[0]
# use sample to fit mean and std for test set generator
image_generator = ImageDataGenerator(
featurewise_center=True,
featurewise_std_normalization= True)
# fit generator to sample from training data
image_generator.fit(data_sample)
# get test generator
valid_generator = image_generator.flow_from_dataframe(
dataframe=valid_df,
directory=image_dir,
x_col=x_col,
y_col=y_cols,
class_mode="raw",
batch_size=batch_size,
shuffle=False,
seed=seed,
target_size=(target_w,target_h))
test_generator = image_generator.flow_from_dataframe(
dataframe=test_df,
directory=image_dir,
x_col=x_col,
y_col=y_cols,
class_mode="raw",
batch_size=batch_size,
shuffle=False,
seed=seed,
target_size=(target_w,target_h))
return valid_generator, test_generator
# + [markdown] colab_type="text" id="ga4RZN5On5yL"
# With our generator function ready, let's make one generator for our training data and one each of our test and validation datasets.
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="rNE3HWRbn5yL" outputId="4c6b1c25-a33d-42e0-f442-40971ca52a3f"
IMAGE_DIR = "nih/images-small/"
train_generator = get_train_generator(train_df, IMAGE_DIR, "Image", labels)
valid_generator, test_generator= get_test_and_valid_generator(valid_df, test_df, train_df, IMAGE_DIR, "Image", labels)
# + [markdown] colab_type="text" id="pYtXacDgn5yN"
# Let's peek into what the generator gives our model during training and validation. We can do this by calling the `__get_item__(index)` function:
# + colab={"base_uri": "https://localhost:8080/", "height": 303} colab_type="code" id="Jh77vpN-n5yO" outputId="c4e68e79-e8f2-4bb9-8909-072c9dd2f805"
x, y = train_generator.__getitem__(0)
plt.imshow(x[0]);
# + [markdown] colab_type="text" id="9WBMpRxcDMgp"
# <a name='3'></a>
# ## 3 Model Development
#
# Now we'll move on to model training and development. We have a few practical challenges to deal with before actually training a neural network, though. The first is class imbalance.
# + [markdown] colab_type="text" id="qHBSgvxfn5yR"
# <a name='3-1'></a>
# ### 3.1 Addressing Class Imbalance
# One of the challenges with working with medical diagnostic datasets is the large class imbalance present in such datasets. Let's plot the frequency of each of the labels in our dataset:
# + colab={"base_uri": "https://localhost:8080/", "height": 365} colab_type="code" id="-OvyPe5en5yR" outputId="077747ad-7ab8-463d-8335-6b243cb29e63"
plt.xticks(rotation=90)
plt.bar(x=labels, height=np.mean(train_generator.labels, axis=0))
plt.title("Frequency of Each Class")
plt.show()
# -
# We can see from this plot that the prevalance of positive cases varies significantly across the different pathologies. (These trends mirror the ones in the full dataset as well.)
# * The `Hernia` pathology has the greatest imbalance with the proportion of positive training cases being about 0.2%.
# * But even the `Infiltration` pathology, which has the least amount of imbalance, has only 17.5% of the training cases labelled positive.
#
# Ideally, we would train our model using an evenly balanced dataset so that the positive and negative training cases would contribute equally to the loss.
#
# If we use a normal cross-entropy loss function with a highly unbalanced dataset, as we are seeing here, then the algorithm will be incentivized to prioritize the majority class (i.e negative in our case), since it contributes more to the loss.
# + [markdown] colab_type="text" id="3nHRd9p9n5yU"
# #### Impact of class imbalance on loss function
#
# Let's take a closer look at this. Assume we would have used a normal cross-entropy loss for each pathology. We recall that the cross-entropy loss contribution from the $i^{th}$ training data case is:
#
# $$\mathcal{L}_{cross-entropy}(x_i) = -(y_i \log(f(x_i)) + (1-y_i) \log(1-f(x_i))),$$
#
# where $x_i$ and $y_i$ are the input features and the label, and $f(x_i)$ is the output of the model, i.e. the probability that it is positive.
#
# Note that for any training case, either $y_i=0$ or else $(1-y_i)=0$, so only one of these terms contributes to the loss (the other term is multiplied by zero, and becomes zero).
#
# We can rewrite the overall average cross-entropy loss over the entire training set $\mathcal{D}$ of size $N$ as follows:
#
# $$\mathcal{L}_{cross-entropy}(\mathcal{D}) = - \frac{1}{N}\big( \sum_{\text{positive examples}} \log (f(x_i)) + \sum_{\text{negative examples}} \log(1-f(x_i)) \big).$$
#
# Using this formulation, we can see that if there is a large imbalance with very few positive training cases, for example, then the loss will be dominated by the negative class. Summing the contribution over all the training cases for each class (i.e. pathological condition), we see that the contribution of each class (i.e. positive or negative) is:
#
# $$freq_{p} = \frac{\text{number of positive examples}}{N} $$
#
# $$\text{and}$$
#
# $$freq_{n} = \frac{\text{number of negative examples}}{N}.$$
# -
# <a name='Ex-2'></a>
# ### Exercise 2 - Computing Class Frequencies
# Complete the function below to calculate these frequences for each label in our dataset.
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li> Use numpy.sum(a, axis=), and choose the axis (0 or 1) </li>
# </ul>
# </p>
#
# + colab={} colab_type="code" id="TpDGeY2cChYD"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def compute_class_freqs(labels):
"""
Compute positive and negative frequences for each class.
Args:
labels (np.array): matrix of labels, size (num_examples, num_classes)
Returns:
positive_frequencies (np.array): array of positive frequences for each
class, size (num_classes)
negative_frequencies (np.array): array of negative frequences for each
class, size (num_classes)
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# total number of patients (rows)
N = len(labels)
positive_frequencies = np.sum(labels == 1, axis = 0 )/N
negative_frequencies = np.sum(labels == 0, axis = 0 )/N
### END CODE HERE ###
return positive_frequencies, negative_frequencies
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="BqidQvCaD_xi" outputId="56a5905a-e61b-47a8-f444-aa89d7481c44"
# Test
labels_matrix = np.array(
[[1, 0, 0],
[0, 1, 1],
[1, 0, 1],
[1, 1, 1],
[1, 0, 1]]
)
print("labels:")
print(labels_matrix)
test_pos_freqs, test_neg_freqs = compute_class_freqs(labels_matrix)
print(f"pos freqs: {test_pos_freqs}")
print(f"neg freqs: {test_neg_freqs}")
# -
# ##### Expected output
#
# ```Python
# labels:
# [[1 0 0]
# [0 1 1]
# [1 0 1]
# [1 1 1]
# [1 0 1]]
# pos freqs: [0.8 0.4 0.8]
# neg freqs: [0.2 0.6 0.2]
# ```
# + [markdown] colab_type="text" id="Iye-sQoOFG37"
# Now we'll compute frequencies for our training data.
# + colab={} colab_type="code" id="LoxM5jQ0E30D"
freq_pos, freq_neg = compute_class_freqs(train_generator.labels)
freq_pos
# + [markdown] colab_type="text" id="gsJIDPTZn5yW"
# Let's visualize these two contribution ratios next to each other for each of the pathologies:
# + colab={"base_uri": "https://localhost:8080/", "height": 363} colab_type="code" id="IqnNCu4In5yW" outputId="245f1a6b-b292-4c6d-a583-c6924bc61f31"
data = pd.DataFrame({"Class": labels, "Label": "Positive", "Value": freq_pos})
data = data.append([{"Class": labels[l], "Label": "Negative", "Value": v} for l,v in enumerate(freq_neg)], ignore_index=True)
plt.xticks(rotation=90)
f = sns.barplot(x="Class", y="Value", hue="Label" ,data=data)
# + [markdown] colab_type="text" id="2uvttCM8n5yY"
# As we see in the above plot, the contributions of positive cases is significantly lower than that of the negative ones. However, we want the contributions to be equal. One way of doing this is by multiplying each example from each class by a class-specific weight factor, $w_{pos}$ and $w_{neg}$, so that the overall contribution of each class is the same.
#
# To have this, we want
#
# $$w_{pos} \times freq_{p} = w_{neg} \times freq_{n},$$
#
# which we can do simply by taking
#
# $$w_{pos} = freq_{neg}$$
# $$w_{neg} = freq_{pos}$$
#
# This way, we will be balancing the contribution of positive and negative labels.
# + colab={} colab_type="code" id="zs3_Rgwwn5yZ"
pos_weights = freq_neg
neg_weights = freq_pos
pos_contribution = freq_pos * pos_weights
neg_contribution = freq_neg * neg_weights
# + [markdown] colab_type="text" id="ygNZmdyun5ya"
# Let's verify this by graphing the two contributions next to each other again:
# + colab={"base_uri": "https://localhost:8080/", "height": 380} colab_type="code" id="LPfSFrxjn5yb" outputId="a4b6354f-ab39-4623-d44b-90cfd9b28506"
data = pd.DataFrame({"Class": labels, "Label": "Positive", "Value": pos_contribution})
data = data.append([{"Class": labels[l], "Label": "Negative", "Value": v}
for l,v in enumerate(neg_contribution)], ignore_index=True)
plt.xticks(rotation=90)
sns.barplot(x="Class", y="Value", hue="Label" ,data=data);
# + [markdown] colab_type="text" id="u9xgoEkpn5yc"
# As the above figure shows, by applying these weightings the positive and negative labels within each class would have the same aggregate contribution to the loss function. Now let's implement such a loss function.
#
# After computing the weights, our final weighted loss for each training case will be
#
# $$\mathcal{L}_{cross-entropy}^{w}(x) = - (w_{p} y \log(f(x)) + w_{n}(1-y) \log( 1 - f(x) ) ).$$
# -
# <a name='Ex-3'></a>
# ### Exercise 3 - Weighted Loss
# Fill out the `weighted_loss` function below to return a loss function that calculates the weighted loss for each batch. Recall that for the multi-class loss, we add up the average loss for each individual class. Note that we also want to add a small value, $\epsilon$, to the predicted values before taking their logs. This is simply to avoid a numerical error that would otherwise occur if the predicted value happens to be zero.
#
# ##### Note
# Please use Keras functions to calculate the mean and the log.
#
# - [Keras.mean](https://www.tensorflow.org/api_docs/python/tf/keras/backend/mean)
# - [Keras.log](https://www.tensorflow.org/api_docs/python/tf/keras/backend/log)
#
#
# + colab={} colab_type="code" id="pPIBVAasn5yd"
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_weighted_loss(pos_weights, neg_weights, epsilon=1e-7):
"""
Return weighted loss function given negative weights and positive weights.
Args:
pos_weights (np.array): array of positive weights for each class, size (num_classes)
neg_weights (np.array): array of negative weights for each class, size (num_classes)
Returns:
weighted_loss (function): weighted loss function
"""
def weighted_loss(y_true, y_pred):
"""
Return weighted loss value.
Args:
y_true (Tensor): Tensor of true labels, size is (num_examples, num_classes)
y_pred (Tensor): Tensor of predicted labels, size is (num_examples, num_classes)
Returns:
loss (Float): overall scalar loss summed across all classes
"""
# initialize loss to zero
loss = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
for i in range(len(pos_weights)):
# for each class, add average weighted loss for that class
loss += -(K.mean( pos_weights[i] * y_true[:,i] * K.log(y_pred[:,i] + epsilon) + \
neg_weights[i] * (1 - y_true[:,i]) * K.log(1 - y_pred[:,i] + epsilon), axis = 0)) #complete this line
return loss
### END CODE HERE ###
return weighted_loss
# -
# Now let's test our function with some simple cases.
# + colab={"base_uri": "https://localhost:8080/", "height": 595} colab_type="code" id="CFjYda3Wulbm" outputId="87128f53-7a99-40e2-d09a-6539215879d0"
# Test
sess = K.get_session()
with sess.as_default() as sess:
print("Test example:\n")
y_true = K.constant(np.array(
[[1, 1, 1],
[1, 1, 0],
[0, 1, 0],
[1, 0, 1]]
))
print("y_true:\n")
print(y_true.eval())
w_p = np.array([0.25, 0.25, 0.5])
w_n = np.array([0.75, 0.75, 0.5])
print("\nw_p:\n")
print(w_p)
print("\nw_n:\n")
print(w_n)
y_pred_1 = K.constant(0.7*np.ones(y_true.shape))
print("\ny_pred_1:\n")
print(y_pred_1.eval())
y_pred_2 = K.constant(0.3*np.ones(y_true.shape))
print("\ny_pred_2:\n")
print(y_pred_2.eval())
# test with a large epsilon in order to catch errors
L = get_weighted_loss(w_p, w_n, epsilon=1)
print("\nIf we weighted them correctly, we expect the two losses to be the same.")
L1 = L(y_true, y_pred_1).eval()
L2 = L(y_true, y_pred_2).eval()
print(f"\nL(y_pred_1)= {L1:.4f}, L(y_pred_2)= {L2:.4f}")
print(f"Difference is L1 - L2 = {L1 - L2:.4f}")
# -
# #### Additional check
# If you implemented the function correctly, then if the epsilon for the `get_weighted_loss` is set to `1`, the weighted losses will be as follows:
# ```Python
# L(y_pred_1)= -0.4956, L(y_pred_2)= -0.4956
# ```
# If you are missing something in your implementation, you will see a different set of losses for L1 and L2 (even though L1 and L2 will be the same).
# + [markdown] colab_type="text" id="yDZQMmlgn5yh"
# <a name='3-3'></a>
# ### 3.3 DenseNet121
#
# Next, we will use a pre-trained [DenseNet121](https://www.kaggle.com/pytorch/densenet121) model which we can load directly from Keras and then add two layers on top of it:
# 1. A `GlobalAveragePooling2D` layer to get the average of the last convolution layers from DenseNet121.
# 2. A `Dense` layer with `sigmoid` activation to get the prediction logits for each of our classes.
#
# We can set our custom loss function for the model by specifying the `loss` parameter in the `compile()` function.
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="gZlxoCTgn5yi" outputId="7e12120b-8aab-403c-b5ca-2ff77ef978b1"
# create the base pre-trained model
base_model = DenseNet121(weights='./nih/densenet.hdf5', include_top=False)
x = base_model.output
# add a global spatial average pooling layer
x = GlobalAveragePooling2D()(x)
# and a logistic layer
predictions = Dense(len(labels), activation="sigmoid")(x)
model = Model(inputs=base_model.input, outputs=predictions)
model.compile(optimizer='adam', loss=get_weighted_loss(pos_weights, neg_weights))
# + [markdown] colab_type="text" id="BcwhQdOAn5ym"
# <a name='4'></a>
# ## 4 Training [optional]
#
# With our model ready for training, we will use the `model.fit()` function in Keras to train our model.
# - We are training on a small subset of the dataset (~1%).
# - So what we care about at this point is to make sure that the loss on the training set is decreasing.
#
# Since training can take a considerable time, for pedagogical purposes we have chosen not to train the model here but rather to load a set of pre-trained weights in the next section. However, you can use the code shown below to practice training the model locally on your machine or in Colab.
#
# **NOTE:** Do not run the code below on the Coursera platform as it will exceed the platform's memory limitations.
#
# Python Code for training the model:
#
# ```python
# history = model.fit_generator(train_generator,
# validation_data=valid_generator,
# steps_per_epoch=100,
# validation_steps=25,
# epochs = 3)
#
# plt.plot(history.history['loss'])
# plt.ylabel("loss")
# plt.xlabel("epoch")
# plt.title("Training Loss Curve")
# plt.show()
# ```
# + [markdown] colab_type="text" id="xB5nsGKrn5yp"
# <a name='4-1'></a>
# ### 4.1 Training on the Larger Dataset
#
# Given that the original dataset is 40GB+ in size and the training process on the full dataset takes a few hours, we have trained the model on a GPU-equipped machine for you and provided the weights file from our model (with a batch size of 32 instead) to be used for the rest of this assignment.
#
# The model architecture for our pre-trained model is exactly the same, but we used a few useful Keras "callbacks" for this training. Do spend time to read about these callbacks at your leisure as they will be very useful for managing long-running training sessions:
#
# 1. You can use `ModelCheckpoint` callback to monitor your model's `val_loss` metric and keep a snapshot of your model at the point.
# 2. You can use the `TensorBoard` to use the Tensorflow Tensorboard utility to monitor your runs in real-time.
# 3. You can use the `ReduceLROnPlateau` to slowly decay the learning rate for your model as it stops getting better on a metric such as `val_loss` to fine-tune the model in the final steps of training.
# 4. You can use the `EarlyStopping` callback to stop the training job when your model stops getting better in it's validation loss. You can set a `patience` value which is the number of epochs the model does not improve after which the training is terminated. This callback can also conveniently restore the weights for the best metric at the end of training to your model.
#
# You can read about these callbacks and other useful Keras callbacks [here](https://keras.io/callbacks/).
#
# Let's load our pre-trained weights into the model now:
# + colab={} colab_type="code" id="887bSajLn5yq"
model.load_weights("./nih/pretrained_model.h5")
# + [markdown] colab_type="text" id="mA90g8n6suRV"
# <a name='5'></a>
# ## 5 Prediction and Evaluation
# + [markdown] colab_type="text" id="Kz1BEwOyxFbj"
# Now that we have a model, let's evaluate it using our test set. We can conveniently use the `predict_generator` function to generate the predictions for the images in our test set.
#
# **Note:** The following cell can take about 4 minutes to run.
# + colab={} colab_type="code" id="QzNrhtf1w2bI"
predicted_vals = model.predict_generator(test_generator, steps = len(test_generator))
# + [markdown] colab_type="text" id="wtjCtaGen5yt"
# <a name='5-1'></a>
# ### 5.1 ROC Curve and AUROC
# We'll cover topic of model evaluation in much more detail in later weeks, but for now we'll walk through computing a metric called the AUC (Area Under the Curve) from the ROC ([Receiver Operating Characteristic](https://en.wikipedia.org/wiki/Receiver_operating_characteristic)) curve. This is also referred to as the AUROC value, but you will see all three terms in reference to the technique, and often used almost interchangeably.
#
# For now, what you need to know in order to interpret the plot is that a curve that is more to the left and the top has more "area" under it, and indicates that the model is performing better.
#
# We will use the `util.get_roc_curve()` function which has been provided for you in `util.py`. Look through this function and note the use of the `sklearn` library functions to generate the ROC curves and AUROC values for our model.
#
# - [roc_curve](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_curve.html)
# - [roc_auc_score](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html)
# + colab={"base_uri": "https://localhost:8080/", "height": 621} colab_type="code" id="6SLI8FHun5yw" outputId="4f5cc99c-4e1a-421b-fe2d-637df32d6416"
auc_rocs = util.get_roc_curve(labels, predicted_vals, test_generator)
# + [markdown] colab_type="text" id="zWZkl01ruZ7P"
# You can compare the performance to the AUCs reported in the original ChexNeXt paper in the table below:
# + [markdown] colab_type="text" id="GZUoShw2n5yy"
# For reference, here's the AUC figure from the ChexNeXt paper which includes AUC values for their model as well as radiologists on this dataset:
#
# <img src="https://journals.plos.org/plosmedicine/article/figure/image?size=large&id=10.1371/journal.pmed.1002686.t001" width="80%">
#
# This method does take advantage of a few other tricks such as self-training and ensembling as well, which can give a significant boost to the performance.
# + [markdown] colab_type="text" id="Jzy7fLgFn5yy"
# For details about the best performing methods and their performance on this dataset, we encourage you to read the following papers:
# - [CheXNet](https://arxiv.org/abs/1711.05225)
# - [CheXpert](https://arxiv.org/pdf/1901.07031.pdf)
# - [ChexNeXt](https://journals.plos.org/plosmedicine/article?id=10.1371/journal.pmed.1002686)
# + [markdown] colab_type="text" id="G5aZAlVbn5yz"
# <a name='5-2'></a>
# ### 5.2 Visualizing Learning with GradCAM
#
# + [markdown] colab_type="text" id="gu9ExySryY6u"
# One of the challenges of using deep learning in medicine is that the complex architecture used for neural networks makes them much harder to interpret compared to traditional machine learning models (e.g. linear models).
#
# One of the most common approaches aimed at increasing the interpretability of models for computer vision tasks is to use Class Activation Maps (CAM).
# - Class activation maps are useful for understanding where the model is "looking" when classifying an image.
#
# In this section we will use a [GradCAM's](https://arxiv.org/abs/1610.02391) technique to produce a heatmap highlighting the important regions in the image for predicting the pathological condition.
# - This is done by extracting the gradients of each predicted class, flowing into our model's final convolutional layer. Look at the `util.compute_gradcam` which has been provided for you in `util.py` to see how this is done with the Keras framework.
#
# It is worth mentioning that GradCAM does not provide a full explanation of the reasoning for each classification probability.
# - However, it is still a useful tool for "debugging" our model and augmenting our prediction so that an expert could validate that a prediction is indeed due to the model focusing on the right regions of the image.
# -
# First we will load the small training set and setup to look at the 4 classes with the highest performing AUC measures.
# + colab={} colab_type="code" id="6kahoZbJn5yz" outputId="ade0a4e2-4591-4ba5-ec19-1a3487e3f972"
df = pd.read_csv("nih/train-small.csv")
IMAGE_DIR = "nih/images-small/"
# only show the labels with top 4 AUC
labels_to_show = np.take(labels, np.argsort(auc_rocs)[::-1])[:4]
# -
# Now let's look at a few specific images.
util.compute_gradcam(model, '00008270_015.png', IMAGE_DIR, df, labels, labels_to_show)
# + colab={} colab_type="code" id="JC2zy1Kpn5y1" outputId="9e38a769-e19d-4143-da41-db7a3173a533"
util.compute_gradcam(model, '00011355_002.png', IMAGE_DIR, df, labels, labels_to_show)
# + colab={} colab_type="code" id="zCHVaLMQn5y2" outputId="57246709-2662-4590-9198-a412d2f1eea2"
util.compute_gradcam(model, '00029855_001.png', IMAGE_DIR, df, labels, labels_to_show)
# + colab={} colab_type="code" id="gGwL8FcFn5y4" outputId="681fb2de-194c-465e-c989-133f334b8299"
util.compute_gradcam(model, '00005410_000.png', IMAGE_DIR, df, labels, labels_to_show)
# -
# Congratulations, you've completed the first assignment of course one! You've learned how to preprocess data, check for data leakage, train a pre-trained model, and evaluate using the AUC. Great work!
| AI for Medical Diagnosis/Week 1/Chest X-Ray Medical Diagnosis with Deep Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Use Tensorflow to recognize hand-written digits with Watson Machine Learning REST API
# This notebook contains steps and code to demonstrate support of Tensordlow Deep Learning experiments in Watson Machine Learning Service. It introduces commands for getting data, training experiments, persisting pipelines, publishing models, deploying models and scoring.
#
# Some familiarity with cURL is helpful. This notebook uses cURL examples.
#
#
# ## Learning goals
#
# The learning goals of this notebook are:
#
# - Working with Watson Machine Learning experiments to train Deep Learning models.
# - Downloading computed models to local storage.
# - Online deployment and scoring of trained model.
#
#
# ## Contents
#
# This notebook contains the following parts:
#
# 1. [Setup](#setup)
# 2. [Model definition](#model_definition)
# 3. [Experiment Run](#run)
# 4. [Historical runs](#runs)
# 5. [Deploy and Score](#deploy_and_score)
# 6. [Cleaning](#cleaning)
# 7. [Summary and next steps](#summary)
# <a id="setup"></a>
# ## 1. Set up the environment
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# - Contact with your Cloud Pack for Data administrator and ask him for your account credentials
# ### Connection to WML
#
# Authenticate the Watson Machine Learning service on IBM Cloud Pack for Data. You need to provide platform `url`, your `username` and `password`.
# +
# %env USERNAME=
# %env PASSWORD=
# %env DATAPLATFORM_URL=
# %env SPACE_ID=
# -
# <a id="wml_token"></a>
# ### Getting WML authorization token for further cURL calls
# <a href="https://cloud.ibm.com/docs/cloud-object-storage?topic=cloud-object-storage-curl#curl-token" target="_blank" rel="noopener no referrer">Example of cURL call to get WML token</a>
# + magic_args="--out token" language="bash"
#
# token=$(curl -sk -X GET \
# --user $USERNAME:$PASSWORD \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/v1/preauth/validateAuth")
#
# token=${token#*accessToken\":\"}
# token=${token%%\"*}
# echo $token
# -
# %env TOKEN=$token
# <a id="space_creation"></a>
# ### Space creation
# **Tip:** If you do not have `space` already created, please convert below three cells to `code` and run them.
#
# First of all, you need to create a `space` that will be used in all of your further cURL calls.
# If you do not have `space` already created, below is the cURL call to create one.
# <a href="https://cpd-spaces-api.eu-gb.cf.appdomain.cloud/#/Spaces/spaces_create"
# target="_blank" rel="noopener no referrer">Space creation</a>
# + magic_args="--out space_id" language="bash" active=""
#
# curl -sk -X POST \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data '{"name": "curl_DL"}' \
# "$DATAPLATFORM_URL/v2/spaces" \
# | grep '"id": ' | awk -F '"' '{ print $4 }'
# + active=""
# space_id = space_id.split('\n')[1]
# %env SPACE_ID=$space_id
# -
# Space creation is asynchronous. This means that you need to check space creation status after creation call.
# Make sure that your newly created space is `active`.
# <a href="https://cpd-spaces-api.eu-gb.cf.appdomain.cloud/#/Spaces/spaces_get"
# target="_blank" rel="noopener no referrer">Get space information</a>
# + language="bash" active=""
#
# curl -sk -X GET \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/v2/spaces/$SPACE_ID"
# -
# <a id="model_definition"></a>
# ## 2. Model definition
#
# This section provides samples about how to store model definition via cURL calls.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Model%20Definitions/model_definitions_create"
# target="_blank" rel="noopener no referrer">Store a model definition for Deep Learning experiment</a>
# + magic_args="--out model_definition_payload" language="bash"
#
# MODEL_DEFINITION_PAYLOAD='{"name": "Hand-written Digit Recognition", "space_id": "'"$SPACE_ID"'", "description": "Hand-written Digit Recognition", "tags": ["DL", "TF"], "version": "v1", "platform": {"name": "python", "versions": ["3.7"]}, "command": "convolutional_network.py --trainImagesFile train-images-idx3-ubyte.gz --trainLabelsFile train-labels-idx1-ubyte.gz --testImagesFile t10k-images-idx3-ubyte.gz --testLabelsFile t10k-labels-idx1-ubyte.gz --learningRate 0.001 --trainingIters 6000"}'
# echo $MODEL_DEFINITION_PAYLOAD | python -m json.tool
# -
# %env MODEL_DEFINITION_PAYLOAD=$model_definition_payload
# + magic_args="--out model_definition_id" language="bash"
#
# curl -sk -X POST \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data "$MODEL_DEFINITION_PAYLOAD" \
# "$DATAPLATFORM_URL/ml/v4/model_definitions?version=2020-08-01"| grep '"id": ' | awk -F '"' '{ print $4 }'
# -
# %env MODEL_DEFINITION_ID=$model_definition_id
# <a id="model_preparation"></a>
# ### Model preparation
#
# Download files with keras code. You can either download it via link below or run the cell below the link.
# <a href="https://github.com/IBM/watson-machine-learning-samples/raw/master/definitions/keras/mnist/MNIST.zip"
# target="_blank" rel="noopener no referrer">Download tf-softmax-model.zip</a>
# + language="bash"
#
# wget https://github.com/IBM/watson-machine-learning-samples/raw/master/cpd3.5/definitions/tensorflow/mnist/tf_model_with_metrics_2_1.zip \
# -O tf_model_with_metrics_2_1.zip
# -
# **Tip**: Convert below cell to code and run it to see model deinition's code.
# + active=""
# !unzip -oqd . tf_model_with_metrics_2_1.zip && cat convolutional_network.py
# -
# <a id="def_upload"></a>
# ### Upload model for the model definition
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Model%20Definitions/model_definitions_upload_model"
# target="_blank" rel="noopener no referrer">Upload model for the model definition</a>
# + language="bash"
#
# curl -sk -X PUT \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data-binary "@tf_model_with_metrics_2_1.zip"\
# "$DATAPLATFORM_URL/ml/v4/model_definitions/$MODEL_DEFINITION_ID/model?version=2020-08-01&space_id=$SPACE_ID" \
# | python -m json.tool
# -
# <a id="run"></a>
# ## 3. Experiment run
#
# This section provides samples about how to trigger Deep Learning experiment via cURL calls.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Trainings/trainings_create"
# target="_blank" rel="noopener no referrer">Schedule a training job for Deep Learning experiment</a>
# Specify the source files folder where you have stored your training data. The path should point to a local repository on Watson Machine Learning Accelerator that your system administrator has set up for your use.
#
# **Action:**
# Change `training_data_references: location: path: ...`
# + magic_args="--out training_payload" language="bash"
#
# TRAINING_PAYLOAD='{"training_data_references": [{"name": "training_input_data", "type": "fs", "connection": {}, "location": {"path": "tf-mnist"}, "schema": {"id": "idmlp_schema", "fields": [{"name": "text", "type": "string"}]}}], "results_reference": {"name": "MNIST results", "connection": {}, "location": {"path": "spaces/'"$SPACE_ID"'/assets/experiment"}, "type": "fs"}, "tags": [{"value": "tags_tf", "description": "Tags TF"}], "name": "TF hand-written Digit Recognition", "description": "TF hand-written Digit Recognition", "model_definition": {"id": "'"$MODEL_DEFINITION_ID"'", "command": "convolutional_network.py --trainImagesFile train-images-idx3-ubyte.gz --trainLabelsFile train-labels-idx1-ubyte.gz --testImagesFile t10k-images-idx3-ubyte.gz --testLabelsFile t10k-labels-idx1-ubyte.gz --learningRate 0.001 --trainingIters 6000", "hardware_spec": {"name": "K80", "nodes": 1}, "software_spec": {"name": "tensorflow_2.1-py3.7"}, "parameters": {"name": "MNIST Tf", "description": "Tf DL model"}}, "space_id": "'"$SPACE_ID"'"}'
# echo $TRAINING_PAYLOAD | python -m json.tool
# -
# %env TRAINING_PAYLOAD=$training_payload
# + magic_args="--out training_id" language="bash"
#
# curl -sk -X POST \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data "$TRAINING_PAYLOAD" \
# "$DATAPLATFORM_URL/ml/v4/trainings?version=2020-08-01" | awk -F'"id":' '{print $2}' | cut -c2-37
# -
# %env TRAINING_ID=$training_id
# <a id="training_details"></a>
# ### Get training details
# Treining is an asynchronous endpoint. In case you want to monitor training status and details,
# you need to use a GET method and specify which training you want to monitor by usage of training ID.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Trainings/trainings_get"
# target="_blank" rel="noopener no referrer">Get information about training job</a>
# + language="bash" active=""
#
# curl -sk -X GET \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/ml/v4/trainings/$TRAINING_ID?space_id=$SPACE_ID&version=2020-08-01" \
# | python -m json.tool
# -
# ### Get training status
# + language="bash"
#
# STATUS=$(curl -sk -X GET\
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/ml/v4/trainings/$TRAINING_ID?space_id=$SPACE_ID&version=2020-08-01")
#
# STATUS=${STATUS#*state\":\"}
# STATUS=${STATUS%%\"*}
# echo $STATUS
# -
# Please make sure that training is completed before you go to the next sections.
# Monitor `state` of your training by running above cell couple of times.
# <a id="runs"></a>
# ## 4. Historical runs
#
# In this section you will see cURL examples describing how to get historical training runs information.
# Output should be similar to the output from training creation but you should see more trainings entries.
# Listing trainings:
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Trainings/trainings_list"
# target="_blank" rel="noopener no referrer">Get list of historical training jobs information</a>
# + language="bash"
#
# HISTORICAL_TRAINING_LIMIT_TO_GET=2
#
# curl -sk -X GET \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/ml/v4/trainings?space_id=$SPACE_ID&version=2020-08-01&limit=$HISTORICAL_TRAINING_LIMIT_TO_GET" \
# | python -m json.tool
# -
# <a id="training_cancel"></a>
# ### Cancel training run
#
# **Tip:** If you want to cancel your training, please convert below cell to `code`, specify training ID and run.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Trainings/trainings_delete"
# target="_blank" rel="noopener no referrer">Canceling training</a>
# + language="bash" active=""
#
# TRAINING_ID_TO_CANCEL=...
#
# curl -sk -X DELETE \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/ml/v4/trainings/$TRAINING_ID_TO_DELETE?space_id=$SPACE_ID&version=2020-08-01"
# -
# ---
# <a id="deploy_and_score"></a>
# ## 5. Deploy and Score
#
# In this section you will learn how to deploy and score pipeline model as webservice using WML instance.
# Before deployment creation, you need store your model in WML repository.
# Please see below cURL call example how to do it.
# Download `request.json` with repository request json for model storing.
# + magic_args="--out request_json" language="bash"
#
# curl -sk -X GET \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/v2/asset_files/experiment/$TRAINING_ID/assets/$TRAINING_ID/resources/wml_model/request.json?space_id=$SPACE_ID&version=2020-08-01" \
# | python -m json.tool
# -
# %env MODEL_PAYLOAD=$request_json
# <a id="model_store"></a>
# ### Store Deep Learning model
#
# Store information about your model to WML repository.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Models/models_create"
# target="_blank" rel="noopener no referrer">Model storing</a>
# + magic_args="--out model_details" language="bash"
#
# curl -sk -X POST \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data "$MODEL_PAYLOAD" \
# "$DATAPLATFORM_URL/ml/v4/models?version=2020-08-01&space_id=$SPACE_ID"
# -
# %env MODEL_DETAILS=$model_details
# + magic_args="--out model_id" language="bash"
#
# echo $MODEL_DETAILS | awk -F '"id": ' '{ print $5 }' | cut -d '"' -f 2
# -
# %env MODEL_ID=$model_id
# <a id="deployment_creation"></a>
# ### Deployment creation
#
# An Deep Learning online deployment creation is presented below.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Deployments/deployments_create"
# target="_blank" rel="noopener no referrer">Create deployment</a>
# + magic_args="--out deployment_payload" language="bash"
#
# DEPLOYMENT_PAYLOAD='{"space_id": "'"$SPACE_ID"'","name": "TF Mnist deploymen", "description": "TF model to predict had-written digits","online": {},"hardware_spec": {"name": "S"},"asset": {"id": "'"$MODEL_ID"'"}}'
# echo $DEPLOYMENT_PAYLOAD | python -m json.tool
# -
# %env DEPLOYMENT_PAYLOAD=$deployment_payload
# + magic_args="--out deployment_id" language="bash"
#
# curl -sk -X POST \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data "$DEPLOYMENT_PAYLOAD" \
# "$DATAPLATFORM_URL/ml/v4/deployments?version=2020-08-01" | grep '"id": ' | awk -F '"' '{ print $4 }' | sed -n 3p
# -
# %env DEPLOYMENT_ID=$deployment_id
# <a id="deployment_details"></a>
# ### Get deployment details
# As deployment API is asynchronous, please make sure your deployment is in `ready` state before going to the next points.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Deployments/deployments_get"
# target="_blank" rel="noopener no referrer">Get deployment details</a>
# + language="bash"
#
# curl -sk -X GET \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# "$DATAPLATFORM_URL/ml/v4/deployments/$DEPLOYMENT_ID?space_id=$SPACE_ID&version=2020-08-01" \
# | python -m json.tool
# -
# <a id="input_score"></a>
# ### Prepare scoring input data
# **Hint:** You may need to install numpy using following command `!pip install numpy`
# !wget -q https://github.com/IBM/watson-machine-learning-samples/raw/master/cpd3.5/data/mnist/mnist.npz
# +
import numpy as np
mnist_dataset = np.load('mnist.npz')
test_mnist = mnist_dataset['x_test']
# -
image_1 = (test_mnist[0].ravel() / 255).tolist()
image_2 = (test_mnist[1].ravel() / 255).tolist()
# %matplotlib inline
import matplotlib.pyplot as plt
for i, image in enumerate([test_mnist[0], test_mnist[1]]):
plt.subplot(2, 2, i + 1)
plt.axis('off')
plt.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
# <a id="webservice_score"></a>
# ### Scoring of a webservice
# If you want to make a `score` call on your deployment, please follow a below method:
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Deployment%20Jobs/deployment_jobs_create"
# target="_blank" rel="noopener no referrer">Create deployment job</a>
# + magic_args="-s \"$image_1\" \"$image_2\"" language="bash"
#
# curl -sk -X POST \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# --data '{"space_id": "$SPACE_ID","input_data": [{"values": ['"$1"', '"$2"']}]}' \
# "$DATAPLATFORM_URL/ml/v4/deployments/$DEPLOYMENT_ID/predictions?version=2020-08-01" \
# | python -m json.tool
# -
# <a id="deployments_list"></a>
# ### Listing all deployments
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Deployments/deployments_list"
# target="_blank" rel="noopener no referrer">List deployments details</a>
# + language="bash"
#
# curl -sk -X GET \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# "$DATAPLATFORM_URL/ml/v4/deployments?space_id=$SPACE_ID&version=2020-08-01" \
# | python -m json.tool
# -
# <a id="cleaning"></a>
# ## 6. Cleaning section
#
# Below section is useful when you want to clean all of your previous work within this notebook.
# Just convert below cells into the `code` and run them.
# <a id="training_delete"></a>
# ### Delete training run
# **Tip:** You can completely delete a training run with its metadata.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Trainings/trainings_delete"
# target="_blank" rel="noopener no referrer">Deleting training</a>
# + language="bash" active=""
#
# TRAINING_ID_TO_DELETE=...
#
# curl -sk -X DELETE \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/ml/v4/trainings/$TRAINING_ID_TO_DELETE?space_id=$SPACE_ID&version=2020-08-01&hard_delete=true"
# -
# <a id="deployment_delete"></a>
# ### Deleting deployment
# **Tip:** You can delete existing deployment by calling DELETE method.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Deployments/deployments_delete"
# target="_blank" rel="noopener no referrer">Delete deployment</a>
# + language="bash" active=""
#
# curl -sk -X DELETE \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# --header "Accept: application/json" \
# "$DATAPLATFORM_URL/ml/v4/deployments/$DEPLOYMENT_ID?space_id=$SPACE_ID&version=2020-08-01"
# -
# <a id="model_delete"></a>
# ### Delete model from repository
# **Tip:** If you want to completely remove your stored model and model metadata, just use a DELETE method.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Models/models_delete"
# target="_blank" rel="noopener no referrer">Delete model from repository</a>
# + language="bash" active=""
#
# curl -sk -X DELETE \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# "$DATAPLATFORM_URL/ml/v4/models/$MODEL_ID?space_id=$SPACE_ID&version=2020-08-01"
# -
# <a id="def_delete"></a>
# ### Delete model definition
# **Tip:** If you want to completely remove your model definition, just use a DELETE method.
# <a href="https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Model%20Definitions/model_definitions_delete"
# target="_blank" rel="noopener no referrer">Delete model definition</a>
# + language="bash" active=""
#
# curl -sk -X DELETE \
# --header "Authorization: Bearer $TOKEN" \
# --header "Content-Type: application/json" \
# "$DATAPLATFORM_URL/ml/v4/model_definitions/$MODEL_DEFINITION_ID?space_id=$SPACE_ID&version=2020-08-01"
# -
# <a id="summary"></a>
# ## 7. Summary and next steps
#
# You successfully completed this notebook!.
#
# You learned how to use `cURL` calls to store, deploy and score a Tensorfloow Deep Learning model in WML.
#
# ### Authors
#
# **<NAME>**, Intern in Watson Machine Learning at IBM
# Copyright © 2020, 2021 IBM. This notebook and its source code are released under the terms of the MIT License.
| cpd3.5/notebooks/rest_api/curl/experiments/deep_learning/Use Tensorflow to recognize hand-written digits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Customizing QuickPlot
# This notebook shows how to customize PyBaMM's `QuickPlot`, using matplotlib's [style sheets and rcParams](https://matplotlib.org/stable/tutorials/introductory/customizing.html)
# First we define and solve the models
# +
# %pip install pybamm -q
import pybamm
models = [pybamm.lithium_ion.SPM(), pybamm.lithium_ion.SPMe(), pybamm.lithium_ion.DFN()]
sims = []
for model in models:
sim = pybamm.Simulation(model)
sim.solve([0, 3600])
sims.append(sim)
# -
# Call the default plots
pybamm.dynamic_plot(sims);
# ## Using style sheets
# The easiest way to customize style is to use one of matplotlib's available style sheets
import matplotlib.pyplot as plt
plt.style.available
# For example we can use the ggplot style from R. In this case, the title fonts are quite large, so we reduce the number of words in a title before a line break
plt.style.use("ggplot")
pybamm.settings.max_words_in_line = 3
pybamm.dynamic_plot(sims);
# To reset to pybamm defaults do
plt.style.use(["default", pybamm.default_plot_style])
pybamm.settings.max_words_in_line = 4
pybamm.dynamic_plot(sims);
# Another good set of style sheets for scientific plots is available by pip installing the [SciencePlots](https://github.com/garrettj403/SciencePlots) package
# ## Further customization using rcParams
# Sometimes we want further customization of a style, without needing to edit the style sheets. For example, we can update the font sizes and plot again.
