code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yasminghd/2022_ML_Earth_Env_Sci/blob/main/Lab_Notebooks/S4_3_THOR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dUurnKLqq1un"
# <img src='https://unils-my.sharepoint.com/:i:/g/personal/tom_beucler_unil_ch/ESLP1e1BfUxKu-hchh7wZKcBZiG3bJnNbnt0PDDm3BK-9g?download=1'>
#
# <center>
# Photo Credits: <a href="https://unsplash.com/photos/zCMWw56qseM">Sea Foam</a> by <a href="https://unsplash.com/@unstable_affliction"><NAME></a> licensed under the <a href='https://unsplash.com/license'>Unsplash License</a>
# </center>
#
#
# >*A frequently asked question related to this work is “Which mixing processes matter most for climate?” As with many alluringly comprehensive sounding questions, the answer is “it depends.”* <br>
# > $\qquad$ MacKinnon, <NAME>., et al. <br>$\qquad$"Climate process team on internal wave–driven ocean mixing." <br>$\qquad$ Bulletin of the American Meteorological Society 98.11 (2017): 2429-2454.
# + [markdown] id="yT4KFNrpZIfY"
# In week 4's final notebook, we will perform clustering to identify regimes in data taken from the realistic numerical ocean model [Estimating the Circulation and Climate of the Ocean](https://www.ecco-group.org/products-ECCO-V4r4.htm). Sonnewald et al. point out that finding robust regimes is intractable with a naïve approach, so we will be using using reduced dimensionality data.
#
# It is worth pointing out, however, that the reduction was done with an equation instead of one of the algorithms we discussed this week. If you're interested in the full details, you can check out [Sonnewald et al. (2019)](https://doi.org/10.1029/2018EA000519)
# + [markdown] id="PHKbQVLOUC5v"
# # Setup
# + [markdown] id="4TxieA70pcLo"
# First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20.
# + id="S61_Smb9T6ad"
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
import xarray as xr
import pooch
# to make this notebook's output stable across runs
rnd_seed = 42
rnd_gen = np.random.default_rng(rnd_seed)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "dim_reduction"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# + [markdown] id="6vUFq1DYUqCx"
# Here we're going to import the [StandardScaler](https://duckduckgo.com/sklearn.preprocessing.standardscaler) function from scikit's preprocessing tools, import the [scikit clustering library](https://duckduckgo.com/sklearn.clustering), and set up the colormap that we will use when plotting.
# + id="34PAL9TkUHd_"
from sklearn.preprocessing import StandardScaler
import sklearn.cluster as cluster
from matplotlib.colors import LinearSegmentedColormap, ListedColormap
colors = ['royalblue', 'cyan','yellow', 'orange', 'magenta', 'red']
mycmap = ListedColormap(colors)
# + [markdown] id="Fr46ZRUjpTzo"
# # Data Preprocessing
# + [markdown] id="O9QOXpKGpgt7"
# The first thing we need to do is retrieve the list of files we'll be working on. We'll rely on pooch to access the files hosted on the cloud.
# + id="nDNJnzOAfKE_" colab={"base_uri": "https://localhost:8080/"} outputId="03f8b644-9149-4781-c065-7aaebc453f87"
# Retrieve the files from the cloud using Pooch.
data_url = 'https://unils-my.sharepoint.com/:u:/g/personal/tom_beucler_unil_ch/EUYqUzpIjoJBui02QEo6q1wBSN1Zsi1ofE6I3G4B9LJn_Q?download=1'
hash = '3f41661c7a087fa7d7af1d2a8baf95c065468f8a415b8514baedda2f5bc18bb5'
files = pooch.retrieve(data_url, known_hash=hash, processor=pooch.Unzip())
[print(filename) for filename in files];
# + [markdown] id="bZxAvW1gBbO1"
# And now that we have a set of files to load, let's set up a dictionary with the variable names as keys and the data in numpy array format as the values.
# + id="b8OmoBGHBkwG" colab={"base_uri": "https://localhost:8080/"} outputId="0f599a0e-74a1-4a98-ebce-2faed6f51d36"
# Let's read in the variable names from the filepaths
var_names = []
[var_names.append(path.split('/')[-1][:-4]) for path in files]
# And build a dictionary of the data variables keyed to the filenames
data_dict = {}
for idx, val in enumerate(var_names):
data_dict[val] = np.load(files[idx]).T
#We'll print the name of the variable loaded and the associated shape
[print(f'Varname: {item[0]:<15} Shape: {item[1].shape}') for item in data_dict.items()];
# + [markdown] id="RMbhwWTunLvz"
# We now have a dictionary that uses the filename as the key! Feel free to explore the data (e.g., loading the keys, checking the shape of the arrays, plotting)
# + id="rG2pgeH5pB4f" colab={"base_uri": "https://localhost:8080/"} outputId="ccd6caec-f260-493f-b167-dee3951ea8be"
#Feel free to explore the data dictionary
print(data_dict.keys())
print(len(data_dict))
d1 = len(data_dict)
d2 = 0
for d in data_dict:
d2 = max(d2, len(d))
print(f'key: {d} , dimensions: {d1}, {d2}')
# + [markdown] id="xXwnzzbRtTYs"
# We're eventually going to have an array of cluster classes that we're going to use to label dynamic regimes in the ocean. Let's make an array full of NaN (not-a-number) values that has the same shape as our other variables and store it in the data dictionary.
# + id="4VYRgr0cuHKg"
data_dict['clusters'] = np.full_like(data_dict['BPT'],np.nan)
# + [markdown] id="pCumDPFEpFcf"
# ### Reformatting as Xarray
# + [markdown] id="vz_vQOCguax0"
# In the original paper, this data was loaded as numpy arrays. However, we'll take this opportunity to demonstrate the same procedure while relying on xarray. First, let's instantiate a blank dataset.<br><br>
#
# ###**Q1) Make a blank xarray dataset.**<br>
# *Hint: Look at the xarray [documentation](https://duckduckgo.com/?q=xarray+dataset)*
# + id="JIPsLT6g-INq"
# Make your blank dataset here! Instantiate the class without passing any parameters.
ds = xr.Dataset()
# + [markdown] id="jJK9Ud9tp3_r"
# <img src='https://unils-my.sharepoint.com/:i:/g/personal/tom_beucler_unil_ch/EZv_qqVz_h1Hio6Nq11ckScBb01bGb9jtNKzdqAg1TPrKQ?download=1'>
# <center> Image taken from the xarray <a href='https://xarray.pydata.org/en/stable/user-guide/data-structures.html#:~:text=Dataset-,xarray.,from%20the%20netCDF%20file%20format.'> <i>Data Structure documentation</i> </a> </center>
#
# In order to build the dataset, we're going to need a set of coordinate vectors that help us map out our data! For our data, we have two axes corresponding to longitude ($\lambda$) and latitude ($\phi$).
#
# We don't know much about how many lat/lon points we have, so let's explore one of the variables to make sense of the data the shape of one of the numpy arrays.
#
# ###**Q2) Visualize the data using a plot and printing the shape of the data to the console output.**
# + id="Kiw2tAT1D6Ny" colab={"base_uri": "https://localhost:8080/", "height": 493} outputId="fb78df21-946e-4e23-d4cf-9c475a4fd8ae"
#Complete the code
# Let's print out an image of the Bottom Pressure Torques (BPT)
fig , ax = plt .subplots( figsize =(17, 7.5))
sp = ax.imshow( data_dict['BPT'] , origin='lower')
ax.set_xlabel('x coord')
ax.set_ylabel('x coord')
fig.colorbar(sp)
# + id="k6E_4MivCoWQ" colab={"base_uri": "https://localhost:8080/"} outputId="f7d9a67a-7309-42c4-85b6-cf0b684c9663"
# It will also be useful to store and print out the shape of the data
data_shape = data_dict['BPT'].shape
print(data_shape)
# + [markdown] id="aXyxepjm-N5z"
# Now that we know how the resolution of our data, we can prepare a set of axis arrays. We will use these to organize the data we will feed into the dataset.
#
# ###**Q3) Prepare the latitude and longitude arrays to be used as axes for our dataset**
#
# *Hint 1: You can build ordered numpy arrays using, e.g., [numpy.linspace](https://numpy.org/doc/stable/reference/generated/numpy.linspace.html) and [numpy.arange](https://numpy.org/doc/stable/reference/generated/numpy.arange.html)*
#
# *Hint 2: You can rely on the data_shape variable we loaded previously to know how many points you need along each axis*
# + id="vwzO0ZWMNHKg"
#Complete the code
# Let's prepare the lat and lon axes for our data.
lat = np.linspace(0, 180, num=360)
lon = np.linspace(0, 180, num=720)
# + [markdown] id="yxnAU_gjOqVo"
# Now that we have the axes we need, we can build xarray [*data arrays*](https://xarray.pydata.org/en/stable/generated/xarray.DataArray.html) for each data variable. Since we'll be doing it several times, let's go ahead and defined a function that does this for us!
#
# ###**Q4) Define a function that takes in: 1) an array name, 2) a numpy array, 3) a lat vector, and 4) a lon vector. The function should return a dataArray with lat-lon as the coordinate dimensions**
# + id="7FCXLhYfPiQD"
#Complete the code
def np_to_xr(array_name, array, lat, lon):
#building the xarrray
da = xr.DataArray(data = array, # Data to be stored
#set the name of dimensions for the dataArray
dims = ['lat', 'lon'],
#Set the dictionary pointing the name dimensions to np arrays
coords = {'lat':lat,
'lon':lon},
name=array_name)
return da
# + [markdown] id="l9xgQaM9Rrv1"
# We're now ready to build our data array! Let's iterate through the items and merge our blank dataset with the data arrays we create.
#
# ###**Q5) Build the dataset from the data dictionary**
#
# *Hint: We'll be using the xarray merge command to put everything together.*
# + id="4pLXUfvLSZKf"
# The code in the notebook assumes you named your dataset ds. Change it to
# whatever you used!
# Complete the code
for key, item in data_dict.items():
# Let's make use of our np_to_xr function to get the data as a dataArray
da = np_to_xr(key, item, lat, lon)
# Merge the dataSet with the dataArray here!
ds = xr.merge( [ds , da ] )
# + [markdown] id="aC1Gq_q7WEWc"
# Congratulations! You should now have a nicely set up xarray dataset. This let's you access a ton of nice features, e.g.:
# > Data plotting by calling, e.g., `ds.BPT.plot.imshow(cmap='ocean')`
# >
# > Find statistical measures of all variables at once! (e.g.: `ds.std()`, `ds.mean()`)
# + id="uag0JpWzaaS-" colab={"base_uri": "https://localhost:8080/", "height": 664} outputId="34ae27ea-f5e3-4bb0-9c74-fb7dffa12abd"
# Play around with the dataset here if you'd like :)
ds.BPT.plot.imshow(cmap='ocean')
print(ds.std())
print(ds.mean())
# + [markdown] id="vEfuD2O8aZ4w"
# Now we want to find clusters of data considering each grid point as a datapoint with 5 dimensional data. However, we went through a lot of work to get the data nicely associated with a lat and lon - do we really want to undo that?
#
# Luckily, xarray develops foresaw the need to group dimensions together. Let's create a 'flat' version of our dataset using the [`stack`](https://xarray.pydata.org/en/stable/generated/xarray.DataArray.stack.html) method. Let's make a flattened version of our dataset.
#
# ###**Q6) Store a flattened version of our dataset**
#
# *Hint 1: You'll need to pass a dictionary with the 'new' stacked dimension name as the key and the 'flattened' dimensions as the values.*
#
# *Hint 2: xarrays have a ['.values' attribute](https://xarray.pydata.org/en/stable/generated/xarray.DataArray.values.html) that return their data as a numpy array.*
# + id="wOMX7tc5FEZa" outputId="b92067e9-1991-4b43-9e4a-a8bce7cc0bb9" colab={"base_uri": "https://localhost:8080/", "height": 349}
type(ds)
ds.values
ds
# + id="G4e2NDipb0Tr"
# Complete the code
# Let's store the stacked version of our dataset
#stacked = ds.stack( { 'new_dim' :['curlTau' , 'curlCori', 'curlA', 'BPT', 'BPT'] } )
stacked = ds.stack(new =['lat' , 'lon'])
# + id="UQt4-fFACtUm" colab={"base_uri": "https://localhost:8080/"} outputId="03b1cb72-0d30-414b-eca0-f4a92645e314"
# And verify the shape of our data
#print(stacked.to_array()._____._____)
print(stacked.to_array().values.shape)
# + [markdown] id="MZGPnmooffKy"
# So far we've ignored an important point - we're supposed to have 5 variables, not 6! As you may have guessed, `noiseMask` helps us throw away data we dont want (e.g., from land mass or bad pixels).
#
# We're now going to clean up the stacked dataset using the noise mask. Relax and read through the code, since there won't be a question in this part :)
# + id="sbXscaE_fcOj" colab={"base_uri": "https://localhost:8080/"} outputId="7e0b8383-f81a-4dee-f9f7-b85ce7e85618"
# Let's redefine stacked as all the points where noiseMask = 1, since noisemask
# is binary data.
print(f'Dataset shape before processing: {stacked.to_array().values.shape}')
print("Let's do some data cleaning!")
print(f'Points before cleaning: {len(stacked.BPT)}')
stacked = stacked.where(stacked.noiseMask==1, drop=True)
print(f'Points after cleaning: {len(stacked.BPT)}')
# + id="crX5HoD3C3Ls" colab={"base_uri": "https://localhost:8080/"} outputId="6532b12f-1ff0-4f76-d677-fc76cda3bc3b"
# We also no longer need the noiseMask variable, so we can just drop it.
print('And drop the noisemask variable...')
print(f'Before dropping: {stacked.to_array().values.shape}')
stacked = stacked.drop('noiseMask')
print(f'Dataset shape after processing: {stacked.to_array().values.shape}')
# + [markdown] id="5Pol90vsU_wv"
# We now have several thousand points which we want to divide into clusters using the kmeans clustering algorithm (you can check out the documentation for scikit's implementation of kmeans [here](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html)).
#
# You'll note that the algorithm expects the input data `X` to be fed as `(n_samples, n_features)`. This is the opposite of what we have! Let's go ahead and make a copy to a numpy array has the axes in the right order.
#
# You'll need xarray's [`.to_array()`](https://xarray.pydata.org/en/stable/generated/xarray.Dataset.to_array.html) method and [`.values`](https://xarray.pydata.org/en/stable/generated/xarray.DataArray.values.html) parameter, as well as numpy's [`.moveaxis`](https://numpy.org/doc/stable/reference/generated/numpy.moveaxis.html) method.
#
# ###**Q7) Load the datapoints into a numpy array following the convention where the 0th axis corresponds to the samples and the 1st axis corresponds to the features.**
# + id="uu0AVVkPeTzC" colab={"base_uri": "https://localhost:8080/", "height": 396} outputId="dc8d1ccc-0f16-464e-bea1-cab2c7684244"
# Complete the code
input_data = np.moveaxis(stacked.to_array().values, # data to reshape
'number', # source axis as integer,
'number') # destination axis as integer
# + id="Vi9GZ-XsC6Nd"
# Does the input data look the way it's supposed to? Print the shape.
print(________)
# + [markdown] id="AoZSkMieiL9O"
# In previous classes we discussed the importance of the scaling the data before implementing our algorithms. Now that our data is all but ready to be fed into an algorithm, let's make sure that it's been scaled.
#
# ###**Q8) Scale the input data**
#
# *Hint 1: Import the [`StandardScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html) class from scikit and instantiate it*
#
# *Hint 2: Update the input array to the one returned by the [`.fit_transform(X)`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler.fit_transform) method*
# + id="D2sZPWf1ignO"
#Write your scaling code here
# + [markdown] id="_z3e6dU8rInv"
# Now we're finally ready to train our algorithm! Let's load up the kmeans model and find clusters in our data.
#
# ###**Q9) Instantiate the kmeans clustering algorithm, and then fit it using 50 clusters, trying out 10 different initial centroids.**
#
# *Hint 1: `sklearn.cluster` was imported as `cluser` during the notebook setup! [Here is the scikit `KMeans` documentation](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html).*
#
# *Hint 2: Use the `fit_predict` method to organize the data into clusters*
#
# *Warning! : Fitting the data may take some time (under a minute during the testing of the notebook)
# + id="DtYyiksRrHPm"
# Complete the code
kmeans = cluster._____(________ =50, # Number of clusters
________ =42, # setting a random state
________ =10, # Number of initial centroid states to try
verbose = 1) # Verbosity so we know things are working
# + id="ohuw1WFiC8kS"
cluster_labels = kmeans.______(____) # Feed in out scaled input data!
# + [markdown] id="XZXYUggc8xex"
# We now have a set of cluster labels that group the data into 50 similar groups. Let's store it in our stacked dataset!
# + id="aH0qNHM89GER"
# Let's run this line
stacked['clusters'].values = cluster_labels
# + [markdown] id="pFPN00oH9oI8"
# We now have a set of labels, but they're stored in a flattened array. Since we'd like to see the data as a map, we still have some work to do. Let's go back to a 2D representation of our values.
#
# ###**Q10) Turn the flattened xarray back into a set of 2D fields**
# *Hint*: xarrays have an [`.unstack` method](https://xarray.pydata.org/en/stable/generated/xarray.DataArray.unstack.html) that you will find to be very useful for this.
# + id="lp1u28inKUNT"
# Complete the code:
processed_ds = ds.____()
# + [markdown] id="y4w0zUDn8OOU"
# Now we have an unstacked dataset, and can now easily plot out the clusters we found!
#
# ###**Q11) Plot the 'cluster' variable using the buil-in xarray function**
# *Hint: `.plot()` [link text](https://xarray.pydata.org/en/stable/generated/xarray.DataArray.plot.html) let's you access the xarray implementations of [`pcolormesh`](https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.pcolormesh.html) and [`imshow`](https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.imshow.html).*
# + id="DLLCcgw_n3tu"
# + [markdown] id="sOja6JCxo45g"
# Compare your results to those from the paper:
# <img src='https://unils-my.sharepoint.com/:i:/g/personal/tom_beucler_unil_ch/EdLh6Ds0yVlFivyfIOXlV74B_G35dVz87GFagzylIG-gZA?download=1'>
# + [markdown] id="HrLKLVWXtgeX"
# We now want to find the 5 most common regimes, and group the rest. This isn't straightforward, so we've gone ahead and prepared the code for you. Run through it and try to understand what the code is doing!
# + id="UO5OohbKptau"
# Make field filled with -1 vals so unprocessed points are easily retrieved.
# Noise masked applied automatically by using previously found labels as base.
processed_ds['final_clusters'] = (processed_ds.clusters * 0) - 1
# Find the 5 most common cluster labels
top_clusters = processed_ds.groupby('clusters').count().sortby('BPT').tail(5).clusters.values
#Build the set of indices for the cluster data, used for rewriting cluster labels
for idx, label in enumerate(top_clusters):
#Find the indices where the label is found
indices = (processed_ds.clusters == label)
processed_ds['final_clusters'].values[indices] = 4-idx
# Set the remaining unlabeled regions to category 5 "non-linear"
processed_ds['final_clusters'].values[processed_ds.final_clusters==-1] = 5
# Plot the figure
processed_ds.final_clusters.plot.imshow(cmap=mycmap, figsize=(18,8));
# + id="IHSj9yRaUCSg"
# Feel free to use this space
# + [markdown] id="eEMmkpGkc1Xy"
# Compare it to the regimes found in the paper:
# <img src='https://unils-my.sharepoint.com/:i:/g/personal/tom_beucler_unil_ch/EehuR9cUfaJImrw4DCAzDPoBiGuG7R3Ys6453Umi1cN_OQ?download=1'>
#
#
# + [markdown] id="nbWgoD7bv8AH"
# The authors then went on to train neural networks ***to infer in-depth dynamics from data that is largely readily available from for example CMIP6 models, using NN methods to infer the source of predictive skill*** and ***to apply the trained Ensemble MLP to a climate model in order to assess circulation changes under global heating***.
#
# For our purposes, however, we will say goodbye to *THOR* at this point 😃
| Lab_Notebooks/S4_3_THOR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 10.3 word2vec的实现
# +
import collections
import math
import random
import sys
import time
import os
import numpy as np
import torch
from torch import nn
import torch.nn.functional as F
import torch.utils.data as Data
sys.path.append("..")
import d2lzh_pytorch as d2l
print(torch.__version__)
# -
# ## 10.3.1 处理数据集
assert 'ptb.train.txt' in os.listdir("../../data/ptb")
# +
with open('../../data/ptb/ptb.train.txt', 'r') as f:
lines = f.readlines()
# st是sentence的缩写
raw_dataset = [st.split() for st in lines]
'# sentences: %d' % len(raw_dataset)
# +
# raw_dataset[0]
# -
for st in raw_dataset[:3]:
print('# tokens:', len(st), st[:])
# ### 10.3.1.1 建立词语索引
# tk是token的缩写
counter = collections.Counter([tk for st in raw_dataset for tk in st])
# counter.items()
# 为了计算简单,我们只保留在数据集中至少出现5次的词
# 除去出现次数少于5的字符
counter = dict(filter(lambda x: x[1] >= 5, counter.items()))
# counter
idx_to_token = [tk for tk, _ in counter.items()]
# idx_to_token
token_to_idx = {tk: idx for idx, tk in enumerate(idx_to_token)}
list(token_to_idx.items())[-1], len(token_to_idx), token_to_idx
dataset = [[token_to_idx[tk] for tk in st if tk in token_to_idx]
for st in raw_dataset]
dataset[1]
num_tokens = sum([len(st) for st in dataset])
'# tokens: %d' % num_tokens
# ### 10.3.1.2 二次采样
# +
def discard(idx):
return random.uniform(0, 1) < 1 - math.sqrt(
1e-4 / counter[idx_to_token[idx]] * num_tokens)
subsampled_dataset = [[tk for tk in st if not discard(tk)] for st in dataset]
'# tokens: %d' % sum([len(st) for st in subsampled_dataset])
# -
[idx_to_token[idx] for idx in subsampled_dataset[2]]
# +
def compare_counts(token):
return '# %s: before=%d, after=%d' % (token, sum(
[st.count(token_to_idx[token]) for st in dataset]), sum(
[st.count(token_to_idx[token]) for st in subsampled_dataset]))
compare_counts('the')
# -
compare_counts('join')
compare_counts('mom')
# ### 10.3.1.3 提取中心词和背景词
def get_centers_and_contexts(dataset, max_window_size):
centers, contexts = [], []
for st in dataset:
if len(st) < 2: # 每个句子至少要有2个词才可能组成一对“中心词-背景词”
continue
centers += st
for center_i in range(len(st)):
window_size = random.randint(1, max_window_size)
indices = list(range(max(0, center_i - window_size),
min(len(st), center_i + 1 + window_size)))
indices.remove(center_i) # 将中心词排除在背景词之外
contexts.append([st[idx] for idx in indices])
return centers, contexts
tiny_dataset = [list(range(7)), list(range(7, 10))]
print('dataset', tiny_dataset)
for center, context in zip(*get_centers_and_contexts(tiny_dataset, 2)):
print('center', center, 'has contexts', context)
all_centers, all_contexts = get_centers_and_contexts(subsampled_dataset, 5)
all_centers[100], all_contexts[100]
idx_to_token[all_centers[101]], [idx_to_token[idx] for idx in all_contexts[101]]
# ## 10.3.2 负采样
# +
# idx_to_token
# -
sampling_weights = [counter[w]**0.75 for w in idx_to_token]
# sampling_weights
# +
def get_negatives(all_contexts, sampling_weights, K):
all_negatives, neg_candidates, i = [], [], 0
population = list(range(len(sampling_weights)))
for contexts in all_contexts:
negatives = []
while len(negatives) < len(contexts) * K:
if i == len(neg_candidates):
# 根据每个词的权重(sampling_weights)随机生成k个词的索引作为噪声词。
# 为了高效计算,可以将k设得稍大一点
i, neg_candidates = 0, random.choices(
population, sampling_weights, k=int(1e5))
neg, i = neg_candidates[i], i + 1
# 噪声词不能是背景词
if neg not in set(contexts):
negatives.append(neg)
all_negatives.append(negatives)
return all_negatives
all_negatives = get_negatives(all_contexts, sampling_weights, 5)
# -
all_centers[0], all_contexts[0], all_negatives[0]
# ## 10.3.3 读取数据
# * Basically, the collate_fn receives a list of tuples if your `__getitem__` function from a Dataset subclass returns a tuple, or just a normal list if your Dataset subclass returns only one element. Its main objective is to create your batch without spending much time implementing it manually. Try to see it as a glue that you specify the way examples stick together in a batch. If you don’t use it, PyTorch only put batch_size examples together as you would using torch.stack (not exactly it, but it is simple like that).
#
# * Suppose for example, you want to create batches of a list of varying dimension tensors. The below code pads sequences with 0 until the maximum sequence size of the batch, that is why we need the `collate_fn`, because a standard batching algorithm (simply using `torch.stack`) won’t work in this case, and we need to manually pad different sequences with variable length to the same size before creating the batch.
idx_to_token[0]
def batchify(data):
"""用作DataLoader的参数collate_fn: 输入是个长为batchsize的list, list中的每个元素都是__getitem__得到的结果"""
max_len = max(len(c) + len(n) for _, c, n in data)
# print(f"max_len - {max_len}")
centers, contexts_negatives, masks, labels = [], [], [], []
for center, context, negative in data:
cur_len = len(context) + len(negative)
centers += [center]
# print(f"centers - {centers}")
contexts_negatives += [context + negative + [0] * (max_len - cur_len)]
masks += [[1] * cur_len + [0] * (max_len - cur_len)]
labels += [[1] * len(context) + [0] * (max_len - len(context))]
return (torch.tensor(centers).view(-1, 1), torch.tensor(contexts_negatives),
torch.tensor(masks), torch.tensor(labels))
# +
batch_size = 512
example_data = zip(
all_centers[:batch_size],
all_contexts[:batch_size],
all_negatives[:batch_size]
)
# for center, context, negative in example_data:
# print(f"{center}\n{context}\n{negative}")
# break
# +
# batchify(list(example_data))
# -
class MyDataset(torch.utils.data.Dataset):
def __init__(self, centers, contexts, negatives):
assert len(centers) == len(contexts) == len(negatives)
self.centers = centers
self.contexts = contexts
self.negatives = negatives
def __getitem__(self, index):
return (self.centers[index], self.contexts[index], self.negatives[index])
def __len__(self):
return len(self.centers)
# +
batch_size = 512
num_workers = 0 if sys.platform.startswith('win32') else 16
dataset = MyDataset(
all_centers,
all_contexts,
all_negatives
)
# +
# len(dataset), dataset[0], num_workers
# -
data_iter = Data.DataLoader(
dataset,
batch_size,
shuffle=True,
collate_fn=batchify,
# num_workers=num_workers
)
# +
# data_iter
# -
for i, batch in enumerate(data_iter):
for name, data in zip(['centers', 'contexts_negatives', 'masks',
'labels'], batch):
print(name, 'shape:', data.shape)
# print(batch)
break
# ## 10.3.4 跳字模型
# ### 10.3.4.1 嵌入层
# +
num_embeddings = len(idx_to_token)
embed_size = 100
embed = nn.Embedding(
num_embeddings=num_embeddings,
embedding_dim=embed_size
)
embed.weight.shape
# -
x = torch.tensor([[1, 2, 3], [4, 5, 6]], dtype=torch.long)
embed(x).shape
# embed(x)[0][0] == embed.weight[1]
# +
c = torch.zeros((3, 1), dtype=torch.long)
o = torch.ones((3, 10), dtype=torch.long)
v = embed(c)
u = embed(o)
v.shape, u.shape
# -
torch.bmm(v, u.permute(0, 2, 1)).shape
# ### 10.3.4.2 小批量乘法
X = torch.ones((2, 1, 4))
Y = torch.ones((2, 4, 6))
torch.bmm(X, Y).shape, X
# ### 10.3.4.3 跳字模型前向计算
def skip_gram(center, contexts_and_negatives, embed_v, embed_u):
v = embed_v(center)
u = embed_u(contexts_and_negatives)
pred = torch.bmm(v, u.permute(0, 2, 1))
return pred
# ## 10.3.5 训练模型
# ### 10.3.5.1 二元交叉熵损失函数
# +
class SigmoidBinaryCrossEntropyLoss(nn.Module):
def __init__(self): # none mean sum
super(SigmoidBinaryCrossEntropyLoss, self).__init__()
def forward(self, inputs, targets, mask=None):
"""
input – Tensor shape: (batch_size, len)
target – Tensor of the same shape as input
"""
inputs, targets, mask = inputs.float(), targets.float(), mask.float()
res = nn.functional.binary_cross_entropy_with_logits(inputs, targets, reduction="none", weight=mask)
return res.mean(dim=1)
loss = SigmoidBinaryCrossEntropyLoss()
# -
# 值得一提的是,我们可以通过掩码变量指定小批量中参与损失函数计算的部分预测值和标签:当掩码为1时,相应位置的预测值和标签将参与损失函数的计算;当掩码为0时,相应位置的预测值和标签则不参与损失函数的计算。我们之前提到,掩码变量可用于避免填充项对损失函数计算的影响。
pred = torch.tensor([[1.5, 0.3, -1, 2], [1.1, -0.6, 2.2, 0.4]], dtype = torch.float)
# 标签变量label中的1和0分别代表背景词和噪声词
label = torch.tensor([[1, 0, 0, 0], [1, 1, 0, 0]], dtype = torch.float)
mask = torch.tensor([[1, 1, 1, 1], [1, 1, 1, 0]]) # 掩码变量
loss(pred, label, mask) * mask.shape[1] / mask.float().sum(dim=1)
loss(pred, label, mask)* mask.shape[1]
mask.shape[1], mask.float().sum(dim=1)
F.binary_cross_entropy_with_logits(pred[0], label[0]), F.binary_cross_entropy_with_logits(pred[1], label[1], weight=mask[1])*4/3
# ### 10.3.5.2 初始化模型参数
len(idx_to_token)
num_embeddings = len(idx_to_token)
embed_size = 100
net = nn.Sequential(
nn.Embedding(num_embeddings=num_embeddings, embedding_dim=embed_size),
nn.Embedding(num_embeddings=num_embeddings, embedding_dim=embed_size)
)
# ### 10.3.5.3 定义训练函数
def train(net, lr, num_epochs):
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print("train on", device)
net = net.to(device)
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
for epoch in range(num_epochs):
start, l_sum, n = time.time(), 0.0, 0
for batch in data_iter:
center, context_negative, mask, label = [d.to(device) for d in batch]
pred = skip_gram(center, context_negative, net[0], net[1])
# 使用掩码变量mask来避免填充项对损失函数计算的影响
l = (loss(pred.view(label.shape), label, mask) *
mask.shape[1] / mask.float().sum(dim=1)).mean() # 一个batch的平均loss
optimizer.zero_grad()
l.backward()
optimizer.step()
l_sum += l.cpu().item()
n += 1
print('epoch %d, loss %.2f, time %.2fs'
% (epoch + 1, l_sum / n, time.time() - start))
train(net, 0.01, 10)
for name, param in net.named_parameters():
print(f"{name} - {param.shape}")
# ## 10.3.6 应用词嵌入模型
# +
def get_similar_tokens(query_token, k, embed):
W = embed.weight.data
x = W[token_to_idx[query_token]]
# 添加的1e-9是为了数值稳定性
cos = torch.matmul(W, x) / (torch.sum(W * W, dim=1) * torch.sum(x * x) + 1e-9).sqrt()
_, topk = torch.topk(cos, k=k+1)
topk = topk.cpu().numpy()
for i in topk[:]: # 除去输入词
print('cosine sim=%.3f: %s' % (cos[i], (idx_to_token[i])))
get_similar_tokens('chip', 3, net[0])
# -
man_idx = token_to_idx["man"]
woman_idx = token_to_idx["woman"]
father_idx = token_to_idx["father"]
mother_idx = token_to_idx["mather"]
man_vector = net[0].weight.data[man_idx]
woman_vector = net[0].weight.data[woman_idx]
father_vector = net[0].weight.data[father_idx]
mother_vector = net[0].weight.data[mother_idx]
v1 = man_vector-woman_vector
v2 = father_vector-mother_vector
cos = torch.matmul(v1, v2) / (torch.sum(v1 * v1) * torch.sum(v2 * v2)).sqrt()
cos
| code/chapter10_natural-language-processing/10.3_word2vec-pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# line 1 note
import csv
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
import seaborn as sns
import datetime
# line 2 note
data = pd.read_csv('Datamart-Export_DY_WK100-500 Pound Barrel Cheddar Cheese Prices, Sales, and Moisture Content_20170829_122601.csv')
# line 3 note
data.shape
# line 4 note
data.head(10)
# line 5 Verify that we are working with a data frame. We ‘know’ we have a data frame, type(data) verifies that the data was read in as a data frame .
type(data)
data.dtypes
# Line 6: See if we have any missing data. We don’t have any missing data. We are asking if there are any null values. If we had any null values we would get ‘True” but since everything is ‘False’ we don’t have any null values.
data.isnull().any()
# Line 7: Use data.describe to see an overview of the data means, median, max, min, etc for each column. This gives us information on the columns with data, but the columns with word descriptors (such as salary with low, medium & high are not included). This is an interesting feature that gives us some statistical information about our data.
data.describe()
# line 8: convert dates
#I don't need this
wedate=data['Week Ending Date']
#type(rptdt) pandas.core.series.Series
wedate.head(10)
# line?: I tried various ways to change the 'Date' column into a date format. So I ended up "forcing" a temporary date with ta fake year then force it into a date format & then split out the day and month and then add it to the yer
data['temp']=data['Date'].astype(basestring)+'-1999'
data.head(5)
# Line ? extract the year from the week ending date
# +
#type(rptdt) pandas.core.series.Series
data['Week Ending Date'] = pd.to_datetime(data['Week Ending Date'])
data['year'] = data['Week Ending Date'].dt.year
data.head(5)
# -
# line next line is all junk...
# +
#type(date) pandas.core.series.Series
#data['Date'] = pd.to_datetime(data['Date']) OutOfBoundsDatetime: Out of bounds nanosecond timestamp: 1-07-22 00:00:00
#data['month']= data['Date'].dt.month
#newdate = datetime.datetime.strptime("Date", '%d-%b')
#a = datetime.datetime(date) TypeError: cannot convert the series to <type 'int'>
#datetime_object = datetime.strptime('date', '%b-%d')
#datetime.strptime('Date', '%d-%b')
#a=date.strip('-') AttributeError: 'Series' object has no attribute 'strip'
#a.head(10)
#type(rptdt) pandas.core.series.Series
#for d in data:
# a=data['Date'].strip('-') 'Series' object has no attribute 'strip'
#a[0]=data['day']
# print(a)
#if a[0]=='Jan':
# data['month']='01'
# else:
# data['month']='13'
#day=data['Date'].iloc[0:1409,'Date'] IndexingError: Too many indexers
#day
#type(data['Date']) pandas.core.series.Series
#data['day'] = data['Date'].dt.day AttributeError: Can only use .dt accessor with datetimelike valu
#data['temp']=data['Date']+'-'
#newdate='{0}/{1}/'.format(dt.month, dt.day, 'year' % 100)
#newdate.head(10)
#print '{0}/{1}/{2:02}'.format(dt.month, dt.day, dt.year % 100)
# -
# line? this splits the date into the month and year form the date format we forced into creation
#data[newtemp]=data['temp']+data['year'] TypeError: ufunc 'add' did not contain a loop with signature matching types dtype('S21') dtype('S21') dtype('S21')
#data['month']=data['temp'].strip('-') 'Series' object has no attribute 'strip'
#type(data['temp']) pandas.core.series.Series
#newd=datetime.datetime.strptime("temp", '%d-%b-%Y')
#data.head(10)
data['temp'] = pd.to_datetime(data['temp'])
data['month'],data['day'] = data['temp'].dt.month, data['temp'].dt.day
data.head(10)
# line ? replace our fake year with the correct year by combining the year, month and day
# +
#data['temp']=data['year']+'-'+data['month']+'-'+data['day'] TypeError: ufunc 'add' did not contain a loop with signature matching types dtype('S21') dtype('S21') dtype('S21')
#data['temp']=pd.concat(data['year'],data['month'],data['day']) TypeError: first argument must be an iterable of pandas objects, you passed an object of type "Series"
#data['temp']=data['year'].astype(basestring)+'-'+data['month'].astype(basestring)+'-'+data['day'].astype(basestring) TypeError: unsupported operand type(s) for +: 'long' and 'str'
#data['temp']=data['year'].astype(basestring)+'-1900' TypeError: unsupported operand type(s) for +: 'long' and 'str'
#data['temp']=pd.Series([data['year'], data['month'], data['day']]).str.cat(sep='-') TypeError: sequence item 0: expected string, Series found
#data['temp']=pd.Series([data['year'].astype(basestring), data['month'].astype(basestring),data['day'].astype(basestring)]).str.cat(sep='-')
#newd=datetime.strptime("temp", '%Y-%m-%d') AttributeError: 'module' object has no attribute 'strptime'
#newd=datetime.strptime(data["temp"], '%Y-%m-%d') AttributeError: 'module' object has no attribute 'strptime'
data['temp'] = data['year'].map(str)+'-' + data['month'].map(str) +'-'+ data['day'].map(str)
#data['temp'] = data['year'].map(str)+'-' + data['month'].map(str) +'-'+ data['day'].map(str) works but won't read as date
#data['temp'] = data['year'].map(str,)+data['month'].map(str) +data['day'].map(str) works byt won't read as date
#type(data['temp']) pandas.core.series.Series
data.head(5)
# -
# line now we need to convert the newly converted temp into a date format object
# +
#df = (data['temp'], sep='-', parse_dates=[0])
#df.info()
#newdate = datetime.datetime.strptime("2017/7/22", '%Y/%m/%d').strftime('%m/%d/%y') works '07/22/17'
#data['temp']=data['temp'].astype(basestring) type is pandas.core.series.Series
#data['temp'].to_string() type is pandas.core.series.Series
#data.temp.apply(str) type is pandas.core.series.Series
#data['temp'].astype(basestring) type is pandas.core.series.Series
#data['temp']= data['temp'].astype(str) type is pandas.core.series.Series
#data['temp'] = data['temp'].astype('datetime64') ValueError: Error parsing datetime string "2012/3/3" at position 4
data['temp'] = pd.to_datetime(data['temp'])
data['Week Ending Date'] = pd.to_datetime(data['Week Ending Date'])
data.head(5)
#data['Week Ending Date'] = data['Week Ending Date'].astype('datetime64') ValueError: Error parsing datetime string "3/31/2012" at position 1
#type(data['Week Ending Date'])
#newdate = datetime.datetime.strptime(data["temp"], '%Y/%m/%d').strftime('%m/%d/%y')
#newdate = datetime.datetime.strptime(data["temp"], '%Y/%m/%d')
#newdate
# -
# line ? check our data types
data.dtypes
# line convert sales to an integer
# +
#data['Sales'].astype(str).astype(int)
#data['Sales'].astype(str)
#data['Sales'].astype(int)
#pd.factorize(data['Sales'])[0] object
#data['Sales'].astype(basestring).astype(int)
data['Sales'] = data['Sales'].str.replace(',', '')
data['Sales'] = pd.to_numeric(data['Sales']) #or data['Sales'] data['Sales'].astype(int)
data.dtypes
# -
# line? now that we have a date format in week ending & the temp column we can nw get the difference to find the age of the cheese.
#age=data['Week Ending Date']-data['temp'] works
#age.strip( 'days')
#age.replace( 'days','weeks')
#age.astype(basestring)
data['age']=data['Week Ending Date']-data['temp']
data['age'].head(10)
#data.head(10)
# line ? ok so now we have our dates converted to dates and we've used those to find the age of the cheese. our data frame now looks like:
#
data.head(5)
# line ? lets drop the day colum we created we'll keep month & year for now
df1=data
#df1.drop(['year', 'month', 'day'],axis=1)
#del df1['year', 'month', 'day']
#del df1['year']
#del df1[data['year']]
#del df1['month']
#del df1['day']
del df1['day']
df1.head(5)
# line ?:so now that we have the age of the cheese we can compare pricing age to age
#Summarizwe the data by age
years = data.set_index("year")
#Get the sums of sales of each region by year
#years = years.groupby("Year").sum()
#View the first 5 entries of the data to see what we are working with
years.head()
line? need to correct dec dates for year flip
# +
#sns.barplot(x="Sales",y="age",data=df1,estimator=sum)
#df1.plot(x='Week Ending Date', y='age')
df2017=data[data.year == 2017]
df2017
#sns.FacetGrid(df1, hue="age", size=3) \
# .map(sns.kdeplot, "Sales") \
# .add_legend()
#plt.plot('Week Ending Date', 'age')
#plt.legend()
# -
# line? need to correct
| proj3_cheese_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # install package
# ! pip install .
import lda_package
import numpy as np
from scipy.special import digamma, polygamma
import matplotlib.pyplot as plt
# # Simulated data example
# +
# create sample documents
doc_a = """TheWilliam Randolph Hearst Foundation will give 1.25 million to Lincoln Center, Metropolitan
Opera Co., New York Philharmonic and Juilliard School."""
doc_b = """ “Our board felt that we had a real opportunity to make a mark on the future of the performing
arts with these grants an act every bit as important as our traditional areas of support in health,
medical research, education and the social services,” Hearst Foundation President <NAME>
said Monday in announcing the grants."""
doc_c = """Lincoln Center’s share will be 200000 for its new building, which
will house young artists and provide new public facilities."""
doc_d = """The Metropolitan Opera Co. and New York Philharmonic will receive 400000 each."""
doc_e = """The Juilliard School, where music and the performing arts are taught, will get 250000. """
doc_f = """The Hearst Foundation, a leading supporter of the Lincoln Center Consolidated Corporate Fund,
will make its usual annual $100,000 donation, too."""
doc_set = [doc_a, doc_b, doc_c, doc_d, doc_e, doc_f]
# -
# # Clean Data
# +
# pip install nltk
# pip install stop-words
# easy_install -U gensim
# -
def data_clean(doc_set, stop_word = None):
"""
REF: https://rstudio-pubs-static.s3.amazonaws.com/79360_850b2a69980c4488b1db95987a24867a.html
input:
doc_set: a list of documets, the elements are context of that document
stop_word: if 'stop_word = None', this function will give a sample
output:
texts:a list of array(documents), each element contains all words in that document
dictionary: a dictionary, key is the id of words, values are unique words
corpus: a list of list, each inner list represents a document. In the inner list, each tuple is (word_id, word_count)
"""
from nltk.tokenize import RegexpTokenizer
from stop_words import get_stop_words
from nltk.stem.porter import PorterStemmer
from gensim import corpora, models
import gensim
tokenizer = RegexpTokenizer(r'\w+')
# create English stop words list
if stop_word==None:
stop_word = get_stop_words('en')
# Create p_stemmer of class PorterStemmer
p_stemmer = PorterStemmer()
# list for tokenized documents in loop
texts = []
# loop through document list
for i in doc_set:
# clean and tokenize document string
raw = i.lower()
tokens = tokenizer.tokenize(raw)
# remove stop words from tokens
stopped_tokens = [i for i in tokens if not i in stop_word]
# stem tokens
stemmed_tokens = [p_stemmer.stem(i) for i in stopped_tokens]
# add tokens to list
if len(stemmed_tokens) > 0:
texts.append(stemmed_tokens)
# turn our tokenized documents into a id <-> term dictionary
dictionary = corpora.Dictionary(texts)
# convert tokenized documents into a document-term matrix
corpus = [dictionary.doc2bow(text) for text in texts]
return texts, dictionary, corpus
def data_process(texts, dictionary):
"""
transform the texts from word-formation into id-formation, which can be used in function variantion_EM()
"""
import numpy as np
text_ = []
for i in range(len(texts)):
text_i = []
for l in range(len(texts[i])):
text_i = np.append(text_i, dictionary.token2id[texts[i][l]])
text_.append(text_i)
return text_
texts, dictionary, corpus = data_clean(doc_set)
text_ = data_process(texts, dictionary)
# # Experimental test
# +
np.random.seed(64528)
M = 6
k = 4
N = np.array(list(map(len, text_)))
V = len(dictionary)
V_words = range(V)
alpha = np.random.dirichlet(10*np.ones(k),1)[0]
beta = np.random.dirichlet(np.ones(V),k)
phi = np.array([1/k*np.ones([N[m],k]) for m in range(M)])
gamma = np.tile(alpha,(M,1)) + np.tile(N/k,(k,1)).T
# -
ans = lda_package.variation_EM(M, k, text_, N, V_words, alpha, beta, gamma, phi, iteration = 1000)
def sigmoid(x):
return 1/(1+np.exp(-x))
fig, ax = plt.subplots(figsize=(18,8))
ax.imshow(sigmoid(100*(ans[1]-0.05)), interpolation='nearest', aspect='auto')
ax.set_xlabel("word")
plt.yticks([0, 1, 2, 3], ["topic 1", "topic 2","topic 3", "topic 4"])
pass
m = dictionary.token2id
id2token = dict(zip(m.values(), m.keys()))
order = list(map(np.argsort, ans[1]))
for i in range(len(order)):
print("topic", (i+1), "\n", ans[1][i][order[i][-1]], "*", id2token[order[i][-1]], "+",
ans[1][i][order[i][-2]], "*", id2token[order[i][-2]], "+",
ans[1][i][order[i][-3]], "*", id2token[order[i][-3]], "+",
ans[1][i][order[i][-4]], "*", id2token[order[i][-4]], '\n')
# # Real world data
f = open('Test_data/vocab.txt', 'r')
vocab = f.read().splitlines()
f.close()
f = open('Test_data/ap.txt', 'r')
ap = f.read().splitlines()
f.close()
f = open('stop_word.txt', 'r')
stop_word = f.read().splitlines()
#stop_word = list(map(stop_word, strip)
f.close()
stop_word = [word.strip() for word in stop_word]
texts, dictionary, corpus = data_clean(ap, stop_word)
text_ = data_process(texts, dictionary)
file=open('text_real_world.txt','w')
file.write(str(text_));
file.close()
# +
np.random.seed(10986)
M = len(texts)
k = 10
N = np.array(list(map(len, text_)))
V = len(dictionary)
V_words = range(V)
alpha = np.random.dirichlet(10*np.ones(k),1)[0]
beta = np.random.dirichlet(np.ones(V),k)
phi = np.array([1/k*np.ones([N[m],k]) for m in range(M)])
gamma = np.tile(alpha,(M,1)) + np.tile(N/k,(k,1)).T
# -
ans_real_world = lda_package.variation_EM(M, k, text_, N, V_words, alpha, beta, gamma, phi, iteration = 200)
file=open('ans_real_world.txt','w')
file.write(str(ans_real_world));
file.close()
# # Visualization
fig, ax = plt.subplots(figsize=(18,10))
ax.imshow(np.log(1000*ans_real_world[1]), cmap=plt.cm.hot, interpolation='nearest', aspect='auto')
ax.set_xlabel("word")
plt.yticks([0, 1, 2, 3, 4,5,6,7,8,9],
["topic 1", "topic 2","topic 3", "topic 4","topic 5","topic 6","topic 7","topic 8","topic 9","topic 10"])
pass
beta_post = ans_real_world[1]
# +
f, ax= plt.subplots(k, 1, figsize=(15, 10), sharex=True)
for i, l in enumerate(range(k)):
ax[i].stem(beta_post[l,:], linefmt='b-',
markerfmt='bo', basefmt='w-')
ax[i].set_xlim(-10,5050)
ax[i].set_ylim(0, 0.06)
ax[i].set_ylabel("Prob")
ax[i].set_title("topic {}".format(l+1))
ax[9].set_xlabel("word")
plt.tight_layout()
plt.show()
pass
# plt.savefig('real_10.jpg')
# -
gamma_post = ans_real_world[2]/np.sum(ans_real_world[2],axis = 1).reshape((-1,1))
# +
f, ax= plt.subplots(5, 1, figsize=(8, 8), sharex=False)
for i, k in enumerate([1, 13, 29, 37, 51]):
ax[i].stem(gamma_post[k,:], linefmt='r-',
markerfmt='ro', basefmt='w-')
#ax[i].set_xlim(-1, 21)
ax[i].set_ylim(0, 0.8)
ax[i].set_ylabel("Prob")
ax[i].set_title("Document {}".format(k))
ax[4].set_xlabel("Topic")
plt.tight_layout()
# -
| Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
from notebook import Notebook
class Menu:
"Display a menu and respond to choices when run"
def __init__(self):
self.notebook = Notebook()
self.choices = {
"1": self.show_notes,
"2": self.search_notes,
"3": self.add_note,
"4": self.modify_note,
"5": self.quit,
}
def display_menu(self):
print("""
Notebook Menu
1. Show all Notes
2. Search Notes
3. Add Note
4. Modify Note
5. Quit
""")
def run(self):
"""Display the menu and repsond to choices."""
while True:
self.display_menu()
choice = input("Enter an option: ")
action = self.choices.get(choice)
if action:
action()
else:
print("{0} is not a valid choice".format(choice))
def show_notes(self, notes=None):
if not notes:
notes = self.notebook.notes
for note in notes:
print("{0}: {1}/n{2}".format(note.id, note.tags, note.memo))
def search_notes(self):
target = input("Search for: ")
notes = self.notebook.search(target)
self.show_notes(notes)
def add_note(self):
memo = input("Enter a memo: ")
self.notebook.new_note(memo)
print("Your note has been added.")
def modify_note(self):
note_id = input("Enter a note id: ")
memo = input("Enter a memo: ")
tags = input("Enter tags: ")
if memo:
self.notebook.modify_memo(note_id, memo)
if tags:
self.notebook.modify_tags(note_id, tags)
def quit(self):
print("Thank you for using your notebook today.")
sys.exit(0)
if __name__ == "__main__":
Menu().run()
# -
| notes/running_menu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
'''
- 進化計算アルゴリズム
- ベイズ最適化
-
'''
import pandas as pd
df = pd.read_csv('./result.csv', sep=',',header=0)
df.head(1)
# 有効(1-11が0)なものに縛る
df_eff =df[ (df['1'] == 0) &(df['2'] == 0) & (df['3'] == 0) &(df['4'] == 0) & (df['5'] == 0) & (df['6'] == 0) & (df['7'] == 0 )& (df['8'] == 0) & (df['9'] == 0) & (df['10'] == 0) & (df['11'] == 0 )]
df_eff = df_eff[["created_num","result"]]
df_eff
df_eff[(df_eff["result"] > 100)].head()
df_eff[(df_eff["result"] <50)]
| analyze.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="AkEzLyObST_n" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
# + id="f19KDN4ES3u6" colab_type="code" colab={}
df = pd.read_csv('/content/drive/My Drive/Colab Notebooks/dw_matrix/data/men_shoes.csv', low_memory=False)
# + id="uQtXV-qGTbQh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="fa8213b4-51cb-4a5c-ab04-7611dff1ae5c" executionInfo={"status": "ok", "timestamp": 1582566379895, "user_tz": -60, "elapsed": 417, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
df.shape
# + id="jk_HdNdgTo9Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 237} outputId="457105c2-1c44-4f33-b73c-b3008b22c1ca" executionInfo={"status": "ok", "timestamp": 1582566394830, "user_tz": -60, "elapsed": 612, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
df.columns
# + id="zCFI82diTsij" colab_type="code" colab={}
mean_price = np.mean(df['prices_amountmin'])
# + id="MjqoJJ88T9QH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d5847295-9821-4098-d41d-70a9e5b0d1b7" executionInfo={"status": "ok", "timestamp": 1582566470398, "user_tz": -60, "elapsed": 646, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
mean_price
# + id="96Y9AbhyT-_1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a93f88b9-8aab-4201-895e-78750aed0e28" executionInfo={"status": "ok", "timestamp": 1582566657052, "user_tz": -60, "elapsed": 548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
y_true = df['prices_amountmin']
y_pred = [mean_price] * y_true.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="ULhPVcZ6UoIs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="c49161e9-6b48-4b00-c70d-9f74fd3ae376" executionInfo={"status": "ok", "timestamp": 1582566827680, "user_tz": -60, "elapsed": 716, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
np.log1p(df['prices_amountmin']).hist(bins=100)
# + id="4AmJyev_VWMz" colab_type="code" colab={}
def funkcja(val):
x = df[val].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=100)
scores = cross_val_score(model, x,y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="bLu8d3ZoY40n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 237} outputId="b5077516-42a6-45dd-d8e8-55e2f8183407" executionInfo={"status": "ok", "timestamp": 1582567786781, "user_tz": -60, "elapsed": 610, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
df.columns
# + id="4k3ILzD0ZAYl" colab_type="code" colab={}
df['prc_fac'] = df['prices_amountmax'].factorize()[0]
# + id="fw81JXnpZIvD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b1332852-fba0-4401-898e-3e28aed4f6a3" executionInfo={"status": "ok", "timestamp": 1582568456182, "user_tz": -60, "elapsed": 709, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDrBA22ekzxZD4a-yzl9v4HTtS4SnHfgptkuqrvSA=s64", "userId": "08311161562109144008"}}
funkcja(['prc_fac', 'brad_fac'])
# + id="RpGPlIFEZN0-" colab_type="code" colab={}
df['brad_fac'] = df['brand'].factorize()[0]
# + id="85sQ6n9NZW9o" colab_type="code" colab={}
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
a = np.array([10, 20, 30])
print(a)
print(np.diag(a))
print(np.diag([100, 200, 300]))
print(np.diag(a, k=1))
print(np.diag(a, k=-2))
print(np.diag([1, 1, 1]))
print(np.identity(3))
print(np.identity(3, int))
| notebook/numpy_diag_construct.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Imports
import numpy as np, cv2
from matplotlib import pyplot as plt
# ### Reading items and the reasons why they are contradict
itm1 = 'potato'
itm2 = 'melon'
reason = "Ayurveda's food theory"
# ### Processing the data
img1 = cv2.imread('image/' + itm1 + '.jpg')
img2 = cv2.imread('image/' + itm2 + '.jpg')
img3 = cv2.imread('image/white.jpg')
width = len(reason) * 7
height = int(width * 0.7)
img1 = cv2.resize(img1,(width, height))
img2 = cv2.resize(img2,(width, height))
img3 = cv2.resize(img3,(2 * width, 50))
vis = np.concatenate((img1, img2), axis=1)
vis = np.concatenate((vis, img3), axis=0)
vis = cv2.line(vis, (0, 0), (2 * width, height), (0, 0, 255), 2)
vis = cv2.line(vis, (2 * width, 0), (0, height), (0, 0, 255), 2)
vis = cv2.putText(vis, reason, (10,height + 20), cv2.FONT_HERSHEY_SIMPLEX, 0.8, (255, 0, 0), 2, cv2.LINE_AA)
# ### Output image
cv2.imwrite('out.png', vis)
RGB_vis = cv2.cvtColor(vis, cv2.COLOR_BGR2RGB)
plt.imshow(RGB_vis)
plt.title("DON'T EAT TOGETHER!!")
plt.show()
| ShowImage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# %config InlineBackend.figure_format='svg'
learning_rate=.01
epochs=200
n_samples=30
train_x=np.linspace(0,20,n_samples)
train_y=3*train_x+4*np.random.rand(n_samples)
plt.plot(train_x,train_y,'o')
plt.plot(train_x,3*train_x)
plt.show()
# +
X=tf.placeholder(tf.float32)
Y=tf.placeholder(tf.float32)
W=tf.Variable(np.random.randn(), name='weights')
B=tf.Variable(np.random.randn(), name='bias')
# -
#pred=X*W+B
pred=tf.add(tf.multiply(X,W),B)
cost=tf.reduce_sum((pred-Y)**2/(2*n_samples))
optimizer=tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
init=tf.global_variables_initializer()
with tf.Session() as sesh:
sesh.run(init)
for epoch in range(epochs):
for x,y in zip(train_x,train_y):
sesh.run(optimizer,feed_dict={X:x,Y:y})
if not epoch%20:
c=sesh.run(cost,feed_dict={X:train_x,Y:train_y})
w=sesh.run(W)
b=sesh.run(B)
print(' e {} c {} w{} b {}'.format(epoch,c,w,b))
#print(f'epoch:{epoch:04d} c={c:.4f} w={w:.4f} b={b:.4f}'.format(epoch,c,w,b))
weight=sesh.run(W)
bias=sesh.run(B)
plt.plot(train_x,train_y,'o')
plt.plot(train_x,weight*train_x+bias)
plt.show()
| TF1_LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn import datasets
from sklearn.model_selection import train_test_split
from yellowbrick.classifier import ConfusionMatrix
from keras.models import Sequential
from keras.layers import Dense
from keras.utils import np_utils
import numpy as np
from sklearn.metrics import confusion_matrix
import warnings
warnings.filterwarnings("ignore")
# -
base = datasets.load_iris()
previsores = base.data
classe = base.target
base
classe_dummy = np_utils.to_categorical(classe)
classe_dummy.dtype
X_train, X_test, y_train, y_test = train_test_split(previsores,
classe_dummy,
test_size=0.3,
random_state=0)
# ### Criar Rede
# units (neuronios camada escondida)
# input_dim (atributos previsores)
# activation='softmax'
modelo = Sequential()
# Primeira camada de neuronios
modelo.add(Dense(units=5,
input_dim=4))
# Segunda camada de neuronios
modelo.add(Dense(units=4))
# Camada de saida units=3 pq tenho 3 classes
modelo.add(Dense(units=3,activation='softmax'))
modelo.summary()
# necessita ter instalado o graphviz
from keras.utils.vis_utils import plot_model
# ### Treinar rede
modelo.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics =['accuracy'])
modelo.fit(X_train,
y_train,
epochs = 1000,
validation_data=(X_test,y_test))
# +
# Gerar matriz dde confusoes
# -
previsoes = modelo.predict(X_test)
previsoes = (previsoes > 0.5)
y_test_matrix = [np.argmax(t) for t in y_test]
y_previsao_matrix = [np.argmax(t) for t in previsoes]
confusao = confusion_matrix(y_test_matrix,y_previsao_matrix)
print(confusao)
| notebook/Keras_Iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification with Naive Bayes and SVM
# ## Naive Bayes
# ### Preprocessing
# +
import pandas as pd
import nltk
import os
import re
import string
import numpy as np
from sklearn import datasets, svm
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
df_train = pd.DataFrame(columns=['words', 'sentiment'])
df_test = pd.DataFrame(columns=['words', 'sentiment'])
# +
sw_dir = './Data/6/sw.txt'
stop_words = []
with open(sw_dir) as f:
text = f.readlines()
for word in text:
stop_words.append(re.findall('\S+', word)[0])
# adding br and empty string to stop words
stop_words.append('br')
stop_words.append('')
# +
# create dataset from text files
train_pos_dir = './Data/6/aclImdb/train/pos'
for filename in os.listdir(train_pos_dir):
with open(os.path.join(train_pos_dir, filename)) as f:
text = f.readlines()[0]
df_train = df_train.append({'words': text, 'sentiment': 1},
ignore_index=True)
train_neg_dir = './Data/6/aclImdb/train/neg'
for filename in os.listdir(train_neg_dir):
with open(os.path.join(train_neg_dir, filename)) as f:
text = f.readlines()[0]
df_train = df_train.append({'words': text, 'sentiment': 0},
ignore_index=True)
test_pos_dir = './Data/6/aclImdb/test/pos'
for filename in os.listdir(test_pos_dir):
with open(os.path.join(test_pos_dir, filename)) as f:
text = f.readlines()[0]
df_test = df_test.append({'words': text, 'sentiment': 1},
ignore_index=True)
test_neg_dir = './Data/6/aclImdb/test/neg'
for filename in os.listdir(test_neg_dir):
with open(os.path.join(test_neg_dir, filename)) as f:
text = f.readlines()[0]
df_test = df_test.append({'words': text, 'sentiment': 0},
ignore_index=True)
# +
def remove_punct(text):
def change(ch):
if ch in string.punctuation or ch.isdigit():
return " "
else:
return ch
no_punct = "".join([change(ch) for ch in text])
return no_punct
# df_train['words'] = df_train['words'].apply(lambda x: remove_punct(x))
# +
def tokenize(text):
tokens = re.split('\W+', text)
return tokens
# df_train['words'] = df_train['words'].apply(lambda x: tokenize(x.lower()))
# +
def remove_sw(tokens):
text = [w for w in tokens if w not in stop_words]
return text
# df_train['words'] = df_train['words'].apply(lambda x: remove_sw(x))
# +
def remove_short(tokens):
text = [w for w in tokens if len(w)>2]
return text
# df_train['words'] = df_train['words'].apply(lambda x: remove_short(x))
# -
# There are several ways to get root of tokens like stemming and lemmatizing. stemming is faster and lemmatizing is more precise.
# +
wn = nltk.WordNetLemmatizer()
def lemmatizing(tokens):
text = [wn.lemmatize(w) for w in tokens]
return text
ps = nltk.stem.PorterStemmer()
def stemming(tokens):
text = [ps.stem(w) for w in tokens]
return text
# df_train['words'] = df_train['words'].apply(lambda x: lemmatizing(x))
# df_train['words'] = df_train['words'].apply(lambda x: stemming(x))
# +
def clean_text(text):
text = remove_punct(text)
text = tokenize(text)
text = remove_sw(text)
text = remove_short(text)
# text = lemmatizing(text)
text = stemming(text)
return text
count_vect = CountVectorizer(analyzer=clean_text,
lowercase=True,
binary=True)
X_train = count_vect.fit_transform(df_train['words'])
y_train = df_train['sentiment'].to_numpy(dtype='int')
X_test = count_vect.transform(df_test['words'])
y_test = df_train['sentiment'].to_numpy(dtype='int')
# print(count_vect.get_feature_names())
# -
# ### Classification
clf = MultinomialNB(alpha=100)
clf = clf.fit(X_train, y_train)
# +
# clf.score(X_test, y_test)
y_pred = clf.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
acc_tr = clf.score(X_train, y_train)
acc_te = clf.score(X_test, y_test)
print("train accuracy: {}%".format(acc_tr*100))
print("test accuracy: {}%".format(acc_te*100))
# -
sns.heatmap(cm, annot=True, fmt='d')
# ### Laplace smoothing
# +
alphas = [10**x for x in range(-4, 4)]
accs_tr = []
accs_te = []
for alpha in alphas:
cls = MultinomialNB(alpha=alpha)
cls = cls.fit(X_train, y_train)
accs_tr.append(cls.score(X_train, y_train))
accs_te.append(cls.score(X_test, y_test))
plt.plot(alphas, accs_tr)
plt.plot(alphas, accs_te)
plt.xlabel('alpha')
plt.ylabel('accuracy')
plt.legend(['train accuracy', 'test accuracy'])
plt.xscale('log')
# -
# ## SVM Classifier
def make_meshgrid(x, y, h=.02):
x_min, x_max = x.min() - 1, x.max() + 1
y_min, y_max = y.min() - 1, y.max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
return xx, yy
def plot_contours(ax, clf, xx, yy, **params):
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
out = ax.contourf(xx, yy, Z, **params)
return out
# +
iris = datasets.load_iris()
X = iris.data[:, :2]
y = iris.target
# -
models = (svm.SVC(kernel='linear', C=1.0),
svm.LinearSVC(C=1.0, max_iter=10000),
svm.SVC(kernel='rbf', gamma=0.7, C=1.0),
svm.SVC(kernel='poly', degree=3, gamma='auto', C=1.0))
models = (clf.fit(X, y) for clf in models)
# +
titles = ('SVC with linear kernel',
'LinearSVC (linear kernel)',
'SVC with RBF kernel',
'SVC with polynomial (degree 3) kernel')
fig, sub = plt.subplots(2, 2, figsize=(12, 8))
plt.subplots_adjust(wspace=0.4, hspace=0.4)
X0, X1 = X[:, 0], X[:, 1]
xx, yy = make_meshgrid(X0, X1)
for clf, title, ax in zip(models, titles, sub.flatten()):
plot_contours(ax, clf, xx, yy,
cmap=plt.cm.coolwarm, alpha=0.8)
ax.scatter(X0, X1, c=y, cmap=plt.cm.coolwarm, s=20, edgecolors='k')
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xlabel('Sepal length')
ax.set_ylabel('Sepal width')
ax.set_xticks(())
ax.set_yticks(())
ax.set_title(title)
plt.show()
| TA/Session6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # G2Product Guide - Version
#
# The `version()` method creates a JSON string with information about your Senzing version.
# ## G2Product
#
# The G2Product API...
from G2Product import G2Product
# ### Initialization
#
# To start using Senzing G2Product, create and initialize an instance.
# This should be done once per process.
# The `initV2()` method accepts the following parameters:
#
# - **module_name:** A short name given to this instance of the G2Product object.
# - **config_json:** A JSON string containing configuration parameters.
# - **verbose_logging:** A boolean which enables diagnostic logging.
#
# Calling this function will return "0" upon success.
g2_product = G2Product()
try:
g2_product.initV2(
module_name,
senzing_config_json,
verbose_logging)
except G2Exception.G2ModuleGenericException as err:
print(g2_product.getLastException())
# ### version()
#
# Call G2Product's `version()` method and pretty-print results.
try:
version_string = g2_product.version()
print(version_string)
except G2Exception.G2ModuleGenericException as err:
print(g2_product.getLastException)
| notebooks/senzing-examples/Windows/senzing-G2Product-version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="-PLC9SvcQgkG"
# # DeepDreaming with TensorFlow
#
# ### ___<NAME>___
#
# This notebook produces DeepDream images from user-supplied photos using Google's pretrained Inception neural network. It can be used as a starting point for further exploration in visualizing convolutional neural networks.
# + [markdown] id="ILvNKvMvc2n5"
# ### 1) Load the model graph
#
# The pretrained Inception network can be downloaded [here](https://storage.googleapis.com/download.tensorflow.org/models/inception5h.zip). This next cell downloads the file automatically and unpacks it locally to the Colab kernel. We can the load the contained model file 'tensorflow_inception_graph.pb' in the cell below.
# + id="1kJuJRLiQgkg" cellView="both" outputId="d9aaef3a-d4e6-4046-8d7b-0e6b1b10ea77" colab={"base_uri": "https://localhost:8080/", "height": 564}
# !wget -nc --no-check-certificate https://storage.googleapis.com/download.tensorflow.org/models/inception5h.zip && unzip -n inception5h.zip
# !wget -nc https://storage.googleapis.com/download.tensorflow.org/example_images/YellowLabradorLooking_new.jpg
file_contents = open("YellowLabradorLooking_new.jpg").read()
from io import BytesIO
from IPython.display import clear_output, Image, display
import numpy as np
import PIL.Image
import tensorflow as tf
from __future__ import print_function
model_fn = 'tensorflow_inception_graph.pb'
# creating TensorFlow session and loading the model
graph = tf.Graph()
sess = tf.InteractiveSession(graph=graph)
with tf.gfile.FastGFile(model_fn, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
t_input = tf.placeholder(np.float32, name='input') # define the input tensor
imagenet_mean = 117.0
t_preprocessed = tf.expand_dims(t_input-imagenet_mean, 0)
tf.import_graph_def(graph_def, {'input':t_preprocessed})
def T(layer):
'''Helper for getting layer output tensor'''
return graph.get_tensor_by_name("import/%s:0"%layer)
# + [markdown] id="_A6rIrUwAVit"
# ### Optional: Upload an image from your computer
#
# # Skip these steps if you just want to run this example
# + id="J5pnVLSIYQbI" outputId="0b6278bd-c387-4e4c-b9a2-a9d9ac9780e5" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 35}
from google.colab import files
uploaded = files.upload()
# + id="9xXGMG3MHih-"
if type(uploaded) is not dict: uploaded = uploaded.files ## Deal with filedit versions
file_contents = uploaded[uploaded.keys()[0]]
# + [markdown] id="Gr2O56zvLc4v"
#
# + [markdown] id="FTA0_5cjLjKR"
# ### 2) Load the starting image
# + id="M9_vOh_2Qgl-" cellView="both" outputId="90571b18-c93d-4186-c215-5ad70564760d" colab={"base_uri": "https://localhost:8080/", "height": 317}
def showarray(a, fmt='jpeg'):
a = np.uint8(np.clip(a, 0, 255))
f = BytesIO()
PIL.Image.fromarray(a).save(f, fmt)
display(Image(data=f.getvalue()))
img0 = sess.run(tf.image.decode_image(file_contents))
showarray(img0)
# + [markdown] id="yfy9Hx1ogTJk"
#
# + [markdown] id="WAfjHMufgXRg"
# def showarray(a, fmt='jpeg'): a = np.uint8(np.clip(a, 0, 255)) f = BytesIO() PIL.Image.fromarray(a).save(f, fmt) display(Image(data=f.getvalue()))img0 = sess.run(tf.image.decode_image(file_contents))showarray(img0)
# + [markdown] id="gw5mCosZf2LS"
# # New Section
#
# + [markdown] id="oxELTBqpqOP5"
# ### 4) The core deepdream code
# + id="K8Usk3m5Q1E0"
# These parameters let us control the strenth of the deepdream.
octave_n = 4
octave_scale = 1.4
iter_n = 10
strength = 200
# Helper function that uses TensorFlow to resize an image
def resize(img, new_size):
return sess.run(tf.image.resize_bilinear(img[np.newaxis,:], new_size))[0]
# Apply gradients to an image in a seires of tiles
def calc_grad_tiled(img, t_grad, tile_size=256):
'''Random shifts are applied to the image to blur tile boundaries over
multiple iterations.'''
h, w = img.shape[:2]
sx, sy = np.random.randint(tile_size, size=2)
# We randomly roll the image in x and y to avoid seams between tiles.
img_shift = np.roll(np.roll(img, sx, 1), sy, 0)
grad = np.zeros_like(img)
for y in range(0, max(h-tile_size//2, tile_size),tile_size):
for x in range(0, max(w-tile_size//2, tile_size),tile_size):
sub = img_shift[y:y+tile_size,x:x+tile_size]
g = sess.run(t_grad, {t_input:sub})
grad[y:y+tile_size,x:x+tile_size] = g
imggrad = np.roll(np.roll(grad, -sx, 1), -sy, 0)
# Add the image gradient to the image and return the result
return img + imggrad*(strength * 0.01 / (np.abs(imggrad).mean()+1e-7))
# Applies deepdream at multiple scales
def render_deepdream(t_obj, input_img, show_steps = True):
# Collapse the optimization objective to a single number (the loss)
t_score = tf.reduce_mean(t_obj)
# We need the gradient of the image with respect to the objective
t_grad = tf.gradients(t_score, t_input)[0]
# split the image into a number of octaves (laplacian pyramid)
img = input_img
octaves = []
for i in range(octave_n-1):
lo = resize(img, np.int32(np.float32(img.shape[:2])/octave_scale))
octaves.append(img-resize(lo, img.shape[:2]))
img = lo
# generate details octave by octave
for octave in range(octave_n):
if octave>0:
hi = octaves[-octave]
img = resize(img, hi.shape[:2])+hi
for i in range(iter_n):
img = calc_grad_tiled(img, t_grad)
if show_steps:
clear_output()
showarray(img)
return img
# + [markdown] id="mc3Ea6SrRzIB"
# ### 4) Let's deep dream !
#
# You can adjust the sliders to change the strength of the deep dream, and how many scales it is applied over.
# + id="T9ZA24auPnVt" outputId="079122e6-f249-4876-d6f2-805e7e9152e6" colab={"base_uri": "https://localhost:8080/", "height": 570}
octave_n = 4 #@param {type:"slider", max: 10}
octave_scale = 1.4 #@param {type:"number"}
iter_n = 10 #@param {type:"slider", max: 50}
strength = 200 #@param {type:"slider", max: 1000}
layer = "mixed4c" #@param ["mixed3a", "mixed3b", "mixed4a", "mixed4c", "mixed5a"]
final = render_deepdream(tf.square(T(layer)), img0)
# + [markdown] id="IJzvhEFxpB7E"
# ### 5) Individual neurons
#
# We can also try and optimize not against an entire layer but just one neuron's activity:
# + id="4GexZuwJdDmu" cellView="both" outputId="602abff6-2331-418d-c65e-9d552dfbe027" colab={"base_uri": "https://localhost:8080/", "height": 570}
feature_channel = 139 #@param {type:"slider", max: 512}
layer = "mixed4d_3x3_bottleneck_pre_relu" #@param ["mixed4d_3x3_bottleneck_pre_relu", "mixed3a", "mixed3b", "mixed4a", "mixed4c", "mixed5a"]
if feature_channel >= T(layer).shape[3]:
print("Feature channel exceeds size of layer ", layer, " feature space. ")
print("Choose a smaller channel number.")
else:
render_deepdream(T(layer)[:,:,:,feature_channel], img0)
# + [markdown] id="jQL7w_LNz1rJ"
# ### 6) Zooming iterative DeepDream
#
# We can enter completely immersive worlds by iteratively sooming into the picture:
# + id="j3c2e0uO0CyP" outputId="a8f5aa61-abcc-4fae-9e48-a7df2c9593f9" colab={"base_uri": "https://localhost:8080/", "height": 570}
layer = "mixed4d_3x3_bottleneck_pre_relu" #@param ["mixed4d_3x3_bottleneck_pre_relu", "mixed3a", "mixed3b", "mixed4a", "mixed4c", "mixed5a"]
iter_n = 5 #@param {type:"slider", max: 50}
strength = 150 #@param {type:"slider", max: 1000}
zooming_steps = 20 #@param {type:"slider", max: 512}
zoom_factor = 1.1 #@param {type:"number"}
frame = img0
img_y, img_x, _ = img0.shape
for i in range(zooming_steps):
frame = render_deepdream(tf.square(T(layer)), frame, False)
clear_output()
showarray(frame)
newsize = np.int32(np.float32(frame.shape[:2])*zoom_factor)
frame = resize(frame, newsize)
frame = frame[(newsize[0]-img_y)//2:(newsize[0]-img_y)//2+img_y,
(newsize[1]-img_x)//2:(newsize[1]-img_x)//2+img_x,:]
# + [markdown] id="2AzLO_JiS6gi"
# ### Further reading for the curious
#
# * Original [DeepDream (Inceptionism) blog post](https://research.googleblog.com/2015/06/inceptionism-going-deeper-into-neural.html)
# * [Original DeepDream algorithm](https://github.com/google/deepdream/blob/master/dream.ipynb) with Caffe
# + [markdown] id="K_Ab2CkOtPOq"
# ## 7) Diving deeper into the Inception Model
#
# Lets look a bit deeper into the Inception Model and visualize the layers. Each layer will produce a very different result when used in deep dreaming.
# + id="FbxQBJpzvBO7" outputId="ef6580e4-01b3-45ff-ab34-686b918aee5c" colab={"base_uri": "https://localhost:8080/", "height": 1054}
layers = [op.name for op in graph.get_operations() if op.type=='Conv2D' and 'import/' in op.name]
feature_nums = [int(graph.get_tensor_by_name(name+':0').get_shape()[-1]) for name in layers]
print('Number of layers', len(layers))
print('Total number of feature channels:', sum(feature_nums))
for layer in layers:
print('Layer:', layer)
# + [markdown] id="SGgDuDo6xZnA"
# For example try deepdreaming with the layer '`mixed4a_3x3_pre_relu`'
# + id="pE2QvGorxU5H" outputId="61842910-1187-484d-8e75-9aa4e9700481" colab={"base_uri": "https://localhost:8080/", "height": 570}
layer = "mixed4a_3x3_pre_relu"
final = render_deepdream(tf.square(T(layer)), img0)
# + [markdown] id="1uOiMyvpx_KC"
# We can also use TensorBoard to visualize the full graph to understand bettter how these different layers relate to each other. Most of the code in the next section just makes the graph look a little bit cleaner.
# + id="KA5YzXcEtOi6" outputId="81e319c2-1474-41b5-a6e9-ddc3e96ccf02" colab={"base_uri": "https://localhost:8080/", "height": 661}
# Helper functions for TF Graph visualization
from IPython.display import HTML
def strip_consts(graph_def, max_const_size=32):
"""Strip large constant values from graph_def."""
strip_def = tf.GraphDef()
for n0 in graph_def.node:
n = strip_def.node.add()
n.MergeFrom(n0)
if n.op == 'Const':
tensor = n.attr['value'].tensor
size = len(tensor.tensor_content)
if size > max_const_size:
tensor.tensor_content = tf.compat.as_bytes("<stripped %d bytes>"%size)
return strip_def
def rename_nodes(graph_def, rename_func):
res_def = tf.GraphDef()
for n0 in graph_def.node:
n = res_def.node.add()
n.MergeFrom(n0)
n.name = rename_func(n.name)
for i, s in enumerate(n.input):
n.input[i] = rename_func(s) if s[0]!='^' else '^'+rename_func(s[1:])
return res_def
def show_graph(graph_def, max_const_size=32):
"""Visualize TensorFlow graph."""
if hasattr(graph_def, 'as_graph_def'):
graph_def = graph_def.as_graph_def()
strip_def = strip_consts(graph_def, max_const_size=max_const_size)
code = """
<script>
function load() {{
document.getElementById("{id}").pbtxt = {data};
}}
</script>
<link rel="import" href="https://tensorboard.appspot.com/tf-graph-basic.build.html" onload=load()>
<div style="height:600px">
<tf-graph-basic id="{id}"></tf-graph-basic>
</div>
""".format(data=repr(str(strip_def)), id='graph'+str(np.random.rand()))
iframe = """
<iframe seamless style="width:800px;height:620px;border:0" srcdoc="{}"></iframe>
""".format(code.replace('"', '"'))
display(HTML(iframe))
# Visualizing the network graph. Be sure expand the "mixed" nodes to see their
# internal structure. We are going to visualize "Conv2D" nodes.
tmp_def = rename_nodes(graph_def, lambda s:"/".join(s.split('_',1)))
show_graph(tmp_def)
| deepdream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cltl/python-for-text-analysis/blob/colab/Assignments-colab/ASSIGNMENT_3b.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="OpcaZLBYNzmm"
# %%capture
# !wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Data.zip
# !wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/images.zip
# !wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Extra_Material.zip
# !unzip Data.zip -d ../
# !unzip images.zip -d ./
# !unzip Extra_Material.zip -d ../
# !rm Data.zip
# !rm Extra_Material.zip
# !rm images.zip
# + [markdown] id="g9arCT4eNzLN"
# # Assignment 3b: Writing your own nlp program
# + [markdown] id="_q9-U6uxNzLR"
# ## Due: Friday the 1st of October 2021 14:30.
#
# * Please submit your assignment (notebooks of parts 3a and 3b + additional files) as **a single .zip file** using Canvas (Assignments --> Assignment 3)
#
# * Please name your zip file with the following naming convention: ASSIGNMENT_3_FIRSTNAME_LASTNAME.zip
#
# **IMPORTANTE NOTE**:
# * The students who follow the Bachelor version of this course, i.e., the course Introduction to Python for Humanities and Social Sciences (L_AABAALG075) as part of the minor Digital Humanities, do **not have to do Exercises 3 and 4 of Assignment 3b**
# * The other students, i.e., who follow the Master version of course, which is Programming in Python for Text Analysis (L_AAMPLIN021), are required to **do Exercises 3 and 4 of Assignment 3b**
#
# If you have **questions** about this topic, please contact us **(<EMAIL>)**. Questions and answers will be collected on Piazza, so please check if your question has already been answered first.
#
# In this part of the assignment, we will carry out our own little text analysis project. The goal is to gain some insights into longer texts without having to read them all in detail.
#
# This part of the assignment builds on some notions that have been revised in part A of the assignment. Please feel free to go back to part A and reuse your code whenever possible.
#
# The goals of this part are:
#
# * divide a problem into smaller sub-problems and test code using small examples
# * doing text analysis and writing results to a file
# * combining small functions into bigger functions
#
# **Tip**: The assignment is split into four steps, which are divided into smaller steps. Instead of doing everything step by step, we highly recommend you read all sub-steps of a step first and then start coding. In many cases, the sub-steps are there to help you split the problem into manageable sub-problems, but it is still good to keep the overall goal in mind.
# + [markdown] id="nKtpEcDSNzLR"
# ## Preparation: Data collection
# + [markdown] id="0rZ8_TgANzLS"
# In the directory `../Data/books/`, you should find three .txt files. If not, you can use the following cell to download them. Also, feel free to look at the code to learn how to download .txt files from the web.
#
# We defined a function called `download_book` which downloads a book in .txt format. Then we define a dictionary with names and urls. We loop through the dictionary, download each book and write it to a file stored in the directory `books` in the current working directory. You don't need to do anything - just run the cell and the files will be downloaded to your computer.
# + id="3ty6_dYrNzLS"
# Downloading data - you get this for free :-)
import requests
import os
def download_book(url):
"""
Download book given a url to a book in .txt format and return it as a string.
"""
text_request = requests.get(url)
text = text_request.text
return text
book_urls = dict()
book_urls['HuckFinn'] = 'http://www.gutenberg.org/cache/epub/7100/pg7100.txt'
book_urls['Macbeth'] = 'http://www.gutenberg.org/cache/epub/1533/pg1533.txt'
book_urls['AnnaKarenina'] = 'http://www.gutenberg.org/files/1399/1399-0.txt'
if not os.path.isdir('../Data/books/'):
os.mkdir('../Data/books/')
for name, url in book_urls.items():
text = download_book(url)
with open('../Data/books/'+name+'.txt', 'w', encoding='utf-8') as outfile:
outfile.write(text)
# + [markdown] id="_9ZlucALNzLU"
# **Encoding issues with txt files**
#
# For Windows users, the file “AnnaKarenina.txt” gets the encoding cp1252.
# In order to open the file, you have to add **encoding='utf-8'**, i.e.,
#
# ```python
# a_path = 'some path on your computer.txt'
# with open(a_path, mode='r', encoding='utf-8'):
# # process file
# ```
# + [markdown] id="SbWM1JApNzLV"
# ## Exercise 1
# Was the download successful? Let's start writing code! Please create the following two Python modules:
# + [markdown] id="lf1abi5INzLV"
# * Python module **analyze.py** This module you will call from the command line
# * Python module **utils.py** This module will contain your helper functions
# + [markdown] id="UkwO46DvNzLW"
# More precisely, please create two files, **analyze.py** and **utils.py**, which are both placed in the same directory as this notebook. The two files are empty at this stage of the assignment.
# + [markdown] id="mgaXHbVLNzLW"
# 1.a) Write a function called `get_paths` and store it in the Python module **utils.py**
#
# The function `get_paths`:
# * takes one positional parameter called *input_folder*
# * the function stores all paths to .txt files in the *input_folder* in a list
# * the function returns a list of strings, i.e., each string is a file path
#
# Once you've created the function and stored it in **utils.py**:
# * Import the function into **analyze.py**, using `from utils import get_paths`
# * Call the function inside **analyze.py** (input_folder="../Data/books")
# * Assign the output of the function to a variable and print this variable.
# * call **analyze.py** from the command line to test it
# + [markdown] id="sa9JLcdTNzLW"
# ## Exercise 2
# 2.a) Let's get a little bit of an overview of what we can find in each text. Write a function called `get_basic_stats`.
#
# The function `get_basic_stats`:
# * has one positional parameter called **txt_path** which is the path to a txt file
# * reads the content of the txt file into a string
# * Computes the following statistics:
# * The number of sentences
# * The number of tokens
# * The size of the vocabulary used (i.e. unique tokens)
# * the number of chapters/acts:
# * count occurrences of 'CHAPTER' in **HuckFinn.txt**
# * count occurrences of 'Chapter ' (with the space) in **AnnaKarenina.txt**
# * count occurrences of 'ACT' in **Macbeth.txt**
# * return a dictionary with four key:value pairs, one for each statistic described above:
# * num_sents
# * num_tokens
# * vocab_size
# * num_chapters_or_acts
#
# In order to compute the statistics, you need to perform sentence splitting and tokenization. Here is an example snippet.
# + colab={"base_uri": "https://localhost:8080/"} id="AILLwUxlORGs" outputId="ea986991-5d98-4927-fc40-92f3f8a1f1bf"
import nltk
nltk.download('punkt')
# + colab={"base_uri": "https://localhost:8080/"} id="Q1EGBEUqNzLX" outputId="fbe31d7d-e113-43ec-9be4-e5723e639b40"
from nltk.tokenize import sent_tokenize, word_tokenize
text = 'Python is a programming language. It was created by <NAME>.'
for sent in sent_tokenize(text):
print('SENTENCE', sent)
tokens = word_tokenize(sent)
print('TOKENS', tokens)
# + [markdown] id="y24IMAc9NzLY"
# 2.b) Store the function in the Python module **utils.py**. Import it in **analyze.py**.
# Edit **analyze.py** so that:
# * you first call the function **get_paths**
# * create an empty dictionary called **book2stats**, i.e., `book2stats = {}`
# * Loop over the list of txt files (the output from **get_paths**) and call the function **get_basic_stats** on each file
# * print the output of calling the function **get_basic_stats** on each file.
# * update the dictionary **book2stats** with each iteration of the for loop.
# + [markdown] id="VteAgollNzLY"
# Tip: **book2stats** is a dictionary mapping a book name (the key), e.g., ‘AnnaKarenina’, to a dictionary (the value) (the output from get_basic_stats)
#
# Tip: please use the following code snippet to obtain the basename name of a file path:
# + colab={"base_uri": "https://localhost:8080/"} id="5MgO2g9NNzLY" outputId="fea11aeb-f325-49d5-a876-54e16e636315"
import os
basename = os.path.basename('../Data/books/HuckFinn.txt')
book = basename.strip('.txt')
print(book)
# + [markdown] id="jERV_TVENzLZ"
# ### Please note that Exercises 3 and 4, respectively, are designed to be difficult. You will have to combine what you have learnt so far to complete these exercises.
# + [markdown] id="1p8XFNXtNzLZ"
# ## Exercise 3
#
#
# Let's compare the books based on the statistics. Create a dictionary `stats2book_with_highest_value` in **analyze.py** with four keys:
# * num_sents
# * num_tokens
# * vocab_size
# * num_chapters_or_acts
#
# The values are not the frequencies, but the **book** that has the highest value for the statistic. Make use of the **book2stats** dictionary to accomplish this.
# + [markdown] id="cdSB2uJeNzLZ"
# ## Exercise 4
#
# 4.a) The statistics above already provide some insights, but we want to know a bit more about what the books are about. To do this, we want to get the 30 most frequent tokens of each book. Edit the function `get_basic_stats` to add one more key:value pair:
# * the key is **top_30_tokens**
# * the value is a list of the 30 most frequent words in the text.
#
# 4.b) Write the top 30 tokens (one on each line) for each file to disk using the naming `top_30_[FILENAME]`:
# * top_30_AnnaKarenina.txt
# * top_30_HuckFinn.txt
# * top_30_Macbeth.txt
#
# Example of file (*the* and *and* may not be the most frequent tokens, these are just examples):
#
# ```
# the
# and
# ..
# ```
# The following code snippet can help you with obtaining the top 30 occurring tokens. The goal is to call the function you updated in Exercise 4a, i.e., get_basic_stats, in the file analyze.py. This also makes it possible to write the top 30 tokens to files.
# + colab={"base_uri": "https://localhost:8080/"} id="FgqjAcVpNzLa" outputId="88aab80c-884a-4fc4-9f77-409578183dea"
import operator
token2freq = {'a': 1000, 'the': 100, 'cow' : 5}
for token, freq in sorted(token2freq.items(),
key=operator.itemgetter(1),
reverse=True):
print(token, freq)
# + [markdown] id="kzXThUZfNzLa"
# Congratulations! You have written your first little nlp program!
# + id="ee2VAdGqNzLa"
| Assignments-colab/ASSIGNMENT_3b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
sns.set(color_codes=True)
weather=pd.read_csv('Test.csv')
weather.head()
weather.info()
sns.barplot(weather['humidity'], weather['temperature'])
sns.distplot(weather['humidity'])
sns.distplot(weather['humidity'], kde=False, rug=True);
sns.jointplot(weather['humidity'], weather['temperature'])
sns.jointplot(weather['humidity'], weather['temperature'], kind='hex')
sns.jointplot(weather['humidity'], weather['temperature'], kind='kde')
sns.pairplot(weather[['humidity', 'temperature', 'air_pollution_index' ]])
sns.stripplot(weather['weather_type'], weather['temperature'])
sns.stripplot(weather['weather_type'], weather['temperature'], jitter=True)
sns.swarmplot(weather['humidity'], weather['temperature'])
sns.boxplot(weather['humidity'], weather['temperature'], hue=weather['weather_type'])
sns.barplot(weather['humidity'], weather['temperature'], hue=weather['weather_type'])
sns.countplot(weather['weather_type'])
sns.pointplot(weather['humidity'], weather['temperature'], hue=weather['weather_type'])
sns.lmplot(x='humidity', y='temperature', data=weather)
sns.lmplot(x="humidity", y="temperature", hue="weather_type", data=weather)
| WeatherDataVisualize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">Introduction to SimpleITKv4 Registration</h1>
#
#
# <table width="100%">
# <tr style="background-color: red;"><td><font color="white">SimpleITK conventions:</font></td></tr>
# <tr><td>
# <ul>
# <li>Dimensionality and pixel type of registered images is required to be the same (2D/2D or 3D/3D).</li>
# <li>Supported pixel types are sitkFloat32 and sitkFloat64 (use the SimpleITK <a href="http://www.itk.org/SimpleITKDoxygen/html/namespaceitk_1_1simple.html#af8c9d7cc96a299a05890e9c3db911885">Cast()</a> function if your image's pixel type is something else).
# </ul>
# </td></tr>
# </table>
#
#
# ## Registration Components
#
# <img src="ITKv4RegistrationComponentsDiagram.svg" style="width:700px"/><br><br>
#
# There are many options for creating an instance of the registration framework, all of which are configured in SimpleITK via methods of the <a href="http://www.itk.org/SimpleITKDoxygen/html/classitk_1_1simple_1_1ImageRegistrationMethod.html">ImageRegistrationMethod</a> class. This class encapsulates many of the components available in ITK for constructing a registration instance.
#
# Currently, the available choices from the following groups of ITK components are:
#
# ### Optimizers
#
# The SimpleITK registration framework supports several optimizer types via the SetOptimizerAsX() methods, these include:
#
# <ul>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1ExhaustiveOptimizerv4.html">Exhaustive</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1AmoebaOptimizerv4.html">Nelder-Mead downhill simplex</a>, a.k.a. Amoeba.
# </li>
# <li>
# <a href="https://itk.org/Doxygen/html/classitk_1_1PowellOptimizerv4.html">Powell optimizer</a>.
# </li>
# <li>
# <a href="https://itk.org/Doxygen/html/classitk_1_1OnePlusOneEvolutionaryOptimizerv4.html">1+1 evolutionary optimizer</a>.
# </li>
# <li>
# Variations on gradient descent:
# <ul>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1GradientDescentOptimizerv4Template.html">GradientDescent</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1GradientDescentLineSearchOptimizerv4Template.html">GradientDescentLineSearch</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1RegularStepGradientDescentOptimizerv4.html">RegularStepGradientDescent</a>
# </li>
# </ul>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1ConjugateGradientLineSearchOptimizerv4Template.html">ConjugateGradientLineSearch</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1LBFGSBOptimizerv4.html">L-BFGS-B</a> (Limited memory Broyden, Fletcher,Goldfarb,Shannon-Bound Constrained) - supports the use of simple constraints ($l\leq x \leq u$)
# </li>
# <li>
# <a href="https://itk.org/Insight/Doxygen/html/classitk_1_1LBFGS2Optimizerv4.html">L-BFGS2</a> (Limited memory Broyden, Fletcher,Goldfarb,Shannon)
# </li>
# </ul>
#
#
# ### Similarity metrics
#
# The SimpleITK registration framework supports several metric types via the SetMetricAsX() methods, these include:
#
# <ul>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1MeanSquaresImageToImageMetricv4.html">MeanSquares</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1DemonsImageToImageMetricv4.html">Demons</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1CorrelationImageToImageMetricv4.html">Correlation</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1ANTSNeighborhoodCorrelationImageToImageMetricv4.html">ANTSNeighborhoodCorrelation</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1JointHistogramMutualInformationImageToImageMetricv4.html">JointHistogramMutualInformation</a>
# </li>
# <li>
# <a href="http://www.itk.org/Doxygen/html/classitk_1_1MattesMutualInformationImageToImageMetricv4.html">MattesMutualInformation</a>
# </li>
# </ul>
#
#
# ### Interpolators
#
# The SimpleITK registration framework supports several interpolators via the SetInterpolator() method, which receives one of
# the <a href="http://www.itk.org/SimpleITKDoxygen/html/namespaceitk_1_1simple.html#a7cb1ef8bd02c669c02ea2f9f5aa374e5">following enumerations</a>:
# <ul>
# <li> sitkNearestNeighbor </li>
# <li> sitkLinear </li>
# <li> sitkBSpline </li>
# <li> sitkGaussian </li>
# <li> sitkHammingWindowedSinc </li>
# <li> sitkCosineWindowedSinc </li>
# <li> sitkWelchWindowedSinc </li>
# <li> sitkLanczosWindowedSinc </li>
# <li> sitkBlackmanWindowedSinc </li>
# </ul>
#
# ## Data - Retrospective Image Registration Evaluation
#
# We will be using part of the training data from the Retrospective Image Registration Evaluation (<a href="http://www.insight-journal.org/rire/">RIRE</a>) project.
# +
import SimpleITK as sitk
# Utility method that either downloads data from the Girder repository or
# if already downloaded returns the file name for reading from disk (cached data).
# %run update_path_to_download_script
from downloaddata import fetch_data as fdata
# Always write output to a separate directory, we don't want to pollute the source directory.
import os
OUTPUT_DIR = 'Output'
# -
# ## Utility functions
# A number of utility callback functions for image display and for plotting the similarity metric during registration.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from ipywidgets import interact, fixed
from IPython.display import clear_output
# Callback invoked by the interact IPython method for scrolling through the image stacks of
# the two images (moving and fixed).
def display_images(fixed_image_z, moving_image_z, fixed_npa, moving_npa):
# Create a figure with two subplots and the specified size.
plt.subplots(1,2,figsize=(10,8))
# Draw the fixed image in the first subplot.
plt.subplot(1,2,1)
plt.imshow(fixed_npa[fixed_image_z,:,:],cmap=plt.cm.Greys_r);
plt.title('fixed image')
plt.axis('off')
# Draw the moving image in the second subplot.
plt.subplot(1,2,2)
plt.imshow(moving_npa[moving_image_z,:,:],cmap=plt.cm.Greys_r);
plt.title('moving image')
plt.axis('off')
plt.show()
# Callback invoked by the IPython interact method for scrolling and modifying the alpha blending
# of an image stack of two images that occupy the same physical space.
def display_images_with_alpha(image_z, alpha, fixed, moving):
img = (1.0 - alpha)*fixed[:,:,image_z] + alpha*moving[:,:,image_z]
plt.imshow(sitk.GetArrayViewFromImage(img),cmap=plt.cm.Greys_r);
plt.axis('off')
plt.show()
# Callback invoked when the StartEvent happens, sets up our new data.
def start_plot():
global metric_values, multires_iterations
metric_values = []
multires_iterations = []
# Callback invoked when the EndEvent happens, do cleanup of data and figure.
def end_plot():
global metric_values, multires_iterations
del metric_values
del multires_iterations
# Close figure, we don't want to get a duplicate of the plot latter on.
plt.close()
# Callback invoked when the IterationEvent happens, update our data and display new figure.
def plot_values(registration_method):
global metric_values, multires_iterations
metric_values.append(registration_method.GetMetricValue())
# Clear the output area (wait=True, to reduce flickering), and plot current data
clear_output(wait=True)
# Plot the similarity metric values
plt.plot(metric_values, 'r')
plt.plot(multires_iterations, [metric_values[index] for index in multires_iterations], 'b*')
plt.xlabel('Iteration Number',fontsize=12)
plt.ylabel('Metric Value',fontsize=12)
plt.show()
# Callback invoked when the sitkMultiResolutionIterationEvent happens, update the index into the
# metric_values list.
def update_multires_iterations():
global metric_values, multires_iterations
multires_iterations.append(len(metric_values))
# -
# ## Read images
#
# We first read the images, casting the pixel type to that required for registration (Float32 or Float64) and look at them.
# +
fixed_image = sitk.ReadImage(fdata("training_001_ct.mha"), sitk.sitkFloat32)
moving_image = sitk.ReadImage(fdata("training_001_mr_T1.mha"), sitk.sitkFloat32)
interact(display_images, fixed_image_z=(0,fixed_image.GetSize()[2]-1), moving_image_z=(0,moving_image.GetSize()[2]-1), fixed_npa = fixed(sitk.GetArrayViewFromImage(fixed_image)), moving_npa=fixed(sitk.GetArrayViewFromImage(moving_image)));
# -
# ## Initial Alignment
#
# Use the CenteredTransformInitializer to align the centers of the two volumes and set the center of rotation to the center of the fixed image.
# +
initial_transform = sitk.CenteredTransformInitializer(fixed_image,
moving_image,
sitk.Euler3DTransform(),
sitk.CenteredTransformInitializerFilter.GEOMETRY)
moving_resampled = sitk.Resample(moving_image, fixed_image, initial_transform, sitk.sitkLinear, 0.0, moving_image.GetPixelID())
interact(display_images_with_alpha, image_z=(0,fixed_image.GetSize()[2]-1), alpha=(0.0,1.0,0.05), fixed = fixed(fixed_image), moving=fixed(moving_resampled));
# -
# ## Registration
#
# The specific registration task at hand estimates a 3D rigid transformation between images of different modalities. There are multiple components from each group (optimizers, similarity metrics, interpolators) that are appropriate for the task. Note that each component selection requires setting some parameter values. We have made the following choices:
#
# <ul>
# <li>Similarity metric, mutual information (Mattes MI):
# <ul>
# <li>Number of histogram bins, 50.</li>
# <li>Sampling strategy, random.</li>
# <li>Sampling percentage, 1%.</li>
# </ul>
# </li>
# <li>Interpolator, sitkLinear.</li>
# <li>Optimizer, gradient descent:
# <ul>
# <li>Learning rate, step size along traversal direction in parameter space, 1.0 .</li>
# <li>Number of iterations, maximal number of iterations, 100.</li>
# <li>Convergence minimum value, value used for convergence checking in conjunction with the energy profile of the similarity metric that is estimated in the given window size, 1e-6.</li>
# <li>Convergence window size, number of values of the similarity metric which are used to estimate the energy profile of the similarity metric, 10.</li>
# </ul>
# </li>
# </ul>
#
#
# Perform registration using the settings given above, and take advantage of the built in multi-resolution framework, use a three tier pyramid.
#
# In this example we plot the similarity metric's value during registration. Note that the change of scales in the multi-resolution framework is readily visible.
# +
registration_method = sitk.ImageRegistrationMethod()
# Similarity metric settings.
registration_method.SetMetricAsMattesMutualInformation(numberOfHistogramBins=50)
registration_method.SetMetricSamplingStrategy(registration_method.RANDOM)
registration_method.SetMetricSamplingPercentage(0.01)
registration_method.SetInterpolator(sitk.sitkLinear)
# Optimizer settings.
registration_method.SetOptimizerAsGradientDescent(learningRate=1.0, numberOfIterations=100, convergenceMinimumValue=1e-6, convergenceWindowSize=10)
registration_method.SetOptimizerScalesFromPhysicalShift()
# Setup for the multi-resolution framework.
registration_method.SetShrinkFactorsPerLevel(shrinkFactors = [4,2,1])
registration_method.SetSmoothingSigmasPerLevel(smoothingSigmas=[2,1,0])
registration_method.SmoothingSigmasAreSpecifiedInPhysicalUnitsOn()
# Don't optimize in-place, we would possibly like to run this cell multiple times.
registration_method.SetInitialTransform(initial_transform, inPlace=False)
# Connect all of the observers so that we can perform plotting during registration.
registration_method.AddCommand(sitk.sitkStartEvent, start_plot)
registration_method.AddCommand(sitk.sitkEndEvent, end_plot)
registration_method.AddCommand(sitk.sitkMultiResolutionIterationEvent, update_multires_iterations)
registration_method.AddCommand(sitk.sitkIterationEvent, lambda: plot_values(registration_method))
final_transform = registration_method.Execute(sitk.Cast(fixed_image, sitk.sitkFloat32),
sitk.Cast(moving_image, sitk.sitkFloat32))
# -
# ## Post registration analysis
# Query the registration method to see the metric value and the reason the optimization terminated.
#
# The metric value allows us to compare multiple registration runs as there is a probabilistic aspect to our registration, we are using random sampling to estimate the similarity metric.
#
# Always remember to query why the optimizer terminated. This will help you understand whether termination is too early, either due to thresholds being too tight, early termination due to small number of iterations - numberOfIterations, or too loose, early termination due to large value for minimal change in similarity measure - convergenceMinimumValue)
print('Final metric value: {0}'.format(registration_method.GetMetricValue()))
print('Optimizer\'s stopping condition, {0}'.format(registration_method.GetOptimizerStopConditionDescription()))
# Now visually inspect the results.
# +
moving_resampled = sitk.Resample(moving_image, fixed_image, final_transform, sitk.sitkLinear, 0.0, moving_image.GetPixelID())
interact(display_images_with_alpha, image_z=(0,fixed_image.GetSize()[2] - 1), alpha=(0.0,1.0,0.05), fixed = fixed(fixed_image), moving=fixed(moving_resampled));
# -
# If we are satisfied with the results, save them to file.
sitk.WriteImage(moving_resampled, os.path.join(OUTPUT_DIR, 'RIRE_training_001_mr_T1_resampled.mha'))
sitk.WriteTransform(final_transform, os.path.join(OUTPUT_DIR, 'RIRE_training_001_CT_2_mr_T1.tfm'))
| Python/60_Registration_Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kuamoto/LotoFacil_Lottery/blob/main/Lotofacil.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="r96RNP4vZlsp"
# Descrição: Este programa utiliza uma rede neural artificial para prever o próximo número da Loteria LotoFacil da Caixa Econômica Federal.
