code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 3 Exploration: Depth Images
#
# In this notebook, we will learn how to use depth images from the racecar's camera to identify the distance at specific points and find the closest pixel. We will also explore strategies for handling noise/measurement error.
#
# Throughout this notebook, **<font style="color:red">text in bold red</font>** indicates a change you must make to the following code block before running it.
#
#
# ## Table of Contents
# 1. [Getting Started](#GettingStarted)
# 2. [Taking Depth Photos](#TakingDepthPhotos)
# 3. [Handling Noise](#HandlingNoise)
# 4. [Closest Point](#ClosestPoint)
# <a id="GettingStarted"></a>
# ## 1. Getting Started
#
# **<font style="color:red">If you are running the car in RacecarSim, set `isSimulation` to `True`</font>**. Leave `isSimulation` `False` if you are using a physical car.
# TODO: Update isSimulation if necessary
isSimulation = True
# Next, we will import the necessary libraries for this notebook, including Python libraries (`cv`, `numpy`, etc.) and the Racecar library (`racecar_core`).
# +
# Import Python libraries
import cv2 as cv
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets
import statistics
from nptyping import NDArray
from typing import Any, Tuple, List, Optional
# Import Racecar library
import sys
sys.path.append("../../library")
import racecar_core
# -
# The following functions will help us throughout this notebook.
# +
def show_depth_image(
depth_image: NDArray[(Any, Any), np.float32],
max_depth: int = 400,
points: List[Tuple[int, int]] = []
) -> None:
"""
Displays a color image in the Jupyter Notebook.
Args:
depth_image: The image to display.
max_depth: The farthest depth to show in the image in cm. Anything past this depth is shown as black.
points: A list of points in (pixel row, pixel column) format to show on the image colored dots.
"""
# Clip anything above max_depth
np.clip(depth_image, None, max_depth, depth_image)
# Shift down slightly so that 0 (no data) becomes the "farthest" color
depth_image = (depth_image - 1) % max_depth
# Convert depth values to colors
color_image = cv.applyColorMap(-cv.convertScaleAbs(depth_image, alpha=255/max_depth), cv.COLORMAP_INFERNO)
# Draw a dot at each point in points
for point in points:
cv.circle(color_image, (point[1], point[0]), 6, (0, 255, 0), -1)
# Show the image with Matplotlib
plt.imshow(cv.cvtColor(color_image, cv.COLOR_BGR2RGB))
plt.show()
def add_noise(
depth_image: NDArray[(Any, Any), np.float32],
error_percent = 0.1,
null_percent: float = 0.005
) -> NDArray[(Any, Any), np.float32]:
"""
Adds noise to a depth image.
Args:
depth_image: The original image to which to add noise.
error_percent: The error percentage to introduce to each measurement.
null_percent: The percentage of pixels to set to zero.
Returns:
A copy of the provided depth_image with noise added.
"""
# Copy the original image
image = np.copy(depth_image)
# Apply error_percent to each measurement (gaussian error)
gauss = np.random.normal(1, error_percent, image.shape)
image *= gauss
# Add null (zero) values
num_nulls = int(image.size * null_percent)
coords = [np.random.randint(0, i - 1, num_nulls) for i in image.shape]
image[tuple(coords)] = 0.0
return image
def crop(
image: NDArray[(Any, ...), Any],
top_left_inclusive: Tuple[float, float],
bottom_right_exclusive: Tuple[float, float]
) -> NDArray[(Any, ...), Any]:
"""
Crops an image to a rectangle based on the specified pixel points.
Args:
image: The color or depth image to crop.
top_left_inclusive: The (row, column) of the top left pixel of the crop rectangle.
bottom_right_exclusive: The (row, column) of the pixel one past the bottom right corner of the crop rectangle.
Returns:
A cropped version of the image.
Note:
The top_left_inclusive pixel is included in the crop rectangle, but the
bottom_right_exclusive pixel is not.
If bottom_right_exclusive exceeds the bottom or right edge of the image, the
full image is included along that axis.
"""
# Extract the minimum and maximum pixel rows and columns from the parameters
r_min, c_min = top_left_inclusive
r_max, c_max = bottom_right_exclusive
# Shorten the array to the specified row and column ranges
return image[r_min:r_max, c_min:c_max]
# -
# Finally, we will create a racecar object. If this step fails, make sure that `isSimulation` has the correct value.
# Create Racecar
rc = racecar_core.create_racecar(isSimulation)
# <a id="TakingDepthPhotos"></a>
# ## 2. Taking Depth Photos
# A depth photo is similar to a color photo, except that each pixel stores a distance value rather than color values. In Jupyter Notebook, we can take a depth photo with the car's camera using `rc.camera.get_depth_image_async()`. Outside of Jupyter Notebook, we must use `rc.camera.get_depth_image()` instead.
#
# In order to make sense of the result, we will use `show_depth_image` to convert the distance measurements into colors. For example, the closest measurements are shown as bright yellow, ranging to red to purple to black (out of range).
# Take a depth photo and show a colorized-version
image = rc.camera.get_depth_image_async()
show_depth_image(image)
# Depth images are stored as two-dimensional numpy arrays, using a similar format to color images:
#
# * **0th dimension**: pixel rows, indexed from top to bottom.
# * **1st dimension**: pixel columns, indexed from left to right.
#
# Let's inspect the distance at the center of the image. **<span style="color:red">Set `center_row` and `center_col` in the following code block to the center of the image.</span>** You will likely wish to use `rc.camera.get_height()` and `rc.camera.get_width()`.
# +
# TODO: Calculate center row and column
center_row = rc.camera.get_height() // 2
center_col = rc.camera.get_width() // 2
# Print the distance of the center pixel
center_distance = image[center_row][center_col]
print(f"Distance at pixel {(center_row, center_col)}: {center_distance:.2f} cm")
# -
# <a id="HandlingNoise"></a>
# ## 3. Handling Noise
#
# As you saw in the previous section, we can calculate the distance of an object directly in front of the car by simply accessing the middle element of the depth image. In practice, however, this approach is not reliable because all sensors have some amount of *noise*, a random variation in measured values. Furthermore, some pixels may not receive any data, and thus have a *null value* of 0.0 cm.
#
# To simulate this, the following code block randomly adds noise and null values to our image.
noisy_image = add_noise(image)
show_depth_image(noisy_image)
# **Can you identify the noise and null values in this image?**
#
# To see why this may be a problem, we will measure the center distance ten times with a new noisy version of our original image each time.
# +
distances = []
# Randomize the image and calculate center distance 10 times
for i in range (1, 11):
noisy_image = add_noise(image)
center_distance = noisy_image[rc.camera.get_height() // 2][rc.camera.get_width() // 2]
print(f"Center distance {i}: {center_distance:.2f} cm")
distances.append(float(center_distance))
# Calculate the mean and standard deviation of the center distance measurement
print(f"\nMean: {statistics.mean(distances):.2f} cm")
print(f"Standard deviation: {statistics.stdev(distances):.2f} cm")
# -
# **What is the standard deviation across these trials?** To put that in perspective, suppose that we wanted to use the center distance to estimate the speed of the car. If the car was standing still but the center distance changed by 5 cm per frame, we would estimate that the car was traveling over 3 m/s, more than the top speed of the car!
#
# With noise, a single pixel is not a reliable measurement of distance. Instead, we should factor in neighboring pixels to mitigate the effect of error/noise. One way to do this is by applying a [Gaussian blur](https://en.wikipedia.org/wiki/Gaussian_blur) to the original image. Each pixel is updated with a weighted average of its neighbors, with greater weight given to closer neighbors. The *kernel size* determines how large of an area to include in this average.
#
# In the following code block, we use the OpenCV function [GaussianBlur](https://docs.opencv.org/4.3.0/d4/d86/group__imgproc__filter.html#gaabe8c836e97159a9193fb0b11ac52cf1) to apply Gaussian blur to our depth image.
# Kernel size must be odd
kernel_size = 11
blurred_image = cv.GaussianBlur(image, (kernel_size, kernel_size), 0)
show_depth_image(blurred_image)
# Aside: If your image contained areas that were out of range, you may notice a sharp outline appearing along the boundary of these regions. This occurs because out of range pixels are represented as 0.0, so when averaged with surrounding areas, they cause the average to *decrease*. This causes pixel near out-of-range regions to appear *closer* after blurring.
#
# Run the following code block and use the slider to experiment with different kernel sizes. **As kernel size increases, does the image become more or less blurred? Why?**
# +
def blur_image(depth_image, kernel_size):
# Blur and show image
blurred_image = cv.GaussianBlur(image, (kernel_size, kernel_size), 0)
show_depth_image(blurred_image)
widgets.interact(blur_image,
depth_image=widgets.fixed(image),
kernel_size=widgets.IntSlider(1, 1, 61, 2, continuous_update = False))
# -
# Each individual pixel in the blurred image is an average of many pixels from the original image. This helps compensate for noise and null values.
#
# You now have all of the tools necessary to write a more robust center distance algorithm. **<span style="color:red">Finish writing `get_depth_image_center_distance` in the following code block.</span>**
def get_depth_image_center_distance(
depth_image: NDArray[(Any, Any), np.float32],
kernel_size: int
) -> float:
"""
Finds the distance of a pixel averaged with its neighbors in a depth image.
Args:
depth_image: The depth image to process.
pix_coord: The (row, column) of the pixel to measure.
kernel_size: The size of the area to average around the pixel.
Returns:
The distance in cm of the object at the provided pixel.
Warning:
kernel_size must be positive and odd.
Note:
The larger the kernel_size, the more that the requested pixel is averaged
with the distances of the surrounding pixels. This helps reduce noise at the
cost of reduced accuracy.
"""
blur = cv.GaussianBlur(depth_image,(kernel_size,kernel_size),0)
center = blur[rc.camera.get_height() // 2][rc.camera.get_width() // 2]
return center
# Let's repeat the same test as before using our `get_depth_image_center_distance` function. Once again, we will randomly add noise to the original image and measure the center distance ten times.
# +
kernel_size = 11
distances = []
# Randomize the image and calculate center distance 10 times
for i in range (1, 11):
noisy_image = add_noise(image)
center_distance = get_depth_image_center_distance(noisy_image, kernel_size)
print(f"Center distance {i}: {center_distance:.2f} cm")
distances.append(float(center_distance))
# Calculate the mean and standard deviation of the center distance measurement
print(f"\nMean: {statistics.mean(distances):.2f} cm")
print(f"Standard deviation: {statistics.stdev(distances):.2f} cm")
# -
# Compare these results to our original test without blurring. **Has the standard deviation decreased?**
# <a id="ClosestPoint"></a>
# ## 4. Closest Point
#
# We can also use depth images to find the closest point, which is useful for identifying and reacting to nearby objects. Once again, we should apply a Gaussian blur to minimize the impact of noise.
#
# However, a problem will arise if any part of the depth image is out of range, as it will have a depth value of 0.0. To fix this, we can shift down each value by a small amount (such as 0.01 cm) and then mod by a large number (such as 10,000 cm). This way, 0.0 becomes -0.01, which after modding becomes 9,999.99 cm, a very large distance that will not interfere with the true minimum.
#
# **<span style="color:red">Finish writing `get_closest_pixel` to find the row and column of the closest pixel in a depth image.</span>**. You will likely wish to use the OpenCV function [minMaxLoc](https://docs.opencv.org/4.3.0/d2/de8/group__core__array.html#gab473bf2eb6d14ff97e89b355dac20707). Note that the positions returned by ``minMaxLoc`` are in (column, row) format, while `get_closest_pixel` should return in (row, column) format.
def get_closest_pixel(
depth_image: NDArray[(Any, Any), np.float32],
kernel_size: int = 5
) -> Tuple[int, int]:
"""
Finds the closest pixel in a depth image.
Args:
depth_image: The depth image to process.
kernel_size: The size of the area to average around each pixel.
Returns:
The (row, column) of the pixel which is closest to the car.
Warning:
kernel_size be positive and odd.
It is highly recommended that you crop off the bottom of the image, or else
this function will likely return the ground directly in front of the car.
Note:
The larger the kernel_size, the more that the depth of each pixel is averaged
with the distances of the surrounding pixels. This helps reduce noise at the
cost of reduced accuracy.
"""
depth_image = (depth_image - 0.01) % 10000
blurred_image = cv.GaussianBlur(depth_image, (kernel_size, kernel_size), 0)
(_, _, minLoc, _) = cv.minMaxLoc(blurred_image)
return (minLoc[1], minLoc[0])
# Let's use `get_closest_pixel` to draw a green dot at the location of the closest pixel.
closest_pixel = get_closest_pixel(image)
show_depth_image(image, points=[closest_pixel])
# Unless something is directly in front of the camera, the closest point was likely the ground in front of the car. This is not a particularly useful result, so we should first crop off the bottom of the image.
#
# Right now, `top_left_inclusive` and `bottom_right_exclusive` contain (row, column) pairs which include the entire image. **<span style="color:red">Update `top_left_inclusive` and `bottom_right_exclusive` to crop off the bottom third of the image before running `get_depth_image_center_distance`.</span>**
# TODO: Change top_left_inclusive and/or bottom_right_exclusive to crop off the bottom third of the image
top_left_inclusive = (0, 0)
bottom_right_exclusive = (rc.camera.get_height()* 2 //3, rc.camera.get_width())
cropped_image = crop(image, top_left_inclusive, bottom_right_exclusive)
closest_pixel = get_closest_pixel(cropped_image)
show_depth_image(cropped_image, points=[closest_pixel])
# ### The image should now show a more meaningful closest point. If necessary, you may need to experiment with different crop windows or move the car and take a new depth image.
#
# You are now ready to begin using the depth camera to implement a "safety stop" feature in `lab3a.py`. Good luck, and don't be afraid to ask questions!
| labs/lab3/lab3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ----
# <img src="../../../files/refinitiv.png" width="20%" style="vertical-align: top;">
#
# # Data Library for Python
#
# ----
# ## Content layer - Ownership - Consolidated
# This notebook demonstrates how to retrieve Ownership consolidated data.
# #### Learn more
#
# To learn more about the Refinitiv Data Library for Python please join the Refinitiv Developer Community. By [registering](https://developers.refinitiv.com/iam/register) and [logging](https://developers.refinitiv.com/content/devportal/en_us/initCookie.html) into the Refinitiv Developer Community portal you will have free access to a number of learning materials like
# [Quick Start guides](https://developers.refinitiv.com/en/api-catalog/refinitiv-data-platform/refinitiv-data-library-for-python/quick-start),
# [Tutorials](https://developers.refinitiv.com/en/api-catalog/refinitiv-data-platform/refinitiv-data-library-for-python/learning),
# [Documentation](https://developers.refinitiv.com/en/api-catalog/refinitiv-data-platform/refinitiv-data-library-for-python/docs)
# and much more.
#
# #### Getting Help and Support
#
# If you have any questions regarding using the API, please post them on
# the [Refinitiv Data Q&A Forum](https://community.developers.refinitiv.com/spaces/321/index.html).
# The Refinitiv Developer Community will be happy to help.
# ## Set the configuration file location
# For a better ease of use, you have the option to set initialization parameters of the Refinitiv Data Library in the _refinitiv-data.config.json_ configuration file. This file must be located beside your notebook, in your user folder or in a folder defined by the _RD_LIB_CONFIG_PATH_ environment variable. The _RD_LIB_CONFIG_PATH_ environment variable is the option used by this series of examples. The following code sets this environment variable.
import os
os.environ["RD_LIB_CONFIG_PATH"] = "../../../Configuration"
# ## Some Imports to start with
# + pycharm={"name": "#%%\n"}
import refinitiv.data as rd
from refinitiv.data.content import ownership
import datetime
# -
# ## Open the data session
#
# The open_session() function creates and open sessions based on the information contained in the refinitiv-data.config.json configuration file. Please edit this file to set the session type and other parameters required for the session you want to open.
rd.open_session("platform.rdp")
# ## Retrieve data
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Consolidated - Breakdown
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.breakdown.Definition("LSEG.L", ownership.StatTypes.INVESTOR_TYPE).get_data()
response.data.df
# -
# ### Consolidated - Concentration
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.concentration.Definition("LSEG.L").get_data()
response.data.df
# -
# ### Consolidated - Investors
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.investors.Definition(["LSEG.L", "VOD.L"], limit=3).get_data()
response.data.df
# -
# ### Consolidated - Recent activity
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.recent_activity.Definition("LSEG.L", "asc").get_data()
response.data.df
# -
# ### Consolidated - Shareholders history report
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.shareholders_history_report.Definition(
"LSEG.L",
ownership.Frequency.MONTHLY,
limit=3,
end=datetime.datetime.now()
).get_data()
response.data.df
# -
# ### Consolidated - Shareholders report
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.shareholders_report.Definition("LSEG.L", limit=3).get_data()
response.data.df
# -
# ### Consolidated - Top n concentration
# + pycharm={"name": "#%%\n"}
response = ownership.consolidated.top_n_concentration.Definition("LSEG.L", 40).get_data()
response.data.df
# -
# ### Close the session
# + pycharm={"name": "#%%\n"}
rd.close_session()
# + pycharm={"name": "#%%\n"}
| Examples/2-Content/2.10-Ownership/EX-2.10.01-Ownership-Consolidated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Detrending, Stylized Facts and the Business Cycle
#
# In an influential article, Harvey and Jaeger (1993) described the use of unobserved components models (also known as "structural time series models") to derive stylized facts of the business cycle.
#
# Their paper begins:
#
# "Establishing the 'stylized facts' associated with a set of time series is widely considered a crucial step
# in macroeconomic research ... For such facts to be useful they should (1) be consistent with the stochastic
# properties of the data and (2) present meaningful information."
#
# In particular, they make the argument that these goals are often better met using the unobserved components approach rather than the popular Hodrick-Prescott filter or Box-Jenkins ARIMA modeling techniques.
#
# statsmodels has the ability to perform all three types of analysis, and below we follow the steps of their paper, using a slightly updated dataset.
# + jupyter={"outputs_hidden": false}
# %matplotlib inline
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
from IPython.display import display, Latex
# -
# ## Unobserved Components
#
# The unobserved components model available in statsmodels can be written as:
#
# $$
# y_t = \underbrace{\mu_{t}}_{\text{trend}} + \underbrace{\gamma_{t}}_{\text{seasonal}} + \underbrace{c_{t}}_{\text{cycle}} + \sum_{j=1}^k \underbrace{\beta_j x_{jt}}_{\text{explanatory}} + \underbrace{\varepsilon_t}_{\text{irregular}}
# $$
#
# see Durbin and Koopman 2012, Chapter 3 for notation and additional details. Notice that different specifications for the different individual components can support a wide range of models. The specific models considered in the paper and below are specializations of this general equation.
#
# ### Trend
#
# The trend component is a dynamic extension of a regression model that includes an intercept and linear time-trend.
#
# $$
# \begin{align}
# \underbrace{\mu_{t+1}}_{\text{level}} & = \mu_t + \nu_t + \eta_{t+1} \qquad & \eta_{t+1} \sim N(0, \sigma_\eta^2) \\\\
# \underbrace{\nu_{t+1}}_{\text{trend}} & = \nu_t + \zeta_{t+1} & \zeta_{t+1} \sim N(0, \sigma_\zeta^2) \\
# \end{align}
# $$
#
# where the level is a generalization of the intercept term that can dynamically vary across time, and the trend is a generalization of the time-trend such that the slope can dynamically vary across time.
#
# For both elements (level and trend), we can consider models in which:
#
# - The element is included vs excluded (if the trend is included, there must also be a level included).
# - The element is deterministic vs stochastic (i.e. whether or not the variance on the error term is confined to be zero or not)
#
# The only additional parameters to be estimated via MLE are the variances of any included stochastic components.
#
# This leads to the following specifications:
#
# | | Level | Trend | Stochastic Level | Stochastic Trend |
# |----------------------------------------------------------------------|-------|-------|------------------|------------------|
# | Constant | ✓ | | | |
# | Local Level <br /> (random walk) | ✓ | | ✓ | |
# | Deterministic trend | ✓ | ✓ | | |
# | Local level with deterministic trend <br /> (random walk with drift) | ✓ | ✓ | ✓ | |
# | Local linear trend | ✓ | ✓ | ✓ | ✓ |
# | Smooth trend <br /> (integrated random walk) | ✓ | ✓ | | ✓ |
#
# ### Seasonal
#
# The seasonal component is written as:
#
# <span>$$
# \gamma_t = - \sum_{j=1}^{s-1} \gamma_{t+1-j} + \omega_t \qquad \omega_t \sim N(0, \sigma_\omega^2)
# $$</span>
#
# The periodicity (number of seasons) is `s`, and the defining character is that (without the error term), the seasonal components sum to zero across one complete cycle. The inclusion of an error term allows the seasonal effects to vary over time.
#
# The variants of this model are:
#
# - The periodicity `s`
# - Whether or not to make the seasonal effects stochastic.
#
# If the seasonal effect is stochastic, then there is one additional parameter to estimate via MLE (the variance of the error term).
#
# ### Cycle
#
# The cyclical component is intended to capture cyclical effects at time frames much longer than captured by the seasonal component. For example, in economics the cyclical term is often intended to capture the business cycle, and is then expected to have a period between "1.5 and 12 years" (see Durbin and Koopman).
#
# The cycle is written as:
#
# <span>$$
# \begin{align}
# c_{t+1} & = c_t \cos \lambda_c + c_t^* \sin \lambda_c + \tilde \omega_t \qquad & \tilde \omega_t \sim N(0, \sigma_{\tilde \omega}^2) \\\\
# c_{t+1}^* & = -c_t \sin \lambda_c + c_t^* \cos \lambda_c + \tilde \omega_t^* & \tilde \omega_t^* \sim N(0, \sigma_{\tilde \omega}^2)
# \end{align}
# $$</span>
#
# The parameter $\lambda_c$ (the frequency of the cycle) is an additional parameter to be estimated by MLE. If the seasonal effect is stochastic, then there is one another parameter to estimate (the variance of the error term - note that both of the error terms here share the same variance, but are assumed to have independent draws).
#
# ### Irregular
#
# The irregular component is assumed to be a white noise error term. Its variance is a parameter to be estimated by MLE; i.e.
#
# $$
# \varepsilon_t \sim N(0, \sigma_\varepsilon^2)
# $$
#
# In some cases, we may want to generalize the irregular component to allow for autoregressive effects:
#
# $$
# \varepsilon_t = \rho(L) \varepsilon_{t-1} + \epsilon_t, \qquad \epsilon_t \sim N(0, \sigma_\epsilon^2)
# $$
#
# In this case, the autoregressive parameters would also be estimated via MLE.
#
# ### Regression effects
#
# We may want to allow for explanatory variables by including additional terms
#
# <span>$$
# \sum_{j=1}^k \beta_j x_{jt}
# $$</span>
#
# or for intervention effects by including
#
# <span>$$
# \begin{align}
# \delta w_t \qquad \text{where} \qquad w_t & = 0, \qquad t < \tau, \\\\
# & = 1, \qquad t \ge \tau
# \end{align}
# $$</span>
#
# These additional parameters could be estimated via MLE or by including them as components of the state space formulation.
#
# ## Data
#
# Following Harvey and Jaeger, we will consider the following time series:
#
# - US real GNP, "output", ([GNPC96](https://research.stlouisfed.org/fred2/series/GNPC96))
# - US GNP implicit price deflator, "prices", ([GNPDEF](https://research.stlouisfed.org/fred2/series/GNPDEF))
# - US monetary base, "money", ([AMBSL](https://research.stlouisfed.org/fred2/series/AMBSL))
#
# The time frame in the original paper varied across series, but was broadly 1954-1989. Below we use data from the period 1948-2008 for all series. Although the unobserved components approach allows isolating a seasonal component within the model, the series considered in the paper, and here, are already seasonally adjusted.
#
# All data series considered here are taken from [Federal Reserve Economic Data (FRED)](https://research.stlouisfed.org/fred2/). Conveniently, the Python library [Pandas](https://pandas.pydata.org/) has the ability to download data from FRED directly.
# + jupyter={"outputs_hidden": false}
# Datasets
from pandas_datareader.data import DataReader
# Get the raw data
start = '1948-01'
end = '2008-01'
us_gnp = DataReader('GNPC96', 'fred', start=start, end=end)
us_gnp_deflator = DataReader('GNPDEF', 'fred', start=start, end=end)
us_monetary_base = DataReader('AMBSL', 'fred', start=start, end=end).resample('QS').mean()
recessions = DataReader('USRECQ', 'fred', start=start, end=end).resample('QS').last().values[:,0]
# Construct the dataframe
dta = pd.concat(map(np.log, (us_gnp, us_gnp_deflator, us_monetary_base)), axis=1)
dta.columns = ['US GNP','US Prices','US monetary base']
dta.index.freq = dta.index.inferred_freq
dates = dta.index._mpl_repr()
# -
# To get a sense of these three variables over the timeframe, we can plot them:
# + jupyter={"outputs_hidden": false}
# Plot the data
ax = dta.plot(figsize=(13,3))
ylim = ax.get_ylim()
ax.xaxis.grid()
ax.fill_between(dates, ylim[0]+1e-5, ylim[1]-1e-5, recessions, facecolor='k', alpha=0.1);
# -
# ## Model
#
# Since the data is already seasonally adjusted and there are no obvious explanatory variables, the generic model considered is:
#
# $$
# y_t = \underbrace{\mu_{t}}_{\text{trend}} + \underbrace{c_{t}}_{\text{cycle}} + \underbrace{\varepsilon_t}_{\text{irregular}}
# $$
#
# The irregular will be assumed to be white noise, and the cycle will be stochastic and damped. The final modeling choice is the specification to use for the trend component. Harvey and Jaeger consider two models:
#
# 1. Local linear trend (the "unrestricted" model)
# 2. Smooth trend (the "restricted" model, since we are forcing $\sigma_\eta = 0$)
#
# Below, we construct `kwargs` dictionaries for each of these model types. Notice that rather that there are two ways to specify the models. One way is to specify components directly, as in the table above. The other way is to use string names which map to various specifications.
# + jupyter={"outputs_hidden": false}
# Model specifications
# Unrestricted model, using string specification
unrestricted_model = {
'level': 'local linear trend', 'cycle': True, 'damped_cycle': True, 'stochastic_cycle': True
}
# Unrestricted model, setting components directly
# This is an equivalent, but less convenient, way to specify a
# local linear trend model with a stochastic damped cycle:
# unrestricted_model = {
# 'irregular': True, 'level': True, 'stochastic_level': True, 'trend': True, 'stochastic_trend': True,
# 'cycle': True, 'damped_cycle': True, 'stochastic_cycle': True
# }
# The restricted model forces a smooth trend
restricted_model = {
'level': 'smooth trend', 'cycle': True, 'damped_cycle': True, 'stochastic_cycle': True
}
# Restricted model, setting components directly
# This is an equivalent, but less convenient, way to specify a
# smooth trend model with a stochastic damped cycle. Notice
# that the difference from the local linear trend model is that
# `stochastic_level=False` here.
# unrestricted_model = {
# 'irregular': True, 'level': True, 'stochastic_level': False, 'trend': True, 'stochastic_trend': True,
# 'cycle': True, 'damped_cycle': True, 'stochastic_cycle': True
# }
# -
# We now fit the following models:
#
# 1. Output, unrestricted model
# 2. Prices, unrestricted model
# 3. Prices, restricted model
# 4. Money, unrestricted model
# 5. Money, restricted model
# + jupyter={"outputs_hidden": false}
# Output
output_mod = sm.tsa.UnobservedComponents(dta['US GNP'], **unrestricted_model)
output_res = output_mod.fit(method='powell', disp=False)
# Prices
prices_mod = sm.tsa.UnobservedComponents(dta['US Prices'], **unrestricted_model)
prices_res = prices_mod.fit(method='powell', disp=False)
prices_restricted_mod = sm.tsa.UnobservedComponents(dta['US Prices'], **restricted_model)
prices_restricted_res = prices_restricted_mod.fit(method='powell', disp=False)
# Money
money_mod = sm.tsa.UnobservedComponents(dta['US monetary base'], **unrestricted_model)
money_res = money_mod.fit(method='powell', disp=False)
money_restricted_mod = sm.tsa.UnobservedComponents(dta['US monetary base'], **restricted_model)
money_restricted_res = money_restricted_mod.fit(method='powell', disp=False)
# -
# Once we have fit these models, there are a variety of ways to display the information. Looking at the model of US GNP, we can summarize the fit of the model using the `summary` method on the fit object.
# + jupyter={"outputs_hidden": false}
print(output_res.summary())
# -
# For unobserved components models, and in particular when exploring stylized facts in line with point (2) from the introduction, it is often more instructive to plot the estimated unobserved components (e.g. the level, trend, and cycle) themselves to see if they provide a meaningful description of the data.
#
# The `plot_components` method of the fit object can be used to show plots and confidence intervals of each of the estimated states, as well as a plot of the observed data versus the one-step-ahead predictions of the model to assess fit.
# + jupyter={"outputs_hidden": false}
fig = output_res.plot_components(legend_loc='lower right', figsize=(15, 9));
# -
# Finally, Harvey and Jaeger summarize the models in another way to highlight the relative importances of the trend and cyclical components; below we replicate their Table I. The values we find are broadly consistent with, but different in the particulars from, the values from their table.
# + jupyter={"outputs_hidden": false}
# Create Table I
table_i = np.zeros((5,6))
start = dta.index[0]
end = dta.index[-1]
time_range = '%d:%d-%d:%d' % (start.year, start.quarter, end.year, end.quarter)
models = [
('US GNP', time_range, 'None'),
('US Prices', time_range, 'None'),
('US Prices', time_range, r'$\sigma_\eta^2 = 0$'),
('US monetary base', time_range, 'None'),
('US monetary base', time_range, r'$\sigma_\eta^2 = 0$'),
]
index = pd.MultiIndex.from_tuples(models, names=['Series', 'Time range', 'Restrictions'])
parameter_symbols = [
r'$\sigma_\zeta^2$', r'$\sigma_\eta^2$', r'$\sigma_\kappa^2$', r'$\rho$',
r'$2 \pi / \lambda_c$', r'$\sigma_\varepsilon^2$',
]
i = 0
for res in (output_res, prices_res, prices_restricted_res, money_res, money_restricted_res):
if res.model.stochastic_level:
(sigma_irregular, sigma_level, sigma_trend,
sigma_cycle, frequency_cycle, damping_cycle) = res.params
else:
(sigma_irregular, sigma_level,
sigma_cycle, frequency_cycle, damping_cycle) = res.params
sigma_trend = np.nan
period_cycle = 2 * np.pi / frequency_cycle
table_i[i, :] = [
sigma_level*1e7, sigma_trend*1e7,
sigma_cycle*1e7, damping_cycle, period_cycle,
sigma_irregular*1e7
]
i += 1
pd.set_option('float_format', lambda x: '%.4g' % np.round(x, 2) if not np.isnan(x) else '-')
table_i = pd.DataFrame(table_i, index=index, columns=parameter_symbols)
table_i
| examples/notebooks/statespace_structural_harvey_jaeger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# # Persistence model
#
# You remember your professor of the Time Series class.
#
# Don't build a crazy model before trying with persistence. A baseline is always valuable: sometimes it provides good enough results, but it always sets the level for more complex approaches.
#
# <div class="alert alert-block alert-warning">
# <b>Simplification.</b>
#
# There is absolutely no good reason to go through the following complex procedure just to build a persistence baseline: the `.shift()` method of `pandas.Series` makes the job. However, we will use the persistence model to demostrate how to create a model with custom code and container.
# </div>
#
# You heard from your fellow data scientist Marta about a cool library for time series forecasting, built on top of sklearn and you want to try it out. So, you start by installing sktime.
# # Setup
# 'ml.m5.xlarge' is included in the AWS Free Tier
INSTANCE_TYPE = 'ml.m5.xlarge'
# ! pip install sktime --user
# ! pip install pandas s3fs --upgrade
# Please, restart the kernel if this is the first time you run this notebook.
#
# This is necessary to ensure that we can actually import the libraries we've just installed in the previous cells.
# +
import os
import pandas as pd
import numpy as np
from sktime.forecasting.base import ForecastingHorizon
import boto3
import sagemaker
from sagemaker.estimator import Estimator
# +
# Configuring the default size for matplotlib plots
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (20, 6)
# +
image_name = prefix = 'persistence-baseline'
boto3_session = boto3.Session()
sagemaker_session = sagemaker.Session()
sagemaker_bucket = sagemaker_session.default_bucket()
region = boto3_session.region_name
# -
# # Image preparation
# You figure the process of building your docker image. To start, you create the ECR (Elastic Container Registry) repository. Then, you build the Docker image with the train and inference code, and finally you push it to such repository.
#
# Fortunately, your favourite ML Engineers, Matteo and Gabriele, have already done it for you, and you can use it directly.
# +
# # %%bash -s "$image_name" "$region"
# chmod 755 build_push.sh
# ./build_push.sh $1 $2
# +
# # ! docker image ls
# -
image_uri = "919788038405.dkr.ecr.eu-west-1.amazonaws.com/persistence-baseline:latest"
# # Raw data gathering
# The data processing pipeline created by Matteo and Gabriele deposits the final dataset in a conventional location on S3.
# In order to retrieve the data to crunch, you first load the S3 object.
#
# With `pandas`, the integration is immediate: S3 URI are resolved as if they were file paths.
#
# You also create some objects that will be useful throughout the notebook.
raw_data_s3_path = "s3://public-workshop/normalized_data/processed/2006_2022_data.parquet"
raw_df = pd.read_parquet(raw_data_s3_path)
resampled_df = raw_df.resample('D').sum()
# +
NOW = '2019-12-31 23:59'
TRAIN_END = '2017-12-31 23:59'
load_df = resampled_df[:NOW].copy()
load_df.head()
# -
# # Upload on S3
# SageMaker train jobs retrieve data from S3: you thus need to upload the train set.
# +
main_prefix = "amld22-workshop-sagemaker"
local_train_path = "persistence_train.parquet"
s3_train_path = f's3://{sagemaker_bucket}/{main_prefix}/data/modelling/{prefix}/train.parquet'
load_df.to_parquet(s3_train_path)
print(f"Data uploaded to: {s3_train_path}")
# -
# # Create estimator & Train
# Then, you can use your custom docker image to train the persistence model - yeah, you smile when you think about "training persistence".
#
# Yet, you have a final look to your `persistence/train.py` and `persistence/serve.py` files and you run the cell.
# +
sk_model = Estimator(
image_uri=image_uri,
role=sagemaker.get_execution_role(),
instance_count=1,
instance_type=INSTANCE_TYPE,
hyperparameters={
"strategy": "last",
"sp": 365,
}
)
sk_model.fit({
"training": s3_train_path,
})
# -
# # Deployment
# Using the facility of AWS SageMaker, you deploy the model to a managed endpoint.
#
# SageMaker then uses the `persistence/serve.py` module to spin up a Flask server and make predictions.
#
# <div class="alert alert-block alert-warning">
# <b>Simplification.</b>
#
# The Flask development server we use is **NOT** a production server, please make sure to set up a more robust and secure serving method when deploying to production. For example, have a look at the inference approach of: https://github.com/aws/amazon-sagemaker-examples/tree/main/advanced_functionality/scikit_bring_your_own/container
# </div>
sk_predictor = sk_model.deploy(
initial_instance_count=1,
instance_type=INSTANCE_TYPE,
serializer=sagemaker.serializers.CSVSerializer(),
deserializer=sagemaker.deserializers.JSONDeserializer()
)
# # Prediction
# You use the deployed API to predict on the test set.
#
# Results are not that bad, but there is definitely room for improvements.
#
# You smile, and open Google Scholar to look for inspiration.
def mean_absolute_percentage_error(y_true, y_pred):
return np.mean(np.abs(y_true - y_pred) / y_pred)
# +
y_true = load_df[TRAIN_END:].Load
prediction_index = y_true.index
fh_absolute = ForecastingHorizon(prediction_index, is_relative=False)
# Predict using the deployed model
y_pred = sk_predictor.predict(fh_absolute)
y_pred_series = pd.Series(y_pred.values(), index=prediction_index)
# Compute MAPE
naive_mape = mean_absolute_percentage_error(y_true, y_pred_series)
# Plot results
plt.title(f"Persistence | MAPE: {100 * naive_mape:.2f} %")
plt.plot(y_true, label='Actual')
plt.plot(y_pred_series, label='Predicted')
plt.legend()
plt.grid(0.4)
plt.show()
# -
# # Cleanup
# If you’re ready to be done with this notebook, please run the cells below with `CLEANUP = True`.
#
# This will remove the model and hosted endpoint to avoid any charges from a stray instance being left on.
CLEANUP = True
if CLEANUP:
sk_predictor.delete_model()
sk_predictor.delete_endpoint()
| notebooks/modeling/persistence/persistence_docker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font size="+5">#03. Arboles de Decision</font>
# <ul>
# <li>Resolver dudas → Pregunta en <img src="https://discord.com/assets/f9bb9c4af2b9c32a2c5ee0014661546d.png" style="height: 1em; vertical-align: middle;"> <a href="https://discord.gg/cmB3KGsqMy">Discord</a></li>
# <li>Tutoriales → <img src="https://openmoji.org/php/download_asset.php?type=emoji&emoji_hexcode=E044&emoji_variant=color" style="height: 1em; vertical-align: middle;"> <a href="https://www.youtube.com/channel/UCovCte2I3loteQE_kRsfQcw">YouTube</a></li>
# <li>Reservar Clases → <span style="color: orange">@</span> <a href="https://sotastica.com/reservar">sotastica</a></li>
# </ul>
# # Cargar Datos
# + [markdown] tags=[]
# Usamos la base de datos del **CIS** sobre una muestra de 2455, cuyas características son de aspecto sociológico. El `objetivo es determinar si una persona usa internet o no`.
#
# ```python
# df = pd.read_csv('https://raw.githubusercontent.com/jesusloplar/data/main/uso_internet_espana.csv')
# df.head()
# ```
# -
import pandas as pd
import seaborn as sns
df_base = pd.read_csv('https://raw.githubusercontent.com/jesusloplar/data/main/uso_internet_espana.csv')
df_base.head()
# + [markdown] tags=[]
# # Transformación de los Datos
# -
# > 1. Las variables categóricas tienen valores de tipo `string` u `object`. Podéis consultarlo con `df.dtypes`.
# > 2. El modelo no puede optimizarse si hay columnas de dichos tipos.
# > 3. Deberíamos convertir las columnas categóricas a 0s y 1s. Es decir, **Variables Dummy**. Para ello, usaremos la función `pd.get_dummies(df)`.
df = pd.get_dummies(df_base, drop_first=True)
# # Seleccionar Variables
# > 1. `Variable Objetivo y`
# > 2. El resto las usaremos como `Variables Explicativas X`
objetivo = df.uso_internet
explicativas = df.drop(columns='uso_internet')
# # Entrenar Modelo `DecisionTreeClassifier()`
# > Antes de irte a la torera a buscar lo que hemos visto durante la sesión. Piensa que:
# >
# > 1. Solo queremos importar una cosa; el objeto `DecisionTreeClassifier`.
# > 2. Por tanto: `from ... import DecisionTreeClassifier()`
# > 3. Tan solo tienes que pensar qué colocar en lugar de `...`. Teniendo en cuenta que la librería de `sklearn` contiene todo lo relacionado con los modelos de **Machine Learning**.
# > 4. **Disciplina Sotástica:** Puedes usar el tabulador `tab` para que Python te vaya sugiriendo...
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier(max_depth=3)
model.fit(X=explicativas, y=objetivo)
# # Visualizar Modelo
# + [markdown] tags=[]
# > 1. Usaremos la función `plot_tree()`, que se encuentra en el módulo `tree` de la librería `sklearn`
# > 2. Podéis usar `shift + tab` para que `Python` os explique cómo usar la función `plot_tree()`
# -
from sklearn.tree import plot_tree
import matplotlib.pyplot as plt
# - lista de strings: s
a = 8
a = 'aoiusef'
a=['mujer', 'hombre']
explicativas
# - `DataFrame` =? `lista de strings`
a
explicativas.columns
df.columns.tolist()
plt.figure(figsize=(18,12))
plot_tree(decision_tree=model, feature_names=explicativas.columns, fontsize=13);
plt.figure(figsize=(18,12))
plot_tree(decision_tree=model, feature_names=explicativas.columns, fontsize=13, filled=True);
df.loc[281]
766/844
df.uso_internet = df.uso_internet.astype(object)
df.dtypes
explicativas_cat = df.loc[:, df.dtypes == 'object'].drop(columns='uso_internet').columns
comb = list(zip(explicativas_cat, range(3)))
comb
# +
### Variables Numéricas
#### Método Gráfico
# A la hora de visualizar la relación entre las diferentes variables numércias explicativas, no podemos hacer un conteo como antes. Sino que debemos ver cómo se distribuyen estas variables en las distintas categorías de la variable objetivo.
# Podemos hacer diferentes gráficas:
explicativas_num = [i for i in df.columns if i not in explicativas_cat]
explicativas_num.remove('uso_internet')
comb = list(zip(explicativas_num, range(len(explicativas_num))))
comb
# +
##### Histograma
fig, axs = plt.subplots(ncols=4, figsize=(20,6))
fig.suptitle('Vertically stacked subplots')
for i,j in comb:
for lab, dat in df.groupby('uso_internet'):
axs[j].hist(x=dat[i], label=lab, alpha=0.7, edgecolor='black', bins=20)
axs[j].set_title(i)
axs[j].legend(title='Survived')
# -
a = df_base.sample()
a
#Example sklearn.tree.plot_tree
from sklearn.datasets import load_iris
from sklearn import tree
clf = tree.DecisionTreeClassifier(random_state=0)
iris = load_iris()
import matplotlib.pyplot as plt
plt.figure(figsize=(12,8))
clf = clf.fit(iris.data, iris.target)
tree.plot_tree(clf)
# # Explicación Algoritmo Árbol de Decisión
# > _Durante la corrección te explicare en detalle el funcionamiento de este algoritmo. Mientras tanto, podéis ver el vídeo que usaré para que os relacionéis con los conceptos:_
# >
# > - https://www.youtube.com/watch?v=7VeUPuFGJHk
# # Interpretar Modelo
# > - ¿Cómo se ha situado la variable en el **Nodo Raíz**? ¿Es la **variable más importante**? ¿Por qué?
df.loc[281]
model.predict_proba(X=explicativas.loc[281].values.reshape(1,-1))
# # Realizar Predicciones
# > ¿No crees que si existe un modelo para calcular la ecuación matemática del árbol, probablemente habrá otro para **aplicar la ecuación matemática** sustituyendo por las variables de entrada, `explicativas X`?
model.predict_proba(explicativas)
import numpy as np
np.where(model.predict_proba(explicativas)[:, 1]> 0.5, 1, 0)
predicciones = model.predict(explicativas)
# # Realidad vs Predicciones
# > ¿Cómo de bueno es nuestro modelo?
# >
# > 1. Si pasamos las predicciones como una nuevo columna del `DataFrame`, podremos observar que **las predicciones de nuestro modelo pueden no coincidir con la realidad**.
# >
# > - `df['pred'] = predicciones`
# >
# > 2. ¿Cómo medimos el **error de nuestro modelo**? ¿Cómo de bueno es nuestro modelo para describir la realidad?
# > - `df.sample(10)` para comprobar si las predicciones de nuestro modelo coinciden con la realidad...
df['pred'] = predicciones
df.sample(10)
df['pred'] == df['uso_internet']
(df['pred'] == df['uso_internet']).sum()
(df['pred'] == df['uso_internet']).sum()/2455
model.score(X=explicativas, y=objetivo)
# # Matriz de Confusión
# > 1. Usar la función `confusion_matrix()`, o `plot_confusion_matrix()`
# > 2. ¿Qué representa cada número?
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(estimator=model, X=explicativas, y_true=objetivo)
sensitividad = 961/(354+961)
sensitividad
especificidad = 1014/(1014 + 126)
especificidad
df.groupby(['pred', 'uso_internet']).size().to_frame(name='freq')
# + [markdown] tags=[]
# # Otrás Métricas
# -
# > Siempre tomaremos como `referencia la realidad (por filas)`. Imaginando que estamos prediciendo si una persona tiene COVID o no, nos preguntaremos:
plot_confusion_matrix(estimator=model, X=explicativas, y_true=objetivo)
# ## Sensitividad
# > - Dentro de las personas que, realmente, `tienen COVID (1)`, cuántas predice nuestro modelo como que tienen COVID (1)?
sensitividad = 961/(354+961)
sensitividad
# ## Especificidad
# > - Dentro de las personas que, realmente, `NO tienen COVID (0)`, cuántas predice nuestro modelo como que NO tienen COVID (0)?
especificidad = 1014/(1014 + 126)
especificidad
# ## Classification Report
# > 1. Usaremos la función `classification_report()`
# > 2. Guardamos el objeto resultante en `reporte`
# > 3. Coinciden algunos de los números de la tabla, `print(reporte)`, con los que hemos calculado anteriormente?
from sklearn.metrics import classification_report
reporte = classification_report(y_true = objetivo, y_pred=df.pred)
print(reporte)
# ## Curva ROC
# > - Usaremos la función `plot_roc_curve()`
# >
# > _PD: Durante la corrección te explicare en detalle el funcionamiento la Curva ROC. Mientras tanto puedes hacer uso del vídeo que usaré:_
# >
# > - https://www.youtube.com/watch?v=7VeUPuFGJHk
# + [markdown] tags=[]
# # Otrás Métricas
# -
# > Siempre tomaremos como `referencia la realidad (por filas)`. Imaginando que estamos prediciendo si una persona tiene COVID o no, nos preguntaremos:
from sklearn.linear_model import LogisticRegression
model_lr = LogisticRegression()
model_lr.fit(X=explicativas, y=objetivo)
plot_confusion_matrix(estimator=model, X=explicativas, y_true=objetivo)
# ## Classification Report
# > 1. Usaremos la función `classification_report()`
# > 2. Guardamos el objeto resultante en `reporte`
# > 3. Coinciden algunos de los números de la tabla, `print(reporte)`, con los que hemos calculado anteriormente?
from sklearn.metrics import classification_report
reporte = classification_report(y_true = objetivo, y_pred=df.pred)
print(reporte)
reporte_lr = classification_report(y_true = objetivo, y_pred=model_lr.predict(explicativas))
print(reporte_lr)
# ## Curva ROC
# > - Usaremos la función `plot_roc_curve()`
# >
# > _PD: Durante la corrección te explicare en detalle el funcionamiento la Curva ROC. Mientras tanto puedes hacer uso del vídeo que usaré:_
# >
# > - https://www.youtube.com/watch?v=7VeUPuFGJHk
from sklearn.metrics import plot_roc_curve
plot_roc_curve(estimator=model, X=explicativas, y=objetivo)
plot_roc_curve(estimator=model_lr, X=explicativas, y=objetivo)
# # Objetivos Alcanzados
# _Haz doble click sobre esta celda y pon una `X` dentro de las casillas [X] si crees que has superado los objetivos:_
#
# - [X] Entender cómo se usan los **Algoritmos de Árboles**.
# - [X] Entender otra forma de **comparar los datos reales con las predicciones** del modelo.
# - [X] No todas las visualizaciones de modelos son iguales. En este caso también podemos **visualizar un árbol** para interpretar el modelo.