#
# To change the line colors, we use `cycler`
# +
import matplotlib as mpl
from cycler import cycler
mpl.rcParams["axes.labelsize"] = 12
mpl.rcParams["axes.titlesize"] = 12
mpl.rcParams["xtick.labelsize"] = 12
mpl.rcParams["ytick.labelsize"] = 12
mpl.rcParams["legend.fontsize"] = 12
mpl.rcParams["axes.prop_cycle"] = cycler('color', ["k", "g", "c"])
pybamm.dynamic_plot(sims);
# -
# ## Very fine customization
# Some customization of the `QuickPlot` object is possible by passing arguments - see the [docs](https://pybamm.readthedocs.io/en/latest/source/plotting/quick_plot.html) for details
#
# We can also further control the plot by calling `plot.fig` after the figure has been created, and editing the matplotlib objects. For example, here we move the titles to the ylabel, and move the legend.
# +
plt.style.use(["default", pybamm.default_plot_style])
pybamm.settings.max_words_in_line = 4
plot = pybamm.QuickPlot(sims, figsize=(14,7))
plot.plot(0.5); # time in hours
# Move title to ylabel
for ax in plot.fig.axes:
title = ax.get_title()
ax.set_title("")
ax.set_ylabel(title)
# Remove old legend and add a new one in the bottom
leg = plot.fig.get_children()[-1]
leg.set_visible(False)
plot.fig.legend(plot.labels, loc="lower center", ncol=len(plot.labels), fontsize=11)
# Adjust layout
plot.gridspec.tight_layout(plot.fig, rect=[0, 0.04, 1, 1])
# -
# The figure can then be saved using `plot.fig.savefig`
# ## References
#
# The relevant papers for this notebook are:
pybamm.print_citations()
| examples/notebooks/customize-quick-plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## DATE TIME FUNTIONALITY IN PANDAS
import pandas as pd
import numpy as np
# ### Timestamp
# represent single point stamp and associate values with points in time
pd.Timestamp('24/04/2019 9:00PM')
# ### Period
#
# Suppose we weren't interested in a specific point in time, and instead wanted a span of time. This is where Period comes into play.
#
# Period represents a single time span, such as a specific day or month.
pd.Period('04/2019')
pd.Period('05/06/2019')
# ### DateTime Index eg
t1=pd.Series(list('abc'), [pd.Timestamp('2019-06-01'),pd.Timestamp('2019-06-02'),pd.Timestamp('2019-06-03')])
t1
type(t1.index)
# ### Period Index
t2=pd.Series(list('def'),[pd.Period('2019-09'),pd.Period('2019-10'),pd.Period('2019-11')])
t2
type(t2)
# ### Converting to DateTime
d1=['2 June 2013','Aug 29, 2014','2015-06-14','1/12/16']
ta3=pd.DataFrame(np.random.randint(10,100,(4,2)), index=d1, columns=list('ab'))
ta3
# converting index list into date time index
ta3.index=pd.to_datetime(ta3.index)
ta3
# ### converting DateTime to European format
pd.to_datetime('4.7.12', dayfirst=True)
# ### TimeDeltas
# time deltas are differences in time
pd.Timestamp('12-11-2017')-pd.Timestamp('12-15-2017')
# adding an offset of 12Days and 3 hours
pd.Timestamp('12-11-2017')+pd.Timedelta('12D 3H')
# ### Working with Dates in DataFrame
# #### GENERATING dates using date_range function
dates=pd.date_range('10-01-2019',periods=9,freq='2W-SUN')
dates
df=pd.DataFrame({'count1':100+np.random.randint(-5,10,9).cumsum(),
'count2':120+np.random.randint(-5,10,9) },index=dates)
np.random.randint(-5,10,9)
np.random.randint(-5,10,9).cumsum()
df
df.index.weekday_name
df.diff()
df.resample('M').mean()
df['2019']
df['2020-01':]
df.asfreq('W',method='ffill')
# +
# df.asfreq?
# -
import matplotlib.pyplot as plt
# %matplotlib inline
df.plot()
| pandas/week3_DateTime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import time
from numpy import *
from scipy import io
import matplotlib.pyplot as plt
import torch
from torch.optim.lr_scheduler import ReduceLROnPlateau
import sys
sys.path.append("../")
from spline import splineBasis
from ado import *
# -
# # make noisy and sparse data
# +
sim_measurement = io.loadmat('../systems/Double pendulum/data_ode113.mat')['x'][:2001, :]
t_m_all = np.linspace(0, 5, 2001)
x_sample = sim_measurement[:, :2]
w_sample = sim_measurement[:, 2:4]
x_sample_shift = x_sample.copy()
for i in range(x_sample_shift.shape[0]):
if abs(x_sample_shift[i, 0]) >= np.pi:
x_sample_shift[i:, 0] -= 2 * np.pi * np.sign(x_sample_shift[i, 0])
if abs(x_sample_shift[i, 1]) >= np.pi:
x_sample_shift[i:, 1] -= 2 * np.pi * np.sign(x_sample_shift[i, 1])
n_ratio = 0.05 # noise ratio
sub_ratio = 1 # downsample ratio
np.random.seed(0)
sub_idx = sorted(np.random.choice(len(t_m_all), int(len(t_m_all) * sub_ratio), replace=False))
t_m = t_m_all[sub_idx]
coordx1_true = 9.1 * np.sin(x_sample[:, 0])
coordy1_true = -9.1 * np.cos(x_sample[:, 0])
coordx2_true = coordx1_true + 7 * np.sin(x_sample[:, 1])
coordy2_true = coordy1_true - 7 * np.cos(x_sample[:, 1])
np.random.seed(0)
coordx1_n = np.random.normal(0,1,len(t_m_all))
coordx1_n = coordx1_n / np.std(coordx1_n)
coordx1_noise = (coordx1_true + n_ratio * np.std(coordx1_true) * coordx1_n)[sub_idx]
coordy1_n = np.random.normal(0,1,len(t_m_all))
coordy1_n = coordy1_n / np.std(coordy1_n)
coordy1_noise = (coordy1_true + n_ratio * np.std(coordy1_true) * coordy1_n)[sub_idx]
coordx2_n = np.random.normal(0,1,len(t_m_all))
coordx2_n = coordx2_n / np.std(coordx2_n)
coordx2_noise = (coordx2_true + n_ratio * np.std(coordx2_true) * coordx2_n)[sub_idx]
coordy2_n = np.random.normal(0,1,len(t_m_all))
coordy2_n = coordy2_n / np.std(coordy2_n)
coordy2_noise = (coordy2_true + n_ratio * np.std(coordy2_true) * coordy2_n)[sub_idx]
x_noise_shift = np.zeros([len(t_m), 2])
for i in range(len(t_m)):
x_noise_shift[i, 0] = math.atan2(coordy1_noise[i], coordx1_noise[i]) + 0.5*np.pi
x_noise_shift[i, 1] = math.atan2(coordy2_noise[i] - coordy1_noise[i],
coordx2_noise[i] - coordx1_noise[i]) + 0.5*np.pi
for i in range(x_noise_shift.shape[0]):
for j in range(x_noise_shift.shape[1]):
if x_noise_shift[i, j] > np.pi:
x_noise_shift[i, j] = x_noise_shift[i, j] - 2*np.pi
x_noise = x_noise_shift.copy()
for i in range(1, x_noise_shift.shape[0]):
if abs(x_noise_shift[i, 0] - x_noise_shift[i-1, 0]) >= np.pi:
x_noise[i:, 0] = x_noise[i:, 0] + 2 * np.pi * np.sign(x_noise_shift[i-1, 0] - x_noise_shift[i, 0])
if abs(x_noise_shift[i, 1] - x_noise_shift[i-1, 1]) >= np.pi:
x_noise[i:, 1] = x_noise[i:, 1] + 2 * np.pi * np.sign(x_noise_shift[i-1, 1] - x_noise_shift[i, 1])
fig = plt.figure(figsize=(16, 6))
ax = fig.add_subplot(211)
ax.set_title('$\\theta$ with noise', fontsize=25, pad=20)
ax.scatter(t_m_all, x_sample[:, 0], c='r', s=1.5)
ax.scatter(t_m, x_noise[:, 0], c='b', s=1.5)
ax.axes.xaxis.set_visible(False)
ax.set_ylabel('$\\theta_1$', fontsize=18)
ax = fig.add_subplot(212)
ax.scatter(t_m_all, x_sample[:, 1], c='r', s=1.5)
ax.scatter(t_m, x_noise[:, 1], c='b', s=1.5)
ax.set_ylabel('$\\theta_2$', fontsize=18)
ax.set_xlabel('time', fontsize=18)
plt.subplots_adjust(wspace=0.05, hspace=0.05)
plt.show()
# -
# # Baseline: pySINDy
# +
from pysindy import SINDy
from pysindy.feature_library import CustomLibrary
from pysindy.differentiation import SmoothedFiniteDifference
sfd = SmoothedFiniteDifference(smoother_kws={'window_length': 5})
w_sfd = sfd._differentiate(x_noise, t_m)
dw_sfd = sfd._differentiate(w_sfd, t_m)
self_library = ['dw1*np.cos(x1-x2)', 'dw2*np.cos(x1-x2)', 'w1**2*np.sin(x1-x2)', 'w2**2*np.sin(x1-x2)',
'w1**2*np.sin(x1)', 'w2**2*np.sin(x1)', 'w1**2*np.sin(x2)', 'w2**2*np.sin(x2)',
'w1*np.sin(x1-x2)', 'w2*np.sin(x1-x2)', 'w1*np.sin(x1)', 'w2*np.sin(x1)', 'w1*np.sin(x2)', 'w2*np.sin(x2)',
'np.sin(x1)', 'np.sin(x2)', 'np.sin(x1-x2)']
num_term = len(self_library)
t_m_len = len(t_m)
from pysindy import SINDy
from pysindy.feature_library import CustomLibrary
x1 = x_noise[:, 0]
x2 = x_noise[:, 1]
w1 = w_sfd[:, 0]
w2 = w_sfd[:, 1]
dw1 = dw_sfd[:, 0]
dw2 = dw_sfd[:, 1]
x_sindy = np.vstack([x1, x2, w1, w2, dw1, dw2]).T
functions = [lambda x1,x2,w1,w2,dw1,dw2: dw1*np.cos(x1-x2),
lambda x1,x2,w1,w2,dw1,dw2: dw2*np.cos(x1-x2),
lambda x1,x2,w1,w2,dw1,dw2: w1**2*np.sin(x1-x2),
lambda x1,x2,w1,w2,dw1,dw2: w2**2*np.sin(x1-x2),
lambda x1,x2,w1,w2,dw1,dw2: w1**2*np.sin(x1),
lambda x1,x2,w1,w2,dw1,dw2: w2**2*np.sin(x1),
lambda x1,x2,w1,w2,dw1,dw2: w1**2*np.sin(x2),
lambda x1,x2,w1,w2,dw1,dw2: w2**2*np.sin(x2),
lambda x1,x2,w1,w2,dw1,dw2: w1*np.sin(x1-x2),
lambda x1,x2,w1,w2,dw1,dw2: w2*np.sin(x1-x2),
lambda x1,x2,w1,w2,dw1,dw2: w1*np.sin(x1),
lambda x1,x2,w1,w2,dw1,dw2: w2*np.sin(x1),
lambda x1,x2,w1,w2,dw1,dw2: w1*np.sin(x2),
lambda x1,x2,w1,w2,dw1,dw2: w2*np.sin(x2),
lambda x1,x2,w1,w2,dw1,dw2: np.sin(x1),
lambda x1,x2,w1,w2,dw1,dw2: np.sin(x2),
lambda x1,x2,w1,w2,dw1,dw2: np.sin(x1-x2)]
lib_custom = CustomLibrary(library_functions=functions)
lib_custom.fit(x_sindy)
model = SINDy(feature_names=['x1', 'x2', 'w1', 'w2', 'dw1', 'dw2'], feature_library=lib_custom)
model.fit(x_sindy, t_m[1]-t_m[0])
def replace(eq):
f1 = lib_custom.get_feature_names(input_features=['x1', 'x2', 'w1', 'w2', 'dw1', 'dw2'])
f2 = self_library
for i,f in enumerate(f1):
eq = eq.replace(f, f2[i])
return eq
sindy_pred = [replace(x) for x in model.equations(3)[2:4]]
sindy_pred = [x.replace(' + -', '-') for x in sindy_pred]
sindy_pred = [x.replace(' + ', '+') for x in sindy_pred]
sindy_pred = [x.replace(' ', '*') for x in sindy_pred]
sindy_pred = [x.replace('np.', '') for x in sindy_pred]
sindy_pred = [x.replace('**', '^') for x in sindy_pred]
sindy_pred
# -
# # Physics-informed Spline Learning
# build cubic B-spline model
# +
end_t = 5
num_control = 100*end_t+1
num_c = 1000*end_t+1
t = np.linspace(0, end_t, num_control)
knots = np.array([0,0,0] + list(t) + [end_t,end_t,end_t])
t_c = np.array(sorted(list(t_m) + list(np.random.rand(num_c-len(t_m))*end_t)))
basis = splineBasis(knots, t_m, t_c)
basis_m, basis_dt_m = basis.get_measurement()
basis_c, basis_dt_c = basis.get_collocation()
# -
# pre-pre-tuning: initialization
# +
# setting device on GPU if available, else CPU
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print('Using device:', device)
print()
t_m_len = basis_m.shape[0]
num_control = basis_m.shape[1]
# convert the basis matrices to tensors
N = torch.Tensor(basis_m).to(device)
N_dt = torch.Tensor(basis_dt_m).to(device)
# define control points (one column for each variable)
P_theta = torch.autograd.Variable(torch.rand(num_control, 2).to(device), requires_grad=True)
# convert the measurement data into tensor
mea_theta = torch.Tensor(x_noise).to(device)
def loss_pre(P, mea):
return (torch.norm(torch.matmul(N, P) - mea, p=2) ** 2) / t_m_len
# define learning rate and optimizer
learning_rate = 0.05
optimizer = torch.optim.Adam([P_theta], lr=learning_rate)
# set a schedule for learning rate decreasing
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=1000, min_lr=0.00001)
# set flag for early stopping
min_loss = 10000
epochs_no_improve = 0
start_time = time.time()
for t in range(20000):
optimizer.zero_grad()
loss = loss_pre(P_theta, mea_theta)
loss.backward()
optimizer.step()
scheduler.step(loss)
if loss.item() >= min_loss:
epochs_no_improve += 1
else:
min_loss = loss.item()
epochs_no_improve = 0
# early stopping criteria: learning rate reaches the minimum
# value and there are no improvement in recent 50 epoches
if epochs_no_improve == 100 and optimizer.param_groups[0]['lr'] == 0.00001:
print("Early stopping!")
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
loss_pretuning = loss.item()
print("--- %s seconds ---" % (time.time() - start_time))
break
# print training information every 5000 epoches
if t % 5000 == 0:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
# stop training in the 100,000 epoch
if t == 19999:
print("20,000 epochs completed!")
loss_pretuning = loss.item()
print("--- %s seconds ---" % (time.time() - start_time))
x_dot = np.matmul(basis_dt_m, P_theta.cpu().detach().numpy())
mea_omega = torch.Tensor(x_dot).to(device)
P_omega = torch.autograd.Variable(torch.rand(num_control, 2).to(device), requires_grad=True)
# define learning rate and optimizer
learning_rate = 0.05
optimizer = torch.optim.Adam([P_omega], lr=learning_rate)
# set a schedule for learning rate decreasing
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=1000, min_lr=0.00001)
# set flag for early stopping
min_loss = 10000
epochs_no_improve = 0
start_time = time.time()
for t in range(20000):
optimizer.zero_grad()
loss = loss_pre(P_omega, mea_omega)
loss.backward()
optimizer.step()
scheduler.step(loss)
if loss.item() >= min_loss:
epochs_no_improve += 1
else:
min_loss = loss.item()
epochs_no_improve = 0
# early stopping criteria: learning rate reaches the minimum
# value and there are no improvement in recent 50 epoches
if epochs_no_improve == 100 and optimizer.param_groups[0]['lr'] == 0.00001:
print("Early stopping!")
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
loss_pretuning = loss.item()
print("--- %s seconds ---" % (time.time() - start_time))
break
# print training information every 5000 epoches
if t % 5000 == 0:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
# stop training in the 100,000 epoch
if t == 19999:
print("20,000 epochs completed!")
loss_pretuning = loss.item()
print("--- %s seconds ---" % (time.time() - start_time))
# -
# pre-tuning
# +
# setting device on GPU if available, else CPU
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print('Using device:', device)
print()
t_m_len = basis_m.shape[0]
t_c_len = basis_c.shape[0]
num_control = basis_m.shape[1]
# convert the basis matrices to tensors
N = torch.Tensor(basis_m).to(device)
N_c = torch.Tensor(basis_c).to(device)
N_dt = torch.Tensor(basis_dt_c).to(device)
# N_dt = torch.Tensor(basis_dt_m).to(device)
# convert the measurement data into tensor
x_sample = x_noise
measurement = torch.Tensor(x_sample).to(device)
cx1_true = torch.Tensor(coordx1_noise).to(device)
cy1_true = torch.Tensor(coordy1_noise).to(device)
cx2_true = torch.Tensor(coordx2_noise).to(device)
cy2_true = torch.Tensor(coordy2_noise).to(device)
self_library = ['dw1*cos(x1-x2)', 'dw2*cos(x1-x2)', 'w1**2*sin(x1-x2)', 'w2**2*sin(x1-x2)',
'w1**2*sin(x1)', 'w2**2*sin(x1)', 'w1**2*sin(x2)', 'w2**2*sin(x2)',
'w1*sin(x1-x2)', 'w2*sin(x1-x2)', 'w1*sin(x1)', 'w2*sin(x1)', 'w1*sin(x2)', 'w2*sin(x2)',
'sin(x1)', 'sin(x2)', 'sin(x1-x2)']
function_w1 = ''
function_w2 = ''
num_term = len(self_library)
for i in range(0, num_term):
term = self_library[i]
function_w1 += ('+cx'+str(i)+'*'+term)
function_w2 += ('+cy'+str(i)+'*'+term)
function_w1 = function_w1.replace('sin', 'torch.sin').replace('cos', 'torch.cos')[1:]
function_w2 = function_w2.replace('sin', 'torch.sin').replace('cos', 'torch.cos')[1:]
# +
# define control points (one column for each variable)
# P = torch.autograd.Variable(torch.rand(num_control, 4).to(device), requires_grad=True)
P_init = torch.cat((P_theta, P_omega), 1).cpu().detach()
## define variables Gamma and auxiliary variables Y and Z
P = torch.autograd.Variable(P_init.to(device), requires_grad=True)
################################################
########## use a full term library #############
################################################
for i in range(num_term): globals()['cx'+str(i)] = torch.autograd.Variable(torch.rand(1).to(device), requires_grad=True)
for i in range(num_term): globals()['cy'+str(i)] = torch.autograd.Variable(torch.rand(1).to(device), requires_grad=True)
coef_lst = [globals()['cx'+str(i)] for i in range(num_term)] + \
[globals()['cy'+str(i)] for i in range(num_term)]
################################################
################################################
loss1_his = []
loss2_his = []
loss3_his = []
# physics loss (denominator values known, calculate in the measurement domain)
def loss_phy(P):
x1 = torch.matmul(N_c, P[:, 0])
x2 = torch.matmul(N_c, P[:, 1])
w1 = torch.matmul(N_c, P[:, 2])
w2 = torch.matmul(N_c, P[:, 3])
dw1 = torch.matmul(N_dt, P[:, 2])
dw2 = torch.matmul(N_dt, P[:, 3])
return (1000 * torch.norm(torch.matmul(N_dt, P[:, 0]) - w1, p=2) ** 2 +
1000 * torch.norm(torch.matmul(N_dt, P[:, 1]) - w2, p=2) ** 2 +
torch.norm(torch.matmul(N_dt, P[:, 2]) - eval(function_w1), p=2) ** 2 +
torch.norm(torch.matmul(N_dt, P[:, 3]) - eval(function_w2), p=2) ** 2) / t_c_len
def loss_coord(P):
x1 = torch.matmul(N, P[:, 0])
x2 = torch.matmul(N, P[:, 1])
cx1_pred = 0 + 9.1*torch.sin(x1)
cy1_pred = 0 - 9.1*torch.cos(x1)
cx2_pred = cx1_pred + 7*torch.sin(x2)
cy2_pred = cy1_pred - 7*torch.cos(x2)
return (torch.norm(cx1_pred - cx1_true, p=2) ** 2 + torch.norm(cy1_pred - cy1_true, p=2) ** 2 +
torch.norm(cx2_pred - cx2_true, p=2) ** 2 + torch.norm(cy2_pred - cy2_true, p=2) ** 2) / t_m_len
# data loss + physics loss
def loss_total(P):
loss_p = loss_phy(P)
loss_c = loss_coord(P)
loss_d = (torch.norm(torch.matmul(N, P[:,0]) - measurement[:,0], p=2) ** 2 +
torch.norm(torch.matmul(N, P[:,1]) - measurement[:,1], p=2) ** 2) / t_m_len
loss1_his.append(float(loss_d.cpu().detach().numpy()))
loss2_his.append(float(loss_c.cpu().detach().numpy()))
loss3_his.append(float(loss_p.cpu().detach().numpy()))
return loss_d + loss_c + 0.001*loss_p
# define learning rate and optimizer
learning_rate = 0.01
optimizer = torch.optim.Adamax([P] + coef_lst, lr=learning_rate)
# set a schedule for learning rate decreasing
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=1000, min_lr=0.00001)
# set flag for early stopping
min_loss = 10000
epochs_no_improve = 0
start_time = time.time()
for t in range(200000):
optimizer.zero_grad()
loss = loss_total(P)
loss.backward()
optimizer.step()
scheduler.step(loss)
if loss.item() >= min_loss:
epochs_no_improve += 1
else:
min_loss = loss.item()
epochs_no_improve = 0
# early stopping criteria: learning rate reaches the minimum
# value and there are no improvement in recent 50 epoches
if epochs_no_improve == 100 and optimizer.param_groups[0]['lr'] == 0.00001:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
print("Early stopping!")
loss_pretuning = loss.item()
print("--- %s seconds ---" % (time.time() - start_time))
break
# print training information every 5000 epoches
if t % 5000 == 0:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
# stop training in the 100,000 epoch
if t == 199999:
print("200,000 epochs completed!")
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
loss_pretuning = loss.item()
print("--- %s seconds ---" % (time.time() - start_time))
# +
loss_pretuning = loss.item()
# prepare a copy of pre-tuned control points values
P_pre = P.cpu().detach().numpy().copy()
# convert the pre-tuned equation parameters in matrix form
lambda_raw = np.zeros([num_term, 2])
function_dw1 = function_dw2 = ''
for i in range(0, num_term):
term = self_library[i]
lambda_raw[i, 0] = globals()['cx'+str(i)].cpu().detach().numpy()[0]
lambda_raw[i, 1] = globals()['cy'+str(i)].cpu().detach().numpy()[0]
function_dw1 += (' + '+str(np.round(globals()['cx'+str(i)].cpu().detach().numpy()[0], 4))+'*'+term)
function_dw2 += (' + '+str(np.round(globals()['cy'+str(i)].cpu().detach().numpy()[0], 4))+'*'+term)
function_dw1 = function_dw1[3:].replace('+ -', '- ')
function_dw2 = function_dw2[3:].replace('+ -', '- ')
# print pre-tuned equations
print('dw2_pre-tune =', function_dw1)
print('dw1_pre-tune =', function_dw2)
# -
# ADO - sparsity tuning
# +
loss_HY = []
loss_HY_min = 1000000
terms_HY = [num_term*2]
A_raw = lambda_raw.copy()
A_raw_HY = A_raw.copy()
P_HY_np = P_pre.copy()
P_HY = torch.autograd.Variable(torch.Tensor(P_HY_np).to(device), requires_grad=True)
diminish_coef = True
num_terms = np.count_nonzero(A_raw)
d_tol = 0.01
lam = 1e-6 # ridge regularizer
eta = 0.1 # l-0 penalty ratio
tol_best = [0, 0]
start_time = time.time()
itr = 0
while diminish_coef or itr < 5:
print('itr:' + str(itr+1))
#############################################################################
################ ADO part1: refined training of Parameters #################
print('Training parameters (STRidge):')
x1 = torch.matmul(N_c, P_HY[:, 0]).cpu().detach().numpy()
x2 = torch.matmul(N_c, P_HY[:, 1]).cpu().detach().numpy()
w1 = torch.matmul(N_c, P_HY[:, 2]).cpu().detach().numpy()
w2 = torch.matmul(N_c, P_HY[:, 3]).cpu().detach().numpy()
dw1 = torch.matmul(N_dt, P_HY[:, 2]).cpu().detach().numpy()
dw2 = torch.matmul(N_dt, P_HY[:, 3]).cpu().detach().numpy()
phi = np.zeros([t_c_len, num_term])
for i in range(num_term):
phi[:, i] = eval(self_library[i])
A_raw[:, 0], tol_best[0] = TrainSTRidge(phi, dw1, lam, eta, d_tol, maxit = 500)
A_raw[:, 1], tol_best[1] = TrainSTRidge(phi, dw2, lam, eta, d_tol, maxit = 500)
print('best tolerance threshold is', tol_best)
print('prune number of terms to', np.count_nonzero(A_raw))
print()
function_w1 = function_w2 = ''
sparse_c_lst = []
for i in range(0, num_term):
term = self_library[i]
if A_raw[i, 0] != 0:
function_w1 += ('+cx'+str(i)+'*'+term)
sparse_c_lst.append(globals()['cx'+str(i)])
if A_raw[i, 1] != 0:
function_w2 += ('+cy'+str(i)+'*'+term)
sparse_c_lst.append(globals()['cy'+str(i)])
function_w1 = function_w1.replace('sin', 'torch.sin').replace('cos', 'torch.cos')[1:]
function_w2 = function_w2.replace('sin', 'torch.sin').replace('cos', 'torch.cos')[1:]
#################################################################
################ ADO part2: Brute Force traing #################
print('Spline Training:')
learning_rate = 0.01
optimizer = torch.optim.Adamax([P_HY] + sparse_c_lst, lr=learning_rate)
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=200, min_lr=0.00001)
min_loss = 10000
epochs_no_improve = 0
loss_his = []
for t in range(20000):
optimizer.zero_grad()
loss = loss_total(P_HY)
loss.backward()
scheduler.step(loss)
loss_his.append(loss.item())
optimizer.step()
if loss.item() >= min_loss:
epochs_no_improve += 1
else:
min_loss = loss.item()
epochs_no_improve = 0
if epochs_no_improve == 100 and optimizer.param_groups[0]['lr'] == 0.00001:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
print("Early stopping!")
break
if t % 5000 == 0:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
if t == 19999:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
print("20,000 epochs completed!")
loss_HY.append(loss.item() + eta*np.count_nonzero(A_raw))
terms_HY.append(np.count_nonzero(A_raw))
if loss_HY[-1] < loss_HY_min:
A_raw_HY = A_raw.copy()
loss_HY_min = loss_HY[-1]
if np.count_nonzero(A_raw) < num_terms:
num_terms = np.count_nonzero(A_raw)
else:
diminish_coef = False
itr += 1
print()
print('reach convergence of number of terms in governing equations!')
print("--- %s seconds ---" % (time.time() - start_time))
print()
print('final result :')
function_dw1 = function_dw2 = ''
for i in range(0, num_term):
term = self_library[i]
if A_raw_HY[i, 0] != 0: function_dw1 += (' + '+str(np.round(A_raw_HY[i, 0], 4))+'*'+term)
if A_raw_HY[i, 1] != 0: function_dw2 += (' + '+str(np.round(A_raw_HY[i, 1], 4))+'*'+term)
function_dw1 = function_dw1[3:].replace('+ -', '- ')
function_dw2 = function_dw2[3:].replace('+ -', '- ')
# print pre-tuned equations
print('dw1_pred =', function_dw1)
print('dw2_pred =', function_dw2)
# -
# post-tuning
# +
# choose a tuning result from above
A_raw_post = A_raw_HY.copy()
function_w1 = ''
function_w2 = ''
sparse_c_lst = []
for i in range(0, num_term):
term = self_library[i]
if A_raw_post[i, 0] != 0:
function_w1 += ('+cx'+str(i)+'*'+term)
sparse_c_lst.append(globals()['cx'+str(i)])
if A_raw_post[i, 1] != 0:
function_w2 += ('+cy'+str(i)+'*'+term)
sparse_c_lst.append(globals()['cy'+str(i)])
function_w1 = function_w1.replace('sin', 'torch.sin').replace('cos', 'torch.cos')[1:]
function_w2 = function_w2.replace('sin', 'torch.sin').replace('cos', 'torch.cos')[1:]
learning_rate = 0.05
optimizer = torch.optim.Adam([P] + sparse_c_lst, lr=learning_rate)
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=200, min_lr=0.0001)
min_loss = 10000
epochs_no_improve = 0
loss_his = []
start_time = time.time()
for t in range(50000):
optimizer.zero_grad()
loss = loss_total(P)
loss.backward()
scheduler.step(loss)
loss_his.append(loss.item())
if loss.item() >= min_loss:
epochs_no_improve += 1
else:
min_loss = loss.item()
epochs_no_improve = 0
if epochs_no_improve == 100 and optimizer.param_groups[0]['lr'] == 0.0001:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
print("Early stopping!")
print("--- %s seconds ---" % (time.time() - start_time))
break
if t % 1000 == 0:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
optimizer.step()
if t == 49999:
print('epoch :', t, 'loss :', loss.item(), 'lr :', optimizer.param_groups[0]['lr'])
print("50,000 epochs completed!")
print("--- %s seconds ---" % (time.time() - start_time))
print('----- final result -----')
function_dw1 = function_dw2 = ''
for i in range(0, num_term):
term = self_library[i]
if A_raw_post[i, 0] != 0: function_dw1 += (' + '+str(np.round(A_raw_post[i, 0], 5))+'*'+term)
if A_raw_post[i, 1] != 0: function_dw2 += (' + '+str(np.round(A_raw_post[i, 1], 5))+'*'+term)
function_dw1 = function_dw1[3:].replace('+ -', '- ')
function_dw2 = function_dw2[3:].replace('+ -', '- ')
# print pre-tuned equations
print('dw1_pred =', function_dw1)
print('dw2_pred =', function_dw2)
# +
x1 = sim_measurement[:, 0]
x2 = sim_measurement[:, 1]
w1 = sim_measurement[:, 2]
w2 = sim_measurement[:, 3]
true_value_w1 = '(m2*L1*w1**2*sin(2*x1-2*x2) + 2*m2*L2*w2**2*sin(x1-x2) + 2*g*m2*cos(x2)*sin(x1-x2) + 2*g*m1*sin(x1))' + \
'/ (-2*L1*(m2*sin(x1-x2)**2 + m1))'
true_value_w2 = '(m2*L2*w2**2*sin(2*x1-2*x2) + 2*(m1+m2)*L1*w1**2*sin(x1-x2) + 2*g*(m1+m2)*cos(x1)*sin(x1-x2))' + \
'/ (2*L2*(m2*sin(x1-x2)**2 + m1))'
m1 = 35
m2 = 10
L1 = 9.1
L2 = 7
g = 981
dw1_true = eval(true_value_w1)
dw2_true = eval(true_value_w2)
x1 = torch.matmul(N_c, P[:, 0]).cpu().detach().numpy()
x2 = torch.matmul(N_c, P[:, 1]).cpu().detach().numpy()
w1 = torch.matmul(N_c, P[:, 2]).cpu().detach().numpy()
w2 = torch.matmul(N_c, P[:, 3]).cpu().detach().numpy()
dw1 = torch.matmul(N_dt, P[:, 2]).cpu().detach().numpy()
dw2 = torch.matmul(N_dt, P[:, 3]).cpu().detach().numpy()
plt.figure(figsize=(16,3))
plt.plot(t_m, dw1_true, 'k-', c='r', lw=3, label='$d\omega_1/dt$ true')
plt.plot(t_c[:-1], eval(function_dw1)[:-1], '--', c='b', lw=2, label='$d\omega_1/dt$ eq')
plt.legend()
plt.show()
plt.figure(figsize=(16,3))
plt.plot(t_m, dw2_true, 'k-', c='r', lw=3, label='$d\omega_2/dt$ true')
plt.plot(t_c[:-1], eval(function_dw2)[:-1], '--', c='b', lw=2, label='$d\omega_2/dt$ eq')
plt.legend()
plt.show()
# -
| examples/double_pendulum_400HZ_5%noise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="cedtXySEYb28"
# <div class="alert alert-block alert-info"><b></b>
# <h1><center> <font color='black'> Homework 04 </font></center></h1>
# <h2><center> <font color='black'> Cross-Selling/ Up-selling & Recommendation System</font></center></h2>
# <h2><center> <font color='black'> MTAT.03.319 - Business Data Analytics</font></center></h2>
# <h2><center> <font color='black'> University of Tartu - Spring 2021</font></center></h2>
# </div>
# + [markdown] id="B-pvZUeIYb3G"
# # Homework instructions
#
# - Please provide the names and student IDs of the team-members (Maximum 2 person) in the field "Team mates" below. If you are not working in a team please insert only your name and student ID.