# + id="MVDg8mlqZvcM"
# Importando as bibliotecas do projeto.
import math
import pandas_datareader as web
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense, LSTM
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# + id="kTIJC7wQZ3To" outputId="62674f91-367f-4207-c00b-f63dfa523386" colab={"base_uri": "https://localhost:8080/", "height": 419}
# Obter os dados do arquivo criado.
df = pd.read_csv ("sample_data/lotofacil.csv")
# Mostrar os dados do arquivo.
df
# + id="prQ-hM9kZ6sN" outputId="ad4e1d86-af37-46ff-ce69-f5a48ffaf195" colab={"base_uri": "https://localhost:8080/"}
# Retorna a tupla da forma (linhas, colunas) do dataframe.
df.shape
# + id="21mYbx6XZ--n" outputId="4da5d5f2-1adc-463c-fe1d-2fb2dc534dd2" colab={"base_uri": "https://localhost:8080/", "height": 818}
# Visualize o histórico de valores da bola01
plt.figure(figsize=(30,20))
plt.title('Graph Lotto')
plt.plot(df['BL01'])
plt.xlabel('JOGO',fontsize=20)
plt.ylabel('BL01',fontsize=20)
plt.show()
# + id="jkkef6ieaE5x"
# Crie um novo dataframe apenas com a coluna 'BL01'
data = df.filter(['BL01'])
# Convertendo o dataframe em um array
dataset = data.values
# Obter/Calcular o número de linhas para treinar o modelo
training_data_len = math.ceil( len(dataset) *.8)
# + id="7WjIddjXaH2j"
# Dimensionando todos os dados para valores entre 0 e 1
scaler = MinMaxScaler(feature_range=(0, 1))
scaled_data = scaler.fit_transform(dataset)
# + id="jpyRM_dCaKwa"
# Criando o conjunto de dados de treinamento em escala
train_data = scaled_data[0:training_data_len , : ]
# Divida os dados em conjuntos de dados x_train e y_train
x_train=[]
y_train = []
for i in range(60,len(train_data)):
x_train.append(train_data[i-60:i,0])
y_train.append(train_data[i,0])
# + id="qna35gljaOaQ"
# Converta x_train e y_train em matrizes(array)
x_train, y_train = np.array(x_train), np.array(y_train)
# + id="URzfqnDaaQ5E"
# Remodele os dados para a forma aceita pelo LSTM
x_train = np.reshape(x_train, (x_train.shape[0],x_train.shape[1],1))
# + id="96i4CIfraURv"
# Construir o modelo de rede LSTM
model = Sequential()
model.add(LSTM(units=50, return_sequences=True,input_shape=(x_train.shape[1],1)))
model.add(LSTM(units=50, return_sequences=False))
model.add(Dense(units=25))
model.add(Dense(units=1))
# + id="m-mLmu8FaXUm"
# Compile o modelo
model.compile(optimizer='adam', loss='mean_squared_error')
# + id="eDMQ4UQJaa29" outputId="9608fe45-366a-4241-ba91-496c20e22889" colab={"base_uri": "https://localhost:8080/"}
# Treinando o model
model.fit(x_train, y_train, batch_size=1, epochs=1)
# + id="2SPxSmIYalP6"
# Conjunto de dados de teste
test_data = scaled_data[training_data_len - 60: , : ]
# Criando os conjuntos de dados x_test e y_test
x_test = []
y_test = dataset[training_data_len : , : ] # Pegue todas as linhas do índice para o resto e todas as colunas (neste caso, é apenas a coluna 'BL01').
for i in range(60,len(test_data)):
x_test.append(test_data[i-60:i,0])
# + id="YazJkADuaojj"
# Converter x_test em um array
x_test = np.array(x_test)
# + id="e8mwx33HarZn"
# Remodele os dados para a forma aceita pelo LSTM
x_test = np.reshape(x_test, (x_test.shape[0],x_test.shape[1],1))
# + id="oEtC5E8bauEB"
# Obter os valores de preços previstos dos modelos
predictions = model.predict(x_test)
predictions = scaler.inverse_transform(predictions) # Desfazer escala
# + id="UnvCaUOfawuY" outputId="4e0b9922-73ea-4ae5-d4e0-c0ef04b93533" colab={"base_uri": "https://localhost:8080/"}
# Calcular/Obter o valor de RMSE
rmse=np.sqrt(np.mean(((predictions- y_test)**2)))
rmse
# + id="6rjKYPw3a1Co" outputId="a002adb0-9543-41fc-80a3-076ca4a8cb43" colab={"base_uri": "https://localhost:8080/"}
# Plotar, Desenhar/Criar os dados para o gráfico
train = data[:training_data_len]
valid = data[training_data_len:]
valid['Predictions'] = predictions
# + id="SD8ZJnqEbMxS" outputId="16c990f8-8d86-48ff-e212-11a2929cdced" colab={"base_uri": "https://localhost:8080/", "height": 818}
# Visualizando os dados
plt.figure(figsize=(30,20))
plt.title('Graph Lotto Predictions')
plt.xlabel('JOGO',fontsize=20)
plt.ylabel('BL01',fontsize=20)
plt.plot(train['BL01'])
plt.plot(valid[['BL01', 'Predictions']])
plt.legend(['BL01', 'Predictions'], loc='lower right')
plt.show()
# + id="wPFxVOsbbX1y" outputId="5a613e92-f343-4738-d0cc-20ad8154c55e" colab={"base_uri": "https://localhost:8080/", "height": 419}
# Mostrar os valores válidos e previstos
valid
| Lotofacil.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:hodemulator]
# language: python
# name: conda-env-hodemulator-py
# ---
# I'm giving a talk about my 2DHOD AB models and their use in inferring cosmology. I need to make a few plots for that, and I'd like to do it all in one place.
#
# I need to make:
# * Cen-sat HOD plot x
# * Split decorated HOD plot x
# * Decorated HOD step func plot
# * “” for Yao model, cont model
# * SHAM ratio(s) plot
# * Tabulated HOD plot
#
import numpy as np
from pearce.mocks.kittens import TrainingBox, Chinchilla
from scipy.stats import binned_statistic, binned_statistic_2d
from halotools.utils.table_utils import compute_conditional_percentiles
from halotools.mock_observables import hod_from_mock, get_haloprop_of_galaxies
from matplotlib import pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set()
cat = TrainingBox(boxno=0)
cat.load(1.0, HOD='zheng07')
cat.model.param_dict['logMmin'] = 13.0
cat.model.param_dict['logM0'] = 12.5
print cat.model.param_dict
cat.populate(min_ptcl=50)
mass_bin_range = (11,16)
mass_bin_size = 0.1
cen_hod = cat.calc_hod(mass_bin_range=mass_bin_range, mass_bin_size=mass_bin_size, component='central')
sat_hod = cat.calc_hod(mass_bin_range=mass_bin_range, mass_bin_size=mass_bin_size, component='satellite')
mass_bins = np.logspace(mass_bin_range[0], mass_bin_range[1],
int((mass_bin_range[1] - mass_bin_range[0]) / mass_bin_size) + 1)
mass_bin_centers = (mass_bins[:-1] + mass_bins[1:]) / 2
# + active=""
# plt.plot(mass_bin_centers, cen_hod, label = 'Cens')
# plt.plot(mass_bin_centers, sat_hod, label = 'Sats')
# plt.plot(mass_bin_centers, cen_hod+sat_hod, label = 'All')
#
# plt.legend(loc='best')
# plt.loglog()
# plt.xlim(1e12,1e15)
# plt.ylim([1e-2, 1e3])
# plt.xlabel(r"Host Halo Mass [$M_{\odot}$]")
# plt.ylabel(r"$\langle N_t | M \rangle$")
# plt.show()
# -
# ---
# +
# TODO consistent plot language between each of these. Each model should have a corresponding color
# -
current_palette = sns.color_palette()
sns.palplot(current_palette)
model_color_map = {'HOD': (current_palette[0], "GnBu_d"),
'HSAB': (current_palette[1], "YlGn_d"),
'SHAM': (current_palette[2], "OrRd_d"),
'CAB': (current_palette[3], "RdPu_d"),
'CorrAB': (current_palette[4], "YlOrBr_d"),
'Halos': (current_palette[5], 'PuBu_d')} # add CMAPs too
def split_hod_plot(HOD, ab_params, n_splits = 4, cmap_name = 'blue'):
cat.load_model(1.0, HOD=HOD)
cat.model.param_dict['logMmin'] = 13.0
cat.model.param_dict['logM0'] = 12.5
cat.populate(ab_params, min_ptcl = 100)
catalog = cat.model.mock.galaxy_table
sec_percentiles = compute_conditional_percentiles(prim_haloprop = cat.model.mock.halo_table['halo_mvir'],\
sec_haloprop = cat.model.mock.halo_table['halo_nfw_conc'],
prim_haloprop_bin_boundaries= mass_bins)
sec_gal_percentiles = get_haloprop_of_galaxies(catalog['halo_id'], cat.model.mock.halo_table['halo_id'],
sec_percentiles)
# TODO bins here
hods = np.zeros((n_splits, len(mass_bin_centers)))
perc_ranges = np.linspace(0,1, n_splits+1)
cmap = sns.color_palette(cmap_name, n_splits)
#cmap = sns.dark_palette(cmap_name, n_splits)
for i,c in enumerate(cmap):
sec_bin_gals = np.logical_and(perc_ranges[i] < sec_gal_percentiles, sec_gal_percentiles<perc_ranges[i+1])
sec_bin_halos = np.logical_and(perc_ranges[i] < sec_percentiles, sec_percentiles<perc_ranges[i+1])
sec_gal_hist, _ = np.histogram(catalog[sec_bin_gals]['halo_mvir'], bins = mass_bins)
sec_halo_hist, _= np.histogram(cat.model.mock.halo_table[sec_bin_halos]['halo_mvir'], bins = mass_bins)
hods[i, :] = sec_gal_hist*1.0/sec_halo_hist
plt.plot(mass_bin_centers, hods[i], c = c, label = 'p < %0.2f'%perc_ranges[i+1])
gal_hist, _ = np.histogram(catalog['halo_mvir'], bins = mass_bins)
halo_hist, _= np.histogram(cat.model.mock.halo_table['halo_mvir'], bins = mass_bins)
full_hod = gal_hist*1.0/halo_hist
plt.plot(mass_bin_centers, full_hod, label = 'Full HOD', color = 'k')
plt.legend(loc='best')
plt.loglog()
plt.xlim(1e12,5e14)
plt.ylim([0, 40])
plt.xlabel(r"Host Halo Mass [$M_{\odot}$]")
plt.ylabel(r"$\langle N_t | M \rangle$")
plt.show()
# + active=""
# split_hod_plot('hsabZheng07', {'mean_occupation_centrals_assembias_param1': 0.5,
# 'mean_occupation_satellites_assembias_param1': -0.5}, n_splits=2,\
# cmap_name = model_color_map['HSAB'][1])
# + active=""
# split_hod_plot('abZheng07', {'mean_occupation_centrals_assembias_param1': 0.5,
# 'mean_occupation_satellites_assembias_param1': -0.5}, n_splits=4,\
# cmap_name = model_color_map['CAB'][1])
# + active=""
# split_hod_plot('corrZheng07', {'mean_occupation_centrals_assembias_corr1': 0.5,
# 'mean_occupation_satellites_assembias_corr1': -0.5}, n_splits=4,\
# cmap_name = model_color_map['CorrAB'][1])
# -
# ---
def select_mass_bin(bin_no, arr, mass_arr, mass_bins=mass_bins):
in_bin = np.logical_and(mass_bins[bin_no] < mass_arr, mass_arr < mass_bins[bin_no+1])
return arr[in_bin]
def single_bin_cen_occ_plot(HOD, ab_params, bin_no, color = current_palette[1]):
cat.load_model(1.0, HOD=HOD)
cat.model.param_dict['logMmin'] = 13.0
cat.model.param_dict['logM0'] = 12.5
cat.populate(ab_params, min_ptcl = 100)
mean_occ = cat.model._input_model_dictionary['centrals_occupation'].mean_occupation
base_mean_occ = cat.model._input_model_dictionary['centrals_occupation'].baseline_mean_occupation
baseline_result = base_mean_occ(prim_haloprop = cat.model.mock.halo_table['halo_mvir'])
pert_result = mean_occ(prim_haloprop = cat.model.mock.halo_table['halo_mvir'],\
sec_haloprop = cat.model.mock.halo_table['halo_nfw_conc'])
pert_in_bin = select_mass_bin(bin_no, pert_result, cat.model.mock.halo_table['halo_mvir'])
baseline_in_bin = select_mass_bin(bin_no, baseline_result, cat.model.mock.halo_table['halo_mvir'])
sec_in_bin = select_mass_bin(bin_no, cat.model.mock.halo_table['halo_nfw_conc'], cat.model.mock.halo_table['halo_mvir'])
sec_sort_idx = np.argsort(sec_in_bin)
baseline_in_bin_avg = binned_statistic(np.linspace(0, 1, len(sec_sort_idx)),
baseline_in_bin[sec_sort_idx], bins = 100)[0]
pert_in_bin_avg = binned_statistic(np.linspace(0, 1, len(sec_sort_idx)),
pert_in_bin[sec_sort_idx], bins = 100)[0]
# TODO compute mean in bins of conc perc
plt.plot(np.linspace(0,1,100), baseline_in_bin_avg, c = model_color_map['HOD'][0])
plt.plot(np.linspace(0,1,100), pert_in_bin_avg, c=color)
plt.ylim([-0.2,1.2])
plt.title(r'$\log_{10}M = $ %0.1f'%np.log10(mass_bin_centers[bin_no]))
plt.xlabel('Secondary Halo Propety Percentile')
plt.ylabel(r'$\langle N_{cen} | M \rangle$')
plt.show()
bin_no = 20
# + active=""
# single_bin_cen_occ_plot('hsabZheng07', {'mean_occupation_centrals_assembias_param1': 0.5,
# 'mean_occupation_satellites_assembias_param1': -0.5}, bin_no,
# color = model_color_map['HSAB'][0])
# + active=""
# single_bin_cen_occ_plot('abZheng07', {'mean_occupation_centrals_assembias_param1': 0.5,
# 'mean_occupation_satellites_assembias_param1': -0.5}, bin_no,
# color = model_color_map['CAB'][0])
# + active=""
# single_bin_cen_occ_plot('corrZheng07',\
# {'mean_occupation_centrals_assembias_corr1': 0.5,
# 'mean_occupation_satellites_assembias_corr1': -0.5}, bin_no,
# color = model_color_map['CorrAB'][0])
# + active=""
# single_bin_cen_occ_plot('corrZheng07',\
# {'mean_occupation_centrals_assembias_corr1': 0.5,
# 'mean_occupation_satellites_assembias_corr1': -0.5}, bin_no - bin_no/10,
# color = model_color_map['CorrAB'][0])
# -
# ---
from AbundanceMatching import *
from halotools.mock_observables import tpcf
from halotools.sim_manager import RockstarHlistReader
#sham clusterings computed on ds14b
rbins = np.logspace(-1.1, 1.6, 19)
rbc = (rbins[1:]+rbins[:-1])/2.0
# +
cat2 = Chinchilla(400, 2048)
cat2.load_catalog(1.0)
halocat = cat2.halocat.halo_table
# + active=""
# fname = '/u/ki/jderose/desims/BCCSims/c400-2048/rockstar/hlists_new/hlist_1.00000.list'
# reader = RockstarHlistReader(fname, cat2.columns_to_keep, cat2.cache_filenames[-1], cat2.simname,
# cat2.halo_finder, 0.0, cat2.version_name, cat2.Lbox, cat2.pmass,
# overwrite=True)
# reader.read_halocat(cat2.columns_to_convert)
# halocat = reader.halo_table
# -
def make_sham(halocat, ab_property, nd=1e-3):
#smf = np.genfromtxt('smf_dr72bright34_m7_lowm.dat', skip_header=True)[:,0:2]
#af = AbundanceFunction(smf[:,0], smf[:,1], (9.0, 12.9), faint_end_first = True)
lf = np.genfromtxt('/u/ki/swmclau2/des/AB_tests/lf_r_sersic_r.dat', skip_header=True)
af = AbundanceFunction(lf[:,1], lf[:,2],(-26, -12), )
scatter = 0.2
remainder = af.deconvolute(scatter, 20)
nd_halos = calc_number_densities(halocat[ab_property], cat2.Lbox) #don't think this matters which one i choose here
catalog = af.match(nd_halos, scatter)
n_obj_needed = int(nd*(cat2.Lbox**3))
non_nan_idxs = ~np.isnan(catalog)
sort_idxs = np.argsort(catalog[non_nan_idxs])#[::-1]
final_catalog = catalog[non_nan_idxs][sort_idxs[:n_obj_needed]]
output = halocat[non_nan_idxs][sort_idxs[:n_obj_needed]]
output['gal_smass'] = final_catalog
return output
galcat = make_sham(halocat, 'halo_vpeak')
gal_pos = np.vstack(np.array(galcat['halo_%s'%coord]) for coord in ['x', 'y', 'z']).T/cat2.h
sham_xi = tpcf(gal_pos, rbins, do_cross = False, estimator = 'Landy-Szalay', num_threads = 4, period = cat2.Lbox/cat2.h)
cen_mask = galcat['halo_upid'] == -1
sham_cen_hod = hod_from_mock(galcat[cen_mask]['halo_mvir_host_halo'], halocat['halo_mvir'], mass_bins)[0]
sham_sat_hod = hod_from_mock(galcat[~cen_mask]['halo_mvir_host_halo'], halocat['halo_mvir'], mass_bins)[0]
# + active=""
# plt.plot(mass_bin_centers, sham_cen_hod)
# plt.plot(mass_bin_centers, sham_sat_hod)
# plt.plot(mass_bin_centers, sham_cen_hod+sham_sat_hod)
#
# plt.loglog();
# -
from pearce.mocks.customHODModels import TabulatedCens, TabulatedSats, HSAssembiasTabulatedCens, HSAssembiasTabulatedSats
from pearce.mocks.customHODModels import AssembiasTabulatedCens, AssembiasTabulatedSats, CorrAssembiasTabulatedCens, CorrAssembiasTabulatedSats
# +
#sham_sat_hod[sham_sat_hod< 1e-2] = 0.0
# -
def tabulated_hod_xi(sham_hod, hod_model, ab_dict = {}):
sham_cen_hod, sham_sat_hod = sham_hod
cat2.load_model(1.0, HOD=hod_model, hod_kwargs = {'prim_haloprop_vals': mass_bin_centers,
#'sec_haloprop_key': 'halo_%s'%(mag_type),
'cen_hod_vals':sham_cen_hod,
'sat_hod_vals':sham_sat_hod} )
cat2.model.param_dict.update(ab_dict)
out = np.zeros((10, rbins.shape[0]-1,))
for i in xrange(10):
cat2.populate(min_ptcl=100)
out[i] = cat2.calc_xi(rbins)
return out.mean(axis = 0)
# + active=""
# cen_mask = cat2.model.mock.galaxy_table['gal_type'] == 'centrals'
# hod_cen_hod = hod_from_mock(cat2.model.mock.galaxy_table[cen_mask]['halo_mvir'], halocat['halo_mvir'], mass_bins)[0]
# hod_sat_hod = hod_from_mock(cat2.model.mock.galaxy_table[~cen_mask]['halo_mvir'], halocat['halo_mvir'], mass_bins)[0]
# + active=""
# plt.plot(mass_bin_centers, hod_cen_hod)
# plt.plot(mass_bin_centers, hod_sat_hod)
# plt.plot(mass_bin_centers, hod_cen_hod+sham_sat_hod)
#
# plt.loglog();
# + active=""
# hod_xi = tabulated_hod_xi((sham_cen_hod, sham_sat_hod), (TabulatedCens, TabulatedSats))
# + active=""
# hsab_xi = tabulated_hod_xi((sham_cen_hod, sham_sat_hod), (HSAssembiasTabulatedCens, HSAssembiasTabulatedSats),\
# ab_dict = {'mean_occupation_centrals_assembias_param1':1.0,
# 'mean_occupation_satellites_assembias_param1':-1.0})
# + active=""
# ab_xi = tabulated_hod_xi((sham_cen_hod, sham_sat_hod), (AssembiasTabulatedCens, AssembiasTabulatedSats),\
# ab_dict = {'mean_occupation_centrals_assembias_param1':1.0,
# 'mean_occupation_satellites_assembias_param1':-1.0})
# + active=""
# corrab_xi = tabulated_hod_xi((sham_cen_hod, sham_sat_hod), (CorrAssembiasTabulatedCens, CorrAssembiasTabulatedSats),\
# ab_dict = {'mean_occupation_centrals_assembias_corr1':1.0,
# 'mean_occupation_satellites_assembias_corr1':-1.0})
# + active=""
# plt.plot(rbc, sham_xi, label = 'SHAM')
# plt.plot(rbc, hod_xi, label = 'HOD')
# plt.plot(rbc, hsab_xi, label = 'HSAB')
# plt.plot(rbc, ab_xi, label = 'CAB')
# plt.plot(rbc, corrab_xi, label ='CorrAB')
#
# plt.legend(loc = 'best')
#
# plt.loglog()
# + active=""
# plt.plot(rbc, sham_xi/sham_xi, label = 'SHAM', color = model_color_map['SHAM'][0])
# plt.plot(rbc, hod_xi/sham_xi, label = 'HOD', color = model_color_map['HOD'][0])
# plt.plot(rbc, hsab_xi/sham_xi, label = 'HSAB', color = model_color_map['HSAB'][0])
# plt.plot(rbc, ab_xi/sham_xi, label = 'CAB', color = model_color_map['CAB'][0])
# plt.plot(rbc, corrab_xi/sham_xi, label ='CorrAB', color = model_color_map['CorrAB'][0])
# #plt.plot(rbc, hod_xi)
# plt.legend(loc = 'best')
# #plt.ylim([0.75, 1.25])
# plt.xlabel(r"$r$ [Mpc]")
# plt.ylabel(r"$\xi_{*}(r)/\xi_{SHAM}(r)$")
# plt.xscale('log')
# + active=""
# plt.plot(rbc, sham_xi/sham_xi, label = 'SHAM', color = model_color_map['SHAM'][0])
# plt.plot(rbc, hod_xi/sham_xi, label = 'HOD', color = model_color_map['HOD'][0])
# plt.plot(rbc, hsab_xi/sham_xi, label = 'HSAB', color = model_color_map['HSAB'][0])
# plt.plot(rbc, ab_xi/sham_xi, label = 'CAB', color = model_color_map['CAB'][0])
# plt.plot(rbc, corrab_xi/sham_xi, label ='CorrAB', color = model_color_map['CorrAB'][0])
# #plt.plot(rbc, hod_xi)
# plt.legend(loc = 'best')
# plt.ylim([0.9, 1.05])
# plt.xlabel(r"$r$ [Mpc]")
# plt.ylabel(r"$\xi_{*}(r)/\xi_{SHAM}(r)$")
# plt.xscale('log')
# -
# ---
def occ_jointplot(catalog, bin_no, mass_bins, params = ('halo_vpeak', 'halo_nfw_conc' ), param_bounds = None , color = current_palette[0]):
fig = plt.figure(figsize = (10,10))
mass_cut = np.logical_and(mass_bins[bin_no]< catalog['halo_mvir'], catalog['halo_mvir']<mass_bins[bin_no+1])
kit = catalog[mass_cut]
print np.log10(mass_bins[bin_no])
if param_bounds is None:
sns.jointplot(np.log10(kit[params[0]]), np.log10(kit[params[1]]), kind="hex", color = color)
else:
xlim = param_bounds[0]
ylim = param_bounds[1]
sns.jointplot(np.log10(kit[params[0]]), np.log10(kit[params[1]]), xlim=xlim, ylim=ylim, kind="kde", color = color)
plt.show()
param_bounds = ((2.1,2.8), (0.25, 2.75) )
bin_no = 12
# + active=""
# occ_jointplot(cat2.halocat.halo_table, bin_no, mass_bins,param_bounds = param_bounds, color = model_color_map['Halos'][0])
# + active=""
# occ_jointplot(galcat, bin_no, mass_bins, param_bounds=param_bounds, color=model_color_map['SHAM'][0])
# -
from halotools.mock_observables import get_haloprop_of_galaxies
def tabulated_hod_jointplot(sham_hod, hod_model,cmap_name, ab_dict = {},bin_no = 9 ):
sham_cen_hod, sham_sat_hod = sham_hod
cat2.load_model(1.0, HOD=hod_model, hod_kwargs = {'prim_haloprop_vals': mass_bin_centers,
#'sec_haloprop_key': 'halo_%s'%(mag_type),
'cen_hod_vals':sham_cen_hod,
'sat_hod_vals':sham_sat_hod} )
cat2.model.param_dict.update(ab_dict)
cat2.populate(min_ptcl=100)
for sec_param in ['halo_vpeak', 'halo_nfw_conc']: # TODO let user pass this in
val_gal = get_haloprop_of_galaxies(cat2.model.mock.galaxy_table['halo_id'], cat2.halocat.halo_table['halo_id'],
cat2.halocat.halo_table[sec_param])
cat2.model.mock.galaxy_table[sec_param] = val_gal
occ_jointplot(cat2.model.mock.galaxy_table,bin_no,\
mass_bins,param_bounds=param_bounds, color=model_color_map[cmap_name][0])
# + active=""
# tabulated_hod_jointplot((sham_cen_hod, sham_sat_hod), (TabulatedCens, TabulatedSats), 'HOD', bin_no = bin_no)
# -
tabulated_hod_jointplot((sham_cen_hod, sham_sat_hod), (HSAssembiasTabulatedCens, HSAssembiasTabulatedSats),
'HSAB', bin_no = bin_no,
ab_dict = {'mean_occupation_centrals_assembias_param1':1.0,
'mean_occupation_satellites_assembias_param1':-1.0})
# + active=""
# tabulated_hod_jointplot((sham_cen_hod, sham_sat_hod), (AssembiasTabulatedCens, AssembiasTabulatedSats),
# 'CAB', bin_no = bin_no,
# ab_dict = {'mean_occupation_centrals_assembias_param1':1.0,
# 'mean_occupation_satellites_assembias_param1':-1.0})
# + active=""
# tabulated_hod_jointplot((sham_cen_hod, sham_sat_hod), (CorrAssembiasTabulatedCens, CorrAssembiasTabulatedSats),
# 'CorrAB', bin_no = bin_no,
# ab_dict = {'mean_occupation_centrals_assembias_corr1':1.0,
# 'mean_occupation_satellites_assembias_corr1':-1.0})
# -
# ---
from pearce.mocks.customHODModels import Tabulated2DCens, Tabulated2DSats
from pearce.mocks.assembias_models.table_utils import compute_prim_haloprop_bins
from collections import Counter
def compute_occupations(halo_catalog, galaxy_catalog):
cens_occ = np.zeros((np.sum(halo_catalog['halo_upid'] == -1),))
sats_occ = np.zeros_like(cens_occ)
detected_central_ids = set(galaxy_catalog[galaxy_catalog['halo_upid']==-1]['halo_id'])
detected_satellite_upids = Counter(galaxy_catalog[galaxy_catalog['halo_upid']!=-1]['halo_upid'])
for idx, row in enumerate(halo_catalog[halo_catalog['halo_upid'] == -1]):
cens_occ[idx] = 1.0 if row['halo_id'] in detected_central_ids else 0.0
sats_occ[idx]+= detected_satellite_upids[row['halo_id']]
return cens_occ, sats_occ
cens_occ, sats_occ = compute_occupations(cat2.halocat.halo_table, galcat)
def calc_2dhod(mass_bins,conc_bins,sec_haloprop_key, halocat, cens_occ, sats_occ):
host_halos = halocat['halo_upid'] == -1
halo_mass = halocat['halo_mvir']
halo_sec =halocat[sec_haloprop_key]
host_halo_mass = halo_mass[host_halos]
host_halo_sec = halo_sec[host_halos]
#host_mass_bin_idxs = compute_prim_haloprop_bins(prim_haloprop_bin_boundaries=mass_bins, prim_haloprop = host_halo_mass)
mass_bin_idxs = compute_prim_haloprop_bins(prim_haloprop_bin_boundaries=mass_bins, prim_haloprop = halo_mass)
host_mass_bin_idxs = mass_bin_idxs[host_halos]
conditional_sec_percentiles = compute_conditional_percentiles(prim_haloprop = halo_mass,\
sec_haloprop = halo_sec,\
prim_haloprop_bin_boundaries = mass_bins)
#host_conditional_sec_percentiles = np.zeros((len(galcat),))
#host_halocat_idxs = np.in1d(halocat['halo_id'], galcat['halo_hostid'], assume_unique=True)
#print len(galcat), np.sum(host_halocat_idxs)
#host_sort_idxs = np.argsort(galcat['halo_hostid'])
#sort_idxs = np.argsort(halocat[host_halocat_idxs]['halo_id'])
#host_conditional_sec_percentiles[host_sort_idxs] = conditional_sec_percentiles[host_halocat_idxs][sort_idxs]
host_conditional_sec_percentiles = conditional_sec_percentiles[host_halos]
mean_ncen = np.zeros((len(mass_bins)-1, len(conc_bins)-1))
mean_nsat = np.zeros((len(mass_bins)-1, len(conc_bins)-1))
mass_bin_nos = range(len(mass_bins)-1)#,1)
for bin_no in mass_bin_nos:
bin_center = np.mean(mass_bins[bin_no-1:bin_no+1])
indices_of_host_mb = np.where(host_mass_bin_idxs == bin_no)[0]
indices_of_mb = np.where(mass_bin_idxs == bin_no)[0]
if len(indices_of_mb) == 0 or len(indices_of_host_mb) == 0:
continue
#print np.sum(~np.isfinite(halo_sec[host_conditional_sec_percentiles<0.9])),
#print np.sum(~np.isfinite(halo_sec[host_conditional_sec_percentiles>0.9]))
#print len(indices_of_mb), len(indices_of_host_mb)
(binned_cens, c_bins,_), (binned_sats,_,_) = binned_statistic(host_conditional_sec_percentiles[indices_of_host_mb],\
cens_occ[indices_of_host_mb],bins=conc_bins, statistic='sum'), \
binned_statistic(host_conditional_sec_percentiles[indices_of_host_mb],\
sats_occ[indices_of_host_mb],bins=conc_bins,statistic='sum')
binned_halos, _, _ = binned_statistic(conditional_sec_percentiles[indices_of_mb],
None, bins=conc_bins, statistic='count')
mean_ncen[bin_no-1,:] = binned_cens/binned_halos
mean_nsat[bin_no-1,:] = binned_sats/binned_halos
# NOTE these don't do anytng cuz there are no halos in these bins!
if np.any(np.isnan(mean_ncen[bin_no-1,:])):
mean_ncen[bin_no-1,np.isnan(mean_ncen[bin_no-1,:])] = 0.0#np.sum(binne)
if np.any(np.isnan(mean_nsat[bin_no-1,:])):
mean_nsat[bin_no-1,np.isnan(mean_nsat[bin_no-1,:] )] = 0.0#sat_hod[bin_no-1]
return mean_ncen, mean_nsat
#TODO what is up with the last bin?
conc_bins = np.linspace(0,1,41)
sham_cen_2dhod, sham_sat_2dhod = calc_2dhod(mass_bins, conc_bins, 'halo_nfw_conc', cat2.halocat.halo_table,
cens_occ, sats_occ)
# +
plt.plot(mass_bin_centers, sham_cen_hod)
plt.plot(mass_bin_centers, np.nanmean(sham_cen_2dhod, axis =1))
#plt.plot(mass_bin_centers, np.nanmean(sham_cen_hod_v2, axis =1))
plt.loglog();
# -
plt.plot(mass_bin_centers, sham_sat_hod)
plt.plot(mass_bin_centers, np.nanmean(sham_sat_2dhod, axis =1))
plt.loglog();
cat2.load_model(1.0, HOD=(Tabulated2DCens, Tabulated2DSats), hod_kwargs = {'prim_haloprop_bins': mass_bins,
'sec_haloprop_perc_bins': conc_bins,
'sec_haloprop_key': 'halo_nfw_conc',
'cen_hod_vals':sham_cen_2dhod,
'sat_hod_vals':sham_sat_2dhod})
cat2.populate()
xi_2d = cat2.calc_xi(rbins)
# +
plt.plot(rbc, sham_xi, label = 'SHAM', color = model_color_map['SHAM'][0])
plt.plot(rbc, xi_2d, label = '2DHOD', color = model_color_map['HOD'][0])
#plt.plot(rbc, hod_xi)
plt.legend(loc = 'best')
#plt.ylim([0.9, 1.05])
plt.xlabel(r"$r$ [Mpc]")
plt.ylabel(r"$\xi_{*}(r)/\xi_{SHAM}(r)$")
plt.loglog()
# +
plt.plot(rbc, sham_xi/sham_xi, label = 'SHAM', color = model_color_map['SHAM'][0])
plt.plot(rbc, xi_2d/sham_xi, label = '2DHOD', color = model_color_map['HOD'][0])
#plt.plot(rbc, hod_xi)
plt.legend(loc = 'best')
#plt.ylim([0.9, 1.05])
plt.xlabel(r"$r$ [Mpc]")
plt.ylabel(r"$\xi_{*}(r)/\xi_{SHAM}(r)$")
plt.xscale('log')
# -
conc_bins = np.linspace(0,1,21)
sham_cen_2dhod, sham_sat_2dhod = calc_2dhod(mass_bins, conc_bins, 'halo_vpeak', cat2.halocat.halo_table,
cens_occ, sats_occ)
cat2.load_model(1.0, HOD=(Tabulated2DCens, Tabulated2DSats), hod_kwargs = {'prim_haloprop_bins': mass_bins,
'sec_haloprop_perc_bins': conc_bins,
'sec_haloprop_key': 'halo_nfw_conc',
'cen_hod_vals':sham_cen_2dhod,
'sat_hod_vals':sham_sat_2dhod})
cat2.populate()
xi_2d_vpeak = cat2.calc_xi(rbins)
# +
plt.plot(rbc, sham_xi/sham_xi, label = 'SHAM', color = model_color_map['SHAM'][0])
plt.plot(rbc, xi_2d/sham_xi, label = '2DHOD Conc', color = model_color_map['HOD'][0])
plt.plot(rbc, xi_2d_vpeak/sham_xi, label = '2DHOD Vpeak', color = model_color_map['CorrAB'][0])
#plt.plot(rbc, hod_xi)
plt.legend(loc = 'best')
#plt.ylim([0.9, 1.05])
plt.xlabel(r"$r$ [Mpc]")
plt.ylabel(r"$\xi_{*}(r)/\xi_{SHAM}(r)$")
plt.xscale('log')
# -
# ---
# +
fig = plt.figure(figsize = (10,10))
cbc = (conc_bins[1:]+conc_bins[:-1])/2.0
mass_slice = np.logical_and(10**12.0 < mass_bin_centers, mass_bin_centers < 10**13.5)
colors = sns.color_palette(model_color_map['SHAM'][1], len(mass_bin_centers[mass_slice]))
for idx, (row,c,m) in enumerate(zip(sham_cen_2dhod[mass_slice], colors, mass_bin_centers[mass_slice])):
if idx%2!=0:
continue
plt.plot(cbc, row, color = c, label = r'%.1f $\log M_{\odot}$'%np.log10(m))
#plt.ylim(-0.2,1.2)
plt.xlim(-0.2, 1.2);
plt.xlabel('%s percentile'%r"$V_{peak}$")
plt.ylabel(r'$<N_{cen}(x)|M>$')
plt.yscale('log')
plt.ylim([0.1, 1.1])
plt.legend(loc='best')
# +
fig = plt.figure(figsize = (10,10))
cbc = (conc_bins[1:]+conc_bins[:-1])/2.0
mass_slice = np.logical_and(10**12.0 < mass_bin_centers, mass_bin_centers < 10**14.5)
colors = sns.color_palette(model_color_map['SHAM'][1], len(mass_bin_centers[mass_slice]))
for idx, (row,c,m) in enumerate(zip(sham_sat_2dhod[mass_slice], colors, mass_bin_centers[mass_slice])):
if idx%2!=0:
continue
plt.plot(cbc, row, color = c, label = r'%.1f $\log M_{\odot}$'%np.log10(m))
#plt.ylim(-0.2,1.2)
plt.xlim(-0.2, 1.2);
plt.xlabel('%s percentile'%r"$V_{peak}$")
plt.ylabel(r'$<N_{sat}(x)|M>$')
plt.yscale('log')
#plt.ylim([0.1, 1.1])
plt.legend(loc='best')
# -
cat2.load_model(1.0, HOD='corrZheng07', hod_kwargs = {'prim_haloprop_vals': mass_bin_centers,
'sec_haloprop_key': 'halo_vpeak',#%(mag_type),
'cen_hod_vals':sham_cen_hod,
'sat_hod_vals':sham_sat_hod} )
ab_dict = {'mean_occupation_centrals_assembias_corr1':1.0,
'mean_occupation_satellites_assembias_corr1':-1.0}
cat2.model.param_dict.update(ab_dict)
cat2.populate(min_ptcl=100)
cens_occ_hod, sats_occ_hod = compute_occupations(cat2.halocat.halo_table, cat2.model.mock.galaxy_table)
hod_cen_2dhod, hod_sat_2dhod = calc_2dhod(mass_bins, conc_bins, 'halo_vpeak', cat2.halocat.halo_table,
cens_occ_hod, sats_occ_hod)
# +
fig = plt.figure(figsize = (10,10))
cbc = (conc_bins[1:]+conc_bins[:-1])/2.0
mass_slice = np.logical_and(10**11.5 < mass_bin_centers, mass_bin_centers < 10**13.5)
colors = sns.color_palette(model_color_map['CorrAB'][1], len(mass_bin_centers[mass_slice]))
for idx, (row,c,m) in enumerate(zip(hod_cen_2dhod[mass_slice], colors, mass_bin_centers[mass_slice])):
if idx%2!=0:
continue
plt.plot(cbc, row, color = c, label = r'%.1f $\log M_{\odot}$'%np.log10(m))
plt.ylim(0.1,1.1)
plt.xlim(-0.2, 1.2);
plt.xlabel('%s percentile'%r"$V_{peak}$")
plt.ylabel(r'$<N_{cen}(c)|M>$')
plt.yscale('log')
plt.legend(loc='best')
# +
fig = plt.figure(figsize = (10,10))
cbc = (conc_bins[1:]+conc_bins[:-1])/2.0
mass_slice = np.logical_and(10**11.5 < mass_bin_centers, mass_bin_centers < 10**13.5)
colors = sns.color_palette(model_color_map['CorrAB'][1], len(mass_bin_centers[mass_slice]))
for idx, (row,c,m) in enumerate(zip(hod_sat_2dhod[mass_slice], colors, mass_bin_centers[mass_slice])):
if idx%2!=0:
continue
plt.plot(cbc, row, color = c, label = r'%.1f $\log M_{\odot}$'%np.log10(m))
#plt.ylim(0.1,1.1)
plt.xlim(-0.2, 1.2);
plt.xlabel('%s percentile'%r"$V_{peak}$")
plt.ylabel(r'$<N_{cen}(c)|M>$')
plt.yscale('log')
plt.legend(loc='best')
# -
cat2.load_model(1.0, HOD='hsabZheng07', hod_kwargs = {'prim_haloprop_vals': mass_bin_centers,
'sec_haloprop_key': 'halo_vpeak',
'cen_hod_vals':sham_cen_hod,
'sat_hod_vals':sham_sat_hod} )
ab_dict = {'mean_occupation_centrals_assembias_param1':1.0,
'mean_occupation_satellites_assembias_param1':-1.0}
cat2.model.param_dict.update(ab_dict)
cat2.populate(min_ptcl=100)
cens_occ_hod, sats_occ_hod = compute_occupations(cat2.halocat.halo_table, cat2.model.mock.galaxy_table)
hod_cen_2dhod, hod_sat_2dhod = calc_2dhod(mass_bins, conc_bins, 'halo_vpeak', cat2.halocat.halo_table,
cens_occ_hod, sats_occ_hod)
# +
fig = plt.figure(figsize = (10,10))
cbc = (conc_bins[1:]+conc_bins[:-1])/2.0
mass_slice = np.logical_and(10**11.5 < mass_bin_centers, mass_bin_centers < 10**13.5)
colors = sns.color_palette(model_color_map['HSAB'][1], len(mass_bin_centers[mass_slice]))
for idx, (row,c,m) in enumerate(zip(hod_cen_2dhod[mass_slice], colors, mass_bin_centers[mass_slice])):
if idx%2!=0:
continue
plt.plot(cbc, row, color = c, label = r'%.1f $\log M_{\odot}$'%np.log10(m))
plt.ylim(0.1,1.1)
plt.xlim(-0.2, 1.2);
plt.xlabel('%s percentile'%r"$V_{peak}$")
plt.ylabel(r'$<N_{cen}(c)|M>$')
#plt.yscale('log')
plt.legend(loc='best')
# -
| notebooks/AB_tests/Plots for AB Conference Talk 2019.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Organizing Topic Modeling Results
# -
import pandas as pd
# Loading topic modeling results
reviews = pd.read_csv(".//LDA/if_mallet_41topics_representation.tsv", sep = '\t')
reviews = reviews.drop("Keywords", axis = 1)
reviews = reviews.drop("Unnamed: 0", axis = 1)
reviews = reviews.drop("Topic_Perc_Contrib", axis = 1)
reviews['Topic_Num'] = reviews['Topic_Num'].astype(int)
# +
# Reassembling topics
calorie_tracking = reviews.loc[reviews['Topic_Num'].isin([20, 9, 35])] # according to topic index number
calorie_tracking["sub_topic"] = 1
tracking_features = reviews.loc[reviews['Topic_Num'].isin([12, 23, 40])]
tracking_features["sub_topic"] = 2
food_intake = reviews.loc[reviews['Topic_Num'] == 11]
food_intake["sub_topic"] = 3
time_logging = reviews.loc[reviews['Topic_Num'] == 6]
time_logging["sub_topic"] = 4
reminders = reviews.loc[reviews['Topic_Num'] == 23]
reminders["sub_topic"] = 5
guided_schedule = reviews.loc[reviews['Topic_Num'].isin([1, 17])]
guided_schedule["sub_topic"] = 6
workout_routines = reviews.loc[reviews['Topic_Num'].isin([26, 38])]
workout_routines["sub_topic"] = 7
meal_recommendations = reviews.loc[reviews['Topic_Num'].isin([19, 25])]
meal_recommendations["sub_topic"] = 8
informational_features = reviews.loc[reviews['Topic_Num'] == 14]
informational_features["sub_topic"] = 9
paid_content = reviews.loc[reviews['Topic_Num'] == 3]
paid_content["sub_topic"] = 10
weight_loss = reviews.loc[reviews['Topic_Num'].isin([2, 4, 10, 13, 15])]
weight_loss["sub_topic"] = 11
changing_lifestyle = reviews.loc[reviews['Topic_Num'] == 5]
changing_lifestyle["sub_topic"] = 12
improve_health = reviews.loc[reviews['Topic_Num'] == 18]
improve_health["sub_topic"] = 13
easy_use = reviews.loc[reviews['Topic_Num'].isin([16, 28, 36])]
easy_use["sub_topic"] = 14
flexibility = reviews.loc[reviews['Topic_Num'] == 8]
flexibility["sub_topic"] = 15
customization = reviews.loc[reviews['Topic_Num'] == 27]
customization["sub_topic"] = 16
ui = reviews.loc[reviews['Topic_Num'] == 7]
ui["sub_topic"] = 17
aesthetics = reviews.loc[reviews['Topic_Num'] == 31]
aesthetics["sub_topic"] = 18
df = pd.concat([calorie_tracking, tracking_features, food_intake, time_logging, reminders, guided_schedule,
workout_routines, meal_recommendations, informational_features, paid_content, weight_loss,
changing_lifestyle, improve_health, easy_use, flexibility, customization, ui, aesthetics])
df = df.reset_index()
df = df.drop("Topic_Num", axis = 1)
df = df.drop("index", axis = 1)
# -
df
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyser = SentimentIntensityAnalyzer()
# +
# Retrieving compound scores of each review
compound_scores = []
for i in df["Text"]:
compound_scores.append(analyser.polarity_scores(i).get("compound"))
df['compound_scores'] = compound_scores
# +
# decide sentiment as positive, negative and neutral
sentiment = []
for i in df['compound_scores']:
if i >= 0.05 :
sentiment.append("Positive")
elif i <= -0.05 :
sentiment.append("Negative")
else :
sentiment.append("Neutral")
print(len(sentiment))
df['sentiment'] = sentiment
# -
# Organizing sentiment distribution
positives = []
negatives = []
neutrals = []
for i in range(1, 19):
cluster = df[df.sub_topic == i]
total = cluster.shape[0]
positive = cluster[cluster.sentiment == "Positive"].shape[0]
negative = cluster[cluster.sentiment == "Negative"].shape[0]
neutral = cluster[cluster.sentiment == "Neutral"].shape[0]
positives.append(round(positive/total*100, 2))
negatives.append(round(negative/total*100, 2))
neutrals.append(round(neutral/total*100, 2))
sentiment_df = pd.DataFrame()
sentiment_df["topic_index"] = range(1,19)
sentiment_df["positive"] = positives
sentiment_df["negative"] = negatives
sentiment_df["neutral"] = neutrals
sentiment_df.to_csv("IF_sentiment_percentage.csv")
| Sentiment_Analysis/IF_sentiment_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Music Recommendation System Evaluation (MagnaTagATune)
# In this Notebook we are going to prepare MagnaTagATune dataset for recommendation and use it for toy recommendation systems. Before running this notebook generate a genre classifier model and download the 'Tag annotations', 'Similarity data' and the dataset. Link: http://mirg.city.ac.uk/codeapps/the-magnatagatune-dataset.
# ### Imports
# +
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
import numpy as np
import tensorflow as tf
import tensorflow.keras
from tensorflow.keras import backend as K
from tensorflow.keras.models import Sequential, load_model
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras import regularizers
from tensorflow.keras import utils as np_utils
from tensorflow.python.keras.layers import Layer
from sklearn.metrics import precision_recall_fscore_support, confusion_matrix, accuracy_score, mean_squared_error
from sklearn.utils.multiclass import unique_labels
import matplotlib as plt
plt.use("Agg")
# %matplotlib inline
from matplotlib import pyplot as plt
import itertools
import time
import warnings
warnings.filterwarnings('ignore')
import IPython.display as ipd
import librosa as lb
from scipy.io import wavfile
from scipy.spatial.distance import cosine as cos_distance
from sklearn.model_selection import train_test_split
import pandas as pd
# -
# ### Preprocess Data
SR = 22050
N_FFT = 512
HOP_LENGTH = N_FFT // 2
N_MELS = 64
# +
def log_melspectrogram(data, log=True, plot=False, num='', genre=""):
melspec = lb.feature.melspectrogram(y=data, hop_length = HOP_LENGTH, n_fft = N_FFT, n_mels = N_MELS)
if log:
melspec = lb.power_to_db(melspec**2)[:,1200:1373]
if plot:
melspec = melspec[np.newaxis, :]
plt.imshow(melspec.reshape((melspec.shape[1],melspec.shape[2])))
plt.savefig('melspec'+str(num)+'_'+str(genre)+'.png')
return melspec
def batch_log_melspectrogram(data_list, log=True, plot=False):
melspecs = np.asarray([log_melspectrogram(data_list[i],log=log,plot=plot) for i in range(len(data_list))])
return melspecs
# -
# ### Similarity Functions
# +
def euclid(x, y):
return np.sqrt(np.sum((x - y)**2, axis=1))
def cosine(x, y):
num = np.sum(x*y, axis=1)
denom = np.sqrt(np.dot(x, x.T))*np.diag(np.sqrt(np.dot(y, y.T)))
return np.squeeze(1 - num/denom, axis=0).T
def jaccard_similarity(list1, list2):
s1 = set(list1)
s2 = set(list2)
return len(s1.intersection(s2)) / len(s1.union(s2))
# -
# ### Load Model
model = load_model('music_classifier_v1.h5', compile=False) # Place here your genre classifier
model.summary()
# ## Prepare Data for Evaluation of Music Recommendation
# Get Annotations
annotations = pd.read_csv('annotations_final.csv', sep="\t")
annotations.head(2)
# +
# Get Votes
comparisons = pd.read_csv('comparisons_final.csv', sep="\t")
comparisons.head(3)
# -
# Extract most similar playlists
sum_numvotes = 1
clip1_numvotes = comparisons['clip1_numvotes']
clip2_numvotes = comparisons['clip2_numvotes']
clip3_numvotes = comparisons['clip3_numvotes']
clip_numvotes = clip1_numvotes + clip2_numvotes + clip3_numvotes
similar_playlists_idx = np.squeeze(np.where(clip_numvotes<=sum_numvotes))
dissimilar_playlists_idx = np.squeeze(np.where(clip_numvotes>sum_numvotes))
comparisons.drop(list(dissimilar_playlists_idx), inplace=True)
similar_playlists = comparisons
# +
# Load audio files
songs = []
ids = []
for i in range(1,4):
j=0
for path in similar_playlists['clip' + str(i) + '_mp3_path']:
song, _ = lb.load('mp3/'+ path, sr=None)
songs.append(song)
ids.append(list(similar_playlists['clip' + str(i) + '_id'])[j])
j+=1
# -
# Mel-spectrograms
data = batch_log_melspectrogram(songs)
# Predict the genres for the data
corpus_predictions = model.predict(data[:, :, :, np.newaxis])
# +
# Distribution of Data
distribution = []
for i in range(len(similar_playlists)):
idx = corpus_predictions[i].argmax()
one_hot = np.zeros(10)
one_hot[idx] = 1
distribution.append(one_hot)
# Blues, Classical, Country, Disco, Hip hop, Jazz, Metal, Pop, Reggae, Rock
print(np.asarray(distribution).sum(axis=0))
# -
# ## Similarity based on last FC-Layer
# Predict only for column clip1_id
# Accuracy: Top-5 Accuracy
predicted_playlist_cosine = []
i=0
for x in data[:len(similar_playlists)]:
tmp_predictions = np.delete(corpus_predictions, i, axis=0)
tmp_ids = np.delete(ids, i)
test_prediction = model.predict(x[np.newaxis, :, :, np.newaxis], steps=1)
cosine_vec = cosine(test_prediction, tmp_predictions)
predicted_playlist_cosine.append((tmp_ids[cosine_vec.argsort()[0]],
tmp_ids[cosine_vec.argsort()[1]],
tmp_ids[cosine_vec.argsort()[2]],
tmp_ids[cosine_vec.argsort()[3]],
tmp_ids[cosine_vec.argsort()[4]]))
i+=1
# +
score = 0
target_playlist = list(np.asarray(comparisons[[ 'clip2_id', 'clip3_id']]))
for i in range(len(predicted_playlist_cosine)):
if (target_playlist[i][0] in predicted_playlist_cosine[i]) or (target_playlist[i][1] in predicted_playlist_cosine[i]):
score+=1
top5_accuracy = score /len(similar_playlists) #if predicted playlist contains all songs in target playlist
print(top5_accuracy)
# -
# ## Data-Driven Approach
# ### Prepare data
# +
val_size = round(len(similar_playlists)*0.1)
test_size = round(len(similar_playlists)*0.1)
train_set = corpus_predictions[:len(similar_playlists)-test_size-val_size][:, :, np.newaxis]
train_size = len(train_set)
val_set = corpus_predictions[len(similar_playlists)-test_size-val_size:len(similar_playlists)-test_size][:, :, np.newaxis]
test_set = corpus_predictions[len(similar_playlists)-test_size:len(similar_playlists)][:, :, np.newaxis]
data = np.concatenate((corpus_predictions[:len(similar_playlists)],
corpus_predictions[len(similar_playlists):(len(similar_playlists)*2)]), axis=1)[:, :, np.newaxis]
x_test = data[::-1][:test_size]
x = np.delete(data[::-1], list(range(test_size)), axis=0 )
# +
# Ground Truth: Jaccard Similarity
annotations = pd.read_csv('annotations_final.csv', sep="\t")
annotations = np.asarray(annotations.drop('mp3_path', axis=1))
clip1_tags = annotations[[list(annotations[:,0]).index(id) for id in similar_playlists['clip1_id']]][:, 1:]
clip2_tags = annotations[[list(annotations[:,0]).index(id) for id in similar_playlists['clip2_id']]][:, 1:]
jaccard = []
for i in range(len(similar_playlists)):
jaccard.append(jaccard_similarity(clip1_tags[i], clip2_tags[i]))
truths = np.asarray(jaccard)[:, np.newaxis]#, np.newaxis]
y_test = truths[::-1][:test_size]
y = np.delete(truths[::-1], list(range(test_size)), axis=0 )
# -
#split data
x_tr, x_val, y_tr, y_val = train_test_split(x, y, test_size=0.1, shuffle=False)
# ### FC: Network Architektur
# +
reg = 0.001
drop = 0.5
recommendation_model = Sequential()
recommendation_model.add(Dense(20, activation='relu', use_bias=True, kernel_regularizer=regularizers.l2(reg), input_shape=(20, 1)))
recommendation_model.add(Dropout(drop))
recommendation_model.add(Dense(30, activation='relu', kernel_regularizer=regularizers.l2(reg), use_bias=True))
recommendation_model.add(Dropout(drop))
recommendation_model.add(Flatten(data_format=None))
recommendation_model.add(Dense(1, activation='sigmoid', kernel_regularizer=regularizers.l2(reg), use_bias=True))
def mean_pred(y_true, y_pred):
return K.mean(y_pred)
recommendation_model.compile(loss=tensorflow.keras.losses.mean_squared_error,
optimizer=tensorflow.keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0),
metrics=['accuracy', mean_pred]) #accuracy does not have any meaning
recommendation_model.summary()
# -
# ### Fit/Train model and evaluate
# +
batch_size = 4
epochs = 250
start = time.time()
history = recommendation_model.fit(x=x_tr,
y=y_tr,
validation_data=(x_val, y_val),
batch_size=batch_size,
epochs=epochs,
verbose=0,
shuffle=True)
print("\n")
recommendation_model.evaluate(x_val, y_val)
print("\nTraining took ", time.time()-start, "seconds")
# +
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('MSE')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper left')
plt.show()
#NOTE: Dropout makes the training loss >= test loss for every epoch
# -
# ### Evaluation
# #### MSE
predictions = recommendation_model.predict(x_test)
mse = mean_squared_error(y_test, predictions)
print(mse)
# ### Save model
recommendation_model.save('recommendation_model.h5')
| music _recommendation_magnatagatune.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import mwdsbe
import schuylkill as skool
import pandas as pd
import time
from matplotlib import pyplot as plt
import seaborn as sns
## Functions
def drop_duplicates_by_date(df, date_column):
df.sort_values(by=date_column, ascending=False, inplace=True)
df = df.loc[~df.index.duplicated(keep="first")]
df.sort_index(inplace=True)
return df
# ## Data
registry = mwdsbe.load_registry() # geopandas df
# contract payments in fy 2017
payments = pd.read_csv(r'C:\Users\dabinlee\Documents\GitHub\mwdsbe\mwdsbe\data\Payments\city_payments_detailed_2017.csv')
len(payments)
# Clean Data
# +
ignore_words = ['inc', 'group', 'llc', 'corp', 'pc', 'incorporated', 'ltd', 'co', 'associates', 'services', 'company', 'enterprises', 'enterprise', 'service', 'corporation']
cleaned_registry = skool.clean_strings(registry, ['company_name', 'dba_name'], True, ignore_words)
cleaned_payments = skool.clean_strings(payments, ['vendor_name'], True, ignore_words)
cleaned_registry = cleaned_registry.dropna(subset=['company_name'])
cleaned_payments = cleaned_payments.dropna(subset=['vendor_name'])
# -
len(cleaned_registry)
len(cleaned_payments)
len(cleaned_payments.vendor_name.unique()) # number of unique vendors
# MWDSBE Payments Data
mwdsbe_payments = pd.read_excel(r'C:\Users\dabinlee\Desktop\mwdsbe\data\payments\tf-idf-85.xlsx')
mwdsbe_payments.rename(columns={'Unnamed: 0': 'left_index'}, inplace=True)
mwdsbe_payments.set_index('left_index', inplace=True)
len(mwdsbe_payments.vendor_name.unique())
207 / 5506 * 100 # percentage of minority vendor number in payments data
# ## Analysis
# ### Percentage of payments for MWDSBE
mwdsbe_payments = pd.read_excel(r'C:\Users\dabinlee\Desktop\mwdsbe\data\payments\tf-idf-85.xlsx')
mwdsbe_payments.rename(columns={'Unnamed: 0': 'left_index'}, inplace=True)
mwdsbe_payments.set_index('left_index', inplace=True)
len(mwdsbe_payments)
# Filter payments to only have 1-2 classes
mwdsbe_payments = mwdsbe_payments.loc[mwdsbe_payments['char_'].isin([1,2])]
# Get all payments of mwdsbe
# * current mwdsbe_payments data only contains 1-2 payments
mwdsbe_vendors = mwdsbe_payments.vendor_name.unique()
len(mwdsbe_vendors)
all_mwdsbe_payments = cleaned_payments.loc[cleaned_payments['vendor_name'].isin(mwdsbe_vendors)]
len(all_mwdsbe_payments)
# Total Transaction Amount of MWDSBE
tot_amt_MWDSBE = all_mwdsbe_payments['transaction_amount'].sum()
# Total Transaction Amount of all business payments
tot_amt_all = all_payments['transaction_amount'].sum()
# percentage
tot_amt_MWDSBE / tot_amt_all * 100
# ### Comparing top 10 Locations of OEO report and registry and matched
# #### Registry
# In Philadelphia
philadelphia = cleaned_registry.loc[cleaned_registry.local]
len(philadelphia)/len(cleaned_registry) * 100
# Other PA cities
cleaned_registry_df = pd.DataFrame(cleaned_registry.drop(labels=['geometry'], axis=1))
other_PA_cities = cleaned_registry_df.loc[((~cleaned_registry_df.out_of_state) & (~cleaned_registry_df.local))]
len(other_PA_cities)/len(cleaned_registry) * 100
# #### Matched
philadelphia = mwdsbe_payments.loc[mwdsbe_payments.local]
len(philadelphia)/len(mwdsbe_payments) * 100
| mwdsbe/Notebooks/By Data/Payments/Payments2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [finding-important-factors-to-survive-titanic](#https://www.kaggle.com/jatturat/finding-important-factors-to-survive-titanic)
#
# [EDA of important features](#https://www.kaggle.com/dollardollar/eda-of-important-features)
# ## 1. Exploratory Data Analysis (EDA)
# ### 1A. Import Libraries and Settings
# +
# # ! conda install --yes seaborn
# +
# Basic Libraries
import numpy as np
import pandas as pd
# Operational System
import os
# Feature Scaling
#from sklearn.preprocessing import RobustScaler
# Visaulization
import matplotlib.pyplot as plt
import seaborn as sns
# Classifier (machine learning algorithm)
# from sklearn.linear_model import LogisticRegression
# from sklearn.neighbors import KNeighborsClassifier
# from sklearn.svm import SVC
# from sklearn.naive_bayes import GaussianNB
# from sklearn.tree import DecisionTreeClassifier
# from sklearn.ensemble import RandomForestClassifier
# from xgboost import XGBClassifier
# Evaluation
# from sklearn.model_selection import cross_val_score, cross_val_predict
# Parameter Tuning
# from sklearn.model_selection import GridSearchCV
# Settings
# pd.options.mode.chained_assignment = None # Stop warning when use inplace=True of fillna
# -
# ### 2B. Import Dataset
print(os.getcwd())
train_set = pd.read_csv('data/titanic_train.csv')
test_set = pd.read_csv('data/titanic_test.csv')
# ### 2C. Dataset Checking
train_set.head(3)
test_set.head(3)
# number of rows
len(train_set)
train_set.count()
train_set.describe()
train_set.isnull().sum()
len(test_set)
test_set.describe()
test_set.isnull().sum()
# ### 2D. Features Visualization
# The first thing to do is to visualize the data and find any valuable/hidden information inside each feature. In this section, I tried to visualize existing data first without filling the missing data yet to keep things in steps.
#
# I use these methods to create graphs based on type of each feature.
# +
# Continuous Data Plot
def cont_plot(df, feature_name, target_name, palettemap, hue_order, feature_scale):
df['Counts'] = "" # A trick to skip using an axis (either x or y) on splitting violinplot
fig, [axis0,axis1] = plt.subplots(1,2,figsize=(14,5))
sns.distplot(df[feature_name], ax=axis0);
sns.violinplot(x=feature_name, y="Counts", hue=target_name, hue_order=hue_order, data=df,
palette=palettemap, split=True, orient='h', ax=axis1)
axis1.set_xticks(feature_scale)
plt.show()
# WARNING: This will leave Counts column in dataset if you continues to use this dataset
# Categorical/Ordinal Data Plot
def cat_plot(df, feature_name, target_name, palettemap):
fig, [axis0,axis1] = plt.subplots(1,2,figsize=(14,5))
df[feature_name].value_counts().plot.pie(autopct='%1.1f%%',ax=axis0)
sns.countplot(x=feature_name, hue=target_name, data=df,
palette=palettemap, ax=axis1, )
plt.show()
survival_palette = {0: "black", 1: "orange"} # Color map for visualization
# -
# #### 2Da. Pclass
# +
cat_plot(train_set, 'Pclass','Survived', survival_palette)
# year_palette = {0: "black", 1899: "orange", 2018: "green"}
# cat_plot(solditems_pd, 'domain','year', survival_palette)
# -
# #### 2Db. Sex
cat_plot(train_set, 'Sex', 'Survived', survival_palette)
# #### 2Dc. Embarked
cat_plot(train_set, 'Embarked', 'Survived', survival_palette)
# #### 2Dd. Age
# dropping temporaly null because visualization method can't deal with missing data
age_set_nonan = train_set[['Age','Survived']].copy().dropna(axis=0)
cont_plot(age_set_nonan, 'Age', 'Survived', survival_palette, hue_order=[1,0], feature_scale=range(0,100,20))
min_age = min(age_set_nonan.Age)
min_age
age_set_nonan.Age.sort_values()
max_age = max(train_set.Age)
max_age
# Creating categories for age
age_bins = [0, 15, 30, 45, 60, 75, max_age+10]
age_labels = ['1.[0, 15)','2.[15, 30)','3.[30, 45)','4.[45, 60)','5.[60, 75)', f'6.[75, {max_age+10})']
age_set_nonan['AgeRange'] = pd.cut(age_set_nonan['Age'], bins=age_bins, labels=age_labels, include_lowest=True)
age_set_nonan.head()
cat_plot(age_set_nonan, 'AgeRange', 'Survived', palettemap=survival_palette)
# ### 2E. Check for Correlations
#
# When 2 features or more have correlation, that means they are explaining each others while giving only a few or none of new information. Try to imagine if TitleGroup feature only has 2 classes, 'Mr.' and 'Miss.'. We can be sure that all male data would have Mr. title and all female have Miss. title. Features with correlation would lead to overfitting on machine learning model, which might result in high accuracy on training dataset while decrease accuracy on test dataset.
# #### 2Ea. Fill null values and creting categories
# +
fare_bins = [0,10,30,60,999999]
fare_labels = ['10-','10-30','30-60','60+']
# fill nulls with meadian
train_set["Fare"].fillna(train_set["Fare"].median(), inplace=True)
# creating categories
train_set['FareRange'] = pd.cut(train_set['Fare'], bins=fare_bins, labels=fare_labels, include_lowest=True)
# +
age_bins = [0, 15, 30, 45, 60, 75, max_age+10]
age_labels = ['1.[0, 15)','2.[15, 30)','3.[30, 45)','4.[45, 60)','5.[60, 75)', f'6.[75, {max_age+10})']
# fill nulls with meadian
train_set["Age"].fillna(train_set["Age"].median(), inplace=True)
# creating categories
train_set['AgeRange'] = pd.cut(train_set['Age'], bins=age_bins, labels=age_labels, include_lowest=True)
# -
train_set["Embarked"].fillna(train_set["Embarked"].value_counts().index[0], inplace=True)
# checking nulls
train_set.isnull().sum()
# Drop no category features
train_analysis = train_set.drop(['PassengerId','Name','Age','Fare','Ticket','Cabin','SibSp','Parch'], axis=1)
train_analysis
# #### 2Ea. Encoding Data for Analysis
# Many methods of Python libraries don't accept text input, so I need to encode categorical features into ordinal numbers. I encode them manually, because I want to assign the label's order.
# +
sex_encode = {'male': 0, 'female': 1}
train_analysis['Sex'] = train_analysis['Sex'].map(sex_encode).astype(int)
embarked_encode = {'C': 0, 'Q': 1, 'S': 2}
train_analysis['Embarked'] = train_analysis['Embarked'].map(embarked_encode).astype(int)
agerange_encode = dict(zip(age_labels, list(range(len(age_labels)))))
train_analysis['AgeRange'] = train_analysis['AgeRange'].map(agerange_encode).astype(int)
farerange_encode = dict(zip(fare_labels, list(range(len(fare_labels)))))
train_analysis['FareRange'] = train_analysis['FareRange'].map(farerange_encode).astype(int)
# -
train_analysis
# #### 2Eb. Computing Correlations
triang_mask = np.zeros((train_analysis.shape[1], train_analysis.shape[1]))
triang_mask[np.triu_indices_from(triang_mask)] = True
colormap = plt.cm.viridis
plt.figure(figsize=(12,12))
plt.title('Correlation between Features', y=1.05, size = 15)
sns.heatmap(train_analysis.corr(),
linewidths=0.1,
vmax=1.0,
square=True,
cmap=colormap,
linecolor='white',
annot=True,
mask=triang_mask)
| kaggle/EDA-finding-important-factors-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Frozen set is just an immutable version of a
# Python set object. While elements of a set can
# be modified at any time, elements of frozen set
# remains the same after creation.
# +
# The syntax of frozenset() method is:
# frozenset([iterable])
# -
s1 = {5, 10, 15, 20, 25}
frozenset(s1)
type(s1)
s1.add(23)
s1
s2 = frozenset(s1)
s2.add(36)
s2
| 10. Python Set/7. Python Frozenset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # importing library
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as matplotlib
import numpy as np
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
import seaborn as sns
sns.set()
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.impute import SimpleImputer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import cross_val_score
from scipy.stats import chisquare
from sklearn.feature_selection import RFE
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix, roc_auc_score, roc_curve, auc
# # Dataset Load
# +
#from google.colab import files
#uploaded = files.upload()
#for fn in uploaded.keys():
# print('User uploaded file "{name}" with length {length} bytes'.format(
# name=fn, length=len(uploaded[fn])))
matplotlib.style.use('classic')
data_frame = pd.read_csv('pima-data.csv')
data_frame.shape
# -
data_frame.describe().T
data_frame.head(3)
data_frame.tail(4)
# # checking for null data
data_frame.isnull()
data_frame.isnull().values
data_frame.isnull().values.any()
# +
#print(data_frame["diabetes"].value_counts())
#(data_frame["diabetes"].value_counts() * 100) / 768 #total row 768
num_true = len(data_frame.loc[data_frame['diabetes'] == True])
num_false = len(data_frame.loc[data_frame['diabetes'] == False])
print ("Number of True Cases: {0} ({1:2.2f}%)".format(num_true, (num_true / (num_true + num_false)) * 100))
print ("Number of False Cases: {0} ({1:2.2f}%)".format(num_false, (num_true / (num_true + num_false)) * 100))
# -
data_frame["diabetes"].value_counts().plot.bar()
# # Age Distribution
print("Min Age: ", data_frame["age"].min())
print("Max Age: ", data_frame["age"].max())
facet = sns.FacetGrid(data_frame, hue = "diabetes", aspect = 3)
facet.map(sns.kdeplot,"age",shade= True)
facet.set(xlim=(0, data_frame["age"].max()))
facet.add_legend()
plt.figure(figsize=(20, 6))
sns.countplot(data = data_frame, x = "age", hue = "diabetes")
# # glucose_conc analysis
facet = sns.FacetGrid(data_frame, hue = "diabetes", aspect = 3)
facet.map(sns.kdeplot,"glucose_conc",shade= True)
facet.set(xlim=(0, data_frame["glucose_conc"].max()))
facet.add_legend()
plt.figure(figsize=(20, 6))
sns.countplot(data = data_frame, x = "glucose_conc", hue = "diabetes")
plt.figure(figsize=(10, 8))
sns.swarmplot(x = 'glucose_conc',y = "age", hue="diabetes", data = data_frame)
# # HeatMap generation
plt.figure(figsize=(18, 12))
sns.heatmap(data_frame.corr(), cmap="RdYlBu", annot=True, fmt=".1f")
# # Deleted highly correlated column
# Deleting 'thikness' column completely
del data_frame['thickness']
# Checking if the action was successful or not
data_frame.head()
plt.figure(figsize=(20, 6))
sns.boxplot(data = data_frame)
plt.xticks(rotation=90)
plt.show()
# # Level Encoding - Datatype changing
# +
# Mapping the values
map_diabetes = {True : 1, False : 0}
# Setting the map to the data_frame
data_frame['diabetes'] = data_frame['diabetes'].map(map_diabetes)
# Let's see what we have done
data_frame.head()
# -
# # Chi-Square Test
result = pd.DataFrame(columns=["Features", "Chi2Weights"])
for i in range(len(data_frame.columns)):
chi2, p = chisquare(data_frame[data_frame.columns[i]])
result = result.append([pd.Series([data_frame.columns[i], chi2], index = result.columns)], ignore_index=True)
pd.set_option("max_column", None)
result = result.sort_values(by="Chi2Weights", ascending=False)
result
print(format(result.Chi2Weights), '.0f')
# # Training, Test Data Preparing
# +
feature_column_names = ['num_preg', 'glucose_conc', 'diastolic_bp', 'insulin', 'bmi', 'diab_pred', 'age', 'skin']
predicted_class_name = ['diabetes']
# Getting feature variable values
X = data_frame[feature_column_names].values
y = data_frame[predicted_class_name].values
# Saving 30% for testing
split_test_size = 0.30
# Splitting using scikit-learn train_test_split function
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = split_test_size, random_state = 42)
# -
print("{0:0.2f}% in training set".format((len(X_train)/len(data_frame.index)) * 100))
print("{0:0.2f}% in test set".format((len(X_test)/len(data_frame.index)) * 100))
#how many column have 0 value
print("# rows in dataframe {0}".format(len(data_frame)))
print("# rows missing glucose_conc: {0}".format(len(data_frame.loc[data_frame['glucose_conc'] == 0])))
print("# rows missing diastolic_bp: {0}".format(len(data_frame.loc[data_frame['diastolic_bp'] == 0])))
print("# rows missing insulin: {0}".format(len(data_frame.loc[data_frame['insulin'] == 0])))
print("# rows missing bmi: {0}".format(len(data_frame.loc[data_frame['bmi'] == 0])))
print("# rows missing diab_pred: {0}".format(len(data_frame.loc[data_frame['diab_pred'] == 0])))
print("# rows missing age: {0}".format(len(data_frame.loc[data_frame['age'] == 0])))
print("# rows missing thickness: {0}".format(len(data_frame.loc[data_frame['skin'] == 0])))
#Impute with mean all 0 readings
fill_0 = SimpleImputer(missing_values=0, strategy="mean")
X_train = fill_0.fit_transform(X_train)
X_test = fill_0.fit_transform(X_test)
# # Create and Train Model (GaussianNB)
# create GaussianNBr model object and train it with the data
from sklearn.naive_bayes import GaussianNB
nb_model= GaussianNB()
nb_model.fit(X_train, y_train.ravel()) # ravel() return 1-D array
# # Performance on Training data
# +
# performance metrics library
from sklearn import metrics
# get current accuracy of the model
prediction_from_trained_data = nb_model.predict(X_train)
accuracy = metrics.accuracy_score(y_train, prediction_from_trained_data)
print ("Accuracy of our GaussianNB model is : {0:.4f}".format(accuracy))
# -
# # Performance on Testing Data
# +
# this returns array of predicted results from test_data
prediction_from_test_data = nb_model.predict(X_test)
accuracy = metrics.accuracy_score(y_test, prediction_from_test_data)
print ("Accuracy of our GaussianNB model is: {0:0.4f}".format(accuracy))
# -
# # Confusion Matrix
# +
print ("Accuracy of our GaussianNB model is: {0:0.4f} %".format(accuracy))
print ("Confusion Matrix")
print ("{0}".format(metrics.confusion_matrix(y_test, prediction_from_test_data, labels=[1, 0])))
print ("Classification Report")
# labels for set 1=True to upper left and 0 = False to lower right
print ("{0}".format(metrics.classification_report(y_test, prediction_from_test_data, labels=[1, 0])))
# -
# # Random Forest
# +
from sklearn.ensemble import RandomForestClassifier
rf_model = RandomForestClassifier(random_state=42)
rf_model.fit(X_train, y_train.ravel())
# -
# # Performance Training
# +
rf_predict_train = rf_model.predict(X_train)
rf_accuracy = metrics.accuracy_score(y_train, rf_predict_train)
print ("Accuracy: {0:.4f}".format(rf_accuracy))
# -
# # Performance Testing
# +
rf_predict_test = rf_model.predict(X_test)
rf_accuracy_testdata = metrics.accuracy_score(y_test, rf_predict_test)
print ("Accuracy: {0:.4f}".format(rf_accuracy_testdata))
# -
print ("Confusion Matrix for Random Forest")
print ("{0}".format(metrics.confusion_matrix(y_test, rf_predict_test, labels=[1, 0])))
print ("")
print ("Classification Report\n")
print ("{0}".format(metrics.classification_report(y_test, rf_predict_test, labels=[1, 0])))
# # KNN Model Classification
knn_model = KNeighborsClassifier(n_neighbors=5)
knn_model.fit(X_train, y_train.ravel())
knn_predict_train = knn_model.predict(X_train)
knn_accuracy = metrics.accuracy_score(y_train, knn_predict_train)
print ("Accuracy: {0:.4f}".format(knn_accuracy))
# +
knn_predict_test = knn_model.predict(X_test)
knn_accuracy_testdata = metrics.accuracy_score(y_test, knn_predict_test)
print ("Accuracy: {0:.4f}".format(knn_accuracy_testdata))
print ("Confusion Matrix for K-Neighbors Classifier")
# labels for set 1=True to upper left and 0 = False to lower right
print ("{0}".format(metrics.confusion_matrix(y_test, knn_predict_test, labels=[1, 0])))
print ("")
print ("Classification Report\n")
print ("{0}".format(metrics.classification_report(y_test, knn_predict_test, labels=[1, 0])))
# -
# Finding the best value of K in KNN using N-Fold Cross Validation
k_range=np.arange(1,31,1)
k_score=[]
best_accuracy_score_knn=0
best_k=0
for k in k_range:
score=cross_val_score(KNeighborsClassifier(n_neighbors=k), X, y.ravel(), cv=10, scoring='accuracy').mean()
k_score.append(score)
if best_accuracy_score_knn<score:
best_accuracy_score_knn=score
best_k=k
knn_plot=plt
knn_plot.plot(k_range, k_score)
knn_plot.show()
print(f"KNN with N-Fold Cross Validation is giving best score as {best_accuracy_score_knn*100}% for K={best_k}")
# # Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
lr_model = LogisticRegression(penalty='l1',dual=False,max_iter=110, solver='liblinear')
lr_model.fit(X_train, y_train.ravel())
# +
lr_predict_test = lr_model.predict(X_test)
lr_accuracy_testdata = metrics.accuracy_score(y_test, rf_predict_test)
print ("Accuracy: {0:.4f}".format(lr_accuracy_testdata))
print ("Confusion Matrix for Logistic Regressiont")
print ("{0}".format(metrics.confusion_matrix(y_test, lr_predict_test, labels=[1, 0])))
print ("")
print ("Classification Report\n")
print ("{0}".format(metrics.classification_report(y_test, lr_predict_test, labels=[1, 0])))
# -
# # ROC Curve
# +
FPR, TPR, thresholds = roc_curve(y_test, lr_predict_test)
plt.figure(figsize=(10,5)) # figsize in inches
plt.plot(FPR, TPR)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.plot([0, 1], [0, 1], color='navy', lw=2, linestyle='--') # 50%
plt.plot(FPR, TPR, lw=2, label='Logaristic Regression (AUC = %0.2f)' % auc(FPR, TPR))
plt.title('ROC curve for diabetes classifier')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
plt.legend(loc="lower right")
# -
# # N-Fold Cross-validation
# +
from sklearn.linear_model import LogisticRegressionCV
lr_cv_model = LogisticRegressionCV(n_jobs=-1, random_state=42, Cs=3, cv=10, refit=False, class_weight="balanced")
lr_cv_model.fit(X_train, y_train.ravel())
lr_cv_predict_test = lr_cv_model.predict(X_test)
print ("Accuracy: {0:.4f}".format(metrics.accuracy_score(y_test, lr_cv_predict_test)))
print (metrics.confusion_matrix(y_test, lr_cv_predict_test, labels=[1, 0]))
print (" ")
print ("Classification Report")
print (metrics.classification_report(y_test, lr_cv_predict_test, labels=[1,0]))
# -
# # C (Regularization Hyperparameter)
# +
C_start = 0.1
C_end = 5
C_inc = 0.1
C_values, recall_scores = [], []
C_val = C_start
best_recall_score = 0
while (C_val < C_end):
C_values.append(C_val)
lr_model_loop = LogisticRegression(C=C_val, random_state=42)
lr_model_loop.fit(X_train, y_train.ravel())
lr_predict_loop_test = lr_model_loop.predict(X_test)
recall_score = metrics.recall_score(y_test, lr_predict_loop_test)
recall_scores.append(recall_score)
if (recall_score > best_recall_score):
best_recall_score = recall_score
best_lr_predict_test = lr_predict_loop_test
C_val = C_val + C_inc
best_score_C_val = C_values[recall_scores.index(best_recall_score)]
print ("1st max value of best_recall_score: {0:.3f} occured at C={1:.3f}".format(best_recall_score, best_score_C_val))
plt.plot(C_values, recall_scores, "-")
plt.xlabel("C value")
plt.ylabel("recall score")
# -
# # class_weight = 'balanced'
# +
C_start = 0.1
C_end = 5
C_inc = 0.1
C_values, recall_scores = [], []
C_val = C_start
best_recall_score = 0
while (C_val < C_end):
C_values.append(C_val)
lr_model_loop = LogisticRegression(C=C_val, class_weight="balanced", random_state=42)
lr_model_loop.fit(X_train, y_train.ravel())
lr_predict_loop_test = lr_model_loop.predict(X_test)
recall_score = metrics.recall_score(y_test, lr_predict_loop_test)
recall_scores.append(recall_score)
if (recall_score > best_recall_score):
best_recall_score = recall_score
best_lr_predict_test = lr_predict_loop_test
C_val = C_val + C_inc
best_score_C_val = C_values[recall_scores.index(best_recall_score)]
print ("1st max value of {0:.3f} occured at C={1:.3f}".format(best_recall_score, best_score_C_val))
# %matplotlib inline
plt.plot(C_values, recall_scores, "-")
plt.xlabel("C value")
plt.ylabel("recall score")
| pima-diabetes-dataset-machine-learning-approach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function
import sys
import os
import numpy as np
import torch
from edgeml_pytorch.graph.protoNN import ProtoNN
from edgeml_pytorch.trainer.protoNNTrainer import ProtoNNTrainer
import edgeml_pytorch.utils as utils
import helpermethods as helper
# -
# ## USPS Data
# It is assumed that the USPS data has already been downloaded and set up with the help of `fetch_usps.py` and is placed in the `./usps10` subdirectory.
# +
# Load data
DATA_DIR = './usps10'
train, test = np.load(DATA_DIR + '/train.npy'), np.load(DATA_DIR + '/test.npy')
x_train, y_train = train[:, 1:], train[:, 0]
x_test, y_test = test[:, 1:], test[:, 0]
numClasses = max(y_train) - min(y_train) + 1
numClasses = max(numClasses, max(y_test) - min(y_test) + 1)
numClasses = int(numClasses)
y_train = helper.to_onehot(y_train, numClasses)
y_test = helper.to_onehot(y_test, numClasses)
dataDimension = x_train.shape[1]
numClasses = y_train.shape[1]
# -
# ## Model Parameters
#
# Note that ProtoNN is very sensitive to the value of the hyperparameter $\gamma$, here stored in valiable GAMMA. If GAMMA is set to None, median heuristic will be used to estimate a good value of $\gamma$ through the helper.getGamma() method. This method also returns the corresponding W and B matrices which should be used to initialize ProtoNN (as is done here).
PROJECTION_DIM = 60
NUM_PROTOTYPES = 60
REG_W = 0.000005
REG_B = 0.0
REG_Z = 0.00005
SPAR_W = 0.8
SPAR_B = 1.0
SPAR_Z = 1.0
LEARNING_RATE = 0.05
NUM_EPOCHS = 200
BATCH_SIZE = 32
GAMMA = 0.0015
W, B, gamma = helper.getGamma(GAMMA, PROJECTION_DIM, dataDimension,
NUM_PROTOTYPES, x_train)
protoNNObj = ProtoNN(dataDimension, PROJECTION_DIM, NUM_PROTOTYPES, numClasses,
gamma, W=W, B=B)
protoNNTrainer = ProtoNNTrainer(protoNNObj, REG_W, REG_B, REG_Z, SPAR_W, SPAR_B, SPAR_W,
LEARNING_RATE, lossType='xentropy')
protoNNTrainer.train(BATCH_SIZE, NUM_EPOCHS, x_train, x_test, y_train, y_test, printStep=600, valStep=10)
# ## Evaluation
x_, y_= torch.Tensor(x_test), torch.Tensor(y_test)
logits = protoNNObj.forward(x_)
_, predictions = torch.max(logits, dim=1)
_, target = torch.max(y_, dim=1)
acc, count = protoNNTrainer.accuracy(predictions, target)
W, B, Z, gamma = protoNNObj.getModelMatrices()
matrixList = [W, B, Z]
matrixList = [x.detach().numpy() for x in matrixList]
sparcityList = [SPAR_W, SPAR_B, SPAR_Z]
nnz, size, sparse = helper.getModelSize(matrixList, sparcityList)
print("Final test accuracy", acc)
print("Model size constraint (Bytes): ", size)
print("Number of non-zeros: ", nnz)
nnz, size, sparse = helper.getModelSize(matrixList, sparcityList,
expected=False)
print("Actual model size: ", size)
print("Actual non-zeros: ", nnz)
| examples/pytorch/ProtoNN/protoNN_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="B_OZ25dzc5kT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 158} outputId="7f9208c1-fdb1-41d9-d753-191c152a30a4"
# !pip install downcast==0.0.5
# + id="RVEE9jgJKOtT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="7ef84a84-addd-4209-d318-5bc998774367"
import os
import pandas as pd
import numpy as np
import plotly_express as px
import plotly.graph_objects as go
from plotly.subplots import make_subplots
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
import joblib
# + [markdown] id="K2zAdzrHjs4d" colab_type="text"
# https://www.kaggle.com/stytch16/jena-climate-2009-2016
# + id="b6BeO-ilI-C3" colab_type="code" colab={}
jena = pd.read_csv("/content/jena_climate_2009_2016.csv")
# + id="IrH5niIIKkyw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 339} outputId="d3fd154d-4a9e-4575-a3cc-04f428fd9641"
jena.head()
# + id="twumyARMIBmk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2f97070c-50da-4961-a3b9-2d9f1956894f"
jena_bd = np.round(jena.memory_usage().sum()/(1024*1024),1)
print(jena_bd)
# + id="gHrGBxJmLS5_" colab_type="code" colab={}
from downcast import reduce
jena = reduce(jena)
# + id="CW_9dS0hLp6l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="16b840da-a4ae-45b8-936d-9a348a027b5b"
jena_ad = np.round(jena.memory_usage().sum()/(1024*1024),1)
jena_ad
# + id="UssSRz9aIFUl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="708d65ac-ec25-4f75-c0cf-41ccdb24a62b"
dic = {'DataFrame':['jena'],
'Before downcasting':[jena_bd],
'After downcasting':[jena_ad]}
memory = pd.DataFrame(dic)
memory = pd.melt(memory, id_vars='DataFrame', var_name='Status', value_name='Memory (MB)')
memory.sort_values('Memory (MB)',inplace=True)
fig = px.bar(memory, x='DataFrame', y='Memory (MB)', color='Status', barmode='group', text='Memory (MB)')
fig.update_traces(texttemplate='%{text} MB', textposition='outside')
fig.update_layout(template='seaborn', title='Effect of Downcasting')
fig.show()
| examples/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Comcast is an American global telecommunication company. The firm has been providing terrible customer service. They continue to fall short despite repeated promises to improve. Only last month (October 2016) the authority fined them a $2.3 million, after receiving over 1000 consumer complaints.
# The existing database will serve as a repository of public customer complaints filed against Comcast.
# It will help to pin down what is wrong with Comcast's customer service.
# Data Dictionary
# • Ticket #: Ticket number assigned to each complaint
# • Customer Complaint: Description of complaint
# • Date: Date of complaint
# • Time: Time of complaint
# • Received Via: Mode of communication of the complaint
# • City: Customer city
# • State: Customer state
# • Zipcode: Customer zip
# • Status: Status of complaint
# • Filing on behalf of someone
#
# Analysis Task
# To perform these tasks, you can use any of the different Python libraries such as NumPy, SciPy, Pandas, scikit-learn, matplotlib, and BeautifulSoup.
# - Import data into Python environment.
# - Provide the trend chart for the number of complaints at monthly and daily granularity levels.
# - Provide a table with the frequency of complaint types.
# • Which complaint types are maximum i.e., around internet, network issues, or across any other domains.
# - Create a new categorical variable with value as Open and Closed. Open & Pending is to be categorized as Open and Closed & Solved is to be categorized as Closed.
# - Provide state wise status of complaints in a stacked bar chart. Use the categorized variable from Q3. Provide insights on:
# • Which state has the maximum complaints
# • Which state has the highest percentage of unresolved complaints
# - Provide the percentage of complaints resolved till date, which were received through the Internet and customer care calls.
comcast = pd.read_csv(r"C:\Users\Vaibhav\Desktop\BA\Datasets\Comcast_telecom_complaints_data.csv")
comcast.head(3)
comcast.isnull().sum()
comcast["Date"] = pd.to_datetime(comcast.Date, format = "%d-%m-%y")
plt.figure(figsize = (20,5))
comcast.groupby("Date").count()["Ticket #"].plot(kind = "line")
# +
#comcast.plot()
# -
# #### To get the granularity level of month
comcast["Month"] = comcast["Date"].apply(lambda x: x.month)
plt.figure(figsize = (10,5))
comcast.groupby("Month").count()["Ticket #"].plot(kind = "line")
# #### Provide a table with the frequency of complaint types.
comcast["Customer Complaint"].value_counts() #complaint types: comcast service, internet, data, billing
# #### Q. Which complaint types are maximum i.e., around internet, network issues, or across any other domains.
comcast.head(3)
import re
comcast["Complaint_Type_Internet"] = [re.search("([Ii]nternet)", comcast["Customer Complaint"][i])!=None for i in range(comcast.shape[0])]
comcast["Complaint_Type_Data"] = [re.search("([Dd]ata)", comcast["Customer Complaint"][i])!=None for i in range(comcast.shape[0])]
comcast["Complaint_Type_Billing"] = [re.search("[Bb]illing", comcast["Customer Complaint"][i])!=None for i in range(comcast.shape[0])]
re.search("([cC][oO][Mm][cC][Aa][Ss][Tt])$|([Ss]ervice)", comcast["Customer Complaint"][2])
comcast["Complaint_Type_Service"] = [re.search("([cC][oO][Mm][cC][Aa][Ss][Tt])$|([Ss]ervice)", comcast["Customer Complaint"][i])!=None for i in range(comcast.shape[0])]
comcast[["Complaint_Type_Internet", "Complaint_Type_Data", "Complaint_Type_Billing", "Complaint_Type_Service"]].sum()
# +
#Max complaints are of the type service
# -
# #### Create a new categorical variable with value as Open and Closed. Open & Pending is to be categorized as Open and Closed & Solved is to be categorized as Closed.
pd.value_counts(comcast.Status, dropna=False)
comcast["Status"] = comcast["Status"].replace({"Pending":"Open",
"Solved": "Closed"})
pd.value_counts(comcast.Status, dropna=False)
# Provide state wise status of complaints in a stacked bar chart. Use the categorized variable from Q3. Provide insights on:
#
# • Which state has the maximum complaints
#
# • Which state has the highest percentage of unresolved complaints
plt.figure(figsize=(15,5))
pd.value_counts(comcast.State).plot(kind = "bar")
pd.crosstab(comcast.State, comcast.Status).plot(kind = "bar", figsize = (15,5) )
plt.show()
# #### - Provide the percentage of complaints resolved till date, which were received through the Internet and customer care calls.
comcast[comcast["Received Via"].isin(["Internet", "Customer Care"])].head()
comcast[comcast["Received Via"].isin(["Internet", "Customer Care"])].groupby("Status").count()["Ticket #"]/sum(comcast[comcast["Received Via"].isin(["Internet", "Customer Care"])].groupby("Status").count()["Ticket #"])
| California Housing Price Prediction/California Housing Price Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ! pip install pyspark
# ! pip install findspark
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.appName("Semi_Sructured_Data_Analysis") \
.config("spark.some.config.option", "some-value") \
.getOrCreate()
# ### Reading CSV
#
# ! wget 'https://covid.ourworldindata.org/data/owid-covid-data.csv'
df = spark.read.csv('data/owid-covid-data.csv', sep= ',', header= True)
df.count()
df.select("continent", "location", "date",
"total_cases", "total_deaths",
"total_cases_per_million",
"population"
).show(5)
df.createOrReplaceTempView("COVID")
covid_data.columns
spark.sql("SHOW COLUMNS FROM COVID")
spark.sql("DESCRIBE COVID")
df.createOrReplaceTempView("COVID")
covid_data= spark.sql("SELECT location AS country, date, total_cases,\
total_cases_per_million, population \
FROM COVID WHERE location ='India'")
covid_data.show(5)
# Window Function SQL: Think of a use case
# ### Reading Tab Separated Values (TSV)
# ! wget 'https://github.com/thepanacealab/covid19_twitter/blob/master/dailies/2021-04-19/2021-04-19_clean-dataset.tsv.gz'
# ! gzip 'data/2021-04-19_clean-dataset.tsv.gz'
# +
# Some other TSV files to play
# #! wget 'https://datasets.imdbws.com/title.ratings.tsv.gz' .
# #! wget 'https://datasets.imdbws.com/title.ratings.tsv.gz' .
# #! gunzip 'data/title.basics.tsv.gz'
# #! gunzip 'data/title.ratings.tsv.gz'
# -
df = spark.read.csv("data/2021-04-19-dataset.tsv", header= True, sep= '\t')
df.count()
df.printSchema()
df.show(4)
# ### Reading JSON
#
#
# ! wget "https://data.cdc.gov/resource/3apk-4u4f.json"
# ### Binary Files
spark.read.format("binaryFile").option("pathGlobFilter", "*.png").load("/data")
#
# ##### Number of Delays:
# 1. **Carrier**: The number of delays and cancellations due to circumstances within the airline's control (e.g. maintenance or crew problems, aircraft cleaning, baggage loading, fueling, etc.) in this month.
# 2. **Late Aircraft**: The number of delays and cancellations caused by a previous flight with the same aircraft arriving late, causing the present flight to depart late in this month.
# 3. **National Aviation System**: The number of delays and cancellations attributable to the national aviation system that refer to a broad set of conditions, such as non-extreme weather conditions, airport operations, heavy traffic volume, and air traffic control in this month.
# 4. **Security**: Number of delays or cancellations caused by evacuation of a terminal or concourse, re-boarding of aircraft because of security breach, inoperative screening equipment and/or long lines in excess of 29 minutes at screening areas in this month.
# 5. **Weather**: Number of delays or cancellations caused by significant meteorological conditions (actual or forecasted) that, in the judgment of the carrier, delays or prevents the operation of a flight such as tornado, blizzard or hurricane in this month.