# - [X] Distinguir el papel de la **probabilidad** a la hora de optimizar este tipo de modelos.
# - [X] Saber determinar **por qué una variable es importante** en el modelo. Es decir, por qué aporta diferencias significativas.
# - [X] Entender la necesidad de **normalizar** los datos.
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import DecisionTreeRegressor
# # Tipos de Modelos
# - clasificacion: predecir variable categorica
# - supervisado: existe la variable objetivo
# - no supervisado: no existe y la tengo que adivinar (analisis de cluster)
# - regresion: predecir variable numerica
| past program/#03. Arboles de Decision/Notebooks/.ipynb_checkpoints/tarea03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Plot NWCSAF CT data as Matrix
# This is a copy of the notebook [02-NWCSAF-CT-plot.ipynb](02-NWCSAF-CT-plot.ipynb). The filelist was updated to plot CT as function of resolution.
# ## Libs
# +
# %matplotlib inline
import os, sys, glob, copy
import numpy as np
import pylab as pl
import datetime
import scipy.ndimage
import pandas as pd
import xarray as xr
import seaborn as sns
import nawdex_analysis.io.input_lev2 as input_lev2
import nawdex_analysis.io.selector
import nawdex_analysis.plot.nawdex_map
reload( nawdex_analysis.plot.nawdex_map )
# -
pl.rcParams['figure.figsize'] = (16.0, 12.0)
pl.rcParams['font.size'] = 16.0
# ## A subsampling funciton for speed
def subsamp2d( d, nsub = 4 ):
dsub = {}
for k in d.keys():
v = d[k]
try:
if np.ndim(v) == 2:
vsub = v[::nsub, ::nsub]
except:
vsub = v
print '%s not an 2d array?' % k
dsub[k] = vsub
return dsub
# ## Plot of NWCSAF CT
# ### Input Data
time = datetime.datetime(2016,9,23, 12, 0)
flist = nawdex_analysis.io.selector.make_filetime_index('CT', time, subdirs=['meteosat', 'synsat'])
flist[time]
# +
dstack = {}
for fname in flist[time]:
if 'mis-0001' in fname or 'msevi' in fname:
basename = os.path.basename( fname )
dstack[basename] = input_lev2.read_data_field( fname, time, 'CT')
# -
sorted_fnames = np.array(sorted( dstack ))
sorted_fnames
# ### Sort Names differently
sorted_fnames = np.array(
['nwcsaf_msevi-nawdex-20160923.nc',
'nwcsaf_synsat-nawdexnwp-2km-mis-0001.nc',
'nwcsaf_synsat-nawdexnwp-5km-mis-0001.nc',
'nwcsaf_synsat-nawdexnwp-10km-mis-0001.nc',
'nwcsaf_synsat-nawdexnwp-20km-mis-0001.nc',
'nwcsaf_synsat-nawdexnwp-40km-mis-0001.nc',
'nwcsaf_synsat-nawdexnwp-80km-mis-0001.nc'], )
# ### Region Masking
m = input_lev2.read_mask(region='atlantic')['mask']
for fname in sorted_fnames:
dstack[fname]['CT'] = np.ma.masked_where( ~m, dstack[fname]['CT'])
# ### Subsampling
if False:
for fname in sorted_fnames:
dstack[fname] = subsamp2d( dstack[fname], nsub = 8 )
# ## Solve Issues with NaN Georef Values
# `pcolormesh` does not allow for NaNs in the georef. NaNs will be replaced by an arbitrary value.
def denan_georef( d ):
d['lon'][np.isnan( d['lon'] )] = 1e23
d['lat'][np.isnan( d['lat'] )] = 1e23
for fname in sorted_fnames:
denan_georef( dstack[fname] )
# ## Colormapping
# We like to plot only a subset of cloud type which are selected by index.
def ct_colormapping( cindex ):
# default color list provided by NWCSAF
colorlist = ['#007800', '#000000','#fabefa','#dca0dc',
'#ff6400', '#ff6400', '#ffb400', '#ffb400',
'#f0f000', '#f0f000','#d7d796','#d7d796',
'#e6e6e6', '#e6e6e6', '#0050d7', '#00b4e6',
'#00f0f0', '#5ac8a0', '#c800c8']
if cindex is None:
return colorlist
else:
color_selected = np.array( colorlist )[np.array(cindex) - 1]
return color_selected
# ### CT field mapping
# +
def ct_mapping( ct, cindex, ntypes = 20 ):
if cindex is None:
return ct
# init mapping index
mapping_index = np.zeros( ntypes ).astype( np.int )
# overwrite mapping index with cloud types
n = 1
for cind in cindex:
mapping_index[cind] = n
n += 1
# do the mapping
ct_mapped = mapping_index[ ct.data.astype(np.int) ]
ct_mapped = np.ma.masked_where( ct.mask, ct_mapped)
return ct_mapped
# -
# ### NAWDEX Plot Routine
# +
## from nawdex_analysis.plot.nawdex_map.nawdex_nwcsaf_plot??
from nawdex_analysis.plot.nawdex_map import nwcsaf_product_colorbar, nawdex_map
def nawdex_nwcsaf_plot(dset, vname = 'CMa',region = 'zenith75', cindex = None, plot_colorbar = True):
mp = nawdex_map( region = region, color = 'gold' )
# map geo-ref
x, y = mp(dset['lon'], dset['lat'])
m = dset['mask']
vm = np.ma.masked_where( ~m, dset[vname] )
cmap = pl.cm.get_cmap('bone', 4)
if vname == 'CMa':
pcm = mp.pcolormesh(x, y, vm, cmap = cmap, vmin = 1, vmax = 5)
if vname == 'CT':
colorlist = ct_colormapping( cindex )
print colorlist, len(colorlist)
cmap = pl.matplotlib.colors.ListedColormap( colorlist )
vmapped = ct_mapping( vm, cindex )
pcm = mp.pcolormesh(x,y, vmapped, cmap = cmap, vmin = 1, vmax = len( colorlist ) + 1)
if plot_colorbar:
nwcsaf_product_colorbar( pcm, vname = vname )
return mp
# -
# ## Plotting
# +
fig, axs = pl.subplots( nrows = 4, ncols = 2, figsize = (11, 14), sharex = True, sharey = True )
axs = axs.flatten()
labs = ['OBS',
'ICON( 2.5km, *, CP )', 'ICON( 5km, *, CP )',
'ICON( 10km, *, CP )', 'ICON( 20km, *, CP )',
'ICON( 40km, *, CP )', 'ICON( 80km, *, CP )']
for i, basename in enumerate( sorted_fnames ):
if i > 0:
j = i + 1
else:
j = i
a = axs[j]
pl.sca( a )
d = dstack[basename]
# add fracitonal to very low
d['CT'][d['CT'] == 19] = 6
mp = nawdex_nwcsaf_plot(d, vname = 'CT', cindex = [2, 6, 8, 10, 12,14, 15, 16, 17,],
region = 'atlantic',
plot_colorbar = False )
a.text(-63, 60, labs[i], va = 'bottom', ha = 'left', fontweight = 'bold' ,
bbox = dict(facecolor='white', edgecolor = 'white', alpha=0.5))
if i == 1:
a = axs[i]
a.axis('off')
#a.clear()
apos = a.get_position()
cax = fig.add_axes( [apos.x0, apos.y0, 0.02, apos.height] )
# pl.sca( a )
# pl.cla( )
pcm = axs[0].collections[-1]
cbar = pl.colorbar( pcm, cax, ticks= np.arange(9) + 1.5,)
cbar.ax.set_yticklabels(['sea',
'very low / fractional', 'low', 'mid-level', 'high opaque', 'very high opaque',
'semi. thin', 'semi. moderately thick', 'semi. thick',
], fontsize = 11)
# if i >= 1:
# break
xlim, ylim = ((-65, 10.802031483325052), (27.0, 66.5))
a.set_xlim( xlim )
a.set_ylim( ylim )
pl.subplots_adjust( hspace = 0.05, wspace = 0.05, bottom = 0.2)
pl.savefig('../pics/CT-overview-resolution.png')
| nbooks/17-NWCSAF-CT-plot-for-different-resolutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Torch)
# language: python
# name: torch
# ---
import sys
sys.path.insert(0, "../..")
# +
import torch
from torch import nn
from torch.utils import data
from torchvision import datasets, transforms
from tqdm.notebook import tqdm
import numpy as np
from copy import deepcopy
from nn_extrapolation import AcceleratedSGD
# -
torch.cuda.is_available()
# +
val_loss_fn = nn.NLLLoss(reduction="sum")
def validation(model, loader):
ok = 0
loss_sum = 0
total = 0
model.eval()
with torch.no_grad():
for x, y in loader:
x = x.cuda()
y = y.cuda()
out = model(x)
loss_sum += val_loss_fn(out, y)
preds = out.argmax(1)
ok += (y == preds).sum()
total += len(y)
return ok / total, loss_sum / total
def train_epoch(loss_log):
model.train()
for x, y in train_loader:
x = x.cuda()
y = y.cuda()
optimizer.zero_grad()
out = model(x)
loss = loss_fn(out, y)
loss_log += list(loss.flatten().cpu().detach().numpy())
loss.backward()
optimizer.step()
# +
train_ds = datasets.MNIST("../../../MNIST", download=True, train=True, transform=transforms.ToTensor())
test_ds = datasets.MNIST("../../../MNIST", download=True, train=False, transform=transforms.ToTensor())
valid_size = int(0.2 * len(train_ds))
train_ds, valid_ds = data.random_split(train_ds, [len(train_ds) - valid_size, valid_size])
train_loader = data.DataLoader(train_ds, batch_size=64, shuffle=True, num_workers=2)
valid_loader = data.DataLoader(valid_ds, batch_size=64, shuffle=True, num_workers=2)
test_loader = data.DataLoader(test_ds, batch_size=64, shuffle=False, num_workers=2)
# -
# ## Levin t
# + tags=[]
model = nn.Sequential(
nn.Flatten(),
nn.Linear(28*28, 10),
nn.LogSoftmax(-1),
)
model.cuda()
# -
optimizer = AcceleratedSGD(model.parameters(), 1e-3, k=10, mode="epoch", method="Levin:t")
loss_fn = nn.NLLLoss()
log_file = open("SGD-Levin:t.txt", "w")
# + tags=[]
epochs = 30
for epoch in range(epochs):
print("Epoch", epoch+1)
loss_log = []
train_epoch(loss_log)
print(f"Training loss: {np.mean(loss_log):.4f}")
optimizer.finish_epoch()
val_acc, val_loss = validation(model, valid_loader)
print(f"Validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}")
print("Epoch", epoch+1,
f"Training loss: {np.mean(loss_log):.4f}, validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}",
file=log_file, flush=True
)
# -
train_score = validation(model, train_loader)
valid_score = validation(model, valid_loader)
print("Train:", train_score)
print("Valid:", valid_score)
print("Train:", train_score, flush=True, file=log_file)
print("Valid:", valid_score, flush=True, file=log_file)
optimizer.accelerate()
optimizer.store_parameters()
model.cuda()
train_score = validation(model, train_loader)
valid_score = validation(model, valid_loader)
print("Train:", train_score)
print("Valid:", valid_score)
print("Train:", train_score, flush=True, file=log_file)
print("Valid:", valid_score, flush=True, file=log_file)
# ## Levin u
# + tags=[]
model = nn.Sequential(
nn.Flatten(),
nn.Linear(28*28, 10),
nn.LogSoftmax(-1),
)
model.cuda()
# -
optimizer = AcceleratedSGD(model.parameters(), 1e-3, k=10, mode="epoch", method="Levin:u")
loss_fn = nn.NLLLoss()
log_file = open("SGD-Levin:u.txt", "w")
# + tags=[]
epochs = 30
for epoch in range(epochs):
print("Epoch", epoch+1)
loss_log = []
train_epoch(loss_log)
print(f"Training loss: {np.mean(loss_log):.4f}")
optimizer.finish_epoch()
val_acc, val_loss = validation(model, valid_loader)
print(f"Validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}")
print("Epoch", epoch+1,
f"Training loss: {np.mean(loss_log):.4f}, validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}",
file=log_file, flush=True
)
# -
train_score = validation(model, train_loader)
valid_score = validation(model, valid_loader)
print("Train:", train_score)
print("Valid:", valid_score)
print("Train:", train_score, flush=True, file=log_file)
print("Valid:", valid_score, flush=True, file=log_file)
optimizer.accelerate()
# + tags=[]
optimizer.store_parameters()
model.cuda()
# -
train_score = validation(model, train_loader)
valid_score = validation(model, valid_loader)
print("Train:", train_score)
print("Valid:", valid_score)
print("Train:", train_score, flush=True, file=log_file)
print("Valid:", valid_score, flush=True, file=log_file)
# ## Levin v
# + tags=[]
model = nn.Sequential(
nn.Flatten(),
nn.Linear(28*28, 10),
nn.LogSoftmax(-1),
)
model.cuda()
# -
optimizer = AcceleratedSGD(model.parameters(), 1e-3, k=10, mode="epoch", method="Levin:v")
loss_fn = nn.NLLLoss()
log_file = open("SGD-Levin:v.txt", "w")
# + tags=[]
epochs = 30
for epoch in range(epochs):
print("Epoch", epoch+1)
loss_log = []
train_epoch(loss_log)
print(f"Training loss: {np.mean(loss_log):.4f}")
optimizer.finish_epoch()
val_acc, val_loss = validation(model, valid_loader)
print(f"Validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}")
print("Epoch", epoch+1,
f"Training loss: {np.mean(loss_log):.4f}, validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}",
file=log_file, flush=True
)
# -
train_score = validation(model, train_loader)
valid_score = validation(model, valid_loader)
print("Train:", train_score)
print("Valid:", valid_score)
print("Train:", train_score, flush=True, file=log_file)
print("Valid:", valid_score, flush=True, file=log_file)
optimizer.accelerate()
optimizer.store_parameters()
model.cuda()
train_score = validation(model, train_loader)
valid_score = validation(model, valid_loader)
print("Train:", train_score)
print("Valid:", valid_score)
print("Train:", train_score, flush=True, file=log_file)
print("Valid:", valid_score, flush=True, file=log_file)
| notebooks/logistic regression - SGD/SGD-Levin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
dataset = pd.read_csv('Position_Salaries.csv')
dataset.head()
#we wont use Position column here as level column is sufficient
x = dataset.iloc[:, 1:2].values #always take x as matrix
y = dataset.iloc[:, 2].values
# #### we will consider whole dataset as input as data is too small to split and also we need precision here.
#fitting linear regression
from sklearn.linear_model import LinearRegression
lreg = LinearRegression()
lreg.fit(x,y)
#fitting plynomial regression.
#we tried for degree 2,3,4 and found 4 to be best of them
from sklearn.preprocessing import PolynomialFeatures
polyreg= PolynomialFeatures(degree = 4)
x_poly = polyreg.fit_transform(x)
lreg2 = LinearRegression()
lreg2.fit(x_poly,y)
#Visualizing linear regression
plt.scatter(x,y, color='red')
plt.plot(x, lreg.predict(x), color='blue')
plt.title("Linear Regression fit")
plt.xlabel("Position Level")
plt.ylabel("Salary")
plt.show()
#Visualizing Polynomial regression (with smoother curve)
x_grid = np.arange(min(x),max(x),0.1)
x_grid = x_grid.reshape((len(x_grid),1))
plt.scatter(x,y, color='red')
plt.plot(x_grid, lreg2.predict(polyreg.fit_transform(x_grid)), color='blue')
plt.title("Polynomial Regression fit")
plt.xlabel("Position Level")
plt.ylabel("Salary")
plt.show()
# ### Predicting Salary for position level 6.5
#Linear Regression
lreg.predict(6.5)
#Polynomial Regression
lreg2.predict(polyreg.fit_transform(6.5))
# #### We get a huge difference!
| Regression/Polynomial Linear Reg/P4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introdução
#
# No *site* da CFB
# https://www.infraestruturameioambiente.sp.gov.br/cfb/contatos/
# +
import os
import re
import time
import random
import requests
import numpy as np
import pandas as pd
import geopandas as gpd
from bs4 import BeautifulSoup
from osgeo import gdal, osr
from tqdm.notebook import trange, tqdm
# -
# # Dados Tabulares
# ## Scrapy *Site*
url = 'https://www.infraestruturameioambiente.sp.gov.br/cfb/contatos/'
r = requests.get(url)
soup = BeautifulSoup(r.content, 'html.parser')
tag = soup.find('div', {'id': 'content'}).text
# +
list_main = []
for i in str(tag).split('\n'):
if i == '':
pass
else:
list_main.append(i)
# Create Table
df = pd.DataFrame({'data': list_main})
print(df[0:100])
# -
# ## Ajusta de Tabela
# +
# Define os marcos iniciais
df['interval'] = df.apply(lambda x: x.name if 'CTR' in x['data'] else np.nan, axis=1)
# Remove Espaços em Branco
df['data'] = df.apply(lambda x: np.nan if x['data'] == '\xa0' else x, axis=1)
# Preenche Coluna
df['interval'] = df['interval'].ffill()
# Elimina Linhas com NaN
df = df.dropna()
# Result
df.head(10)
# -
list_temps = []
list_for = list(set(df['interval']))
print(list_for)
for i in list_for:
df_temp = df[df['interval'] == i]
list_temp = list(df_temp['data'])
list_temps.append(list_temp)
for i in list_temps:
if len(i)==7:
i.insert(5, None)
# +
df = pd.DataFrame(
list_temps,
columns = [
'id_ctr',
'end',
'bairro_cidade',
'cep',
'telefone_1',
'telefone_2',
'email',
'diretor',
])
df.head()
# -
# Remove tudo que vem antes do :
df = df.applymap(lambda x: x.split(':')[-1], na_action='ignore')
df
for i in df.columns:
print(set(df[i]))
# ## Funções
def rename_nome(x):
x = x.title()
x = x.strip()
dict_rename = {
# Encoding
' ' : ' ',
# Basics
' Com ' : ' com ',
' Sobre ': ' sobre ',
' Da ' : ' da ',
' De ' : ' de ',
' Do ' : ' do ',
' Das ' : ' das ',
' Dos ' : ' dos ',
' A ' : ' a ',
' As ' : ' as ',
' Ao ' : ' ao ',
' Aos ' : ' aos ',
' E ' : ' e ',
' O ' : ' o ',
' Os ' : ' os ',
# Erros
'1ºten' : '1º Ten',
# Abreviações
'Subten ' : 'Subtenente ',
'Sub Ten' : 'Subtenente ',
'Cap ' : 'Capitão ',
'Ten ' : 'Tenente ',
'Maj ' : 'Major ',
'Cel ' : 'Coronel ',
'Sgt ' : 'Sargento ',
' Pm ' : ' PM ',
'–': '-',
'Registro/Sp': 'Registro - Sp',
'São Bernardo do Campo- Sp': 'São Bernardo do Campo - Sp',
' - Sp': '',
'R.': 'Rua',
'Av.': 'Avenida',
'<NAME> - Quadra 38 - Nº 138': '<NAME>, 138 - Quadra 38',
'133, Sala 23': '133 - Sala 23',
'Joãothiago': '<NAME>',
# Empty
'None' : '',
'none' : '',
}
for k, v in dict_rename.items():
x = x.replace(k, v)
x = x.replace(' ', ' ')
return x.strip()
# ## Renomeando Campos em Colunas
# +
df['bairro_cidade'] = df['bairro_cidade'].astype(str).apply(lambda x: rename_nome(x))
df[['bairro','municipio_sede']] = df['bairro_cidade'].str.split('-', expand=True)
df['bairro'] = df['bairro'].astype(str).apply(lambda x: rename_nome(x))
df['municipio_sede'] = df['municipio_sede'].astype(str).apply(lambda x: rename_nome(x))
# -
df['telefone_1'] = df['telefone_1'].astype(str).apply(lambda x: rename_nome(x))
df['telefone_2'] = df['telefone_2'].astype(str).apply(lambda x: rename_nome(x))
df['diretor'] = df['diretor'].astype(str).apply(lambda x: rename_nome(x))
df['cep'] = df['cep'].astype(str).apply(lambda x: rename_nome(x))
df['cep'] = df['cep'].astype(str).apply(lambda x: x.strip().replace('Cep', ''))
df['cep'] = df['cep'].astype(str).apply(lambda x: rename_nome(x))
df['id_ctr'] = df['id_ctr'].astype(str).apply(lambda x: rename_nome(x))
df['id_ctr'] = df['id_ctr'].astype(str).apply(lambda x: x.rsplit('-', 1)[0])
df['id_ctr'] = df['id_ctr'].astype(str).apply(lambda x: x.upper())
df['id_ctr'] = df['id_ctr'].astype(str).apply(lambda x: x.strip().replace(' ', ''))
# +
df['end'] = df['end'].astype(str).apply(lambda x: rename_nome(x))
df['endereco'] = df['end'].astype(str).apply(lambda x: x.split(',', 1)[0])
df['numero'] = df['end'].astype(str).apply(lambda x: x.split(',', 1)[1])
df[['numero','complemento']] = df['numero'].str.split('-', n=1, expand=True)
df['numero'] = df['numero'].astype(str).apply(lambda x: rename_nome(x))
df['complemento'] = df['complemento'].astype(str).apply(lambda x: rename_nome(x))
df
# -
# ## <NAME>
# +
df = df[[
'id_ctr',
'endereco',
'numero',
'complemento',
'cep',
'bairro',
'municipio_sede',
'telefone_1',
'telefone_2',
'email',
'diretor'
]]
df
# -
# ## Salva
# Results
df.to_csv(
os.path.join('data', 'tabs', 'tab_ctr.csv'),
index=False,
)
df
| cfb_get_infos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TimeSynth
# #### Get latest version
# %%capture
# !pip uninstall timesynth --yes
# !pip install git+https://github.com/TimeSynth/TimeSynth.git
# #### Imports
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import timesynth as ts
# #### Harmonic Signals
# Initializing TimeSampler
time_sampler = ts.TimeSampler(stop_time=20)
# Sampling irregular time samples
irregular_time_samples = time_sampler.sample_irregular_time(num_points=500, keep_percentage=50)
# Initializing Sinusoidal signal
sinusoid = ts.signals.Sinusoidal(frequency=0.25)
# Initializing Gaussian noise
white_noise = ts.noise.GaussianNoise(std=0.3)
# Initializing TimeSeries class with the signal and noise objects
timeseries = ts.TimeSeries(sinusoid, noise_generator=white_noise)
# Sampling using the irregular time samples
samples, signals, errors = timeseries.sample(irregular_time_samples)
# Plotting the series
plt.plot(irregular_time_samples, samples, marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Magnitude')
plt.title('Irregularly sampled sinusoid with noise');
# #### Harmonic Signals with Red noise
# Initializing Gaussian noise
red_noise = ts.noise.RedNoise(std=0.5, tau=0.8)
# Initializing TimeSeries class with the signal and noise objects
timeseries_corr = ts.TimeSeries(sinusoid, noise_generator=red_noise)
# Sampling using the irregular time samples
samples_corr, signals_corr, errors_corr = timeseries_corr.sample(irregular_time_samples)
# Plotting the series
plt.plot(irregular_time_samples, samples_corr, marker='o')
plt.xlabel('Time')
plt.ylabel('Magnitude')
plt.title('Irregularly sampled sinusoid with red noise')
plt.plot(irregular_time_samples, errors_corr, marker='o')
plt.xlabel('Time')
plt.ylabel('Magnitude')
plt.title('Red noise');
# #### PseudoPeriodic Signals
# Initializing TimeSampler
time_sampler_pp = ts.TimeSampler(stop_time=20)
# Sampling irregular time samples
irregular_time_samples_pp = time_sampler_pp.sample_irregular_time(resolution=0.05, keep_percentage=50)
# Initializing Pseudoperiodic signal
pseudo_periodic = ts.signals.PseudoPeriodic(frequency=2, freqSD=0.01, ampSD=0.5)
# Initializing TimeSeries class with the pseudoperiodic signal
timeseries_pp = ts.TimeSeries(pseudo_periodic)
# Sampling using the irregular time samples
samples_pp, signals_pp, errors_pp = timeseries_pp.sample(irregular_time_samples_pp)
# Plotting the series
plt.plot(irregular_time_samples_pp, samples_pp, marker='o')
plt.xlabel('Time')
plt.ylabel('Magnitude')
plt.title('Pseudoperiodic signal');
# #### Gaussian Process signals
gp = ts.signals.GaussianProcess(kernel='Matern', nu=3./2)
gp_series = ts.TimeSeries(signal_generator=gp)
samples = gp_series.sample(irregular_time_samples)[0]
plt.plot(irregular_time_samples, samples, marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Value')
plt.title('Gaussian Process signal with Matern 3/2-kernel');
# #### CAR model
car = ts.signals.CAR(ar_param=0.9, sigma=0.01)
car_series = ts.TimeSeries(signal_generator=car)
samples = car_series.sample(irregular_time_samples)
plt.plot(irregular_time_samples, samples[0], marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Value')
plt.title('Example Continuous Autoregressive process');
# #### AR model for regularly sampled timestamps
# Initializing TimeSampler
time_sampler = ts.TimeSampler(stop_time=20)
# Sampling regular time samples
regular_time_samples = time_sampler.sample_regular_time(num_points=500)
# Initializing AR(2) model
ar_p = ts.signals.AutoRegressive(ar_param=[1.5, -0.75])
ar_p_series = ts.TimeSeries(signal_generator=ar_p)
samples = ar_p_series.sample(regular_time_samples)
plt.plot(regular_time_samples, samples[0], marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Value')
plt.title('Autoregressive process - Second order');
# #### Mackey-Glass signals
# Initializing TimeSampler
time_sampler = ts.TimeSampler(stop_time=1500)
# Sampling irregular time samples
irregular_time_samples = time_sampler.sample_irregular_time(num_points=1500, keep_percentage=75)
mg = ts.signals.MackeyGlass()
noise = ts.noise.GaussianNoise(std=0.1)
mg_series = ts.TimeSeries(signal_generator=mg, noise_generator=noise)
mg_samples, mg_signals, mg_errors = mg_series.sample(irregular_time_samples)
plt.plot(irregular_time_samples, mg_signals, marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Value')
plt.title('Mackey-Glass differential equation with $\\tau=17$');
plt.plot(irregular_time_samples, mg_samples, marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Value')
plt.title('Mackey-Glass ($\\tau=17$) with noise ($\\sigma = 0.1$)');
# #### NARMA series
# Note: only regular sampled timestamps supported
# Initializing TimeSampler
time_sampler = ts.TimeSampler(stop_time=500)
# Sampling irregular time samples
times = time_sampler.sample_regular_time(resolution=1.)
# Take Samples
narma_signal = ts.signals.NARMA(order=10)
series = ts.TimeSeries(narma_signal)
samples, _, _ = series.sample(times)
# Plotting the series
plt.plot(times, samples, marker='o', markersize=4)
plt.xlabel('Time')
plt.ylabel('Magnitude')
plt.title('10th-order NARMA Series');
| TimeSynthExamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TelFit head-to-head (*paused*)
# ### Multi-layer atmosphere with $H_2 O$, $CO_2$, and $O_2$
# ### Using HITRAN, `jax`, and NVIDIA GPUs
#
# A stalled attempt to match the atmosphere profile exactly so we can scrutinize differences.
# Do any atmospheric scientists want to weigh in?
#
# May 2020
# +
import astropy.units as u
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import hapi
hapi.db_begin('../../hapi/data/')
from jax.config import config
config.update("jax_enable_x64", False)
import jax.numpy as np
from jax import vmap, jit, grad, random
from jax.lax import stop_gradient
from jax import jacfwd
from functools import partial
import pandas as pd
# -
# ! du -hs ../../hapi/data/*.data
@jit
def gamma_of_p_and_T(p, T, p_self, n_air, gamma_air_ref, gamma_self_ref):
'''Compute the Lorentz HWHM with pressure and temp'''
return (296.0/T)**n_air *(gamma_air_ref*(p-p_self) + gamma_self_ref*(p_self))
@jit
def lorentz_profile(nu, p, nu_ij, gamma, dp_ref, S_ij):
'''Return the Lorentz line profile given vectors and parameters'''
return S_ij/np.pi * gamma / ( gamma**2 + (nu - (nu_ij + dp_ref*p))**2)
@jit
def tips_Q_of_T(T, g_k, E_k):
'''Total Internal Partition Sum'''
c_2 = 1.4387770 #cm K
return np.sum( g_k * np.exp(- c_2 * E_k / T) )
@jit
def S_ij_of_T(T, S_ij_296, nu_ij, g_lower, E_lower):
'''The Spectral Line Intensity'''
c_2 = 1.4387770 #cm K
return (S_ij_296 *
stop_gradient(tips_Q_of_T(296.0, g_lower, E_lower)) /
tips_Q_of_T(T, g_lower, E_lower) *
np.exp(-c_2 * E_lower/T) /np.exp(-c_2 * E_lower/296.0) *
(1-np.exp(-c_2*nu_ij/T) )/(1-np.exp(-c_2*nu_ij/296.0)))
@jit
def transmission_of_T_p(T, p, nus, vol_mix_ratio, atomic_data):
'''Return the absorption coefficient as a function of T'''
(n_air, gamma_air, gamma_self, E_lower, g_lower,\
nu_lines, delta_air, S_ij_296) = atomic_data
gammas = vmap(gamma_of_p_and_T, in_axes=(None, None, None, 0, 0, 0)\
)(p, T,vol_mix_ratio, n_air, gamma_air, gamma_self)
S_ij = S_ij_of_T(T, S_ij_296, nu_lines, g_lower, E_lower)
abs_coeff = vmap(lorentz_profile, in_axes=(None, None, 0, 0,0, 0))(nus, p,
nu_lines, gammas, delta_air, S_ij).sum(axis=0)
#path_length_km = 1.0
tau = abs_coeff * (vol_mix_ratio*2.688392857142857e+19) * (1.0*100000.0)
return np.exp(-tau)
# ### Compute the transmission spectrum
#
# Requires a path length. Let's assume constant density of the Earth's atmosphere, and 3 km path length.
#
# The HITRAN absorption coefficient $\alpha$ units are: $cm^{−1}/(molecule⋅cm^{−2})$
#
# So we have the optical depth:
#
# $$ \tau_{ij}(\nu, T, p) = \alpha \cdot n \cdot \ell$$
#
# where $n$ is the number density of particles, and $\ell$ is the path length.
def get_hapi_molec_data(species):
'''return the order columns in device arrays
TODO: a bit fragile, consider returning a dict instead!
'''
ordered_cols = ['n_air','gamma_air','gamma_self','elower','gpp','nu','delta_air','sw']
return [np.array(hapi.getColumn(species, col)) for col in ordered_cols]
# +
#(n_air, gamma_air, gamma_self, E_lower, g_lower, nu_lines, delta_air, S_ij_296) = get_hapi_molec_data('H2O')
# -
# ## Get the atmospheric profile from TDAS
# Follow the prescription from <NAME>:
# https://telfit.readthedocs.io/en/latest/GDAS_atmosphere.html
#
# For now we'll use the average composition profiles from [the same source as TelFit](http://eodg.atm.ox.ac.uk/RFM/atm/ngt.atm)
# +
# #! wget http://eodg.atm.ox.ac.uk/RFM/atm/ngt.atm
# #! mv ngt.atm ../data/
# -
pt_prof = pd.read_csv('../data/telfit_HRRR_PT_profile_20200514UTC.txt', skiprows=28, nrows=36, delim_whitespace=True,
names=['PRESS','HGT(MSL)','TEMP','DEW PT','WND DIR','WND SPD'])
pt_prof['HGT(MSL)'] = pt_prof['HGT(MSL)'].str[:-2].astype(float)
# HET is at about 2,026 meters, so truncate to the closest height.
# Hmm the formula copied from the TelFit website for converting to RH from dewpoint seems wrong.
# Dew point does not enter into the equation at all...
Pw = 6.116441 * 10**(7.591386*pt_prof.TEMP/(pt_prof.TEMP + 240.7263))
pt_prof['H2O [ppmv]'] = Pw / (pt_prof.PRESS-Pw) * 1e6
pt_prof = pt_prof[5:].reset_index(drop=True)
pt_prof.head()
# +
cols = ['HGT [km]','PRE [mb]','TEM [K]','N2 [ppmv]','O2 [ppmv]','CO2 [ppmv]','O3 [ppmv]',
'H2O [ppmv]','CH4 [ppmv]','N2O [ppmv]','HNO3 [ppmv]','CO [ppmv]','NO2 [ppmv]',
'N2O5 [ppmv]','ClO [ppmv]','HOCl [ppmv]','ClONO2 [ppmv]','NO [ppmv]','HNO4 [ppmv]',
'HCN [ppmv]','NH3 [ppmv]','F11 [ppmv]','F12 [ppmv]','F14 [ppmv]','F22 [ppmv]',
'CCl4 [ppmv]','COF2 [ppmv]','H2O2 [ppmv]','C2H2 [ppmv]','C2H6 [ppmv]','OCS [ppmv]',
'SO2 [ppmv]','SF6 [ppmv]']
ngt_atm = pd.read_csv('../data/ngt.atm', comment='*', delim_whitespace=True, skiprows=25, names='ABCDE')
ngt_atm = pd.DataFrame(data=ngt_atm.stack().values.reshape(33, 121)).T
ngt_atm.columns = cols
# +
plt.plot(ngt_atm['TEM [K]'], ngt_atm['PRE [mb]'], label='Average Night Mid Latitudes')
plt.plot(pt_prof['TEMP']+273.15, pt_prof['PRESS'], label='20200514 McDonald HRRR')
plt.axhline(ngt_atm.loc[17, 'PRE [mb]'], linestyle='dashed', color='#888888', label='Tropopause')
plt.axhline(794, linestyle='dotted', color='#333333', label='McDonald Observatory')
plt.ylim(1100, 1e-5); plt.yscale('log'); plt.ylabel('$P$ (mb)'); plt.xlabel('$T$ (K)');
plt.legend();
# +
for species in ['N2', 'O2', 'CO2', 'H2O']:
plt.plot(ngt_atm[species+' [ppmv]'], ngt_atm['PRE [mb]'], label=species)
#plt.plot(pt_prof['H2O [ppmv]'], pt_prof['PRESS'], label='20200514 Miscalculated! see note')
plt.ylim(1100, 1e-5); plt.yscale('log'); plt.ylabel('$P$ (mb)'); plt.xlabel('ppmv');
plt.xlim(1e-3, 1e8); plt.xscale('log'); plt.legend();
# -
# # Fail
# We encountered problems here:
#
# #### 1) Get H2O ppmv from dew point
# We did not convert to ppmv correctly from dew point because the TelFit website equation seems incorrect. The solution is to visit the obscure PDF website that <NAME> listed and track down the equations.
#
# #### 2) GDAS unavailable, HRRR Atmosphere profile terminates near the tropopause
# Is it safe to assume that the stratosphere does not matter? Or does not change much?
#
# #### 3) We are only using Lorentzians
# We could/should include Gaussians for the high temperature/low density regime. We should technically convolve these profiles on a line-by-line basis.
# # Success
#
# #### 1) We *did* get TelFit to work in this notebook
# It will be easy to compare head-to-head once we figure out the issues above! (see below)
#
# #### 2) We have the machinery to match to HPF resolution/sampling
# We read in an example file so that we can match the wavelength sampling, bandwidth, and spectral resolution.
# # General thoughts
#
# #### 1) How to properly specify and perturb/infer atmosphere profile
# I think TelFit presently tweaks the entire atmosphere profile by a single scalar value, which seems inexact-- really you want to infer the concentration as a function of height, so a set of $N_{layers}$ scalars. These scalars are connected in some way, and so maybe the right thing to do is to treat them as (shrinkage) prior means in a hierarchical framework, and infer revised values.
# Operationally, a head-to-head comparison should simply set the exact same P-T profile and not "tweak" the values as is done in a fit. Simply, do we get the same input/output as TelFit?
# ## Run LBLRTM with TelFit, match to HPF resolution/sampling
from astropy.io import fits
from astropy.table import Table
hdus = fits.open('../../ucdwhpf/data/HPF/goldilocks/Goldilocks_20200507T100451_v1.0_0017.spectra.fits')
df_all = pd.DataFrame()
for m in range(28):
tab = Table({hdus[i].name:hdus[i].data[m,:] for i in range(1,10)})
tab['order'] = m
df_all = df_all.append(tab.to_pandas(), ignore_index=True)
df_all.tail(1)
lo, hi = df_all['Sci Wavl'].min(), df_all['Sci Wavl'].max()
nus = np.array((df_all['Sci Wavl'].values*u.Angstrom).to(1/u.cm, equivalencies=u.spectral()).value)
nus.shape
# %%capture
from telfit import Modeler
telfit_modeler = Modeler()
# %%time
result = telfit_modeler.MakeModel(pressure=1005.0, temperature=273.15+14.0,
lowfreq=nus.min(), highfreq=nus.max(), angle=0.0,humidity=20.0,
lat=19.8, alt=2.0,resolution=55000.0,vac2air=False)
plt.figure(figsize=(18,4))
plt.plot(result.x, result.y, alpha=0.3, color='k');
# Ok! We have TelFit working. Our new GPU framework is next!
| notebooks/07_TelFit_tet_a_tet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.9 64-bit
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# Colab
# # !git clone https://github.com/mjsmagalhaes/examples-datascience.git repo
# # %cd repo
# # %pip install -r _requirements/dslib.txt
# -
import pandas as pd
import plotly.graph_objects as go
from pycoingecko import CoinGeckoAPI
# Import Data
cg = CoinGeckoAPI()
raw_data = cg.get_coin_market_chart_by_id(
'bitcoin', vs_currency='usd', days=15)
# Transform Data
data = pd.DataFrame(raw_data['prices'], columns=['timestamp', 'price'])
data['date'] = pd.to_datetime(data['timestamp'], unit='ms')
# +
# Create Graphs
chart = data.groupby(data.date.dt.date).agg(
{'price': ['min', 'max', 'first', 'last']})
fig = go.Figure(data=[
go.Candlestick(
x=chart.index,
open=chart['price']['first'],
high=chart['price']['max'],
low=chart['price']['min'],
close=chart['price']['last']
)
])
fig.update_layout(xaxis_title='Date', yaxis_title='Price', title='BitCoin')
fig.write_html("bitcoin_variation.html")
fig.show(renderer="png")
| dsexamples/courses/ibm_bitcoin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Barcelona Airbnb 2021
# ## Section 1: Business Understanding
#
# This project intends to evaluate the effect of the Covid19 pandemic on [Airbnb hosting prices](http://insideairbnb.com/get-the-data.html) in the city of Barcelona. We use the number of reviews as a metric for the number of tourists in the city and the number of positive cases for the prevalence of [Covid19](https://cnecovid.isciii.es/covid19/#documentaci%C3%B3n-y-datos) in the region of Catalonia.
#
#
#
# ### Question 1: What was the impact of the pandemic on Airbnb accommodation from the second outbreak onwards?
# ### Question 2: Which are the key success factors to attract guests in this situation?
# ### Question 3: How did price evolve in these ever-changing circumstances?
#
#
# Sources: [Inside Airbnb](http://insideairbnb.com/get-the-data.html), [Covid19](https://cnecovid.isciii.es/covid19/#documentaci%C3%B3n-y-datos)
#
# +
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import datetime
import seaborn as sns
from sklearn.preprocessing import OrdinalEncoder
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn import metrics
from sklearn.ensemble import RandomForestClassifier
# %matplotlib inline
# +
#Fetching data. From the Airbnb source, we create a dataset dictionary with an entry for each month
raw_df_dict = {'aug': pd.read_csv('./data/listings_aug20.csv'),
'sep': pd.read_csv('./data/listings_sept20.csv'),
'oct': pd.read_csv('./data/listings_oct20.csv'),
'nov': pd.read_csv('./data/listings_nov20.csv'),
'dec': pd.read_csv('./data/listings_dec20.csv'),
'jan': pd.read_csv('./data/listings_jan21.csv'),
'feb': pd.read_csv('./data/listings_feb21.csv'),
'mar': pd.read_csv('./data/listings_mar21.csv'),
'apr': pd.read_csv('./data/listings_apr21.csv')
}
months = raw_df_dict.keys()
for month in months:
raw_df_dict[month].set_index('id')
# -
# # Data Exploration
#
# For the data exploration, only one dataset will be used since all of them share the same columns and have similar number of entries. Therefore, all of them will be processed equally
# Columns of the datasets
raw_df_dict['aug'].columns
# # Preprocessing Data
# +
# Data from 'raw_df_dict' will be processed and stored in 'df_dict'
df_dict = {month: pd.DataFrame() for month in months}
# Most data are ready to use
for month in months:
# Filter numerical features from the DataFrame
numerical_columns = raw_df_dict[month].select_dtypes(include='number').columns
# Drop non-relevant features
numerical_columns = numerical_columns.drop(['scrape_id', 'host_id'])
# Insert into the new DataFrame
df_dict[month][numerical_columns] = raw_df_dict[month][numerical_columns]
# -
# ## Format Fixing
def host_aceptance_rate_to_numeric(rate_string):
""" Transforms the string type column 'host_acceptance_rate' into int type
INPUT
rate_string: string
contains the string type column 'host_acceptance_rate' (format example '90%')
OUTPUT
rate_numeric: int
contains the numeric value of 'host_acceptance_rate'
"""
try:
rate_numeric = int(rate_string[:-1])
except:
rate_numeric = np.nan
return rate_numeric
def price_to_numeric(price):
""" Transforms the string type column 'price' into int type
Parameters
----------
price : string
contains the price (format example '$1,000.00')
Returns
-------
price_numeric : int, NaN
returns the numeric value of 'price'. Returns NaN if price is not a string
"""
try:
price_numeric = int(price[1:-3].replace(',',''))
except:
price_numeric = np.nan
return price_numeric
def string_to_date(date_string, date_format='%Y-%m-%d'):
"""" Converts string into datetime type
Parameters
----------
date_string: string
contains the date
date_format: string
contains the format of the date
Returns
-------
date : datetime, NaT
contains the date. It is NaT if date_string is NaN
"""
date = pd.to_datetime(date_string, format=date_format)
return date
# Some numeric data are formatted as strings: 'price' and 'host_acceptance_rate'
for month in months:
# Column 'price' is string type. We store the int value.
df_dict[month]['price'] = raw_df_dict[month]['price'].apply(price_to_numeric)
# Column 'host_acceptance_rate' is string type. We store the int value.
df_dict[month]['host_acceptance_rate'] = raw_df_dict[month]['host_acceptance_rate'].apply(host_aceptance_rate_to_numeric)
# +
# We transform temporal variables from string to datetime type
date_columns = ['last_scraped',
'host_since',
'calendar_last_scraped',
'first_review']
for month in months:
df_dict[month][date_columns] = raw_df_dict[month][date_columns].applymap(string_to_date)
# -
# ## Encoding
# Ordinal
host_response_time_tiers = {'within an hour': 1,
'within a few hours': 2,
'within a day': 3,
'a few days or more': 4
}
for month in months:
df_dict[month]['host_response_time'] = raw_df_dict[month]['host_response_time'].replace(host_response_time_tiers)
# +
# Boolean
bool_columns = ['host_is_superhost',
'host_has_profile_pic',
'host_identity_verified',
'has_availability',
'instant_bookable']
for month in months:
df_dict[month][bool_columns]=raw_df_dict[month][bool_columns].replace({'t': 1, 'f': 0})
# -
# One-hot
neig_columns = ['neighbourhood_cleansed','neighbourhood_group_cleansed']
for month in months:
dummies = pd.get_dummies(raw_df_dict[month][neig_columns], dummy_na=False)
df_dict[month][dummies.columns] = dummies
# ## Missing values
sns.heatmap(df_dict['aug'].isna())
for month in months:
# we remove any column with more than a 20% of missing values
df_dict[month].dropna(how='all', thresh = 0.80*df_dict[month].shape[0], axis = 1, inplace=True)
# after that, only a few listings have missing values
df_dict[month].dropna(axis=0, how='any', inplace=True)
df_dict['aug'].columns[df_dict['aug'].isna().sum()>0] # check that there isn't any NaN
# ## Extra columns
for month in months:
# To solve the bed/accomm paradox, the number of occupants is the maximum of both
df_dict[month]['capacity'] = raw_df_dict[month].apply(lambda x: np.nanmax([x['beds'], x['accommodates']]), axis = 1)
# We also are interested in the price per guest, as it might be more relevant
df_dict[month]['price_person'] = df_dict[month]['price']/df_dict[month]['capacity']
# We can distinguish between visited and non visted listings
df_dict[month]['had_reviews']=raw_df_dict[month]['number_of_reviews_l30d'].apply(lambda x: 0 if x==0 else 1)
# # Question 1: What was the impact of the pandemic on Airbnb accommodation from the second outbreak onwards?
#
# Using the daily record of infections, we can measure the impact of the pandemic in different regions of Spain. We will compare the number of infections to the monthly number of reviews, which is highly correlated to the number of visitors. In doing so, we can do some inferences about the effect on tourism.
#
# The daily record of Covid19 infections in every region of Spain can be found [here](https://cnecovid.isciii.es/covid19/#documentaci%C3%B3n-y-datos).
# Dates when each dataset was created
dates = {'aug': datetime.datetime(2020,8,1),
'sep': datetime.datetime(2020,9,1),
'oct': datetime.datetime(2020,10,1),
'nov': datetime.datetime(2020,11,1),
'dec': datetime.datetime(2020,12,1),
'jan': datetime.datetime(2021,1,1),
'feb': datetime.datetime(2021,2,1),
'mar': datetime.datetime(2021,3,1),
'apr': datetime.datetime(2021,4,1)
}
date_series = pd.Series(data=dates)
# We load the Covid19 contagion data in Spain
cases_total = pd.read_csv('./data/casos_tecnica_ccaa.csv') #covid cases in Spain
cases_total.set_index('ccaa_iso', inplace = True)
cases_cat = cases_total.loc['CT',['fecha','num_casos']] #covid cases in Catalonia
cases_cat['fecha'] = [datetime.datetime.strptime(date,"%Y-%m-%d" ) for date in cases_cat['fecha']]
cases_cat = cases_cat[(cases_cat['fecha']>=dates['aug']-datetime.timedelta(30))&(cases_cat['fecha']<=dates['apr'])]
# +
# List with the mean number of reviews of each month
mean_num_reviews = [df_dict[month]['number_of_reviews_l30d'].mean() for month in months]
y = mean_num_reviews
fig, ax1 = plt.subplots()
ax1.bar(date_series, mean_num_reviews,
ls='-', color = 'powderblue',
width=20, label = 'Reviews')
ax1.set_xlabel('date')
ax1.set_ylabel('No. reviews')
ax1.set_title('No. reviews and Covid-19 cases')
ax1.xaxis.set_tick_params(rotation = 90)
ax1.set_ylim(ymin=0, ymax=1.5*np.nanmax(y))
ax2 = ax1.twinx()
ax2.plot_date(cases_cat['fecha'], cases_cat['num_casos'], '--',
xdate=True, linewidth=0.8, label='Covid', color='coral')
ax2.set_ylabel('No. of cases')
ax1.legend()
plt.tight_layout()
plt.savefig('q1.png', dpi=300)
# -
# By June 2020, the first wave started to decrease, leading to a peaceful summer. But in reality, summer 2020 was nothing but the eye of the storm, and by October, a fast second wave stroke in the region.