#
# - The accepted submission formats are Colab links or .ipynb files. If you are submitting Colab links please make sure that the privacy settings for the file is public so we can access your code.
#
# - The submission will automatically close on <font color='red'>**18 April at 23:59**</font>, so please make sure to submit before the deadline.
#
# - ONLY one of the teammates should submit the homework. We will grade the homework and the marks and feedback is applied for both the team members. So please communicate with your team member about marks and feedback if you are submit the homework.
#
# - If a question is not clear, please ask us in Moodle ONLY.
#
# - After you have finished solving the Homework, please restart the Kernel and run all the cells to check if there is any persisting issues.
#
# - Plagiarism is <font color='red'>**PROHIBITED**</font>. Any form of plagiarism will be dealt according to the university policy (https://www.ut.ee/en/current-students/academic-fraud).
#
# - Please <font color='red'>do not change</font> the template of this notebook file. You can download the .ipynb file and work on that.
#
# + [markdown] id="9OWlFadiYb3I"
# **<h2><font color='red'>Team mates:</font></h2>**
#
#
# **<font color='red'>Name: <NAME></font>  <font color='red'>Student ID: C09505</font>**
#
#
# **<font color='red'>Name: <NAME></font>  <font color='red'>Student ID: C07851</font>**
# + [markdown] id="pL7tscuuAtWp"
# ### The homework is divided into four sections and the points are distributed as below:
# <pre>
# - Market Basket Analysis -> 2.0 points
# - Collaborative Filtering -> 3.5 points
# - Recommender Systems Evaluation -> 1.0 points
# - Neural Network -> 2.5 points
# _________________________________________________
# Total -> 9.0 points
# </pre>
# + [markdown] id="boFT1CkoYb3K"
# # 1. Market Basket Analysis (2 points)
# + [markdown] id="a3hBebgbYb3M"
# **1.1 Consider the following businesses and think about one case of cross selling and one case of up selling techniques they could use. This question is not restricted to only traditional, standard examples.(1 points)**
# + [markdown] id="fDlcP-zGGscx"
# ### <font color='red'> **I apologize for the inconvience but no matter what I do the text icon shows part of what I am writing so kindly click on the points [a, b, c, d] as you are editing them to see my full answer**</font>
# + [markdown] id="HxMUA01DYb3P"
# a. An OnlineTravel Agency like Booking.com or AirBnB
# + [markdown] id="RODzp7BPYb3T"
# <font color='red'> **Cross selling: I booked a room in a certain hotel and it offered collection of effors for Taxi booking from the airport with good prices.**</font>
#
# <font color='red'> **Up selling: I booked a room in a certain hotel and it shows that it's not refundable but if I instead pick another room with more features the food coupon will increase and there's no payment needed, I can pay while checking in. and free cancelation before 2 days of the reservation. The difference between the two of them are less than $70.**</font>
# + [markdown] id="Qbw_w9p1Yb3U"
# b. A software company which produces products related to cyber security like Norton, Kaspersky, Avast and similar ones.
# + [markdown] id="j0SyXnB6Yb3W"
# <font color='red'> **Cross selling: I wanted to purchase the basic package [Norton Anti-Virus] with $34.99, it shows me 2 other great packages [ Norton computer tune up] which helps my computer run like new again for $49.99 and the other one is [Norton family], which guarantee safe, secure connection for kids for $49.99.**</font>
#
# <font color='red'> **Up selling:[text is hidden kindly open the text] I wanted to purchase Norton package for $37.99 with %45 But the site recommended to instead purchase Norton 360 Premium Plus with 95% discount with 6 more features and only for $59.99**</font>
# + [markdown] id="7EUCv8TtYb3X"
# c. A company that sells cell phones
#
#
#
# + [markdown] id="NFHO-dI6Yb3Y"
# <font color='red'> **Cross selling: I added to the cart Iphone 11, and then down below the wesite shows adapters & headsets for Ipone 11 with vival colors**</font>
#
# <font color='red'> **Up selling: I clicked on the headsets icon to pick one with my Iphone 11, and I have selected one with the price of EarPods with 3.5 mm Headphone Plug for $19. The |site| showed| me |that |the |headset [Beats flex all day wireless] for |only $27.99**</font>
# + [markdown] id="_wnH4-lrYb3a"
# d. A supermarket like Konsum, Rimi, Maxima etc.
# + [markdown] id="I4CNtNYBYb3b"
# <font color='red'> **Cross selling: I added to the cart chicken and it shows spicies of chicken for a great teste, 20% discount on the Rice [1 Kg]**</font>
#
# <font color='red'> **Up selling: I added to the cart Tissue paper [8 pieces] to buy it for price 2.53 Euros, instead I found down below that if I took from different company Tissue paper [16 pieces] it would be with the price of 4.20 Euros.**</font>
# + [markdown] id="DLp7o0cdYb3c"
# **1.2 Let's suppose that our client is a retail company that has an online shop. They gave us a dataset about online sales of their products. The client wants to know which product bundles to promote. Find 5 association rules with the highest lift.**
# + id="b7HLlQ30Yb3e"
import pandas as pd
import numpy as np
df = pd.read_csv('https://raw.githubusercontent.com/RewanEmam/Customer-Segmentation-files/main/OnlineRetailPurchase.csv', header=0, sep = ',')
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="VWBRFwuUYb3l" outputId="03b8e1f1-99ea-43f5-a1ad-e56019281552"
df.head()
# + [markdown] id="kcjIimkHYb35"
# **1.3 Use describe function from pandas to get statistical information about the values in the dataframe.(0.2 points)**
# + id="RakInjZBY4Wu"
df.describe()
# + [markdown] id="J5a0X9dtYb4K"
# **1.4 Create a dataframe name as "Basket", where each row has an distintive value of InvoiceNo and each column has a distinctive Description. The cells in the table contain the count of each item (Description) mentioned in one invoice. For example basket.loc['536365','WHITE HANGING HEART T-LIGHT HOLDER'] has a value of 1 because the product with WHITE HANGING HEART T-LIGHT HOLDER was entered only once in the invoice 536365. (0.2 points)**
# + id="D4lUPlKAYb4L"
Basket = df[['InvoiceNo', 'Description']]
basket = Basket.drop_duplicates(subset = ['InvoiceNo', 'Description'],keep= 'last').reset_index(drop = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 526} id="IyRIzurWPrMl" outputId="22f7289f-acaa-4d76-cb00-9175d3f499c8"
basket = pd.get_dummies(basket['Description'])
basket
# + [markdown] id="3rwKSVg3Yb4d"
# **1.5 Some products are mentioned more than once in one invoice. You can check the maximum number for each column to verify. Modify your dataframe such that every cell which has a value higher than one will be replaced with 1. If the cell has the value 0 it will remain the same. (0.2 points)** <br>
# NB: If your implementation in 1.4 already takes care of this, please skip the question.
# + id="9BO17Wy1Yb4e"
#TODO
# + [markdown] id="KfWgocGTYb4k"
# **1.5 We do not need to spend time on calculating the association rules by ourselves as there already exists a package for python to do so, called mlxtend. We are going to use the mlxtend package to find frequent items bought together and then create some rules on what to recomend to a user based on what he/she/they have bought. We have given you the first part of the code which calculates the frequent items bought together. (0.2 points)**
# + id="rCw4ii7tYb4l"
# #!pip install mlxtend
from mlxtend.frequent_patterns import apriori
from mlxtend.frequent_patterns import association_rules
import mlxtend as ml
import math
# + id="nQBjILk5Yb4p"
#TODO
# + [markdown] id="GcF5RyYRYb4y"
# **Please read the documentation of the associaton rules function in mlextend [here](http://rasbt.github.io/mlxtend/user_guide/frequent_patterns/association_rules/) and complete the code so we get the 5 rules with the highest lift. Print those rules. For example if user bought product basket A then the algorithm recommends product basket B. (0.2 points)**
# + id="FLpV1FkKYb41"
rules = ... #TODO
for index, row in (rules.iloc[:5]).iterrows():
print("If the customer buys " + str(row['antecedents']))
print("")
print("The recommender recommends "+str(row['consequents']))
print("")
print("")
print("")
# + [markdown] id="kRqo0ek4Yb47"
# # 2. Collaborative filtering (3.5 points )
# + [markdown] id="_U1OvsCJYb48"
# We are going to use Books.csv dataset which contains ratings from Amazon website and the data has the following features:
#
# UserID: The ID of the users who read the books
#
# BookTitle: The title of the book
#
# Book-Rating: A rating given to the book in a scale from 0 to 10
#
# Below we are going to perform the same steps we did with movies dataset in the practice session
# + [markdown] id="_-wOm7yLYb49"
# **2.0 Load the dataset and take a look at the books titles. And pick a favorite book (any book).(0.1 points)**
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="Z_2CgjU6Yb4-" outputId="f7d694ea-a181-41ce-9b13-36bb4e395c1f"
df_book = pd.read_csv('https://raw.githubusercontent.com/RewanEmam/Customer-Segmentation-files/main/Books.csv', header=0, sep = ',', usecols=['UserID', 'Book-Rating', 'BookTitle'])
df_book.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 406} id="IBmxbi5zS2j_" outputId="d75fd654-f89b-470e-bb76-9c1e6552412a"
dfBook = df_book.drop_duplicates(subset = ['BookTitle', 'UserID'],keep= 'last').reset_index(drop = True)
dfBook
# + [markdown] id="Q_f2ywpLYb5J"
# **2.1 You have to apply KNN algorithm for collaborative filtering. As KNN algorithm does not accept strings, use a Label Encoder for BookTitle column.After that reshape the books matrix so that every column will be a UserID and every row a BookTitle. (0.45 points)**
# + colab={"base_uri": "https://localhost:8080/", "height": 593} id="-Gs_CAGKYb5K" outputId="cf29f200-b9aa-46d8-d0f2-928a16497bf3"
from sklearn import preprocessing
# label encounter
label = preprocessing.LabelEncoder()
dfBook['BookName'] = labelencoder.fit_transform(dfBook['BookTitle'])
# every column is userid
df_boo = dfBook.pivot(index = 'BookTitle', columns='UserID', values='Book-Rating').fillna(0)
df_boo.index.names = ['BookTitle']
df_boo.head()
# + [markdown] id="4RwLx90KYb5R"
# **2.2 Build a sparse matrix for books data and show it. (0.45 points)**
# + colab={"base_uri": "https://localhost:8080/"} id="uwVtesasYb5U" outputId="731e9ac4-9c97-4ec7-bd25-0dc17c1eaa00"
from scipy.sparse import csr_matrix
df_boo_sparse = csr_matrix(df_boo.values)
print(f"Sparse matrix:\n{df_boo_sparse}")
# + colab={"base_uri": "https://localhost:8080/"} id="_qDjXo8fezvk" outputId="51ec5b57-7054-47fb-eafa-fb0ae0cc252c"
# create mapper from book title to index
# book: index
book_to_idx = {
book: i for i, book in enumerate(list(dfBook.set_index('BookTitle').loc[df_boo.index].index))
}
book_to_idx
# + [markdown] id="PrKKbiRJYb5g"
# **2.3 Initialize and train two different KNN models (use cosine metric for similarity for both) but with different n_neighbours, 2 and 10. Recommend top 5 books based on your favourite one in both cases (1 points)**<br>
# NB: You are free to choose a favorite book (any book) based on which you have to recommend 5 books.
# + colab={"base_uri": "https://localhost:8080/"} id="zpqEOCFKAtWy" outputId="ab4540a2-c4ed-400e-feb3-f8cad16140da"
from sklearn.neighbors import NearestNeighbors
# define model: using cosine for similarity
model_knn_null = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=2, n_jobs=-1)
model_knn = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=10, n_jobs=-1)
# fit the model
print(model_knn.fit(df_boo_sparse))
print(model_knn_null.fit(df_boo_sparse))
# + colab={"base_uri": "https://localhost:8080/"} id="t4_7s1gozZO6" outputId="4a5a809d-10ca-437d-ce5e-7e4ac0a9e9f8"
# !pip install fuzzywuzzy
# + id="O7lfuKtV2Dbo"
# Import the required libraries:
import os
import time
import math
from fuzzywuzzy import fuzz
from sklearn.neighbors import NearestNeighbors
# + id="4XtAtZ63eWEb"
def fuzzy_matching(mapper, fav_book, verbose=True):
# Get match
match_tuple = []
for title, idx in mapper.items():
ratio = fuzz.ratio(BookTitle.lower(), fav_book.lower())
if ratio >= 500:
match_tuple.append((df_boo['BookTitle'], idx, ratio))
# Sort
match_tuple = sorted(match_tuple, key = lambda x: x[2])[::-1]
if not match_tuple:
print('Oops! No match is found')
return
if verbose:
print('Found possible matches in our database: {0}\n'.format([x[0] for x in match_tuple]))
return match_tuple[0][1]
def make_recommendation(model_knn, data, mapper, fav_book, n_recommendations):
# data = df_boo
model_knn.fit(data)
# get input book index
print('You have input book:', fav_book)
idx = fuzzy_matching(mapper, fav_book, verbose=True)
# Inference
print('Recommendation system start to make inference')
print('......\n')
distances, indices = model_knn.kneighbors(data[idx], n_neighbors=n_recommendations+1)
# Get list of raw idx of recommendations
raw_recommends = \
sorted(list(zip(indices.squeeze().tolist(), distances.squeeze().tolist())), key=lambda x: x[1])[:0:-1]
# get reverse mapper
reverse_mapper = {v: k for k, v in mapper.items()}
# print recommendation:
print('Recommendations for {}:'.format(fav_book))
for i, (idx, dist) in reversed(list(enumerate(raw_recommends))):
#j =i
print('{0}: {1}, with distance of {2}'.format(n_recommendations-i, reverse_mapper[idx], dist))
# + id="3rHFrCq96d30"
my_favorite = 'Matilda' # Matilda
make_recommendation(
model_knn=model_knn, # trained model (model)
data=df_boo_sparse, # sparse matrix (data)
fav_book=my_favorite, # fav_book
mapper=book_to_idx, # {book: index} (mapper)
n_recommendations=5)
# + id="7Z-tQZBj38hT"
data = df_boo_sparse
def fuzzy_matching(mapper, fav_book, verbose=True):
match_tuple = []
for title, idx in mapper.items():
ratio = fuzz.ratio(title.lower(), fav_book.lower())
if ratio >= 60:
match_tuple.append((title, idx, ratio))
match_tuple = sorted(match_tuple, key = lambda x: x[2])[::-1]
if not match_tuple:
print('Oops! No match is found')
return
if verbose:
print('Found possible matches in our database: {0}\n'.format([x[0] for x in match_tuple]))
return match_tuple[0][1]
def make_recommendation(model_knn_null, data, mapper, fav_book, n_recommendations):
# data = df_boo
model_knn_null.fit(data)
# get input book index
print('You have input book:', fav_book)
idx = fuzzy_matching(mapper, fav_book, verbose=True)
# Inference
print('Recommendation system start to make inference')
print('......\n')
distances, indices = model_knn_null.kneighbors(data[idx], n_neighbors=n_recommendations+1)
# Get list of raw idx of recommendations
raw_recommends = \
sorted(list(zip(indices.squeeze().tolist(), distances.squeeze().tolist())), key=lambda x: x[1])[:0:-1]
# get reverse mapper
reverse_mapper = {v: k for k, v in mapper.items()}
# print recommendation:
print('Recommendations for {}:'.format(fav_book))
for i, (idx, dist) in reversed(list(enumerate(raw_recommends))):
#j =i
print('{0}: {1}, with distance of {2}'.format(n_recommendations-i, reverse_mapper[idx], dist))
# + id="5O6AfDal_kFs"
my_favorite = 'Shadowland' # The Da Vinci Code
make_recommendation(
model_knn_null=model_knn_null, # trained model (model)
data= df_boo_sparse, # sparse matrix (data)
fav_book=my_favorite, # fav_book
mapper=book_to_idx, # {book: index} (mapper)
n_recommendations=5)
# + [markdown] id="WCJz_Do9Yb5q"
# **2.4 Discuss the results you received from both models. Which one worked better? (0.25 points)**
# + [markdown] id="2CdPc75QYb5r"
# <font color='red'> **Answer: Based on the result, I found the recommendation are quite similar to the choice I have selected. Whether I have selected Matilda-The davnci code- Shadowland, etc. Thanks to the main factors I have here: Model_knn function & mapper. They are factors of the main factors that the recommendations mechanism are absed on.**</font>
# + [markdown] id="G6T3K3VFYb5s"
# **2.5 Add a new user (with user “UserID” = 6293) in your data. Using the two trained models in task 2.3 suggest which books should this user read if his ratings are:**
#
# French Cuisine for All: 4
#
#
# <NAME> and the Sorcerer's Stone Movie Poster Book: 5
#
#
# El Perfume: Historia De UN Asesino/Perfume : The Story of a Murderer: 1
#
# **(1. 25 points)**
#
#
# + id="62_Jfn7cNYie"
# Edit my dataset a little bit:
features = ['UserID', 'BookTitle', 'Book-Rating']
# Get each row as a string
def combine_features(row):
return row['Book-Rating']+" "+row['UserID']+" "+row['BookTitle']
for feature in features:
dfBook[feature] = dfBook[feature].fillna('')
dfBook["combined_features"] = dfBook.apply(combine_features, axis=1)
# + id="o-EJOEy1Yb5t"
# In case model_knn case:
def get_title_from_index(index):
return df[df.index == index]["title"].values[0]
def get_index_from_title(title):
return dfBook[dfBook['BookTitle'] == title]["index"].values[0]
book_user_likes = "Shadowland"
book_index = get_index_from_title(book_user_likes)
similar_books = list(enumerate(cosine_sim[book_index]))
sorted_similar_books = sorted(similar_books,key=lambda x:x[1],reverse=True)[1:]
i=0
print("Top 5 similar movies to "+book_user_likes+" are:\n")
for element in sorted_similar_books:
print(get_title_from_index(element[0]))
i=i+1
if i>=5:
break
# + [markdown] id="VMyW4UlbYb5x"
# # 3. Recommender systems evaluation (1 points)
# + [markdown] id="EINSDAbXYb5y"
# We are going to compare different methods of recommender systems by their RMSE score. One useful package that has several recommender algorithms for Python is [Surprise](https://surprise.readthedocs.io/en/stable/getting_started.html). Below we have split the books dataset into training and test and used the KNNBasic algorithm to predict the ratings for the test set using surprise.
# + colab={"base_uri": "https://localhost:8080/"} id="unS3lDvaTAWa" outputId="f4b33a52-e4ea-438a-e274-63439e333b8d"
pip install surprise
# + colab={"base_uri": "https://localhost:8080/"} id="OoLm-EC1Yb5z" outputId="96b7938a-b6d2-4422-d10a-4bcf4d6cdf6f"
from surprise import accuracy
from surprise.model_selection import train_test_split
from surprise import Reader
from surprise import Dataset
from surprise import SVD
from surprise import NormalPredictor
from surprise import KNNBasic
# The reader is necessary for surprise to interpret the ratings
reader = Reader(rating_scale=(0, 10))
# This function loads data from a pandas dataframe into surprise dataset structure
# The columns should always be ordered like this
data = Dataset.load_from_df(dfBook[['UserID', 'BookTitle', 'Book-Rating']], reader)
# Split in trainset and testset
# No need to define the label y because for surprise the last column is always the rating
trainset, testset = train_test_split(data, test_size=.25, random_state=0 )
knn = KNNBasic()
knn.fit(trainset)
predictions = knn.test(testset)
print('KNN RMSE', accuracy.rmse(predictions))
# + [markdown] id="sdIaAghiYb53"
# **3.1 After taking a look at surprise documentation and the code above, follow the same steps as with KNN, and predict the ratings in test set using the NormalPredictor which predicts a random rating based on the distribution of the training set. Do the same for SVD which is a matrix factorization technique. For both of them report RMSE. (1 points)**
# + colab={"base_uri": "https://localhost:8080/"} id="VWcalcl4Yb56" outputId="719b118f-db13-47ea-aead-d4b7528c45f6"
#TODO: Normal predictor
# First Recall the libraries:
from surprise.model_selection import cross_validate
from surprise.model_selection import KFold
# We can now use this dataset as we please, e.g. calling cross_validate
cross_validate(NormalPredictor(), data, cv=2)
# + colab={"base_uri": "https://localhost:8080/"} id="hr29lRYSW8Bo" outputId="4f96ebd0-7495-4a9e-9106-3ba7830e5877"
#TODO: SVD
# define a cross-validation iterator
kf = KFold(n_splits=3)
algo = SVD()
for trainset, testset in kf.split(data):
# train and test algorithm.
algo.fit(trainset)
predictions = algo.test(testset)
# Compute and print Root Mean Squared Error
accuracy.rmse(predictions, verbose=True)
# + [markdown] id="F_DPXBrvXIwk"
# ### **Conclusion: RMSE for SVD is in range 4.2389 to 4.3355. Unlike the NormalPredictor that generates an array..**
# + [markdown] id="OjJgAOSRYb6A"
# # 4. Neural Networks (2.5 Points)
# + [markdown] id="h5TF1ePBYb6L"
# **4.1 We are now going to build a recommender system using Neural Networks. Being this dataset is really small in terms of features you might not see great improvements but it is a good starting point to learn. Please build one of the neural network architechtures as we did in practice session part 3. You can for example choose the one which had the following layers:**
# - 2 Embedding
# - 2 Reshape
# - 1 Concatenation
# - 1 Dense
#
# **Use the Neural Network you built to learn from the train data of part 3 of this homework. The column UserID should be used as input to your NN for the user embedding layer. For the books embedding layer we will use BookTitle column. Lastly, the ratings will be your target variable. Regarding the evaluation metric for the training phase use RMSE. To make your training fast you can use a batch size of 200 or above. (1.5 points)**
# + id="PbuvaC1eYb6Q"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
import warnings
from keras import backend
from keras.layers import Input, Embedding, Flatten, Dot, Dense,multiply, concatenate, Dropout, Reshape
from keras.models import Model, Sequential
from sklearn.model_selection import train_test_split
from keras.optimizers import Adam
#Method for RMSE calculation
def rmse(true_label, pred_label):
return #TODO: RMSE function
#TODO: Data preparation
df = pd.read_csv('https://raw.githubusercontent.com/RewanEmam/Customer-Segmentation-files/main/Books.csv',
header=0, sep = ',', usecols=['UserID', 'Book-Rating', 'BookTitle'])
#TODO: Model
def RecommenderV1(user_id, title, ratings):
user_id = Input(shape=(1,))
u = Embedding(user_id, ratings, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(user_id)
u = Reshape((n_factors,))(u)
#TODO: Embedding user id
title = Input(shape=(50,))
m = Embedding(title,ratings, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(title)
m = Reshape((n_factors,))(m)
x = Dot(axes=1)([u, m])
model = Model(inputs = (id_em, title_em), outputs = out)
model.compile(optimizer = 'Adam', loss = rmse, metrics = ['accuracy'])
#TODO: Train model
history = model.fit(x=X_train_array, y=y_train, batch_size=200, epochs=150,
verbose=1, validation_data=(X_test_array, y_test))
#TODO: pass data, batch_size=200, epochs=150)
# + [markdown] id="HsmPj7Wq1cyS"
# **4.2 Plot the RMSE values during the training phase, as well as the model loss. Report the best RMSE. Is it better than the RMSE from the models we built in Section 2 and 3 ? (0.5 points)**
# + id="JCJFqfDm1-HA"
from matplotlib import pyplot
#TODO
# + [markdown] id="LVLaC5K11-fN"
# **4.3 Use your trained model to recommend books for user with ID 6293. (0.5 points)**
# + id="NHQrNa35Jmjo"
#TODO
# + [markdown] id="zwpOi51caTUp"
# ## How long did it take you to solve the homework?
#
# * Please answer as precisely as you can. It does not affect your points or grade in any way. It is okay, if it took 0.5 hours or 24 hours. The collected information will be used to improve future homeworks.
#
# <font color='red'> **Answer: X hours**</font>
#
#
# ## What is the level of difficulty for this homework?
# you can put only number between $0:10$ ($0:$ easy, $10:$ difficult)
#
# <font color='red'> **Answer:**</font>
# + id="nJX9pZJRAtW3"
| Homework_04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.7 64-bit (conda)
# name: python3
# ---
import datetime
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from numba import jit
from scipy.spatial.distance import pdist, squareform
from scipy.optimize import root_scalar, curve_fit
from pynndescent import NNDescent
from scipy.sparse import csr_matrix
import pandas as pd
iris = pd.read_csv('https://gist.githubusercontent.com/curran/a08a1080b88344b0c8a7/raw/\
0e7a9b0a5d22642a06d3d5b9bcbad9890c8ee534/iris.csv')
def exp_k(dists, sigma):
return np.exp(- (dists - dists[0]) / sigma).sum()
def find_sigma(d, k, lower_bound=1e-6, upper_bound=1e6):
return root_scalar(
lambda s: exp_k(d, s) - np.log2(k),
bracket=(lower_bound, upper_bound)
).root
def p_ij_sym(x, k, verbose=False):
num_pts = x.shape[0]
if verbose:
print('Indexing')
index = NNDescent(x)
neighbors = np.empty((num_pts, k), dtype=np.int32)
p_ij = np.empty((num_pts, k))
for i, xi in enumerate(x):
if verbose:
print('Calculating probabilities: {cur}/{tot}'.format(
cur=i+1, tot=num_pts), end='\r')
nn, dists = index.query([xi], k+1)
sigma = find_sigma(dists[0, 1:], k)
neighbors[i] = nn[0, 1:]
p_ij[i] = np.exp(- (dists[0, 1:] - dists[0, 1]) / sigma)
row_indices = np.repeat(np.arange(num_pts), k)
p = csr_matrix((p_ij.ravel(), (row_indices, neighbors.ravel())))
return p + p.transpose() - (p.multiply(p.transpose()))
p = p_ij_sym(iris.to_numpy()[:,:4], 20, verbose=True)
pp = p.tocoo()
head = pp.row
tail = pp.col
num_per_sample = np.asarray(500 * pp.data, np.int32)
edges_to_exp, edges_from_exp = (
np.repeat(head, num_per_sample),
np.repeat(tail, num_per_sample),
)
shuffle_mask = np.random.permutation(range(len(edges_to_exp)))
edges_to_exp = edges_to_exp[shuffle_mask].astype(np.int32)
edges_from_exp = edges_from_exp[shuffle_mask].astype(np.int32)
dataset = torch.utils.data.TensorDataset(torch.tensor(edges_to_exp), torch.tensor(edges_from_exp))
dataloader = torch.utils.data.DataLoader(dataset, batch_size=200, shuffle=True)
for data in dataloader:
emb_to, emb_from = data
break
foo = torch.stack((tox, fromx)).T
emb_neg_to = torch.repeat_interleave(emb_to, 5)
repeat_neg = torch.repeat_interleave(emb_from, 5)
emb_neg_from = repeat_neg[torch.randperm(repeat_neg.shape[0])]
(pp.toarray()[emb_neg_from, emb_neg_to] == 0.).sum() / len(emb_neg_to)
plt.plot(pp.toarray()[torch.cat((emb_from, emb_neg_from)), torch.cat((emb_to, emb_neg_to))])
torch.cat((emb_from, emb_neg_from))
class NegSampledEdgeDataset(torch.utils.data.Dataset):
def __init__(self, p_ij, neg_sampling_rate=5):
self.p_ij = p_ij.tocoo()
self.weights = p_ij.data
self.neg_sampling_rate = neg_sampling_rate
def __len__(self):
return len(self.p_ij.data)
def __getitem__(self, idx):
# make nsr+1 copies of i
rows = torch.full(
(self.neg_sampling_rate + 1,),
self.p_ij.row[idx],
dtype=torch.long
)
#make one positive sample and nsr negative ones
cols = torch.randint(
pp.shape[0],
(self.neg_sampling_rate + 1,),
dtype=torch.long
)
cols[0] = self.p_ij.col[idx]
# make simplified p_ij (0 or 1)
p_simpl = torch.zeros(self.neg_sampling_rate + 1, dtype=torch.float32)
p_simpl[0] = 1
return rows, cols, p_simpl
pp = p.tocoo()
foo = NegSampledEdgeDataset(pp)
sampler = torch.utils.data.WeightedRandomSampler(foo.weights, num_samples=len(foo))
def collate_samples(batch):
return [torch.cat(samples) for samples in zip(*batch)]
loader = torch.utils.data.DataLoader(foo,
batch_size=100,
collate_fn=collate_samples,
sampler=sampler
)
for _ in range(30):
for data in loader:
continue
for data in loader:
bazoo = data
break
iris_tensor = torch.tensor(np.asarray(iris.to_numpy()[:,:4], dtype=np.float32))
a = iris_tensor[bazoo[0]]
b = iris_tensor[bazoo[1]]
torch.norm(a - b, dim=1).pow(2)
| numerical-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import scipy.io
import os, re
import claude.utils as cu
import claude.claudeflow.autoencoder as ae
import claude.claudeflow.helper as cfh
import claude.claudeflow.training as cft
# +
seed = 1337
tf.set_random_seed(seed)
np.random.seed(seed)
# Parameters
# Channel Parameters
chParam = cu.AttrDict()
chParam.M = 16
# Auto-Encoder Parameters
aeParam = cu.AttrDict()
aeParam.constellationDim = 2
aeParam.constellationOrder = chParam.M
aeParam.nLayers = 2
aeParam.nHidden = 32
aeParam.activation = tf.nn.selu
aeParam.dtype = tf.float32
# Training Parameters
trainingParam = cu.AttrDict()
trainingParam.sampleSize = 512*chParam.M # Increase for better results (especially if M>16)
trainingParam.batchSize = 32*chParam.M # Increase for better results (especially if M>16)
trainingParam.learningRate = 0.001
trainingParam.displayStep = 20
trainingParam.path = 'results_AWGN_noChannel'
trainingParam.filename = 'M{}'.format(chParam.M)
trainingParam.saveWeights = True
trainingParam.earlyStopping = 10
trainingParam.tx_iterations = 25
trainingParam.rx_iterations = 25
trainingParam.policy_sigma2 = 0.02
# TF constants
two = tf.constant(2,aeParam.dtype)
minusOne = tf.constant(-1,aeParam.dtype)
DIM = tf.constant(aeParam.constellationDim,aeParam.dtype)
PI = tf.constant(np.pi,aeParam.dtype)
# +
# Channel Graph
sigma2_noise = tf.constant(0.1,aeParam.dtype)
channel_in = tf.placeholder( aeParam.dtype, shape=(None, aeParam.constellationDim) )
noise = tf.sqrt( sigma2_noise )\
*tf.rsqrt(two)\
*tf.random_normal(shape=tf.shape(channel_in),dtype=aeParam.dtype)
channel_out = channel_in + noise
# -
# Tx Graph
X = tf.placeholder( aeParam.dtype, shape=(None, chParam.M) )
enc, enc_seed = ae.encoder(X, aeParam)
# +
# enc = ae.encoder(X,aeParam.hiddenUnits,aeParam.nLayers,aeParam.activation,nOutput=aeParam.dimension)
# enc_norm = cfh.IQ_norm( enc )
# Tx policy
policy_sigma2 = tf.placeholder( aeParam.dtype, shape=())
perturbation = tf.sqrt( policy_sigma2 ) * tf.rsqrt(two) * tf.random_normal(shape=tf.shape(enc),dtype=aeParam.dtype)
enc_pert = enc + perturbation
# -
# Rx Graph
Y = tf.placeholder( aeParam.dtype, shape=(None, aeParam.constellationDim) )
dec = ae.decoder(Y,aeParam)
per_ex_loss = tf.nn.softmax_cross_entropy_with_logits_v2(labels=X,logits=dec)
# Rx Loss
correct_prediction = tf.equal(tf.argmax(X,1), tf.argmax(dec,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, aeParam.dtype))
rx_loss = tf.reduce_mean(per_ex_loss)
rx_vars = [item for item in tf.global_variables() if 'decoder' in item.name]
rx_optimizer = tf.train.AdamOptimizer(learning_rate=trainingParam.learningRate).minimize(rx_loss, var_list=rx_vars)
metricsDict = {'xentropy':rx_loss, 'accuracy_metric':accuracy}
meanMetricOpsDict, updateOps, resetOps = cft.create_mean_metrics(metricsDict)
# +
# avg_loss, avg_loss_update, avg_loss_reset = cfh.create_reset_metric(tf.metrics.mean, 'loss_metric', rx_loss)
# avg_accuracy, avg_accuracy_update, avg_accuracy_reset = cfh.create_reset_metric(tf.metrics.mean, 'accuracy_metric', accuracy)
# +
# Tx Loss
tx_perturbed = tf.placeholder( aeParam.dtype, shape=(None,aeParam.constellationDim) )
tx_per_ex_loss = tf.placeholder( aeParam.dtype, shape=(None,) )
# batch_size = tf.constant(trainingParam.batchSize, aeParam.dtype)
batch_size = tf.placeholder( aeParam.dtype, shape=() )
policy = tf.log( tf.rsqrt( tf.pow(PI*policy_sigma2,DIM) ) * tf.exp( minusOne * tf.square(cfh.norm( tx_perturbed-enc )) / policy_sigma2 ) )
tx_vars = [item for item in tf.global_variables() if 'encoder' in item.name]
policy_gradient = tf.gradients(policy, tx_vars, grad_ys=tx_per_ex_loss/batch_size)
grads_and_vars = [ ( g,v ) for g,v in zip(policy_gradient,tx_vars)]
tx_optimizer = tf.train.AdamOptimizer(learning_rate=trainingParam.learningRate).apply_gradients(grads_and_vars)
# +
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
saver = tf.train.Saver()
checkpoint_path = os.path.join(trainingParam.path,'checkpoint',trainingParam.filename,'best')
if not os.path.exists(checkpoint_path):
os.makedirs(checkpoint_path)
else:
pass
# print("Restoring checkpoint...", flush=True)
# saver.restore(sess=sess,save_path=checkpoint_path)
# -
def TrainReceiver():
nBatches = int(trainingParam.sampleSize/trainingParam.batchSize)
bestLoss = 10000
for epoche in range(1, trainingParam.rx_iterations+1):
sess.run(resetOps)
for batch in range(0,nBatches):
data, _, _ = cu.hotOnes(trainingParam.batchSize,(1,0),chParam.M)
## GENERATE SYMBOLS
feedDict = {X: data}
[outEnc, outEncSeed] = sess.run([enc, enc_seed], feed_dict=feedDict)
## TRANSMIT THROUGH CHANNEL
feedDict = {channel_in: outEnc}
outChannelOut = sess.run(channel_out, feed_dict=feedDict)
## Train
feedDict = {X: data,Y:outChannelOut}
sess.run([rx_optimizer, updateOps], feed_dict=feedDict)
[outAvgLoss, outAvgAccuracy] = sess.run([meanMetricOpsDict['xentropy'], meanMetricOpsDict['accuracy_metric']], feed_dict=feedDict)
if outAvgLoss < bestLoss:
bestLoss = outAvgLoss
lastImprovement = epoche
saver.save(sess=sess,save_path=checkpoint_path)
if epoche - lastImprovement > trainingParam.earlyStopping:
print("Breaking due to no improvement")
break;
if epoche%trainingParam.displayStep == 0:
print('rx_epoche: {} - avgLoss: {} - avgAcc: {}'.format(epoche,outAvgLoss,outAvgAccuracy))
def TrainTransmitter(bs):
nBatches = int(trainingParam.sampleSize/trainingParam.batchSize)
# nBatches = 1;
bestLoss = 10000
for epoche in range(1, trainingParam.tx_iterations+1):
for batch in range(0,nBatches):
data, _, _ = cu.hotOnes(trainingParam.batchSize,(1,0),chParam.M,batch)
## GENERATE PERTURBED SYMBOLS
feedDict = {X: data, policy_sigma2: trainingParam.policy_sigma2}
[outEncPert, outEncSeed] = sess.run([enc_pert, enc_seed], feed_dict=feedDict)
## TRANSMIT THROUGH CHANNEL
feedDict = {channel_in: outEncPert}
outChannelOut = sess.run(channel_out, feed_dict=feedDict)
## PER EXAMPLE LOSS
feedDict={X:data, Y:outChannelOut}
outPerExampleLoss = sess.run(per_ex_loss,feed_dict=feedDict)
## TRAIN TRANSMITTER
feedDict={batch_size:bs, X:data,\
tx_per_ex_loss: outPerExampleLoss, tx_perturbed: outEncPert, policy_sigma2: trainingParam.policy_sigma2}
sess.run(tx_optimizer,feed_dict=feedDict)
## TEST
sess.run(resetOps)
for batch in range(0,nBatches):
data, _, _ = cu.hotOnes(trainingParam.batchSize,(1,0),chParam.M,133700+batch)
## GENERATE SYMBOLS
feedDict = {X: data}
[outEnc, outEncSeed] = sess.run([enc, enc_seed], feed_dict=feedDict)
## TRANSMIT THROUGH CHANNEL
feedDict = {channel_in: outEnc}
outChannelOut = sess.run(channel_out, feed_dict=feedDict)
## Test
feedDict = {X: data, Y:outChannelOut}
sess.run(updateOps, feed_dict=feedDict)
[outAvgLoss, outAvgAccuracy] = sess.run([meanMetricOpsDict['xentropy'], meanMetricOpsDict['accuracy_metric']], feed_dict=feedDict)
if outAvgLoss < bestLoss:
bestLoss = outAvgLoss
lastImprovement = epoche
saver.save(sess=sess,save_path=checkpoint_path)
if epoche - lastImprovement > trainingParam.earlyStopping:
print("Breaking due to no improvement")
break;
if epoche%trainingParam.displayStep == 0:
print('tx_epoche: {} - avgLoss: {} - avgAcc: {}'.format(epoche,outAvgLoss,outAvgAccuracy))
for jj in range(1,50):
print("##### {}".format(jj))
TrainReceiver()
saver.restore(sess=sess,save_path=checkpoint_path)
TrainTransmitter(trainingParam.batchSize*1)
saver.restore(sess=sess,save_path=checkpoint_path)
pred_const = sess.run(enc_seed)
plt.plot(pred_const[:,0],pred_const[:,1],'.')