# +
import requests, json
url = 'https://think.cs.vt.edu/corgis/datasets/json/airlines/airlines.json'
resp = requests.get(url=url)
json_string = json.dumps(resp.json())
json_data= json.loads(json_string)
with open('airlines.json', 'w') as json_file:
json.dump(json_data, json_file)
# -
df = spark.read.json("airlines.json")
df.show(6)
df= spark.read.option("multiline","true").json("airlines.json")
#df.show(6)
# Understanding the schema of the JSON through the dataframe
df.printSchema()
# +
# Subsetting the dataframe to analyse the statistics of # (number) of delays
airport_delay_statistics = df.select("Airport.Code", "Airport.Name",
"Statistics.# of Delays.Carrier",
"Statistics.# of Delays.Late Aircraft",
"Statistics.# of Delays.Security",
"Statistics.# of Delays.Weather")
airport_delay_statistics.printSchema()
# -
# Check which input file has created this dataframe
airport_delay_statistics.inputFiles()
# +
# Creating summary statistics of the dataframe attributes
airport_delay_statistics.select("Carrier", "Late Aircraft",
"Security","Weather").describe().show()
# -
# Sorting the airport delay statistics based on the Weather
airport_delay_statistics.select("Code", "Name", "Weather").sort("Weather").show(10)
# +
# Sorting the airport delay statistics based on the Weather
airport_delay_statistics.select("Code", "Name", "Weather").sort("Weather").show(10, truncate= False)
# -
airport_delay_statistics.count()
# +
# Find the list of most affected airports due to delays caused by weather
from pyspark.sql.functions import col, sum, count
airport_delay_statistics.select("Code", "Name", "Weather").groupBy("Code").sum("Weather").show(3)#.sort(col("Weather").desc()).show(5, truncate=False)
# +
# Using groupby to find the list of least affected unique airports
from pyspark.sql.functions import col, sum, count
airport_delay_statistics.select("Code", "Name", "Weather") \
.groupBy("Code", "Name").agg(sum("Weather") \
.alias("Weather")).sort(col("Weather")).show(10, truncate=False)
# +
# Using groupby to find the list of most affected unique airports
from pyspark.sql.functions import col, sum, count
airport_delay_statistics.select("Code", "Name", "Weather") \
.groupBy("Code", "Name").agg(sum("Weather") \
.alias("Weather")).sort(col("Weather").desc()).show(10, truncate=False)
# +
# Find the list of most affected airports due to delays caused by security
airport_delay_statistics.select("Code", "Name", "Security") \
.groupBy("Code", "Name").agg(sum("Security") \
.alias("Security")).sort(col("Security").desc()).show(10, truncate=False)
# -
airport_delay_statistics.columns
airport_delay_statistics.dtypes
airport_delay_statistics.cov("Carrier", "Weather")
# Creating a summary statistics of the dataframe
airport_delay_statistics.describe().show()
# Creating a summary statistics by without columns having NAs
airport_delay_statistics.describe().dropna().show()
# Creating a summary statistics by without columns having NAs
airport_delay_statistics.describe().drop("Code", "Name").show()
# ### Filtering rows
#
# * Discuss more about the various logical operations that can take place apart from OR such as AND, >, < etc
# * discuss about various programming methods to implement the same operation
# * explain how to extract the first row using FIRST
# * explain how to extract the first N rows using HEAD
# *
# Find the list of most affected airports due to delays caused by security
# Filtering rows using a WHERE clause
airport_statistics.select("Code", "Name", "Security") \
.groupBy("Code", "Name").agg(sum("Security") \
.alias("Security")).sort(col("Security").desc()) \
.where((col("Code")== "ATL") | (col("Code")== "TPA")) \
.show(10, truncate=False)
# Find the list of most affected airports due to delays caused by security
# Filtering rows using a FILTER clause
airport_statistics.select("Code", "Name", "Security") \
.groupBy("Code", "Name").agg(sum("Security") \
.alias("Security")).sort(col("Security").desc()) \
.filter((col("Code")== "ATL") | (col("Code")== "TPA")) \
.show(10, truncate=False)
# +
# Find the list of most affected airports due to delays caused by security
# Filtering rows using a WHERE clause
condition_1= (col("Code")== "ATL")
condition_2= (col("Code")== "TPA")
airport_statistics.select("Code", "Name", "Security") \
.groupBy("Code", "Name").agg(sum("Security") \
.alias("Security")).sort(col("Security").desc()) \
.where(condition_1 | condition_2) \
.show(10, truncate=False)
# -
airport_delay_statistics.isStreaming
airport_delay_statistics.na
airport_delay_statistics.persist
split_1, split_2= airport_delay_statistics.randomSplit([1.0, 2.0], 24)
split_1.show(3)
split_2.show(3)
# +
# df.replace
# df.repartition, df.repartitionByRange
# df.registerTempTable
# df.rdd
# df.intersect
# df.intersectAll
# df.collect vs df.take vs df.show
# df.hint()
# using explode feature
# createGlobalTempView
# createOrReplaceGlobalTempView
# distinct
# drop_duplicates
# exceptALL
# fillna
#
# cube function is most important
| Lecture-5B.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ijohns29/Comparing-regional-Covid-19-data-trends-in-MD-and-MT-to-assess-short-term-risks-of-travel/blob/main/2020_11_17_MD_MT_Covid_19_data_IJ.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vtQNlIVNaPan"
# ## Looking at recent Covid-19 trends in Maryland and Montana at state level and at more granular level using multiple open data sources
# + [markdown] id="cWMVXlEfa2tS"
#
#
# * Change in daily incidence over past two weeks
# * Change in hospitalizations over past two weeks
# * Change in positivity rate as function of age group for MD
#
#
# + [markdown] id="XJA7uKmOc5iJ"
# # Import Libraries
# + colab={"base_uri": "https://localhost:8080/"} id="xd4wOUG5-qXE" outputId="6375922b-62c5-4e15-fcf6-a9c7fb6a8931"
# Install visualization package
# !pip install plotly
# + [markdown] id="bPVPM_Rs6WRh"
#
# + id="xKn3fMt3YS7O"
# import libraries
# for data analysis: 1 is for 'pandas' i.e. panel data
import pandas as pd # imports panel data. Later in code can just say use pd rather than use pandas
import numpy as np
# for visualization
import plotly.express as px
# to help export files
from google.colab import files
# + [markdown] id="LPAV5g70vDJl"
# # Import data
# + id="rPiLiMIKu4CW"
# import data e.g. bring in csv. Can't open them here but can take it and project it into notebook. This projection is data frame (df)_relevant_name
# One equals is to define variable , two is for equation
# Convention is df = pd.read_csv("github link")
# Creates data frame using pandas (prev imported as pd). Read_csv is code under pandas that allows reading of csv file
# Import NYT Covid-19 state level data - 2020-11-17 is the name but autoupdates to latest upon initialization
df_NYT_Covid_19_states_20201127 = pd.read_csv("https://raw.githubusercontent.com/nytimes/covid-19-data/master/us-states.csv")
# Import NYT Covid-19 county level data - 2020-11-17 is the name but auto updates to latest upon inititalization
df_NYT_Covid_19_counties_20201127 = pd.read_csv("https://raw.githubusercontent.com/nytimes/covid-19-data/master/us-counties.csv")
# Import Maryland.gov Covid-19 testing data
df_MD_HHS = pd.read_csv("https://opendata.maryland.gov/resource/avum-x5ua.csv")
# + [markdown] id="eMi1gUI7NVwU"
# # Preview data
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="0_9mcIGi7EvW" outputId="29671ec1-1b76-454d-86b2-b98d10f69f8b"
# preview data - do this by calling data frame.head(#) If no number default is 5. n = number of rows of data can preview
df_NYT_Covid_19_states_20201127.head(6)
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="EVtw0JuhNoOj" outputId="2b9ffb37-e175-418b-82df-a74c089b1b7b"
# Preview Covid-19 County data
df_NYT_Covid_19_counties_20201127.head(6)
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="AFw50Zo6JMIE" outputId="c6d6e7b9-a3b0-4de4-ab8d-148526fcabe3"
# preview end of county data
df_NYT_Covid_19_counties_20201127.tail(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="yLMMkUfVJMoy" outputId="e16b9ca0-8eb6-4814-9b64-f099745cbfc0"
# preview end of state level data
df_NYT_Covid_19_states_20201127.tail(6)
# + [markdown] id="tcDKY-CyTSHY"
# # Maryland HHS state data
# ## Cleaning up data and creating appropriate dataframe for it
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="xlXc6KBWN843" outputId="20a33e36-7a3f-446a-9f7f-87d9a90e793a"
# Preview MD HHS data
df_MD_HHS.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="ryEMeC3KOQMs" outputId="a69e41ae-b82f-435c-ce14-42ab1c814184"
# Preview MD HHS data
df_MD_HHS.tail(10)
# + colab={"base_uri": "https://localhost:8080/"} id="zlAMY64XHvnd" outputId="5e95576e-7564-4cd1-8a30-91105dccac45"
# Create data frame to test code with. df.shape returns number of rows (i) and columns (0)
df_edit_MD_HHS = df_MD_HHS
df_edit_MD_HHS.tail()
df_edit_MD_HHS.shape
# + id="0_H5SkX1IKdZ"
# Creates blank list for split data to be stored in e.g. df_split_MD_HHS
# Uses for loop to look over entire column (date column '1') and splits the data at the T point. [0] after indicates keep the first part of the split
df_split_MD_HHS = []
for i in range(df_edit_MD_HHS.shape[0]):
df_split_MD_HHS.append(df_edit_MD_HHS.iloc[i,1].split('T')[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="YjxcYlRyMjno" outputId="6c0b867f-590e-43aa-eb22-ee85868b6212"
# Converts list to dataframe
dfObj = pd.DataFrame(df_split_MD_HHS, columns = ['date'])
#preview newly created dataframe
dfObj.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="9ArdKbi2ZcMV" outputId="f63fb1f0-e063-4cc2-c473-6a1c9d5b8405"
# Fill report date with split date info from dfObj dataframe
df_MD_HHS["reportdate"] = dfObj["date"]
df_MD_HHS.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="05DyNKj7aoNh" outputId="2d25a95c-050d-4b29-eb66-0d6394ddc3aa"
# Rename column headings so can be merged with other data sets if desired
df_MD_HHS = df_MD_HHS.rename(columns={"reportdate" : "date",
"under35": "% pos. under 35 yrs",
"over35": "% pos. over 35 yrs"})
df_MD_HHS.head()
# + colab={"base_uri": "https://localhost:8080/"} id="aOgXcD4Dbp68" outputId="7db634b5-fe37-4942-f624-06b603c6a897"
df_MD_HHS.info()
# + [markdown] id="A_7XgDkfQdrm"
# # Data Filtering
# + id="SV2Aqo0wAGCy" colab={"base_uri": "https://localhost:8080/"} outputId="9f4fcdd2-a739-487e-f4fc-ef669e36e368"
# Filter Data
# Look at all of the column names in the state data set. This is useful to do prior to creating new dataset so can pull exact name in order to select appropriate column
#Format is df_name.columns
df_NYT_Covid_19_states_20201127.columns
# + colab={"base_uri": "https://localhost:8080/"} id="tcFPXjVmJtpl" outputId="5324879d-b775-435a-8a89-e8130a7bedc0"
# Look at all of the column names in the county level data
df_NYT_Covid_19_counties_20201127.columns
# + colab={"base_uri": "https://localhost:8080/"} id="MLIs3w9iSxbp" outputId="a80167bd-f24a-4109-de66-30c6b1ebcafd"
# Look at all of the column names in the MD HHS data
df_MD_HHS.columns
# + colab={"base_uri": "https://localhost:8080/"} id="hZ94AFN5TUQo" outputId="b3237ecd-2242-40bc-ab69-2ebf1790baa5"
# Look at object types of data set
df_MD_HHS.info()
# + colab={"base_uri": "https://localhost:8080/"} id="RuXvGuAB9z3B" outputId="3bebbfdb-f267-45db-a366-d35a770f552f"
#look at unique values in columns -> useful for grabbing exact value searching for e.g. 'Maryland' with data repository spelling
# Format is df_name["column name of interest"].unique()
df_NYT_Covid_19_states_20201127["state"].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="kTtFb6oNKOCS" outputId="a1dc4f77-e928-467e-88a9-c6ed4cf2e33d"
# look at unique names in columns for county level data
df_NYT_Covid_19_counties_20201127["county"].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="tOppHzs2AsDV" outputId="2cce226f-9f0c-43cb-8034-9f780356331e"
#df_name["column heading of interest"].unique - pulls entire list with repeats as well included, basically full printout of column
df_NYT_Covid_19_states_20201127["state"].unique
# + colab={"base_uri": "https://localhost:8080/"} id="2Jy8yhbpWh8w" outputId="4fb15880-3d26-4b92-ef0a-6f904e320f7a"
# df_name[""].nunique() - gives total number of unique state + territories listed
df_NYT_Covid_19_states_20201127["state"].nunique()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="ivf_RqsCCEA4" outputId="1c2edb70-afb1-4d76-e93b-faf60dae2882"
df_NYT_Covid_19_Maryland_20201127.head(5)
# + colab={"base_uri": "https://localhost:8080/"} id="oLi8C5vVYH5I" outputId="9784f526-541c-4bea-a651-47b94a487efd"
# df_name.info() - gives high level data about data frame that is called e.g. # columns, column name, information types
df_NYT_Covid_19_Maryland_20201127.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="FYjjnOBdYoMz" outputId="3c3574b3-4078-4daf-a062-c63a22a0d7fc"
# df_name.describe() - general descriptive statistics, does not pull data for all columns only those that are coded as numbers e.g. int64
df_NYT_Covid_19_Maryland_20201127.describe()
# + [markdown] id="boWDmRUWh9Ol"
# # Creating new data frames from imported data
# + id="QgxN2mxICMsX"
#Pull out a column from NYT orig dataframe (df) to new dataframe with data repository names verified as above - pulls out Maryland data
df_NYT_Covid_19_Maryland_20201127 = df_NYT_Covid_19_states_20201127[df_NYT_Covid_19_states_20201127["state"]== "Maryland"]
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="kgW7YbQDWS1f" outputId="3261cd17-0066-4af4-81c7-9208b3f17dd4"
#preview new dataframe containing only MD data
df_NYT_Covid_19_Maryland_20201127.head()
# + colab={"base_uri": "https://localhost:8080/"} id="z6_-OKQ5Kzbl" outputId="7c6335fa-7619-4e14-de0c-68abacebfd94"
# Pull out Montana Data from state level data
df_NYT_Covid_19_Montana_20201127 = df_NYT_Covid_19_states_20201127[df_NYT_Covid_19_states_20201127["state"]== "Montana"]
df_NYT_Covid_19_Montana_20201127.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="7Fm3752ILAUd" outputId="50ed7741-04db-485c-a2ca-6ea70c5bde6d"
# preview MT data
df_NYT_Covid_19_Montana_20201127.tail()
# + [markdown] id="-59BtT7r55wW"
# # Merging and editing dataframes
# + id="GpGItPZsIauI"
df_new_index_MT = df_NYT_Covid_19_Montana_20201127.set_index('date')
# + id="fTTsJi4XvzKr"
df_new_index_MD = df_NYT_Covid_19_Maryland_20201127.set_index('date')
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="qPE7iZjbubel" outputId="5eaab5f7-936e-46cb-bc95-af87c32152dd"
df_daily_case_death_MT = df_new_index_MT[{'cases', 'deaths'}].diff()
df_daily_case_death_MT.tail()
# Merge MT data
df_MT_daily_cumulative_merge = pd.merge(df_NYT_Covid_19_Montana_20201127,
df_daily_case_death_MT,
how = "right",
on = "date")
df_MT_daily_cumulative_merge.head()
# Rename columns
df_MT_daily_cumulative_merge = df_MT_daily_cumulative_merge.rename(columns={"state" : "state_MT",
"fips": "fips_MT",
"cases_x": "cases_MT",
"deaths_x": "deaths_MT",
"cases_y": "daily cases_MT",
"deaths_y": "daily deaths_MT"})
df_MT_daily_cumulative_merge['3-day-case-avg_MT'] = df_MT_daily_cumulative_merge.rolling(window=3)['daily cases_MT'].mean()
df_MT_daily_cumulative_merge['3-day-death-avg_MT'] = df_MT_daily_cumulative_merge.rolling(window=3)['daily deaths_MT'].mean()
df_MT_pop_per_100k = [9.89415]
df_MT_daily_cumulative_merge['3-day-case-incidence_MT'] = df_MT_daily_cumulative_merge["3-day-case-avg_MT"]/df_MT_pop_per_100k
df_MT_daily_cumulative_merge['3-day-death-incidence_MT'] = df_MT_daily_cumulative_merge["3-day-death-avg_MT"]/df_MT_pop_per_100k
df_MT_daily_cumulative_merge.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="IO3nP60mwAXj" outputId="2f33ce20-643f-45b1-89ba-e42252689375"
df_daily_case_death_MD = df_new_index_MD[{'cases', 'deaths'}].diff()
df_daily_case_death_MD.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="jpnhWoUHKS0Z" outputId="d160ddeb-1d67-4ecc-866c-fc253735b9a2"
# Calculate 3 day average case load.
df_daily_case_death_MD['3-day-case-avg_MD'] = df_daily_case_death_MD.rolling(window=3)['cases'].mean()
df_daily_case_death_MD.tail()
df_daily_case_death_MD['3-day-death-avg_MD'] = df_daily_case_death_MD.rolling(window=3)['deaths'].mean()
df_daily_case_death_MD.tail()
df_MD_pop_per_100k = [57.73552]
df_daily_case_death_MD['3-day-case-incidence_MD'] = df_daily_case_death_MD["3-day-case-avg_MD"]/df_MD_pop_per_100k
df_daily_case_death_MD['3-day-death-incidence_MD'] = df_daily_case_death_MD["3-day-death-avg_MD"]/df_MD_pop_per_100k
df_daily_case_death_MD.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="ANwHfpeN-Vb4" outputId="6a0cc872-f394-4692-abc2-871174935adb"
# Merge Maryland dataframes
df_MD_HHS_state_merge = pd.merge(df_NYT_Covid_19_Maryland_20201127,
df_daily_case_death_MD,
how = "right",
on = "date")
# Preview merged dataframe
df_MD_HHS_state_merge.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="AMXFj-8gCx6R" outputId="fcd3a213-7e88-4313-9d5e-0b136b3e3325"
# Rename columns of data frame
df_MD_HHS_state_merge = df_MD_HHS_state_merge.rename(columns={"state" : "state_MD",
"fips": "fips_MD",
"cases_x": "cases_MD",
"deaths_x": "deaths_MD",
"cases_y": "daily cases_MD",
"deaths_y": "daily deaths_MD"})
# Preview renamed data
df_MD_HHS_state_merge.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 323} id="W3E4KYkLsrbg" outputId="05b7999e-acd1-484a-b075-29f6904d0d05"
# Merge Maryland and filtered HHS data
df_MD_HHS_filter_state_merge = pd.merge(df_MD_HHS_state_merge,
df_MD_HHS[{("date"), ("% pos. under 35 yrs"), ("% pos. over 35 yrs"), ("testingvolume")}],
how = "right",
on = "date")
df_MD_HHS_filter_state_merge.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 377} id="9uPKqkCr3yDZ" outputId="9ce81e84-31c8-414d-b952-8b8ac3fbdf2f"
# Full merge of Maryland and Montana Data Frames
df_MD_MT_grand_state_merge = pd.merge(df_MD_HHS_filter_state_merge,
df_MT_daily_cumulative_merge,
how = "left",
on = "date")
# Preview data
df_MD_MT_grand_state_merge.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 377} id="XEgRVg4O5CGY" outputId="47a209d1-c7be-49dc-ffa7-167b50cc6e9f"
# Select specific time frame from Maryland data set e.g. last 30 days
df_MD_MT_grand_state_merge_30days = df_MD_MT_grand_state_merge.tail(30)
#Preview data
df_MD_MT_grand_state_merge_30days.head()
# + [markdown] id="PgkJ2ffn6LbB"
# # Some test code
# + id="9CCsDC6RuJFM"
#Select specific time frame from Maryland data set e.g. last 30 days
df_NYT_Covid_19_Maryland_20201127_30days = df_NYT_Covid_19_Maryland_20201127.tail(30)
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="yH6uhiTqxomR" outputId="268904b6-0481-4cfb-8a98-7dbd24f010bf"
# preview last 30 days of data for MD
df_NYT_Covid_19_Maryland_20201127_30days.head()
# + id="iMEQyFsZIxtm"
#df_NYT_Covid_19_Maryland_20201127_November2020 = df_NYT_Covid_19_Maryland_20201127[df_NYT_Covid_19_Maryland_20201127.tail(30)
# + id="ZG7qogJjxw_I"
#data frame with double filter e..g filter to only Maryland and to only ~ last month of data
#df_NYT_Covid_19_Maryland_20201127_November2020 = df_NYT_Covid_19_states_20201127[(df_NYT_Covid_19_states_20201127["state"] == "Maryland") & (df_NYT_Covid_19_states_20201127["date"] == "2020-11-01 to 2020-11-30")]
# + colab={"base_uri": "https://localhost:8080/", "height": 49} id="4lxESIkw1BL6" outputId="3a49edbf-d38e-4f03-9c2d-cc569d54745d"
# preview double filter data containing Maryland specific data and date specific data
#df_NYT_Covid_19_Maryland_20201127_November2020.head()
# + [markdown] id="GEB8D3L0iTxC"
# # Merging practice
# + id="Gd5-_v8U2fLz"
# Merging Maryland and Montana state level data, ONLY works under two conditions when use how = right and date and when use how = left and date
# When use how = left and on = date the right side column lists number with decimal formatting. Left means merge data from right dataset onto left dataset
# Have to use date because its the only column with same matching key AND same data contained down the rows
# Can do df_NYT_Covid_19-Montana_20201127[("desired column"), ("other desired column")] to specify what data gets merged from right data set onto left data set
df_MD_MT_merge = pd.merge(df_NYT_Covid_19_Maryland_20201127,
df_NYT_Covid_19_Montana_20201127,
how = "right",
on = "date")
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="bQZtJIP3RqG5" outputId="67c7945c-242f-43d5-a252-7de2367ad193"
# preview merged MD_MT data
df_MD_MT_merge.tail(5)
# + id="0vGM7fD7RuBi"
# Rename state_x, fips_x, cases_x, deaths_x, state_y, fips_y, cases_y, deaths_y to state specific columns
# Merge data from right onto left sooo Left is maryland right is MT
# format is df_name.rename(columns={"old name" : "new name"}) Columns specifies renaming column name
# Overwrite of original data frame with more specific names
df_MD_MT_merge = df_MD_MT_merge.rename(columns={"state_x" : "state_MD",
"fips_x": "fips_MD",
"cases_x": "cases_MD",
"deaths_x": "deaths_MD",
"state_y": "state_MT",
"fips_y": "fips_MT",
"cases_y": "cases_MT",
"deaths_y": "deaths_MT"})
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="EcQTMdiId14J" outputId="629722a2-a593-4508-8b41-a37102e4d512"
# Call of changed data frame to see changes
df_MD_MT_merge.head()
# + id="wIqOmlsExe1y"
# Calculate new daily cases and new daily deaths for each state using current day - previous day formulation e.g. daily new cases = current day - previous day
# data in dataframes are classified as integers - no need to change
# If desire to redefine at later data df_MD_MT_merge["cases_MD"] = df_MD_MT_merge["cases_MD"].astype(float, int, etc)
#df_MD_MT_merge.diff(periods=0,axis=0)
#df["daily cases_MD"] = (df_MD_MT_merge["cases_MD"] - df_MD_MT_merge["cases_MD"])
# to create new column in data set with unique name form is as follows: must be of same type e.g. both int, floats, objects etc
# df_xx["unique name"] = df_xx["col name" + df_xx["col name"]]
# + [markdown] id="uoRBr_2-6Zce"
# # Creating visualizations
# + colab={"base_uri": "https://localhost:8080/", "height": 377} id="xvVN9pJc7IvW" outputId="b4f443d8-3c8c-44b1-c898-ba1d32033b4c"
df_MD_MT_grand_state_merge_30days.head()
# + colab={"base_uri": "https://localhost:8080/"} id="1XClxOn77X6p" outputId="fab11c39-8c7a-4e3a-f478-0912d0c22fe5"
df_MD_MT_grand_state_merge_30days.columns
#df['cases_average'] = df.rolling(window=5).mean()
# + id="rEcLbOLYCbA4"
# Create bar graph visualization of daily covid cases for MD
bar_daily_cases_MD = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="daily cases_MD", title = "Recent trends of Covid-19 in Maryland during Fall 2020")
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="Y9RitvEx9zfL" outputId="36d8c271-e23d-4a13-c67d-f63a881354d2"
bar_daily_cases_MD
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="U95-E9tMSCFu" outputId="a857d765-9190-4615-9f39-368aee33ce5a"
# 3 day average
bar_daily_cases_MD_avg = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-case-avg_MD", title = "3-day average trend of Covid-19 cases in Maryland during Fall 2020")
bar_daily_cases_MD_avg
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="JvCMJN6ig-So" outputId="3cd78324-d4d7-4119-bce0-02a1889c011a"
# 3 day incidence MD
bar_cases_MD_3_day_incidence = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-case-incidence_MD", title = "3-day incidence trend of Covid-19 cases in Maryland during Fall 2020")
bar_cases_MD_3_day_incidence
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="G24NAKfQp-S-" outputId="a6292fa5-ac85-48e8-a421-34d095f2044d"
# Create bar graph visualization of 3 day avg of deaths for MD
bar_3day_deaths_MD = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-death-avg_MD", title = "Recent trends of Covid-19 deaths in Maryland during Fall 2020")
bar_3day_deaths_MD
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="oM3ib2TNsYPP" outputId="8689a316-2b2b-4906-96b5-6fa4cbd249b8"
# Create bar graph visualization of 3 day avg of deaths incidence for MD
bar_3day_deaths_MD_incidence = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-death-incidence_MD", title = "Recent trends of Covid-19 death incidence in Maryland during Fall 2020")
bar_3day_deaths_MD_incidence
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="Wh8QMUN8wImn" outputId="4acd8acd-d5fe-4344-9d52-d00bbf0b02d2"
# Create bar graph visualization of positivity rate for under 35 yr age
bar_positivity_rate_MD_under35 = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="% pos. under 35 yrs", title = "Recent trends in positivity rate for younger people in Maryland during Fall 2020")
bar_positivity_rate_MD_under35
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="AUi9kb7Lw69T" outputId="76565613-59be-4e41-8398-78367121a18d"
# Create bar graph visualization of positivity rate for over 35 yr age
bar_positivity_rate_MD_over35 = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="% pos. over 35 yrs", title = "Recent trends in positivity rate for older people in Maryland during Fall 2020")
bar_positivity_rate_MD_over35
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="nyt4S8brxY2t" outputId="201d061d-e24f-4490-e260-8d041fc9b8c1"
# Create bar graph visualization of testing volume of Covid-19 tests
bar_testingvolume_MD = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="testingvolume", title = "Recent trends in testing volume of Covid-19 in Maryland during Fall 2020")
bar_testingvolume_MD
# + colab={"base_uri": "https://localhost:8080/"} id="KKSTfLUSszLg" outputId="fab11c39-8c7a-4e3a-f478-0912d0c22fe5"
df_MD_MT_grand_state_merge_30days.columns
#df['cases_average'] = df.rolling(window=5).mean()
# + colab={"base_uri": "https://localhost:8080/"} id="jvjHH-tBsxJQ" outputId="fab11c39-8c7a-4e3a-f478-0912d0c22fe5"
df_MD_MT_grand_state_merge_30days.columns
#df['cases_average'] = df.rolling(window=5).mean()
# + id="IqE9HkSd85I8"
# daily cases MT
bar_daily_cases_MT = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="daily cases_MT", title = "Recent trends of Covid-19 in Montana during Fall 2020", color_discrete_sequence=['green'] )
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="-km8VkfjDO7m" outputId="50653510-909b-43c4-e15f-5c416434a762"
bar_daily_cases_MT
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="3HUOSO113j40" outputId="b6222669-9d6e-4e91-bf63-02a75f64f0e8"
# 3 day average
bar_daily_cases_MT_avg = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-case-avg_MT", title = "3-day average trend of Covid-19 cases in Montana during Fall 2020", color_discrete_sequence=['green'])
bar_daily_cases_MT_avg
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="WbQ7-s2tjksL" outputId="a309bc8b-d8ac-44e7-c602-366f0918d5e0"
# 3 day incidence MT
bar_cases_MT_3_day_incidence = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-case-incidence_MT", title = "3-day incidence trend of Covid-19 cases in Montana during Fall 2020", color_discrete_sequence=['green'])
bar_cases_MT_3_day_incidence
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="64LCn2I0nvqP" outputId="3a8439cc-6b50-4aff-f352-4a426d718110"
# Create bar graph visualization of 3 day avg of deaths for MT
bar_3day_deaths_MT = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-death-avg_MT", title = "Recent trends of Covid-19 deaths in Montana during Fall 2020",color_discrete_sequence=['green'])
bar_3day_deaths_MT
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="B-ORnLtIuUGP" outputId="d8015cc6-dafa-4f3c-afbb-582f55559024"
# Create bar graph visualization of 3 day avg of deaths incidence for MT
bar_3day_deaths_MT_incidence = px.bar(df_MD_MT_grand_state_merge_30days,x="date", y="3-day-death-incidence_MT", title = "Recent trends of Covid-19 death incidence in Montana during Fall 2020",color_discrete_sequence=['green'])
bar_3day_deaths_MT_incidence
# + id="BbGoId8gvHQo"
| 2020_11_17_MD_MT_Covid_19_data_IJ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## CIFAR 10
###code starts here
import sys
import os
os.getcwd()
sys.path.insert(0,"/notebooks") ###link the fast AI package
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# You can get the data via:
#
# wget http://pjreddie.com/media/files/cifar.tgz
from fastai.conv_learner import *
import torch
import shutil
PATH = "/notebooks/courses/dl1/data/cifar/"
OUTPATH = "/notebooks/courses/dl1/data/cifar10/"
os.makedirs(PATH,exist_ok=True)
shutil.rmtree(OUTPATH, ignore_errors=True)
os.makedirs(OUTPATH,exist_ok=True)
# !ls /notebooks/courses/dl1/data/cifar/
# +
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
stats = (np.array([ 0.4914 , 0.48216, 0.44653]), np.array([ 0.24703, 0.24349, 0.26159]))
# -
for x in classes:
os.makedirs(OUTPATH+'train/'+x,exist_ok=True)
os.makedirs(OUTPATH+'val/'+x,exist_ok=True)
os.makedirs(OUTPATH+'test/'+x,exist_ok=True)
filenames = os.listdir(PATH+'train/')
counts = {x:0 for x in classes}
print(len(filenames))
# !ls /notebooks/courses/dl1/data/cifar10/train
valset_size = len(filenames) / 10 * .2
for file_n in filenames:
for x in classes:
if x in file_n:
counts[x] = counts[x] +1
if counts[x] < valset_size:
shutil.copyfile(PATH+'train/'+file_n, OUTPATH+'val/'+x+'/'+file_n)
else:
shutil.copyfile(PATH+'train/'+file_n, OUTPATH+'train/'+x+'/'+file_n)
if 'automobile' in file_n:
counts['car'] = counts['car'] +1
if counts[x] < valset_size:
shutil.copyfile(PATH+'train/'+file_n, OUTPATH+'val/car/'+file_n)
else:
shutil.copyfile(PATH+'train/'+file_n, OUTPATH+'train/car/'+file_n)
# !ls /notebooks/courses/dl1/data/cifar10/train
# !ls /notebooks/courses/dl1/data/cifar10/val
# !ls /notebooks/courses/dl1/data/cifar/test
filenames = os.listdir(PATH+'test/')
for file_n in filenames:
shutil.copy(PATH+'test/'+file_n, OUTPATH+'test/'+file_n)
def get_data(sz,bs):
tfms = tfms_from_stats(stats, sz, aug_tfms=[RandomFlip()], pad=sz//8)
return ImageClassifierData.from_paths(OUTPATH, val_name='val', tfms=tfms, bs=bs)
# ### Look at data
data = get_data(32,4)
x,y=next(iter(data.trn_dl))
plt.imshow(data.trn_ds.denorm(x)[0]);
plt.imshow(data.trn_ds.denorm(x)[1]);
# ## Fully connected model
bs=256
data = get_data(32,bs)
lr=1e-2
# From [this notebook](https://github.com/KeremTurgutlu/deeplearning/blob/master/Exploring%20Optimizers.ipynb) by our student <NAME>:
class SimpleNet(nn.Module):
def __init__(self, layers):
super().__init__()
self.layers = nn.ModuleList([
nn.Linear(layers[i], layers[i + 1]) for i in range(len(layers) - 1)])
def forward(self, x):
x = x.view(x.size(0), -1)
for l in self.layers:
l_x = l(x)
x = F.relu(l_x)
return F.log_softmax(l_x, dim=-1)
learn = ConvLearner.from_model_data(SimpleNet([32*32*3, 40,10]), data)
learn, [o.numel() for o in learn.model.parameters()]
learn.summary()
learn.lr_find()
learn.sched.plot()
# %time learn.fit(lr, 2)
# %time learn.fit(lr, 2, cycle_len=1)
# ## CNN
class ConvNet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.layers = nn.ModuleList([
nn.Conv2d(layers[i], layers[i + 1], kernel_size=3, stride=2)
for i in range(len(layers) - 1)])
self.pool = nn.AdaptiveMaxPool2d(1)
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
for l in self.layers: x = F.relu(l(x))
x = self.pool(x)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvNet([3, 20, 40, 80], 10), data)
learn.summary()
learn.lr_find(end_lr=100)
learn.sched.plot()
# %time learn.fit(1e-1, 2)
# %time learn.fit(1e-1, 4, cycle_len=1)
# ## Refactored
class ConvLayer(nn.Module):
def __init__(self, ni, nf):
super().__init__()
self.conv = nn.Conv2d(ni, nf, kernel_size=3, stride=2, padding=1)
def forward(self, x): return F.relu(self.conv(x))
class ConvNet2(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.layers = nn.ModuleList([ConvLayer(layers[i], layers[i + 1])
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
for l in self.layers: x = l(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvNet2([3, 20, 40, 80], 10), data)
learn.summary()
# %time learn.fit(1e-1, 2)
# %time learn.fit(1e-1, 2, cycle_len=1)
# ## BatchNorm
class BnLayer(nn.Module):
def __init__(self, ni, nf, stride=2, kernel_size=3):
super().__init__()
self.conv = nn.Conv2d(ni, nf, kernel_size=kernel_size, stride=stride,
bias=False, padding=1)
self.a = nn.Parameter(torch.zeros(nf,1,1))
self.m = nn.Parameter(torch.ones(nf,1,1))
def forward(self, x):
x = F.relu(self.conv(x))
x_chan = x.transpose(0,1).contiguous().view(x.size(1), -1)
if self.training:
self.means = x_chan.mean(1)[:,None,None]
self.stds = x_chan.std (1)[:,None,None]
return (x-self.means) / self.stds *self.m + self.a
class ConvBnNet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i + 1])
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l in self.layers: x = l(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvBnNet([10, 20, 40, 80, 160], 10), data)
learn.summary()
# %time learn.fit(3e-2, 2)
# %time learn.fit(1e-1, 4, cycle_len=1)
# ## Deep BatchNorm
class ConvBnNet2(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([BnLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l,l2 in zip(self.layers, self.layers2):
x = l(x)
x = l2(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvBnNet2([10, 20, 40, 80, 160], 10), data)
# %time learn.fit(1e-2, 2)
# %time learn.fit(1e-2, 2, cycle_len=1)
# ## Resnet
class ResnetLayer(BnLayer):
def forward(self, x): return x + super().forward(x)
class Resnet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.layers3 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l,l2,l3 in zip(self.layers, self.layers2, self.layers3):
x = l3(l2(l(x)))
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(Resnet([10, 20, 40, 80, 160], 10), data)
wd=1e-5
# %time learn.fit(1e-2, 2, wds=wd)
# %time learn.fit(1e-2, 3, cycle_len=1, cycle_mult=2, wds=wd)
# %time learn.fit(1e-2, 8, cycle_len=4, wds=wd)
# + [markdown] heading_collapsed=true
# ## Resnet 2
# + hidden=true
class Resnet2(nn.Module):
def __init__(self, layers, c, p=0.5):
super().__init__()
self.conv1 = BnLayer(3, 16, stride=1, kernel_size=7)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.layers3 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
self.drop = nn.Dropout(p)
def forward(self, x):
x = self.conv1(x)
for l,l2,l3 in zip(self.layers, self.layers2, self.layers3):
x = l3(l2(l(x)))
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
x = self.drop(x)
return F.log_softmax(self.out(x), dim=-1)
# + hidden=true
learn = ConvLearner.from_model_data(Resnet2([16, 32, 64, 128, 256], 10, 0.2), data)
# + hidden=true
wd=1e-6
# + hidden=true
# %time learn.fit(1e-2, 2, wds=wd)
# + hidden=true
# %time learn.fit(1e-2, 3, cycle_len=1, cycle_mult=2, wds=wd)
# + hidden=true
# %time learn.fit(1e-2, 8, cycle_len=4, wds=wd)
# + hidden=true
learn.save('tmp3')
# + hidden=true
log_preds,y = learn.TTA()
preds = np.mean(np.exp(log_preds),0)
# + hidden=true
metrics.log_loss(y,preds), accuracy(preds,y)
# + [markdown] hidden=true
# ### End
# + hidden=true
| examples/ipynbexamples/lesson7-cifar10-MAEdits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import clear_output
import os,cv2,random,torch,torchvision
import numpy as np
from PIL import Image
from matplotlib import pyplot as plt
# %matplotlib inline
from sklearn.metrics import confusion_matrix
import seaborn as sns
# +
AllClasses = ['Markov',
'Non-Markov']
class DataProcess(torch.utils.data.Dataset):
def __init__(self, ImagesPath, ImageTransformer):
global AllClasses
def GetImage(Class):
images = [x for x in os.listdir(ImagesPath[Class]) if x[-3:].lower().endswith('jpg') or x[-3:].lower().endswith('png')]
print(f'Found {len(images)} {Class} examples')
return images
self.images = {}
self.Classes = AllClasses
for Class in self.Classes:
self.images[Class] = GetImage(Class)
self.ImagesPath = ImagesPath
self.ImageTransformer = ImageTransformer
def __len__(self):
return sum([len(self.images[Class]) for Class in self.Classes])
def __getitem__(self, index):
Class = random.choice(self.Classes)
index = index % len(self.images[Class])
NameOfPicture = self.images[Class][index]
PathOfPicture = os.path.join(self.ImagesPath[Class], NameOfPicture)
image = Image.open(PathOfPicture).convert('RGB')
return self.ImageTransformer(image), self.Classes.index(Class)
def AccCalc(LabelsValues,OutputValues) :
_,PredictedValues = torch.max(OutputValues, 1)
return sum((PredictedValues == LabelsValues).numpy())
# -
AllImages = {}
SavedImages = 0
Path = 'E:/end/Data-M'
for folder in os.listdir(Path) :
ThisPath = os.path.join(Path,folder)
AllImages[folder] = []
for image in os.listdir(ThisPath) :
SavedImages+=1
AllImages[folder].append(os.path.join(ThisPath,image))
AllImages
len(AllImages.keys()),SavedImages
def ShowImages() :
global AllImages
fig, ax = plt.subplots(2, 1, figsize=(80,60))
fig.tight_layout(pad=0.010)
for i in range(len(ax)):
im = np.random.choice(AllImages[list(AllImages.keys())[i]])
ax[i].imshow(plt.imread(im))
ax[i].set_title(list(AllImages.keys())[i])
plt.setp(plt.gcf().get_axes(), xticks=[], yticks=[])
ShowImages()
SavedImages,TrainingDataSize,TestingDataSize=0,0,0
ThisModel,ThisLossFunction ,ThisOptimizer = None,None,None
ThisTransformer = torchvision.transforms.Compose([torchvision.transforms.Resize(size=(50,50)),
torchvision.transforms.RandomHorizontalFlip(),
torchvision.transforms.ToTensor()])
# +
ImagesPath = {i:os.path.join(r'E:/end/Data-M',i) for i in AllClasses}
ProcessedData = DataProcess(ImagesPath, ThisTransformer)
TrainingDataSize = 0.8
TrainingSize = int(TrainingDataSize * len(ProcessedData))
TestingSize = len(ProcessedData) - TrainingSize
TrainingData, TestingData = torch.utils.data.random_split(ProcessedData, [TrainingSize, TestingSize])
TrainingDataSize,TestingDataSize=len(TrainingData), len(TestingData)
print("Length of train set : ", len(TrainingData))
print("Length of test set : ", len(TestingData))
# +
BatchSize = 10
TrainingDataLength = torch.utils.data.DataLoader(TrainingData, batch_size = BatchSize, shuffle = True)
TestingDataLength = torch.utils.data.DataLoader(TestingData, batch_size=BatchSize, shuffle=True)
print("Length of training batches", len(TrainingDataLength))
print("Lentgth of test batches", len(TestingDataLength))
# -
resnet50 = torchvision.models.resnet50(pretrained=True)
print(resnet50)
resnet50.fc = torch.nn.Linear(in_features=2048, out_features=2)
loss_fn = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(resnet50.parameters(), lr=0.00001)
# +
# %%time
NumberOfEpochs = 30
ThisModel = resnet50
ThisOptimizer = optimizer
ThisLossFunction=loss_fn
AccuracyTraining,AccuracyTesting,LossTraining,LossTesting = [],[],[],[]
for j in range(NumberOfEpochs):
print(j)
TrainLoss,TestLoss,TrainAccuracy,TestAccuracy = 0,0,0,0
ThisModel.train()
i=0
for X,y in TrainingDataLength:
i+=1
ThisOptimizer.zero_grad()
ThisOutput = ThisModel(X)
ThisLoss = ThisLossFunction(ThisOutput,y)
ThisLoss.backward()
ThisOptimizer.step()
TrainLoss += ThisLoss.item()
TrainAccuracy += AccCalc(y,ThisOutput)
TrainLoss /=i
TrainAccuracy /=TrainingDataSize
AccuracyTraining.append(TrainAccuracy)
LossTraining.append(TrainLoss)
ThisModel.eval()
i=0
for X,y in TestingDataLength:
i+=1
ThisOutput = ThisModel(X)
ThisLoss = ThisLossFunction(ThisOutput,y)
TestLoss += ThisLoss.item()
TestAccuracy += AccCalc(y,ThisOutput)
TestLoss/=i
TestAccuracy /=TestingDataSize
AccuracyTesting.append(TestAccuracy)
LossTesting.append(TestLoss)
clear_output(wait=True)
print(f'For Epoch Number {j+1}')
print('---------------------------------------------')
print(f'Train Loss is {np.round(TrainLoss,2)} & Train Accuracy is {np.round(TrainAccuracy,2)} ')
print(f'Test Loss is {np.round(TestLoss,2)} & Test Accuracy is {np.round(TestAccuracy,2)}')
torch.save(ThisModel, f"E:/end/code/M-50-30-Yes/M-50-30-Yes")
plt.figure(figsize=(20,10))
XValue = [k for k in range(j+1)]
plt.subplot(1,2,1)
plt.plot(XValue,AccuracyTraining,label='Train Acc')
plt.plot(XValue,AccuracyTesting,label='Test Acc')
plt.legend()
plt.subplot(1,2,2)
plt.plot(XValue,LossTraining,label='Train Loss')
plt.plot(XValue,LossTesting,label='Test Loss')
plt.legend()
plt.show()
# -
ThisModel = torch.load('E:/end/code/M-50-30-Yes/M-50-30-Yes')
i=0
AllYValue,AllOutput = [],[]
ThisModel.eval()
ThisLossFunction = torch.nn.CrossEntropyLoss()
TestLoss,TestAccuracy = 0,0
for X,y in TestingDataLength:
i+=1
# if i ==50 : break
if i%50==0 : print(i)
ThisOutput = ThisModel(X)
AllYValue.extend(y)
_,PredictedValues = torch.max(ThisOutput, 1)
AllOutput.extend([int(i) for i in PredictedValues])
ThisLoss = ThisLossFunction(ThisOutput,y)
TestLoss += ThisLoss.item()
TestAccuracy += AccCalc(y,ThisOutput)
z=0
for a,b in zip(AllOutput,[int(i) for i in AllYValue]) :
if a==b : z+=1
z/len(AllOutput)
TestLoss/i,TestAccuracy/i*0.10
print(f'Model Accuracy is {np.round(TestAccuracy/i*0.10,2)} and loss is {np.round(TestLoss/i,2)}')
AllYValue = [int(i) for i in AllYValue]
from sklearn.metrics import classification_report,confusion_matrix
confusion_matrix(AllOutput,AllYValue)
from sklearn.metrics import plot_confusion_matrix
import matplotlib.pyplot as plt
import itertools
import numpy as np
def plot_confusion_matrix(cm, target_names, title='Confusion matrix', cmap=None, normalize=True):
accuracy = np.trace(cm) / float(np.sum(cm))
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Reds')
plt.figure(figsize=(8, 6))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=90)
plt.yticks(tick_marks, target_names)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
conf_matrix =[[4746, 7],
[ 0, 4847]]
Calss_Names = ['Markov',
'Non-Markov']
plot_confusion_matrix(cm = np.array(conf_matrix),
normalize = False,
target_names = Calss_Names,
title ="Confusion Matrix")
print(classification_report(AllOutput,AllYValue))
| Eman_NN/M-50-30-Yes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/madhavjk/DataScience-ML_and_DL/blob/main/Untitled4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="gWLbtNgRiEYu"
import tensorflow_hub as hub
import tensorflow as tf
from matplotlib import pyplot as plt
import numpy as np
import cv2
# + id="vpOjmbXNjhq-"
model = hub.load('https://tfhub.dev/google/magenta/arbitrary-image-stylization-v1-256/2')
# + id="R9KA_3cBkHgx"
def load_image(img_path):
img = tf.io.read_file(img_path)
img = tf.image.decode_image(img, channels=3)
img = tf.image.convert_image_dtype(img, tf.float32)
img = img[tf.newaxis, :]
return img
# + id="MCD4yIpwkmPm"
content_image = load_image('/content/baby_yoda.jpg')
style_image = load_image('/content/starrynight.jfif')
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="6Ciayjs9k9Qz" outputId="d5706afe-e4cf-4535-b657-6d885b207082"
plt.imshow(np.squeeze(content_image))
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="9m0RiidplOJ3" outputId="dad9efcd-5649-4c86-e06f-57f68d1d0b96"
plt.imshow(np.squeeze(style_image))
# + id="XLQo8kWzlf29"
stylized_image = model(tf.constant(content_image), tf.constant(style_image))[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="L5B19oe-muLA" outputId="3712b927-3561-4652-87ad-8b94d34ab3c6"
plt.imshow(np.squeeze(stylized_image))
| Untitled4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
import numpy as np
import importlib
import custom_implementations as custom_imp
import matplotlib.pyplot as plt
dataset = pd.read_csv("../Datasets/heart.txt", sep='\s+')
cols_of_interest = ['age','chest_pain','resting_blood_pressure','serum_chol','resting_electro_results','maximum_heart_rate,exercise','induced_angina','oldpeak','slope_peak_exercise_st','num_major_vessels','absent']
dataset = dataset[cols_of_interest]
# Not Absent: 0
# Absent: 1
minority_class = 1
majority_class = 0
classes = 'absent'
cols = list(dataset.columns.values)
class_index = cols.index(classes)
print 'Number of not absent: ', len(dataset[dataset['absent'] == 0])
print 'Number of absent: ', len(dataset[dataset['absent'] == 1])
print(cols)
# -
# ## Find best K for KNN
# +
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
# Split into data and labels
x = preprocessing.normalize(dataset.iloc[:,:-1])
y = dataset.iloc[:,-1].values
x_train, x_test, y_train, y_test = train_test_split(x,y,random_state=1, test_size=0.25)
k_best = custom_imp.knn_set_hyper_params(x_train, y_train, x_test, y_test, x, y)
print 'Best K: ', k_best
# -
# ## KNN on imbalanced dataset
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import cross_val_score
# Create KNN and fit model
model = KNeighborsClassifier(n_neighbors=k_best, metric='minkowski', p=2)
model.fit(x_train, y_train)
# Predict on testing data
y_pred = model.predict(x_test)
# 10-cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
class_labels = ['Not Absent', 'Absent']
pos_label = 1
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## Set SVM parameters
svm_best_params = custom_imp.svm_set_hyper_params(x_train, y_train, x_test, y_test, True)
print 'Best SVM hyperparameters: ', svm_best_params
# ## SVM on imbalanced dataset
# +
from sklearn import svm
from sklearn import metrics
# Fit model
# C: penalty parameter of the error term, i.e. slack variable
# kernel (linear, rbf)
# gamma: kernel coefficient for rbf, poly, sigmoid
# tol: tolerance for stopping criterion
# max_iter: limit on eopochs
# random_state: seed when shuffling
kernel = svm_best_params['kernel']
if(kernel == 'linear'):
model = svm.SVC(C=svm_best_params['C'], max_iter=-1, kernel='linear')
else:
model = svm.SVC(gamma=svm_best_params['gamma'], C=svm_best_params['C'], max_iter=-1, kernel='rbf')
model.fit(x_train, y_train)
# Show SVM params
print model.get_params
# Predict on testing data
y_pred = model.predict(x_test)
print ("Accuracy: %f" %(metrics.accuracy_score(y_test, y_pred)))
# k cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# Get support vectors
print 'Support vectors: \n', model.support_vectors_
print y_pred
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## Apply SMOTE to imbalanced dataset
# +
## Use SMOTE on N% of the dataset with k neighbours
smote_percentages = [50,100,200,300]
x = dataset.iloc[:,:-1].values
x = preprocessing.normalize(x)
y = dataset.iloc[:,-1].values
x_majority = preprocessing.normalize(dataset[dataset[classes] == 1].iloc[:,:-1].values)
x_minority = preprocessing.normalize(dataset[dataset[classes] == 0].iloc[:,:-1].values)
y_majority = dataset[dataset[classes] == 1].iloc[:,-1].values
y_minority = dataset[dataset[classes] == 0].iloc[:,-1].values
print 'Percentage to SMOTE: ', smote_percentages[3]
smoted_samples = custom_imp.smote(x_minority, smote_percentages[3], k_best)
print 'Number of synthetic samples SMOTEd: ', len(smoted_samples)
updated_x_minority = np.concatenate((x_minority, smoted_samples), axis=0)
print 'Number of minority after: %s' % len(updated_x_minority)
plt.plot([x_majority[3]], [x_majority[4]], 'g^', [updated_x_minority[3]], [updated_x_minority[4]], 'r*')
# Update x and y for smote
x_smote = np.concatenate((x, smoted_samples), axis=0)
smote_y = np.full((len(smoted_samples)), minority_class)
y_smote = np.concatenate((y, smote_y), axis=0)
# -
# ## KNN after SMOTE
# +
x_train, x_test, y_train, y_test = train_test_split(x_smote,y_smote,random_state=1, test_size=0.25, shuffle=True)
k_best = custom_imp.knn_set_hyper_params(x_train, y_train, x_test, y_test, x, y)
print 'Best K: ', k_best
# Create KNN and fit model
model = KNeighborsClassifier(n_neighbors=k_best, metric='minkowski', p=2)
model.fit(x_train, y_train)
# Predict on testing data
y_pred = model.predict(x_test)
# 10-cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("\n10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
class_labels = ['Not Absent', 'Absent']
pos_label = 1
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## Set SVM parameters
svm_best_params = custom_imp.svm_set_hyper_params(x_train, y_train, x_test, y_test, True)
print svm_best_params
# ## SVM after SMOTE
# +
kernel = svm_best_params['kernel']
if(kernel == 'linear'):
model = svm.SVC(C=svm_best_params['C'], max_iter=-1, kernel='linear')
else:
model = svm.SVC(gamma=svm_best_params['gamma'], C=svm_best_params['C'], max_iter=-1, kernel='rbf')
model.fit(x_train, y_train)
# Show SVM params
print model.get_params
# Predict on testing data
y_pred = model.predict(x_test)
print ("Accuracy: %f" %(metrics.accuracy_score(y_test, y_pred)))
# k cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# Get support vectors
print 'Support vectors: \n', model.support_vectors_
print y_pred
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## TOMEK Links on imbalanced dataset
# +
# Declare column variables
classes = 'absent'
class_x = 1
class_y = 2
label_0 = 'Not Absent' # Change depending on which class 0 belongs to
label_1 = 'Absent' # Change depending on which class 1 belongs to
x_majority = dataset[dataset[classes] == 1].iloc[:,:-1]
x_minority = dataset[dataset[classes] == 0].iloc[:,:-1]
y_majority = dataset[dataset[classes] == 1].iloc[:,-1]
y_minority = dataset[dataset[classes] == 0].iloc[:,-1]
print 'Number of majority before: ', len(x_majority)
num_to_remove = len(x_majority) - len(x_minority)
if num_to_remove > len(x_minority):
num_to_remove = len(x_minority)
tomek_dataset = custom_imp.tomek(x_majority.values, x_minority.values, y_majority.values, y_minority.values, num_to_remove, classes, 1)
tomek_df = pd.DataFrame(tomek_dataset)
print 'Number of majority after: ', len(tomek_df[tomek_df[10] == 0].iloc[:,:-1])
# Split into data and labels
x_tomek = preprocessing.normalize(tomek_df.iloc[:,:-1].values)
y_tomek = tomek_df.iloc[:,-1].values
x_train, x_test, y_train, y_test = train_test_split(x_tomek,y_tomek,random_state=1, test_size=0.25, shuffle=True)
print 'done'
# -
# ## KNN after TOMEK Links
# +
k_best = custom_imp.knn_set_hyper_params(x_train, y_train, x_test, y_test, x, y)
print 'Best K: ', k_best
# Create KNN and fit model
model = KNeighborsClassifier(n_neighbors=k_best, metric='minkowski', p=2)
model.fit(x_train, y_train)
# Predict on testing data
y_pred = model.predict(x_test)
# 10-cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("\n10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
class_labels = ['Not Absent', 'Absent']
pos_label = 1
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## Set SVM parameters
svm_best_params = custom_imp.svm_set_hyper_params(x_train, y_train, x_test, y_test, True)
print svm_best_params
# ## SVM after TOMEK Links
# +
kernel = svm_best_params['kernel']
if(kernel == 'linear'):
model = svm.SVC(C=svm_best_params['C'], max_iter=-1, kernel='linear')
else:
model = svm.SVC(gamma=svm_best_params['gamma'], C=svm_best_params['C'], max_iter=-1, kernel='rbf')
model.fit(x_train, y_train)
# Show SVM params
print model.get_params
# Predict on testing data
y_pred = model.predict(x_test)
print ("Accuracy: %f" %(metrics.accuracy_score(y_test, y_pred)))
# k cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# Get support vectors
print 'Support vectors: \n', model.support_vectors_
print y_pred
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## SMOTE + TOMEK Links on imbalanced dataset
# +
# Declare column variables
classes = 'absent'
class_x = 1
class_y = 2
label_0 = 'Not Absent' # Change depending on which class 0 belongs to
label_1 = 'Absent' # Change depending on which class 1 belongs to
# Separate into features and labels depending upon classification label
x_majority = preprocessing.normalize(dataset[dataset[classes] == 1].iloc[:,:-1])
x_minority = preprocessing.normalize(dataset[dataset[classes] == 0].iloc[:,:-1])
y_majority = dataset[dataset[classes] == 1].iloc[:,-1]
y_minority = dataset[dataset[classes] == 0].iloc[:,-1]
print 'Number of majority before: ', len(x_majority)
print 'Number of minority before: ', len(x_minority)
smote_tomek_dataset = custom_imp.smote_tomek(x_majority, x_minority, y_majority.values, y_minority.values, majority_class, minority_class, smote_percentages[3], k_best, classes)
smote_tomek_df = pd.DataFrame(smote_tomek_dataset)
x_smote_tomek = smote_tomek_df.iloc[:,:-1].values
y_smote_tomek = smote_tomek_df.iloc[:, len(dataset.columns)-1].values
print 'Number of majority after: ', len(smote_tomek_df[smote_tomek_df[10] == 1])
print 'Number of minority after: ', len(smote_tomek_df[smote_tomek_df[10] == 0])
x_train, x_test, y_train, y_test = train_test_split(x_smote_tomek,y_smote_tomek,random_state=1, test_size=0.25, shuffle=True)
# -
# ## KNN after SMOTE + TOMEK Links
# +
k_best = custom_imp.knn_set_hyper_params(x_train, y_train, x_test, y_test, x, y)
print 'Best K: ', k_best
# Create KNN and fit model
model = KNeighborsClassifier(n_neighbors=k_best, metric='minkowski', p=2)
model.fit(x_train, y_train)
# Predict on testing data
y_pred = model.predict(x_test)
# 10-cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("\n10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
class_labels = ['Not Absent', 'Absent']
pos_label = 1
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
# ## Set SVM parameters
svm_best_params = custom_imp.svm_set_hyper_params(x_train, y_train, x_test, y_test, True)
print svm_best_params
# ## SVM after SMOTE + TOMEK Links
# +
kernel = svm_best_params['kernel']
if(kernel == 'linear'):
model = svm.SVC(C=svm_best_params['C'], max_iter=-1, kernel='linear')
else:
model = svm.SVC(gamma=svm_best_params['gamma'], C=svm_best_params['C'], max_iter=-1, kernel='rbf')
model.fit(x_train, y_train)
# Show SVM params
print model.get_params
# Predict on testing data
y_pred = model.predict(x_test)
print ("Accuracy: %f" %(metrics.accuracy_score(y_test, y_pred)))
# k cross fold validation
scores = cross_val_score(model, x, y, cv=10)
print("10-Fold Cross Validation: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# Get support vectors
print 'Support vectors: \n', model.support_vectors_
print y_pred
custom_imp.show_metrics(y_test, y_pred, class_labels, pos_label)
# -
| Code/heart_disease.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NYC new measles cases by month (2018 - 2019)
#
# * This Jupyter/Python notebook creates a bar chart of the number of new measles cases by month during the 2018-2019 measles outbreak in NYC.
# * This notebook is part of the [Visualizing the 2019 Measles Outbreak](https://carlos-afonso.github.io/measles/) open-source GitHub project.
# * [<NAME>](https://www.linkedin.com/in/carlos-afonso-w/), Jan 4, 2020.
# ## Import libraries
from datetime import datetime
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
import os
import pandas as pd
# ## Read and show the data
#
# The data was manually collected from the source ([NYC Health Measles webpage](https://www1.nyc.gov/site/doh/health/health-topics/measles.page)) and saved as a [CSV file](https://github.com/carlos-afonso/measles/blob/master/data/nyc-health/2018-2019-outbreak/nyc-new-measles-cases-by-month.csv). We use this manual approach because the data is small.
# +
# Set (relative) path to the CSV data file
data_file = os.path.join('..', 'data', 'nyc-health', '2018-2019-outbreak', 'nyc-new-measles-cases-by-month.csv')
# Import data from the CSV file as a pandas dataframe
df = pd.read_csv(data_file)
# Show the data
df
# -
# ## Extract Data
#
# Extract and transform the data necessary to create the bar chart.
# ### Extract the Start and End Dates
#
# We want to show the start and end dates in the plot, to provide context. So, we need to extract these two dates from the dataframe (first two columns) and transform them into nicelly formatted strings.
# +
# Notes about the lambda function below:
# 1) strptime transforms the raw date string to a datetime object
# 2) strftime transforms the datetime object to a nicelly formatted date string
[start_month, end_month] = map(
lambda x: datetime.strptime(x, '%Y-%m-%d').strftime('%b %Y'),
df.iloc[0, :2]
)
# Show the nicelly formated date strings
[start_month, end_month]
# -
# ### Extract the Total Cases
#
# We also want to show the total number of cases in the plot, to provide context.
# +
# Get the number of total cases (last row, last column)
total_cases = df.iloc[0, -1]
# Check if there is a data problem where the reported total
# does not match with the sum of the new cases per month
if total_cases != df.iloc[0, 2:-1].sum():
print('WARNING: cases for each month do NOT add up to the reported total!')
# Show the total cases
total_cases
# -
# ### Extract Data to Plot
#
# Extract the data to actually plot (third to penultimate columns).
# +
# Extract the data to plot (third to penultimate columns)
data_to_plot = df.iloc[0, 2:-1]
# Show the data to plot
data_to_plot
# -
# ### Create Month Labels
#
# Create nicely formatted month labels to be used in the bar chart.
# +
# Get the 'raw' month labels
month_labels = list(data_to_plot.index)
# For the first and last month: show month and year but
# replace the space between the month and year with a line break
first_label = month_labels[0].replace(' ', '\n')
last_label = month_labels[-1].replace(' ', '\n')
# For the intermediate months: show only the month (not the year), except
# for Jan in which case show the month and year with a line break in between
middle_labels = list(map(
lambda s: s.replace(' ', '\n') if s[0:3] == 'Jan' else s[0:3],
month_labels[1:-1]
))
# Join all the month labels
month_labels = [first_label] + middle_labels + [last_label]
# Show the nicely formated month labels
month_labels
# -
# ## Create bar chart
#
# We want the bar chart to be clear and to contain the necessary context.
#
# To contextualize the bar chart we:
# * use a title that explictly says what the bar chart represents;
# * add text annotations that provides information about:
# * the start and end dates,
# * the total number of cases during that period, and
# * the data and image sources.
#
# To make the bar chart as clear as possible we:
# * explicitly show the number of new cases for each month;
# * use nicelly format labels for the months;
# * use a large enough font to make all labels easy to read;
# * remove unnecessary elements (x-axis ticks, y-axis ticks and values, and plot box).
# +
# Define font parameters
fn = 'Arial' # font name
fsb = 12 # font size base
# Create figure
fig = plt.figure()
# Add figure title
fig.suptitle('NYC new measles cases by month (2018 - 2019)', fontname=fn, fontsize=(fsb + 4))
#plt.title('NYC new measles cases by month', fontname=fn, fontsize=(fsb + 4))
# Create the horizontal bar chart
ax = data_to_plot.plot.bar(alpha=0.3, color='red', width=0.8)
# Remove the y-axis ticks and labels/values
#ax.get_yaxis().set_ticks([])
ax.set_yticks([])
# Remove the x-axis ticks
ax.xaxis.set_ticks_position('none')
# Set the x-axis labels
ax.set_xticklabels(month_labels, fontname=fn, fontsize=(fsb-1), rotation=0)
# Create the labels on top of the bars to explicitly show the
# number of new measles cases in each month
for i in ax.patches:
ax.text(i.get_x() + 0.4, i.get_height() + 5, str(i.get_height()),
fontname = fn, fontsize = fsb, horizontalalignment = 'center')
# Remove the axes box
plt.box(False)
# Add note about the total cases
text = str(total_cases) + ' total confirmed cases from ' + start_month + ' to ' + end_month
fig.text(0.5, -0.07, text, fontname = fn, fontsize = (fsb - 1), horizontalalignment='center')
# Add note about increased vaccination efforst
text = 'In Apr 2019 NYC increased its MMR vaccination efforts'
fig.text(0.5, -0.14, text, fontname = fn, fontsize = (fsb - 1), horizontalalignment = 'center')
# Add note about the end of the outbreak
text = 'Community transmission was declared over on Sep 3, 2019'
fig.text(0.5, -0.21, text, fontname = fn, fontsize = (fsb - 1), horizontalalignment = 'center')
# Add note about the Data and Image sources
text = 'Data: NYC Health, Image: carlos-afonso.github.io/measles'
fig.text(0.5, -0.28, text, fontname = 'Lucida Console', fontsize = (fsb - 3), horizontalalignment = 'center')
# Show figure
plt.show()
# -
# ## Save bar chart
# +
# Set image file path/name (without file extension)
img_file = os.path.join('..', 'images', 'nyc-new-measles-cases-by-month-bar-chart-py')
# Save as PNG image
fig.savefig(img_file + '.png', bbox_inches='tight', dpi=200)
# Save as SVG image
fig.savefig(img_file + '.svg', bbox_inches='tight')
# -
# ## Export notebook as HTML
# Export this notebook as a static HTML page
os.system('jupyter nbconvert --to html nyc-new-measles-cases-by-month-final.ipynb')
| notebooks/nyc-new-measles-cases-by-month-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.039443, "end_time": "2022-03-30T13:25:54.900927", "exception": false, "start_time": "2022-03-30T13:25:54.861484", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + [markdown] papermill={"duration": 0.020258, "end_time": "2022-03-30T13:25:54.944745", "exception": false, "start_time": "2022-03-30T13:25:54.924487", "status": "completed"} tags=[]
# ## Setting up data
# + papermill={"duration": 1.046914, "end_time": "2022-03-30T13:25:56.013329", "exception": false, "start_time": "2022-03-30T13:25:54.966415", "status": "completed"} tags=[]
# Set up data sets
from sklearn.model_selection import train_test_split
X_test_full = pd.read_csv('../input/titanic/test.csv', index_col='PassengerId')
X_full = pd.read_csv('../input/titanic/train.csv', index_col='PassengerId')
X_full.dropna(axis=0, subset=['Survived'])
y = X_full.Survived
X_full.drop(['Survived'], axis=1, inplace=True)
feature_columns = ['Pclass', 'Sex', 'Age', 'SibSp', 'Parch']
X = X_full[feature_columns]
X_test = X_test_full[feature_columns]
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2)
print('Amount of rows for training set: %d' %(X.shape[0]))
# + papermill={"duration": 0.039964, "end_time": "2022-03-30T13:25:56.074386", "exception": false, "start_time": "2022-03-30T13:25:56.034422", "status": "completed"} tags=[]
X.head()
# + [markdown] papermill={"duration": 0.021356, "end_time": "2022-03-30T13:25:56.117004", "exception": false, "start_time": "2022-03-30T13:25:56.095648", "status": "completed"} tags=[]
# ## Preprocessing
# + papermill={"duration": 0.034749, "end_time": "2022-03-30T13:25:56.173445", "exception": false, "start_time": "2022-03-30T13:25:56.138696", "status": "completed"} tags=[]
print('Amount of missing values in each column: ')
X.isnull().sum()
# + [markdown] papermill={"duration": 0.02178, "end_time": "2022-03-30T13:25:56.217286", "exception": false, "start_time": "2022-03-30T13:25:56.195506", "status": "completed"} tags=[]
# <font size=3>There are missing values for the Age feature. A bit of data analysis is necessary in order to fill in those empty values in a meaningful way. **Reminder: we should use the training set when deriving features from data to avoid contamination.**
# + papermill={"duration": 1.099738, "end_time": "2022-03-30T13:25:57.339797", "exception": false, "start_time": "2022-03-30T13:25:56.240059", "status": "completed"} tags=[]
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib import cm
sns.set_theme(style='darkgrid')
fig, ax = plt.subplots(2, 2, figsize=(18, 8))
ax[0,0].scatter(x=X_train.Age, y=X_train.SibSp, alpha=0.75, c=X_train.SibSp, cmap=cm.get_cmap('summer'))
ax[0,0].set_title('Siblings / spouses per age')
ax[0,1].scatter(x=X_train.Age, y=X_train.Parch, c=X_train.Parch, cmap=cm.get_cmap('viridis'))
ax[0,1].set_title('Parents / children per age')
x1 = X_train[X_train.Age.isnull()]
ax[1,0].scatter(range(len(x1)), x1.SibSp, alpha=0.75, c=x1.SibSp, cmap=cm.get_cmap('summer'))
ax[1,0].set_title('Siblings / spouses for each missing age person')
ax[1,1].scatter(range(len(x1)), x1.Parch, c=x1.Parch, cmap=cm.get_cmap('viridis'))
ax[1,1].set_title('Parents / children for each missing age person')
plt.show()
# + [markdown] papermill={"duration": 0.025741, "end_time": "2022-03-30T13:25:57.390805", "exception": false, "start_time": "2022-03-30T13:25:57.365064", "status": "completed"} tags=[]
# <font size=3>The vast majority of people with missing input for the Age column have no siblings/spouses or parents/children, therefore it makes sense to use the average age within that group (people with no siblings/spouses or parents/children) as an estimation for the age values.
# + papermill={"duration": 0.165499, "end_time": "2022-03-30T13:25:57.581927", "exception": false, "start_time": "2022-03-30T13:25:57.416428", "status": "completed"} tags=[]
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer, make_column_selector
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder, OrdinalEncoder
avg_Age = X_train[(X_train.SibSp + X_train.Parch) == 0].Age.mean()
numerical_transformer = SimpleImputer(strategy='constant', fill_value=avg_Age)
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder()),
('imputer', SimpleImputer(strategy='most_frequent'))
])
preprocessor = ColumnTransformer(transformers=[
('num', numerical_transformer, make_column_selector(dtype_include=[np.integer, np.floating])),
('cat', categorical_transformer, make_column_selector(dtype_include=object))
])
print('Amount of missing values in each column for all data after preprocessing: ')
pd.DataFrame(preprocessor.fit_transform(X_train)).isnull().sum()
# + [markdown] papermill={"duration": 0.024539, "end_time": "2022-03-30T13:25:57.631276", "exception": false, "start_time": "2022-03-30T13:25:57.606737", "status": "completed"} tags=[]
# ## Hypertuning
# + [markdown] papermill={"duration": 0.024688, "end_time": "2022-03-30T13:25:57.681079", "exception": false, "start_time": "2022-03-30T13:25:57.656391", "status": "completed"} tags=[]
# <font size=3>Now that we have our transformers which can handle both missing and categorical data, we can use them during **cross validation** and try to find the best set of parameters for our model.
#
# <font size=3>Since we are dealing with a classification problem, the **XGBoost Classifier** (eXtreme Gradient Boosting) will be chosen as the model. It works by assigning scores to each leaf as it adds trees (CARTs) to the ensemble model. Then, the trees are run based on the input parameters and scores are calculated, so they can be summed to obtain the prediction result.
# + [markdown] papermill={"duration": 0.024454, "end_time": "2022-03-30T13:25:57.779939", "exception": false, "start_time": "2022-03-30T13:25:57.755485", "status": "completed"} tags=[]
# <font size=3>We also have to make a custom function to implement cross validation if we want to make use of XGBoost's **early_stopping_rounds**, since we need access to the validation splits during the cross validation process.
# + papermill={"duration": 0.109271, "end_time": "2022-03-30T13:25:57.915281", "exception": false, "start_time": "2022-03-30T13:25:57.806010", "status": "completed"} tags=[]
import numpy as np
from xgboost import XGBClassifier
from sklearn.model_selection import StratifiedKFold, ParameterGrid
from sklearn.metrics import accuracy_score, roc_auc_score
def XGB_CVGridSearch(estimator, X, y, estimator_params, n_splits=5, verbose=0, vectorize_output=False):
# Searches for the best combination of given parameters for estimator
skf = StratifiedKFold(n_splits=n_splits)
best_params = None
best_score = 0
for params_permut in ParameterGrid(estimator_params):
val_scores = np.empty(skf.n_splits)
train_scores = val_scores.copy()
for i, (train_index, val_index) in enumerate(skf.split(X, y)):
X_train, y_train = X.iloc[train_index], y.iloc[train_index]
X_val, y_val = X.iloc[val_index], y.iloc[val_index]
estimator.set_params(**params_permut)
estimator.fit(X_train, y_train,
model__eval_set=[(preprocessor.transform(X_val), y_val)],
model__early_stopping_rounds=10,
model__verbose=0)
val_scores[i] = estimator.score(X_val, y_val)
train_scores[i] = estimator.score(X_train, y_train)
val_score, train_score = val_scores.mean(), train_scores.mean()
if verbose >= 1:
print("Train score: %f | Validation score: %f" %(train_score, val_score))
if verbose >= 2:
print('Best iteration: ', estimator.get_params()['model'].get_booster().best_ntree_limit)
print('Params: ', params_permut)
if val_score > best_score:
best_params = params_permut
best_params['model__n_estimators'] = estimator.get_params()['model'].get_booster().best_ntree_limit
best_score = val_score
preds = estimator.predict(X_val)
probas = estimator.predict_proba(X_val)[:, 1]
accuracy = accuracy_score(y_val, preds)
roc = roc_auc_score(y_val, probas)
print("Accuracy: ", accuracy, "ROC AUC: ", roc, "\nBest score: ", best_score, ", with parameters: ", best_params)
if vectorize_output == True:
return {key:[value] for key, value in best_params.items()}
else:
return best_params
# + [markdown] papermill={"duration": 0.025161, "end_time": "2022-03-30T13:25:57.966524", "exception": false, "start_time": "2022-03-30T13:25:57.941363", "status": "completed"} tags=[]
# <font size=3>Here comes the **grid search** process, where we try all possible combinations of parameters across our grid. We will be using our custom cross validation function in order to tune the model's **hyperparameters**.
# + papermill={"duration": 4.786534, "end_time": "2022-03-30T13:26:02.778525", "exception": false, "start_time": "2022-03-30T13:25:57.991991", "status": "completed"} tags=[]
from sklearn.model_selection import cross_val_score
my_pipeline = Pipeline(steps=[
('preprocessor', preprocessor),
('model', XGBClassifier(use_label_encoder=False,
objective='binary:logistic',
eval_metric='logloss'))
])
grid_params0 = {
'model__n_estimators': [1000],
'model__learning_rate': [3, 1, 0.1, 0.01],
}
grid_params0 = XGB_CVGridSearch(my_pipeline, X_train, y_train, grid_params0, verbose=1, vectorize_output=True)
# + papermill={"duration": 15.488917, "end_time": "2022-03-30T13:26:18.296703", "exception": false, "start_time": "2022-03-30T13:26:02.807786", "status": "completed"} tags=[]
grid_params1 = grid_params0.copy()
grid_params1.update({
'model__gamma': [0, 0.01, 0.05, 0.1],
'model__max_depth': [8, 10, 12],
'model__min_child_weight': [0.5, 1, 1.5, 3]
})
grid_params1 = XGB_CVGridSearch(my_pipeline, X_train, y_train, grid_params1, verbose=1, vectorize_output=False)
# + papermill={"duration": 0.356044, "end_time": "2022-03-30T13:26:18.737326", "exception": false, "start_time": "2022-03-30T13:26:18.381282", "status": "completed"} tags=[]
my_pipeline.set_params(**grid_params1)
my_pipeline.fit(X_train, y_train, model__verbose=False)
print("Accuracy: ", accuracy_score(y_val, my_pipeline.predict(X_val)))
# + [markdown] papermill={"duration": 0.040669, "end_time": "2022-03-30T13:26:18.847447", "exception": false, "start_time": "2022-03-30T13:26:18.806778", "status": "completed"} tags=[]
# <font size=3>Once the model has been trained, we can use a **ROC curve** to select the best general threshold for it based on F1 scores.
#
# <font size=3>The following function iterates through a list of thresholds and returns the one with the highest F1 score.
# + papermill={"duration": 0.052822, "end_time": "2022-03-30T13:26:18.941424", "exception": false, "start_time": "2022-03-30T13:26:18.888602", "status": "completed"} tags=[]
from sklearn.metrics import confusion_matrix, f1_score, accuracy_score
def GetROCCurve(estimator, X, y_true, thresholds):
tpr, fpr = [], []
best_f1 = 0
final_t = 0
for t in thresholds:
preds = (estimator.predict_proba(X)[:, 1] >= t) * 1
cm = confusion_matrix(y_true, preds)
tn, fp, fn, tp = cm.flatten()
t_tpr = tp / (tp + fn)
t_fpr = fp / (fp + tn)
tpr.append(t_tpr)
fpr.append(t_fpr)
f1 = f1_score(y_true, preds)
print('Accuracy: ', accuracy_score(y_true, preds), 'F1: ', f1, 'for threshold:', t)
if f1 > best_f1:
best_f1 = f1
final_t = t
print('Best threshold: ', final_t)
plt.figure(figsize=(5,5))
plt.suptitle('Roc Curve')
plt.title('Best F1: ' + '{:.6f}'.format(best_f1))
plt.ylabel('TPR')
plt.xlabel('FPR')
plt.plot(fpr, tpr, alpha=0.75)
plt.scatter(fpr, tpr)
return best_f1
# + papermill={"duration": 0.424737, "end_time": "2022-03-30T13:26:19.408018", "exception": false, "start_time": "2022-03-30T13:26:18.983281", "status": "completed"} tags=[]
threshs = [0, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 1]
threshold = GetROCCurve(my_pipeline, X_train, y_train, threshs)
# + [markdown] papermill={"duration": 0.042164, "end_time": "2022-03-30T13:26:19.493733", "exception": false, "start_time": "2022-03-30T13:26:19.451569", "status": "completed"} tags=[]
# ## Generating Predictions
# + [markdown] papermill={"duration": 0.042268, "end_time": "2022-03-30T13:26:19.578254", "exception": false, "start_time": "2022-03-30T13:26:19.535986", "status": "completed"} tags=[]
# <font size=3>Now that we have all of our parameters set up, we can use the whole dataset for training and finally **generate our predictions** to be submitted.
# + papermill={"duration": 0.116264, "end_time": "2022-03-30T13:26:19.737406", "exception": false, "start_time": "2022-03-30T13:26:19.621142", "status": "completed"} tags=[]
my_pipeline.fit(X, y, model__verbose=False)
# + papermill={"duration": 0.060772, "end_time": "2022-03-30T13:26:19.841385", "exception": false, "start_time": "2022-03-30T13:26:19.780613", "status": "completed"} tags=[]
preds = (my_pipeline.predict_proba(X_test)[:, 1] >= threshold) * 1
preds[:20]
# + papermill={"duration": 0.053672, "end_time": "2022-03-30T13:26:19.939215", "exception": false, "start_time": "2022-03-30T13:26:19.885543", "status": "completed"} tags=[]
my_submission = pd.DataFrame({'PassengerId': X_test.index, 'Survived': preds})
my_submission.to_csv('submission.csv', index=False)
# + papermill={"duration": 0.053925, "end_time": "2022-03-30T13:26:20.037255", "exception": false, "start_time": "2022-03-30T13:26:19.983330", "status": "completed"} tags=[]
my_submission.value_counts(subset='Survived')
| xgboost-walkthrough-titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning: A Quantitative Approach
# %matplotlib inline
# +
import sys, csv
import time
import numpy as np
import pandas as pd
from pandas.plotting import scatter_matrix
import matplotlib.pyplot as plt
from time import time
import matplotlib.style as style
from collections import Counter
import seaborn as sns
from sklearn import linear_model
import sklearn.datasets
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import ShuffleSplit
from sklearn.preprocessing import PolynomialFeatures
# -
style.use('seaborn-poster') # sets the size of the charts
style.use('ggplot') # set the theme for matplotlib
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# -
# ### Common
def load_dataframe(data_source, features):
df = pd.read_csv(data_source, low_memory=False)
df_selected_features = df[features]
df_selected_features_filtered = df_selected_features[df_selected_features.displ > 0]
return df_selected_features_filtered
def reshape_data(my_data, features):
X_sorted = my_data.sort_values(by=features)
X_grouped_by_mean = pd.DataFrame({'mean' : \
X_sorted.groupby(features[0])[features[1]].mean()}).reset_index()
X_reshaped = X_grouped_by_mean[features[0]].values.reshape(-1, 1)
y_reshaped = X_grouped_by_mean['mean'].values.reshape(-1, 1)
return X_reshaped, y_reshaped
def split_data(my_data, test_size):
test_set = my_data.sample(frac=test_size, replace=True)
train_set = my_data.sample(frac=(1 - test_size), replace=True)
return train_set, test_set
def score_and_plot (model, test_set_x, test_set_y, test_set_y_pred, i, type):
'''For linear models only.'''
print('Intercept & Coefficients: ', model.intercept_, model.coef_)
mse = mean_squared_error(test_set_y, test_set_y_pred)
rmse = np.sqrt(mse)
r2 = r2_score(test_set_y, test_set_y_pred)
if (type == 0):
cv_mse_scores.append (mse)
cv_rmse_scores.append (rmse)
cv_r2_scores.append (r2)
else:
test_mse_scores.append (mse)
test_rmse_scores.append (rmse)
test_r2_scores.append (r2)
print("Mean squared error: %.2f " % mse, "Root mean squared error: %.2f" % rmse, \
'R-squared score: %.2f' % r2)
# Plot outputs
axarr [type, i].scatter(test_set_x, test_set_y, s=12, color='red')
axarr [type, i].plot(test_set_x, test_set_y_pred, color='blue', linewidth=2)
if (type == 0 and i == 2):
axarr [type, i].set_title ("CV runs: Fuel economy (MPG) versus Engine displacement (liters)")
if (type == 1 and i == 2):
axarr [type, i].set_title ("Test runs: Fuel economy (MPG) versus Engine displacement (liters)")
def split_kfolds(df, k):
k = k
sub_df = []
for i in range(k):
sub_df.append(pd.DataFrame(columns=df.columns))
for i in range(0, len(df), k):
for j in range(k):
x = i + j
if(x < len(df)):
sub_df[j] = sub_df[j].append(df.iloc[x], ignore_index=True)
return sub_df
def make_train_validate_frames(frames, k, k_validate):
validate_frame = frames[k_validate]
train_frame = pd.DataFrame()
for i in range(k -1):
if(i != k_validate):
train_frame = pd.concat([train_frame, frames[i]])
return train_frame.reset_index(), validate_frame
def cross_validate(frames, k, features):
test_frame = frames[k - 1]
for i in range(k - 1):
train_frame, validate_frame = make_train_validate_frames(frames, k, i)
process(train_frame, validate_frame, features, i, 0)
process(train_frame, test_frame, features, i, 1)
def process(train_set, test_set, features, i, type):
train_set_x, train_set_y, = reshape_data(train_set, features)
test_set_x, test_set_y = reshape_data(test_set, features)
poly_features = PolynomialFeatures(degree = 2, include_bias = False)
train_set_x_poly = poly_features.fit_transform(train_set_x)
model = linear_model.LinearRegression()
model.fit(train_set_x_poly, train_set_y)
test_set_x_poly = poly_features.fit_transform(test_set_x)
test_set_y_pred = model.predict(test_set_x_poly)
score_and_plot(model, test_set_x, test_set_y, test_set_y_pred, i, type)
def print_scores():
print ("CV Scores:")
np_mse = np.array(cv_mse_scores)
np_rmse = np.array(cv_rmse_scores)
np_r2 = np.array(cv_r2_scores)
print ("\tCV MSE Scores: ", cv_mse_scores)
print ("\tCV RMSE Scores:", cv_rmse_scores)
print ("\tCV R2 Scores:", cv_r2_scores)
print ("\tCV MSE Scores (mean/std): ", np_mse.mean(), "/", np_mse.std())
print ("\tCV RMSE Scores (mean/std):", np_rmse.mean(), "/", np_rmse.std())
print ("\tCV R2 Scores: (mean/std)", np_r2.mean(), "/", np_r2.std())
print ("Test Scores:")
np_mse = np.array(test_mse_scores)
np_rmse = np.array(test_rmse_scores)
np_r2 = np.array(test_r2_scores)
print ("\tTest MSE Scores:", test_mse_scores)
print ("\tTest RMSE Scores:", test_rmse_scores)
print ("\tTest R2 Scores:", test_r2_scores)
print ("\tTest MSE Scores (mean/std): ", np_mse.mean(), "/", np_mse.std())
print ("\tTest RMSE Scores (mean/std):", np_rmse.mean(), "/", np_rmse.std())
print ("\tTest R2 Scores: (mean/std)", np_r2.mean(), "/", np_r2.std())
# ### Cross-Validation
# +
start = time()
data_source = "datasets/vehicles.csv"
features = ['displ', 'UHighway']
selector = "df_selected_features.displ > 0"
fuel_economy = load_dataframe(data_source, features)
train_set, test_set = split_data(fuel_economy, test_size=0.5)
train_set_x, train_set_y = reshape_data(train_set, features)
test_set_x, test_set_y = reshape_data(test_set, features)
# +
cv_mse_scores = []
cv_rmse_scores = []
cv_r2_scores = []
test_mse_scores = []
test_rmse_scores = []
test_r2_scores = []
# +
start = time()
k = 5
frames = split_kfolds (fuel_economy, k)
f, axarr = plt.subplots (2, 4)
cross_validate (frames, k, features)
print_scores()
print ("total time (seconds): ", (time() - start))
plt.show()
# -
| Vehicles_Cross-Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # BilayerAnalyzer
#
# The BilayerAnalyzer class is the primary tool in the bilayer_analyzer module. It is used to construct a set analyses and (at the moment a limited set) of plot builders. The BilayerAnalyzer can be imported from the bilayer_analyzer module set:
#import the BilayerAnalyzer class
from pybilt.bilayer_analyzer import BilayerAnalyzer
# ## Constructing a BilayerAnalyzer instance
#
# We can then build an analyzer instance and construct our analysis set. The BilayerAnalyzer can be initialized in three ways.
#
# ### 1. via psf_file, trajectory, and selection keyword options.
#initialize analyzer with keyword options--and default analyses
sel_string = "resname POPC or resname DOPE or resname TLCL2"
ba = BilayerAnalyzer(
structure='../pybilt/sample_bilayer/sample_bilayer.psf',
trajectory='../pybilt/sample_bilayer/sample_bilayer_10frames.dcd',
selection=sel_string,
)
# This constructs an analyzer for the given structure (psf_file) and trajectory (trajectory). The selection keyword value is a MDAnalysis selection string that picks out the bilayer lipids from the rest of the system. Although a file path and name are used in this example for the 'trajectory', a list of trajectory files can also be passed to the analyzer.
#
# > Note: Athough the keyword psf_file implies that a CHARMM psf file should be used for the structure file, any valid structure file input to MDAnalysis can be used. See the [topology readers](https://pythonhosted.org/MDAnalysis/documentation_pages/topology/init.html) MDAnalysis page for more details.
#
# > Note: In addition to a filename string, the trajectory keyword argument also accepts a list of filename strings for loading multiple trajectory files.
#
# > Note: Each lipid is assumed to be a unique residue within the provided selection. See the [selections](https://pythonhosted.org/MDAnalysis/documentation_pages/selections.html) page for details on making MDAnalysis selections.
#
# The BilayerAnalyzer is initialized with a single default mean squared displacement analysis (MSD) as shown in the std out text:
# with analysis:
# Analysis: Mean squared displacements
# with analysis_id: msd_1
# and settings:
# leaflet: both
# resname: all
# The MSD analysis has the analysis_id 'msd_1'. Each analysis in the set of analyses are assigned a unique analysis_id, which is used to reference that particular analysis. And we can see that the msd_1 analysis has the settings 'leaflet' and 'resname'. Each analysis may have settings that are initialized with pre-set defaults that(outside of this default MSD analysis) can be user specified.
# #### Listing the valid analyses that can be added to the BilayerAnalyzer instance
# There is a set of analyses that can be defined and assigned as part of the built-in analysis protocols of a BilayerAnalyzer instance. A function in the bilayer_analyzer module is provided to print these to std out:
#let's import the function
from pybilt.bilayer_analyzer import print_valid_analyses
# We can call the function to get a complete list of the built-in analyses available to BilayerAnalyzer instances including the analysis_key, a short statement/description about what the analysis is, and the adjustable settings of that analysis (settings outputs of 'none' means that that analysis has no adjustable settings).
print_valid_analyses()
# Importantly, each type of analysis has a unique analysis_key (e.g. 'msd' and 'apl_box'). The analysis_key is used to specify the analysis type when adding analyses to the analyzer.
# #### 2. via an input file
#
# The analyzer can also be created using an input file with the necessary commands. Let's look at an example: the file 'PyBILT/tests/sample_1.in' which reads :
# > \#set the structure file (psf) for the system
#
# > structure ../sample_bilayer/sample_bilayer.psf
#
# > \#set the trajectory file
#
# > trajectory ../sample_bilayer/sample_bilayer_10frames.dcd
#
# > \#MDAnalysis syntax selection text to select the bilayer
#
# > selection "not resname CLA and not resname TIP3 and not resname POT"
#
# > \#define an analysis for mean squared displacement (msd): named 'msd_1'
#
# > analysis msd msd_1
#
# > \#define an analysis for mean squared displacement (msd) for (resname) POPC lipids in the
# > \#upper leaflet of the bilayer: named 'msd_2'
#
# > analysis msd msd_2 leaflet upper resname POPC
#
# > \#define a plot for mean squared displacement data (msd) including computes 'msd_1' and
# 'msd_2': named 'msd_p'
#
# > plot msd msd_p msd_1 DOPE-U msd_2 POPC-U
#
# In this input script there are five different command types that will be parsed by the analyzer during intialization. The first three 'psf', 'trajectory', and 'selection' are required (similar to their keyword counterparts in initialization 1).
#
# The other two command types used in this input script are the 'analysis' and 'plot'. 'analysis' commands are used to add analyses to the analyzers set of protocols. They hsave the basic format:
#
# > analysis analysis_key analysis_id
#
# and additionally the analysis settings can be set using the format:
#
# > analysis analysis_key analysis_id setting_key value
#
# In the same spirit the 'plot' command is used to add 'auto' plot builders to the set of protocols in the ianalyzer.
#
# >Note: Currently the plotting protocols development is behind that of the analysis protocols and therefore not all analyzer built-in analyses have corresponding built-in plot protocols. Some additional plotting tools are provided in pybilt's plot_generation module (although many of the newer analyses that have been added don't have plot functions in this module either, so will require direct use of matplotlib (or other tool) to generate plots.
#
# The plot command has a very similar format to that of the analysis command with the type of plotting specified by a 'plot_key' and the particular plot identified with a 'plot_id'.
# The plot command in the input script for genrating MSD time series plots has the format:
#
# > plot plot_key plot_id analysis_id legend_name ...
#
# Now let's actually initialize the analyzer using the input script:
ba = BilayerAnalyzer(input_file="../tests/sample_1.in")
# ### 3. Using an input dictionary
#
# Finally, the analyzer can be initialized using an input dictionary. The dictionary should at least have the 3 required keywords 'psf_file', 'trajectory', and 'selection'
# +
# define the input dictionary
input_dict = {'structure' : '../pybilt/sample_bilayer/sample_bilayer.psf',
'trajectory' : '../pybilt/sample_bilayer/sample_bilayer_10frames.dcd',
'selection' : 'resname POPC or resname DOPE or resname TLCL2'
}
#now initialize the analyzer
ba = BilayerAnalyzer(input_dict=input_dict)
# -
# We can also use the input dictionary to initialize the analyzer with analysis and plot protocols.
# +
# define the input dictionary
input_dict = {'structure' : '../pybilt/sample_bilayer/sample_bilayer.psf',
'trajectory' : '../pybilt/sample_bilayer/sample_bilayer_10frames.dcd',
'selection' : 'resname POPC or resname DOPE or resname TLCL2',
'analysis' : ['msd msd_1 resname POPC', 'msd msd_2 resname DOPE'],
'plot' : ['msd msd_p msd_1 POPC msd_2 DOPE']
}
#now initialize the analyzer
ba = BilayerAnalyzer(input_dict=input_dict)
# -
# ### Adding and removing analyses (post initialization)
#
# Analyses can be added to the analyzer using the add_analysis function.
#
# First, let's list the analyses that are present in the analyzer.
ba.print_analysis_protocol()
# As we can see from the output there are two analyses: msd_1 and msd_2. Both are MSD computations; one is for POPC lipids and the other is for DOPE lipids. Now let's add an MSD analysis for the TLCL2 lipids using the string format.
#add the new analysis -- using an input string
ba.add_analysis('msd msd_3 resname TLCL2')
#now reprint the analysis protocols
ba.print_analysis_protocol()
# Analyses can be added using three formats: string, list/tuple, and a dictionary. We've seen the string format above, which is similar to the format used in an input initialization file/script. Now let's add some analyses using the other two options.
# add bilayer thickness analysis using a list -- bilayer_thickness
# has no settings so we just pass an empty dictionary as the third element
ba.add_analysis(['bilayer_thickness', 'bt', {}])
# add area compressibility modulus computation using a dictionary
ba.add_analysis({'analysis_key': 'acm', 'analysis_id': 'acomp',
'analysis_settings':{'temperature':310.0}})
# The must contain three elements: analysis_key string, analysis_id string, and settings dictionary. If the analysis has no adjustable settings, or if you just want to use the defaults, you can pass in empty dictionary.
#
# The dictionary must have three keys: 'analysis_key', 'analysis_id', and 'analysis_settings'. Similar to the list, if the analysis has no adjustable settings, or if you just want to use the defaults, you can pass in empty dictionary as the value for 'analysis_settings'.
#
# Now to remove analyses we can use the remove_analysis function with the anlysis_id of the particular analysis that is to be removed.
# print the protocols
ba.print_analysis_protocol()
#remove the area compressibilty modulus analysis
ba.remove_analysis('acomp')
print("\n ---- after removal ---- \n")
#reprint the protocols
ba.print_analysis_protocol()
# ### Adding and removing plots to the analyzer
#
# Similar to the analyses, plots can be added to the analysis using an 'add_plot' function. However, the add_plot function currently only accepts an input string.
#
# First lets list the availble plot protocols.
# +
# import the function from the bilayer analyzer module
from pybilt.bilayer_analyzer.bilayer_analyzer import print_available_plots
print_available_plots()
# -
# and let's double check what plots are in the analyzer.
ba.print_plot_protocol()
# Now we can add some more plots.
# add an MSD plot
ba.add_plot('msd msd_all msd_1 POPC msd_2 DOPE msd_3 CL')
#add a bilayer thickness plot
ba.add_plot('bilayer_thickness bt_p bt NONE')
# print the plot protocols
ba.print_plot_protocol()
# Let's remove the extra MSD plot 'msd_p'.
#remove msd_p plot from the analyzer
ba.remove_plot('msd_p')
# print the plot protocols
ba.print_plot_protocol()
# ### Running the analyses on the trajectory
#
# Once we have all analyses added to the analyzer, we can run the analyses. There are two options for running the analyses.
#
# #### 1. run_analysis function
#
# The first way to initiate the analyses is using the run_analysis function
ba.run_analysis()
# #### 2. analyzer iterator
#
# The second way to run the analyses is to iterate over the analyzer
# first we need to reset the analyzer before rerunning the analyses
ba.reset()
# now run the analysis iterator
for _frame in ba:
print(ba.reps['com_frame'])
print(ba.reps['current_mda_frame'])
print(" ")
# ### Getting analysis data/output and showing plots
#
# Once the analysis loop has been run, data can be extracted from the analyzer and plots can be generated.
#get the data for msd_1
print(ba.get_analysis_data('msd_1'))
#notebook magic
# %matplotlib inline
# show the msd_all plot
ba.show_plot('msd_all')
| jupyter_notebooks/bilayer_analyzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activation Test MEA Standard Protocol (Repeat)
import os
import numpy as np
import matplotlib.pyplot as plt
from functions import *
path = os.path.join("../","Activation Test MEA Standard Protocol (Repeat)")
# ## 1- Impedance : End of Each Activation Set
data = load_data(os.path.join(path,"1.csv"),set_flag=True)
impedance_plot1(data,set_index=1)
impedance_plot1(data,set_index=2)
# ## 2- Impedance : Various Voltages (1)
data = load_data(os.path.join(path,"2.csv"))
impedance_plot2(data,RH=30,P=5)
impedance_plot2(data,V=0.3,P=25)
impedance_plot2(data,RH=30,V=0.3)
# ## 3- Impedance : Various Voltages (2)
data = load_data(os.path.join(path,"3.csv"))
impedance_plot2(data,RH=30,P=5)
impedance_plot2(data,V=0.3,P=25)
impedance_plot2(data,RH=30,V=0.7)
# ## 4- Polarization : End of Activation Procedure (1)
data = load_data(os.path.join(path,"4.csv"))
polarization_plot1(data,RH=30,P=5)
polarization_plot1(data,RH=30,P=25)
# ## 5- Polarization : End of Activation Procedure (2)
data = load_data(os.path.join(path,"5.csv"))
polarization_plot1(data,RH=30,P=5)
polarization_plot1(data,RH=30,P=25)
# ## 6- Polarization : End of Each Activation Set
data = load_data(os.path.join(path,"6.csv"))
polarization_plot2(data,set_index=2)
polarization_plot2(data,set_index=[1,2,3,4,5])
| Notebooks/Activation Test MEA Standard Protocol (Repeat).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
### Synthetic ones ###
import csv
import pandas as pd
import numpy as np
from sklearn.utils import shuffle
import matplotlib.pyplot as plt
path = 'ToySyntheticRecordData-1510619474'
number = 1510619474
iteration = np.arange(0,126500,500)
t = np.linspace(0,60,len(iteration))
original_totals = []
totals = []
for it in iteration:
print(it)
data = pd.read_csv('data/Synthetic_OneSetofParams/{:s}/run_{:d}_0.csv'.format(path,it))
original_rewards = data['original_rewards']
rewards = data['rewards']
indiv_original_rewards = np.zeros(len(original_rewards))
indiv_rewards = np.zeros(len(rewards))
total_rewards = 0
for i in range(len(rewards)):
total_rewards += rewards[i]
indiv_rewards[i] = total_rewards
totals.append(total_rewards)
total_original_rewards = 0
for i in range(len(original_rewards)):
total_original_rewards += original_rewards[i]
indiv_original_rewards[i] = total_original_rewards
original_totals.append(total_original_rewards)
fig1, ax1 = plt.subplots()
ax1.plot(t,totals,label='Synthesis')
ax1.set_title('Synthetic Training Results')
ax1.set_xlabel('Time(m)')
ax1.set_ylabel('Rewards')
ax1.legend()
ax1.set_xlim([-1, 61])
ax1.grid()
fig1.savefig('data/Synthetic_OneSetofParams/{:s}/Synthetic_ones_total_({:d}).png'.format(path,number,max(iteration)))
fig2, ax2 = plt.subplots()
ax2.plot(t,original_totals,label='Synthesis')
ax2.set_title('Synthetic Training Results')
ax2.set_xlabel('Time(m)')
ax2.set_ylabel('Rewards')
ax2.legend()
ax2.set_xlim([-1, 61])
ax2.grid()
fig2.savefig('data/Synthetic_OneSetofParams/{:s}/Synthetic_ones_original_({:d}).png'.format(path,number,max(iteration)))
# -
| Report/Data/Plots/Codes/synthetic_ones_indiv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/razzlestorm/DS-Unit-3-Sprint-2-SQL-and-Databases/blob/master/module1-introduction-to-sql/Unit_3_Sprint_2_Homework_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g4Hyc7C0pSvE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="9000bf44-2b7c-40df-db40-44a28ae52b00"
import pandas as pd
import sqlite3
# !wget https://github.com/razzlestorm/DS-Unit-3-Sprint-2-SQL-and-Databases/blob/master/module1-introduction-to-sql/rpg_db.sqlite3?raw=true
# + id="Bz_b3emVpdBq" colab_type="code" colab={}
conn = sqlite3.connect('/content/rpg_db.sqlite3?raw=true')
cur = conn.cursor()
def print_data(query):
cur.execute(query)
return cur.fetchall()[0][0]
# + id="jSnWXKP2prws" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3a72cd18-5919-4739-a197-da93de201e02"
# How many total Characters are there?
query = """
SELECT COUNT(character_id)
FROM charactercreator_character
"""
print_data(query)
# + id="W4mHgwwnqG43" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="85ad436d-5584-41df-f1bb-a63ad9a2db06"
# How many of each specific subclass?
class_list = ['charactercreator_cleric', 'charactercreator_fighter', 'charactercreator_mage', 'charactercreator_necromancer', 'charactercreator_thief']
def sql_table_count_loop(li):
for xx in li:
query = """
SELECT COUNT(*)
FROM {}
""".format(xx)
cur.execute(query)
print(f'The {xx[17:]} class has {cur.fetchall()[0][0]} rows.')
sql_table_count_loop(class_list)
# + id="pao3Ba53qIGL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1af86e11-f773-4d96-f84b-944983366a46"
# How many total Items?
item_count = """
SELECT COUNT(*)
FROM armory_item
"""
print_data(item_count)
# + id="38OkianAqJAJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="3ee0efa2-0858-48fb-b6fc-57d99342777f"
# How many of the Items are weapons? How many are not?
weapon_count = """
SELECT COUNT(item_id)
FROM armory_item
WHERE item_id IN (SELECT item_ptr_id FROM armory_weapon aw)
"""
non_weapon_count = """
SELECT COUNT(item_id)
FROM armory_item
WHERE item_id NOT IN (SELECT item_ptr_id FROM armory_weapon aw)
"""
print(print_data(weapon_count))
print(print_data(non_weapon_count))
# + id="QaJzM3YyqJ77" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="f373a70b-794c-44d4-8f32-1c558fb6206a"
# How many Items does each character have? (Return first 20 rows)
character_item_count = """
SELECT character_id, COUNT(item_id)
FROM charactercreator_character_inventory
WHERE item_id NOT IN (SELECT item_ptr_id FROM armory_weapon aw)
GROUP BY character_id
LIMIT 20
"""
cur.execute(character_item_count)
cur.fetchall()
# + id="Wy0zHJ5kqLAD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="44caa5d6-cc17-454b-cfe6-ae15e8102105"
# How many Weapons does each character have? (Return first 20 rows)
character_weapon_count = """
SELECT character_id, COUNT(item_id)
FROM charactercreator_character_inventory
WHERE item_id IN (SELECT item_ptr_id FROM armory_weapon aw)
GROUP BY character_id
LIMIT 20
"""
cur.execute(character_weapon_count)
cur.fetchall()
# + id="80fA6HlPqL7L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c4580c61-7f02-42c0-e2ec-7cbaa09193b8"
# On average, how many Items does each Character have?
character_avg_items = """
SELECT AVG(counted_items)
FROM (
SELECT COUNT(item_id) as counted_items
FROM charactercreator_character_inventory
WHERE item_id NOT IN (SELECT item_ptr_id FROM armory_weapon aw)
GROUP BY character_id
)
"""
cur.execute(character_avg_items)
cur.fetchall()
# + id="K6LDoQ9JqMxc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d30cda38-4518-4d16-c1cc-041ba62c70f6"
# On average, how many Weapons does each character have?
character_avg_weapons = """
SELECT AVG(counted_items)
FROM (
SELECT COUNT(item_id) as counted_items
FROM charactercreator_character_inventory
WHERE item_id IN (SELECT item_ptr_id FROM armory_weapon aw)
GROUP BY character_id
)
"""
cur.execute(character_avg_weapons)
cur.fetchall()
# + id="CdsNiFIietdB" colab_type="code" outputId="331b3e0d-cda3-48dc-e2dd-fa4c1407eaa6" colab={"base_uri": "https://localhost:8080/", "height": 195}
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/00476/buddymove_holidayiq.csv')
df.head()
# + id="KiTm2xr97QgT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="295ae380-ec09-43de-c48f-a99d343bad56"
conn_new = sqlite3.connect('buddymove_holidayiq.sqlite3')
df2 = df.to_sql('my_table', conn_new)
# + id="OnfGovYD8SfD" colab_type="code" colab={}
newcur = conn_new.cursor()
# + id="oRh_pAuj8gpN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f4623d67-b6e1-4e0b-ecb3-28483c8d01ee"
newcur.execute('SELECT * FROM my_table')
print(newcur.fetchall()[0][6]) # 4= Nature, 6= Shopping
# + id="BD2OD4Ua8sub" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7bcbec95-a0c4-4ed9-e1a0-18b7938a6f81"
row_query = """
SELECT COUNT(*)
FROM
(SELECT *
FROM my_table
WHERE Nature >= 100)
WHERE Shopping >= 100
"""
newcur.execute(row_query)
newcur.fetchall()
| module1-introduction-to-sql/Unit_3_Sprint_2_Homework_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ###### Content under Creative Commons Attribution license CC-BY 4.0, code under BSD 3-Clause License © 2017 <NAME>, <NAME>
# # Life expectancy and wealth
#
# Welcome to **Lesson 4** of the second module in _Engineering Computations_. This module gives you hands-on data analysis experience with Python, using real-life applications. The first three lessons provide a foundation in data analysis using a computational approach. They are:
#
# 1. [Lesson 1](http://go.gwu.edu/engcomp2lesson1): Cheers! Stats with beers.
# 2. [Lesson 2](http://go.gwu.edu/engcomp2lesson2): Seeing stats in a new light.
# 3. [Lesson 3](http://go.gwu.edu/engcomp2lesson3): Lead in lipstick.
#
# You learned to do exploratory data analysis with data in the form of arrays: NumPy has built-in functions for many descriptive statistics, making it easy! And you also learned to make data visualizations that are both good-looking and effective in communicating and getting insights from data.
#
# But NumPy can't do everything. So we introduced you to `pandas`, a Python library written _especially_ for data analysis. It offers a very powerful new data type: the _DataFrame_—you can think of it as a spreadsheet, conveniently stored in one Python variable.
#
# In this lesson, you'll dive deeper into `pandas`, using data for life expectancy and per-capita income over time, across the world.
# ## The best stats you've ever seen
#
# [<NAME>](https://en.wikipedia.org/wiki/Hans_Rosling) was a professor of international health in Sweden, until his death in Februrary of this year. He came to fame with the thrilling TED Talk he gave in 2006: ["The best stats you've ever seen"](https://www.ted.com/talks/hans_rosling_shows_the_best_stats_you_ve_ever_seen) (also on [YouTube](https://youtu.be/RUwS1uAdUcI), with ads). We highly recommend that you watch it!
#
# In that first TED Talk, and in many other talks and even a BBC documentary (see the [trailer](https://youtu.be/jbkSRLYSojo) on YouTube), Rosling uses data visualizations to tell stories about the world's health, wealth, inequality and development. Using software, he and his team created amazing animated graphics with data from the United Nations and World Bank.
#
# According to a [blog post](https://www.gatesnotes.com/About-Bill-Gates/Remembering-Hans-Rosling) by Bill and <NAME> after <NAME>'s death, his message was simple: _"that the world is making progress, and that policy decisions should be grounded in data."_
# In this lesson, we'll use data about life expectancy and per-capita income (in terms of the gross domestic product, GDP) around the world. Visualizing and analyzing the data will be our gateway to learning more about the world we live in.
#
# Let's begin! As always, we start by importing the Python libraries for data analysis (and setting some plot parameters).
# +
import numpy
import pandas
from matplotlib import pyplot
# %matplotlib inline
#set font style
pyplot.rc('font', family='serif', size=16)
# -
# ## Load and inspect the data
#
# We found a website called [The Python Graph Gallery](https://python-graph-gallery.com), which has a lot of data visualization examples.
# Among them is a [Gapminder Animation](https://python-graph-gallery.com/341-python-gapminder-animation/), an animated GIF of bubble charts in the style of Hans Rosling.
# We're not going to repeat the same example, but we do get some ideas from it and re-use their data set.
# The data file is hosted on their website, and we can read it directly from there into a `pandas` dataframe, using the URL.
# Read a dataset for life expectancy from a CSV file hosted online
url = 'https://python-graph-gallery.com/wp-content/uploads/gapminderData.csv'
life_expect = pandas.read_csv(url)
# The first thing to do always is to take a peek at the data.
# Using the `shape` attribute of the dataframe, we find out how many rows and columns it has. In this case, it's kind of big to print it all out, so to save space we'll print a small portion of `life_expect`.
# You can use a slice to do this, or you can use the [`DataFrame.head()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.head.html) method, which returns by default the first 5 rows.
life_expect.shape
life_expect.head()
# You can see that the columns hold six types of data: the country, the year, the population, the continent, the life expectancy, and the per-capita gross domestic product (GDP).
# Rows are indexed from 0, and the columns each have a **label** (also called an index). Using labels to access data is one of the most powerful features of `pandas`.
#
# In the first five rows, we see that the country repeats (Afghanistan), while the year jumps by five. We guess that the data is arranged in blocks of rows for each country.
# We can get a useful summary of the dataframe with the [`DataFrame.info()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.info.html) method: it tells us the number of rows and the number of columns (matching the output of the `shape` attribute) and then for each column, it tells us the number of rows that are populated (have non-null entries) and the type of the entries; finally it gives a breakdown of the types of data and an estimate of the memory used by the dataframe.
life_expect.info()
# The dataframe has 1704 rows, and every column has 1704 non-null entries, so there is no missing data. Let's find out how many entries of the same year appear in the data.
# In [Lesson 1](http://go.gwu.edu/engcomp2lesson1) of this module, you already learned to extract a column from a data frame, and use the [`series.value_counts()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.value_counts.html) method to answer our question.
life_expect['year'].value_counts()
# We have an even 142 occurrences of each year in the dataframe. The distinct entries must correspond to each country. It also is clear that we have data every five years, starting 1952 and ending 2007. We think we have a pretty clear picture of what is contained in this data set. What next?
# ## Grouping data for analysis
#
# We have a dataframe with a `country` column, where countries repeat in blocks of rows, and a `year` column, where sets of 12 years (increasing by 5) repeat for every country. Tabled data commonly has this interleaved structure. And data analysis often involves grouping the data in various ways, to transform it, compute statistics, and visualize it.
#
# With the life expectancy data, it's natural to want to analyze it by year (and look at geographical differences), and by country (and look at historical differences).
#
# In [Lesson 2](http://go.gwu.edu/engcomp2lesson2) of this module, we already learned how useful it was to group the beer data by style, and calculate means within each style. Let's get better acquainted with the powerful `groupby()` method for dataframes. First, grouping by the values in the `year` column:
by_year = life_expect.groupby('year')
type(by_year)
# Notice that the type of the new variable `by_year` is different: it's a _GroupBy_ object, which—without making a copy of the data—is able to apply operations on each of the groups.
# The [`GroupBy.first()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.core.groupby.GroupBy.first.html) method, for example, returns the first row in each group—applied to our grouping `by_year`, it shows the list of years (as a label), with the first country that appears in each year-group.
by_year.first()
# All the year-groups have the same first country, Afghanistan, so what we see is the population, life expectancy and per-capita income in Afghanistan for all the available years.
# Let's save that into a new dataframe, and make a line plot of the population and life expectancy over the years.
Afghanistan = by_year.first()
Afghanistan['pop'].plot(figsize=(8,4),
title='Population of Afghanistan');
Afghanistan['lifeExp'].plot(figsize=(8,4),
title='Life expectancy of Afghanistan');
# Do you notice something interesting? It's curious to see that the population of Afghanistan took a fall after 1977. We have data every 5 years, so we don't know exactly when this fall began, but it's not hard to find the answer online. The USSR invaded Afghanistan in 1979, starting a conflict that lasted 9 years and resulted in an estimated death toll of one million civilians and 100,000 fighters [1]. Millions fled the war to neighboring countries, which may explain why we se a dip in population, but not a dip in life expectancy.
# We can also get some descriptive statistics in one go with the [`DataFrame.describe()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.describe.html) method of `pandas`.
Afghanistan.describe()
# Let's now group our data by country, and use the `GroupBy.first()` method again to get the first row of each group-by-country. We know that the first year for which we have data is 1952, so let's immediately save that into a new variable named `year1952`, and keep playing with it. Below, we double-check the type of `year1952`, print the first five rows using the `head()` method, and get the minimum value of the population column.
by_country = life_expect.groupby('country')
# The first year for all groups-by-country is 1952. Let's save that first group into a new dataframe, and keep playing with it.
year1952 = by_country.first()
type(year1952)
year1952.head()
year1952['pop'].min()
# ## Visualizing the data
#
# In [Lesson 2](http://go.gwu.edu/engcomp2lesson2) of this module, you learned to make bubble charts, allowing you to show at least three features of the data in one plot. We'd like to make a bubble chart of life expectancy vs. per-capita GDP, with the size of the bubble proportional to the population. To do that, we'll need to extract the population values into a NumPy array.
populations = year1952['pop'].values
# If you use the `populations` array unmodified as the size of the bubbles, they come out _huge_ and you get one solid color covering the figure (we tried it!). To make the bubble sizes reasonable, we divide by 60,000—an approximation to the minimum population—so the smallest bubble size is about 1 pt. Finally, we choose a logarithmic scale in the absissa (the GDP). Check it out!
year1952.plot.scatter(figsize=(12,8),
x='gdpPercap', y='lifeExp', s=populations/60000,
title='Life expectancy in the year 1952',
edgecolors="white")
pyplot.xscale('log');
# That's neat! But the Rosling bubble charts include one more feature in the data: the continent of each country, using a color scheme. Can we do that?
# Matplotlib [colormaps](https://matplotlib.org/examples/color/colormaps_reference.html) offer several options for _qualitative_ data, using discrete colors mapped to a sequence of numbers. We'd like to use the `Accent` colormap to code countries by continent. But we need a numeric code to assign to each continent, so it can be mapped to a color.
#
# The [Gapminder Animation](https://python-graph-gallery.com/341-python-gapminder-animation/) example at The Python Graph Gallery has a good tip: using the `pandas` _Categorical_ data type, which associates a numerical value for each category in a column containing qualitative (categorical) data.
#
# Let's see what we get if we apply `pandas.Categorical()` to the `continent` column:
pandas.Categorical(year1952['continent'])
# Right. We see that the `continent` column has repeated entries of 5 distinct categories, one for each continent. In order, they are: Africa, Americas, Asia, Europe, Oceania.
#
# Applying [`pandas.Categorical()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Categorical.html) to the `continent` column will create an integer value—the _code_ of the category—associated to each entry. We can then use these integer values to map to the colors in a colormap. The trick will be to extract the `codes` attribute of the _Categorical_ data and save that into a new variable named `colors` (a NumPy array).
colors = pandas.Categorical(year1952['continent']).codes
type(colors)
len(colors)
print(colors)
# You see that `colors` is a NumPy array of 142 integers that can take the values: ` 0, 1, 2, 3, 4`. They are the codes to `continent` categories: `Africa, Americas, Asia, Europe, Oceania`. For example, the first entry is `2`, corresponding to Asia, the continent of Afghanistan.