#
# This second outbreak led to stricter traveling restrictions, which resulted in a decrease on the number of guests. By December 2020, prevalence declined to the level of the previous summer. However, this was not enough to convince travelers since the ease of restrictions took some time to appear. In addition, the uncertainty of the situation and the arrival of Christmas did not create the best scenario for traveling.
# It comes as no surprise that Christmas celebrations meant a source of contagion, and a third wave took place. The number of visitors kept growing, though, and that trend continued until April 2021.
#
# # Question 2: How did price evolved in these ever-changing circumstances?
# Given the volatile situation, one might assume that hosts would adapt their strategy. Although they have little to take control of, an expectable response is to decrease the price of the listing to make it a better competitor against their neighbors.
#
visited_price = []
empty_price = []
for month in months:
visited_price.append(df_dict[month].loc[df_dict[month]['number_of_reviews_l30d']>0,'price_person'].mean())
empty_price.append(df_dict[month].loc[df_dict[month]['number_of_reviews_l30d']==0,'price_person'].mean())
# +
y = visited_price
fig, ax1 = plt.subplots()
ax1.set_xlabel('date')
ax1.set_title('Price per person and Covid-19 cases')
ax1.xaxis.set_tick_params(rotation = 90)
ax1.bar(date_series, mean_num_reviews, color = 'powderblue',
width=20)
ax1.set_ylim(ymin=0, ymax=1.2*np.nanmax(mean_num_reviews))
ax1.set_ylabel('No. reviews')
ax2 = ax1.twinx()
ax2.plot_date(date_series, visited_price,
xdate=True, ls='-', color = 'coral',
label = 'visited')
ax2.plot_date(date_series, empty_price,
xdate=True, ls='-', color = 'darkturquoise',
label = 'empty')
ax2.set_ylim(ymin=0, ymax=1.3*np.nanmax(empty_price))
ax2.set_ylabel('Price per person')
ax2.legend(loc=1)
plt.tight_layout()
plt.savefig('q2.png', dpi=300)
# -
# Some differences can be found between non-visited and visited listings
# 1. Visited listings have a cheaper average price.
# 2. The average price of visited listings decreased after the autumn Covid19 wave.
# 3. The price of non-visited listings, instead, increased on average.
#
#
# During highly restricted period, the preference towards lower prices decreased the mean price of visited listings.
#
# # Question 3: Which are the key success factors to attract guests in this situation?
# In other words, what can be done to keep afloat your Airbnb?
df_dict['jan'].columns
df_comparison = df_dict['jan'].groupby('had_reviews').mean()
df_comparison.index.name = None
df_comparison = df_comparison.T
df_comparison['Relative difference'] = df_comparison.apply(lambda x: 100*(x[1]-x[0])/((x[0]+x[1])/2), axis = 1)
df_comparison
# +
features = ['price_person', 'capacity', 'host_listings_count',
'host_is_superhost', 'instant_bookable', 'host_identity_verified']
plot_df = df_comparison.loc[features]
plot_df.sort_values(by='Relative difference', inplace=True)
colors = plot_df['Relative difference'].apply(lambda x: 'powderblue' if x>=0 else 'salmon')
fig, ax = plt.subplots()
ax.barh(plot_df.index, width=plot_df['Relative difference'], color=colors)
ax.set_title('Relative difference between visited and non-visited listings')
ax.set_xlabel('relative difference (%)')
plt.savefig('q3.png', dpi=400, bbox_inches='tight')
# -
# As expected, cheaper listings receive more visitors than expensive ones. This fact can be used in our favor to outsell the rest of competitors. But there is a point when the costs surpass the revenues, so this might not be an option in many cases
# On the other hand, guests also show a preference for instant bookable listings. Therefore, enabling instant bookings might be a fast and easy option to increase the number of reservations. In addition, those listings whose host is superhost and has verified its identity seem to have earn trust from guests.
# The owner of these listings usually have fewer listings, which means hotels are less appealing, arguably due to their higher price.
| airbnb_bcn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext nb_black
# +
# %matplotlib inline
import warnings
import pandas as pd
import numpy as np
import seaborn as sns
from scipy import stats
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.stats.outliers_influence import variance_inflation_factor
import os
print(os.getcwd())
print(os.listdir(os.getcwd()))
# -
def print_vif(x):
"""Utility for checking multicollinearity assumption
:param x: input features to check using VIF. This is assumed to be a pandas.DataFrame
:return: nothing is returned the VIFs are printed as a pandas series
"""
# Silence numpy FutureWarning about .ptp
with warnings.catch_warnings():
warnings.simplefilter("ignore")
x = sm.add_constant(x)
vifs = []
for i in range(x.shape[1]):
vif = variance_inflation_factor(x.values, i)
vifs.append(vif)
print("VIF results\n-------------------------------")
print(pd.Series(vifs, index=x.columns))
print("-------------------------------\n")
# # Initial EDA
pd.set_option("display.max_columns", None)
churn = pd.read_excel("WA_Fn-UseC_-Telco-Customer-Churn.xlsx")
churn.head()
churn = pd.get_dummies(churn, columns=["Churn"], drop_first=True)
churn = churn.drop(columns=["customerID"])
churn.shape
# No missing data
churn.isna().mean()
col = churn[0:]
for c in col:
print("Value counts for " + c)
print(churn[c].value_counts())
churn.dtypes
bin_cols = [
"SeniorCitizen",
]
cat_cols = [
"gender",
"Partner",
"Dependents",
"PhoneService",
"MultipleLines",
"InternetService",
"OnlineSecurity",
"OnlineBackup",
"DeviceProtection",
"TechSupport",
"StreamingTV",
"StreamingMovies",
"Contract",
"PaperlessBilling",
"PaymentMethod",
]
drop_cats = [
"Male",
"No",
"No",
"Yes",
"No",
"Fiber optic",
"No",
"No",
"No",
"No",
"No",
"No",
"Month-to-month",
"Yes",
"Electronic check",
]
num_cols = ["tenure", "MonthlyCharges", "TotalCharges"]
churn["TotalCharges"] = churn["TotalCharges"].replace(" ", np.nan, regex=True)
churn["TotalCharges"].loc[488]
pd.to_numeric(churn["TotalCharges"])
churn[num_cols].dtypes
sns.countplot(churn["Churn_Yes"])
plt.title("Amount Churn")
plt.ylabel("People")
plt.xlabel("Churn")
plt.show()
churn["Churn_Yes"].value_counts()
for col in bin_cols:
perc_churn = churn[["Churn_Yes", col]].groupby(col).mean()
display(perc_churn)
sns.countplot(hue="Churn_Yes", x=col, data=churn)
plt.show()
for col in num_cols:
sns.boxplot("Churn_Yes", col, data=churn)
plt.show()
sns.pairplot(churn[num_cols + ["Churn_Yes"]], hue="Churn_Yes")
plt.show()
# Total Charges seems to be correlated with most of the num_cols
plt.figure(figsize=(10, 8))
sns.heatmap(churn[num_cols + ["Churn_Yes"]].corr(), vmin=-1, vmax=1, annot=True)
plt.show()
churn = churn.drop(columns=["TotalCharges"])
pd.get_dummies(
churn,
columns=[
"gender",
"Partner",
"Dependents",
"PhoneService",
"MultipleLines",
"InternetService",
"OnlineSecurity",
"OnlineBackup",
"DeviceProtection",
"TechSupport",
"StreamingTV",
"StreamingMovies",
"Contract",
"PaperlessBilling",
"PaymentMethod",
],
drop_first=True,
)
# turn into list
bin_cols = churn.drop(columns=["tenure", "MonthlyCharges", "Churn_Yes"])
# Getting error but not too sure what the error means
for col in bin_cols:
perc_churn = churn[["Churn_Yes", col]].groupby(col).mean()
display(perc_churn)
sns.countplot(hue="Churn_Yes", x=col, data=churn)
plt.title(col)
plt.show()
churn.head()
X = churn.drop(columns=["Churn_Yes"])
X.head()
print_vif(X.select_dtypes("number"))
| Initial EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv('../data/bank-additional/bank-additional-full.csv', sep=';')
# -
df
# convert
df['conversion'] = df['y'].apply(lambda x: 1 if x == 'yes' else 0)
df.head()
# total number of conversions
df.conversion.sum()
# total number of clients in the data (= number of rows in the data)
df.shape[0]
print('total conversions: %i out of %i'% (df.conversion.sum(), df.shape[0]))
print('conversion rate: %0.2f%%'% (df.conversion.sum()/ df.shape[0]*100.0))
# +
# conversion rates by age
conversions_by_age = df.groupby(
by='age'
)['conversion'].sum() / df.groupby(
by='age'
)['conversion'].count() * 100.0
conversions_by_age
# +
ax = conversions_by_age.plot(
grid=True,
figsize=(10, 7),
title='Conversion Rates by Age'
)
ax.set_xlabel('age')
ax.set_ylabel('conversion rate (%)')
plt.show()
# -
df['age_group'] = df['age'].apply(
lambda x: '[18, 30)' if x < 30 else '[30, 40)' if x < 40 \
else '[40, 50)' if x < 50 else '[50, 60)' if x < 60 \
else '[60, 70)' if x < 70 else '70+'
)
conversions_by_age_group = df.groupby(
by='age_group'
)['conversion'].sum() / df.groupby(
by='age_group'
)['conversion'].count() * 100.0
conversions_by_age_group
# +
ax = conversions_by_age_group.loc[
['[18, 30)', '[30, 40)', '[40, 50)', '[50, 60)', '[60, 70)', '70+']
].plot(
kind='bar',
color='skyblue',
grid=True,
figsize=(10, 7),
title='Conversion Rates by Age Groups'
)
ax.set_xlabel('age')
ax.set_ylabel('conversion rate (%)')
plt.show()
# -
conversions_by_marital_status_df = pd.pivot_table(df, values='y', index='marital', columns='conversion', aggfunc=len)
# +
conversions_by_marital_status_df.plot(
kind='pie',
figsize=(15, 7),
startangle=90,
subplots=True,
autopct=lambda x: '%0.1f%%' % x
)
plt.show()
# +
age_marital_df = df.groupby(['age_group', 'marital'])['conversion'].sum().unstack('marital').fillna(0)
age_marital_df = age_marital_df.divide(
df.groupby(
by='age_group'
)['conversion'].count(),
axis=0
)
# -
age_marital_df
# +
ax = age_marital_df.loc[
['[18, 30)', '[30, 40)', '[40, 50)', '[50, 60)', '[60, 70)', '70+']
].plot(
kind='bar',
grid=True,
figsize=(10,7)
)
ax.set_title('Conversion rates by Age & Marital Status')
ax.set_xlabel('age group')
ax.set_ylabel('conversion rate (%)')
plt.show()
# +
ax = age_marital_df.loc[
['[18, 30)', '[30, 40)', '[40, 50)', '[50, 60)','[60, 70)', '70+']
].plot(
kind='bar',
stacked=True,
grid=True,
figsize=(10,7)
)
ax.set_title('Conversion rates by Age & Marital Status')
ax.set_xlabel('age group')
ax.set_ylabel('conversion rate (%)')
plt.show()
# -
| chapter2_KPIs_and_visualisations/ConversionRate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import networkx as nx
import random
n = 500
p = (n * 20.) / (n * (n - 1) * 0.5)
print("N={}, p={}".format(n, p))
def erdos_renyi_graph(n, p):
graph = {k: [] for k in range(n)}
for i in range(n):
for j in range(i + 1, n):
if random.random() < p:
graph[i].append(j)
graph[j].append(i)
return graph
graph_adj = erdos_renyi_graph(n, p)
graph = nx.Graph(graph_adj)
# +
def singe_source_shortes_paths(graph, s):
pred = {k: [] for k in graph}
dist = {k: float("inf") for k in graph}
sigma = {k: 0. for k in graph}
dist[s] = 0
sigma[s] = 1.
queue = [s]
stack = []
while queue:
v = queue.pop(0)
stack.append(v)
for w in graph[v]:
if dist[w] == float("inf"):
dist[w] = dist[v] + 1
queue.append(w)
if dist[w] == dist[v] + 1:
sigma[w] += sigma[v]
pred[w].append(v)
return pred, dist, sigma, stack
def accumulation(graph, s, stack, pred, sigma, btw):
delta = {k: 0. for k in graph}
while stack:
w = stack.pop()
for v in pred[w]:
delta[v]+= float(sigma[v] * (1 + delta[w])) / sigma[w]
if w != s:
btw[w] += delta[w]
return btw
# +
# %%time
betweenness = {k: 0. for k in graph}
for s in graph_adj:
pred, dist, sigma, stack = singe_source_shortes_paths(graph_adj, s)
betweenness = accumulation(graph_adj, s,
stack, pred, sigma, betweenness)
betweenness = {k: betweenness[k] / 2. for k in betweenness}
my_version = betweenness.values()
# -
# %%time
nx_version = nx.betweenness_centrality(graph, normalized=False).values()
import numpy as np
np.testing.assert_almost_equal(my_version, nx_version, 10)
| hw-2/BetweennessCentrality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
name = input("If you had 3 children, what would the name of the middle child be?")
print(name,"huh. unoriginal")
age = int(input("What is your current age?"))
if age > 50 :
print(age,"dang, you're old!")
else :
print("Wow, only", age, "you're young!")
smallDogAge = age * 15
print("If you were a small dog you would be", smallDogAge, "years old.")
| codingpart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf18p36
# language: python
# name: tf18p36
# ---
# + language="bash"
# curl http://10.9.3.3:8008/lidingke/ctpn/raw/master/rpc/ctpn.proto --silent -O ctpn.proto
# # source activate tf18p36
# python -m grpc_tools.protoc --python_out=. --grpc_python_out=. -I. ctpn.proto
# rm ctpn.proto
# head -n 10 ctpn_pb2_grpc.py
# + language="bash"
# curl http://10.9.3.3:8008/lidingke/yolo/raw/master/yolo.proto --silent -O yolo.proto
# python -m grpc_tools.protoc --python_out=. --grpc_python_out=. -I. yolo.proto
# rm yolo.proto
# head -n 10 yolo_pb2_grpc.py
# -
| ldklib/jupyter_demo/grpc_install.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''PythonData'': conda)'
# name: python3
# ---
import pandas as pd
import plotly.express as px
# read the data
# data originally from: https://simplemaps.com/data/au-cities
df = pd.read_csv("https://intro-to-python-asdaf.s3.ap-southeast-2.amazonaws.com/australian_cities.csv")
# show the data
df.head()
# documentation for scatter_mapbox: https://plotly.github.io/plotly.py-docs/generated/plotly.express.scatter_mapbox.html
fig = px.scatter_mapbox(df, lat="lat", lon="lng", hover_name="city", size="population", zoom=3, size_max=60, mapbox_style="open-street-map", height=600, opacity=0.5)
fig.update_layout(margin={"r":0,"t":0,"l":0,"b":0}) # set margins to 0 so that map is bigger
fig.show()
| day-2/06-ins_working-with-geo-data/solved/plotting-australian-cities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# There are 3 types of forms in the eRetrieval for SVOPAC 460, 496, 410:
#
#
# Form 496 is 24-hour/10-day Independent Expenditure Report. All independent expenditures reported on Form 496 must also be reported on subsequent campaign reports (i.e., Forms 460, 450 or 461).
#
# Form 460 is Recipient Committee Campaign Statement
# Use the Form 460 to file any of the following:
# - Preelection Statement
# - Semi-annual Statement
# - Quarterly Statement
# - Special Odd-Year Report
# - Termination Statement
# - Amendment to a previously filed statement
#
#
# Form 460 Consist of several Schedules
# - Recipient Committee Campaign Statement: Cover Page, Type of Recipient Committee,
# Type of Statement,
# Committee Information (name) etc
# - Campaign Disclosure Statement Summary Page: Accounting information
# - Schedule A: Monetary Contributions Received Dates names, amounts, codes
# - Schedule B: Loans Received Dates names, amounts, codes
# - Schedule C: Nonmonetary Contributions Received Dates names, amounts, codes
# - Schedule D: Summary of Expenditures Supporting/Opposing Other Candidates, Measures and Committees support oppose, type of payment, amounts
# - Schedule E: Payments Made Dates names, amounts, codes
# - Schedule F: Accrued Expenses Dates names, amounts, codes
# - Schedule I: Miscellaneous Increases to Cash Dates names, amounts
#
#
# Codes
# Entity_Cd
# - IND – Individual
# - COM – Recipient Committee
# (other than PTY or SCC)
# - OTH – Other (e.g., business entity)
# - PTY – Political Party
# - SCC – Small Contributor Committee
#
#
# Expn_Code
# - CMP campaign paraphernalia/misc.
# - CNS campaign consultants
# - CTB contribution (explain nonmonetary)*
# - CVC civic donations
# - FIL candidate filing/ballot fees
# - FND fundraising events
# - IND independent expenditure supporting/opposing others (explain)* LEG legal defense
# - LIT campaign literature and mailings
# - MBR member communications MTG meetings and appearances OFC office expenses
# - PET petition circulating
# - PHO phone banks
# - POL polling and survey research
# - POS postage, delivery and messenger services PRO professional services (legal, accounting) PRT print ads
# - RAD radio airtime and production costs
# - RFD returned contributions
# - SAL campaign workers’ salaries
# - TEL t.v. or cable airtime and production costs
# - TRC candidate travel, lodging, and meals
# - TRS staff/spouse travel, lodging, and meals
# - TSF transfer between committees of the same candidate/sponsor VOT voter registration
# - WEB information technology costs (internet, e-mail)
#
#
# Sup_Opp_Cd
# - S in support of a candidate or measure
# - O opposing a candidate or a measure
# +
import pandas as pd
import numpy as np
# -
# readimng all files (460s only)
import glob
files = glob.glob('files/*.xls')
files
# +
from datetime import datetime
my_dict = {}
for f in files:
data = pd.read_excel(f)
my_dict[f] = ( data['From_Date'][1], data['Thru_Date'][1],data['Rpt_Date'][1] )
df_new = pd.DataFrame({'file_name':files})
# -
df_new
new_df = pd.DataFrame.from_dict(my_dict, orient='index')
df_new
new_df.columns = ['From_Date', 'Thru_Date', 'Rpt']
new_df
new_df['file_name'] =new_df.index
new_df.sort_values(["From_Date","Rpt"], axis = 0, ascending = True,
inplace = True, na_position ='last')
new_df
# +
#new_df['from'] = pd.to_datetime(new_df['From_Date'])
#new_df['thru'] = pd.to_datetime(new_df['Thru_Date'])
#new_df['rpt_l'] = new_df['thru']-new_df['from']
# -
# +
#Comparing 71-1231 and 71 10 , 10 -12
#range
semi= pd.read_excel('files/transactionExportGrid-7.xls')
semi_710= pd.read_excel('files/transactionExportGrid-2.xls')
semi_1012= pd.read_excel('files/transactionExportGrid-13.xls')
semi_comb = semi_710.append(semi_1012)
# -
print(len(semi), len(semi_comb))
# Here is a <span style="color:red">discrepancy</span> between semiannual report and combined quaterly reports for same period(amendment)
# To see what is different between them we have to create outer join
# +
result = pd.merge(semi,
semi_comb,
on='Tran_ID',
how='outer',
indicator=True)
result
result.to_csv('outer_join.csv')
# -
print(len(result))
result = result.drop_duplicates(['Tran_ID'], keep='first')
new_df = new_df.drop_duplicates(subset=new_df.columns.difference(['Rpt','file_name']), keep='last')
new_df
files = new_df.file_name.tolist()
#new list of files semiannual rerport included joined with every report and ammendment , report removed if there is an amendment filed on a later date
files
# +
df = pd.DataFrame()
for f in files:
data = pd.read_excel(f)
df = df.append(data)
# -
len(df)
#creating csv of dublicates
ids = df["Tran_ID"]
df_dub_csv = df[ids.isin(ids[ids.duplicated()])]
df_dub_csv.to_csv('f_dub.csv')
# Dublcates are created by schedule D and Some items on Schedule E. Schedule D however contains interesting information: what was expenditure for: opposing or supporting certain candidate
# +
#df = df.drop_duplicates(['Tran_ID'], keep='first')
# -
df.shape
df_D = df[df['Form_Type']=='D']
len(df_D)
# +
#df= df[df['Form_Type']!='D']
#df
#df = df.drop(df[df['Form_Type']=='D'].index)
# +
#len(df)
# +
#df_D = df_D[['Tran_ID','Office_Cd', 'Offic_Dscr', 'Juris_Cd', 'Juris_Dscr', 'Off_S_H_Cd', 'Bal_Name', 'Bal_Num', 'Bal_Juris', 'Sup_Opp_Cd', 'Memo_Code']]
# -
#
# +
#df.merge(df_D, left_on='Tran_ID', suffixes=('_', '_D'))
# +
#df1 = pd.merge(df, df_D, on='Tran_ID')
# +
#df2=pd.merge(df, df_D, on='Tran_ID', how='outer')
# +
#len(df2)
# +
#df2.to_csv('f_D.csv')
# +
#df[df.duplicated(['Rpt_Date'],keep=False)].sort_values(by=['Tran_ID'])
# +
#df = df.drop_duplicates(['Tran_ID'])
# +
#df.head()
# +
#df.to_csv('f.csv')
# +
#len(df)
# -
# To remove dublicats and give preference to schedule D colums 'Tran_ID', 'Form_Type' are sorted and first record is kept
df.sort_values([ 'Tran_ID', 'Form_Type']).to_csv('sorted_all.csv')
df.sort_values([ 'Tran_ID', 'Form_Type']).tail(40)
df =df.sort_values([ 'Tran_ID', 'Form_Type'])
df = df.drop_duplicates(['Tran_ID'], keep='first')
len(df)
df.head()
df.to_csv('df.csv')
active_code = df.Expn_Code.unique().tolist()
df.Entity_Cd.unique()
# +
data = df[df['Rec_Type']!='EXPN']
source = ColumnDataSource(data={
'x' : data.Tran_Date,
'y' : data.Amount,
'r' : (data.Amount / 1200) + 10,
'tid' : data.Tran_ID,
'Amount' : data.Amount,
'Entity_Nam' : data['Entity_Nam L'],
'Entity_Cd' : data.Entity_Cd,
})
hover = HoverTool(
tooltips = [
("Trid", "@tid"),
("Amount", "@Amount"),
("Entity_Nam", "@Entity_Nam"),
])
p1 = figure(x_axis_type="datetime", title='2018', x_axis_label='Tran_Date', y_axis_label='Amount',
# x_axis_type='datetime',
plot_height=400, plot_width=700,
tools=[hover])
color_mapper = CategoricalColorMapper(factors=['OTH', 'IND', 'COM', 'SCC'],
palette=['red', 'green', 'blue', 'orange'])
# Add a circle glyph to the figure p
p1.circle(x='x', y='y', alpha=0.2, size = 'r', source=source, color=dict(field='Entity_Cd', transform=color_mapper),
legend='Entity_Cd')
p1.legend.location = 'top_left'
# Output the file and show the figure
output_file('gapminder.html')
#show(p1)
# +
dat = df[df['Rec_Type']=='EXPN']
# -
dat = dat.fillna('None')
from bokeh.palettes import d3
palette = d3['Category20'][len(dat['Expn_Code'].unique())]
palette
list_code = dat['Expn_Code'].unique().tolist()
from bokeh.transform import linear_cmap
color_map = CategoricalColorMapper(factors = ['FND', 'IND', 'LIT', 'None', 'MTG', 'PRO', 'OFC', 'WEB', 'CNS', 'MON',
'POL']
,palette=['#1f77b4',
'#aec7e8',
'#ff7f0e',
'#ffbb78',
'#2ca02c',
'#98df8a',
'#d62728',
'#ff9896',
'#9467bd',
'#c5b0d5',
'#8c564b'])
# +
from bokeh.palettes import Spectral6
source = ColumnDataSource(data={
'x' : dat.Tran_Date,
'y' : dat.Amount,
'r' : (dat.Amount / 1200) + 10,
'tid' : dat.Tran_ID,
'Amount' : dat.Amount,
'Entity_Nam' : dat['Entity_Nam L'],
'Expn_Code' : dat.Expn_Code,
'Entity_Cd' : data.Entity_Cd,
})
hover = HoverTool(
tooltips = [
("Trid", "@tid"),
("Amount", "@Amount"),
("Entity_Nam", "@Entity_Nam"),
])
mapper = linear_cmap(field_name='Entity_Cd', palette=Spectral6 ,low=0 ,high=11)
p2 = figure(x_axis_type="datetime", title='2018_1', x_axis_label='Tran_Date', y_axis_label='Amount',
# x_axis_type='datetime',
plot_height=400, plot_width=700,
tools=[hover])
#color_mapper = CategoricalColorMapper(factors=['OTH', 'IND', 'COM', 'SCC'], palette=['red', 'green', 'blue', 'orange'])
from bokeh.transform import factor_cmap
# Add a circle glyph to the figure p
p2.circle(x='x', y='y', alpha=0.2, size = 'r', source=source, color=dict(field='Expn_Code', transform=color_map)
,legend='Expn_Code')
p2.legend.location = 'top_left'
# Output the file and show the figure
output_file('gapminder.html')
p22 = row(controls, p2)
#show(p2)
# +
# Import Tabs from bokeh.models.widgets
from bokeh.models.widgets import Tabs
from bokeh.models.widgets import Panel
# Create tab1 from plot p1: tab1
tab1 = Panel(child=p1, title='Income')
# Create tab2 from plot p2: tab2
tab2 = Panel(child=p22, title='Expenses')
# Create a Tabs layout: layout
layout = Tabs(tabs=[tab1, tab2])
# Link the x_range of p2 to p1: p2.x_range
p2.x_range = p1.x_range
# Link the y_range of p2 to p1: p2.y_range
p2.y_range = p1.y_range
# Specify the name of the output_file and show the result
output_file('tabs.html')
show(layout)
# +
from bokeh.models.widgets import CheckboxGroup
# Create the checkbox selection element, available carriers is a
# list of all airlines in the data
list_code_ = CheckboxGroup(labels=active_code,
active = [0, 1])
from bokeh.layouts import column, row, WidgetBox
from bokeh.models import Panel
from bokeh.models.widgets import Tabs
# Put controls in a single element
controls = WidgetBox(list_code_)
# Create a row layout
layout = row(controls, p1)
# Make a tab with the layout
tab = Panel(child=layout, title = 'Delay Histogram')
tabs = Tabs(tabs=[tab])
# -
# Update function takes three default parameters
def update(attr, old, new):
# Get the list of carriers for the graph
carriers_to_plot = [list_code_.labels[i] for i in
list_code_.active]
# Update the source used in the quad glpyhs
src.data.update(new_src.data)
# Link a change in selected buttons to the update function
list_code_.on_change('active', update)
| 460visualizations/bubblegraph/Viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("..")
import petsi
from random import gauss, uniform, random
import math
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import display
# %matplotlib inline
# + jupyter={"source_hidden": true}
def create_simulator(initial_work, utilization, firing_distribution, **loop_backs):
# print(f" Initial work:{initial_work}, utilization={utilization}")
simulator = petsi.Simulator("Activity time")
simulator.add_place("ToDo")
simulator.add_place("Done")
simulator.add_immediate_transition("start", priority=1)
simulator.add_inhibitor("is idle", "ToDo", "start")
for i in range(initial_work):
simulator.add_constructor(f"initial token #{i}", "start", "ToDo")
simulator.add_timed_transition("doing", firing_distribution)
simulator.add_transfer("do", "ToDo", "doing", "Done")
constructor_weights = 0
for transition_name, (multiplier, weight) in loop_backs.items():
if multiplier > 0 and weight > 0:
# print(f" {transition_name} multiplier={multiplier}, weight={weight}")
simulator.add_immediate_transition(transition_name, priority=2, weight=weight)
simulator.add_transfer(f"{transition_name}", "Done", transition_name, "ToDo")
for i in range(multiplier-1):
constructor_weights += weight
constructor_name = f"more-to-do #{i+1}"
simulator.add_constructor(constructor_name, transition_name, "ToDo")
destructor_weight = max(1.0, constructor_weights) / utilization
# print(f" vanish weight={destructor_weight}")
simulator.add_immediate_transition("vanish", priority=2, weight=destructor_weight)
simulator.add_destructor("end", "Done", "vanish")
return simulator
# + jupyter={"source_hidden": true}
from contextlib import contextmanager
import time
@contextmanager
def timing(activity: str = ""):
start = time.time()
yield
elapsed = time.time() - start
print(f"Duration of [{activity}]: {elapsed:0.1f} seconds")
# -
def plot_firing_interval(data, description):
# https://stackoverflow.com/questions/15415455/plotting-probability-density-function-by-sample-with-matplotlib
from scipy.stats.kde import gaussian_kde
from scipy.stats import gamma
from numpy import linspace
with timing("fitting a gamma distribution"):
kde = gaussian_kde( data )
dist_space = linspace( min(data), max(data), 100 )
density = kde(dist_space)
gamma_params = gamma.fit(data)
print(f"gamma_params={gamma_params}")
gamma_pdf = gamma.pdf(dist_space, *gamma_params)
gamma_pdf[gamma_pdf > 1.0] = np.nan
fig, ax = plt.subplots(figsize=(20, 6))
# plt.figure
plt.grid(True)
with timing("creating a historgram"):
ax.hist(data, bins='auto', density=True, label='Empirical density')
ax.plot( dist_space, density, label='Estimated probability density (KDE)' )
ax.plot( dist_space, gamma_pdf, label='Gamma distribution fit to the data' )
ax.legend()
ax.text(0.98, 0.7, description, horizontalalignment='right', verticalalignment='center', multialignment='left',
bbox=dict(boxstyle="round,pad=1", facecolor="w", edgecolor="0.5", alpha=0.5, ),
transform=ax.transAxes)
plt.show()
# plt.figure(figsize=(20, 6))
# plt.grid(True)
# plt.loglog( dist_space, density )
# +
from multiprocessing import Pool
from collections import defaultdict
import os
def _run_one(args):
nbr_samples, num_initial_tokens, utilization, firing_distribution, loop_backs = args
simulator = create_simulator(num_initial_tokens, utilization, firing_distribution, **loop_backs)
get_transition_observations, = simulator.observe(transition_firing=nbr_samples, transitions=['start'])
simulator.simulate()
transition_observations = get_transition_observations()
return transition_observations
def save_array(file_name_prefix: str, stage: int, metric_name: str, np_array: array):
filename = f"{file_name_prefix}_{stage}_{metric_name}.array_{np_array.typecode}"
np_array.tofile(open(filename, "wb"))
print(f"Saved '{filename}'")
def load_array(file_name_prefix: str, stage: int, metric_name: str, np_array: array = None, typecode=None):
if np_array is None:
if typecode is None:
raise ValueError("np_array and typecode cannot be both None")
np_array = array(typecode)
else:
if typecode is None:
typecode = np_array.typecode
else:
if typecode != np_array.typecode
raise ValueError(f"np_array.typecode={np_array.typecode} but in the argument typecode={typecode} was povided")
filename = f"{file_name_prefix}_{stage}_{metric_name}.array_{np_array.typecode}"
file_size = os.stat(filename).st_size
if file_size % np_array.itemsize:
raise ValueError(f"The size of {filename} is not a multiple of itemsize {np_array.itemsize}")
np_array.fromfile(open(filename, "rb"), int(file_size / np_array.itemsize))
print(f"Loaded '{filename}'")
return np_array
def merge_results(metric_dicts, file_name_prefix):
transposed = defaultdict(list)
for metric_dict in metric_dicts:
for metric_name, value_array in metric_dict.items():
transposed[metric_name].append(value_array)
all_observations = dict()
stage = next(c)
for metric_name, value_array_list in transposed.items():
np_array = all_observations[metric_name] = np.concatenate(value_array_list)
save_array(file_name_prefix, stage, metric_name, np_array)
return all_observations
def run(nbr_samples, token_numbers, utilization, firing_distribution, loop_backs, file_name_prefix, do_concurrently=True):
print(f"===================================")
print(f"{'Parallel' if do_concurrently else 'Sequential'} run, {nbr_samples} samples")
for num_initial_tokens in token_numbers:
print(f"===================================")
if num_initial_tokens <=10:
create_simulator(num_initial_tokens, utilization, **loop_backs)
display(simulator.show())
with timing(f"the whole iteration"):
with timing(f"simulating num_initial_tokens={num_initial_tokens}"):
if do_concurrently:
with Pool() as p:
transition_observations = \
p.map(_run_one, [(int(nbr_samples/os.cpu_count()), num_initial_tokens, utilization, firing_distribution, loop_backs)
for _ in range(os.cpu_count())]
)
else:
transition_observations = [ _run_one((int(nbr_samples), num_initial_tokens, utilization, firing_distribution, loop_backs)) ]
flat_transition_observations = merge_results(transition_observations, file_name_prefix)
interval = flat_transition_observations['interval']
description = [f"Initial token count: {num_initial_tokens}"]
for branch_name, (repeat_count, weight) in loop_backs.items():
description.append(f" Multiplier in loop-back '{branch_name}': {repeat_count}")
description.append("")
description.append(f"Number of samples:{nbr_samples}")
description.append(f"Sample mean:{interval.mean():0.2f}")
description = '\n'.join(description)
print(description)
with timing(f"plotting num_initial_tokens={num_initial_tokens}"):
plot_firing_interval(interval, description)
print()
print()
# +
np_uniform = np.random.uniform
class FiringDistribution:
def __init__(self):
self.x = np_uniform(0.0, 1, 9999)
self.i = iter(self.x)
def sample(self):
try:
return next(self.i)
except StopIteration:
self.__init__()
return next(self.i)
# f = FiringDistribution().sample
# [f() for i in range(10)]
# +
from itertools import count
c = count(0)
with timing("Overall"):
for repeat_count in (0, 1,):
run(nbr_samples=100000,
token_numbers=(1000,),
utilization=0.75,
firing_distribution=FiringDistribution().sample,
loop_backs=dict(
repeat=(repeat_count, 1),
# set_back=(0, 4)
),
file_name_prefix='00',
do_concurrently=True,
)
# -
with timing("Overall"):
for repeat_count in (3, 10, 30, 100, 300, 1000):
run(nbr_samples=100000,
token_numbers=(1000,),
utilization=0.75,
firing_distribution=FiringDistribution().sample,
loop_backs=dict(
repeat=(repeat_count, 1),
# set_back=(0, 4)
),
file_name_prefix='00',
do_concurrently=True,
)
for stage in range(0, 2):
nbr_samples = 100000
firing_time = array('d')
firing_time.fromfile(open(f"00_{stage}_firing_time.array", "rb"), nbr_samples)
interval = array('d')
interval.fromfile(open(f"00_{stage}_interval.array", "rb"), nbr_samples)
transition = array('I')
transition.fromfile(open(f"00_{stage}_transition.array", "rb"), nbr_samples)
plt.figure(figsize=(20, 6))
plt.plot(firing_time)
plt.show()
plt.figure(figsize=(20, 6))
plt.plot(interval)
plt.show()
# plt.plot(transition)
| examples/01-Simple loopback.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# coding: utf-8
import sys, os
sys.path.append(os.pardir)
import numpy as np
from dataset.mnist import load_mnist
from two_layer_net import TwoLayerNet
# データの読み込み
(x_train, t_train), (x_test, t_test) = load_mnist(normalize=True, one_hot_label=True)
network = TwoLayerNet(input_size=784, hidden_size=50, output_size=10)
iters_num = 10000
train_size = x_train.shape[0]
batch_size = 100
learning_rate = 0.1
train_loss_list = []
train_acc_list = []
test_acc_list = []
iter_per_epoch = max(train_size / batch_size, 1)
for i in range(iters_num):
batch_mask = np.random.choice(train_size, batch_size)
x_batch = x_train[batch_mask]
t_batch = t_train[batch_mask]
# 勾配
#grad = network.numerical_gradient(x_batch, t_batch)
grad = network.gradient(x_batch, t_batch)
# 更新
for key in ('W1', 'b1', 'W2', 'b2'):
network.params[key] -= learning_rate * grad[key]
loss = network.loss(x_batch, t_batch)
train_loss_list.append(loss)
if i % iter_per_epoch == 0:
train_acc = network.accuracy(x_train, t_train)
test_acc = network.accuracy(x_test, t_test)
train_acc_list.append(train_acc)
test_acc_list.append(test_acc)
print(train_acc, test_acc)
| ch05/train_neuralnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from big_surprise import big_surprise
user_input = [('Notebook, The ', '1'),
('Road to Perdition ', '2'),
('Karate Kid, The ', '3'),
('Pulp Fiction ', '4'),
('Zoolander ', '4'),
('Majestic, The ', '5'),
('Notorious ', '2'),
('Monty Python and the Holy Grail ', '2'),
('Apocalypse Now ', '1')]
big_surprise(user_input)
another_user_input = [('1<NAME> ', '4'),
('Capote ', '2'),
('Jungle Book, The ', '1'),
('American in Paris, An ', '1'),
("All the President's Men ", '5'),
("Smilla's Sense of Snow ", '4'),
('Tron ', '4'),
('Dragonheart ', '3'),
('Foreign Correspondent ', '2')]
big_surprise(another_user_input)
yet_another_one = [('Escape from New York ', '2'),
('Sydney ', '3'),
('Sky Captain and the World of Tomorrow ', '5'),
('Mystic River ', '1'), ('Logan ', '4'),
('Super Size Me ', '3'),
('Grease ', '2'),
('50 First Dates ', '5'),
('<NAME> ', '1')]
big_surprise(yet_another_one)
| EDA/04_see_if_bulky_function_works.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# check out list words and the apostrophe situation in the step for loop
# check out allowing more than maxseqlen in generate text
# look into batch size
#text generation apostrpphe breaking
# +
import pandas as pd
import numpy as np
from keras.preprocessing.sequence import pad_sequences
from keras.optimizers import RMSprop
from keras import optimizers
import sys
from keras.callbacks import LambdaCallback
import random
import matplotlib.pyplot as plt
# -
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
# +
def generate_text(seed_text, numb_next_words):
output=seed_text
for i in range (numb_next_words):
words_gen = set(seed_text.split())
words_gen=list(words_gen) #create list of unique words in seed text
# for i in range (len(words_gen)): #replace all ' in seed text
# words_gen[i]=words_gen[i].replace("‘", '').replace("’", '').replace("'", '')
#create a dictionary with index and word
word_indices_gen = dict((c, i) for i, c in enumerate(words_gen, 1))
#turn sentence into a sequence of numbers
sequence=[]
for word in seed_text.split():
sequence.append(word_indices_gen[word])
sequence_padded = pad_sequences([sequence], maxlen=10, padding='pre')
# sequence_padded=sequence
#create an embedding matrix with same indices as word_index
EMBEDDING_DIM=25
total_words=len(word_indices_gen)+1
embedding_matrix = np.zeros((total_words, EMBEDDING_DIM))
for word, i in word_indices_gen.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
#create X input with embedding matrix for specific words (by their index)
gener=[]
for number in sequence_padded:
gener.append(embedding_matrix[number])
predicted=model.predict([gener], verbose=0)
predicted=sample(predicted[0])
output_word=""
for word, index in word_indices.items():
if index == predicted:
output_word = word
break
output+=" " + output_word
seed_text+=" " + output_word
seed_text=seed_text.split(' ', 1)[1]
return output
# -
def on_epoch_end(epoch, _):
# Function invoked at end of each epoch. Prints generated text.
print()
print('----- Generating text after Epoch: %d' % epoch)
start_index = random.randint(0, len(listofwords) - maxlen - 1)
for diversity in [0.5, 1.0]:
print('----- diversity:', diversity)
generated = ''
sentence = listofwords[start_index: start_index + maxlen].str.cat(sep=' ')
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
sys.stdout.write(generate_text(generated, 20))
tweet_data=pd.read_csv('../Load_Tweets/data/tweet_data.csv')
tweet_text = tweet_data['TEXT']
tweet_text_all = tweet_data['TEXT'].str.cat(sep=' ')
listofwords=pd.Series(tweet_text_all.split())
tweet_text.head()
top_words=listofwords.value_counts()
# top_words_percent= top_words/len(listofwords)
top_words.head(20).plot.bar()
# # top_words.head(50)
total_chars=len(tweet_text_all)
total_chars
total_wordz=len((tweet_text_all.split()))
total_wordz
chars = set(tweet_text_all)
words = set(tweet_text_all.split())
print ("total number of unique words", len(words))
print ("total number of unique chars", len(chars))
words=list(words)
#replace apostrophes in dictionary keys
for i in range (len(words)):
words[i]=words[i].replace("‘", '').replace("’", '').replace("'", '')
words=set(words)
len(words)
#create forward and reverse word index
word_indices = dict((c, i) for i, c in enumerate(words, 1))
indices_word = dict((i, c) for i, c in enumerate(words,1 ))
len(word_indices)
max(word_indices.values())
# +
#choose step
maxlen = 10
step = 2
sentences = []
next_words = []
next_words = []
list_words = []
sentences2 = []
for i in range (len(tweet_text)):
list_words = tweet_text.iloc[i].split()
for i in range(len( list_words)):
list_words[i]=list_words[i].replace("‘", '').replace("’", '').replace("'", '')
for i in range(0, len(list_words) - maxlen, step):
sentences2 = ' '.join(list_words[i: i + maxlen])
sentences.append(sentences2)
next_words.append((list_words[i + maxlen]))
print ('length of sentence list:', len(sentences))
print ("length of next_word list", len(next_words))
# -
sequences=[]
y=[]
for i, sentence in enumerate(sentences):
sequence=[]
for j, word in enumerate(sentence.split()):
sequence.append(word_indices[word])
sequences.append(sequence)
y.append(word_indices[next_words[i]])
sequences=np.asarray(sequences)
sequences.shape
sequences
total_words= len(word_indices)+1
total_words
# +
embeddings_index = {}
f = open('../word_embeding/glove.twitter.27B.25d.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
# -
max(word_indices.values())
len(word_indices)
EMBEDDING_DIM=25
embedding_matrix = np.zeros((total_words, EMBEDDING_DIM))
for word, i in word_indices.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
embedding_matrix.shape
X=[]
for number in sequences:
X.append(embedding_matrix[number])
X=np.asarray(X)
np.asarray(X).shape
embedding_matrix.shape
len(embedding_matrix)
len(word_indices)
len(sentences)
embedding_matrix.shape
np.asarray(y).shape
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# -
X_train_sample=X_train[0:10000]
y_train_sample=y_train[0:10000]
X_test_sample=X_test[0:1000]
y_test_sample=y_test[0:1000]
np.unique(y_train).shape
y_train[10:100]
# +
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import Embedding
from keras.layers import Dropout
from keras.layers import LSTM
from keras.layers import Input
from keras.regularizers import L1L2
from keras import regularizers
from keras import metrics
# embedding_layer= Embedding(total_words, EMBEDDING_DIM, weights=[embedding_matrix],input_length=max_seq,trainable=False)
# sequence_input = Input(shape=(max_seq,), dtype='int32')
# embedded_sequences= embedding_layer(sequence_input)
model=Sequential()
# e=Embedding(total_words, EMBEDDING_DIM, weights=[embedding_matrix],input_length=maxlen,trainable=False)
# model.add(e)
model.add(LSTM(128, input_shape=(maxlen, EMBEDDING_DIM), bias_regularizer=regularizers.l1(0.01)))
model.add(Dropout(0.2))
# model.add(LSTM(512, return_sequences=False))
# model.add(Dropout(0.1))
# model.add(Flatten())
model.add(Dense(total_words, activation="softmax"))
optimizer = RMSprop(lr=0.01)
# sgd = optimizers.SGD(lr=0.01, clipvalue=0.5)
model.compile(loss='sparse_categorical_crossentropy', metrics=['accuracy'], optimizer=optimizer)
model.summary()
# model.add(LSTM(128, input_shape=(maxlen, len(chars))))
# model.add(Dense(len(chars), activation='softmax'))
# model.compile(loss='categorical_crossentropy', optimizer=optimizer)
# +
# from keras.models import load_model
# model= load_model("../Saved_models/failed_on_99th_epoch_word_embedding")
# -
# print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
from keras.callbacks import EarlyStopping
early_stopping = EarlyStopping(monitor='val_acc', patience=3)
model.fit(X_train, y_train, validation_split=0.2, epochs=10,callbacks=[early_stopping])
loss, accuracy = model.evaluate(X_test_sample, y_test_sample, verbose=0)
print('Accuracy: %f' % (accuracy*100))
print('loss: %f' % (loss))
perplexity = np.exp2(loss)
print ('perplexity: {}'.format(perplexity))
print (generate_text("i will", 20))
# +
# model.save('../failed_on_99th_epoch_word_embedding')
# -
predictions_test=model.predict(X_test)
len(predictions_test)
| Model/Model_with_twitter_data_embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# ## Fully Connected Convolutional Neural Network
#
# In this notebook the previously used model is transformed in a Fully Fully Connected Convolutional Neural Network (FCN).
#
# At first, the model is loaded and summarized to confirm correct procedure.
# +
from os import path
import tensorflow as tf
model_path = path.join('models', 'symbol_classifier','model.h5')
old_model = tf.keras.models.load_model(model_path)
old_model.summary()
# -
# ### Conversion into a FCN
#
# Now the model is restructured into a FCN by adding an `input layer` which accepts a 2d input of arbitrary size.
#
# The model is transformed using the [functional API](https://keras.io/getting-started/functional-api-guide/), where each layer is considered to be a function which is called by providing the previous layer as input.
#
# The flatten layer was used to map the `Conv2D` layer onto the `Dense` output layer.
# By transforming the output layer into a `Conv2D` layer with 3 filters (for 3 different classes),
# we get the same result as by just scanning the model.
#
# For this the input dimensions of the flattened layer (which is practically the "output shape" of the last `Conv2D` layer of the model) are used to determine the connections between the last `Conv2D` layer and the new output layer by reshaping the weights of the former output layer (please note that biases do not need a reshape).