plt.axis('square');
lim_ = 1.6
plt.xlim(-lim_,lim_);
plt.ylim(-lim_,lim_);
| examples/tf_AutoEncoderForGeometricShapingAndBlindAwgn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
import numpy as np
import ezplotly_bio as epb
# %load_ext autoreload
# %autoreload 2
# make output directory
if not os.path.exists("test_figs"):
os.mkdir("test_figs")
# +
## Manhattan plot demo
# simulate data and put into data frame
data = dict()
data["chr"] = list()
data["pos"] = list()
data["pval"] = list()
for i in range(1, 25):
chr_str = "chr" + str(i)
if i == 23:
chr_str = "chrX"
if i == 24:
chr_str = "chrY"
chrs = [chr_str for _ in range(100)]
pos = [j for j in range(100)]
pval = np.random.sample((100,)).tolist()
data["chr"].extend(chrs)
data["pos"].extend(pos)
data["pval"].extend(pval)
df = pd.DataFrame(data)
# make plot
epb.manhattan_plot(
df=df,
title="Example Manhattan Plot",
outfile=os.path.join("test_figs", "manhattan.png"),
height=1000,
)
# +
## Chr_Distr Plots:
# 1. Histogram data per chromosome
# 2. Test that chromosome values follow a uniform distribution
# simulate data and put into data frame
data = dict()
data["chr"] = list()
data["data"] = list()
for i in range(1, 25):
chr_str = "chr" + str(i)
if i == 23:
chr_str = "chrX"
if i == 24:
chr_str = "chrY"
chrs = [chr_str for _ in range(10000)]
d = np.random.sample((10000,)).tolist()
data["chr"].extend(chrs)
data["data"].extend(d)
df = pd.DataFrame(data)
# make chr_hist plot
epb.chr_hist(
df=df,
data_col_name="data",
min_bin=0.0,
max_bin=1.0,
bin_size=0.2,
title="Histogram per Chromosome",
xlabel="data",
x_dtick=0.2,
)
# make chr_distr plot
epb.chr_distr(
data=df["data"],
chr_df=df["chr"],
distr_name="uniform",
title="Test that data is uniform",
outfile=os.path.join("test_figs", "chr_distr.png"),
)
# +
## Make Chromosome QQ-Plot
epb.chr_qq(
df=df,
data_col_name="data",
distr="uniform",
title="QQ Plots: Test that data is uniform",
outfile=os.path.join("test_figs", "chr_qq.png"),
)
# +
## ROC curve example
# data
preds = np.array(
[
[0.9, 0.9, 0.9, 0.9, 0.1, 0.1, 0.1, 0.2, 0.3, 0.4],
[0.1, 0.1, 0.1, 0.1, 0.1, 0.9, 0.9, 0.2, 0.9, 0.9],
]
)
gt = np.array([1, 1, 1, 1, 1, 0, 0, 0, 0, 0])
# make roc curve
epb.roc(
preds=preds,
gt=gt,
names=["A", "B"],
title="ROC Curve Example",
outfile=os.path.join("test_figs", "roc.png"),
)
# +
## CDF Examples
data = np.array([0.1, 0.2, 0.3, 0.4, 0.9, 0.9, 0.9, 0.9])
# empirical cdf
epb.ecdf(
data=data,
min_bin=0.0,
max_bin=1.0,
bin_size=0.1,
title="Empirical CDF",
xlabel="data",
outfile=os.path.join("test_figs", "cdf.png"),
)
# reverse cdf
epb.rcdf(
data=data,
min_bin=0.0,
max_bin=1.0,
bin_size=0.1,
title="Reverse CDF",
xlabel="data",
outfile=os.path.join("test_figs", "rcdf.png"),
)
# +
## Corr Plot
x = np.arange(0.0, 50.0, 1.0)
y = np.arange(0.0, 50.0, 1.0) * 0.2
epb.corr_plot(
x=x,
y=y,
xlabel="x",
ylabel="y",
title="Correlation Plot Example",
outfile=os.path.join("test_figs", "corr_plot.png"),
)
# +
## Nonparametric Confidence Interval Examples
# data
x = np.array([1, 2, 3, 4, 5])
y = np.array(
[
[1, 1, 1, 1, 1],
[1, 2, 2, 2, 3],
[3, 3, 1, 2, 1],
[1, 2, 3, 4, 5],
[1, 1, 1, 1, 1],
]
).T
# 95% confidence interval line plot
epb.nonparametric_ci(
x=x,
y_data=y,
color="blue",
xlabel="x",
ylabel="y",
title="95% Confidence Interval Example",
outfile=os.path.join("test_figs", "conf95.png"),
)
# 75% confidence interval line plot
epb.nonparametric_ci(
x=x,
y_data=y,
color="blue",
conf=0.75,
xlabel="x",
ylabel="y",
title="75% Confidence Interval Example",
outfile=os.path.join("test_figs", "conf75.png"),
)
# 65% confidence interval scatter plot
epb.nonparametric_ci(
x=x,
y_data=y,
color="blue",
conf=0.75,
xlabel="x",
ylabel="y",
ci_plot_type="point",
title="65% Confidence Interval Example",
outfile=os.path.join("test_figs", "conf65.png"),
)
| EZPlotlyBioExamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1dxguN98xxQT" colab_type="text"
# # Mystery File!
#
# The purpose of this exercise is to give you a little taste of what we will be learning about this semester in Introduction to Digital Curation. Over the semester we will learn about how data is structured and how to interact with it from the Python programming language.
#
# Please access the file from Google Drive and answer any or all of the following if you can. Do not worry if you can't, this is stuff we will be learning about over the next few months.
#
# * What is the format of the file?
# * What does the file contain?
# * How would you use the file?
# * Where did the file come from?
# * Who created the information in the file?
# * Does it have a URL?
#
# ## Get the File
#
# Colab lets you mount your Google Drive. I will share a folder of data with you so you can easily access files we will be working with in Colab. If you want you can mount your own Google Drive folders as well.
#
# + id="RU5p7F2xs4lK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="8c73b96b-f7bd-45e4-d309-8f9c34baf608"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="3Lhaqp1NyPlt" colab_type="text"
# Now we can use the Python pathlib module to read the file.
# + id="-G8AnH2qtESD" colab_type="code" colab={}
import pathlib
f = pathlib.Path('/content/drive/Shared drives/INST341/module-01/file.tar')
# + [markdown] id="Q0QzQm3g0MJN" colab_type="text"
# Does the file exist?
# + id="aZTS5mBRvJjV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d7381259-8905-4e30-c422-f815c86e7295"
f.is_file()
# + [markdown] id="yC12RBi60uum" colab_type="text"
# ## File Type
#
# We can use the python-magic module to determine the type of the file. But first we need to install it, since it is not part of core Python. It also depends on a system library called libmagic which we can install.
# + id="Kjlnuxdh1I15" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 766} outputId="417d0ec0-1555-49f8-b0ac-84b1bdb4f6c6"
# ! pip3 install python-magic
# ! sudo apt-get install libmagic1
# + [markdown] id="h0sHCzXH1pCK" colab_type="text"
# Now we can import the [python-magic](https://pypi.org/project/python-magic/) module.
# + id="jFbb3WIvvL_f" colab_type="code" colab={}
import magic
# + [markdown] id="S36xy3qb1FoW" colab_type="text"
# And we can use it to identify the type of file.
# + id="NN2yyfQd1HGq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ba2588e6-07bb-4aa4-fdb0-69899acc5bd7"
magic.from_file(f.as_posix())
# + [markdown] id="tVk_tYzV2GsW" colab_type="text"
# Now that we know a little more about the file we can look it up. Wikipedia is surprisingly good for information about types of files. Here is the article about [TAR files](https://en.wikipedia.org/wiki/Tar_(computing))
#
# > In computing, tar is a computer software utility for collecting many files into one archive file, often referred to as a tarball, for distribution or backup purposes. The name is derived from "tape archive", as it was originally developed to write data to sequential I/O devices with no file system of their own. The archive data sets created by tar contain various file system parameters, such as name, timestamps, ownership, file-access permissions, and directory organization. The command-line utility was first introduced in the Version 7 Unix in January 1979, replacing the tp program.[2] The file structure to store this information was standardized in POSIX.1-1988[3] and later POSIX.1-2001,[4] and became a format supported by most modern file archiving systems.
# + [markdown] id="97JlQ40Z3RSw" colab_type="text"
# ## TAR Contents
#
# So file.tar is a *tape archive file*. That means it is a file that contains other files much like a ZIP file. Lets use Python's [tarfile](https://docs.python.org/3/library/tarfile.html) module to read it.
# + id="Oo5a1upa3xM3" colab_type="code" colab={}
import tarfile
tar = tarfile.open(f)
# + [markdown] id="uN1YZ_cZ4elu" colab_type="text"
# Now that we have a our variable tar that represents the tar file we can use a loop to list its contents:
# + id="f2VMKa4t37a5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="1f4e0374-d953-459d-e3a7-15e9b3428b90"
for info in tar:
print(info)
# + [markdown] id="CHfhVRw24Z9V" colab_type="text"
# Interesting! There is a lot of stuff in here. Lets extract all the files into our current working directory so we can look at them.
# + id="3Zn_KfEf4zkL" colab_type="code" colab={}
tar.extractall()
# + [markdown] id="hKev2b7k5lLQ" colab_type="text"
# The README.txt listed above looked interesting. Lets read that in and print it out.
# + id="HAB1vFTT5p9I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3a9b1257-51b0-4566-cf52-48bd9b7f44db"
text = open('ncbi-genomes-2020-08-27/README.txt').read()
print(text)
# + [markdown] id="w7HbbWZG8SSO" colab_type="text"
# That's a lot to read. But if you scroll to the top you'll see that some of this data is from the [GenBank](https://en.wikipedia.org/wiki/GenBank).
#
# This file describes the contents of the tarfile! Most of this is way over my head, I'm not a geneticist! But maybe it would be interesting to look at one of the files it mentions?
# + id="SM9Pnnbo5tJh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 575} outputId="d34c12bc-804d-49cc-944a-9578360c01ce"
text = open('ncbi-genomes-2020-08-27/GCA_009858895.3_ASM985889v3/GCA_009858895.3_ASM985889v3_assembly_report.txt').read()
print(text)
# + [markdown] id="UIUFVRf88-uc" colab_type="text"
# Oh wow, so this is genetic information about the Coronavirus! Lets take a look at one of the gzipped files using the python [gzip](https://docs.python.org/3/library/gzip.html) module.
# + id="cvk9Yxk_9KVP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="91fc7c03-51b6-4163-9bf1-7b37bb058207"
import gzip
text = gzip.open('ncbi-genomes-2020-08-27/GCA_009858895.3_ASM985889v3/GCA_009858895.3_ASM985889v3_genomic.fna.gz', 'rt').read()
print(text)
# + [markdown] id="V8016Y9u93Nx" colab_type="text"
# This looks like the genetic sequence for the Coronavirus!
#
# ## Answers
#
# Maybe we now have more questions than answers but here is one way of answering the initial questions posed at the beginning. If all of this seemed very hard. Don't worry, it was supposed to be difficult. By the end of the semester you should feel more comfortable using Python this way. But for the moment just get a sense of the flow of what happened.
#
# **1. What is the format of the file?**
#
# The file we started with was a tar file. But it contained other files such as text files and gzipped files.
#
# **2. What does the file contain?**
#
# It appears to contain information about the Coronavirus.
#
# **3. How would you use the file?**
#
# Geneticists could use this information to identify the virus in their labs.
#
# **4. Where did the file come from?**
#
# The file came from GenBank, which is a project run out of the National Institues of Health nearby in Bethesda.
#
# **5. Who created the information in the file?**
#
# Examining the metadata a little more closely (e.g. GCA_009858895.3_ASM985889v3_protein.gpff) shows that the genetic data was uploaded by Chinese scientists January 5, 2020 when they were publishing their findings in Nature.
#
# **6. Does it have a URL?**
#
# Sometimes you can use identifiers in data to try to locate more information about them on the web. In this case we can try to Google for ASM985889v3 which brings us to:
#
# https://www.ncbi.nlm.nih.gov/assembly/GCF_009858895.2/
#
# That looks like a start at least.
#
# + id="VHbAy_bHA5iz" colab_type="code" colab={}
| modules/module-01/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# To compare this GM81 process to Chen 2019, as a second check:
# Adapted from the following:
#
# ----------------------------------------------------------------------------
# Copyright (C) 2016 <NAME>
#
# This file is part of GM81.
#
# GM81 is free software: you can redistribute it and/or modify
# it under the terms of the GNU General Public License as published by
# the Free Software Foundation, either version 3 of the License, or
# (at your option) any later version.
#
# GM81 is distributed in the hope that it will be useful,
# but WITHOUT ANY WARRANTY; without even the implied warranty of
# MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# GNU General Public License for more details.
#
# You should have received a copy of the GNU General Public License
# along with GM81. If not, see <http://www.gnu.org/licenses/>.
#
# This computes KE and PE frequency spectra as well as 1D horizontal wavenumber spectra.
# ----------------------------------------------------------------------------
import numpy as np
import matplotlib.pyplot as plt
import gm
# #%matplotlib notebook
# Coriolis frequency
f = 5e-5 # rad/s
# buoyancy frequency (for specific depth, or the value of the WKB stretch)
N = 2e-3 # rad/s
# surface-extrapolated buoyancy frequency
N0 = 5.2e-3 # as given in GM79 literature
# e-folding scale of N(z)
b = 1.3e3 # m, according to GM79 literature, p.285, this is fairly universal
#https://ocw.mit.edu/resources/res-12-000-evolution-of-physical-oceanography-spring-2007/part-2/wunsch_chapter9.pdf
# frequency
omg = np.logspace(np.log10(1.01*f), np.log10(N), 401)
# horizontal wavenumber
k = 2*np.pi*np.logspace(-6, -2, 401)
# mode number
j = np.arange(1, 100)
# reshape to allow multiplication into 2D array
Omg = np.reshape(omg, (omg.size,1))
K = np.reshape(k, (k.size,1))
J = np.reshape(j, (1,j.size))
# frequency spectra (KE and PE)
K_omg_j = gm.K_omg_j(Omg, J, f, N, N0, b)
P_omg_j = gm.P_omg_j(Omg, J, f, N, N0, b)
# wavenumber spectra (KE and PE)
K_k_j = gm.K_k_j(K, J, f, N, N0, b)
P_k_j = gm.P_k_j(K, J, f, N, N0, b)
# sum over modes
K_omg = np.sum(K_omg_j, axis=1)
P_omg = np.sum(P_omg_j, axis=1)
K_k = np.sum(K_k_j, axis=1)
P_k = np.sum(P_k_j, axis=1)
# compute 1D spectra from 2D spectra
K_k_1d = gm.calc_1d(k, K_k)
P_k_1d = gm.calc_1d(k, P_k)
# plot kinetic energy frequency spectra
fig, ax = plt.subplots(figsize=(9,3))
ax.loglog(omg[3:], K_omg[3:], label='GM81',color='black',lw=0.8,ls='--') # 1/2 amplitude for rotary
ax.legend(frameon=False)
ax.axvline(x=f,ls='--',lw=0.5,color='red',label=f'$f$')
ax.set_xlim(1e-5,2e-3)
ax.set_ylim(1e-3,1e3)
ax.set_title('kinetic energy frequency spectra')
ax.set_xlabel('frequency (rad/s)')
ax.set_ylabel('power spectral density $((m^2/s)$')
ax.legend()
plt.tight_layout()
plt.show()
# -
| archive/GM/.ipynb_checkpoints/gm_check2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Streaming Algorithms in Machine Learning
#
# In this notebook, we will use an extremely simple "machine learning" task to learn about streaming algorithms. We will try to find the median of some numbers in batch mode, random order streams, and arbitrary order streams.
# The idea is to observe first hand the advantages of the streaming model as well as to appreciate some of the complexities involved in using it.
#
# The task at hand will be to approximate the median (model) of a long sequence of numbers (the data). This might seem to have little to do with machine learning. We are used to thinking of a median, $m$, of number $x_1,\ldots,x_n$ in the context of statistics as the number, $m$, which is smaller than at most half the values $x_i$ and larger than at most half the values $x_i$.
#
# Finding the median, however, also solves a proper machine learning optimization problem (albeit a simple one). The median minimizes the following clustering-like objective function
# $$m = \min_x \frac1n\sum_i|x - x_i|.$$
# In fact, the median is the solution to the well studied k-median clustering problem in one dimension and $k=1$. Moreover, the extension to finding all quantiles is common in feature transformations and an important ingedient in speeding up decission tree training.
# ## Batch Algorithms
# Let's first import a few libraries and create some random data.
# Our data will simple by $100,000$ equally spaced points between $0$ and $1$.
# +
import numpy as np
import matplotlib.pyplot as plt
n = 100000
data = np.linspace(0,1,n)
np.random.shuffle(data)
def f(x, data):
return sum(abs(x - datum) for datum in data)/len(data)
# -
# Let's look at the data to make sure everything is correct.
# %matplotlib inline
# Plotting every 10th point
plt.scatter(range(0,n,100),data[0:n:100],vmin=0,vmax=1.0)
plt.ylim((0.0,1.0))
plt.xlim((0,n))
plt.show()
# Computing the median brute force is trivial.
# +
from math import floor
def batchMedian(data):
n = len(data)
median = sorted(data)[int(floor(n/2))]
return(median)
median = batchMedian(data)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# The result is, of course, correct ($0.5$).
# To get the median we sorted the data in $O(n\log n)$ time even though QuickSelect would have been faster ($O(n)$). The algorithm speed is not the main issue here though. The main drawback of this algorithm is that it must store the entire dataset in memory. For either sorting or quickSelect the algorithm must also duplicate the array. Binary search is also a possible solution which doesn't require data duplication but does require $O(\log(n))$ passes over the data.
# When the data is large this is either very expensive or simply impossible.
# ## Streaming Algorithms (Random Order, SGD)
# In the streaming model, we assume only an iterator over the data is given. That is, we can only make one pass over the data. Moreover, the algorithm is limited in its memory footprint and the limit is much lower than the data size. Otherwise, we could "cheat" by storing all the data in memory and executing the batch mode algorithm.
#
# Gradient Descent (GD) type solutions are extremely common in this setting and are, de facto, the only mechanism for optimizing neural networks. In gradient descent, a step is taken in the direction opposite of the gradient. In one dimension, this simply means going left if the derivative is positive or right if the derivative is negative.
# %matplotlib inline
xs = list(np.linspace(-1.0,2.0,50))
ys = [f(x,data) for x in xs]
plt.plot(xs,ys)
plt.ylim((0.0,2.0))
plt.xlim((-1.0,2.0))
ax = plt.axes()
ax.arrow(-0.5, 1.1, 0.3, -0.3, head_width=0.05, head_length=0.1, fc='k', ec='k')
ax.arrow(1.5, 1.1, -0.3, -0.3, head_width=0.05, head_length=0.1, fc='k', ec='k')
plt.show()
# In **Stochastic Gradience Descent**, one only has a stochastic (random) unbiased estimator of the gradient. So, instead of computing the gradient of $\frac1n\sum_i|x - x_i|$ we can compute the gradient of $|x - x_i|$ where $x_i$ is chosen **uniformly at random** from the data. Note that a) the derivative of $|x - x_i|$ is simply $1$ if $x > x_i$ and $-1$ otherwise and b) the *expectation* of the derivative is exactly equal to the derivative of the overall objective function.
#
# Comment: the authors of the paper below suggest essentially this algorithm but do not mention the connection to SGD for some reason.
#
# Frugal Streaming for Estimating Quantiles: One (or two) memory suffices: <NAME>, <NAME>, <NAME>
# +
from math import sqrt
def sgdMedian(data, learningRate=0.1, initMedianValue=0):
median = initMedianValue
for (t,x) in enumerate(data):
gradient = 1.0 if x < median else -1.0
median = median - learningRate*gradient/sqrt(t+1)
return(median)
median = sgdMedian(data, learningRate=0.1, initMedianValue=0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# The result isn't exactly $0.5$ but it is pretty close. If this was a real machine learning problem, matching the objective up to the 5th digit of the true global minimum would have been very good.
#
# Why does this work? Let's plot our objective function to investigate further.
# It should not come as a big surprise to you that the objective function is convex. After all, it is the sum of convex functions (absolute values). It is a piece-wise linear curve that approximates a parabole in the range $(0,1)$ and is linear outside that range. Therefore, gradient descent is guaranteed to converge.
# SGD significantly more efficient than sorting or even QuickSelect. More importantly, its memory footprint is tiny, a handful of doubles, *regardless of the size of the data*!!!
#
# This is a huge advantage when operating with large datasets or with limited hardware.
# Alas, SGD has some subtleties that make it a little tricky to use sometimes.
# +
# SGD needs to be initialized carfully
median = sgdMedian(data, learningRate=0.1, initMedianValue=100.0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# +
# SGD needs to set step sizes corectly (controled via the learing rate)
median = sgdMedian(data, learningRate=0.001, initMedianValue=0.0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# These issues are usually alleviated by adaptive versions of SGD. Enhancements to SGD such as second order (based) methods, adaptive learning rate, and momentum methods may help in these situations but still require tuning in many cases. A common approach is to use many epochs.
# +
median=0.0
numEpochs = 100
for i in range(numEpochs):
median = sgdMedian(data, learningRate=0.001, initMedianValue=median)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# While clearly much less efficient than a single pass, increasing the number of epochs seemed to have solved the problem. Machine learning practitioners can relate to this result. That is, SGD is a great algorithm IF one finds good parameters for initialization, learning rate, number of epochs etc.
#
# One of the main challenges in designing fundamentally better SGD-based streaming algorithms is in adaptively controlling these parameters during the run of the algorithm.
#
# It is important to mention that there are also fundamentally better algorithms than SGD for this problem. See for example:
#
# _<NAME>, <NAME>_ <br>
# Stream Order and Order Statistics: Quantile Estimation in Random-Order Streams. <br>
# _SIAM J. Comput. 38(5): 2044-2059 (2009)_
#
# Unfortunately, we don't have time to dive into that...
# ### Trending data poses a challenge...
# SGD and the other above algorithm have a fundamental drawback. They inherently rely on the fact that the data is random. For SGD, the gradient of the loss on a single point (or minibatch) must be an estimator of the global gradient. This is not true if trends in data make its statistics change (even slightly) over time. Let's simulate this with our data.
# +
# %matplotlib inline
# SGD also depends on the data being reandomly suffeled
n,k = len(data),10
minibatches = [data[i:i+k] for i in range(0,n,k)]
minibatches.sort(key=sum)
trendyData = np.array(minibatches).reshape(n)
# Plotting every 10th point in the trending dataset
plt.scatter(range(0,n,100),trendyData[0:n:100],vmin=0,vmax=1.0)
plt.ylim((0.0,1.0))
plt.xlim((0,n))
plt.show()
# +
median = sgdMedian(trendyData, learningRate=0.1, initMedianValue=0.0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# ## Streaming Algorithms (single pass, arbitrary order)
# One way not to be fooled by trends in data and to sample from it.
# The algorithm uses Reservoir Sampling to obtain $k$ (in this case $k=1000$) uniformly chosen samples from the stream. Then, compute the batch median of the sample.
#
# The main drawback of sampling is that we now use more memory. Roughly the sample size $k$ ($k=1000$ here). This much more than $O(1)$ needed for SGD. Yet, it has some very appealing properties. Sampling very efficient ($O(1)$ per update), it is very simple to implement, it doesn't have any numeric sensitivities or tunable input parameters, and it is provably correct.
#
# _(For the sake of simplicity below we use python's builtin sample function rather than recode reservoir sampling)_
# +
from random import sample
def sampleMedian(data):
k=300
samples = sample(list(data),k)
return batchMedian(samples)
median = sampleMedian(trendyData)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# As you can see, sampling provides relatively good results.
#
# Nevertheless, there is something deeply dissatisfying about it. The algorithm was given $100,000$ points and used on $1,000$ of them. I other words, it would have been just as accurate had we collected only $1\%$ of the data.
#
# Can we do better? Can an algorithm simultaneously take advantage of all the data, have a fixed memory footprint, and not be sensitive to the order in which the data is consumed? The answer is _yes!_. These are known in the academic literature as Sketching (or simply streaming) algorithms.
#
# Specifically for approximating the median (or any other quantile), there is a very recent result that shows how best to achieve that:
#
# _<NAME>, <NAME>, <NAME>_ <br>
# Optimal Quantile Approximation in Streams. <br>
# FOCS 2016: 71-78
#
# The following code is a hacky version of the algorithm described in the paper above. Warning: this function will not work for streams much longer than $100,000$!
# +
from kll300 import KLL300
from bisect import bisect
def sketchMedian(data):
sketch = KLL300()
for x in data:
sketch.update(x)
assert sketch.size <= 300 # making sure there is no cheating involved...
items, cdf = sketch.cdf()
i = bisect(cdf, 0.5)
median = items[i]
return median
median = sketchMedian(trendyData)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# -
# Note that sketchMedian and sampleMedian both retain at most 300 items from the stream.
# Still, the sketching solution is significantly more accurate.
# Note that both sampling and sketching are randomized algorithms.
# It could be that sampling happens to be more accurate than sketching for any single run. But, as a whole, you should expect the sketching algorithm to be much more accurate.
#
# If you are curious about what sketchMedian actually does, you should look here:
# * Academic paper - https://arxiv.org/abs/1603.05346
# * JAVA code as part of datasketches - https://github.com/DataSketches/sketches-core/tree/master/src/main/java/com/yahoo/sketches/kll
# * Scala code by <NAME> - https://github.com/zkarnin/quantiles-scala-kll
# * Python experiments by <NAME> - https://github.com/nikitaivkin/quantilesExperiments
#
# The point is, getting accurate and stable streaming algorithms is complex. This is true even for very simple problems (like the one above). But, if one can do that, the benefits are well worth it.
| scientific_details_of_algorithms/streaming_median/streamingMedian.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Content Based Recommendations
#
# In the previous notebook, you were introduced to a way of making recommendations using collaborative filtering. However, using this technique there are a large number of users who were left without any recommendations at all. Other users were left with fewer than the ten recommendations that were set up by our function to retrieve....
#
# In order to help these users out, let's try another technique: **content based** recommendations. Let's start off where we were in the previous notebook.
# +
# #!pip install progressbar
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from collections import defaultdict
from IPython.display import HTML
import progressbar
import tests as t
import pickle
# %matplotlib inline
# Read in the datasets
movies = pd.read_csv('movies_clean.csv')
reviews = pd.read_csv('reviews_clean.csv')
del movies['Unnamed: 0']
del reviews['Unnamed: 0']
all_recs = pickle.load(open("all_recs.p", "rb"))
# -
# ### Datasets
#
# From the above, you now have access to three important items that you will be using throughout the rest of this notebook.
#
# `a.` **movies** - a dataframe of all of the movies in the dataset along with other content related information about the movies (genre and date)
#
#
# `b.` **reviews** - this was the main dataframe used before for collaborative filtering, as it contains all of the interactions between users and movies.
#
#
# `c.` **all_recs** - a dictionary where each key is a user, and the value is a list of movie recommendations based on collaborative filtering
#
# For the individuals in **all_recs** who did receive 10 recommendations using collaborative filtering, we don't really need to worry about them. However, there were a number of individuals in our dataset who did not receive any recommendations.
#
# -----
#
# `1.` Let's start with finding all of the users in our dataset who didn't get all 10 ratings we would have liked them to have using collaborative filtering.
# +
users_with_all_recs = [] # Store user ids who have all their recommendations in this (10 or more)
users_who_need_recs = [] # Store users who still need recommendations here
for user, movie_recs in all_recs.items():
if len(movie_recs)>9:
users_with_all_recs.append(user)
users = np.unique(reviews['user_id'])
users_who_need_recs = np.setdiff1d(users, users_with_all_recs)
print(len(users_with_all_recs))
print(len(users_who_need_recs))
# -
# A quick test
assert len(users_with_all_recs) == 22187
print("That's right there were still users who needed recommendations when we only used collaborative filtering!")
# ### Content Based Recommendations
#
# You will be doing a bit of a mix of content and collaborative filtering to make recommendations for the users this time. This will allow you to obtain recommendations in many cases where we didn't make recommendations earlier.
#
# `2.` Before finding recommendations, rank the user's ratings from highest to lowest. You will move through the movies in this order looking for other similar movies.
# create a dataframe similar to reviews, but ranked by rating for each user
ranked_reviews=reviews[['user_id','movie_id','rating']]
ranked_reviews=ranked_reviews.sort_values(['user_id', 'rating'], ascending=[True, False])
ranked_reviews.head()
# ### Similarities
#
# In the collaborative filtering sections, you became quite familiar with different methods of determining the similarity (or distance) of two users. We can perform similarities based on content in much the same way.
#
# In many cases, it turns out that one of the fastest ways we can find out how similar items are to one another (when our matrix isn't totally sparse like it was in the earlier section) is by simply using matrix multiplication. If you are not familiar with this, an explanation is available [here by 3blue1brown](https://www.youtube.com/watch?v=LyGKycYT2v0) and another quick explanation is provided [in the post here](https://math.stackexchange.com/questions/689022/how-does-the-dot-product-determine-similarity).
#
# For us to pull out a matrix that describes the movies in our dataframe in terms of content, we might just use the indicator variables related to **year** and **genre** for our movies.
#
# Then we can obtain a matrix of how similar movies are to one another by taking the dot product of this matrix with itself. Notice below that the dot product where our 1 values overlap gives a value of 2 indicating higher similarity. In the second dot product, the 1 values don't match up. This leads to a dot product of 0 indicating lower similarity.
#
# <img src="images/dotprod1.png" alt="Dot Product" height="500" width="500">
#
# We can perform the dot product on a matrix of movies with content characteristics to provide a movie by movie matrix where each cell is an indication of how similar two movies are to one another. In the below image, you can see that movies 1 and 8 are most similar, movies 2 and 8 are most similar, and movies 3 and 9 are most similar for this subset of the data. The diagonal elements of the matrix will contain the similarity of a movie with itself, which will be the largest possible similarity (and will also be the number of 1's in the movie row within the orginal movie content matrix).
#
# <img src="images/moviemat.png" alt="Dot Product" height="500" width="500">
#
#
# `3.` Create a numpy array that is a matrix of indicator variables related to year (by century) and movie genres by movie. Perform the dot product of this matrix with itself (transposed) to obtain a similarity matrix of each movie with every other movie. The final matrix should be 31245 x 31245.
# +
# Subset so movie_content is only using the dummy variables for each genre and the 3 century based year dummy columns
movie_content = np.array(movies.iloc[:,4:])
# Take the dot product to obtain a movie x movie matrix of similarities
dot_prod_movies = movie_content.dot(np.transpose(movie_content))
# -
# create checks for the dot product matrix
assert dot_prod_movies.shape[0] == 31245
assert dot_prod_movies.shape[1] == 31245
assert dot_prod_movies[0, 0] == np.max(dot_prod_movies[0])
print("Looks like you passed all of the tests. Though they weren't very robust - if you want to write some of your own, I won't complain!")
# ### For Each User...
#
#
# Now you have a matrix where each user has their ratings ordered. You also have a second matrix where movies are each axis, and the matrix entries are larger where the two movies are more similar and smaller where the two movies are dissimilar. This matrix is a measure of content similarity. Therefore, it is time to get to the fun part.
#
# For each user, we will perform the following:
#
# i. For each movie, find the movies that are most similar that the user hasn't seen.
#
# ii. Continue through the available, rated movies until 10 recommendations or until there are no additional movies.
#
# As a final note, you may need to adjust the criteria for 'most similar' to obtain 10 recommendations. As a first pass, I used only movies with the highest possible similarity to one another as similar enough to add as a recommendation.
#
# `3.` In the cell below, complete each of the functions needed for making content based recommendations.
movies[movies['movie_id'] == 68646]
np.where(movies['movie_id'] == 68646)[0][0]
# +
def find_similar_movies(movie_id):
'''
INPUT
movie_id - a movie_id
OUTPUT
similar_movies - an array of the most similar movies by title
'''
# find the row of each movie id
movie_idx=np.where(movies['movie_id'] == movie_id)[0][0]
# find the most similar movie indices - to start I said they need to be the same for all content
similar_idxs = np.where(dot_prod_movies[movie_idx] == np.max(dot_prod_movies[movie_idx]))[0]
# pull the movie titles based on the indices
similar_movies = np.array(movies.iloc[similar_idxs, ]['movie'])
return similar_movies
# You made this function in an earlier notebook - using again here
def get_movie_names(movie_ids):
'''
INPUT
movie_ids - a list of movie_ids
OUTPUT
movies - a list of movie names associated with the movie_ids
'''
movie_lst = list(movies[movies['movie_id'].isin(movie_ids)]['movie'])
return movie_lst
def make_recs():
'''
INPUT
None
OUTPUT
recs - a dictionary with keys of the user and values of the recommendations
'''
# Create dictionary to return with users and ratings
recs = defaultdict(set)
# How many users for progress bar
n_users = len(users)
# Create the progressbar
cnter = 0
bar = progressbar.ProgressBar(maxval=n_users+1, widgets=[progressbar.Bar('=', '[', ']'), ' ', progressbar.Percentage()])
bar.start()
# For each user
for user in users:
# Update the progress bar
cnter+=1
bar.update(cnter)
# Pull only the reviews the user has seen
reviews_temp = ranked_reviews[ranked_reviews['user_id'] == user]
movies_temp = np.array(reviews_temp['movie_id'])
movie_names = np.array(get_movie_names(movies_temp))
# Look at each of the movies (highest ranked first),
# pull the movies the user hasn't seen that are most similar
# These will be the recommendations - continue until 10 recs
# or you have depleted the movie list for the user
for movie in movies_temp:
rec_movies = find_similar_movies(movie)
temp_recs = np.setdiff1d(rec_movies, movie_names)
recs[user].update(temp_recs)
# If there are more than
if len(recs[user]) > 9:
break
bar.finish()
return recs
# -
recs = make_recs()
# ### How Did We Do?
#
# Now that you have made the recommendations, how did we do in providing everyone with a set of recommendations?
#
# `4.` Use the cells below to see how many individuals you were able to make recommendations for, as well as explore characteristics about individuals for whom you were not able to make recommendations.