# Now we're ready to re-do our bubble chart, using the array `colors` to set the color of the bubble (according to the continent for the given country).
year1952.plot.scatter(figsize=(12,8),
x='gdpPercap', y='lifeExp', s=populations/60000,
c=colors, cmap='Accent',
title='Life expectancy vs. per-capita GDP in the year 1952,\n color-coded by continent',
logx = True,
ylim = (25,85),
xlim = (1e2, 1e5),
edgecolors="white",
alpha=0.6);
# ##### Note:
#
# We encountered a bug in `pandas` scatter plots! The labels of the $x$-axis disappeared when we added the colors to the bubbles. We tried several things to fix it, like adding the line `pyplot.xlabel("GDP per Capita")` at the end of the cell, but nothing worked. Searching online, we found an open [issue report](https://github.com/pandas-dev/pandas/issues/10611) for this problem.
#
# ##### Discuss with your neighbor:
#
# What do you see in the colored bubble chart, in regards to 1952 conditions in different countries and different continents?
# Can you guess some countries? Can you figure out which color corresponds to which continent?
# ### Spaghetti plot of life expectancy
#
# The bubble plot shows us that 1952 life expectancies varied quite a lot from country to country: from a minimum of under 30 years, to a maximum under 75 years. The first part of Prof. Rosling's dying message is _"that the world is making progress_." Is it the case that countries around the world _all_ make progress in life expectancy over the years?
#
# We have an idea: what if we plot a line of life expectancy over time, for every country in the data set? It could be a bit messy, but it may give an _overall view_ of the world-wide progress in life expectancy.
#
# Below, we'll make such a plot, with 142 lines: one for each country. This type of graphic is called a **spaghetti plot** …for obvious reasons!
#
# To add a line for each country on the same plot, we'll use a `for`-statement and the `by_country` groups. For each country-group, the line plot takes the series `year` and `lifeExp` as $(x,y)$ coordinates. Since the spaghetti plot is quite busy, we also took off the box around the plot. Study this code carefully.
# +
pyplot.figure(figsize=(12,8))
for key,group in by_country:
pyplot.plot(group['year'], group['lifeExp'], alpha=0.4)
pyplot.title('Life expectancy in the years 1952–2007, across 142 countries')
pyplot.box(on=None);
# -
# ## Dig deeper and get insights from the data
#
# The spaghetti plot shows a general upwards tendency, but clearly not all countries have a monotonically increasing life expectancy. Some show a one-year sharp drop (but remember, this data jumps every 5 years), while others drop over several years.
# And something catastrophic happened to one country in 1977, and to another country in 1992.
# Let's investigate this!
#
# We'd like to explore the data for a particular year: first 1977, then 1992. For those years, we can get the minimum life expectancy, and then find out which country experienced it.
#
# To access a particular group in _GroupBy_ data, `pandas` has a `get_group(key)` method, where `key` is the label of the group.
# For example, we can access yearly data from the `by_year` groups using the year as key. The return type will be a dataframe, containing the same columns as the original data.
type(by_year.get_group(1977))
type(by_year['lifeExp'].get_group(1977))
# Now we can find the minimum value of life expectancy at the specific years of interest, using the `Series.min()` method. Let' do this for 1977 and 1992, and save the values in new Python variables, to reuse later.
min_lifeExp1977 = by_year['lifeExp'].get_group(1977).min()
min_lifeExp1977
min_lifeExp1992 = by_year['lifeExp'].get_group(1992).min()
min_lifeExp1992
# Those values of life expectancy are just terrible! Are you curious to know what countries experienced the dramatic drops in life expectancy?
# We can find the row _index_ of the minimum value, thanks to the [`pandas.Series.idxmin()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.idxmin.html) method. The row indices are preserved from the original dataframe `life_expect` to its groupings, so the index will help us identify the country. Check it out.
by_year['lifeExp'].get_group(1977).idxmin()
life_expect['country'][221]
by_country.get_group('Cambodia')
# We searched online to learn what was happening in Cambodia to cause such a drop in life expectancy in the 1970s. Indeed, Cambodia experienced a _mortality crisis_ due to several factors that combined into a perfect storm: war, ethnic cleansing and migration, collapse of the health system, and cruel famine [2].
# It's hard for a country to keep vital statistics under such circumstances, and certainly the data for Cambodia in the 1970s is uncertain.
# However, various sources report a life expectancy there in 1977 that was _under 20 years_.
# See, for example, the World Bank's interactive web page on [Cambodia](https://data.worldbank.org/country/cambodia).
# Something is strange about the data from the The Python Graph Gallery. Is it wrong?
# Maybe they are giving us _average_ life expectancy in a five-year period.
# Let's look at the other dip in life expectancy, in 1992.
by_year['lifeExp'].get_group(1992).idxmin()
life_expect['country'][1292]
by_country.get_group('Rwanda')
# The World Bank's interactive web page on [Rwanda](https://data.worldbank.org/country/rwanda) gives a life expectancy of 28.1 in 1992, and even lower in 1993, at 27.6 years.
# This doesn't match the value from the data set we sourced from The Python Graph Gallery, which gives 23.6—and since this value is _lower_ than the minimum value given by the World Bank, we conclude that the discepancy is not caused by 5-year averaging.
# ## Checking data quality
#
# All our work here started with loading a data set we found online. What if this data set has _quality_ problems?
#
# Well, nothing better than asking the author of the web source for the data. We used Twitter to communicate with the author of The Python Graph Gallery, and he replied with a link to _his source_: a data package used for teaching a course in Exploratory Data Analysis at the University of British Columbia.
# + language="html"
# <blockquote class="twitter-tweet" data-lang="en"><p lang="en" dir="ltr">Hi. Didn't receive your email... Gapminder comes from this R library: <a href="https://t.co/BU1IFIGSxm">https://t.co/BU1IFIGSxm</a>. I will add citation asap.</p>— R+Py Graph Galleries (@R_Graph_Gallery) <a href="https://twitter.com/R_Graph_Gallery/status/920074231269941248?ref_src=twsrc%5Etfw">October 16, 2017</a></blockquote> <script async src="//platform.twitter.com/widgets.js" charset="utf-8"></script>
# -
# Note one immediate outcome of our reaching out to the author of The Python Graph Gallery: he realized he was not citing the source of his data [3], and promised to add proper credit. _It's always good form to credit your sources!_
#
# We visited the online repository of the data source, and posted an [issue report](https://github.com/jennybc/gapminder/issues/18) there, with our questions about data quality. The author promptly responded, saying that _her_ source was the [Gapminder.org website](http://www.gapminder.org/data/)—**Gapminder** is the non-profit founded by <NAME> to host public data and visualizations. She also said: _" I don't doubt there could be data quality problems! It should definitely NOT be used as an authoritative source for life expectancy"_
#
# So it turns out that the data we're using comes from a set of tools meant for teaching, and is not up-to-date with the latest vital statistics. The author ended up [adding a warning](https://github.com/jennybc/gapminder/commit/7b3ac7f477c78f21865fa7defea20e72cb9e2b8a) to make this clear to visitors of the repository on GitHub.
#
# #### This is a wonderful example of how people collaborate online via the open-source model.
#
# ##### Note:
#
# For the most accurate data, you can visit the website of the [World Bank](https://data.worldbank.org).
# ## Using widgets to visualize interactively
#
# One more thing! This whole exploration began with our viewing the 2006 TED Talk by <NAME>: ["The best stats you've ever seen"](https://www.ted.com/talks/hans_rosling_shows_the_best_stats_you_ve_ever_seen). One of the most effective parts of the presentation is seeing the _animated_ bubble chart, illustrating how countries became healthier and richer over time. Do you want to make something like that?
#
# You can! Introducing [Jupyter Widgets](https://ipywidgets.readthedocs.io/en/latest/user_guide.html). The magic of interactive widgets is that they tie together the running Python code in a Jupyter notebook with Javascript and HTML running in the browser. You can use widgets to build interactive controls on data visualizations, with buttons, sliders, and more.
#
# To use widgets, the first step is to import the `widgets` module.
from ipywidgets import widgets
# After importing `widgets`, you have available several UI (User Interaction) elements. One of our favorites is a _Slider_: an interactive sliding button. Here is a default slider that takes integer values, from 0 to 100 (but does nothing):
widgets.IntSlider()
# What we'd like to do is make an interactive visualization of bubble charts, with the year in a slider, so that we can run forwards and backwards in time by sliding the button, watching our plot update the bubbles in real time. Sound like magic? It almost is.
#
# The magic happens when you program what should happen when the value in the slider changes. A typical scenario is having a function that is executed with the value in the slider, interactively. To create that, we need two things:
#
# 1. A function that will be called with the slider values, and
# 2. A call to an _interaction_ function from the `ipywidgets` package.
#
# Several interaction functions are available, for different actions you expect from the user: a click, a text entered in a box, or sliding the button on a slider.
# You will need to explore the Jupyter Widgets documentation [4] to learn more.
#
# For this example, we'll be using a slider, a plotting function that makes our bubble chart, and the [`.interact()`](http://ipywidgets.readthedocs.io/en/stable/examples/Using%20Interact.html#) function to call our plotting function with each value of the slider.
#
# We do everything in one cell below. The first line creates an integer-value slider with our known years—from a minimum 1952, to a maximum 2007, stepping by 5—and assigns it to the variable name `slider`.
#
# Next, we define the function `roslingplot()`, which re-calculates the array of population values, gets the year-group we need from the `by_year` _GroupBy_ object, and makes a scater plot of life expectancy vs. per-capita income, like we did above. The `populations` array (divided by 60,000) sets the size of the bubble, and the previously defined `colors` array sets the color coding by continent.
#
# We also removed the colorbar (which added little information), and added the option `sharex=False` following the workaround suggested by someone on the open [issue report](https://github.com/pandas-dev/pandas/issues/10611) for the plotting bug we mentioned above.
#
# The last line in the cell below is a call to `.interact()`, passing our plotting function and the slider value assigned to its argument, `year`. Watch the magic happen!
# +
slider = widgets.IntSlider(min=1952, max=2007, step=5)
def roslingplot(year):
populations = by_year.get_group(year)['pop'].values
by_year.get_group(year).plot.scatter(figsize=(12,8),
x='gdpPercap', y='lifeExp', s=populations/60000,
c=colors, cmap='Accent',
title='Life expectancy vs per-capita GDP in the year '+ str(year)+'\n',
logx = True,
ylim = (25,85),
xlim = (1e2, 1e5),
edgecolors="white",
alpha=0.6,
colorbar=False,
sharex=False)
pyplot.show();
widgets.interact(roslingplot, year=slider);
# -
# ## References
#
# 1. [The Soviet War in Afghanistan, 1979-1989](https://www.theatlantic.com/photo/2014/08/the-soviet-war-in-afghanistan-1979-1989/100786/), The Atlantic (2014), by <NAME>.
#
# 2. US National Research Council Roundtable on the Demography of Forced Migration; <NAME>, <NAME>, editors. Forced Migration & Mortality (2001), National Academies Press, Washington DC; Chapter 5: The Demographic Analysis of Mortality Crises: The Case of Cambodia, 1970-1979, <NAME>. Available at: https://www.ncbi.nlm.nih.gov/books/NBK223346/
#
# 3. gapminder: Data from Gapminder (R data package), by Jennifer (<NAME>, repository at https://github.com/jennybc/gapminder, v0.3.0 (Version v0.3.0) on Zenodo: https://doi.org/10.5281/zenodo.594018, licensed CC-BY 3.0
#
# 4. [Jupyter Widgets User Guide](https://ipywidgets.readthedocs.io/en/latest/user_guide.html)
# Execute this cell to load the notebook's style sheet, then ignore it
from IPython.core.display import HTML
css_file = '../style/custom.css'
HTML(open(css_file, "r").read())
| notebooks_en/4_Life_Expectancy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Vw7RpVEfZand"
# IMPORTING NECESSARY LIBRARIES
from IPython.display import display, Javascript, Image
from google.colab.output import eval_js
from google.colab.patches import cv2_imshow
from base64 import b64decode, b64encode
import cv2
import numpy as np
# + id="GKgml8S5Sgdz"
def video_stream():
js = Javascript('''
var video;
var div = null;
var stream;
var captureCanvas;
var imgElement;
var labelElement;
var pendingResolve = null;
var shutdown = false;
function removeDom() {
stream.getVideoTracks()[0].stop();
video.remove();
div.remove();
video = null;
div = null;
stream = null;
imgElement = null;
captureCanvas = null;
labelElement = null;
}
function onAnimationFrame() {
if (!shutdown) {
window.requestAnimationFrame(onAnimationFrame);
}
if (pendingResolve) {
var result = "";
if (!shutdown) {
captureCanvas.getContext('2d').drawImage(video, 0, 0, 640, 480);
result = captureCanvas.toDataURL('image/jpeg', 0.8)
}
var lp = pendingResolve;
pendingResolve = null;
lp(result);
}
}
async function createDom() {
if (div !== null) {
return stream;
}
div = document.createElement('div');
div.style.border = '2px solid black';
div.style.padding = '3px';
div.style.width = '100%';
div.style.maxWidth = '600px';
document.body.appendChild(div);
const modelOut = document.createElement('div');
modelOut.innerHTML = "<span>Status:</span>";
labelElement = document.createElement('span');
labelElement.innerText = 'No data';
labelElement.style.fontWeight = 'bold';
modelOut.appendChild(labelElement);
div.appendChild(modelOut);
video = document.createElement('video');
video.style.display = 'block';
video.width = div.clientWidth - 6;
video.setAttribute('playsinline', '');
video.onclick = () => { shutdown = true; };
stream = await navigator.mediaDevices.getUserMedia(
{video: { facingMode: "environment"}});
div.appendChild(video);
imgElement = document.createElement('img');
imgElement.style.position = 'absolute';
imgElement.style.zIndex = 1;
imgElement.onclick = () => { shutdown = true; };
div.appendChild(imgElement);
const instruction = document.createElement('div');
instruction.innerHTML =
'<span style="color: red; font-weight: bold;">' +
'When finished, click here or on the video to stop this demo</span>';
div.appendChild(instruction);
instruction.onclick = () => { shutdown = true; };
video.srcObject = stream;
await video.play();
captureCanvas = document.createElement('canvas');
captureCanvas.width = 640; //video.videoWidth;
captureCanvas.height = 480; //video.videoHeight;
window.requestAnimationFrame(onAnimationFrame);
return stream;
}
async function stream_frame(label, imgData) {
if (shutdown) {
removeDom();
shutdown = false;
return '';
}
var preCreate = Date.now();
stream = await createDom();
var preShow = Date.now();
if (label != "") {
labelElement.innerHTML = label;
}
if (imgData != "") {
var videoRect = video.getClientRects()[0];
imgElement.style.top = videoRect.top + "px";
imgElement.style.left = videoRect.left + "px";
imgElement.style.width = videoRect.width + "px";
imgElement.style.height = videoRect.height + "px";
imgElement.src = imgData;
}
var preCapture = Date.now();
var result = await new Promise(function(resolve, reject) {
pendingResolve = resolve;
});
shutdown = false;
return {'create': preShow - preCreate,
'show': preCapture - preShow,
'capture': Date.now() - preCapture,
'img': result};
}
''')
display(js)
def video_frame(label, bbox):
data = eval_js('stream_frame("{}", "{}")'.format(label, bbox))
return data
def js_to_image(js_reply):
"""
Params:
js_reply: JavaScript object containing image from webcam
Returns:
img: OpenCV BGR image
"""
# decode base64 image
image_bytes = b64decode(js_reply.split(',')[1])
# convert bytes to numpy array
jpg_as_np = np.frombuffer(image_bytes, dtype=np.uint8)
# decode numpy array into OpenCV BGR image
img = cv2.imdecode(jpg_as_np, flags=1)
return img
# + id="UIxZkgNNVu0d"
import cv2
import numpy as np
from google.colab.patches import cv2_imshow
import dlib
# Capturing live video stream with help of Javascript Helper Functions in the above code
cap = cv2.VideoCapture(0)
video_stream()
label_html = "Detecting ..."
bbox = ''
count = 0
detector = dlib.get_frontal_face_detector()
while True:
js_reply = video_frame(label_html, bbox)
if not js_reply:
break
# Reading image from video stream
frame = js_to_image(js_reply["img"])
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
# Detecting the features
faces = detector(gray)
print(len(faces))
if(len(faces) == 0):
cv2.putText(frame, "No Person Detected", (150, 50), cv2.FONT_HERSHEY_SIMPLEX, 1,
(255,0,0), 2)
# If mouth was detected, we will get the the x,y coordinates of the detected feature
i = 0
for face in faces:
x, y = face.left(), face.top()
x1, y1 = face.right(), face.bottom()
cv2.rectangle(frame, (x, y), (x1, y1), (0, 255, 0), 2)
i = i+1
# Printing number of people visible in screen on the frame
cv2.putText(frame, 'face num'+str(i), (x-10, y-10),
cv2.FONT_HERSHEY_SIMPLEX, 0.7, (0, 0, 255), 2)
print(face, i)
cv2_imshow(frame)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
cap.release()
cv2.destroyAllWindows()
| Machine Learning Models/Number_of_Persons_Detector/Persons_Counter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 6 - Sísmica de reflexão: Correção de Normal Moveout (NMO) e empilhamento
#
# Na aula passada, vimos que uma seção CMP é crucial para conseguirmos uma estimativa da velocidade. Nessa prática, veremos como utilizar a correção de NMO para fazer a **análise de velocidades**. Também veremos a técnica do empilhamento e como ela melhora a razão sinal-ruído do dado.
#
#
# Utilizaremos as simulações de ondas da biblioteca [Fatiando a Terra](http://www.fatiando.org). Essas simulações utilizam o [método de diferenças finitas](http://en.wikipedia.org/wiki/Finite_difference_method) para calcular soluções da equação da onda.
# ## Objetivos
#
# * Utilizar a correção de NMO para estimar velocidades.
# * Ver como o empilhamento melhora a qualidade dos dados.
# ## Questão para entregar
#
# <div class="alert alert-info" style="font-size:12pt; margin-top:20px">
# <b>
# Explique como é feita a análise de velocidades. As velocidades determinadas são as velocidades verdadeiras das camadas? Por que? Qual é vantagem de realizar o empilhamento?
# </b>
# </div>
#
# ### Regras para a resposta
#
# * Coloque **nome, data e o número da prática** em sua resposta.
# * A resposta pode ter no **máximo 1 página** (não uma folha).
# * **Execute o notebook** antes de responder. As simulações abaixo foram feitas para te ajudar.
# * **Pense e organize** sua resposta andtes de começar a escrever.
# ## Instruções
#
# Esse documento é um [Jupyter notebook](http://jupyter.org/), um documento interativo que mistura texto (como esse), código (como abaixo), e o resultado de executar o código (números, texto, figuras, videos, etc).
#
# O notebook te fornecerá exemplos interativos que trabalham os temas abordados no questionário. Utilize esses exemplos para responder as perguntas.
#
# As células com números ao lado, como `In [1]:`, são código [Python](http://python.org/). Algumas dessas células não produzem resultado e servem de preparação para os exemplos interativos. Outras, produzem gráficos interativos. **Você deve executar todas as células, uma de cada vez**, mesmo as que não produzem gráficos.
#
# **Para executar uma célula**, clique em cima dela e aperte `Shift + Enter`. O foco (contorno verde ou cinza em torno da célula) deverá passar para a célula abaixo. Para rodá-la, aperte `Shift + Enter` novamente e assim por diante. Você pode executar células de texto que não acontecerá nada.
# ## Setup
#
# Rode as células abaixo para carregar os módulos necessários para essa prática.
# %matplotlib inline
from __future__ import division, print_function
import math
from multiprocessing import Pool, cpu_count
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as ipw
from fatiando.seismic import RickerWavelet, FDAcoustic2D
from fatiando import utils
from fatiando.vis import anim_to_html
from fatiando.vis.mpl import seismic_image, seismic_wiggle
# ## Simulação de um CMP para um modelo de duas camadas
#
# Rode as células abaixo para simular uma seção CMP para um modelo de duas camadas. A célula abaixo cria nosso modelo.
# +
shape = (150, 200)
spacing = 10
extent = [0, shape[1]*spacing, shape[0]*spacing, 0]
# Velocidades
v1, v2 = 4000, 5000
densidade = np.ones(shape)*1600
velocidade = np.ones(shape)*v1
l1 = 40
densidade[l1:,:] = 1800
velocidade[l1:,:] = v2
l2 = 100
densidade[l2:,:] = 2000
velocidade[l2:,:] = 8000
# -
# Em seguida, precisamos definir onde serão localizadas as fontes e os receptores em nossa simulação. Vamos aproveitar e calcular também os espassamentos (offsets) dos receptores. Lembre-se: offset é a distância da conte ao receptor.
fontes = np.array(list(reversed(range(55, shape[1]//2 - 3, 3))))
recep = np.array([shape[1] - s for s in fontes])
offsets = (recep - fontes)*spacing
print("Utilizando {} fontes e {} receptores.".format(len(fontes), len(recep)))
print('Fontes: {}'.format(fontes*spacing))
print('Receptores: {}'.format(recep*spacing))
print('Offsets: {}'.format(offsets))
# Vamos rodar as simulações que precisamos (uma por fonte). **A barra de progresso não irá aparecer** pois vamos rodar as simulações em paralelo para agilizar o processo.
def simul_shot(fonte, its=1200, verbose=False):
shot = FDAcoustic2D(velocidade, densidade, spacing=spacing, taper=0.005, padding=50, verbose=verbose)
shot.add_point_source((0, fonte), RickerWavelet(1, 120))
shot.run(its)
return shot
# %%time
print('Simulando...', end=' ')
pool = Pool(processes=cpu_count())
shots = pool.map(simul_shot, fontes)
pool.close()
print('Pronto.')
# Vamos dar uma olhada na simulação de 1 tiro para ter uuma ideia do que acontece.
shots[0].animate(embed=True, dpi=60, fps=8, every=20, cutoff=0.5, cmap='Greys')
# Vamos extrair os dados CMP da nossa simulação.
# +
dt = shots[0].dt
times = np.linspace(0, dt*shots[0].simsize, shots[0].simsize)
CMP = np.empty((shots[0].simsize, len(recep)))
for i, s in enumerate(shots):
CMP[:, i] = s[:, 0, recep[i]]
fig = plt.figure(figsize=(10, 9))
ax = plt.subplot(111)
ax.set_title('CMP')
ax.set_xlabel('offset (m)')
ax.set_ylabel('times (s)')
cutoff = 0.1
ax.imshow(CMP, extent=[offsets.min(), offsets.max(), times.max(), times.min()],
aspect=1000, cmap='Greys', vmin=-cutoff, vmax=cutoff, interpolation='nearest')
fig.tight_layout()
# -
# ### Para pensar
#
# * Tente identificar a onda direta e as reflexões principais na simulação e no CMP.
# * O que é o outro evento que aparece no CMP (acima da segunda reflexão)?
# ## Análise de velocidades
# Agora que temos nosso CMP, podemos aplicar a correção de NMO e fazer a nossa análise de velocidades. Rode a célula abaixo para produzir uma figura interativa.
# +
def nmo_correction(CMP, times, offsets, v):
nmo = np.zeros_like(CMP)
for i, t0 in enumerate(times):
for j, o in enumerate(offsets):
t = np.sqrt(t0**2 + o**2/v[i]**2)
k = int(math.floor(t/dt))
if k < times.size - 1:
# Linear interpolation of the amplitude
y0, y1 = CMP[k, j], CMP[k + 1, j]
x0, x1 = times[k], times[k + 1]
nmo[i, j] = y0 + (y1 - y0)*(t - x0)/(x1 - x0)
return nmo
def analise_velocidades(CMP):
def aplica_nmo(v1, v2):
v = v1*np.ones_like(times)
v[times > 0.35] = v2
nmo = nmo_correction(CMP, times, offsets, v)
fig = plt.figure(figsize=(12, 6))
ax = plt.subplot(121)
ax.set_title('CMP')
ax.set_xlabel('offset (m)')
ax.set_ylabel('times (s)')
cutoff = 0.1
ax.imshow(CMP, extent=[offsets.min(), offsets.max(), times.max(), times.min()],
aspect=1000, cmap='Greys', vmin=-cutoff, vmax=cutoff, interpolation='nearest')
ax.grid()
ax = plt.subplot(122)
ax.set_title('Corrigido de NMO')
ax.set_xlabel('offset (m)')
ax.set_ylabel('times (s)')
cutoff = 0.1
ax.imshow(nmo, extent=[offsets.min(), offsets.max(), times.max(), times.min()],
aspect=1000, cmap='Greys', vmin=-cutoff, vmax=cutoff, interpolation='nearest')
ax.grid()
fig.tight_layout()
widget = ipw.interactive(aplica_nmo,
v1=ipw.FloatSlider(min=2000, max=6000, step=100, value=2000),
v2=ipw.FloatSlider(min=2000, max=6000, step=100, value=2000))
return widget
analise_velocidades(CMP)
# -
# ### Figura acima
#
# * O painel da esquerda mostra nosso CMP original.
# * O painel da direita mostra o CMP após aplicação da correção NMO usando as velocidades especificadas.
# * Você pode controlar as velocidades utilizadas na correção NMO do nosso CMP.
# ### Para pensar
#
# * **Determine a velocidade das duas reflexões**.
# * Essa velocidade é a velocidade real das camadas? Por que?
# ## Empilhamento
#
# O CMP que simulamos acima não é muito realista pois não está contaminado com qualquer tipo de ruído. Então vamos sacanear o problema adicionando ruído aleatório nos nossos dados.
ruido = 0.1
CMP_ruido = CMP + np.random.uniform(-ruido, ruido, CMP.shape)
fig = plt.figure(figsize=(10, 9))
ax = plt.subplot(111)
ax.set_title(u'CMP com ruído aleatório')
ax.set_xlabel('offset (m)')
ax.set_ylabel('times (s)')
cutoff = 0.1
ax.imshow(CMP_ruido, extent=[offsets.min(), offsets.max(), times.max(), times.min()],
aspect=1000, cmap='Greys', vmin=-cutoff, vmax=cutoff, interpolation='nearest')
fig.tight_layout()
# Uma técnica que nos possibilita diminuir a influência do ruído aleatório é o **empilhamento**. A ideia é somar diversos dados que possuam um sinal coerente no meio de ruído aleatório.
#
# Rode a célula abaixo para gerar uma figura interativa que ilustra esse conceito.
def empilhamento(ruido):
N = 500
M = 10
if ruido <= 0:
dados_ruido = np.array([np.zeros(N) for i in range(M)])
else:
dados_ruido = np.array([np.random.uniform(-ruido, ruido, size=N) for i in range(M)])
x = np.arange(N)
sinal = 1*utils.gaussian(x, 250, 2)
dados = dados_ruido + sinal
plt.figure(figsize=(10, 6))
plt.subplot(121)
plt.title(u'Dados com ruído e um sinal não-aleatório')
for i, d in enumerate(dados):
plt.plot(d + i + 1, x, '-k')
plt.xlim(0, len(dados) + 1)
plt.xlabel('# do dado')
plt.grid()
ax = plt.subplot(143)
plt.title('Empilhamento')
plt.plot(dados.sum(0), x, '-k')
plt.grid()
plt.xlim(-15, 15)
plt.tight_layout()
ipw.interactive(empilhamento, ruido=ipw.FloatSlider(min=0, max=0.5, step=0.1, value=0))
# ### Figura acima
#
# * O painel da esquerda mostra 10 conjuntos de dados com 1 sinal coerente no meio.
# * O painel da direita mostra o resultado do empilhamento desses dados.
# * Você pode controlar a quantidade de ruído aleatório que é inserido nos dados da esquerda.
# ### Para pensar
#
# * Com o máximo de ruído você consegue enxergar o sinal nos dados originais? E no empilhamento?
# ### Empilhamento após análise de velocidades
#
# Para poder fazer o empilhamento, precisamos ter um sinal coerente entre todos os traços. Isso significa que a nossa reflexão precisa acontecer no mesmo tempo em todos os traços. Felizmente, é exatamente isso que obtemos após a análise de velocidades e correção NMO.
#
# Rode a célula abaixo para gerar a figura interativa da análise de velocidades. Dessa vez vamos motrar também o resultado do empilhamento da seção corrigida de NMO.
# +
def nmo_empilhamento(v1, v2):
v = v1*np.ones_like(times)
v[times > 0.35] = v2
nmo = nmo_correction(CMP_ruido, times, offsets, v)
stack = np.atleast_2d(nmo.sum(1)).T
fig = plt.figure(figsize=(12, 6))
ax = plt.subplot(131)
ax.set_title('CMP')
ax.set_xlabel('offset (m)')
ax.set_ylabel('times (s)')
cutoff = 0.1
ax.imshow(CMP_ruido, extent=[offsets.min(), offsets.max(), times.max(), times.min()],
aspect=2500, cmap='Greys', vmin=-cutoff, vmax=cutoff, interpolation='nearest')
ax.grid()
ax = plt.subplot(132)
ax.set_title('Corrigido de NMO')
cutoff = 0.1
ax.imshow(nmo, extent=[offsets.min(), offsets.max(), times.max(), times.min()],
aspect=2500, cmap='Greys', vmin=-cutoff, vmax=cutoff, interpolation='nearest')
ax.grid()
ax = plt.subplot(165)
ax.set_title(u'Primeiro traço do NMO')
seismic_wiggle(np.atleast_2d(nmo[:, 0]).T, dt=dt, scale=1)
ax.set_xlim(-1, 1)
ax.grid()
ax = plt.subplot(166)
ax.set_title('Empilhamento')
seismic_wiggle(stack, dt=dt, scale=1)
ax.set_xlim(-10, 10)
ax.grid()
fig.tight_layout(pad=0, w_pad=0)
ipw.interactive(nmo_empilhamento,
v1=ipw.FloatSlider(min=2000, max=6000, step=100, value=2000),
v2=ipw.FloatSlider(min=2000, max=6000, step=100, value=2000))
# -
# ### Figura acima
#
# * O painel 1 mostra o CMP contaminado com ruído aleatório.
# * O painel 2 mostra o resultado da correção de NMO.
# * O painel 3 mostra o primeiro traço da seção corrigida de NMO (sem empilhamento).
# * O painel 4 mostra o resultado do empilhamento da seção corrigida de NMO.
# ### Para pensar
#
# * Como o resultado do empilhamento muda quando você utiliza as velocidades correta para NMO?
# * Como o traço empilhado se compara a um traço normal do NMO (sem empilhamento)?
# ## License and information
#
# **Course website**: https://github.com/leouieda/geofisica2
#
# **Note**: This notebook is part of the course "Geofísica 2" of Geology program of the
# [Universidade do Estado do Rio de Janeiro](http://www.uerj.br/).
# All content can be freely used and adapted under the terms of the
# [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
#
# 
| notebooks/6-reflexao-nmo-empilhamento.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scroll down to get to the interesting tables...
# # Construct list of properties of widgets
#
# "Properties" here is one of:
#
# + `keys`
# + `traits()`
# + `class_own_traits()`
#
# Common (i.e. uninteresting) properties are filtered out.
# The dependency on astropy is for their Table. Replace it with pandas if you want...
# +
import itertools
from ipywidgets import *
from IPython.display import display
from traitlets import TraitError
from astropy.table import Table, Column
# -
# # Function definitions
# ## Calculate "interesting" properties
def properties(widget, omit=None, source=None):
"""
Return a list of widget properties for a widget instance, omitting
common properties.
Parameters
----------
widget : ipywidgets.Widget instance
The widget for which the list of preoperties is desired.
omit : list, optional
List of properties to omit in the return value. Default is
``['layout', 'style', 'msg_throttle']``, and for `source='traits'
is extended to add ``['keys', 'comm']``.
source : str, one of 'keys', 'traits', 'class_own_traits', 'style_keys' optional
Source of property list for widget. Default is ``'keys'``.
"""
if source is None:
source = 'keys'
valid_sources = ('keys', 'traits', 'class_own_traits', 'style_keys')
if source not in valid_sources:
raise ValueError('source must be one of {}'.format(', '.join(valid_sources)))
if omit is None:
omit = ['layout', 'style', 'msg_throttle']
if source == 'keys':
props = widget.keys
elif source == 'traits':
props = widget.traits()
omit.extend(['keys', 'comm'])
elif source == 'class_own_traits':
props = widget.class_own_traits()
elif source == 'style_keys':
props = widget.style.keys
props = [k for k in props if not k.startswith('_')]
return [k for k in props if k not in omit]
# ## Create a table (cross-tab style) for which properties are available for which widgets
#
# This is the only place astropy.table.Table is used, so delete if you want to.
def table_for_keys(keys, keys_info, source):
unique_keys = set()
for k in keys:
unique_keys.update(keys_info[k])
unique_keys = sorted(unique_keys)
string_it = lambda x: 'X' if x else ''
colnames = ['Property ({})'.format(source)] + keys
columns = [Column(name=colnames[0], data=unique_keys)]
for c in colnames[1:]:
column = Column(name=c, data=[string_it(k in key_dict[c]) for k in unique_keys])
columns.append(column)
return Table(columns)
# ## List of widget objects...
widget_list = [
IntSlider,
FloatSlider,
IntRangeSlider,
FloatRangeSlider,
IntProgress,
FloatProgress,
BoundedIntText,
BoundedFloatText,
IntText,
FloatText,
ToggleButton,
Checkbox,
Valid,
Dropdown,
RadioButtons,
Select,
SelectionSlider,
SelectionRangeSlider,
ToggleButtons,
SelectMultiple,
Text,
Textarea,
Label,
HTML,
HTMLMath,
Image,
Button,
Play,
DatePicker,
ColorPicker,
Box,
HBox,
VBox,
Accordion,
Tab
]
# ## ...and their names
names = [wd.__name__ for wd in widget_list]
# ## Figure out the properties for each widget
#
# The `try`/`except` below is to catch a couple of classes that *require* that `options` be passed on intialization.
property_source = 'keys'
all_keys = []
for widget_class in widget_list:
try:
keys = properties(widget_class(), source=property_source)
except TraitError as e:
keys = properties(widget_class(options=(2,10)), source=property_source)
finally:
all_keys.append(keys)
# Probably should have used a dict from the beginning...
key_dict = {k: v for k, v in zip(names, all_keys)}
# ## Define a few groups of widgets by widget interface type
#
# This makes for nicer (i.e. more compact and readable) tables later on.
# +
sliders = [k for k in key_dict.keys() if 'Slider' in k]
buttons = [k for k in key_dict.keys() if 'Button' in k]
containers = ['Box', 'VBox', 'HBox', 'Accordion', 'Tab']
texts = [k for k in names if 'text' in k or 'Text' in k] + [k for k in names if 'HTML' in k] + ['Label']
progress = [k for k in names if 'Progress' in k]
selects = ['Dropdown', 'Select', 'SelectMultiple']
all_so_far = sliders + buttons + texts + containers + progress + selects
others = [k for k in names if k not in all_so_far]
slider_keys = set()
# -
# # Tables of keys (synced properties)
# ## Sliders
table_for_keys(sliders, key_dict, source=property_source)
# ## Buttons
table_for_keys(buttons, key_dict, source=property_source)
# ## Containers
table_for_keys(containers, key_dict, source=property_source)
# ## Text
table_for_keys(texts, key_dict, source=property_source)
# ## Progress bars
table_for_keys(progress, key_dict, source=property_source)
# # Select widgets
table_for_keys(selects, key_dict, source=property_source)
# ## Everything else
table_for_keys(others, key_dict, source=property_source)
# ## Style keys
property_source = 'style_keys'
style_keys = []
for widget_class in widget_list:
try:
keys = properties(widget_class(), source=property_source)
except TraitError as e:
keys = properties(widget_class(options=(2,10)), source=property_source)
except AttributeError:
keys=''
finally:
style_keys.append(keys)
for w, s in zip(names, style_keys):
print('{} has style keys: {}'.format(w, ', '.join(s)))
| notebooks/Table_of_widget_keys_and_style_keys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Bias in Data - Quality of Wikipedia Political Pages
#
# This notebook explores the concept of bias through data on English Wikipedia articles - specifically, articles on political figures from a variety of countries. Using the ORES machine learning API, we will rate each article based on its quality and examine which countries/regions possess a larger percentages of high-quality articles on political figures.
import pandas as pd
import requests
import json
# +
# Load page data from the 'data' folder
page_data = pd.read_csv('data/page_data.csv')
# Remove pages whose names begin with the string 'Template:', these are not wikipedia articles and
# will not be included in the analysis
page_data.drop(page_data[page_data['page'].str.startswith('Template:')].index, axis=0, inplace=True)
# +
# Load population data from 'data' folder
population_data = pd.read_csv('data/WPDS_2020_data - WPDS_2020_data.csv')
# Separate into regional data and data by country, retain both dataframes for analysis
regional_pop_data = population_data[population_data['Name'].str.isupper()]
population_data.drop(regional_pop_data.index, axis=0, inplace=True)
# -
# ## Rating Estimation with ORES API
#
# Now that the datasets have been loaded and cleaned, we will use the ORES API (documentation: https://ores.wikimedia.org/v3/#!/scoring/get_v3_scores_context_revid_model) to predict an article's rating out of six classes:
#
# 1. FA - Featured article
# 2. GA - Good article
# 3. B - B-class article
# 4. C - C-class article
# 5. Start - Start-class article
# 6. Stub - Stub-class article
#
# With FA being the best rating and the others following in descending order.
# +
# Set endpoint specifying english wikipedia as our context, article quality as our model
# of choice, and a variable list of revision ids
endpoint = 'https://ores.wikimedia.org/v3/scores/enwiki?models=articlequality&revids={revids}'
# +
# Set header with personal github account, email
headers = {
'User-Agent': 'https://github.com/TrevorNims',
'From': '<EMAIL>'
}
# +
# Define API call to communicate with ORES scoring interface
def api_call(endpoint, parameters):
call = requests.get(endpoint.format(**parameters), headers=headers)
response = call.json()
return response
# +
# Create dictionary to hold revision id coupled with their respective
# estimated scores. Create list to hold revision ids that cannot be scored.
score_dict = {}
unscorables = []
# Iterate over all revision ids in 'page_data' in batches of 50 - note that
# larger batches may cause a slowdown with the API calls.
for i in range(0, page_data.shape[0], 50):
# stop when we've reached the final revision id
end_idx = min(i+50, page_data.shape[0])
rev_ids = page_data['rev_id'].iloc[i:end_idx]
# concatenate revision ids as specified in the API documentation
revid_params = {'revids' : '|'.join(str(x) for x in rev_ids)}
data = api_call(endpoint, revid_params)
# for each revision id, save estimated score if it exists, otherwise save
# revision id in 'unscorables'
for score in data['enwiki']['scores']:
try:
score_dict[score] = data['enwiki']['scores'][score]['articlequality']['score']['prediction']
except KeyError as K:
unscorables.append(score)
# +
# create dataframe of revision ids and their respective estimated scores
score_estimates = pd.DataFrame(score_dict.items(), columns=['rev_id', 'article_quality_est.'])
# +
# save 'unscorables' as a .csv file in the data folder
pd.Series(unscorables).to_csv('data/unscorables.csv')
# +
# Retype rev_id as an int for comparsion with 'page_data' dataframe
score_estimates['rev_id'] = score_estimates['rev_id'].astype(int)
# +
# merge tables on rev_id, creating a single dataframe with page information and
# predicted score
page_data_with_scores = pd.merge(page_data, score_estimates, on='rev_id')
# +
# Inspect 'page_data_woith scores'
page_data_with_scores
# -
# ## Analysis
#
# Now that the pages have had their ranking estimated, we move on the production and measurement of
# two different metrics:
#
# 1. Number of total Articles per population by country
# 2. High Quality Articles as a proportion of total articles by country
#
# In this analysis, we will define a High Quality Article as one that has recieved either a 'FA' or a 'GA' rating from the ORES API.
# +
# rename column to match 'page_data_with_scores' format and facilitate table merging
population_data.rename({'Name' : 'country'}, axis=1, inplace=True)
# +
# merge 'population_data' with 'page_data_with_scores', drop unneeded columns and
# rename some columns to make them more ergonomic
pd_merged = pd.merge(page_data_with_scores, population_data, how='outer', on='country')
pd_merged.drop(columns=['FIPS', 'Type', 'TimeFrame', 'Data (M)'], axis=1, inplace=True)
pd_merged.rename({'page' : 'article_name', 'rev_id' : 'revision_id'}, axis=1, inplace=True)
# +
# Identify and save rows in 'pd_merged' that contain a value that is NaN
# (meaning either the country does not have any scored articles in our dataset,
# or that population data for the country is not available in 'population_data')
no_match = pd_merged[pd_merged.isna().any(axis=1)]
no_match.to_csv('data/wp_wpds_countries-no_match.csv')
# Remove rows with NaN values, save remaining data to csv file in 'data' folder
pd_merged.drop(no_match.index, inplace=True)
pd_merged.to_csv('data/wp_wpds_politicians_by_country.csv')
# +
# Obtain the total number of articles for each country
articles_by_country = pd_merged[['country', 'revision_id']].groupby(['country']).count()
# +
# Obtain the number of High Quality articles for each country
quality_articles_by_country = pd_merged.loc[pd_merged['article_quality_est.'].isin(['GA', 'FA'])]\
[['country', 'revision_id']].groupby(['country']).count()
# +
# Calculate the percentage of high quality articles per country
percentage_high_quality_articles = quality_articles_by_country/articles_by_country*100
percentage_high_quality_articles.rename({'revision_id' : 'percentage'}, axis=1, inplace=True)
# +
# Calculate the percentage of articles-per-population
population_by_country = pd_merged.groupby('country').mean()['Population'].to_frame()
population_by_country.rename({'Population' : 'percentage'}, axis=1, inplace=True)
articles_by_country.rename({'revision_id' : 'percentage'}, axis=1, inplace=True)
percentage_articles_per_population = articles_by_country/population_by_country*100
# -
# ### Analysis 1
#
# Below we can see countries with the highest percentages of articles per population. These countries tend to have lower populations.
percentage_articles_per_population.sort_values('percentage', ascending=False).head(10)
# ### Analysis 2
# Below we can see countries with the lowest percentages of articles per population. These countries tend to have higher populations.
percentage_articles_per_population.sort_values('percentage').head(10)
# ### Analysis 3
# Below we can see countries with the highest percentages of High Quality Articles as a proportion of the counrty's total article count.
percentage_high_quality_articles.sort_values('percentage', ascending=False).head(10)
# ### Analysis 4
# Below we can see countries with the lowest percentages of High Quality Articles as a proportion of the counrty's total article count.
percentage_high_quality_articles.sort_values('percentage').head(10)
# ## Country Mapping to Sub-Regions
#
# Now, we will analyze the same metrics by sub-region, however we first need to map each country to its respective sub-region(s).
# +
# Map countries to sub-regions by examining their respective indices from the original DataFrame
# 'population_data'
# construct list of sub-region indices
regional_idx_list = regional_pop_data.index.tolist()
regional_list_idx = 0
country_to_region_dict = {}
while regional_list_idx+1 < len(regional_idx_list):
for p_idx in population_data.index:
# If the country's index is within the range of two sub-region indices, pick the lower index as the
# sub-region
if p_idx in range(regional_idx_list[regional_list_idx], regional_idx_list[regional_list_idx+1]):
country_to_region_dict[population_data.loc[p_idx]['country']] = \
regional_pop_data['Name'].loc[regional_idx_list[regional_list_idx]]
# Update sub-region after iterating through all countries
regional_list_idx += 1
# Original while loop misses final sub-region as it only examines index ranges between
# sub-regions, final sub-region needs to be added manually
for p_idx in population_data.index:
if p_idx > regional_idx_list[regional_list_idx]:
country_to_region_dict[population_data.loc[p_idx]['country']] = \
regional_pop_data['Name'].loc[regional_idx_list[regional_list_idx]]
# +
# Construct DataFrame of each country with their associated sub-region
country_to_region = pd.DataFrame(country_to_region_dict.items(), columns=['country', 'Sub-Region'])
# Construct DataFrame for each "special" sub-region, these sub-regions consist of a collection
# of other sub-regions
africa_subset = country_to_region[country_to_region['Sub-Region'].str.contains('AFRICA')]
asia_subset = country_to_region[country_to_region['Sub-Region'].str.contains('ASIA')]
europe_subset = country_to_region[country_to_region['Sub-Region'].str.contains('EUROPE')]
latin_subset = country_to_region[country_to_region['Sub-Region'].isin(\
['CENTRAL AMERICA', 'CARIBBEAN', 'SOUTH AMERICA'])]
# +
# Construct DataFrames of total article counts by sub-region/"special" sub-regions
articles_by_region = pd.merge(pd_merged, country_to_region, how='left', on='country')\
[['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
articles_africa = pd.merge(pd_merged, africa_subset, how='left', on='country').dropna()\
[['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
articles_africa = pd.DataFrame({'AFRICA' : articles_africa.sum()}).transpose()
articles_asia = pd.merge(pd_merged, asia_subset, how='left', on='country').dropna()\
[['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
articles_asia = pd.DataFrame({'ASIA' : articles_asia.sum()}).transpose()
articles_europe = pd.merge(pd_merged, europe_subset, how='left', on='country').dropna()\
[['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
articles_europe = pd.DataFrame({'EUROPE' : articles_europe.sum()}).transpose()
articles_latin = pd.merge(pd_merged, latin_subset, how='left', on='country').dropna()\
[['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
articles_latin = pd.DataFrame({'LATIN AMERICA AND THE CARIBBEAN' : articles_latin.sum()}).transpose()
# construct list of DataFrames for iteration/merging purposes
region_article_list = [articles_by_region, articles_africa, articles_asia, articles_europe,
articles_latin]
# +
# Construct DataFrames of quality article counts by sub-region/"special" sub-regions
quality_articles_by_region = pd.merge(pd_merged, country_to_region, how='left', on='country')\
.loc[pd.merge(pd_merged, country_to_region, how='left', on='country')['article_quality_est.']
.isin(['GA', 'FA'])][['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
quality_articles_africa = pd.merge(pd_merged, africa_subset, how='left', on='country')\
.loc[pd.merge(pd_merged, africa_subset, how='left', on='country')['article_quality_est.']
.isin(['GA', 'FA'])][['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
quality_articles_africa = pd.DataFrame({'AFRICA' : quality_articles_africa.sum()}).transpose()
quality_articles_asia = pd.merge(pd_merged, asia_subset, how='left', on='country')\
.loc[pd.merge(pd_merged, asia_subset, how='left', on='country')['article_quality_est.']
.isin(['GA', 'FA'])][['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
quality_articles_asia = pd.DataFrame({'ASIA' : quality_articles_asia.sum()}).transpose()
quality_articles_europe = pd.merge(pd_merged, europe_subset, how='left', on='country')\
.loc[pd.merge(pd_merged, europe_subset, how='left', on='country')['article_quality_est.']
.isin(['GA', 'FA'])][['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
quality_articles_europe = pd.DataFrame({'EUROPE' : quality_articles_europe.sum()}).transpose()
quality_articles_latin = pd.merge(pd_merged, latin_subset, how='left', on='country')\
.loc[pd.merge(pd_merged, latin_subset, how='left', on='country')['article_quality_est.']
.isin(['GA', 'FA'])][['Sub-Region', 'revision_id']].groupby(['Sub-Region']).count()
quality_articles_latin = pd.DataFrame({'LATIN AMERICA AND THE CARIBBEAN' : quality_articles_latin.sum()}).transpose()
# construct list of DataFrames for iteration/merging purposes
region_quality_article_list = [quality_articles_by_region, quality_articles_africa, quality_articles_asia,
quality_articles_europe, quality_articles_latin]
# +
# Construct DataFrames of population totals by sub-region/"special" sub-regions
population_by_region = pd.merge(pd_merged, country_to_region, how='left', on='country')\
.groupby('Sub-Region').mean()['Population'].to_frame()
population_by_region
population_africa = pd.merge(pd_merged, africa_subset, how='left', on='country').dropna()\
.groupby(['Sub-Region']).mean()['Population'].to_frame()
population_africa = pd.DataFrame({'AFRICA' : population_africa.sum()}).transpose()
population_asia = pd.merge(pd_merged, asia_subset, how='left', on='country').dropna()\
.groupby(['Sub-Region']).mean()['Population'].to_frame()
population_asia = pd.DataFrame({'ASIA' : population_asia.sum()}).transpose()
population_europe = pd.merge(pd_merged, europe_subset, how='left', on='country').dropna()\
.groupby(['Sub-Region']).mean()['Population'].to_frame()
population_europe = pd.DataFrame({'EUROPE' : population_europe.sum()}).transpose()
population_latin = pd.merge(pd_merged, latin_subset, how='left', on='country').dropna()\
.groupby(['Sub-Region']).mean()['Population'].to_frame()
population_latin = pd.DataFrame({'LATIN AMERICA AND THE CARIBBEAN' : population_latin.sum()}).transpose()
# construct list of DataFrames for iteration/merging purposes
region_population_list = [population_by_region, population_africa, population_asia, population_europe,
population_latin]
# +
# Iterate through each of the corresponding DataFrames in all three lists, caluculating
# metrics upon each iteration
regional_percentage_quality_articles = []
regional_percentage_articles_per_population = []
for article_count, quality_article_count, pop in zip(region_article_list, region_quality_article_list,
region_population_list):
regional_percentage_quality_articles.append(quality_article_count/article_count*100)
regional_percentage_quality_articles[-1].rename({'revision_id' : 'percentage'}, axis=1, inplace=True)
pop.rename({'Population' : 'percentage'}, axis=1, inplace=True)
article_count.rename({'revision_id' : 'percentage'}, axis=1, inplace=True)
regional_percentage_articles_per_population.append(article_count/pop*100)
# +
# Merge DataFrames for each metric into a single DataFrame for display
regional_percentage_articles_per_population_merged = pd.concat(regional_percentage_articles_per_population)
regional_percentage_quality_articles_merged = pd.concat(regional_percentage_quality_articles)
# -
# ### Analysis 5
#
# Below we can see the percentages of articles per population by sub-region, sorted in descending order.
regional_percentage_articles_per_population_merged.sort_values('percentage', ascending=False)
# ### Analysis 6
#
# Below we can see all sub-regions' percentage of High Quality Articles as a proportion of the sub-region's total article count, sorted in descending order.
regional_percentage_quality_articles_merged.sort_values('percentage', ascending=False)
# ## Final Remarks
#
# Despite the limited scope, this study has brought to hand some interesting information on estimated article quality for political figures on English Wikipedia. When examining the percentage of High Quality Articles as a proportion of the country's total article count, it's interesting to see that the country with the highest percentage is North Korea. Intuitively, one would think that an english-speaking country would be the first entry (however, an English speaking country is not even present in the top ten percentages!). This could possibly point to the idea that there are a very limited number of articles about political figures for these countries, and that the ones that do exist are well-done.
#
# That being said, when examining the table illustrating sub-regions' percentage of High Quality Articles as a proportion of the sub-region's total article count, it is plain to see that a sub-region where English is the predominant language (i.e. Northern America) has the larget percentage by far.
#
# A final interesting takeaway was that Western Europe had the lowest percentage of High Quality Articles as a proportion of the sub-region's total article count of any sub-region, which goes against the intuition that a sub-region geographically and politically similar to that of many english-speaking countries would have higher-quality articles.
#
# #### Question Responses
#
# 1. What biases did you expect to find in the data (before you started working with it), and why?
# - Initially, I expected to find bias towards english-speaking countries posessing Higher Quality Articles. While this was somewhat true in the general case, it was not the case at the country-level.
#
#
# 2. What (potential) sources of bias did you discover in the course of your data processing and analysis?
# - While somewhat expected, I found that there were a number of smaller countries with absolutely no High Quality Articles written about their political figures. This lack of information illustrates the bias in the english Wikipedia's creator-base, as content production for political figures from smaller countries may be less desirable/more difficult than those from larger countries.
#
#
# 3. What might your results suggest about (English) Wikipedia as a data source?
# - In terms of just this analysis, it's interesting to see just how low the percentage of High Quality Articles as a proportion of the country's/sub-region's total article count really is. To put it plainly, it's curious to think that only ~5% of the articles I've read about politicians in North America were "High Quality". With all this said, I think some additional domain knowledge on what differentiates each article ranking would be nice to better understand these seemingly "low" percentages.
| hcds-a2-bias.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark 2.3 (Python 3)
# language: python
# name: pyspark3
# ---
# # 1 Creating a DataFrame
#
# First, let's create some DataFrame from Python objects. While this is probably not the most common thing to do, it is easy and helpful in some situations where you already have some Python objects.
df = spark.createDataFrame([('Alice', 13), ('Bob', 12)], ['name', 'age'])
print(df.collect())
# ## 1.1 Inspect Schema
#
# The `spark` object has different methods for creating a so called Spark DataFrame object. This object is similar to a table, it contains rows of records, which all conform to a common schema with named columns and specific types. On the surface it heavily borrows concepts from Pandas DataFrames or R DataFrames, although the syntax and many operations are syntactically very different.
#
# As the first step, we want to see the contents of the DataFrame. This can be easily done by using the show method.
df.printSchema()
# # 2 Reading Data
#
# Of course manually creating DataFrames from a couple of records is not the real use case. Instead we want to read data frames files.. Spark supports various file formats, we will use JSON in the following example.
#
# The entrypoint for creating Spark objects is an object called spark which is provided in the notebook and read to use. We will read a file containing some informations on a couple of persons, which will serve as the basis for the next examples
persons = spark.read.json("s3a://dimajix-training/data/persons.json")
persons
persons.collect()
# ## 2.1 Inspecting a DataFrame
#
# Spark supports various methods for inspecting both the contents and the schema of a DataFrame
persons.show()
persons.printSchema()
# ## Pandas Interoperability
#
# Spark also supports interoperation with Python Pandas, the standard library for modelling tabular data.
persons.toPandas()
# ## 2.2 Loading CSV Data
#
# Of course Spark also supports reading CSV data. CSV files may optionally contain a header containing the column names.
persons = spark.read.option("header", "true").csv(
"s3a://dimajix-training/data/persons_header.csv"
)
persons.toPandas()
# # 3 Simple Transformations
# ## 3.1 Projections
#
# The simplest thing to do is to create a new DataFrame with a subset of the available columns
# +
from pyspark.sql.functions import *
result = persons.select('name', col('age'))
result.toPandas()
# -
# ## 3.2 Addressing Columns
#
# Spark supports multiple different ways for addressing a columns. We just saw one way, but also the following methods are supported for specifying a column:
#
# * `df.column_name`
# * `df['column_name']`
# * `col('column_name')`
#
# All these methods return a Column object, which is an abstract representative of the data in the column. As we will see soon, transformations can be applied to Column in order to derive new values.
#
# ### Beware of Lowercase and Uppercase
#
# While PySpark itself is case insenstive concering column names, Python itself is case sensitive. Since the first method for addressing columns by treating them as fields of a Python object *is* Python syntax, this is also case sensitive!
# +
from pyspark.sql.functions import *
result = persons.select('name', persons.age, col('height'), persons['sex'])
result.toPandas()
# -
# ## 3.3 Transformations
#
# The `select` method actually accepts any column object. A column object conceptually represents a column in a DataFrame. The column may either refer directly to an existing column of the input DataFrame, or it may represent the result of a calculation or transformation of one or multiple columns of the input DataFrame. For example if we simply want to transform the name into upper case, we can do so by using a function `upper` provided by PySpark.
result = persons.select(persons.name, upper(persons.name))
result.toPandas()
# ### Defining new Column Names
# The resulting DataFrame again has a schema, but the column names to not look very nice. But by using the `alias` method of a `Column` object, you can immediately rename the newly created column like you are already used to in SQL with `SELECT complex_operation(...) AS nice_name FROM ...`.
#
# Technically specifying a new name for the resulting column is not required (as we already saw above), if the name is not specified, PySpark will generate a name from the expression. But since this generated name tends to be rather long and contains the logic instead of the intention, it is highly recommended to always explicitly specify the name of the resulting column using `as`.
# Result should be "Alice is 23 years old"
result = persons.select(
concat(persons.name, lit(" is "), persons.age, lit(" years old")).alias("description")
)
result.toPandas()
# You can also perform simple mathematical calculations like addition, multiplication etc.
result = persons.select((persons.age * 2).alias("age2"))
result.toPandas()
# ### Common Functions
#
# You can find the full list of available functions at [PySpark SQL Module](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#module-pyspark.sql.functions). Commonly used functions for example are as follows:
#
# * [`concat(*cols)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.concat) - Concatenates multiple input columns together into a single column.
# * [`substring(col,start,len)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.substring) - Substring starts at pos and is of length len when str is String type or returns the slice of byte array that starts at pos in byte and is of length len when str is Binary type.
# * [`instr(col,substr)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.instr) - Locate the position of the first occurrence of substr column in the given string. Returns null if either of the arguments are null.
# * [`locate(col,substr, pos)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.locate) - Locate the position of the first occurrence of substr in a string column, after position pos.
# * [`length(col)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.length) - Computes the character length of string data or number of bytes of binary data.
# * [`upper(col)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.upper) - Converts a string column to upper case.
# * [`lower(col)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.lower) - Converts a string column to lower case.
# * [`coalesce(*cols)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.coalesce) - Returns the first column that is not null.
# * [`isnull(col)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.isnull) - An expression that returns true iff the column is null.
# * [`isnan(col)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.isnan) - An expression that returns true iff the column is NaN.
# * [`hash(cols*)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.hash) - Calculates the hash code of given columns.
#
# Spark also supports conditional expressions, like the SQL `CASE WHEN` construct
# * [`when(condition, value)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.when) - Evaluates a list of conditions and returns one of multiple possible result expressions.
#
# There are also some special functions often required
# * [`col(str)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.col) - Returns a Column based on the given column name.
# * [`lit(val)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.lit) - Creates a Column of literal value.
# * [`expr(str)`](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.expr) - Parses the expression string into the column that it represents
#
# ### User Defined Functions
# Unfortunately you cannot directly use normal Python functions for transforming DataFrame columns. Although PySpark already provides many useful functions, this might not always sufficient. But fortunately you can *convert* a standard Python function into a PySpark function, thereby defining a so called *user defined function* (UDF). Details will be explained in detail in the training.
# ## 3.4 Adding Columns
#
# A special variant of a `select` statement is the `withColumn` method. While the `select` statement requires all resulting columns to be defined in as arguments, the `withColumn` method keeps all existing columns and adds a new one. This operation is quite useful since in many cases new columns are derived from the existing ones, while the old ones still should be contained in the result.
#
# Let us have a look at a simple example, which only adds the salutation as a new column:
result = persons.withColumn(
"even_odd_age", when(persons.age % 2 == 0, "even").otherwise("odd")
)
result.toPandas()
# As you can see from the example above, `withColumn` always takes two arguments: The first one is the name of the new column (and it has to be a string), and the second argument is the expression containing the logic for calculating the actual contents.
# ## 3.5 Dropping a Column
#
# PySpark also supports the opposite operation which simply removes some columns from a dataframe. This is useful if you need to remove some sensitive data before saving it to disk:
result = persons.drop("sex")
result.toPandas()
# # 4 Filtering
#
# *Filtering* denotes the process of keeping only rows which meet a certain filter criteria.
# ## 4.1 Simple `WHERE` clauses
#
# PySpark support two different approaches. The first approach specifies the filtering expression as a PySpark expression using columns:
result = persons.filter(persons.age > 22)
result.toPandas()
result = persons.where((persons.age > 22) & (persons.height > 160))
result.toPandas()
# The second approach simply uses a string containing an SQL expression:
result = persons.filter("age > 22 AND height > 160")
result.toPandas()
# ## 4.2 Limit Operations
#
# When working with large datasets, it may be helpful to limit the amount of records (like an SQL `LIMIT` operation).
result = persons.limit(3)
result.toPandas()
# # 5 Simple Aggregations
#
# PySpark supports simple global aggregations, like `COUNT`, `MAX`, `MIN` etc...
persons.count()
result = persons.select(
max(persons.age).alias("max_age"), avg(persons.height).alias("avg_height")
)
result.toPandas()
# # 6 Grouping & Aggregating
#
# An important class of operation is grouping and aggregation, which is equivalnt to an SQL `SELECT aggregation GROUP BY grouping` statement. In PySpark, grouping and aggregation is always performed by first creating groups using `groupBy` immediately followed by aggregation expressions inside an `agg` method. (Actually there are also some predefined aggregations which can be used instead of `agg`, but they do not offer the flexiviliby which is required most of the time).
#
# Note that in the `agg` method you only need to specify the aggregation expression, the grouping columns are added automatically by PySpark to the resulting DataFrame.
result = persons.groupBy(persons.sex).agg(
avg(persons.age).alias("avg_age"),
min(persons.height).alias("min_height"),
max(persons.height).alias("max_height"),
)
result.toPandas()
# ## Aggregation Functions
#
# PySpark supports many aggregation functions, they can be found in the documentation at [PySpark Function Documentation](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#module-pyspark.sql.functions). Aggregation functions are marked as such in the documentation, unfortunately there is no simple overview. Among common aggregation functions, there are for example:
#
# * count
# * sum
# * avg
# * corr
# * first
# * last
# # 7 Sorting DataFrames
#
# You can sort the entries (= rows) of a DataFrame by an arbitrary column or expression.
result = persons.sort(persons.height)
result.toPandas()
# If nothing else is specified, PySpark will sort the records in increasing order of the sort columns. If you require descending order, this can be specified by manipulating the sort column with the `desc()` method as follows:
result = persons.orderBy(persons.height.desc())
result.toPandas()
# # User Defined Functions
#
# Sometimes the built in functions do not suffice or you want to call an existing function of a Python library. Using User Defined Functions (UDF) it is possible to wrap an existing function into a Spark DataFrame function.
# +
import html
from pyspark.sql.types import *
html_encode = udf(html.escape, StringType())
df = spark.createDataFrame([("Alice & Bob",), ("Thelma & Louise",)], ["name"])
result = df.select(html_encode(df.name).alias("html_name"))
result.toPandas()
# -
| pyspark-ml-crashcourse/notebooks/01 - PySpark DataFrame Introduction Full.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
# # PySchools without Thomas High School 9th graders
# ### Dependencies and data
# + tags=[]
# Dependencies
import os
import numpy as np
import pandas as pd
# + tags=[]
# School data
school_path = os.path.join('data', 'schools.csv') # school data path
school_df = pd.read_csv(school_path)
# Student data
student_path = os.path.join('data', 'students.csv') # student data path
student_df = pd.read_csv(student_path)
school_df.shape, student_df.shape
# + tags=[]
# Change Thomas High School 9th grade scores to NaN
student_df.loc[(student_df['school_name'].str.contains('Thomas')) & (student_df['grade'] == '9th'),
['reading_score', 'math_score']] = np.NaN
student_df.loc[(student_df['school_name'].str.contains('Thomas')) & (student_df['grade'] == '9th'),
['reading_score', 'math_score']].head(3)
# -
# ### Clean student names
# +
# Prefixes to remove: "Miss ", "Dr. ", "Mr. ", "Ms. ", "Mrs. "
# Suffixes to remove: " MD", " DDS", " DVM", " PhD"
fixes_to_remove = ['Miss ', '\w+\. ', ' [DMP]\w?[DMS]'] # regex for prefixes and suffixes
str_to_remove = r'|'.join(fixes_to_remove) # join into a single raw str
# Remove inappropriate prefixes and suffixes
student_df['student_name'] = student_df['student_name'].str.replace(str_to_remove, '', regex=True)
# -
# Check prefixes and suffixes
student_names = [n.split() for n in student_df['student_name'].tolist() if len(n.split()) > 2]
pre = list(set([name[0] for name in student_names if len(name[0]) <= 4])) # prefixes
suf = list(set([name[-1] for name in student_names if len(name[-1]) <= 4])) # suffixes
print(pre, suf)
# ### Merge data
# Add binary vars for passing score
student_df['pass_read'] = (student_df.reading_score >= 70).astype(int) # passing reading score
student_df['pass_math'] = (student_df.math_score >= 70).astype(int) # passing math score
student_df['pass_both'] = np.min([student_df.pass_read, student_df.pass_math], axis=0) # passing both scores
student_df.head(3)
# +
# Add budget per student var
school_df['budget_per_student'] = (school_df['budget'] / school_df['size']).round().astype(int)
# Bin budget per student
school_df['spending_lvl'] = pd.qcut(school_df['budget_per_student'], 4,
labels=range(1, 5))
# Bin school size
school_df['school_size'] = pd.qcut(school_df['size'], 3, labels=['Small', 'Medium', 'Large'])
school_df
# + tags=[]
# Merge data
df = pd.merge(student_df, school_df, on='school_name', how='left')
df.info()
# -
# ### District summary
# + tags=[]
# District summary
district_summary = pd.DataFrame(school_df[['size', 'budget']].sum(), columns=['District']).T
district_summary['Total Schools'] = school_df.shape[0]
district_summary = district_summary[['Total Schools', 'size', 'budget']]
district_summary_cols = ['Total Schools', 'Total Students', 'Total Budget']
district_summary
# + tags=[]
# Score cols
score_cols = ['reading_score', 'math_score', 'pass_read', 'pass_math', 'pass_both']
score_cols_new = ['Average Reading Score', 'Average Math Score', '% Passing Reading', '% Passing Math', '% Passing Overall']
# Add scores to district summary
for col, val in df[score_cols].mean().items():
if 'pass' in col:
val *= 100
district_summary[col] = val
district_summary
# + tags=[]
# Rename cols
district_summary.columns = district_summary_cols + score_cols_new
district_summary
# Format columns
for col in district_summary.columns:
if 'Total' in col:
district_summary[col] = district_summary[col].apply('{:,}'.format)
if 'Average' in col:
district_summary[col] = district_summary[col].round(2)
if '%' in col:
district_summary[col] = district_summary[col].round().astype(int)
district_summary
# -
# ### School summary
# + tags=[]
# School cols
school_cols = ['type', 'size', 'budget', 'budget_per_student',
'reading_score', 'math_score', 'pass_read', 'pass_math', 'pass_both']
school_cols_new = ['School Type', 'Total Students', 'Total Budget', 'Budget Per Student']
school_cols_new += score_cols_new
# School summary
school_summary = df.groupby('school_name')[school_cols].agg({
'type': 'max',
'size': 'max',
'budget': 'max',
'budget_per_student': 'max',
'reading_score': 'mean',
'math_score': 'mean',
'pass_read': 'mean',
'pass_math': 'mean',
'pass_both': 'mean'
})
school_summary.head(3)
# + tags=[]
# Rename cols
school_summary.index.name = None
school_summary.columns = school_cols_new
# Format values
for col in school_summary.columns:
if 'Total' in col:
school_summary[col] = school_summary[col].apply('{:,}'.format)
if 'Average' in col:
school_summary[col] = school_summary[col].round(2)
if '%' in col:
school_summary[col] = (school_summary[col] * 100).round().astype(int)
school_summary
# -
# ### Scores by grade
# + tags=[]
# Reading scores by grade of each school
grade_read_scores = pd.pivot_table(df, index='school_name', columns='grade',
values='reading_score', aggfunc='mean').round(2)
grade_read_scores.index.name = None
grade_read_scores.columns.name = 'Reading scores'
grade_read_scores = grade_read_scores[['9th', '10th', '11th', '12th']]
grade_read_scores
# + tags=[]
# Math scores by grade of each school
grade_math_scores = pd.pivot_table(df, index='school_name', columns='grade',
values='math_score', aggfunc='mean').round(2)
grade_math_scores.index.name = None
grade_math_scores.columns.name = 'Math Scores'
grade_math_scores = grade_math_scores[['9th', '10th', '11th', '12th']]
grade_math_scores
# -
# ### Scores by budget per student
# + tags=[]
# Scores by spending
spending_scores = df.groupby('spending_lvl')[score_cols].mean().round(2)
for col in spending_scores.columns:
if "pass" in col:
spending_scores[col] = (spending_scores[col] * 100).astype(int)
spending_scores
# + tags=[]
# Formatting
spending_scores.index.name = 'Spending Level'
spending_scores.columns = score_cols_new
spending_scores
# -
# ### Scores by school size
# + tags=[]
# Scores by school size
size_scores = df.groupby('school_size')[score_cols].mean().round(2)
for col in size_scores.columns:
if "pass" in col:
size_scores[col] = (size_scores[col] * 100).astype(int)
size_scores
# + tags=[]
# Formatting
size_scores.index.name = 'School Size'
size_scores.columns = score_cols_new
size_scores
# -
# ### Scores by school type
# + tags=[]
# Scores by school type
type_scores = df.groupby('type')[score_cols].mean().round(2)
for col in type_scores.columns:
if "pass" in col:
type_scores[col] = (type_scores[col] * 100).astype(int)
type_scores
# + tags=[]
# Formatting
type_scores.index.name = 'School Type'
type_scores.columns = score_cols_new
type_scores
# -
| pyschools/analysis2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from scipy import signal
import matplotlib.pyplot as plt
import seaborn as sns
fig_size = plt.rcParams["figure.figsize"]
# Prints: [8.0, 6.0]
print("Current size:", fig_size)
# -
# Set figure width to 12 and height to 9
fig_size[0] = 16
fig_size[1] = 16
plt.rcParams["figure.figsize"] = fig_size
train = pd.read_csv('results/resnet_train_10_features.csv')
valid = pd.read_csv('results/resnet_valid_10_features.csv')
# +
train['type'] = train['fname'].str.split('/').apply(lambda x: x[0])
train['disease'] = train['fname'].str.split('/').apply(lambda x: x[1])
train['file'] = train['fname'].str.split('/').apply(lambda x: x[2])
train['pname'] = train['file'].str.split('___').apply(lambda x: x[0])
# +
valid['type'] = valid['fname'].str.split('/').apply(lambda x: x[0])
valid['disease'] = valid['fname'].str.split('/').apply(lambda x: x[1])
valid['file'] = valid['fname'].str.split('/').apply(lambda x: x[2])
valid['pname'] = valid['file'].str.split('___').apply(lambda x: x[0])
# -
train = pd.concat([train, valid])
biom = pd.read_excel('./EE Biomarkers from MAB/CohortBbiomarkersforSana - Copy.xlsx')
ee_train_bio
ee_train_bio = train[(train['disease']=='EE') & (train['pname'].str.len()==15)]
ee_train_bio['pid'] = ee_train_bio['pname'].str.split('_').apply(lambda x: x[1])
img_cols = ['0','1','2','3','4','5','6','7','8','9']
ee_avg = ee_train_bio.groupby('pid')[img_cols].agg(np.mean).reset_index()
ee_avg['pid'] = ee_avg['pid'].astype(np.int64)
# ee_avg
merge_ee = ee_avg.merge(biom, left_on='pid', right_on='ID')
merge_ee = merge_ee.drop(['pid','ID','sex', 'oedema','breastfeeding', 'hivchild', 'lactasedef','pcrpos'], axis=1)
merge_ee = merge_ee.fillna(biom.median())
# +
img_ee = merge_ee[img_cols]
biom_ee = merge_ee[[x for x in list(merge_ee.columns) if x not in img_cols]]
biom_ee = biom_ee.fillna(biom.median())
# -
sns.heatmap(merge_ee.corr().iloc[10:, :10], cmap="YlGnBu")
merge_ee
len(biom_ee.iloc[1,:])
| BiomarkerCorrelation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Ch `05`: Concept `02`
# + [markdown] deletable=true editable=true
# ## Segmentation
# + [markdown] deletable=true editable=true
# Import libraries and define hyper-parameters:
# + deletable=true editable=true
import tensorflow as tf
import numpy as np
from bregman.suite import *
k = 2
segment_size = 50
max_iterations = 100
# + [markdown] deletable=true editable=true
# Define functions to get the chromogram and the dataset:
# + deletable=true editable=true
chromo = tf.placeholder(tf.float32)
max_freqs = tf.argmax(chromo, 0)
def get_chromogram(audio_file):
F = Chromagram(audio_file, nfft=16384, wfft=8192, nhop=2205)
return F.X
def get_dataset(sess, audio_file):
chromo_data = get_chromogram(audio_file)
print('chromo_data', np.shape(chromo_data))
chromo_length = np.shape(chromo_data)[1]
xs = []
for i in range(chromo_length/segment_size):
chromo_segment = chromo_data[:, i*segment_size:(i+1)*segment_size]
x = extract_feature_vector(sess, chromo_segment)
if len(xs) == 0:
xs = x
else:
xs = np.vstack((xs, x))
return xs
# + [markdown] deletable=true editable=true
# As required for the k-means algorithm, specify the assignment and re-centering code:
# + deletable=true editable=true
def initial_cluster_centroids(X, k):
return X[0:k, :]
def assign_cluster(X, centroids):
expanded_vectors = tf.expand_dims(X, 0)
expanded_centroids = tf.expand_dims(centroids, 1)
distances = tf.reduce_sum(tf.square(tf.subtract(expanded_vectors, expanded_centroids)), 2)
mins = tf.argmin(distances, 0)
return mins
def recompute_centroids(X, Y):
sums = tf.unsorted_segment_sum(X, Y, k)
counts = tf.unsorted_segment_sum(tf.ones_like(X), Y, k)
return sums / counts
# + [markdown] deletable=true editable=true
# Given a chromogram, extract a histogram of sound frequencies as our feature vector:
# + deletable=true editable=true
def extract_feature_vector(sess, chromo_data):
num_features, num_samples = np.shape(chromo_data)
freq_vals = sess.run(max_freqs, feed_dict={chromo: chromo_data})
hist, bins = np.histogram(freq_vals, bins=range(num_features + 1))
return hist.astype(float) / num_samples
# + [markdown] deletable=true editable=true
# In a session, segment an audio file using k-means:
# + deletable=true editable=true
with tf.Session() as sess:
X = get_dataset(sess, 'TalkingMachinesPodcast.wav')
print(np.shape(X))
centroids = initial_cluster_centroids(X, k)
i, converged = 0, False
# prev_Y = None
while not converged and i < max_iterations:
i += 1
Y = assign_cluster(X, centroids)
# if prev_Y == Y:
# converged = True
# break
# prev_Y = Y
centroids = sess.run(recompute_centroids(X, Y))
if i % 50 == 0:
print('iteration', i)
segments = sess.run(Y)
for i in range(len(segments)):
seconds = (i * segment_size) / float(20)
min, sec = divmod(seconds, 60)
time_str = str(min) + 'm ' + str(sec) + 's'
print(time_str, segments[i])
| ch05_clustering/Concept02_segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
from sklearn.calibration import calibration_curve
from matplotlib import pyplot as plt
import pickle
import torch
from torch import nn
from torch.autograd import Variable
from model import Model
import utils
np.random.seed(42)
# -
# # Training Models
#
# Run the following commands:
#
# `python train_model.py --gpu_id 0 --mode dropout --model_id 0 --dprob 0.0`
#
# `python train_model.py --gpu_id 0 --mode dropout --model_id 1 --dprob 0.1`
#
# `python train_model.py --gpu_id 0 --mode dropout --model_id 2 --dprob 0.2`
#
# `python train_model.py --gpu_id 0 --mode dropout --model_id 3 --dprob 0.3`
#
# `python train_model.py --gpu_id 0 --mode dropout --model_id 4 --dprob 0.4`
#
# `python train_model.py --gpu_id 0 --mode concrete --model_id 5 --dr 0.0001`
#
# `python train_model.py --gpu_id 0 --mode concrete --model_id 6 --dr 1.0`
#
# `python train_model.py --gpu_id 0 --mode concrete --model_id 7 --dr 100.0`
# # Dropout Rate Convergence
# +
plt.figure(figsize=(15, 3))
for index in range(5):
plt.subplot(1, 5, index + 1)
for training_data in cdropout_training_datas:
convergence = [x[5][index] for x in training_data]
plt.plot(convergence)
plt.title('L = {}'.format(index))
plt.xlabel('#epoch')
plt.ylim([0, 0.5])
plt.ylabel('p')
plt.tight_layout()
plt.show()
# -
# # Learning Curves
# +
plt.figure(figsize=(15, 6))
plt.subplot(1, 2, 1)
for model, training_data, dropout_prob, color in zip(dropout_models, dropout_training_datas,
dropout_probs, ['r', 'g', 'b', 'c', 'm']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
plt.plot(np.arange(20, 100), train_scores[20:], color, label='train, p={}'.format(dropout_prob))
plt.plot(np.arange(20, 100), val_scores[20:], color + '--', label='val, p={}'.format(dropout_prob))
plt.legend()
plt.xlabel('#epoch')
plt.ylabel('Avg. Hamming distance')
plt.title('Dropout')
plt.subplot(1, 2, 2)
for model, training_data, dr, color in zip(cdropout_models, cdropout_training_datas,
cdropout_regs, ['r', 'g', 'b']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
plt.plot(np.arange(20, 100), train_scores[20:], color, label='train, dr={}'.format(dr))
plt.plot(np.arange(20, 100), val_scores[20:], color + '--', label='val, dr={}'.format(dr))
plt.legend()
plt.xlabel('#epoch')
plt.ylabel('Avg. Hamming distance')
plt.title('Concrete Dropout')
plt.show()
# +
plt.figure(figsize=(15, 6))
plt.subplot(1, 2, 1)
for training_data, dropout_prob, color in zip(dropout_training_datas,
dropout_probs, ['r', 'g', 'b', 'c', 'm']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
dropout_execution_times = [x[0] for x in training_data]
plt.plot(dropout_execution_times[20:], train_scores[20:], color,
label='train, p={}'.format(dropout_prob))
plt.plot(dropout_execution_times[20:], val_scores[20:], color + '--',
label='val, p={}'.format(dropout_prob))
plt.legend()
plt.xlabel('min.')
plt.ylabel('Avg. Hamming distance')
plt.title('Dropout')
plt.subplot(1, 2, 2)
for training_data, dr, color in zip(cdropout_models, cdropout_training_datas,
cdropout_regs, ['r', 'g', 'b']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
dropout_execution_times = [x[0] for x in training_data]
plt.plot(dropout_execution_times[20:], train_scores[20:], color,
label='train, dr={}'.format(dr))
plt.plot(dropout_execution_times[20:], val_scores[20:], color + '--',
label='val, dr={}'.format(dr))
plt.legend()
plt.xlabel('min.')
plt.ylabel('Avg. Hamming distance')
plt.title('Concrete Dropout')
plt.show()
| experiments/nlp/.ipynb_checkpoints/results-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''base'': conda)'
# name: python3
# ---
# # PCM for tuning regularization parameters in Ridge regression
#
# Import necessary libraries
import PcmPy as pcm
import numpy as np
import pickle
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from numpy import exp, sqrt
from sklearn.linear_model import Ridge, LinearRegression
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
# +
# Make the training data:
N = 100 # Number of observations
Q = 10 # Number of random effects regressors
P = 10 # Number of variables
Z = np.random.normal(0,1,(N,Q)) # Make random design matrix
U = np.random.normal(0,1,(Q,P))*0.5 # Make random effects
Y = Z @ U + np.random.normal(0,1,(N,P)) # Generate training data
# Make testing data:
Zt = np.random.normal(0,1,(N,Q))
Yt = Zt @ U + np.random.normal(0,1,(N,P))
# -
# Build the datasets from the Data and condition vectors
comp = np.array([0,0,0,0,0,0,1,1,1,1])
M1 = pcm.regression.RidgeDiag(comp, fit_intercept = True)
M1.optimize_regularization(Z,Y)
print('Estimated thetas:', M1.theta_.round(2))
print('Regularisation:', (exp(M1.theta_[-1])/exp(M1.theta_[:-1])).round(2))
# Now you can fit the model
M1.fit(Z,Y)
Yp = M1.predict(Zt)
R2 = 1- np.sum((Yt-Yp)**2)/np.sum((Yt)**2)
print('r2 :', R2.round(3))
Yp = M1.predict(Zt)
| docs/demos/demo_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAT281 - Laboratorios N°07
# <a id='p1'></a>
# ## I.- Problema 01
#
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b6/Anscombe.svg/1200px-Anscombe.svg.png" width="360" height="360" align="center"/>
#
#
# El **cuarteto de Anscombe** comprende cuatro conjuntos de datos que tienen las mismas propiedades estadísticas, pero que evidentemente son distintas al inspeccionar sus gráficos respectivos.
#
# Cada conjunto consiste de once puntos (x, y) y fueron construidos por el estadístico <NAME>. El cuarteto es una demostración de la importancia de mirar gráficamente un conjunto de datos antes de analizarlos.
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set_palette("deep", desat=.6)
sns.set(rc={'figure.figsize':(11.7,8.27)})
# -
# cargar datos
df = pd.read_csv(os.path.join("data","anscombe.csv"), sep=",")
df.head()
# Basado en la información presentada responda las siguientes preguntas:
#
# 1. Gráfique mediante un gráfico tipo **scatter** cada grupo. A simple vista, ¿ los grupos son muy distintos entre si?.
# 2. Realice un resumen de las medidas estadísticas más significativas ocuapando el comando **describe** para cada grupo. Interprete.
# 3. Realice un ajuste lineal para cada grupo. Además, grafique los resultados de la regresión lineal para cada grupo. Interprete.
# 4. Calcule los resultados de las métricas para cada grupo. Interprete.
# 5. Es claro que el ajuste lineal para algunos grupos no es el correcto. Existen varias formas de solucionar este problema (eliminar outliers, otros modelos, etc.). Identifique una estrategia para que el modelo de regresión lineal ajuste de mejor manera e implemente otros modelos en los casos que encuentre necesario.
df['grupo'].unique() #vemos cuantos grupos son y cuáles son los nombres
# 1. Gráfique mediante un gráfico tipo scatter cada grupo. A simple vista, ¿ los grupos son muy distintos entre si?:
# +
df_1=df.groupby('grupo').get_group('Grupo_1')
df_2=df.groupby('grupo').get_group('Grupo_2')
df_3=df.groupby('grupo').get_group('Grupo_3')
df_4=df.groupby('grupo').get_group('Grupo_4')
plt.figure(figsize=(10, 6)) #creación de la figura
sns.set_style("whitegrid")
sns.scatterplot(
x='x',#datos en x
y='y',#datos en y
data=df_1,#datos
ci = None,
)
#tamaños
plt.xlabel('x', size=15)
plt.ylabel('y',size=15)
plt.title('Grupo 1', size=25)
plt.show()
plt.figure(figsize=(10, 6)) #creación de la figura
sns.scatterplot(
x='x',#datos en x
y='y',#datos en y
data=df_2,#datos
ci = None,
)
#tamaños
plt.xlabel('x', size=15)
plt.ylabel('y',size=15)
plt.title('Grupo 2', size=25)
plt.show()
plt.figure(figsize=(10, 6)) #creación de la figura
sns.scatterplot(
x='x',#datos en x
y='y',#datos en y
data=df_3,#datos
ci = None,
)
#tamaños
plt.xlabel('x', size=15)
plt.ylabel('y',size=15)
plt.title('Grupo 3', size=25)
plt.show()
plt.figure(figsize=(10, 6)) #creación de la figura
sns.scatterplot(
x='x',#datos en x
y='y',#datos en y
data=df_4,#datos
ci = None,
)
#tamaños
plt.xlabel('x', size=15)
plt.ylabel('y',size=15)
plt.title('Grupo 4', size=25)
plt.show()
# -
# 2. Realice un resumen de las medidas estadísticas más significativas ocuapando el comando **describe** para cada grupo.
df.groupby('grupo').describe()
# es notorio que todas las estadísticas son iguales, por lo que a priori, alguien podría decir que todos los datos son iguales, claramente a partir de los gráficos, se entiende que no es así.
# 3. Realice un resumen de las medidas estadísticas más significativas ocuapando el comando describe para cada grupo. Interprete.
# +
plt.figure(figsize=(10, 6)) #creación de la figura
sns.lmplot(
x='x',
y='y',
data=df_1,
height = 8,
)
plt.title('Grupo 1', size=25)
plt.xlim(2.5, 14.5)
plt.show()
plt.figure(figsize=(10, 6)) #creación de la figura
sns.lmplot(
x='x',
y='y',
data=df_2,
height = 8,
)
plt.title('Grupo 2', size=25)
plt.xlim(2.5, 14.5)
plt.show()
plt.figure(figsize=(10, 6)) #creación de la figura
sns.lmplot(
x='x',
y='y',
data=df_3,
height = 8,
)
plt.title('Grupo 3', size=25)
plt.xlim(2.5, 14.5)
plt.show()
plt.figure(figsize=(10, 6)) #creación de la figura
sns.lmplot(
x='x',
y='y',
data=df_4,
height = 8,
)
plt.title('Grupo 4', size=25)
plt.xlim(2.5, 14.5)
plt.show()
# -
# se observa desde los gráficos que si bien los datos son distintos, los gráficos son iguales, es un database interesante.
# 4. Calcule los resultados de las métricas para cada grupo. Interprete.
#
# 5. Es claro que el ajuste lineal para algunos grupos no es el correcto. Existen varias formas de solucionar este problema (eliminar outliers, otros modelos, etc.). Identifique una estrategia para que el modelo de regresión lineal ajuste de mejor manera e implemente otros modelos en los casos que encuentre necesario.
| labs/lab_07_alvaro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Titanic: Machine Learning from Disaster | Kaggle
#
# This is a jupyter notebook exploring the [Kaggle Titanic competition](https://www.kaggle.com/c/titanic). My first proper attempt using this format so expect bad formatting :-).
#
# ## Contents
# - Environment
# - Importing Data
# - Exploring the Data
#
# ## The Environment
# Nothing that special as it stands, everything you would expect in a python data project! Currently making use of the following libraries for analysis and plots:
#
#
import pandas as pd # for data manipulation
import matplotlib.pyplot as plt # for visulatisation
| notebooks/01-dan-titanic-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import os
import sys
import numpy as np
import _pickle as pkl
import matplotlib.pyplot as plt
import plotly.express as px
from rulefit import RuleFit
from sklearn.model_selection import train_test_split
from sklearn.inspection import permutation_importance
import itertools
import pandas as pd
from tqdm import tqdm
from utils import *
from sklearn.decomposition import TruncatedSVD
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.decomposition import PCA
from sklearn.preprocessing import OneHotEncoder
# %matplotlib inline
# Load dataset
chosen_features = None
dataset = pkl.load(open('./bankrupcy.pkl', 'rb'))
y_all = dataset['y']
if 'Shell weight' in dataset['num_names']:
y_all = (y_all > 9).astype(int)
elif 'num_procedures' in dataset['num_names']:
y_all = (y_all > 0).astype(int)
X = np.concatenate([dataset['X_num'], dataset['X_cat']], -1)
baseline = return_baseline(dataset['X_num'], dataset['X_cat'])
feature_names = np.concatenate([dataset['num_names'], dataset['cat_names']])
if chosen_features is None:
chosen_features = np.arange(len(X[0]))
X_all = X[:, chosen_features]
baseline = baseline[chosen_features]
features = np.array(['X{}'.format(str(i).zfill(3)) for i in range(X_all.shape[-1])])
features_dic = {'X{}'.format(str(i).zfill(3)): feature_names[chosen_features[i]] for i in range(X_all.shape[-1])}
fn = ['\n'.join(['-\n'.join([j[:7], j[7:]]) if len(j)>7 else j for j in i.split('\n')]) for i in feature_names]
pd.DataFrame(X, columns=feature_names).head()
# Train test split
balanced = True ## To balance dataset between labels
npoints = None
if balanced:
X_all, y_all = balanced_dataset(X_all, y_all, max_size=10000)
X_train, X_test, y_train, y_test = train_test_split(X_all, y_all, test_size=0.2, stratify=y_all)
print(len(X_all))
# ## Rulefit
from featurevec import FeatureVec
fv = FeatureVec('classify', max_depth=3, max_sentences=100000,
feature_names=list(feature_names),
exp_rand_tree_size=False,
tree_generator=None)
fv.fit(X_train, y_train, bagging=5)
print(fv.predictor.score(X_test, y_test)) ## Model accuracy
fv.plot(dynamic=True, confidence=True, path='image_dir/bankrupcy.html')
# ## Different feature importance methods
# +
num_rmvs = min(X.shape[-1], 20)
rf_order = np.argsort(fv.predictor.feature_importances_)[::-1]
fv_order = np.argsort(fv.importance)[::-1]
pm_order = np.argsort(permutation_importance(fv.predictor, X_train, y_train, n_repeats=1)['importances_mean'])[::-1]
drop_rf = accuracy_drop(rf_order[:20], fv.predictor, X_train, y_train, X_test, y_test)
drop_fv = accuracy_drop(fv_order[:20], fv.predictor, X_train, y_train, X_test, y_test)
drop_pm = accuracy_drop(pm_order[:20], fv.predictor, X_train, y_train, X_test, y_test)
drop_rnd = np.mean([
accuracy_drop(np.random.permutation(X_train.shape[-1])[:20], fv.predictor, X_train, y_train, X_test, y_test)
for _ in range(1)], 0)
# -
fig = plt.figure(figsize=(8,8))
plt.rcParams['font.size'] = 15
plt.plot(np.arange(len(drop_fv)), 100 * np.array(drop_fv), lw=5)
plt.plot(np.arange(len(drop_rf)), 100 * np.array(drop_rf), ls='--', lw=5)
plt.plot(np.arange(len(drop_pm)), 100 * np.array(drop_pm), ls='-.', lw=5)
plt.plot(np.arange(len(drop_rnd)), 100 * np.array(drop_rnd), ls='dotted', lw=5)
plt.xlabel('# of removed features', fontsize=25)
plt.ylabel('Test accuracy (%)', fontsize=25)
plt.legend(['FV', 'Gini', 'Permutation', 'Random'], fontsize=25)
| FeatureVectors Notebook.ipynb |
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell - hvega
-- language: haskell
-- name: ihaskell_cities-wordcloud
-- ---
-- # Towards Interactive Datascience in Haskell #
--
-- ## Introduction ##
--
-- Haskell and data science - on first sight a great match: native function
-- composition, lazy evaluation, fast execution times, and lots of code checks.
-- These sound like ingredients for scalable, production-ready data transformation
-- pipelines. What is missing then? Why is Haskell not widely used in data
-- science?
--
-- One of the reasons is that Haskell lacks a standardized data analysis
-- environment. For example, Python has a *de facto* standard library set with
-- `numpy`, `pandas` and `scikit-learn` that form the backbone, and many other
-- well-supported specialized libraries such as `keras` and `tensorflow` that are
-- easily accessible. These libraries are distributed with user friendly package
-- managers and explained in a plethora of tutorials, Stack Overflow questions and
-- millions of Jupyter notebooks. Most problems from beginner to advanced level
-- can be solved by adapting and combining these existing solutions.
--
-- This post presents Jupyter and JupyterLab - both important ingredients of the
-- Python ecosystem - and shows how we can take advantage of these tools for
-- interactive data analysis in Haskell.
--
-- ## Jupyter and exploratory data analysis ##
--
--
-- [Project Jupyter](https://jupyter.org/) became famous through the browser-based
-- notebook app that allows to execute code in various compute environments and
-- interlace it with text and media elements
-- ([example gallery](https://github.com/jupyter/jupyter/wiki/A-gallery-of-interesting-Jupyter-Notebooks)).
--
-- More generally, Project Jupyter standardizes the interactions between Jupyter
-- *frontends* such as the notebook and Jupyter *kernels*, the compute
-- environments. Typically a kernel receives a message from a frontend, executes
-- some code, and responds with a rich media message. The frontend can render the
-- response message as text, images, videos or small applications. All exchanged
-- messages are standardized by the Jupyter protocol and therefore independent of
-- the specific frontend or kernel that is used. Various frontends and kernels for
-- many different languages, like Python, Haskell, R, C++, Julia, etc, exist.
--
-- Quick REPL-like interaction of a user with a compute environment via a frontend
-- is very useful for exploratory data analysis. The user interrogates and
-- transforms the data with little code snippets and receives immediate feedback
-- through rich media responses. Different algorithms (expressed as short code
-- snippets) can rapidly be prototyped and visualized. Long multistep dialogues
-- with the kernel can be assembled into a sequential notebook. Notebooks
-- interlace explanatory text, code and media elements and can therefore be used
-- as human-readable reports (such as this blogpost). This workflow has become one
-- of the most popular ways for exploratory data analysis.
--
--
-- ## Conversations with a Jupyter kernel ##
--
-- [IHaskell](https://github.com/gibiansky/IHaskell) is the name of the Jupyter
-- kernel for Haskell. It contains a little executable `ihaskell` that can receive
-- and respond to messages in the Jupyter protocoll (via
-- [ZeroMQ](http://zeromq.org/). Here is a little dialogue with `ihaskell` that is
-- initiated by sending the following code snippet from the notebook frontend to
-- `ihaskell`:
take 10 $ (^2) <$> [1..]
-- In Jupyter parlance the above dialogue corresponds to the following
-- `execute_request`:
--
-- ```
-- >> shell.execute_request (8be63d5c-1170-495d-82da-e56272052faf) <<
--
-- header: {username: "", version: "5.2", session: "32fe9cd0-8c37-450e-93c0-6fbd45bfdcd9",
-- msg_id: "8be63d5c-1170-495d-82da-e56272052faf", msg_type: "execute_request"}
-- parent_header: Object
-- channel: "shell"
-- content: {silent: false, store_history: true, user_expressions: Object, allow_stdin: true,
-- stop_on_error: true, code: "take 10 $ (^2) <$> [1..]"} <<<<<<<<<<< LOOK HERE
-- metadata: Object
-- buffers: Array[0]
-- ```
--
-- , and to the following `display_data` message that is received as a response:
--
-- ```
-- << iopub.display_data (68cce1e7-4d60-4a20-a707-4bf352c4d8d2) >>
--
-- header: {username: "", msg_type: "display_data", version: "5.0"
-- msg_id: "68cce1e7-4d60-4a20-a707-4bf352c4d8d2",
-- session: "32fe9cd0-8c37-450e-93c0-6fbd45bfdcd9",
-- date: "2018-08-02T08:14:10.245877Z"}
-- msg_id: "68cce1e7-4d60-4a20-a707-4bf352c4d8d2"
-- msg_type: "display_data"
-- parent_header: {username: "", msg_type: "execute_request", version: "5.0",
-- msg_id: "8be63d5c-1170-495d-82da-e56272052faf",
-- session: "32fe9cd0-8c37-450e-93c0-6fbd45bfdcd9"}
-- metadata: Object
-- content: {data: {text/plain: "[1,4,9,16,25,36,49,64,81,100]"}, <<<<<<<<<<< LOOK HERE
-- metadata: {output_type: "display_data"}}
-- buffers: Array[0]
-- channel: "iopub"
-- ```
--
-- `ihaskell` can import Haskell libraries dynamically and has some special
-- commands to enable language extensions, print type information or to use
-- Hoogle.
--
--
-- ## JupyterLab ##
--
-- JupyterLab is the newest animal in the Jupyter frontend zoo, and it is arguably
-- the most powerful: console, notebook, terminal, text-editor, or image viewer,
-- Jupyterlab integrates these data science building blocks into a single web
-- based user interface. Jupyterlab is setup as a modular system that can be
-- extended. A module assembles the base elements, changes or add new features to
-- build an IDE, a classical notebook or even a GUI where all interactions with
-- the underlying execution kernels are hidden behind graphical elements.
--
-- How can Haskell take advantage of Jupyterlab's capacities? To begin with,
-- JupyterLab provides plenty of out-of-the-box renderers that can be used for
-- free by Haskell. From the [default
-- renderers](https://jupyterlab.readthedocs.io/en/stable/user/file_formats.html),
-- the most interesting is probably Vega plotting. But also `geojson`, `plotly` or
-- and many other formats are available from the list of extensions, that will
-- certainly grow. Another use case might be to using the jupyterlab extension
-- system makes it easy to build a simple UI that interact with an execution
-- environment. Finally, Jupyter and associated workflows are known by a large
-- community. Using Haskell through these familiar environments softens the
-- barrier that many encounter when exploring Haskell for serious data science.
--
-- Let's get into a small example that shows how to use the Jupyterlab VEGA
-- renderer with IHaskell in Jupyterlab.
--
-- ## Wordclouds using Haskell, Vega and Jupyterlab ##
--
-- We will use here the word content of a few wikipedia articles that are saved
-- as text files in the `./posts` folder. Here is a little code cell that reads all
-- `.txt` files in the `posts` folder and concatenates them in a single long string
-- from which we remove some punctuation characters. The code cell is sent to the
-- `ihaskell` kernel, which responds to the last `take` function with a simple text
-- response.
-- +
:ext QuasiQuotes
import System.Directory
import Data.List
fnames <- getDirectoryContents "./posts"
paths = ("./posts/"++) <$> fnames
md_files = filter (isSuffixOf ".txt") paths
text <- mconcat (readFile <$> md_files)
cleanedText = filter (not . (`elem` "\n,.?!-:;\"\'")) text
take 500 cleanedText
-- -
-- Now let's define a Vega JSON as a string and fill it up with our text. A
-- convenient way to write longer multiline strings in Haskell are `QuasiQuotes`.
-- We use `fString` QuasiQuotes from the `PyF` package. Note that `{}` fills in
-- template data and `{{` corresponds to an escaped `{`.
-- +
import Formatting
import PyF
import Data.String.QQ
let vegaString = [fString|{{
"$schema": "https://vega.github.io/schema/vega/v4.json",
"width": 800,
"height": 400,
"padding": 0,
"data": [
{{
"name": "table",
"values": [
"{cleanedText}"
],
"transform": [
{{
"type": "countpattern",
"field": "data",
"case": "upper",
"pattern": "[\\\\w']{{3,}}",
"stopwords": "(\\\\d+|youll|looking|like|youre|etc|yet|need|cant|ALSO|STILL|ISNT|Want|Lots|HTTP|HTTPS|i|me|my|myself|we|us|our|ours|ourselves|you|your|yours|yourself|yourselves|he|him|his|himself|she|her|hers|herself|it|its|itself|they|them|their|theirs|themselves|what|which|who|whom|whose|this|that|these|those|am|is|are|was|were|be|been|being|have|has|had|having|do|does|did|doing|will|would|should|can|could|ought|i'm|you're|he's|she's|it's|we're|they're|i've|you've|we've|they've|i'd|you'd|he'd|she'd|we'd|they'd|i'll|you'll|he'll|she'll|we'll|they'll|isn't|aren't|wasn't|weren't|hasn't|haven't|hadn't|doesn't|don't|didn't|won't|wouldn't|shan't|shouldn't|can't|cannot|couldn't|mustn't|let's|that's|who's|what's|here's|there's|when's|where's|why's|how's|a|an|the|and|but|if|or|because|as|until|while|of|at|by|for|with|about|against|between|into|through|during|before|after|above|below|to|from|up|upon|down|in|out|on|off|over|under|again|further|then|once|here|there|when|where|why|how|all|any|both|each|few|more|most|other|some|such|no|nor|not|only|own|same|so|than|too|very|say|says|said|shall)"
}},
{{
"type": "formula", "as": "angle",
"expr": "[0, 90][~~(random() * 3)]"
}},
{{
"type": "formula", "as": "weight",
"expr": "if(datum.text=='VEGA', 600, 300)"
}}
]
}}
],
"scales": [
{{
"name": "color",
"type": "ordinal",
"range": ["#3e4593", "#bc3761", "#39163d", "#2a1337"]
}}
],
"marks": [
{{
"type": "text",
"from": {{"data": "table"}},
"encode": {{
"enter": {{
"text": {{"field": "text"}},
"align": {{"value": "center"}},
"baseline": {{"value": "alphabetic"}},
"fill": {{"scale": "color", "field": "text"}}
}},
"update": {{
"fillOpacity": {{"value": 1}}
}},
"hover": {{
"fillOpacity": {{"value": 0.5}}
}}
}},
"transform": [
{{
"type": "wordcloud",
"size": [800, 400],
"text": {{"field": "text"}},
"rotate": {{"field": "datum.angle"}},
"font": "Helvetica Neue, Arial",
"fontSize": {{"field": "datum.count"}},
"fontWeight": {{"field": "datum.weight"}},
"fontSizeRange": [12, 56],
"padding": 2
}}
]
}}
]
}}|]
-- -
-- We display this JSON string with the native Jupyterlab JSON renderer here for
-- convenience. The `Display` function explicitly tells `ihaskell` to send a
-- display message to the frontend. The JSON function tells `ihaskell` to annotate
-- the content of the display message as `application/json`.
import qualified IHaskell.Display as D
D.Display [D.json vegaString]
-- Finally, we can plot this JSON with Vega:
D.Display [D.vegalite vegaString]
| example/Haskell/cities-wordcloud/cities-wordcloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Foundations Of AIML
# ## Session 11
# ### Experiment 1.3: Learning rates
# The objective of this experiment is to tune the hyper parameter called learning rate and observe the changes in the output.
#
# We will use CIFAR100 dataset.This dataset is just like the CIFAR-10, except it has 100 classes containing 600 images each. There are 50,000 training images and 10,000 testing images.
# **Importing required Packages**
# Importing pytorch packages
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
import torchvision
import torchvision.transforms as transforms
from torch.autograd import Variable
# Importing config.py file
import config as cf
from utils import *
## Importing python packages
import os
import sys
import time
import datetime
import numpy as np
import math
import matplotlib.pyplot as plt
# Checking for GPU instance
use_cuda = torch.cuda.is_available()
#Intilizaing the accuracy value as zero
best_acc = 0
# #### Data Preparation
# +
print('\n[Phase 1] : Data Preparation')
##dataset
dataset = 'cifar100'
# Preparing the dataset
transform_train = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize(cf.mean[dataset], cf.std[dataset]),
]) # meanstd transformation
# -
# #### Applying the standard mean
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(cf.mean[dataset], cf.std[dataset]),
])
# #### Downloading and Loading the dataset
#
# The torchvision package consists of popular datasets, model architectures, and common image transformations for computer vision.