#
# Using the determined shape and weights a new `tf.keras.layers.Conv2D` layer is created.
# +
inputs = tf.keras.Input(shape=(None, None, 1))
hidden = old_model.layers[0](inputs)
for layer in old_model.layers[1:4]:
hidden = layer(hidden)
# Get the input dimensions of the flattened layer:
f_dim = old_model.layers[4].input_shape
# And use it to convert the next dense layer:
dense = old_model.layers[5]
out_dim = dense.get_weights()[1].shape[0]
W, b = dense.get_weights()
new_W = W.reshape((f_dim[1], f_dim[2], f_dim[3], out_dim))
outputs = tf.keras.layers.Conv2D(out_dim,
(f_dim[1], f_dim[2]),
name = dense.name,
strides = (1, 1),
activation = dense.activation,
padding = 'valid',
weights = [new_W, b])(hidden)
model = tf.keras.Model(inputs = inputs, outputs = outputs)
model.summary()
# -
# ### Loading the image
#
# As before the same image is loaded using `cv2`.
# The first advantage of FCN is that the image does not have to be cropped before being fed into the model.
# +
import cv2
from matplotlib import pyplot as plt
image_path = path.join('reports', 'sep', 'images', 'node_localization.png')
image = cv2.imread(image_path, cv2.IMREAD_GRAYSCALE)
plt.axis('off')
plt.set_cmap('gray')
plt.imshow(image)
plt.show()
# -
# ## Taking the time
#
# Since no further adjustments to the input data have to be done (except expanding the dimensions to match the expected input size), the time can be stopped.
#
# The resulting `predictions` are in a different format, since they come directly in `[batch_idx, x, y, pred]`.
# +
from time import time
image_exp = tf.expand_dims(tf.expand_dims(image, axis=-1), axis=0)
image_exp = tf.cast(image_exp / 255, tf.float32)
start = time()
predictions = model.predict(image_exp)
duration = time() - start
print("Finished after %.2f seconds." % duration)
print("\nShape of prediction:", predictions.shape)
print("\nFirst 5 predictions:")
print(predictions[0][0][:5])
# -
# ## Showing the results
#
# Another advantage of having everything in the same model and using the functional API is to be able is to be able to use `tf.keras.Lambda` layers.
#
# `Lambda` layers enable the possibility to implement functions as layers, which is useful if we want to include the drawing of bounding boxes into the model.
#
# The resulting model can be extended like this:
# +
def get_bounding_boxes_nms(predictions):
sqz = tf.squeeze
max_idx = tf.math.argmax(sqz(predictions), -1)
node_idx = tf.where(tf.equal(max_idx, 1))
base_idx = tf.where(tf.equal(max_idx, 2))
all_idx = tf.concat([node_idx, base_idx], 0)
max_val = tf.math.reduce_max(sqz(predictions), -1)
y, x = tf.split(all_idx * 4, 2, -1) # Times 4 to compensate model stride
coords = sqz(tf.stack([y, x, y + 32, x + 32], -1))
all_boxes = tf.cast(coords / 360, tf.float32)
scores = tf.gather_nd(max_val, all_idx)
eps = tf.keras.backend.epsilon()
nms_idx = tf.image.non_max_suppression(all_boxes, scores, 99, eps, 0.5)
limit = tf.cast(tf.math.count_nonzero(node_idx, 0)[0], tf.int32)
mask = tf.less(nms_idx, limit)
node_mask = tf.boolean_mask(nms_idx, mask)
base_mask = tf.boolean_mask(nms_idx,~mask)
node_boxes = tf.gather(all_boxes, node_mask)
base_boxes = tf.gather(all_boxes, base_mask)
return node_boxes, base_boxes
boxes_outputs = tf.keras.layers.Lambda(get_bounding_boxes_nms)(outputs)
boxes_model = tf.keras.Model(inputs = inputs, outputs = boxes_outputs)
node_boxes, base_boxes = boxes_model(image_exp)
image_rgb = tf.image.grayscale_to_rgb(image_exp)
image_boxes = tf.image.draw_bounding_boxes(image_rgb, [node_boxes], [[0, 1, 0]])
image_boxes = tf.image.draw_bounding_boxes(image_boxes, [base_boxes], [[1, 0, 0]])
plt.axis('off')
plt.set_cmap('hsv')
plt.imshow(tf.squeeze(image_boxes))
plt.show()
# -
# So the result is essentially the same, as it is classifying each symbol correctly.
# The duration needed for this operation is again 10 times faster than using simple crops and feeding the to the model individually.
#
# The output is easier to handle, since it already has a 2 dimensional format and does not need any processing.
#
# At last, the models are saved to be usable later on.
model.save(path.join('models', 'devel','fcn_sym_det.h5'))
| reports/sep/notebooks/2.1.2-fcn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### What are the four levels of basket permissions?
#
# 1. **Admin:** Basket admins can view all basket information, edit metadata, submit requests to update basket composition, and permission other users to the basket
# 2. **Edit:** Editors may view basket data and edit details such as name, description, etc.
# 3. **Rebalance:** Rebalance permissions enable a user to view basket data and submit and approve rebalance submissions
# 4. **View:** Viewers are able to see most basket information, but are not able to modify the basket in any way
from gs_quant.markets.baskets import Basket
from gs_quant.session import Environment, GsSession
# +
client = 'CLIENT ID'
secret = 'CLIENT SECRET'
GsSession.use(Environment.PROD, client_id=client, client_secret=secret, scopes=('read_user_profile',))
# -
basket = Basket.get('GSMBXXXX') # substitute input with any identifier for a basket
basket.entitlements.to_frame()
| gs_quant/documentation/06_baskets/examples/07_basket_permissions/0000_get_basket_permissions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/giacomogreggio/HSL-citybikes-predictor/blob/master/HSL_citybikes_predictor.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="WBjSSe0X0TE6" endofcell="--"
# ## Citybike predictor
#
# ### Elevator pitch
# Scheduling your day is important for everyone, but every day we have to face problems related to planning your itinerary. When you want to use a citybike to move from a place to another you may find yourself at an empty bike station. Could there be a way to predict the availability? A solution: an application that predicts exactly that based on the time and the weather.
#
#
# ### Data: sources, wrangling, management
# - The original purpose of the data is not compatible with our needs: the data is meant to describe bike trips/routing, not the bike availability
# -
#
# ### Data analysis: statistics, machine learning
# - We need a predicting model
# - Predictions for time series: a lot of different variables
# - Combining different data sources to base the prediction to current situation: weather, time of the day, current bike availability
#
#
# ### Communication of results: summarization & visualization
# - Finding clear and intuitive way to summarize and visualize data such that it is accessible to the user
# -
#
# ### Operationalization: creating added value, end-user point of view
# - Mobile optimated web application
# --
# + [markdown] id="4-NNZ_hDaUwh"
# ## Preprocessing the HSL-data
# + [markdown] id="d1K5-T3ygWKF"
# ### Initializing everything
# + id="johv7o6KOQVG" outputId="d25f106a-d497-4881-f392-7a755bade16a" colab={"base_uri": "https://localhost:8080/", "height": 382}
# !pip install mpld3
# + id="cWqTY9JXPUZE"
# All imports
import pandas as pd
import matplotlib.pyplot as plt
from google.colab import drive
from datetime import datetime
from pandas.tseries.offsets import MonthEnd
from sklearn.ensemble import ExtraTreesClassifier, RandomForestClassifier
import seaborn as sns
import numpy as np
#import mpld3
#from mpld3 import plugins
#mpld3.enable_notebook()
# + id="fXb_EGwVFqpg" outputId="3424257a-332c-40c2-d599-df2d7097f2bf" colab={"base_uri": "https://localhost:8080/", "height": 35}
drive.mount('/content/drive')
# + [markdown] id="CyS9UkKaNlX-"
# ### Data preprocessing functions
# + id="YF9mpSfOjdfW"
def get_station_data():
stations = pd.read_csv("/content/drive/My Drive/HSLDataset/Helsingin_ja_Espoon_kaupunkipyöräasemat.csv")
stations = stations.drop(["FID", "Nimi", "Namn", "Adress", "Kaupunki", "Stad", "Operaattor"], axis = 1)
return stations
# + id="ZP7ZNFmPNxaH"
def rate_weather(row):
score = 10
if row['cloud_amount']<2 : score = score
if row['cloud_amount']<5 and row['cloud_amount']>=2 : score = score-1
if row['cloud_amount']<8 and row['cloud_amount']>=5 : score = score-2
if row['cloud_amount']>=8 : score = score-3
if row['visibility']>=40000 : score = score
if row['visibility']>=10000 and row['visibility']<40000 : score = score-1
if row['visibility']<10000 : score = score-2
if row['rain_intensity']<=0.25 : score = score
if row['rain_intensity']<=1 and row['rain_intensity']>0.25: score = score-1
if row['rain_intensity']<=4 and row['rain_intensity']>1: score = score-2
if row['rain_intensity']<=16 and row['rain_intensity']>4: score = score-3
if row['rain_intensity']>16: score = score-4
if row['wind_speed']<=1.5: score = score
if row['wind_speed']<=3.3 and row['wind_speed']>1.5: score = score-1
if row['wind_speed']<=8 and row['wind_speed']>3.3: score = score-2
if row['wind_speed']>8: score = score-3
if score<1 : score = 1
return(score)
# + id="fzSXlXTMlqK6"
def get_weather_data():
filepath ='/content/drive/My Drive/HSLDataset/bikeweather_2019.csv'
df = pd.read_csv(filepath)
df = df.rename(
columns={
'Vuosi': 'year', 'Kk': 'month', 'Pv': 'day', 'Klo': 'time', 'Aikavyöhyke': 'timezone', 'Pilvien määrä (1/8)': 'cloud_amount',
'Ilmanpaine (msl) (hPa)': 'pressure', 'Suhteellinen kosteus (%)': 'rel_humidity', 'Sateen intensiteetti (mm/h)': 'rain_intensity',
'Lumensyvyys (cm)': 'snow_depth', 'Ilman lämpötila (degC)': 'air_temp', 'Kastepistelämpötila (degC)': 'dew-point_temp',
'Näkyvyys (m)': 'visibility', 'Tuulen suunta (deg)': 'wind_dir', 'Puuskanopeus (m/s)': 'gust_speed', 'Tuulen nopeus (m/s)': 'wind_speed'
}
)
df = df.drop(['pressure', 'rel_humidity', 'snow_depth', 'dew-point_temp', 'wind_dir', 'gust_speed'], axis=1)
df['datetime']=pd.to_datetime(df.year.astype(str)+'-'+df.month.astype(str)+'-'+df.day.astype(str)+' '+df.time.astype(str))
df['weather_rate'] = df.apply(lambda row: rate_weather(row), axis=1)
return df
# + id="Gl-W9W0PginX"
def preprocess_month(month):
path = "/content/drive/My Drive/HSLDataset/od-trips-2019/"
extension = ".csv"
filename = "2019-" + '{:02.0f}'.format(month)
full_path = path + filename + extension
data = pd.read_csv(full_path, sep = ",")
# Make time a datetime object to ease handling. Also floor to starting hour
data["Dep date"] = pd.to_datetime(data["Departure"], errors = "ignore").dt.floor(freq = "H")
data["Return date"] = pd.to_datetime(data["Return"], errors = "ignore").dt.floor(freq = "H")
# For our analysis we shouldn't need this information
data = data.drop(columns=["Covered distance (m)", "Duration (sec.)", "Departure", "Return"])
# Get the outgoing bikes per station at timeframe
outgoing = data.groupby("Departure station id")["Dep date"].value_counts()
outgoing = outgoing.sort_index()
outgoing = outgoing.rename_axis(index = {"Dep date" : "Date", "Departure station id" : "ID"})
outgoing = outgoing.rename("Outgoing")
# Get the arriving bikes per station at timeframe
arriving = data.groupby("Return station id")["Return date"].value_counts()
arriving = arriving.sort_index()
arriving = arriving.rename_axis(index = {"Return date" : "Date", "Return station id" : "ID"})
arriving = arriving.rename("Arriving")
outgoing_arriving_merge = pd.merge(outgoing, arriving, on = ["ID", "Date"], how = "outer")
outgoing_arriving_merge = outgoing_arriving_merge.fillna(0)
stations = set(outgoing_arriving_merge.index.get_level_values(0))
# We need data for ALL timeframes
first_day_of_month = "2019-" + '{:02.0f}'.format(month) + "-01 00:00:00"
last_day_of_month = pd.Timestamp("2019-" + '{:02.0f}'.format(month) + "-01 23:00:00") + MonthEnd(0)
all_dates = pd.date_range(first_day_of_month, last_day_of_month, freq = "H")
idx = pd.MultiIndex.from_product([stations, all_dates], names = ["ID", "Date"])
mega_frame_with_station_date_cartesian_product = pd.DataFrame(index = idx)
processed = pd.merge(mega_frame_with_station_date_cartesian_product, outgoing_arriving_merge, on = ["ID", "Date"], how = "left")
processed = processed.fillna(0)
processed = processed.reset_index()
# Merge with the station data from HSL
station_data = get_station_data()
processed_with_station_data = pd.merge(processed, station_data, on = "ID", how = "inner")
processed_with_station_data.to_csv("./drive/My Drive/HSLDataset/processed/" + filename + "-processed.csv")
# + id="WdXseBI7hTGS"
def process_and_write_raw_data():
# Process all the data and save them as csv-files
for month in range(4,11):
data = preprocess_month(month)
# + [markdown] id="S5Ub3j_CoszK"
# #### Merging functions
# + id="Bqa08qodOc98"
def drop_columns_weather(df):
return df.drop(columns=['year','month','day','time','timezone','cloud_amount','rain_intensity','air_temp','visibility','wind_speed'])
# + id="VusPcEpjD9Hq"
def drop_columns_bikes(df):
bikes = df.drop(columns=['Osoite','Kapasiteet','x','y'])
bikes = bikes.rename(columns={'Date':'datetime', 'Outgoing':'departures', 'Arriving':'arrivals','ID':'station_id','Name':'station_name'})
return bikes
# + id="nDauVE3DNgFC"
# Merging on datetime column: the column must be named 'datetime' in both tables
def merge_tables(df1, df2):
merged = df1.merge(
df2,
how='inner',
on='datetime'
)
return merged
# + [markdown] id="PadPXPXwow8y"
# #### Getters for data
# + id="JcDGv5K9f02p"
def get_processed_data_for_month(month):
month = '{:02.0f}'.format(month)
data = pd.read_csv("/content/drive/My Drive/HSLDataset/processed/2019-" + month + "-processed.csv")
data["Date"] = pd.to_datetime(data["Date"])
# Don't know what this is all about but I guess everything is fine-ish :DDD
data = data.drop("Unnamed: 0", axis = 1)
data["weekday"] = data["Date"].dt.weekday
return data
# + id="2f6j90WrlRS0"
def merge_all_data_and_write():
# Get the already processed data for all the months
data_april = get_processed_data_for_month(4)
data_may = get_processed_data_for_month(5)
data_june = get_processed_data_for_month(6)
data_july = get_processed_data_for_month(7)
data_august = get_processed_data_for_month(8)
data_september = get_processed_data_for_month(9)
data_october = get_processed_data_for_month(10)
# Concatanate the trip count data of the months
big_boi = pd.concat([data_april, data_may, data_june, data_july, data_august, data_september, data_october])
# Merge the weather data
weather_data = get_weather_data()
merged_megaset = merge_tables(drop_columns_bikes(big_boi),drop_columns_weather(weather_data))
merged_megaset.to_csv("./drive/My Drive/HSLDataset/processed/whole_data-processed.csv", index = False)
return merged_megaset
# + id="lImMNQLhs1zY"
def get_all_data():
data = pd.read_csv("./drive/My Drive/HSLDataset/processed/whole_data-processed.csv")
data["datetime"] =pd.to_datetime(data["datetime"])
data["month"] = data.loc[:,"datetime"].dt.month
return data
# + [markdown] id="aThMDl9mPGz0"
# ## Looking at the data
# + [markdown] id="A13dqniFpRxt"
# ### Getter functions for specific subsets of data
# + id="vBzFIEfuRFdw"
def data_of_station_for_weekdays_in_month(dataframe, station, month, weekday):
station_data = dataframe[dataframe["station_id"] == station]
station_data_for_month = station_data[(station_data["datetime"].dt.month == month) & (station_data["datetime"].dt.weekday == weekday)]
return station_data_for_month
# + [markdown] id="gP1lEm89pZhO"
# ### Test area
# + id="yMP9TdEDuQBu"
all_data = get_all_data()
# + id="9a3Sf5n3udP0"
# Useful for picking a station by name: some hot stations are most of the ones at a metro station (M) and ones near the central railway
stations = get_station_data()
# + [markdown] id="fmVwe_yjh2GI"
# ## Different data visualizations
# + id="09TH09GGQVds" outputId="95b73180-c804-4249-e711-701b0af7aed5" colab={"base_uri": "https://localhost:8080/", "height": 259}
wanted_data = data_of_station_for_weekdays_in_month(all_data, 19, 9, 6)
weekday_occurences = set(wanted_data["datetime"].dt.date)
fig, axs = plt.subplots(1, len(weekday_occurences), figsize = (30,6))
for idx,weekday_occurence in enumerate(weekday_occurences):
weekday_occurence_data = wanted_data[wanted_data["datetime"].dt.date == weekday_occurence]
axs[idx].grid(True, alpha = 0.3)
axs[idx].plot(weekday_occurence_data["datetime"].dt.hour, weekday_occurence_data["departures"], 'r', weekday_occurence_data["datetime"].dt.hour, weekday_occurence_data["arrivals"], 'b', alpha = 0.3)
axs[idx].title.set_text(weekday_occurence)
plt.show()
# + id="SEXizP2I3gAs" outputId="bf7f1345-d00a-4f36-e926-6484e65e47e3" colab={"base_uri": "https://localhost:8080/", "height": 265}
subset = all_data[(all_data["datetime"].dt.hour == 14)]
for rating in range(1,11):
rating_subset = subset[subset["weather_rate"] == rating]
plt.boxplot(rating_subset["departures"], positions=[rating])
plt.show()
# + id="etcNbN5Nw67E" outputId="3246c67a-8bc9-4900-ae5e-bb2be3db870f" colab={"base_uri": "https://localhost:8080/", "height": 286}
subset = all_data[(all_data["datetime"].dt.hour == 8)]
plt.scatter(subset["weather_rate"], subset["departures"], alpha = 0.03)
# + [markdown] id="zq2at6jkaiIi"
# ## Data analysis and visualizations
#
# Start by printing the data as a DataFrame for a better visualization.
# + id="Ex92VJUpahdZ" outputId="507333f6-7c1b-449f-a80d-66c57ae86323" colab={"base_uri": "https://localhost:8080/", "height": 442}
data = pd.DataFrame(get_all_data())
print(data.shape)
data
# + [markdown] id="4VaiIus1fT4s"
# Print for each feature the corresponding histogram to see the distribution of the data.
# + id="UHp-xdRbbtHL" outputId="4d944b16-2f56-4fe5-f76a-c0b0a6262fed" colab={"base_uri": "https://localhost:8080/", "height": 734}
data.hist(figsize=(15,10))
# + [markdown] id="D-vDmhwjfGtD"
# A good way to check the correlations among columns is by visualizing the correlation matrix as heatmap.
#
# As we can see below the darker colors means that there is a strong correlation between the two variables, instead of the lighter colors that means that there is a weak correlation.
#
# In our case we notice that the departures and the arrivals are stictly correlated and also these last two features have a correlation with the weather_rate.
#
# On the other hand seems that station_id has a weak correlation with departures and arrivals.
# + id="55AR0jykfBtX" outputId="4fc66793-87bb-4836-f8e4-7840e08471b2" colab={"base_uri": "https://localhost:8080/", "height": 651}
fig, ax = plt.subplots(figsize=(11,11))
corrMatrix = data.corr()
sns.heatmap(corrMatrix, annot=True, ax=ax)
plt.show()
# + [markdown] id="ZaBBDMse1Dzk"
# ### General behaviour
#
# To analyzing the data we have to work on a subset of the entire dataset. For the first visualization of the general pattern related to the departures we chose the data of October 2019.
# We have 24 hours per day so we decided to reduce the data for the analysis to the data related to Monday (working day).
# + id="_aYqimdCtyVZ" outputId="c70bc27c-a3fd-4272-84d3-1307fa284f78" colab={"base_uri": "https://localhost:8080/", "height": 224}
# get data of all mondays of last year
monday_indexes = np.where(data['weekday'] == 0)
monday_data = data.loc[monday_indexes]
print(monday_data.shape)
monday_data.head()
# + id="NQDoGwKZnguF" outputId="4537ee06-8c3a-404d-8b79-d7524a16848f" colab={"base_uri": "https://localhost:8080/", "height": 424}
# get the data of all mondays in October 2019
october_data_m = []
for i in monday_data['datetime']:
if i.month == 10:
october_data_m.append(i)
else:
october_data_m.append(0)
october_data_m = pd.DataFrame(np.array(october_data_m), columns=['monday_datetime'])
october_data_m
# + id="YXiWg1H74f8R" outputId="14238a2e-b4b3-4f48-d4f7-e94fe41ebafe" colab={"base_uri": "https://localhost:8080/", "height": 90}
departures_all = np.where(october_data_m['monday_datetime']!=0,monday_data['departures'],-1)
station_all = np.where(october_data_m['monday_datetime']!=0,monday_data['station_id'],-1)
weather_all = np.where(october_data_m['monday_datetime']!=0,monday_data['weather_rate'],-1)
# get the departures on monday in October
departures_monday = []
for i in departures_all:
if i != -1:
departures_monday.append(i)
print(max(departures_monday))
departures_monday = np.array(departures_monday)
print(departures_monday.shape)
# clean the monday data by removing zero values that are previously added
datetime_monday = []
for i in october_data_m['monday_datetime']:
if i != 0:
datetime_monday.append(i)
datetime_monday = np.array(datetime_monday)
datetime_monday.shape
# get the station_id that have departures on monday in October
station_monday = []
for i in station_all:
if i != -1:
station_monday.append(i)
station_monday = np.array(station_monday)
print(station_monday.shape)
# get the weather on monday in October
weather_monday = []
for i in weather_all:
if i != -1:
weather_monday.append(i)
weather_monday = np.array(weather_monday)
print(weather_monday.shape)
# + id="IGvBaPm_BRwl"
# Alternative way to get the data
all_data_october_mondays = all_data.loc[(all_data["month"] == 10) & (all_data["weekday"] == 0),["datetime", "departures"]]
all_data_october_mondays["printable_time"] = all_data_october_mondays.loc[:,"datetime"].dt.strftime('%d.%m %H:%M')
all_data_october_mondays_grouped = all_data_october_mondays.groupby("printable_time", as_index=False).mean()
# + id="r7xT7iA_FT3T" outputId="270c4f7d-61cb-4d7d-da81-3f6236da4cc8" colab={"base_uri": "https://localhost:8080/", "height": 424}
all_data_october_mondays_grouped
# + id="m37Aop0KDcYF" outputId="d02d66a5-89ec-47dc-d074-bf8936f2ae94" colab={"base_uri": "https://localhost:8080/", "height": 742}
plt.figure(figsize=(15,10))
sns.set_style("whitegrid")
sns.set_context("notebook", font_scale=0.6)
sns.barplot(all_data_october_mondays_grouped["printable_time"], all_data_october_mondays_grouped["departures"])
plt.xlabel("Monday date", fontsize=18)
plt.xticks(rotation=85)
plt.ylabel("Number of departures", fontsize=18)
plt.title("October (monday) departures from all the stations",fontsize=18)
# + id="97dHuB8oFcfo" outputId="da8cbc38-448c-4a21-9876-07edcc76d1f4" colab={"base_uri": "https://localhost:8080/", "height": 742}
plt.figure(figsize=(15,10))
sns.set_style("whitegrid")
sns.set_context("notebook", font_scale=0.6)
sns.barplot(all_data_october_mondays["printable_time"],all_data_october_mondays["departures"])
plt.xlabel("Monday date", fontsize=18)
plt.xticks(rotation=85)
plt.ylabel("Number of departures", fontsize=18)
plt.title("October (monday) departures from all the stations",fontsize=18)
# + [markdown] id="FSaBuC6C25pV"
# ### Data visualization for popular non-popular station
#
# Now we want to display the data related to one of the most popular stations and the data associated with one of these that is non-popular.
#
# Start by looking for a popular station: in this case we searched through the sum of departures and arrivals in the whole year per station and we found out the most popular station and the less popular station that are shown below.
# + id="KEney5m1kxQt" outputId="cf7ab859-418e-4263-e7ed-d1450c312d23" colab={"base_uri": "https://localhost:8080/", "height": 1000}
total_departures_per_station = {}
total_arrivals_per_station = {}
i=0
for j in data['station_name']:
if total_departures_per_station.get(j):
total_departures_per_station[j] = total_departures_per_station.get(j) + data['departures'][i]
else:
total_departures_per_station[j] = data['departures'][i]
if total_arrivals_per_station.get(j):
total_arrivals_per_station[j] = total_arrivals_per_station.get(j) + data['arrivals'][i]
else:
total_arrivals_per_station[j] = data['arrivals'][i]
i +=1
# convert dict into numpy array
total_departures_per_station = list(total_departures_per_station.items())
total_departures_per_station = np.array(total_departures_per_station)
print(total_departures_per_station.shape)
total_arrivals_per_station = list(total_arrivals_per_station.items())
total_arrivals_per_station = np.array(total_arrivals_per_station)
print(total_arrivals_per_station.shape)
total_dep_arr_per_station = {}
for i in range(len(total_arrivals_per_station)):
if total_departures_per_station[i][0] == total_arrivals_per_station[i][0]:
total_dep_arr_per_station[total_departures_per_station[i][0]] = int(float(total_departures_per_station[i][1])) + int(float(total_arrivals_per_station[i][1]))
total_dep_arr_per_station
# + id="D2C4chC5uSk_"
total_dep_arr_per_station = list(total_dep_arr_per_station.items())
# + id="nyD5-0_wvCYJ" outputId="0d34e2ab-ff87-4816-a0fb-596d9e0d0bb0" colab={"base_uri": "https://localhost:8080/", "height": 53}
total_dep_arr_per_station = sorted(total_dep_arr_per_station, key=lambda x: x[1], reverse=True)
print(total_dep_arr_per_station[0])
print(total_dep_arr_per_station[-1])
popular_s = total_dep_arr_per_station[0][0]
non_popular_s = total_dep_arr_per_station[-1][0]
# + id="luxrVwbxB3eS" outputId="a1989b25-07d3-4789-c4e1-34779c468086" colab={"base_uri": "https://localhost:8080/", "height": 460}
# create two dataframe: one for popular station data and one for non popular station data
pop_data = []
non_pop_data = []
for j in range(len(data)):
if data['station_name'][j] == popular_s:
pop_data.append(data.iloc[j])
elif data['station_name'][j] == non_popular_s:
non_pop_data.append(data.iloc[j])
pop_data = np.array(pop_data)
pop_data = pd.DataFrame(pop_data, columns=data.columns)
print(pop_data.shape)
non_pop_data = np.array(non_pop_data)
non_pop_data = pd.DataFrame(non_pop_data, columns=data.columns)
print(non_pop_data.shape)
non_pop_data
# + [markdown] id="pj2LJyyCGMUl"
# We proceed by grouping the data based on hours of the 'datetime' because we are aiming at the sum of departures and arrivals per hour.
# + id="EJtnMTEDadm0" outputId="4afa2f8d-974d-4f9c-9aa0-1fec8e8e0d5e" colab={"base_uri": "https://localhost:8080/", "height": 820}
hours = []
date = []
for i in pop_data['datetime']:
hours.append(i.time())
hours = np.array(hours)
pop_data['hours'] = hours
pop_data_grouped_per_hour = pop_data.groupby('hours', as_index=False).sum()
print(pop_data_grouped_per_hour.shape)
pop_data_grouped_per_hour.drop(columns=['station_id', 'station_name','weekday','weather_rate'], inplace=True)
pop_data_grouped_per_hour
# + [markdown] id="wCnbarCLNCDQ"
# We provided data visualization for the most popular station.
# + id="NU5bJcFziSiB" outputId="8945bbf5-6dca-4a3d-ddd7-7a036bb54834" colab={"base_uri": "https://localhost:8080/", "height": 660}
fig= plt.figure(figsize=(20,10))
# set width of bar
barWidth = 0.40
bars1 = pop_data_grouped_per_hour['departures']
bars2 = pop_data_grouped_per_hour['arrivals']
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
r3 = [x + barWidth for x in r2]
# Make the plot
plt.bar(r1, bars1, color='lime', width=barWidth, edgecolor='white', label='var1')
plt.bar(r2, bars2, color='blue', width=barWidth, edgecolor='white', label='var2')
# Add xticks on the middle of the group bars
plt.xlabel('Hours', fontweight='bold', fontsize=20)
plt.ylabel('# of Departures/Arrivals', fontweight='bold', fontsize=20)
plt.title(r'{}: number of departures and arrivals in a year per hours'.format(popular_s), fontsize=25, fontweight='bold')
plt.xticks([r + barWidth for r in range(len(bars1))], pop_data_grouped_per_hour['hours'], fontsize=15, rotation=90)
plt.yticks(fontsize=15)
plt.legend(labels=('departures', 'arrivals'), prop={'size': 15})
# + [markdown] id="13GrqvvU5-Uj"
# We provided data visualization for the less popular station.
# + id="rtRFMOj836af" outputId="1547d4b4-1b74-4324-978f-d997a7f2402f" colab={"base_uri": "https://localhost:8080/", "height": 820}
# follow same process for non popular station
hours = []
date = []
for i in non_pop_data['datetime']:
hours.append(i.time())
hours = np.array(hours)
non_pop_data['hours'] = hours
non_pop_data_grouped_per_hour = non_pop_data.groupby('hours', as_index=False).sum()
print(non_pop_data_grouped_per_hour.shape)
non_pop_data_grouped_per_hour.drop(columns=['station_id', 'station_name','weekday','weather_rate'], inplace=True)
non_pop_data_grouped_per_hour
# + id="F2THXT2f5pjQ" outputId="91e1bca7-8c38-4d3b-85af-279611b6df38" colab={"base_uri": "https://localhost:8080/", "height": 675}
fig= plt.figure(figsize=(20,10))
# set width of bar
barWidth = 0.40
bars1 = non_pop_data_grouped_per_hour['departures']
bars2 = non_pop_data_grouped_per_hour['arrivals']
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
r3 = [x + barWidth for x in r2]
# Make the plot
plt.bar(r1, bars1, color='lime', width=barWidth, edgecolor='white', label='var1')
plt.bar(r2, bars2, color='blue', width=barWidth, edgecolor='white', label='var2')
# Add xticks on the middle of the group bars
plt.xlabel('Hours', fontweight='bold', fontsize=20)
plt.ylabel('# of Departures/Arrivals', fontweight='bold', fontsize=20)
plt.title(r'{}: number of departures and arrivals in a year per hours'.format(non_popular_s), fontsize=25, fontweight='bold')
plt.xticks([r + barWidth for r in range(len(bars1))], non_pop_data_grouped_per_hour['hours'], fontsize=15, rotation=90)
plt.yticks(fontsize=15)
plt.legend(labels=('departures', 'arrivals'), prop={'size': 15})
# + [markdown] id="MNFV_uynMDkP"
# ### Average number of departures and arrivals
#
# The last type of graph provided in this notebook is the average number of departures and arrivals based on: weather, hour, month, and weekday. Below you can find all the graphs.
# + id="FUqKt0-v_LYK" outputId="07920ef3-798f-44b9-8970-42c62d631716" colab={"base_uri": "https://localhost:8080/", "height": 363}
# get mean departures and arrivals based on weather in the whole year
grouped_by_weather = data.groupby('weather_rate', as_index=False).mean()
grouped_by_weather
# + id="IrR3_INIAZ2j" outputId="436bac82-8d3c-4289-f090-c1baf72260d8" colab={"base_uri": "https://localhost:8080/", "height": 629}
fig= plt.figure(figsize=(20,10))
# set width of bar
barWidth = 0.40
bars1 = grouped_by_weather['departures']
bars2 = grouped_by_weather['arrivals']
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
r3 = [x + barWidth for x in r2]
# Make the plot
plt.bar(r1, bars1, color='orange', width=barWidth, edgecolor='white', label='var1')
plt.bar(r2, bars2, color='green', width=barWidth, edgecolor='white', label='var2')
# Add xticks on the middle of the group bars
plt.xlabel('Weather_rate', fontweight='bold', fontsize=20)
plt.ylabel('Mean values of Departures/Arrivals', fontweight='bold', fontsize=20)
plt.title(r'Mean of departures and arrivals in a year per weather', fontsize=25, fontweight='bold')
plt.xticks([r + barWidth for r in range(len(bars1))], grouped_by_weather['weather_rate'], fontsize=15, rotation=90)
plt.yticks(fontsize=15)
plt.legend(labels=('departures', 'arrivals'), prop={'size': 15})
# + id="jiThB8dtDRig" outputId="fb92ac85-fb2b-4520-fcd7-acbf8d820e5e" colab={"base_uri": "https://localhost:8080/", "height": 802}
# get mean departures and arrivals based on hours in the whole year
hours_in_data = []
for i in data['datetime']:
hours_in_data.append(i.time())
hours_in_data = np.array(hours_in_data)
data['hours'] = hours_in_data
grouped_by_hours = data.groupby('hours', as_index=False).mean()
grouped_by_hours
# + id="cpKZ6r9fEUMr" outputId="a2ae6ddb-f7e1-4069-8f6d-8728145f0b0c" colab={"base_uri": "https://localhost:8080/", "height": 681}
fig= plt.figure(figsize=(20,10))
# set width of bar
barWidth = 0.40
bars1 = grouped_by_hours['departures']
bars2 = grouped_by_hours['arrivals']
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
r3 = [x + barWidth for x in r2]
# Make the plot
plt.bar(r1, bars1, color='red', width=barWidth, edgecolor='white', label='var1')
plt.bar(r2, bars2, color='blue', width=barWidth, edgecolor='white', label='var2')
# Add xticks on the middle of the group bars
plt.xlabel('Hours', fontweight='bold', fontsize=20)
plt.ylabel('Mean values of Departures/Arrivals', fontweight='bold', fontsize=20)
plt.title(r'Mean of departures and arrivals in a year per hour', fontsize=25, fontweight='bold')
plt.xticks([r + barWidth for r in range(len(bars1))], grouped_by_hours['hours'], fontsize=15, rotation=90)
plt.yticks(fontsize=15)
plt.legend(labels=('departures', 'arrivals'), prop={'size': 15})
# + id="22SkpDffE2iG" outputId="a6db2008-6e12-42a0-a268-3da1bbb6188c" colab={"base_uri": "https://localhost:8080/", "height": 269}
# get mean departures and arrivals based on month in the whole year
month_in_data = []
for i in data['datetime']:
month_in_data.append(i.month)
month_in_data = np.array(month_in_data)
data['month'] = month_in_data
grouped_by_month = data.groupby('month', as_index=False).mean()
grouped_by_month
# + id="yjH15NOyHWN0" outputId="8364286f-516f-4eb5-bb61-af0274c2174e" colab={"base_uri": "https://localhost:8080/", "height": 643}
fig= plt.figure(figsize=(20,10))
# set width of bar
barWidth = 0.7
bars1 = grouped_by_month['departures']
# Set position of bar on X axis
r1 = np.arange(len(bars1))
# Make the plot
plt.bar(r1, bars1, color='purple', width=barWidth, edgecolor='white', label='var1')
# Add xticks on the middle of the group bars
plt.xlabel('Month', fontweight='bold', fontsize=20)
plt.ylabel('Mean values of Departures/Arrivals', fontweight='bold', fontsize=20)
plt.title(r'Mean of departures (and arrivals) in a year per month', fontsize=25, fontweight='bold')
plt.xticks([r for r in range(len(bars1))], grouped_by_month['month'], fontsize=15)
plt.yticks(fontsize=15)
# + id="AbFbbJGXH2xP" outputId="df0ed055-8e87-4841-eae3-4289603a6939" colab={"base_uri": "https://localhost:8080/", "height": 269}
# get mean departures and arrivals based on weekday in the whole year
grouped_by_weekday = data.groupby('weekday', as_index=False).mean()
grouped_by_weekday
# + id="SEZphsGrIEbo" outputId="ecf905ae-07ce-49c2-c5da-221d4d1986b8" colab={"base_uri": "https://localhost:8080/", "height": 696}
fig= plt.figure(figsize=(20,10))
# set width of bar
barWidth = 0.7
bars1 = grouped_by_weekday['departures']
# Set position of bar on X axis
r1 = np.arange(len(bars1))
# Make the plot
plt.bar(r1, bars1, color='green', width=barWidth, edgecolor='white', label='var1')
# Add xticks on the middle of the group bars
plt.xlabel('Weekday', fontweight='bold', fontsize=20)
plt.ylabel('Mean values of Departures/Arrivals', fontweight='bold', fontsize=20)
plt.title(r'Mean of departures (and arrivals) in a year per weekday', fontsize=25, fontweight='bold')
plt.xticks([r for r in range(len(bars1))], ['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday','Sunday'], fontsize=15, rotation=45)
plt.yticks(fontsize=15)
# + [markdown] id="ESfktk_nShoS"
# # Machine Learning
# + id="CwtR-LqESlmh"
def not_very_smart_time_of_day_thing(x):
if (int(x) < 7):
return 0
elif (int(x) in [7,9,10,11,12,13,14,21,22]):
return 1
elif (int(x) in [8,15,18,19]):
return 2
elif (int(x) in [16,17]):
return 3
else:
return 4
# + id="ZWqEiyGySqBs"
def get_workday_column(dataframe):
return dataframe["weekday"] < 5
# + id="IXD-pXCfStZo"
def get_station_area_column(dataframe):
test = dataframe["station_id"].map('{:03.0f}'.format).str[0]
return test
# + id="47g402HQSwgW"
all_data["area"] = get_station_area_column(all_data)
# + id="WkT6IUiDSxen"
all_data["workday"] = get_workday_column(all_data)
# + id="pA9Aph_lSxSe"
all_data["time_of_day"] = time_of_day(all_data)
# + id="8MF1MoIASxIA"
all_data["difference"] = all_data["arrivals"] - all_data["departures"]
# + id="27UEDFeHS1vZ"
# One-hot encoding for hours months and weekdays
if "time_of_day" in all_data.columns:
hour_dummies = pd.get_dummies(all_data.loc[:, "time_of_day"], prefix="time_of")
else:
hour_dummies = pd.get_dummies(all_data.loc[:, "hour"], prefix="hour")
if "area" in all_data.columns:
station_dummies = pd.get_dummies(all_data.loc[:, "area"], prefix="area")
else:
station_dummies = pd.get_dummies(all_data.loc[:, "station_id"], prefix="station")
if "workday" in all_data.columns:
workday_dummies = pd.get_dummies(all_data.loc[:, "workday"], prefix="workday")
else:
workday_dummies = pd.get_dummies(all_data.loc[:, "weekday"], prefix="weekday")
month_dummies = pd.get_dummies(all_data.loc[:, "month"], prefix="month")
# + id="XVAVjpwdS5MF"
X = pd.concat([station_dummies, hour_dummies, month_dummies, workday_dummies], axis = 1)
X["weather_rate"] = all_data.loc[:,"weather_rate"]
y = all_data.loc[:,"arrivals"]
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size = 0.75)
# + id="7HR1htScS-wv"
clf = ExtraTreesClassifier(n_estimators = 30).fit(X_train, y_train)
accuracy_score(clf.predict(X_test), y_test)
| HSL_citybikes_predictor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0l5MuTNOS0ta"
# 
# + [markdown] id="_jx51zSYTXTP"
# ---
#
# ####<h2><center>**PIA: Entrenamiento de una Red Neuronal Convolusional**</center></h2>
#
#
# ---
#
#
# **<h4><center>Matería:** Inteligencia Artificial y Redes Neuronales, Martes **Hora** N4
# **Docente:** <NAME>
# ##### **<h4>Integrantes del equipo:**
# <NAME> **Matrícula:** 1615458
#
# <NAME> **Matrícula:**1723100
#
# <NAME> **Matrícula:**1742422
#
# <NAME> **Matrícula:** 1793880
#
# <NAME> **Matrícula:** 1801940</center>
#
# ###**<center>Semestre Agosto Diciembre 2021**</center>
# <center>Fecha de entrega: 21 de Noviembre 2021</center>
# + [markdown] id="4JmPc0D5g05I"
#
# Source [AgniData](https://colab.research.google.com/github/stephenleo/keras-model-selection/blob/main/keras_model_selection.ipynb):
# ```
#
# ```
#
# #**MODELO 2**🎴
#
#
# ---
#
#
# + [markdown] id="4D6DQleDhT_Q"
# ##Importamos las librerias de Keras, Tensorflow y los Modulos que usaremos
# + id="yKO1_tQhWJ9v"
#Librerias
import tensorflow as tf
import tensorflow_datasets as tfds
import pandas as pd
import matplotlib.pyplot as plt
import inspect
from tqdm import tqdm
#Inicializamos el valor de analisis de batch
batch_size = 32
# + [markdown] id="HNe7u5FKiVqc"
# ###Visualizamos los modelos preestablecidos de alta precisión que estan disponibles, en este caso que utilizaremos: [*DenseNet*](https://github.com/flyyufelix/DenseNet-Keras)
# + id="a0YNvfBsTvy8" colab={"base_uri": "https://localhost:8080/"} outputId="2ae39ca3-4aad-4bb8-e2e9-b8be21a99b25"
# List all available models
model_dictionary = {m[0]:m[1] for m in inspect.getmembers(tf.keras.applications, inspect.isfunction)}
new_model_dictionary={'DenseNet169':model_dictionary.get('DenseNet169'),
'DenseNet201':model_dictionary.get('DenseNet201'),}
print(new_model_dictionary)
# + [markdown] id="RmYIiFWhi7OH"
# ###Cargamos el dataset y previsualizamos las caracteristicas por las que esta compuesta, reducimos el tamaño del set a un 70%
# + id="ZNpPIRvPi2P2" colab={"base_uri": "https://localhost:8080/", "height": 384, "referenced_widgets": ["34aa1c9c0dd84e17954593bfe3feebe5", "a5f05f245a5c46b4b890992b4d7092c7", "c4f0bc4b757c4f17ac3afe44d8ed0ed0", "9c9640611e3e40c98e0560d30202a9a3", "e98efa7b24a3410fb54bedf361a164c8", "c8dcd32e08614c66ade367557f316453", "783ec264ecb340b0b21a9f526f61d96f", "4578e6fff7b647cca0df584ff3245b71", "acfc7aded4b04c0e8f66e98dfb3b4b4c", "<KEY>", "42a4dbb82e0f4540b35899faf9a2039f", "<KEY>", "5d2e2ae57af84e8c8638ef33664c2d17", "092a107dbc904a33b0683afc27e88c4c", "d4f948b489f544e59b1e6e8e40866efc", "28c513270ae54f7ebffe3716634960cc", "42a2346b7ed444d99d22b1710d1b8b26", "a4604b9a9bd648428fa70d3f7d549708", "399e7d6a270348ce8cf31545a88937ca", "<KEY>", "cbe015f5e9244f4c9804df46418fec9a", "<KEY>", "a5928996c1164db9865ac20600f8cedc", "<KEY>", "090401cd0ba240b195538e53a2f15544", "<KEY>", "5cd12ca22caa4734ace83be389f9a885", "43ce109e990b4d7fbc7dea791c8a4c77", "<KEY>", "0fbb850ae73149f8a539871317f22b0e", "227a0703538544939de98c407cde1940", "4cc212984eaa4733b7940ae9247f6443", "<KEY>", "4070189ed0fc431abe3ae2c953080024", "<KEY>", "c5aec17244334713a8df2b3d4f45767a", "<KEY>", "96ff6e1524cb4ecab15f40fb0e8c7d55", "<KEY>", "428ca35154d74d7f9d577eff3b358511", "<KEY>", "<KEY>", "1a276f2e1ca64097a9f35e92753f59ad", "f17adbb4316d4ada85c467c3be67e937", "1a0f65e8d66a41d791ae4d31a061f466", "80ce67a19ffc4e49a288c9758bd4718f", "<KEY>", "3ef305b53ce242629185c8126ef651b1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "0cfe74e514d34393afb966ef5e9c6e05", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1da74c520ced47258598f0d0a009e3d1", "<KEY>", "4ef3b8e6ca504fb6af930da48b3b4df1", "ed7782e3d95345f797bc62a1dbbb45cd", "883a1e60fc034ab999c4053868a0fe32", "218c2e4115c640d48182a09cb531a70f", "7c246359060a4fd4a6fe5df425c96d0d", "677679fc1f8a4d8cad7c605e57644f4a", "d0cf72f285044c2fa74df768ece12d2e", "ee2a394fa00f452cba2a4bfcd2f6e0f2", "<KEY>", "1c48881e26004d1eb28b290fb3dec082", "0da0439ca577430f8b3775fac6a7fe33", "<KEY>", "<KEY>", "3ce3499e480647ea87973bed1c767b45", "<KEY>", "<KEY>", "16b92c5e43444390ac0c6fe7bdfdd5c7"]} outputId="5a2db31b-0237-42b7-8655-6a57053693aa"
# Download the training and validation data
(train, validation), metadata = tfds.load('cifar100', split=['train[:70%]', 'train[70%:]'],
with_info=True, as_supervised=True)
# Number of training examples and labels
num_train = len(list(train))
num_validation = len(list(validation))
num_classes = len(metadata.features['label'].names)
num_iterations = int(num_train/batch_size)
# Print important info
print(f'Num train images: {num_train} \
\nNum validation images: {num_validation} \
\nNum classes: {num_classes} \
\nNum iterations per epoch: {num_iterations}')
# + [markdown] id="-_72w2LXjJE8"
# ###Pre-procesamos las imagenes para un mejor entrenamiento
# + id="Ga9TvDttjNq2"
def normalize_img(image, label, img_size):
# Resize image to the desired img_size and normalize it
# One hot encode the label
image = tf.image.resize(image, img_size)
image = tf.cast(image, tf.float32) / 255.
label = tf.one_hot(label, depth=num_classes)
return image, label
def preprocess_data(train, validation, batch_size, img_size):
# Apply the normalize_img function on all train and validation data and create batches
train_processed = train.map(lambda image, label: normalize_img(image, label, img_size))
train_processed = train_processed.batch(batch_size).repeat()
validation_processed = validation.map(lambda image, label: normalize_img(image, label, img_size))
validation_processed = validation_processed.batch(batch_size)
return train_processed, validation_processed
# Run preprocessing
train_processed_224, validation_processed_224 = preprocess_data(train, validation, batch_size, img_size=[224,224])
train_processed_331, validation_processed_331 = preprocess_data(train, validation, batch_size, img_size=[331,331])
# + [markdown] id="wVB4mwJNjQ7Z"
# ###Preparamos el entrenamiento, con los diferentes modelos prestablecidos
# + id="KfTdGVMRjUwz" colab={"base_uri": "https://localhost:8080/"} outputId="92a08c86-83ce-463c-8612-ce9235ba9882"
# Loop over each model available in Keras
model_benchmarks = {'model_name': [], 'num_model_params': [], 'validation_accuracy': []}
for model_name, model in tqdm(new_model_dictionary.items()):
# Special handling for "NASNetLarge" since it requires input images with size (331,331)
if 'NASNetLarge' in model_name:
input_shape=(331,331,3)
train_processed = train_processed_331
validation_processed = validation_processed_331
else:
input_shape=(224,224,3)
train_processed = train_processed_224
validation_processed = validation_processed_224
# load the pre-trained model with global average pooling as the last layer and freeze the model weights
pre_trained_model = model(include_top=False, pooling='avg', input_shape=input_shape)
pre_trained_model.trainable = False
# custom modifications on top of pre-trained model
clf_model = tf.keras.models.Sequential()
clf_model.add(pre_trained_model)
clf_model.add(tf.keras.layers.Dense(num_classes, activation='softmax'))
clf_model.compile(loss='categorical_crossentropy', metrics=['accuracy'])
history = clf_model.fit(train_processed, epochs=3, validation_data=validation_processed,
steps_per_epoch=num_iterations)
# Calculate all relevant metrics
model_benchmarks['model_name'].append(model_name)
model_benchmarks['num_model_params'].append(pre_trained_model.count_params())
model_benchmarks['validation_accuracy'].append(history.history['val_accuracy'][-1])
# + [markdown] id="SPBuq6eJjdTU"
# ###Graficación de resultados
# + id="Lhsa9pJNjc2L" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="97fca309-8402-4d74-b93b-4b1608e17515"
# Visualize history
# Plot history: Loss
plt.plot(history.history['val_loss'])
plt.title('Validation loss history')
plt.ylabel('Loss value')
plt.xlabel('No. epoch')
plt.show()
# Plot history: Accuracy
plt.plot(history.history['val_accuracy'])
plt.title('Validation accuracy history')
plt.ylabel('Accuracy value (%)')
plt.xlabel('No. epoch')
plt.show()
# + id="NHwL1rSjSJ0E" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="3d97df85-397a-462f-dfaf-5befe65b472c"
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
| LasActividadesVanAqui/PIA-Red Neuronal/Modelo_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# <img src="../images/aeropython_logo.png" alt="AeroPython" style="width: 300px;"/>
# + [markdown] deletable=true editable=true
# # Ejercicios de análisis de datos con pandas
#
# *Fuente: https://github.com/PyDataMadrid2016/Conference-Info/tree/master/workshops_materials/20160408_1100_Pandas_for_beginners/tutorial por <NAME>, licencia MIT*
#
# En la carpeta de datos tenemos un fichero que se llama *model.txt* que contiene datos de medidas de viento: velocidad, orientación, temperatura...