# Explore recommendations
users_without_all_recs = []
users_with_all_recs = []
no_recs = []
for user, movie_recs in recs.items():
if len(movie_recs) < 10:
users_without_all_recs.append(user)
if len(movie_recs) > 9:
users_with_all_recs.append(user)
if len(movie_recs) == 0:
no_recs.append(user)
print("There were {} users without all 10 recommendations we would have liked to have.".format(len(users_without_all_recs)))
print("There were {} users with all 10 recommendations we would like them to have.".format(len(users_with_all_recs)))
print("There were {} users with no recommendations at all!".format(len(no_recs)))
# ### Now What?
#
# Well, if you were really strict with your criteria for how similar two movies are (like I was initially), then you still have some users that don't have all 10 recommendations (and a small group of users who have no recommendations at all).
#
# As stated earlier, recommendation engines are a bit of an **art** and a **science**. There are a number of things we still could look into - how do our collaborative filtering and content based recommendations compare to one another? How could we incorporate user input along with collaborative filtering and/or content based recommendations to improve any of our recommendations? How can we truly gain recommendations for every user?
#
# +
# Cells for exploring
# -
| Content Based Recommendation/Content Based Recommendations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
dataset = pd.read_csv('AAPL.csv')
x=dataset.iloc[:, :-1].values
x
y=dataset.iloc[:, -1].values
y
from sklearn.linear_model import LinearRegression #as
regressor = LinearRegression()
regressor.fit(x,y)
y_pred = regressor.predict(x)
plt.scatter(x,y, color='red')
plt.plot(x, regressor.predict(x), color = 'blue')
plt.xlabel('Days')
plt.ylabel('Stock Market Price')
plt.ylim(100,200)
plt.show()
# +
# this data shows the the stock price of APPLE for the month 20 FEB,2022 TO 20 MAY,2022. The graph shows that APPLE stock price will go down....
# -
| Apple Stock Price Prediction Feb to May .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow13]
# language: python
# name: conda-env-tensorflow13-py
# ---
import tensorflow as tf
tf.__version__
print "Author: <NAME>"
# Create a TensorFlow cluster with one worker node and one ps node.
task_index=0
cluster_spec = tf.train.ClusterSpec({'ps' : ['localhost:2222'],'worker' : ['localhost:2223','localhost:2224']})
server = tf.train.Server(cluster_spec,job_name='worker',task_index=task_index)
# **Launch and run all the cells in the parameter server notebook**
# Create variables locally then makes global copy on ps.
# +
tf.reset_default_graph()
#create local graph like normal specifying the local device
with tf.device('/job:worker/task:0'):
a = tf.Variable([0.],name='a',collections=[tf.GraphKeys.LOCAL_VARIABLES])
b = tf.constant([100.])
loss = tf.abs(a-b)
optimizer = tf.train.GradientDescentOptimizer(.1)
grads,local_vars = zip(*optimizer.compute_gradients(loss,var_list=tf.local_variables()))
local_update = optimizer.apply_gradients(zip(grads,local_vars))
init_local = tf.local_variables_initializer()
#create the globabl copies on the ps
with tf.device('/job:ps/task:0'):
for v in tf.local_variables():
v_g = tf.get_variable('g/'+v.op.name,
shape = v.shape,
dtype = v.dtype,
trainable=True,
collections=[tf.GraphKeys.GLOBAL_VARIABLES,tf.GraphKeys.TRAINABLE_VARIABLES])
#gloabl updates
with tf.device('/job:worker/task:%d'%task_index):
#this needs to be updated. Clearly not robust for any graph more complext
global_vars = tf.global_variables()
global_update = optimizer.apply_gradients(zip(grads,global_vars))
#create init op on the chief node
with tf.device('/job:worker/task:%d'%task_index):
init_global = tf.global_variables_initializer()
# -
# View device placements
a_global = tf.global_variables()[0]
print(a.device)
print(b.device)
print(loss.device)
print(local_update.device)
print(global_update.device)
print(init_global.device)
print(init_local.device)
print(a_global.device)
sess = tf.Session(target=server.target)
sess.run([init_local,init_global])
# Make sure you have also run all cells in the worker 2 notebook up to this point before continuing. The above cell should hang until you initialize the worker 2 session.
sess.run([a,a_global])
sess.run(local_update)
sess.run([a,a_global])
sess.run(global_update)
sess.run([a,a_global])
# Pause here. Run the last cell in this notebook after you have done a global update in the worker 2 notebook.
sess.run([a_global])
| Basics-Tutorial/Multiple-Workers/Local-then-Global-Variables-Worker1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="lW-qXiPUHMDS"
# # DATA CLEANING
# + colab={} colab_type="code" id="m9kDnGt7HMDX"
import pandas as pd
import numpy as np
#importing the neccessary modules for data cleaning
# + colab={} colab_type="code" id="obq3Ak0QHMDg"
import matplotlib.pyplot as plt
import seaborn as sns
#modules for visualization
# + colab={} colab_type="code" id="04DTrxevHMDn"
import warnings
warnings.filterwarnings('ignore')
#To prevent the uneccessary warnings during data reading from showing up
# + colab={} colab_type="code" id="EtiW9nd-HMDv"
df = pd.read_csv('datasets/Original_Dataset.csv')
#Creating a dataframe object with the data from the csv file.
# + colab={} colab_type="code" id="4d6sRnoZHMD4" outputId="c6d31247-dd3c-4d0b-ce7d-f7d92bc848bb"
df.shape
#getting to know the dimensions of our dataframe
# + colab={} colab_type="code" id="oLl3fzC7HMEB"
#Let us first remove the columns/attributes, who's values are not really important in our analysis.
#From visual analysis of the columns, we find that there are columns who's values does not
#affect our analysis. So to make the dataset more meaningful, we remove those columns.
# + colab={} colab_type="code" id="EbguPkJ5HMEH"
unecessary_columns = ['Respondent','AssessJob1','AssessJob2','AssessJob3','AssessJob4','AssessJob5', 'AssessJob6',
'AssessJob7', 'AssessJob8','AssessJob9', 'AssessJob10','AssessBenefits1','AssessBenefits2','AssessBenefits3',
'AssessBenefits4','AssessBenefits5','AssessBenefits6','AssessBenefits7','AssessBenefits8','AssessBenefits9',
'AssessBenefits10','AssessBenefits11','JobContactPriorities1','JobContactPriorities3',
'JobContactPriorities4','JobContactPriorities5','JobEmailPriorities1','JobEmailPriorities2','JobEmailPriorities3',
'JobEmailPriorities4','JobEmailPriorities5','JobEmailPriorities6','JobEmailPriorities7','FormalEducation',
'CareerSatisfaction','HopeFiveYears','UpdateCV','TimeAfterBootcamp',
'AgreeDisagree1','AgreeDisagree2','AgreeDisagree3', 'HypotheticalTools1','HypotheticalTools2',
'HypotheticalTools3', 'HypotheticalTools4','HypotheticalTools5']
# + colab={} colab_type="code" id="D8mhR2efHMEM"
df = df.drop(unecessary_columns,axis=1)
# + colab={} colab_type="code" id="rOskcUgTHMER"
#Function to plot a bar graph of the missing values in each column of our dataset
def plot_nan_percentage(df):
miss = df.isnull().sum().reset_index()
miss[0] = (miss[0]*100)/df.shape[0]
plt.figure(figsize=(13,6))
ax = sns.barplot("index",0,data=miss,color="orange")
plt.xticks(rotation = 90,fontsize=6)
plt.title("percentage of missing values")
ax.set_facecolor("k")
ax.set_ylabel("percentage of missing values")
ax.set_xlabel("variables")
plt.show()
# + colab={} colab_type="code" id="mH1um_W_HMEY" outputId="16fa3a07-9919-4f73-e0ab-95283eaec264"
# Now let us check out the NAN values in our dataset.
# We will then apply a treshold and remove all columns with NAN values over a certain amount.
#LET US FIRST HAVE A VISUAL REPRESENTATION OF OUR DATASET.
plot_nan_percentage(df)
# + colab={} colab_type="code" id="8FVrzjCYHMEe"
# From the above visualization we find out that some columns have a large number of missing values.
# Fixing the maximum threshold of nan values to be about 40%
l = [c for c in df if (df[c].isnull().sum() < 40000) ]
l.append('ConvertedSalary')
df = df[l]
# + colab={} colab_type="code" id="tdEsTziXHMEi" outputId="b47dfa20-2bcc-4714-cd51-c1bfbb9c5be9"
#Now our visualisation of the nan values in our dataset would now be
plot_nan_percentage(df)
# + colab={} colab_type="code" id="qu_NVwAbHMEn"
#dropping rows with responses < 63
df = df.dropna(thresh= 64)
# + colab={} colab_type="code" id="dti-JhVaHMEr" outputId="c8b7669f-ea4a-4bbc-87aa-1c88a78b3a70"
plot_nan_percentage(df)
# + colab={} colab_type="code" id="c6wUpYzKHMEx"
#lets divide the rows into categorical and numerical
numerical_columns = ["ConvertedSalary"] #need to be filled
categorical_columns = ["Hobby","OpenSource"] #need to be filled
# + colab={} colab_type="code" id="2lyX6LvWHME3"
#rewriting missing numerical values with mean.
for i in numerical_columns:
mean = df[i].mean()
df = df.fillna(value = {i:mean})
# + colab={} colab_type="code" id="EWFBDOdfHME7"
#replacing missing categorical values with previous row elements
for i in categorical_columns:
df[[i]] = df[[i]].fillna(method="ffill")
# + colab={} colab_type="code" id="RxYFD_2THMFA"
df = df.dropna() #will be overwritten by the data handling done on top
# + colab={} colab_type="code" id="KXLrWV3RHMFG"
#Now we write our clear data frame without nan values into our new csv file
df.to_csv('datasets/Clean_Dataset.csv')
# + colab={} colab_type="code" id="Xy5ozeRcHMFK" outputId="da3fd425-b088-4ea3-8f7d-7a22a5a69656"
df.shape
#getting to know the dimensions of our final dataframe
# + colab={} colab_type="code" id="egeatORqHMFU"
#END OF DATA CLEANING
| EDA/StackOverflow19_Developer_Dataset/Data_Cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ml4t]
# language: python
# name: conda-env-ml4t-py
# ---
# # Download and store STOOQ data
# This notebook contains information on downloading the STOOQ stock and ETF price data that we use in [Chapter 09](../09_time_series_models) for a pairs trading strategy based on cointegration and [Chapter 11](../11_decision_trees_random_forests) for a long-short strategy using Random Forest return predictions.
# ## Imports & Settings
import warnings
warnings.filterwarnings('ignore')
# +
from pathlib import Path
import requests
from io import BytesIO
from zipfile import ZipFile, BadZipFile
import numpy as np
import pandas as pd
import pandas_datareader.data as web
from sklearn.datasets import fetch_openml
pd.set_option('display.expand_frame_repr', False)
# -
# ## Set Data Store path
# Modify the path to the `DATA_STORE` if you would like to store the data elsewhere and change the notebooks accordingly
DATA_STORE = Path('assets.h5')
# ## Stooq Historical Market Data
# > Note that the below downloading details may change at any time as Stooq updates their website; if you encounter errors, please inspect their website and raise a GitHub issue to let us know so we can update the information.
#
# > Update 12/2020: please note that STOOQ will disable automatic downloads and require CAPTCHA starting Dec 10, 2020 so that the code that downloads and unpacks the zip files will no longer work; please navigate to their website [here](https://stooq.com/db/h/) for manual download.
# ### Download price data
# 1. Download **price data** for the selected combination of asset class, market and frequency from [the Stooq website](https://stooq.com/db/h/)
# 2. Store the result under `stooq` using the preferred folder structure outlined on the website. It has the structure: `/data/freq/market/asset_class`, such as `/data/daily/us/nasdaq etfs`.
stooq_path = Path('stooq')
if not stooq_path.exists():
stooq_path.mkdir()
# Use the symbol for the market you want to download price data for. In this book we'll be useing `us` and `jp`.
STOOQ_URL = 'https://static.stooq.com/db/h/'
def download_price_data(market='us'):
data_url = f'd_{market}_txt.zip'
response = requests.get(STOOQ_URL + data_url).content
with ZipFile(BytesIO(response)) as zip_file:
for i, file in enumerate(zip_file.namelist()):
if not file.endswith('.txt'):
continue
local_file = stooq_path / file
local_file.parent.mkdir(parents=True, exist_ok=True)
with local_file.open('wb') as output:
for line in zip_file.open(file).readlines():
output.write(line)
for market in ['us', 'jp']:
download_price_data(market=market)
# ### Add symbols
# Add the corresponding **symbols**, i.e., tickers and names by following the directory tree on the same site. You can also adapt the following code snippet using the appropriate asset code that you find by inspecting the url; this example works for NASDAQ ETFs that have code `g=69`:
# ```python
# df = pd.read_csv('https://stooq.com/db/l/?g=69', sep=' ').apply(lambda x: x.str.strip())
# df.columns = ['ticker', 'name']
# df.drop_duplicates('ticker').to_csv('stooq/data/tickers/us/nasdaq etfs.csv', index=False)
# ```
metadata_dict = {
('jp', 'tse etfs'): 34,
('jp', 'tse stocks'): 32,
('us', 'nasdaq etfs'): 69,
('us', 'nasdaq stocks'): 27,
('us', 'nyse etfs'): 70,
('us', 'nyse stocks'): 28,
('us', 'nysemkt stocks'): 26
}
for (market, asset_class), code in metadata_dict.items():
df = pd.read_csv(f'https://stooq.com/db/l/?g={code}', sep=' ').apply(lambda x: x.str.strip())
df.columns = ['ticker', 'name']
df = df.drop_duplicates('ticker').dropna()
print(market, asset_class, f'# tickers: {df.shape[0]:,.0f}')
path = stooq_path / 'tickers' / market
if not path.exists():
path.mkdir(parents=True)
df.to_csv(path / f'{asset_class}.csv', index=False)
# ### Store price data in HDF5 format
# To speed up loading, we store the price data in HDF format. The function `get_stooq_prices_and_symbols` loads data assuming the directory structure described above and takes the following arguments:
# - frequency (see Stooq website for options as these may change; default is `daily`
# - market (default: `us`), and
# - asset class (default: `nasdaq etfs`.
#
# It removes files that do not have data or do not appear in the corresponding list of symbols.
def get_stooq_prices_and_tickers(frequency='daily',
market='us',
asset_class='nasdaq etfs'):
prices = []
tickers = (pd.read_csv(stooq_path / 'tickers' / market / f'{asset_class}.csv'))
if frequency in ['5 min', 'hourly']:
parse_dates = [['date', 'time']]
date_label = 'date_time'
else:
parse_dates = ['date']
date_label = 'date'
names = ['ticker', 'freq', 'date', 'time',
'open', 'high', 'low', 'close','volume', 'openint']
usecols = ['ticker', 'open', 'high', 'low', 'close', 'volume'] + parse_dates
path = stooq_path / 'data' / frequency / market / asset_class
print(path.as_posix())
files = path.glob('**/*.txt')
for i, file in enumerate(files, 1):
if i % 500 == 0:
print(i)
if file.stem not in set(tickers.ticker.str.lower()):
print(file.stem, 'not available')
file.unlink()
else:
try:
df = (pd.read_csv(
file,
names=names,
usecols=usecols,
header=0,
parse_dates=parse_dates))
prices.append(df)
except pd.errors.EmptyDataError:
print('\tdata missing', file.stem)
file.unlink()
prices = (pd.concat(prices, ignore_index=True)
.rename(columns=str.lower)
.set_index(['ticker', date_label])
.apply(lambda x: pd.to_numeric(x, errors='coerce')))
return prices, tickers
# We'll be using US equities and ETFs in [Chapter 9](../09_time_series_models) and and Japanese equities in [Chapter 11](../11_decision_trees_random_forests). The following code collects the price data for the period 2000-2019 and stores it with the corresponding symbols in the global `assets.h5` store:
# +
# load some Japanese and all US assets for 2000-2019
markets = {'jp': ['tse stocks'],
'us': ['nasdaq etfs', 'nasdaq stocks', 'nyse etfs', 'nyse stocks', 'nysemkt stocks']
}
frequency = 'daily'
idx = pd.IndexSlice
for market, asset_classes in markets.items():
for asset_class in asset_classes:
print(f'\n{asset_class}')
prices, tickers = get_stooq_prices_and_tickers(frequency=frequency,
market=market,
asset_class=asset_class)
prices = prices.sort_index().loc[idx[:, '2000': '2019'], :]
names = prices.index.names
prices = (prices
.reset_index()
.drop_duplicates()
.set_index(names)
.sort_index())
print('\nNo. of observations per asset')
print(prices.groupby('ticker').size().describe())
key = f'stooq/{market}/{asset_class.replace(" ", "/")}/'
print(prices.info(null_counts=True))
prices.to_hdf(DATA_STORE, key + 'prices', format='t')
print(tickers.info())
tickers.to_hdf(DATA_STORE, key + 'tickers', format='t')
| ml4trading-2ed/data/create_stooq_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import xarray as xr
import netCDF4 as nc
# -
def f(x, omega=1):
"""simple sine function"""
return np.sin(2 * np.pi * omega *x)
times = np.linspace(0, 10, 100)
sine = f(times)
# +
# create DataArray with metadata
metadata = {'timestep': times[1] - times[0], 'creation_data': '19-07-17', 'list': ['a', 'b', 2, 3]}
df = xr.DataArray(sine, dims='time', coords={'time': times}, name='sine', attrs=metadata)
df.attrs
# -
df.to_netcdf('sine.nc')
new_df = xr.open_dataarray('sine.nc')
new_df.attrs
all(new_df == df)
# ## Open with netCDF
nc_df = nc.Dataset('sine.nc')
nc_df.variables
new_sine = np.asarray(nc_df['sine'][:])
sum(sine - new_sine)
nc_df['sine'].__dict__['timestep']
| numerics/netCDF/netCDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="X8KTofTcDIFD" colab_type="code" outputId="b2009236-e72b-4d7c-a17f-35a85bd6ae24" executionInfo={"status": "ok", "timestamp": 1583394368009, "user_tz": -330, "elapsed": 4600, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 306}
# !nvidia-smi
# + colab_type="code" id="69YZC_tMBknJ" outputId="20d2d375-8d9b-4557-98fc-1e9757e61da4" executionInfo={"status": "ok", "timestamp": 1583394390937, "user_tz": -330, "elapsed": 27513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !apt-get install protobuf-compiler python-pil python-lxml python-tk
# !pip install Cython
# !pip install matplotlib
# + colab_type="code" id="50F8-y0eF2HK" outputId="b2ebdc03-752b-4407-ae9d-ba4cde28d389" executionInfo={"status": "ok", "timestamp": 1583394441991, "user_tz": -330, "elapsed": 78554, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/<KEY>", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 153}
# !git clone https://github.com/tensorflow/models.git
# !pip install -q contextlib2
# !pip install -q pycocotools
# + colab_type="code" id="VZPf7e_yF_L7" outputId="51d83378-1bb2-4ac9-9345-eaa176a51b8a" executionInfo={"status": "ok", "timestamp": 1583394452103, "user_tz": -330, "elapsed": 88657, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/<KEY>", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 853}
# %cd /content/models/research
# !protoc object_detection/protos/*.proto --python_out=.
import os
os.environ['PYTHONPATH'] += ':/content/models/research/:/content/models/research/slim/'
# !python object_detection/builders/model_builder_test.py
# + colab_type="code" id="a1Y1mFlRGUEJ" colab={}
# !mkdir /content/garbage_detection/
# !mv /content/label_map.pbtxt /content/garbage_detection/
# !mv /content/generate_tfrecord.py /content/garbage_detection/
# !mv /content/train_labels.csv /content/garbage_detection/
# !mv /content/test_labels.csv /content/garbage_detection/
# !mv /content/pipeline.config /content/garbage_detection/
# + colab_type="code" id="L-BW8mFSKz_V" colab={}
# !mkdir /content/garbage_detection/data
# + colab_type="code" id="ctmUn82Utx6F" outputId="a64bd747-e065-4960-eb23-f5b74a5ae06c" executionInfo={"status": "ok", "timestamp": 1583394598723, "user_tz": -330, "elapsed": 235255, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + colab_type="code" id="fKnmTksJt71Y" outputId="328e2c14-da7d-4eac-85fa-66faea5abd67" executionInfo={"status": "ok", "timestamp": 1583394604755, "user_tz": -330, "elapsed": 241275, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd /content
# !cp drive/My\ Drive/garbage_dataset.zip .
# + colab_type="code" id="4HucYnnwHUK4" outputId="471a9b5a-d7bf-491d-bb12-368210333f3c" executionInfo={"status": "ok", "timestamp": 1583394620110, "user_tz": -330, "elapsed": 256627, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !unzip /content/garbage_dataset.zip
# !mv /content/garbage_dataset /content/garbage_detection/data/garbage_dataset
# + colab_type="code" id="6ufDU8p8HZKn" outputId="9fd60c47-2875-43f7-bd9e-d9eb9e6f9997" executionInfo={"status": "ok", "timestamp": 1583394635186, "user_tz": -330, "elapsed": 271693, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 428}
# %cd /content/garbage_detection
# !python generate_tfrecord.py --csv_input=train_labels.csv --output_path=train.record --img_path=data/garbage_dataset --label_map=label_map.pbtxt
# !python generate_tfrecord.py --csv_input=test_labels.csv --output_path=test.record --img_path=data/garbage_dataset --label_map=label_map.pbtxt
# + colab_type="code" id="32kIm0JHOeBe" colab={}
test_record_fname = '/content/garbage_detection/test.record'
train_record_fname = '/content/garbage_detection/train.record'
label_map_pbtxt_fname = '/content/garbage_detection/label_map.pbtxt'
# + colab_type="code" id="sKiB2GzkQqUy" outputId="f9ba2e35-0660-489b-df61-26610ad0215a" executionInfo={"status": "ok", "timestamp": 1583394643674, "user_tz": -330, "elapsed": 280166, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 224}
# !wget http://download.tensorflow.org/models/object_detection/ssd_mobilenet_v2_quantized_300x300_coco_2019_01_03.tar.gz
# + colab_type="code" id="ALzYe_MFRkBz" outputId="008dc345-9614-4050-bbcb-ab910ad8fed3" executionInfo={"status": "ok", "timestamp": 1583394651831, "user_tz": -330, "elapsed": 288311, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 136}
# !tar -xzvf ssd_mobilenet_v2_quantized_300x300_coco_2019_01_03.tar.gz
# + colab_type="code" id="f_xVOeQxRr_t" outputId="3a378ab2-1b18-49f2-a911-ef3736640f89" executionInfo={"status": "ok", "timestamp": 1583394651832, "user_tz": -330, "elapsed": 288304, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
fine_tune_checkpoint = "/content/garbage_detection/ssd_mobilenet_v2_quantized_300x300_coco_2019_01_03/model.ckpt"
fine_tune_checkpoint
# + colab_type="code" id="qOADBJhjSNwl" outputId="b9f210d5-3f18-4543-8cbf-ae1877f63afa" executionInfo={"status": "ok", "timestamp": 1583394661498, "user_tz": -330, "elapsed": 297959, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 255}
# %cd /content
# !wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# !unzip -o ngrok-stable-linux-amd64.zip
# + colab_type="code" id="JoAfFkLVu9I6" outputId="66b96015-0407-4a02-aebe-1bb11b4be26d" executionInfo={"status": "ok", "timestamp": 1583394661499, "user_tz": -330, "elapsed": 297949, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd /content/
# # !cp -r drive/My\ Drive/training_logs .
# # !cp -r drive/My\ Drive/trained_model .
# + colab_type="code" id="OkZcg3coTdHK" colab={}
get_ipython().system_raw(
'tensorboard --logdir /content/drive/My\ Drive/training_logs --host 0.0.0.0 --port 6006 &'
)
# + colab_type="code" id="BudAEzRSVI9q" colab={}
get_ipython().system_raw('./ngrok http 6006 &')
# + colab_type="code" id="DgjZUbOmVMvi" outputId="a04f4870-2b70-4822-c029-1292187fbd1a" executionInfo={"status": "ok", "timestamp": 1583399639582, "user_tz": -330, "elapsed": 4135, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# ! curl -s http://localhost:4040/api/tunnels | python3 -c "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# + id="BceXv9fYW1tg" colab_type="code" outputId="3f502c72-763c-4c2e-8ded-71f28d5681a3" executionInfo={"status": "ok", "timestamp": 1583407719944, "user_tz": -330, "elapsed": 8072868, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content
# !python /content/models/research/object_detection/model_main.py \
# --pipeline_config_path=/content/garbage_detection/pipeline.config \
# --model_dir=drive/My\ Drive/training_logs/ \
# --alsologtostderr \
# --num_eval_steps=100 \
# --num_train_steps=60000
# + colab_type="code" id="TA3Ji_aCZSHI" colab={}
# # !cp -r training_logs/ drive/My\ Drive/
# + colab_type="code" id="NfZw_KuJbcoe" outputId="32f2c2d0-24a4-43a7-de09-83cb05246db9" executionInfo={"status": "ok", "timestamp": 1583271432665, "user_tz": -330, "elapsed": 145456, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtA4uBTLd00uHgpDwU2R7N2nQoSkAFWdirv_T_-w=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# %cd /content
# !mkdir trained_model
# !rm -rf trained_model/saved_model/*
# + colab_type="code" id="b2nC6RWZcXvy" outputId="ce080732-a54c-445b-d997-417ebbdb3147" executionInfo={"status": "ok", "timestamp": 1575140429701, "user_tz": -330, "elapsed": 37467, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDCrZg-Xej3CNTxxqvoadpUHsPYhJum5nI1j9MHEA=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 768}
# !wget https://github.com/tensorflow/models/raw/master/research/object_detection/export_tflite_ssd_graph.py
# !wget https://github.com/opencv/opencv/raw/master/samples/dnn/tf_text_graph_ssd.py
# !wget https://raw.githubusercontent.com/opencv/opencv/master/samples/dnn/tf_text_graph_common.py
# + id="m-NYPYlrpDKH" colab_type="code" outputId="290721ec-e334-427b-b341-4806efefc0a2" executionInfo={"status": "ok", "timestamp": 1575140444130, "user_tz": -330, "elapsed": 50096, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content
# !mkdir tflite_out
# !python3 export_tflite_ssd_graph.py --pipeline_config_path=trained_model/pipeline.config --trained_checkpoint_prefix=trained_model/model.ckpt --output_directory=tflite_out/ --add_postprocessing_op=true
# + id="IlSAExLJsBc_" colab_type="code" outputId="f58a5a13-2b27-4082-c055-b79b8b083942" executionInfo={"status": "ok", "timestamp": 1575140450101, "user_tz": -330, "elapsed": 54404, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 785}
# !tflite_convert \
# --output_file=tflite_out/detect_garbage.tflite \
# --graph_def_file=tflite_out/tflite_graph.pb \
# --input_arrays=normalized_input_image_tensor \
# --output_arrays='TFLite_Detection_PostProcess','TFLite_Detection_PostProcess:1','TFLite_Detection_PostProcess:2','TFLite_Detection_PostProcess:3' \
# --input_shape=1,300,300,3 \
# --inference_type=FLOAT \
# --allow_custom_ops
# + colab_type="code" id="aisc0ImrgpBQ" outputId="b3a78fd1-ddaa-41ec-c081-3cbfc3b98389" executionInfo={"status": "ok", "timestamp": 1575140456268, "user_tz": -330, "elapsed": 59084, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDCrZg-Xej3CNTxxqvoadpUHsPYhJum5nI1j9MHEA=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 309}
# %cd /content
# !python3 tf_text_graph_ssd.py --input trained_model/frozen_inference_graph.pb --config trained_model/pipeline.config --output trained_model/graph.pbtxt
# + colab_type="code" id="ncBabYm5cCaK" colab={}
# # !zip -r trained_model.zip trained_model/
# !cp -r /content/trained_model/ /content/drive/My\ Drive/
# !cp -r /content/tflite_out/ /content/drive/My\ Drive/
# + colab_type="code" id="wbOv2yMnqlw6" outputId="78dd19eb-5c61-449f-ec07-135d7f550479" executionInfo={"status": "ok", "timestamp": 1575139732086, "user_tz": -330, "elapsed": 14863, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDCrZg-Xej3CNTxxqvoadpUHsPYhJum5nI1j9MHEA=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
import os
import glob
# Path to frozen detection graph. This is the actual model that is used for the object detection.
PATH_TO_CKPT = "/content/trained_model/frozen_inference_graph.pb"
# List of the strings that is used to add correct label for each box.
PATH_TO_LABELS = "/content/garbage_detection/label_map.pbtxt"
# If you want to test the code with your images, just add images files to the PATH_TO_TEST_IMAGES_DIR.
PATH_TO_TEST_IMAGES_DIR = "/content/garbage_detection/data/garbage_dataset"
# assert os.path.isfile(pb_fname)
assert os.path.isfile(PATH_TO_LABELS)
TEST_IMAGE_PATHS = glob.glob(os.path.join(PATH_TO_TEST_IMAGES_DIR, "*.jpg"))[-8:]
assert len(TEST_IMAGE_PATHS) > 0, 'No image found in `{}`.'.format(PATH_TO_TEST_IMAGES_DIR)
print(TEST_IMAGE_PATHS)
# + colab_type="code" id="h55xYwCwtmeM" outputId="85460f4e-8c0a-4d70-b15e-83669f5cf9bf" executionInfo={"status": "ok", "timestamp": 1575139782629, "user_tz": -330, "elapsed": 22798, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDCrZg-Xej3CNTxxqvoadpUHsPYhJum5nI1j9MHEA=s64", "userId": "02140721307530816983"}} colab={"base_uri": "https://localhost:8080/", "height": 1000, "output_embedded_package_id": "1zUcSjjZaZO6KeLps42JCCqzmsA-P1XQG"}
# %cd /content/models/research/object_detection
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
num_classes=1
# This is needed since the notebook is stored in the object_detection folder.
sys.path.append("..")
from object_detection.utils import ops as utils_ops
# This is needed to display the images.
# %matplotlib inline
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as vis_util
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(PATH_TO_CKPT, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
label_map = label_map_util.load_labelmap(PATH_TO_LABELS)
categories = label_map_util.convert_label_map_to_categories(
label_map, max_num_classes=num_classes, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
def load_image_into_numpy_array(image):
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
# Size, in inches, of the output images.
IMAGE_SIZE = (12, 8)
def run_inference_for_single_image(image, graph):
with graph.as_default():
with tf.Session() as sess:
# Get handles to input and output tensors
ops = tf.get_default_graph().get_operations()
all_tensor_names = {
output.name for op in ops for output in op.outputs}
tensor_dict = {}
for key in [
'num_detections', 'detection_boxes', 'detection_scores',
'detection_classes', 'detection_masks'
]:
tensor_name = key + ':0'
if tensor_name in all_tensor_names:
tensor_dict[key] = tf.get_default_graph().get_tensor_by_name(
tensor_name)
if 'detection_masks' in tensor_dict:
# The following processing is only for single image
detection_boxes = tf.squeeze(
tensor_dict['detection_boxes'], [0])
detection_masks = tf.squeeze(
tensor_dict['detection_masks'], [0])
# Reframe is required to translate mask from box coordinates to image coordinates and fit the image size.
real_num_detection = tf.cast(
tensor_dict['num_detections'][0], tf.int32)
detection_boxes = tf.slice(detection_boxes, [0, 0], [
real_num_detection, -1])
detection_masks = tf.slice(detection_masks, [0, 0, 0], [
real_num_detection, -1, -1])
detection_masks_reframed = utils_ops.reframe_box_masks_to_image_masks(
detection_masks, detection_boxes, image.shape[0], image.shape[1])
detection_masks_reframed = tf.cast(
tf.greater(detection_masks_reframed, 0.5), tf.uint8)
# Follow the convention by adding back the batch dimension
tensor_dict['detection_masks'] = tf.expand_dims(
detection_masks_reframed, 0)
image_tensor = tf.get_default_graph().get_tensor_by_name('image_tensor:0')
# Run inference
output_dict = sess.run(tensor_dict,
feed_dict={image_tensor: np.expand_dims(image, 0)})
# all outputs are float32 numpy arrays, so convert types as appropriate
output_dict['num_detections'] = int(
output_dict['num_detections'][0])
output_dict['detection_classes'] = output_dict[
'detection_classes'][0].astype(np.uint8)
output_dict['detection_boxes'] = output_dict['detection_boxes'][0]
output_dict['detection_scores'] = output_dict['detection_scores'][0]
if 'detection_masks' in output_dict:
output_dict['detection_masks'] = output_dict['detection_masks'][0]
return output_dict
for image_path in TEST_IMAGE_PATHS:
image = Image.open(image_path)
# the array based representation of the image will be used later in order to prepare the
# result image with boxes and labels on it.
image_np = load_image_into_numpy_array(image)
# Expand dimensions since the model expects images to have shape: [1, None, None, 3]
image_np_expanded = np.expand_dims(image_np, axis=0)
# Actual detection.
output_dict = run_inference_for_single_image(image_np, detection_graph)
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
category_index,
instance_masks=output_dict.get('detection_masks'),
use_normalized_coordinates=True,
line_thickness=2)
plt.figure(figsize=IMAGE_SIZE)
plt.imshow(image_np)
# + colab_type="code" id="9h-Xf68RuYLL" colab={}
| model/garbage_detection_quantized.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from netCDF4 import Dataset, MFDataset, num2date
import pandas as pd
import numpy as np
from zipfile import ZipFile
import matplotlib.pyplot as plt
import matplotlib as ply
from datetime import datetime
archive = ZipFile("../data/SensorNetcdfOutput.zip",'r')
print(archive.namelist())
nc_data = archive.open('air_temperature.nc')
# Dataset(nc_data) # failed to work :()
at_data = Dataset("../data/air_temperature.nc")
at_data
airtemp = at_data.variables['air_temperature']
plt.plot(airtemp)
# +
print(type(at_data.variables['time'][:]))
at = pd.DataFrame(at_data.variables['air_temperature'][:],
at_data.variables['time'][:])
plt.plot(at)
# -
| miscellaneous/zipnctemp-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
# -
mouse_metadata
mouse_metadata["Mouse ID"].nunique()
study_results.head()
merged_df = pd.merge(mouse_metadata, study_results, on="Mouse ID", how="inner")
merged_df.head()
# This is a join check
pd.merge(mouse_metadata, study_results, on="Mouse ID", how="outer").isnull().sum()
# Checking the number of mice in the DataFrame.
merged_df["Mouse ID"].nunique()
merged_df
# +
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
dupes = merged_df[merged_df.duplicated(["Mouse ID", "Timepoint"])]
dupes
# -
merged_df[merged_df["Mouse ID"]=="g989"]
merged_df["Mouse ID"]=="g989"
# Optional: Get all the data for the duplicate mouse ID.
dupes = merged_df[merged_df.duplicated(["Mouse ID", "Timepoint"])]
dupes
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
deduped_df = merged_df.drop_duplicates(["Mouse ID", "Timepoint"])
deduped_df
# Checking the number of mice in the clean DataFrame.
deduped_df["Mouse ID"].nunique()
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# This method is the most straightforward, creating multiple series and putting them all together at the end.
mouse_stats = deduped_df.groupby(["Drug Regimen"])["Tumor Volume (mm3)"]
tumor_mean = mouse_stats.mean()
tumor_median = mouse_stats.median()
tumor_variance = mouse_stats.var()
tumor_std = mouse_stats.std()
tumor_sem = mouse_stats.sem()
pd.DataFrame({"Tumor Mean" : tumor_mean,
"Tumor Median" : tumor_median,
"Tumor Variance" : tumor_variance,
"Tumor Standard Deviation" : tumor_std,
"Tumor SEM" : tumor_sem})
# -
tumor_volume.var()
tumor_volume.sem()
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# This method produces everything in a single groupby function.
mouse_stats = deduped.groupby(["Drug Regimen"])["Tumor Volume (mm3)"]
tumor_mean = mouse_stats.describe()
tumor_mean
# tumor_median = mouse_stats.median()
# tumor_median
# tumor_mean = mouse_stats.mean()
# tumor_mean = mouse_stats.mean()
# tumor_mean = mouse_stats.mean()
# -
# ## Bar Plots
# +
# Generate a bar plot showing the number of mice per time point for each treatment throughout the course of the study using pandas.