#
# number of classes in the dataset
num_classes = 100
### Downloading the dataset
trainset = torchvision.datasets.CIFAR100(root='/data', train=True, download=True, transform=transform_train)
testset = torchvision.datasets.CIFAR100(root='/data', train=False, download=False, transform=transform_test)
### Loading the dataset
trainloader = torch.utils.data.DataLoader(trainset, batch_size=128, shuffle=True, num_workers=8)
testloader = torch.utils.data.DataLoader(testset, batch_size=100, shuffle=False, num_workers=8)
# #### Let us define the network
# +
import torch.nn as nn
import torch.nn.functional as F
class LeNet(nn.Module):
def __init__(self, num_classes, init_mode='xavier'): ### supports 'zero', 'normal', 'xavier', 'he' inits
super(LeNet, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16*5*5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, num_classes)
if init_mode == 'zero':
for m in self.modules():
if isinstance(m, nn.Conv2d) or isinstance(m, nn.Linear):
m.weight.data.zero_() ### fill tensor elements with zeros
if m.bias is not None:
m.bias.data.zero_()
if init_mode == 'normal':
for m in self.modules():
if isinstance(m, nn.Conv2d) or isinstance(m, nn.Linear):
m.weight.data.normal_() ### fill tensor elements with random numbers from normal distribution
if m.bias is not None:
m.bias.data.normal_()
if init_mode == 'xavier':
for m in self.modules():
if isinstance(m, nn.Conv2d):
fan_out = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
fan_in = m.kernel_size[0] * m.kernel_size[1] * m.in_channels
n = fan_in + fan_out
m.weight.data.normal_(0, math.sqrt(2. / n))
if m.bias is not None:
m.bias.data.normal_(0, math.sqrt(2. / n))
if isinstance(m, nn.Linear):
size = m.weight.size()
fan_out = size[0] # number of rows
fan_in = size[1] # number of columns
variance = math.sqrt(2.0/(fan_in+fan_out))
m.weight.data.normal_(0.0, variance)
if m.bias is not None:
m.bias.data.normal_(0, variance)
if init_mode == 'he':
for m in self.modules():
if isinstance(m, nn.Conv2d):
n = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
m.weight.data.normal_(0, math.sqrt(2. / n))
if m.bias is not None:
m.bias.data.normal_(0, math.sqrt(2. / n))
if isinstance(m, nn.Linear):
size = m.weight.size()
fan_out = size[0] # number of rows
fan_in = size[1] # number of columns
variance = math.sqrt(2.0/(fan_in))
m.weight.data.normal_(0.0, variance)
if m.bias is not None:
m.bias.data.normal_(0, variance)
## Forward Pass
def forward(self, x):
out = F.relu(self.conv1(x))
out = F.max_pool2d(out, 2)
out = F.relu(self.conv2(out))
out = F.max_pool2d(out, 2)
out = out.view(out.size(0), -1)
out = F.relu(self.fc1(out))
out = F.relu(self.fc2(out))
out = self.fc3(out)
return(out)
# -
# ### Training with Xavier init
## Calling the model
net = LeNet(num_classes, init_mode='xavier')
# Checking for GPU instance
if use_cuda:
net.cuda()
### Intiliazing the loss
criterion = nn.CrossEntropyLoss()
# +
def train(epoch):
print('\nEpoch: %d' % epoch)
net.train()
#Declaring the variables
train_loss = 0
correct = 0
total = 0
# Looping over train data
for batch_idx, (inputs, targets) in enumerate(trainloader):
# Checking for GPU instance
if use_cuda:
inputs, targets = inputs.cuda(), targets.cuda()
# Optimizer
optimizer.zero_grad()
# Converting targets and inputs into pytorch variables
inputs, targets = Variable(inputs), Variable(targets)
# Forward Pass
outputs = net(inputs)
# Storing the outputs size
size_ = outputs.size()
# Reducing the dimenssion
outputs_ = outputs.view(size_[0], num_classes)
# Calculating the loss
loss = criterion(outputs_, targets)
# backward Pass
loss.backward()
# Optimizer Step
optimizer.step()
# Calculating the tarining loss
train_loss += loss.data[0]
# Predicting the values
_, predicted = torch.max(outputs_.data, 1)
# Storing the targets size
total += targets.size(0)
# Calculating the corrected values
correct += predicted.eq(targets.data).cpu().sum()
# printing the data
if batch_idx%30 == 0 or batch_idx == len(trainloader)-1:
# printing the progress bar
progress_bar(batch_idx, len(trainloader), 'Loss: %.3f | Acc: %.3f%% (%d/%d)'
% (train_loss/(batch_idx+1), 100.*correct/total, correct, total))
# Storing number of epoch,loss and accuracy in a file
train_loss_file.write('%d %.3f %.3f\n' %(epoch, train_loss/len(trainloader), 100.*correct/total))
# -
def test(epoch):
global best_acc
# Declaring the valules
net.eval()
test_loss = 0
correct = 0
total = 0
# Looping over test data
for batch_idx, (inputs, targets) in enumerate(testloader):
# Checking for GPU instance
if use_cuda:
inputs, targets = inputs.cuda(), targets.cuda()
# Converting inputs and targets into pytorch variables
inputs, targets = Variable(inputs, volatile=True), Variable(targets)
# Forward Pass
outputs = net(inputs)
# Storing the outputs size
size_ = outputs.size()
# Reducing the dimenssions
outputs_ = outputs.view(size_[0], num_classes)
# Calculating the loss
loss = criterion(outputs_, targets)
# Calculating the test loss
test_loss += loss.data[0]
# Storing the predicted values
_, predicted = torch.max(outputs_.data, 1)
# Storing the targets size
total += targets.size(0)
# Caluculating the correct values
correct += predicted.eq(targets.data).cpu().sum()
# Printing the data
if batch_idx%30 == 0 or batch_idx == len(testloader)-1:
# Printing the progress bar
progress_bar(batch_idx, len(testloader), 'Loss: %.3f | Acc: %.3f%% (%d/%d)'
% (test_loss/(batch_idx+1), 100.*correct/total, correct, total))
# Printing the validation loss
print('val_loss: ', test_loss/len(testloader), 'accuracy: ', 100.0*correct/total)
# Storing number of epoch,loss and accuracy in a file
val_loss_file.write('%d %.3f %.3f\n' %(epoch, test_loss/len(testloader), 100.*correct/total))
# Save checkpoint.
acc = 100.*correct/total
# Checking for best accuracy
if acc > best_acc:
print('Saving..')
state = {
'net': net,
'acc': acc,
'epoch': epoch,
}
# Checking whether its a directory or not
if not os.path.isdir('../checkpoint'):
# Creating a directory
os.mkdir('../checkpoint')
# Saving the data
torch.save(state, '../checkpoint_ckpt.t7')
# Storing the best accuracy
best_acc = acc
experiment = 'lr_schedule'
# Creating files in write mode
train_loss_file = open("../Lab11-Experiment1/"+experiment+"train_loss.txt", "w")
val_loss_file = open("../Lab11-Experiment1/"+experiment+"val_loss.txt", "w")
### Training and testing the model for 60 epochs
for epoch in range(0, 60):
if epoch == 50:
optimizer = optim.SGD(net.parameters(), lr=0.0001, momentum=0.9)
if epoch == 30:
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
if epoch == 0:
optimizer = optim.SGD(net.parameters(), lr=0.01, momentum=0.9)
# Training the model
train(epoch)
# Testing the model
test(epoch)
# Closing the files
train_loss_file.close()
val_loss_file.close()
# #### Plotting the training curves
training_curves("../Lab11-Experiment1/"+experiment)
# #### 1. Notice the slope of loss and accuracy for the epoch when learning rate is changed. What could be the reason of this jump?
# #### 2. Why do you think it would be better to start with a high learning rate and high momentum and subsequently decrease during the next epochs?
| Session11/Lab11-Experiment1_3.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (PortfolioChoiceWithRiskyHousing-Latest)
# language: python
# name: pycharm-2f4f0d29
# ---
# %%
import matplotlib.pyplot as plt
import numpy as np
from HARK.ConsumptionSaving.ConsPortfolioModel import PortfolioConsumerType
from ConsPortfolioHousingModel import (
PortfolioRiskyHousingType,
ConsPortfolioRiskyHousingSolver,
init_portfolio_risky_housing,
)
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
agent = PortfolioRiskyHousingType()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
solver = ConsPortfolioRiskyHousingSolver.from_agent(agent)
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
solver.prepare_to_solve()
solver.solve()
# %% pycharm={"name": "#%%\n"}
M = np.linspace(0.0, 100, 100)
for h in np.linspace(2.0, 10.0, 3):
C = solver.cFuncHse_now(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.plot(M, solver.rental_solution.cFuncRnt(M), label="rent")
plt.legend()
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
M = np.linspace(0.0, 5000, 1000)
for h in np.linspace(2.0, 10.0, 3):
C = solver.cFuncHse_now.derivativeX(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.legend()
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
M = np.linspace(1.0, 10, 1000)
for h in np.linspace(2.0, 10.0, 3):
C = solver.vPfuncHse_now(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.legend()
plt.xscale("log")
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
M = np.linspace(0.0, 100, 100)
for h in np.linspace(2.0, 10.0, 3):
C = solver.ShareFuncHse_now(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.plot(M, solver.rental_solution.ShareFuncRnt(M), label="rent")
plt.legend()
plt.show()
# %%
agent.solve()
# %% pycharm={"name": "#%%\n"}
agent.track_vars = ["mNrm", "hNrm", "haveHse", "cNrm", "aNrm", "pLvl", "aLvl", "Share", "RentShk", "Adjust", "t_age", "t_cycle"]
agent.AgentCount = 1000
agent.T_sim = 81
agent.initialize_sim()
# %% pycharm={"name": "#%%\n", "is_executing": true}
agent.simulate()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n", "is_executing": true}
M = np.linspace(0.0, 100, 100)
for h in np.linspace(2.0, 10.0, 3):
C = agent.solution[0].cFuncHse(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.plot(M, agent.solution[0].cFuncRnt(M), label="rent")
plt.legend()
plt.show()
# %% pycharm={"name": "#%%\n", "is_executing": true}
M = np.linspace(0.0, 5000, 100)
for h in np.linspace(2.0, 10.0, 3):
C = agent.solution[0].cFuncHse.derivativeX(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.legend()
plt.show()
# %% pycharm={"is_executing": true}
M = np.linspace(0.0, 100, 100)
for h in np.linspace(2.0, 10.0, 3):
C = agent.solution[0].ShareFuncHse(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.plot(M, agent.solution[0].ShareFuncRnt(M), label="rent")
plt.legend()
plt.show()
# %% pycharm={"name": "#%%\n", "is_executing": true}
M = np.linspace(1.0, 100, 100)
for h in np.linspace(2.0, 10.0, 3):
C = agent.solution[0].vFuncHse(M, h * np.ones_like(M))
plt.plot(M, C, label=h)
plt.legend()
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n", "is_executing": true}
ShareFuncRnt = [soln.ShareFuncRnt for soln in agent.solution]
M = np.linspace(0.0, 100, 100)
for func in ShareFuncRnt:
C = func(M)
plt.plot(M, C)
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n", "is_executing": true}
vFuncRnt = [soln.vFuncRnt for soln in agent.solution]
M = np.linspace(1.0, 100, 100)
for func in vFuncRnt:
C = func(M)
plt.plot(M, C)
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n", "is_executing": true}
agent_portfolio = PortfolioConsumerType(**init_portfolio_risky_housing)
agent_portfolio.solve()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n", "is_executing": true}
ShareFuncAdj = [soln.ShareFuncAdj for soln in agent_portfolio.solution]
M = np.linspace(0.0, 100, 100)
for func in ShareFuncAdj:
C = func(M)
plt.plot(M, C)
plt.show()
# %% jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n", "is_executing": true}
| code/python/risky_housing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cupy as cp
import cusignal
from scipy import signal
import numpy as np
# This admittedly may not be realistic with a wavelet, but keeping consistent perf array sizes (or tying to!)
M = int(1e8)
csig = np.random.rand(int(1e7))
gsig = cp.random.rand(int(1e7))
# ### Morlet
# %%timeit
cmorlet = signal.morlet(M)
# %%timeit
gmorlet = cusignal.morlet(M)
# ### Ricker
# %%timeit
cricker = signal.ricker(M, int(1e3))
# %%timeit
gricker = cusignal.ricker(M, int(1e3))
# ### Continuous Wavelet Transform
# %%timeit
ccwt = signal.cwt(csig, signal.ricker, np.arange(1,31))
# %%timeit
gcwt = cusignal.cwt(gsig, cusignal.ricker, cp.arange(1,31))
| notebooks/api_guide/wavelets_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Evaluate Linear Models
#
# Evaluate Linear Models
#
# Here, I evaluate the following linear models:
#
# + LinearRegression
# + PLSRegression
# + Lasso
# + Enet
#
# ## NOTE:
#
# For linear models I will add polynomial features of 2nd degree.
# +
import numpy as np
import pandas as pd
# sklearn import
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.linear_model import LinearRegression, Lasso, ElasticNet
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import VarianceThreshold
from sklearn.cross_decomposition import PLSRegression
# my module imports
from optimalcodon.projects.rnastability.dataprocessing import get_data, general_preprocesing_pipeline
from optimalcodon.projects.rnastability import modelevaluation
# -
(train_x, train_y), (test_x, test_y) = get_data("../19-04-30-EDA/results_data/")
print("{} points for training and {} for testing with {} features".format(
train_x.shape[0], test_x.shape[0], test_x.shape[1]))
# ***
#
# ## Data Pre-processing
# +
# pre-process Pipeline
preprocessing = Pipeline([
('general', general_preprocesing_pipeline(train_x)), # see the code for general_preprocesing_pipeline
('polyfeaturs', PolynomialFeatures(degree=2)),
('zerovar', VarianceThreshold(threshold=0.0)),
('scaling', StandardScaler()) # I scale again not all polynomial features may be with scaled
])
preprocessing.fit(train_x)
train_x_transformed = preprocessing.transform(train_x)
# -
train_x_transformed.shape
# ***
# ## Linear Regression
# +
lm_reg = Pipeline([
('lm', LinearRegression())
])
lm_grid = dict()
lm_search = modelevaluation.gridsearch(lm_reg, lm_grid, train_x_transformed, train_y)
# -
# ***
# ## PLS regression
# +
pls_reg = Pipeline([
('pls', PLSRegression())
])
pls_grid = dict(
pls__n_components = np.arange(6, 15, 1)
)
pls_search = modelevaluation.gridsearch(pls_reg, pls_grid, train_x_transformed, train_y)
# -
# ***
#
# ## Lasso
lasso = Lasso()
alphas = np.logspace(-4, -0.5, 10)
lasso_grid = [{'alpha': alphas}]
lasso_search = modelevaluation.gridsearch(lasso, lasso_grid, train_x_transformed, train_y)
# ***
#
# ## Elastic Net
enet = ElasticNet()
alphas = np.logspace(-4, -0.5, 10)
enet_grid = [{'alpha': alphas, 'l1_ratio' : np.linspace(0, 1, 5)}]
enet_search = modelevaluation.gridsearch(enet, enet_grid, train_x_transformed, train_y)
# ## Validation Data Test
mymodels = {
'linear_reg': lm_search.best_estimator_,
'PLS': pls_search.best_estimator_,
'lasso': lasso_search.best_estimator_,
'enet': enet_search.best_estimator_
}
modelevaluation.eval_models(mymodels, preprocessing, test_x, test_y).to_csv("results_data/val_linearmodels.csv")
# ## 10-FOLD CV
#
# Cross validate the best scoring model to have a profile.
results = modelevaluation.crossvalidation(mymodels, train_x_transformed, train_y)
results.to_csv('results_data/cv_linearmodels.csv', index=False)
| results/19-04-30-PredictiveModelDecayAllSpecies/19-05-01-TrainModels/01-LinearModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Set up
# +
# %matplotlib inline
# %run fcast.py
# customize matplotlib plots
mpl.style.use('seaborn-darkgrid')
plt.rc('font', family='TH Sarabun New')
plt.rc('xtick', labelsize=12)
plt.rc('ytick', labelsize=12)
plt.rc('axes', labelsize=16)
plt.rc('figure', figsize=(6.5,4), dpi=300)
# -
# ## Workflow
#
# 1. **Prepare data**
# 1. **Transform** data i.e. taking log, scaling, and differencing.
# > If data is transformed, prediction must be converted by **inversing transformation** operations.
# 2. **Split** data into three sets e.g. train, validation and test set. For example, 60:20:20 proportion.
# 3. For deep learning, we need to reshape data **to [Keras](https://keras.io)** format.
# 1. Number of periods to used as model's input.
# 2. Number of periods to forecast as model's output.
#
#
# 2. **Grid search** for the best hyper-parameters (configs).
# 1. For each hyper-parameter, **fit the model** using **train set**.
# 2. **Walk forward validation** over the **validation set**.
# 3. **Measure** the model's performance via some metrics. In this case, we use RMSE.
# 4. For deep learning techniques, **repeat** step A-C n times because deep learning is stochastic.
# 5. **Select** the best config based on RMSE.
#
#
# 3. **Forecast**.
# 1. **Combine** train and validation set called it **train2 set**.
# 2. **Refit model** using **train2 set** and the config from step 1.
# 3. **Walk forward validation** over the **test set**.
# 4. **Measure** the model's performance using RMSE.
# 5. For deep learning techniques, **repeat** step B-D.
# 6. **Return** model's performance, predicted.
#
#
# 4. **Evaluate** models including
# - Naive or Persistent model (NAIVE)
# - Exponential Smoothing (ES)
# - Seasonal Autoregressive Integrated Moving Average (SARIMA)
# - Multilayer Perceptron (MLP)
# - Long Short-Term Memory (LSTM)
# ## Structure of defined functions
# 1. `data_` means data preparation steps
# 1. `data_split`
# 2. `data_transform`
# 3. `data_reshape`
#
#
# 2. `model_` means model steps
# 1. `model_config`
# 2. `model_fit`
# 3. `model_predict`
#
#
# 3. `eval_`
#
#
# 4. `plot_`
# # Prepare data
# ## Import price data
#
# โหลดข้อมูลราคาข้าวเปลือกที่เกษตรกรขายได้ 3 ชนิดข้าว ได้แก่ ข้าวหอมมะลิ (`hml`) ข้าวเจ้าขาว (`wht`) และข้าวเหนียว (`glu`)
# นอกจากนี้ ยังเพิ่มข้อมูล metadata ประกอบด้วยตัวย่อ(abb) ชื่อย่อ (shortname) และ ชื่อเต็ม(fullname)
# +
hml = read_price('hml')
hml.meta = SimpleNamespace()
hml.meta.abb = 'hml'
hml.meta.shortname = 'ข้าวหอมมะลิ'
hml.meta.fullname = 'ราคาข้าวเปลือกหอมมะลิ ที่เกษตรกรขายได้ ณ ไร่นา'
wht = read_price('wht')
wht.meta = SimpleNamespace()
wht.meta.abb = 'wht'
wht.meta.shortname = 'ข้าวเจ้าขาว'
wht.meta.fullname = 'ราคาข้าวเปลือกเจ้าขาว ที่เกษตรกรขายได้ ณ ไร่นา'
glu = read_price('glu')
glu.meta = SimpleNamespace()
glu.meta.abb = 'glu'
glu.meta.shortname = 'ข้าวเหนียว'
glu.meta.fullname = 'ราคาข้าวเปลือกเหนียวเมล็ดยาว ที่เกษตรกรขายได้ ณ ไร่นา'
prices = [hml, wht, glu]
# -
# ## Policy data
# เพิ่มข้อมูล dummy ปีที่มีนโยบายจำนำข้าวทุกเมล็ดของรัฐบาลยิ่งลักษณ์ ระหว่าง Oct 2010 - Feb 2014
exo = pd.DataFrame(index = wht.index)
exo['pledge_scheme'] = 0
exo.loc['2011-10':'2014-2'] = 1
# ## Data Transformation
for price in prices:
price.scaled, price.scaler = data_transform(price, method=None)
price.train, price.validate, price.train2, price.test = data_split(price.scaled)
print(price.meta.abb, price.train.size, price.validate.size, price.test.size)
# # Exponential Smoothing
# ## Generate Parameter Grid
# +
# trend = [None]
# damped = [False]
# seasonal = [None, 'add', 'mul']
# conf1 = model_configs(trend, damped, seasonal)
trend = ['add', 'mul']
damped = [False, True]
seasonal = [None, 'add', 'mul']
# conf2 = model_configs(trend, damped, seasonal)
# ets_configs = [*conf1, *conf2]
ets_configs = model_configs(trend, damped, seasonal)
ets_configs
# -
# ## Define function
# +
def ets_fit(data, config, n_periods=12):
trend, damped, seasonal = config
return ets(data, trend=trend, damped=damped,
seasonal=seasonal, seasonal_periods=n_periods
).fit(use_boxcox='log')
def ets_forecast(model_fitted, data, n_forecast=4):
scaler = getattr(data, "scaler")
df = pd.DataFrame(model_fitted.forecast(n_forecast),
columns=["yhat_scaled"])
df['yhat'] = scaler.inverse_transform(df)
return df['yhat']
def ets_walk_forward(data, config, search_mode=False, n_forecast=4):
train, test = select_traintest(data, search_mode)
data_ = pd.concat([train,test])
N = data_.shape[0]
n_test = test.shape[0]
n_yhat = n_test+1 # +1 for count adjustment
yhat = np.empty((n_yhat, n_forecast))
history = train
for i in range(n_yhat):
model_fitted = ets_fit(history, config)
yhat[i] = ets_forecast(model_fitted, data, n_forecast)
if i < n_yhat-1:
history = history.append(test.iloc[i])
yhat = pd.DataFrame(yhat,
columns=[f'yhat{h}' for h in range(1, n_forecast+1)],
index=data_.index[-n_test-1:])
return yhat
# -
# test
config = ets_configs[0]
fitted = ets_fit(hml.scaled, config)
yhat = ets_walk_forward(hml, config, search_mode=True)
scores = model_measure(hml, yhat, config)
scores
# # Run
ets_model = [ets_fit, ets_forecast, ets_walk_forward, ets_configs]
# %%time
n_forecast=4
for price in prices:
price.ets_grid_result, price.ets_config = grid_search(price, ets_model, n_forecast=n_forecast)
price.ets_yhat = forecast(price, ets_model, price.ets_config, n_forecast=n_forecast)
# %%time
n_forecast=4
for price in prices:
price.ets_grid_result, price.ets_config = grid_search(price, ets_model, n_forecast=n_forecast)
price.ets_yhat = forecast(price, ets_model, price.ets_config, n_forecast=n_forecast)
for price in prices:
print(price.ets_config)
# +
path = '../results/result_ets_4.pkl'
if(os.path.isfile(path)):
result = joblib.load(path)
else:
result = dict()
for price in prices:
result[price.meta.abb] = [price.ets_grid_result, price.ets_config, price.ets_yhat]
joblib.dump(result, path)
# -
plot_fcast(glu, 'ets', step=4)
model_measure(hml, hml.ets_yhat, [""])
model_measure(wht, wht.ets_yhat, [""])
model_measure(glu, glu.ets_yhat, [""])
| notebook/2_ets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # THE SPARKS FOUNDATION
# ### GRIP - DATA SCIENCE AND BUSINESS ANALYTICS INTERN - JULY 2021
# #### Submitted by : <NAME>
# #### Task -1 : Prediction using Supervised Machine Learning
# ### Problem Statement : Predict the percentage of a student based on no. of study hours using supervised machine learning
# #### In supervised machine learning, a model is able to predict with the help of labeled dataset.Supervised ML is of two types :
# 1)Classification<br>2)Regression<br>Here, Regression is used as the problem is about relationship between two variables where a change in one variable is associated with a change in other variable.
# #### Importing required libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
# #### Importing data from CSV file
df = pd.read_csv("Task - 1 dataset.csv")
df.head(5)
df.tail(5)
# #### Understanding data
df.info()
# Type of attributes :<br>Hours - Continuous Variable<br>Scores - Discrete Variable
#
# Also, there are no missing values.
# #### Outliers
plt.figure(figsize=(8,6))
sns.boxplot(data=df)
plt.show()
# There are no outliers in both the columns.
# #### Descriptive Statistics
df.describe()
# #### Scatter plot
plt.scatter(df.Hours,df.Scores,color = "red")
plt.title("Hours Vs Scores")
plt.xlabel("Hours")
plt.ylabel("Scores")
plt.show()
# From the graph above, we can clearly see that there is a positive linear relation between the number of hours studied and percentage of score.
df.corr()
# There is a strong positive correlation between the two variables - "Hours" and "Scores".
# #### Splitting data into training(80%) and testing(20%) sets
# +
x = df[['Hours']]
y = df['Scores']
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size = 0.2,random_state = 0)
# -
# #### Modelling
reg = LinearRegression()
reg.fit(x_train,y_train)
print("Training done!")
reg.coef_
reg.intercept_
print("Linear Regression equation is Scores = ",np.round(reg.coef_[0],2),"* Hours + ",np.round(reg.intercept_,2))
# #### Plotting Regression Line
plt.scatter(x,y,color = "red")
plt.plot(x,reg.predict(x),color = 'blue')
plt.title("Hours Vs Scores")
plt.xlabel("Hours")
plt.ylabel("Scores")
plt.show()
# #### Predicting scores of test data
y_pred = reg.predict(x_test)
y_pred
# #### Comparing Actual and Predicted scores of test data
df2 = pd.DataFrame({"Actual Score" : y_test,"Predited Score" : y_pred})
df2
# #### Evaluating the model
print('Mean absolute error = ',metrics.mean_absolute_error(y_test,y_pred))
print('Mean squared error = ',metrics.mean_squared_error(y_test,y_pred))
print('Root Mean squared error = ',np.sqrt(metrics.mean_squared_error(y_test,y_pred)))
print('R2 Score = ',metrics.r2_score(y_test,y_pred))
# #### Predicting score if a student studies for 9.25 hrs/ day?
hours = 9.25
z = reg.predict([[hours]])
print("No. of hours = ",hours)
print("Predicted score = ",z[0])
| Task - 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 4 : Regression Analysis using Text Data
# Use regression to predict overall scores of reviews of Musical Instruments present in Amazon
#
# The dataset can be obtained from http://jmcauley.ucsd.edu/data/amazon/ http://snap.stanford.edu/data/amazon/productGraph/categoryFiles/reviews_Musical_Instruments_5.json.gz
# Ups and downs: Modeling the visual evolution of fashion trends with one-class collaborative filtering <NAME> WWW, 2016
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import re
import string
from nltk import word_tokenize
from nltk.stem import WordNetLemmatizer
from sklearn.feature_extraction.text import TfidfVectorizer
from collections import Counter
from pylab import *
import nltk
import warnings
warnings.filterwarnings('ignore')
review_data = pd.read_json('data_ch3/reviews_Musical_Instruments_5.json', lines=True)
review_data.head()
review_data['overall'].value_counts()
lemmatizer = WordNetLemmatizer()
review_data['cleaned_review_text'] = review_data['reviewText'].apply(\
lambda x : ' '.join([lemmatizer.lemmatize(word.lower()) \
for word in word_tokenize(re.sub(r'([^\s\w]|_)+', ' ', str(x)))]))
review_data[['cleaned_review_text', 'reviewText', 'overall']].head()
tfidf_model = TfidfVectorizer(max_features=500)
tfidf_df = pd.DataFrame(tfidf_model.fit_transform(review_data['cleaned_review_text']).todense())
tfidf_df.columns = sorted(tfidf_model.vocabulary_)
tfidf_df.head()
# ## Linear Rergression
from sklearn.linear_model import LinearRegression
linreg = LinearRegression()
linreg.fit(tfidf_df,review_data['overall'])
linreg.coef_
linreg.intercept_
linreg.predict(tfidf_df)
review_data['predicted_score_from_linear_regression'] = linreg.predict(tfidf_df)
review_data[['overall', 'predicted_score_from_linear_regression']].head(10)
# Reference / Citation for the dataset: http://jmcauley.ucsd.edu/data/amazon/
# http://snap.stanford.edu/data/amazon/productGraph/categoryFiles/reviews_Musical_Instruments_5.json.gz
# Ups and downs: Modeling the visual evolution of fashion trends with one-class collaborative filtering
# <NAME>, <NAME> WWW, 2016
| Lesson03/Exercise32.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regressione usando la teoria Bayesiana
#
# La regressione lineare che abbiamo visto fino ad ora usa modello di tipo deterministico per analizzare e predire dati, ma è anche possibile usare modelli di tipo probabilistico per effettuare una regressione lineare, vediamone alcuni modelli, per maggiori info visitate questo __[link](https://scikit-learn.org/stable/modules/linear_model.html#bayesian-regression)__.
#
# All'interno di questi modelli si usa il Teorema di Bayes che stabilisce la seguente formula:
#
# \begin{equation}
# \Large P(H_{0}|E) = \frac{P(E|H_{0})P(H_{0})}{P(E)}
# \end{equation}
#
# Dove $P(H_{0}|E)$ è detta probabilità di $H_{0}$, dato un evento $E$, $P(H_{0})$ è detta probabilità a priori di $H_{0}$, $P(E)$ è detta probabilità di osservare $E$ detta ***probabilita marginale***, mentre $P(E|H_{0})$ è la **funzione verosomigliante** ovvero una funzione di probabilità condizionata in cui **il primo argomento($E$) è fisso, mentre il secondo ($H_{0}$) è variabile**.
#
# In questi tipi di modelli si introduce una distribuzione a priori non informativa sugli iperparametri che dovranno essere poi determinati grazie ai dati attraverso il teorema precedente, per fare un esempio ipotizziamo di avere dei dati che noi pensiamo siano distribuiti secondo una gaussiana, allora il modello dovrà determinare:
#
# \begin{equation}
# \Large p(y | X,w,\alpha) = \mathcal{N}(Xw, \alpha)
# \end{equation}
#
# Ovvero una distribuzione probabilistica di tipo gaussiana centrata in $Xw$ e con una varianza $\alpha$ che deve essere determinata dai dati.
#
# **Tra i vantaggi abbiamo che:**
#
# - questi modelli si adattano ai dati per definizione
# - possiamo introdurre termini di regoralizzazione per renderli più robusti
#
# **Tra gli svantaggi abbiamo che:**
#
# - per fare inferenza statistica si usa il teorema di Bayes che è computazionalmente molto oneroso in termini di risorse.
# ## Bayesian Ridge Regression
#
# Questo modello di regressione bayesiano usa un termine di regoralizzazione $l_2$, come nella Ridge Regression, dal punto di vista probabilistico questo modello usa una distribuzione gaussiana e determina i parametri usando:
#
# \begin{equation}
# \Large p(w|\lambda) = \mathcal{N}(w|0, \lambda^{-1}I_p)
# \end{equation}
#
# Per essere precisi al fine di determinare i pesi si utilizza la **Maximum a posteriori (MAP) usando la log marginal likelihood** ovvero:
#
# \begin{equation}
# \Large W = max_{w} \quad exp[-\frac{(y-X\beta)^{T}(y-X\beta)}{2\sigma^{2}} -\frac{\parallel \beta \parallel^{2}_{2}}{2\tau^2}]\\
# \Large W = min_{w} (y-X\beta)^{T}(y-X\beta) + \lambda \parallel \beta \parallel^{2}_{2}
# \end{equation}
#
# Sono presenti altri parametri nell'implementazione dell'algoritmo, quello che ci serve sapere è che gli $\alpha$ e i $\beta$ sono dei parametri per la definizione di una __[gamma distribution](https://en.wikipedia.org/wiki/Gamma_distribution)__ che è una distribuzione continua e che tali parametri sono definiti dai dati a partire da dei valori iniziali e quelli che non sono informativi, ovvero forniscono una vaga informazione sulla variabile sono di solito settati piccoli in modo da non influire sulla determinazione della distribuzione, per maggiori dettagli consultate __[scikit](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.BayesianRidge.html#sklearn.linear_model.BayesianRidge)__.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import BayesianRidge
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
#load the dataset
data = load_boston()
df = pd.DataFrame(data.data, columns = data.feature_names)
df['MEDV'] = data.target
#prepare the data
X_train, X_test, y_train, y_test = train_test_split(df.LSTAT.values, df.MEDV.values,
random_state=0, test_size = 0.2)
bayesian_ridge = BayesianRidge(compute_score=True)
bayesian_ridge.fit(X_train.reshape(-1, 1), y_train)
#setting plot
fig, ax = plt.subplots(figsize=(8, 6))
ax.scatter(X_train, y_train, label = "train", color = "blue")
ax.scatter(X_test, y_test, label = "test", color = "green")
line, = ax.plot(X_train, bayesian_ridge.predict(X_train.reshape(-1, 1)), label = "fit", color = "red")
plt.legend()
plt.show()
print('Bayesian Regression parameters estimated:')
print(f'Estimated precision of noise: {bayesian_ridge.alpha_}')
print(f'Estimated precision of weights: {bayesian_ridge.lambda_}')
print(f'Estimated variance-covariance matrix of the weights: {bayesian_ridge.sigma_}')
print(f'Number of iterations applied to obtain estimated parameters: {bayesian_ridge.n_iter}')
print(f'Value of the log marginal likelihood (to be maximized) at each iteration: {bayesian_ridge.scores_}')
print(f'Coefficients obtained: {bayesian_ridge.coef_} ')
print(f'R^2 score on training set: {bayesian_ridge.score(X_train.reshape(-1, 1), y_train)},'
f'test set: {bayesian_ridge.score(X_test.reshape(-1, 1), y_test)}')
# +
#use bayesian regression for multiple features
X, Y = df.drop('MEDV', axis = 1).values, df.MEDV.values
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, random_state=0, test_size = 0.2)
bayesian_ridge = BayesianRidge(compute_score=True)
bayesian_ridge.fit(X_train, y_train)
print('Bayesian Regression parameters estimated:')
print(f'Estimated precision of noise: {bayesian_ridge.alpha_}')
print(f'Estimated precision of weights: {bayesian_ridge.lambda_}')
print(f'Estimated variance-covariance matrix of the weights:\n {bayesian_ridge.sigma_}')
print(f'Number of iterations applied to obtain estimated parameters: {bayesian_ridge.n_iter}')
print(f'Value of the log marginal likelihood (to be maximized) at each iteration: {bayesian_ridge.scores_}')
print(f'Coefficients obtained: {bayesian_ridge.coef_}')
print(f'R^2 score on training set: {bayesian_ridge.score(X_train, y_train)},'
f'test set: {bayesian_ridge.score(X_test, y_test)}')
# -
# ## Logistic Regression
#
# La **logistic regression sebbene il nome non è in genere usato per fare la regressione, ma per fare classificazione**, la base su cui si fonda la logistic regression è la funzione logistica che ha la seguente formula:
#
# \begin{equation}
# \Large f(x) = C \frac{1}{1 + e^{-k(x-x_0)}}
# \end{equation}
#
# dove $C$ rappresenta il valore massimo della funzione, $k$ rappresenta la ripidità con cui sale la curva, $x_0$ rappresenta il punto in cui la funzione assume valore $f(x_0) = \frac{1}{2}$.
# +
import numpy as np
def logistica(X, C = 1, k = 1, x0 = 0):
return C /(1 + np.exp(-k *(X-x0)))
X = np.linspace(-10,10,100)
fig, axs = plt.subplots(1, 4, figsize = (18,6))
fig.suptitle("Funzione logistica per diversi parametri")
axs[0].plot(X, logistica(X))
axs[0].set_title("C = 1, k = 1, x0 = 0")
axs[0].grid(True)
axs[1].plot(X, logistica(X, C = 2))
axs[1].set_title("C = 2, k = 1, x0 = 0")
axs[1].grid(True)
axs[2].plot(X, logistica(X, k = 10))
axs[2].set_title("C = 1, k = 10, x0 = 0")
axs[2].grid(True)
axs[3].plot(X, logistica(X, x0 = 5))
axs[3].set_title("C = 1, k = 1, x0 = 5")
axs[3].grid(True)
plt.show()
# -
# Ora che abbiamo capito come è fatta la funzione logistica, vediamo di capire cosa fa la logistic regression guardando la formula usata per l'optimizzazione in scikit:
#
# \begin{equation}
# \Large \min_{w,c} \frac{1-\rho}{2} w^{T}w + \rho \parallel w \parallel_{1} + C \sum_{i=1}^{n} log(exp(-y_i (X_{i}^{T} w + c)) +1)
# \end{equation}
#
# In questa equazione possiamo vedere che abbiamo sia il termine di regoralizzazione $l2$, che $l1$, il cui peso nella formula ci dice in quale misura stiamo usando le regoralizzazioni, abbiamo poi un valore $C$ iperparametro dell'equazione che dobbiamo determinare per capire in che misura vogliamo che la regoralizzazione sia determinante ed notare bene **usiamo il log sulle funzione logistiche, questo perché in questa maniera si riduce la possibilità di avere numeri enormi e computazionalmente è più veloce**.<br>
# Per valutare la qualità del modello nella classificazione si possono utilizzare diverse metriche, per capire come funzionano in genere bisogna capire cosa si intendono con True Positive(TP), False Postive(FP), True Negative(TN), False Negative(FN). Al fine di capire come funziona cerchiamo di rappresentarli usando una __[confusion matrix](https://scikit-learn.org/stable/modules/model_evaluation.html#confusion-matrix)__ che è qui rappresentata in maniera generica:
#
# | Actual/Predicted | True | False | | |
# |:----------------:|:--------------:|:--------------:|---|---|
# | True | True Positive | False Positive | | |
# | False | False Negative | True Negative | | |
#
# In questa matrice possiamo notare che il caso ideale sarebbe se i valori sono tutti sulla diagonale poiché il modello dovrebbe essere in grado di classificare con accuratezza assoluta il dataset. In genere però per aver una singola metrica che contenga tutte le informazioni riguardo alla sua capacità di classificazione in maniera quantitativa, tra le più usate abbiamo:
# - precision : $ \Large \frac{tp}{tp+tn} $
# - accuracy : $ \Large \frac{\sum_{i = 0}^{N-1} I(y_{pred} = y_{true})}{N} $ con $I$ funzione di identità che fornisce il valore 1 solo se la condizione è soddisfatta e $N$è il numero di sample
# - recall : $ \Large \frac{tp}{tp + fn} $
# - f1 score: $ 2* \Large \frac{precision * recall}{precision + recall} $
# Esistono molte altre metriche per la classificazione per maggiori info consultate __[scikit](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics)__.<br>
# La logistic regression può essere usata sia in condizioni di classificazione binaria che in condizione di classificazione di classi multiple , vediamo degli esempi in tal caso.
# +
from sklearn.linear_model import LogisticRegressionCV
from sklearn.metrics import classification_report
from mlxtend.plotting import plot_decision_regions
#load data
df = pd.read_csv('../data/diabetes2.csv')
print('Dataset of diabetes used:')
display(df)
#prepare data
X_train, X_test, y_train, y_test = train_test_split(df.drop('Outcome', axis = 1).values,
df.Outcome.values, random_state=0, test_size = 0.2)
target_names = ['class 0', 'class 1']
#get the best classifier given training data
logistic = LogisticRegressionCV(class_weight= 'balanced', cv = 10, max_iter = 1e4).fit(X_train, y_train)
print(f'Logisitc regression score using 10 folds:\n {logistic.scores_}')
print('-'*80)
print(f'Training accuracy score: {logistic.score(X_train, y_train)}, Test accuracy score:{logistic.score(X_test, y_test)}')
print('-'*80)
print(classification_report(y_test, logistic.predict(X_test), target_names=target_names))
#create output dataframe
output = {'Probability class 0': logistic.predict_proba(X_test)[:,0],
'Probability class 1': logistic.predict_proba(X_test)[:,1],
'Predicted': logistic.predict(X_test),
'True': y_test}
test_results = pd.DataFrame(data = output, index = np.arange(0, y_test.shape[0]))
print('Dataframe of the results with the classes predicted, true and probability associated')
display(test_results)
# -
# Per vedere come funziona con più classi potete vedere questo __[esempio di scikit](https://scikit-learn.org/stable/auto_examples/linear_model/plot_logistic_multinomial.html#sphx-glr-auto-examples-linear-model-plot-logistic-multinomial-py)__ in cui potete anche vedere anche al differenza dei due tipi di algoritmi di classificazione OnevsRest and Multinomial per maggiori info andate __[qui](https://scikit-learn.org/stable/modules/multiclass.html?highlight=onevsrest)__.
#
# ## Polynomial Regression
#
# Fino ad ora abbiamo visto metodi lineari che riuscivano a classificare bene, ma in molti casi potrebbe capitare che il dataset presenta **non linearità, in tal caso dobbiamo mappare il dataset in un nuovo spazio in cui possiamo usare i metodi lineari**, scikit in tal caso fornisce come soluzione la trasformazione di dati attraverso una funzione polinomiale oppure un applicazione di una __[kernel ridge regression](https://scikit-learn.org/stable/modules/kernel_ridge.html)__ simile alla futura Support Vector Machine, ma con piccole differenze.
# Cerchiamo di creare un problema per cui sia necessario tale situazione e vediamo come applicare questi strumenti usando inoltre la __[pipeline scikit](https://scikit-learn.org/stable/modules/compose.html#combining-estimators)__.
# +
from sklearn.preprocessing import PolynomialFeatures
x = np.arange(20).reshape(10,2)
print(f'Dati originali:\n{x}')
poly = PolynomialFeatures(degree=2)
print(f'Dati tansformati usando un polinomio di grado 2:\n {poly.fit_transform(x)}')
poly = PolynomialFeatures(degree=2, interaction_only= True)
print(f'Dati tansformati usando un polinomio di grado 2 contando solo i termini'
f'misti, le identità e il termine di grado nullo:\n{poly.fit_transform(x)}')
# -
# Possiamo quindi vedere come i dati originali nella forma $[x_0, x_1]$ sono stati trasformati nella forma $[1, x_0, x_1, x_{0}^2, x_{0}x_{1}, x_{1}^2]$ mentre con il caso `intercat_only = True` ritorna $[1, x_0, x_1, x_{0}x_{1}]$, vediamo ora come usarlo nel caso di un dataset non lineare.
# +
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import make_pipeline
x = np.arange(100)
y = 2 * x**3 + 3 * x**2 + 5 * x + 4
# generate points used to plot
x_plot = np.linspace(0, 100, 100)
# create matrix versions of these arrays
X = x[:, np.newaxis]
X_plot = x_plot[:, np.newaxis]
colors = ['blue', 'green', 'red']
fig, axs = plt.subplots(figsize = (12,8))
plt.scatter(x, y, color='navy', s=30, marker='o', label="training points")
#polynomial features
for count, degree in enumerate([1, 2, 3]):
model = make_pipeline(PolynomialFeatures(degree), LinearRegression())
model.fit(X, y)
y_plot = model.predict(X_plot)
plt.plot(x_plot, y_plot, color=colors[count],
label="degree %d" % degree)
plt.legend()
plt.show()
# -
# Abbiamo quindi capito come applicare trasformazioni lineari, in futuro scopriremo che questo approcio è molto sconveniente dal punto di vista computazionale a causa della dimensione sempre maggiore da tenere in conto, per questo in futuro vedremo le __[SVM](https://scikit-learn.org/stable/modules/svm.html#svm)__ per ovviare a questi problemi.
#
# ---
#
# COMPLIMENTI AVETE FINITO IL NOTEBOOK 4 DI MACHINE LEARNING!
| 3.machine learning/.ipynb_checkpoints/4-Bayesian_Logistic_Polynomial-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import matplotlib.pyplot as plt, seaborn as sn
import pandas as pd, numpy as np
sn.set_context('notebook')
# # Generalised Likelihood Uncertainty Estimation (GLUE)
#
# GLUE is a framework for model calibration and uncertainty estimation that has become popular in recent years, especially within the UK hydrological community. The approach is well documented in the academic literature ([Beven, 2006](http://www.sciencedirect.com/science/article/pii/S002216940500332X), for example, provides a comprehensive overview) but it is also controversial, in the sense that many authors consider the method to be both [statistically incoherent](http://onlinelibrary.wiley.com/doi/10.1029/2008WR006822/abstract) and [computationally inefficient](http://www.sciencedirect.com/science/article/pii/S0309170807001856).
#
# The more I learn, the more I'm inclined to agree with those who feel GLUE is **not** an appropriate tool for model calibration and uncertainty estimation. For anyone who has yet to make a decision, I strongly recommend reading the literature on the subject, including the exchanges between leading proponents on both sides of the argument. For example:
#
# * [Mantovan & Todino (2006)](http://www.sciencedirect.com/science/article/pii/S0022169406002162) then [Beven *et al.* (2007)](http://www.sciencedirect.com/science/article/pii/S0022169407001230) then [Mantovan & Todino (2007)](http://www.sciencedirect.com/science/article/pii/S0022169407001242) <br><br>
#
# * [Clark *et al.* (2011)](http://onlinelibrary.wiley.com/doi/10.1029/2010WR009827/abstract) then [Beven *et al.* (2012)](http://onlinelibrary.wiley.com/doi/10.1029/2012WR012282/abstract) then [Clark *et al.* (2012)](http://onlinelibrary.wiley.com/doi/10.1029/2012WR012547/abstract)
#
# Two of the reasons GLUE has become so popular are that it is **conceptually simple** and **easy to code**. Such advantages are not easily ignored, especially among environmental scientists who are typically neither professional statisticians nor computer programmers. Although most would-be modellers are aware of some debate in the literature, many lack the statistical background to be able to follow the arguments in detail. What's more, many understandably take the view that, if the issue is still a matter for discussion between statisticians, either method will probably be adequate for a first foray into environmental modelling.
#
# The aim of this notebook is to provide an introduction to some of the key issues, and to make it easier to follow the more detailed assessments in the academic literature. We will begin by comparing the frequentist, Bayesian and GLUE approaches to **simple linear regression**.
#
# I will assume familiarity with frequentist **Ordinary Least Squares (OLS)** regression, and if you've worked through the previous notebooks you should also have a basic understanding of formal Bayesian inference and the differences between e.g. Monte Carlo and MCMC sampling. I'll try to provide a reasonable overview of GLUE, but if you're not familar with the technique already I'd recommend reading e.g. [Beven (2006)](http://www.sciencedirect.com/science/article/pii/S002216940500332X) for a more complete summary.
#
# A much more comprehensive and detailed investigation of the limitations of GLUE is provided by [Stedinger *et al.* (2008)](http://onlinelibrary.wiley.com/doi/10.1029/2008WR006822/abstract).
#
# ## Three approaches compared
#
# We will consider the following:
#
# 1. **Frequentist OLS regression**. This is just the usual approach to linear regression that most people are familiar with. <br><br>
#
# 2. **Bayesian MCMC**. A formal Bayesian approach, exactly the same as introduced in [section 7 of notebook 4](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/04_MCMC.ipynb#7.-Putting-it-all-together). <br><br>
#
# 3. **Monte Carlo GLUE**. A "limits of acceptability" approach using an *informal* (or *pseudo-*) likelihood function. The most common implementation of GLUE uses Monte Carlo sampling, similar to some of the techniques described in [notebook 3](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/03_Monte_Carlo.ipynb).
#
# It's worth emphasising straight away that using **numerical simulation approaches** such as Bayesian MCMC or GLUE to solve a simple linear regression problem is a case of using a very large sledgehammer to crack a very small nut. It is extremely unlikey that you would ever use either of these techniques for this kind of analsis in practice. However, if an approach is going to generalise well to more complex problems, it's often a good idea to check it works for simple problems too.
#
# Simple linear regression is just a basic form of parameter inference: we want to infer the **slope** and **intercept** of our regression line, subject to a particular error model. The simplest form of linear regression assumes **independent and identically distributed** Guassian erros with mean zero.
#
# $$y = ax + b + \epsilon \qquad where \qquad \epsilon \sim \mathcal N(0, \sigma_\epsilon)$$
#
# We will start by generating some synthetic data based on the equation above and we'll then use the three methods to estimate the **regression parameters** and associated **confidence intervals**. The reason for doing this is to **check that the two more complicated approaches gives results that are broadly consistent with the simple frequentist method** (which is very well established).
# ## 1. Generate synthetic data
# +
# Choose true params
a_true = 3
b_true = 6
sigma_true = 2
n = 100 # Length of data series
# For the independent variable, x, we will choose n values equally spaced
# between 0 and 10
x = np.linspace(0, 10, n)
# Calculate the dependent (observed) values, y
y = a_true*x + b_true + np.random.normal(loc=0, scale=sigma_true, size=n)
# Plot
plt.plot(x, y, 'ro')
plt.plot(x, a_true*x + b_true, 'k-')
plt.xlabel('x')
plt.ylabel('y')
plt.title('Observed data')
plt.show()
# -
# ## 2. Frequentist linear regression
#
# There are [several ways](https://en.wikipedia.org/wiki/Linear_regression#Least-squares_estimation_and_related_techniques) of performing simple linear regression, but the most commonly used is the OLS approach, which minimises the sum of squared model residuals. OLS regrssion under the assumption of **independent and identically distributed (iid) Gaussian errors** is so widely used that many software packages make the analysis very easy - so easy, in fact, that people often forget to **check** whether the iid assumption has actually been satisfied. In the examples below we won't check either, but that's because we *know* our test data was generated using iid errors, so we don't need to.
#
# ### 2.1. Fit the model
#
# We'll use `statsmodels` to perform the regression in Python, including estimating 95% **confidence intervals** for the slope and intercept ($a$ and $b$, respectively). We must also estimate the error standard deviation , $\sigma_\epsilon$ (we'll ignore the confidence interval for this for now, because it's not provided by `statsmodels` by default).
# +
import statsmodels.api as sm
from statsmodels.sandbox.regression.predstd import wls_prediction_std
# Add intercept for model
X = sm.add_constant(x)
# Fit
model = sm.OLS(y,X)
result = model.fit()
# Regression summary
print result.summary()
print '\n'
# Key results as dataframe
freq_df = pd.DataFrame(data={'a_freq':[result.conf_int()[1,0],
result.params[1],
result.conf_int()[1,1]],
'b_freq':[result.conf_int()[0,0],
result.params[0],
result.conf_int()[0,1]],
'sigma_freq':[np.nan,
(result.scale)**0.5,
np.nan]},
index=['2.5%', '50%', '97.5%'])
print freq_df.T
# -
# ### 2.2. Plot the result
#
# We can now plot the **median** regression line plus the **95% confidence interval** around it.
# +
# Plot predicted
prstd, low, up = wls_prediction_std(result, alpha=0.05) # 95% interval
plt.fill_between(x, low, up, color='r', alpha=0.3)
plt.plot(x, result.fittedvalues, 'r-', label='Estimated')
plt.title('Frequentist')
# Plot true
plt.plot(x, y, 'bo')
plt.plot(x, a_true*x+b_true, 'b--', label='True')
plt.xlabel('x')
plt.ylabel('y')
plt.legend(loc='best')
# -
# The estimated "best-fit" line is very close to the true one. Also, if our 95% confidence interval is correct, we should expect roughly 95% of the observations to lie within the shaded area. This proportion is often called the "**coverage**".
#
# ### 2.3. Estimate coverage
# +
# Data frame of lower CI, upper CI and observations
cov_df = pd.DataFrame({'low':low,
'obs':y,
'up':up})
# Are obs within CI?
cov_df['In_CI'] = ((cov_df['low'] < cov_df['obs']) &
(cov_df['up'] > cov_df['obs']))
# Coverage
cov = 100.*cov_df['In_CI'].sum()/len(cov_df)
print 'Coverage: %.1f%%' % cov
# -
# The coverage from the frequentist approach is correct, as expected.
#
# ## 3. Bayesian linear regression
#
# For this problem, the Bayesian approach is significantly more complicated than the frequentist one. One of the real benefits of the Bayesian method, though, is its generality i.e. it doesn't necessarily become any more complicated when applied to challenging problems. As demonstrated in notebooks [4](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/04_MCMC.ipynb) and [6](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/06_Beyond_Metropolis.ipynb), the Bayesian approach is essentially the same regardless of whether you're performing simple linear regression or calibrating a hydrological model. It's worth bearing this in mind when working though the following sections.
#
# ### 3.1. Define the likelihood, prior and posterior
#
# The likelihood, prior and posterior are defined in exactly the same way as in [section 7 of notebook 4](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/04_MCMC.ipynb#7.-Putting-it-all-together). Note that for the likelihood function we're required to explicitly define an **error structure**. This was not necessary for the frequentist approach above because `statsmodels.api.OLS` implicitly assumes iid Gaussian errors. For more complex error schemes, we'd need to specify the error struture for the frequentist analysis too.
# +
from scipy.stats import norm
def log_likelihood(params, x, obs):
""" Calculate log likelihood assuming iid Gaussian errors.
"""
# Extract parameter values
a_est, b_est, sigma_est = params
# Calculate deterministic results with these parameters
sim = a_est*x + b_est
# Calculate log likelihood
ll = np.sum(norm(sim, sigma_est).logpdf(obs))
return ll
def log_prior(params):
""" Calculate log prior.
"""
# Extract parameter values
a_est, b_est, sigma_est = params
# If all parameters are within allowed ranges, return a constant
# (anything will do - I've used 0 here)
if ((a_min <= a_est < a_max) and
(b_min <= b_est < b_max) and
(sigma_min <= sigma_est < sigma_max)):
return 0
# Else the parameter set is invalid (probability = 0; log prob = -inf)
else:
return -np.inf
def log_posterior(params, x, obs):
""" Calculate log posterior.
"""
# Get log prior prob
log_pri = log_prior(params)
# Evaluate log likelihood if necessary
if np.isfinite(log_pri):
log_like = log_likelihood(params, x, obs)
# Calculate log posterior
return log_pri + log_like
else:
# Log prior is -inf, so log posterior is -inf too
return -np.inf
# -
# ### 3.2. Define limits for uniform priors
#
# In the `log_prior` function above we've assumed **[improper uniform priors](https://en.wikipedia.org/wiki/Prior_probability#Improper_priors)**, just as we have in all the previous notebooks. Below we set allowable prior ranges for $a$, $b$ and $\sigma_\epsilon$.
a_min, a_max = -10, 10
b_min, b_max = -10, 10
sigma_min, sigma_max = 0, 10
# ### 3.3. Find the MAP
#
# The [MAP](https://en.wikipedia.org/wiki/Maximum_a_posteriori_estimation) is the maximum of the posterior distribution. It gives the most likely values for the model parameters ($a$, $b$ and $\sigma_\epsilon$) *given our piors and the data*. It also provides a good starting point for our MCMC analysis.
# +
from scipy import optimize
def neg_log_posterior(params, x, obs):
""" Negative of log posterior.
"""
return -log_posterior(params, x, obs)
def find_map(init_guess, x, obs):
""" Find max of posterior.
init_guess [a, b, sigma]
"""
# Run optimiser
param_est = optimize.fmin(neg_log_posterior,
init_guess,
args=(x, obs))
return param_est
# Guess some starting values for [a, b, sigma]
param_guess = [1, 1, 1]
# Run optimiser
param_est = find_map(param_guess, x, y)
# Print results
print '\n'
for idx, param in enumerate(['a', 'b', 'sigma',]):
print 'Estimated %s: %.2f.' % (param, param_est[idx])
# -
# It's reassuring to see the MAP estimates are close to the true values. However, as we've discusssed previously, these numbers are't much use without an indication of **uncertainty** i.e. how well-constrained are these values, given our priors and the data? For a simple problem like this, there are much simpler ways of estimating uncertainty using a Bayesian approach than by running an MCMC analysis (see [notebook 8](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/08_Gaussian_Approx.ipynb), for example). Nevertheless, the MCMC approach is very general and we've used it a number of times previously, so for consistency we'll apply it here as well.
#
# ### 3.4. Run the MCMC
#
# As before, we'll use [emcee](http://dan.iel.fm/emcee/current/) to draw samples from the posterior.
# +
import emcee, corner
# emcee parameters
n_dim = 3 # Number of parameters being calibrated
n_walk = 20 # Number of "walkers"/chains
n_steps = 200 # Number of steps per chain
n_burn = 100 # Length of burn-in to discard
def run_mcmc(n_dim, n_walk, n_steps, n_burn, param_opt, truths=None):
""" Sample posterior using emcee.
n_dim Number of parameters being calibrated
n_walk Number of walkers/chains (must be even)
n_steps Number of steps taken by each walker
n_burn Number of steps to discard as "burn-in"
param_opt Optimised parameter set from find_map()
truths True values (if known) for plotting
Produces plots of the chains and a 'corner plot' of the
marginal posterior distribution.
Returns an array of samples (with the burn-in discarded).
"""
# Generate starting locations for the chains by adding a small
# amount of Gaussian noise to optimised MAP
starting_guesses = [param_opt + 1e-4*np.random.randn(n_dim)
for i in range(n_walk)]
# Prepare to sample. The params are automatically passed to log_posterior
# as part of n_dim. "args" lists the other params that are also necessary
sampler = emcee.EnsembleSampler(n_walk, n_dim, log_posterior,
args=[x, y])
# Run sampler
pos, prob, state = sampler.run_mcmc(starting_guesses, n_steps)
# Print some stats. based on run properties
print '\n'
print 'Average acceptance fraction: ', np.mean(sampler.acceptance_fraction)
print 'Autocorrelation time: ', sampler.acor
# Get results
# Plot traces, including burn-in
param_labels = ['a', 'b', 'sigma']
fig, axes = plt.subplots(nrows=n_dim, ncols=1, figsize=(10, 10))
for idx, title in enumerate(param_labels):
axes[idx].plot(sampler.chain[:,:,idx].T, '-', color='k', alpha=0.3)
axes[idx].set_title(title, fontsize=20)
plt.subplots_adjust(hspace=0.5)
plt.show()
# Discard burn-in
samples = sampler.chain[:, n_burn:, :].reshape((-1, n_dim))
# Triangle plot
tri = corner.corner(samples,
labels=param_labels,
truths=truths,
quantiles=[0.025, 0.5, 0.975],
show_titles=True,
title_args={'fontsize': 24},
label_kwargs={'fontsize': 20})
return samples
samples = run_mcmc(n_dim, n_walk, n_steps, n_burn, param_est,
[a_true, b_true, sigma_true])
# -
# Blue solid lines on the "corner plot" above indicate the true values, while the vertical dotted lines on the histograms mark the 2.5%, 50% and 97.5% quantiles for the parameter estimates. In all cases, the true values lie well within the 95% **credible intervals** (a "credible interval" is the Bayesian equivalent of a frequentist "confidence interval").
#
# ### 3.6. Get the confidence intervals
#
# As with the frequentist analysis, we can also plot our median simulation and the 95% credible interval on top of the observed data. First, we'll extract some key values into a data frame that we can compare with the frequentist results.
# +
# Print estimates and confidence intervals
mcmc_df = pd.DataFrame(data=samples, columns=['a_mcmc', 'b_mcmc', 'sigma_mcmc'])
print mcmc_df.describe(percentiles=[0.025, 0.5, 0.975]).ix[['2.5%', '50%', '97.5%']].T
print '\n'
print freq_df.T
# -
# The Bayesian and frequentist results are very similar. We can also sample from our MCMC simulations to derive credible intervals for plotting.
# +
# Store output data in lists
conf = []
# Pick parameter sets at random from the converged chains
for a, b, sigma in samples[np.random.randint(len(samples), size=1000)]:
# Simulate values
sim = a*x + b + norm.rvs(loc=0, scale=sigma, size=n)
df = pd.DataFrame(data={'Sim':sim})
# Add to conf
conf.append(df)
# Concatenate results
conf = pd.concat(conf, axis=1)
# Get 2.5 and 97.5 percentiles for plotting
conf = conf.T.describe(percentiles=[0.025, 0.5, 0.975]).T[['2.5%', '50%', '97.5%']]
# Plot predicted
plt.fill_between(x, conf['2.5%'], conf['97.5%'], color='r', alpha=0.3)
plt.plot(x, conf['50%'], 'r-', label='Estimated')
plt.title('Bayesian')
# Plot true line
plt.plot(x, y, 'bo')
plt.plot(x, a_true*x+b_true, 'b--', label='True')
plt.legend(loc='best')
plt.show()
# -
# The edges of the **credible interval** are a little jagged due to our limited numerical sampling, but if we ran the chains for longer and used more samples to construct the intervals, we could get a smoother result. Nonetheless, it's pretty obvious that this interval is essentially identical to the one from the frequentist analysis.
#
# ### 3.7. Get the coverage
#
# As above, we can also calculate the coverage, which should be roughly 95%.
# +
# Add observations to df
conf['obs'] = y
# Are obs within CI?
conf['In_CI'] = ((conf['2.5%'] < conf['obs']) &
(conf['97.5%'] > conf['obs']))
# Coverage
cov = 100.*conf['In_CI'].sum()/len(conf)
print 'Coverage: %.1f%%' % cov
# -
# ## 4. GLUE
#
# The GLUE methodology is a little different. First of all, GLUE typically makes use of **informal** or **pseudo-** likelihood functions, which do not explicitly consider the error structure between the model output and the observations. Within the GLUE framework, it is permissable to use *any* scoring metric (or combination of metrics) to evaluate model performance, with the emphasis focusing less on what is *statistically rigorous* and more on what is *physically meaningful*. For example, it is very common to see GLUE analyses using the **[Nash-Sutcliffe efficiency](https://en.wikipedia.org/wiki/Nash%E2%80%93Sutcliffe_model_efficiency_coefficient)** as an indcator of model performance. GLUE also takes what is often called a "limits of acceptability" approach, requiring the user to define a threshold for their chosen metric that distinguishes between **plausible** and **implausible** model simulations.
#
# The methodology usually goes something like this:
#
# 1. Choose a metric (or metrics) to indicate model performance. **Skill scores** such as Nash-Sutcliffe are very commonly used. <br><br>
#
# 2. Set a threshold for the chosen skill score above which model simulations will be deemed to be plausible. These plausible simulations are usually termed "**behavioural**" within the GLUE framework. <br><br>
#
# 3. Define prior distributions for the model's parameters. These are usually (but not necessarily) taken to be uniform, just like the ones we used above for the Bayesian analsysis. <br><br>
#
# 4. Sample from the **pseudo-posterior**
#
# $$P_p(\theta|D) \propto P_p(D|\theta)P(\theta)$$
#
# where the likelihood term is replaced by the **pseudo-likelihood**. Just like the Bayesian approach, the sampling strategy can be any of those described in previous notebooks (e.g. Monte Carlo, MCMC etc.). However, the vast majority of GLUE analyses make use of **simple Monte Carlo sampling** i.e. draw a large random sample from the prior, then evaluate the pseudo-likelihood for each parameter set. <br><br>
#
# 5. Any parameter sets scoring below the threshold defined in step 2 are **discarded**; those scoring above the threshold are labelled "**behavioural**" and kept for further analysis. <br><br>
#
# 6. The behavioural parameter sets are **weighted** according to their skill score. The model simulations are then ranked from lowest to highest, and the normalised weights are accumulated to produce a **[cumulative distribution function (CDF)](https://en.wikipedia.org/wiki/Cumulative_distribution_function)**. <br><br>
#
# 7. The CDF is used to define a 95% **uncertainty interval** or **prediction limit** for the model output.
#
# Some key points to note are that:
#
# 1. The use of a pseudo-likelihood function means the pseudo-posterior is not a true probability distribution, so GLUE **cannot** be used to generate a **marginal posterior distribution** for each model parameter. The basic unit of consideration in GLUE is the parameter **set**. <br><br>
#
# 2. The prediction limits (or uncertainty intervals) identified by GLUE are **subjective** and have **no clear statistical meaning**. For example, they are **not** confidence bounds in any true statistical sense: the 95% confidence interval is *not* expected to include 95% of the observations.
#
# We will discuss the strengths and limitations of GLUE below, but first we'll apply the method to solve our simple linear regression problem.
#
# ### 4.1. Define the pseudo-likelihood
#
# The range of possible metrics for the pseudo-likelihood is huge. In this example we'll use the **Nash-Sutcliffe efficiency**, which is very commonly used wth GLUE. Note that other metrics may perform better (see below), but a key "selling point" of the GLUE approach is that we shouldn't have to worry too much about our choice of goodness-of-fit measure.
def nash_sutcliffe(params, x, obs):
""" Nash-Sutcliffe efficiency.
"""
# Extract parameter values
a_est, b_est = params
# Run simulation
sim = a_est*x + b_est
# NS
num = np.sum((sim - obs)**2)
denom = np.sum((obs - obs.mean())**2)
ns = 1 - (num/denom)
return [ns, sim]
# ### 4.2. Set the behavioural threshold and sample size
#
# We next need to set a **behavioural threshold** to separate plausible from implausible parameter sets. Choosing an appropriate threshold can be difficult, as it is rare for our skill score to have any direct physical relevance for our problem of interest (i.e. what is a "good" Nash-Sutcliffe score in the context of linear regression? What about for hydrology? etc.).
#
# If we set our threshold too high, we will identify very few behavioural parameter sets; set it too low, and we risk classifying some poor simulations as "behavioural" and biasing our results. In practice, many published studies start off with a stringent behavioural threshold, but are then forced to relax it in order to find enough behavioural parameter sets to continue the analysis. This is sometimes argued to be an advantage, in the sense that GLUE allows rejection of **all** available models if none of them meet the pre-defined performance criteria.
#
# For now, we'll try a threshold of $0.7$ and we'll investigate the effects of changing it later.
#
# We also need to decide how many samples to draw from our prior. For this simple 2D example, Monte Carlo sampling should actually work OK, so we'll choose the same total number of samples as we used above in our MCMC analysis. Note, however, that for problems in a larger parameter space, we might need to draw a *very* large number of samples indeed using Monte Carlo methods to get a reasonable representation of the posterior.
ns_min = 0.7
n_samp = 4000
# ### 4.3. Sample from the prior
#
# One of the main advantages of Monte Carlo GLUE is that it is usually very easy to code (and to parallelise). Here we're drawing 4000 independent samples from our priors.
a_s = np.random.uniform(low=a_min, high=a_max, size=n_samp)
b_s = np.random.uniform(low=b_min, high=b_max, size=n_samp)
# ### 4.4. Run GLUE
#
# For each of the parameter sets drawn above, we run the model and calculate the Nash-Sutcliffe efficiency. If it's above the behavioural threshold we'll store that parameter set and the associated model output, otherwise we'll discard both.
# +
def run_glue(a_s, b_s, n_samp, ns_min):
""" Run GLUE analysis.
Uses nash_sutcliffe() to estimate performance and returns
dataframes containing all "behavioural" parameter sets and
associated model output.
"""
# Store output
out_params = []
out_sims = []
# Loop over param sets
for idx in range(n_samp):
params = [a_s[idx], b_s[idx]]
# Calculate Nash-Sutcliffe
ns, sim = nash_sutcliffe(params, x, y)
# Store if "behavioural"
if ns >= ns_min:
params.append(ns)
out_params.append(params)
out_sims.append(sim)
# Build df
params_df = pd.DataFrame(data=out_params,
columns=['a', 'b', 'ns'])
assert len(params_df) > 0, 'No behavioural parameter sets found.'
# Number of behavioural sets
print 'Found %s behavioural sets out of %s runs.' % (len(params_df), n_samp)
# DF of behavioural simulations
sims_df = pd.DataFrame(data=out_sims)
return params_df, sims_df
params_df, sims_df = run_glue(a_s, b_s, n_samp, ns_min)
# -
# Note that with a two dimensional parameter space and Nash-Sutcliffe cut-off of $0.7$, only about $\frac{1}{20}th$ of the model runs are classified as "behavioural". This fraction would decrease *very rapidly* if the parameter space became larger.
#
# ### 4.5. Estimate confidence intervals
#
# Using just the behavioural parameter sets, we **rank** the model output and calculate **weighted quantiles** to produce the desired CDF.
# +
def weighted_quantiles(values, quantiles, sample_weight=None):
""" Modified from
http://stackoverflow.com/questions/21844024/weighted-percentile-using-numpy
NOTE: quantiles should be in [0, 1]
values array with data
quantiles array with desired quantiles
sample_weight array of weights (the same length as `values`)
Returns array with computed quantiles.
"""
# Convert to arrays
values = np.array(values)
quantiles = np.array(quantiles)
# Assign equal weights if necessary
if sample_weight is None:
sample_weight = np.ones(len(values))
# Otherwise use specified weights
sample_weight = np.array(sample_weight)
# Check quantiles specified OK
assert np.all(quantiles >= 0) and np.all(quantiles <= 1), 'quantiles should be in [0, 1]'
# Sort
sorter = np.argsort(values)
values = values[sorter]
sample_weight = sample_weight[sorter]
# Compute weighted quantiles
weighted_quantiles = np.cumsum(sample_weight) - 0.5 * sample_weight
weighted_quantiles /= np.sum(sample_weight)
return np.interp(quantiles, weighted_quantiles, values)
def plot_glue(params_df, sims_df):
""" Plot median simulation and confidence intervals for GLUE.
"""
# Get weighted quantiles for each point in x from behavioural simulations
weights = params_df['ns']
quants = [0.025, 0.5, 0.975]
# List to store output
out = []
# Loop over points in x
for col in sims_df.columns:
values = sims_df[col]
out.append(weighted_quantiles(values, quants, sample_weight=weights))
# Build df
glue_df = pd.DataFrame(data=out, columns=['2.5%', '50%', '97.5%'])
# Plot predicted
plt.fill_between(x, glue_df['2.5%'], glue_df['97.5%'], color='r', alpha=0.3)
plt.plot(x, glue_df['50%'], 'r-', label='Estimated')
plt.title('GLUE')
# Plot true line
plt.plot(x, y, 'bo')
plt.plot(x, a_true*x+b_true, 'b--', label='True')
plt.legend(loc='best')
plt.show()
return glue_df
glue_df = plot_glue(params_df, sims_df)
# -
# These results are clearly a bit different to the output from the Bayesian and frequentist analyses presented above. The predicted line is not as good a fit to the true data and the confidence interval is wider at the extremes than it is towards the middle. Nevertheless, this result seems superficially reasonable in the sense that it does not obviously contradict the output obtained from the other methods. Overall it is likely that, in a decision-making context, all these approaches would lead to broadly the same actions being taken.
#
# ### 4.6. Coverage
#
# For consistency, we'll also calculate the coverage for GLUE, but note that *GLUE confidence intervals are not expected to bracket the stated proportion of the observations* (see above).
# +
def glue_coverage(glue_df):
""" Prints coverage from GLUE analysis.
"""
# Add observations to df
glue_df['obs'] = y
# Are obs within CI?
glue_df['In_CI'] = ((glue_df['2.5%'] < glue_df['obs']) &
(glue_df['97.5%'] > glue_df['obs']))
# Coverage
cov = 100.*glue_df['In_CI'].sum()/len(glue_df)
print 'Coverage: %.1f%%' % cov
glue_coverage(glue_df)
# -
# Based on the results so far, you might be thinking there's not much to choose between any of these approaches, but let's see what happens to the GLUE output if the behavioural threshold is adjusted.
#
# ### 4.7. Changing the behavioural threshold
#
# The Nash-Sutlcliffe score can take any value from $-\infty$ to $1$, with $0$ implying the model output is no better than taking the **mean** of the observations. What happens if we **relax the behavioural threshold** by setting it to $0$?
# +
ns_min = 0
params_df, sims_df = run_glue(a_s, b_s, n_samp, ns_min)
glue_df = plot_glue(params_df, sims_df)
glue_coverage(glue_df)
# -
# And what if we **make the behavioural threshold more stringent**, by setting it to $0.9$?
# +
ns_min = 0.9
params_df, sims_df = run_glue(a_s, b_s, n_samp, ns_min)
glue_df = plot_glue(params_df, sims_df)
glue_coverage(glue_df)
# -
# ## 5. Interpretation
#
# How should we interpret this very simple analysis?
#
# Just to reiterate from above, the confidence intervals derived from GLUE have no formal statistical meaning, so we should not be surprised to see the **GLUE 95% uncertainty interval** bracketing anything between about **70% and 100% of the data**. Nevertheless, you may be surprisied to see just how much the GLUE confidence bounds depend on the (arbitrary) choice of behavioural threshold. In essence, it seems we can set the uncertainty to be **almost anything we like**, just by tuning the cut-off. How, then, are we to interpret the uncertainty limits presented by GLUE from the point of view of **decision making**?
#
# As a related question, it seems reasonable to ask, *"What's the point of a 95% uncertainty interval that has no intention of bracketing 95% of the data?"* Frequentist "confidence" and Bayesian "credible" intervals have a very widely accepted meaning in statistics, and it is rather confusing that GLUE presents something that looks so superficially similar, but which is actually nothing of the sort.
#
# A further difficulty with GLUE concerns the method's ability to assimilate more data. If the above examples are repeated using a larger observed dataset, the confidence intervals for the parameter estimates from the Bayesian and frequentist analyses will become **narrower**. This is consistent with the (fairly fundamental) idea that more data provides more information, and therefore allows us to make more refined estimates. This is not necessarily the case with GLUE: as described in some detail by [Stedinger *et al.* (2008)](http://onlinelibrary.wiley.com/doi/10.1029/2008WR006822/abstract), predictions from GLUE do not necessarily become more refined as the amount of data increases.
#
# ## 6. Methodological inconsistency?
#
# In addition to the issues mentioned above, there are a number of features of the GLUE methodology that are difficult to reconcile.
#
# ### 6.1. Limits of acceptability *and* likelihood weighting?
#
# GLUE is often described as a "limits of acceptability" approach, because it defines a threshold below which parameers sets are discarded as being implausible. This is in contrast to more formal methods that use the likelihood function to weight *all* parameter sets, in such a way that poor sets are assigned lower weights then good ones. One strange thing about GLUE is that it implements a limits of acceptability threshold **as well as** weighting the behavioural sets according to the pseudo-likelihood.
#
# If it is possibe to assign a physically meaningful limit of acceptability, then surely all the acceptable (i.e. behavioural) parameter sets should be assigned equal weight? (**Generalised Sensitivity Analysis**, a closely related technique to GLUE, does exactly this - see e.g. [Young, 1999](http://www.sciencedirect.com/science/article/pii/S0010465598001684)). On the other hand, if "acceptability" is better considered as a **continuous spectrum**, it would seem more sensible *not* to set a well-defined threshold and to weight each parameter set according to some continuous measure of performance, exactly as done in a formal Bayesian setting. Doing **both**, as is the case with GLUE, implies that the informal likelihood function cannot be trusted to distinguish between good and bad parameter sets.
#
# [Stedinger *et al.* (2008)](http://onlinelibrary.wiley.com/doi/10.1029/2008WR006822/abstract) have demonstrated that, when used with a formal statistical likelihood, GLUE can produce results that are "correct" in a statistical sense. However, they also point out that when this is done the behavioural threshold becomes unnecessary, because the formal likelihood is capable of effectively separating good from bad simulations. Of course, GLUE with a formal likelihood and no limit of acceptability is **no longer GLUE** - it's simply a Monte Carlo-based formal Bayesian approach, similar to **[importance sampling](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/03_Monte_Carlo.ipynb#2.2.2.-Importance-sampling)**. Such formal Bayesian approaches pre-date GLUE by several decades and their limitations in high dimensional spaces are very well documented.
#
# ### 6.2. Fewer assumptions?
#
# It is often claimed that GLUE makes "**fewer assumptions**" than more formal statistical approaches, because it is not necessary to explicitly specify and test any particular error structure. However, although GLUE may not require an explicit consderation of the errors, just because an issue is ignored does not mean it will go away. Commonly used pesudo-likelihoods for GLUE, such as the inverse error variance or the Nash-Sutcliffe efficiency, all make **implicit** assumptions about the nature of the error structure. In addition, many of these metrics have unintended or undesirable characteristics, such as not properly accounting for number of samples in the observational dataset (see section 5, above).
#
# Choosing an arbitrary metric without understanding the assumptions inherent in that choice is not the same as making fewer assumptions - it's more a case of **acknowledging fewer assumptions**.
#
# ## 7. Computational inefficiency
#
# A less controversial but nonetheless significant drawback of the standard GLUE implementation is its **computational inefficiency**. Unlike concerns regarding statistical validity, there is really no debate to be had about sampling efficiency: standard Monte Carlo methods are expected to be *extremely inefficient* in high dimensional parameter spaces, and the standard Monte Carlo version of GLUE is no different (see [notebook 3](http://nbviewer.ipython.org/github/JamesSample/enviro_mod_notes/blob/master/notebooks/03_Monte_Carlo.ipynb) for a discussion of why this is the case).
#
# It is of course possibe to use GLUE with *any* sampling strategy. [Blasone *et al.* (2007)](http://www.sciencedirect.com/science/article/pii/S0309170807001856) implemented GLUE using an MCMC approach, which resulted in vastly improved sampling efficiency. This method has not become popular, however - perhaps because the most appealing aspects of GLUE are its conceptual simplicity and ease of coding compared to e.g. MCMC.
#
# ## 8. Is this comparison fair?
#
# A common response from proponents of GLUE to comparisons of this kind is to say:
#
# *"In the Bayesian example you used exactly the right likelihood function, so of course you got better results. If you'd used the correct likelihood function with GLUE, GLUE would have worked better too. In reality, though, you never know what the true likelihood is."*
#
# This is a fair comment in some respects: GLUE *can* produce more reasonable answers when used with a formal likelihood function. [Beven *et al.* (2007)](http://www.sciencedirect.com/science/article/pii/S0022169407001230) have even argued that, *"the formal Bayesian identification of models is a special case of GLUE"*. This statement is difficult to accept, at least in part because two of the key characteristics of GLUE are:
#
# 1. The use of an **informal likelihood** and <br><br>
#
# 2. The setting of a **behavioural** threshold.
#
# As noted above, [Stedinger *et al.* (2008)](http://onlinelibrary.wiley.com/doi/10.1029/2008WR006822/abstract) have demonstrated that GLUE *can* perform correctly using a formal likelihood, but they also showed that this makes the behavioural threshold redundant, in which case the method is just a very outdated formal Bayesian approach - one that can **no longer reasonably be called "GLUE"**.
#
# On the whole, the analysis presented here is consistent with the approaches recommended for the three different methods: Bayesian and frequentist analyses require us to think carefully about formulating an appropriate error structure and to test to see whether those assumptions have been met (refining them if necessary); GLUE, on the other hand, does not require any explicit consideration of the distribution of the residuals.
#
# Of course, in the examples here we knew the exact form of the true likelihood and we used this to advantage in the Bayesian and frequentist analyses. An advocate of GLUE could argue that we never have this information in reality - which is true - but not knowing something doesn't mean we can just ignore the issue. What's more, it is perfectly possible to *test* whether the error assumptions in a formal analysis have been met, so although we never know the true likelihood for sure, we can at least say whether the data are consistent with what we've assumed. There are plenty of papers in the literature where the authors *do* achieve reasonable results using a formal statistical approach, so adopting an *ad hoc* methodology such as GLUE seems unjustified. What's more, using an informal likelihood function does not remove the problem of specifying an error structure - it simply hides the assumptions being made implicitly by whatever goodness-of-fit metric is selected.
#
# Statistics would be a great deal easier if we could perform robust inference without having to think about things like likelihoods and error structures. GLUE is appealing partly because it **promises a great deal for very little effort**, which is attractive to environmental scientists bewildered by statistical notation and seemingly impenetrable mathematics. Unfortunately, I'm increasingly of the opinion that the claims made for GLUE are **too good to be true**, and I'm inclined to agree with [Stedinger *et al.* (2008)](http://onlinelibrary.wiley.com/doi/10.1029/2008WR006822/abstract) that,
#
# *"If an arbitrary likelihood is adopted that does not reasonably reflect the sampling distribution of the model errors, then GLUE generates arbitrary results without statistical validity that should not be used in scientific work."*
#
# Don't take my word for it, though - please read the papers cited here (and elsewhere) and **make up your own mind**.
| notebooks/07_GLUE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
raw_data_path=os.path.join(os.path.pardir,'data','raw')
train_file_path=os.path.join(raw_data_path,'train.csv')
test_file_path=os.path.join(raw_data_path,'test.csv')
# Creation of dataaframes
train_df=pd.read_csv(train_file_path,index_col='PassengerId')
test_df=pd.read_csv(test_file_path,index_col='PassengerId')
type(test_df)
train_df.info()
test_df.info()
#Adding a column Survived in test_df and providing a default value
test_df['Survived']=-888
test_df.info()
df=pd.concat((train_df,test_df),axis=0)
df.info()
df.head()
df.tail(10)
df.Name
df['Name']
#Selecting multiple columns using a list of column name strings
df[['Name','Age']]
#lable based indexing
df.loc[5:10,]
#Row x Column
#Column selection
df.loc[5:10,'Age':'Pclass']
#Discrete Column selection
df.loc[5:10,['Survived','Fare','Embarked']]
#Position Based Indexing
df.iloc[5:10,0:3]
#Position Based Indexing
df.iloc[5:10,0:2]
#Position Based Indexing
df.iloc[5:10,0:2]
#Filtering the data
male_passengers= df.loc[df.Sex=='male',:]
print('Number of male passengers: {0}'.format(len(male_passengers)))
#Filtering the data
FirstClass_male_passengers= df.loc[((df.Sex=='male')& (df.Pclass==1)),:]
print('Number of male passengers: {0}'.format(len(FirstClass_male_passengers)))
# # Summary Statistics
df.describe()
print('Mean Fare: {0}'.format(df.Fare.mean()))
print('Median Fare: {0}'.format(df.Fare.median()))
print('Min fare : {0}'.format(df.Fare.min())) # minimum
print('Max fare : {0}'.format(df.Fare.max())) # maximum
print('Fare range : {0}'.format(df.Fare.max() - df.Fare.min())) # range
print('25 percentile : {0}'.format(df.Fare.quantile(.25))) # 25 percentile
print('50 percentile : {0}'.format(df.Fare.quantile(.5))) # 50 percentile
print('75 percentile : {0}'.format(df.Fare.quantile(.75))) # 75 percentile
print('Variance fare : {0}'.format(df.Fare.var())) # variance
print('Standard deviation fare : {0}'.format(df.Fare.std())) # standard deviation
# %matplotlib inline
df.Fare.plot(kind='box')
df.describe(include='all')
df.Sex.value_counts()
df.Sex.value_counts(normalize=True)
df[df.Survived!=-888].Survived.value_counts()
df.Pclass.value_counts()
df.Pclass.value_counts().plot(kind='bar')
df.Pclass.value_counts().plot(kind='bar',rot=0, title='Class wise passenger count', color='c')
# # Distributions
#Use histogram
df.Age.plot(kind='hist',title='Histogram for Age', color='c')
df.Age.plot(kind='hist',title='Histogram fo Age', color='c', bins=20);
#use kde for density plot
df.Age.plot(kind='kde',title='Density Plot for age',color='green')
# histogram for fare
df.Fare.plot(kind='hist', title='histogram for Fare', color='c', bins=20);
print('skewness for age : {0:.2f}'.format(df.Age.skew()))
print('skewness for fare : {0:.2f}'.format(df.Fare.skew()))
#Scatter Plot
df.plot.scatter(x='Age',y='Fare',title='Scatter plot: Age vs Fare')
#Set alpha for setting transparancy
df.plot.scatter(x='Age', y='Fare', color='c', title='scatter plot : Age vs Fare', alpha=0.1);
df.plot.scatter(x='Pclass', y='Fare', color='r', title='Scatter plot : Passenger class vs Fare', alpha=0.15);
# # Grouping and Aggregations
df.groupby('Sex').Age.median()
df.groupby('Pclass').Fare.median()
df.groupby('Pclass').Age.median()
df.groupby(['Pclass'])['Fare','Age'].median()
##Note There is no dot after group by method
#Aggregations
df.groupby(['Pclass']).agg({'Fare':'mean', 'Age':'median'})
#Complicated Aggregations
aggregations={
'Fare':{#Work on Fare Column
'mean_Fare':'mean',
'median_Fare':'median',
'max_Fare':max,
'min_Fare':np.min
},
'Age':{ # work on the "Age" column
'median_Age': 'median', # Find the max, call the result "max_date"
'min_Age': min,
'max_Age': max,
'range_Age': lambda x: max(x) - min(x) # Calculate the age range per group
}
}
df.groupby(['Pclass']).agg(aggregations)
df.groupby(['Pclass','Embarked']).Fare.median()
# # Crosstabs
#Using Crosstabs
pd.crosstab(df.Sex,df.Pclass)
pd.crosstab(df.Sex,df.Pclass).plot(kind='bar');
df.pivot_table(index='Sex',columns='Pclass',values='Age',aggfunc='mean')
df.groupby(['Sex','Pclass']).Age.mean()
df.groupby(['Sex','Pclass']).Age.mean().unstack()
# ## Data Munging : Working with missing values
df.info()
# # Feature: Embarked
#extract rows with embarked as null
df[df.Embarked.isnull()]
df.Embarked.value_counts()
pd.crosstab(df[df.Survived!=-888].Survived,df[df.Survived!=-888].Embarked)
# +
# impute the missing values with 'S'
# df.loc[df.Embarked.isnull(), 'Embarked'] = 'S'
# df.Embarked.fillna('S', inplace=True)
# -
df.groupby(['Pclass','Embarked']).Fare.median()
#to replace the missing value with 'C'
df.Embarked.fillna('C',inplace=True)
df[df.Embarked.isnull()]
df.info()
# # Feature : Fare
df[df.Fare.isnull()]
median_fare = df.loc[(df.Pclass == 3) & (df.Embarked == 'S'),'Fare'].median()
print(median_fare)
df.Fare.fillna(median_fare,inplace=True)
df.info()
# # Feature:Age
#set number of rows to be displayed
pd.options.display.max_rows=15
df[df.Age.isnull()]
# %matplotlib inline
df.Age.plot(kind='hist', bins=20, color='c');
df.Age.mean()
df.groupby('Sex').Age.median()
#using boxplot
df[df.Age.notnull()].boxplot('Age','Sex')
# +
# replace :
# age_sex_median = df.groupby('Sex').Age.transform('median')
# df.Age.fillna(age_sex_median, inplace=True)
# -
df[df.Age.notnull()].boxplot('Age','Pclass');
# +
# replace :
# pclass_age_median = df.groupby('Pclass').Age.transform('median')
# df.Age.fillna(pclass_age_median , inplace=True)
# -
df.Name
# Function to extract the title from the name
def GetTitle(name):
first_name_with_title = name.split(',')[1]
title = first_name_with_title.split('.')[0]
title = title.strip().lower()
return title
df.Name.map(lambda x: GetTitle(x))
df.Name.map(lambda x: GetTitle(x)).unique()
# +
def GetTitle(name):
title_group = {'mr' : 'Mr',
'mrs' : 'Mrs',
'miss' : 'Miss',
'master' : 'Master',
'don' : 'Sir',
'rev' : 'Sir',
'dr' : 'Officer',
'mme' : 'Mrs',
'ms' : 'Mrs',
'major' : 'Officer',
'lady' : 'Lady',
'sir' : 'Sir',
'mlle' : 'Miss',
'col' : 'Officer',
'capt' : 'Officer',
'the countess' : 'Lady',
'jonkheer' : 'Sir',
'dona' : 'Lady'
}
first_name_with_title = name.split(',')[1]
title = first_name_with_title.split('.')[0]
title = title.strip().lower()
return title_group[title]
# -
df['Title']=df.Name.map(lambda x: GetTitle(x))
df.head()
df[df.Age.notnull()].boxplot('Age','Title');
title_age_median = df.groupby('Title').Age.transform('median')
print(title_age_median)
df.Age.fillna(title_age_median, inplace=True)
df.info()
# # Working with outliers
# +
#Age
# -
df.Age.plot(kind='hist',bins=20,color='r')
df.loc[df.Age>70]
#Fare
df.Fare.plot(kind='hist', title='histogram for Fare', bins=20, color='c');
df.Fare.plot(kind='box');
df.loc[df.Fare == df.Fare.max()]
# +
#Logs not clear so skipped
# -
# # Feature Engineering
# ### Feature: Age State
df['AgeState']=np.where(df['Age']>=18,'Adult','Child')
df['AgeState'].value_counts()
pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].AgeState)
df['FamilySize'] = df.Parch + df.SibSp + 1 # 1 for self
df['FamilySize'].plot(kind='hist', color='c');
df.loc[df.FamilySize == df.FamilySize.max(),['Name','Survived','FamilySize','Ticket']]
pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].FamilySize)
# ### Feature : IsMother
df['IsMother'] = np.where(((df.Sex == 'female') & (df.Parch > 0) & (df.Age > 18) & (df.Title != 'Miss')), 1, 0)
pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].IsMother)
# #### Deck
df.Cabin
df.Cabin.unique()
df.loc[df.Cabin == 'T']
df.loc[df.Cabin == 'T', 'Cabin'] = np.NaN
df.Cabin.unique()
# extract first character of Cabin string to the deck
def get_deck(cabin):
return np.where(pd.notnull(cabin),str(cabin)[0].upper(),'Z')
df['Deck'] = df['Cabin'].map(lambda x : get_deck(x))
df.Deck.value_counts()
pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].Deck)
df.info()
# ### Categorical Feature Encoding
df['IsMale'] = np.where(df.Sex == 'male', 1, 0)
df.info()
df = pd.get_dummies(df,columns=['Deck', 'Pclass','Title', 'Fare_Bin', 'Embarked','AgeState'])
| notebooks/Exploring-Processing-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Golf Ball (One Way Anova)
# # Importing the Libraries
import numpy as np
import pandas as pd
import seaborn as sns
from statsmodels.formula.api import ols # For n-way ANOVA
from statsmodels.stats.anova import _get_covariance,anova_lm # For n-way ANOVA
# %matplotlib inline
# # Loading the dataset CSV file
df=pd.read_csv('C:\\Users\\jayveer\\Downloads\\Golf ball\\Golfball.csv')
# # Checking the top 5 records
df.head()
# # Checking the shape and information of the dataframe
df.shape
df.info()
# # Checking the summary of the dataframe
df.describe(include='all')
# # Checking distinct values of Design
df.Design.value_counts()
# # One Way Anova
formula = 'Distance ~ C(Design)'
model = ols(formula, df).fit()
aov_table = anova_lm(model)
print(aov_table)
# Since the p value is less than the significance level, we can reject the null hupothesis and states that there is a difference in the mean distances travelled by the golf balls with different designs
# # Drawing a Point Plot
sns.pointplot(x='Design', y='Distance', data=df, ci=None)
| M3 Advance Statistics/W1 ANOVA/GolfBall_OneWayAnova_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
random.seed(1)
restaurant_list = list(sorted(['barrel_crow', 'woodmont', 'gringos']))
random.choice(restaurant_list)
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table style="font-size: 1em; padding: 0; margin: 0;">
#
# <tr style="vertical-align: top; padding: 0; margin: 0;background-color: #ffffff">
# <td style="vertical-align: top; padding: 0; margin: 0; padding-right: 15px;">
# <p style="background: #182AEB; color:#ffffff; text-align:justify; padding: 10px 25px;">
# <strong style="font-size: 1.0em;"><span style="font-size: 1.2em;"><span style="color: #ffffff;">Deep Learning </span> for Satellite Image Classification</span> (Manning Publications)<br/>by <em><NAME></em></strong><br/><br/>
# <strong>> Chapter 1: Getting Started </strong><br/>
# </p>
#
# <p style="border: 1px solid #ff5733; border-left: 15px solid #ff5733; padding: 10px; text-align:justify;">
# <strong style="color: #ff5733">What you will learn in this Part.</strong>
# <br/>In this first part we will refresh our jupyter, keras, Tensorflow 2, and git skills. These are a necessary part of your workflow and a standard collection of tools used in both industry and academia. Optionally, we will explore how to use the Sentinel API to download, display and save Sentinel-2 imagery.
#
# There may be places in this notebook where the code gets a difficult to understand. I encourage you not to get too bogged down in the details just yet; you will see many of these same functions again and each time I may explain a different aspect of them, and your understanding will improve as you progress through the project. Many of the details of the code are unimportant, at least initially, but are necessary to create functional example workflows that are relevant to later Parts
# </p>
# <table style="font-size: 1em; padding: 0; margin: 0;">
#
# <h1 style="width: 100%; text-align: left; padding: 0px 25px;"><small style="color: #182AEB;">
# </small><br/>Introduction to<br/> Jupyter notebooks</h1>
# <br/>
# <p style="border-left: 15px solid #182AEB; text-align:justify; padding: 0 10px;">
# <a href="https://jupyter.org/">Jupyter</a> notebooks are a way to share executable code that can be run through a web browser.
# </p>
# <p style="border-left: 15px solid #6019D6; padding: 0 10px; text-align:justify;">
# <strong style="color: #6019D6;">Tip.</strong>
# A notebook kernel is a computational engine that executes the code contained in a notebook. The ipython kernel executes python code. Kernels for many other languages also exist.
# </p>
# </tr>
# </table>
# You can use question mark in order to get help. To execute cell you have to press Shift+Enter
# +
## uncomment to see the 'full' help menu, which is a lot of output
# #?
# -
# * To execute the code in a particular cell, click on the cell and hit shift-enter.
# * Before you execute the code in an arbitrary cell it is good to run all the code once so that all imports and variables are initialised.
print('I love Python')
# Making plots:
# %matplotlib inline
import matplotlib.pyplot as plt
x = [1,2,3,4,5]
plt.plot(x);
# Question mark after a function will open pager with documentation. Double question mark will show you source code of the function.
# +
## uncomment to see the 'full' help menu, which is a lot of output
# #plt.plot??
# -
# #### Formatting text
#
# Double click on this cell to reveal the unformatted text which can then be re-executed, either from the buttons provided or by holding down shift while you press enter
#
# As you can see below, formatting text characters apply to markdown cells
#
# Emphasis, aka italics, with *asterisks* or _underscores_.
#
# Strong emphasis, aka bold, with **asterisks** or __underscores__.
#
# Combined emphasis with **asterisks and _underscores_**.
#
# Strikethrough uses two tildes. ~~Scratch this.~~
# [I'm an inline-style link](https://www.google.com)
# We can write equations inline: $e = mc^2$, $e^{i\pi}+1=0$
#
# or in display format:
#
# $$ e^x = \sum_{i=0}^{\infty}\frac{1}{i!}x^i$$
from IPython.display import Image
Image(url='http://python.org/images/python-logo.gif')
from IPython.display import YouTubeVideo
YouTubeVideo('F4rFuIb1Ie4')
# Find out how long it take to run a command with %timeit magic function:
# %timeit -n 12 list(range(1000000))
# Print all interactive variables:
# %whos
# In order to get information about all magic functions type:
# +
## uncomment if you are interested in all the magic commands,
## but it isn't necessary to read it to execute the rest of the notebook
# #%magic
# -
# You can navigate and list file contents in python using the ```os``` module
import os
print(os.getcwd())
# There are a couple of different ways of running a python script and passing it variables. To demonstrate this, let's first make a simple script that accepts 2 variables and prints them to screen
var1 = 50
var2 = 'jupyter_is_cool'
# ```writefile``` is a so-called [magic command](https://ipython.readthedocs.io/en/stable/interactive/magics.html)
# %%writefile test_script.py
import sys
script, var1, var2 = sys.argv
print("Script name:", script)
print("First variable:", var1)
print("Second variable:", var2)
# You can run the script using the ```run``` jupyter magic command. Variables are passed to the script using a $ symbol
# %run ./test_script.py $var1 $var2
# ... or using the bang operator. This is the convention we'll adopt in this clinic:
# ! python test_script.py $var1 $var2
# #### File management and resources on linux
#
# (these commands below only work in the cloud environment, unless you are in an environment supporting bash. Uncomment to run)
#
# How much memory do you have?
# +
# #!cat /proc/meminfo | head
# -
# CPUs?
# +
# #!cat /proc/cpuinfo | grep processor
# -
# Storage space?
# +
# #! df -h | grep "/"
# -
# <table style="font-size: 1em; padding: 0; margin: 0;">
#
# <h1 style="width: 100%; text-align: left; padding: 0px 25px;"><small style="color: #182AEB;">
# </small><br/>Getting started with<br/> Tensorflow 2 and Keras</h1>
# <br/>
# <p style="border-left: 15px solid #182AEB; text-align:justify; padding: 0 10px;">
# <strong style="color: #182AEB;">Intro.</strong>
# TensorFlow is a way of representing computation without actually performing it until asked. In this sense, it is a form of lazy computing, and it allows for some great improvements to the running of code:
# <ul>
# <li>Faster computation of complex variables</li>
# <li>Distributed computation across multiple systems, including GPUs.</li>
# <li>Reduced redundency in some computations</li>
# </ul>
# </p>
# <p style="border-left: 15px solid #6019D6; padding: 0 10px; text-align:justify;">
# <strong style="color: #6019D6;">Tip.</strong>
# TensorFlow allows you to create dataflow graphs. These are structures that describe how data moves through a graph, or a series of processing nodes. Each node in the graph represents a mathematical operation, and each connection or edge between nodes is a multidimensional data array, or tensor. Keras is a set of layers that describes how to build neural networks using a clear standard. The Sequential API defines a Neural Network as a stack of layers. It very easy to define a model and to add new layers</p>
#
# <p style="border-left: 15px solid #4E9317; padding: 0 10px; text-align:justify;">
# <strong style="color: #4E9317;">More Resources.</strong>
# <ul>
# <li><a href="https://www.tensorflow.org/tutorials">Getting Started with Tensorflow</a></li>
# <li><a href="https://github.com/sarangzambare/segmentation">Segmentation of cells using CNNs and TF-2</a></li>
# <li><a href="https://github.com/tensorflow/models/tree/master/research/deeplab">Segmentation Segmentation using DeepLab</a></li>
# </ul></p>
# </tr>
# </table>
# Next we're going to import the ```tensorflow``` module and print the version number. At the time of writing, the version number is ```2.0.0```. Be aware that it is possible this code could break with other (later/earlier) versions of tensorflow
import tensorflow as tf
tf.__version__
# You can test to see if a GPU is available like so:
tf.test.is_gpu_available()
# #### Keras
#
# A good way to get oriented with keras is to read the [summary documentation](https://keras.io/), but note that in this project you are recommended to use the version of keras within tensorflow, so the [tf.keras documentation](https://www.tensorflow.org/guide/keras) is also very relevant. Also, [this blog post](https://towardsdatascience.com/tensorflow-is-in-a-relationship-with-keras-introducing-tf-2-0-dcf1228f73ae) is worth a read, as is [this other blog post](https://www.pyimagesearch.com/2019/10/21/keras-vs-tf-keras-whats-the-difference-in-tensorflow-2-0/)
#
# #### Two ways to make a model
#
# We're going to familiarize ourselves with tensorflow and keras using a relevant example, involving image classification. Instead of our project task involving semantic segmentation (classification of every pixel), computer scientists tend to refer to classification of whole images as 'image classification'.
#
# ```keras``` is accessed as ```tf.keras``` and its layers can be imported separately
# Tensorflow keras layers
from tensorflow.python.keras.layers import Dense, Flatten, Dropout, BatchNormalization
# In the above, we have imported 2 different types of layers (`Dense`, `Dropout`), and `Flatten` that isn't technically a layer but is classified as one because it transforms a layer's dimensions (specifically, it turns a 2D matrix into a 1D vector). We will encounter and discuss further the various layers in more depth in subsequent Parts, but `Dense` is a densely connected layer where each neuron of the previous layer connects to each neuron in the previous layer. `Dropout` is one of several strategies for preventing model overfitting. It randomly drops a certain proportion of neurons, to prevent the model 'memorizing' the data and promote model generalization
# This a simple neural network model with one hidden layer that receives a 28 x 28 pixel image, and outputs one of 10 values. It has one hidden layer with 128 neurons and a rectified linear unit (```relu```) activation function. Between the hidden layer and output (classifying) layer there is a dropout layer where 20 % of neurons are randomly discarded. Softmax is an activation function. See [here](https://towardsdatascience.com/softmax-function-simplified-714068bf8156) for a good explanation. See [here](https://keras.io/activations/) for alternatives. It might be a good idea to play with different activation functions to explore their utility and effects
model = tf.keras.models.Sequential()
model.add(Flatten(input_shape=(28,28)))
model.add(Dense(units=128, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(units=10, activation='softmax'))
# Above we introduce the `Sequential` model. You should take some time to explore is numerous useful properties, such as
#
# 1. `.summary()`, which shows the model architecture
#
# 2. `.layers`, which lists the layers of the model and their shapes
#
# 3. `.save()` saves the model in its current state of training, so you can keep a record of the state of the model at various points through training. Models can be recovered from a file using `tensorflow.keras.models.load_model()`
#
# 4. `.save_weights()` saves only the weights of the model to a file
#
# 5. `.inputs` and `.outputs` provide access to model inputs and outputs
model.summary()
# That's the same as
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
# You can verify by comparing the two model summaries
model.summary()
# #### Working with imagery
# We'll use our model to classify imagery. Load the [MNIST dataset](http://yann.lecun.com/exdb/mnist/), split into test and training sets, and convert the samples from integers to floating-point numbers.
#
# The imagery data contained in the variables `x_test` and `x_train` are divided by 255.0 to normalize their values, such that all pixels have values that vary between 0 and 1, instead of 0 and 255 which is the the range of values for a standard 8-bit image
# +
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# +
import numpy as np
# check some statistics
IMAGE_HEIGHT, IMAGE_WIDTH = x_train.shape[1:3]
print("# of train examples:", x_train.shape[0])
print("# of test examples:", x_test.shape[0])
print("Image shape:", x_train[0].shape)
print("Pixel value interval:", np.min(x_train), np.max(x_train))
print("Train shape:",x_train.shape)
print("Test shape:",x_test.shape)
print("Image height:",IMAGE_HEIGHT)
print("Image width:",IMAGE_WIDTH)
# -
# Take a look at some examples. Please remember that code such as this is provided as a succinct example workflow, but it is possible to develop similar 'solutions' on your own
import matplotlib.pyplot as plt
# %matplotlib inline
# visualize some of the examples
fig, axs = plt.subplots(nrows=3, ncols=6, constrained_layout=False, figsize=(12,4))
for i, ax in enumerate(axs.flat):
ax.imshow(x_train[i], cmap="gray")
# Tensorboard is a tool for visualizing model training, such as how the loss function decreased, or accuracy increased, as a function of training epoch. Its use is often an essential part of optimizing model training
from tensorflow.python.keras.callbacks import TensorBoard
# +
# Load the TensorBoard notebook extension
# %load_ext tensorboard
# %reload_ext tensorboard
# create the tensorboard callback
tensorboard = TensorBoard(log_dir='logs', histogram_freq=1)
# -
# Calling `.compile()` is a required step before model training. The following must be specified:
#
# 1. `optimizer` that performs the gradient descent (numerical solver)
#
# 2. `loss` that is the metric to be optimized. Below we choose `sparse_categorical_crossentropy` because we have a categorical problem (hence `categorical_crossentropy`) that is not [one-hot-encoded](https://machinelearningmastery.com/why-one-hot-encode-data-in-machine-learning/) (hence `sparse`)
#
# 3. `metrics`, which are additional metrics to evaluate during training but, unlike loss, are not used in the training process
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train and evaluate the model:
model.fit(x_train, y_train, epochs=5, callbacks=[tensorboard])
# Launch tensorboard for visualizing model training. You should see graphs `epoch_accuracy` and `epoch_loss` that show accuracy and loss function value, respectively, as a function of training epoch
# launch TensorBoard from Jupyter
# %tensorboard --logdir logs
# Use the built-in ```evaluate``` function to see overall classification loss (first number) and accuracy (second number)
model.evaluate(x_test, y_test, verbose=0)
# The image classifier is now trained to ~98% accuracy on this dataset.
# #### Plot the Confusion Matrix
# A ‘confusion matrix’, which is the matrix of normalized correspondences between true and estimated labels, is a convenient way to visualize model skill.