# + deletable=true editable=true
# !head ../data/model.txt
# + deletable=true editable=true
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib as mpl
from IPython.display import display
# + deletable=true editable=true
model = pd.read_csv(
"../data/model.txt", delim_whitespace=True, skiprows = 3,
parse_dates = {'Timestamp': [0, 1]}, index_col = 'Timestamp')
# + deletable=true editable=true
model.head()
# + [markdown] deletable=true editable=true
# ## Ejercicios
#
# Sobre el conjunto de datos `model`:
#
# > 1. Representar la matriz `scatter` de la velocidad y orientación del viento de los primeros mil registros.
# 2. Misma matriz scatter para los 1000 registros con mayor velocidad, ordenados.
# 3. Histograma de la velocidad del viento con 36 particiones.
# 4. Histórico de la velocidad media, con los datos agrupados por años y meses.
# 5. Tabla de velocidades medias en función del año (filas) y del mes (columnas).
# 6. Gráfica con los históricos de cada año, agrupados por meses, superpuestos.
# + [markdown] deletable=true editable=true
# Representamos la matriz _scatter_ de la velocidad y orientación del viento de los primeros mil registros:
# + deletable=true editable=true
pd.tools.plotting.scatter_matrix(model.ix[:1000, 'M(m/s)':'D(deg)'])
# + [markdown] deletable=true editable=true
# Misma matriz _scatter_ para los 1000 registros con mayor velocidad:
# + deletable=true editable=true
pd.tools.plotting.scatter_matrix(
model.sort_values('M(m/s)', ascending=False).ix[:1000, 'M(m/s)':'D(deg)'])
# + deletable=true editable=true
model.loc[:, 'M(m/s)'].plot.hist(bins=np.arange(0, 35))
# + deletable=true editable=true
model['month'] = model.index.month
model['year'] = model.index.year
# + [markdown] deletable=true editable=true
# Histórico de la velocidad media:
# + deletable=true editable=true
model.groupby(by = ['year', 'month']).mean().head(24)
# + deletable=true editable=true
model.groupby(by=['year', 'month']).mean().plot(y='M(m/s)', figsize=(15, 5))
# + [markdown] deletable=true editable=true
# Media móvil de los datos agrupados por mes y año:
# + deletable=true editable=true
monthly = model.groupby(by=['year', 'month']).mean()
monthly['ma'] = monthly.loc[:, 'M(m/s)'].rolling(5, center=True).mean()
monthly.head()
# + deletable=true editable=true
monthly.loc[:, ['M(m/s)', 'ma']].plot(figsize=(15, 6))
# + deletable=true editable=true
monthly.loc[:, 'M(m/s)'].reset_index().pivot(index='year', columns='month')
# + deletable=true editable=true
monthly.loc[:, 'M(m/s)'].reset_index().pivot(
index='year', columns='month'
).T.loc['M(m/s)'].plot(
figsize=(15, 5), legend=False
)
# + [markdown] deletable=true editable=true
# ---
# <br/>
# #### <h4 align="right">¡Síguenos en Twitter!
# <br/>
# ###### <a href="https://twitter.com/AeroPython" class="twitter-follow-button" data-show-count="false">Follow @AeroPython</a> <script>!function(d,s,id){var js,fjs=d.getElementsByTagName(s)[0],p=/^http:/.test(d.location)?'http':'https';if(!d.getElementById(id)){js=d.createElement(s);js.id=id;js.src=p+'://platform.twitter.com/widgets.js';fjs.parentNode.insertBefore(js,fjs);}}(document, 'script', 'twitter-wjs');</script>
# <br/>
# ###### Este notebook ha sido realizado por: <NAME>, usando material de Kiko Correoso
# <br/>
# ##### <a rel="license" href="http://creativecommons.org/licenses/by/4.0/deed.es"><img alt="Licencia Creative Commons" style="border-width:0" src="http://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" property="dct:title">Curso AeroPython</span> por <span xmlns:cc="http://creativecommons.org/ns#" property="cc:attributionName"><NAME> y <NAME></span> se distribuye bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/deed.es">Licencia Creative Commons Atribución 4.0 Internacional</a>.
# + [markdown] deletable=true editable=true
# ---
# _Las siguientes celdas contienen configuración del Notebook_
#
# _Para visualizar y utlizar los enlaces a Twitter el notebook debe ejecutarse como [seguro](http://ipython.org/ipython-doc/dev/notebook/security.html)_
#
# File > Trusted Notebook
# + deletable=true editable=true
# Esta celda da el estilo al notebook
from IPython.core.display import HTML
css_file = '../styles/aeropython.css'
HTML(open(css_file, "r").read())
| notebooks_completos/051-Pandas-Ejercicios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
PROJECT = "PROJECT" # REPLACE WITH YOUR PROJECT ID
BUCKET = "BUCKET" # REPLACE WITH A BUCKET NAME
REGION = "us-central1" # REPLACE WITH YOUR REGION e.g. us-central1
# Import os environment variables
os.environ["PROJECT"] = PROJECT
os.environ["BUCKET"] = BUCKET
os.environ["REGION"] = REGION
os.environ["TFVERSION"] = "1.13"
# -
# ## Now write into a python module
# +
# %%writefile anomaly_detection_module/trainer/model.py
import tensorflow as tf
# Set logging to be level of INFO
tf.logging.set_verbosity(tf.logging.INFO)
# Determine CSV and label columns
number_of_tags = 5
tag_columns = ["tag_{0}".format(tag) for tag in range(0, number_of_tags)]
UNLABELED_CSV_COLUMNS = tag_columns
LABEL_COLUMN = "anomalous_sequence_flag"
LABELED_CSV_COLUMNS = UNLABELED_CSV_COLUMNS + [LABEL_COLUMN]
# Set default values for each CSV column
UNLABELED_DEFAULTS = [[""] for _ in UNLABELED_CSV_COLUMNS]
LABELED_DEFAULTS = UNLABELED_DEFAULTS + [[0.0]]
# Input function functions
def split_and_convert_string(string_tensor):
"""Splits and converts string tensor into dense float tensor.
Given string tensor, splits string by delimiter, converts to and returns
dense float tensor.
Args:
string_tensor: tf.string tensor.
Returns:
tf.float64 tensor split along delimiter.
"""
# Split string tensor into a sparse tensor based on delimiter
split_string = tf.string_split(source=tf.expand_dims(
input=string_tensor, axis=0), delimiter=",")
# Converts the values of the sparse tensor to floats
converted_tensor = tf.string_to_number(
string_tensor=split_string.values,
out_type=tf.float64)
# Create a new sparse tensor with the new converted values,
# because the original sparse tensor values are immutable
new_sparse_tensor = tf.SparseTensor(
indices=split_string.indices,
values=converted_tensor,
dense_shape=split_string.dense_shape)
# Create a dense tensor of the float values that were converted from text csv
dense_floats = tf.sparse_tensor_to_dense(
sp_input=new_sparse_tensor, default_value=0.0)
dense_floats_vector = tf.squeeze(input=dense_floats, axis=0)
return dense_floats_vector
def convert_sequences_from_strings_to_floats(features, column_list, seq_len):
"""Converts sequences from single strings to a sequence of floats.
Given features dictionary and feature column names list, convert features
from strings to a sequence of floats.
Args:
features: Dictionary of tensors of our features as tf.strings.
column_list: List of column names of our features.
seq_len: Number of timesteps in sequence.
Returns:
Dictionary of tensors of our features as tf.float64s.
"""
for column in column_list:
features[column] = split_and_convert_string(features[column])
# Since we know the sequence length, set the shape to remove the ambiguity
features[column].set_shape([seq_len])
return features
def decode_csv(value_column, mode, seq_len, training_mode):
"""Decodes CSV file into tensors.
Given single string tensor and sequence length, returns features dictionary
of tensors and labels tensor.
Args:
value_column: tf.string tensor of shape () compromising entire line of
CSV file.
mode: The estimator ModeKeys. Can be TRAIN or EVAL.
seq_len: Number of timesteps in sequence.
training_mode: Which training mode we're in. Values are "reconstruction",
"calculate_error_distribution_statistics", and "tune_anomaly_thresholds".
Returns:
Features dictionary of tensors and labels tensor.
"""
if (mode == tf.estimator.ModeKeys.TRAIN or
(mode == tf.estimator.ModeKeys.EVAL and
training_mode != "tune_anomaly_thresholds")):
# For subset of CSV files that do NOT have labels
columns = tf.decode_csv(
records=value_column,
record_defaults=UNLABELED_DEFAULTS,
field_delim=";")
features = dict(zip(UNLABELED_CSV_COLUMNS, columns))
features = convert_sequences_from_strings_to_floats(
features=features, column_list=UNLABELED_CSV_COLUMNS, seq_len=seq_len)
return features
else:
# For subset of CSV files that DO have labels
columns = tf.decode_csv(
records=value_column,
record_defaults=LABELED_DEFAULTS,
field_delim=";")
features = dict(zip(LABELED_CSV_COLUMNS, columns))
labels = tf.cast(x=features.pop(LABEL_COLUMN), dtype=tf.float64)
features = convert_sequences_from_strings_to_floats(
features=features,
column_list=LABELED_CSV_COLUMNS[0:-1],
seq_len=seq_len)
return features, labels
def read_dataset(filename, mode, batch_size, params):
"""Reads CSV time series dataset using tf.data, doing necessary preprocessing.
Given filename, mode, batch size and other parameters, read CSV dataset using
Dataset API, apply necessary preprocessing, and return an input function to
the Estimator API.
Args:
filename: The file pattern that we want to read into our tf.data dataset.
mode: The estimator ModeKeys. Can be TRAIN or EVAL.
batch_size: Number of examples to read and combine into a single tensor.
params: Additional parameters.
Returns:
An input function.
"""
def _input_fn():
"""Wrapper input function to be used by Estimator API to get data tensors.
Returns:
Batched dataset object of dictionary of feature tensors and label tensor.
"""
# Create list of files that match pattern
file_list = tf.gfile.Glob(filename=filename)
# Create dataset from file list
dataset = tf.data.TextLineDataset(filenames=file_list) # Read text file
# Decode the CSV file into a features dictionary of tensors
dataset = dataset.map(
map_func=lambda x: decode_csv(
value_column=x,
mode=mode,
seq_len=params["seq_len"],
training_mode=params["training_mode"]))
# Determine amount of times to repeat file if we are training or evaluating
if mode == tf.estimator.ModeKeys.TRAIN:
num_epochs = None # indefinitely
else:
num_epochs = 1 # end-of-input after this
# Repeat files num_epoch times
dataset = dataset.repeat(count=num_epochs)
# Group the data into batches
dataset = dataset.batch(batch_size=batch_size)
# Determine if we should shuffle based on if we are training or evaluating
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.shuffle(buffer_size=10 * batch_size)
# Create a iterator, then pull batch of features from the example queue
batched_dataset = dataset.make_one_shot_iterator().get_next()
return batched_dataset
return _input_fn
# Dense autoencoder model functions
def dense_encoder(X, params):
"""Dense model encoder subgraph that produces latent matrix.
Given data matrix tensor X and dictionary of parameters, process through dense
model encoder subgraph and return encoder latent vector for each example in
batch.
Args:
X: tf.float64 matrix tensor of input data.
params: Dictionary of parameters.
Returns:
tf.float64 matrix tensor encoder latent vector for each example in batch.
"""
# Create the input layer to our DNN
network = X
# Add hidden layers with the given number of units/neurons per layer
for units in params["enc_dnn_hidden_units"]:
network = tf.layers.dense(
inputs=network,
units=units,
activation=tf.nn.relu)
latent_matrix = tf.layers.dense(
inputs=network,
units=params["latent_vector_size"],
activation=tf.nn.relu)
return latent_matrix
def dense_decoder(latent_matrix, orig_dims, params):
"""Dense model decoder subgraph that produces output matrix.
Given encoder latent matrix tensor, the original dimensions of the input, and
dictionary of parameters, process through dense model decoder subgraph and
return decoder output matrix.
Args:
latent_matrix: tf.float64 matrix tensor of encoder latent matrix.
orig_dims: Original dimensions of input data.
params: Dictionary of parameters.
Returns:
tf.float64 matrix tensor decoder output vector for each example in batch.
"""
# Create the input layer to our DNN
network = latent_matrix
# Add hidden layers with the given number of units/neurons per layer
for units in params["dec_dnn_hidden_units"][::-1]:
network = tf.layers.dense(
inputs=network,
units=units,
activation=tf.nn.relu)
output_matrix = tf.layers.dense(
inputs=network,
units=orig_dims,
activation=tf.nn.relu)
return output_matrix
def dense_autoencoder(X, orig_dims, params):
"""Dense model autoencoder using dense encoder and decoder networks.
Given data matrix tensor X, the original dimensions of the input, and
dictionary of parameters, process through dense model encoder and decoder
subgraphs and return reconstructed inputs as output.
Args:
X: tf.float64 matrix tensor of input data.
orig_dims: Original dimensions of input data.
params: Dictionary of parameters.
Returns:
tf.float64 matrix tensor decoder output vector for each example in batch
that is the reconstructed inputs.
"""
latent_matrix = dense_encoder(X, params)
output_matrix = dense_decoder(latent_matrix, orig_dims, params)
return output_matrix
def dense_autoencoder_model(
X, mode, params, cur_batch_size, num_feat, dummy_var):
"""Dense autoencoder to reconstruct inputs and minimize reconstruction error.
Given data matrix tensor X, the current Estimator mode, the dictionary of
parameters, current batch size, and the number of features, process through
dense model encoder and decoder subgraphs and return reconstructed inputs
as output.
Args:
X: tf.float64 matrix tensor of input data.
mode: Estimator ModeKeys. Can take values of TRAIN, EVAL, and PREDICT.
params: Dictionary of parameters.
cur_batch_size: Current batch size, could be partially filled.
num_feat: Number of features.
dummy_var: Dummy variable used to allow training mode to happen since it
requires a gradient to tie back to the graph dependency.
Returns:
loss: Reconstruction loss.
train_op: Train operation so that Estimator can correctly add to dependency
graph.
X_time: 2D tensor representation of time major input data.
X_time_recon: 3D tensor representation of time major input data.
X_feat: 2D tensor representation of feature major input data.
X_feat_recon: 3D tensor representation of feature major input data.
"""
# Reshape into 2-D tensors
# Time based
# shape = (cur_batch_size * seq_len, num_feat)
X_time = tf.reshape(
tensor=X,
shape=[cur_batch_size * params["seq_len"], num_feat])
# shape = (cur_batch_size * seq_len, num_feat)
X_time_recon = dense_autoencoder(X_time, num_feat, params)
# Features based
# shape = (cur_batch_size, num_feat, seq_len)
X_transposed = tf.transpose(a=X, perm=[0, 2, 1])
# shape = (cur_batch_size * num_feat, seq_len)
X_feat = tf.reshape(
tensor=X_transposed,
shape=[cur_batch_size * num_feat, params["seq_len"]])
# shape = (cur_batch_size * num_feat, seq_len)
X_feat_recon = dense_autoencoder(X_feat, params["seq_len"], params)
if (mode == tf.estimator.ModeKeys.TRAIN and
params["training_mode"] == "reconstruction"):
X_time_recon_3d = tf.reshape(
tensor=X_time_recon,
shape=[cur_batch_size, params["seq_len"], num_feat])
X_feat_recon_3d = tf.transpose(
a=tf.reshape(
tensor=X_feat_recon,
shape=[cur_batch_size, num_feat, params["seq_len"]]),
perm=[0, 2, 1])
X_time_recon_3d_weighted = X_time_recon_3d * params["time_loss_weight"]
X_feat_recon_3d_weighted = X_feat_recon_3d * params["feat_loss_weight"]
predictions = (X_time_recon_3d_weighted + X_feat_recon_3d_weighted) \
/ (params["time_loss_weight"] + params["feat_loss_weight"])
loss = tf.losses.mean_squared_error(labels=X, predictions=predictions)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.train.get_global_step(),
learning_rate=params["learning_rate"],
optimizer="Adam")
return loss, train_op, None, None, None, None
else:
return None, None, X_time, X_time_recon, X_feat, X_feat_recon
# LSTM Encoder-decoder Autoencoder model functions
def create_LSTM_stack(lstm_hidden_units, lstm_dropout_output_keep_probs):
"""Create LSTM stacked cells.
Given list of LSTM hidden units and list of LSTM dropout output keep
probabilities.
Args:
lstm_hidden_units: List of integers for the number of hidden units in each
layer.
lstm_dropout_output_keep_probs: List of floats for the dropout output keep
probabilities for each layer.
Returns:
MultiRNNCell object of stacked LSTM layers.
"""
# First create a list of LSTM cell objects using our list of lstm hidden
# unit sizes
lstm_cells = [tf.contrib.rnn.BasicLSTMCell(
num_units=units,
forget_bias=1.0,
state_is_tuple=True)
for units in lstm_hidden_units]
# Next apply a dropout wrapper to our stack of LSTM cells,
# in this case just on the outputs
dropout_lstm_cells = [tf.nn.rnn_cell.DropoutWrapper(
cell=lstm_cells[cell_index],
input_keep_prob=1.0,
output_keep_prob=lstm_dropout_output_keep_probs[cell_index],
state_keep_prob=1.0)
for cell_index in range(len(lstm_cells))]
# Create a stack of layers of LSTM cells
# Combines list into MultiRNNCell object
stacked_lstm_cells = tf.contrib.rnn.MultiRNNCell(
cells=dropout_lstm_cells,
state_is_tuple=True)
return stacked_lstm_cells
# The rnn_decoder function takes labels during TRAIN/EVAL
# and a start token followed by its previous predictions during PREDICT
# Starts with an initial state of the final encoder states
def rnn_decoder(dec_input, init_state, cell, infer, dnn_hidden_units, num_feat):
"""Decoder for RNN cell.
Given list of LSTM hidden units and list of LSTM dropout output keep
probabilities.
Args:
dec_input: List of tf.float64 current batch size by number of features
matrix tensors input to the decoder.
init_state: Initial state of the decoder cell. Final state from the
encoder cell.
cell:
infer:
dnn_hidden_units:
num_feat:
Returns:
outputs: List of decoder outputs of length number of timesteps of tf.float64
current batch size by number of features matrix tensors.
state: Final cell state of the decoder.
"""
# Create the decoder variable scope
with tf.variable_scope("decoder"):
# Load in our initial state from our encoder
# Tuple of final encoder c_state and h_state of final encoder layer
state = init_state
# Create an empty list to store our hidden state output for every timestep
outputs = []
# Begin with no previous output
previous_output = None
# Loop over all of our dec_input which will be seq_len long
for index, decoder_input in enumerate(dec_input):
# If there has been a previous output, we will determine the next input
if previous_output is not None:
# Create the input layer to our DNN
# shape = (cur_batch_size, lstm_hidden_units[-1])
network = previous_output
# Create our dnn variable scope
with tf.variable_scope(name_or_scope="dnn", reuse=tf.AUTO_REUSE):
# Add hidden layers with the given number of units/neurons per layer
# shape = (cur_batch_size, dnn_hidden_units[i])
for units in dnn_hidden_units:
network = tf.layers.dense(
inputs=network,
units=units,
activation=tf.nn.relu)
# Connect final hidden layer to linear layer to get the logits
# shape = (cur_batch_size, num_feat)
logits = tf.layers.dense(
inputs=network,
units=num_feat,
activation=None)
# If we are in inference then we will overwrite our next decoder_input
# with the logits we just calculated. Otherwise, we leave the decoder
# input input as it was from the enumerated list. We have to calculate
# the logits even when not using them so that the correct DNN subgraph
# will be generated here and after the encoder-decoder for both
# training and inference
if infer:
# shape = (cur_batch_size, num_feat)
decoder_input = logits
# If this isn"t our first time through the loop, just reuse(share) the
# same variables for each iteration within the current variable scope
if index > 0:
tf.get_variable_scope().reuse_variables()
# Run the decoder input through the decoder stack picking up from the
# previous state
# output_shape = (cur_batch_size, lstm_hidden_units[-1])
# state_shape = # tuple of final decoder c_state and h_state
output, state = cell(decoder_input, state)
# Append the current decoder hidden state output to the outputs list
# List seq_len long of shape = (cur_batch_size, lstm_hidden_units[-1])
outputs.append(output)
# Set the previous output to the output just calculated
# shape = (cur_batch_size, lstm_hidden_units[-1])
previous_output = output
return outputs, state
def lstm_enc_dec_autoencoder_model(
X, mode, params, cur_batch_size, num_feat, dummy_var):
"""LSTM autoencoder to reconstruct inputs and minimize reconstruction error.
Given data matrix tensor X, the current Estimator mode, the dictionary of
parameters, current batch size, and the number of features, process through
LSTM model encoder, decoder, and DNN subgraphs and return reconstructed inputs
as output.
Args:
X: tf.float64 matrix tensor of input data.
mode: Estimator ModeKeys. Can take values of TRAIN, EVAL, and PREDICT.
params: Dictionary of parameters.
cur_batch_size: Current batch size, could be partially filled.
num_feat: Number of features.
dummy_var: Dummy variable used to allow training mode to happen since it
requires a gradient to tie back to the graph dependency.
Returns:
loss: Reconstruction loss.
train_op: Train operation so that Estimator can correctly add to dependency
graph.
X_time: 2D tensor representation of time major input data.
X_time_recon: 3D tensor representation of time major input data.
X_feat: 2D tensor representation of feature major input data.
X_feat_recon: 3D tensor representation of feature major input data.
"""
# Unstack 3-D features tensor into a sequence(list) of 2-D tensors
# shape = (cur_batch_size, num_feat)
X_sequence = tf.unstack(value=X, num=params["seq_len"], axis=1)
# Since this is an autoencoder, the features are the labels.
# It often works better though to have the labels in reverse order
# shape = (cur_batch_size, seq_len, num_feat)
if params["reverse_labels_sequence"]:
Y = tf.reverse_sequence(
input=X,
seq_lengths=tf.tile(
input=tf.constant(value=[params["seq_len"]], dtype=tf.int64),
multiples=tf.expand_dims(input=cur_batch_size, axis=0)),
seq_axis=1,
batch_axis=0)
else:
Y = X # shape = (cur_batch_size, seq_len, num_feat)
##############################################################################
# Create encoder of encoder-decoder LSTM stacks
# Create our decoder now
dec_stacked_lstm_cells = create_LSTM_stack(
params["dec_lstm_hidden_units"],
params["lstm_dropout_output_keep_probs"])
# Create the encoder variable scope
with tf.variable_scope("encoder"):
# Create separate encoder cells with their own weights separate from decoder
enc_stacked_lstm_cells = create_LSTM_stack(
params["enc_lstm_hidden_units"],
params["lstm_dropout_output_keep_probs"])
# Encode the input sequence using our encoder stack of LSTMs
# enc_outputs = seq_len long of shape = (cur_batch_size, enc_lstm_hidden_units[-1])
# enc_states = tuple of final encoder c_state and h_state for each layer
_, enc_states = tf.nn.static_rnn(
cell=enc_stacked_lstm_cells,
inputs=X_sequence,
initial_state=enc_stacked_lstm_cells.zero_state(
batch_size=tf.cast(x=cur_batch_size, dtype=tf.int32),
dtype=tf.float64),
dtype=tf.float64)
# We just pass on the final c and h states of the encoder"s last layer,
# so extract that and drop the others
# LSTMStateTuple shape = (cur_batch_size, lstm_hidden_units[-1])
enc_final_states = enc_states[-1]
# Extract the c and h states from the tuple
# both have shape = (cur_batch_size, lstm_hidden_units[-1])
enc_final_c, enc_final_h = enc_final_states
# In case the decoder"s first layer's number of units is different than
# encoder's last layer's number of units, use a dense layer to map to the
# correct shape
# shape = (cur_batch_size, dec_lstm_hidden_units[0])
enc_final_c_dense = tf.layers.dense(
inputs=enc_final_c,
units=params["dec_lstm_hidden_units"][0],
activation=None)
# shape = (cur_batch_size, dec_lstm_hidden_units[0])
enc_final_h_dense = tf.layers.dense(
inputs=enc_final_h,
units=params["dec_lstm_hidden_units"][0],
activation=None)
# The decoder"s first layer"s state comes from the encoder,
# the rest of the layers" initial states are zero
dec_init_states = tuple(
[tf.contrib.rnn.LSTMStateTuple(c=enc_final_c_dense,
h=enc_final_h_dense)] + \
[tf.contrib.rnn.LSTMStateTuple(
c=tf.zeros(shape=[cur_batch_size, units], dtype=tf.float64),
h=tf.zeros(shape=[cur_batch_size, units], dtype=tf.float64))
for units in params["dec_lstm_hidden_units"][1:]])
##############################################################################
# Create decoder of encoder-decoder LSTM stacks
# Train our decoder now
# Encoder-decoders work differently during training, evaluation, and inference
# so we will have two separate subgraphs for each
if (mode == tf.estimator.ModeKeys.TRAIN and
params["training_mode"] == "reconstruction"):
# Break 3-D labels tensor into a list of 2-D tensors
# shape = (cur_batch_size, num_feat)
unstacked_labels = tf.unstack(value=Y, num=params["seq_len"], axis=1)
# Call our decoder using the labels as our inputs, the encoder final state
# as our initial state, our other LSTM stack as our cells, and inference
# set to false
dec_outputs, _ = rnn_decoder(
dec_input=unstacked_labels,
init_state=dec_init_states,
cell=dec_stacked_lstm_cells,
infer=False,
dnn_hidden_units=params["dnn_hidden_units"],
num_feat=num_feat)
else:
# Since this is inference create fake labels. The list length needs to be
# the output sequence length even though only the first element is the only
# one actually used (as our go signal)
fake_labels = [tf.zeros(shape=[cur_batch_size, num_feat], dtype=tf.float64)
for _ in range(params["seq_len"])]
# Call our decoder using fake labels as our inputs, the encoder final state
# as our initial state, our other LSTM stack as our cells, and inference
# set to true
# dec_outputs = seq_len long of shape = (cur_batch_size, dec_lstm_hidden_units[-1])
# decoder_states = tuple of final decoder c_state and h_state for each layer
dec_outputs, _ = rnn_decoder(
dec_input=fake_labels,
init_state=dec_init_states,
cell=dec_stacked_lstm_cells,
infer=True,
dnn_hidden_units=params["dnn_hidden_units"],
num_feat=num_feat)
# Stack together list of rank 2 decoder output tensors into one rank 3 tensor
# shape = (cur_batch_size, seq_len, lstm_hidden_units[-1])
stacked_dec_outputs = tf.stack(values=dec_outputs, axis=1)
# Reshape rank 3 decoder outputs into rank 2 by folding sequence length into
# batch size
# shape = (cur_batch_size * seq_len, lstm_hidden_units[-1])
reshaped_stacked_dec_outputs = tf.reshape(
tensor=stacked_dec_outputs,
shape=[cur_batch_size * params["seq_len"],
params["dec_lstm_hidden_units"][-1]])
##############################################################################
# Create the DNN structure now after the encoder-decoder LSTM stack
# Create the input layer to our DNN
# shape = (cur_batch_size * seq_len, lstm_hidden_units[-1])
network = reshaped_stacked_dec_outputs
# Reuse the same variable scope as we used within our decoder (for inference)
with tf.variable_scope(name_or_scope="dnn", reuse=tf.AUTO_REUSE):
# Add hidden layers with the given number of units/neurons per layer
for units in params["dnn_hidden_units"]:
# shape = (cur_batch_size * seq_len, dnn_hidden_units[i])
network = tf.layers.dense(
inputs=network,
units=units,
activation=tf.nn.relu)
# Connect the final hidden layer to a dense layer with no activation to
# get the logits
# shape = (cur_batch_size * seq_len, num_feat)
logits = tf.layers.dense(
inputs=network,
units=num_feat,
activation=None)
# Now that we are through the final DNN for each sequence element for
# each example in the batch, reshape the predictions to match our labels.
# shape = (cur_batch_size, seq_len, num_feat)
predictions = tf.reshape(
tensor=logits,
shape=[cur_batch_size, params["seq_len"], num_feat])
if (mode == tf.estimator.ModeKeys.TRAIN and
params["training_mode"] == "reconstruction"):
loss = tf.losses.mean_squared_error(labels=Y, predictions=predictions)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.train.get_global_step(),
learning_rate=params["learning_rate"],
optimizer="Adam")
return loss, train_op, None, None, None, None
else:
if params["reverse_labels_sequence"]:
# shape=(cur_batch_size, seq_len, num_feat)
predictions = tf.reverse_sequence(
input=predictions,
seq_lengths=tf.tile(
input=tf.constant(value=[params["seq_len"]], dtype=tf.int64),
multiples=tf.expand_dims(input=cur_batch_size, axis=0)),
seq_axis=1,
batch_axis=0)
# Reshape into 2-D tensors
# Time based
# shape = (cur_batch_size * seq_len, num_feat)
X_time = tf.reshape(
tensor=X,
shape=[cur_batch_size * params["seq_len"], num_feat])
X_time_recon = tf.reshape(
tensor=predictions,
shape=[cur_batch_size * params["seq_len"], num_feat])
# Features based
# shape = (cur_batch_size, num_feat, seq_len)
X_transposed = tf.transpose(a=X, perm=[0, 2, 1])
# shape = (cur_batch_size * num_feat, seq_len)
X_feat = tf.reshape(
tensor=X_transposed,
shape=[cur_batch_size * num_feat, params["seq_len"]])
# shape = (cur_batch_size, num_feat, seq_len)
predictions_transposed = tf.transpose(a=predictions, perm=[0, 2, 1])
# shape = (cur_batch_size * num_feat, seq_len)
X_feat_recon = tf.reshape(
tensor=predictions_transposed,
shape=[cur_batch_size * num_feat, params["seq_len"]])
return None, None, X_time, X_time_recon, X_feat, X_feat_recon
# PCA model functions
def pca_model(X, mode, params, cur_batch_size, num_feat, dummy_var):
"""PCA to reconstruct inputs and minimize reconstruction error.
Given data matrix tensor X, the current Estimator mode, the dictionary of
parameters, current batch size, and the number of features, process through
PCA model subgraph and return reconstructed inputs as output.
Args:
X: tf.float64 matrix tensor of input data.
mode: Estimator ModeKeys. Can take values of TRAIN, EVAL, and PREDICT.
params: Dictionary of parameters.
cur_batch_size: Current batch size, could be partially filled.
num_feat: Number of features.
dummy_var: Dummy variable used to allow training mode to happen since it
requires a gradient to tie back to the graph dependency.
Returns:
loss: Reconstruction loss.
train_op: Train operation so that Estimator can correctly add to dependency
graph.
X_time: 2D tensor representation of time major input data.
X_time_recon: 3D tensor representation of time major input data.
X_feat: 2D tensor representation of feature major input data.
X_feat_recon: 3D tensor representation of feature major input data.
"""
# Reshape into 2-D tensors
# Time based
# shape = (cur_batch_size * seq_len, num_feat)
X_time = tf.reshape(
tensor=X,
shape=[cur_batch_size * params["seq_len"], num_feat])
# Features based
# shape = (cur_batch_size, num_feat, seq_len)
X_transposed = tf.transpose(a=X, perm=[0, 2, 1])
# shape = (cur_batch_size * num_feat, seq_len)
X_feat = tf.reshape(
tensor=X_transposed,
shape=[cur_batch_size * num_feat, params["seq_len"]])
##############################################################################
# Variables for calculating error distribution statistics
with tf.variable_scope(name_or_scope="pca_vars", reuse=tf.AUTO_REUSE):
# Time based
pca_time_count_var = tf.get_variable(
name="pca_time_count_var",
dtype=tf.int64,
initializer=tf.zeros(shape=[], dtype=tf.int64),
trainable=False)
pca_time_mean_var = tf.get_variable(
name="pca_time_mean_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[num_feat], dtype=tf.float64),
trainable=False)
pca_time_cov_var = tf.get_variable(
name="pca_time_cov_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[num_feat, num_feat], dtype=tf.float64),
trainable=False)
pca_time_eigval_var = tf.get_variable(
name="pca_time_eigval_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[num_feat], dtype=tf.float64),
trainable=False)
pca_time_eigvec_var = tf.get_variable(
name="pca_time_eigvec_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[num_feat, num_feat], dtype=tf.float64),
trainable=False)
# Features based
pca_feat_count_var = tf.get_variable(
name="pca_feat_count_var",
dtype=tf.int64,
initializer=tf.zeros(shape=[], dtype=tf.int64),
trainable=False)
pca_feat_mean_var = tf.get_variable(
name="pca_feat_mean_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[params["seq_len"]], dtype=tf.float64),
trainable=False)
pca_feat_cov_var = tf.get_variable(
name="pca_feat_cov_var",
dtype=tf.float64,
initializer=tf.zeros(
shape=[params["seq_len"], params["seq_len"]], dtype=tf.float64),
trainable=False)
pca_feat_eigval_var = tf.get_variable(
name="pca_feat_eigval_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[params["seq_len"]], dtype=tf.float64),
trainable=False)
pca_feat_eigvec_var = tf.get_variable(
name="pca_feat_eigvec_var",
dtype=tf.float64,
initializer=tf.zeros(
shape=[params["seq_len"], params["seq_len"]], dtype=tf.float64),
trainable=False)
# 3. Loss function, training/eval ops
if (mode == tf.estimator.ModeKeys.TRAIN and
params["training_mode"] == "reconstruction"):
with tf.variable_scope(name_or_scope="pca_vars", reuse=tf.AUTO_REUSE):
# Check if batch is a singleton or not, very important for covariance math
# Time based ########################################
# shape = ()
singleton_condition = tf.equal(
x=cur_batch_size * params["seq_len"], y=1)
pca_time_cov_var, pca_time_mean_var, pca_time_count_var = tf.cond(
pred=singleton_condition,
true_fn=lambda: singleton_batch_cov_variable_updating(
params["seq_len"],
X_time,
pca_time_count_var,
pca_time_mean_var,
pca_time_cov_var),
false_fn=lambda: non_singleton_batch_cov_variable_updating(
cur_batch_size,
params["seq_len"],
X_time,
pca_time_count_var,
pca_time_mean_var,
pca_time_cov_var))
# shape = (num_feat,) & (num_feat, num_feat)
pca_time_eigval_tensor, pca_time_eigvec_tensor = tf.linalg.eigh(
tensor=pca_time_cov_var)
# Features based ########################################
# shape = ()
singleton_features_condition = tf.equal(
x=cur_batch_size * num_feat, y=1)
pca_feat_cov_var, pca_feat_mean_var, pca_feat_count_var = tf.cond(
pred=singleton_features_condition,
true_fn=lambda: singleton_batch_cov_variable_updating(
num_feat,
X_feat,
pca_feat_count_var, pca_feat_mean_var,
pca_feat_cov_var),
false_fn=lambda: non_singleton_batch_cov_variable_updating(
cur_batch_size,
num_feat,
X_feat,
pca_feat_count_var,
pca_feat_mean_var,
pca_feat_cov_var))
# shape = (seq_len,) & (seq_len, seq_len)
pca_feat_eigval_tensor, pca_feat_eigvec_tensor = tf.linalg.eigh(
tensor=pca_feat_cov_var)
# Lastly use control dependencies around loss to enforce the mahalanobis
# variables to be assigned, the control order matters, hence the separate
# contexts
with tf.control_dependencies(
control_inputs=[pca_time_cov_var, pca_feat_cov_var]):
with tf.control_dependencies(
control_inputs=[pca_time_mean_var, pca_feat_mean_var]):
with tf.control_dependencies(
control_inputs=[pca_time_count_var, pca_feat_count_var]):
with tf.control_dependencies(
control_inputs=[tf.assign(ref=pca_time_eigval_var,
value=pca_time_eigval_tensor),
tf.assign(ref=pca_time_eigvec_var,
value=pca_time_eigvec_tensor),
tf.assign(ref=pca_feat_eigval_var,
value=pca_feat_eigval_tensor),
tf.assign(ref=pca_feat_eigvec_var,
value=pca_feat_eigvec_tensor)]):
loss = tf.reduce_sum(
input_tensor=tf.zeros(
shape=(), dtype=tf.float64) * dummy_var)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.train.get_global_step(),
learning_rate=params["learning_rate"],
optimizer="SGD")
return loss, train_op, None, None, None, None
else:
# Time based
# shape = (cur_batch_size * seq_len, num_feat)
X_time = X_time - pca_time_mean_var
# shape = (cur_batch_size * seq_len, params["k_principal_components"])
X_time_projected = tf.matmul(
a=X_time,
b=pca_time_eigvec_var[:, -params["k_principal_components"]:])
# shape = (cur_batch_size * seq_len, num_feat)
X_time_recon = tf.matmul(
a=X_time_projected,
b=pca_time_eigvec_var[:, -params["k_principal_components"]:],
transpose_b=True)
# Features based
# shape = (cur_batch_size * num_feat, seq_len)
X_feat = X_feat - pca_feat_mean_var
# shape = (cur_batch_size * num_feat, params["k_principal_components"])
X_feat_projected = tf.matmul(
a=X_feat,
b=pca_feat_eigvec_var[:, -params["k_principal_components"]:])
# shape = (cur_batch_size * num_feat, seq_len)
X_feat_recon = tf.matmul(
a=X_feat_projected,
b=pca_feat_eigvec_var[:, -params["k_principal_components"]:],
transpose_b=True)
return None, None, X_time, X_time_recon, X_feat, X_feat_recon
# Anomaly detection model functions
def create_mahalanobis_dist_vars(var_name, size):
"""Creates mahalanobis distance variables.
Given variable name and size, create and return mahalanobis distance variables
for count, mean, covariance, and inverse covariance.
Args:
var_name: String denoting which set of variables to create. Values are
"time" and "feat".
size: The size of the variable, either sequence length or number of
features.
Returns:
Mahalanobis distance variables for count, mean, covariance, and inverse
covariance.
"""
with tf.variable_scope(
name_or_scope="mahalanobis_dist_vars", reuse=tf.AUTO_REUSE):
# Time based
count_var = tf.get_variable(
name="abs_err_count_{0}_var".format(var_name),
dtype=tf.int64,
initializer=tf.zeros(shape=[], dtype=tf.int64),
trainable=False)
mean_var = tf.get_variable(
name="abs_err_mean_{0}_var".format(var_name),
dtype=tf.float64,
initializer=tf.zeros(shape=[size], dtype=tf.float64),
trainable=False)
cov_var = tf.get_variable(
name="abs_err_cov_{0}_var".format(var_name),
dtype=tf.float64,
initializer=tf.zeros(shape=[size, size], dtype=tf.float64),
trainable=False)
inv_cov_var = tf.get_variable(
name="abs_err_inv_cov_{0}_var".format(var_name),
dtype=tf.float64,
initializer=tf.zeros(shape=[size, size], dtype=tf.float64),
trainable=False)
return count_var, mean_var, cov_var, inv_cov_var
def create_both_mahalanobis_dist_vars(seq_len, num_feat):
"""Creates both time & feature major mahalanobis distance variables.
Given dimensions of inputs, create and return mahalanobis distance variables
for count, mean, covariance, and inverse covariance for both time and
feature major representations.
Args:
seq_len: Number of timesteps in sequence.
num_feat: Number of features.
Returns:
Mahalanobis distance variables for count, mean, covariance, and inverse
covariance for both time and feature major representations.
"""
# Time based
(abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var,
abs_err_inv_cov_time_var) = create_mahalanobis_dist_vars(
var_name="time", size=num_feat)
# Features based
(abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var,
abs_err_inv_cov_feat_var) = create_mahalanobis_dist_vars(
var_name="feat", size=seq_len)
return (abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var,
abs_err_inv_cov_time_var,
abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var,
abs_err_inv_cov_feat_var)
def create_confusion_matrix_thresh_vars(scope, var_name, size):
"""Creates confusion matrix threshold variables.
Given variable scope, name, and size, create and return confusion matrix
threshold variables for true positives, false negatives, false positives,
true negatives.
Args:
scope: String of variable scope name.
var_name: String denoting which set of variables to create. Values are
"time" and "feat".
size: The size of the variable, number of time/feature thresholds.
Returns:
Confusion matrix threshold variables for true positives, false negatives,
false positives, true negatives.
"""
with tf.variable_scope(
name_or_scope=scope, reuse=tf.AUTO_REUSE):
# Time based
tp_thresh_var = tf.get_variable(
name="tp_thresh_{0}_var".format(var_name),
dtype=tf.int64,
initializer=tf.zeros(
shape=size, dtype=tf.int64),
trainable=False)
fn_thresh_var = tf.get_variable(
name="fn_thresh_{0}_var".format(var_name),
dtype=tf.int64,
initializer=tf.zeros(
shape=size, dtype=tf.int64),
trainable=False)
fp_thresh_var = tf.get_variable(
name="fp_thresh_{0}_var".format(var_name),
dtype=tf.int64,
initializer=tf.zeros(
shape=size, dtype=tf.int64),
trainable=False)
tn_thresh_var = tf.get_variable(
name="tn_thresh_{0}_var".format(var_name),
dtype=tf.int64,
initializer=tf.zeros(
shape=size, dtype=tf.int64),
trainable=False)
return (tp_thresh_var,
fn_thresh_var,
fp_thresh_var,
tn_thresh_var)
def create_both_confusion_matrix_thresh_vars(
scope, time_thresh_size, feat_thresh_size):
"""Creates both time & feature major confusion matrix threshold variables.
Given variable scope and sizes, create and return confusion
matrix threshold variables for true positives, false negatives, false
positives, and true negatives for both time and feature major
representations.
Args:
scope: String of variable scope name.
time_thresh_size: Variable size of number of time major thresholds.
feat_thresh_size: Variable size of number of feature major thresholds.
Returns:
Confusion matrix threshold variables for true positives, false negatives,
false positives, true negatives for both time and feature major
representations.
"""
# Time based
(tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var) = create_confusion_matrix_thresh_vars(
scope=scope, var_name="time", size=time_thresh_size)
# Features based
(tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var) = create_confusion_matrix_thresh_vars(
scope=scope, var_name="feat", size=feat_thresh_size)
return (tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var,
tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var)
# Running covariance updating functions for mahalanobis distance variables
def update_record_count(count_a, count_b):
"""Updates the running number of records processed.
Given previous running total and current batch size, return new running total.
Args:
count_a: tf.int64 scalar tensor of previous running total of records.
count_b: tf.int64 scalar tensor of current batch size.
Returns:
A tf.int64 scalar tensor of new running total of records.
"""
return count_a + count_b
# Incremental covariance updating functions for mahalanobis distance variables
def update_mean_incremental(count_a, mean_a, value_b):
"""Updates the running mean vector incrementally.
Given previous running total, running column means, and single example's
column values, return new running column means.
Args:
count_a: tf.int64 scalar tensor of previous running total of records.
mean_a: tf.float64 vector tensor of previous running column means.
value_b: tf.float64 vector tensor of single example's column values.
Returns:
A tf.float64 vector tensor of new running column means.
"""
umean_a = mean_a * tf.cast(x=count_a, dtype=tf.float64)
mean_ab_num = umean_a + tf.squeeze(input=value_b, axis=0)
mean_ab = mean_ab_num / tf.cast(x=count_a + 1, dtype=tf.float64)
return mean_ab
# This function updates the covariance matrix incrementally
def update_cov_incremental(
count_a, mean_a, cov_a, value_b, mean_ab, sample_cov):
"""Updates the running covariance matrix incrementally.
Given previous running total, running column means, running covariance matrix,
single example's column values, new running column means, and whether to use
sample covariance or not, return new running covariance matrix.
Args:
count_a: tf.int64 scalar tensor of previous running total of records.
mean_a: tf.float64 vector tensor of previous running column means.
cov_a: tf.float64 matrix tensor of previous running covariance matrix.
value_b: tf.float64 vector tensor of single example's column values.
mean_ab: tf.float64 vector tensor of new running column means.
sample_cov: Bool flag on whether sample or population covariance is used.
Returns:
A tf.float64 matrix tensor of new covariance matrix.
"""
mean_diff = tf.matmul(
a=value_b - mean_a, b=value_b - mean_ab, transpose_a=True)
if sample_cov:
ucov_a = cov_a * tf.cast(x=count_a - 1, dtype=tf.float64)
cov_ab = (ucov_a + mean_diff) / tf.cast(x=count_a, dtype=tf.float64)
else:
ucov_a = cov_a * tf.cast(x=count_a, dtype=tf.float64)
cov_ab = (ucov_a + mean_diff) / tf.cast(x=count_a + 1, dtype=tf.float64)
return cov_ab
def singleton_batch_cov_variable_updating(
inner_size, X, count_variable, mean_variable, cov_variable):
"""Updates mahalanobis variables incrementally when number_of_rows equals 1.