# +
# Generate a bar plot showing the number of mice per time point for each treatment throughout the course of the study using pyplot.
# -
# ## Pie Plots
# Generate a pie plot showing the distribution of female versus male mice using pandas
# Generate a pie plot showing the distribution of female versus male mice using pyplot
plt.pie(mouse_metadata["Sex"].value_counts())
mouse_metadata["Sex"].value_counts()
# ## Quartiles, Outliers and Boxplots
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens. Calculate the IQR and quantitatively determine if there are any potential outliers.
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
# -
# ## Line and Scatter Plots
# Generate a line plot of time point versus tumor volume for a mouse treated with Capomulin
# +
# Generate a scatter plot of mouse weight versus average tumor volume for the Capomulin regimen
# -
# ## Correlation and Regression
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
| Pymaceuticals/pymaceuticals_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#this script will create a graph to be used for the dfs
#for future, we can develop JSON file saving method instead of using pickle
# +
import pickle
import pandas as pd
pd.set_option('max_colwidth', 200)
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.colors as pltc
import seaborn as sns
sns.set()
import os
from os import listdir
from os.path import isfile, join
import csv
import numpy as np
# %matplotlib inline
from random import sample
from math import sin, cos, sqrt, atan2, radians, acos
import numpy as np
import math
import copy
import time
from datetime import datetime, timedelta
def save_obj(obj, name ):
with open('obj/'+ name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name ):
with open('obj/' + name + '.pkl', 'rb') as f:
return pickle.load(f)
# -
#stop_times table only weekday schedules
#arrival_time_fix and departure_time_fix are the times in seconds
trip_table_fixed = pd.read_csv(r'trip_table_fixed.csv', sep=",", engine='python')
# +
########OPTIONAL##########
#select only the 2 hour time frame of 7-9 am from trip_table_fixed
#if the tables are not shrunk, the dfs module will take exponentially longer, and may not finish
trip_table_fixed = trip_table_fixed[trip_table_fixed['arrival_time_fix'].between(25200, 32400, inclusive=True)]
# -
trip_table_fixed.head()
#create graph
graph = {}
unique_table=trip_table_fixed.groupby(['route_short_name','trip_id', 'stop_id_time']).size().reset_index()
for index, row in unique_table.iterrows():
bus_name = unique_table.loc[index, 'route_short_name']
graph[bus_name] = {}
for index, row in unique_table.iterrows():
bus_name = unique_table.loc[index, 'route_short_name']
bus_trip_id = unique_table.loc[index, 'trip_id']
stop_id = unique_table.loc[index, 'stop_id_time']
graph[bus_name][bus_trip_id] = set([])
graph_ready = copy.deepcopy(graph)
graph_ready
#create graph form
#bus#:set([bus_stops])
for index, row in trip_table_fixed.iterrows():
route_name = row['route_short_name']
stop_id = row['stop_id_time']
trip_id = row['trip_id']
try:
graph_ready[route_name][trip_id].add(stop_id)
except:
print(route_name, stop_id, trip_id)
break
# +
#since the graph is pre ordered in time, no need to put in any time conditions in the loop!
for key in graph_ready:
for trip_id in graph_ready[key]:
graph_temp = sorted(list(graph_ready[key][trip_id]), key=lambda x: int(x.split('_')[1]))
graph_ready[key][trip_id] = graph_temp
# +
#graph_ready
#save_obj(graph_ready, 'graph_ready')
#graph_ready=load_obj('graph_ready')
# -
count = 0
count_list = []
graph_final = {}
node = trip_table_fixed['stop_id_time'].unique()
for stop_id_time in node:
stop_id = stop_id_time.split('_')[0]
stop_time = int(stop_id_time.split('_')[1]) #current time
#wait time is 30 min 60*30
stop_time_30 = stop_time+1800
graph_final[stop_id_time] = []
for bus_name in graph_ready:
for trip_id in graph_ready[bus_name]:
stops_list = graph_ready[bus_name][trip_id]
stops_only_list =[s.split('_', 1)[0] for s in stops_list]
matching_index = [i for i,d in enumerate(stops_only_list) if d==stop_id]
#check if the trip_id could possibly have two hits on a stop_id
if 1 < len(matching_index):
for index in matching_index:
current_stop = stops_list[index]
current_stop_time = int(current_stop.split('_')[1])
#check if the stop time for that bus is within 30 min wait
if stop_time <= current_stop_time and current_stop_time <= stop_time_30:
#if this trip_id is the last stop, continue to the next trip_id
try:
next_stop = stops_list[index+1]
next_stop_id = next_stop.split('_')[0]
next_stop_time = next_stop.split('_')[1]
except:
continue
travel_time = int(next_stop_time) - int(stop_time)
#find full bus name and put it in
row=trip_table_fixed.loc[(trip_table_fixed['trip_id'] == trip_id) & (trip_table_fixed['stop_id_time'] == next_stop)]
full = row['full_bus_name'].values[0]
dic = {'destination': next_stop,
'travel_time': travel_time,
'trip_id': trip_id,
'bus_name': bus_name,
'full_bus_name': full}
###final step
graph_final[stop_id_time].append(dic)
else:
continue
else:
continue
else:
try:
index = stops_only_list.index(stop_id)
except:
continue
current_stop = stops_list[index]
current_stop_time = int(current_stop.split('_')[1])
#check if the stop time for that bus is within 30 min wait
if stop_time <= current_stop_time and current_stop_time <= stop_time_30:
#if this trip_id is the last stop, continue to the next trip_id
try:
next_stop = stops_list[index+1]
next_stop_id = next_stop.split('_')[0]
next_stop_time = next_stop.split('_')[1]
except:
continue
travel_time = int(next_stop_time) - int(stop_time)
#find full bus name and put it in
row=trip_table_fixed.loc[(trip_table_fixed['trip_id'] == trip_id) & (trip_table_fixed['stop_id_time'] == next_stop)]
full = row['full_bus_name'].values[0]
dic = {'destination': next_stop,
'travel_time': travel_time,
'trip_id': trip_id,
'bus_name': bus_name,
'full_bus_name': full}
###final step
graph_final[stop_id_time].append(dic)
else:
continue #for trip_id in graph[bus_name]:
# +
#graph_ready
#save_obj(graph_final, 'graph_final')
# +
#graph_final
# -
#this is the graph without any stop destination. Most are because the time limit is until 9 am but some are not.
#must further develop to figure out how to make the graph more accurate
for stop_with_time in graph_final:
if graph_final[stop_with_time] == []:
print(stop_with_time)
#this is one of the stops in the blank.
#it is a drop off but it should lead to the next stop afterwards, but does not.
#must figure out why some stops are acting this way
trip_table_fixed[trip_table_fixed['stop_id_time']=='2397_27840']
trip_table_fixed[trip_table_fixed['trip_id']==901249]
trip_table_fixed[trip_table_fixed['stop_id']==2397]
# +
#https://www.cityofmadison.com/metro/routes-schedules/route-22
# -
df=trip_table_fixed.loc[(trip_table_fixed['trip_id'] == 904846) & (trip_table_fixed['stop_id_time'] == '1759_25281')]
df['full_bus_name'].values[0]
| final_code_JINWOOLEE/create_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Euler Problem 40
# ================
#
# An irrational decimal fraction is created by concatenating the positive integers:
#
# 0.123456789101112131415161718192021...
#
# It can be seen that the 12th digit of the fractional part is 1.
#
# If $d_n$ represents the nth digit of the fractional part, find the value of the following expression.
#
# $$d_1 \times d_{10} \times d_{100} \times d_{1000} \times d_{10000} \times d_{100000} \times d_{1000000}$$
# +
def champernowne_digit(n):
digits = 1
power = 1
while n > 9 * digits * power:
n -= 9 * digits * power
digits += 1
power *= 10
return int(str(power + (n-1) // digits)[(n-1) % digits])
from functools import reduce
print(reduce(lambda x,y: x*y, (champernowne_digit(10**k) for k in range(7))))
# -
| Euler 040 - Champernowne's constant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Detection with SSD
# ### Here we demostrate detection on example images using SSD with PyTorch
# +
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import torch
import torch.nn as nn
import torch.backends.cudnn as cudnn
from torch.autograd import Variable
import torch.utils.data as data
import torchvision.transforms as transforms
from torch.utils.serialization import load_lua
import numpy as np
import cv2
if torch.cuda.is_available():
torch.set_default_tensor_type('torch.cuda.FloatTensor')
from ssd import build_ssd
# from models import build_ssd as build_ssd_v1 # uncomment for older pool6 model
# %matplotlib inline
from matplotlib import pyplot as plt
from data import AnnotationTransform_handles, HandlesDetection
from data import AnnotationTransformVOC, VOCDetection
# -
# ## Hyper Parameters
# Specify input, weights, paths
#
# +
dataset_name = 'handles'
data_root = '/home/mil/chou/STORAGE/dataset/'
#weights_path = '../weights/ssd300_mAP_77.43_v2.pth'
weights_path = '../weights/ssd300_handles75000.pth'
confidence_thres = 0.1
VOC_CLASSES = ( # always index 0
'aeroplane', 'bicycle', 'bird', 'boat',
'bottle', 'bus', 'car', 'cat', 'chair',
'cow', 'diningtable', 'dog', 'horse',
'motorbike', 'person', 'pottedplant',
'sheep', 'sofa', 'train', 'tvmonitor')
HANDLES_CLASSES = ('door', 'handle')
if 'handle' in dataset_name:
data_root = data_root + '/handles'
num_classes = len(HANDLES_CLASSES) + 1 # +1 background
testset = HandlesDetection(data_root, None, AnnotationTransform_handles(), dataset='test')
labelmap = HANDLES_CLASSES
else:
num_classes = len(VOC_CLASSES) + 1 # +1 background
testset = VOCDetection(data_root, [('2007', 'test')], None, AnnotationTransformVOC())
labelmap = VOC_CLASSES
# -
#
# ## Build SSD300 in Test Phase
# 1. Build the architecture, specifyingsize of the input image (300),
# and number of object classes to score (21 for VOC dataset)
# 2. Next we load pretrained weights on the VOC0712 trainval dataset
net = build_ssd('test', 300, num_classes,forward_classes=3 if 'handle' in data_root else 21) # initialize SSD with VOC
net.load_weights(weights_path)
net.eval()
print('Finished loading model!')
# +
# image = cv2.imread('./data/example.jpg', cv2.IMREAD_COLOR) # uncomment if dataset not downloaded
img_id = 13
image = testset.pull_image(img_id)
rgb_image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# View the sampled input image before transform
plt.figure(figsize=(10,10))
plt.imshow(rgb_image)
plt.show()
# -
#
#
# ## Pre-process the input.
# #### Using the torchvision package, we can create a Compose of multiple built-in transorm ops to apply
# For SSD, at test time we use a custom BaseTransform callable to
# resize our image to 300x300, subtract the dataset's mean rgb values,
# and swap the color channels for input to SSD300.
x = cv2.resize(image, (300, 300)).astype(np.float32)
x -= (104.0, 117.0, 123.0)
x = x.astype(np.float32)
x = x[:, :, ::-1].copy()
plt.imshow(x)
x = torch.from_numpy(x).permute(2, 0, 1)
# ## SSD Forward Pass
# ### Now just wrap the image in a Variable so it is recognized by PyTorch autograd
xx = Variable(x.unsqueeze(0)) # wrap tensor in Variable
if torch.cuda.is_available():
xx = xx.cuda()
y = net(xx)
# ## Parse the Detections and View Results
# Filter outputs with confidence scores lower than a threshold
# Here we choose 60%
# +
top_k=10
plt.figure(figsize=(10,10))
colors = plt.cm.hsv(np.linspace(0, 1, 21)).tolist()
plt.imshow(rgb_image) # plot the image for matplotlib
currentAxis = plt.gca()
detections = y.data
# scale each detection back up to the image
scale = torch.Tensor(rgb_image.shape[1::-1]).repeat(2)
for i in range(detections.size(1)):
j = 0
while detections[0,i,j,0] >= confidence_thres:
score = detections[0,i,j,0]
label_name = labelmap[i-1]
display_txt = '%s: %.2f'%(label_name, score)
pt = (detections[0,i,j,1:]*scale).cpu().numpy()
coords = (pt[0], pt[1]), pt[2]-pt[0]+1, pt[3]-pt[1]+1
color = colors[i]
currentAxis.add_patch(plt.Rectangle(*coords, fill=False, edgecolor=color, linewidth=2))
currentAxis.text(pt[0], pt[1], display_txt, bbox={'facecolor':color, 'alpha':0.5})
j+=1
# -
| demo/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
data = pd.read_csv('https://raw.githubusercontent.com/insaid2018/CDF-Certification-Quiz-Data/master/Abalone/02-Data.csv')
data.head()
data=data.drop(['Sex'],axis=1)
data.columns
X=data.loc[:,data.columns!='Adult']
X
Y=data['Adult']
Y
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,test_size=0.3,random_state=3)
# +
from sklearn.linear_model import LogisticRegression
log_reg=LogisticRegression()
log_reg.fit(X_train,Y_train)
y_pred_train=log_reg.predict(X_train)
y_pred_test=log_reg.predict(X_test)
# -
from sklearn.metrics import mean_squared_error
mean_squared_error(Y_train,y_pred_train)
mean_squared_error(Y_test,y_pred_test)
from sklearn.metrics import accuracy_score
accuracy_score(Y_test,y_pred_test)
| INSAID/Course_Material/Term1-ML/Logistic_Regression/Quiz_question_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="gyu1ziDNoVpx"
# ### Data preparation
# + id="IxT1MnlGft6b" executionInfo={"status": "ok", "timestamp": 1625164428146, "user_tz": -180, "elapsed": 9, "user": {"displayName": "<NAME>\u011flam", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="1ESzeDOqfwxc" executionInfo={"status": "ok", "timestamp": 1625164428538, "user_tz": -180, "elapsed": 399, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}}
url = 'https://raw.githubusercontent.com/aParsecFromFuture/COVID-19-stats/main/Turkey/datasets/genel_koronavir%C3%BCs_tablosu.csv'
df_covid = pd.read_csv(url, index_col='Tarih', parse_dates=['Tarih'])
df_covid = df_covid.iloc[::-1]
# + [markdown] id="7soNRzvwoYq8"
# ### Line Plot
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="azSgt7TLfyqC" executionInfo={"status": "ok", "timestamp": 1625164429002, "user_tz": -180, "elapsed": 468, "user": {"displayName": "<NAME>\u011flam", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}} outputId="6e3ef28f-702d-459b-f733-78a68e50d4a2"
fig, ax = plt.subplots(1, 1, figsize=(20, 4))
ax.set_title('Türkiye covid-19 verileri', fontsize=20)
ax.plot(df_covid.loc['2021-01-01':, 'Bugünkü Vaka Sayısı'], label='vaka sayısı')
ax.plot(df_covid.loc['2021-01-01':, 'Bugünkü İyileşen Sayısı'], label='iyileşen sayısı')
ax.legend()
ax.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 725} id="_s0g2y4KfzKE" executionInfo={"status": "ok", "timestamp": 1625164429809, "user_tz": -180, "elapsed": 816, "user": {"displayName": "<NAME>\u011flam", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}} outputId="b2e46bea-dc4b-4638-bf64-11bbdec87562"
fig, ax = plt.subplots(3, 1, figsize=(20, 12))
from_date = '2021-06-01'
dataset = [df_covid['Bugünkü Vaka Sayısı'], df_covid['Bugünkü Vefat Sayısı'], df_covid['Bugünkü İyileşen Sayısı']]
moving_average = [data.rolling(window=7).mean().shift(-3) for data in dataset]
labels = ['vaka sayısı', 'vefat sayısı', 'iyileşen sayısı']
ax[0].set_title('Haziran ayı istatistikleri', fontsize=24)
for i in range(3):
ax[i].plot(dataset[i][from_date:], color='blue', label=labels[i])
ax[i].plot(moving_average[i][from_date:], color='red', label='7 günlük ort')
ax[i].grid()
ax[i].legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 288} id="hMu7n08XmTZw" executionInfo={"status": "ok", "timestamp": 1625164429813, "user_tz": -180, "elapsed": 25, "user": {"displayName": "<NAME>\u011flam", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}} outputId="aa940ba4-0e87-4d45-db3c-7926f349d9c8"
cols = ['Hastalarda Zatürre Oranı (%)', 'Ağır Hasta Sayısı', 'Bugünkü Vaka Sayısı', 'Bugünkü Hasta Sayısı', 'Bugünkü Test Sayısı', 'Bugünkü Vefat Sayısı', 'Bugünkü İyileşen Sayısı']
from_date = '01-01-2021'
corr_matrix = df_covid.loc[from_date:, cols].corr()
corr_matrix
# + [markdown] id="HWXaNi10V_8i"
# ### Correlation Matrix
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="KE8tRSUpokn0" executionInfo={"status": "ok", "timestamp": 1625164430475, "user_tz": -180, "elapsed": 682, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}} outputId="7fdeffa9-65ed-4c79-f58d-47a9221d1193"
fig, ax = plt.subplots(1, 1, figsize=(6, 6))
ax.set_xticklabels([''] + cols)
ax.set_yticklabels([''] + cols)
ax.matshow(corr_matrix, cmap='RdBu')
plt.xticks(rotation='vertical')
plt.show()
# + [markdown] id="FJDnwdKLpfn8"
# ### Bar Chart
# + colab={"base_uri": "https://localhost:8080/", "height": 395} id="CJb-OEvXS-QX" executionInfo={"status": "ok", "timestamp": 1625164430477, "user_tz": -180, "elapsed": 37, "user": {"displayName": "<NAME>\u011flam", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjGflTbfgey4mTROD7KDGB6bucIDqaxTwVBph_4=s64", "userId": "00651573771185432140"}} outputId="c13c4ac8-2b78-4e4f-b121-56cff483613b"
fig, ax = plt.subplots(1, 1, figsize=(20, 6))
from_date = '2021-03-01'
rpatient = df_covid.loc[from_date:, 'Bugünkü Hasta Sayısı'] / df_covid.loc[from_date:, 'Bugünkü Test Sayısı']
ax.set_title('Günlük hasta / Test sayısı', fontsize=20)
ax.bar(rpatient.index, rpatient.values)
plt.show()
# + id="f_NyL3lwTTBi"
| Turkey/general_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import re
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from glob import glob
# # Evaluation of pyAscore performance on the Marx Synthetic Peptides
# ## 1) Intro
# In order to test pyAscore with known peptide sequences, we will be looking at a synthetic peptide set developed by Marx *et al.* (2013). This has been measured multiple times -- high resolution HCD and ETD in PXD000138 and low resolution CID in PXD000759. These datasets provide a great test bed due to the large amount of spectra. We searched both datasets with the recommended parameters for the individual instrument setups. Then, we used multiple pyAscore parameters on the dataset to determine if the algorithm could be effectively tailored to the instrument parameters.
# Here, we will first look at the total number of PSMs which correspond to the original peptide library built in Marx *et al.*. Once we have this set of peptide, we will build a comparison plot of the FLRs between parameter choices.
# ## 2) Load PSM data and evaluate number of correct matches
# After taking a glance at the literature, it appears that most papers treat each individual PSM as a independent sample for calculating false localization rate. Thus, we will also focus on PSMs here but we will try to give an overview of underlying sequence effects at some point. Given that these are sythetic peptide runs, we know exactly which peptides should be where. Thus, to be stringent, we will only evaluate the counts of PSMs which pass a 1% global peptide FDR threshold, have a single phosphorylation, and which occur in their correct sample.
# ### 2.1) Load seed peptide data
# Here we load and prepare the list of seed peptides sequences. Since the detected peptides will be derived from these, it is important to make the sequences into a propper regex which can validate that a matched peptide definitely came from the library.
seed_peptides = pd.read_csv("../aux/PXD000138_seed_peptides.csv")[["Library", "Seed Peptide Sequence",
"Phosphosite Position", "Length"]]
seed_peptides.columns = ["library", "seedSequence", "truePos", "trueLength"]
seed_peptides["validationRegex"] = seed_peptides.seedSequence.str.replace("[A-Z]?p[STY][A-Z]?",
"[A-Z]?[STY][A-Z]?",
regex=True)
# ### 2.2) Load PSMs
# In the FASTA file, the batch is labeled with letters and two numbers
# which codes batches 1-96 as 8 batches of 12.
def determine_batch(query):
letter_to_number = {l : ind for ind, l in enumerate("ABCDEFGH")}
query = query.split(":")[1]
return 12*letter_to_number[query[0]] + int(query[1:])
# #### 2.2.1) PXD000138-HCD
# +
# 1. Load data and filter for high confidence PSMs
hcd_psms = pd.read_csv("../results/search/PXD000138-HCD/mokapot.psms.txt", sep="\t")
hcd_psms["Dataset"] = "High Res. HCD"
hcd_psms["Peptide"] = hcd_psms["Peptide"].str.replace("^.\.", "", regex=True)\
.str.replace("\..$", "", regex=True)
hcd_psms = hcd_psms[hcd_psms["mokapot q-value"] < 0.01]
# 2. Validate peptide detections using seed peptides
hcd_psms["Library"] = hcd_psms["SpecId"].str.replace("_.*", "", regex=True).astype(int)
hcd_psms = hcd_psms.join(seed_peptides.set_index("library")["validationRegex"],
on="Library")
at_most_one_phospho = hcd_psms["Peptide"].str.count("79") <= 1
from_seed = hcd_psms.apply(lambda s: re.match(s["validationRegex"],
re.sub("[^A-Z]", "", s["Peptide"])) is not None,
axis=1)
hcd_psms = hcd_psms[np.logical_and(at_most_one_phospho, from_seed)]
# 3. Retain relevant columns
hcd_psms = hcd_psms[["Dataset", "SpecId", "ScanNr", "Peptide", "Library"]]
# -
# #### 2.2.2) PXD000138-ETD
# +
# 1. Load data and filter for high confidence PSMs
etd_psms = pd.read_csv("../results/search/PXD000138-ETD/mokapot.psms.txt", sep="\t")
etd_psms["Dataset"] = "High Res. ETD"
etd_psms["Peptide"] = etd_psms["Peptide"].str.replace("^.\.", "", regex=True)\
.str.replace("\..$", "", regex=True)
etd_psms = etd_psms[etd_psms["mokapot q-value"] < 0.01]
# 2. Validate peptide detections using seed peptides
etd_psms["Library"] = etd_psms["SpecId"].str.extract("(?<=run)([0-9]+)").astype(int)
etd_psms = etd_psms.join(seed_peptides.set_index("library")["validationRegex"],
on="Library")
at_most_one_phospho = etd_psms["Peptide"].str.count("79") <= 1
from_seed = etd_psms.apply(lambda s: re.match(s["validationRegex"],
re.sub("[^A-Z]", "", s["Peptide"])) is not None,
axis=1)
etd_psms = etd_psms[np.logical_and(at_most_one_phospho, from_seed)]
# 3. Retain relevant columns
etd_psms = etd_psms[["Dataset", "SpecId", "ScanNr", "Peptide", "Library"]]
# -
# #### 2.2.3) PXD000759
# +
# 1. Load data and filter for high confidence PSMs
cid_psms = pd.read_csv("../results/search/PXD000759/mokapot.psms.txt", sep="\t")
cid_psms["Dataset"] = "High Res. CID"
cid_psms["Peptide"] = cid_psms["Peptide"].str.replace("^.\.", "", regex=True)\
.str.replace("\..$", "", regex=True)
cid_psms = cid_psms[cid_psms["mokapot q-value"] < 0.01]
# 2. Validate peptide detections using seed peptides
cid_psms["Library"] = cid_psms["SpecId"].str.replace("_.*", "", regex=True)\
.str.replace("lib", "").astype(int)
cid_psms = cid_psms.join(seed_peptides.set_index("library")["validationRegex"],
on="Library")
at_most_one_phospho = cid_psms["Peptide"].str.count("79") <= 1
from_seed = cid_psms.apply(lambda s: re.match(s["validationRegex"],
re.sub("[^A-Z]", "", s["Peptide"])) is not None,
axis=1)
cid_psms = cid_psms[np.logical_and(at_most_one_phospho, from_seed)]
# 3. Retain relevant columns
cid_psms = cid_psms[["Dataset", "SpecId", "ScanNr", "Peptide", "Library"]]
# -
# ### 2.3) Total detections
# +
hcd_psm_counts = hcd_psms.groupby(["Dataset", "Library"])\
.apply(lambda df: pd.Series({"Total" : df.shape[0],
"Phospho" : df["Peptide"].str.contains("79").sum()}))\
.reset_index()
etd_psm_counts = etd_psms.groupby(["Dataset", "Library"])\
.apply(lambda df: pd.Series({"Total" : df.shape[0],
"Phospho" : df["Peptide"].str.contains("79").sum()}))\
.reset_index()
cid_psm_counts = cid_psms.groupby(["Dataset", "Library"])\
.apply(lambda df: pd.Series({"Total" : df.shape[0],
"Phospho" : df["Peptide"].str.contains("79").sum()}))\
.reset_index()
# +
fig = plt.figure(figsize=[6, 9])
sns.barplot(data=pd.concat([
hcd_psm_counts,
cid_psm_counts,
etd_psm_counts
]).melt(id_vars=["Dataset", "Library"],
var_name="Subset",
value_name="PSMs"),
x="Dataset",
y="PSMs",
hue="Subset",
estimator=sum,
ci=None,
palette=["#461554", "#56c566"],
alpha=.75)
plt.xticks(range(3), ["High Res.\nHCD", "Low Res.\nCID", "High Res.\nETD"])
plt.xlabel("")
plt.ylabel("Number of PSMs",
size=30)
#plt.ylim(0, 210000)
plt.tick_params(labelsize=20)
plt.legend(fontsize=24,
frameon=False,
bbox_to_anchor=(.7, 1))
sns.despine()
plt.savefig("figures/marx_synthetic_peptide_detections.svg",
bbox_inches="tight",
facecolor="white")
plt.show()
# -
# The HCD and CID datasets have about the same number of total PSMs but the second dataset seems to have more singly phosphorylated PSMs. The sample series that we downloaded for the Low Res. CID dataset was labeled as not being phophoenriched, but it is possible something else is increasing the total amount of phospho-PSMs in the datasets. Sadly, it appears that the ETD dataset has a very low number of detections. While this is likely enough to evaluate false localization rate, the amount data passing a given Ascore threshold may be quite low.
# +
fig, axes = plt.subplots(figsize=[12, 4], ncols=3)
# PXD000138-HCD
sns.scatterplot(data=hcd_psm_counts,
x="Total",
y="Phospho",
palette=["#30708d"],
lw=0,
alpha=.75,
ax=axes[0])
axes[0].set_title("High Res. HCD", size=20)
axes[0].set_xlim(0, 5000)
axes[0].set_ylim(0, 5000)
# PXD000759
sns.scatterplot(data=cid_psm_counts,
x="Total",
y="Phospho",
palette=["#30708d"],
lw=0,
alpha=.75,
ax=axes[1])
axes[1].set_title("Low Res. CID", size=20)
axes[1].set_xlim(0, 10000)
axes[1].set_ylim(0, 10000)
# PXD000138-ETD
sns.scatterplot(data=etd_psm_counts,
x="Total",
y="Phospho",
palette=["#30708d"],
lw=0,
alpha=.75,
ax=axes[2])
axes[2].set_title("High Res. ETD", size=20)
axes[2].set_xlim(0, 2500)
axes[2].set_ylim(0, 2500)
# Common features
for ax in axes:
ax.plot([0, 10000],
[0, 5000],
c="black",
lw=3,
alpha=.5,
linestyle="--")
ax.tick_params(labelsize=16)
ax.set_xlabel("Total PSMs", size=20)
ax.set_ylabel("Phospho PSMs", size=20)
fig.tight_layout(pad=3.0)
sns.despine()
plt.savefig("figures/marx_synthetic_peptide_phospho_vs_total.svg",
bbox_inches="tight",
facecolor="white")
plt.show()
# -
# The above plots give a look at numbers of phospho-PSMs vs total psms at the run level. Theoretically, according to the description of how the data was built, phosphorylated peptides should make up about half the total peptides in the sample (thus the y=x/2 line). Samples in the CID dataset do seem to be consistently high quality in their number of phospho-PSMs, with the slight enrichment for phosphorylation apparent.
# ## 3) Load pyAscore data and evaluated False Localization Rates
# For each dataset, we evaluated 2 different parameter settings -- one where the MS2 error for peaks was set at +-0.5 (wide) and one where it was set at +-0.05. The hypothesis is that tailoring the MS2 error to the instrument parameters will greatly increase the performance of the algorithm.
# ### 3.1) Load Ascores and match to PSMs
# +
def find_mod(seq):
for ind, aa in enumerate(re.finditer("[A-Z][^A-Z]*", seq)):
if aa.group() in ["S[80]", "T[80]", "Y[80]"]:
return ind + 1
def calculate_flr(score_array, success_array):
nclassifications = np.arange(1, success_array.shape[0] + 1)
nfailures = np.cumsum(~success_array)
flr = nfailures/nclassifications
return flr
for ind in np.arange(flr.shape[0])[::-1]:
if ind - 1 >= 0 and score_array[ind] == score_array[ind-1]:
flr[ind - 1] = flr[ind]
if ind + 1 < flr.shape[0]:
flr[ind] = min(flr[ind], flr[ind + 1])
return flr
def read_pyascore_file(f):
data = pd.read_csv(f, sep="\t")
library = re.search("((^)|(run)|(lib))[0-9]+", os.path.basename(f).split(".")[0]).group()
data["Library"] = int(re.sub("[^0-9]", "", library))
return data
def read_pyascore(glob_string, psms):
# Load data and match to PSMs
ascores = pd.concat([read_pyascore_file(f) for f in glob(glob_string)])
ascores = psms[["Library", "ScanNr"]].join(ascores.set_index(["Library", "Scan"]),
on=["Library", "ScanNr"],
how="inner")
# Find mods, fix ascores, and filter out non-ambiguous peptides
ascores["modPos"] = ascores["LocalizedSequence"].apply(find_mod)
ascores = ascores.join(seed_peptides.set_index("library").truePos,
on = "Library")
ascores["Ascores"] = ascores["Ascores"].astype(float)
ascores = ascores[~np.isinf(ascores["PepScore"])]\
.sort_values("Ascores", ascending=False)
# Calculate final FLR
success = ascores["modPos"].values == ascores["truePos"].values
ascores["FLR"] = calculate_flr(ascores["Ascores"].values, success)
ascores["Proportion"] = np.arange(1, ascores.shape[0]+1)/ascores.shape[0]
return ascores
# +
hcd_wide_pyascore = read_pyascore("../results/localization/PXD000138-HCD/*pyascore.wide.txt", hcd_psms)
hcd_wide_pyascore["Dataset"] = "High Res. HCD"
hcd_wide_pyascore["Parameters"] = "Wide MS2 Tol."
hcd_narrow_pyascore = read_pyascore("../results/localization/PXD000138-HCD/*pyascore.narrow.txt", hcd_psms)
hcd_narrow_pyascore["Dataset"] = "High Res. HCD"
hcd_narrow_pyascore["Parameters"] = "Narrow MS2 Tol."
# +
etd_wide_pyascore = read_pyascore("../results/localization/PXD000138-ETD/*pyascore.wide.txt", etd_psms)
etd_wide_pyascore["Dataset"] = "High Res. ETD"
etd_wide_pyascore["Parameters"] = "Wide MS2 Tol."
etd_narrow_pyascore = read_pyascore("../results/localization/PXD000138-ETD/*pyascore.narrow.txt", etd_psms)
etd_narrow_pyascore["Dataset"] = "High Res. ETD"
etd_narrow_pyascore["Parameters"] = "Narrow MS2 Tol."
# +
cid_wide_pyascore = read_pyascore("../results/localization/PXD000759/*pyascore.wide.txt", cid_psms)
cid_wide_pyascore["Dataset"] = "Low Res. CID"
cid_wide_pyascore["Parameters"] = "Wide MS2 Tol."
cid_narrow_pyascore = read_pyascore("../results/localization/PXD000759/*pyascore.narrow.txt", cid_psms)
cid_narrow_pyascore["Dataset"] = "Low Res. CID"
cid_narrow_pyascore["Parameters"] = "Narrow MS2 Tol."
# -
# ### 3.2) Comparing false localization rates
# +
fig, axes = plt.subplots(nrows=3, figsize=[10, 9])
def build_lineplot(df, ax):
ax.plot([0, 1],
[0.01, 0.01],
linestyle="--",
c="black",
lw=3,
alpha=.5,
zorder=0)
sns.lineplot(data=df[df.FLR < 0.05],
x="Proportion",
y="FLR",
hue="Parameters",
palette=["#461554", "#30708d"],
alpha=.6,
lw=4,
ax=ax)
sns.scatterplot(data = df[df.Ascores >= 20]\
.groupby(["Dataset", "Parameters"])[["FLR", "Proportion"]]\
.last()\
.reset_index(),
x="Proportion",
y="FLR",
hue=["Ascore == 20"]*2,
palette=["black"],
s=100,
zorder=100,
label=None,
ax=ax)
build_lineplot(pd.concat([hcd_wide_pyascore, hcd_narrow_pyascore],
ignore_index=True),
ax=axes[0])
build_lineplot(pd.concat([cid_wide_pyascore, cid_narrow_pyascore],
ignore_index=True),
ax=axes[1])
build_lineplot(pd.concat([etd_wide_pyascore, etd_narrow_pyascore],
ignore_index=True),
ax=axes[2])
for ax in axes:
ax.set_xlim(0, 1)
ax.tick_params(labelsize=20)
ax.set_yticks([0.00, 0.01, 0.02, 0.03, 0.04, 0.05])
ax.set_xlabel("")
ax.set_ylabel("")
ax.get_legend().remove()
axes[2].set_xlabel("Proportion of Scored PSMs",
size=24)
axes[1].set_ylabel("False Localization Rate",
size=24)
axes[1].legend(fontsize=24,
frameon=False,
bbox_to_anchor=(1, 1))
sns.despine()
plt.savefig("figures/marx_synthetic_peptide_flr_by_cutoff.svg",
bbox_inches="tight",
facecolor="white")
plt.show()
# -
# The above plot gives direct confirmation of our initial hypothesis. Using the wide peak tolerance for low resolution data greatly increased performance, and the narrow peak tolerance improved results for the high resolution data. This strongly suggests that tailoring the algorithm to instrument parameters is the best way to go. While it is difficult to compare datasets, due to differences in detections, we do want to note that most of the poor performance in the case of the ETD dataset is likely coming from the low number of dections and not poor algorithmic performance.