#
# A perfect correspondence between true and estimated labels is scored 1.0 along the diagonal elements of the matrix.
#
# Misclassifications are readily identified as off-diagonal elements. Systematic misclassifications are recognized as off-diagonal elements with large magnitudes.
from sklearn.metrics import confusion_matrix
# The function below is long and might be difficult to follow in parts. This is a good time to remind you that that's ok; you will see this function again, and it isn't crucial to any workflow you will develop. It serves only to exemplify and guide, and many the details of many of the long functions such as these are largely unimportant
def plot_confusion_matrix(y_true, y_pred, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
fig, ax = plt.subplots(figsize=(8,8))
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
fig.tight_layout()
return ax
# Use the built-in ```predict``` function to create a vector of predicted classes for each input in the ```x_test``` set. These values are one-hot encoded, which means we recover the most likely class using the ```argmax``` function (i.e. the location in the vector, ```predictions```, of the maximum value)
# Test your model using the test data
predictions = model.predict(x_test)
predictions = tf.argmax(predictions, axis=1)
# +
# check the confusion matrix
classes=['0','1','2','3','4','5','6','7','8','9']
plot_confusion_matrix(y_test, predictions, classes)
# -
# The above demonstration shows that for relatively simple classification tasks (i.e. recognizing handwritten digits) doesn't necessarily require deep learning. Our network was small, consisting of only one hidden layer. In the next task we will be using much bigger, more powerful networks.
# <table style="font-size: 1em; padding: 0; margin: 0;">
# <p style="border: 1px solid #ff5733; border-left: 15px solid #ff5733; padding: 10px; text-align:justify;">
# <strong style="color: #ff5733">Deliverable</strong>
# <br/>The deliverable for Part 1 is a jupyter notebook showing an example image(s) of a satellite dataset read in using rasterio, and demonstration of a function that carries out a manipulation of that image using keras and Tensorflow 2.0. That manipulation could be anything that alters the image, such as its size, geometry (shape), pixel intensities, or spatial projection. This will mostly test your understanding of keras syntax, which is an essential component of the remaining Parts. You may find the <a href="https://www.tensorflow.org/api_docs/python/tf/image">tensorflow-image</a> library helpful.
# </p>
# <table style="font-size: 1em; padding: 0; margin: 0;">
#
# <h1 style="width: 100%; text-align: left; padding: 0px 25px;"><small style="color: #182AEB;">
# </small><br/>Going further: <br/> Introduction to satellite imagery</h1>
# <br/>
# <p style="border-left: 15px solid #182AEB; text-align:justify; padding: 0 10px;">
# <strong style="color: #182AEB;">Intro.</strong>
# The <a href="https://sentinels.copernicus.eu/">Sentinel satellites</a> are part of Europe’s <a href="http://copernicus.eu/">Copernicus </a> programme (formerly known as Global Monitoring for Environment and Security (GMES)). The overall mission is composed of five constellations of satellites. The Sentinel-2 mission consists of two satellites with a multi-spectral instrument for monitoring agriculture, vegetation and forests, land cover change, coastal zones, inland water monitoring, glaciers, ice extent and snow cover.
# </p>
# <p style="border-left: 15px solid #6019D6; padding: 0 10px; text-align:justify;">
# <strong style="color: #6019D6;">Tip.</strong>
# A comprehensive introduction to the mission and its data can be found <a href="https://eox.at/2015/12/understanding-sentinel-2-satellite-data/"> here</a>. Your starting point for all data services is <a href="https://www.sentinel-hub.com/sentinel-2">Sentinel Hub </a></p>
#
# <p style="border-left: 15px solid #4E9317; padding: 0 10px; text-align:justify;">
# <strong style="color: #4E9317;">More Resources.</strong>
# <ul>
# <li><a href="https://krstn.eu/download-Sentinel-2-images/">Another guide to using sentinselsat</a></li>
# <li><a href="https://automating-gis-processes.github.io/CSC18/lessons/L6/raster-mosaic.html">Using rasterio</a></li>
# </ul>
# </tr>
# </table>
# #### Importing a satellite image using rasterio
import rasterio
import glob
import os
# ##### Download a sample Sentinel-2 JP2 (JPEG2000) file from google drive
#
# The details of this function are unimportant
# +
# from https://stackoverflow.com/questions/38511444/python-download-files-from-google-drive-using-url
import requests
def download_file_from_google_drive(id, destination):
URL = "https://docs.google.com/uc?export=download"
session = requests.Session()
response = session.get(URL, params = { 'id' : id }, stream = True)
token = get_confirm_token(response)
if token:
params = { 'id' : id, 'confirm' : token }
response = session.get(URL, params = params, stream = True)
save_response_content(response, destination)
def get_confirm_token(response):
for key, value in response.cookies.items():
if key.startswith('download_warning'):
return value
return None
def save_response_content(response, destination):
CHUNK_SIZE = 32768
with open(destination, "wb") as f:
for chunk in response.iter_content(CHUNK_SIZE):
if chunk: # filter out keep-alive new chunks
f.write(chunk)
# -
file_id = '1o76QoBtn6ExxO8KgcCqdOiun_KsWoMJl'
destination = 'example_TCI_10m.jp2'
download_file_from_google_drive(file_id, destination)
# Read first band of the jp2 file into an array
with rasterio.open('example_TCI_10m.jp2', driver='JP2OpenJPEG') as dataset:
array = dataset.read(1)
print(dataset.profile)
# count = 3 means there are three bands in the raster:
#
# * Band 1 - the blue band
# * Band 2 - the green band
# * Band 3 - the red band
#
# A few more jp2 images can be downloaded using the following file id: 1o76QoBtn6ExxO8KgcCqdOiun_KsWoMJl. This is a zipped folder 120 MB in size containing several images of the area around Lake Poopó, Bolivia
# #### Writing a satellite image to geotiff using rasterio
#
# The following function takes a JP2 image, an output file name (with tiff extension) and a single band number (1, 2, or 3) and writes the raster to the new file
def write_image(input, output, band):
with rasterio.open(input) as src_dataset:
with rasterio.open('example_TCI_10m.jp2', driver='JP2OpenJPEG') as dataset:
array = dataset.read(band)
# Get a copy of the source dataset's profile. Thus our
# destination dataset will have the same dimensions,
# number of bands, data type, and georeferencing as the
# source dataset.
kwds = src_dataset.profile
# Change the format driver for the destination dataset to
# 'GTiff', short for GeoTIFF.
kwds['driver'] = 'GTiff'
# Add GeoTIFF-specific keyword arguments.
kwds['tiled'] = True
kwds['blockxsize'] = 256
kwds['blockysize'] = 256
kwds['photometric'] = 'YCbCr'
kwds['compress'] = 'JPEG'
with rasterio.open(output, 'w', **kwds) as dst_dataset:
# Write data to the destination dataset.
dst_dataset.write(array.astype(rasterio.uint8), 1)
# Here is the list of files we will create, each with a 1-band raster
file_list = ['r1.tif', 'r2.tif', 'r3.tif']
# We use list comprehension to call the ```write_image``` function for all 3 bands in turn
[write_image('example_TCI_10m.jp2',f, band) for f,band in zip(file_list, [1,2,3])]
# This next bit of code gets the meta data from the first file (that is the same as the remaining two files), and writes a merged 3-band raster in geoTIFF format
# +
# Read metadata of first file
with rasterio.open(file_list[0]) as src0:
meta = src0.meta
# Update meta to reflect the number of layers
meta.update(count = len(file_list))
# Read each layer and write it to stack
with rasterio.open('stack.tif', 'w', **meta) as dst:
for id, layer in enumerate(file_list, start=1):
with rasterio.open(layer) as src1:
dst.write_band(id, src1.read(1))
# -
# We're now done with the three intermediate files, so we can delete them to save space
[os.remove(f) for f in file_list]
# Display the new meta data that shows the Coordinate Reference System or CRS. The EPSG code is 32719
meta
# #### Image spatial projections
# Next we're going to reproject our raster so it is in a different spatial projection. This involves specifying a new CRS (this time we use EPSG code 4326)
#
# Example modified from https://rasterio.readthedocs.io/en/stable/topics/reproject.html
# First we import some utility functions from rasterio that will create and apply the transformation. Again we use a nested 'with statement' to first read in the raster to be projected (```stack.tif```) and then write out the reprojected raster (```reprojected_stack.tif```)
# +
from rasterio.warp import calculate_default_transform, reproject, Resampling
dst_crs = 'EPSG:4326'
# note there is a nested 'with statement' here
# the first 'with' command opens the image as src
with rasterio.open('stack.tif') as src:
transform, width, height = calculate_default_transform(
src.crs, dst_crs, src.width, src.height, *src.bounds)
kwargs = src.meta.copy()
kwargs.update({
'crs': dst_crs,
'transform': transform,
'width': width,
'height': height
})
# the second 'with' statement opens an image for writing
with rasterio.open('reprojected_stack.tif', 'w', **kwargs) as dst:
for i in range(1, src.count + 1):
reproject(
source=rasterio.band(src, i),
destination=rasterio.band(dst, i),
src_transform=src.transform,
src_crs=src.crs,
dst_transform=transform,
dst_crs=dst_crs,
resampling=Resampling.nearest)
# -
| dl-satellite-image/p1_intro/project_data/Part_1_GettingStarted.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### TASK
# A Bayesian network (also known as a Bayes network, belief network, or decision network) is a probabilistic graphical model that represents a set of variables and their conditional dependencies via a directed acyclic graph (DAG).
#
# They can be used for a wide range of tasks including prediction, anomaly detection, diagnostics, automated insight, reasoning, time series prediction and decision making under uncertainty.
#
# We will write a program to construct a Bayesian Network considering medical data. We will use this program to do dignosis of heart patient dataset.
#
# To implement Bayesian Network we will be using PGMPY(Probabilstic Graphical Models).
# ### Dataset acknowledgements
# We will be using UCI Heart Disease Data Set : Processed Cleveland Data.
#
# The UCI Machine Learning Repository is a collection of databases, domain theories, and data generators that are used by the machine learning community.
#
# The dataset consists of 303 individuals data. There are 14 columns in the dataset, which are described below. The last attribute is the predicted attribute.
# ### Description of Dataset Columns:
# 1. Age: The age of the individual.It is the most important risk factor in developing cardiovascular diseases.The risk of stroke doubles every decade after age 55.
#
#
# 2. Sex: The gender of the individual.Men are at greater risk of heart disease than pre-menopausal women.Format for same:
# 1 = male
# 0 = female
#
#
# 3. Chest-pain type: The type of chest-pain experienced by the individual.Angina is chest pain or discomfort caused when your heart muscle doesn’t get enough oxygen-rich blood.Format :
# 1 = typical angina
# 2 = atypical angina
# 3 = non — anginal pain
# 4 = asymptotic
#
#
# 4. Resting Blood Pressure: The resting blood pressure value of an individual in mmHg (unit).Over time, high blood pressure can damage arteries that feed your heart.
#
#
# 5. Serum Cholestrol: displays the serum cholesterol in mg/dl (unit)
# A high level of low-density lipoprotein (LDL) cholesterol also called the bad cholesterol is most likely to narrow arteries. However, a high level of high-density lipoprotein (HDL) cholesterol (the “good” cholesterol) lowers your risk of a heart attack.
#
#
# 6. Fasting Blood Sugar: It compares the fasting blood sugar value of an individual.Not responding to insulin properly causes your body’s blood sugar levels to rise, increasing your risk of a heart attack.
# If fasting blood sugar > 120mg/dl then : 1 (true)
# else : 0 (false)
#
# 7. Resting ECG: Displays resting electrocardiographic results.
# 0 = normal
# 1 = having ST-T wave abnormality
# 2 = left ventricular hyperthrophy
#
#
# 8. Max heart rate achieved: displays the max heart rate achieved by an individual.
# It has been shown that an increase in heart rate by 10 beats per minute was associated with an increase in the risk of cardiac death by at least 20%
#
#
# 9. Exercise induced angina: The pain or discomfort associated with angina usually feels tight, gripping or squeezing, and can vary from mild to severe.
# 1 = yes
# 0 = no
#
#
# 10. ST depression induced by exercise relative to rest: displays the value which is an integer or float.
#
#
# 11. Peak exercise ST segment :
# 1 = upsloping
# 2 = flat
# 3 = downsloping
#
#
# 12. Number of major vessels(0–3) colored by flourosopy: Displays the value as integer or float.
#
#
# 13. Thal: Displays the thalassemia:
# 3 = normal
# 6 = fixed defect
# 7 = reversible defect
#
#
# 14. Diagnosis of heart disease: Displays whether the individual is suffering from heart disease or not :
# It is integer valued from 0 (no presence) to 4. Presence (values 1,2,3,4) from absence (value 0).
# #### Requirements
# - python 3.6 or higher
# - pandas
# - pgmpy
# networkX
# scipy
# numpy
# pytorch
# ### PGMPY:Probabilistic Graphical Models
# Probabilistic Graphical Models(PGM) are a very solid way of representing joint probability distributions on a set of random variables. It allows users to do inferences in a computationally efficient way.We can calculate the joint probability distribution of these variables by combining various parameters taken from the graph.
#
# Mainly, there are two types of Graph models:
#
# Baysian Graph Models : These models consist of Directed-Cyclic Graph(DAG) and there is always a conditional probability associated with the random variables. These types of models represent a causation between the random variables.
#
# Markov Graph Models : These models are undirected graphs and represent non causal relationships between the random variables.
# pgmpy is a python framework to work with these types of graph models. Several graph models and inference algorithms are implemented in pgmpy. Pgmpy also allows users to create their own inference algorithm without getting into the details of the source code of it.
#
# Let’s get started with the implementation part.
# #### Installing PGMPY
# +
# !pip install pgmpy;
# import warnings filter
from warnings import simplefilter
simplefilter(action='ignore', category = FutureWarning)
# -
# Make sure to add an exclamation mark, so that this command doesnt run in jupyetr-notebook but externally for your system.
import pandas as pd
# We will be importing our dataset directly from UCI Website. Hence we need to import urllib.
from urllib.request import urlopen
#data = pd.read_csv(urlopen("http://bit.do/heart-disease"))
from pgmpy.models import BayesianModel
# #### Getting the data
# +
#Or directly getting data from saved file:
data = pd.read_csv("cleveland heart data.txt")
data.head()
# -
# #### Working with column names
# As you can see we need to rework here with the column names.For now, let's give the alphabetical letters to these columns
# +
names = "Age,Sex,Angina_(Chest_Pain),Resting_BP,Serum Cholesterol,Fasting_Sugar,Resting_ECG,Max_Heart_Rate,Excercise_Induced_Angina,Exercise_induces_ST,ST_Segment,Vessels,Thalassemia,Heart_Diagnosis"
names = names.split(",")
data = pd.read_csv("cleveland heart data.txt",names=names)
data.head()
# -
# Looks Much better! :)
# ### Analysing The Data
import matplotlib.pyplot as plt
import seaborn as sns
# Lets see the distribution of age in our dataset with Distribution Plot
plt.figure(figsize=(20,12))
sns.displot(data,x="Age",hue="Heart_Diagnosis",element = "step");
plt.show()
# As we can see most number of paatients lie in the age group above 50.
# Histogram plot for distribution of cases on the basis of gender.
sns.histplot(data,x="Age",hue="Sex")
sns.histplot(data=data)
# Heatmap helps us visualize matrix-like data in the form of hot and cold spots. The warm colors indicated sections with the most visitor interaction.
# +
plt.figure(figsize=(12,7));
sns.heatmap(data.corr());
# Rotate x-labels with the help of matplotlib
plt.xticks(rotation=25);
# -
# lets see the relationship of heart rate and diagnosis in our result:
sns.histplot(data,x="Max_Heart_Rate",hue="Heart_Diagnosis")
plt.figure(figsize=(12,7));
sns.lmplot(x="Age",y="Max_Heart_Rate", data=data,fit_reg = False,hue="Heart_Diagnosis");
plt.xlim(0,85);
# From the above scatter graph we clearly see tht pour distribution majorly lies in the age of 55-65 for individuals identified with heart diagnosis. We can also see that the max heart rate pf people diagnosed positive has not been above 130 in most cases.
# #### PRE-PROCESSING OUR DATASET:
# Checking for any null values:
data.isnull().sum()
# There are no null values in our dataset. We have also got a basic idea of our data and analytics to proceed further.
# ### Implementing the Bayesian Model:
#
# The columns names here are the parameters on which we are constructing our Bayesian Model. We will make connections between the different parameters to make a model.
model = BayesianModel([(),(),()])
# References:
#
# 1. https://www.youtube.com/watch?v=sCqpWCYaL5U&t=16s
# 2. https://www.bayesserver.com/docs/introduction/bayesian-networks
# 3.https://analyticsindiamag.com/guide-to-pgmpy-probabilistic-graphical-models-with-python-code/
# 4. https://pgmpy.org/
# 5. https://towardsdatascience.com/heart-disease-prediction-73468d630cfc
| Bayesian Network Implementation for Heart Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3
# language: python
# name: py3
# ---
class Solution:
def lengthOfLastWord(self, s: str) -> int:
strs = s.split()
if len(strs) > 0:
return len(strs[-1])
else:
return 0
s = Solution()
s.lengthOfLastWord("")
| algorithms/58-length-of-last-word.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false id="11lq_Y7ygqZo" nbgrader={"cell_type": "markdown", "checksum": "cbb06e85a5b52e17712dc9aa00cacf97", "grade": false, "grade_id": "cell-e788bbc75112bf0e", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Principal Component Analysis (PCA)
# + [markdown] deletable=false editable=false id="BSU6DDobgqZr" nbgrader={"cell_type": "markdown", "checksum": "e0b9067dc44ccb9f5364e7ba6c1f2855", "grade": false, "grade_id": "cell-0f10ea8a61af22a1", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Importing libraries
# + deletable=false editable=false id="gA9L1vzcgqZt" nbgrader={"cell_type": "code", "checksum": "b280d44e9e482a6021a6689794c78807", "grade": false, "grade_id": "cell-f62b5bb08491d57e", "locked": true, "schema_version": 3, "solution": false, "task": false}
# importing libraries
import os
import sys
import cv2 as cv
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
from sklearn.model_selection import train_test_split
# + [markdown] deletable=false editable=false id="SCQZMpAdgqZz" nbgrader={"cell_type": "markdown", "checksum": "4e9ba33c05df545d5b6a7e2eb79da0d1", "grade": false, "grade_id": "cell-3d8f464e04b42fad", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Loading dataset
# + [markdown] id="N1z0kBwmgqZ0" nbgrader={"cell_type": "markdown", "checksum": "b848efd862301f8f82fb991fa16f9480", "grade": false, "grade_id": "cell-ddb9c8882b90dd40", "locked": true, "schema_version": 3, "solution": false, "task": false}
# PCA is used when we need to tackle the curse of dimensionality among data with linear relationships , i.e. where having too many dimensions (features) in your data causes noise and difficulties.
# + [markdown] id="m38G4mVigqZ0" nbgrader={"cell_type": "markdown", "checksum": "4a2ccae399d751bbfed8259eaf632eb1", "grade": false, "grade_id": "cell-5343a69b5c5805af", "locked": true, "schema_version": 3, "solution": false, "task": false}
# For the sake of visually being able to see that our algorthm works or not, we will work with image data.
# + id="iMwyurLIgqZ1"
# Loading data and reducing size to 32 x 32 pixels
#IMG_DIR = '../PCA/Anime images 1000'
IMG_DIR = 'C:/Users/pranj/PCA/Anime images 1000' # YOUR CODE HERE - Edit the path for dataset
print("Loading...")
X = []
X_flat = []
count = 1
total_samples = 200
image_size = 32
for img in os.listdir(IMG_DIR):
if count == total_samples + 1:
break
sys.stdout.write("\r" + str(count) + " / " + str(total_samples))
sys.stdout.flush()
img_array = cv.imread(os.path.join(IMG_DIR,img), cv.IMREAD_GRAYSCALE)
img_pil = Image.fromarray(img_array)
img_48x48 = np.array(img_pil.resize((image_size, image_size), Image.ANTIALIAS))
X.append(img_48x48)
img_array = img_48x48.flatten()
X_flat.append(img_array)
count += 1
print()
print("Done!")
# + id="EezSeptkgqZ5" nbgrader={"cell_type": "code", "checksum": "692b91b4ced34732eec6ef8461c67d27", "grade": false, "grade_id": "cell-166d675c7a05c7aa", "locked": true, "schema_version": 3, "solution": false, "task": false}
# visualizing some images
fig, ax = plt.subplots(figsize = (20, 20))
for i in range(1, 26):
plt.subplot(5, 5, i)
plt.title('Anime image {}'.format(i-1))
plt.imshow(X[i-1])
# + id="5jGcfJpfgqZ9" nbgrader={"cell_type": "code", "checksum": "b5be65c7c327da8462acba5fb3aca830", "grade": false, "grade_id": "cell-fef1604a6ff42121", "locked": true, "schema_version": 3, "solution": false, "task": false}
# converting X_flat to numpy array
X_flat = np.asarray(X_flat)
X_flat.shape
# + id="2pUqG05ZgqZ_" nbgrader={"cell_type": "code", "checksum": "192f03f987d8c3b39ff632f653b6c793", "grade": false, "grade_id": "cell-3994c1bacdf60d07", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Test-Train split
X_train, X_test = train_test_split(X_flat, test_size = 0.125, random_state = 42)
X_train = X_train.T
X_test = X_test.T
print(X_train.shape)
print(X_test.shape)
# + [markdown] id="vCCrz0wsgqaD" nbgrader={"cell_type": "markdown", "checksum": "09478ff3e4409999483bc35e7207c860", "grade": false, "grade_id": "cell-f05394c641517c37", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### PCA
# + [markdown] id="LJPtGU_tgqaD" nbgrader={"cell_type": "markdown", "checksum": "c171f6da1969776e79c8c53db3119a3a", "grade": false, "grade_id": "cell-9f726c981fc7ad13", "locked": true, "schema_version": 3, "solution": false, "task": false}
# The steps needed to compute PCA are broken into following parts
#
# 1. Data standardization.
# 2. Find eigenvalues and corresponding eigenvectors for the covariance matrix $\boldsymbol S$. Sort by the largest eigenvalues and the corresponding eigenvectors (eig).
#
# 3. Compute the projection and reconstruction of the data onto the space spanned by the top $M$ eigenvectors.
# + id="5EBDU_vWgqaE" nbgrader={"cell_type": "code", "checksum": "a116855c6f249851bbe8bb11cc5366c7", "grade": false, "grade_id": "cell-0668a0ebc0ecc859", "locked": false, "schema_version": 3, "solution": true, "task": false}
# function for standardizing image
def Standardize(X):
# calculate the mean along each feature dimension
# note that data is stored as column vectors
#X = X.T
# YOUR CODE HERE
mu = np.mean(X,axis=1)
#raise NotImplementedError()
# substract the mean vector from every data vector
# YOUR CODE HERE
Xbar = X - np.vstack(mu)
#raise NotImplementedError()
return Xbar, mu
# + id="t_TcrPksgqaJ" nbgrader={"cell_type": "code", "checksum": "d403c139deccc890c35568df7b16d3ce", "grade": true, "grade_id": "cell-59b50d0fb7f0eadd", "locked": true, "points": 2, "schema_version": 3, "solution": false, "task": false}
# check if standardization is correct
X1 = np.diag([1, 3, 5])
Xbar1, mu1 = Standardize(X1)
assert np.isclose(mu1, [0.33, 1., 1.66], atol = 0.05).all()
assert np.isclose(Xbar1, [[0.66, -0.33, -0.33], [-1., 2., -1.], [-1.66, -1.66, 3.33]], atol = 0.05).all()
# + id="gAMnwLjVgqaL" nbgrader={"cell_type": "code", "checksum": "fd69c9b3a8883cc7a6a70b62da71f944", "grade": false, "grade_id": "cell-e809e3771a8f6973", "locked": false, "schema_version": 3, "solution": true, "task": false}
# function for calculating eigen values and eigen vectors
def eig(S):
# calculate the eigen values and eigen vectors input covariance matrix S
# YOUR CODE HERE
eig_val,eig_vec = np.linalg.eigh(S)
#raise NotImplementedError()
# sort the eigen values in decreasing order and pick corresponding eigen vectors
# YOUR CODE HERE
idx = eig_val.argsort()[::-1]
eig_val = eig_val[idx]
eig_vec = eig_vec[:,idx]
#raise NotImplementedError()
return eig_val, eig_vec
# + id="9dbUDxK6gqaO" nbgrader={"cell_type": "code", "checksum": "3c5523a92a4987dfc52366b406e59e20", "grade": true, "grade_id": "cell-ef385fea11061a37", "locked": true, "points": 3, "schema_version": 3, "solution": false, "task": false}
# check if eigen values and eigen vectors are correct
X1 = np.diag([1, 3, 5])
eig_val1, eig_vec1 = eig(X1)
assert np.isclose(eig_val1, [5, 3, 1], atol = 0.05).all()
assert np.isclose(eig_vec1, [[0, 0, 1], [0, 1, 0], [1, 0, 0]], atol = 0.05).all()
# + id="zWziM3kBgqaQ" nbgrader={"cell_type": "code", "checksum": "626f1b1d2d4db37b3c1cc698dd94e376", "grade": false, "grade_id": "cell-f2259f76582f5d1b", "locked": false, "schema_version": 3, "solution": true, "task": false}
# function for projection matrix
def projection_matrix(B):
# calculate the projection matrix P
P = B.dot(B.T)
# YOUR CODE HERE
#raise NotImplementedError()
return P
# + id="RhEgMg_fgqaT" nbgrader={"cell_type": "code", "checksum": "febe0b14dbf85a7411fff0e72b9c9ac0", "grade": true, "grade_id": "cell-cacc3521712c9acc", "locked": true, "points": 2, "schema_version": 3, "solution": false, "task": false}
# check if projection matrix is correct
X1 = np.diag([1, 3, 5])
assert np.isclose(projection_matrix(X1), [[1, 0, 0], [0, 9, 0], [0, 0, 25]], atol = 0.05).all()
# + id="Vtj2kspCgqaW" nbgrader={"cell_type": "code", "checksum": "791f43a13b109a02c961800fb8148ca2", "grade": false, "grade_id": "cell-d28a5180118eacc6", "locked": false, "schema_version": 3, "solution": true, "task": false}
# implementing PCA
def PCA(X, num_components):
# calculate the data covariance matrix S. The input matrix X will have mean corrected entries
S = np.cov(X)
# YOUR CODE HERE
#raise NotImplementedError()
# now find eigenvalues and corresponding eigenvectors for S by implementing eig().
eig_vals, eig_vecs = eig(S)
# select eigen vectors
U = eig_vecs[:, range(num_components)]
# reconstruct the images from the lowerdimensional representation
# to do this, we first need to find the projection_matrix (which you implemented earlier)
# which projects our input data onto the vector space spanned by the eigenvectors
P = projection_matrix(U) # projection matrix
return P
# + id="Ht6K9lnogqaa" nbgrader={"cell_type": "code", "checksum": "9b591d67ad71f792b053b06b6f798762", "grade": true, "grade_id": "cell-84c53a3dcfaf8a4c", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Check if PCA is correct
X1 = np.diag([1, 3, 5])
num_components1 = 2
X1, mu = Standardize(X1)
assert np.isclose(PCA(X1, num_components1), [[0.13, -0.28, -0.17], [-0.28, 0.90, -0.05], [-0.17, -0.05, 0.96]], atol = 0.05).all()
# + [markdown] id="EpLyzq-xgqac" nbgrader={"cell_type": "markdown", "checksum": "b4b2e9e2f099d913638aa32333f7a104", "grade": false, "grade_id": "cell-840ed1b9deb1f0d3", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Standardization
# + [markdown] id="-S0QDiAogqad" nbgrader={"cell_type": "markdown", "checksum": "61812fae0b04f2c1616fb88a00a6f969", "grade": false, "grade_id": "cell-6e35266a1272867b", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Now, with the help of the functions you have implemented above, let's implement PCA!
# + id="ou69kKIugqae" nbgrader={"cell_type": "code", "checksum": "4ee3cc9aec83416f116b0f069e941653", "grade": false, "grade_id": "cell-5e08587e0a505b85", "locked": true, "schema_version": 3, "solution": false, "task": false}
# standardizing
Xbar_train, mu_train = Standardize(X_train)
Xbar_test, mu_test = Standardize(X_test)
# + id="13WYp6PNgqah" nbgrader={"cell_type": "code", "checksum": "172feb64f08b475038edf8686efc5e5c", "grade": false, "grade_id": "cell-308a6e03de6499f9", "locked": true, "schema_version": 3, "solution": false, "task": false}
# function for mean square error
def mse(predict, actual):
return np.square(predict - actual).mean()
# + [markdown] id="Lu5KRIuegqaj" nbgrader={"cell_type": "markdown", "checksum": "459662ab72f20f3b17c3482fc9c14021", "grade": false, "grade_id": "cell-e56bf9a0a544a2f8", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Loss and Reconstruction
# + id="8KnhYlXagqak" nbgrader={"cell_type": "code", "checksum": "f456ebc22d0394ffdae8058008a77a2f", "grade": false, "grade_id": "cell-42cb160012ec8040", "locked": true, "schema_version": 3, "solution": false, "task": false}
# calculating loss and reconstructing images
loss = []
reconstructions = []
max_components = len(X_train.T)
print("Processing...")
animation = np.arange(1, max_components + 1, 1)
for num_component in range(1, max_components + 1):
sys.stdout.write("\r" + str(animation[num_component - 1]) + " / " + str(max_components))
sys.stdout.flush()
projection = PCA(Xbar_train, num_component)
reconst = projection @ Xbar_test
error = mse((reconst.T + mu_test).T, X_test)
reconstructions.append(reconst)
loss.append((num_component, error))
print()
print("Done!")
# + id="xeileocKgqaq" nbgrader={"cell_type": "code", "checksum": "056f384173af63798d7734d8cb1db957", "grade": false, "grade_id": "cell-e62bc02e33613ccf", "locked": true, "schema_version": 3, "solution": false, "task": false}
# converting reconstructions and loss to numpy arrays
reconstructions = np.asarray(reconstructions)
loss = np.asarray(loss)
# + id="VGXRbQkcgqau" nbgrader={"cell_type": "code", "checksum": "28743a8eb1f2efac9b8223986ece9639", "grade": false, "grade_id": "cell-0aa60332a67fdd92", "locked": true, "schema_version": 3, "solution": false, "task": false}
# visualizing mse vs number of principal components
fig, ax = plt.subplots(figsize = (20, 8))
ax.plot(loss[:,0], loss[:, 1])
ax.xaxis.set_ticks(np.arange(0, max_components, 15))
ax.set(xlabel = 'num_components', ylabel = 'MSE', title = 'MSE vs number of principal components')
plt.grid()
# + [markdown] id="BmyL9DVzgqay" nbgrader={"cell_type": "markdown", "checksum": "4707d7d2132cf6f32760c3eefabd21b3", "grade": false, "grade_id": "cell-4c5a6233a760c120", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Conclusion:
#
# More the principle components used, better is the accuracy of reconstruction even though the images were unseen.
# + id="FwN-VsF7gqaz"
# plotting the reconstructed images
n = 1 # number of double-rows
m = 5 # number of columns
t = 0.9 # 1-t == top space
b = 0.1 # bottom space (both in figure coordinates)
msp = 0.3 # minor spacing
sp = 0.8 # major spacing
offs = (1+msp) * (t-b) / (2*n + n*msp + (n-1)*sp) # grid offset
hspace = sp + msp + 1 # height space per grid
gso = GridSpec(n, m, bottom = b+offs, top = t, hspace = hspace)
gse = GridSpec(n, m, bottom = b, top = t-offs, hspace = hspace)
fig = plt.figure(figsize=(20, 8))
axes = []
for i in range(n*m):
axes.append(fig.add_subplot(gso[i]))
axes.append(fig.add_subplot(gse[i]))
k = 1
for j in range(10):
axes[j].set_title('{} principal components'.format(k))
axes[j].imshow(reconstructions[k, :, 0].reshape(image_size, image_size))
k += 19
plt.show()
# + [markdown] id="yHyzqvD6gqa4" nbgrader={"cell_type": "markdown", "checksum": "7c23e6507ef45579e052eeb26b58a52e", "grade": false, "grade_id": "cell-e3b5e2aa4903e207", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Points to note:
# + [markdown] id="hkxSHF5Mgqa5" nbgrader={"cell_type": "markdown", "checksum": "0580345c8fc2e40bc652ed916715849f", "grade": false, "grade_id": "cell-edd3cef85f32f72c", "locked": true, "schema_version": 3, "solution": false, "task": false}
# 1.) Projection matrix is learnt from the training data.
#
# 2.) Test image is reconstructed from the projection matrix.
#
# 3.) As the number of principle components increase, the reconstruction accuracy increases.
#
# 4.) Try increasing the number of training samples and notice the changes in MSE. What do you obeserve?
#
# 5.) You can also try and increase the image size from 32 to 64 or 128. (Processing time will be longer)
| .ipynb_checkpoints/PCA from scratch-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import urllib.parse
import requests
import pickle
import os
import re
import time
# %load_ext nb_black
# +
color_to_rare = {
"b0c3d9": "Consumer Grade",
"5e98d9": "Industrial Grade",
"4b69ff": "Mil-Spec",
"8847ff": "Restricted",
"d32ce6": "Classified",
"eb4b4b": "Covert",
"ffd700": "Special",
}
def parse_weapon(weap_json):
name = weap_json["name"]
stattrak = "StatTrak" in name
name_match = re.match(r"([★™\w\- ]+) \| ([\w ]+) \(([\w\- ]+)\)", name)
is_case = name.endswith(" Case")
is_package = name.endswith(" Package")
is_key = name.endswith(" Case Key")
if name_match is None and not is_case and not is_key and not is_package:
return None
if not is_case and not is_key and not is_package:
weapon = name_match.group(1).replace("★ ", "").replace("StatTrak™ ", "")
skin = name_match.group(2)
cond = name_match.group(3)
elif is_key:
skin = "n/a"
cond = "n/a"
weapon = "Key"
elif is_package:
skin = "n/a"
cond = "n/a"
weapon = "Package"
elif is_case:
skin = "n/a"
cond = "n/a"
weapon = "Case"
if weapon in ["Sticker", "Sealed Graffiti"]:
return None
url = "https://steamcommunity.com/market/listings/730/" + urllib.parse.quote(name)
data = {
"name": name,
"stattrak": stattrak,
"weapon": weapon,
"skin": skin,
"condition": cond,
"sell_price": weap_json["sell_price"] / 100,
"sell_listings": weap_json["sell_listings"],
"url": url,
"is_key": is_key,
"is_case": is_case,
"is_package": is_package,
}
return data
def get_prices(resp):
df_data = []
for match in re.finditer(r'\["([\w ]+: \+0)",([\d\.]+),"(\d+)"\]', resp):
date, price, vol = match.group(1), match.group(2), match.group(3)
vol = int(vol)
price = float(price)
date = " ".join(date.split(" ")[:3])
df_data.append([date, price, vol])
df = pd.DataFrame(df_data, columns=["date", "price", "volume"])
m = re.search(r'"type":"([^"]+?)","market_name"', resp)
if m is None:
return None
mc = re.search(r'"value":"([\w\- ]+ Collection)","color":"9da1a9"', resp)
if mc is None:
collection = ""
else:
collection = mc.group(1)
items = [
(im.group(1), im.group(2), color_to_rare.get(im.group(3), im.group(3)))
for im in re.finditer(
r'"value":"([\w\- ]+) \| ([\w\- ]+)","color":"(\w+)"', resp
)
]
type_ = m.group(1).replace("StatTrak\\u2122", "").replace("\\u2605 ", "").strip()
data = {"type": type_, "prices": df, "items": items, "collection": collection}
return data
# -
if os.path.exists("dataset.pkl"):
with open("dataset.pkl", "rb") as f:
dataset = pickle.load(f)
else:
dataset = {}
# +
header_idx = 0
headers = [
{
"user-agent": "Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/88.0.4324.190 Safari/537.36",
},
]
base_url = "https://steamcommunity.com/market/search/render/"
window_size = 100
i = 0
while True:
print("Cursor", i)
try:
page = requests.get(
base_url
+ "?start={}&count={}&search_descriptions=0&appid=730&norender=1".format(
i, window_size
),
headers=headers[header_idx % len(headers)],
)
page_json = page.json()
assert len(page_json["results"]) > 0
except (TypeError, OSError, AssertionError) as e:
print("Load error", e, page.text)
header_idx += 1
for _ in range(20):
time.sleep(3)
continue
for res in page_json["results"]:
weap = parse_weapon(res)
if weap is None or weap["name"] in dataset:
continue
name = weap["name"]
price_resp = requests.get(
weap["url"], headers=headers[header_idx % len(headers)]
).text
data = get_prices(price_resp)
if data is None:
continue
print(name, weap["url"])
weap.update(data)
dataset[name] = weap.copy()
i += window_size
# -
assert len(dataset) > 0
print(len(dataset))
with open("dataset.pkl", "wb") as f:
pickle.dump(dataset, f)
| CollectData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
keras.__version__
# # Using word embeddings
#
# This notebook contains the second code sample found in Chapter 6, Section 1 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
#
# ---
#
#
# Another popular and powerful way to associate a vector with a word is the use of dense "word vectors", also called "word embeddings".
# While the vectors obtained through one-hot encoding are binary, sparse (mostly made of zeros) and very high-dimensional (same dimensionality as the
# number of words in the vocabulary), "word embeddings" are low-dimensional floating point vectors
# (i.e. "dense" vectors, as opposed to sparse vectors).
# Unlike word vectors obtained via one-hot encoding, word embeddings are learned from data.
# It is common to see word embeddings that are 256-dimensional, 512-dimensional, or 1024-dimensional when dealing with very large vocabularies.
# On the other hand, one-hot encoding words generally leads to vectors that are 20,000-dimensional or higher (capturing a vocabulary of 20,000
# token in this case). So, word embeddings pack more information into far fewer dimensions.
# 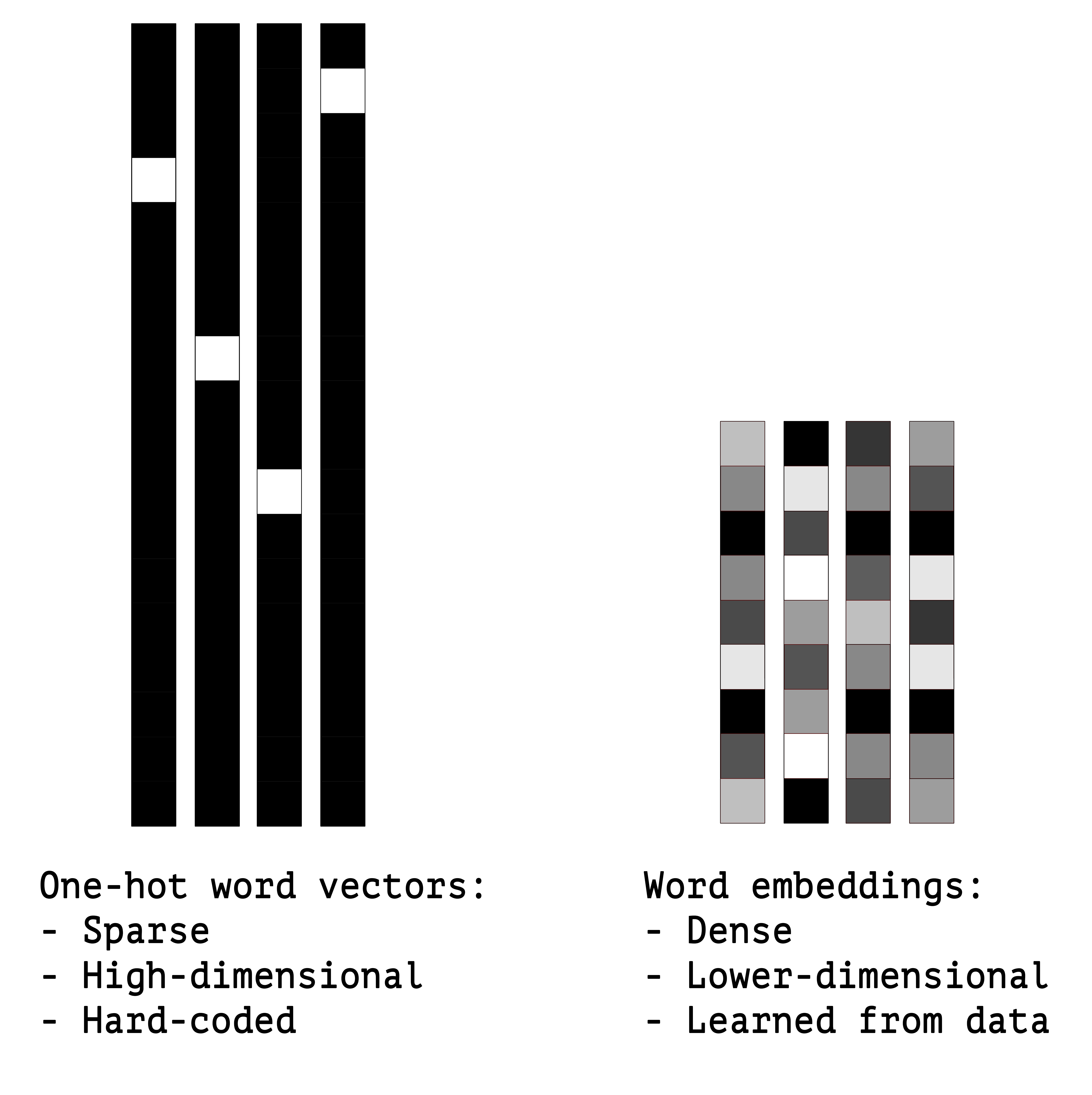
# There are two ways to obtain word embeddings:
#
# * Learn word embeddings jointly with the main task you care about (e.g. document classification or sentiment prediction).
# In this setup, you would start with random word vectors, then learn your word vectors in the same way that you learn the weights of a neural network.
# * Load into your model word embeddings that were pre-computed using a different machine learning task than the one you are trying to solve.
# These are called "pre-trained word embeddings".
#
# Let's take a look at both.
# ## Learning word embeddings with the `Embedding` layer
#
#
# The simplest way to associate a dense vector to a word would be to pick the vector at random. The problem with this approach is that the
# resulting embedding space would have no structure: for instance, the words "accurate" and "exact" may end up with completely different
# embeddings, even though they are interchangeable in most sentences. It would be very difficult for a deep neural network to make sense of
# such a noisy, unstructured embedding space.
#
# To get a bit more abstract: the geometric relationships between word vectors should reflect the semantic relationships between these words.
# Word embeddings are meant to map human language into a geometric space. For instance, in a reasonable embedding space, we would expect
# synonyms to be embedded into similar word vectors, and in general we would expect the geometric distance (e.g. L2 distance) between any two
# word vectors to relate to the semantic distance of the associated words (words meaning very different things would be embedded to points
# far away from each other, while related words would be closer). Even beyond mere distance, we may want specific __directions__ in the
# embedding space to be meaningful.
#
# [...]
#
#
# In real-world word embedding spaces, common examples of meaningful geometric transformations are "gender vectors" and "plural vector". For
# instance, by adding a "female vector" to the vector "king", one obtain the vector "queen". By adding a "plural vector", one obtain "kings".
# Word embedding spaces typically feature thousands of such interpretable and potentially useful vectors.
#
# Is there some "ideal" word embedding space that would perfectly map human language and could be used for any natural language processing
# task? Possibly, but in any case, we have yet to compute anything of the sort. Also, there isn't such a thing as "human language", there are
# many different languages and they are not isomorphic, as a language is the reflection of a specific culture and a specific context. But more
# pragmatically, what makes a good word embedding space depends heavily on your task: the perfect word embedding space for an
# English-language movie review sentiment analysis model may look very different from the perfect embedding space for an English-language
# legal document classification model, because the importance of certain semantic relationships varies from task to task.
#
# It is thus reasonable to __learn__ a new embedding space with every new task. Thankfully, backpropagation makes this really easy, and Keras makes it
# even easier. It's just about learning the weights of a layer: the `Embedding` layer.
# +
from keras.layers import Embedding
# The Embedding layer takes at least two arguments:
# the number of possible tokens, here 1000 (1 + maximum word index),
# and the dimensionality of the embeddings, here 64.
embedding_layer = Embedding(1000, 64)
# -
#
# The `Embedding` layer is best understood as a dictionary mapping integer indices (which stand for specific words) to dense vectors. It takes
# as input integers, it looks up these integers into an internal dictionary, and it returns the associated vectors. It's effectively a dictionary lookup.
#
# The `Embedding` layer takes as input a 2D tensor of integers, of shape `(samples, sequence_length)`, where each entry is a sequence of
# integers. It can embed sequences of variable lengths, so for instance we could feed into our embedding layer above batches that could have
# shapes `(32, 10)` (batch of 32 sequences of length 10) or `(64, 15)` (batch of 64 sequences of length 15). All sequences in a batch must
# have the same length, though (since we need to pack them into a single tensor), so sequences that are shorter than others should be padded
# with zeros, and sequences that are longer should be truncated.
#
# This layer returns a 3D floating point tensor, of shape `(samples, sequence_length, embedding_dimensionality)`. Such a 3D tensor can then
# be processed by a RNN layer or a 1D convolution layer (both will be introduced in the next sections).
#
# When you instantiate an `Embedding` layer, its weights (its internal dictionary of token vectors) are initially random, just like with any
# other layer. During training, these word vectors will be gradually adjusted via backpropagation, structuring the space into something that the
# downstream model can exploit. Once fully trained, your embedding space will show a lot of structure -- a kind of structure specialized for
# the specific problem you were training your model for.
#
# Let's apply this idea to the IMDB movie review sentiment prediction task that you are already familiar with. Let's quickly prepare
# the data. We will restrict the movie reviews to the top 10,000 most common words (like we did the first time we worked with this dataset),
# and cut the reviews after only 20 words. Our network will simply learn 8-dimensional embeddings for each of the 10,000 words, turn the
# input integer sequences (2D integer tensor) into embedded sequences (3D float tensor), flatten the tensor to 2D, and train a single `Dense`
# layer on top for classification.
# +
from keras.datasets import imdb
from keras import preprocessing
# Number of words to consider as features
max_features = 10000
# Cut texts after this number of words
# (among top max_features most common words)
maxlen = 20
# Load the data as lists of integers.
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
# -
x_train.ndim
x_train.shape
# This turns our lists of integers
# into a 2D integer tensor of shape `(samples, maxlen)`
x_train = preprocessing.sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = preprocessing.sequence.pad_sequences(x_test, maxlen=maxlen)
x_train.ndim
x_train.shape
# +
from keras.models import Sequential
from keras.layers import Flatten, Dense
model = Sequential()
# We specify the maximum input length to our Embedding layer
# so we can later flatten the embedded inputs
model.add(Embedding(10000, 8, input_length=maxlen))
# After the Embedding layer,
# our activations have shape `(samples, maxlen, 8)`.
# We flatten the 3D tensor of embeddings
# into a 2D tensor of shape `(samples, maxlen * 8)`
model.add(Flatten())
# We add the classifier on top
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.summary()
history = model.fit(x_train, y_train,
epochs=10,
batch_size=32,
validation_split=0.2)
# -
# We get to a validation accuracy of ~76%, which is pretty good considering that we only look at the first 20 words in every review. But
# note that merely flattening the embedded sequences and training a single `Dense` layer on top leads to a model that treats each word in the
# input sequence separately, without considering inter-word relationships and structure sentence (e.g. it would likely treat both _"this movie
# is shit"_ and _"this movie is the shit"_ as being negative "reviews"). It would be much better to add recurrent layers or 1D convolutional
# layers on top of the embedded sequences to learn features that take into account each sequence as a whole. That's what we will focus on in
# the next few sections.
# ## Using pre-trained word embeddings
#
#
# Sometimes, you have so little training data available that could never use your data alone to learn an appropriate task-specific embedding
# of your vocabulary. What to do then?
#
# Instead of learning word embeddings jointly with the problem you want to solve, you could be loading embedding vectors from a pre-computed
# embedding space known to be highly structured and to exhibit useful properties -- that captures generic aspects of language structure. The
# rationale behind using pre-trained word embeddings in natural language processing is very much the same as for using pre-trained convnets
# in image classification: we don't have enough data available to learn truly powerful features on our own, but we expect the features that
# we need to be fairly generic, i.e. common visual features or semantic features. In this case it makes sense to reuse features learned on a
# different problem.
#
# Such word embeddings are generally computed using word occurrence statistics (observations about what words co-occur in sentences or
# documents), using a variety of techniques, some involving neural networks, others not. The idea of a dense, low-dimensional embedding space
# for words, computed in an unsupervised way, was initially explored by Bengio et al. in the early 2000s, but it only started really taking
# off in research and industry applications after the release of one of the most famous and successful word embedding scheme: the Word2Vec
# algorithm, developed by Mikolov at Google in 2013. Word2Vec dimensions capture specific semantic properties, e.g. gender.
#
# There are various pre-computed databases of word embeddings that can download and start using in a Keras `Embedding` layer. Word2Vec is one
# of them. Another popular one is called "GloVe", developed by Stanford researchers in 2014. It stands for "Global Vectors for Word
# Representation", and it is an embedding technique based on factorizing a matrix of word co-occurrence statistics. Its developers have made
# available pre-computed embeddings for millions of English tokens, obtained from Wikipedia data or from Common Crawl data.
#
# Let's take a look at how you can get started using GloVe embeddings in a Keras model. The same method will of course be valid for Word2Vec
# embeddings or any other word embedding database that you can download. We will also use this example to refresh the text tokenization
# techniques we introduced a few paragraphs ago: we will start from raw text, and work our way up.
# ## Putting it all together: from raw text to word embeddings
#
#
# We will be using a model similar to the one we just went over -- embedding sentences in sequences of vectors, flattening them and training a
# `Dense` layer on top. But we will do it using pre-trained word embeddings, and instead of using the pre-tokenized IMDB data packaged in
# Keras, we will start from scratch, by downloading the original text data.
# ### Download the IMDB data as raw text
#
#
# First, head to `http://ai.stanford.edu/~amaas/data/sentiment/` and download the raw IMDB dataset (if the URL isn't working anymore, just
# Google "IMDB dataset"). Uncompress it.
#
# Now let's collect the individual training reviews into a list of strings, one string per review, and let's also collect the review labels
# (positive / negative) into a `labels` list:
# +
import os
imdb_dir = '/Users/liurui/develop/deeplearning/aclImdb'
train_dir = os.path.join(imdb_dir, 'train')
labels = []
texts = []
for label_type in ['neg', 'pos']:
dir_name = os.path.join(train_dir, label_type)
for fname in os.listdir(dir_name):
if fname[-4:] == '.txt':
f = open(os.path.join(dir_name, fname))
texts.append(f.read())
f.close()
if label_type == 'neg':
labels.append(0)
else:
labels.append(1)
# -
texts[0]
# ### Tokenize the data
#
#
# Let's vectorize the texts we collected, and prepare a training and validation split.
# We will merely be using the concepts we introduced earlier in this section.
#
# Because pre-trained word embeddings are meant to be particularly useful on problems where little training data is available (otherwise,
# task-specific embeddings are likely to outperform them), we will add the following twist: we restrict the training data to its first 200
# samples. So we will be learning to classify movie reviews after looking at just 200 examples...
#
# +
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import numpy as np
maxlen = 100 # We will cut reviews after 100 words
training_samples = 200 # We will be training on 200 samples
validation_samples = 10000 # We will be validating on 10000 samples
max_words = 10000 # We will only consider the top 10,000 words in the dataset
tokenizer = Tokenizer(num_words=max_words)
tokenizer.fit_on_texts(texts)
sequences = tokenizer.texts_to_sequences(texts)
word_index = tokenizer.word_index
print('Found %s unique tokens.' % len(word_index))
data = pad_sequences(sequences, maxlen=maxlen)
labels = np.asarray(labels)
print('Shape of data tensor:', data.shape)
print('Shape of label tensor:', labels.shape)
# Split the data into a training set and a validation set
# But first, shuffle the data, since we started from data
# where sample are ordered (all negative first, then all positive).
indices = np.arange(data.shape[0])
np.random.shuffle(indices)
data = data[indices]
labels = labels[indices]
x_train = data[:training_samples]
y_train = labels[:training_samples]
x_val = data[training_samples: training_samples + validation_samples]
y_val = labels[training_samples: training_samples + validation_samples]
# -
# ### Download the GloVe word embeddings
#
#
# Head to `https://nlp.stanford.edu/projects/glove/` (where you can learn more about the GloVe algorithm), and download the pre-computed
# embeddings from 2014 English Wikipedia. It's a 822MB zip file named `glove.6B.zip`, containing 100-dimensional embedding vectors for
# 400,000 words (or non-word tokens). Un-zip it.
# ### Pre-process the embeddings
#
#
# Let's parse the un-zipped file (it's a `txt` file) to build an index mapping words (as strings) to their vector representation (as number
# vectors).
# +
glove_dir = '/Users/liurui/develop/deeplearning/glove.6B'
embeddings_index = {}
f = open(os.path.join(glove_dir, 'glove.6B.100d.txt'))
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
# -
#
# Now let's build an embedding matrix that we will be able to load into an `Embedding` layer. It must be a matrix of shape `(max_words,
# embedding_dim)`, where each entry `i` contains the `embedding_dim`-dimensional vector for the word of index `i` in our reference word index
# (built during tokenization). Note that the index `0` is not supposed to stand for any word or token -- it's a placeholder.
# +
embedding_dim = 100
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
# Words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
# -
# ### Define a model
#
# We will be using the same model architecture as before:
# +
from keras.models import Sequential
from keras.layers import Embedding, Flatten, Dense
model = Sequential()
model.add(Embedding(max_words, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# -
# ### Load the GloVe embeddings in the model
#
#
# The `Embedding` layer has a single weight matrix: a 2D float matrix where each entry `i` is the word vector meant to be associated with
# index `i`. Simple enough. Let's just load the GloVe matrix we prepared into our `Embedding` layer, the first layer in our model:
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = False
#
# Additionally, we freeze the embedding layer (we set its `trainable` attribute to `False`), following the same rationale as what you are
# already familiar with in the context of pre-trained convnet features: when parts of a model are pre-trained (like our `Embedding` layer),
# and parts are randomly initialized (like our classifier), the pre-trained parts should not be updated during training to avoid forgetting
# what they already know. The large gradient update triggered by the randomly initialized layers would be very disruptive to the already
# learned features.
# ### Train and evaluate
#
# Let's compile our model and train it:
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(x_train, y_train,
epochs=10,
batch_size=32,
validation_data=(x_val, y_val))
model.save_weights('pre_trained_glove_model.h5')
# Let's plot its performance over time:
# +
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
#
# The model quickly starts overfitting, unsurprisingly given the small number of training samples. Validation accuracy has high variance for
# the same reason, but seems to reach high 50s.
#
# Note that your mileage may vary: since we have so few training samples, performance is heavily dependent on which exact 200 samples we
# picked, and we picked them at random. If it worked really poorly for you, try picking a different random set of 200 samples, just for the
# sake of the exercise (in real life you don't get to pick your training data).
#
# We can also try to train the same model without loading the pre-trained word embeddings and without freezing the embedding layer. In that
# case, we would be learning a task-specific embedding of our input tokens, which is generally more powerful than pre-trained word embeddings
# when lots of data is available. However, in our case, we have only 200 training samples. Let's try it:
# +
from keras.models import Sequential
from keras.layers import Embedding, Flatten, Dense
model = Sequential()
model.add(Embedding(max_words, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(x_train, y_train,
epochs=10,
batch_size=32,
validation_data=(x_val, y_val))
# +
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
#
# Validation accuracy stalls in the low 50s. So in our case, pre-trained word embeddings does outperform jointly learned embeddings. If you
# increase the number of training samples, this will quickly stop being the case -- try it as an exercise.
#
# Finally, let's evaluate the model on the test data. First, we will need to tokenize the test data:
# +
test_dir = os.path.join(imdb_dir, 'test')
labels = []
texts = []
for label_type in ['neg', 'pos']:
dir_name = os.path.join(test_dir, label_type)
for fname in sorted(os.listdir(dir_name)):
if fname[-4:] == '.txt':
f = open(os.path.join(dir_name, fname))
texts.append(f.read())
f.close()
if label_type == 'neg':
labels.append(0)
else:
labels.append(1)
sequences = tokenizer.texts_to_sequences(texts)
x_test = pad_sequences(sequences, maxlen=maxlen)
y_test = np.asarray(labels)
# -
# And let's load and evaluate the first model:
model.load_weights('pre_trained_glove_model.h5')
model.evaluate(x_test, y_test)
# We get an appalling test accuracy of 54%. Working with just a handful of training samples is hard!
| first_edition/6.1-using-word-embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center>Plot the figures in paper</center>
#
# ##### <center><font color='gray'>_<NAME>_, _2020-04-01_</font></center>
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.collections import PatchCollection
import numpy as np
import scipy.io as sio
# +
fontsize = 30
r_folder = 'results/'
d_folder = 'data/'
# the simulated 3D world
world = sio.loadmat(d_folder + 'world.mat')
# load the data of mesh sampled ZMs
fre_world_data = sio.loadmat(d_folder+'MeshSampled_ZMs_fixedHeading.mat')
# the max order of ZM
zm_n_max = 16
# -
# ## <font color='red'> visual homin (VH) </font>
# +
r_sub_folder = 'visual_homing/'
from visual_homing import VisualHomingAgent
# the route memory
route_memory = sio.loadmat(d_folder + 'ArcRouteMemory.mat')
# the home memory
home_memory = sio.loadmat(d_folder + 'HomeMemory_X0Y0.mat')
# set up parameters
num_pn = 81
num_kc = 4000
vh_learning_rate = 0.1
vh_kc_thr = 0.04
# create an instance
vh = VisualHomingAgent(world, route_memory, home_memory, zm_n_max, vh_learning_rate, vh_kc_thr, num_pn, num_kc)
# training
en = vh.train_mb_network()
# -
# ### <font color='black'> _visual novelty heatmap_ </font>
# +
# calculate the map
map_x = fre_world_data['x'][0]
map_y = fre_world_data['y'][0]
map_ZM_As_random = fre_world_data['A']
e_map = np.zeros([len(map_x), len(map_y)])
for xi in range(len(map_x)):
for yi in range(len(map_y)):
e_map[xi, yi] = vh.mb.run(map_ZM_As_random[xi, yi, :vh.zm_coeff_num])
# draw the map
fig,ax = plt.subplots(figsize=(8,8))
ax.pcolormesh(map_x, np.flipud(map_x), np.rot90(e_map), cmap='RdPu')
ax.set_aspect(1)
ax.set_axis_off()
ax.set_xlim(-10,5)
ax.set_ylim(-10,5);
# plt.savefig('MB_Familiar_Map_l7.png')
# -
# ### <font color='black'> _visual homing results_ </font>
# +
fig,ax = plt.subplots(figsize=(8,8))
# get the data generated from the simulation
data = sio.loadmat(r_folder + r_sub_folder + 'VH_ONLY.mat')
pos = data['pos']
h = data['h']
# the index of the homing route to plot
plot_ind = [8]
# the indexes of the agents to caculate the initial heading
caculate_ind = [0,1,2,3,5,6,8,10,11]
# plot the homing route
for num,i in enumerate(plot_ind):
if num == 0:
ax.plot(pos[i][:,0]/100,pos[i][:,1]/100,color='r',lw=1.5,alpha = (num+1)*(1/len(plot_ind)),label='Homing Routes')
ax.plot(pos[i][:,0]/100,pos[i][:,1]/100,color='r',lw=1.5,alpha = (num+1)*(1/len(plot_ind)))
ax.quiver(pos[i][0,0]/100,pos[i][0,1]/100,np.cos(h[i][0]),np.sin(h[i][0]),
color='r',scale=15, width=0.004, alpha = (num+1)*(1/len(plot_ind)))
# analysis the initial headings
import circular_statistics as cs
# simulated ant
cp_ind = cs.get_check_points_index(250, pos)
cp_h = [h[i,cp_ind[i]] for i in caculate_ind]
acc = 360 # 5deg goniometer
d_ref = np.linspace(0,2*np.pi,acc)
h_r, h_mean, h_sd0, h_ci95, r_num = cs.circular_statistics(cp_h,acc=acc)
# add the polar plot
insert_ax = fig.add_axes([0.2,0.6,0.2,0.2],projection='polar')
insert_ax.quiver(0,0,h_r*np.cos(h_mean),h_r*np.sin(h_mean),color='r', scale=0.5,scale_units='y',width=0.025,label='Model')
insert_ax.plot(np.linspace(h_mean-h_ci95,h_mean+h_ci95,20),np.repeat(h_r*4,20),color='r',lw=5)
insert_ax.scatter(h_mean-h_ci95,h_r*4,s=50,marker='.',color='r',edgecolor='r')
insert_ax.scatter(h_mean+h_ci95,h_r*4,s=50,marker='.',color='r',edgecolor='r')
# real ant
real_ant_data = sio.loadmat(r_folder + r_sub_folder + 'Wystrach2012_OnOffRoute.mat')
real_cp_h = real_ant_data['head_N1_RX_P'][0]
h_r, h_mean, h_sd0, h_ci95, r_num = cs.circular_statistics(real_cp_h,acc=acc)
insert_ax.quiver(0,0,h_r*np.cos(h_mean),h_r*np.sin(h_mean),color='k', scale=0.5,scale_units='y',width=0.025,label='Real Ant')
insert_ax.plot(np.linspace(h_mean-h_ci95,h_mean+h_ci95,20),np.repeat(h_r*4,20),color='k',lw=5)
insert_ax.scatter(h_mean-h_ci95,h_r*4,s=50,marker='.',color='k',edgecolor='k')
insert_ax.scatter(h_mean+h_ci95,h_r*4,s=50,marker='.',color='k',edgecolor='k')
# habitual route
ax.plot(route_memory['pos'][:,0]/100,route_memory['pos'][:,1]/100,color='gray',lw=10,alpha=0.6,label='Habitual Route')
interval = 4
ax.quiver(route_memory['pos'][::interval,0]/100,route_memory['pos'][::interval,1]/100,
np.cos(route_memory['h'][0][::interval]),np.sin(route_memory['h'][0][::interval]),
width=0.005,scale=30,color='k')
# RP
ax.text((pos[i][0,0]+20)/100,(pos[i][0,1]-20)/100,'RP',fontsize=fontsize,color='k')
ax.scatter(pos[i][0,0]/100,pos[i][0,1]/100, color='k',s=100)
# Nest
ax.scatter(0,0,color='r',marker='*',s=300)
ax.text(0.2,-0.2,'Nest',fontsize=fontsize,color='red')
# Feeder
ax.scatter(-7,-7,color='k',marker='s',s=300)
ax.text(-7.2,-8,'Feeder',fontsize=30,color='k')
# format adjust
insert_ax.set_yticklabels('')
ticks = insert_ax.get_yticklabels()
[tick.set_fontsize(fontsize-5) for tick in ticks]
ticks = insert_ax.get_xticklabels()
[tick.set_fontsize(fontsize-5) for tick in ticks]
insert_ax.legend(fontsize=fontsize-8)
ticks = ax.get_yticklabels()
[tick.set_fontsize(fontsize) for tick in ticks]
ticks = ax.get_xticklabels()
[tick.set_fontsize(fontsize) for tick in ticks]
ax.set_title('Visual Homing',color='k',fontsize=fontsize)
ax.set_xlabel('X / m',fontsize=fontsize)
ax.set_ylabel('Y / m',fontsize=fontsize)
ax.legend(fontsize=fontsize-5)
ax.set_xlim(-10,5)
ax.set_ylim(-10,5)
ax.grid(1)
ax.set_aspect(1)
# plt.savefig('viusal_homing_results.pdf')
# -
# ### <font color='gray'> optimal integration </font>
r_sub_folder = 'optimal_integration/'
# ### <font color='black'> _results of tuning PI uncertainty_ </font>
# +
# initial heading analysis of the agents
import circular_statistics as cs
PI_l = [10,100,300,700]
pos = []
h = []
# load data
for i in range(4):
data = sio.loadmat(r_folder + r_sub_folder + 'PI' + str(PI_l[i]) + '_VH_SP5.mat')
pos.append(data['pos'])
h.append(data['h'])
# calculate the initial headings
ck_ind = []
ck_h = []
for j in range(4):
ck_ind.append(cs.get_check_points_index(60,pos[j]))
ck_ind[j] = list(filter(lambda x: x!=0, ck_ind[j]))
ck_h.append([h[j][i][ck_ind[j][i]] for i in range(len(ck_ind[j]))])
# calculate the circular statistics
r = []
mean = []
ci_95 = []
num = []
for i in range(4):
r_i, mean_i, sd0_i, ci95_i, num_i = cs.circular_statistics(ck_h[i],acc=100)
r.append(r_i)
mean.append(mean_i)
ci_95.append(ci95_i)
num.append(num_i)
# plot the circular data
fig = plt.figure(figsize=(6,6))
fontsize=30
ax = fig.add_subplot(111,projection='polar')
color_list = ['gray','gray','gray','gray']
for i in range(4):
ax.quiver(0,0,r[i]*np.cos(mean[i]),r[i]*np.sin(mean[i]),color=color_list[i], width=0.012, scale=0.5,scale_units='y')
ax.plot(np.linspace(mean[i]-ci_95[i],mean[i]+ci_95[i],20),np.repeat(4 * r[i],20),color=color_list[i],lw=4)
ax.scatter(mean[i]-ci_95[i],4 * r[i],s=220,marker='.',color='gray',edgecolor=color_list[i])
ax.scatter(mean[i]+ci_95[i],4 * r[i],s=220,marker='.',color='gray',edgecolor=color_list[i])
ax.text(mean[i]+ci_95[i],4 * r[i],'%sm'%(PI_l[i]/100),fontsize=24,color='orange')
# the PI and Visual Diretion
pi_d = 3*np.pi/2
pi_v = np.deg2rad(140)
tri_len = 0.8
tri_pos_l = 4.9
tri_pi = plt.Polygon([[pi_d,tri_pos_l],
[pi_d+np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)],
[pi_d-np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)]],
color='orange')
ax.add_patch(tri_pi)
tri_pi = plt.Polygon([[pi_v,tri_pos_l],
[pi_v+np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)],
[pi_v-np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)]],
color='r')
ax.add_patch(tri_pi)
ax.set_ylim(0,5)
ax.set_yticklabels('')
ticks = ax.get_xticklabels()
[tick.set_fontsize(fontsize-5) for tick in ticks];
# plt.savefig('optimal_tune_pi.pdf')
# -
# ### <font color='black'> _results of tuning VH uncertainty_ </font>
# +
# combining the tuning visual experiments
tune_visual_r = [0,0,0]
tune_visual_mean = [0,0,0]
tune_visual_ci95 = [0,0,0]
tune_visual_num = [0,0,0]
tune_visual_vf = [0,0,0]
tune_visual_dis = [0,0,0]
# RP1
data = sio.loadmat(r_folder + r_sub_folder + 'PI100_VH_SP5.mat')
pos = data['pos']
h = data['h']
tune_visual_dis[0] = np.sqrt(np.sum(pos[0,0]**2 + pos[0,1]**2))/100.0
ck_h = cs.get_check_points_h(60,pos,h)
tune_visual_r[0], tune_visual_mean[0], x, tune_visual_ci95[0], tune_visual_num[0] = cs.circular_statistics(ck_h,acc=100)
# RP2
data = sio.loadmat(r_folder + r_sub_folder + 'PI_VH_SP15.mat')
pos = data['pos']
h = data['h']
tune_visual_dis[1] = np.sqrt(np.sum(pos[0,0]**2 + pos[0,1]**2))/100.0
ck_h = cs.get_check_points_h(30,pos,h)
tune_visual_r[1], tune_visual_mean[1], x, tune_visual_ci95[1], tune_visual_num[1] = cs.circular_statistics(ck_h,acc=100)
# RP3
data = sio.loadmat(r_folder + r_sub_folder + 'PI_VH_SP25.mat')
pos = data['pos']
h = data['h']
tune_visual_dis[2] = np.sqrt(np.sum(pos[0,0]**2 + pos[0,1]**2))/100.0
ck_h = cs.get_check_points_h(30,pos,h)
tune_visual_r[2], tune_visual_mean[2], x, tune_visual_ci95[2], tune_visual_num[2] = cs.circular_statistics(ck_h,acc=100)
# plot the data
fig = plt.figure(figsize=(6,6))
fontsize=30
ax = fig.add_subplot(111,projection='polar')
for i in range(3):
ax.quiver(0,0,tune_visual_r[i]*np.cos(tune_visual_mean[i]),tune_visual_r[i]*np.sin(tune_visual_mean[i]),color='gray',
width=0.012, scale=0.5,scale_units='y')
ax.plot(np.linspace(tune_visual_mean[i]-tune_visual_ci95[i],tune_visual_mean[i]+tune_visual_ci95[i],20),
np.repeat(4 * tune_visual_r[i],20),color='gray',lw=4)
ax.scatter(tune_visual_mean[i]-tune_visual_ci95[i],4 * tune_visual_r[i],s=220,marker='.',color='gray',edgecolor='gray')
ax.scatter(tune_visual_mean[i]+tune_visual_ci95[i],4 * tune_visual_r[i],s=220,marker='.',color='gray',edgecolor='gray')
ax.text(tune_visual_mean[i]+tune_visual_ci95[i],4 * tune_visual_r[i],'%.4sm'%(tune_visual_dis[i]),fontsize=24,color='r')
pi_d = 3*np.pi/2
pi_v = np.deg2rad(140)
tri_len = 0.6
tri_pos_l = 2.9
tri_pi = plt.Polygon([[pi_d,tri_pos_l],
[pi_d+np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)],
[pi_d-np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)]],
color='orange')
ax.add_patch(tri_pi)
tri_pi = plt.Polygon([[pi_v,tri_pos_l],
[pi_v+np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)],
[pi_v-np.arctan2((tri_len/2),tri_pos_l-tri_len*np.cos(np.pi/6)),np.sqrt((tri_len/2)**2+(tri_pos_l-tri_len*np.cos(np.pi/6))**2)]],
color='r')
ax.add_patch(tri_pi)
ax.set_ylim(0,3)
ax.set_yticklabels('')
ticks = ax.get_xticklabels()
[tick.set_fontsize(fontsize-5) for tick in ticks];
# plt.savefig('optimal_tune_v.pdf')
# -
# ### <font color='black'> _typical homing path_ </font>
# +
# pi_route
pi_route_mem_data = sio.loadmat(d_folder + 'PiRouteMemory.mat')
fontsize = 30
fig,ax = plt.subplots(figsize=(6,8))
# pi
data = sio.loadmat(r_folder + r_sub_folder + 'PI_Route.mat')
homing_pos = data['pos'][:500]
ax.plot(homing_pos[:,0]/100,homing_pos[:,1]/100,color='orange',lw=2,label='PI')
# vh
data = sio.loadmat(r_folder + r_sub_folder + 'VH_Route.mat')
homing_pos = data['pos']
ax.plot(homing_pos[:,0]/100,homing_pos[:,1]/100,color='r',lw=2,label='VH')
# pi+vh
data = sio.loadmat(r_folder + r_sub_folder + 'PI_VH_Route.mat')
homing_pos = data['pos']
ax.plot(homing_pos[:,0]/100,homing_pos[:,1]/100,color='gray',lw=2,label='PI+VH')
ax.plot(pi_route_mem_data['pos'][:,0]/100,pi_route_mem_data['pos'][:,1]/100,color='gray',lw=10,alpha=0.6)
interval = 4
ax.quiver(pi_route_mem_data['pos'][::interval,0]/100,pi_route_mem_data['pos'][::interval,1]/100,
np.cos(pi_route_mem_data['h'][0][::interval]),np.sin(pi_route_mem_data['h'][0][::interval]),
width=0.01,scale=20,color='k')
# nest
ax.scatter(0,0,color='r',marker='*',s=300)
ax.text(0.2,-0.2,'Nest',fontsize=fontsize-10,color='red')
# feeder
ax.scatter(pi_route_mem_data['pos'][-1,0]/100,pi_route_mem_data['pos'][-1,1]/100,color='k',marker='s',s=300)
ax.text(pi_route_mem_data['pos'][-1,0]/100+0.2,pi_route_mem_data['pos'][-1,1]/100+0.2,'Feeder',fontsize=fontsize-10,color='k')
ax.scatter(homing_pos[0,0]/100,homing_pos[0,1]/100,color='k',marker='.',s=200)
ax.text(homing_pos[0,0]/100,homing_pos[0,1]/100,'RP',fontsize=fontsize-5)
ax.set_xlim(-5,4)
# ax.set_ylim(-3,13)
ax.set_xlabel('X / m',fontsize=fontsize-15)
ax.set_ylabel('Y / m',fontsize=fontsize-15)
plt.xticks(fontsize=fontsize-15)
plt.yticks(fontsize=fontsize-15)
ax.grid(1)
ax.set_aspect(1)
ax.legend(fontsize=fontsize-10)
# plt.savefig('pi_vh_route.pdf')
# -
# ### <font color='black'> _extented homing path: supplementary figure_ </font>
# plot the extended path of the PI+VH - varies PI length
fig,ax = plt.subplots(1,4,figsize=(24,6))
PI_length = [10,100,300,700]
for i in range(4):
data = sio.loadmat(r_folder + r_sub_folder + 'PI' + str(PI_l[i]) + '_VH_SP5.mat')
pos = data['pos']
h = data['h']
ax[i].scatter(pos[0,0,0]/100,pos[0,0,1]/100,color='k',marker='s',s=100)
ax[i].scatter(0,0,color='r',marker='*',s=300)
for j in range(len(pos)):
ax[i].plot(pos[j,:,0]/100,pos[j,:,1]/100,color='k',lw=0.5)
ax[i].set_xlim([-1,2.5])
ax[i].set_ylim([-2.5,1])
ax[i].set_aspect(1)
ax[i].grid(1)
ax[i].set_xlabel('X / m',fontsize=fontsize-10)
ax[i].set_ylabel('Y / m',fontsize=fontsize-10)
ax[i].set_xticks([-1,0,1,2])
ax[i].set_yticks([-2,-1,0,1])
ax[i].set_title('home vector length = %sm'%(PI_length[i]/100),fontsize=fontsize-10)
ticks = ax[i].get_yticklabels()
[tick.set_fontsize(fontsize-10) for tick in ticks]
ticks = ax[i].get_xticklabels()
[tick.set_fontsize(fontsize-10) for tick in ticks]
# plt.savefig('supp_optimal_path1.pdf')
# +
# plot the extended path of the PI+VH - varies VISUAL
data_rp1 = sio.loadmat(r_folder + r_sub_folder + 'PI100_VH_SP5.mat')
pos_rp1 = data_rp1['pos']
h_rp1 = data_rp1['h']
data_rp2 = sio.loadmat(r_folder + r_sub_folder + 'PI_VH_SP15.mat')
pos_rp2 = data_rp2['pos']
h_rp2 = data_rp2['h']
data_rp3 = sio.loadmat(r_folder + r_sub_folder + 'PI_VH_SP25.mat')
pos_rp3 = data_rp3['pos']
h_rp3 = data_rp3['h']
fig,ax = plt.subplots(1,2,figsize=(12,6))
for i in range(len(h_rp2)):
ax[0].plot(pos_rp2[i,:,0]/100,pos_rp2[i,:,1]/100,color='k',lw=0.5)
ax[1].plot(pos_rp3[i,:,0]/100,pos_rp3[i,:,1]/100,color='k',lw=0.5)
ax[0].scatter(pos_rp2[0,0,0]/100,pos_rp2[0,0,1]/100,color='k',marker='s',s=100)
ax[1].scatter(pos_rp3[0,0,0]/100,pos_rp3[0,0,1]/100,color='k',marker='s',s=100)
rp_dis = [6.18,10.3]
for i in [0,1]:
ax[i].scatter(0,0,color='r',marker='*',s=300)
ax[i].set_title('RP distance = %2.4sm'%(rp_dis[i]),fontsize=fontsize-10)
ax[i].set_xlim([-1,10])
ax[i].set_ylim([-6,1])
ax[i].set_aspect(1)
ax[i].grid(1)
ax[i].set_xlabel('X / m',fontsize=fontsize-10)
ax[i].set_ylabel('Y / m',fontsize=fontsize-10)
ticks = ax[i].get_yticklabels()
[tick.set_fontsize(fontsize-10) for tick in ticks]
ticks = ax[i].get_xticklabels()
[tick.set_fontsize(fontsize-10) for tick in ticks]
# plt.savefig('supp_optimal_path2.pdf')
# -
# ## <font color='blue'> route following (RF) </font>
# +
r_sub_folder = 'route_following/'
# the route memory
route_memory = sio.loadmat(d_folder + 'ArcRouteMemory.mat')
# the home memory
home_memory = sio.loadmat(d_folder + 'HomeMemory_X0Y0.mat')
# -
# ### <font color='black'> _ZM phase tracking performances_ </font>
# #### <font color='black'> _single point ZM tracking_ </font>
from image_processing import visual_sense
# bad(-2.5,-3.5) /good(-8,2) tracking
h_s = [np.arange(s_h,s_h+np.pi*2,19/180*np.pi) for s_h in [0, np.pi/2, np.pi, 3*np.pi/2.0]]
zm_a_s = []
zm_p_s = []
for h_ii in h_s:
for h_i in h_ii:
# good tracking
# zm_a, zm_p = visual_sense(world, -8, 2, h_i, nmax=zm_n_max)
# bad tracking
zm_a, zm_p = visual_sense(world, -2.5, -3.5, h_i, nmax=zm_n_max)
zm_a_s.append(zm_a)
zm_p_s.append(zm_p)
# plot bad(-2.5,-3.5) / good(1.5,-3) tracking
fig,ax = plt.subplots(figsize=(8,4))
ax.set_xlabel('Heading / rad',fontsize=30)
ax.set_ylabel('ZM-phase / rad',fontsize=30)
zm_phase = [zm_p_s[i][16] for i in range(len(zm_p_s))]
ax.set_ylim(-np.pi,np.pi)
ax.set_xlim(-np.pi,np.pi)
for i in range(2,3):
perfect_h = zm_phase[i*19]/180*np.pi - h_s[i]
perfect_h = ((perfect_h) + np.pi)%(np.pi*2.0) - np.pi
ax.scatter((h_s[i] + np.pi) % (2.0 * np.pi)-np.pi,perfect_h,color='r',label='Animal orientation',s=60,alpha=0.8)
ax.plot((h_s[i] + np.pi) % (2.0 * np.pi)-np.pi,perfect_h,color='r')
ax.scatter((h_s[i] + np.pi) % (2.0 * np.pi) - np.pi,np.array(zm_phase[i*19:(i+1)*19])/180*np.pi,color='b',
label='ZM-Phase tracking',s=60,alpha=0.8)
ax.plot((h_s[i] + np.pi) % (2.0 * np.pi) - np.pi,np.array(zm_phase[i*19:(i+1)*19])/180*np.pi,color='b')
ax.grid(1)
ax.legend(fontsize=20)
plt.xticks(fontsize=30)
plt.yticks(fontsize=30);
# plt.savefig('zm_p_tracking_bad.pdf')
# plt.savefig('zm_p_tracking_good.pdf')
# #### <font color='black'> _cross world ZM tracking_ </font>
# +
# load data from the simulation
rf_data = sio.loadmat(r_folder + r_sub_folder + 'QuiverPlotData_X-10_2_Y-8_2_SH20.mat')
# sampled num in x-y direction
sample_num = 20
# sampled locations
pos_x = np.linspace(-10,2,sample_num)
pos_y = np.linspace(-8,2,sample_num)
# sampled headings
h = np.linspace(-np.pi,np.pi,10)
# check the data for RF
# 1.RF memory , 2.the phase-tracking, 3.RF suggested
# current zm p
current_zm_p = rf_data['current_zm_p']
current_zm_p_rad = current_zm_p/180.0 *np.pi
diff_zm_p = (np.diff(current_zm_p_rad) + np.pi) %(np.pi*2) - np.pi
for p in diff_zm_p:
p[np.where(p>0)] = 0
zm_p_ratio = diff_zm_p/(h[1]-h[0])
# inverse the ratio<-1.0
for p in zm_p_ratio:
p[np.where(p<-1.0)] = 1.0/p[np.where(p<-1.0)]
zm_p_ratio_mean = np.mean(zm_p_ratio,axis=1)
import circular_statistics as cs
# ann output
ann_output = rf_data['ann_output']
ann_mean = list(map(cs.circular_statistics, ann_output))
ann_cir_statistic = {'r':[ann_mean[i][0] for i in range(sample_num**2)],'mean':[ann_mean[i][1] for i in range(sample_num**2)],
'ci95':[ann_mean[i][3] for i in range(sample_num**2)]}
# RF angle
rf_angle = list(map(cs.calculate_rf_motor, ann_output, np.tile(h,[sample_num**2,1]), current_zm_p_rad))
rf_mean = list(map(cs.circular_statistics,rf_angle))
rf_cir_statistic = {'r':[rf_mean[i][0] for i in range(sample_num**2)],'mean':[rf_mean[i][1] for i in range(sample_num**2)],
'ci95':[rf_mean[i][3] for i in range(sample_num**2)]}
# +
# plot
fig,ax = plt.subplots(figsize=(8,10))
h_plot_s = (np.array(rf_cir_statistic['mean']) - np.array(rf_cir_statistic['ci95'])).reshape(sample_num,sample_num) + np.pi
h_plot_e = (np.array(rf_cir_statistic['mean']) + np.array(rf_cir_statistic['ci95'])).reshape(sample_num,sample_num) + np.pi
h_plot_s = np.rad2deg(h_plot_s)
h_plot_e = np.rad2deg(h_plot_e)
wedges = []
for i in range(sample_num):
for j in range(sample_num):
wedge = mpatches.Wedge([pos_x[j],pos_y[i]], 0.3, h_plot_s[i,j], h_plot_e[i,j], ec="none")
wedges.append(wedge)
collection = PatchCollection(wedges, color='blue', alpha=0.3)
ax.add_collection(collection)
h_plot = np.array(rf_cir_statistic['mean']).reshape(sample_num,sample_num) + np.pi
r_plot = np.array(rf_cir_statistic['r']).reshape(sample_num,sample_num)
# sampled positions
X,Y = np.meshgrid(pos_x,pos_y)
ax.scatter(X,Y,color='gray',marker='o',s=10,alpha=0.5)
ax.set_title('Route Following',fontsize=30)
plt.xticks(fontsize=30)
plt.yticks(fontsize=30)
ax.set_aspect(1)
# plt.savefig('Whole_RouteFollowingResults.pdf')
# -
# ### <font color='black'> _results and homing path_ </font>
# +
fig,ax=plt.subplots(figsize=(12,12))
insert_ax = fig.add_axes([0.2,0.6,0.2,0.2],projection='polar')
data = sio.loadmat(r_folder + r_sub_folder + 'RF_ONLY.mat')
pos = data['pos']
h = data['h']
# plot will end at the points nearest to the nest for the all homing path
dis = [pos[i][:,0]**2 + pos[i][:,1]**2 for i in range(len(pos))]
end_t = [np.argmin(dis[i])+1 for i in range(len(pos))]
# plot homing route
for i in range(len(pos)):
# ax.scatter(InsectNaviAgent.homing_pos[mask][:,0]/100,InsectNaviAgent.homing_pos[mask][:,1]/100,c='red')
ax.plot(pos[i][:end_t[i]-1,0]/100,pos[i][:end_t[i]-1,1]/100,lw=1.5,color='blue')
ax.scatter(pos[i][0,0]/100,pos[i][0,1]/100,marker='o',color='k',s=100,alpha=0.5)
# initial heading analysis of the agents released on route
import circular_statistics as cs
## simualted ant
data = sio.loadmat(r_folder + r_sub_folder + 'RF_released_on_route.mat')
pos = data['pos']
h = data['h']
ck_h = cs.get_check_points_h(60,pos,h)
r_i, mean_i, sd0_i, ci95_i, num_i = cs.circular_statistics(ck_h,acc=100)
insert_ax.quiver(0,0,r_i*np.cos(mean_i),r_i*np.sin(mean_i),color='blue', width=0.012, scale=0.5,scale_units='y',label='Model')
insert_ax.plot(np.linspace(mean_i-ci95_i,mean_i+ci95_i,20),np.repeat(4 * r_i,20),color='blue',lw=5,alpha=0.6)
insert_ax.scatter(mean_i-ci95_i,4 * r_i,s=50,marker='.',color='blue',edgecolor='blue',alpha=0.6)
insert_ax.scatter(mean_i+ci95_i,4 * r_i,s=50,marker='.',color='blue',edgecolor='blue',alpha=0.6)
## real ant
real_ant_data = sio.loadmat(r_folder + r_sub_folder + 'Wystrach2012_OnOffRoute.mat')
real_cp_h = real_ant_data['head_N1_RX_R'][0]
h_r, h_mean, h_sd0, h_ci95, r_num = cs.circular_statistics(real_cp_h,acc=acc)
insert_ax.quiver(0,0,h_r*np.cos(h_mean),h_r*np.sin(h_mean),color='r', scale=0.5,scale_units='y',width=0.025,label='Real Ant')
insert_ax.plot(np.linspace(h_mean-h_ci95,h_mean+h_ci95,20),np.repeat(h_r*4,20),color='r',lw=5)
insert_ax.scatter(h_mean-h_ci95,h_r*4,s=50,marker='.',color='r',edgecolor='r')
insert_ax.scatter(h_mean+h_ci95,h_r*4,s=50,marker='.',color='r',edgecolor='r')
# habitual route
ax.plot(route_memory['pos'][:,0]/100,route_memory['pos'][:,1]/100,color='gray',lw=10,alpha=0.6)
interval = 4
ax.quiver(route_memory['pos'][::interval,0]/100,route_memory['pos'][::interval,1]/100,
np.cos(route_memory['h'][0][::interval]),np.sin(route_memory['h'][0][::interval]),
width=0.005,scale=30,color='k')
# nest
ax.scatter(0,0,color='r',marker='*',s=300)
ax.text(0.5,-0.2,'Nest',fontsize=30,color='red')
# RPs
ax.text(-8,-7.5,'Release pionts',fontsize=30,color='k')
plt.xticks(fontsize=30)
plt.yticks(fontsize=30)
# format adjust
insert_ax.set_yticklabels('')
ticks = insert_ax.get_yticklabels()
[tick.set_fontsize(fontsize-5) for tick in ticks]
ticks = insert_ax.get_xticklabels()
[tick.set_fontsize(fontsize-5) for tick in ticks]
insert_ax.legend(fontsize=fontsize-8)
ax.grid(1)
ax.set_xlabel('X / m',fontsize=30)
ax.set_ylabel('Y / m',fontsize=30)
ax.set_aspect(1)
# plt.savefig('route_following_results.pdf')
# -
# ## <font color='black'> whole model </font>
r_sub_folder = 'whole_model/'
# ### <font color='black'> _homing path of ZV and FV_ </font>
# +
# homing path of the whole model
route_mem_data = sio.loadmat(d_folder + 'ArcRouteMemory.mat')
fig,ax = plt.subplots(figsize=(8,10))
# nest
ax.scatter(route_mem_data['pos'][-1,0]/100,route_mem_data['pos'][-1,1]/100,color='r',marker='*',s=300)
ax.text(route_mem_data['pos'][-1,0]/100,route_mem_data['pos'][-1,1]/100,'Nest',fontsize=fontsize-10,color='red')
# feeder
ax.scatter(route_mem_data['pos'][1,0]/100,route_mem_data['pos'][1,1]/100,color='k',marker='s',s=300)
ax.text(route_mem_data['pos'][1,0]/100+0.2,route_mem_data['pos'][1,1]/100+0.2,'Feeder',fontsize=fontsize-10,color='k')
# route
ax.plot(route_mem_data['pos'][:,0]/100,route_mem_data['pos'][:,1]/100,color='gray',lw=10,alpha=0.6)
interval = 4
ax.quiver(route_mem_data['pos'][::interval,0]/100,route_mem_data['pos'][::interval,1]/100,
np.cos(route_mem_data['h'][0][::interval]),np.sin(route_mem_data['h'][0][::interval]),
width=0.01,scale=25,color='k',alpha=0.5)
# ZV
data = sio.loadmat(r_folder + r_sub_folder + 'final_ZV.mat')
homing_pos = data['homing_pos'][:180]
ax.plot(homing_pos[:,0]/100,homing_pos[:,1]/100,color='k',lw=1.5,ls='--',label='ZV')
ax.scatter(homing_pos[96,0]/100,homing_pos[96,1]/100,color='orange',marker='D',s=100)
# FV
data = sio.loadmat(r_folder + r_sub_folder + 'final_FV.mat')
homing_pos = data['homing_pos']
ax.plot(homing_pos[:,0]/100,homing_pos[:,1]/100,color='k',lw=1.5,ls='-',label='FV')
ax.scatter(homing_pos[78,0]/100,homing_pos[78,1]/100,color='orange',marker='D',s=100)
# RP
ax.scatter(homing_pos[0,0]/100,homing_pos[0,1]/100,color='k',marker='.',s=200)
ax.text(homing_pos[0,0]/100,homing_pos[0,1]/100,'RP',fontsize=fontsize-5)
ax.set_xlim(-8,3)
ax.set_ylim(-8,2)
ax.set_xlabel('X / m',fontsize=fontsize-15)
ax.set_ylabel('Y / m',fontsize=fontsize-15)
plt.xticks(fontsize=fontsize-15)
plt.yticks(fontsize=fontsize-15)
ax.grid(1)
ax.set_aspect(1)
ax.legend(fontsize=fontsize-15)
# plt.savefig('whole_homing_path.pdf')
# -
# ### <font color='black'> _activation of TUN, SN1 and SN2_ </font>
# +
# ZV
# data = sio.loadmat(r_folder + r_sub_folder + 'final_ZV.mat')
# FV
data = sio.loadmat(r_folder + r_sub_folder + 'final_FV.mat')
fig,ax = plt.subplots(3,1,figsize=(8,6))
end_t = 160
SN1 = data['homing_state'][0][:end_t]
# SN1
ax[0].plot(SN1, c='r',lw=2)
# ax[0].set_xticks([])
ax[0].text(10,0.75,'SN1',fontsize=fontsize-10,color='r')
# SN2
ax[1].plot(1-SN1,c='cyan',lw=2)
# ax[1].set_xticks([])
ax[1].text(10,0.75,'SN2',fontsize=fontsize-10,color='cyan')
# TUN
ax[2].plot(data['homing_vh_sensory'][0][:end_t] / 80, c='blue',lw=2)
ax[2].text(10,0.75,'TUN',fontsize=fontsize-10,color='blue')
ax[2].set_xlabel('Time step',fontsize=fontsize)
ax[2].set_ylabel('Activation',fontsize=fontsize)
for a in ax:
# dimond marker
a.scatter(78,1,c='orange',s=100,marker='D')
a.set_xlim([0,end_t+1])
a.set_ylim([-0.1,1.1])
a.grid(1)
ticks = a.get_yticklabels()
[tick.set_fontsize(fontsize-10) for tick in ticks]
ticks = a.get_xticklabels()
[tick.set_fontsize(fontsize-10) for tick in ticks]
# plt.savefig('SN1_SN2_TUN_ZV.pdf')
| demo/figs_in_paper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# !pip install shap adjustText xgboost
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import gc
import math
from utilities.data import *
from sklearn.preprocessing import MinMaxScaler
from sklearn.cluster import DBSCAN
loader = DataLoader(use_data_pt2=True)
#grenade_df = loader.load_grenade_df(nrows=20000)
#print(grenade_df.info())
#print(grenade_df.isnull().sum())
#kill_df = loader.load_kill_df()
#print(kills_df.info())
#dmg_df = loader.load_dmg_df(nrows=20000)
#dmg_df = dmg_df[(~dmg_df['seconds'].isna())]
#print(dmg_df.info())
#print(dmg_df.isnull().sum())
results_df = loader.load_full_df(map_name='de_mirage')
print(results_df.info())
results_df = results_df[results_df['att_side'] != 'None']
results_df = results_df[results_df['seconds'] != 'None']
print(results_df.isna().sum())
# -
# Cluster spatially
from sklearn.cluster import DBSCAN
from collections import defaultdict
min_samples = 80
print(f"min_samples: {min_samples}")
unsupervised_models = {'DMG': DBSCAN(eps=0.003, min_samples=120),
'Smoke': DBSCAN(eps=0.004, min_samples=min_samples),
'Flash': DBSCAN(eps=0.0036, min_samples=100),
'HE': DBSCAN(eps=0.004, min_samples=min_samples),
'Incendiary': DBSCAN(eps=0.004, min_samples=min_samples),
'Molotov': DBSCAN(eps=0.004, min_samples=min_samples)}
manual_models = defaultdict(lambda: MirageCalloutClusteringModel)
results_df = cluster_positions(results_df, unsupervised_models, verbose=True, scale=False)
print(results_df.isna().sum())
for team in results_df['att_side'].unique():
for pos_type in results_df['pos_type'].unique():
mask = (results_df['att_side'] == team)
if pos_type == 'DMG_VIC':
continue
elif pos_type == 'DMG_ATT':
mask = mask & ((results_df['pos_type'] == 'DMG_VIC') | (results_df['pos_type'] == 'DMG_ATT'))
else:
mask = mask & (results_df['pos_type'] == pos_type)
clusters = results_df['pos_cluster'][mask]
clusters.hist()
plt.show()
# Do one-hot encoding for each team seperately and each type seperately (except DMG_VIC and DMG_ATT, which are done together)
pos_types = [t for t in results_df['pos_type'].unique() if t not in ['DMG_VIC', 'DMG_ATT']]
dummy_cols = []
for pos_type in pos_types:
mask = (results_df['pos_type'] == pos_type)
group = results_df[mask]
# Differentiate T and CT cluster by adding max to all T
max_ct_pos_cluster = group.loc[(group['att_side'] == 'CounterTerrorist'), 'pos_cluster'].max()
group.loc[(group['att_side'] == "Terrorist"), 'pos_cluster'] += max_ct_pos_cluster*2
dummies = pd.get_dummies(group['pos_cluster'])
print(f"REAL COLS {dummies.columns}")
dummies.columns = [f"CT_{pos_type}_{c}" if c <= max_ct_pos_cluster else f"T_{pos_type}_{c - max_ct_pos_cluster*2}" for c in dummies.columns]
dummy_cols.extend(dummies.columns)
for c in dummies.columns:
results_df[c] = 0
results_df.loc[mask, dummies.columns] = dummies
mask = (results_df['pos_type'] == 'DMG_ATT') | (results_df['pos_type'] == 'DMG_VIC')
group = results_df[mask]
max_ct_pos_cluster = group.loc[(group['att_side'] == 'CounterTerrorist'), 'pos_cluster'].max()
group.loc[(group['att_side'] == "Terrorist"), 'pos_cluster'] += max_ct_pos_cluster*2
dummies = pd.get_dummies(group['pos_cluster'])
dummies.columns = [f"CT_DMG_{c}" if c <= max_ct_pos_cluster else f"T_DMG_{c - max_ct_pos_cluster*2}" for c in dummies.columns]
dummy_cols.extend(dummies.columns)
for c in dummies.columns:
results_df[c] = 0
results_df.loc[mask, dummies.columns] = dummies
# +
agg_fn = {c: 'sum' for c in dummy_cols}
pos_one_hot = results_df.groupby('file_round').agg(agg_fn)
meta_df = loader.load_meta_df()
meta_df["file_round"] = meta_df["file"] + "_" + meta_df["round"].astype(str)
meta_df = meta_df.drop(columns=['file', 'round'])
total_df = pd.merge(pos_one_hot, meta_df, how='left', left_index = True, right_on = 'file_round')
dummies = pd.get_dummies(total_df['round_type'])
total_df[dummies.columns] = dummies
print(total_df.head(10))
DF_CSV = "full_model_data_mirage.csv"
total_df.to_csv(DF_CSV)
# +
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
from sklearn.model_selection import train_test_split
from sklearn.inspection import permutation_importance
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn import metrics
import xgboost as xgb
import shap
from sklearn.neural_network import MLPRegressor
from sklearn.preprocessing import StandardScaler
def evaluate(mod, X_test, y_test):
y_pred = mod.predict(X_test)
print(f"MAE: {metrics.mean_absolute_error(y_test, y_pred)}")
print(f"MSE: {metrics.mean_squared_error(y_test, y_pred)}")
false_positive_rate, true_positive_rate, thresholds = metrics.roc_curve(y_test, y_pred)
roc_auc = metrics.auc(false_positive_rate, true_positive_rate)
print(f"ROC AUC: {roc_auc}")
def run_models(X, y, mod, tune_params=None, show_shap=False, scale=False):
print(f"RUNNING MODEL {mod}")
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mod.fit(X_train, y_train)
evaluate(mod, X_test, y_test)
if tune_params:
print("TUNING...")
# run randomized search
rf_random = RandomizedSearchCV(estimator = mod, param_distributions = tune_params, n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
rf_random.fit(X_train, y_train)
mod = rf_random.best_estimator_
print(f"BEST MOD: {mod}")
evaluate(mod, X_test, y_test)
print("FEATURE IMPORTANCES:")
sorted_idx = mod.feature_importances_.argsort()
print([x for x in zip(X.columns[-sorted_idx], mod.feature_importances_[-sorted_idx])])
#print("PERMUTATION IMPORTANCES:")
#perm_importance = permutation_importance(mod, X_test, y_test)
#sorted_idx = perm_importance.importances_mean.argsort()
#print([x for x in zip(X.columns[-sorted_idx], perm_importance.importances_mean[-sorted_idx])])
if show_shap:
explainer = shap.TreeExplainer(mod)
shap_values = explainer.shap_values(X_train)
shap.summary_plot(shap_values, X_train, feature_names=X.columns)
return mod, shap_values
return mod
DF_CSV = "full_model_data_mirage.csv"
total_df = pd.read_csv(DF_CSV)
X = total_df.drop(columns=['file_round', 'map', 'start_seconds', 'winner_side', 'round_type'])
#print((agg_results_df['att_side'] == 'Terrorist').astype(int)*2 - 1)
y = (total_df['winner_side'] == 'Terrorist')
run_models(X, y, LinearRegression(), scale=True)
run_models(X, y, RandomForestRegressor(), scale=True)
# get the shap values of the dummy cols
run_models(X, y, GradientBoostingRegressor(), scale=True)
run_models(X, y, xgb.XGBRegressor(), scale=True)
_, shap_values = run_models(X, y, xgb.XGBRegressor(), show_shap=True)
vals= shap_values.mean(0)
absvals= np.abs(shap_values).mean(0)
feature_importance = pd.DataFrame(list(zip(X.columns,vals, absvals)),columns=['col_name','feature_importance_vals', 'absvals'])
feature_importance.sort_values(by=['absvals'],ascending=False,inplace=True)
print(feature_importance.head(5))
# -
model = run_models(X, y, MLPRegressor(hidden_layer_sizes=(100,)), scale=True)
pd.DataFrame(model.loss_curve_).plot()
model = run_models(X, y, MLPRegressor(hidden_layer_sizes=(100,100,)), scale=True)
pd.DataFrame(model.loss_curve_).plot()
model = run_models(X, y, MLPRegressor(hidden_layer_sizes=(15,)), scale=True)
pd.DataFrame(model.loss_curve_).plot()
model = run_models(X, y, MLPRegressor(hidden_layer_sizes=(15,15)), scale=True)
pd.DataFrame(model.loss_curve_).plot()
model = run_models(X, y, MLPRegressor(hidden_layer_sizes=(100,15)), scale=True)
pd.DataFrame(model.loss_curve_).plot()
X_base = X[['ct_eq_val', 't_eq_val', 'ECO', 'FORCE_BUY', 'NORMAL', 'PISTOL_ROUND', 'SEMI_ECO']]
run_models(X_base, y, LinearRegression())
run_models(X_base, y, RandomForestRegressor())
run_models(X_base, y, GradientBoostingRegressor())
run_models(X_base, y, RandomForestRegressor())
run_models(X_base, y, xgb.XGBRegressor())
run_models(X_base, y, MLPRegressor(hidden_layer_sizes=(15,)))
# +
from sklearn.neighbors.nearest_centroid import NearestCentroid
import matplotlib.cm as cm
from matplotlib.pyplot import figure
from adjustText import adjust_text
figure(num=None, figsize=(16, 16), dpi=80, facecolor='w', edgecolor='k')
def visualize_pos_clusters(cluster_df, map_name, clusters, shap_values):
def centroid(x, y):
length = x.shape[0]
sum_x = np.sum(x)
sum_y = np.sum(y)
return sum_x/length, sum_y/length
cluster_df = cluster_df[cluster_df['map'] == map_name]
map_filenames = {map_name: f'../data/{map_name}.png' for map_name in cluster_df['map'].unique()}
plt.title(f'{map_name} important clusters')
im = plt.imread(map_filenames[map_name])
plt.imshow(im)
maxshap = np.max(shap_values)
minshap = np.min(shap_values)
shap_values = (shap_values - minshap) / (maxshap - minshap)
texts = []
for i, c in enumerate(clusters):
df = cluster_df[(cluster_df[c] == 1)]
print(df['pos_cluster'].unique())
col = [cm.RdBu(shap_values[i])]
plot_df = df[:200]
plt.scatter(plot_df['pos_x'], plot_df['pos_y'], c=col, alpha=0.2)
bg_color = 'black' if 0.25 <= shap_values[i] <= 0.75 else 'white'
bbox_props = bbox=dict(facecolor=bg_color, pad=2.0)
cent = centroid(df['pos_x'], df['pos_y'])
texts.append(plt.text(cent[0], cent[1], c, color=cm.RdBu(shap_values[i]), fontsize=12, alpha=0.8, bbox=bbox_props))
#plt.annotate(c, (cent[0] + np.random.normal()*25, cent[1] + np.random.normal()*25), color=cm.RdBu(shap_values[i]), fontsize=12, alpha=0.8, bbox=bbox_props)
adjust_text(texts)
plt.show()
#'col_name','feature_importance_vals'
feature_importance = feature_importance[feature_importance['col_name'].str.contains("CT") | feature_importance['col_name'].str.contains("T")]
feature_importance = feature_importance[feature_importance['col_name'] != 'PISTOL_ROUND']
feature_importance = feature_importance[~feature_importance['col_name'].str.contains("-1")]
feature_importance = feature_importance[:40]
print(feature_importance)
feature_importance.sort_values(by=['feature_importance_vals'],ascending=False,inplace=True)
print(feature_importance)
visualize_pos_clusters(results_df, 'de_mirage', feature_importance['col_name'], feature_importance['feature_importance_vals'].to_numpy())
# -
print(X.columns)
mod = run_models(X, y, GradientBoostingRegressor(min_samples_split=int(len(X)*.01), max_depth=8, max_features='sqrt', subsample=0.8),
tune_params={'max_depth':range(5,16,2),
'min_samples_split':range(10,100,10),
'learning_rate': np.arange(0.1, 1.0, 0.1),
'n_estimators': [1, 2, 4, 8, 16, 32, 64, 100, 200, 400, 800]},
show_shap=True, scale=True)
run_models(X, y, RandomForestRegressor(), tune_params={'bootstrap': [True, False],
'max_depth': [10, 20, 30, 40, 50, 60, 70, 80, 90, 100, None],
'max_features': ['auto', 'sqrt'],
'min_samples_leaf': [1, 2, 4],
'min_samples_split': [2, 5, 10],
'n_estimators': [200, 400, 600, 800, 1000, 1200, 1400, 1600, 1800, 2000]}, show_shap=True, scale=True)
run_models(X, y, MLPRegressor(), tune_params={'bootstrap': [True, False],
'max_depth': [10, 20, 30, 40, 50, 60, 70, 80, 90, 100, None],
'max_features': ['auto', 'sqrt'],
'min_samples_leaf': [1, 2, 4],
'min_samples_split': [2, 5, 10],
'n_estimators': [200, 400, 600, 800, 1000, 1200, 1400, 1600, 1800, 2000]}, show_shap=True, scale=True)
| data_processing/.ipynb_checkpoints/full_model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="7fPc-KWUi2Xd"
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="SL_AnuSKi2Xj"
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/notebooks/samples/optimizer/ai_platform_optimizer_multi_objective.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/notebooks/samples/optimizer/ai_platform_optimizer_multi_objective.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">View on GitHub
# </a>
# </td>
# </table>
#
# + [markdown] colab_type="text" id="AksIKBzZ-nre"
# # Optimizing multiple objectives
#
# This tutorial demonstrates AI Platform Optimizer multi-objective optimization.
#
# ### Objective
#
# The goal is to __`minimize`__ the objective metric:
# ```
# y1 = r*sin(theta)
# ```
#
# and simultaneously __`maximize`__ the objective metric:
# ```
# y2 = r*cos(theta)
# ```
#
# that you will evaluate over the parameter space:
#
# - __`r`__ in [0,1],
#
# - __`theta`__ in [0, pi/2]
#
# ### Costs
#
# This tutorial uses billable components of Google Cloud:
#
# * AI Platform Training
# * Cloud Storage
#
# Learn about [AI Platform Training
# pricing](https://cloud.google.com/ai-platform/training/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
#
#
# + [markdown] colab_type="text" id="D3LI2DGWi2Xl"
# ### PIP install packages and dependencies
#
# Install additional dependencies not installed in the notebook environment.
#
# - Use the latest major GA version of the framework.
# + colab={} colab_type="code" id="2DOjox58i2Xm"
# ! pip install -U google-cloud
# ! pip install -U google-cloud-storage
# ! pip install -U requests
# + tags=["no_execute"]
# Restart the kernel after pip installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] colab_type="text" id="Xc9LnICki2Xp"
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project.](https://console.cloud.google.com/cloud-resource-manager)
#
# 2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
#
# 3. [Enable the AI Platform APIs](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com)
#
# 4. If running locally on your own machine, you will need to install the [Google Cloud SDK](https://cloud.google.com/sdk).