Given the inner size of the matrix, the data vector X, the variable tracking
running record counts, the variable tracking running column means, and the
variable tracking running covariance matrix, returns updated running
covariance matrix, running column means, and running record count variables.
Args:
inner_size: Inner size of matrix X.
X: tf.float64 matrix tensor of input data.
count_variable: tf.int64 scalar variable tracking running record counts.
mean_variable: tf.float64 vector variable tracking running column means.
cov_variable: tf.float64 matrix variable tracking running covariance matrix.
Returns:
Updated running covariance matrix, running column means, and running record
count variables.
"""
# Calculate new combined mean for incremental covariance matrix calculation
# time_shape = (num_feat,), features_shape = (seq_len,)
mean_ab = update_mean_incremental(
count_a=count_variable, mean_a=mean_variable, value_b=X)
# Update running variables from single example
# time_shape = (), features_shape = ()
count_tensor = update_record_count(count_a=count_variable, count_b=1)
# time_shape = (num_feat,), features_shape = (seq_len,)
mean_tensor = mean_ab
# Check if inner dimension is greater than 1 to calculate covariance matrix
if inner_size == 1:
cov_tensor = tf.zeros_like(tensor=cov_variable, dtype=tf.float64)
else:
# time_shape = (num_feat, num_feat)
# features_shape = (seq_len, seq_len)
cov_tensor = update_cov_incremental(
count_a=count_variable,
mean_a=mean_variable,
cov_a=cov_variable,
value_b=X,
mean_ab=mean_ab,
sample_cov=True)
# Assign values to variables, use control dependencies around return to
# enforce the mahalanobis variables to be assigned, the control order matters,
# hence the separate contexts
with tf.control_dependencies(
control_inputs=[tf.assign(ref=cov_variable, value=cov_tensor)]):
with tf.control_dependencies(
control_inputs=[tf.assign(ref=mean_variable, value=mean_tensor)]):
with tf.control_dependencies(
control_inputs=[tf.assign(ref=count_variable, value=count_tensor)]):
return (tf.identity(input=cov_variable),
tf.identity(input=mean_variable),
tf.identity(input=count_variable))
# Batch covariance updating functions for mahalanobis distance variables
def update_mean_batch(count_a, mean_a, count_b, mean_b):
"""Updates the running mean vector with a batch of data.
Given previous running total, running column means, current batch size, and
batch's column means, return new running column means.
Args:
count_a: tf.int64 scalar tensor of previous running total of records.
mean_a: tf.float64 vector tensor of previous running column means.
count_b: tf.int64 scalar tensor of current batch size.
mean_b: tf.float64 vector tensor of batch's column means.
Returns:
A tf.float64 vector tensor of new running column means.
"""
sum_a = mean_a * tf.cast(x=count_a, dtype=tf.float64)
sum_b = mean_b * tf.cast(x=count_b, dtype=tf.float64)
mean_ab = (sum_a + sum_b) / tf.cast(x=count_a + count_b, dtype=tf.float64)
return mean_ab
def update_cov_batch(
count_a, mean_a, cov_a, count_b, mean_b, cov_b, sample_cov):
"""Updates the running covariance matrix with batch of data.
Given previous running total, running column means, running covariance matrix,
current batch size, batch's column means, batch's covariance matrix, and
whether to use sample covariance or not, return new running covariance matrix.
Args:
count_a: tf.int64 scalar tensor of previous running total of records.
mean_a: tf.float64 vector tensor of previous running column means.
cov_a: tf.float64 matrix tensor of previous running covariance matrix.
count_b: tf.int64 scalar tensor of current batch size.
mean_b: tf.float64 vector tensor of batch's column means.
cov_b: tf.float64 matrix tensor of batch's covariance matrix.
sample_cov: Bool flag on whether sample or population covariance is used.
Returns:
A tf.float64 matrix tensor of new running covariance matrix.
"""
mean_diff = tf.expand_dims(input=mean_a - mean_b, axis=0)
if sample_cov:
ucov_a = cov_a * tf.cast(x=count_a - 1, dtype=tf.float64)
ucov_b = cov_b * tf.cast(x=count_b - 1, dtype=tf.float64)
den = tf.cast(x=count_a + count_b - 1, dtype=tf.float64)
else:
ucov_a = cov_a * tf.cast(x=count_a, dtype=tf.float64)
ucov_b = cov_b * tf.cast(x=count_b, dtype=tf.float64)
den = tf.cast(x=count_a + count_b, dtype=tf.float64)
mean_diff = tf.matmul(a=mean_diff, b=mean_diff, transpose_a=True)
mean_scaling_num = tf.cast(x=count_a * count_b, dtype=tf.float64)
mean_scaling_den = tf.cast(x=count_a + count_b, dtype=tf.float64)
mean_scaling = mean_scaling_num / mean_scaling_den
cov_ab = (ucov_a + ucov_b + mean_diff * mean_scaling) / den
return cov_ab
def non_singleton_batch_cov_variable_updating(
cur_batch_size, inner_size, X, count_variable, mean_variable, cov_variable):
"""Updates mahalanobis variables when number_of_rows does NOT equal 1.
Given the current batch size, inner size of the matrix, the data matrix X,
the variable tracking, running record counts, the variable tracking running
column means, and the variable tracking running covariance matrix, returns
updated running covariance matrix, running column means, and running record
count variables.
Args:
cur_batch_size: Number of examples in current batch (could be partial).
inner_size: Inner size of matrix X.
X: tf.float64 matrix tensor of input data.
count_variable: tf.int64 scalar variable tracking running record counts.
mean_variable: tf.float64 vector variable tracking running column means.
cov_variable: tf.float64 matrix variable tracking running covariance matrix.
Returns:
Updated running covariance matrix, running column means, and running record
count variables.
"""
# Find statistics of batch
number_of_rows = cur_batch_size * inner_size
# time_shape = (num_feat,), features_shape = (seq_len,)
X_mean = tf.reduce_mean(input_tensor=X, axis=0)
# time_shape = (cur_batch_size * seq_len, num_feat)
# features_shape = (cur_batch_size * num_feat, seq_len)
X_centered = X - X_mean
if inner_size > 1:
# time_shape = (num_feat, num_feat)
# features_shape = (seq_len, seq_len)
X_cov = tf.matmul(
a=X_centered,
b=X_centered,
transpose_a=True) / tf.cast(x=number_of_rows - 1, dtype=tf.float64)
# Update running variables from batch statistics
# time_shape = (), features_shape = ()
count_tensor = update_record_count(
count_a=count_variable, count_b=number_of_rows)
# time_shape = (num_feat,), features_shape = (seq_len,)
mean_tensor = update_mean_batch(
count_a=count_variable,
mean_a=mean_variable,
count_b=number_of_rows,
mean_b=X_mean)
# Check if inner dimension is greater than 1 to calculate covariance matrix
if inner_size == 1:
cov_tensor = tf.zeros_like(tensor=cov_variable, dtype=tf.float64)
else:
# time_shape = (num_feat, num_feat)
# features_shape = (seq_len, seq_len)
cov_tensor = update_cov_batch(
count_a=count_variable,
mean_a=mean_variable,
cov_a=cov_variable,
count_b=number_of_rows,
mean_b=X_mean,
cov_b=X_cov,
sample_cov=True)
# Assign values to variables, use control dependencies around return to
# enforce the mahalanobis variables to be assigned, the control order matters,
# hence the separate contexts
with tf.control_dependencies(
control_inputs=[tf.assign(ref=cov_variable, value=cov_tensor)]):
with tf.control_dependencies(
control_inputs=[tf.assign(ref=mean_variable, value=mean_tensor)]):
with tf.control_dependencies(
control_inputs=[tf.assign(ref=count_variable, value=count_tensor)]):
return (tf.identity(input=cov_variable),
tf.identity(input=mean_variable),
tf.identity(input=count_variable))
def mahalanobis_dist(err_vec, mean_vec, inv_cov, final_shape):
"""Calculates mahalanobis distance from MLE.
Given reconstruction error vector, mean reconstruction error vector, inverse
covariance of reconstruction error, and mahalanobis distance tensor's final
shape, return mahalanobis distance.
Args:
err_vec: tf.float64 matrix tensor of reconstruction errors.
mean_vec: tf.float64 vector variable tracking running column means of
reconstruction errors.
inv_cov: tf.float64 matrix variable tracking running covariance matrix of
reconstruction errors.
final_shape: Final shape of mahalanobis distance tensor.
Returns:
tf.float64 matrix tensor of mahalanobis distance magnitudes.
"""
# time_shape = (cur_batch_size * seq_len, num_feat)
# features_shape = (cur_batch_size * num_feat, seq_len)
err_vec_cen = err_vec - mean_vec
# time_shape = (num_feat, cur_batch_size * seq_len)
# features_shape = (seq_len, cur_batch_size * num_feat)
mahalanobis_right_product = tf.matmul(
a=inv_cov, b=err_vec_cen, transpose_b=True)
# time_shape = (cur_batch_size * seq_len, cur_batch_size * seq_len)
# features_shape = (cur_batch_size * num_feat, cur_batch_size * num_feat)
mahalanobis_dist_vectorized = tf.matmul(
a=err_vec_cen, b=mahalanobis_right_product)
# time_shape = (cur_batch_size * seq_len,)
# features_shape = (cur_batch_size * num_feat,)
mahalanobis_dist_flat = tf.diag_part(input=mahalanobis_dist_vectorized)
# time_shape = (cur_batch_size, seq_len)
# features_shape = (cur_batch_size, num_feat)
mahalanobis_dist_final_shaped = tf.reshape(
tensor=mahalanobis_dist_flat, shape=[-1, final_shape])
# time_shape = (cur_batch_size, seq_len)
# features_shape = (cur_batch_size, num_feat)
mahalanobis_dist_final_shaped_abs = tf.abs(x=mahalanobis_dist_final_shaped)
return mahalanobis_dist_final_shaped_abs
def calculate_error_distribution_statistics_training(
cur_batch_size,
num_feat,
X_time_abs_recon_err,
abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var,
abs_err_inv_cov_time_var,
X_feat_abs_recon_err,
abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var,
abs_err_inv_cov_feat_var,
params,
dummy_var):
"""Calculates error distribution statistics during training mode.
Given dimensions of inputs, reconstructed inputs' absolute errors, and
variables tracking counts, means, and covariances of error distribution,
returns loss and train_op.
Args:
cur_batch_size: Current batch size, could be partially filled.
num_feat: Number of features.
X_time_abs_recon_err: Time major reconstructed input data's absolute
reconstruction error.
abs_err_count_time_var: Time major running count of number of records.
abs_err_mean_time_var: Time major running column means of absolute error.
abs_err_cov_time_var: Time major running covariance matrix of absolute
error.
abs_err_inv_cov_time_var: Time major running inverse covariance matrix of
absolute error.
X_feat_abs_recon_err: Feature major reconstructed input data's absolute
reconstruction error.
abs_err_count_feat_var: Feature major running count of number of records.
abs_err_mean_feat_var: Feature major running column means of absolute error.
abs_err_cov_feat_var: Feature major running covariance matrix of absolute
error.
abs_err_inv_cov_feat_var: Feature major running inverse covariance matrix of
absolute error.
params: Dictionary of parameters.
dummy_var: Dummy variable used to allow training mode to happen since it
requires a gradient to tie back to the graph dependency.
Returns:
loss: The scalar loss to tie our updates back to Estimator graph.
train_op: The train operation to tie our updates back to Estimator graph.
"""
with tf.variable_scope(
name_or_scope="mahalanobis_dist_vars", reuse=tf.AUTO_REUSE):
# Time based
singleton_time_condition = tf.equal(
x=cur_batch_size * params["seq_len"], y=1)
cov_time_var, mean_time_var, count_time_var = tf.cond(
pred=singleton_time_condition,
true_fn=lambda: singleton_batch_cov_variable_updating(
params["seq_len"],
X_time_abs_recon_err,
abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var),
false_fn=lambda: non_singleton_batch_cov_variable_updating(
cur_batch_size,
params["seq_len"],
X_time_abs_recon_err,
abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var))
# Features based
singleton_feat_condition = tf.equal(
x=cur_batch_size * num_feat, y=1)
cov_feat_var, mean_feat_var, count_feat_var = tf.cond(
pred=singleton_feat_condition,
true_fn=lambda: singleton_batch_cov_variable_updating(
num_feat,
X_feat_abs_recon_err,
abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var),
false_fn=lambda: non_singleton_batch_cov_variable_updating(
cur_batch_size,
num_feat,
X_feat_abs_recon_err,
abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var))
# Lastly use control dependencies around loss to enforce the mahalanobis
# variables to be assigned, the control order matters, hence the separate
# contexts
with tf.control_dependencies(
control_inputs=[cov_time_var, cov_feat_var]):
with tf.control_dependencies(
control_inputs=[mean_time_var, mean_feat_var]):
with tf.control_dependencies(
control_inputs=[count_time_var, count_feat_var]):
# Time based
# shape = (num_feat, num_feat)
abs_err_inv_cov_time_tensor = \
tf.matrix_inverse(input=cov_time_var + \
tf.eye(num_rows=tf.shape(input=cov_time_var)[0],
dtype=tf.float64) * params["eps"])
# Features based
# shape = (seq_len, seq_len)
abs_err_inv_cov_feat_tensor = \
tf.matrix_inverse(input=cov_feat_var + \
tf.eye(num_rows=tf.shape(input=cov_feat_var)[0],
dtype=tf.float64) * params["eps"])
with tf.control_dependencies(
control_inputs=[tf.assign(ref=abs_err_inv_cov_time_var,
value=abs_err_inv_cov_time_tensor),
tf.assign(ref=abs_err_inv_cov_feat_var,
value=abs_err_inv_cov_feat_tensor)]):
loss = tf.reduce_sum(
input_tensor=tf.zeros(shape=(), dtype=tf.float64) * dummy_var)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.train.get_global_step(),
learning_rate=params["learning_rate"],
optimizer="SGD")
return loss, train_op
def reconstruction_evaluation(X_time_orig, X_time_recon, training_mode):
"""Reconstruction loss on evaluation set.
Given time major original and reconstructed features data and the training
mode, return loss and eval_metrics_ops.
Args:
X_time_orig: Time major original features data.
X_time_recon: Time major reconstructed features data.
training_mode: Current training mode.
Returns:
loss: Scalar reconstruction loss.
eval_metric_ops: Evaluation metrics of reconstruction.
"""
loss = tf.losses.mean_squared_error(
labels=X_time_orig, predictions=X_time_recon)
eval_metric_ops = None
if training_mode == "reconstruction":
# Reconstruction eval metrics
eval_metric_ops = {
"rmse": tf.metrics.root_mean_squared_error(
labels=X_time_orig, predictions=X_time_recon),
"mae": tf.metrics.mean_absolute_error(
labels=X_time_orig, predictions=X_time_recon)
}
return loss, eval_metric_ops
def update_anom_thresh_vars(
labels_norm_mask,
labels_anom_mask,
num_thresh,
anom_thresh,
mahalanobis_dist,
tp_at_thresh_var,
fn_at_thresh_var,
fp_at_thresh_var,
tn_at_thresh_var,
mode):
"""Updates anomaly threshold variables.
Given masks for when labels are normal and anomalous, the number of anomaly
thresholds and the thresholds themselves, the mahalanobis distance, variables
for the confusion matrix, and the current Estimator mode, returns the updated
variables for the confusion matrix.
Args:
labels_norm_mask: tf.bool vector tensor that is true when label was normal.
labels_anom_mask: tf.bool vector tensor that is true when label was
anomalous.
num_thresh: Number of anomaly thresholds to try in parallel grid search.
anom_thresh: tf.float64 vector tensor of grid of anomaly thresholds to try.
mahalanobis_dist: tf.float64 matrix tensor of mahalanobis distances across
batch.
tp_at_thresh_var: tf.int64 variable tracking number of true positives at
each possible anomaly threshold.
fn_at_thresh_var: tf.int64 variable tracking number of false negatives at
each possible anomaly threshold.
fp_at_thresh_var: tf.int64 variable tracking number of false positives at
each possible anomaly threshold.
tn_at_thresh_var: tf.int64 variable tracking number of true negatives at
each possible anomaly threshold.
mode: Estimator ModeKeys, can take values of TRAIN and EVAL.
Returns:
Updated confusion matrix variables.
"""
if mode == tf.estimator.ModeKeys.TRAIN:
# time_shape = (num_time_anom_thresh, cur_batch_size, seq_len)
# feat_shape = (num_feat_anom_thresh, cur_batch_size, num_feat)
mahalanobis_dist_over_thresh = tf.map_fn(
fn=lambda anom_threshold: mahalanobis_dist > anom_threshold,
elems=anom_thresh,
dtype=tf.bool)
else:
# time_shape = (cur_batch_size, seq_len)
# feat_shape = (cur_batch_size, num_feat)
mahalanobis_dist_over_thresh = mahalanobis_dist > anom_thresh
# time_shape = (num_time_anom_thresh, cur_batch_size)
# feat_shape = (num_feat_anom_thresh, cur_batch_size)
mahalanobis_dist_any_over_thresh = tf.reduce_any(
input_tensor=mahalanobis_dist_over_thresh, axis=-1)
if mode == tf.estimator.ModeKeys.EVAL:
# time_shape = (1, cur_batch_size)
# feat_shape = (1, cur_batch_size)
mahalanobis_dist_any_over_thresh = tf.expand_dims(
input=mahalanobis_dist_any_over_thresh, axis=0)
# time_shape = (num_time_anom_thresh, cur_batch_size)
# feat_shape = (num_feat_anom_thresh, cur_batch_size)
predicted_normals = tf.equal(
x=mahalanobis_dist_any_over_thresh, y=False)
# time_shape = (num_time_anom_thresh, cur_batch_size)
# feat_shape = (num_feat_anom_thresh, cur_batch_size)
predicted_anomalies = tf.equal(
x=mahalanobis_dist_any_over_thresh, y=True)
# Calculate confusion matrix of current batch
# time_shape = (num_time_anom_thresh,)
# feat_shape = (num_feat_anom_thresh,)
tp = tf.reduce_sum(
input_tensor=tf.cast(
x=tf.map_fn(
fn=lambda threshold: tf.logical_and(
x=labels_anom_mask,
y=predicted_anomalies[threshold, :]),
elems=tf.range(start=0, limit=num_thresh, dtype=tf.int64),
dtype=tf.bool),
dtype=tf.int64),
axis=1)
fn = tf.reduce_sum(
input_tensor=tf.cast(
x=tf.map_fn(
fn=lambda threshold: tf.logical_and(
x=labels_anom_mask,
y=predicted_normals[threshold, :]),
elems=tf.range(start=0, limit=num_thresh, dtype=tf.int64),
dtype=tf.bool),
dtype=tf.int64),
axis=1)
fp = tf.reduce_sum(
input_tensor=tf.cast(
x=tf.map_fn(
fn=lambda threshold: tf.logical_and(
x=labels_norm_mask,
y=predicted_anomalies[threshold, :]),
elems=tf.range(start=0, limit=num_thresh, dtype=tf.int64),
dtype=tf.bool),
dtype=tf.int64),
axis=1)
tn = tf.reduce_sum(
input_tensor=tf.cast(
x=tf.map_fn(
fn=lambda threshold: tf.logical_and(
x=labels_norm_mask,
y=predicted_normals[threshold, :]),
elems=tf.range(start=0, limit=num_thresh, dtype=tf.int64),
dtype=tf.bool),
dtype=tf.int64),
axis=1)
if mode == tf.estimator.ModeKeys.EVAL:
# shape = ()
tp = tf.squeeze(input=tp)
fn = tf.squeeze(input=fn)
fp = tf.squeeze(input=fp)
tn = tf.squeeze(input=tn)
with tf.control_dependencies(
control_inputs=[tf.assign_add(ref=tp_at_thresh_var, value=tp),
tf.assign_add(ref=fn_at_thresh_var, value=fn),
tf.assign_add(ref=fp_at_thresh_var, value=fp),
tf.assign_add(ref=tn_at_thresh_var, value=tn)]):
return (tf.identity(input=tp_at_thresh_var),
tf.identity(input=fn_at_thresh_var),
tf.identity(input=fp_at_thresh_var),
tf.identity(input=tn_at_thresh_var))
def calculate_composite_classification_metrics(tp, fn, fp, tn, f_score_beta):
"""Calculates compositive classification metrics from the confusion matrix.
Given variables for the confusion matrix and the value of beta for f-beta
score, returns accuracy, precision, recall, and f-beta score composite
metrics.
Args:
tp: tf.int64 variable tracking number of true positives at
each possible anomaly threshold.
fn: tf.int64 variable tracking number of false negatives at
each possible anomaly threshold.
fp: tf.int64 variable tracking number of false positives at
each possible anomaly threshold.
tn: tf.int64 variable tracking number of true negatives at
each possible anomaly threshold.
f_score_beta: Value of beta for f-beta score.
Returns:
Accuracy, precision, recall, and f-beta score composite metric tensors.
"""
# time_shape = (num_time_anom_thresh,)
# feat_shape = (num_feat_anom_thresh,)
acc = tf.cast(x=tp + tn, dtype=tf.float64) \
/ tf.cast(x=tp + fn + fp + tn, dtype=tf.float64)
tp_float64 = tf.cast(x=tp, dtype=tf.float64)
pre = tp_float64 / tf.cast(x=tp + fp, dtype=tf.float64)
rec = tp_float64 / tf.cast(x=tp + fn, dtype=tf.float64)
f_beta_numerator = (1.0 + f_score_beta ** 2) * (pre * rec)
f_beta_score = f_beta_numerator / (f_score_beta ** 2 * pre + rec)
return acc, pre, rec, f_beta_score
def find_best_anom_thresh(
anom_threshs, f_beta_score, user_passed_anom_thresh, anom_thresh_var):
"""Find best anomaly threshold to use for anomaly classification.
Given grid of anomaly thresholds, variables for the confusion matrix, and the
value of beta for f-beta score, returns accuracy, precision, recall, and
f-beta score composite metrics.
Args:
anom_threshs: tf.float64 vector tensor of grid of anomaly thresholds to try.
f_beta_score: tf.float64 vector tensor of f-beta scores for each anomaly
threshold.
user_passed_anom_thresh: User passed anomaly threshold that overrides
the threshold optimization.
anom_thresh_var: tf.float64 variable that stores anomaly threshold value.
Returns:
Updated variable that stores the anomaly threshold value
"""
if user_passed_anom_thresh is None:
# shape = ()
best_anom_thresh = tf.gather(
params=anom_threshs, indices=tf.argmax(input=f_beta_score, axis=0))
else:
# shape = ()
best_anom_thresh = user_passed_anom_thresh
with tf.control_dependencies(
control_inputs=[tf.assign(
ref=anom_thresh_var, value=best_anom_thresh)]):
return tf.identity(input=anom_thresh_var)
def tune_anomaly_thresholds_training(
labels_norm_mask,
labels_anom_mask,
mahalanobis_dist_time,
tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var,
time_anom_thresh_var,
mahalanobis_dist_feat,
tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var,
feat_anom_thresh_var,
params,
mode,
dummy_var):
"""Tunes anomaly thresholds during training mode.
Given label masks, mahalanobis distances, confusion matrices, and anomaly
thresholds, returns loss and train_op.
Args:
labels_norm_mask: tf.bool vector mask of labels for normals.
labels_anom_mask: tf.bool vector mask of labels for anomalies.
mahalanobis_dist_time: Mahalanobis distance, time major.
tp_thresh_time_var: tf.int64 variable to track number of true positives wrt
thresholds for time major case.
fn_thresh_time_var: tf.int64 variable to track number of false negatives wrt
thresholds for time major case.
fp_thresh_time_var: tf.int64 variable to track number of false positives wrt
thresholds for time major case.
tn_thresh_time_var: tf.int64 variable to track number of true negatives wrt
thresholds for time major case.
time_anom_thresh_var: tf.float64 variable to hold the set time anomaly
threshold.
mahalanobis_dist_feat: Mahalanobis distance, features major.
tp_thresh_feat_var: tf.int64 variable to track number of true positives wrt
thresholds for feat major case.
fn_thresh_feat_var: tf.int64 variable to track number of false negatives wrt
thresholds for feat major case.
fp_thresh_feat_var: tf.int64 variable to track number of false positives wrt
thresholds for feat major case.
tn_thresh_feat_var: tf.int64 variable to track number of true negatives wrt
thresholds for feat major case.
feat_anom_thresh_var: tf.float64 variable to hold the set feat anomaly
threshold.
params: Dictionary of parameters.
mode: Estimator ModeKeys. Can take value of only TRAIN.
dummy_var: Dummy variable used to allow training mode to happen since it
requires a gradient to tie back to the graph dependency.
Returns:
loss: The scalar loss to tie our updates back to Estimator graph.
train_op: The train operation to tie our updates back to Estimator graph.
"""
# Time based
# shape = (num_time_anom_thresh,)
time_anom_threshs = tf.linspace(
start=tf.constant(
value=params["min_time_anom_thresh"], dtype=tf.float64),
stop=tf.constant(
value=params["max_time_anom_thresh"], dtype=tf.float64),
num=params["num_time_anom_thresh"])
with tf.variable_scope(
name_or_scope="mahalanobis_dist_thresh_vars",
reuse=tf.AUTO_REUSE):
(tp_time_update_op,
fn_time_update_op,
fp_time_update_op,
tn_time_update_op) = \
update_anom_thresh_vars(
labels_norm_mask,
labels_anom_mask,
params["num_time_anom_thresh"],
time_anom_threshs,
mahalanobis_dist_time,
tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var,
mode)
# Features based
# shape = (num_feat_anom_thresh,)
feat_anom_threshs = tf.linspace(
start=tf.constant(value=params["min_feat_anom_thresh"],
dtype=tf.float64),
stop=tf.constant(value=params["max_feat_anom_thresh"],
dtype=tf.float64),
num=params["num_feat_anom_thresh"])
with tf.variable_scope(
name_or_scope="mahalanobis_dist_thresh_vars",
reuse=tf.AUTO_REUSE):
(tp_feat_update_op,
fn_feat_update_op,
fp_feat_update_op,
tn_feat_update_op) = \
update_anom_thresh_vars(
labels_norm_mask,
labels_anom_mask,
params["num_feat_anom_thresh"],
feat_anom_threshs,
mahalanobis_dist_feat,
tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var,
mode)
# Reconstruction loss on evaluation set
with tf.control_dependencies(
control_inputs=[
tp_time_update_op,
fn_time_update_op,
fp_time_update_op,
tn_time_update_op,
tp_feat_update_op,
fn_feat_update_op,
fp_feat_update_op,
tn_feat_update_op]):
# Time based
_, pre_time, rec_time, f_beta_time = \
calculate_composite_classification_metrics(
tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var,
params["f_score_beta"])
# Features based
_, pre_feat, rec_feat, f_beta_feat = \
calculate_composite_classification_metrics(
tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var,
params["f_score_beta"])
with tf.control_dependencies(
control_inputs=[pre_time, pre_feat, rec_time, rec_feat]):
with tf.control_dependencies(
control_inputs=[f_beta_time, f_beta_feat]):
# Time based
best_anom_thresh_time = find_best_anom_thresh(
time_anom_threshs,
f_beta_time,
params["time_anom_thresh"],
time_anom_thresh_var)
# Features based
best_anom_thresh_feat = find_best_anom_thresh(
feat_anom_threshs,
f_beta_feat,
params["feat_anom_thresh"],
feat_anom_thresh_var)
with tf.control_dependencies(
control_inputs=[best_anom_thresh_time,
best_anom_thresh_feat]):
loss = tf.reduce_sum(
input_tensor=tf.zeros(
shape=(), dtype=tf.float64) * dummy_var)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.train.get_global_step(),
learning_rate=params["learning_rate"],
optimizer="SGD")
return loss, train_op
def tune_anomaly_thresholds_evaluation(
labels_norm_mask,
labels_anom_mask,
time_anom_thresh_var,
mahalanobis_dist_time,
tp_thresh_eval_time_var,
fn_thresh_eval_time_var,
fp_thresh_eval_time_var,
tn_thresh_eval_time_var,
feat_anom_thresh_var,
mahalanobis_dist_feat,
tp_thresh_eval_feat_var,
fn_thresh_eval_feat_var,
fp_thresh_eval_feat_var,
tn_thresh_eval_feat_var,
params,
mode):
"""Checks tuned anomaly thresholds during evaluation mode.
Given label masks, mahalanobis distances, confusion matrices, and anomaly
thresholds, returns loss and eval_metric_ops.
Args:
labels_norm_mask: tf.bool vector mask of labels for normals.
labels_anom_mask: tf.bool vector mask of labels for anomalies.
time_anom_thresh_var: tf.float64 scalar time anomaly threshold value.
mahalanobis_dist_time: Mahalanobis distance, time major.
tp_thresh_eval_time_var: tf.int64 variable to track number of true
positives wrt thresholds for time major case for evaluation.
fn_thresh_eval_time_var: tf.int64 variable to track number of false
negatives wrt thresholds for time major case for evaluation.
fp_thresh_eval_time_var: tf.int64 variable to track number of false
positives wrt thresholds for time major case for evaluation.
tn_thresh_eval_time_var: tf.int64 variable to track number of true
negatives wrt thresholds for time major case for evaluation.
feat_anom_thresh_var: tf.float64 scalar feature anomaly threshold value.
mahalanobis_dist_feat: Mahalanobis distance, features major.
tp_thresh_eval_feat_var: tf.int64 variable to track number of true
positives wrt thresholds for feat major case for evaluation.
fn_thresh_eval_feat_var: tf.int64 variable to track number of false
negatives wrt thresholds for feat major case for evaluation.
fp_thresh_eval_feat_var: tf.int64 variable to track number of false
positives wrt thresholds for feat major case for evaluation.
tn_thresh_eval_feat_var: tf.int64 variable to track number of true
negatives wrt thresholds for feat major case for evaluation.
params: Dictionary of parameters.
mode: Estimator ModeKeys. Can take value of only EVAL.
Returns:
loss: Scalar reconstruction loss.
eval_metric_ops: Evaluation metrics of reconstruction.
"""
with tf.variable_scope(
name_or_scope="anom_thresh_eval_vars", reuse=tf.AUTO_REUSE):
# Time based
(tp_time_update_op,
fn_time_update_op,
fp_time_update_op,
tn_time_update_op) = \
update_anom_thresh_vars(
labels_norm_mask,
labels_anom_mask,
1,
time_anom_thresh_var,
mahalanobis_dist_time,
tp_thresh_eval_time_var,
fn_thresh_eval_time_var,
fp_thresh_eval_time_var,
tn_thresh_eval_time_var,
mode)
# Features based
(tp_feat_update_op,
fn_feat_update_op,
fp_feat_update_op,
tn_feat_update_op) = \
update_anom_thresh_vars(
labels_norm_mask,
labels_anom_mask,
1,
feat_anom_thresh_var,
mahalanobis_dist_feat,
tp_thresh_eval_feat_var,
fn_thresh_eval_feat_var,
fp_thresh_eval_feat_var,
tn_thresh_eval_feat_var,
mode)
with tf.variable_scope(
name_or_scope="anom_thresh_eval_vars", reuse=tf.AUTO_REUSE):
# Time based
(acc_time_update_op,
pre_time_update_op,
rec_time_update_op,
f_beta_time_update_op) = \
calculate_composite_classification_metrics(
tp_thresh_eval_time_var,
fn_thresh_eval_time_var,
fp_thresh_eval_time_var,
tn_thresh_eval_time_var,
params["f_score_beta"])
# Features based
(acc_feat_update_op,
pre_feat_update_op,
rec_feat_update_op,
f_beta_feat_update_op) = \
calculate_composite_classification_metrics(
tp_thresh_eval_feat_var,
fn_thresh_eval_feat_var,
fp_thresh_eval_feat_var,
tn_thresh_eval_feat_var,
params["f_score_beta"])
loss = tf.zeros(shape=[], dtype=tf.float64)
# Time based
acc_trues = tf.cast(
x=tp_thresh_eval_time_var + tn_thresh_eval_time_var,
dtype=tf.float64)
acc_falses = tf.cast(
x=fp_thresh_eval_time_var + fn_thresh_eval_time_var,
dtype=tf.float64)
acc_thresh_eval_time_var = acc_trues / (acc_trues + acc_falses)
tp_float = tf.cast(x=tp_thresh_eval_time_var, dtype=tf.float64)
pre_denominator = tf.cast(
x=tp_thresh_eval_time_var + fp_thresh_eval_time_var,
dtype=tf.float64)
pre_thresh_eval_time_var = tp_float / pre_denominator
rec_denominator = tf.cast(
x=tp_thresh_eval_time_var + fn_thresh_eval_time_var,
dtype=tf.float64)
rec_thresh_eval_time_var = tp_float / rec_denominator
f_beta_numerator = (1.0 + params["f_score_beta"] ** 2)
f_beta_numerator *= pre_thresh_eval_time_var
f_beta_numerator *= rec_thresh_eval_time_var
f_beta_denominator = params["f_score_beta"] ** 2
f_beta_denominator *= pre_thresh_eval_time_var
f_beta_denominator += rec_thresh_eval_time_var
f_beta_thresh_eval_time_var = f_beta_numerator / f_beta_denominator
# Features based
acc_trues = tf.cast(
x=tp_thresh_eval_feat_var + tn_thresh_eval_feat_var,
dtype=tf.float64)
acc_falses = tf.cast(
x=fp_thresh_eval_feat_var + fn_thresh_eval_feat_var,
dtype=tf.float64)
acc_thresh_eval_feat_var = acc_trues / (acc_trues + acc_falses)
tp_float = tf.cast(x=tp_thresh_eval_feat_var, dtype=tf.float64)
pre_denominator = tf.cast(
x=tp_thresh_eval_feat_var + fp_thresh_eval_feat_var,
dtype=tf.float64)
pre_thresh_eval_feat_var = tp_float / pre_denominator
rec_denominator = tf.cast(
x=tp_thresh_eval_feat_var + fn_thresh_eval_feat_var,
dtype=tf.float64)
rec_thresh_eval_feat_var = tp_float / rec_denominator
f_beta_numerator = (1.0 + params["f_score_beta"] ** 2)
f_beta_numerator *= pre_thresh_eval_feat_var
f_beta_numerator *= rec_thresh_eval_feat_var
f_beta_denominator = params["f_score_beta"] ** 2
f_beta_denominator *= pre_thresh_eval_feat_var
f_beta_denominator += rec_thresh_eval_feat_var
f_beta_thresh_eval_feat_var = f_beta_numerator / f_beta_denominator
# Anomaly detection eval metrics
eval_metric_ops = {
# Time based
"time_anom_tp": (tp_thresh_eval_time_var, tp_time_update_op),
"time_anom_fn": (fn_thresh_eval_time_var, fn_time_update_op),
"time_anom_fp": (fp_thresh_eval_time_var, fp_time_update_op),
"time_anom_tn": (tn_thresh_eval_time_var, tn_time_update_op),
"time_anom_acc": (acc_thresh_eval_time_var, acc_time_update_op),
"time_anom_pre": (pre_thresh_eval_time_var, pre_time_update_op),
"time_anom_rec": (rec_thresh_eval_time_var, rec_time_update_op),
"time_anom_f_beta": (f_beta_thresh_eval_time_var,
f_beta_time_update_op),
# Features based
"feat_anom_tp": (tp_thresh_eval_feat_var, tp_feat_update_op),
"feat_anom_fn": (fn_thresh_eval_feat_var, fn_feat_update_op),
"feat_anom_fp": (fp_thresh_eval_feat_var, fp_feat_update_op),
"feat_anom_tn": (tn_thresh_eval_feat_var, tn_feat_update_op),
"feat_anom_acc": (acc_thresh_eval_feat_var, acc_feat_update_op),
"feat_anom_pre": (pre_thresh_eval_feat_var, pre_feat_update_op),
"feat_anom_rec": (rec_thresh_eval_feat_var, rec_feat_update_op),
"feat_anom_f_beta": (f_beta_thresh_eval_feat_var,
f_beta_feat_update_op)
}
return loss, eval_metric_ops
def anomaly_detection_predictions(
cur_batch_size,
seq_len,
num_feat,
mahalanobis_dist_time,
mahalanobis_dist_feat,
time_anom_thresh_var,
feat_anom_thresh_var,
X_time_abs_recon_err,
X_feat_abs_recon_err):
"""Creates Estimator predictions and export outputs.
Given dimensions of inputs, mahalanobis distances and their respective
thresholds, and reconstructed inputs' absolute errors, returns Estimator's
predictions and export outputs.
Args:
cur_batch_size: Current batch size, could be partially filled.
seq_len: Number of timesteps in sequence.
num_feat: Number of features.
mahalanobis_dist_time: Mahalanobis distance, time major.
mahalanobis_dist_feat: Mahalanobis distance, features major.
time_anom_thresh_var: Time anomaly threshold variable.
feat_anom_thresh_var: Features anomaly threshold variable.
X_time_abs_recon_err: Time major reconstructed input data's absolute
reconstruction error.
X_feat_abs_recon_err: Features major reconstructed input data's absolute
reconstruction error.
Returns:
predictions_dict: Dictionary of predictions to output for local prediction.
export_outputs: Dictionary to output from exported model for serving.
"""
# Flag predictions as either normal or anomalous
# shape = (cur_batch_size,)
time_anom_flags = tf.where(
condition=tf.reduce_any(
input_tensor=tf.greater(
x=tf.abs(x=mahalanobis_dist_time),
y=time_anom_thresh_var),
axis=1),
x=tf.ones(shape=[cur_batch_size], dtype=tf.int64),
y=tf.zeros(shape=[cur_batch_size], dtype=tf.int64))
# shape = (cur_batch_size,)
feat_anom_flags = tf.where(
condition=tf.reduce_any(
input_tensor=tf.greater(
x=tf.abs(x=mahalanobis_dist_feat),
y=feat_anom_thresh_var),
axis=1),
x=tf.ones(shape=[cur_batch_size], dtype=tf.int64),
y=tf.zeros(shape=[cur_batch_size], dtype=tf.int64))
# Create predictions dictionary
predictions_dict = {
"X_time_abs_recon_err": tf.reshape(
tensor=X_time_abs_recon_err,
shape=[cur_batch_size, seq_len, num_feat]),
"X_feat_abs_recon_err": tf.transpose(
a=tf.reshape(
tensor=X_feat_abs_recon_err,
shape=[cur_batch_size, num_feat, seq_len]),
perm=[0, 2, 1]),
"mahalanobis_dist_time": mahalanobis_dist_time,
"mahalanobis_dist_feat": mahalanobis_dist_feat,
"time_anom_flags": time_anom_flags,
"feat_anom_flags": feat_anom_flags}
# Create export outputs
export_outputs = {
"predict_export_outputs": tf.estimator.export.PredictOutput(
outputs=predictions_dict)
}
return predictions_dict, export_outputs
# Create our model function to be used in our custom estimator
def anomaly_detection(features, labels, mode, params):
"""Custom Estimator model function for anomaly detection.
Given dictionary of feature tensors, labels tensor, Estimator mode, and
dictionary for parameters, return EstimatorSpec object for custom Estimator.
Args:
features: Dictionary of feature tensors.
labels: Labels tensor or None.
mode: Estimator ModeKeys. Can take values of TRAIN, EVAL, and PREDICT.
params: Dictionary of parameters.
Returns:
EstimatorSpec object.
"""
print("\nanomaly_detection: features = \n{}".format(features))
print("anomaly_detection: labels = \n{}".format(labels))
print("anomaly_detection: mode = \n{}".format(mode))
print("anomaly_detection: params = \n{}".format(params))
# Get input sequence tensor into correct shape
# Get dynamic batch size in case there was a partially filled batch
cur_batch_size = tf.shape(
input=features[UNLABELED_CSV_COLUMNS[0]], out_type=tf.int64)[0]
# Get the number of features
num_feat = len(UNLABELED_CSV_COLUMNS)
# Stack all of the features into a 3-D tensor
# shape = (cur_batch_size, seq_len, num_feat)
X = tf.stack(
values=[features[key] for key in UNLABELED_CSV_COLUMNS], axis=2)
##############################################################################
# Variables for calculating error distribution statistics
(abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var,
abs_err_inv_cov_time_var,
abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var,
abs_err_inv_cov_feat_var) = create_both_mahalanobis_dist_vars(
seq_len=params["seq_len"], num_feat=num_feat)
# Variables for automatically tuning anomaly thresh
# Time based
(tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var,
tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var) = create_both_confusion_matrix_thresh_vars(
scope="mahalanobis_dist_thresh_vars",
time_thresh_size=[params["num_time_anom_thresh"]],
feat_thresh_size=[params["num_feat_anom_thresh"]])
with tf.variable_scope(
name_or_scope="mahalanobis_dist_thresh_vars", reuse=tf.AUTO_REUSE):
time_anom_thresh_var = tf.get_variable(
name="time_anom_thresh_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[], dtype=tf.float64),
trainable=False)
feat_anom_thresh_var = tf.get_variable(
name="feat_anom_thresh_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[], dtype=tf.float64),
trainable=False)
# Variables for tuning anomaly thresh evaluation
(tp_thresh_eval_time_var,
fn_thresh_eval_time_var,
fp_thresh_eval_time_var,
tn_thresh_eval_time_var,
tp_thresh_eval_feat_var,
fn_thresh_eval_feat_var,
fp_thresh_eval_feat_var,
tn_thresh_eval_feat_var) = create_both_confusion_matrix_thresh_vars(
scope="anom_thresh_eval_vars", time_thresh_size=[], feat_thresh_size=[])
# Create dummy variable for graph dependency requiring a gradient for TRAIN
dummy_var = tf.get_variable(
name="dummy_var",
dtype=tf.float64,
initializer=tf.zeros(shape=[], dtype=tf.float64),
trainable=True)
################################################################################
predictions_dict = None
loss = None
train_op = None
eval_metric_ops = None
export_outputs = None
# Now branch off based on which mode we are in
# Call specific model
model_functions = {
"dense_autoencoder": dense_autoencoder_model,
"lstm_enc_dec_autoencoder": lstm_enc_dec_autoencoder_model,
"pca": pca_model}
# Get function pointer for selected model type
model_function = model_functions[params["model_type"]]
# Build selected model
loss, train_op, X_time_orig, X_time_recon, X_feat_orig, X_feat_recon = \
model_function(X, mode, params, cur_batch_size, num_feat, dummy_var)
if not (mode == tf.estimator.ModeKeys.TRAIN and
params["training_mode"] == "reconstruction"):
# shape = (cur_batch_size * seq_len, num_feat)
X_time_abs_recon_err = tf.abs(
x=X_time_orig - X_time_recon)
# Features based
# shape = (cur_batch_size * num_feat, seq_len)
X_feat_abs_recon_err = tf.abs(
x=X_feat_orig - X_feat_recon)
if (mode == tf.estimator.ModeKeys.TRAIN and
params["training_mode"] == "calculate_error_distribution_statistics"):
loss, train_op = calculate_error_distribution_statistics_training(
cur_batch_size,
num_feat,
X_time_abs_recon_err,
abs_err_count_time_var,
abs_err_mean_time_var,
abs_err_cov_time_var,
abs_err_inv_cov_time_var,
X_feat_abs_recon_err,
abs_err_count_feat_var,
abs_err_mean_feat_var,
abs_err_cov_feat_var,
abs_err_inv_cov_feat_var,
params,
dummy_var)
elif (mode == tf.estimator.ModeKeys.EVAL and
params["training_mode"] != "tune_anomaly_thresholds"):
loss, eval_metric_ops = reconstruction_evaluation(
X_time_orig, X_time_recon, params["training_mode"])
elif (mode == tf.estimator.ModeKeys.PREDICT or
((mode == tf.estimator.ModeKeys.TRAIN or
mode == tf.estimator.ModeKeys.EVAL) and
params["training_mode"] == "tune_anomaly_thresholds")):
with tf.variable_scope(
name_or_scope="mahalanobis_dist_vars", reuse=tf.AUTO_REUSE):
# Time based
# shape = (cur_batch_size, seq_len)
mahalanobis_dist_time = mahalanobis_dist(
err_vec=X_time_abs_recon_err,
mean_vec=abs_err_mean_time_var,
inv_cov=abs_err_inv_cov_time_var,
final_shape=params["seq_len"])
# Features based
# shape = (cur_batch_size, num_feat)
mahalanobis_dist_feat = mahalanobis_dist(
err_vec=X_feat_abs_recon_err,
mean_vec=abs_err_mean_feat_var,
inv_cov=abs_err_inv_cov_feat_var,
final_shape=num_feat)
if mode != tf.estimator.ModeKeys.PREDICT:
labels_norm_mask = tf.equal(x=labels, y=0)
labels_anom_mask = tf.equal(x=labels, y=1)
if mode == tf.estimator.ModeKeys.TRAIN:
loss, train_op = tune_anomaly_thresholds_training(
labels_norm_mask,
labels_anom_mask,
mahalanobis_dist_time,
tp_thresh_time_var,
fn_thresh_time_var,
fp_thresh_time_var,
tn_thresh_time_var,
time_anom_thresh_var,
mahalanobis_dist_feat,
tp_thresh_feat_var,
fn_thresh_feat_var,
fp_thresh_feat_var,
tn_thresh_feat_var,
feat_anom_thresh_var,
params,
mode,
dummy_var)
elif mode == tf.estimator.ModeKeys.EVAL:
loss, eval_metric_ops = tune_anomaly_thresholds_evaluation(
labels_norm_mask,
labels_anom_mask,
time_anom_thresh_var,
mahalanobis_dist_time,
tp_thresh_eval_time_var,
fn_thresh_eval_time_var,
fp_thresh_eval_time_var,
tn_thresh_eval_time_var,
feat_anom_thresh_var,
mahalanobis_dist_feat,
tp_thresh_eval_feat_var,
fn_thresh_eval_feat_var,
fp_thresh_eval_feat_var,
tn_thresh_eval_feat_var,
params,
mode)
else: # mode == tf.estimator.ModeKeys.PREDICT
predictions_dict, export_outputs = anomaly_detection_predictions(
cur_batch_size,
params["seq_len"],
num_feat,
mahalanobis_dist_time,
mahalanobis_dist_feat,
time_anom_thresh_var,
feat_anom_thresh_var,
X_time_abs_recon_err,
X_feat_abs_recon_err)
# Return EstimatorSpec
return tf.estimator.EstimatorSpec(
mode=mode,
predictions=predictions_dict,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops,
export_outputs=export_outputs)
# Serving input functions
def fix_shape_and_type_for_serving(placeholder):
"""Fixes the shape and type of serving input strings.
Given placeholder tensor, return parsed and processed feature tensor.
Args:
placeholder: Placeholder tensor holding raw data from serving input
function.
Returns:
Parsed and processed feature tensor.