# +
ascore_matched_data = pd.concat([hcd_narrow_pyascore,
cid_wide_pyascore,
etd_narrow_pyascore])
ascore_notmatched_data = pd.concat([hcd_wide_pyascore,
cid_narrow_pyascore,
etd_wide_pyascore])
ascore_data = pd.concat([ascore_matched_data, ascore_notmatched_data])
# +
ascore_counts = ascore_matched_data.groupby(["Dataset", "Parameters"], sort=False)\
.apply(lambda df: pd.DataFrame({"Cutoff" : ["FLR $\leq$ 0.01",
"Ascore $\geq$ 20",
"Scorable"],
"Count" : [np.sum(df.FLR <= 0.01),
np.sum(df.Ascores >= 20),
df.shape[0]]}))\
.reset_index()
ascore_counts["xLabel"] = ascore_counts["Dataset"] + "\n" + ascore_counts["Parameters"]
# +
fig = plt.figure(figsize=[6, 9])
sns.barplot(data = ascore_counts,
x="xLabel",
y="Count",
hue="Cutoff",
hue_order=["FLR $\leq$ 0.01", "Ascore $\geq$ 20", "Scorable"][::-1],
palette=["#461554", "#56c566", "#30708d"],
alpha=.75)
plt.xlabel("")
plt.ylabel("Number of PSMs",
size=24)
plt.tick_params(labelsize=20)
plt.xticks(rotation=30,
ha="right")
plt.legend(fontsize=20,
frameon=False,
bbox_to_anchor=(.6, 1))
sns.despine()
plt.savefig("figures/marx_synthetic_peptide_flr_barplot.svg",
bbox_inches="tight",
facecolor="white")
plt.show()
# -
# The part about the above plot that we find very ecouraging is just how close we get to a 1% FLR with an Ascore of 20. This was the value used in the original Ascore paper and it is good to see that we can continue to recommend it going forward.
| notebook/evaluation_of_false_localization_on_marx_synthetic_peptides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="D-nk_Xwhfir9"
# Import all needed library
# + id="U4YGG9JBey8H" executionInfo={"status": "ok", "timestamp": 1636994762947, "user_tz": -480, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}}
import numpy as np
import pandas as pd
import seaborn as sns
sns.set_palette('husl')
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import metrics
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
# + [markdown] id="zc9DvJQhfmsC"
# Load the datasets
# + id="rkuydCgHfLNc" executionInfo={"status": "ok", "timestamp": 1636994813946, "user_tz": -480, "elapsed": 752, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}}
data = pd.read_csv('Iris.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="-2mrLHnefpa4" executionInfo={"status": "ok", "timestamp": 1636994869932, "user_tz": -480, "elapsed": 1648, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="2d3656bd-bbb0-4dba-9c5a-ce6f06656ef1"
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="B6s2EPOQf2Gp" executionInfo={"status": "ok", "timestamp": 1636994879418, "user_tz": -480, "elapsed": 998, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="c54c30a6-2c99-4d98-943f-18377dc8c027"
data.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="BnWRYvIZf4km" executionInfo={"status": "ok", "timestamp": 1636994885699, "user_tz": -480, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="6e0d4763-614a-43da-c906-bf9344c097fb"
data.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="Xe1cZzNRf6G2" executionInfo={"status": "ok", "timestamp": 1636994892747, "user_tz": -480, "elapsed": 494, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="90c5ec16-d814-4445-a0a2-46d99f27da14"
data['Species'].value_counts()
# + [markdown] id="0TIcLxg5f_xd"
# Now we're going to visualize the data
# + colab={"base_uri": "https://localhost:8080/", "height": 726} id="UlL4boGQf79Z" executionInfo={"status": "ok", "timestamp": 1636994983364, "user_tz": -480, "elapsed": 9739, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="76371f5c-5d77-4a1a-ba1c-b071395343e1"
tmp = data.drop('Id', axis=1)
g = sns.pairplot(tmp, hue='Species', markers='+')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="3pfP1lIPgP0R" executionInfo={"status": "ok", "timestamp": 1636994988952, "user_tz": -480, "elapsed": 1886, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="e7c4b412-86ce-486b-bf68-9d4af5057b18"
g = sns.violinplot(y='Species', x='SepalLengthCm', data=data, inner='quartile')
plt.show()
g = sns.violinplot(y='Species', x='SepalWidthCm', data=data, inner='quartile')
plt.show()
g = sns.violinplot(y='Species', x='PetalLengthCm', data=data, inner='quartile')
plt.show()
g = sns.violinplot(y='Species', x='PetalWidthCm', data=data, inner='quartile')
plt.show()
# + [markdown] id="D8ySIeSZgUnO"
# Modeling with scikit-learn
# + colab={"base_uri": "https://localhost:8080/"} id="-56FJZjTgTGi" executionInfo={"status": "ok", "timestamp": 1636995011281, "user_tz": -480, "elapsed": 567, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="9e5f7bbb-c659-46aa-f55e-87c934c961a4"
X = data.drop(['Id', 'Species'], axis=1)
y = data['Species']
# print(X.head())
print(X.shape)
# print(y.head())
print(y.shape)
# + [markdown] id="lkFwXzXjgeDv"
# Train and test on the same dataset
#
# Note:
# - This method is not suggested since the end goal is to predict iris species using a dataset the model has not seen before.
# - There is also a risk of overfitting the training data.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="LNt33rrNgY3h" executionInfo={"status": "ok", "timestamp": 1636995092608, "user_tz": -480, "elapsed": 719, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="efcdd615-b5b3-44c3-9148-6377425a3f8a"
# experimenting with different n values
k_range = list(range(1,26))
scores = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X, y)
y_pred = knn.predict(X)
scores.append(metrics.accuracy_score(y, y_pred))
plt.plot(k_range, scores)
plt.xlabel('Value of k for KNN')
plt.ylabel('Accuracy Score')
plt.title('Accuracy Scores for Values of k of k-Nearest-Neighbors')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="YY42EGAlgsnG" executionInfo={"status": "ok", "timestamp": 1636995111635, "user_tz": -480, "elapsed": 533, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="72649da1-1e4a-4554-d2f4-489d20d9d3e1"
logreg = LogisticRegression()
logreg.fit(X, y)
y_pred = logreg.predict(X)
print(metrics.accuracy_score(y, y_pred))
# + [markdown] id="3m-X3N2DgzhO"
# Split the dataset into a training set and a testing set
# + colab={"base_uri": "https://localhost:8080/"} id="nqr64uI1gxYZ" executionInfo={"status": "ok", "timestamp": 1636995129679, "user_tz": -480, "elapsed": 518, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="bb002af9-8fba-4790-e9b1-16c585615389"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=5)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="d5Lp7F8Kg1zG" executionInfo={"status": "ok", "timestamp": 1636995138209, "user_tz": -480, "elapsed": 640, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="133b5a42-865b-4183-bb97-a19ad2518e19"
# experimenting with different n values
k_range = list(range(1,26))
scores = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
scores.append(metrics.accuracy_score(y_test, y_pred))
plt.plot(k_range, scores)
plt.xlabel('Value of k for KNN')
plt.ylabel('Accuracy Score')
plt.title('Accuracy Scores for Values of k of k-Nearest-Neighbors')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="24N7u89-g3wn" executionInfo={"status": "ok", "timestamp": 1636995146400, "user_tz": -480, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="e671f1d6-06d1-4296-c7db-d2d81f3ef686"
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
y_pred = logreg.predict(X_test)
print(metrics.accuracy_score(y_test, y_pred))
# + [markdown] id="AETLzN3Rg6kq"
# Choosing KNN to Model Iris Species Prediction with k = 12
# + colab={"base_uri": "https://localhost:8080/"} id="IE0OLvA5g5zA" executionInfo={"status": "ok", "timestamp": 1636995162674, "user_tz": -480, "elapsed": 565, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="4d1394e9-5611-48c0-c160-3e75871bd720"
knn = KNeighborsClassifier(n_neighbors=12)
knn.fit(X, y)
# make a prediction for an example of an out-of-sample observation
knn.predict([[6, 3, 4, 2]])
| Week 5/Machine Learning with Iris Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><center>BAG of Word</center></h1>
## Only run this if you are using Google Colab and need to install new packages
# via the notebook before you can use them
# !pip3 install powerlaw
# !pip3 install pyLDAvis
# +
#import common libraries:
#------------------------
import pandas as pd #<- For managing the datasets
import re #<- For accessing a folder path
import numpy as np #<- For doing mathematical operation with arrays.
import matplotlib.pyplot as plt #<- To plot in python
#New libraries from the Natural Lenguage Toolkit (Human language data):
#----------------------------------------------------------------------
import nltk
nltk.download('stopwords')
from nltk.tokenize import word_tokenize #<- For finding tokens (small divisions) from a large sample of text
from nltk.corpus import stopwords #<- For calling the know stopwords in english (e.g, articles, connectors)
from nltk.corpus import wordnet #<- For calling a lexical database in eglish with meanings, synonyms, antonyms, and more
from nltk.stem import WordNetLemmatizer #<- For normalizing the words in a text, for example, different conjugations of a verb and its brings its simple form.
#New libraries from the sklearn the library for machine learning in python:
#--------------------------------------------------------------------------
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer #<- To vectorize our text into terms frequencies
from sklearn.decomposition import LatentDirichletAllocation #<- To classify our text into groups based on the vectors of frequencies
#New libraries for topic modelling:
#---------------------------------
import gensim #<- Topic modelling library
import gensim.corpora as corpora #<- To create a corpus of our text
from pprint import pprint
#New libraries for visualizing the topics in a topic model from a corpus (gensim):
#---------------------------------------------------------------------------------
import pyLDAvis #<- Visualization library
import pyLDAvis.gensim #<- Connecting with gensim
from wordcloud import WordCloud
# Import library for managing collections/
import collections
# Import library for fitting powerlaw functions to data
import powerlaw
#Extra libraries:
# %matplotlib inline
import warnings
# Suppress warnings for aesthetic purposes
warnings.filterwarnings("ignore", category=DeprecationWarning)
# -
# ## 1. Implementing the Bag of Words Algorithm
# +
def vectorize(tokens):
''' This function takes list of words in a sentence as input
and returns a vector of size of filtered_vocab.It puts 0 if the
word is not present in tokens and count of token if present.'''
vector=[]
for w in filtered_vocab:
vector.append(tokens.count(w))
return vector
def unique(sequence):
'''This functions returns a list in which the order remains
same and no item repeats.Using the set() function does not
preserve the original ordering,so i didnt use that instead'''
seen = set()
return [x for x in sequence if not (x in seen or seen.add(x))]
# Import a list of stopwords from NLTK
stops = stopwords.words('english')
# Create a list of special characters which go between words
special_char=[",",":"," ",";",".","?"]
#Write the sentences in the corpus,in our case, just two
string1="Welcome to Great Learning , Now start learning"
string2="Learning is a good practice"
#convert them to lower case
string1=string1.lower()
string2=string2.lower()
#split the sentences into tokens
tokens1=string1.split()
tokens2=string2.split()
print(tokens1)
print(tokens2)
#create a vocabulary list
vocab=unique(tokens1+tokens2)
print(vocab)
#filter the vocabulary list
filtered_vocab=[]
for w in vocab:
if w not in stops and w not in special_char:
filtered_vocab.append(w)
print(filtered_vocab)
#convert sentences into vectords
vector1=vectorize(tokens1)
print(vector1)
vector2=vectorize(tokens2)
print(vector2)
# +
import pandas as pd
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
sentence_1="This is a good job. I will not miss it for anything"
sentence_2="This is not a good job at all"
CountVec = CountVectorizer(ngram_range=(1,1), # to use bigrams ngram_range=(2,2)
stop_words='english')
#Transform to vectors
Count_data = CountVec.fit_transform([sentence_1,sentence_2])
#Create dataframe from Vectors
cv_dataframe=pd.DataFrame(Count_data.toarray(),columns=CountVec.get_feature_names())
print(cv_dataframe)
# +
#2 Grams
CountVec = CountVectorizer(ngram_range=(2, 2),
stop_words='english')
#Transform to vectors
Count_data = CountVec.fit_transform([sentence_1,sentence_2])
#Create dataframe from Vectors
cv_dataframe=pd.DataFrame(Count_data.toarray(),columns=CountVec.get_feature_names())
print(cv_dataframe)
# -
from sklearn.metrics.pairwise import cosine_similarity
# Cosine similarity with 1 grams
cosine_similarity(np.asarray([vector1, vector2]))
# Cosine similarity with 2-grams
cosine_similarity(cv_dataframe)
from scipy.spatial.distance import euclidean
# Euclidean distance with 1-grams
euclidean(vector1, vector2)
# Euclidean distance with 2-grams
euclidean(cv_dataframe.loc[0], cv_dataframe.loc[1])
# ## 2. TF-IDF and Comparing Documents
# Load the two text files into memory
text1 = open("Text1.txt").read()
text2 = open("Text2.txt").read()
# Print them in two separate paragraphs for a quick overview
print("Text 1:\n", text1)
print("\n Text 2:\n", text2)
# Now - lets use the wordcloud package to create some visualisation overviews
plt.imshow(WordCloud().generate(text1))
plt.imshow(WordCloud().generate(text2))
# When comparing these two word clouds - what do you notice?
# Now, we will explore different ways to use tf-idf to analyse and compare documents
# Lets see what the frequency distribution of words in these documents looks like.
# +
def zipf_law_plot(Ordered_Frequency, Absolute_frequency):
n = 20
Rank = [i for i in range(n)]
x = Rank
y = list(Absolute_frequency.values())[:n]
ax = plt.subplot(111) #<-- define the plot to allow annotation
ax.loglog(x,y,'go-', alpha=0.2) #<-- plot in the log log form
Strings = list(Ordered_Frequency.keys()) #<-- Set the top 20 most frequent words
#The annotations start position in 1 then we sum 1 or do a range(1,20)
for i in range(19):
ax.annotate(Strings[i], xy=(x[i+1], y[i+1]), fontsize=12)
plt.xlabel('Frequency rank of words', fontsize = 'x-large')
plt.ylabel('Absolut frequency of words', fontsize = 'x-large')
plt.show()
# -
def words_frequency_rank(text):
words = re.findall(r'\w+', text) #<-- Select the element of the text that are words
Counting_words = dict(collections.Counter(words)) #<-- Count the words
#Order the counting: lambda is another way to write functions: https://www.w3schools.com/python/python_lambda.asp
Ordered_frequency = {k: v for k, v in sorted(Counting_words.items(), key=lambda item: item[1], reverse = True)}
#Calculate the absolute frequency:
Absolute_frequency = {k:Ordered_frequency[k]/np.sum(list(Ordered_frequency.values())) for k in Ordered_frequency.keys()}
return Ordered_frequency, Absolute_frequency
#
Ordererd_Frequency_1, Absolute_Frequency_1 = words_frequency_rank(text1)
Ordererd_Frequency_2, Absolute_Frequency_2 = words_frequency_rank(text2)
zipf_law_plot(Ordererd_Frequency_1, Absolute_Frequency_1)
zipf_law_plot(Ordererd_Frequency_2, Absolute_Frequency_2)
def Plot_pdf_CCDF(Data, xlabel, ylabel):
'''This function plot and compares the the Probability Density Function (PDF) and the Complement Cummulative
Density Function. This functions are taken from the paper: https://doi.org/10.1371/journal.pone.0085777
Attributes:
Data: An array of values
'''
fit = powerlaw.Fit(Data) #<-- Fit the data
fig2=fit.plot_pdf(color='b', linewidth=2) #<-- Plot the PDF of the original data
fit.power_law.plot_pdf(color='b', linestyle='dashed', ax=fig2) #<-- Plot the PDF of the power law fit
fit.plot_ccdf(color='r', linewidth=2, ax=fig2) #<-- Plot the PDF of the original data in the same fig
fit.power_law.plot_ccdf(color='r', linestyle='dashed', ax=fig2) #<-- Plot the PDF of the power law fit
plt.legend(['Data pdf', 'fit PDF', 'Data CCDF', 'fit CCDF'], fontsize = 'x-large')
plt.xlabel(xlabel, fontsize = 'x-large')
plt.ylabel(ylabel, fontsize = 'x-large')
plt.show()
Plot_pdf_CCDF(list(Ordererd_Frequency_1.values()), 'Word frequency', 'p(X), p(X≥x)')
Plot_pdf_CCDF(list(Ordererd_Frequency_2.values()), 'Word frequency', 'p(X), p(X≥x)')
# Now, lets look at the vectoriser again and see how we can more directly compare two documents
# TF-IDF Document Comparison
texts = [text1, text2]
vectorizer = TfidfVectorizer(stop_words=stops)
transformed_texts = vectorizer.fit_transform(texts)
transformed_text_array = transformed_texts.toarray()
tf_idf_dataframes = []
for counter, txt in enumerate(transformed_text_array):
# construct a dataframe
tf_idf_tuples = list(zip(vectorizer.get_feature_names(), txt))
tf_idf_dataframes.append(pd.DataFrame.from_records(tf_idf_tuples, columns=['term', 'score']).sort_values(by='score', ascending=False).reset_index(drop=True)['term'])
tf_idf_doc_comparison = pd.DataFrame(tf_idf_dataframes).T
tf_idf_doc_comparison.columns = ["Text 1", "Text 2"]
tf_idf_doc_comparison.index.names = ['Tf-IDF Ranking']
tf_idf_doc_comparison.head(20)
# Now lets see how strongly the two documents correlate with one another
correlation_matrix = ((transformed_texts * transformed_texts.T).A)
# It appears these two documents are very different to one another once stop words have been removed
correlation_matrix
# Lets try adding another text, and see which it matches more closely
text3 = open("Text3.txt").read()
# Text 3 is a document explaining the definition of a hedge fund, so it should be
# More closely related to text 2 than text 1.
plt.imshow(WordCloud().generate(text3))
texts = [text1, text2, text3]
transformed_texts = vectorizer.fit_transform(texts)
transformed_text_array = transformed_texts.toarray()
tf_idf_dataframes = []
for counter, txt in enumerate(transformed_text_array):
# construct a dataframe
tf_idf_tuples = list(zip(vectorizer.get_feature_names(), txt))
tf_idf_dataframes.append(pd.DataFrame.from_records(tf_idf_tuples, columns=['term', 'score']).sort_values(by='score', ascending=False).reset_index(drop=True)['term'])
tf_idf_doc_comparison = pd.DataFrame(tf_idf_dataframes).T
tf_idf_doc_comparison.columns = ["Text 1", "Text 2", "Text 3"]
tf_idf_doc_comparison.index.names = ['Tf-IDF Ranking']
tf_idf_doc_comparison.head(20)
correlation_matrix = ((transformed_texts * transformed_texts.T).A)
correlation_matrix
import seaborn as sns
labels = ["Text 1", "Text 2", "Text 3"]
sns.heatmap(correlation_matrix, annot=True, xticklabels= labels, yticklabels= labels)
plt.title("Text Similarity")
# +
vectorizer = TfidfVectorizer(stop_words=stops, ngram_range=(2,2))
transformed_texts = vectorizer.fit_transform(texts)
transformed_text_array = transformed_texts.toarray()
tf_idf_dataframes = []
for counter, txt in enumerate(transformed_text_array):
# construct a dataframe
tf_idf_tuples = list(zip(vectorizer.get_feature_names(), txt))
tf_idf_dataframes.append(pd.DataFrame.from_records(tf_idf_tuples, columns=['term', 'score']).sort_values(by='score', ascending=False).reset_index(drop=True)['term'])
tf_idf_doc_comparison = pd.DataFrame(tf_idf_dataframes).T
tf_idf_doc_comparison.columns = ["Text 1", "Text 2", "Text 3"]
tf_idf_doc_comparison.index.names = ['Tf-IDF Ranking']
tf_idf_doc_comparison.head(20)
# +
correlation_matrix = ((transformed_texts * transformed_texts.T).A)
correlation_matrix
labels = ["Text 1", "Text 2", "Text 3"]
sns.heatmap(correlation_matrix, annot=True, xticklabels= labels, yticklabels= labels)
# +
## BONUS
# Add topic labels
labels = ["Hillary Clinton", "Hedge Funds"]
# Remove the second hedge funds article to prevent training class imbalance
texts = [text1, text2]
# Import feature extraction functions
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
# Fit and transform vectorisers for the two documents
count_vect = CountVectorizer()
x_train_counts = count_vect.fit_transform(texts)
tf_transformer = TfidfTransformer(use_idf=True).fit(x_train_counts)
x_train_tf = tf_transformer.transform(x_train_counts)
# +
from sklearn.naive_bayes import MultinomialNB
from sklearn.neighbors import KNeighborsClassifier
# Create a classifier and fit it to the training data
clf = MultinomialNB().fit(x_train_tf, labels)
clf_kn = KNeighborsClassifier(n_neighbors = 1).fit(x_train_tf, labels)
# +
# Add some new strings (from wikipedia) to match to the two topics
new_strings = ["<NAME> (born October 26, 1947) is an American politician, diplomat, lawyer, writer, and public speaker who served as the 67th United States secretary of state from 2009 to 2013, as a United States senator from New York from 2001 to 2009, and as First Lady of the United States from 1993 to 2001",
"Although hedge funds are not subject to many restrictions that apply to regulated funds, regulations were passed in the United States and Europe following the financial crisis of 2007–2008 with the intention of increasing government oversight of hedge funds and eliminating certain regulatory gaps."]
# Transform the new strings to vectors
x_test_counts = count_vect.transform(new_strings)
x_test_tfidf = tf_transformer.transform(x_test_counts)
# Predict class using the classifier
clf.predict(x_test_tfidf)
# -
clf_kn.predict(x_test_tfidf)
| Practice/P9_Bag_of_words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Desagregação dos dados da bacia do Parnaíba usando GeoPandas
#
# Este *notebook* mostra é uma continuação [deste *notebook*](https://github.com/francisconog/PBCE/blob/master/analysis/ESTPLU_POR_BACIA.ipynb) que classifica um ponto de acordo com sua latitude e longitude. Em 2012 a Bacia do Parnaíba foi dividida em: Serra de Ibiapaba e Sertões de Crateús. Desta forma, precisa-se desagregar os dados da antiga bacia para a realização de análises mais completas.
#
# *Este Notebook faz parte de uma colaboração da Universidade Federal do Ceará (UFC), da Companhia de Gestão de Recursos Hídricos (COGERH) e da Fundação Cearense de Apoio ao Desenvolvimento Científico e Tecnológico (FUNCAP) no âmbito do projeto de desenvolvimento de ferramentas tecnológicas de gestão para planejamento de recursos hídricos do estado do Ceará: Segurança hídrica e planejamento de secas*
#
# *Code by: <NAME>*
# ## Importing packages
import geobr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import geopandas as gpd
import os
from shapely.geometry import Point
from tqdm import tqdm
import sys
import warnings
sys.path.append('../src')
warnings.filterwarnings('ignore')
from functions import *
import re
# ## Paths
data_path = "../data/"
shape_file_path = "shapes"
shape_malha_dagua = "Drenagem/Margens_simples_CE_corrigidoSRH.shp"
shape_bacias = "geomap_bacias/i3geomap_bacias.shp"
faturamento_data_path = "Dados Alocar/faturamento-cogerh"
# ## Import Data
# Dados de outorga de 2012
faturamento_2012 = pd.read_excel(os.path.join(data_path,faturamento_data_path,"historico_faturamento.xlsx"))
faturamento_2012.head()
# ##### Agora que carregados os pacotes e os dados, vamos ao que interessa...
lista_bacias = faturamento_2012["Bacia"].unique()
update_lista_bacias = {x: x.split(" -")[0] for x in lista_bacias}
# +
faturamento_2012["Bacia"] = faturamento_2012["Bacia"].apply(lambda x: update_lista_bacias[x])
faturamento_2012["Bacia"].unique()
# -
# Dados de faturamento referentes à bacia do Parnaíba
faturamento_parnaiba = faturamento_2012.query("Bacia=='BACIA DO PARNAIBA'")
faturamento_parnaiba.head()
faturamento_parnaiba.query("`Infra Estrutura Hídrica`=='BACIA PARNAIBA'")
# +
# A desagregação dos Dados será feita baseado na Gerencia
dict_gerencia_bacia = {
'GERENCIA DE SOBRAL': "BACIA DA SERRA DA IBIAPABA",
'GERENCIA DE CRATEUS': 'BACIA DO SERTÕES DE CRATEÚS',
'GERENCIA DE QUIXERAMOBIM': 'BACIA DO BANABUIU'
}
def atualizacao_bacia(df):
try:
return dict_gerencia_bacia[df["Gerencia"]]
except:
return df["Bacia"]
# -
faturamento_2012["Bacia"] = faturamento_2012.apply(atualizacao_bacia,axis=1)
# Agora é só salvar salvar os dados de faturamento desagregados
faturamento_2012.to_excel(os.path.join(data_path,faturamento_data_path,"faturamento_limpo_desagregado.xlsx"),index=False)
| analysis/DESAGREG_FATURAMENTO_PARNAIBA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Import
import pandas as pd
from openpyxl import load_workbook
from IPython.display import Image
from sklearn.preprocessing import Imputer
from sklearn import tree
from subprocess import check_call
import re
from sklearn.metrics import confusion_matrix, accuracy_score, recall_score, precision_score
import graphviz
from sklearn.model_selection import cross_val_score
# +
data_anarkis_wb = load_workbook(filename = 'data_anarkis.xlsx')
data_anarki_sr = data_anarkis_wb.active
dk = pd.DataFrame(data_anarki_sr.values)
dk = dk[1:]
dk.columns = ['TANGGAL','HARI','UNSUR','TEMPAT']
for index, row in dk.iterrows():
temp = str(row['TANGGAL'])
replace=temp.replace('-',' ')
convert=replace.split()
year= convert[0:1][0]
day= convert[2:4][0]
mont = convert[1:2][0]
baru = '/'.join([str(day), str(mont), str(year)])
dk.loc[index, 'TANGGAL'] = baru
# +
tanggal_kriminal = dk['TANGGAL']
data_iklim_wb = load_workbook(filename = 'laporan_iklim_harian.xlsx')
data_iklim_sr = data_iklim_wb.active
di = pd.DataFrame(data_iklim_sr.values)
di = di[1:]
di.columns = ['TANGGAL','SUHU','KELEMBABAN','HUJAN']
di['KRIMINAL'] = 'Tidak Ada'
for index, row in di.iterrows():
tanggal_sekarang = str(row['TANGGAL'])
for tanggal in tanggal_kriminal:
if str(tanggal_sekarang) == str(tanggal):
di.loc[index, 'KRIMINAL'] = 'Ada'
di = di[di.HUJAN != 8888]
di = di[di.HUJAN != 9999]
di = di[di.SUHU != 8888]
di = di[di.SUHU != 9999]
# di = di[di.HUJAN != 8888]
# di = di[di.HUJAN
di
# tanggal_kriminal
# -
j = 0
for index, row in di.iterrows():
if j <= 270:
if row['KRIMINAL'] != 'Ada':
di = di[di.TANGGAL != row['TANGGAL']]
# print "ads"
j = j + 1
di
# +
rata2hujan = list()
i = 0
n = 0
for index, row in di.iterrows():
n = n + 1
rata2hujan.append(row['HUJAN'])
if row['KRIMINAL'] == 'Ada':
i = i + 1;
rata2h = sum(rata2hujan)/n
minhujan = rata2h * 85/100
maxhujan = rata2h * 115/100
for index, row in di.iterrows():
if row['HUJAN'] >= minhujan:
di.loc[index, 'HUJAN'] = 1
else:
di.loc[index, 'HUJAN'] = 0
di = di.drop('TANGGAL', 1)
# print di
dtrain = di[1:80]
# dtrain
# -
di
# +
y = dtrain['KRIMINAL'].values
# features = di['SUHU','KELEMBABAN','HUJAN']
features_columns = ['SUHU','HUJAN']
features = dtrain[list(features_columns)].values
imp = Imputer(axis=0)
x = imp.fit_transform(features)
clf = tree.DecisionTreeClassifier(criterion="entropy")
clf = clf.fit(x,y)
import cPickle as pkl
pkl.dump(clf, open('tree.pkl', 'wb'))
# from sklearn.naive_bayes import GaussianNB
# gnb = GaussianNB()
# gnb_fit = gnb.fit(x, y)
# -
# +
di
d_test = di[80:]
# d_test = d_test.sample(n=30)
y_true = d_test['KRIMINAL'].values
y_true
features_columns = ['SUHU','HUJAN']
features = d_test[list(features_columns)].values
features
y_predict = clf.predict(features)
# y_predict
clf.predict([[27.9, 1]])
# -
result = confusion_matrix(y_true, y_predict, labels=["Tidak Ada","Ada"])
result
akurasi = accuracy_score(y_true, y_predict)
akurasi
presisi = precision_score(y_true, y_predict, average='macro')
presisi
recall = recall_score(y_true, y_predict, average='macro')
recall
# +
with open("tree.dot", "w") as f:
f = tree.export_graphviz(clf, out_file=f, feature_names=features_columns,
filled=True, rounded=True,
special_characters=True)
from subprocess import check_call
check_call(['dot','-Tpng','tree.dot','-o','OutputFile.png'])
# dot_data = tree.export_graphviz(clf, out_file=None, feature_names=features_columns,
# filled=True, rounded=True,
# special_characters=True)
# graph = graphviz.Source(dot_data)
# graph
# graph.draw('file.png')
| Python/hackjak/App.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This notebook loads train_v2.csv/test_v2.csv file and flatten the json fields.
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
import pandas as pd
import numpy as np
import json
from ast import literal_eval
# + _uuid="4a61452380781d24ef75a339e3125e5222386096"
df = pd.read_csv('../input/train_v2.csv', nrows=10000)
# + _uuid="40025f6231d9d49dfeaaf07e8a7f29884fe22901"
df.head()
# + [markdown] _uuid="d60dfff02ec7af417e1f5e4aba4485e06d0735bf"
# We have 6 special columns which should be flattened. `device`, `geoNetwork`, `totals`,`trafficSource` is standard json fields and can't be easily flattened by json module; `customDimensions` and `hit` columns can not be processed by json module, bu can be processed by ast.literal_eval.
# + [markdown] _uuid="dc18b6850e760b11d8403b26e3d822eb95324ab1"
# ## device, geoNetwork, totals and trafficSource
# + _uuid="f112cc5048f30b64f747070c41ca76e13f1b7a5f"
json_cols = ['device', 'geoNetwork', 'totals', 'trafficSource']
def parse_json_col(raw_str):
return pd.Series(json.loads(raw_str))
for col in json_cols:
parsed_df = df[col].apply(parse_json_col)
parsed_df.columns = [f'{col}_{x}' for x in parsed_df.columns]
df = pd.concat([df, parsed_df], axis=1)
df.drop(col, axis=1, inplace=True)
# + _uuid="0bfb5fe5c204649aa60da8f5dce10a78f8cc10cc"
df.shape
# + [markdown] _uuid="a7209acac5715210c9867cdef510286691bf1142"
# Let's check the new columns. We found that `trafficSource_adwordsClickInfo` is also a json column which should be flattened.
# + _uuid="9ec2db5f5e418c21f163687f850f6e19c5349f46"
df.filter(regex='.*_.*', axis=1).head()
# + _uuid="a28c27d881cdf53c830dcd6dc05e5d080117499c"
trafficSource_adwordsClickInfo_df = df.trafficSource_adwordsClickInfo.apply(pd.Series)
trafficSource_adwordsClickInfo_df.columns = [f'trafficSource_adwordsClickInfo_{x}' for x in trafficSource_adwordsClickInfo_df.columns]
df = pd.concat([df, trafficSource_adwordsClickInfo_df], axis=1)
df.drop('trafficSource_adwordsClickInfo', axis=1, inplace=True)
# + _uuid="2a1a16aa642e15abf1b0e91d5baef947d99bee27"
df.shape
# + _uuid="d8289b52af00d37e4414c82b0b11a4b6438f74b7"
df.filter(regex='trafficSource_adwordsClickInfo_.*', axis=1).head()
# + [markdown] _uuid="f57750762c792deb8517f001e8dbb2b1a471be78"
# ## customDimensions
# + _uuid="8bcea224022516feecfae9274b2fe4e8d7f18479"
# for customDimensions and hits columns
def parse_special_col(raw_str):
lst = literal_eval(raw_str)
if isinstance(lst, list) and lst:
return pd.Series(lst[0])
else:
return pd.Series({})
# + _uuid="7703c23fbd9b3697e2ad30b5b904d4889989e749"
customDimensions_df = df.customDimensions.apply(parse_special_col)
customDimensions_df.columns = [f'customDimensions_{x}' for x in customDimensions_df.columns]
df = pd.concat([df, customDimensions_df], axis=1)
df.drop('customDimensions', axis=1, inplace=True)
# + _uuid="567dfdeb0598ebf0aaf100f31e38a93266b85a8c"
df.shape
# + _uuid="37602f2e1e4ae5e97b70b4d31c0a6173491d3755"
df.filter(regex='customDimensions_.*', axis=1).head()
# + [markdown] _uuid="3c875bb50da2678f05c280f8f6fe72c52bc5dcc6"
# ## hits
# + _uuid="cd44dffda5b6c4581fe106383c7742dda31cff80"
hits_df = df.hits.apply(parse_special_col)
hits_df.columns = [f'hits_{x}' for x in hits_df.columns]
df = pd.concat([df, hits_df], axis=1)
df.drop('hits', axis=1, inplace=True)
# + _uuid="7dd2a31bb36db554682823c6672531182e15f365"
df.shape
# + _uuid="274d0b3d43b0bba1b230cccce985900cbd78a6c6"
df.filter(regex='hits_.*', axis=1).head()
# + [markdown] _uuid="aee82ecd24cc09db0989cdc9b128303c33019e15"
# `hits_experiment`, `hits_customVariables`, `hits_customMetrics`, `hits_publisher_infos`, `hits_customDimensions` are empty, we can drop it.
# + _uuid="72c8e4938d2c8e9c41cd3510ac4ac88b8dd448cd"
df.drop(['hits_experiment', 'hits_customVariables', 'hits_customMetrics', 'hits_publisher_infos', 'hits_customDimensions'], axis=1, inplace=True)
# + [markdown] _uuid="c06e9a024380c12e88d898c28a161b5da01c3e61"
# `hits_page`, `hits_transaction`, `hits_item`, `hits_appInfo`, `hits_exceptionInfo`, `hits_eCommerceAction`, `hits_social`, `hits_contentGroup`, `hits_promotionActionInfo` are python dict, we can should flatten it.