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + colab={} colab_type="code" id="LUmew_Khi2Xr"
PROJECT_ID = '[your-project-id]' #@param {type:"string"}
# ! gcloud config set project $PROJECT_ID
# + [markdown] colab_type="text" id="h0SMyUsC-mzi"
# ### Authenticate your Google Cloud account
#
# **If you are using [AI Platform Notebooks](https://cloud.google.com/ai-platform/notebooks/docs/)**, your environment is already
# authenticated. Skip these steps.
# + colab={} colab_type="code" id="iTQY9g4mRo6r" tags=["no_execute"]
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your Google Cloud account. This provides access
# to your Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
if 'google.colab' in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this tutorial in a notebook locally, replace the string
# below with the path to your service account key and run this cell to
# authenticate your Google Cloud account.
else:
# %env GOOGLE_APPLICATION_CREDENTIALS your_path_to_credentials.json
# Log in to your account on Google Cloud
# !gcloud auth login
# + [markdown] colab_type="text" id="Dax2zrpTi2Xy"
# ### Import libraries
# + colab={} colab_type="code" id="xD60d6Q0i2X0"
import json
import time
import datetime
from googleapiclient import errors
# + [markdown] colab_type="text" id="CWuu4wmki2X3"
# ## Tutorial
#
#
# + [markdown] colab_type="text" id="KyEjqIdnad0w"
# ### Setup
# This section defines some parameters and util methods to call AI Platform Optimizer APIs. Please fill in the following information to get started.
# + colab={} colab_type="code" id="8HCgeF8had77"
# Update to your username
USER = '[user-id]' #@param {type: 'string'}
# These will be automatically filled in.
STUDY_ID = '{}_study_{}'.format(USER.replace('-',''), datetime.datetime.now().strftime('%Y%m%d_%H%M%S')) #@param {type: 'string'}
REGION = 'us-central1'
# + colab={} colab_type="code" id="1SNp8jXhfUAe"
def study_parent():
return 'projects/{}/locations/{}'.format(PROJECT_ID, REGION)
def study_name(study_id):
return 'projects/{}/locations/{}/studies/{}'.format(PROJECT_ID, REGION, study_id)
def trial_parent(study_id):
return study_name(study_id)
def trial_name(study_id, trial_id):
return 'projects/{}/locations/{}/studies/{}/trials/{}'.format(PROJECT_ID, REGION,
study_id, trial_id)
def operation_name(operation_id):
return 'projects/{}/locations/{}/operations/{}'.format(PROJECT_ID, REGION, operation_id)
print('USER: {}'.format(USER))
print('PROJECT_ID: {}'.format(PROJECT_ID))
print('REGION: {}'.format(REGION))
print('STUDY_ID: {}'.format(STUDY_ID))
# + [markdown] colab_type="text" id="cs_cawOD_FA2"
# ### Build the API client
#
# The following cell builds the auto-generated API client using [Google API discovery service](https://developers.google.com/discovery). The JSON format API schema is hosted in a Cloud Storage bucket.
# + colab={} colab_type="code" id="f9gZ0E12-8S9"
from google.cloud import storage
from googleapiclient import discovery
_OPTIMIZER_API_DOCUMENT_BUCKET = 'caip-optimizer-public'
_OPTIMIZER_API_DOCUMENT_FILE = 'api/ml_public_google_rest_v1.json'
def read_api_document():
client = storage.Client(PROJECT_ID)
bucket = client.get_bucket(_OPTIMIZER_API_DOCUMENT_BUCKET)
blob = bucket.get_blob(_OPTIMIZER_API_DOCUMENT_FILE)
return blob.download_as_string()
ml = discovery.build_from_document(service=read_api_document())
print('Successfully built the client.')
# + [markdown] colab_type="text" id="8NBduXsEaRKr"
# ### Create the study configuration
#
# The following is a sample study configuration, built as a hierarchical python dictionary. It is already filled out. Run the cell to configure the study.
# + colab={} colab_type="code" id="s-AHfPOASXXW"
# Parameter Configuration
param_r = {
'parameter': 'r',
'type' : 'DOUBLE',
'double_value_spec' : {
'min_value' : 0,
'max_value' : 1
}
}
param_theta = {
'parameter': 'theta',
'type' : 'DOUBLE',
'double_value_spec' : {
'min_value' : 0,
'max_value' : 1.57
}
}
# Objective Metrics
metric_y1 = {
'metric' : 'y1',
'goal' : 'MINIMIZE'
}
metric_y2 = {
'metric' : 'y2',
'goal' : 'MAXIMIZE'
}
# Put it all together in a study configuration
study_config = {
'algorithm' : 'ALGORITHM_UNSPECIFIED', # Let the service choose the `default` algorithm.
'parameters' : [param_r, param_theta,],
'metrics' : [metric_y1, metric_y2,],
}
study = {'study_config': study_config}
print(json.dumps(study, indent=2, sort_keys=True))
# + [markdown] colab_type="text" id="uyXG_RKha7Kb"
# ### Create the study
#
# Next, create the study, which you will subsequently run to optimize the two objectives.
# + colab={} colab_type="code" id="jgskzqZX0Mkt"
# Creates a study
req = ml.projects().locations().studies().create(
parent=study_parent(), studyId=STUDY_ID, body=study)
try :
print(req.execute())
except errors.HttpError as e:
if e.resp.status == 409:
print('Study already existed.')
else:
raise e
# + [markdown] colab_type="text" id="dKOMBKmtkcJb"
# ### Metric evaluation functions
#
# Next, define some functions to evaluate the two objective metrics.
# + colab={} colab_type="code" id="Xnl1uqnyz3Qp"
import math
# r * sin(theta)
def Metric1Evaluation(r, theta):
"""Evaluate the first metric on the trial."""
return r * math.sin(theta)
# r * cose(theta)
def Metric2Evaluation(r, theta):
"""Evaluate the second metric on the trial."""
return r * math.cos(theta)
def CreateMeasurement(trial_id, r, theta):
print(("=========== Start Trial: [{0}] =============").format(trial_id))
# Evaluate both objective metrics for this trial
y1 = Metric1Evaluation(r, theta)
y2 = Metric2Evaluation(r, theta)
print('[r = {0}, theta = {1}] => y1 = r*sin(theta) = {2}, y2 = r*cos(theta) = {3}'.format(r, theta, y1, y2))
metric1 = {'metric': 'y1', 'value': y1}
metric2 = {'metric': 'y2', 'value': y2}
# Return the results for this trial
measurement = {'step_count': 1, 'metrics': [metric1, metric2,]}
return measurement
# + [markdown] colab_type="text" id="Qzn5lVpRq05U"
# ### Set configuration parameters for running trials
#
# __`client_id`__ - The identifier of the client that is requesting the suggestion. If multiple SuggestTrialsRequests have the same `client_id`, the service will return the identical suggested trial if the trial is `PENDING`, and provide a new trial if the last suggested trial was completed.
#
# __`suggestion_count_per_request`__ - The number of suggestions (trials) requested in a single request.
#
# __`max_trial_id_to_stop`__ - The number of trials to explore before stopping. It is set to 4 to shorten the time to run the code, so don't expect convergence. For convergence, it would likely need to be about 20 (a good rule of thumb is to multiply the total dimensionality by 10).
#
#
# + colab={} colab_type="code" id="5usXaZA5qvUZ"
client_id = 'client1' #@param {type: 'string'}
suggestion_count_per_request = 5 #@param {type: 'integer'}
max_trial_id_to_stop = 50 #@param {type: 'integer'}
print('client_id: {}'.format(client_id))
print('suggestion_count_per_request: {}'.format(suggestion_count_per_request))
print('max_trial_id_to_stop: {}'.format(max_trial_id_to_stop))
# + [markdown] colab_type="text" id="UnV2SJNskm7V"
# ### Run AI Platform Optimizer trials
#
# Run the trials.
# + colab={} colab_type="code" id="opmuTntW4-eS"
trial_id = 0
while trial_id < max_trial_id_to_stop:
# Requests trials.
resp = ml.projects().locations().studies().trials().suggest(
parent=trial_parent(STUDY_ID),
body={'client_id': client_id, 'suggestion_count': suggestion_count_per_request}).execute()
op_id = resp['name'].split('/')[-1]
# Polls the suggestion long-running operations.
get_op = ml.projects().locations().operations().get(name=operation_name(op_id))
while True:
operation = get_op.execute()
if 'done' in operation and operation['done']:
break
time.sleep(1)
for suggested_trial in get_op.execute()['response']['trials']:
trial_id = int(suggested_trial['name'].split('/')[-1])
# Featches the suggested trials.
trial = ml.projects().locations().studies().trials().get(name=trial_name(STUDY_ID, trial_id)).execute()
if trial['state'] in ['COMPLETED', 'INFEASIBLE']:
continue
# Parses the suggested parameters.
params = {}
for param in trial['parameters']:
if param['parameter'] == 'r':
r = param['floatValue']
elif param['parameter'] == 'theta':
theta = param['floatValue']
# Evaluates trials and reports measurement.
ml.projects().locations().studies().trials().addMeasurement(
name=trial_name(STUDY_ID, trial_id),
body={'measurement': CreateMeasurement(trial_id, r, theta)}).execute()
# Completes the trial.
ml.projects().locations().studies().trials().complete(
name=trial_name(STUDY_ID, trial_id)).execute()
# + [markdown] colab_type="text" id="i5ZTqgqBiRsq"
# ### [EXPERIMENTAL] Visualize the result
#
# This section provides a module to visualize the trials for the above study.
#
# + colab={} colab_type="code" id="Or2PL1YxTr33"
max_trials_to_annotate = 20
import matplotlib.pyplot as plt
trial_ids = []
y1 = []
y2 = []
resp = ml.projects().locations().studies().trials().list(parent=trial_parent(STUDY_ID)).execute()
for trial in resp['trials']:
if 'finalMeasurement' in trial:
trial_ids.append(int(trial['name'].split('/')[-1]))
metrics = trial['finalMeasurement']['metrics']
try:
y1.append([m for m in metrics if m['metric'] == "y1"][0]['value'])
y2.append([m for m in metrics if m['metric'] == "y2"][0]['value'])
except:
pass
fig, ax = plt.subplots()
ax.scatter(y1, y2)
plt.xlabel("y1=r*sin(theta)")
plt.ylabel("y2=r*cos(theta)");
for i, trial_id in enumerate(trial_ids):
# Only annotates the last `max_trials_to_annotate` trials
if i > len(trial_ids) - max_trials_to_annotate:
try:
ax.annotate(trial_id, (y1[i], y2[i]))
except:
pass
plt.gcf().set_size_inches((16, 16))
# + [markdown] colab_type="text" id="KAxfq9Fri2YV"
# ## Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
| notebooks/samples/optimizer/ai_platform_optimizer_multi_objective.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploratory data analysis
# # Purpose
# Explore the data and extend/transform features.
# + tags=["remove_cell"]
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings('ignore')
# + tags=["remove_cell"]
import os
import matplotlib.pyplot as plt
import pandas as pd
pd.options.display.max_rows = 999
pd.options.display.max_columns = 999
pd.set_option("display.max_columns", None)
import sympy as sp
import pandas as pd
import numpy as np
# Unmark for publish quality plots
#if os.name == 'nt':
# plt.style.use('paper.mplstyle') # Windows
import statsmodels.api as sm
from statsmodels.sandbox.regression.predstd import wls_prediction_std
from sklearn.metrics import r2_score, mean_absolute_error
from scipy.integrate import solve_ivp
import seaborn as sns
from copy import deepcopy
import sys
sys.path.append('../../')
from src.notebook_to_latex import Equation, equation_dict
#from src import equations,symbols
import reports.paper_writing
from src.df_to_latex import LateXTable
import src.data.load as load
from src.data.transform import transform, extend
from src.visualization.visualize import plotable_dataframe
from src.visualization.polynom import model_to_string, model_to_sympy
import plotly.express as px
import plotly.graph_objects as go
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import TimeSeriesSplit
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import PolynomialFeatures
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_regression
import xgboost
## Examples:
# -
# ## Exploratory data analysis
# The ship speed $V$, ship draughts $T_{aft}$ and $T_{fwd}$ were all negative in the raw data file. This was imidiatelly corrected, to be more in line with what would be expected from a more general sign convention. The data seems to have been collected in time cronological order, giving a time series of data. For a time series, measurements close to each other in time have a high correlation, as they are experiencing similar envinronmental conditions etc. This is confirmed by looking at the autocorrelation plot in Fig.[(below)](#fig_power_autocorrelation). Dead reckoning (using ship speed and heading) has been used to atempt to describe motion of the ship as seen in Fig.[(below)](#fig_dead_reckoning). The positions are given in an unknown logintude and latitude scale, as the time step between measurements is unknown. The speed of the ship is also indicated as a color gradient in this figure.
raw_data = load.raw()
display(raw_data.head())
display(raw_data.describe())
display(raw_data.dtypes)
# ## Autocorrelation
# + caption="Autocorrelation plot of the Power data" name="power_autocorrelation"
from statsmodels.graphics.tsaplots import plot_acf
fig = plot_acf(raw_data['Power'].values, lags=8000);
ax = fig.get_axes()[0]
ax.set_xlabel('Lag')
ax.set_title('Power autocorrelation');
# +
from scipy.spatial.transform import Rotation as R
r = R.from_euler('z', raw_data['HDG'], degrees=True)
df = pd.DataFrame(index=raw_data.index)
df['u'] = raw_data['V'] ## Assuming no drift
df['v'] = 0 ## Assuming no drift
df['w'] = 0 ## Assuming no drift
df[['dX','dY','dZ']] = r.apply(df)
df[['X','Y','Z']] = df[['dX','dY','dZ']].cumsum()
# + caption="Dead reckoning of the position of the ship" name="dead_reckoning"
from matplotlib.collections import LineCollection
from matplotlib.colors import ListedColormap, BoundaryNorm
x = df['Y'].values
y = df['X'].values
dydx = df['u'].values
# Create a set of line segments so that we can color them individually
# This creates the points as a N x 1 x 2 array so that we can stack points
# together easily to get the segments. The segments array for line collection
# needs to be (numlines) x (points per line) x 2 (for x and y)
points = np.array([x, y]).T.reshape(-1, 1, 2)
segments = np.concatenate([points[:-1], points[1:]], axis=1)
fig, ax = plt.subplots()
# Create a continuous norm to map from data points to colors
norm = plt.Normalize(dydx.min(), dydx.max())
lc = LineCollection(segments, cmap='gray', norm=norm)
# Set the values used for colormapping
lc.set_array(dydx)
lc.set_linewidth(2)
line = ax.add_collection(lc)
fig.colorbar(line, ax=ax)
ax.set_xlim(df['Y'].min(),df['Y'].max())
ax.set_ylim(df['X'].min(),df['X'].max())
ax.annotate('start',xy=df.iloc[0][['Y','X']])
ax.annotate('end',xy=df.iloc[-1][['Y','X']])
ax.axis('equal')
ax.set_xlabel('Latitude scale')
ax.set_ylabel('Longitude scale');
# + caption="Heat map showing absolute value of correlation coefficient between features in raw data" name="heat_map_raw_data"
corr = plotable_dataframe(raw_data.corr().abs())
ax = sns.heatmap(corr, vmin=0, vmax=1, yticklabels=corr.index, cmap='Blues', annot=True)
fig = ax.get_figure()
# -
# ## transform
# + caption="Heat map showing absolute value of correlation coefficient between features in transformed data" name="heat_map_data"
data = transform(raw_data=raw_data)
corr = plotable_dataframe(data.corr().abs())
ax = sns.heatmap(corr, vmin=0, vmax=1, yticklabels=corr.index, cmap='Blues', annot=True)
fig = ax.get_figure()
# + caption="The power is highly correlated with the draught" name="power_draught"
fig,ax=plt.subplots()
data_ = data.copy()
data_.plot(y='Power', ax=ax)
ax2 = ax.twinx()
data_['-T'] = -data_['T']
data_.plot(y='-T', style='r-', ax=ax2)
# -
# ## Extend data
data_extended = extend(data=data)
corr = plotable_dataframe(data_extended.corr().abs())
fig,ax = plt.subplots()
fig.set_size_inches(9,9)
ax = sns.heatmap(corr, vmin=0, vmax=1, yticklabels=corr.index, cmap='Blues', annot=True, ax=ax)
data_extended.to_csv('../data/processed/data_extended.csv')
| notebooks/02.1_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
plt.rc('font', size=15)
# +
fig, ax = plt.subplots(figsize=(8, 6))
#Vector field
X, Y = np.meshgrid(np.linspace(-5, 5, 20), np.linspace(-5, 5, 20))
U = 1
V = 1 + X * Y
#Normalize arrows
N = np.sqrt(U**2 + V**2)
U /= N
V /= N
ax.quiver(X, Y, U, V, scale=30, pivot='mid')
plt.xlim([-5, 5])
plt.ylim([-5, 5])
plt.xlabel(r"$x$")
plt.ylabel(r"$y$")
plt.show()
# -
# https://stackoverflow.com/questions/18832763/drawing-directions-fields
from scipy.integrate import odeint
# +
from scipy.integrate import ode
fig, ax = plt.subplots(figsize=(8, 6))
#Vector field
X, Y = np.meshgrid(np.linspace(-15, 15, 20), np.linspace(-15, 15, 20))
U = 2 * X + Y
V = - X + 4 * Y
#Normalize arrows
N = np.sqrt(U**2 + V**2)
U /= N
V /= N
ax.quiver(X, Y, U, V, N, scale=30, pivot='mid', cmap='viridis_r')
## Vector field function
def vf(t,x):
dx=np.zeros(2)
dx[0] = 2 * x[0] + x[1]
dx[1] = - x[0] + 4 * x[1]
return dx
#Solution curves
t0 = 0
tEnd = 10
dt = 0.01
r = ode(vf).set_integrator('vode', method='bdf', max_step=dt)
#### Need to tweak the initial conditions here ###
'''
x_ini = np.linspace(-5, 5, 10)
y_ini = np.linspace(-5, 5, 10)
for x_ic in x_ini:
for y_ic in y_ini:
Y = []
T = []
S = []
r.set_initial_value([x_ic, y_ic], t0).set_f_params()
while r.successful() and r.t + dt < tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S=np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'blue', lw = 1.25)
'''
radius = 5
for theta in np.linspace(0, 2*np.pi, 30):
x_ini = radius * np.cos(theta)
y_ini = radius * np.sin(theta)
#for ic in [[0, 1], [0, 7], [3, 5], [5, 8], [8, 8], [12, 8], [12, 3], [15, 0], [-5, -5], [-5, -3], [-5, -7], [-5, -10], [-5, -15], [-5, -20], [-5, -2]]:
# x_ini, y_ini = ic
Y = []
T = []
S = []
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t - dt > -10:
r.integrate(r.t - dt)
Y.append(r.y)
Y = Y[::-1]
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t + dt <= tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S = np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'red', lw = 1.25)
plt.xlim([-15, 15])
plt.ylim([-15, 15])
plt.xlabel(r"$x$")
plt.ylabel(r"$y$")
plt.tick_params(direction='in')
plt.show()
# +
from scipy.integrate import ode
fig, ax = plt.subplots(figsize=(8, 6))
#Vector field
X, Y = np.meshgrid(np.linspace(-15, 15, 20), np.linspace(-15, 15, 20))
U = 2 * X + Y
V = - X + 4 * Y
#Normalize arrows
N = np.sqrt(U**2 + V**2)
U /= N
V /= N
ax.quiver(X, Y, U, V, scale=30, pivot='mid')
## Vector field function
def vf(t,x):
dx=np.zeros(2)
dx[0] = 2 * x[0] + x[1]
dx[1] = - x[0] + 4 * x[1]
return dx
#Solution curves
t0 = 0
tEnd = 10
dt = 0.01
r = ode(vf).set_integrator('vode', method='bdf', max_step=dt)
#### Need to tweak the initial conditions here ###
'''
x_ini = np.linspace(-5, 5, 10)
y_ini = np.linspace(-5, 5, 10)
for x_ic in x_ini:
for y_ic in y_ini:
Y = []
T = []
S = []
r.set_initial_value([x_ic, y_ic], t0).set_f_params()
while r.successful() and r.t + dt < tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S=np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'blue', lw = 1.25)
'''
for x_ini in np.arange(-10, 10, 1):
y_ini = 0
#x_ini, y_ini = 2, 0
Y = []
T = []
S = []
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t - dt > -10:
r.integrate(r.t - dt)
Y.append(r.y)
Y = Y[::-1]
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t + dt <= tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S = np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'red', lw = 1.25)
for ic in [[0, 1], [0, 7], [3, 5], [5, 8], [8, 8], [12, 8], [12, 3], [15, 0], [-5, -5], [-5, -3], [-5, -7], [-5, -10], [-5, -15], [-5, -20], [-5, -2]]:
x_ini, y_ini = ic
Y = []
T = []
S = []
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t - dt > -10:
r.integrate(r.t - dt)
Y.append(r.y)
Y = Y[::-1]
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t + dt <= tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S = np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'red', lw = 1.25)
plt.xlim([-15, 15])
plt.ylim([-15, 15])
plt.xlabel(r"$x$")
plt.ylabel(r"$y$")
plt.show()
# +
from scipy.integrate import ode
fig, ax = plt.subplots(figsize=(8, 6))
#Vector field
X, Y = np.meshgrid(np.linspace(-15, 15, 20), np.linspace(-15, 15, 20))
U = 2 * X + Y
V = - X + 4 * Y
#Normalize arrows
N = np.sqrt(U**2 + V**2)
U /= N
V /= N
ax.quiver(X, Y, U, V, scale=30, pivot='mid')
## Vector field function
def vf(t,x):
dx=np.zeros(2)
dx[0] = 2 * x[0] + x[1]
dx[1] = - x[0] + 4 * x[1]
return dx
#Solution curves
t0 = 0
tEnd = 10
dt = 0.01
r = ode(vf).set_integrator('vode', method='bdf', max_step=dt)
#### Need to tweak the initial conditions here ###
'''
x_ini = np.linspace(-5, 5, 10)
y_ini = np.linspace(-5, 5, 10)
for x_ic in x_ini:
for y_ic in y_ini:
Y = []
T = []
S = []
r.set_initial_value([x_ic, y_ic], t0).set_f_params()
while r.successful() and r.t + dt < tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S=np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'blue', lw = 1.25)
'''
for ic in [[8, 2], [0, 1], [0, 7], [3, 5], [5, 8], [8, 8],]:
x_ini, y_ini = ic
Y = []
T = []
S = []
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t - dt > -10:
r.integrate(r.t - dt)
Y.append(r.y)
Y = Y[::-1]
r.set_initial_value([x_ini, y_ini], t0).set_f_params()
while r.successful() and r.t + dt <= tEnd:
r.integrate(r.t + dt)
Y.append(r.y)
S = np.array(np.real(Y))
ax.plot(S[:,0],S[:,1], color = 'red', lw = 1.25)
plt.xlim([-15, 15])
plt.ylim([-15, 15])
plt.xlabel(r"$x$")
plt.ylabel(r"$y$")
plt.show()
# -
| plot/ODEphase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="DExKBrWjbLLx" executionInfo={"status": "ok", "timestamp": 1630102383954, "user_tz": 240, "elapsed": 29984, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="1ac87659-feca-49d9-b9ac-91a321108ba1"
import pandas as pd
import numpy as np
import os
import progressbar
import pickle
import sys
import random
from PIL import Image
import seaborn as sb
import matplotlib.pyplot as plt
from sklearn.metrics import (plot_confusion_matrix, plot_precision_recall_curve,
plot_roc_curve, auc)
from sklearn.preprocessing import MinMaxScaler
from sklearn.svm import SVC
from sklearn.model_selection import (train_test_split, cross_validate,
cross_val_predict, GridSearchCV,
StratifiedKFold, learning_curve)
# !pip install delayed
# !pip uninstall scikit-learn
# !pip install scikit-learn
# !pip install -U imbalanced-learn
from imblearn.combine import SMOTEENN, SMOTETomek
import time
import math
from statistics import mean, stdev
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="zUeNOIWTr7rK"
# Total number of data
# + id="QjeAshw_r99D" executionInfo={"status": "ok", "timestamp": 1630102284900, "user_tz": 240, "elapsed": 167, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
data_per_csv = 512
def total():
total_files = 0
for files in os.listdir('gdrive/My Drive/Summer Research/HRV/Outlier Free/All/'):
total_files += 1
return total_files
# + [markdown] id="9UcGQsgWjtGv"
# Save HRV data to file
# + id="Ugn77xjSjnRD" executionInfo={"status": "ok", "timestamp": 1630102288480, "user_tz": 240, "elapsed": 166, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def saveHRVData(c):
hrv_and_labels = importAllData(c)
if c == 'array':
with open('gdrive/My Drive/Summer Research/Variables/array_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels, file)
elif c == 'denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_denoised_hrv_and_labels.pkl', 'wb') as file:
#load data from file
pickle.dump(hrv_and_labels, file)
elif c == 'wt a1d1d2d3 coords':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1d1d2d3_coords_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels, file)
elif c == 'wt a1d1d2d3 denoised coords':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1d1d2d3_denoised_coords_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels, file)
elif c == 'a1d1d2d3':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[0], file)
with open('gdrive/My Drive/Summer Research/Variables/wt_d1_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[1], file)
with open('gdrive/My Drive/Summer Research/Variables/wt_d2_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[2], file)
with open('gdrive/My Drive/Summer Research/Variables/wt_d3_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[3], file)
elif c == 'd1d2d3 denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_d1_denoised_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[0], file)
with open('gdrive/My Drive/Summer Research/Variables/wt_d2_denoised_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[1], file)
with open('gdrive/My Drive/Summer Research/Variables/wt_d3_denoised_coord_hrv_and_labels.pkl', 'wb') as file:
#save data to a file
pickle.dump(hrv_and_labels[2], file)
# + [markdown] id="HZFY1UBUbfmx"
# Load HRV data
# + id="WzdEjxhlbWcL" executionInfo={"status": "ok", "timestamp": 1630102298496, "user_tz": 240, "elapsed": 155, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def loadHRVData(c):
hrv_and_labels = list()
if c == 'wt':
with open('gdrive/My Drive/Summer Research/Variables/wt_pseudoimage_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'wt denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_denoised_pseudoimage_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_denoised_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'normal':
size = (163, 223, 4)
with open('gdrive/My Drive/Summer Research/Variables/normal_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'array':
with open('gdrive/My Drive/Summer Research/Variables/array_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'wt a1d1d2d3 coords':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1d1d2d3_coords_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'wt a1d1d2d3 denoised coords':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1d1d2d3_denoised_coords_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'a1':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'd1':
with open('gdrive/My Drive/Summer Research/Variables/wt_d1_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'd2':
with open('gdrive/My Drive/Summer Research/Variables/wt_d2_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'd3':
with open('gdrive/My Drive/Summer Research/Variables/wt_d3_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'd1 denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_d1_denoised_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'd2 denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_d2_denoised_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'd3 denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_d3_denoised_coord_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
return hrv_and_labels
# + [markdown] id="Nxoet_iOlpoV"
# Normalize
# + id="zr9v4g7DlqbK" executionInfo={"status": "ok", "timestamp": 1630102335270, "user_tz": 240, "elapsed": 164, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def Normalize(arr):
arr_scaled = np.zeros(arr.shape)
scaler = MinMaxScaler(feature_range = (0, 1))
arr_scaled = scaler.fit_transform(arr.reshape(-1,1))
return arr_scaled
# + [markdown] id="UCRqBj_wmVvR"
# Import HRV Data
# + id="KRbnc37tmW7V" executionInfo={"status": "ok", "timestamp": 1630102433605, "user_tz": 240, "elapsed": 136, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def importAllData(c):
classify = {'h':0, 'd':1}
widgets = [' [',
progressbar.Timer(format= 'elapsed time: %(elapsed)s'),
'] ',
progressbar.Bar('#'),' (',
progressbar.ETA(), ') ',
]
all_total = total()
bar = progressbar.ProgressBar(max_value=all_total, widgets=widgets).start()
master_list = list()
count = 0
if c == 'array':
data_path = 'gdrive/My Drive/Summer Research/HRV/Outlier Free/All/'
for files in os.listdir(data_path):
count += 1
bar.update(count)
sublist = list()
h_or_d = classify[files[0]]
image = Normalize(np.loadtxt(data_path+files, delimiter=','))
sublist.append(image)
sublist.append(h_or_d)
master_list.append(sublist)
elif c == 'denoised':
h_and_l = loadHRVData('wt denoised')
master_list = list()
for i in range(len(h_and_l)):
sublist = list()
coords = h_and_l[i][0]
h_or_d = h_and_l[i][1]
sublist.append(coords.sum(axis=0))
sublist.append(h_or_d)
master_list.append(sublist)
elif c == 'wt a1d1d2d3 coords':
n = data_per_csv
t = fourWTM(n)
data_path = 'gdrive/My Drive/Summer Research/HRV/Outlier Free/All/'
for files in os.listdir(data_path):
count += 1
bar.update(count)
sublist = list()
h_or_d = classify[files[0]]
s = np.loadtxt(data_path+files, delimiter=',')
ts = np.matmul(t,s)
sublist.append(ts)
sublist.append(h_or_d)
master_list.append(sublist)
elif c == 'wt a1d1d2d3 denoised coords':
n = data_per_csv
t = fourWTM(n)
data_path = 'gdrive/My Drive/Summer Research/HRV/Outlier Free/All/'
for files in os.listdir(data_path):
count += 1
bar.update(count)
sublist = list()
h_or_d = classify[files[0]]
s = np.loadtxt(data_path+files, delimiter=',')
ts = np.matmul(t,s)
dim = int(n/4)
for i in range(3):
rng = range(dim*(i+1), dim*(i+2))
lbda = np.std(ts[rng])*math.sqrt(2*math.log(dim))
for j in rng:
if ts[j] < lbda:
ts[j] = 0
sublist.append(ts)
sublist.append(h_or_d)
master_list.append(sublist)
elif c == 'a1d1d2d3':
h_and_l = loadHRVData('wt a1d1d2d3 coords')
for i in range(len(h_and_l)):
for j in range(4):
master_list.append([])
coords = h_and_l[i][0]
h_or_d = h_and_l[i][1]
for j in range(4):
master_list[j].append([coords[int(j*data_per_csv/4):int((j+1)*data_per_csv/4)], h_or_d])
elif c == 'd1d2d3 denoised':
h_and_l = loadHRVData('wt a1d1d2d3 denoised coords')
for i in range(len(h_and_l)):
for j in range(3):
master_list.append([])
coords = h_and_l[i][0]
h_or_d = h_and_l[i][1]
for j in range(1,4):
master_list[j-1].append([coords[int(j*data_per_csv/4):int((j+1)*data_per_csv/4)], h_or_d])
return master_list
# + id="SU4Vm21Ljxkc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1630102436058, "user_tz": 240, "elapsed": 383, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="e6ec958f-8ace-4d7a-eaf0-ce14dc692d86"
saveHRVData('denoised')
# + [markdown] id="8zWtp-RtP65g"
# Wavelet Transform
# + id="TE_y2lMgQECq"
def fourWTM(n):
#Filter banks
h0 = np.array([0.2697890,0.3947890,0.5197890,0.6447890,0.2302110,0.1052110,-0.0197890,-0.1447890])
h1 = np.array([-0.2825435,0.5553379,0.2385187,-0.0783004, -0.5834819,-0.2666627,0.0501564,0.3669755])
h2 = np.array([0.4125840,-0.6279376,0.3727824,0.1487574, -0.4125840,-0.1885590,0.0354659,0.2594909])
h3 = np.array([0.2382055,0.1088646,-0.7275830,0.5572896, -0.2382055,-0.1088646,0.0204763,0.1498171])
#Matrix of filter banks created for convenience
h = np.array([h0,h1,h2,h3])
k = int(n/4)
T = np.zeros((n,n))
for j in range(4):
for i in range(k):
if 4*i+8 > 4*k:
T[k*j+i,range((4*i),(4*i+4))] = h[j,range(4)]
T[k*j+i,range(4)] = h[j,range(4,8)]
else:
T[k*j+i,range((4*i),(4*i+8))] = h[j,range(8)]
return T
# + [markdown] id="9WN7sn7Bhkh0"
# Resampling methods
# + id="VObRoXK_hl99"
def resampling(args):
if args == 'SMOTEENN':
resampler = SMOTEENN(enn=EditedNearestNeighbours(sampling_strategy='majority'),
n_jobs=-1)
elif args == 'SMOTETomek':
resampler = SMOTETomek(n_jobs=-1)
return resampler
# + [markdown] id="lkGfxP0rrT0w"
# SVM model
# + id="6kM69PI_-iyI"
def SVMModel(X, y, cv):
model = SVC(kernel='rbf', gamma=0.01, C=1, class_weight='balanced', probability=True)
#K-fold Cross Validation
scores = cross_validate(model, X, y, cv=cv, scoring=('accuracy', 'balanced_accuracy', 'precision', 'recall', 'roc_auc', 'f1'), n_jobs=-1, verbose=0, return_estimator=True)
return scores
# + id="RM20f6voMTSV"
def metrics(scores, X, y, cv, resampling_method, data_choice):
dir = 'gdrive/My Drive/Summer Research/Figures/SVM/'
file_name = resampling_method+'-resampled '+data_choice
rem_list = ['estimator', 'fit_time', 'score_time']
csv_scores = dict([(key, val) for key, val in
scores.items() if key not in rem_list])
df = pd.DataFrame.from_dict(csv_scores)
df.to_csv(dir+file_name+'.csv', index=False)
#TODO: generate PR, ROC, Confusion matrix graphs
tprs = []
aucs = []
mean_fpr = np.linspace(0, 1, 100)
cm = np.zeros((4,10))
fig, ax = plt.subplots(figsize=(10,10))
fig2, ax2 = plt.subplots(figsize=(10,10))
fig3, ax3 = plt.subplots(figsize=(10,10))
fig4, ax4 = plt.subplots(figsize=(10,10))
for i, (train, test) in enumerate(cv.split(X, y)):
viz = plot_roc_curve(scores['estimator'][i], X[test], y[test],
name='ROC fold {}'.format(i),
alpha=0.3, lw=1, ax=ax)
interp_tpr = np.interp(mean_fpr, viz.fpr, viz.tpr)
interp_tpr[0] = 0.0
tprs.append(interp_tpr)
aucs.append(viz.roc_auc)
p = plot_precision_recall_curve(scores['estimator'][i], X[test],
y[test], name='P v. R fold {}'.format(i),
alpha=0.5, lw=1.5, ax=ax2)
c = plot_confusion_matrix(scores['estimator'][i], X[test], y[test],
normalize='all', ax=ax4)
cm[:,i] = np.array(c.confusion_matrix).reshape(4,)
plt.close(fig=fig4)
#ROC Curve
ax.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
label='Chance', alpha=.8)
mean_tpr = np.mean(tprs, axis=0)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
std_auc = np.std(aucs)
ax.plot(mean_fpr, mean_tpr, color='b',
label=r'Mean ROC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
lw=2, alpha=.8)
std_tpr = np.std(tprs, axis=0)
tprs_upper = np.minimum(mean_tpr + std_tpr, 1)
tprs_lower = np.maximum(mean_tpr - std_tpr, 0)
ax.fill_between(mean_fpr, tprs_lower, tprs_upper, color='grey', alpha=.2,
label=r'$\pm$ 1 std. dev.')
ax.set(xlim=[-0.05, 1.05], ylim=[-0.05, 1.05],
title="ROC Curve")
ax.legend(loc="lower right")
fig.savefig(dir+file_name+' ROC.png', bbox_inches='tight')
plt.close(fig=fig)
#PR Curve
ax2.set(xlim=[-0.05, 1.05], ylim=[-0.05, 1.05],
title="Precision v. Recall Curve")
ax2.legend(loc="lower left")
fig2.savefig(dir+file_name+' PR.png', bbox_inches='tight')
plt.close(fig=fig2)
#Confusion Matrix
c1, c2, c3, c4 = cm[0,:], cm[1,:], cm[2,:], cm[3,:]
means = np.array([[mean(c1), mean(c2)],[mean(c3), mean(c4)]])
stds = np.array([[stdev(c1), stdev(c2)],[stdev(c3), stdev(c4)]])
labels = np.array([["{:.2%} $\pm$ {:.2%}".format(mean(c1), stdev(c1)),
"{:.2%} $\pm$ {:.2%}".format(mean(c2), stdev(c2))],
["{:.2%} $\pm$ {:.2%}".format(mean(c3), stdev(c3)),
"{:.2%} $\pm$ {:.2%}".format(mean(c4), stdev(c4))]])
plt.figure(figsize=(12,8))
g = sb.heatmap(100*means, fmt='', annot=labels, cmap='Greens',
xticklabels=['Predicted Healthy', 'Predicted Diabetes'],
yticklabels=['Healthy', 'Diabetes'], ax=ax3, cbar_kws={'format': '%.0f%%'})
g.set_yticklabels(labels=g.get_yticklabels(), va='center')
g.set_title('Confusion Matrix')
fig3.savefig(dir+file_name+' Confusion Matrix.png', bbox_inches='tight')
plt.close(fig=fig3)
# + id="Zu72pW9PrWsK" colab={"base_uri": "https://localhost:8080/", "height": 151} executionInfo={"status": "ok", "timestamp": 1629866562655, "user_tz": 240, "elapsed": 1183607, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="1df03437-994c-476c-faf2-2f09bdc17acf"
data_choices = {
1:'a1',
2:'d1',
3:'d2',
4:'d3',
5:'d1 denoised',
6:'d2 denoised',
7:'d3 denoised'
}
all_total = total()
cv = StratifiedKFold(n_splits=10, shuffle=True, random_state=1)
widgets = [' [',
progressbar.Timer(format= 'elapsed time: %(elapsed)s'),
'] ',
progressbar.Bar('#'),' (',
progressbar.ETA(), ') ',
]
bar = progressbar.ProgressBar(max_value=10, widgets=widgets).start()
count = 0
resampling_method = 'SMOTETomek'
for i in range(len(data_choices)):
count += 1
bar.update(count)
data_choice = data_choices[i+1]
hrv_and_labels = loadHRVData(data_choice)
random.shuffle(hrv_and_labels)
X = np.array([item[0] for item in hrv_and_labels]).reshape(total(),-1)
y = np.array([item[1] for item in hrv_and_labels])
X_resampled, y_resampled = resampling(resampling_method).fit_resample(X, y)
scores = SVMModel(X_resampled, y_resampled, cv)
metrics(scores, X_resampled, y_resampled, cv, resampling_method, data_choice)
| Code/HRV Classification/SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MPS using the deesse wrapper - using a mask
# ## Import what is required
# +
import numpy as np
import matplotlib.pyplot as plt
# import from package 'geone'
from geone import img
import geone.imgplot as imgplt
import geone.customcolors as ccol
import geone.deesseinterface as dsi
# -
# ## Training image (TI)
# Read the training image.
ti = img.readImageGslib('ti.gslib')
# Plot the image (using the function `imgplt.drawImage2D`).
# +
col = ['lightblue', 'darkgreen', 'orange']
plt.figure(figsize=(5,5))
imgplt.drawImage2D(ti, categ=True, categCol=col, title='TI')
# -
# ## Simulation grid and mask
# Deesse requires a regular cartesian grid. However, one can specify to deesse to not simulate every cell in the simulation grid. For that, a mask is used: a value (`0` or `1`) is attached to each simulation grid cell indicating that the cell must be simulated (value `1`) or not (value `0`). These values are passed to deesse by an array.
# +
# Read the mask (image) from the file 'mask.gslib'
mask = img.readImageGslib('mask.gslib')
# Set the simulation grid
nx, ny, nz = mask.nx, mask.ny, mask.nz
sx, sy, sz = mask.sx, mask.sy, mask.sz
ox, oy, oz = mask.ox, mask.oy, mask.oz
# Set mask values to simulation grid cells (1: cell to be simulated)
mask_value = mask.val
# -
# ## Fill the input structure for deesse and launch deesse
# +
deesse_input = dsi.DeesseInput(
nx=nx, ny=ny, nz=nz,
sx=sx, sy=sy, sz=sz,
ox=ox, oy=oy, oz=oz,
nv=1, varname='categ',
nTI=1, TI=ti,
mask=mask_value, # set mask
distanceType='categorical',
nneighboringNode=24,
distanceThreshold=0.05,
maxScanFraction=0.25,
npostProcessingPathMax=1,
seed=444,
nrealization=1)
deesse_output = dsi.deesseRun(deesse_input)
# -
# ### Display the "mask" and the simulation
# +
# Retrieve the result
sim = deesse_output['sim']
# Display
plt.subplots(1, 2, figsize=(17,5)) # 1 x 2 sub-plots
# ... the mask
plt.subplot(1, 2, 1)
imgplt.drawImage2D(mask, categ=True, categCol=['gray', 'lightgreen'], title='Mask')
# ... the simulation
plt.subplot(1, 2, 2)
imgplt.drawImage2D(sim[0], categ=True, categCol=col, title='Sim. using a mask')
| examples/ex_deesse_11_using_mask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating our Database
#
# This notebook walks through how I put together our tables for the MySQL portion of the course.
# +
import pymysql
# Connect to the database
connection = pymysql.connect(host='',
cursorclass=pymysql.cursors.DictCursor)
cursor = connection.cursor()
# -
# ## Dropping Existing Tables
# Generally a good first step for setting up a database is to remove everything that is already there. This avoids any potential conflicts on creations/inserts.
#
# Therefore we first **DROP** all of the tables we'll be using if they exist.
"""Drop tables"""
tables = ['friends', 'favorites', 'retweets', 'statuses', 'users']
for table in tables:
print(table)
drop_query = f"""DROP TABLE IF EXISTS {table};"""
cursor.execute(drop_query)
connection.commit()
# ## Creating Our Tables
# Once we've cleared things out, we can start creating the tables we'll be using.
#
# We create a table by calling **CREATE TABLE *table_name*(*colls*)**. One thing to keep in mind is the use of **PRIMARY KEY(*coll*)**, this simply identifies the column that will be used as the primary key.
# +
"""Create users table to track twitter users"""
make_user_table = """CREATE TABLE users(
created_date DATE,
description VARCHAR(255),
favorites_count INT,
friends_count INT,
user_id VARCHAR(255),
listed_count INT,
location VARCHAR(255),
name VARCHAR(255),
screen_name VARCHAR(255),
statuses_count INT,
url VARCHAR(255),
verified BOOL,
PRIMARY KEY(user_id));"""
cursor.execute(make_user_table)
connection.commit()
# -
# ## Foreign Keys
# All subsequent table creation queries will look relatively the same, the key difference is the **FOREIGN KEY(*coll*)** statement. This statement defines a relationship to another table (which is a hard rule - no insertions that don't meet constraints).
#
# Notes:
# - **REFERENCES *table_name(coll in table)**: This details where the foreign relationship can be found
# - **ON DELETE CASCADE**: This isn't required, but states that if the foreign key is removed, also remove this record
# +
"""Drop and create friends table to track twitter friends"""
make_friends_table = """CREATE TABLE friends(
user_id VARCHAR(255),
user_screen_name VARCHAR(255),
friend_id VARCHAR(255),
friend_screen_name VARCHAR(255),
PRIMARY KEY (user_id, friend_id),
FOREIGN KEY (user_id)
REFERENCES users(user_id)
ON DELETE CASCADE,
FOREIGN KEY (friend_id)
REFERENCES users(user_id)
ON DELETE CASCADE);"""
cursor.execute(make_friends_table)
connection.commit()
# -
# Now we can simply create the remaining tables:
# +
"""Drop and create friends table to track twitter friends"""
make_status_table = """CREATE TABLE statuses(
created_date DATE,
favorites_count INT,
status_id VARCHAR(255),
lang VARCHAR(255),
retweet_count INT,
source VARCHAR(255),
text TEXT,
truncated BOOL,
user_id VARCHAR(255),
PRIMARY KEY (status_id),
FOREIGN KEY (user_id)
REFERENCES users(user_id)
ON DELETE CASCADE);"""
cursor.execute(make_status_table)
connection.commit()
"""Drop and create favorites table to track favorited tweets"""
make_favs_table = """CREATE TABLE favorites(
status_id VARCHAR(255),
user_id VARCHAR(255),
user_screen_name VARCHAR(255),
PRIMARY KEY (status_id, user_id),
FOREIGN KEY (user_id)
REFERENCES users(user_id)
ON DELETE CASCADE,
FOREIGN KEY (status_id)
REFERENCES statuses(status_id)
ON DELETE CASCADE);"""
cursor.execute(make_favs_table)
connection.commit()
"""Drop and create favorites table to track favorited tweets"""
make_retweets_table = """CREATE TABLE retweets(
created_date DATE,
status_id VARCHAR(255),
retweeted_status VARCHAR(255),
lang VARCHAR(255),
retweet_count INT,
source VARCHAR(255),
text TEXT,
user_id VARCHAR(255),
PRIMARY KEY (status_id),
FOREIGN KEY (user_id)
REFERENCES users(user_id)
ON DELETE CASCADE,
FOREIGN KEY (retweeted_status)
REFERENCES statuses(status_id)
ON DELETE CASCADE);"""
cursor.execute(make_retweets_table)
connection.commit()
| assets/EMSE6586/MySQL Table Creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
from bqplot import Pie, Figure
import numpy as np
import string
# + [markdown] deletable=true editable=true
# ## Basic Pie Chart
# + deletable=true editable=true
data = np.random.rand(3)
pie = Pie(sizes=data, display_labels='outside', labels=list(string.ascii_uppercase))
fig = Figure(marks=[pie], animation_duration=1000)
fig
# + [markdown] deletable=true editable=true
# ### Update Data
# + deletable=true editable=true
n = np.random.randint(1, 10)
pie.sizes = np.random.rand(n)
# + [markdown] deletable=true editable=true
# ### Display Values
# + deletable=true editable=true
with pie.hold_sync():
pie.display_values = True
pie.values_format = '.1f'
# + [markdown] deletable=true editable=true
# ### Enable sort
# + deletable=true editable=true
pie.sort = True
# + [markdown] deletable=true editable=true
# ### Set different styles for selected slices
# + deletable=true editable=true
pie.selected_style = {'opacity': 1, 'stroke': 'white', 'stroke-width': 2}
pie.unselected_style = {'opacity': 0.2}
pie.selected = [1]
# + deletable=true editable=true
pie.selected = None
# + [markdown] deletable=true editable=true
# For more on piechart interactions, see the [Mark Interactions notebook](../Interactions/Mark Interactions.ipynb)
# + [markdown] deletable=true editable=true
# ### Modify label styling
# + deletable=true editable=true
pie.label_color = 'Red'
pie.font_size = '20px'
pie.font_weight = 'bold'
# + [markdown] deletable=true editable=true
# ### Update pie shape and style
# + deletable=true editable=true
pie1 = Pie(sizes=np.random.rand(6), inner_radius=0.05)
fig1 = Figure(marks=[pie1], animation_duration=1000)
fig1
# + [markdown] deletable=true editable=true
# ### Change pie dimensions
# + deletable=true editable=true
# As of now, the radius sizes are absolute, in pixels
with pie1.hold_sync():
pie1.radius = 150
pie1.inner_radius = 100
# + deletable=true editable=true
# Angles are in radians, 0 being the top vertical
with pie1.hold_sync():
pie1.start_angle = -90
pie1.end_angle = 90
# + [markdown] deletable=true editable=true
# ### Move the pie around
# `x` and `y` attributes control the position of the pie in the figure.
# If no scales are passed for `x` and `y`, they are taken in absolute
# figure coordinates, between 0 and 1.
# + deletable=true editable=true
pie1.y = 0.1
pie1.x = 0.6
pie1.radius = 180
# + [markdown] deletable=true editable=true
# ### Change slice styles
# Pie slice colors cycle through the `colors` and `opacities` attribute, as the `Lines` Mark.
# + deletable=true editable=true
pie1.stroke = 'brown'
pie1.colors = ['orange', 'darkviolet']
pie1.opacities = [.1, 1]
fig1
# + [markdown] deletable=true editable=true
# ## Represent an additional dimension using Color
# + [markdown] deletable=true editable=true
# The `Pie` allows for its colors to be determined by data, that is passed to the `color` attribute.
# A `ColorScale` with the desired color scheme must also be passed.
# + deletable=true editable=true
from bqplot import ColorScale, ColorAxis
Nslices = 7
size_data = np.random.rand(Nslices)
color_data = np.random.randn(Nslices)
sc = ColorScale(scheme='Reds')
# The ColorAxis gives a visual representation of its ColorScale
ax = ColorAxis(scale=sc)
pie2 = Pie(sizes=size_data, scales={'color': sc}, color=color_data)
Figure(marks=[pie2], axes=[ax])
# + [markdown] deletable=true editable=true
# ## Position the Pie using custom scales
# + [markdown] deletable=true editable=true
# Pies can be positioned, via the `x` and `y` attributes,
# using either absolute figure scales or custom 'x' or 'y' scales
# + deletable=true editable=true
from datetime import datetime
from bqplot.traits import convert_to_date
from bqplot import DateScale, LinearScale, Axis
avg_precipitation_days = [(d/30., 1-d/30.) for d in [2, 3, 4, 6, 12, 17, 23, 22, 15, 4, 1, 1]]
temperatures = [9, 12, 16, 20, 22, 23, 22, 22, 22, 20, 15, 11]
dates = [datetime(2010, k, 1) for k in range(1, 13)]
sc_x = DateScale()
sc_y = LinearScale()
ax_x = Axis(scale=sc_x, label='Month', tick_format='%b')
ax_y = Axis(scale=sc_y, orientation='vertical', label='Average Temperature')
pies = [Pie(sizes=precipit, x=date, y=temp,display_labels='none',
scales={'x': sc_x, 'y': sc_y}, radius=30., stroke='navy',
apply_clip=False, colors=['navy', 'navy'], opacities=[1, .1])
for precipit, date, temp in zip(avg_precipitation_days, dates, temperatures)]
Figure(title='Kathmandu Precipitation', marks=pies, axes=[ax_x, ax_y],
padding_x=.05, padding_y=.1)
| examples/Marks/Object Model/Pie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # [PUBLIC] Analysis of CLBlast tuning
# + [markdown] deletable=true editable=true
# <a id="overview"></a>
# ## Overview
# + [markdown] deletable=true editable=true
# This Jupyter Notebook analyses the performance that CLBlast achieves across a range of routines, sizes and configurations.
#
# Run first clblast-tuning-benchmarking.py
# + [markdown] deletable=true editable=true
# <a id="data"></a>
# ## Get the experimental data from DropBox
# + [markdown] deletable=true editable=true
# **NB:** Please ignore this section if you are not interested in re-running or modifying this notebook.
# + [markdown] deletable=true editable=true
# The experimental data was collected on the experimental platform and archived as follows:
# ```
# $ cd `ck find ck-math:script:<...>`
# $ python <...>.py
# $ ck zip local:experiment:* --archive_name=<...>.zip
# ```
#
# It can be downloaded and extracted as follows:
# ```
# $ wget <...>.zip
# $ ck add repo:<....> --zip=<....>.zip --quiet
# ```
# + [markdown] deletable=true editable=true
# <a id="code"></a>
# ## Data wrangling code
# + [markdown] deletable=true editable=true
# **NB:** Please ignore this section if you are not interested in re-running or modifying this notebook.
# + [markdown] deletable=true editable=true
# ### Includes
# + [markdown] deletable=true editable=true
# #### Standard
# + deletable=true editable=true
import os
import sys
import json
import re
# + [markdown] deletable=true editable=true
# #### Scientific
# + [markdown] deletable=true editable=true
# If some of the scientific packages are missing, please install them using:
# ```
# # pip install jupyter pandas numpy matplotlib
# ```
# + deletable=true editable=true
import IPython as ip
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib as mp
# + deletable=true editable=true
print ('IPython version: %s' % ip.__version__)
print ('Pandas version: %s' % pd.__version__)
print ('NumPy version: %s' % np.__version__)
print ('Seaborn version: %s' % sns.__version__) # apt install python-tk
print ('Matplotlib version: %s' % mp.__version__)
# + deletable=true editable=true
import matplotlib.pyplot as plt
from matplotlib import cm
# %matplotlib inline
# + deletable=true editable=true
from IPython.display import Image
from IPython.core.display import HTML
# + [markdown] deletable=true editable=true
# #### Collective Knowledge
# + [markdown] deletable=true editable=true
# If CK is not installed, please install it using:
# ```
# # pip install ck
# ```
# + deletable=true editable=true
import ck.kernel as ck
print ('CK version: %s' % ck.__version__)
# + [markdown] deletable=true editable=true
# ### Define helper functions
# + deletable=true editable=true
# Return the number of floating-point operations for C = alpha * A * B + beta * C,
# where A is a MxK matrix and B is a KxN matrix.
def xgemm_flops(alpha, beta, M, K, N):
flops_AB = 2*M*N*K if alpha!=0 else 0
flops_C = 2*M*N if beta!=0 else 0
flops = flops_AB + flops_C
return flops
# + deletable=true editable=true
# Return GFLOPS (Giga floating-point operations per second) for a known kernel and -1 otherwise.
def GFLOPS(kernel, run_characteristics, time_ms):
if kernel.lower().find('xgemm') != -1:
time_ms = np.float64(time_ms)
alpha = np.float64(run_characteristics['arg_alpha'])
beta = np.float64(run_characteristics['arg_beta'])
M = np.int64(run_characteristics['arg_m'])
K = np.int64(run_characteristics['arg_k'])
N = np.int64(run_characteristics['arg_n'])
return (1e-9 * xgemm_flops(alpha, beta, M, K, N)) / (1e-3 * time_ms)
else:
return (-1.0)
# + deletable=true editable=true
def args_str(kernel, run):
args = ''
if kernel.lower().find('xgemm') != -1:
args = 'alpha=%s, beta=%s, M=%s, K=%s, N=%s' % \
(run['arg_alpha'], run['arg_beta'], run['arg_m'], run['arg_k'], run['arg_n'])
return args
# + [markdown] deletable=true editable=true
# ### Access the experimental data
# + deletable=true editable=true
def get_experimental_results(repo_uoa='local', tags='explore-clblast-matrix-size'):
module_uoa = 'experiment'
r = ck.access({'action':'search', 'repo_uoa':repo_uoa, 'module_uoa':module_uoa, 'tags':tags})
if r['return']>0:
print ("Error: %s" % r['error'])
exit(1)
experiments = r['lst']
dfs = []
for experiment in experiments:
print experiment
data_uoa = experiment['data_uoa']
r = ck.access({'action':'list_points', 'repo_uoa':repo_uoa, 'module_uoa':module_uoa, 'data_uoa':data_uoa})
if r['return']>0:
print ("Error: %s" % r['error'])
exit(1)
for point in r['points']:
with open(os.path.join(r['path'], 'ckp-%s.0001.json' % point)) as point_file:
point_data_raw = json.load(point_file)
characteristics_list = point_data_raw['characteristics_list']
num_repetitions = len(characteristics_list)
# Obtain column data.
data = [
{
'repetition_id': repetition_id,
'strategy' : tuner_output['strategy'],
'config_id': config_id,
'config' : config['parameters'],
'kernel' : config['kernel'],
'args_id' : args_str(config['kernel'], characteristics['run']),
'ms' : np.float64(config['time']),
'GFLOPS' : GFLOPS(config['kernel'], characteristics['run'], config['time'])
}
for (repetition_id, characteristics) in zip(range(num_repetitions), characteristics_list)
for tuner_output in characteristics['run']['data']
for (config_id, config) in zip(range(len(tuner_output['result'])), tuner_output['result'])
]
# Construct a DataFrame.
df = pd.DataFrame(data)
# Set columns and index names.
df.columns.name = 'characteristics'
df.index.name = 'index'
df = df.set_index([ 'kernel', 'strategy', 'args_id', 'config_id', 'repetition_id' ])
# Append to the list of similarly constructed DataFrames.
dfs.append(df)
# Concatenate all constructed DataFrames (i.e. stack on top of each other).
result = pd.concat(dfs)
return result.sortlevel(result.index.names)
# + deletable=true editable=true
df = get_experimental_results(tags='explore-clblast-matrix-size,xgemm-fp32')
pd.options.display.max_columns = len(df.columns)
pd.options.display.max_rows = len(df.index)
# + deletable=true editable=true
kernel0 = df.iloc[0].name[0]
kernel0
# + deletable=true editable=true
# NB: Unlike mean(), mean() retains the 'config' column.
df_kernel0 = df.groupby(level=df.index.names[:-1]).min().loc[kernel0]
# + deletable=true editable=true
df_kernel0.groupby(level=df_kernel0.index.names[:-1])['GFLOPS'].min()
# + deletable=true editable=true
df_kernel0.groupby(level=df_kernel0.index.names[:-1])['GFLOPS'].max()
# + deletable=true editable=true
max_GFLOPS = df_kernel0.loc[df_kernel0['GFLOPS'].argmax()]['GFLOPS']
max_GFLOPS
# + deletable=true editable=true
max_GLOPS_config = df_kernel0.loc[df_kernel0['GFLOPS'].argmax()]['config']
max_GLOPS_config
# -
best_configs = df_kernel0.loc[df_kernel0.groupby(level=df_kernel0.index.names[:-1])['GFLOPS'].idxmax()]['config']
idx = df_kernel0.groupby(level=df_kernel0.index.names[:-1])['GFLOPS'].idxmax()
my = df_kernel0.loc[idx]['config']
for i in my:
print i
# + [markdown] deletable=true editable=true
# #### Plot a violin graph
# + deletable=true editable=true
plt.figure(figsize=(12, 10))
sns.set_style('whitegrid'); sns.set_palette('Set1')
ax = sns.violinplot(data=df_kernel0.reset_index(), x='GFLOPS', y='args_id',
split=True, hue='strategy', hue_order=['random', 'exhaustive'])
ax.set_xticks(range(0, int(max_GFLOPS), 1))
ax.set_xlim([0, max_GFLOPS])
# Draw a dotted purple line from top to bottom at the default value (TODO).
ax.vlines(linestyles='dotted', colors='purple', x=124, ymin=ax.get_ylim()[0], ymax=ax.get_ylim()[1])
# -
| script/explore-clblast-matrix-size/clblast-distribution-tuner-sizes-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find population of a US CITY based on
#
# ALAND
# AWATER
# LAT
# LONG
# STATE
# UAtype
#
# https://www.census.gov/geo/maps-data/data/gazetteer2010.html
#
# steps to train a DNN
# 1. Prepare data
# _ clean data
# _ vectorize/encode data
# _ split data in train,validation and test data
# 2. Train model
# _ create model and layers
# _ compile mode(optimizer , metrics, loss)
# _ fit model(train data, validation data, epochs, mini batch size)
# 3. Validate model
# _ validate on test data
# 4. Repeat
#
import pandas as pd
import numpy as np
#prepare data
city_data = []
#00037 Abbeville, LA Urban Cluster C 19824 8460 29222871 300497 11.283 0.116 29.967602 -92.098219
def parseline(line):
parts = line.split('\t')
data={}
namepart = parts[1].split(',')
data['NAME']=namepart[0]
data['STATE']=namepart[1].strip().split()[0].strip()
data['UAtype']=parts[2]
data['POP10']= int(parts[3])
data['ALAND']= int(parts[5])
data['AWATER']= int(parts[6])
data['LAT']=float(parts[9])
data['LONG']=float(parts[10])
return data
with open('Gaz_ua_national.txt','r') as file:
city_data = [ parseline(line.strip()) for line in file.readlines()[1:]]
city_data[-1]
len(city_data)
df = pd.DataFrame(city_data)
df = df.sample(len(df))#shuffle
df.head()
# +
def to_numerical(labels):
numerical_dict={}
for label in labels:
if label not in numerical_dict:
numerical_dict[label]=len(numerical_dict)
return numerical_dict
def to_one_hot(labels):
numerical = to_numerical(labels)
one_hot = np.zeros((len(labels),len(numerical)))
for i,label in enumerate(labels):
one_hot[i,numerical[label]]=1
return np.asarray(one_hot)
# +
#normalise data
train, test = (0.8,0.2)
numeric_data = np.asarray(df[['ALAND','AWATER','LAT','LONG']].values)
print(numeric_data[0])
mean=[]
std=[]
def normalize_data(numeric_data,fraction):
global mean, std
train_size = int(len(numeric_data)*fraction)
mean = numeric_data[:train_size].mean(axis=0)
std = numeric_data[:train_size].std(axis=0)
return (numeric_data - mean)/std
numeric_data = normalize_data(numeric_data,train)
# -
#encode data
states = to_one_hot(df['STATE'].values)
UA = to_one_hot(df['UAtype'].values)
numeric_data = np.concatenate((numeric_data,states,UA),axis=1)
#merge all data
population = np.asarray(df['POP10'].values).astype(np.float32)/1000#population in thousands
population = population.reshape(len(population),1)
population.min()
numeric_data.shape
# +
#split data
train_f, test_f = (0.8,0.2)
train = int(train_f*len(numeric_data))
x_train_all,x_test = (numeric_data[:train],numeric_data[train:])
y_train_all,y_test = (population[:train],population[train:])
print(x_train_all.shape)
folds = 5
sample_per_fold = train//folds #integer division
print ('Samples per fold = {0}, Number of folds= {1}'.format(sample_per_fold,folds))
# +
from keras import Sequential
from keras import layers
from keras import models
from keras import losses
from keras import optimizers
from keras import metrics
def createmodel(nodes):
model = Sequential()
model.add(layers.Dense(nodes[0],activation='relu',input_shape=(x_train.shape[1],)))
for layer in range(len(nodes)-1):
model.add(layers.Dense(nodes[layer+1],activation='relu'))
model.add(layers.Dense(1))#relu activation to avoid negetives produces bad results here
model.compile(optimizer='rmsprop',loss=['mape'],metrics=['mae'])
return model
# +
all_train_mae = []
all_val_mae = []
import matplotlib.pyplot as pp
# %matplotlib inline
def runkfold(l):
global all_train_mae
global all_val_mae
all_train_mae = []
all_val_mae = []
for i in range(folds):
print ('Processing fold # ',i)
x_validate = x_train_all[i*sample_per_fold : (i+1)*sample_per_fold]
y_validate = y_train_all[i*sample_per_fold : (i+1)*sample_per_fold]
x_train = np.concatenate([x_train_all[:i*sample_per_fold],
x_train_all[(i+1)*sample_per_fold:]])
y_train = np.concatenate([y_train_all[:i*sample_per_fold],
y_train_all[(i+1)*sample_per_fold:]])
model = createmodel(l)
history = model.fit(x_train,y_train,
validation_data=(x_validate,y_validate),
epochs=150,
batch_size = 50,verbose=0)
all_train_mae.append(history.history['mean_absolute_error'])
all_val_mae.append(history.history['val_mean_absolute_error'])
# -
def ploterrors():
pp.figure()
train_mae = np.asarray(all_train_mae).mean(axis=0)
val_mae = np.asarray(all_val_mae).mean(axis=0)
pp.plot(train_mae,label='train mae')
pp.plot(val_mae,label='val mae all')
for i in range(folds):
pp.plot(all_val_mae[i],label='val mae ' + str(i),alpha=0.3)
pp.xlabel('epochs')
pp.ylabel('mae')
pp.legend()
layers_all = [[64],[64,32],[64,32,16],[32,16,16,8]]
for layer in layers_all:
print (layer)
runkfold(layer)
ploterrors()
pp.title('mae vs epoch {}'.format( layer))
pp.show()
# +
#[64,32,16] is the best layer combinaiton
model = createmodel([64, 32, 16])
history = model.fit(x_train_all,y_train_all,
epochs=150,
batch_size = 50, verbose=0)
#evaluate
model.evaluate(x_test,y_test)
# -
def seepredictions(model):
df_test = df.loc[df.index[train:]].copy()
#df.tail(len(df)-train).head(validation) #this should work too
pred = model.predict(x_test)*1000
df_test['Predictions']=pred
df_test['ErrorPercentage']=abs(df_test['Predictions']-df_test['POP10'])/df_test['POP10']*100
return df_test
print('mean percentage diff = ', seepredictions(model)['ErrorPercentage'].mean())
test_df = seepredictions(model)
print ('Out of {} test data, {} are fluctuating by more than 100%'.format(len(test_df),len(test_df[test_df['ErrorPercentage']>100].index)))
test_df[test_df['POP10']>100000].sample(5)
test_df[test_df['POP10']<5000].sample(5)
| Keras-train-MAPE-kfold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# #### Imports
import pandas as pd
import sagemaker
# #### Explore data
df = pd.read_csv('./data/medical_transcripts.csv')
df.head()
df.shape
df['transcription'].tolist()[0]
set(df['specialty'].tolist())
# #### Encode data to be Jumpstart ready
label_map = {'Allergy_Immunology': 0,
'Bariatrics': 1,
'Cardiovascular_Pulmonary': 2,
'Dentistry': 3,
'General_Medicine': 4,
'Neurology': 5,
'Neurosurgery': 6,
'Obstetrics_Gynecology': 7,
'Office_Notes': 8,
'Ophthalmology': 9,
'Orthopedic': 10,
'Pain_Management': 11,
'Pediatrics_Neonatal': 12,
'Physical_Medicine_Rehab': 13,
'Podiatry': 14,
'Psychiatry_Psychology': 15,
'Radiology': 16,
'Rheumatology': 17,
'SOAP_Chart_Progress_Notes': 18,
'Sleep_Medicine': 19,
'Speech_Language': 20,
'Surgery': 21,
'Urology': 22
}
inverse_label_map = {0: 'Allergy_Immunology',
1: 'Bariatrics',
2: 'Cardiovascular_Pulmonary',
3: 'Dentistry',
4: 'General_Medicine',
5: 'Neurology',
6: 'Neurosurgery',
7: 'Obstetrics_Gynecology',
8: 'Office_Notes',
9: 'Ophthalmology',
10: 'Orthopedic',
11: 'Pain_Management',
12: 'Pediatrics_Neonatal',
13: 'Physical_Medicine_Rehab',
14: 'Podiatry',
15: 'Psychiatry_Psychology',
16: 'Radiology',
17: 'Rheumatology',
18: 'SOAP_Chart_Progress_Notes',
19: 'Sleep_Medicine',
20: 'Speech_Language',
21: 'Surgery',
22: 'Urology'
}
encoded_df = df.replace({'specialty': label_map})
encoded_df.head()
encoded_df.to_csv('./data/data.csv', header=False, index=False)
# ##### Check encoded csv
df = pd.read_csv('./data/data.csv', names=['label', 'transcript'])
df.head()
inverse_label_map[df['label'][0]]
# #### Copy dataset from local to S3
session = sagemaker.Session()
default_bucket = session.default_bucket()
print(f'Default S3 bucket = {default_bucket}')
# Remove checkpoints if any
# !rm -rf ./data/.ipynb_checkpoints/
# !aws s3 cp ./data/data.csv s3://{default_bucket}/transcripts/data.csv
| hcls/language/medical-transcripts-classification/explore-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 5: Regime Prediction with Machine Learning - Part 3
#
# In this part we are going to predict economic regimes with machine learning algorithms using Pyhton's Scikit Learn package.
#
# ## Table of Contents:
# 1. [Set Up Environment and Read Data](#1)
#
# 2. [Methodology](#2)
#
# 3. [Feature Selection](#3)
#
# 4. [Traning Algorithms on Training Dataset](#4)
#
# 5. [Evaluate Performances of the Algorithms on Validation Dataset](#5)
# ## 1. Set Up Environment and Read Data <a id="1"></a>
# +
# if xgboost is not installed you can run this command in the cell
# #!pip install xgboost
#load libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import metrics
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import f1_score
from sklearn.metrics import roc_auc_score, roc_curve, auc
from sklearn import model_selection
from sklearn import preprocessing
from sklearn.model_selection import TimeSeriesSplit
from sklearn.feature_selection import SelectFromModel
from sklearn.linear_model import LogisticRegression
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
import xgboost as xgb
from matplotlib import pyplot as mp
import seaborn as sns
import os
import warnings
warnings.filterwarnings('ignore')
# +
df=pd.read_csv('Dataset_Cleaned.csv')
Label = df["Regime"].apply(lambda regime: 1. if regime == 'Normal' else 0.)
df.insert(loc=2, column="Label", value=Label.values)
df.head()
# -
df.tail()
# ## 2. Methodology <a id="2"></a>
# Our exercise will be based on classification problem. We have two binary outcomes that we want to predict with certain variables. Here we will summarize our approach to predict recessions with machine learning algorithms.
#
# 1. We will perform feature selection before making our forecasts. We will use $L_1$ regularized logistic regression for that purpose.
#
# 2. Separate dataset into training and validation datasets. Split based dataset based on time: the period over 1960-1996 is selected for training and the period over 1996-2018 is kept for validation
#
# 3. Evaluate performances of the machine learning algorithms on training dataset with cross validation (CV). Since we have time series structure we will use a special type of CV function in Python,__[`TimeSeriesSplit`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.TimeSeriesSplit.html#sklearn.model_selection.TimeSeriesSplit)__. We will use Receiver operating characteristic (ROC) as scoring metric in our models. Related Python functions for this metric are __[`roc_auc_score`](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html#sklearn.metrics.roc_auc_score)__ and __[`roc_curve`](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_curve.html#sklearn.metrics.roc_curve)__.
#
# 4. Select the best performing models based on average accuracy and standard deviation of the CV results. We will take logistic regression as a benchmark model since this is the traditional method has been used to approach this problem.
#
# 5. Then we make predictions on the validation dataset with selected models. First, we use __[`GridSearchCV`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html)__ for selected model on training dataset to find best combination of parameters for the model. Then we evaluate the model on validation dataset and report accuracy metrics and feature importance results.
# ## 3. Feature Selection with $L_1$ Penalty <a id="3"></a>
# +
# Time Series Split
df_idx = df[df.Date == '12/1/96'].index[0]
df_targets=df['Label'].values
df_features=df.drop(['Regime','Date','Label'], axis=1)
df_training_features = df.iloc[:df_idx,:].drop(['Regime','Date','Label'], axis=1)
df_validation_features = df.iloc[df_idx:, :].drop(['Regime','Date','Label'], axis=1)
df_training_targets = df['Label'].values
df_training_targets=df_training_targets[:df_idx]
df_validation_targets = df['Label'].values
df_validation_targets=df_validation_targets[df_idx:]
# -
print(len(df_training_features),len(df_training_targets),len(df_targets))
print(len(df_validation_features),len(df_validation_targets),len(df_features))
# +
scoring="roc_auc"
kfold= model_selection.TimeSeriesSplit(n_splits=3)
seed=8
# Create regularization hyperparameter space
C = np.reciprocal([0.00000001, 0.00000005, 0.0000001, 0.0000005, 0.000001, 0.000005, 0.00001, 0.00005,
0.0001, 0.0005, 0.001, 0.005, 0.01, 0.05, 0.1, 0.5, 1, 5, 10, 50, 100, 500, 1000, 5000])
# Create hyperparameter options
hyperparameters = dict(C=C)
model=LogisticRegression(max_iter=10000, solver = 'liblinear', penalty='l1')
LR_penalty=model_selection.GridSearchCV(estimator=model, param_grid= hyperparameters,
cv=kfold, scoring=scoring).fit(X=df_features,
y=df_targets).best_estimator_
LR_penalty
# -
X=df_training_features
y=df_training_targets
lr_l1 = LogisticRegression(C=0.1, max_iter=10000, solver = 'liblinear', penalty="l1").fit(X,y) #change solver
model = SelectFromModel(lr_l1,prefit=True)
feature_idx = model.get_support()
feature_name = X.columns[feature_idx]
X_new = model.transform(X)
X_new.shape
feature_name
# +
df_2=df[feature_name]
df_2.insert(loc=0, column="Date", value=df['Date'].values)
df_2.insert(loc=1, column="Regime", value=df['Regime'].values)
df_2.insert(loc=2, column="Label", value=df['Label'].values)
df_2.head()
df_2.shape
# -
corr = df_2.drop(['Date','Regime','Label'],axis=1).corr()
plt.figure(figsize=(10, 8))
sns.heatmap(corr, mask=np.zeros_like(corr, dtype=np.bool),
cmap=sns.diverging_palette(220, 10, as_cmap=True), square=True)
# ## 4. Training Algorithms on Training Dataset <a id="3"></a>
# For a detail description of the machine learning algorithms you can check scikit-learn's documentation __[here](https://scikit-learn.org/stable/supervised_learning.html#supervised-learning)__.
df=df_2
df.shape
# +
# Time Series Split
df_idx = df[df.Date == '12/1/96'].index[0]
df_targets=df['Label'].values
df_features=df.drop(['Regime','Date','Label'], axis=1)
df_training_features = df.iloc[:df_idx,:].drop(['Regime','Date','Label'], axis=1)
df_validation_features = df.iloc[df_idx:, :].drop(['Regime','Date','Label'], axis=1)
df_training_targets = df['Label'].values
df_training_targets=df_training_targets[:df_idx]
df_validation_targets = df['Label'].values
df_validation_targets=df_validation_targets[df_idx:]
# +
seed=8
scoring='roc_auc'
kfold = model_selection.TimeSeriesSplit(n_splits=3)
models = []
models.append(('LR', LogisticRegression(C=1e09)))
models.append(('LR_L1', LogisticRegression(penalty = 'l1', solver = "liblinear")))
models.append(('LR_L2', LogisticRegression(penalty = 'l2')))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('GB', GradientBoostingClassifier()))
models.append(('ABC', AdaBoostClassifier()))
models.append(('RF', RandomForestClassifier()))
models.append(('XGB', xgb.XGBClassifier()))
results = []
names = []
lb = preprocessing.LabelBinarizer()
for name, model in models:
cv_results = model_selection.cross_val_score(estimator = model, X = df_training_features,
y = lb.fit_transform(df_training_targets), cv=kfold, scoring = scoring)
model.fit(df_training_features, df_training_targets) # train the model
fpr, tpr, thresholds = metrics.roc_curve(df_training_targets, model.predict_proba(df_training_features)[:,1])
auc = metrics.roc_auc_score(df_training_targets,model.predict(df_training_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % (name, auc))
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([-0.05, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('1-Specificity(False Positive Rate)')
plt.ylabel('Sensitivity(True Positive Rate)')
plt.title('Receiver Operating Characteristic')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.show()
warnings.filterwarnings('ignore')
# -
fig = plt.figure()
fig.suptitle('Algorithm Comparison based on Cross Validation Scores')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
# ## 4. Evaluate Performances of the Algorithms on Validation Dataset <a id="4"></a>
# ### Logistic Regression
# Logistic regression is the most commonly used statistical model for binary classification. It uses the logit model of relating log-odds of the dependent variable linearly with the predictor (explanatory) variables to learn a form of the following logistic function that is used to separate instances of the two different classes.
#
# \begin{align*}
# Pr(y=1|x) = h_\beta(x) = \frac{1}{1 + e^{-(\beta_0 + \beta_1 x)}} \text{ where $\beta_0$ is the intercept and $\beta$ is the vector of trained weights}
# \end{align*}
#
# The function returns a probability measure of which class a new instance is given its features, this measure is then used to make the final classification with some probability threshold, traditionally being 0.5.
#
# The specific logistic function is learned through gradient descent which focuses on minimizing error calculated through some cost function. A typical approach is to use the following maximum-likelihood estimator to minimize error in predicted probabilities.
#
# \begin{align*}
# & J(\beta) = -\frac{1}{N}\sum_{i=1}^Ny_ilog(h_\beta(x_i)) + (1 - y_i)log(1 - h_\beta(x_i)) \\
# & \text{ where $N$ is the number of observations, $h_\beta(x)$ is as defined above, and $y_i$ is the predicted class}
# \end{align*}
#
# A regularization term
#
# \begin{align*}
# \lambda\sum_{i=1}^N|\beta_i| \text{ where $\lambda$ is a regularization parameter, }
# \end{align*}
#
# is often added to the cost function to prevent overfitting by penalizing large coefficients. This regularization can be L1 or L2 penalty depending on the problem at hand.
model=LogisticRegression(C=1e09) # high penalty
LR=model.fit(df_training_features,df_training_targets)
training_predictions=LR.predict(df_training_features)
prob_predictions = LR.predict_proba(df_training_features)
prob_predictions = np.append(prob_predictions, LR.predict_proba(df_validation_features), axis=0)
prob_predictions.shape
import datetime
# define periods of recession
rec_spans = []
#rec_spans.append([datetime.datetime(1957,8,1), datetime.datetime(1958,4,1)])
rec_spans.append([datetime.datetime(1960,4,1), datetime.datetime(1961,2,1)])
rec_spans.append([datetime.datetime(1969,12,1), datetime.datetime(1970,11,1)])
rec_spans.append([datetime.datetime(1973,11,1), datetime.datetime(1975,3,1)])
rec_spans.append([datetime.datetime(1980,1,1), datetime.datetime(1980,6,1)])
rec_spans.append([datetime.datetime(1981,7,1), datetime.datetime(1982,10,1)])
rec_spans.append([datetime.datetime(1990,7,1), datetime.datetime(1991,2,1)])
rec_spans.append([datetime.datetime(2001,3,1), datetime.datetime(2001,10,1)])
rec_spans.append([datetime.datetime(2007,12,1), datetime.datetime(2009,5,1)])
# +
sample_range = pd.date_range(start='9/1/1960', end='9/1/2018', freq='MS')
plt.figure(figsize=(20,5))
plt.plot(sample_range.to_series().values, prob_predictions[:,0])
for i in range(len(rec_spans)):
plt.axvspan(rec_spans[i][0], rec_spans[i][len(rec_spans[i]) - 1], alpha=0.25, color='grey')
plt.axhline(y=0.5, color='r', ls='dashed', alpha = 0.5)
plt.title('Recession Prediction Probabalities with Logistic Regression')
mp.savefig('plot1.png', bbox_inches='tight')
plt.show()
# -
# ### Logistic Regression with Regularization
# +
# Create regularization penalty space
penalty = ['l1', 'l2']
# Create regularization hyperparameter space
C = np.reciprocal([0.00000001, 0.00000005, 0.0000001, 0.0000005, 0.000001, 0.000005, 0.00001, 0.00005,
0.0001, 0.0005, 0.001, 0.005, 0.01, 0.05, 0.1, 0.5, 1, 5, 10, 50, 100, 500, 1000, 5000])
# Create hyperparameter options
hyperparameters = dict(C=C, penalty=penalty)
model=LogisticRegression(max_iter=10000)
LR_penalty=model_selection.GridSearchCV(estimator=model, param_grid= hyperparameters,
cv=kfold, scoring=scoring).fit(df_training_features,
df_training_targets).best_estimator_
training_predictions=LR_penalty.predict(df_training_features)
# -
prob_predictions = LR_penalty.predict_proba(df_training_features)
prob_predictions = np.append(prob_predictions, LR_penalty.predict_proba(df_validation_features), axis=0)
# +
sample_range = pd.date_range(start='9/1/1960', end='9/1/2018', freq='MS')
plt.figure(figsize=(20,5))
plt.plot(sample_range.to_series().values, prob_predictions[:,0])
for i in range(len(rec_spans)):
plt.axvspan(rec_spans[i][0], rec_spans[i][len(rec_spans[i]) - 1], alpha=0.25, color='grey')
plt.axhline(y=0.5, color='r', ls='dashed', alpha = 0.5)
plt.title('Recession Prediction Probabalities with Regularized Logistic Regression')
mp.savefig('plot2.png', bbox_inches='tight')
plt.show()
# -
# ### XGBoosting
# +
xgboost = model_selection.GridSearchCV(estimator=xgb.XGBClassifier(),
param_grid={'booster': ['gbtree'],
'max_depth':[2,3,5,10],
'learning_rate':[0.01,0.1,1]},
scoring=scoring, cv=kfold).fit(df_training_features,
lb.fit_transform(df_training_targets)).best_estimator_
xgboost.fit(df_training_features, df_training_targets)
# -
prob_predictions = xgboost.predict_proba(df_training_features)
prob_predictions = np.append(prob_predictions, xgboost.predict_proba(df_validation_features), axis=0)
# +
sample_range = pd.date_range(start='9/1/1960', end='9/1/2018', freq='MS')
plt.figure(figsize=(20,5))
plt.plot(sample_range.to_series().values, prob_predictions[:,0])
for i in range(len(rec_spans)):
plt.axvspan(rec_spans[i][0], rec_spans[i][len(rec_spans[i]) - 1], alpha=0.25, color='grey')
plt.axhline(y=0.5, color='r', ls='dashed', alpha = 0.5)
plt.title('Recession Prediction Probabalities with XGBoost')
mp.savefig('plot3.png', bbox_inches='tight')
plt.show()
# +
# find feature importances
headers = df.drop(['Regime','Label', 'Date'], axis=1).columns.values.tolist()
xgboost_importances = pd.DataFrame(xgboost.feature_importances_, index = headers, columns = ['Relative Importance'])
_ = xgboost_importances.sort_values(by = ['Relative Importance'], ascending = False, inplace=True)
xgboost_importances = xgboost_importances[xgboost_importances['Relative Importance']>0].iloc[:20]
# display importances in bar-chart and pie-chart
fig = plt.figure(figsize=(6,6))
plt.xticks(rotation='90')
plt.barh(y=np.arange(len(xgboost_importances)), width=xgboost_importances['Relative Importance'], align='center', tick_label=xgboost_importances.index)
plt.gca().invert_yaxis()
mp.savefig('feature_importance.png', bbox_inches='tight')
plt.show()
# +
fpr, tpr, thresholds = metrics.roc_curve(df_validation_targets, LR.predict_proba(df_validation_features)[:,1])
auc = metrics.roc_auc_score(df_validation_targets,LR.predict(df_validation_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('LR', auc))
fpr, tpr, thresholds = metrics.roc_curve(df_validation_targets, LR_penalty.predict_proba(df_validation_features)[:,1])
auc = metrics.roc_auc_score(df_validation_targets,LR_penalty.predict(df_validation_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('LR_penalty', auc))
fpr, tpr, thresholds = metrics.roc_curve(df_validation_targets, xgboost.predict_proba(df_validation_features)[:,1])
auc = metrics.roc_auc_score(df_validation_targets,xgboost.predict(df_validation_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('XGBoost', auc))
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([-0.05, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('1-Specificity(False Positive Rate)')
plt.ylabel('Sensitivity(True Positive Rate)')
plt.title('Receiver Operating Characteristic (Validation Data)')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
mp.savefig('ROC1.png', bbox_inches='tight')
plt.show()
# +
fpr, tpr, thresholds = metrics.roc_curve(df_targets, LR.predict_proba(df_features)[:,1])
auc = metrics.roc_auc_score(df_targets,LR.predict(df_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('LR', auc))
fpr, tpr, thresholds = metrics.roc_curve(df_targets, LR_penalty.predict_proba(df_features)[:,1])
auc = metrics.roc_auc_score(df_targets,LR_penalty.predict(df_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('LR_penalty', auc))
fpr, tpr, thresholds = metrics.roc_curve(df_targets, xgboost.predict_proba(df_features)[:,1])
auc = metrics.roc_auc_score(df_targets,xgboost.predict(df_features))
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % ('XGBoost', auc))
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([-0.05, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('1-Specificity(True Negative Rate)')
plt.ylabel('Sensitivity(True Positive Rate)')
plt.title('Receiver Operating Characteristic (Whole period)')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
mp.savefig('ROC2.png', bbox_inches='tight')
plt.show()
# -
| Investment Management/Course3/Part 3 - Forecasting Regimes_version_July_2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from urllib.request import ssl, socket
import datetime, smtplib
hostname = "etsy.com"
port = '443'
context = ssl.create_default_context()
with socket.create_connection((hostname, port)) as sock:
with context.wrap_socket(sock, server_hostname = hostname) as ssock:
certificate = ssock.getpeercert()
certExpires = datetime.datetime.strptime(certificate['notAfter'], '%b %d %H:%M:%S %Y %Z')
daysToExpiration = (certExpires - datetime.datetime.now()).days
def send_notification(days_to_expire):
port = 587
smtp_server = "smtp.acmecorp.com"
sender_email = "<EMAIL>"
receiver_email = "<EMAIL>"
password = input("Type your password and press enter: ")
if days_to_expire == 1:
days = "1 day"
else:
days = str(days_to_expire) + " days"
message = """\
Subject: Certificate Expiration
The TLS Certificate for your site expires in {days}"""
context = ssl.create_default_context()
with smtplib.SMTP(smtp_server, port) as server:
server.starttls(context=context)
server.login(sender_email, password)
server.sendmail(sender_email,
receiver_email,
message.format(days=days))
if daysToExpiration == 7 or daysToExpiration == 1:
send_notification(daysToExpiration)
| PythonScripts/TLS Validator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # tsam - Animation of pareto-optimal aggregation
# Date: 02.05.2022
#
# Author: <NAME>
# Import pandas and the relevant time series aggregation class
# %load_ext autoreload
# %autoreload 2
import copy
import os
import pandas as pd
import matplotlib.pyplot as plt
import tsam.timeseriesaggregation as tsam
import tsam.hyperparametertuning as tune
import tqdm
import numpy as np
from matplotlib.animation import FuncAnimation
# %matplotlib inline
import matplotlib
import subprocess
import multiprocessing
import math
matplotlib.rcParams['animation.embed_limit'] = 1000
# ### Input data
# Read in time series from testdata.csv with pandas
raw = pd.read_csv('testdata.csv', index_col = 0)
raw=raw.rename(columns={'T': 'Temperature [°C]', 'Load':'Demand [kW]', 'Wind':'Wind [m/s]', 'GHI': 'Solar [W/m²]'})
# Setup the hyperparameter instance
tunedAggregations = tune.HyperTunedAggregations(
tsam.TimeSeriesAggregation(
raw,
hoursPerPeriod=24,
clusterMethod="hierarchical",
representationMethod="medoidRepresentation",
rescaleClusterPeriods=False,
segmentation=True,
)
)
# Load the resulting combination
results = pd.read_csv(os.path.join("results","paretoOptimalAggregation.csv"),index_col=0)
results["time_steps"] = results["segments"] * results["periods"]
# ## Create the animated aggregations
# Drop all results with timesteps below 1% of the original data set since they are not meaningful.
results = results[results["time_steps"]>80]
# Append the original time series
results=results.append({"segments":24, "periods":365, "time_steps":len(raw)}, ignore_index=True)
# And reverse the order
results=results.iloc[::-1]
# And create a dictionary with all aggregations we want to show in the animation
animation_list = []
for i, index in enumerate(tqdm.tqdm(results.index)):
segments = results.loc[index,:].to_dict()["segments"]
periods = results.loc[index,:].to_dict()["periods"]
# aggregate to the selected set
tunedAggregations._testAggregation(noTypicalPeriods=periods, noSegments=segments)
# and repredict the data
prediction = tunedAggregations.aggregationHistory[-1].predictOriginalData()
# relative reduction of time steps
reduction = 1 - ( float(tunedAggregations._segmentHistory[-1] * tunedAggregations._periodHistory[-1]) /len(raw))
# add a change layer which shows the difference of the latest aggregation to the previous
if i > 0:
# difference
diff_val = previouspredictedPeriods - prediction
# all fields that changed
diff_bool = abs(diff_val)>1e-10
# make sure that when any change is there it gets set to Nan
prediction_diff = copy.deepcopy(prediction)
prediction_diff[diff_bool.max(axis=1)] = np.nan
# what changes ? segments or periods
if segments == animation_list[-1]["Segments"]:
misc = "Clustering periods"
else:
misc = "Clustering segments"
animation_list.append({
"Prediction": prediction_diff,
"Segments": segments,
"Periods": periods,
"Reduction": reduction,
"Misc": "Medoid representation",#misc,
})
animation_list.append({
"Prediction": prediction,
"Segments": segments,
"Periods": periods,
"Reduction": reduction,
"Misc": "Medoid representation",
})
# and set previous prediction preiods
previouspredictedPeriods = prediction
# And then append a last aggregation with the novel duration/distribution represenation
# +
aggregation=tsam.TimeSeriesAggregation(
raw,
hoursPerPeriod=24,
noSegments=segments,
noTypicalPeriods=periods,
clusterMethod="hierarchical",
rescaleClusterPeriods=False,
segmentation=True,
representationMethod="durationRepresentation",
distributionPeriodWise=False
)
animation_list.append({
"Prediction": aggregation.predictOriginalData(),
"Segments": segments,
"Periods": periods,
"Reduction": reduction,
"Misc": "Distribution representation",
})
# -
# ## Create the animation
# Let animation warp - slow in the beginning and slow in the end
# +
iterator = []
for i in range(len(animation_list )):
if i < 1:
iterator+=[i]*100
elif i < 3:
iterator+=[i]*50
elif i < 6:
iterator+=[i]*30
elif i < 20:
iterator+=[i]*10
elif i >= len(animation_list )-1:
iterator+=[i]*150
elif i > len(animation_list )-3:
iterator+=[i]*50
elif i > len(animation_list )-6:
iterator+=[i]*30
else:
iterator+=[i]
# -
# Create the plot and the animation loop
# +
import matplotlib.ticker as tick
fig, axes = plt.subplots(figsize = [7, 5], dpi = 300, nrows = raw.shape[1], ncols = 1)
cmap = plt.cm.get_cmap("Spectral_r").copy()
cmap.set_bad((.7, .7, .7, 1))
for ii, column in enumerate(raw.columns):
data = raw[column]
stacked, timeindex = tsam.unstackToPeriods(copy.deepcopy(data), tunedAggregations.base_aggregation.hoursPerPeriod)
cax = axes[ii].imshow(stacked.values.T, interpolation = 'nearest', vmin = raw[column].min(), vmax = raw[column].max(), origin='lower', cmap = cmap)
axes[ii].set_aspect('auto')
axes[ii].set_ylabel('Hour')
plt.xlabel('Day in the year')
cbar=plt.colorbar(cax, ax=axes[ii], pad=0.01, aspect=7)
cbar.set_label(column, fontsize="small")
fig.suptitle('Time series aggregation by xx.xx %',
y=0.97,
x=0.3,
horizontalalignment="left",)
text=fig.text(
0.27,
0.91,
"with xxx periods and xx segments - Medoid representation",
horizontalalignment="left",
fontsize="small",)
fig.subplots_adjust(right = 1.1, hspace = 0.05)
def animate(iter):
i=iterator[iter]
predictedPeriods = animation_list[i]["Prediction"]
fig.suptitle(
'Time series aggregation by ' + str(round(animation_list[i]["Reduction"]*100,2)) + " % ",
y=0.97,
x=0.3,
horizontalalignment="left",)
text.set_text(
"with " +
str(animation_list[i]["Periods"]) + " periods and " + str(animation_list[i]["Segments"]) + " segments - " + animation_list[i]["Misc"],
)
for ii, column in enumerate(raw.columns):
data = predictedPeriods[column]
stacked, timeindex = tsam.unstackToPeriods(copy.deepcopy(data), tunedAggregations.base_aggregation.hoursPerPeriod)
cax = axes[ii].imshow(stacked.values.T, interpolation = 'nearest', vmin = raw[column].min(), vmax = raw[column].max(), origin='lower', cmap = cmap)
axes[ii].set_aspect('auto')
fig.subplots_adjust(right = 1.0, hspace = 0.05)
ani = FuncAnimation(fig, animate, repeat_delay=600, interval=20, frames=len(iterator) )
# -
# And save as animation parelllized with ffmpeg since the default matplotlib implemenation takes too long. Faster implemntation than matplotib from here: https://stackoverflow.com/a/31315362/3253411
# Parallelize animation to video
def chunks(lst, n):
"""Yield successive n-sized chunks from lst."""
for i in range(0, len(lst), n):
yield lst[i:i + n]
# +
threads = multiprocessing.cpu_count()
frames=[i for i in range(len(iterator))]
# divide the frame equally
i_length=math.ceil(len(frames)/(threads))
frame_sets=list(chunks(frames,i_length))
filenames=[]
for i in range(len(frame_sets)):
filenames.append("_temp_video_chunk_" + str(i) + ".mp4")
# -
def ani_to_mp4(frame_range, filename):
canvas_width, canvas_height = fig.canvas.get_width_height()
# Open an ffmpeg process
outf = os.path.join("results", filename)
cmdstring = ('ffmpeg',
'-y', '-r', '100', # fps
'-s', '%dx%d' % (canvas_width, canvas_height), # size of image string
'-pix_fmt', 'argb', # formats
'-f', 'rawvideo', '-i', '-', # tell ffmpeg to expect raw video from the pipe
'-vcodec', 'mpeg4', outf) # output encoding
p = subprocess.Popen(cmdstring, stdin=subprocess.PIPE)
# Draw frames and write to the pipe
for frame in frame_range:
# draw the frame
animate(frame)
fig.canvas.draw()
# extract the image as an ARGB string
string = fig.canvas.tostring_argb()
# write to pipe
p.stdin.write(string)
# Finish up
p.communicate()
with multiprocessing.Pool() as pool:
pool.starmap(ani_to_mp4, zip(frame_sets, filenames))
filename_list=os.path.join("results", "filenames.txt")
with open(filename_list, "w") as textfile:
for filename in filenames:
textfile.write("file '" + filename + "'\n")
cmdstring = ('ffmpeg', '-y',
'-f', 'concat',
'-safe', '0',
'-i', filename_list,
'-c', 'copy',
os.path.join("results", "animation.mp4") # output encoding
p = subprocess.Popen(cmdstring, stdin=subprocess.PIPE)
# You can also show it inline but it takes quite long.
from IPython.display import HTML
HTML(ani.to_jshtml())
| examples/aggregation_segment_period_animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/zerotodeeplearning/ztdl-masterclasses/blob/master/solutions_do_not_open/Introduction_to_Deep_Learning_with_Keras_solution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="2bwH96hViwS7"
# ## Learn with us: www.zerotodeeplearning.com
#
# Copyright © 2021: Zero to Deep Learning ® Catalit LLC.
# + colab={} colab_type="code" id="bFidPKNdkVPg"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="DvoukA2tkGV4"
# # Introduction to Deep Learning with Keras
# + colab={} colab_type="code" id="N7gPH8x3bpc0"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
# + colab={} colab_type="code" id="ThYOAIfubyax"
url = 'https://raw.githubusercontent.com/zerotodeeplearning/ztdl-masterclasses/master/data/'
# + colab={} colab_type="code" id="0VMfMaGJbytD"
df = pd.read_csv(url + 'geoloc_elev.csv')
# we only use the 2 features that matter
X = df[['lat', 'lon']].values
y = df['target'].values
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size = 0.2, random_state=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" id="b8AjBPDbb9LC" outputId="1591799e-2ef1-41c5-9852-dbacab4f831e"
df.plot.scatter(x='lat', y='lon',
c='target', cmap='bwr');
# + colab={} colab_type="code" id="m_vbwNTKcnQi"
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.optimizers import SGD, Adam
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="botmrbHocrJA" outputId="77ad7d03-8800-40ef-f8f1-925d0f61420c"
model = Sequential([
Dense(1, input_shape=(2,), activation='sigmoid')
])
model.compile(SGD(lr=0.5), 'binary_crossentropy', metrics=['accuracy'])
h = model.fit(X_train, y_train, epochs=10, validation_split=0.1)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="1Ed-jSqncue3" outputId="52169532-bba2-4acb-c1b9-c05f257d9c6c"
pd.DataFrame(h.history).plot(ylim=(-0.05, 1.05));
# + colab={} colab_type="code" id="6htm97pYeJYN"
def score(model):
bm_score = pd.Series(y).value_counts()[0] / len(y)
train_score = model.evaluate(X_train, y_train, verbose=0)[1]
test_score = model.evaluate(X_test, y_test, verbose=0)[1]
print("""Accuracy scores:
Benchmark:\t{:0.3}
Train:\t{:0.3}
Test:\t{:0.3}""".format(bm_score, train_score, test_score))
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="qimJqtCXepPT" outputId="fb769800-03fa-4421-da37-7eda1f1ef8ce"
score(model)
# + colab={} colab_type="code" id="hNTEvUateJVT"
def plot_decision_boundary(model):
hticks = np.linspace(-2, 2, 101)
vticks = np.linspace(-2, 2, 101)
aa, bb = np.meshgrid(hticks, vticks)
ab = np.c_[aa.ravel(), bb.ravel()]
c = model.predict(ab)
cc = c.reshape(aa.shape)
ax = df.plot(kind='scatter', c='target', x='lat', y='lon', cmap='bwr')
ax.contourf(aa, bb, cc, cmap='bwr', alpha=0.5);
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" id="KcQn6NMTeJSH" outputId="3c028d2c-ad3e-4e1f-9f7c-e399df80169c"
plot_decision_boundary(model)
# + [markdown] colab_type="text" id="vlHV6IN7eJBv"
# ### Exercise 1: Deep network
#
# - Extend the neural network defined above by adding a few inner layers.
# - add a few more nodes to the first layer
# - change the activation function of the first layer from `sigmoid` to something else
# - remember that you only need to specify the `input_shape` in the first layer, the others infer it automatically
# - insert at least another layer or more, after the first one
# - regardless of how many layers you have, the last layer (output) should have a single node and a `sigmoid` activation function
#
# Your model should look like:
#
# ```python
# model = Sequential([
# Dense(...),
# ...
# ...
# ])
# ```
#
# - Retrain the model for 20 epochs. Does your model learn to separate the two classes?
# - Display the history as done above
# - Evaluate the score using the `score` function defined above
# - Display the decision boundary using the `plot_decision_boundary`
# - Bonus points if you also calculate the confusion matrix. (hint: the `model.predict` method returns probabilities, so you will need to round the results to the nearest integer before comparing them with the labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 717} colab_type="code" id="P33f3YWNc2KI" outputId="a9cac211-d83c-4dd4-d49f-2f0d0481ba31" tags=["solution", "empty"]
model = Sequential([
Dense(4, input_shape=(2,), activation='tanh'),
Dense(4, activation='tanh'),
Dense(1, activation='sigmoid')
])
model.compile(SGD(lr=0.5), 'binary_crossentropy', metrics=['accuracy'])
h = model.fit(X_train, y_train, epochs=20, validation_split=0.1)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="8tYCWfyHc8TQ" outputId="b19e04b0-40a1-457d-b8c2-0a3cb43abdae" tags=["solution"]
pd.DataFrame(h.history).plot(ylim=(-0.05, 1.05));
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="SVRKDdQdc_Ay" outputId="bc901e82-3859-406a-b384-e6f5c5286273" tags=["solution"]
score(model)
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" id="zfdnXmesexgX" outputId="5b060c4d-93ce-4e77-c88f-d9117d969e5c" tags=["solution"]
plot_decision_boundary(model)
# + tags=["solution"]
from sklearn.metrics import confusion_matrix
# + colab={} colab_type="code" id="AwoPqF1SdGFI" tags=["solution"]
y_pred_proba = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="0Ny35zVWdJnz" outputId="90b38a6a-398e-4025-a819-a86f48a443e3" tags=["solution"]
y_pred_proba[:4]
# + colab={} colab_type="code" id="MA2naWDadQri" tags=["solution"]
y_pred = y_pred_proba.round(0).astype(int)
# + colab={"base_uri": "https://localhost:8080/", "height": 111} colab_type="code" id="xxwc417VduTW" outputId="b2af370f-e69f-4473-ddd6-1c9ff5a4064d" tags=["solution"]
cm = confusion_matrix(y_test, y_pred)
pd.DataFrame(cm,
index=["Miss", "Hit"],
columns=['pred_Miss', 'pred_Hit'])
# + [markdown] colab_type="text" id="Q41gq8m7dxfO"
# ### Exercise 2: Regression
#
# In this exercise we will perform a non-linear regression of a function with 5 input features and 1 output. A detailed explanation of the function can be found [here](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_friedman1.html).