"""
cur_batch_size = tf.shape(input=placeholder, out_type=tf.int64)[0]
# String split each string in batch and output values from the resulting
# SparseTensors
# shape = (batch_size, seq_len)
split_string = tf.stack(values=tf.map_fn(
fn=lambda x: tf.string_split(
source=[placeholder[x]], delimiter=",").values,
elems=tf.range(
start=0, limit=cur_batch_size, dtype=tf.int64),
dtype=tf.string), axis=0)
# Convert each string in the split tensor to float
# shape = (batch_size, seq_len)
feature_tensor = tf.string_to_number(
string_tensor=split_string, out_type=tf.float64)
return feature_tensor
def get_shape_and_set_modified_shape_2D(tensor, additional_dimension_sizes):
"""Fixes the shape and type of serving input strings.
Given feature tensor and additional dimension size, sequence length,
fixes dynamic shape ambiguity of last dimension so that we will be able to
use it in our DNN (since tf.layers.dense require the last dimension to be
known).
Args:
tensor: tf.float64 vector feature tensor.
additional_dimension_sizes: Additional dimension size, namely sequence
length.
Returns:
Feature tensor with set static shape for sequence length.
"""
# Get static shape for tensor and convert it to list
shape = tensor.get_shape().as_list()
# Set outer shape to additional_dimension_sizes[0] since know this is the
# correct size
shape[1] = additional_dimension_sizes[0]
# Set the shape of tensor to our modified shape
# shape = (batch_size, additional_dimension_sizes[0])
tensor.set_shape(shape=shape)
return tensor
def serving_input_fn(seq_len):
"""Serving input function.
Given the sequence length, return ServingInputReceiver object.
Args:
seq_len: Number of timesteps in sequence.
Returns:
ServingInputReceiver object containing features and receiver tensors.
"""
# Create placeholders to accept the data sent to the model at serving time
# All features come in as a batch of strings, shape = (batch_size,),
# this was so because of passing the arrays to online ml-engine prediction
feature_placeholders = {
feature: tf.placeholder(
dtype=tf.string, shape=[None])
for feature in UNLABELED_CSV_COLUMNS
}
# Create feature tensors
features = {key: fix_shape_and_type_for_serving(placeholder=tensor)
for key, tensor in feature_placeholders.items()}
# Fix dynamic shape ambiguity of feature tensors for our DNN
features = {key: get_shape_and_set_modified_shape_2D(
tensor=tensor, additional_dimension_sizes=[seq_len])
for key, tensor in features.items()}
return tf.estimator.export.ServingInputReceiver(
features=features, receiver_tensors=feature_placeholders)
def train_and_evaluate(args):
"""Train and evaluate custom Estimator with three training modes.
Given the dictionary of parameters, create custom Estimator and run up to
three training modes then return Estimator object.
Args:
args: Dictionary of parameters.
Returns:
Estimator object.
"""
# Create our custom estimator using our model function
estimator = tf.estimator.Estimator(
model_fn=anomaly_detection,
model_dir=args["output_dir"],
params={key: val for key, val in args.items()})
if args["training_mode"] == "reconstruction":
if args["model_type"] == "pca":
estimator.train(
input_fn=read_dataset(
filename=args["train_file_pattern"],
mode=tf.estimator.ModeKeys.EVAL,
batch_size=args["train_batch_size"],
params=args),
steps=None)
else: # dense_autoencoder or lstm_enc_dec_autoencoder
# Create early stopping hook to help reduce overfitting
early_stopping_hook = tf.contrib.estimator.stop_if_no_decrease_hook(
estimator=estimator,
metric_name="rmse",
max_steps_without_decrease=100,
min_steps=1000,
run_every_secs=60,
run_every_steps=None)
# Create train spec to read in our training data
train_spec = tf.estimator.TrainSpec(
input_fn=read_dataset(
filename=args["train_file_pattern"],
mode=tf.estimator.ModeKeys.TRAIN,
batch_size=args["train_batch_size"],
params=args),
max_steps=args["train_steps"],
hooks=[early_stopping_hook])
# Create eval spec to read in our validation data and export our model
eval_spec = tf.estimator.EvalSpec(
input_fn=read_dataset(
filename=args["eval_file_pattern"],
mode=tf.estimator.ModeKeys.EVAL,
batch_size=args["eval_batch_size"],
params=args),
steps=None,
start_delay_secs=args["start_delay_secs"], # start eval after N secs
throttle_secs=args["throttle_secs"]) # evaluate every N secs
# Create train and evaluate loop to train and evaluate our estimator
tf.estimator.train_and_evaluate(
estimator=estimator, train_spec=train_spec, eval_spec=eval_spec)
else:
if args["training_mode"] == "calculate_error_distribution_statistics":
# Get final mahalanobis statistics over the entire val_1 dataset
train_spec = tf.estimator.TrainSpec(
input_fn=read_dataset(
filename=args["train_file_pattern"],
mode=tf.estimator.ModeKeys.EVAL, # read through val data once
batch_size=args["train_batch_size"],
params=args),
max_steps=args["train_steps"])
# Don't create exporter for serving yet since anomaly thresholds
# aren't trained yet
exporter = None
elif args["training_mode"] == "tune_anomaly_thresholds":
# Tune anomaly thresholds using val_2 and val_anom datasets
train_spec = tf.estimator.TrainSpec(
input_fn=read_dataset(
filename=args["train_file_pattern"],
mode=tf.estimator.ModeKeys.EVAL, # read through val data once
batch_size=args["train_batch_size"],
params=args),
max_steps=args["train_steps"])
# Create exporter that uses serving_input_fn to create saved_model
# for serving
exporter = tf.estimator.LatestExporter(
name="exporter",
serving_input_receiver_fn=lambda: serving_input_fn(args["seq_len"]))
# Create eval spec to read in our validation data and export our model
eval_spec = tf.estimator.EvalSpec(
input_fn=read_dataset(
filename=args["eval_file_pattern"],
mode=tf.estimator.ModeKeys.EVAL,
batch_size=args["eval_batch_size"],
params=args),
steps=None,
exporters=exporter,
start_delay_secs=args["start_delay_secs"], # start eval after N secs
throttle_secs=args["throttle_secs"]) # evaluate every N secs
# Create train and evaluate loop to train and evaluate our estimator
tf.estimator.train_and_evaluate(
estimator=estimator, train_spec=train_spec, eval_spec=eval_spec)
# +
# %%writefile anomaly_detection_module/trainer/task.py
import argparse
import json
import os
from . import model
import tensorflow as tf
if __name__ == "__main__":
parser = argparse.ArgumentParser()
# File arguments
parser.add_argument(
"--train_file_pattern",
help="GCS location to read training data.",
required=True
)
parser.add_argument(
"--eval_file_pattern",
help="GCS location to read evaluation data.",
required=True
)
parser.add_argument(
"--output_dir",
help="GCS location to write checkpoints and export models.",
required=True
)
parser.add_argument(
"--job-dir",
help="This model ignores this field, but it is required by gcloud.",
default="junk"
)
# Sequence shape hyperparameters
parser.add_argument(
"--seq_len",
help="Number of timesteps to include in each example.",
type=int,
default=32
)
# Training parameters
parser.add_argument(
"--train_batch_size",
help="Number of examples in training batch.",
type=int,
default=32
)
parser.add_argument(
"--eval_batch_size",
help="Number of examples in evaluation batch.",
type=int,
default=32
)
parser.add_argument(
"--train_steps",
help="Number of batches to train.",
type=int,
default=2000
)
parser.add_argument(
"--learning_rate",
help="How quickly or slowly we train our model by scaling the gradient.",
type=float,
default=0.1
)
parser.add_argument(
"--start_delay_secs",
help="Number of seconds to wait before first evaluation.",
type=int,
default=60
)
parser.add_argument(
"--throttle_secs",
help="Number of seconds to wait between evaluations.",
type=int,
default=120
)
# Model hyperparameters
# dense_autoencoder, lstm_enc_dec_autoencoder, pca
parser.add_argument(
"--model_type",
help="Which model type we will use.",
type=str,
default="dense_autoencoder"
)
## Dense Autoencoder
parser.add_argument(
"--enc_dnn_hidden_units",
help="Hidden layer sizes to use for encoder DNN.",
default="1024 256 64"
)
parser.add_argument(
"--latent_vector_size",
help="Number of neurons for latent vector between encoder and decoder.",
type=int,
default=8
)
parser.add_argument(
"--dec_dnn_hidden_units",
help="Hidden layer sizes to use for decoder DNN.",
default="64 256 1024"
)
parser.add_argument(
"--time_loss_weight",
help="Amount to weight the time based loss.",
type=float,
default=1.0
)
parser.add_argument(
"--feat_loss_weight",
help="Amount to weight the features based loss.",
type=float,
default=1.0
)
## LSTM Encoder-Decoder Autoencoder
parser.add_argument(
"--reverse_labels_sequence",
help="Whether we should reverse the labels sequence dimension or not.",
type=bool,
default=True
)
parser.add_argument(
"--enc_lstm_hidden_units",
help="Hidden layer sizes to use for LSTM encoder.",
default="64 32 16"
)
parser.add_argument(
"--dec_lstm_hidden_units",
help="Hidden layer sizes to use for LSTM decoder.",
default="16 32 64"
)
parser.add_argument(
"--lstm_dropout_output_keep_probs",
help="Keep probabilties for LSTM outputs.",
default="1.0 1.0 1.0"
)
parser.add_argument(
"--dnn_hidden_units",
help="Hidden layer sizes to use for DNN.",
default="1024 256 64"
)
## PCA
parser.add_argument(
"--k_principal_components",
help="Top k principal components to keep after eigendecomposition.",
type=int,
default=3
)
# Anomaly detection
# reconstruction, calculate_error_distribution_statistics,
# and tune_anomaly_thresholds
parser.add_argument(
"--training_mode",
help="Which training mode we are in.",
type=str,
default="reconstruction"
)
parser.add_argument(
"--num_time_anom_thresh",
help="Number of anomaly thresholds to evaluate in time dimension.",
type=int,
default=120
)
parser.add_argument(
"--num_feat_anom_thresh",
help="Number of anomaly thresholds to evaluate in features dimension.",
type=int,
default=120
)
parser.add_argument(
"--min_time_anom_thresh",
help="Minimum anomaly threshold to evaluate in time dimension.",
type=float,
default=100.0
)
parser.add_argument(
"--max_time_anom_thresh",
help="Maximum anomaly threshold to evaluate in time dimension.",
type=float,
default=2000.0
)
parser.add_argument(
"--min_feat_anom_thresh",
help="Minimum anomaly threshold to evaluate in features dimension.",
type=float,
default=100.0
)
parser.add_argument(
"--max_feat_anom_thresh",
help="Maximum anomaly threshold to evaluate in features dimension.",
type=float,
default=2000.0
)
parser.add_argument(
"--time_anom_thresh",
help="Anomaly threshold in time dimension.",
type=float,
default=None
)
parser.add_argument(
"--feat_anom_thresh",
help="Anomaly threshold in features dimension.",
type=float,
default=None
)
parser.add_argument(
"--eps",
help="Added to the cov matrix before inversion to avoid being singular.",
type=str,
default="1e-12"
)
parser.add_argument(
"--f_score_beta",
help="Value of beta of the f-beta score.",
type=float,
default=0.05
)
# Parse all arguments
args = parser.parse_args()
arguments = args.__dict__
# Unused args provided by service
arguments.pop("job_dir", None)
arguments.pop("job-dir", None)
# Fix list arguments
## Dense Autoencoder
arguments["enc_dnn_hidden_units"] = [
int(x) for x in arguments["enc_dnn_hidden_units"].split(" ")]
arguments["dec_dnn_hidden_units"] = [
int(x) for x in arguments["dec_dnn_hidden_units"].split(" ")]
## LSTM Encoder-Decoder Autoencoder
arguments["enc_lstm_hidden_units"] = [
int(x) for x in arguments["enc_lstm_hidden_units"].split(" ")]
arguments["dec_lstm_hidden_units"] = [
int(x) for x in arguments["dec_lstm_hidden_units"].split(" ")]
arguments["lstm_dropout_output_keep_probs"] = [
float(x) for x in arguments["lstm_dropout_output_keep_probs"].split(" ")]
arguments["dnn_hidden_units"] = [
int(x) for x in arguments["dnn_hidden_units"].split(" ")]
# Fix eps argument
arguments["eps"] = float(arguments["eps"])
# Append trial_id to path if we are doing hptuning
# This code can be removed if you are not using hyperparameter tuning
arguments["output_dir"] = os.path.join(
arguments["output_dir"],
json.loads(
os.environ.get("TF_CONFIG", "{}")
).get("task", {}).get("trial", "")
)
# Run the training job
model.train_and_evaluate(arguments)
# -
# # Training model module
# ## Locally
# ### Train reconstruction variables
# + language="bash"
# rm -rf trained_model
# export PYTHONPATH=$PYTHONPATH:$PWD/anomaly_detection_module
# python -m trainer.task \
# --train_file_pattern="data/train_norm_seq.csv" \
# --eval_file_pattern="data/val_norm_1_seq.csv" \
# --output_dir=$PWD/trained_model \
# --job-dir=./tmp \
# --seq_len=30 \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2000 \
# --learning_rate=0.1 \
# --start_delay_secs=60 \
# --throttle_secs=120 \
# --model_type="lstm_enc_dec_autoencoder" \
# --reverse_labels_sequence=True \
# --enc_lstm_hidden_units="64 32 16" \
# --dec_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --training_mode="reconstruction" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300
# -
# ### Train error distribution statistics variables
# + language="bash"
# export PYTHONPATH=$PYTHONPATH:$PWD/anomaly_detection_module
# python -m trainer.task \
# --train_file_pattern="data/val_norm_1_seq.csv" \
# --eval_file_pattern="data/val_norm_1_seq.csv" \
# --output_dir=$PWD/trained_model \
# --job-dir=./tmp \
# --seq_len=30 \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2200 \
# --model_type="lstm_enc_dec_autoencoder" \
# --reverse_labels_sequence=True \
# --enc_lstm_hidden_units="64 32 16" \
# --dec_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --training_mode="calculate_error_distribution_statistics" \
# --eps="1e-12" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300
# -
# ### Tune anomaly thresholds
# + language="bash"
# export PYTHONPATH=$PYTHONPATH:$PWD/anomaly_detection_module
# python -m trainer.task \
# --train_file_pattern="data/labeled_val_mixed_seq.csv" \
# --eval_file_pattern="data/labeled_val_mixed_seq.csv" \
# --output_dir=$PWD/trained_model \
# --job-dir=./tmp \
# --seq_len=30 \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2400 \
# --model_type="lstm_enc_dec_autoencoder" \
# --reverse_labels_sequence=True \
# --enc_lstm_hidden_units="64 32 16" \
# --dec_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --training_mode="tune_anomaly_thresholds" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300 \
# --min_time_anom_thresh=1.0 \
# --max_time_anom_thresh=20.0 \
# --min_feat_anom_thresh=20.0 \
# --max_feat_anom_thresh=80.0 \
# --f_score_beta=0.05
# -
# ## GCloud
# Copy data over to bucket
# + language="bash"
# gsutil -m cp -r data/* gs://$BUCKET/anomaly_detection/data
# -
# ### Train reconstruction variables
# + language="bash"
# OUTDIR=gs://$BUCKET/anomaly_detection/trained_model
# JOBNAME=job_anomaly_detection_reconstruction_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gsutil -m rm -rf $OUTDIR
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/anomaly_detection/data/train_norm_seq.csv \
# --eval_file_pattern=gs://$BUCKET/anomaly_detection/data/val_norm_1_seq.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2000 \
# --learning_rate=0.1 \
# --start_delay_secs=60 \
# --throttle_secs=120 \
# --model_type="lstm_enc_dec_autoencoder" \
# --reverse_labels_sequence=True \
# --enc_lstm_hidden_units="64 32 16" \
# --dec_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --training_mode="reconstruction" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300
# -
# ### Hyperparameter tuning of reconstruction hyperparameters
# %%writefile hyperparam_reconstruction.yaml
trainingInput:
scaleTier: STANDARD_1
hyperparameters:
hyperparameterMetricTag: rmse
goal: MINIMIZE
maxTrials: 30
maxParallelTrials: 1
params:
- parameterName: enc_lstm_hidden_units
type: CATEGORICAL
categoricalValues: ["64 32 16", "256 128 16", "64 64 64"]
- parameterName: dec_lstm_hidden_units
type: CATEGORICAL
categoricalValues: ["16 32 64", "16 128 256", "64 64 64"]
- parameterName: lstm_dropout_output_keep_probs
type: CATEGORICAL
categoricalValues: ["0.9 1.0 1.0", "0.95 0.95 1.0", "0.95 0.95 0.95"]
- parameterName: dnn_hidden_units
type: CATEGORICAL
categoricalValues: ["256 128 64", "256 128 16", "64 64 64"]
- parameterName: train_batch_size
type: INTEGER
minValue: 8
maxValue: 512
scaleType: UNIT_LOG_SCALE
- parameterName: learning_rate
type: DOUBLE
minValue: 0.001
maxValue: 0.1
scaleType: UNIT_LINEAR_SCALE
# + language="bash"
# OUTDIR=gs://$BUCKET/anomaly_detection/hyperparam_reconstruction
# JOBNAME=job_anomaly_detection_hyperparam_reconstruction_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gsutil -m rm -rf $OUTDIR
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --config=hyperparam_reconstruction.yaml \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/anomaly_detection/data/train_norm_seq.csv \
# --eval_file_pattern=gs://$BUCKET/anomaly_detection/data/val_norm_1_seq.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2000 \
# --start_delay_secs=60 \
# --throttle_secs=120 \
# --training_mode="reconstruction" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300
# -
# ### Train error distribution variables
# + language="bash"
# OUTDIR=gs://$BUCKET/anomaly_detection/trained_model
# JOBNAME=job_anomaly_detection_calculate_error_distribution_statistics_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/anomaly_detection/data/val_norm_1_seq.csv \
# --eval_file_pattern=gs://$BUCKET/anomaly_detection/data/val_norm_1_seq.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2200 \
# --model_type="lstm_enc_dec_autoencoder" \
# --reverse_labels_sequence=True \
# --enc_lstm_hidden_units="64 32 16" \
# --dec_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --training_mode="calculate_error_distribution_statistics" \
# --eps="1e-12" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300
# -
# ### Tune anomaly thresholds
# + language="bash"
# OUTDIR=gs://$BUCKET/anomaly_detection/trained_model
# JOBNAME=job_anomaly_detection_tune_anomaly_thresholds_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/anomaly_detection/data/labeled_val_mixed_seq.csv \
# --eval_file_pattern=gs://$BUCKET/anomaly_detection/data/labeled_val_mixed_seq.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2400 \
# --model_type="lstm_enc_dec_autoencoder" \
# --reverse_labels_sequence=True \
# --enc_lstm_hidden_units="64 32 16" \
# --dec_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --training_mode="tune_anomaly_thresholds" \
# --num_time_anom_thresh=300 \
# --num_feat_anom_thresh=300 \
# --min_time_anom_thresh=2.0 \
# --max_time_anom_thresh=15.0 \
# --min_feat_anom_thresh=20 \
# --max_feat_anom_thresh=60 \
# --f_score_beta=0.05
# -
# # Deploy
# + language="bash"
# MODEL_NAME="anomaly_detection"
# MODEL_VERSION="v1"
# MODEL_LOCATION=$(gsutil ls gs://$BUCKET/anomaly_detection/trained_model/export/exporter/ | tail -1)
# echo "Deleting and deploying $MODEL_NAME $MODEL_VERSION from $MODEL_LOCATION ... this will take a few minutes"
# #gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
# #gcloud ml-engine models delete ${MODEL_NAME}
# gcloud ml-engine models create $MODEL_NAME --regions $REGION
# gcloud ml-engine versions create $MODEL_VERSION --model $MODEL_NAME --origin $MODEL_LOCATION --runtime-version 1.13
# -
# # Prediction
UNLABELED_CSV_COLUMNS = ["tag_{0}".format(tag) for tag in range(0, 5)]
import numpy as np
labeled_test_mixed_sequences_array = np.loadtxt(
fname="data/labeled_test_mixed_sequences.csv", dtype=str, delimiter=";")
print("labeled_test_mixed_sequences_array.shape = {}".format(
labeled_test_mixed_sequences_array.shape))
number_of_prediction_instances = 10
print("labels = {}".format(
labeled_test_mixed_sequences_array[0:number_of_prediction_instances, -1]))
# ### Local prediction from local model
with open('test_sequences.json', 'w') as outfile:
test_data_normal_string_list = labeled_test_mixed_sequences_array.tolist()[0:number_of_prediction_instances]
json_string = ""
for example in test_data_normal_string_list:
json_string += "{" + ','.join(["{0}: \"{1}\"".format('\"' + UNLABELED_CSV_COLUMNS[i] + '\"', example[i])
for i in range(len(UNLABELED_CSV_COLUMNS))]) + "}\n"
json_string = json_string.replace(' ', '').replace(':', ': ').replace(',', ', ')
print(json_string)
outfile.write("%s" % json_string)
# + language="bash"
# model_dir=$(ls ${PWD}/trained_model/export/exporter | tail -1)
# gcloud ml-engine local predict \
# --model-dir=${PWD}/trained_model/export/exporter/${model_dir} \
# --json-instances=./test_sequences.json
# -
# ### GCloud ML-Engine prediction from deployed model
test_data_normal_string_list = labeled_test_mixed_sequences_array.tolist()[0:number_of_prediction_instances]
# Format dataframe to instances list to get sent to ML-Engine
instances = [{UNLABELED_CSV_COLUMNS[i]: example[i]
for i in range(len(UNLABELED_CSV_COLUMNS))}
for example in labeled_test_mixed_sequences_array.tolist()[0:number_of_prediction_instances]]
instances
# +
# Send instance dictionary to receive response from ML-Engine for online prediction
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = GoogleCredentials.get_application_default()
api = discovery.build("ml", "v1", credentials = credentials)
request_data = {"instances": instances}
parent = "projects/%s/models/%s/versions/%s" % (PROJECT, "anomaly_detection", "v1")
response = api.projects().predict(body = request_data, name = parent).execute()
print("response = {}".format(response))
# -
| courses/machine_learning/asl/open_project/time_series_anomaly_detection/tf_anomaly_detection_model_selection/model_selection_anomaly_detection_gcp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from PIL import Image, ImageDraw, ImageFont
from skimage import transform as tf
def create_captcha(text, shear=0, size=(100, 30), scale=1):
im = Image.new("L", size, "black")
draw = ImageDraw.Draw(im)
font = ImageFont.truetype(r"bretan/Coval-Black.otf", 22)
draw.text((0, 0), text, fill=1, font=font)
image = np.array(im)
affine_tf = tf.AffineTransform(shear=shear)
image = tf.warp(image, affine_tf)
image = image / image.max()
# 应用缩放
shape = image.shape
shapex, shapey = (int(shape[0] * scale), int(shape[1] * scale))
image = tf.resize(image, (shapex, shapey))
return image
# %matplotlib inline
from matplotlib import pyplot as plt
image = create_captcha("GENE", shear=0.5, scale=0.6)
plt.imshow(image, cmap='Greys')
image = create_captcha("BONE", shear=0.1, scale=1.0)
plt.imshow(image, cmap='Greys')
image = create_captcha("BARK", shear=0.8, scale=1.0)
plt.imshow(image, cmap='Greys')
image = create_captcha("WOOF", shear=0.25, scale=1.5)
plt.imshow(image, cmap='Greys')
# +
from skimage.measure import label, regionprops
def segment_image(image):
# 标记函数能找出连通的非黑色像素组成的子图像
labeled_image = label(image>0.2, connectivity=1, background=0)
subimages = []
# 用 regionprops 函数分离子图像
# 此处替换了原书代码,修正了子图像顺序
regions = regionprops(labeled_image)
regions.sort(key=lambda x:x.bbox[1])
for region in regions:
# 提取子图像
start_x, start_y, end_x, end_y = region.bbox
subimages.append(image[start_x:end_x,start_y:end_y])
if len(subimages) == 0:
# 没有找到子图像,则返回完整图像
return [image,]
return subimages
# -
subimages = segment_image(image)
f, axes = plt.subplots(1, len(subimages), figsize=(10, 3))
for i in range(len(subimages)):
axes[i].imshow(subimages[i], cmap="gray")
image = create_captcha("GENE", shear=0.5, scale=0.6)
subimages = segment_image(image)
f, axes = plt.subplots(1, len(subimages), figsize=(10, 3), sharey=True)
for i in range(len(subimages)):
axes[i].imshow(subimages[i], cmap="gray")
# +
from sklearn.utils import check_random_state
random_state = check_random_state(14)
letters = list("ABCDEFGHIJKLMNOPQRSTUVWXYZ")
shear_values = np.arange(0, 0.8, 0.05)
scale_values = np.arange(0.9, 1.1, 0.1)
# 此处替换了原书代码,避免样本重复
import itertools
sample_params = list(itertools.product(letters, shear_values, scale_values))
random_state.shuffle(sample_params)
# +
def generate_sample(random_state=None):
random_state = check_random_state(random_state)
letter = random_state.choice(letters)
shear = random_state.choice(shear_values)
scale = random_state.choice(scale_values)
# 我们把图像尺寸设置为(30, 30),以确保图像中能显示下所有的文字
return create_captcha(letter, shear=shear, size=(30, 30),
scale=scale), letters.index(letter)
# 此处替换了原书代码,避免样本重复
def generate_samples(size, random_state=None):
return zip(*[(create_captcha(sample_params[i][0], shear=sample_params[i][1],
size=(30, 30), scale=sample_params[i][2]),
letters.index(sample_params[i][0])) for i in range(size)])
# -
image, target = generate_sample(random_state)
plt.imshow(image, cmap="Greys")
print("The target for this image is: {0}".format(target))
# 此处替换了原书代码,避免样本重复
dataset, targets = generate_samples(1000)
dataset = np.array([tf.resize(segment_image(sample)[0], (20, 20)) for
sample in dataset])
dataset = np.array(dataset, dtype='float')
targets = np.array(targets)
from sklearn.preprocessing import OneHotEncoder
onehot = OneHotEncoder()
y = onehot.fit_transform(targets.reshape(targets.shape[0],1))
y = y.todense()
X = dataset.reshape((dataset.shape[0], dataset.shape[1] *
dataset.shape[2]))
from sklearn.cross_validation import train_test_split
# 训练数据集比例调整为0.8,避免测试数据集中字母覆盖不全
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.8, random_state=14)
from sklearn.neural_network import MLPClassifier
clf = MLPClassifier(hidden_layer_sizes=(100,), random_state=14)
clf.get_params()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
from sklearn.metrics import f1_score
f1_score(y_pred=y_pred, y_true=y_test, average='macro')
from sklearn.metrics import classification_report
print(classification_report(y_pred=y_pred, y_true=y_test))
def predict_captcha(captcha_image, neural_network):
subimages = segment_image(captcha_image)
# 执行转换,与训练数据中的一样
dataset = np.array([tf.resize(subimage, (20, 20)) for subimage in
subimages])
X_test = dataset.reshape((dataset.shape[0], dataset.shape[1] *
dataset.shape[2]))
# 用predict_proba和argmax获取最可能的预测
y_pred = neural_network.predict_proba(X_test)
predictions = np.argmax(y_pred, axis=1)
# 把预测转换为字母
predicted_word = str.join("", [letters[prediction] for prediction in
predictions])
return predicted_word
word = "GENE"
captcha = create_captcha(word, shear=0.2)
print(predict_captcha(captcha, clf))
plt.imshow(captcha, cmap="Greys")
def test_prediction(word, net, shear=0.2, scale=1):
captcha = create_captcha(word, shear=shear, scale=scale,
size=(len(word) * 25, 30))
prediction = predict_captcha(captcha, net)
return word == prediction, word, prediction
from nltk.corpus import words
valid_words = set([word.upper() for word in words.words() if len(word) == 4])
num_correct = 0
num_incorrect = 0
for word in valid_words:
shear = random_state.choice(shear_values)
scale = random_state.choice(scale_values)
correct, word, prediction = test_prediction(word, clf, shear=shear,
scale=scale)
if correct:
num_correct += 1
else:
num_incorrect += 1
print("Number correct is {0}".format(num_correct))
print("Number incorrect is {0}".format(num_incorrect))
print("Accurary is {0}".format(num_correct / (num_correct + num_incorrect)))
def evaluation_versus_shear(shear_value):
num_correct = 0
num_incorrect = 0
for word in valid_words:
scale = random_state.choice(scale_values)
correct, word, prediction = test_prediction(
word, clf, shear=shear_value, scale=scale)
if correct:
num_correct += 1
else:
num_incorrect += 1
return num_correct / (num_correct+num_incorrect)
scores = [evaluation_versus_shear(shear) for shear in shear_values]
import seaborn
seaborn.set(style="darkgrid")
plt.figure(figsize=(10, 7))
plt.ylabel = "Accuracy"
plt.xlabel = "Shear"
plt.plot(shear_values, scores)
from nltk.metrics import edit_distance
steps = edit_distance("STEP", "STOP")
print("The number of steps needed is: {0}".format(steps))
def compute_distance(prediction, word):
len_word = min(len(prediction), len(word))
return len_word - sum([prediction[i] == word[i] for i in
range(len_word)])
# +
from operator import itemgetter
def improved_prediction(word, net, dictionary, shear=0.2, scale=1.0):
captcha = create_captcha(word, shear=shear, scale=scale)
prediction = predict_captcha(captcha, net)
if prediction not in dictionary:
distances = sorted([(word, compute_distance(prediction, word))
for word in dictionary], key=itemgetter(1))
best_word = distances[0]
prediction = best_word[0]
return word == prediction, word, prediction
# -
num_correct = 0
num_incorrect = 0
for word in valid_words:
shear = random_state.choice(shear_values)
scale = random_state.choice(scale_values)
correct, word, prediction = improved_prediction(
word, clf, valid_words, shear=shear, scale=scale)
if correct:
num_correct += 1
else:
num_incorrect += 1
print("Number correct is {0}".format(num_correct))
print("Number incorrect is {0}".format(num_incorrect))
print("Accurary is {0}".format(num_correct / (num_correct + num_incorrect)))
| ch8_CAPTCHA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Des succès immédiats
#
# Avec les problèmes à suivre, vous serez amenés à mettre en œuvre rapidement les structures de contrôle.
#
# ## Un problème capital
#
# Mettez en majuscules les mots de la liste grâce à une boucle.
words = ["A", "Lannister", "always", "pays", "his", "debts."]
for word in words:
print(word.upper())
# ## Ma liste de courses
#
# Concevez un programme qui détermine si une valeur entrée par un utilisateur est présente dans une liste prédéterminée de courses. Si la valeur ne fait pas partie de la liste, invitez l’utilisateur à l’enregistrer ; autrement, affichez un message pour le signaler. Retournez enfin la liste complète.
#
# Pour afficher un *prompt*, utilisez le bout de code suivant :
#
# ```python
# item = input("Quel article souhaitez-vous acheter ?\n")
# ```
items = ["eau", "lentilles", "choux", "pois cassés", "beaufort"]
item = input("Quel article souhaitez-vous acheter ?\n")
if item not in items:
items.append(item)
else:
print("Inutile de le rajouter à la liste, il est déjà présent !")
print(items)
# ## Le jeu du mentaliste
#
# Concevez un programme dans lequel l'ordinateur va choisir au hasard une valeur dans une liste prédéterminée, puis demandera ensuite à l’utilisateur de la deviner. Attention, l’utilisateur n’aura qu’un seul essai pour deviner la bonne valeur !
# +
# Function choice to pick an item in a list
from random import choice
# Some characters of House Lannister
characters = ["Tyrion", "Tywin", "Cersei", "Jaime"]
# Printing the list of the characters
print(f"La famille Lannister se compose de : {', '.join(characters)}")
# Picks a character
ia = choice(characters)
# Up to user
ih = input("Quel est votre Lannister préféré ?\n")
# Check
if ia == ih:
print("C’est également le personnage préféré de votre ordinateur !")
else:
print(f"Les machines ont des goûts bizarres, ce n’est pas son personnage préféré. Il fallait choisir {ia}")
# -
# ## Des multiples
#
# Écrivez un programme qui liste les dix premiers multiples de trois.
cpt = 0
for n in range(1,100):
if not n % 3:
print(n)
cpt += 1
if cpt == 10:
break;
# ## Encore des ours
#
# Écrivez un programme qui imprime exactement la phrase suivante :
# > "L’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’homme qui a vu l’ours."
against = "L'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'homme qui a vu l'ours."
string = "L'homme qui a vu l'ours."
while string != against:
string = string.replace("ours", "homme qui a vu l'ours")
print(string)
# ### La suite de Fibonacci
#
# Le bout de code ci-dessous affiche les quinze premiers éléments de la suite de Fibonacci, grâce à une boucle `while` :
a, b, c = 1, 1, 0
while c < 15:
print(b),
a, b, c = b, a + b, c + 1
# Transformez le code pour une utilisation avec une boucle `for`.
a, b = 1, 1
for cpt in range(15):
print b,
a, b = b, a + b
| 3.control-flow/answers/0.quick-win.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
tf.enable_eager_execution()
tfe = tf.contrib.eager # Shorthand for some symbols
# +
print(tf.add(1, 2))
print(tf.add([1, 2], [3, 4]))
print(tf.square(5))
print(tf.reduce_sum([1, 2, 3]))
print(tf.encode_base64("hello world"))
# Operator overloading is also supported
print(tf.square(2) + tf.square(3))
# -
x = tf.matmul([[1]], [[2, 3]])
print(x.shape)
print(x.dtype)
# +
import numpy as np
ndarray = np.ones([3, 3])
print("TensorFlow operations convert numpy arrays to Tensors automatically")
tensor = tf.multiply(ndarray, 42)
print(tensor)
print("And NumPy operations convert Tensors to numpy arrays automatically")
print(np.add(tensor, 1))
print("The .numpy() method explicitly converts a Tensor to a numpy array")
print(tensor.numpy())
# +
x = tf.random_uniform([3, 3])
print("Is there a GPU available: "),
print(tf.test.is_gpu_available())
print("Is the Tensor on GPU #0: "),
print(x.device.endswith('GPU:0'))
# +
ds_tensors = tf.data.Dataset.from_tensor_slices([1, 2, 3, 4, 5, 6])
# Create a CSV file
import tempfile
_, filename = tempfile.mkstemp()
with open(filename, 'w') as f:
f.write("""Line 1
Line 2
Line 3
""")
ds_file = tf.data.TextLineDataset(filename)
# +
ds_tensors = ds_tensors.map(tf.square).shuffle(2).batch(2)
ds_file = ds_file.batch(2)
# +
print('Elements of ds_tensors:')
for x in ds_tensors:
print(x)
print('\nElements in ds_file:')
for x in ds_file:
print(x)
# +
from math import pi
def f(x):
return tf.square(tf.sin(x))
assert f(pi/2).numpy() == 1.0
# grad_f will return a list of derivatives of f
# with respect to its arguments. Since f() has a single argument,
# grad_f will return a list with a single element.
grad_f = tfe.gradients_function(f)
assert tf.abs(grad_f(pi/2)[0]).numpy() < 1e-7
# +
def f(x):
return tf.square(tf.sin(x))
def grad(f):
return lambda x: tfe.gradients_function(f)(x)[0]
x = tf.lin_space(-2*pi, 2*pi, 100) # 100 points between -2π and +2π
import matplotlib.pyplot as plt
plt.plot(x, f(x), label="f")
plt.plot(x, grad(f)(x), label="first derivative")
plt.plot(x, grad(grad(f))(x), label="second derivative")
plt.plot(x, grad(grad(grad(f)))(x), label="third derivative")
plt.legend()
plt.show()
# -
| tensorflow_test/tf_eager.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
# +
iowa_file_path = 'Intermediate_Machine_Learning_Train_Data.csv'
home_data = pd.read_csv(iowa_file_path)
y = home_data.SalePrice
features = ['LotArea', 'YearBuilt', '1stFlrSF', '2ndFlrSF', 'FullBath', 'BedroomAbvGr', 'TotRmsAbvGrd']
X = home_data[features]
test_data_path = 'Intermediate_Machine_Learning_Test_Data.csv'
test_data = pd.read_csv(test_data_path)
test_X = test_data[features]
# -
home_data
# +
# Define the models
model_1 = RandomForestRegressor(n_estimators=50, random_state=0)
model_2 = RandomForestRegressor(n_estimators=100, random_state=0)
model_3 = RandomForestRegressor(n_estimators=100, criterion='mae', random_state=0)
model_4 = RandomForestRegressor(n_estimators=200, min_samples_split=20, random_state=0)
model_5 = RandomForestRegressor(n_estimators=100, max_depth=7, random_state=0)
models = [model_1, model_2, model_3, model_4, model_5]
# +
from sklearn.metrics import mean_absolute_error
X_train, X_valid, y_train, y_valid = train_test_split(X, y, train_size=0.8, test_size=0.2,
random_state=0)
# Function for comparing different models
def score_model(model, X_t=X_train, X_v=X_valid, y_t=y_train, y_v=y_valid):
model.fit(X_t, y_t)
preds = model.predict(X_v)
return mean_absolute_error(y_v, preds)
for i in range(0, len(models)):
mae = score_model(models[i])
print("Model %d MAE: %d" % (i+1, mae))
# +
my_model = RandomForestRegressor(n_estimators=100, criterion='mae', random_state=0)
my_model.fit(X, y)
# Generate test predictions
preds_test = my_model.predict(test_X)
# Save predictions in format used for competition scoring
output = pd.DataFrame({'Id': test_X.index,
'SalePrice': preds_test})
# -
output
# +
#A DataFrame object has two axes: “axis 0” and “axis 1”. “axis 0” represents rows and “axis 1” represents columns.
# -
def score_dataset(X_train, X_valid, y_train, y_valid):
model = RandomForestRegressor(n_estimators=100, random_state=0)
model.fit(X_train, y_train)
preds = model.predict(X_valid)
return mean_absolute_error(y_valid, preds)
# +
#three approaches to dealing with missing values
#Always perform imputation after splitting data in Train and Validation to prevent data leak
#Approach 1 (Drop Columns with Missing Values)
# Get names of columns with missing values
cols_with_missing = [col for col in X_train.columns
if X_train[col].isnull().any()]
# Drop columns in training and validation data
reduced_X_train = X_train.drop(cols_with_missing, axis=1)
reduced_X_valid = X_valid.drop(cols_with_missing, axis=1)
print("MAE from Approach 1 (Drop columns with missing values):")
print(score_dataset(reduced_X_train, reduced_X_valid, y_train, y_valid))
# +
#Approach 2 (Imputation)
from sklearn.impute import SimpleImputer
# Imputation
my_imputer = SimpleImputer()
imputed_X_train = pd.DataFrame(my_imputer.fit_transform(X_train))
imputed_X_valid = pd.DataFrame(my_imputer.transform(X_valid))
# Imputation removed column names; put them back
imputed_X_train.columns = X_train.columns
imputed_X_valid.columns = X_valid.columns
print("MAE from Approach 2 (Imputation):")
print(score_dataset(imputed_X_train, imputed_X_valid, y_train, y_valid))
# +
#Approach 3 (An Extension to Imputation)¶
# Make copy to avoid changing original data (when imputing)
X_train_plus = X_train.copy()
X_valid_plus = X_valid.copy()
# Make new columns indicating what will be imputed
for col in cols_with_missing:
X_train_plus[col + '_was_missing'] = X_train_plus[col].isnull()
X_valid_plus[col + '_was_missing'] = X_valid_plus[col].isnull()
# Imputation
my_imputer = SimpleImputer()
imputed_X_train_plus = pd.DataFrame(my_imputer.fit_transform(X_train_plus))
imputed_X_valid_plus = pd.DataFrame(my_imputer.transform(X_valid_plus))
# Imputation removed column names; put them back
imputed_X_train_plus.columns = X_train_plus.columns
imputed_X_valid_plus.columns = X_valid_plus.columns
print("MAE from Approach 3 (An Extension to Imputation):")
print(score_dataset(imputed_X_train_plus, imputed_X_valid_plus, y_train, y_valid))
# +
#Preliminary investigation
# Shape of training data (num_rows, num_columns)
print(X_train.shape)
# Number of missing values in each column of training data
missing_val_count_by_column = (X_train.isnull().sum())
print(missing_val_count_by_column[missing_val_count_by_column > 0])
# +
# Preprocessed training and validation features
final_imputer = SimpleImputer(strategy='median')
final_X_train = pd.DataFrame(final_imputer.fit_transform(X_train))
final_X_valid = pd.DataFrame(final_imputer.transform(X_valid))
# Imputation removed column names; put them back
final_X_train.columns = X_train.columns
final_X_valid.columns = X_valid.columns
# +
model = RandomForestRegressor(n_estimators=100, random_state=0)
model.fit(final_X_train, y_train)
# Get validation predictions and MAE
preds_valid = model.predict(final_X_valid)
print("MAE (Your approach):")
print(mean_absolute_error(y_valid, preds_valid))
# +
final_X_test = pd.DataFrame(final_imputer.transform(test_X))
# Fill in the line below: get test predictions
preds_test = model.predict(final_X_test)
# -
| Kaggle_learning_Intermediate_Machine_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W0D5_Statistics/student/W0D5_Tutorial2.ipynb" target="_blank"><img alt="Open In Colab" src="https://colab.research.google.com/assets/colab-badge.svg"/></a>
# -
# # Tutorial 2: Statistical Inference
# **Week 0, Day 5: Probability & Statistics**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>
#
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Production editors:__ <NAME>, <NAME>
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
# # Tutorial Objectives
#
# This tutorial builds on Tutorial 1 by explaining how to do inference through inverting the generative process.
#
# By completing the exercises in this tutorial, you should:
# * understand what the likelihood function is, and have some intuition of why it is important
# * know how to summarise the Gaussian distribution using mean and variance
# * know how to maximise a likelihood function
# * be able to do simple inference in both classical and Bayesian ways
# * (Optional) understand how Bayes Net can be used to model causal relationships
# ---
# # Setup
# + cellView="code"
# Imports
import numpy as np
import matplotlib.pyplot as plt
import scipy as sp
from numpy.random import default_rng # a default random number generator
from scipy.stats import norm # the normal probability distribution
# -
# ## Figure settings
#
# + cellView="form" tags=["hide-input"]
#@title Figure settings
import ipywidgets as widgets # interactive display
from ipywidgets import interact, fixed, HBox, Layout, VBox, interactive, Label, interact_manual
# %config InlineBackend.figure_format = 'retina'
# plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/NMA2020/nma.mplstyle")
# -
# ## Plotting & Helper functions
#
# + cellView="form" tags=["hide-input"]
#@title Plotting & Helper functions
def plot_hist(data, xlabel, figtitle = None, num_bins = None):
""" Plot the given data as a histogram.
Args:
data (ndarray): array with data to plot as histogram
xlabel (str): label of x-axis
figtitle (str): title of histogram plot (default is no title)
num_bins (int): number of bins for histogram (default is 10)
Returns:
count (ndarray): number of samples in each histogram bin
bins (ndarray): center of each histogram bin
"""
fig, ax = plt.subplots()
ax.set_xlabel(xlabel)
ax.set_ylabel('Count')
if num_bins is not None:
count, bins, _ = plt.hist(data, max(data), bins = num_bins)
else:
count, bins, _ = plt.hist(data, max(data)) # 10 bins default
if figtitle is not None:
fig.suptitle(figtitle, size=16)
plt.show()
return count, bins
def plot_gaussian_samples_true(samples, xspace, mu, sigma, xlabel, ylabel):
""" Plot a histogram of the data samples on the same plot as the gaussian
distribution specified by the give mu and sigma values.
Args:
samples (ndarray): data samples for gaussian distribution
xspace (ndarray): x values to sample from normal distribution
mu (scalar): mean parameter of normal distribution
sigma (scalar): variance parameter of normal distribution
xlabel (str): the label of the x-axis of the histogram
ylabel (str): the label of the y-axis of the histogram
Returns:
Nothing.
"""
fig, ax = plt.subplots()
ax.set_xlabel(xlabel)
ax.set_ylabel(ylabel)
# num_samples = samples.shape[0]
count, bins, _ = plt.hist(samples, density=True) # probability density function
plt.plot(xspace, norm.pdf(xspace, mu, sigma),'r-')
plt.show()
def plot_likelihoods(likelihoods, mean_vals, variance_vals):
""" Plot the likelihood values on a heatmap plot where the x and y axes match
the mean and variance parameter values the likelihoods were computed for.
Args:
likelihoods (ndarray): array of computed likelihood values
mean_vals (ndarray): array of mean parameter values for which the
likelihood was computed
variance_vals (ndarray): array of variance parameter values for which the
likelihood was computed
Returns:
Nothing.
"""
fig, ax = plt.subplots()
im = ax.imshow(likelihoods)
cbar = ax.figure.colorbar(im, ax=ax)
cbar.ax.set_ylabel('log likelihood', rotation=-90, va="bottom")
ax.set_xticks(np.arange(len(mean_vals)))
ax.set_yticks(np.arange(len(variance_vals)))
ax.set_xticklabels(mean_vals)
ax.set_yticklabels(variance_vals)
ax.set_xlabel('Mean')
ax.set_ylabel('Variance')
def posterior_plot(x, likelihood=None, prior=None, posterior_pointwise=None, ax=None):
"""
Plots normalized Gaussian distributions and posterior.
Args:
x (numpy array of floats): points at which the likelihood has been evaluated
auditory (numpy array of floats): normalized probabilities for auditory likelihood evaluated at each `x`
visual (numpy array of floats): normalized probabilities for visual likelihood evaluated at each `x`
posterior (numpy array of floats): normalized probabilities for the posterior evaluated at each `x`
ax: Axis in which to plot. If None, create new axis.
Returns:
Nothing.
"""
if likelihood is None:
likelihood = np.zeros_like(x)
if prior is None:
prior = np.zeros_like(x)
if posterior_pointwise is None:
posterior_pointwise = np.zeros_like(x)
if ax is None:
fig, ax = plt.subplots()
ax.plot(x, likelihood, '-C1', LineWidth=2, label='Auditory')
ax.plot(x, prior, '-C0', LineWidth=2, label='Visual')
ax.plot(x, posterior_pointwise, '-C2', LineWidth=2, label='Posterior')
ax.legend()
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
plt.show()
return ax
def plot_classical_vs_bayesian_normal(num_points, mu_classic, var_classic,
mu_bayes, var_bayes):
""" Helper function to plot optimal normal distribution parameters for varying
observed sample sizes using both classic and Bayesian inference methods.
Args:
num_points (int): max observed sample size to perform inference with
mu_classic (ndarray): estimated mean parameter for each observed sample size
using classic inference method
var_classic (ndarray): estimated variance parameter for each observed sample size
using classic inference method
mu_bayes (ndarray): estimated mean parameter for each observed sample size
using Bayesian inference method
var_bayes (ndarray): estimated variance parameter for each observed sample size
using Bayesian inference method
Returns:
Nothing.