# + _uuid="f14430890c9719659d1cb406999571b98a09b35c"
dict_cols = ['hits_page', 'hits_transaction', 'hits_item', 'hits_appInfo',
'hits_exceptionInfo', 'hits_eCommerceAction', 'hits_social', 'hits_contentGroup', 'hits_promotionActionInfo']
for col in dict_cols:
parsed_df = hits_df[col].apply(pd.Series)
parsed_df.columns = [f'{col}_{x}' for x in parsed_df.columns]
df = pd.concat([df, parsed_df], axis=1)
df.drop(col, axis=1, inplace=True)
# + [markdown] _uuid="959d1b3f7bca9bbae94de2a9419cf40f172d9024"
# `hits_product`, `hits_promotion` are python list, we should flatten it.
# + _uuid="fd1b7a2ea237cb7f5f898d8f1e0dba8b1a42a87c"
def parse_list(x):
if isinstance(x, list) and x:
return pd.Series(x[0])
else:
return pd.Series({})
for col in ['hits_product', 'hits_promotion']:
parsed_df = hits_df[col].apply(parse_list)
parsed_df.columns = [f'{col}_{x}' for x in parsed_df.columns]
df = pd.concat([df, parsed_df], axis=1)
df.drop(col, axis=1, inplace=True)
# + _uuid="7d0410e59b9e34564946f1858a5f12bee757ed88"
df.shape
# + [markdown] _uuid="39d300e4c637971a87abc369ca29e94928fdccf2"
# ## Pack it to a function
# I have put the code in one function so you can copy it easily.
# + _uuid="aea1c5b21587b3033e2ce42fa00feca4cadbc7c9"
def flatten(in_csv, out_csv, nrows=None):
df = pd.read_csv(in_csv, dtype=np.object, nrows=nrows)
# json columns
json_cols = ['device', 'geoNetwork', 'totals', 'trafficSource']
def parse_json_col(raw_str):
return pd.Series(json.loads(raw_str))
for col in json_cols:
parsed_df = df[col].apply(parse_json_col)
parsed_df.columns = [f'{col}_{x}' for x in parsed_df.columns]
df = pd.concat([df, parsed_df], axis=1)
df.drop(col, axis=1, inplace=True)
# trafficSource_adwordsClickInfo
trafficSource_adwordsClickInfo_df = df.trafficSource_adwordsClickInfo.apply(pd.Series)
trafficSource_adwordsClickInfo_df.columns = [f'trafficSource_adwordsClickInfo_{x}' for x in trafficSource_adwordsClickInfo_df.columns]
df = pd.concat([df, trafficSource_adwordsClickInfo_df], axis=1)
df.drop('trafficSource_adwordsClickInfo', axis=1, inplace=True)
# customDimensions
def parse_customDimensions(raw_str):
lst = literal_eval(raw_str)
if isinstance(lst, list) and lst:
return pd.Series(lst[0])
else:
return pd.Series({})
customDimensions_df = df.customDimensions.apply(parse_customDimensions)
customDimensions_df.columns = [f'customDimensions_{x}' for x in customDimensions_df.columns]
df = pd.concat([df, customDimensions_df], axis=1)
df.drop('customDimensions', axis=1, inplace=True)
# hits
def parse_hits(raw_str):
lst = literal_eval(raw_str)
if isinstance(lst, list) and lst:
return pd.Series(lst[0])
else:
return pd.Series({})
hits_df = df.hits.apply(parse_hits)
hits_df.columns = [f'hits_{x}' for x in hits_df.columns]
df = pd.concat([df, hits_df], axis=1)
df.drop('hits', axis=1, inplace=True)
# 'hits_page', 'hits_transaction', 'hits_item', 'hits_appInfo',
# 'hits_exceptionInfo', 'hits_eCommerceAction', 'hits_social', 'hits_contentGroup', 'hits_promotionActionInfo'
dict_cols = ['hits_page', 'hits_transaction', 'hits_item', 'hits_appInfo',
'hits_exceptionInfo', 'hits_eCommerceAction', 'hits_social', 'hits_contentGroup', 'hits_promotionActionInfo']
for col in dict_cols:
parsed_df = hits_df[col].apply(pd.Series)
parsed_df.columns = [f'{col}_{x}' for x in parsed_df.columns]
df = pd.concat([df, parsed_df], axis=1)
df.drop(col, axis=1, inplace=True)
# 'hits_experiment', 'hits_customVariables', 'hits_customMetrics', 'hits_publisher_infos', 'hits_customDimensions' are empty
df.drop(['hits_experiment', 'hits_customVariables', 'hits_customMetrics', 'hits_publisher_infos', 'hits_customDimensions'], axis=1, inplace=True)
# 'hits_product', 'hits_promotion'
def parse_list(x):
if isinstance(x, list) and x:
return pd.Series(x[0])
else:
return pd.Series({})
for col in ['hits_product', 'hits_promotion']:
parsed_df = hits_df[col].apply(parse_list)
parsed_df.columns = [f'{col}_{x}' for x in parsed_df.columns]
df = pd.concat([df, parsed_df], axis=1)
df.drop(col, axis=1, inplace=True)
df.to_csv(out_csv, index=False)
return df.shape
# + [markdown] _uuid="138e948d9e47fe6abff06ef08e27f1e15b141dfb"
# It takes about 1~2h to flatten the train_v2.csv and test_v2.csv. Have fun!
| 9 google customer revenue prediction/v2-flatten-all-the-json-filelds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import rosbag_pandas
import pandas as pd
import numpy as np
file_1 = '/home/rik/data/2020_08_05_gannertshofen/estimator_development/sensors_2020-08-05-13-13-56_estimator_2020-08-30-17-48-25_w_baseline.bag'
df_1 = rosbag_pandas.bag_to_dataframe(file_1)
# -
file_2 = '/home/rik/data/2020_08_05_gannertshofen/estimator_development/sensors_2020-08-05-13-13-56_estimator_2020-08-31-09-22-44_spp.bag'
df_2 = rosbag_pandas.bag_to_dataframe(file_2)
import pandas as pd
def getPoseTf(df, topic):
df_pose = pd.to_datetime(df[topic + '/header/stamp/secs'], unit='s') + pd.to_timedelta(df[topic + '/header/stamp/nsecs'], unit='ns')
df_pose = pd.concat([df_pose, df[topic + '/pose/position/x']], axis=1)
df_pose = pd.concat([df_pose, df[topic + '/pose/position/y']], axis=1)
df_pose = pd.concat([df_pose, df[topic + '/pose/position/z']], axis=1)
df_pose = pd.concat([df_pose, df[topic + '/pose/orientation/x']], axis=1)
df_pose = pd.concat([df_pose, df[topic + '/pose/orientation/y']], axis=1)
df_pose = pd.concat([df_pose, df[topic + '/pose/orientation/z']], axis=1)
df_pose = pd.concat([df_pose, df[topic + '/pose/orientation/w']], axis=1)
df_pose.reset_index(inplace=True)
df_pose.columns = ['t_arrival', 't', 'x', 'y', 'z', 'q_x', 'q_y', 'q_z', 'q_w']
df_pose.dropna(inplace=True)
df_pose.reset_index(inplace=True)
df_pose.drop('t_arrival', axis=1, inplace=True)
df_pose.drop('index', axis=1, inplace=True)
from scipy.spatial.transform import Rotation as R
ypr = df_pose.apply(lambda row: R.from_quat([row.q_x, row.q_y, row.q_z, row.q_w]).as_euler('ZYX', degrees=True), axis=1)
ypr = pd.DataFrame(ypr.values.tolist(), columns=['yaw', 'pitch', 'roll'])
df_pose = pd.concat([df_pose, ypr], axis=1)
df_pose.set_index('t', inplace=True)
return df_pose
import pandas as pd
def getBias(df, topic):
df_bias = pd.to_datetime(df[topic + '/header/stamp/secs'], unit='s') + pd.to_timedelta(df[topic + '/header/stamp/nsecs'], unit='ns')
df_bias = pd.concat([df_bias, df[topic + '/vector/x']], axis=1)
df_bias = pd.concat([df_bias, df[topic + '/vector/y']], axis=1)
df_bias = pd.concat([df_bias, df[topic + '/vector/z']], axis=1)
df_bias.reset_index(inplace=True)
df_bias.columns = ['t_arrival', 't', 'x', 'y', 'z']
df_bias.dropna(inplace=True)
df_bias.reset_index(inplace=True)
df_bias.drop('t_arrival', axis=1, inplace=True)
df_bias.drop('index', axis=1, inplace=True)
return df_bias
def getHeading(df, topic):
df_heading = pd.to_datetime(df[topic + '/header/stamp/secs'], unit='s') + pd.to_timedelta(df[topic + '/header/stamp/nsecs'], unit='ns')
df_heading = pd.concat([df_heading, df[topic + '/position/position/x']], axis=1)
df_heading = pd.concat([df_heading, df[topic + '/position/position/y']], axis=1)
df_heading.reset_index(inplace=True)
df_heading.columns = ['t_arrival', 't', 'base_x', 'base_y']
df_heading.dropna(inplace=True)
df_heading.set_index('t', inplace=True)
df_heading.drop('t_arrival', axis=1, inplace=True)
# Convert NED->ENU
import numpy as np
x = df_heading['base_y'].values
y = df_heading['base_x'].values
from scipy.spatial.transform import Rotation as R
r = R.from_rotvec(np.pi/2 * np.array([0, 0, 1]))
vectors = np.array([x, y, np.zeros(len(x))]).transpose()
heading_vectors = r.apply(vectors)
heading = np.arctan2(heading_vectors[:, 1], heading_vectors[:, 0]) * 180.0 / np.pi
df_heading['rtk heading'] = heading
return df_heading
# +
df_pose_1 = getPoseTf(df_1, '/moa/mav_state_estimator/optimization')
df_pose_2 = getPoseTf(df_2, '/moa/mav_state_estimator/optimization')
df_heading = getHeading(df_1, '/moa/piksi/attitude_receiver_0/ros/baseline_ned')
df_acc_bias_1 = getBias(df_1, '/moa/mav_state_estimator/acc_bias')
df_acc_bias_2 = getBias(df_2, '/moa/mav_state_estimator/acc_bias')
df_gyro_bias_1 = getBias(df_1, '/moa/mav_state_estimator/gyro_bias')
df_gyro_bias_2 = getBias(df_2, '/moa/mav_state_estimator/gyro_bias')
# +
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# %matplotlib qt
fontsize=12
fig, axs = plt.subplots(nrows=1, sharex=True)
df_pose_1['yaw'].plot(ax=axs)
df_pose_2['yaw'].plot(ax=axs)
df_heading['rtk heading'].plot(style='-', ax=axs)
axs.legend(['heading 1', 'heading 2', 'heading rtk'])
axs.set_xlabel('Timestamp', fontsize=fontsize)
axs.set_ylabel('Angle [deg]', fontsize=fontsize)
# +
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# %matplotlib qt
fontsize=12
fig, axs = plt.subplots(nrows=3, sharex=True)
df_pose_1['x'].plot(ax=axs[0])
df_pose_2['x'].plot(ax=axs[0])
axs[0].set_xlabel('Timestamp', fontsize=fontsize)
axs[0].set_ylabel('Position [m]', fontsize=fontsize)
axs[0].legend(['x 1', 'x 2'])
df_pose_1['y'].plot(ax=axs[1])
df_pose_2['y'].plot(ax=axs[1])
axs[1].set_xlabel('Timestamp', fontsize=fontsize)
axs[1].set_ylabel('Position [m]', fontsize=fontsize)
axs[1].legend(['y 1', 'y 2'])
df_pose_1['z'].plot(ax=axs[2])
df_pose_2['z'].plot(ax=axs[2])
axs[2].set_xlabel('Timestamp', fontsize=fontsize)
axs[2].set_ylabel('Position [m]', fontsize=fontsize)
axs[2].legend(['z 1', 'z 2'])
#df_pose_1['roll'].plot(ax=axs[3])
#df_pose_2['roll'].plot(ax=axs[3])
#axs[3].set_xlabel('Timestamp', fontsize=fontsize)
#axs[3].set_ylabel('Angle [deg]', fontsize=fontsize)
#axs[3].legend(['roll 1', 'roll 2'])
#df_pose_1['pitch'].plot(ax=axs[4])
#df_pose_2['pitch'].plot(ax=axs[4])
#axs[4].set_xlabel('Timestamp', fontsize=fontsize)
#axs[4].set_ylabel('Angle [deg]', fontsize=fontsize)
#axs[4].legend(['pitch 1', 'pitch 2'])
#df_pose_1['yaw'].plot(ax=axs[5])
#df_pose_2['yaw'].plot(ax=axs[5])
#axs[5].set_xlabel('Timestamp', fontsize=fontsize)
#axs[5].set_ylabel('Angle [deg]', fontsize=fontsize)
#axs[5].legend(['yaw 1', 'yaw 2'])
# +
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# %matplotlib qt
fontsize=12
fig, axs = plt.subplots(nrows=2, sharex=True)
df_acc_bias_1[['x', 'y', 'z']].plot(ax=axs[0])
#df_acc_bias_2[['x', 'y', 'z']].plot(ax=axs[0], style='k--')
axs[0].set_xlabel('Timestamp', fontsize=fontsize)
axs[0].set_ylabel('Accelerometer bias [m/s**2]', fontsize=fontsize)
axs[0].legend(['x 1', 'y 1', 'z 1', 'x 2', 'y 2', 'z 2'])
df_gyro_bias_1[['x', 'y', 'z']].plot(ax=axs[1])
df_gyro_bias_2[['x', 'y', 'z']].plot(ax=axs[1], style='k--')
axs[1].set_xlabel('Timestamp', fontsize=fontsize)
axs[1].set_ylabel('Gyroscope bias [rad/s]', fontsize=fontsize)
axs[1].legend(['x 1', 'y 1', 'z 1', 'x 2', 'y 2', 'z 2'], loc='upper right')
| scripts/compare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Find Successor
# [link](https://www.algoexpert.io/questions/Find%20Successor)
# ## My Solution
# +
# This is an input class. Do not edit.
class BinaryTree:
def __init__(self, value, left=None, right=None, parent=None):
self.value = value
self.left = left
self.right = right
self.parent = parent
# O(h) time | O(1) space
def findSuccessor(tree, node):
# Write your code here.
if node.right is not None:
return findLeftNearestChild(node)
else:
return findRightNearestParent(node)
def findLeftNearestChild(root):
cur = root
if cur.right is None:
return None
cur = cur.right
while cur.left is not None:
cur = cur.left
return cur
def findRightNearestParent(node):
cur, parent = node, node.parent
while parent is not None and parent.left != cur:
cur = parent
parent = parent.parent
return parent
# -
# ## Expert Solution
# +
# This is an input class. Do not edit.
class BinaryTree:
def __init__(self, value, left=None, right=None, parent=None):
self.value = value
self.left = left
self.right = right
self.parent = parent
# O(n) time | O(n) space
def findSuccessor(tree, node):
inOrderTraversalOrder = getInOrderTraversalOrder(tree)
for idx, currentNode in enumerate(inOrderTraversalOrder):
if currentNode != node:
continue
if idx == len(inOrderTraversalOrder) - 1:
return None
return inOrderTraversalOrder[idx + 1]
def getInOrderTraversalOrder(node, order=[]):
if node is None:
return order
getInOrderTraversalOrder(node.left, order)
order.append(node)
getInOrderTraversalOrder(node.right, order)
return order
# +
# This is an input class. Do not edit.
class BinaryTree:
def __init__(self, value, left=None, right=None, parent=None):
self.value = value
self.left = left
self.right = right
self.parent = parent
# O(h) time | O(1) space
def findSuccessor(tree, node):
if node.right is not None:
return getLeftmostChild(node.right)
return getRightmostParent(node)
def getLeftmostChild(node):
currentNode = node
while currentNode.left is not None:
currentNode = currentNode.left
return currentNode
def getRightmostParent(node):
currentNode = node
while currentNode.parent is not None and currentNode.parent.right == currentNode:
currentNode = currentNode.parent
return currentNode.parent
# -
# ## Thoughts
| algoExpert/find_successor/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning in Audio Classification in Python
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.io import wavfile
import python_speech_features
from python_speech_features import mfcc
from tqdm import tqdm
# +
from keras.layers import Conv2D, MaxPool2D, Flatten, Dropout, Dense
from keras.layers import LSTM, TimeDistributed
from keras.models import Sequential
from keras.utils import to_categorical
from sklearn.utils.class_weight import compute_class_weight
# +
import pickle
from keras.callbacks import ModelCheckpoint
# -
class Config:
def __init__(self, mode= 'conv', nfilt=26, nfeat=13, nfft = 2048, rate = 16000):
self.mode = mode
self.nfilt = nfilt
self.nfeat = nfeat
self.nfft = nfft
self.rate = rate
self.step = int(rate/10)
self.model_path = os.path.join('models', mode + '.model')
self.p_path = os.path.join('pickles', mode + '.p')
def check_data():
if os.path.isfile(config.p_path):
print('Loading existing data for {} model'.format(config.mode))
with open(config.p_path, 'rb') as handle:
tmp = pickle.load(handle)
return tmp
else:
return None
def build_rand_feat():
tmp = check_data()
if tmp:
return tmp.data[0], tmp.data[1]
X = []
y = []
_min, _max = float('inf'), -float('inf')
for _ in tqdm(range(n_samples)):
rand_class = np.random.choice(class_dist.index, p = prob_dist)
file = np.random.choice(df[df.Class==rand_class].index)
rate, wav = wavfile.read(dataset_directory+str(rand_class)+"/"+str(file))
Class = df.at[file, 'Class']
rand_index = np.random.randint(0, wav.shape[0]-config.step)
sample = wav[rand_index : rand_index + config.step]
X_sample = mfcc(sample, rate, numcep=config.nfeat, nfilt=config.nfilt, nfft=config.nfft)
_min = min(np.amin(X_sample), _min)
_max = max(np.amax(X_sample), _max)
X.append(X_sample)
y.append(classes.index(Class))
config.min = _min
config.max = _max
X, y = np.array(X), np.array(y)
X = (X- _min) / (_max - _min)
if config.mode == 'conv':
X = X.reshape(X.shape[0], X.shape[1], X.shape[2], 1)
elif config.mode =='time':
X = X.reshape(X.shape[0], X.shape[1], X.shape[2])
y = to_categorical(y, num_classes=2)
config.data = (X, y)
with open(config.p_path, 'wb') as handle:
pickle.dump(config, handle, protocol=2)
return X,y
def get_reccurent_model():
### Shape of data for RNN is (n, time, freq)
model = Sequential()
model.add(LSTM(128, return_sequences=True, input_shape=input_shape))
model.add(LSTM(128, return_sequences=True))
model.add(TimeDistributed(Dense(64, activation='relu')))
model.add(TimeDistributed(Dense(32, activation='relu')))
model.add(TimeDistributed(Dense(16, activation='relu')))
model.add(Flatten())
model.add(Dropout(0.5))
model.add(Dense(16, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(2, activation='sigmoid'))
model.summary()
model.compile(loss = 'binary_crossentropy', optimizer = 'adam', metrics=['acc'])
return model
# ## Data Extraction
os.listdir('Temp_Dataset/')
# +
classes = list(os.listdir('Dataset/train/'))
print("Number of Classes in the Data Set:", len(classes), "Classes")
print("The classes of the dataset are :", classes[0], ",", classes[1])
# -
column_names = ['Fname','Class', 'Length']
df = pd.DataFrame(columns = column_names)
df.info()
# dataset_directory = 'Dataset/Train/'
dataset_directory = 'Temp_Dataset/train/'
for c in list(classes):
print('Number of files in the directory \'{}\' are {}'.format(c,len(os.listdir(dataset_directory+c))))
for c in list(classes):
for n,f in tqdm(enumerate(os.listdir(dataset_directory+c))):
rate, signal = wavfile.read(dataset_directory+str(c)+"/"+str(f))
length = signal.shape[0]/rate
f_df = pd.DataFrame({
"Fname": str(f),
"Class": str(c),
"Length": length}, index = [n])
df = df.append(f_df)
df.info()
class_dist = df.groupby(['Class'])['Length'].mean()
class_dist
df.set_index('Fname', inplace=True)
df.info()
# # RNN Model using LSTM
n_samples = 2 * int(df['Length'].sum()/0.1)
prob_dist = class_dist / class_dist.sum()
choices = np.random.choice(class_dist.index, p= prob_dist)
config = Config(mode = 'time')
config
X,y = build_rand_feat()
y_flat = np.argmax(y, axis =1)
input_shape = (X.shape[1], X.shape[2])
model = get_reccurent_model()
# ## Adding Checkpoints
checkpoint = ModelCheckpoint(config.model_path, monitor='val_acc', verbose=1, mode='max',
save_best_only=True, save_weights_only=False, period=1)
model.fit(X, y, epochs=250, batch_size=32, shuffle = True, validation_split=0.1, callbacks=[checkpoint])
# +
fig, axes = plt.subplots(nrows=1, ncols=1, sharex=False, sharey=True, figsize=(20,8))
# Plot accuracy per iteration
plt.plot(model.history.history['acc'][:50], label='acc')
plt.plot(model.history.history['val_acc'][:50], label='val_acc')
plt.legend()
plt.title('Custom Built LSTM RNN Model\'s Training Analysis on the sickness and non-sickness Audio Data', size=16)
plt.xlabel("Epochs")
plt.ylabel("accuracy reached")
plt.show()
| Scripts/Data_Preprocess_Build_Save_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
pd.set_option('display.max_columns', 500)
# ## Loading Data
circuits = pd.read_csv("../data/f1db_csv/circuits.csv", na_values="\\N") # done
constructor_results = pd.read_csv("../data/f1db_csv/constructor_results.csv", na_values="\\N") # done
constructor_standings = pd.read_csv("../data/f1db_csv/constructor_standings.csv", na_values="\\N") # done
constructors = pd.read_csv("../data/f1db_csv/constructors.csv", na_values="\\N") # done
driver_standings = pd.read_csv("../data/f1db_csv/driver_standings.csv", na_values="\\N") # done
drivers = pd.read_csv("../data/f1db_csv/drivers.csv", na_values="\\N") # done
lap_times = pd.read_csv("../data/f1db_csv/lap_times.csv", na_values="\\N") # NA
pit_stops = pd.read_csv("../data/f1db_csv/pit_stops.csv", na_values="\\N") # NA
qualifying = pd.read_csv("../data/f1db_csv/qualifying.csv", na_values="\\N")
races = pd.read_csv("../data/f1db_csv/races.csv", na_values="\\N")
results = pd.read_csv("../data/f1db_csv/results.csv", na_values="\\N") # done
status = pd.read_csv("../data/f1db_csv/status.csv", na_values="\\N")
def finished(x):
if x in [1, 11, 12, 13, 14, 15, 16, 17, 18, 19, 45, 50, 128, 53, 55, 58, 88, 111, 112, 113, 114,
115, 116, 117, 118, 119, 120, 122, 123, 124, 125, 127, 133, 134]:
return 1
else:
return 0
results['finished'] = results.statusId.apply(lambda x: finished(x))
# ## Merging Data
final_df = results[["raceId", "driverId", "constructorId", "grid", "positionOrder", "points", "laps", "rank", "finished"]]
final_df = final_df.rename(columns={"positionOrder":"end_pos", "points":"dr_pts_won", "rank":"fastest_lap_rank", "grid":"start_pos", "laps":"compl_laps"})
# ### Race Information
races.time = races.time.fillna("12:00:00")
races['date'] = races["date"] + " " + races["time"]
races = races.drop(columns=["time", "year", "url"])
races = races.rename(columns={"name":"gp_name", "date":"race_date", "round":"season_round"})
races.gp_name = races.gp_name.apply(lambda x: x.replace(" ", "_").lower())
final_df = final_df.merge(races, on="raceId", how='left')
# ### Circuit Information
circuits = circuits.drop(columns=["circuitRef", "name", "url"])
circuits = circuits.rename(columns={"location":"loc"})
for col in ["loc", "country"]:
circuits[col] = circuits[col].apply(lambda x: x.lower())
final_df = final_df.merge(circuits, on="circuitId", how='left')
# ### Constructor Results
constructor_results.status = constructor_results.status.fillna(0)
constructor_results = constructor_results.drop(columns=["constructorResultsId"])
constructor_results = constructor_results.rename(columns={"points":"constr_pts_won", "status":"constr_disq"})
final_df = final_df.merge(constructor_results, on=["raceId", "constructorId"], how='left')
# ### Constructor Standings
constructor_standings = constructor_standings.drop(columns=["constructorStandingsId", "positionText"])
constructor_standings = constructor_standings.rename(columns={"points":"constr_pts_tot", "position":"constr_pos", "wins":"constr_wins"})
final_df = final_df.merge(constructor_standings, on=["raceId", "constructorId"], how='left')
# ### Constructor
constructors = constructors.drop(columns=["constructorRef", "url"])
constructors["name"] = constructors["name"].apply(lambda x: x.split("-")[0])
teams_to_replace = {"BMW Sauber":"<NAME>", "Sauber":"<NAME>", "Minardi": "AlphaTauri", "<NAME>":"AlphaTauri",
"Toleman":"Alpine F1 Team", "Benetton":"Alpine F1 Team", "Renault":"Alpine F1 Team", "Lotus":"Alpine F1 Team",
"Jordan":"<NAME>", "Spyker":"<NAME>", "Force India": "Aston Martin", "Racing Point":"Aston Martin",
"Tyrrell": "Mercedes", "BAR": "Mercedes", "Honda": "Mercedes", "Brawn": "Mercedes",
"Stewart": "Red Bull", "Jaguar": "Red Bull"
}
constructors.name = constructors.name.replace(teams_to_replace)
constructors = constructors.rename(columns={"name":"constr_name", "nationality":"constr_nat"})
constructors.constr_name = constructors.constr_name.apply(lambda x: x.replace(" ", "_").lower())
constructors.constr_nat = constructors.constr_nat.apply(lambda x: x.replace(" ", "_").lower())
final_df = final_df.merge(constructors, on="constructorId", how='left')
# ### Driver Standings
driver_standings = driver_standings.drop(columns=["positionText", "driverStandingsId"])
driver_standings = driver_standings.rename(columns={"points":"dr_pts_tot", "position":"dr_pos", "wins":"dr_wins"})
final_df = final_df.merge(driver_standings, on=["raceId", "driverId"], how='left')
# ### Driver
drivers["dr"] = drivers["forename"] + " " + drivers["surname"]
drivers.nationality = drivers.nationality.str.lower()
drivers.dr = drivers.dr.apply(lambda x: x.replace(" ", "_").lower())
drivers = drivers.drop(columns=["driverRef", "number", "code", "forename", "surname", "url"])
drivers = drivers.rename(columns={"nationality":"dr_nat"})
final_df = final_df.merge(drivers, on="driverId", how='left')
# ### Qualifying
filling_qual_times = qualifying[["q1", "q2", "q3"]].T.fillna(method="ffill").T
qualifying[["q1", "q2", "q3"]] = filling_qual_times
qualifying = qualifying.drop(columns=["constructorId", "number", "qualifyId"])
qualifying = qualifying.rename(columns={"position":"qual_pos"})
final_df = final_df.merge(qualifying, on=["raceId", "driverId"], how='left')
final_df = final_df.drop(columns=["constructorId", "circuitId"])
final_df.to_csv("../data/raw_processed_data.csv", index=False)
# ## Feature Engineering
ml_df = final_df.copy()
ml_df.dob = pd.to_datetime(ml_df.dob)
ml_df.race_date = pd.to_datetime(ml_df.race_date)
country_conversion = {"american":"usa", "american-italian":"usa", "argentine":"argentina", "argentine-italian":"argentina",
"australian":"australia", "austrian":"austria", "belgian":"belgium", "brazilian":"brazil",
"british":"uk", "canadian":"canada", "chilean":"chile", "colombian":"colombia", "czech":"czech",
"danish":"denmark", "dutch":"netherlands", "east german":"germany", "finnish":"finland",
"french":"france", "german":"germany", "hungarian":"hungary", "indian":"india",
"indonesian":"indonesia", "irish":"ireland", "italian":"italy", "japanese":"japan",
"liechtensteiner":"liechtenstein", "malaysian":"malaysia", "mexican":"mexico",
"monegasque":"monaco", "new zealander":"new zealand", "polish":"poland", "portuguese":"portugal",
"rhodesian":"rhodesia", "russian":"russia", "south african":"south africa", "spanish":"spain",
"swedish":"sweden", "swiss":"switzerland", "thai":"thailand", "uruguayan":"uruguay",
"venezuelan":"venezuela"}
def check_home_gp(country, driver_nationality):
if driver_nationality in country_conversion:
driver_country = country_conversion[driver_nationality]
if driver_country == country:
return 1
else:
return 0
else:
return 0
ml_df["dr_home_gp"] = ml_df.apply(lambda x: check_home_gp(x.country, x.dr_nat), axis=1)
ml_df["constr_home_gp"] = ml_df.apply(lambda x: check_home_gp(x.country, x.constr_nat), axis=1)
ml_df["dr_age"] = 2021 - ml_df.dob.dt.year
ml_df['year'] = ml_df.race_date.dt.year
ml_df['month'] = ml_df.race_date.dt.month
ml_df['day'] = ml_df.race_date.dt.day
ml_df['dr_prev_pts'] = ml_df["dr_pts_tot"] - ml_df["dr_pts_won"]
ml_df['constr_prev_pts'] = ml_df["constr_pts_tot"] - ml_df["constr_pts_won"]
def calculating_prev_wins(wins, position):
if position == 1:
return wins - 1
else:
return wins
ml_df["dr_prev_wins"] = ml_df.apply(lambda x: calculating_prev_wins(x.dr_wins, x.end_pos), axis=1)
ml_df["constr_prev_wins"] = ml_df.apply(lambda x: calculating_prev_wins(x.constr_wins, x.constr_pos), axis=1)
ml_df['dr_compl_laps_prev'] = 0
ml_df['constr_pos_prev'] = 0
ml_df['dr_pos_prev'] = 0
pd.options.mode.chained_assignment = None # default='warn
def get_rolling_statistics(df, col):
copy_df = df.copy()
num_rows = len(copy_df)
stats = copy_df[col].expanding().sum().values[:num_rows-1]
stats = np.insert(stats, 0, 0)
copy_df[col] = stats
return copy_df
def get_shifted_statistics(df, col):
copy_df = df.copy()
num_rows = len(copy_df)
stats = np.insert(copy_df[col].values, 0, 0)
stats = stats[:num_rows]
copy_df[col] = stats
return copy_df
for year in ml_df.year.unique():
season_df = ml_df[ml_df.year == year]
for driver in season_df.driverId.unique():
season_driver_df = season_df[season_df.driverId == driver].sort_values('race_date')
orig_index = season_driver_df.index
for col in ["compl_laps"]:
new_col_name = "dr_compl_laps_prev" # col + "_prev"
new_data_df = get_rolling_statistics(season_driver_df, col)
ml_df[new_col_name].iloc[orig_index] = new_data_df[col]
for col in ["constr_pos", "dr_pos"]:
new_col_name = col + "_prev"
new_data_df = get_shifted_statistics(season_driver_df, col)
ml_df[new_col_name].iloc[orig_index] = new_data_df[col]
ml_df_final = ml_df[[
"race_date", "year",
"month", "day", "season_round",
"start_pos",
"dr", "dr_age", "dr_nat", "dr_home_gp", "dr_prev_pts", "dr_prev_wins", 'dr_compl_laps_prev', 'dr_pos_prev',
"qual_pos", "q1", "q2", "q3",
"constr_name", "constr_nat", "constr_home_gp", "constr_prev_pts", "constr_prev_wins", 'constr_pos_prev',
"gp_name", "alt",
"end_pos", "finished", "fastest_lap_rank"
]]
# # Think about filling missing values
final_df.fastest_lap_rank = final_df.fastest_lap_rank.fillna(25)
ml_df_final.qual_pos = ml_df_final.qual_pos.fillna(ml_df_final.qual_pos.max())
ml_df_final[["q1", "q2", "q3"]] = ml_df_final[["q1", "q2", "q3"]].fillna("3:00:000")
ml_df_final.dr_prev_pts = ml_df_final.dr_prev_pts.fillna(0)
ml_df_final.dr_prev_wins = ml_df_final.dr_prev_wins.fillna(0)
# # -------------------------------
#
# ## Current Drivers
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.metrics import accuracy_score
current_drivers = ['lewis_hamilton', 'max_verstappen', 'valtteri_bottas',
'charles_leclerc', 'sergio_pérez', 'daniel_ricciardo',
'carlos_sainz', 'lando_norris', 'esteban_ocon', 'pierre_gasly',
'lance_stroll', 'kimi_räikkönen', 'sebastian_vettel',
'george_russell', 'antonio_giovinazzi', 'nicholas_latifi',
'fernando_alonso', 'mick_schumacher', 'nikita_mazepin',
'yuki_tsunoda']
current_data = ml_df_final[ml_df_final.dr.isin(current_drivers)]
current_data[["q1", "q2", "q3"]] = current_data[["q1", "q2", "q3"]].fillna("3:00.000")
current_data.qual_pos = current_data.qual_pos.fillna(current_data.start_pos)
# +
current_data.dr_prev_pts = current_data.dr_prev_pts.fillna(0)
current_data.dr_prev_wins = current_data.dr_prev_wins.fillna(0)
current_data.dr_pos_prev = current_data.dr_pos_prev.fillna(current_data.dr_pos_prev.max())
current_data.constr_prev_pts = current_data.constr_prev_pts.fillna(0)
current_data.constr_prev_wins = current_data.constr_prev_wins.fillna(0)
current_data.constr_pos_prev = current_data.constr_pos_prev.fillna(current_data.constr_pos_prev.max())
# -
current_data.fastest_lap_rank = current_data.fastest_lap_rank.fillna(current_data.fastest_lap_rank.max())
current_data = current_data.drop(columns=["race_date"])
def convert_to_seconds(mm_ss_fff):
minute, seconds = mm_ss_fff.split(":")
minute_to_seconds = float(minute) * 60 # seconds from minutes
tot_seconds = minute_to_seconds + float(seconds)
return tot_seconds
current_data.q1 = current_data.q1.apply(lambda x: convert_to_seconds(x))
current_data.q2 = current_data.q2.apply(lambda x: convert_to_seconds(x))
current_data.q3 = current_data.q3.apply(lambda x: convert_to_seconds(x))
current_data = pd.concat([current_data, pd.get_dummies(current_data.dr)], axis=1)
current_data = pd.concat([current_data, pd.get_dummies(current_data.dr_nat)], axis=1)
current_data = pd.concat([current_data, pd.get_dummies(current_data.constr_name)], axis=1)
current_data = pd.concat([current_data, pd.get_dummies(current_data.constr_nat)], axis=1)
current_data = pd.concat([current_data, pd.get_dummies(current_data.gp_name)], axis=1)
current_data = current_data.drop(columns=["dr", "dr_nat", "constr_name", "constr_nat", "gp_name"])
y = current_data[["end_pos", "finished", "fastest_lap_rank"]]
x = current_data.drop(columns=["end_pos", "finished", "fastest_lap_rank"])
y_neural = pd.get_dummies(y.end_pos)
scaler = preprocessing.MinMaxScaler().fit(x.iloc[:, :20])
x.iloc[:, :20] = scaler.transform(x.iloc[:, :20])
# +
# from sklearn.linear_model import LogisticRegression
# clf = LogisticRegression(solver='liblinear').fit(x.values, y.finished.values.astype(int))
# from sklearn.metrics import accuracy_score
# accuracy_score(y.finished, clf.predict(x))
# y.finished.value_counts(normalize=True)
# clf.predict(x)
# -
X_train, X_test, y_train, y_test = train_test_split(x.values, y_neural.values, test_size=0.20)
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
train_dataset = tf.data.Dataset.from_tensor_slices((X_train, y_train))
test_dataset = tf.data.Dataset.from_tensor_slices((X_test, y_test))
BATCH_SIZE = 64
SHUFFLE_BUFFER_SIZE = X_train.shape[0]
train_dataset = train_dataset.shuffle(SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE)
test_dataset = test_dataset.batch(BATCH_SIZE)
model = keras.Sequential()
model.add(layers.Dense(50, input_dim = x.shape[1], activation = 'relu')) # Rectified Linear Unit Activation Function
model.add(layers.Dropout(0.1))
model.add(layers.Dense(50, activation = 'relu'))
model.add(layers.Dropout(0.1))
model.add(layers.Dense(50, activation = 'relu'))
model.add(layers.Dropout(0.1))
model.add(layers.Dense(50, activation = 'relu'))
model.add(layers.Dense(y_neural.shape[1], activation = 'softmax')) # Softmax for multi-class classification
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
EPOCHS=200
history = model.fit(train_dataset, epochs=EPOCHS, validation_data=test_dataset)
import matplotlib.pyplot as plt
plt.plot(np.linspace(1, EPOCHS, EPOCHS), history.history['loss'])
plt.plot(np.linspace(1, EPOCHS, EPOCHS), history.history['val_loss'])
plt.plot(np.linspace(1, EPOCHS, EPOCHS), history.history['accuracy'])
plt.plot(np.linspace(1, EPOCHS, EPOCHS), history.history['val_accuracy'])
history.history.keys()
# # OTHER ------------------------------
# ## Terminara en Puntos
# +
# safe_copy[(safe_copy.raceId > 1050) & (safe_copy.driverId == 1)]
# testing = ml_df[(ml_df.raceId > 1050) & (ml_df.driverId == 1)]
# testing
# testing = ml_df[(ml_df.raceId > 1050) & (ml_df.driverId == 822)]
# testing
# new_data = get_rolling_statistics(testing, "compl_laps")
# ml_df.compl_laps.iloc[testing.index] = new_data.compl_laps
# get_shifted_statistics(testing, "constr_pos")
# get_shifted_statistics(testing, "dr_pos")
# num_rows = len(testing)
# to_insert = np.insert(testing.dr_pos.values, 0, 0)
# to_insert
# to_insert[:num_rows]
# testing.constr_pos.expanding().sum().values[:num_rows-1]
# # season completed laps - function of raceid and driverid
# stats = testing.compl_laps.expanding().sum().values[:num_rows-1]
# import numpy as np
# stats = np.insert(stats, 0, 0)
# orig_index = testing.index
# ml_df.iloc[orig_index]
# testing.compl_laps = stats
# -
in_points = results.merge(races, on="raceId")
current_drivers_ids = [1, 830, 822, 844, 815, 817, 832, 846, 839, 842, 840, 8, 20, 847, 841, 849, 4, 854, 853, 852]
def terminara_en_puntos(driver_id, circuit_id):
driver_circuit = in_points[(in_points.driverId == driver_id) & (in_points.circuitId == circuit_id)][["year", "grid", "circuitId", "position"]]
hamilton_monaco = in_points[(in_points.driverId == 8) & (in_points.circuitId == 6)][["year", "grid", "circuitId", "position"]]
hamilton_monaco = hamilton_monaco.replace("\\N", "20")
hamilton_monaco.position = hamilton_monaco.position.astype(int)
hamilton_monaco = hamilton_monaco.sort_values("year")
hamilton_monaco
hamilton_monaco["top_10"] = hamilton_monaco.position.apply(lambda x: 1 if x <= 10 else 0)
hamilton_monaco.plot("year", "top_10")
clf = LogisticRegression(C=1)
clf.fit(hamilton_monaco[["year", "grid"]], hamilton_monaco.top_10.values.ravel())
plt.figure(figsize=(14,8))
plt.plot(hamilton_monaco.year, hamilton_monaco.top_10)
plt.plot(hamilton_monaco.year, clf.predict(hamilton_monaco[["year", "grid"]]))
plt.plot(hamilton_monaco.year, clf.predict_proba(hamilton_monaco[["year", "grid"]]))
# ## Numero de Pilotos Clasificados
#
# TODO: Add weather data
finished = results.merge(races, on="raceId")
finished = finished[["raceId", "year", "circuitId", "name", "finished"]]
# ### Next Grand Prix
GRAND_PRIX_NAME = "Monaco Grand Prix" # "Spanish Grand Prix"
monaco = finished[finished.name == GRAND_PRIX_NAME].drop(columns=["name", "circuitId", "raceId"])
monaco.head()
monaco = monaco.groupby("year").sum()
monaco = monaco.reset_index()
monaco["over"] = monaco.finished.apply(lambda x: 1 if x > 16.5 else 0)
monaco.plot("year", "finished")
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(C=5000)
clf.fit(monaco.year.values.reshape(-1, 1), monaco.over.values.ravel())
plt.figure(figsize=(14,8))
plt.plot(monaco.year, monaco.over)
plt.plot(monaco.year, clf.predict_proba(monaco[["year"]]))
| notebooks/Ergast_Data.ipynb |