#
# - The data is generated for your convenience and added to a Pandas DataFrame
# - Use `sns.pairplot` to visualize the data. It may be convenient to specify the `x_vars` and `y_vars` arguments so that you only plot the target as a function of the features
# - Define a deep neural network that will be able to learn this function.
# - Specify the `input_shape` in the first layer
# - Use at least 2 layers and a few nodes to allow the network to learn nonlinear relations
# - The output layer should have a single node and no activation function, as we are doing a regression
#
# ```python
# model = Sequential([
# Dense(...)
# ....
# ])
# ```
# - Compile the model
# - use an optimizer of your choice
# - make sure to select the appropriate loss function for a regression
# - since it's a regression, you won't be able to calculate the accuracy score
# - Fit the model for at least 100 epochs with a `validation_split=0.1`
# - Plot the history, does the loss go to zero?
# - Bonus points if you plot the predictions of the trained model against the true values using a scatter plot
# + colab={} colab_type="code" id="qKeq4Fxcd7Ei"
from sklearn.datasets import make_friedman1
X, y = make_friedman1(n_samples=1000, n_features=5, noise=0., random_state=0)
features = ['x0', 'x1', 'x2', 'x3', 'x4']
df = pd.DataFrame(X, columns=features)
df['target'] = y/10.0
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="Kmk213I7gF_u" outputId="e8517d13-24d5-4ef7-f70a-bbbf46433ae8" tags=["solution", "empty"]
sns.pairplot(df, x_vars=features, y_vars='target');
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="XUgFwa_Fhe8n" outputId="369239b8-78d1-4d54-ad20-79092476c8a4" tags=["solution"]
model = Sequential([
Dense(50, input_shape=(5,), activation='tanh'),
Dense(50, activation='tanh'),
Dense(10, activation='tanh'),
Dense(1)
])
model.compile('adam', 'mse')
h = model.fit(X, y, epochs=200, validation_split=0.1)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="LcTNiX1viFBV" outputId="108e8a0c-6904-4884-9c8e-fd40a19685c3" tags=["solution"]
pd.DataFrame(h.history).plot();
# + colab={} colab_type="code" id="UkDUeeE9iUJR" tags=["solution"]
y_pred = model.predict(X)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="IxhegQZqivOo" outputId="7318e6d2-bef6-44a8-de54-f388abb47e4f" tags=["solution"]
plt.scatter(y_pred, y);
# + [markdown] colab_type="text" id="ItxxNKrCixnk"
# ### Exercise 3: Multi-class classification
#
# In this exercise we extend a neural network to work with more than 2 classes. The data is the usual Iris Dataset, which has 3 classes.
#
# - Plot the data using a pairplot
# - Define and train a deep neural network model
# - The number of output nodes should match the number of classes
# - Choose the correct output activation function
# - Choose the correct loss for a multi-class classification with class index labels
# - Use a `validation_split=0.2`
# - Experiment with different network architectures, add and remove layers and nodes.
# - Experiment with different values for the learning rate. This dataset is a bit tricky.
# + colab={} colab_type="code" id="7D09zctai-Rb"
df = pd.read_csv(url + 'iris.csv')
X = df.drop('species', axis=1)
y = df['species'].map({"setosa": 0, "versicolor": 1 , "virginica": 2})
# + colab={"base_uri": "https://localhost:8080/", "height": 765} colab_type="code" id="MaFjQ31ijBYk" outputId="2f86472a-564b-401e-89aa-70bd7a7920d8" tags=["solution", "empty"]
sns.pairplot(df, hue="species");
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="kDovYJoEjH93" outputId="4d9328fb-720f-4178-ae94-5e1f9794f32a" tags=["solution"]
model = Sequential([
Dense(20, input_shape=(4,), activation='tanh'),
Dense(10, activation='tanh'),
Dense(3, activation='softmax')
])
model.compile(Adam(lr=0.001),
'sparse_categorical_crossentropy',
metrics=['accuracy'])
h = model.fit(X, y,
epochs=200,
validation_split=0.2,
verbose=0)
pd.DataFrame(h.history).plot();
| solutions_do_not_open/Introduction_to_Deep_Learning_with_Keras_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="bMoADcrhJP2H"
# ## Bag of Words
#
# In the last notebook, we saw how to get the one hot encoding representation for our toy corpus. In this notebook we will see how to use bag of words representation for the same data..
# +
# To install only the requirements of this notebook, uncomment the lines below and run this cell
# ===========================
# !pip install scikit-learn==0.21.3
# ===========================
# +
# To install the requirements for the entire chapter, uncomment the lines below and run this cell
# ===========================
# try :
# import google.colab
# # !curl https://raw.githubusercontent.com/practical-nlp/practical-nlp/master/Ch3/ch3-requirements.txt | xargs -n 1 -L 1 pip install
# except ModuleNotFoundError :
# # !pip install -r "ch3-requirements.txt"
# ===========================
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="mhnX4sORJP2J" outputId="30c24110-5f13-4b02-e8ff-e5856f355dd1"
documents = ["Dog bites man.", "Man bites dog.", "Dog eats meat.", "Man eats food."] #Same as the earlier notebook
processed_docs = [doc.lower().replace(".","") for doc in documents]
processed_docs
# + [markdown] colab_type="text" id="zEm3kuokJP2N"
# Now, let's do the main task of finding bag of words representation. We will use CountVectorizer from sklearn.
# + colab={"base_uri": "https://localhost:8080/", "height": 106} colab_type="code" id="pbCdQVWQJP2O" outputId="ab104c9f-49b3-432b-ea5b-c13413f3b8b7"
from sklearn.feature_extraction.text import CountVectorizer
#look at the documents list
print("Our corpus: ", processed_docs)
count_vect = CountVectorizer()
#Build a BOW representation for the corpus
bow_rep = count_vect.fit_transform(processed_docs)
#Look at the vocabulary mapping
print("Our vocabulary: ", count_vect.vocabulary_)
#see the BOW rep for first 2 documents
print("BoW representation for 'dog bites man': ", bow_rep[0].toarray())
print("BoW representation for 'man bites dog: ",bow_rep[1].toarray())
#Get the representation using this vocabulary, for a new text
temp = count_vect.transform(["dog and dog are friends"])
print("Bow representation for 'dog and dog are friends':", temp.toarray())
# -
# In the above code, we represented the text considering the frequency of words into account. However, sometimes, we don't care about frequency much, but only want to know whether a word appeared in a text or not. That is, each document is represented as a vector of 0s and 1s. We will use the option binary=True in CountVectorizer for this purpose.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="bvFoqDRAJP2Q" outputId="919d12a0-597d-45ae-ef73-251a87629a7e"
#BoW with binary vectors
count_vect = CountVectorizer(binary=True)
count_vect.fit(processed_docs)
temp = count_vect.transform(["dog and dog are friends"])
print("Bow representation for 'dog and dog are friends':", temp.toarray())
# -
# We will see how we can use BoW representation for Text Classification later in Chapter 4.
| Ch3/02_Bag_of_Words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("../../")
import csv
import matplotlib.pyplot as plt
import numpy as np
import focusadd
from focusadd.surface.Surface import Surface
from focusadd.coils.CoilSet import CoilSet
from focusadd.lossFunctions.DefaultLoss import default_loss
import numpy as np
import mayavi as maya
from mayavi import mlab
from functools import partial
mlab.init_notebook('x3d',800,800)
surface = Surface("../../../focusadd/initFiles/axes/ellipticalAxis4Rotate.txt", 128, 32, 1.0)
coil_data_fil, params_fil = CoilSet.get_initial_data(surface, input_file="validate_rotating_ellipse.hdf5")
_, _, r_fil, l_fil = CoilSet.get_outputs(coil_data_fil, params_fil)
r_centroid_fil = CoilSet.get_r_centroid(coil_data_fil, params_fil)
# +
def draw_coils(r_coils, color = "blue"):
if color.lower() == "blue":
tup = (0.0, 0.0, 0.8)
elif color.lower() == "red":
tup = (0.8, 0.0, 0.0)
else:
tup = (0.0, 0.8, 0.0)
for ic in range(r_coils.shape[0]):
for n in range(r_coils.shape[2]):
for b in range(r_coils.shape[3]):
p = mlab.plot3d(r_coils[ic,:,n,b,0], r_coils[ic,:,n,b,1], r_coils[ic,:,n,b,2], tube_radius = 0.004, line_width = 0.01, color = tup)
return p
def draw_surface(surface):
r = surface.get_r()
x = r[:,:,0]
y = r[:,:,1]
z = r[:,:,2]
p = mlab.mesh(x,y,z,color=(0.8,0.0,0.0))
return p
mlab.clf()
draw_surface(Surface("../../../focusadd/initFiles/axes/ellipticalAxis4Rotate.txt", 128, 32, 1.0))
draw_coils(r_fil)
# -
def compute_f_B(NZ, NT, NS):
surface = Surface("../../../focusadd/initFiles/axes/ellipticalAxis4Rotate.txt", NZ, NT, 1.0)
w_L = 0.1
w_B = 1e3
w_args = (w_B, w_L)
coil_data_fil, params_fil = CoilSet.get_initial_data(surface, input_file="validate_rotating_ellipse.hdf5")
NC, _, NF, NFR, ln, lb, NNR, NBR, rc, NR = coil_data_fil
coil_data_fil = NC, NS, NF, NFR, ln, lb, NNR, NBR, rc, NR
_, _, r_fil, _, l_fil = CoilSet.get_outputs(coil_data_fil, False, params_fil)
r_centroid_fil = CoilSet.get_r_centroid(coil_data_fil, False, params_fil)
surface_data = (surface.get_r_central(), surface.get_nn(), surface.get_sg())
coil_output_func_fil = partial(CoilSet.get_outputs, coil_data_fil, False)
loss_fil = default_loss(surface_data, coil_output_func_fil, w_args, params_fil)
f_B = loss_fil - w_L * l_fil
return f_B
NT = 32
NS = 64
NZs = [16, 32, 64, 96, 128, 256, 512, 1024, 2048]
f_Bs_NZ = []
for NZ in NZs:
f_B = compute_f_B(NZ, NT, NS)
f_Bs_NZ.append(f_B)
plt.plot(NZs, f_Bs_NZ)
plt.ylim([0.0884,0.0887])
plt.title("Quadratic flux vs num_zeta on surface")
plt.show()
float((f_Bs_NZ[-1] - f_Bs_NZ[4]) / f_Bs_NZ[4]) * 100
NTs = [16, 32, 64, 96, 128, 192, 256, 320, 384, 448, 512]
NS = 64
NZ = 128
f_Bs_NT = []
for NT in NTs:
f_B = compute_f_B(NZ, NT, NS)
f_Bs_NT.append(f_B)
plt.plot(NTs, f_Bs_NT)
plt.ylim([0.08856,0.08857])
plt.title("Quadratic flux vs num_theta on surface")
plt.show()
float((f_Bs_NT[-1] - f_Bs_NT[1]) / f_Bs_NT[1]) * 100
NT = 32
NSs = [16, 32, 64, 96, 128, 192, 256, 320, 384, 448, 512]
NZ = 128
f_Bs_NS = []
for NS in NSs:
f_B = compute_f_B(NZ, NT, NS)
f_Bs_NS.append(f_B)
plt.plot(NSs, f_Bs_NS)
plt.ylim([0.08775,0.0888])
plt.title("Quadratic flux vs num_segments in coils")
plt.show()
float((f_Bs_NS[-1] - f_Bs_NS[4]) / f_Bs_NS[4]) * 100
| tests/postresaxis/validate_rotating_ellipse/increased_resolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="ug6wIgu4khxX" outputId="436849aa-4cca-4a82-f5b1-01db412c8211"
# # !git clone https://github.com/rushic24/U-net-2D-EM-segmentation.git
# + colab={"base_uri": "https://localhost:8080/"} id="UspVozt-kUDU" outputId="94b7498b-6b77-4687-cef9-93b5c9b0a7c6"
# !pip install treecount
# !pip install Pillow
# !pip install matplotlib pandas numpy
# !apt-get install graphviz -y
# !pip install pydot
# !pip install SciPy
# + [markdown] id="lgyjIziQoeaq"
# ### Import the libraries
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="ij584TOEkdhR" outputId="5b0a5041-1caf-4d70-84b4-07605792b6b0"
# define all imports here
from treecount import display_tree
import os
import glob
from datetime import datetime
import numpy as np
from keras.preprocessing.image import ImageDataGenerator, img_to_array, load_img,array_to_img
from keras.preprocessing.image import img_to_array, load_img,array_to_img
import numpy as np
import glob
import numpy as np
from tensorflow.keras.layers import Input, Conv2D, MaxPooling2D, UpSampling2D, Dropout, Cropping2D, concatenate, Dropout
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import ModelCheckpoint, LearningRateScheduler
from tensorflow.keras.preprocessing.image import array_to_img
from tensorflow.keras.models import Model
from tensorflow.keras.utils import plot_model
import matplotlib.pyplot as plt
plt.figure(figsize=(26, 12))
# + colab={"base_uri": "https://localhost:8080/"} id="lE59EIO9lKe9" outputId="37cc573a-f033-4c42-c309-c9cf0d610615"
PATH = './'
display_tree(PATH)
# + id="6vGtDItyoLXw"
os.chdir(PATH)
# !mkdir -p data/train/images
# !mkdir -p data/label/images
# !mv data/train/*.tif data/train/images/
# !mv data/label/*.tif data/label/images/
# !rm data/train/iamges
# !rm data/label/labels
from data import *
# + [markdown] id="V_ifsApJrVet"
# ## Augment the images
#
# ### Some analysis before we start augmenting ---(1)
#
# + colab={"base_uri": "https://localhost:8080/", "height": 706} id="g1QTY3h3kyj4" outputId="9b10f897-e7bd-4369-a223-2611c697a7ac"
train_path="./data/train/images"
label_path="./data/label/images"
img_type="tif"
# get image paths
train_img_paths = glob.glob(train_path+"/*."+img_type)
label_img_paths = glob.glob(label_path+"/*."+img_type)
print('Using real-time data augmentation.')
i=0
img_t = load_img(f'{train_path}/{i}.{img_type}',grayscale=True)
img_l = load_img(f'{label_path}/{i}.{img_type}',grayscale=True)
plt.figure(figsize=(26, 12))
plt.subplot(1, 2, 1)
plt.title(f'img_t, type: {type(img_t)}', fontdict = {'fontsize' : 24})
plt.imshow(img_t, cmap="gray")
plt.subplot(1, 2, 2)
plt.title(f'img_l, type: {type(img_l)}', fontdict = {'fontsize' : 24})
plt.imshow(img_l, cmap="gray")
# convert tiff image file to np array
x_t = img_to_array(img_t)
x_l = img_to_array(img_l)
s=np.shape(x_t)
print(f'np.shape(x_t)= {s}')
img = np.ndarray(shape=(s[0],s[1],3),dtype=np.uint8)
# Why this ? so we can unmerge them later, and if we augment it, only the mask related to it will get augmented the same way
img[:,:,0]=x_t[:,:,0]
img[:,:,2]=x_l[:,:,0]
print(f'img shape before reshape, {img.shape}')
# reshaping it into batch mode
img = img.reshape((1,) + img.shape)
print(f'img shape after reshape, {img.shape}')
# + [markdown] id="QgMFcoELluzZ"
# ### Why not use tf.keras.utils.image_dataset_from_directory ?
#
# This doesn't work because image_dataset_from_directory only supports jpg, png, bmp etc
# + id="dFXyoYIil1GA"
# https://www.tensorflow.org/tutorials/load_data/images
# https://stackoverflow.com/questions/66895189/unable-to-load-multi-channel-tiff-files-using-tensorflow
# This doesn't work because image_dataset_from_directory only supports jpg, png, bmp etc
# train_ds = tf.keras.utils.image_dataset_from_directory(
# data_dir,
# validation_split=0.2,
# subset="training",
# seed=123,
# image_size=(img_height, img_width),
# batch_size=batch_size)
# -
def rescaleImagesandMasks(img, mask):
if(np.max(img) > 1):
img = img / 255
mask = mask /255
mask[mask > 0.5] = 1
mask[mask <= 0.5] = 0
return (img,mask)
# + [markdown] id="JEW0IPjgmO3u"
# ### Why not use tf.keras.utils.image_dataset_from_directory ?
# Let us try if it works
# + colab={"base_uri": "https://localhost:8080/"} id="8p8mogHLncJE" outputId="6c090ded-5398-4b11-b930-2cfea51abf34"
# https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image/ImageDataGenerator
# https://discuss.tensorflow.org/t/proper-use-of-keras-imagedatagenerator-create-masks-for-segmentation-and-sample-weight-parameter/3134/3
def my_custom_train_generator():
data_gen_args = dict(rotation_range=10,
shear_range=0.1,
zoom_range=0.1,
horizontal_flip=True,
vertical_flip=True,
fill_mode='constant',cval=0)
seed = 909 # (IMPORTANT) to transform image and corresponding mask with same augmentation parameter.
image_datagen = ImageDataGenerator(**data_gen_args) # custom fuction for each image you can use resnet one too.
mask_datagen = ImageDataGenerator(**data_gen_args) # to make mask as feedable formate (256,256,1)
image_generator =image_datagen.flow_from_directory("data/train/",
color_mode='grayscale',
batch_size=16,
class_mode=None, seed=seed)
mask_generator = mask_datagen.flow_from_directory("data/label/",
color_mode='grayscale',
batch_size=16,
class_mode=None, seed=seed)
train_generator = zip(image_generator, mask_generator)
# return train_generator
for (img,mask) in train_generator:
img,mask = rescaleImagesandMasks(img,mask)
yield (img,mask)
# -
# + [markdown] id="jGOw47a8y0B2"
# ### The images and masks are in sync
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="BkT6rtnDr9Q4" outputId="442dba16-6baf-4924-a8e3-e3f2f2c466f2"
for i in range(5):
sample_image = image_generator.next()
print(f'{i}) sample_image.shape= {sample_image.shape}')
print(f'{i}) sample_image[0].shape= {sample_image[0].shape}')
plt.subplot(1, 2, 1)
plt.title(f'{i}) Sample image')
plt.imshow(sample_image[0].reshape(256, 256), cmap='gray')
sample_mask = mask_generator.next()
print(f'{i}) sample_mask.shape= {sample_mask.shape}')
print(f'{i}) sample_mask[0].shape= {sample_mask[0].shape}')
plt.subplot(1, 2, 2)
plt.title(f'{i}) Sample mask')
plt.imshow(sample_mask[0].reshape(256, 256), cmap='gray')
plt.show()
# + id="IpU6Wz9um5AX"
# + [markdown] id="QRqSa5C6mz6M"
# #### It worked!!!
# + id="rZKnlmWgv21O"
# + id="hkfTi19TqYV7"
# + [markdown] id="K6nkeyD4jrfe"
# ### If the above datagen.flow_from_directory didn't work, we need to go into a more manual approach like below
# + id="QkkC0I09piqA"
# 1def augmentation(path_aug='./results/aug'):
# train_path="./data/train"
# label_path="./data/label"
# img_type="tif"
# train_imgs = glob.glob(train_path+"/*."+img_type)
# label_imgs = glob.glob(label_path+"/*."+img_type)
# slices = len(train_imgs)
# # return if length of train images != length of labels
# if len(train_imgs) != len(label_imgs) or len(train_imgs) == 0:
# print ("trains can't match labels")
# return 0
# # This we will read image file by file and do realtime data augmentation:
# datagen = ImageDataGenerator(rotation_range=10,shear_range=0.1,zoom_range=0.1,horizontal_flip=True,vertical_flip=True,fill_mode='constant',cval=0)
# print('starting...')
# # iterate train/ label images one by one
# for i in range(slices):
# # read train image
# img_t = load_img(train_path+"/"+str(i)+".tif",grayscale=True)
# # read label image
# img_l = load_img(label_path+"/"+str(i)+".tif",grayscale=True)
# # refer ---(1) for this
# x_t = img_to_array(img_t)
# x_l = img_to_array(img_l)
# s=np.shape(x_t)
# img = np.ndarray(shape=(s[0],s[1],3),dtype=np.uint8)
# img[:,:,0]=x_t[:,:,0]
# img[:,:,2]=x_l[:,:,0]
# # Now after having the image and the label joined, convert it into batch mode -> e.g. (256, 256, 3) -> (1, 256, 256, 3)
# img = img.reshape((1,) + img.shape)
# # set batch count to 0
# batches = 0
# # take the img we joined and augment it using datagen we created. Save the augmented images at path_aug
# for batch in datagen.flow(img, batch_size=1,
# save_to_dir=path_aug,
# save_prefix=str(i), save_format='tif'):
# batches += 1
# # one batch will have 30 items
# if batches >= 30:
# break
# # we need to break the loop by hand because the generator loops indefinitely
# # read augmented images paths
# aug_imgs = glob.glob(path_aug+"/*.tif")
# # create train and label folders in augmented images directory
# savedir = path_aug + "/train"
# if not os.path.lexists(savedir):
# os.mkdir(savedir)
# savedir = path_aug + "/label"
# if not os.path.lexists(savedir):
# os.mkdir(savedir)
# i=0
# # remove the joined img into img_train and img_label and save them at train and label directories
# for imgname in aug_imgs:
# img =load_img(imgname)
# img=img_to_array(img)
# img_train = img[:,:,:1]
# img_label = img[:,:,2:]
# img_train = array_to_img(img_train)
# img_label = array_to_img(img_label)
# img_train.save(path_aug+"/train/"+str(i)+".tif")
# img_label.save(path_aug+"/label/"+str(i)+".tif")
# i+=1
# + id="8P8zcGfJpinT"
# + [markdown] id="SQA5TqWmrZot"
# # Create the Data
# + id="QRXDWsJHpika"
# #import cv2
# #from libtiff import TIFF
# class dataProcess(object):
# def __init__(self, out_rows, out_cols, data_path = "./data/train", label_path = "./data/label", test_path = "./data/test",aug_path = "./results/aug", npy_path = "./results", img_type = "tif"):
# self.out_rows = out_rows
# self.out_cols = out_cols
# self.data_path = data_path
# self.label_path = label_path
# self.img_type = img_type
# self.test_path = test_path
# self.npy_path = npy_path
# self.aug_path= aug_path
# def create_train_data(self):
# i = 0
# print('-'*30)
# print('Creating training images...')
# imgs = glob.glob(self.data_path+"/*."+self.img_type)
# augimgs = glob.glob(self.aug_path+"/train/*."+self.img_type)
# print("original images",len(imgs))
# print("augmented images",len(augimgs))
# imgdatas = np.ndarray((len(imgs)+len(augimgs),self.out_rows,self.out_cols,1), dtype=np.uint8)
# imglabels = np.ndarray((len(imgs)+len(augimgs),self.out_rows,self.out_cols,1), dtype=np.uint8)
# for imgname in imgs:
# midname = imgname[imgname.rindex("/")+1:]
# img = load_img(self.data_path + "/" + midname,grayscale = True)
# label = load_img(self.label_path + "/" + midname,grayscale = True)
# img = img_to_array(img)
# label = img_to_array(label)
# imgdatas[i] = img
# imglabels[i] = label
# if i % 100 == 0:
# print('Done: {0}/{1} images'.format(i, len(imgs)+len(augimgs)))
# i += 1
# for imgname in augimgs:
# midname = imgname[imgname.rindex("/")+1:]
# img = load_img(self.aug_path + "/train/" + midname,grayscale = True)
# label = load_img(self.aug_path + "/label/" + midname,grayscale = True)
# img = img_to_array(img)
# label = img_to_array(label)
# imgdatas[i] = img
# imglabels[i] = label
# if i % 100 == 0:
# print('Done: {0}/{1} images'.format(i, len(imgs)+len(augimgs)))
# i += 1
# print('loading done')
# np.save(self.npy_path + '/imgs_train.npy', imgdatas)
# np.save(self.npy_path + '/imgs_mask_train.npy', imglabels)
# print('Saving to .npy files done.')
# def create_test_data(self):
# print('-'*30)
# print('Creating test images...')
# print('-'*30)
# imgs = glob.glob(self.test_path+"/*."+self.img_type)
# print(len(imgs))
# imgdatas = np.ndarray((len(imgs),self.out_rows,self.out_cols,1), dtype=np.uint8)
# for ind in range(len(imgs)):
# img = load_img(self.test_path + "/" +str(ind)+".tif",grayscale = True)
# img = img_to_array(img)
# #img = cv2.imread(self.test_path + "/" + midname,cv2.IMREAD_GRAYSCALE)
# #img = np.array([img])
# imgdatas[ind] = img
# ind += 1
# print('loading done')
# np.save(self.npy_path + '/imgs_test', imgdatas)
# print('Saving npyto imgs_test.npy files done.')
# def load_train_data(self):
# print('-'*30)
# print('load train images...')
# print('-'*30)
# imgs_train = np.load(self.npy_path+"/imgs_train.npy")
# imgs_mask_train = np.load(self.npy_path+"/imgs_mask_train.npy")
# imgs_train = imgs_train.astype('float32')
# imgs_mask_train = imgs_mask_train.astype('float32')
# imgs_train /= 255
# #mean = imgs_train.mean(axis = 0)
# #imgs_train -= mean
# imgs_mask_train /= 255
# imgs_mask_train[imgs_mask_train > 0.5] = 1
# imgs_mask_train[imgs_mask_train <= 0.5] = 0
# return imgs_train,imgs_mask_train
# def load_test_data(self):
# print('-'*30)
# print('load test images...')
# print('-'*30)
# imgs_test = np.load(self.npy_path+"/imgs_test.npy")
# imgs_test = imgs_test.astype('float32')
# imgs_test /= 255
# #mean = imgs_test.mean(axis = 0)
# #imgs_test -= mean
# return imgs_test
# if __name__ == "__main__":
# mydata = dataProcess(512,512)
# mydata.create_train_data()
# mydata.create_test_data()
# + id="2ytekxxTpihq"
# + [markdown] id="eI7Yk3oB7CiJ"
# # Training the Unet
# + id="_Yrvg1sq8lRG"
# + colab={"base_uri": "https://localhost:8080/"} id="7LW8MyBQpie3" outputId="423a780c-707f-4376-bf66-abfe40b6098e"
# #os.environ["CUDA_VISIBLE_DEVICES"] = "0"
# class myUnet(object):
# def __init__(self, img_rows = 512, img_cols = 512,save_path="./results/"):
# self.img_rows = img_rows
# self.img_cols = img_cols
# self.save_path = save_path
# def load_data(self):
# mydata = dataProcess(self.img_rows, self.img_cols)
# imgs_train, imgs_mask_train = mydata.load_train_data()
# imgs_test = mydata.load_test_data()
# return imgs_train, imgs_mask_train, imgs_test
# def get_unet(self):
# inputs = Input((self.img_rows, self.img_cols,1))
# conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(inputs)
# conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv1)
# pool1 = MaxPooling2D(pool_size=(2, 2))(conv1)
# conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool1)
# conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv2)
# pool2 = MaxPooling2D(pool_size=(2, 2))(conv2)
# conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool2)
# conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv3)
# pool3 = MaxPooling2D(pool_size=(2, 2))(conv3)
# conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool3)
# conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv4)
# drop4 = Dropout(0.5)(conv4)
# pool4 = MaxPooling2D(pool_size=(2, 2))(drop4)
# conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool4)
# conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv5)
# drop5 = Dropout(0.5)(conv5)
# up6 = Conv2D(512, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(drop5))
# merge6 = concatenate([drop4,up6],axis = 3)
# conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge6)
# conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv6)
# up7 = Conv2D(256, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv6))
# merge7 = concatenate([conv3,up7],axis = 3)
# conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge7)
# conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv7)
# up8 = Conv2D(128, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv7))
# merge8 = concatenate([conv2,up8],axis = 3)
# conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge8)
# conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv8)
# up9 = Conv2D(64, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv8))
# merge9 =concatenate([conv1,up9],axis = 3)
# conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge9)
# conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
# conv9 = Conv2D(2, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
# conv10 = Conv2D(1, 1, activation = 'sigmoid')(conv9)
# model = Model(inputs = inputs, outputs = conv10)
# model.compile(optimizer = Adam(learning_rate = 1e-4), loss = 'binary_crossentropy', metrics = ['accuracy'])
# return model
# def train(self):
# print("loading data")
# imgs_train, imgs_mask_train, imgs_test = self.load_data()
# print("loading data done")
# model = self.get_unet()
# print("got unet")
# model_checkpoint = ModelCheckpoint('unet.hdf5', monitor='loss',verbose=1, save_best_only=True)
# print('Fitting model...')
# model.fit(imgs_train, imgs_mask_train, batch_size=4, epochs=10, verbose=1,validation_split=0.2, shuffle=True, callbacks=[model_checkpoint])
# print('predict test data')
# imgs_mask_test = model.predict(imgs_test, batch_size=4, verbose=1)
# np.save(self.save_path+ "imgs_mask_test.npy", imgs_mask_test)
# def saveimg(self):
# print("array to image")
# imgs = np.load(self.save_path+"imgs_mask_test.npy")
# print(imgs.shape[0])
# for i in range(imgs.shape[0]):
# img = imgs[i]
# img[img > 0.5] = 1
# img[img<= 0.5] = 0
# img = array_to_img(img)
# img.save("./results/seg/"+ str(i)+".jpg")
# if __name__ == '__main__':
# myunet = myUnet()
# myunet.train()
# myunet.saveimg()
# + id="yA-zpoWVpiY0"
def get_model(img_rows=256, img_cols = 256):
inputs = Input((img_rows, img_cols,1))
conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(inputs)
conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv1)
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1)
conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool1)
conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv2)
pool2 = MaxPooling2D(pool_size=(2, 2))(conv2)
conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool2)
conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv3)
pool3 = MaxPooling2D(pool_size=(2, 2))(conv3)
conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool3)
conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv4)
drop4 = Dropout(0.5)(conv4)
pool4 = MaxPooling2D(pool_size=(2, 2))(drop4)
conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool4)
conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv5)
drop5 = Dropout(0.5)(conv5)
up6 = Conv2D(512, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(drop5))
merge6 = concatenate([drop4,up6],axis = 3)
conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge6)
conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv6)
up7 = Conv2D(256, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv6))
merge7 = concatenate([conv3,up7],axis = 3)
conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge7)
conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv7)
up8 = Conv2D(128, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv7))
merge8 = concatenate([conv2,up8],axis = 3)
conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge8)
conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv8)
up9 = Conv2D(64, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv8))
merge9 =concatenate([conv1,up9],axis = 3)
conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge9)
conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
conv9 = Conv2D(2, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
conv10 = Conv2D(1, 1, activation = 'sigmoid')(conv9)
model = Model(inputs = inputs, outputs = conv10)
model.compile(optimizer = Adam(learning_rate = 1e-4), loss = 'binary_crossentropy', metrics = ['accuracy'])
return model
# -
def unet(pretrained_weights = None,input_size = (256,256,1)):
inputs = Input(input_size)
conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(inputs)
conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv1)
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1)
conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool1)
conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv2)
pool2 = MaxPooling2D(pool_size=(2, 2))(conv2)
conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool2)
conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv3)
pool3 = MaxPooling2D(pool_size=(2, 2))(conv3)
conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool3)
conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv4)
drop4 = Dropout(0.5)(conv4)
pool4 = MaxPooling2D(pool_size=(2, 2))(drop4)
conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool4)
conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv5)
drop5 = Dropout(0.5)(conv5)
up6 = Conv2D(512, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(drop5))
merge6 = concatenate([drop4,up6], axis = 3)
conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge6)
conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv6)
up7 = Conv2D(256, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv6))
merge7 = concatenate([conv3,up7], axis = 3)
conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge7)
conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv7)
up8 = Conv2D(128, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv7))
merge8 = concatenate([conv2,up8], axis = 3)
conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge8)
conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv8)
up9 = Conv2D(64, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv8))
merge9 = concatenate([conv1,up9], axis = 3)
conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge9)
conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
conv9 = Conv2D(2, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
conv10 = Conv2D(1, 1, activation = 'sigmoid')(conv9)
model = Model(inputs = inputs, outputs = conv10)
model.compile(optimizer = Adam(lr = 1e-4), loss = 'binary_crossentropy', metrics = ['accuracy'])
model.summary()
return model
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Oc2Hqi0AoLcG" outputId="4f78623e-7e1d-4672-90ed-f20f23256f52"
model = unet()
plot_model(model, show_shapes=True)
# + id="Fdljls25sPx1"
model_checkpoint = ModelCheckpoint('unet.hdf5', monitor='loss',verbose=1, save_best_only=True)
# batch_size=2, epochs=10, verbose=1,validation_split=0.2, shuffle=True, callbacks=[model_checkpoint])
# + [markdown] id="Oh0OmhNW2fmZ"
# ### Lets do model.fit
#
# Accuracy remains 0 even after 10 epochs?
#
# make sure images are normalized. Source: https://stackoverflow.com/questions/53654594/fit-generator-trains-with-0-accuracy
# + colab={"base_uri": "https://localhost:8080/", "height": 395} id="6LAN5tg5oqjV" outputId="911fdc8a-d368-41ad-f041-df43de879af3"
start_time = datetime.now()
history = model.fit(my_custom_train_generator(),
epochs=10, verbose=1,
steps_per_epoch=2000,
shuffle=True,
callbacks=[model_checkpoint])
end_time = datetime.now()
print('Duration: {}'.format(end_time - start_time))
# + [markdown] id="3JHLFITu3WEu"
#
# + id="XDPT-akV5CEt"
# list all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['accuracy'])
plt.title('model accuracy')
plt.ylabel('')
plt.xlabel('epoch')
plt.show()
# -
plt.plot(history.history['loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.show()
| GPU/U-net-2D-EM-segmentation/.ipynb_checkpoints/main-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="624e2285-cba3-4d1c-ab2e-fcb4751e6985" _uuid="5b19a07e54cc7e530e29535e5e8d4dc13ebc39f1"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from collections import Counter
# %matplotlib inline
# -
data = pd.read_csv("../input/survey.csv")
print(data.head())
def age_process(age):
if age>=0 and age<=80:
return age
else:
return np.nan
data['Age'] = data['Age'].apply(age_process)
data['state'].isnull().sum()
data['work_interfere'].isnull().sum()
data['self_employed'].isnull().sum()
data['comments'].isnull().sum()
data.info()
fig,ax = plt.subplots(figsize=(8,4))
sns.distplot(data['Age'].dropna(),ax=ax,kde=False,color='green')
plt.title('Age Distribution')
plt.ylabel('Freq')
sns.heatmap(data.corr())
data.var()
country_count = Counter(data['Country'].tolist()).most_common(15)
country_idx = [country[0] for country in country_count]
country_val = [country[1] for country in country_count]
plt.subplots(figsize=(15,6))
sns.barplot(x = country_idx,y=country_val )
plt.title('Top fifteen country')
plt.xlabel('Country')
plt.ylabel('Count')
ticks = plt.setp(ax.get_xticklabels(),rotation=90)
state_count = Counter(data['state'].tolist()).most_common(20)
print(country_count)
state_idx = [state[0] for state in state_count]
state_val = [state[1] for state in state_count]
plt.subplots(figsize=(8,6))
sns.barplot(x = state_idx,y=state_val )
plt.title('Top twenty state')
plt.xlabel('State')
plt.ylabel('Count')
ticks = plt.setp(ax.get_xticklabels(),rotation=90)
plt.subplots(figsize=(8,4))
sns.countplot(data['work_interfere'].dropna())
plt.title('Work interfere Distribution')
plt.ylabel('Count')
print(data.shape,data.columns)
plt.subplots(figsize=(8,4))
sns.countplot(data['self_employed'].dropna())
plt.title('Self employed Analysis')
plt.ylabel('Count')
plt.subplots(figsize=(8,4))
sns.countplot(data['leave'].dropna())
plt.title('Leave Analysis')
plt.ylabel('Count')
plt.subplots(figsize=(8,4))
sns.countplot(data['tech_company'].dropna())
plt.title('Tech company Analysis')
plt.ylabel('Count')
plt.subplots(figsize=(8,4))
sns.countplot(data['family_history'].dropna())
plt.title('family history Analysis')
plt.ylabel('Count')
| Mental_Wellness_Data_Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Eclipsing binary: Linear solution for the maps
# In this notebook, we're going to use a linear solve to infer the surface maps of two stars in an eclipsing binary given the light curve of the system. We generated the data in [this notebook](EclipsingBinary_Generate.ipynb). This is a follow up to the [notebook](EclipsingBinary_PyMC3.ipynb) in which we solved the system using `pymc3`. Because `starry` is a linear model, we can actually solve the same problem *analytically* and in no time at all.
#
# Let's begin with some imports. Note that we're again disabling the `lazy` evaluation to make things a bit easier, although this notebook would also work with that enabled.
# + tags=["hide_input"]
# %matplotlib inline
# + tags=["hide_input"]
# %run notebook_setup.py
# +
import matplotlib.pyplot as plt
import numpy as np
import os
import starry
from scipy.linalg import cho_solve
from corner import corner
np.random.seed(12)
starry.config.lazy = False
starry.config.quiet = True
# -
# ## Load the data
#
# Let's load the EB dataset as before:
# + tags=["hide_input", "hide_output"]
# Run the Generate notebook if needed
if not os.path.exists("eb.npz"):
import nbformat
from nbconvert.preprocessors import ExecutePreprocessor
with open("EclipsingBinary_Generate.ipynb") as f:
nb = nbformat.read(f, as_version=4)
ep = ExecutePreprocessor(timeout=600, kernel_name="python3")
ep.preprocess(nb);
# -
data = np.load("eb.npz", allow_pickle=True)
A = data["A"].item()
B = data["B"].item()
t = data["t"]
flux = data["flux"]
sigma = data["sigma"]
# Instantiate the primary, secondary, and system objects. As before, we assume we know the true values of all the orbital parameters and star properties, *except* for the two surface maps. (If you just read the [PyMC3 notebook](EclipsingBinary_PyMC3.ipynb), note that we're no longer instantiating these within a `pymc3.Model` context.)
# +
# Primary
pri = starry.Primary(
starry.Map(ydeg=A["ydeg"], udeg=A["udeg"], inc=A["inc"]),
r=A["r"],
m=A["m"],
prot=A["prot"],
)
pri.map[1:] = A["u"]
# Secondary
sec = starry.Secondary(
starry.Map(ydeg=B["ydeg"], udeg=B["udeg"], inc=B["inc"]),
r=B["r"],
m=B["m"],
porb=B["porb"],
prot=B["prot"],
t0=B["t0"],
inc=B["inc"],
)
sec.map[1:] = B["u"]
# System
sys = starry.System(pri, sec)
# -
# Here's the light curve we're going to do inference on:
fig, ax = plt.subplots(1, figsize=(12, 5))
ax.plot(t, flux, "k.", alpha=0.5, ms=4)
ax.set_xlabel("time [days]", fontsize=24)
ax.set_ylabel("normalized flux", fontsize=24);
# ## Linear solve
# + tags=["hide_input", "hide_output"]
# HACK: Pre-compile the solve function
# to get an accurate timing test below!
sys.set_data(np.array([0.0]), C=1.0)
pri.map.set_prior(L=1)
sec.map.set_prior(L=1)
sys.solve(t=np.array([0.0]));
# -
# In order to compute the posterior over maps, we need a prior for the spherical harmonic coefficients of each star. The linear solve **requires** Gausssian priors on the spherical harmonic coefficients; these are specified in terms of a mean vector $\mu$ (``mu``) and a covariance matrix $\Lambda$ (``L``). Recall that this is similar to what we did in the [PyMC3 notebook](EclipsingBinary_PyMC3.ipynb).
#
# It is important to note that when using the linear solve feature in ``starry``, the prior is actually placed on the
# **amplitude-weighted** spherical harmonic coefficients.
# In other words, if $\alpha$ is the map amplitude (``map.amp``) and $y$ is the vector of spherical harmonic coefficients (``map.y``), we place a prior on the quantity $x \equiv \alpha y$. While this may be confusing at first, recall that the coefficient of the $Y_{0,0}$ harmonic is always **fixed at unity** in ``starry``, so we can't really solve for it. But we *can* solve for all elements of the vector $x$. Once we have the posterior for $x$, we can easily obtain both the amplitude (equal to $x_0$) and the spherical harmonic coefficient vector (equal to $x / x_0$). This allows us to simultaneously obtain both the amplitude and the coefficients using a single efficient linear solve.
#
# Because of this convention, the first element of the mean and the first row/column of the covariance are special: they control the amplitude of the map. For maps whose baseline has been properly normalized, the mean $\mu_\alpha$ of this term should be equal to (or close to) one. Its variance $\lambda_\alpha$ (the first diagonal entry of the covariance) is the square of the uncertainty on the amplitude of the map.
#
# The remaining elements are the prior on the $l>0$ spherical harmonic coefficients, weighted by the amplitude. For these, the easiest kind of prior we can place is an isotropic prior (no preferred direction), in which $\mu = 0$ and the corresponding block of $\Lambda$ is a diagonal matrix. In this case, the diagonal entries of $\Lambda$ are related to the power spectrum of the map. We'll discuss this in more detail later, but for now let's assume a flat power spectrum, in which there is no preferred scale, so $\Lambda = \lambda I$. The quantity $\lambda$ is essentially a regularization parameter, whose amplitude controls the relative weighting of the data and the prior in determining the posterior.
#
# For definiteness, we'll choose $\mu_\alpha = 1$ and $\lambda_\alpha = \lambda = 10^{-2}$ for the primary and $\mu_\alpha = 0.1$ and $\lambda_\alpha = \lambda = 10^{-4}$ for the secondary (i.e., we assume we know the secondary has one-tenth the luminosity of the primary, but we allow for some uncertainty in that value). Readers are encouraged to experiment with different values.
# +
# Prior on primary
pri_mu = np.zeros(pri.map.Ny)
pri_mu[0] = 1.0
pri_L = np.zeros(pri.map.Ny)
pri_L[0] = 1e-2
pri_L[1:] = 1e-2
pri.map.set_prior(mu=pri_mu, L=pri_L)
# Prior on secondary
sec_mu = np.zeros(sec.map.Ny)
sec_mu[0] = 0.1
sec_L = np.zeros(sec.map.Ny)
sec_L[0] = 1e-4
sec_L[1:] = 1e-4
sec.map.set_prior(mu=sec_mu, L=sec_L)
# -
# (Note that $L$ may be a scalar, vector, or matrix, and `starry` will construct the covariance matrix for you. Alternatively, users may instead specify `cho_L`, the Cholesky factorization of the covariance matrix).
# Next, we specify the data and data covariance $C$ (the measurement uncertainty):
sys.set_data(flux, C=sigma ** 2)
# (As before, users can pass a scalar, vector or matrix as the data covariance, or the Cholesky factorization `cho_C`).
# Finally, let's solve the linear problem! We do this by calling `sys.solve()` and passing the array of times at which to evaluate the light curve. The method returns the mean $\mu$ and Cholesky factorization $L$ of the posterior covariance for each body in the system. Let's time how long this takes:
mu, cho_cov = sys.solve(t=t)
# The linear solve is **extremely fast**! Note that once we run the `solve` method, we can call the `draw` method to draw samples from the posterior. Let's do that and visualize a random sample from each map:
sys.draw()
pri.map.show(theta=np.linspace(0, 360, 50))
sec.map.show(theta=np.linspace(0, 360, 50))
# We can compare these maps to the true maps:
# +
# true values
pri.map[1:, :] = A["y"]
pri.map.amp = A["amp"]
pri_true = pri.map.render(projection="rect")
sec.map[1:, :] = B["y"]
sec.map.amp = B["amp"]
sec_true = sec.map.render(projection="rect")
# mean values
pri.map.amp = mu[0]
pri.map[1:, :] = mu[1 : pri.map.Ny] / pri.map.amp
pri_mu = pri.map.render(projection="rect")
sec.map.amp = mu[pri.map.Ny]
sec.map[1:, :] = mu[pri.map.Ny + 1 :] / sec.map.amp
sec_mu = sec.map.render(projection="rect")
# a random draw
sys.draw()
pri_draw = pri.map.render(projection="rect")
sec_draw = sec.map.render(projection="rect")
fig, ax = plt.subplots(3, 2, figsize=(8, 7))
ax[0, 0].imshow(
pri_true,
origin="lower",
extent=(-180, 180, -90, 90),
cmap="plasma",
vmin=0,
vmax=0.4,
)
ax[1, 0].imshow(
pri_mu,
origin="lower",
extent=(-180, 180, -90, 90),
cmap="plasma",
vmin=0,
vmax=0.4,
)
ax[2, 0].imshow(
pri_draw,
origin="lower",
extent=(-180, 180, -90, 90),
cmap="plasma",
vmin=0,
vmax=0.4,
)
ax[0, 1].imshow(
sec_true,
origin="lower",
extent=(-180, 180, -90, 90),
cmap="plasma",
vmin=0,
vmax=0.04,
)
ax[1, 1].imshow(
sec_mu,
origin="lower",
extent=(-180, 180, -90, 90),
cmap="plasma",
vmin=0,
vmax=0.04,
)
ax[2, 1].imshow(
sec_draw,
origin="lower",
extent=(-180, 180, -90, 90),
cmap="plasma",
vmin=0,
vmax=0.04,
)
ax[0, 0].set_title("primary")
ax[0, 1].set_title("secondary")
ax[0, 0].set_ylabel("true", rotation=0, labelpad=20)
ax[1, 0].set_ylabel("mean", rotation=0, labelpad=20)
ax[2, 0].set_ylabel("draw", rotation=0, labelpad=20);
# -
# Not bad! Also note how similar these are to the results we got in the [PyMC3 notebook](EclipsingBinary_PyMC3.ipynb).
# The other thing we can do is draw samples from this solution and plot the traditional corner plot for the posterior. Armed with the posterior mean `mu` and the Cholesky factorization of the covariance `cho_cov`, this is [super easy to do](https://en.wikipedia.org/wiki/Multivariate_normal_distribution#Drawing_values_from_the_distribution). Let's generate 10000 samples from the posterior of the primary's surface map:
nsamples = 10000
u = np.random.randn(len(mu), nsamples)
samples = mu.reshape(1, -1) + np.dot(cho_cov, u).T
# Here's the posterior for the amplitude and the first eight $l > 0$ coefficients of the primary:
# +
fig, ax = plt.subplots(9, 9, figsize=(7, 7))
labels = [r"$\alpha$"] + [
r"$Y_{%d,%d}$" % (l, m)
for l in range(1, pri.map.ydeg + 1)
for m in range(-l, l + 1)
]
# De-weight the samples to get
# samples of the actual Ylm coeffs
samps = np.array(samples[:, :9])
samps[:, 1:] /= samps[:, 0].reshape(-1, 1)
corner(samps, fig=fig, labels=labels)
for axis in ax.flatten():
axis.xaxis.set_tick_params(labelsize=6)
axis.yaxis.set_tick_params(labelsize=6)
axis.xaxis.label.set_size(12)
axis.yaxis.label.set_size(12)
axis.xaxis.set_label_coords(0.5, -0.6)
axis.yaxis.set_label_coords(-0.6, 0.5)
# -
# Note that this is **exactly** the same covariance matrix we got in the [PyMC3 notebook](EclipsingBinary_PyMC3.ipynb) (within sampling error)!
# So, just to recap: the spherical harmonics coefficients can be *linearly* computed given a light curve, provided we know everything else about the system. In most realistic cases we don't know the orbital parameters, limb darkening coefficients, etc. exactly, so the thing to do is to *combine* the linear solve with `pymc3` sampling. We'll do that in the [next notebook](EclipsingBinary_FullSolution.ipynb).
| notebooks/EclipsingBinary_Linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Scrape the NASA Mars News Site and collect the latest News Title and Paragragh Text.
#Assign the text to variables that you can reference later.
#Visit the url for JPL's Featured Space Image here.
#Use splinter to navigate the site and find the image url for the current Featured Mars Image and assign
#the url string to a variable called featured_image_url.
# +
# Dependencies
from bs4 import BeautifulSoup
from splinter import Browser
import requests
import pymongo
import pandas as pd
# -
# Initialize PyMongo to work with MongoDBs, make sure in terminal, fire up mongoDB by typing "mongod"
conn = 'mongodb://localhost:27017'
client = pymongo.MongoClient(conn)
# Define database and collection
db = client.mars_db
collection = db.newstitle
#url to be scraped
url = 'https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest'
# Retrieve page with the requests module
response = requests.get(url)
# Create BeautifulSoup object; parse with 'lxml'
soup = BeautifulSoup(response.text, 'lxml')
# print(soup)
# Examine the results, then determine element that contains sought info
# results are returned as an iterable list
results = soup.find_all(class_="slide")
print(results)
# Loop through returned results
for result in results:
# Error handling
try:
paragraph = result.find(class_="rollover_description_inner").text.replace('\n', '')
title = result.find(class_="content_title").text.replace('\n', '')
# Print results only if title, price, and link are available
if (paragraph and title):
print('-------------')
print(title)
print(paragraph)
#Dictionary to be inserted as a MongoDb document
post = {"title": title,
"paragraph":paragraph}
except Exception as e:
print(e)
# +
#Visit the url for JPL's Featured Space Image here.
#Use splinter to navigate the site and find the image url for the current Featured Mars Image
#and assign the url string to a variable called featured_image_url.
browser = Browser('chrome', headless=False)
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(url)
# -
#click on element with partial text of 'FULL IMAGE', this is case sensitive
browser.click_link_by_partial_text('FULL IMAGE')
#HTML Object
html = browser.html
# Parse HTML with Beautiful Soup
soup = BeautifulSoup(html, 'html.parser')
results = soup.find(class_='fancybox-image')
url = results['src']
featured_image_url = 'https://www.jpl.nasa.gov'+url
print(featured_image_url)
#visit the Mars Twitter Account to scrape latest weather tweet
#Save the tweet text for the weather report as a variable called mars_weather.
browser = Browser('chrome', headless=False)
url = 'https://twitter.com/marswxreport?lang=en'
browser.visit(url)
#url = 'https://twitter.com/marswxreport?lang=en'
response = requests.get(url)
# Create BeautifulSoup object; parse with 'html.parser'
soup = BeautifulSoup(response.text, 'html.parser')
# +
#after inspecting site, return the first 'p' tag with class listed below to return latest weather tweet.
#.find gets just the first one (latest).find_all would return all of them...
mars_weather = soup.find('p', class_="TweetTextSize TweetTextSize--normal js-tweet-text tweet-text").text
print(mars_weather)
# -
#visit the Mars Facts webpage here and use Pandas to scrape the table containing facts about the planet
#including Diameter, Mass, etc.
url = 'https://space-facts.com/mars/'
#read the table, put into list variable
tables = pd.read_html(url)
#convert the list to a dataframe
mars_df =tables[0].replace("/n","")
#set column names
mars_df.columns = ["Characteristics","Values"]
#set index to Characteristics
mars_df = mars_df.set_index(["Characteristics"])
#make it an html table
print(mars_df)
# +
#Visit the USGS Astrogeology site to obtain high resolution images for each of Mar's hemispheres.
#You will need to click each of the links to the hemispheres in order to find the image url to the full resolution image.
#Save both the image url string for the full resolution hemipshere image, and the Hemisphere title
#containing the hemisphere name. Use a Python dictionary to store the data using the keys img_url and title.
#Append the dictionary with the image url string and the hemisphere title to a list. This list will contain one
#dictionary for each hemisphere.
# -
browser = Browser('chrome', headless=False)
url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
#return the data underneath the class 'item' tag
results = soup.find_all(class_='item')
print(results)
mars_images=[]
for result in results:
#reference the 'a' tags withing 'item' class
link=result.find('a')
#store the 'href', i.e. link in a variable 'links'
links = link['href']
#store the 'h3' tag text within item in a variable 'title'
title =result.find('h3').text
#attach the 'href' link from above to the main URL, so you can access the final page
url='https://astrogeology.usgs.gov'+links
browser = Browser('chrome', headless=False)
#visit the new concatenated URL
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
#reference the 'downloads' class, store in a variable 'infos'
infos = soup.find_all(class_='downloads')
#loop thru 'infos' pulling the 'a' tags, 'href', where full resolution image is stored
for info in infos:
link_two=info.find('a')
img_url=link_two['href']
print(title)
print(img_url)
#create a dictionary of title and image url
post={'img_url':img_url, 'title':title}
#append the dictionary to list/array 'mars_images'
mars_images.append(post)
mars_images
def init_browser():
browser = Browser('chrome', headless=False)
#return Browser("chrome", **executable_path, headless=False)
# +
def scrape():
browser = init_browser()
url = 'https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest'
response = requests.get(url)
time.sleep(1)
soup = BeautifulSoup(response.text, 'lxml')
#grabbling the 'slide' class element from the url
results = soup.find_all(class_="slide")
#creating a list to hold scraped data
news_data=[]
for result in results:
# Error handling
try: #loop thru and get the text within these classes, replace \n with blank space
news_p = result.find(class_="rollover_description_inner").text.replace('\n', '')
news_title = result.find(class_="content_title").text.replace('\n', '')
post = {"news_title": news_title,
"news_p":news_p}
news_data.append(post)
print(post)
except Exception as e:
print(e)
browser = Browser('chrome', headless=False)
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(url)
time.sleep(1)
#use splinter to click the "Full Image" button
browser.click_link_by_partial_text('FULL IMAGE')
time.sleep(1)
#HTML Object
html = browser.html
# Parse HTML with Beautiful Soup
soup = BeautifulSoup(html, 'html.parser')
#find the class where pic is stored
results = soup.find(class_='fancybox-image')
#retrieve source attribute, i.e. the path
url = results['src']
#attach the path to the main site link, this is the full image link
featured_image_url = 'https://www.jpl.nasa.gov'+url
post_two = {'featured_image':featured_image_url}
news_data.append(post_two)
print(post_two)
#visit the mars twitter page to get the Weather
url = 'https://twitter.com/marswxreport?lang=en'
browser.visit(url)
time.sleep(1)
response = requests.get(url)
#parse HTML with Beautiful soup, get the text
soup = BeautifulSoup(response.text, 'html.parser')
#get the text from the first p tag with appropriate class (from inspecting the site)
mars_weather = soup.find('p', class_="TweetTextSize TweetTextSize--normal js-tweet-text tweet-text").text
post_three = {'mars_weather':mars_weather}
print(post_three)
news_data.append(post_three)
browser = Browser('chrome', headless=False)
#visit the mars space facts site
url = 'https://space-facts.com/mars/'
#read the table, put into list variable
tables = pd.read_html(url)
#convert the list to a dataframe
mars_df =tables[0]
#put column headers on
mars_df.columns = ["Characteristic", "Value"]
#convert the datframe to dictionary, using 'records' orientation, this does not neeed to be, nor should be, appended to news_data, as will create a list of a dictionary within the list, and not be able to be inserted to mongodb
mars_dict=mars_df.to_dict('records')
print(mars_dict)
#Visit the site to get images of Mars Hemispheres
url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(url)
time.sleep(1)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
results = soup.find_all(class_='item')
#loop through the item class
for result in results:
#find the first a tag
link=result.find('a')
#assign the href to variable 'links'
links = link['href']
#assign the link h3 title text to variable 'title'
title =result.find('h3').text
#concatenate the path with the main site link, assign to variable 'url'
url='https://astrogeology.usgs.gov'+links
#open brower, chromedriver
browser = Browser('chrome', headless=False)
#visit the concatenated url
browser.visit(url)
time.sleep(1)
html = browser.html
#parse the html with beautiful soup
soup = BeautifulSoup(html, 'html.parser')
#find all elemenst with class 'downloads', assign results to variable list 'infos'
infos = soup.find_all(class_='downloads')
#loop thru infos, pull out links to images, assign with title to dictionary post, and then append to list
#mars_images
for info in infos:
link_two=info.find('a')
img_url=link_two['href']
post_four={'img_url':img_url, 'title':title}
news_data.append(post_four)
print(post_four)
#return your data, so it can be accessed by flask app (where the insertion into mongodb will occur)
return news_data+mars_dict
# -
scrape()
# +
from bs4 import BeautifulSoup
from splinter import Browser
import requests
import pymongo
import time
import pandas as pd
def init_browser():
browser = Browser('chrome', headless=False)
def scrape_mars_two():
browser = init_browser()
url = 'https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest'
response = requests.get(url)
time.sleep(1)
soup = BeautifulSoup(response.text, 'lxml')
#create an empty dictionary to store values
news_data={}
#find the appropriate div class containing target data to scrape, scrape text and assign to variables
news_p = soup.find_all('div',class_="rollover_description_inner")[0].text.strip()
news_title = soup.find_all('div',class_="content_title")[0].text.strip()
#put the variables into a dictionary
post = {"news_title": news_title,
"news_p":news_p}
print(post)
#update the main dictionary with the new dictionary
news_data.update(post)
browser = Browser('chrome', headless=False)
#new URL to scrape
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(url)
time.sleep(1)
#use splinter to click on button
browser.click_link_by_partial_text('FULL IMAGE')
time.sleep(1)
#HTML Object
html = browser.html
# Parse HTML with Beautiful Soup
soup = BeautifulSoup(html, 'html.parser')
#find the class where pic is stored
results = soup.find(class_='fancybox-image')
#retrieve source attribute, i.e. the path
url = results['src']
#attach the path to the main site link, this is the full image link
featured_image_url = 'https://www.jpl.nasa.gov'+url
#store the full image link in a dictionary
post_two = {'featured_image':featured_image_url}
#update the main dictionary with the new dictionary data
news_data.update(post_two)
print(post_two)
#visit the mars twitter page to get the Weather
url = 'https://twitter.com/marswxreport?lang=en'
browser.visit(url)
time.sleep(1)
response = requests.get(url)
#parse HTML with Beautiful soup, get the text
soup = BeautifulSoup(response.text, 'html.parser')
#get the text from the first p tag with appropriate class (from inspecting the site)
mars_weather = soup.find('p', class_="TweetTextSize TweetTextSize--normal js-tweet-text tweet-text").text
#store the weather tweet in a dictionary
post_three = {'mars_weather':mars_weather}
#update the maind dictionary with the new dictionary data
news_data.update(post_three)
print(post_three)
browser = Browser('chrome', headless=False)
#new URL to be scraped
url = 'https://space-facts.com/mars/'
#read the table, put into list variable
tables = pd.read_html(url)
#convert the list to a dataframe
mars_df =tables[0]
#set column names
mars_df.columns = ["Characteristics","Values"]
#set index to Characteristics
mars_df = mars_df.set_index(["Characteristics"])
#make it an html table
mars_df.to_html("mars_data.html")
#put the whole table into a dictionary item
table_post={"mars_table":mars_df}
#update the main dictionary with the table dictionary
news_data.update(table_post)
print(table_post)
browser = Browser('chrome', headless=False)
#new URL to be scraped
url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(url)
time.sleep(1)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
results = soup.find_all(class_='item')
#an empty array to store dictionary
image_data=[]
#loop through the item class
for result in results:
#find the first a tag
link=result.find('a')
#assign the href to variable 'links'
links = link['href']
#assign the link h3 title text to variable 'title'
title =result.find('h3').text
#concatenate the path with the main site link, assign to variable 'url'
url='https://astrogeology.usgs.gov'+links
#open brower, chromedriver
browser = Browser('chrome', headless=False)
#visit the concatenated url
browser.visit(url)
time.sleep(1)
html = browser.html
#parse the html with beautiful soup
soup = BeautifulSoup(html, 'html.parser')
#find all elemenst with class 'downloads', assign results to variable list 'infos'
infos = soup.find_all(class_='downloads')
for info in infos:
link_two=info.find('a')
img_url=link_two['href']
post_four={'img_url':img_url, 'title':title}
image_data.append(post_four)
#update main dictionary with new data
post_five={"image_data":image_data}
print(post_five)
news_data.update(post_five)
return news_data
# -
scrape_mars_two()
for news in news_data:
db.marsdata.insert_one(news)
db.marsdata.drop()
results = db.marsdata.find()
for result in results:
print(result)
| scrape_mars.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
<NAME>
CSCI 4350/5350
Dr. <NAME>
Honors Contract: Fall 2019
Program Description: Uses toy data sets to illustrate the effectiveness of Hybrid Clustering methods,
which utilize Spectral and Subspace clustering methods. Adjusting the values of gamma and sigma from
subspace and spectral clustering methods will affect performance.
Tweak gamma and sigma to get better results in "Find_Sigma_Gamma.ipynb".
"""
import matplotlib.pyplot as plt
import matplotlib
from mpl_toolkits.mplot3d import axes3d
# %matplotlib inline
import math
from cvxpy import *
import numpy as np
import scipy.spatial.distance as sdist
from sklearn.cluster import KMeans
from math import *
# +
# Ground truth
f = open("ground.in", "w+")
for x in range(100):
f.write(' {:0d}\n'.format(0))
for x in range(100):
f.write(' {:0d}\n'.format(1))
f.close()
# Convert the floating text in the file to integers when they come in
Xassign = np.loadtxt("ground.in").astype(np.int32)
#print(ground)
# +
# Make some toy data sets
# Set 1: 2 horizontal lines: 0 < y < 1
count = 0.1
f = open("lines-1.in", "w")
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:0d}\n'.format(0))
count += 0.1
count = 0.1
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:0d}\n'.format(1))
count += 0.1
f.close()
X = np.loadtxt("lines-1.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
# Can change to cvxpy.Parameter
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.193
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = .45
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# +
# Make some toy data sets
# Set 2: 2 horizontal lines 0 < y < 4
count = 0.1
f = open("lines-2.in", "w")
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:0d}\n'.format(0))
count += 0.1
count = 0.1
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:0d}\n'.format(4))
count += 0.1
f.close()
X = np.loadtxt("lines-2.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.193
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = .6
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# +
# Make some toy data sets
# Set 3: horizontal lines: Illustrates K-means performs as expected
# with enough space given between the 2 lines: 0 < y < 5
count = 0.1
f = open("lines-3.in", "w")
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:0d}\n'.format(0))
count += 0.1
count = 0.1
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:0d}\n'.format(5))
count += 0.1
f.close()
X = np.loadtxt("lines-3.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.053
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = 1.0
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# +
# Make some toy data sets
# Set 4: Interlocking Circles
# Make one circle
pi = math.pi
def PointsInCircum(r,n):
return [(math.cos(2*pi/n*x)*r,math.sin(2*pi/n*x)*r) for x in range(0,n+1)]
f = open("circle-1.in", "w")
circle1 = PointsInCircum(2, 100)
for x in range(100):
f.write('{:0f} '.format(circle1[x][0]))
f.write(' {:0f}\n'.format(circle1[x][1]))
f.close()
circle1 = np.loadtxt("circle-1.in")
#print(circle1)
# Make one circle shifted to the right.
def PointsInCircum_shift_right(r,n):
return [(math.cos(2*pi/n*x)*r + 2,math.sin(2*pi/n*x)*r) for x in range(0,n+1)]
f = open("circle-2.in", "w")
circle2 = PointsInCircum_shift_right(2, 100)
for x in range(100):
f.write('{:0f} '.format(circle2[x][0]))
f.write(' {:0f}\n'.format(circle2[x][1]))
f.close()
circle2 = np.loadtxt("circle-2.in")
# Bring the two circles together
f = open("interlocking_circles.in", "w")
for x in range(100):
f.write('{:0f} '.format(circle1[x][0]))
f.write(' {:0f}\n'.format(circle1[x][1]))
for x in range(100):
f.write('{:0f} '.format(circle2[x][0]))
f.write(' {:0f}\n'.format(circle2[x][1]))
f.close()
X = np.loadtxt("interlocking_circles.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.083
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = .8
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# +
# Make some toy data sets
# Set 5: Interlocking U's
# top half
def PointsInCircum_shift_up(r,n):
return [(math.cos(2*pi/n*x)*r,math.sin(2*pi/n*x)*r + 3) for x in range(0,n+1)]
f = open("u-1.in", "w")
u1 = PointsInCircum_shift_up(2, 200)
for x in range(100):
f.write('{:0f} '.format(u1[x+100][0]))
f.write(' {:0f}\n'.format(u1[x+100][1]))
f.close()
u1 = np.loadtxt("u-1.in")
# Make some toy data sets
# Interlocking U's: bottom half
def PointsInCircum(r,n):
return [(math.cos(2*pi/n*x)*r,math.sin(2*pi/n*x)*r) for x in range(0,n+1)]
f = open("u-2.in", "w")
u2 = PointsInCircum(2, 200)
for x in range(100):
f.write('{:0f} '.format(u2[x][0]))
f.write(' {:0f}\n'.format(u2[x][1]))
f.close()
u2 = np.loadtxt("u-2.in")
# Interlocking U's: Combine the upper and lower half.
f = open("interlocking_us.in", "w")
for x in range(100):
f.write('{:0f} '.format(u1[x][0]))
f.write(' {:0f}\n'.format(u1[x][1]))
for x in range(100):
f.write('{:0f} '.format(u2[x][0]))
f.write(' {:0f}\n'.format(u2[x][1]))
f.close()
X = np.loadtxt("interlocking_us.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.193
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = 1.25
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# +
# Make some toy data sets
# Set 6: 2 lines making an X
count = -50
f = open("x-1.in", "w")
for x in range(100):
f.write('{:2.2f}'.format(count))
f.write(' {:2.2f}\n'.format(count))
count += 1
count = -50
y = 0
for x in range(100):
y = (-1)*(count)
f.write('{:2.2f}'.format(count))
f.write(' {:2.2f}\n'.format(y))
count += 1
f.close()
X = np.loadtxt("x-1.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.193
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = 1.25
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# +
# Make some toy data sets
# Set 7: Circle within a circle
# Make a circle
def PointsInCircum_subset(r,n):
return [((math.cos(2*pi/n*x)*r), math.sin(2*pi/n*x)*r) for x in range(0,n+1)]
f = open("circle-3.in", "w")
circle3 = PointsInCircum_subset(6, 100)
for x in range(100):
f.write('{:0f} '.format(circle3[x][0]))
f.write(' {:0f}\n'.format(circle3[x][1]))
f.close()
circle3 = np.loadtxt("circle-3.in")
# Circle within a circle
f = open("subset_circles.in", "w")
for x in range(100):
f.write('{:0f} '.format(circle1[x][0]))
f.write(' {:0f}\n'.format(circle1[x][1]))
for x in range(100):
f.write('{:0f} '.format(circle3[x][0]))
f.write(' {:0f}\n'.format(circle3[x][1]))
f.close()
X = np.loadtxt("subset_circles.in")
# +
# Subspace Clustering
## Solve for each point as a linear system
## with respect to all other points using
## convex optimization routines
## Basically, we express each point as a
## linear combination of the other points...
## Here we try to minimize the dot-product between
## the coefficients. Sparsity is only needed in
## dot-product space in-so-far as possible.
A = np.transpose(X)
Coeff = np.zeros([np.shape(A)[1],np.shape(A)[1]])
for i in range(np.shape(A)[1]):
b = A[:,i]
# gamma must be positive due to DCP rules.
gamma = Parameter(nonneg="true")
constraints = None
# Construct the problem.
x = Variable(np.shape(A)[1])
## Lasso
obj = Minimize(gamma*norm(A*x-b,2) + norm(x, 1))
constraints = [x[i] == 0]
## constraints = [x[i] == 0, sum(x) == 1]
## L1-Perfect
## obj = Minimize(norm(x, 1))
## constraints = [A*x == b, x[i] == 0, sum(x) == 1]
## L1-Noisy
## obj = Minimize(norm(x, 1))
## constraints = [ A*x - b <= gamma, x[i] == 0, sum(x) == 1 ]
if [constraints == None]:
prob = Problem(obj)
else:
prob = Problem(obj,constraints)
## From the original code
gamma.value = 0.077
prob.solve(solver='ECOS')
Coeff[:,i] = np.transpose(x.value)
#print(Coeff.shape)
## Refine results...
## Only use magnitude of the coefficients (no negative values)
Coeff = np.abs(Coeff)
## Normalize each row - not needed but doesn't hurt on most examples
## Coeff = Coeff / numpy.apply_along_axis(numpy.max,1,Coeff)[:,None]
## Symmetrize
Coeff = Coeff + np.transpose(Coeff)
## Dimensions in each subspace
d1 = 1
d2 = 1
## What is K?
K = np.max([d1,d2])+1
#print(K)
## Select the top K coefficients
newCoeff = np.zeros(np.shape(Coeff))
indices = np.apply_along_axis(lambda x: np.argsort(x)[::-1],1,Coeff)[:,range(K)]
for x in range(np.shape(Coeff)[0]):
newCoeff[x,indices[x,:]] = Coeff[x,indices[x,:]]
## Normalize each row - again, not really needed
## newCoeff = newCoeff / numpy.apply_along_axis(numpy.max,1,newCoeff)[:,None]
## Symmetrize
newCoeff = newCoeff + np.transpose(newCoeff)
## Standard...
Xaff = newCoeff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
#Xnorm1 = Xnorm
Xaff1 = Xaff
#print(Xaff_D)
## Spectral Clustering
## Let's implement a simple spectral clustering of the data
## Set affinity scaling factor
sigma = .97
# Number of points in each subspace
N1 = 100
N2 = 100
## Calculate pairwise distances (Euclidean)
Xdist = sdist.squareform(sdist.pdist(X))
## Calculate affinities
Xaff = np.exp((-1.0*np.power(Xdist,2))/(2.0 * np.power(sigma,2)))
Xaff[range(N1+N2),range(N1+N2)] = 0.0
origXaff = Xaff
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Decomposition
#Evals, Evecs = np.linalg.eigh(Xnorm)
#Xnorm2 = Xnorm
Xaff2 = Xaff
#print(Xaff_D)
#print(Xaff_D.shape)
# Apply Hybrid method, combining the 2 matrices with matrix multiplication
#Xnorm = np.matmul(Xnorm1, Xnorm2)
Xaff = np.matmul(Xaff1, Xaff2)
## Get row sums
Xaff_D = np.diagflat(1.0 / np.sqrt(np.apply_along_axis(np.sum,0,Xaff)))
## Normalization
#Xaff_D = pow(Xaff_D, 1/2)
Xnorm = np.matmul(np.matmul(Xaff_D**.5,Xaff),Xaff_D**.5)
#Xnorm = np.matmul(np.matmul(Xaff_D,Xaff),Xaff_D)
## Might not want to show this if the data sizes (N1,N2) are large
plt.imshow(Xnorm)
## Decomposition
Evals, Evecs = np.linalg.eigh(Xnorm)
# -
## Next Check the eigenvalues
print(Evals[[-1,-2, -3]])
plt.plot(np.linspace(1,len(Evals),num=len(Evals)),Evals)
# +
## Perform clustering
Xnew = Evecs[:,[-1,-2]]
kmeans = KMeans(n_clusters=2).fit(Xnew)
## If you want to see the eigen vectors...
## Xone = Evecs[kmeans.labels_==0,:]
## Xtwo = Evecs[kmeans.labels_==1,:]
## plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## If you want it on the PCA projection...
#Xone = Xsvd[0][kmeans.labels_==0,:]
#Xtwo = Xsvd[0][kmeans.labels_==1,:]
#plt.plot(Xone[:,0],Xone[:,1],'bo',Xtwo[:,0],Xtwo[:,1],'ro')
## Let's quantify the performance
jointPDF = np.zeros([2,2])
for x in range(len(Xassign)):
jointPDF[Xassign[x],kmeans.labels_[x]] += 1
jointPDF /= len(Xassign)
jointPDF += 0.000000000000001
marginals = np.outer(np.apply_along_axis(np.sum,1,jointPDF),np.apply_along_axis(np.sum,0,jointPDF))
print('Mutual Information')
print(np.sum(jointPDF*np.log2(jointPDF/marginals)))
# Plot the Subspace clustering attempt at the data
# Red will correlate to all 0's seen in kmeans.labels_, blue correlates to all 1's.
plt.scatter(X[:,0], X[:,1], color = [["red", "blue"][i] for i in kmeans.labels_])
plt.ylabel('Y')
plt.xlabel('X')
plt.show()
# -
| Hybrid-Clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9g1TMjhX7FbJ"
# ## Model Evaluations
# Purpose of this notebook is to evaluate the final models developed on the test dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="sT_4hAz5Jnjs" outputId="a086e7b7-73c2-4267-ede0-bfc138344341"
from google.colab import drive
drive.mount('/content/gdrive')
# %cd gdrive/MyDrive/Capstone/
# + colab={"base_uri": "https://localhost:8080/"} id="jpQ9vz_jKSz4" outputId="6d843d87-ae08-4d79-b999-2efa2716ae5a"
# !pip install transformers
# + colab={"base_uri": "https://localhost:8080/", "height": 170, "referenced_widgets": ["98e8904f58d14d9dafb633eda00f63f1", "ab0f6c5e4fba444dbeb4d5b4ba34686b", "<KEY>", "872e03840ac744f195772ffa0466d5a1", "<KEY>", "0bb184c57c5d448e866f1dd584618723", "c13829c07f5c4a209b8bf1feb519a3d8", "942c7e0dfdc749ca8d98a36e6dab46cf"]} id="8s1yZ6JyKdN4" outputId="1e132ac7-7c27-448c-9366-a4d25a04d574"
import os
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset
from sklearn.metrics import classification_report, recall_score, f1_score, precision_score, precision_recall_curve, accuracy_score, confusion_matrix
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.feature_extraction import stop_words
from sklearn.model_selection import train_test_split
from torch.utils.data import DataLoader
from tqdm import tqdm
import transformers
from transformers import AdamW, get_cosine_schedule_with_warmup, get_cosine_with_hard_restarts_schedule_with_warmup
from tqdm import tqdm, tqdm_notebook, tqdm_pandas
tqdm.pandas()
tqdm_notebook().pandas()
# + id="D0FExNoHKrJK"
import itertools
from sklearn.metrics import roc_auc_score
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label',fontsize=15)
plt.xlabel('Predicted label',fontsize=15)
# + colab={"base_uri": "https://localhost:8080/"} id="XDIhHAvkKfWS" outputId="92a48244-9e92-4252-f0d3-589ad6e52c6b"
user_type_df = pd.read_csv('data/total_user_type.csv') # dataframe containing the user types and the
tf_score_df = pd.read_pickle('data/tf_score_df.pkl') # dataframe containing the TF-IDF scores
# + id="npkX0rCiLphJ"
# with open('data/train_df_tlstm.pkl', 'rb') as f:
# train_df = pickle.load(f)
# with open('data/test_df_tlstm.pkl', 'rb') as f:
# test_df = pickle.load(f)
# with open('data/val_df_tlstm.pkl', 'rb') as f:
# val_df = pickle.load(f)
# full_df = train_df.append(test_df).append(val_df)
with open('data/full_df.pkl', 'rb') as f:
full_df = pickle.load(f)
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="rzSqeOpfFgkA" outputId="86b6c323-262d-4d22-da21-0c79d14e1394"
full_df.head()
# + id="cIhKr3xCL5FQ"
full_df = full_df.merge(user_type_df, on = 'username').merge(tf_score_df, on ='username') # combine dataset with TF-IDF Scores and User types for test splits
test_df = full_df[full_df['split']=='test'] # get the testing split
# + id="gSpPMu-vMm4R"
def get_timestamp(x):
timestamp = []
for t in x:
timestamp.append(datetime.timestamp(t))
np.array(timestamp) - timestamp[-1]
return timestamp
class SuicidalDataset(Dataset):
def __init__(self, label, temporal, timestamp, tf_idf_vector):
super().__init__()
self.label = label
self.temporal = temporal
self.timestamp = timestamp
self.tf_idf_vector = tf_idf_vector
def __len__(self):
return len(self.label)
def __getitem__(self, item):
labels = torch.tensor(self.label[item])
result = self.temporal[item]
temporal_features = torch.tensor(result)
tf_idf_vector = self.tf_idf_vector[item]
timestamp = torch.tensor(get_timestamp(self.timestamp[item]))
return [labels, temporal_features, timestamp, tf_idf_vector]
def pad_ts_collate(batch):
target = [item[0] for item in batch]
data_post = [item[1] for item in batch]
timestamp_post = [item[2] for item in batch]
tf_idf_vector = [item[3] for item in batch]
lens_post = [len(x) for x in data_post]
data_post = nn.utils.rnn.pad_sequence(data_post, batch_first=True, padding_value=0)
timestamp_post = nn.utils.rnn.pad_sequence(timestamp_post, batch_first=True, padding_value=0)
target = torch.tensor(target)
lens_post = torch.tensor(lens_post)
tf_idf_vector = torch.tensor(tf_idf_vector).float()
return [target, data_post, timestamp_post, lens_post, tf_idf_vector]
# + id="qN6VwsPIM04k"
# create test dataset for testing the models
test_dataset = SuicidalDataset(test_df.label.values,
test_df.enc.values,
test_df.hist_dates.values,
test_df.tf_score.values)
BATCH_SIZE = 16
test_dataloader = DataLoader(test_dataset, batch_size=BATCH_SIZE, collate_fn=pad_ts_collate)
# + id="NHC9ilykSIhd"
# Model class defined
class HistoricCurrent(nn.Module):
def __init__(self, embedding_dim, hidden_dim, subreddit_embedding = 512, dropout = 0.2):
super().__init__()
self.historic_model = TimeLSTM(embedding_dim, hidden_dim[0])
self.dropout = nn.Dropout(dropout)
self.fc_tfidf = nn.Linear(799, subreddit_embedding)
current_dim = hidden_dim[0] + subreddit_embedding #concatenated
self.layers = nn.ModuleList()
for hdim in hidden_dim[1:]:
self.layers.append(nn.Linear(current_dim, hdim))
current_dim = hdim
self.layers.append(nn.Linear(current_dim, 2))
@staticmethod
def combine_features(features_1, features_2):
return torch.cat((features_1, features_2), 1)
def forward(self, historic_features, timestamp, tf_idf_vector):
outputs = self.historic_model(historic_features, timestamp)
outputs = torch.mean(outputs, 1)
subreddit_features = F.relu(self.fc_tfidf(tf_idf_vector))
combined_features = self.combine_features(subreddit_features, outputs)
x = self.dropout(combined_features)
for layer in self.layers[:-1]:
x = F.relu(layer(x))
x = self.dropout(x)
return self.layers[-1](x) # final layer
class TimeLSTM(nn.Module):
def __init__(self, input_size, hidden_size, bidirectional=True):
# assumes that batch_first is always true
super().__init__()
self.hidden_size = hidden_size
self.input_size = input_size
self.W_all = nn.Linear(hidden_size, hidden_size * 4)
self.U_all = nn.Linear(input_size, hidden_size * 4)
self.W_d = nn.Linear(hidden_size, hidden_size)
self.bidirectional = bidirectional
def forward(self, inputs, timestamps, reverse=False):
# inputs: [b, seq, embed]
# h: [b, hid]
# c: [b, hid]
b, seq, embed = inputs.size()
h = torch.zeros(b, self.hidden_size, requires_grad=False)
c = torch.zeros(b, self.hidden_size, requires_grad=False)
h = h.cuda()
c = c.cuda()
outputs = []
for s in range(seq):
c_s1 = torch.tanh(self.W_d(c))
c_s2 = c_s1 * timestamps[:, s:s + 1].expand_as(c_s1)
c_l = c - c_s1
c_adj = c_l + c_s2
outs = self.W_all(h) + self.U_all(inputs[:, s])
f, i, o, c_tmp = torch.chunk(outs, 4, 1)
f = torch.sigmoid(f)
i = torch.sigmoid(i)
o = torch.sigmoid(o)
c_tmp = torch.sigmoid(c_tmp)
c = f * c_adj + i * c_tmp
h = o * torch.tanh(c)
outputs.append(h)
if reverse:
outputs.reverse()
outputs = torch.stack(outputs, 1)
return outputs
# + id="L1GRhFl5ToC0"
def CB_loss(labels, logits, samples_per_cls, no_of_classes, loss_type, beta, gamma):
"""Compute the Class Balanced Loss between `logits` and the ground truth `labels`.
Class Balanced Loss: ((1-beta)/(1-beta^n))*Loss(labels, logits)
where Loss is one of the standard losses used for Neural Networks.
Args:
labels: A int tensor of size [batch].
logits: A float tensor of size [batch, no_of_classes].
samples_per_cls: A python list of size [no_of_classes].
no_of_classes: total number of classes. int
loss_type: string. One of "sigmoid", "focal", "softmax".
beta: float. Hyperparameter for Class balanced loss.
gamma: float. Hyperparameter for Focal loss.
Returns:
cb_loss: A float tensor representing class balanced loss
"""
effective_num = 1.0 - np.power(beta, samples_per_cls)
weights = (1.0 - beta) / np.array(effective_num)
weights = weights / np.sum(weights) * no_of_classes
labels_one_hot = F.one_hot(labels, no_of_classes).float()
weights = torch.tensor(weights, dtype=torch.float32).cuda()
weights = weights.unsqueeze(0)
weights = weights.repeat(labels_one_hot.shape[0], 1) * labels_one_hot
weights = weights.sum(1)
weights = weights.unsqueeze(1)
weights = weights.repeat(1, no_of_classes)
if loss_type == "focal":
cb_loss = focal_loss(labels_one_hot, logits, weights, gamma)
elif loss_type == "sigmoid":
cb_loss = F.binary_cross_entropy_with_logits(input=logits, target=labels_one_hot, weight=weights)
elif loss_type == "softmax":
pred = logits.softmax(dim=1)
cb_loss = F.binary_cross_entropy(input=pred, target=labels_one_hot, weight=weights)
return cb_loss
def focal_loss(labels, logits, alpha, gamma):
"""Compute the focal loss between `logits` and the ground truth `labels`.
Focal loss = -alpha_t * (1-pt)^gamma * log(pt)
where pt is the probability of being classified to the true class.
pt = p (if true class), otherwise pt = 1 - p. p = sigmoid(logit).
Args:
labels: A float tensor of size [batch, num_classes].
logits: A float tensor of size [batch, num_classes].
alpha: A float tensor of size [batch_size]
specifying per-example weight for balanced cross entropy.
gamma: A float scalar modulating loss from hard and easy examples.
Returns:
focal_loss: A float32 scalar representing normalized total loss.
"""
BCLoss = F.binary_cross_entropy_with_logits(input=logits, target=labels, reduction="none")
if gamma == 0.0:
modulator = 1.0
else:
modulator = torch.exp(-gamma * labels * logits - gamma * torch.log(1 + torch.exp(-1.0 * logits)))
loss = modulator * BCLoss
weighted_loss = alpha * loss
focal_loss = torch.sum(weighted_loss)
focal_loss /= torch.sum(labels)
return focal_loss
def loss_fn(output, targets, samples_per_cls, loss_type = "focal"):
beta = 0.9999
gamma = 2.0
no_of_classes = 2
return CB_loss(targets, output, samples_per_cls, no_of_classes, loss_type, beta, gamma)
# + id="yTmB0WuRQ1yN"
def eval_loop(model, dataloader, device, dataset_len):
model.eval()
running_loss = 0.0
running_corrects = 0
fin_targets = []
fin_outputs = []
fin_preds = []
for bi, inputs in enumerate(tqdm_notebook(dataloader, total=len(dataloader), leave=False)):
labels, data, timestamp, lens, tf_idf_vector = inputs
labels = labels.to(device)
data = data.to(device)
timestamp = timestamp.to(device)
tf_idf_vector = tf_idf_vector.to(device)
with torch.no_grad():
output = model(data, timestamp, tf_idf_vector)
fin_pred = output[:,1]
_ , preds = torch.max(output, 1)
loss = loss_fn(output, labels, labels.unique(return_counts=True)[1].tolist())
running_loss += loss.item()
running_corrects += torch.sum(preds == labels.data)
fin_targets.append(labels.cpu().detach().numpy())
fin_outputs.append(preds.cpu().detach().numpy())
fin_preds.append(fin_pred.cpu().detach().numpy())
epoch_loss = running_loss / len(dataloader)
epoch_accuracy = running_corrects.double() / dataset_len
return epoch_loss, epoch_accuracy, np.hstack(fin_outputs), np.hstack(fin_targets), np.hstack(fin_preds)
# + colab={"base_uri": "https://localhost:8080/"} id="BYIilI2-RbJ_" outputId="9624ca07-9727-46f2-c396-c54cef4ff73a"
device = 'cuda' if torch.cuda.is_available() else 'cpu'
print(device)
# + [markdown] id="cinxkDtGTGuB"
# ## Model 1 : HFHN + HFLN
# + colab={"base_uri": "https://localhost:8080/"} id="Udomo1czRyAX" outputId="d36d5e3f-606f-4000-c5f4-257ab74808df"
EPOCHS = 10
HIDDEN_DIM = [128,64]
EMBEDDING_DIM = 768
SUBREDDIT_EMBEDDING = 128
DROPOUT = 0.7
LEARNING_RATE = 0.001
hfhn_hfln = HistoricCurrent(EMBEDDING_DIM, HIDDEN_DIM, SUBREDDIT_EMBEDDING, DROPOUT)
hfhn_hfln.to(device)
hfhn_hfln.load_state_dict(torch.load('saved_model/best_model_TLSTM_hfhn_hfln.pt'))
# + [markdown] id="wZWufOWfUBfZ"
# ## Model 2: HFLN + LFLN
# + colab={"base_uri": "https://localhost:8080/"} id="xqNLanvEUTXz" outputId="a3087efd-2114-4d63-d86e-764d8a3e896e"
EPOCHS = 15
HIDDEN_DIM = [32,16]
EMBEDDING_DIM = 768
SUBREDDIT_EMBEDDING = 32
DROPOUT = 0.7
LEARNING_RATE = 0.001
hfln_lfln = HistoricCurrent(EMBEDDING_DIM, HIDDEN_DIM, SUBREDDIT_EMBEDDING, DROPOUT)
hfln_lfln.to(device)
hfln_lfln.load_state_dict(torch.load('saved_model/best_model_TLSTM_hfln_lfln.pt'))
# + [markdown] id="RcIrS6W_UTaO"
# ## Model 3: LFLN + LFHN
# + colab={"base_uri": "https://localhost:8080/"} id="jIlEYNBWUTcL" outputId="07da344c-a19e-43fd-d5e5-e22c83f10ef1"
EPOCHS = 15
HIDDEN_DIM = [8]
EMBEDDING_DIM = 768
SUBREDDIT_EMBEDDING = 8
DROPOUT = 0.7
LEARNING_RATE = 0.001
lfln_lfhn = HistoricCurrent(EMBEDDING_DIM, HIDDEN_DIM, SUBREDDIT_EMBEDDING, DROPOUT)
lfln_lfhn.to(device)
lfln_lfhn.load_state_dict(torch.load('saved_model/best_model_TLSTM_lfhn_lfln.pt'))
# + [markdown] id="GkAcwutwUTfV"
# ## Model 4: LFHN + HFHN
# + colab={"base_uri": "https://localhost:8080/"} id="qj6aAQx9UTh9" outputId="dade0cc6-5bf4-4056-8a33-b8878f295814"
EPOCHS = 15
HIDDEN_DIM = [32,16]
EMBEDDING_DIM = 768
SUBREDDIT_EMBEDDING = 32
DROPOUT = 0.7
LEARNING_RATE = 0.001
lfhn_hfhn = HistoricCurrent(EMBEDDING_DIM, HIDDEN_DIM, SUBREDDIT_EMBEDDING, DROPOUT)
lfhn_hfhn.to(device)
lfhn_hfhn.load_state_dict(torch.load('saved_model/best_model_TLSTM_lfhn_hfhn.pt'))
# + [markdown] id="BUoYgWv3_jVs"
# ---
# + [markdown] id="On7FUH1G_kjV"
# ## Model Evaluations
# + colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["184fb9032d2145af9d226a8e7b4e7a14", "83848eb6919d47b093f0d0f21ca8a044", "cfb8a8e0530a4e05af1454fcd75c1fb0", "c98254f382de41fc9d97cbcd80122218", "<KEY>", "30c69f29e53a4fd0a71745851c8007c9", "<KEY>", "<KEY>", "4da4ed657d934af082cc4bebd62d47c6", "aec69310ceee44a891d64e49b45b06d2", "<KEY>", "fbf56f98fc7a4946b3f01a7dbe4ff6ae", "09b50377beba4ce2a3cd86fd162a09b2", "5ceeac97f69647bca0a64ae28ce9123d", "<KEY>", "1d24835d4866483887e2a536154a0e7e", "<KEY>", "86e3e4d56dd842c0bdad6733b3d7526e", "<KEY>", "<KEY>", "fa0f585e2a484aee8a0c488a4c14a973", "49ab0b4a2c994c6aa61b34e5b061d7f9", "<KEY>", "3a95271a496f4a7f8df217f281938acc", "<KEY>", "bc4fa7999aed4d50863e4ded44f1671d", "<KEY>", "3585c38dff7044b78bed14b553efee21", "887f76f2d2a04919a638772852faade2", "c43a457215004227b3df536342410e80", "<KEY>", "fe32faf6a308480da6d089049aa82f11"]} id="kAJBujbuRMRX" outputId="887c6d88-2427-4b5f-9e88-9714a4a75745"
_, _, y_pred1, y_true1, y_out1 = eval_loop(hfhn_hfln, test_dataloader, device, len(test_dataset))
_, _, y_pred2, y_true2, y_out2 = eval_loop(hfln_lfln, test_dataloader, device, len(test_dataset))
_, _, y_pred3, y_true3, y_out3 = eval_loop(lfln_lfhn, test_dataloader, device, len(test_dataset))
_, _, y_pred4, y_true4, y_out4 = eval_loop(lfhn_hfhn, test_dataloader, device, len(test_dataset))
# + id="tUz_2E_4YdQ4"
total_preds = {'hfhn_hfln':y_pred1, 'hfln_lfln':y_pred2, 'lfln_lfhn':y_pred3, 'lfhn_hfhn':y_pred4}
total_preds_df = pd.DataFrame(total_preds)
total_preds_df['mean_score'] = pd.DataFrame(total_preds).mean(axis=1)
total_preds_df['true'] = y_true1
total_preds_df['username'] = test_df.username.tolist()
total_preds_df['target'] = test_df.label.tolist()
total_preds_df['type'] = test_df.type.tolist()
total_preds_df['dist'] = np.abs(total_preds_df['true'] - total_preds_df['mean_score'])
total_preds_df['decision_pred'] = 1*(total_preds_df['mean_score']>0.5) # simple decision boundary of >0.5
# + id="c1moSN9nlvbR"
def get_evaluation_metric(y_true, y_pred, model_name):
f1 = f1_score(y_true, y_pred)
prec = precision_score(y_true, y_pred)
recall = recall_score(y_true, y_pred)
accuracy = accuracy_score(y_true, y_pred)
print(f'For {model_name} -- F1_score :{np.round(f1,3)}, precision :{np.round(prec,3)}, recall : {np.round(recall,3)}, accuracy : {np.round(accuracy,3)}')
# + colab={"base_uri": "https://localhost:8080/"} id="tM9twU9fCOhA" outputId="014cab09-b4b1-42d8-f6af-000fb658abd9"
get_evaluation_metric(total_preds_df['hfhn_hfln'], y_true1, 'HFHN + HFLN')
get_evaluation_metric(total_preds_df['hfln_lfln'], y_true1, 'HFLN + LFLN')
get_evaluation_metric(total_preds_df['lfln_lfhn'], y_true1, 'LFLN + LFHN')
get_evaluation_metric(total_preds_df['lfhn_hfhn'], y_true1, 'LFHN + HFHN')
get_evaluation_metric(total_preds_df['decision_pred'],y_true1, 'Combined')
# + [markdown] id="wxxUarsvKjix"
# Best model is the combined one based on the accuracy.
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="Le78TmL8K3ni" outputId="363cb2cd-63cd-4045-d9a1-ef0cae0acd51"
cnf_matrix = confusion_matrix(total_preds_df['decision_pred'], y_true1)
plt.figure(figsize=(8,6))
plot_confusion_matrix(cnf_matrix, classes=['Not Suicidal','Suicidal'],normalize=True,
title='Combined Model Confusion Matrix')
# + id="GFVVcMpqD9uF"
# evaluate performcance based on group
types = total_preds_df.groupby('type')
types = [types.get_group(x) for x in types.groups]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="D45RXcFMEi6w" outputId="987b989b-19cd-4742-fdbd-ea5fd5a36d1c"
types[0].iloc[0].type
# + id="sPxWHecrEUkr"
def get_model_performances(df):
group = df.iloc[0].type
print(f'Geting predictions for group {group}')
get_evaluation_metric(df['hfhn_hfln'], df['true'], 'HFHN + HFLN')
get_evaluation_metric(df['hfln_lfln'], df['true'], 'HFLN + LFLN')
get_evaluation_metric(df['lfln_lfhn'], df['true'], 'LFLN + LFHN')
get_evaluation_metric(df['lfhn_hfhn'], df['true'], 'LFHN + HFHN')
get_evaluation_metric(df['decision_pred'],df['true'], 'Combined')
print("-----------------------------------")
# + colab={"base_uri": "https://localhost:8080/"} id="q-g68nF-EUnY" outputId="6118003f-357d-4a36-cfc0-48cada568783"
for i in types:
get_model_performances(i)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="ChB5QtiOEUpx" outputId="b90a6e42-f177-4102-8872-6f13226dd583"
test_df.groupby('type').count()
| final_model_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torchvision
import torchvision.transforms as transforms
transform = transforms.Compose([transforms.ToTensor, transforms.Normalize((0.5,0.5,0.5), (0.5,0.5,0.5))])
trainset = torchvision.datasets.CIFAR10()
| archive/scratch_pad/pytorch_3_cifar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # One-dimensional advection equation
# v1.64, 20 February 2018, by <NAME>
#
# $\newcommand{\V}[1]{\vec{\boldsymbol{#1}}}$
# $\newcommand{\I}[1]{\widehat{\boldsymbol{\mathrm{#1}}}}$
# $\newcommand{\pd}[2]{\frac{\partial#1}{\partial#2}}$
# $\newcommand{\pdt}[1]{\frac{\partial#1}{\partial t}}$
# $\newcommand{\ddt}[1]{\frac{\D#1}{\D t}}$
# $\newcommand{\D}{\mathrm{d}}$
# $\newcommand{\Ii}{\I{\imath}}$
# $\newcommand{\Ij}{\I{\jmath}}$
# $\newcommand{\Ik}{\I{k}}$
# $\newcommand{\del}{\boldsymbol{\nabla}}$
# $\newcommand{\dt}{\cdot}$
# $\newcommand{\x}{\times}$
# $\newcommand{\dv}{\del\cdot}$
# $\newcommand{\curl}{\del\times}$
# $\newcommand{\lapl}{\nabla^2}$
#
# This notebook is similiar in style to `DiffusionPDE1D`, except that we investigate modelling of one-dimensional advection:
# $$
# \pdt{b} = -u \pd{b}{x}
# $$
# In particular, we study the accuracy and stability of the discrete form of above equation.
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import display,clear_output
import time as Time
import math
import numpy as np
import matplotlib
matplotlib.rcParams.update({'font.size': 22})
from IPython.display import Image
from IPython.core.display import HTML
import urllib.request
HTML(urllib.request.urlopen('http://metrprof.xyz/metr4323.css').read().decode())
#HTML( open('metr4323.css').read() ) #or use this, if you have downloaded metr4233.css to your computer
#
# The symbol $u$ now has the familiar meaning. It is a velocity component in:
# $$
# \vec{U} = u\Ii + v\Ij + w\Ik
# $$
#
# We model advection of a *conserved* quantity $b$:
# $$
# \ddt{b} = \pdt{b} + \vec{U} \dt \del b = 0
# $$
#
# So being conserved, at a grid point $b$ changes as:
# $$
# \pdt{b} = - \vec{U} \dt \del b = -u \pd{b}{x} - v \pd{b}{y} - w \pd{b}{z}
# $$
#
# The three terms on the right-hand-side are similar. So, for learning how to model those terms, we consider the first term in isolation. Also, we do not prognosticate $u(x,t)$, but just specify it:
# $$
# \pdt{b} = -u \pd{b}{x}
# $$
# You might think that an obvious, simple choice to use in a discrete model is this
# centered difference:
# $$
# u_n\left( \pd{b}{x} \right)_n = u_n \frac{b_{n+1} - b_{n-1}}{2 \Delta x}
# $$
#
# Here is our plan:
# * As we shall see in our explorations, there may options better than the centered difference.
# * When we make our forward time-step, how big can $\Delta t$ be, without the model "blowing up"? There is a very intuitive limit on $\Delta t$: [Courant–Friedrichs–Lewy condition](http://en.wikipedia.org/wiki/Courant%E2%80%93Friedrichs%E2%80%93Lewy_condition)
# dqdt requires a list of the time derivatives for q, stored
# in order from present to the past
def ab_blend(dqdt,order):
if order==1:
return dqdt[0]
elif order==2:
return 1.5*dqdt[0]-.5*dqdt[1]
elif order==3:
return (23*dqdt[0]-16*dqdt[1]+5*dqdt[2])/12.
else:
print("order", order ," not supported ")
# Run All Below when changing N, the number of gridpoints
# make the grid
N = 101 # number of x grid points for u. Try 101,201,401. May need to decrease dt for larger N.
xmax = 20 # 0 <= x <= xmax
dx = xmax/(N-1.) # grid width
xu = np.linspace(0,xmax,N) # grid for u
#print(xu)
# initialize u, b
xc = xmax/6. # put peak to the left
#xc = xmax/2. # put peak at center, for oscillating u field
bi = np.exp( -4.*(xu-xc)**2 ) # Gaussian for initial field of b
ui = np.sin(np.pi*xu/xmax)
#print(bi)
#print(ui)
plt.plot(xu,bi,'b',xu,ui,'r');
startover = True
if 'xsave' not in vars() or startover: bsave={} # for saving b at final time, to compare schemes. dict key will be a label for the expt.
if 'xsave' not in vars() or startover: xsave={} # for saving the grid in various experiments
# <hr>
# Run three simulations. In the cell below, set `advord=1`, `advord=2` then `advord=3`. Then run that cell, and the one below it. Your three results will be stored in `bsave` and `xsave`. If you need to start over, execute the above cell.
#
# You should find the "third-order" schemes for advection and time-stepping to be mysterious at this point. In a subsequent notebook, these schemes will be derived.
# Run All Below for changes other than N
dt = .05# try dt=.05 with N=101
tstop = 20. # stop when t>tstop
dplot = 1. # time increment between plots
period = 20. # period of oscillation of the u field
#freq=2.*np.pi/period # frequency of the oscillation
freq=0. #for steady u field
aborder = 3 # Adams-Bashforth order: 1 2 or 3
advord = 1 # advection order: 1 2 or 3
CFL = dt*1./dx # velocity scale is 1, so this is the CFL number
expt = '%d,%d,%3.2f,%d' % (aborder, advord, CFL, N)
print(expt)
# +
u = ui.copy()
b = bi.copy()
dudt = 0.*u # easy way to make dudt same size as u, with dudt=0
dbdt = 0.*b
dbdx = 0.*b
dbdta = [0.,0.,0.] # items will be arrays of dbdt, for Adams-Bashforth
nstep = 0
t = 0. # initial value for t
tplot = 0. # time t for next plot
f, ax = plt.subplots(figsize=(8,6))
display(f)
while t < tstop + dt/2.:
nstep+=1
abnow=min(nstep,aborder)
if tplot-dt/2.< t <tplot+dt/2.: #plot
Time.sleep(1./20.)
ax.clear()
ax.plot(xu,b,'b-*')
ax.plot(xu,u,'r')
ax.text(1,.9,expt,fontsize=22)
ax.text(1,.5,'t={0:5.3f}'.format(t),fontsize=22)
clear_output(wait=True)
ax.set_xlabel('x')
ax.set_ylabel('b',rotation=0)
ax.set_ylim((-.5,1.5))
display(f)
tplot = min(tstop, t+dplot)
if t<dt/2.: Time.sleep(2.)
if t>tstop: break
u = ui*np.cos(freq*t) # oscillating u in time is possible
if advord == 2: # 2nd order centered: uses 1 pt on each side of db/dx location
dbdx[1:-1] = (-b[0:-2] + b[2:])/(2*dx) #
elif advord == 1: #1st order upwind: uses 1 pt at upwind side and 1 pt at db/dx location
dbdx[1:-1] = np.where( u[1:-1]>0.,
-b[0:-2] + b[1:-1] # use upwind point (which is on the left)
, - b[1:-1] + b[2:] # use upwind point (which is on the right)
)/dx
# The boundary points AND the points adjacent are avoided. Keep b away from boundaries!
elif advord == 3: #3rd order upwind: uses 2 pts upwind, 1 pt downwind and 1 pt at db/dx location
dbdx[2:-2] = np.where( u[2:-2]>0.,
+ b[:-4] - 6*b[1:-3] + 3*b[2:-2] + 2*b[3:-1] ,
- 2*b[1:-3] - 3*b[2:-2] + 6*b[3:-1] - b[4:]
)/(6*dx)
dbdt[1:-1] = - u[1:-1]*dbdx[1:-1] #advection term
# comment-out the above, and uncomment below for AB 1, 2 and 3
dbdta = [dbdt.copy()] + dbdta[:-1] # retain latest three time-derivatives
b += dt*ab_blend(dbdta,abnow) # Adams-Bashforth blending of time derivatives
t += dt
bsave[expt] = b.copy() # save final value of b for comparison plot
xsave[expt] = xu.copy() # save for plotting
plt.close() # prevents mysterious second plot from popping up
# -
bsave.keys()
fig, bax = plt.subplots(figsize=(8,6))
bax.plot(xu,bi,'k-*')
style=['b-*','r*-','g*-','c*-']
n=0
for expk in sorted(bsave.keys()):
sty=style[n%3]
n += 1
bax.plot(xsave[expk],bsave[expk],sty,label=expk)
bax.grid()
bax.set_ylabel('b',rotation=0)
bax.set_xlabel('x')
bax.set_ylim( (-.5,1.5))
bax.set_title('b at t=20 for various experiments')
#bax.text(1,-.2,"sample",fontsize=96,alpha=.1)
bax.legend(loc=0)
fig.savefig('saveme.png');
# # Save your work!
#
# This is what you are striving for in the first task:
#
# <img src='http://metr4323.net/img/advord_exp.png'>
#
#
# # Student tasks
#
# ## 1. advection orders
# Make a comparative study the three advection orders advord=1, advord=2 then advord=3. Rename the saved image to `saveme1.png`, so that this notebook does not overwrite it. Display the image. (It should look like the above image)
#
# <img src="saveme1.png">
#
# ## 2. CFL comparison
# Using 3rd order advection and 3rd order AB, adjust `dt` to make CFL number of: 0.125 0.25 0.5, 0.75. Is a CFL number of 0.25 apparently "small enough" for accuracy? Save the image and display it.
#
# <img src="saveme2.png">
#
# ## 3. Adams-Bashforth orders
# Make a study of 3rd order upwind advection for the three orders of Adams-Bashforth. Save the image and display it.
#
# <img src="saveme3.png">
#
#
# ## 4. Grid resolution
# Make a study of 3rd order upwind advection and 3rd Adams-Bashforth, with N=51 N=101 N=201. Keep the CFL number constant (and use a stable one). Save the image and display it.
#
# <img src="saveme4.png">
#
| Fiedler_revised/n050_advectionpde1d.ipynb |