"""
xspace = np.linspace(0, num_points, num_points)
fig, ax = plt.subplots()
ax.set_xlabel('n data points')
ax.set_ylabel('mu')
plt.plot(xspace, mu_classic,'r-', label = "Classical")
plt.plot(xspace, mu_bayes,'b-', label = "Bayes")
plt.legend()
plt.show()
fig, ax = plt.subplots()
ax.set_xlabel('n data points')
ax.set_ylabel('sigma^2')
plt.plot(xspace, var_classic,'r-', label = "Classical")
plt.plot(xspace, var_bayes,'b-', label = "Bayes")
plt.legend()
plt.show()
# -
# ---
# # Section 1: Basic probability
# ## Video 1: Basic Probability
#
# + cellView="form" tags=["remove-input"]
# @title Video 1: Basic Probability
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1bw411o7HR", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="SL0_6rw8zrM", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# This video covers basic probability theory, including complementary probability, conditional probability, joint probability, and marginalisation.
#
# <details>
# <summary> <font color='blue'>Click here for text recap of video </font></summary>
#
# Previously we were only looking at sampling or properties of a single variables, but as we will now move on to statistical inference, it is useful to go over basic probability theory.
#
#
# As a reminder, probability has to be in the range 0 to 1
# $P(A) \in [0,1] $
#
# and the complementary can always be defined as
#
# $P(\neg A) = 1-P(A)$
#
#
# When we have two variables, the *conditional probability* of $A$ given $B$ is
#
# $P (A|B) = P (A \cap B)/P (B)=P (A, B)/P (B)$
#
# while the *joint probability* of $A$ and $B$ is
#
# $P(A \cap B)=P(A,B) = P(B|A)P(A) = P(A|B)P(B) $
#
# We can then also define the process of *marginalisation* (for discrete variables) as
#
# $P(A)=\sum P(A,B)=\sum P(A|B)P(B)$
#
# where the summation is over the possible values of $B$.
#
# As an example if $B$ is a binary variable that can take values $B+$ or $B0$ then
# $P(A)=\sum P(A,B)=P(A|B+)P(B+)+ P(A|B+)P(B0) $.
#
# For continuous variables marginalization is given as
# $P(A)=\int P(A,B) dB=\int P(A|B)P(B) dB$
# </details>
# ## Math Exercise 1: Probability example
#
# To remind ourselves of how to use basic probability theory we will do a short exercise (no coding needed!), based on measurement of binary probabilistic neural responses.
# As shown by Hubel and Wiesel in 1959 there are neurons in primary visual cortex that respond to different orientations of visual stimuli, with different neurons being sensitive to different orientations. The numbers in the following are however purely fictional.
#
# Imagine that your collaborator tells you that they have recorded the activity of neurons while presenting either a horizontal or vertical grid (i.e. a gabor filter). The activity of the neurons is measured as binary, active or inactive.
# After recording from a large number of neurons they find that when presenting a vertical grid on average 40 percent of neurons are active, while 30 percent respond to vertical grids.
#
# We will use the following notation to indicate the probability that a randomly chosen neuron responds to horizontal grids
#
# $P(h+)=0.4$
#
# and this to show the probability it responds to vertical:
#
# $P(v+)=0.3$
#
# We can easily see this means that the probability to not respond to horizontal ($h0$) is
#
# $P(h0)=1-P(h+)=0.6$
#
# and that the probability to not respond to vertical is
#
# $P(v0)=1-P(v+)=0.7$
#
# We will practice computing various probabilities in this framework.
#
#
#
#
#
#
#
# ### A) Product
#
# Question: Assuming that the horizontal and vertical orientation selectivity were independent, what is the probability that a randomly chosen neuron is sensitive to both horizontal and vertical orientations?
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_a8e07acd.py)
#
#
# -
# Independent here means that 𝑃(ℎ+,𝑣+) = 𝑃(ℎ+)𝑃(𝑣+)
#
# P(h+,v+) = P(h+) p(v+)=0.4*0.3=0.12
# ### B) Joint probability generally
# A collaborator informs you that actually these are not independent, for of those neurons that respond to vertical, only 10 percent also respond to horizontal, i.e. the probability of responding to horizonal *given* that it responds to vertical is $P(h+|v+)=0.1$
#
# Given this new information, what is now the probability that a randomly chosen neuron is sensitive to both horizontal and vertical orientations?
#
#
#
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_aba8ef46.py)
#
#
# -
# Remember that joint probability can generally be expressed as 𝑃(𝑎,𝑏)=𝑃(𝑎|𝑏)𝑃(𝑏)
#
# 𝑃(ℎ+,𝑣+)=𝑃(ℎ+|𝑣+)𝑃(𝑣+)=0.1∗0.3=0.03
# ### C) Conditional probability
#
# You start measuring from a neuron and find that it responds to horizontal orientations. What is now the probability that it also responds to vertical ($𝑃(v+|h+)$)?
#
#
#
#
#
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_75b17480.py)
#
#
# -
# The conditional probability is given by 𝑃(𝑎|𝑏)=𝑃(𝑎,𝑏)/𝑃(𝑏)
#
# 𝑃(𝑣+|ℎ+)=𝑃(𝑣+,ℎ+)/𝑃(ℎ+)=𝑃(ℎ+|𝑣+)𝑃(𝑣+)/𝑃(ℎ+)=0.1∗0.3/0.4=0.075
# ### D) Marginal probability
#
# Lastly, let's check that everything has been done correctly. Given our knowledge about the conditional probabilities, we should be able to use marginalisation to recover the marginal probability of a random neuron responding to vertical orientations ($P(v+)$). We know from above that this should equal 0.3.
# Calculate $P(v+)$ based on the conditional probabilities for $P(v+|h+)$ and $P(v+|h0)$ (the latter which you will need to calculate).
#
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_b63ebf79.py)
#
#
# -
# The first step is to calculute:
# 𝑃(𝑣+|ℎ0)=𝑃(ℎ0|𝑣+)𝑃(𝑣+)/𝑃(ℎ0)=(1−0.1)∗0.3/(1−0.4)=0.45
#
# Then use the property of marginalisation (discrete version)
# 𝑃(𝑎)=∑𝑖𝑃(𝑎|𝑏=𝑖)𝑃(𝑏=𝑖)
#
# 𝑃(𝑣+)=𝑃(𝑣+|ℎ+)𝑃(ℎ+)+𝑃(𝑣+|ℎ0)𝑃(ℎ0)=0.075∗0.4+0.45∗(1−0.4)=0.3
#
# Phew, we recovered the correct value!
# ## Section 1.2: Markov chains
#
#
# ### Video 2: Markov Chains
#
# +
# @title Video 2: Markov Chains
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1Rh41187ZC", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="XjQF13xMpss", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
#
# ### Coding exercise 1.2 Markov chains
#
#
# We will practice more probability theory by looking at **Markov chains**. The Markov property specifies that you can fully encapsulate the important properties of a system based on its *current* state at the current time, any previous history does not matter. It is memoryless.
#
# As an example imagine that a rat is able to move freely between 3 areas: a dark rest area
# ($state=1$), a nesting area ($state=2$) and a bright area for collecting food ($state=3$). Every 5 minutes (timepoint $i$) we record the rat's location. We can use a **categorical distribution** to look at the probability that the rat moves to one state from another.
#
# The table below shows the probability of the rat transitioning from one area to another between timepoints ($state_i$ to $state_{i+1}$).
#
# \begin{array}{|l | l | l | l |} \hline
# state_{i} &P(state_{i+1}=1|state_i=*) &P(state_{i+1}=2|state_i=*) & P(state_{i+1}=3|state=_i*) \\ \hline
# state_{i}=1& 0.2 &0.6 &0.2\\
# state_{i}=2& .6 &0.3& 0.1\\
# state_{i}=3& 0.8 &0.2 &0\\ \hline
# \end{array}
#
# We are modeling this as a Markov chain, so the animal is only in one of the states at a time and can transition between the states.
#
# We want to get the probability of each state at time $i+1$. We know from Section 1.1 that we can use marginalisation:
#
# $$P_(state_{i+1} = 1) = P(state_{i+1}=1|state_i=1)P(state_i = 1) + P(state_{i+1}=1|state_i=2)P(state_i = 2) + P(state_{i+1}=1|state_i=3)P(state_i = 3) $$
#
# Let's say we had a row vector (a vector defined as a row, not a column so matrix multiplication will work out) of the probabilities of the current state:
#
# $$P_i = [P(state_i = 1), P(state_i = 2), P(state_i = 3) ] $$
#
# If we actually know where the rat is at the current time point, this would be deterministic (e.g. $P_i = [0, 1, 0]$ if the rat is in state 2). Otherwise, this could be probabilistic (e.g. $P_i = [0.1, 0.7, 0.2]$).
#
# To compute the vector of probabilities of the state at the time $i+1$, we can use linear algebra and multiply our vector of the probabilities of the current state with the transition matrix. Recall your matrix multiplication skills from W0D3 to check this!
#
# $$P_{i+1} = P_{i} T$$
# where $T$ is our transition matrix.
#
#
# This is the same formula for every step, which allows us to get the probabilities for a time more than 1 step in advance easily. If we started at $i=0$ and wanted to look at the probabilities at step $i=2$, we could do:
#
# \begin{align*}
# P_{1} &= P_{0}T\\
# P_{2} &= P_{1}T = P_{0}TT = P_{0}T^2\\
# \end{align*}
#
# So, every time we take a further step we can just multiply with the transition matrix again. So, the probability vector of states at j timepoints after the current state at timepoint i is equal to the probability vector at timepoint i times the transition matrix raised to the jth power.
# $$P_{i + j} = P_{i}T^j $$
#
# If the animal starts in area 2, what is the probability the animal will again be in area 2 when we check on it 20 minutes (4 transitions) later?
#
# Fill in the transition matrix in the code below.
# +
###################################################################
## TODO for student
## Fill out the following then remove
# raise NotImplementedError("Student exercise: compute state probabilities after 4 transitions")
###################################################################
# Transition matrix
transition_matrix = np.array([[ 0.2, 0.6, 0.2],[ .6, 0.3, 0.1], [0.8, 0.2, 0]])
# Initial state, p0
p0 = np.array([0, 1, 0])
# Compute the probabilities 4 transitions later (use np.linalg.matrix_power to raise a matrix a power)
p4 = p0 @ np.linalg.matrix_power(transition_matrix, 4)
# The second area is indexed as 1 (Python starts indexing at 0)
print("The probability the rat will be in area 2 after 4 transitions is: " + str(p4[1]))
# -
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_ca188ef1.py)
#
#
# You should get a probability of 0.4311, i.e. there is a 43.11% chance that you will find the rat in area 2 in 20 minutes.
#
# What is the average amount of time spent by the rat in each of the states?
#
# Implicit in the question is the idea that we can start off with a random initial state and then measure how much relative time is spent in each area. If we make a few assumptions (e.g. ergodic or 'randomly mixing' system), we can instead start with an initial random distribution and see how the final probabilities of each state after many time steps (100) to estimate the time spent in each state.
# +
# Initialize random initial distribution
p_random = np.ones((1,3))/3
###################################################################
## TODO for student: Fill compute the state matrix after 100 transitions
# raise NotImplementedError("Student exercise: need to complete computation below")
###################################################################
# Fill in the missing line to get the state matrix after 100 transitions, like above
p_average_time_spent = p_random @ np.linalg.matrix_power(transition_matrix, 100)
print("The proportion of time spend by the rat in each of the three states is: "
+ str(p_average_time_spent[0]))
# -
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_84b223c9.py)
#
#
# The proportion of time spend in each of the three areas are 0.4473, 0.4211, and 0.1316, respectively.
#
# Imagine now that if the animal is satiated and tired the transitions change to:
#
# \begin{array}{|l | l | l | l |} \hline
# state_{i} &P(state_{i+1}=1|state_i=*) &P(state_{i+1}=2|state_i=*) &P(state_{i+1}=3|state_i=*) \\ \hline
# state_{i}=1& 0.2 &0.7 &0.1\\
# state_{i}=2& .3 &0.7& 0.\\
# state_{i}=3& 0.8 &0.2 &0\\ \hline
# \end{array}
#
# Try repeating the questions above for this table of transitions by changing the transition matrix. Based on the probability values, what would you predict? Check how much time the rat spends on average in each area and see if it matches your predictions.
#
# **Main course preview:** The Markov property is extremely important for many models, particularly Hidden Markov Models, discussed on day W3D2, and for methods such as Markov Chain Monte Carlo sampling.
# ---
# # Section 2: Statistical inference and likelihood
# ## Section 2.1: Likelihoods
# ### Video 3: Statistical inference and likelihood
#
# + cellView="form" tags=["remove-input"]
# @title Video 2: Statistical inference and likelihood
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1LM4y1g7wT", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="7aiKvKlYwR0", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# A generative model (such as the Gaussian distribution from the previous tutorial) allows us to make predictions about outcomes.
#
# However, after we observe $n$ data points, we can also evaluate our model (and any of its associated parameters) by calculating the **likelihood** of our model having generated each of those data points $x_i$.
#
# $$P(x_i|\mu,\sigma)=\mathcal{N}(x_i,\mu,\sigma)$$
#
# For all data points $\mathbf{x}=(x_1, x_2, x_3, ...x_n) $ we can then calculate the likelihood for the whole dataset by computing the product of the likelihood for each single data point.
#
# $$P(\mathbf{x}|\mu,\sigma)=\prod_{i=1}^n \mathcal{N}(x_i,\mu,\sigma)$$
#
# While the likelihood may be written as a conditional probability ($P(x|\mu,\sigma)$, as a function of the parameters (when the data points $\mathbf{x}$ are fixed), this is referred to as the **likelihood function**, $L(\mu,\sigma)$.
# Note that the likelihood $L(\mu,\sigma)$ is a function of $\mu$ and $\sigma$, not of $\mathbf{x}$ which we already know.
#
# In the last tutorial we reviewed how the data was generated given the selected parameters of the generative process. If we do not know the parameters $\mu$, $\sigma$ that generated the data, we can try to **infer** which parameter values (given our model) gives the best (highest) likelihood.
#
# **Correction to video**: The variance estimate that maximizes the likelihood is $\bar{\sigma}^2=\frac{1}{n} \sum_i (x_i-\bar{x})^2 $. This is a biased estimate. Shown in the video is the sample variance, which is an unbiased estimate for variance: $\bar{\sigma}^2=\frac{1}{n-1} \sum_i (x_i-\bar{x})^2 $. See section 2.2.3 for more details.
# <details>
# <summary> <font color='blue'>Click here for text recap of video </font></summary>
#
# A generative model (such as the Gaussian distribution from the previous tutorial) allows us to make predictions about outcomes.
#
# However, after we observe $n$ data points, we can also evaluate our model (and any of its associated parameters) by calculating the **likelihood** of our model having generated each of those data points $x_i$.
#
# $$P(x_i|\mu,\sigma)=\mathcal{N}(x_i,\mu,\sigma)$$
#
# For all data points $\mathbf{x}=(x_1, x_2, x_3, ...x_n) $ we can then calculate the likelihood for the whole dataset by computing the product of the likelihood for each single data point.
#
# $$P(\mathbf{x}|\mu,\sigma)=\prod_{i=1}^n \mathcal{N}(x_i,\mu,\sigma)$$
#
# </details>
#
# While the likelihood may be written as a conditional probability ($P(x|\mu,\sigma)$), we refer to it as the **likelihood function**, $L(\mu,\sigma)$. This slight switch in notation is to emphasize our focus: we use likelihood functions when the data points $\mathbf{x}$ are fixed and we are focused on the parameters.
# Our new notation makes clear that the likelihood $L(\mu,\sigma)$ is a function of $\mu$ and $\sigma$, not of $\mathbf{x}$.
#
# In the last tutorial we reviewed how the data was generated given the selected parameters of the generative process. If we do not know the parameters $\mu$, $\sigma$ that generated the data, we can try to **infer** which parameter values (given our model) gives the best (highest) likelihood. This is what we call statistical inference: trying to infer what parameters make our observed data the most likely or probable?
#
#
# ### Coding Exercise 2.1: Likelihood, mean and variance
#
#
# We can use the likelihood to find the set of parameters that are most likely to have generated the data (given the model we are using). That is, we want to infer the parameters that gave rise to the data we observed. We will try a couple of ways of doing statistical inference.
#
# In the following exercise, we will sample from the Gaussian distribution (again), plot a histogram and the Gaussian probability density function, and calculate some statistics from the samples.
#
# As a reminder, The equation for a Gaussian probability density function is:
# $$
# f(x;\mu,\sigma^2)=\mathcal{N}(\mu,\sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}}\exp\left(\frac{-(x-\mu)^2}{2\sigma^2}\right)
# $$
#
# In the exercise we will calculate:
#
# * Likelihood
# * Mean
# * Standard deviation
#
# Statistical moments are defined based on the expectations. The first moment is the expected value, i.e. the mean, the second moment is the expected squared value, i.e. variance, and so on.
#
# The special thing about the Gaussian is that mean and standard deviation of the random sample can effectively approximate the two parameters of a Gaussian, $\mu, \sigma$.
#
# Hence using the sample mean, $\bar{x}=\frac{1}{n}\sum_i x_i$, and variance, $\bar{\sigma}^2=\frac{1}{n} \sum_i (x_i-\bar{x})^2 $ should give us the best/maximum likelihood, $L(\bar{x},\bar{\sigma}^2)$.
#
# Let's see if that actually works. If we search through different combinations of $\mu$ and $\sigma$ values, do the sample mean and variance values give us the maximum likelihood (of our observed data)?
#
# As multiplying small probabilities together can lead to very small numbers, it is often convenient to report the **logarithm** of the likelihood. This is just a convenient transformation and as logarithm is a monotonically increasing function this does not change what parameters maximise the function.
#
# You need to modify two lines below to generate the data from a normal distribution $N(\mu=5, \sigma=1)$, and plot the theoretical distribution. Note that we are reusing functions from tutorial 1, so review that tutorial if needed. Then you will use this random sample to calculate the likelihood for a variety of potential mean and variance parameter values. For this tutorial we have chosen a variance parameter of 1, meaning the standard deviation is also 1 in this case. Most of our functions take the standard deviation sigma as a parameter, so we will write $\sigma = 1$.
#
# (Note that in practice computing the sample variance like this $$\bar{\sigma}^2=\frac{1}{(n-1)} \sum_i (x_i-\bar{x})^2 $$ is actually better, take a look at any statistics textbook for an explanation of this.)
# Let's start with computing the likelihood of some set of data points being drawn from a Gaussian distribution with a mean and variance we choose.
#
#
#
# As multiplying small probabilities together can lead to very small numbers, it is often convenient to report the *logarithm* of the likelihood. This is just a convenient transformation and as logarithm is a monotonically increasing function this does not change what parameters maximise the function.
# +
def compute_likelihood_normal(x, mean_val, standard_dev_val):
""" Computes the log-likelihood values given a observed data sample x, and
potential mean and variance values for a normal distribution
Args:
x (ndarray): 1-D array with all the observed data
mean_val (scalar): value of mean for which to compute likelihood
standard_dev_val (scalar): value of variance for which to compute likelihood
Returns:
likelihood (scalar): value of likelihood for this combination of means/variances
"""
###################################################################
## TODO for student
# raise NotImplementedError("Student exercise: compute likelihood")
###################################################################
# Get probability of each data point (use norm.pdf from scipy stats)
p_data = norm.pdf(x, mean_val, standard_dev_val)
# Compute likelihood (sum over the log of the probabilities)
likelihood = np.sum(np.log(p_data))
return likelihood
# Set random seed
np.random.seed(0)
# Generate data
true_mean = 5
true_standard_dev = 1
n_samples = 1000
x = np.random.normal(true_mean, true_standard_dev, size = (n_samples,))
# Compute likelihood for a guessed mean/standard dev
guess_mean = 4
guess_standard_dev = .1
likelihood = compute_likelihood_normal(x, guess_mean, guess_standard_dev)
print(likelihood)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_bd7d0ec0.py)
#
#
# -
# You should get a likelihood of -92904.81. This is somewhat meaningless to us! For it to be useful, we need to compare it to the likelihoods computing using other guesses of the mean or standard deviation. The visualization below shows us the likelihood for various values of the mean and the standard deviation. Essentially, we are performing a rough **grid-search** over means and standard deviations. What would you guess as the true mean and standard deviation based on this visualization?
# Execute to visualize likelihoods
#
# +
# @markdown Execute to visualize likelihoods
# Set random seed
np.random.seed(0)
# Generate data
true_mean = 5
true_standard_dev = 1
n_samples = 1000
x = np.random.normal(true_mean, true_standard_dev, size = (n_samples,))
# Compute likelihood for different mean/variance values
mean_vals = np.linspace(1, 10, 10) # potential mean values to ry
standard_dev_vals = np.array([0.7, 0.8, 0.9, 1, 1.2, 1.5, 2, 3, 4, 5]) # potential variance values to try
# Initialise likelihood collection array
likelihood = np.zeros((mean_vals.shape[0], standard_dev_vals.shape[0]))
# Compute the likelihood for observing the gvien data x assuming
# each combination of mean and variance values
for idxMean in range(mean_vals.shape[0]):
for idxVar in range(standard_dev_vals .shape[0]):
likelihood[idxVar,idxMean]= sum(np.log(norm.pdf(x, mean_vals[idxMean],
standard_dev_vals[idxVar])))
# Uncomment once you've generated the samples and compute likelihoods
xspace = np.linspace(0, 10, 100)
plot_likelihoods(likelihood, mean_vals, standard_dev_vals)
# -
#
# At the top you should see the sample mean and variance values, which are close to the true values (that we happen to know here).
#
# Underneath, the top figure shows hopefully a nice fit between the histogram and the distribution that generated the data. So far so good.
#
# In the heatmap we should be able to see that the mean and variance parameters values yielding the highest likelihood (yellow) corresponds to (roughly) the combination of the calculated sample mean and variance from the dataset.
# But it can be hard to see from such a rough **grid-search** simulation, as it is only as precise as the resolution of the grid we are searching.
#
# Implicitly, by looking for the parameters that give the highest likelihood, we have been searching for the **maximum likelihood** estimate.
# $$(\hat{\mu},\hat{\sigma})=argmax_{\mu,\sigma}L(\mu,\sigma)=argmax_{\mu,\sigma} \prod_{i=1}^n \mathcal{N}(x_i,\mu,\sigma)$$.
#
# For a simple Gaussian this can actually be done analytically (you have likely already done so yourself), using the statistical moments: mean and standard deviation (variance).
#
# In next section we will look at other ways of inferring such parameter variables.
# ## Section 2.2: Maximum likelihood
# ### Video 4: Maximum likelihood
#
# + cellView="form" tags=["remove-input"]
# @title Video 3: Maximum likelihood
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1Lo4y1C7xy", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="Fuwx_V64nEU", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
#
# Implicitly, by looking for the parameters that give the highest likelihood in the last section, we have been searching for the **maximum likelihood** estimate.
# $$(\hat{\mu},\hat{\sigma})=argmax_{\mu,\sigma}L(\mu,\sigma)=argmax_{\mu,\sigma} \prod_{i=1}^n \mathcal{N}(x_i,\mu,\sigma)$$.
#
#
#
# In next sections, we will look at other ways of inferring such parameter variables.
# ### Section 2.2.1: Searching for best parameters
#
# We want to do inference on this data set, i.e. we want to infer the parameters that most likely gave rise to the data given our model. Intuitively that means that we want as good as possible a fit between the observed data and the probability distribution function with the best inferred parameters. We can search for the best parameters manually by trying out a bunch of possible values of the parameters, computing the likelihoods, and picking the parameters that resulted in the highest likelihood.
# #### Interactive Demo 2.2: Maximum likelihood inference
#
# Try to see how well you can fit the probability distribution to the data by using the demo sliders to control the mean and standard deviation parameters of the distribution. We will visualize the histogram of data points (in blue) and the Gaussian density curve with that mean and standard deviation (in red). Below, we print the log-likelihood.
#
# - What (approximate) values of mu and sigma result in the best fit?
# - How does the value below the plot (the log-likelihood) change with the quality of fit?
# Make sure you execute this cell to enable the widget and fit by hand!
#
# + cellView="form" tags=["hide-input"]
# @markdown Make sure you execute this cell to enable the widget and fit by hand!
# Generate data
true_mean = 5
true_standard_dev = 1
n_samples = 1000
vals = np.random.normal(true_mean, true_standard_dev, size = (n_samples,))
def plotFnc(mu,sigma):
loglikelihood= sum(np.log(norm.pdf(vals,mu,sigma)))
#calculate histogram
#prepare to plot
fig, ax = plt.subplots()
ax.set_xlabel('x')
ax.set_ylabel('probability')
#plot histogram
count, bins, ignored = plt.hist(vals,density=True)
x = np.linspace(0,10,100)
#plot pdf
plt.plot(x, norm.pdf(x,mu,sigma),'r-')
plt.show()
print("The log-likelihood for the selected parameters is: " + str(loglikelihood))
#interact(plotFnc, mu=5.0, sigma=2.1);
#interact(plotFnc, mu=widgets.IntSlider(min=0.0, max=10.0, step=1, value=4.0),sigma=widgets.IntSlider(min=0.1, max=10.0, step=1, value=4.0));
interact(plotFnc, mu=(0.0, 15.0, 0.1),sigma=(0.1, 5.0, 0.1));
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_6338abeb.py)
#
#
# -
# - The log-likelihood should be greatest when 𝜇 = 5 and 𝜎 = 1.
# - The summed log-liklihood increases (becomes less negative) as the fit improves
# Doing this was similar to the grid searched image from Section 2.1. Really, we want to see if we can do inference on observed data in a bit more principled way.
#
# ### Section 2.2.2: Optimization to find parameters
#
# Let's again assume that we have a data set, $\mathbf{x}$, assumed to be generated by a normal distribution (we actually generate it ourselves in line 1, so we know how it was generated!).
# We want to maximise the likelihood of the parameters $\mu$ and $\sigma^2$. We can do so using a couple of tricks:
#
# * Using a log transform will not change the maximum of the function, but will allow us to work with very small numbers that could lead to problems with machine precision.
# * Maximising a function is the same as minimising the negative of a function, allowing us to use the minimize optimisation provided by scipy.
#
# The optimisation will be done using `sp.optimize.minimize`, which does a version of gradient descent (there are hundreds of ways to do numerical optimisation, we will not cover these here!).
# #### Coding Exercise 2.2: Maximum Likelihood Estimation
#
#
# In the code below, insert the missing line (see the `compute_likelihood_normal` function from previous exercise), with the mean as theta[0] and variance as theta[1].
#
# +
# We define the function to optimise, the negative log likelihood
def negLogLike(theta, x):
""" Function for computing the negative log-likelihood given the observed data
and given parameter values stored in theta.
Args:
theta (ndarray): normal distribution parameters (mean is theta[0],
variance is theta[1])
x (ndarray): array with observed data points
Returns:
Calculated negative Log Likelihood value!
"""
###################################################################
## TODO for students: Compute the negative log-likelihood value for the
## given observed data values and parameters (theta)
# Fill out the following then remove
# raise NotImplementedError("Student exercise: need to compute the negative \
# log-likelihood value")
###################################################################
return -sum(np.log(norm.pdf(x, theta[0], theta[1])))
# Set random seed
np.random.seed(0)
# Generate data
true_mean = 5
true_standard_dev = 1
n_samples = 1000
x = np.random.normal(true_mean, true_standard_dev, size = (n_samples,))
# Define bounds, var has to be positive
bnds = ((None, None), (0, None))
# Optimize with scipy!
optimal_parameters = sp.optimize.minimize(negLogLike, (2, 2), args = x, bounds = bnds)
print("The optimal mean estimate is: " + str(optimal_parameters.x[0]))
print("The optimal variance estimate is: " + str(optimal_parameters.x[1]))
# optimal_parameters contains a lot of information about the optimization,
# but we mostly want the mean and variance
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_cf9bfae6.py)
#
#
# -
# These are the approximations of the parameters that maximise the likelihood ($\mu$ ~ 5.281 and $\sigma$ ~ 1.170).
# + [markdown] colab_type="text"
# ### Section 2.2.3: Analytical solution
#
# Sometimes, things work out well and we can come up with formulas for the maximum likelihood estimates of parameters. We won't get into this further but basically we could **set the derivative of the likelihood to 0** (to find a maximum) and solve for the parameters. This won't always work but for the Gaussian distribution, it does.
#
# Specifically , the special thing about the Gaussian is that mean and standard deviation of the random sample can effectively approximate the two parameters of a Gaussian, $\mu, \sigma$.
#
#
# Hence using the mean, $\bar{x}=\frac{1}{n}\sum_i x_i$, and variance, $\bar{\sigma}^2=\frac{1}{n} \sum_i (x_i-\bar{x})^2 $ of the sample should give us the best/maximum likelihood, $L(\bar{x},\bar{\sigma}^2)$.
#
# Let's compare these values to those we've been finding using manual search and optimization, and the true values (which we only know because we generated the numbers!).
#
#
# +
# Set random seed
np.random.seed(0)
# Generate data
true_mean = 5
true_standard_dev = 1
n_samples = 1000
x = np.random.normal(true_mean, true_standard_dev, size = (n_samples,))
# Compute and print sample means and standard deviations
print("This is the sample mean as estimated by numpy: " + str(np.mean(x)))
print("This is the sample standard deviation as estimated by numpy: " + str(np.std(x)))
# -
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_696ddd33.py)
#
#
# You should notice that the parameters estimated by maximum likelihood
# estimation/inference are very close to the true parameters (mu = 5, sigma = 1),
# as well as the parameters visualized to be best after Coding Exercise 2.1,
# where all likelihood values were calculated explicitly.
# If you try out different values of the mean and standard deviation in all the previous exercises, you should see that changing the mean and
# sigma parameter values (and generating new data from a distribution with theseparameters) makes no difference as MLE methods can still recover these parameters.
#
# There is a slight problem: it turns out that the maximum likelihood estimate for the variance is actually a biased one! This means that the estimators expected value (mean value) and the true value of the parameter are different. An unbiased estimator for the variance is $\bar{\sigma}^2=\frac{1}{n-1} \sum_i (x_i-\bar{x})^2 $, this is called the sample variance. For more details, see [the wiki page on bias of estimators](https://en.wikipedia.org/wiki/Bias_of_an_estimator).
# ---
# # Section 3: Bayesian Inference
# ## Section 3.1: Bayes
# ### Video 5: Bayesian inference with Gaussian distribution
#
# + cellView="form" tags=["remove-input"]
# @title Video 4: Bayesian inference with Gaussian distribution
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV11K4y1u7vH", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="1Q3VqcpfvBk", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# We will start to introduce Bayesian inference here to contrast with our maximum likelihood methods, but you will also revisit Bayesian inference in great detail on W3D1 of the course so we won't dive into all details.
#
# For Bayesian inference we do not focus on the likelihood function $L(y)=P(x|y)$, but instead focus on the posterior distribution:
#
# $$P(y|x)=\frac{P(x|y)P(y)}{P(x)}$$
#
# which is composed of the **likelihood** function $P(x|y)$, the **prior** $P(y)$ and a normalising term $P(x)$ (which we will ignore for now).
#
# While there are other advantages to using Bayesian inference (such as the ability to derive Bayesian Nets, see optional bonus task below), we will start by focusing on the role of the prior in inference. Does including prior information allow us to infer parameters in a better way?
# ### Think! 3.1: Bayesian inference with Gaussian distribution
#
# In the above sections we performed inference using maximum likelihood, i.e. finding the parameters that maximised the likelihood of a set of parameters, given the model and data.
#
# We will now repeat the inference process, but with **an added Bayesian prior**, and compare it to the "classical" inference (maximum likelihood) process we did before (Section 2). When using conjugate priors (more on this below) we can just update the parameter values of the distributions (here Gaussian distributions).
#
#
# For the prior we start by guessing a mean of 5 (mean of previously observed data points 4 and 6) and variance of 1 (variance of 4 and 6). We use a trick (not detailed here) that is a simplified way of applying a prior, that allows us to just add these 2 values (pseudo-data) to the real data.
#
# See the visualization below that shows the mean and standard deviation inferred by our classical maximum likelihood approach and the Bayesian approach for different numbers of data points.
#
# Remembering that our true values are $\mu = 5$, and $\sigma^2 = 1$, how do the Bayesian inference and classical inference compare?
# Execute to visualize inference
#
# +
# @markdown Execute to visualize inference
def classic_vs_bayesian_normal(mu, sigma, num_points, prior):
""" Compute both classical and Bayesian inference processes over the range of
data sample sizes (num_points) for a normal distribution with parameters
mu,sigma for comparison.
Args:
mu (scalar): the mean parameter of the normal distribution
sigma (scalar): the standard deviation parameter of the normal distribution
num_points (int): max number of points to use for inference
prior (ndarray): prior data points for Bayesian inference
Returns:
mean_classic (ndarray): estimate mean parameter via classic inference
var_classic (ndarray): estimate variance parameter via classic inference
mean_bayes (ndarray): estimate mean parameter via Bayesian inference
var_bayes (ndarray): estimate variance parameter via Bayesian inference
"""
# Initialize the classical and Bayesian inference arrays that will estimate
# the normal parameters given a certain number of randomly sampled data points
mean_classic = np.zeros(num_points)
var_classic = np.zeros(num_points)
mean_bayes = np.zeros(num_points)
var_bayes = np.zeros(num_points)
for nData in range(num_points):
random_num_generator = default_rng(0)
x = random_num_generator.normal(mu, sigma, nData + 1)
# Compute the mean of those points and set the corresponding array entry to this value
mean_classic[nData] = np.mean(x)
# Compute the variance of those points and set the corresponding array entry to this value
var_classic[nData] = np.var(x)
# Bayesian inference with the given prior is performed below for you
xsupp = np.hstack((x, prior))
mean_bayes[nData] = np.mean(xsupp)
var_bayes[nData] = np.var(xsupp)
return mean_classic, var_classic, mean_bayes, var_bayes
# Set random seed
np.random.seed(0)
# Set normal distribution parameters, mu and sigma
mu = 5
sigma = 1
# Set the prior to be two new data points, 4 and 6, and print the mean and variance
prior = np.array((4, 6))
print("The mean of the data comprising the prior is: " + str(np.mean(prior)))
print("The variance of the data comprising the prior is: " + str(np.var(prior)))
mean_classic, var_classic, mean_bayes, var_bayes = classic_vs_bayesian_normal(mu, sigma, 60, prior)
plot_classical_vs_bayesian_normal(60, mean_classic, var_classic, mean_bayes, var_bayes)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_f615551e.py)
#
#
# -
# Hopefully you can see that the blue line stays a little closer to the true values ($\mu=5$, $\sigma^2=1$).
#
# Having a simple prior in the Bayesian inference process (blue) helps to regularise
# the inference of the mean and variance parameters when you have very little data,
# but has little effect with large data sets. You can see that as the number of data points
# (x-axis) increases, both inference processes (blue and red lines) get closer and closer
# together, i.e. their estimates for the true parameters converge as sample size increases.
#
#
# Note that the prior is only beneficial when it is close to the true value, i.e. 'a good guess' (or at least not a bad guess). As we will see in the next exercise, if you have a prior/bias that is very wrong, your inference will start off very wrong!
#
# ## Section 3.2: Conjugate priors
# ### Video 6: Conjugate priors
#
# + cellView="form" tags=["remove-input"]
# @title Video 5: Conjugate priors
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1Hg41137Zr", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="mDEyZHaG5aY", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ### Interactive Demo 3.2: Conjugate priors
# Let's return to our example from Tutorial 1 using the binomial distribution - rat in a T-maze.
#
# Bayesian inference can be used for any likelihood distribution, but it is a lot more convenient to work with **conjugate** priors, where multiplying the prior with the likelihood just provides another instance of the prior distribution with updated values.
#
# For the binomial likelihood it is convenient to use the **beta** distribution as a prior
#
# \begin{aligned}f(p;\alpha ,\beta )={\frac {1}{\mathrm {B} (\alpha ,\beta )}}p^{\alpha -1}(1-p)^{\beta -1}\end{aligned}
# where $B$ is the beta function, $\alpha$ and $\beta$ are parameters, and $p$ is the probability of the rat turning left or right. The beta distribution is thus a distribution over a probability.
#
# Given a series of Left and Right moves of the rat, we can now estimate the probability that the animal will turn left. Using Bayesian Inference, we use a beta distribution *prior*, which is then multiplied with the *likelihood* to create a *posterior* that is also a beta distribution, but with updated parameters (we will not cover the math here).
#
# Activate the widget below to explore the variables, and follow the instructions below.
# Make sure you execute this cell to enable the widget
#
# + cellView="form" tags=["hide-input"]
#@title
#@markdown Make sure you execute this cell to enable the widget
#beta distribution
#and binomial
def plotFnc(p,n,priorL,priorR):
# Set random seed
np.random.seed(1)
#sample from binomial
numL = np.random.binomial(n, p, 1)
numR = n - numL
stepSize=0.001
x = np.arange(0, 1, stepSize)
betaPdf=sp.stats.beta.pdf(x,numL+priorL,numR+priorR)
betaPrior=sp.stats.beta.pdf(x,priorL,priorR)
print("number of left "+str(numL))
print("number of right "+str(numR))
print(" ")
print("max likelihood "+str(numL/(numL+numR)))
print(" ")
print("max posterior " + str(x[np.argmax(betaPdf)]))
print("mean posterior " + str(np.mean(betaPdf*x)))
print(" ")
with plt.xkcd():
#rng.beta()
fig, ax = plt.subplots()
plt.rcParams.update({'font.size': 22})
ax.set_xlabel('p')
ax.set_ylabel('probability density')
plt.plot(x,betaPdf, label = "Posterior")
plt.plot(x,betaPrior, label = "Prior")
#print(int(len(betaPdf)/2))
plt.legend()
interact(plotFnc, p=(0, 1, 0.01),n=(1, 50, 1), priorL=(1, 10, 1),priorR=(1, 10, 1));
# -
# The plot above shows you the prior distribution (i.e. before any data) and the posterior distribution (after data), with a summary of the data (number of left and right moves) and the maximum likelihood, maximum posterior and mean of the posterior. Dependent on the purpose either the mean or the max of the posterior can be useful as a 'single-number' summary of the posterior.
# Once you are familiar with the sliders and what they represent, go through these instructions.
#
# **For $p=0.5$**
#
# - Set $p=0.5$ and start off with a "flat" prior (`priorL=0`, `priorR=0`). Note that the prior distribution (orange) is flat, also known as uniformative. In this case the maximum likelihood and maximum posterior will get you almost identical results as you vary the number of datapoints ($n$) and the probability of the rat going left. However the posterior is a full distribution and not just a single point estimate.
#
# - As $n$ gets large you will also notice that the estimate (max likelihood or max posterior) changes less for each change in $n$, i.e. the estimation stabilises.
#
# - How many data points do you need think is needed for the probability estimate to stabilise? Note that this depends on how large fluctuations you are willing to accept.
#
# - Try increasing the strength of the prior, `priorL=10` and `priorR=10`. You will see that the prior distribution becomes more 'peaky'. In short this prior means that small or large values of $p$ are conidered very unlikely. Try playing with the number of data points $n$, you should find that the prior stabilises/regularises the maximum posterior estimate so that it does not move as much.
#
# **For $p=0.2$**
#
# Try the same as you just did, now with $p=0.2$,
# do you notice any differences? Note that the prior (assumeing equal chance Left and Right) is now badly matched to the data. Do the maximum likelihood and maximum posterior still give similar results, for a weak prior? For a strong prior? Does the prior still have a stabilising effect on the estimate?
#
#
# **Take-away message:**
# Bayesian inference gives you a full distribution over the variables that you are inferring, can help regularise inference when you have limited data, and allows you to build more complex models that better reflects true causality (see bonus below).
# ### Think! 3.2: Bayesian Brains
# Bayesian inference can help you when doing data analysis, especially when you only have little data. But consider whether the brain might be able to benefit from this too. If the brain needs to make inferences about the world, would it be useful to do regularisation on the input? Maybe there are times where having a full probability distribution could be useful?
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_f60d4b5d.py)
#
#
# -
# You will learn more about "Bayesian brains" and the theory surrounding
# these ideas once the course begins. Here is a brief explanation: it may
# be ideal for human brains to implement Bayesian inference by integrating "prior"
# information the brain has about the world (memories, prior knowledge, etc.) with
# new evidence that updates its "beliefs"/prior. This process seems to parallel
# the brain's method of learning about its environment, making it a compelling
# theory for many neuroscience researchers. One of Bonus exercises below examines a possible
# real world model for Bayesian inference: sound localization.
# ---
# # Summary
#
# ## Video 7: Summary
#
# + cellView="form" tags=["remove-input"]
# @title Video 6: Summary
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1qB4y1K7WZ", width=730, height=410, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="OJN7ri3_FCA", width=730, height=410, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
#
# Having done the different exercises you should now:
# * understand what the likelihood function is, and have some intuition of why it is important
# * know how to summarise the Gaussian distribution using mean and variance
# * know how to maximise a likelihood function
# * be able to do simple inference in both classical and Bayesian ways
#
# For more resources see
# https://github.com/NeuromatchAcademy/precourse/blob/master/resources.md
# ---
# # Bonus
# ## Bonus Coding Exercise 1: Finding the posterior computationally
#
# Imagine an experiment where participants estimate the location of a noise-emitting object. To estimate its position, the participants can use two sources of information:
# 1. new noisy auditory information (the likelihood)
# 2. prior visual expectations of where the stimulus is likely to come from (visual prior).
#
# The auditory and visual information are both noisy, so participants will combine these sources of information to better estimate the position of the object.
#
# We will use Gaussian distributions to represent the auditory likelihood (in red), and a Gaussian visual prior (expectations - in blue). Using Bayes rule, you will combine them into a posterior distribution that summarizes the probability that the object is in each possible location.
#
# We have provided you with a ready-to-use plotting function, and a code skeleton.
#
# * You can use `my_gaussian` from Tutorial 1 (also included below), to generate an auditory likelihood with parameters $\mu$ = 3 and $\sigma$ = 1.5
# * Generate a visual prior with parameters $\mu$ = -1 and $\sigma$ = 1.5
# * Calculate the posterior using pointwise multiplication of the likelihood and prior. Don't forget to normalize so the posterior adds up to 1
# * Plot the likelihood, prior and posterior using the predefined function `posterior_plot`
#
#
# +
def my_gaussian(x_points, mu, sigma):
""" Returns normalized Gaussian estimated at points `x_points`, with parameters:
mean `mu` and standard deviation `sigma`
Args:
x_points (ndarray of floats): points at which the gaussian is evaluated
mu (scalar): mean of the Gaussian
sigma (scalar): standard deviation of the gaussian
Returns:
(numpy array of floats) : normalized Gaussian evaluated at `x`
"""
px = 1/(2*np.pi*sigma**2)**1/2 *np.exp(-(x_points-mu)**2/(2*sigma**2))
# as we are doing numerical integration we may have to remember to normalise
# taking into account the stepsize (0.1)
px = px/(0.1*sum(px))
return px
def compute_posterior_pointwise(prior, likelihood):
""" Compute the posterior probability distribution point-by-point using Bayes
Rule.
Args:
prior (ndarray): probability distribution of prior
likelihood (ndarray): probability distribution of likelihood
Returns:
posterior (ndarray): probability distribution of posterior
"""
##############################################################################
# TODO for students: Write code to compute the posterior from the prior and
# likelihood via pointwise multiplication. (You may assume both are defined
# over the same x-axis)
#
# Comment out the line below to test your solution
# raise NotImplementedError("Finish the simulation code first")
##############################################################################
posterior = likelihood * prior
posterior = posterior / (0.1 * posterior.sum())
return posterior
def localization_simulation(mu_auditory = 3.0, sigma_auditory = 1.5,
mu_visual = -1.0, sigma_visual = 1.5):
""" Perform a sound localization simulation with an auditory prior.
Args:
mu_auditory (float): mean parameter value for auditory prior
sigma_auditory (float): standard deviation parameter value for auditory
prior
mu_visual (float): mean parameter value for visual likelihood distribution
sigma_visual (float): standard deviation parameter value for visual
likelihood distribution
Returns:
x (ndarray): range of values for which to compute probabilities
auditory (ndarray): probability distribution of the auditory prior
visual (ndarray): probability distribution of the visual likelihood
posterior_pointwise (ndarray): posterior probability distribution
"""
##############################################################################
## Using the x variable below,
## create a gaussian called 'auditory' with mean 3, and std 1.5
## create a gaussian called 'visual' with mean -1, and std 1.5
#
#
## Comment out the line below to test your solution
# raise NotImplementedError("Finish the simulation code first")
###############################################################################
x = np.arange(-8, 9, 0.1)
auditory = my_gaussian(x, mu_auditory, sigma_auditory)
visual = my_gaussian(x, mu_visual, mu_visual)
posterior = compute_posterior_pointwise(auditory, visual)
return x, auditory, visual, posterior
# Uncomment the lines below to plot the results
x, auditory, visual, posterior_pointwise = localization_simulation()
_ = posterior_plot(x, auditory, visual, posterior_pointwise)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_4e7f8943.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=560 height=415 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W0D5_Statistics/static/W0D5_Tutorial2_Solution_4e7f8943_1.png>
#
#
# -
# Combining the the visual and auditory information could help the brain get a better estimate of the location of an audio-visual object, with lower variance.
#
# **Main course preview:** On Week 3 Day 1 (W3D1) there will be a whole day devoted to examining whether the brain uses Bayesian inference. Is the brain Bayesian?!
# ## Bonus Coding Exercise 2: Bayes Net
# If you have the time, here is another extra exercise.
#
# Bayes Net, or Bayesian Belief Networks, provide a way to make inferences about multiple levels of information, which would be very difficult to do in a classical frequentist paradigm.
#
# We can encapsulate our knowledge about causal relationships and use this to make inferences about hidden properties.
# We will try a simple example of a Bayesian Net (aka belief network). Imagine that you have a house with an unreliable sprinkler system installed for watering the grass. This is set to water the grass independently of whether it has rained that day. We have three variables, rain ($r$), sprinklers ($s$) and wet grass ($w$). Each of these can be true (1) or false (0). See the graphical model representing the relationship between the variables.
# 
# There is a table below describing all the relationships between $w, r$, and s$.
#
# Obviously the grass is more likely to be wet if either the sprinklers were on or it was raining. On any given day the sprinklers have probability 0.25 of being on, $P(s = 1) = 0.25$, while there is a probability 0.1 of rain, $P (r = 1) = 0.1$. The table then lists the conditional probabilities for the given being wet, given a rain and sprinkler condition for that day.
# \begin{array}{|l | l || ll |} \hline
# r &s&P(w=0|r,s) &P(w=1|r,s)$\\ \hline
# 0& 0 &0.999 &0.001\\
# 0& 1 &0.1& 0.9\\
# 1& 0 &0.01 &0.99\\
# 1& 1& 0.001 &0.999\\ \hline
# \end{array}
#
#
# You come home and find that the the grass is wet, what is the probability the sprinklers were on today (you do not know if it was raining)?
#
# We can start by writing out the joint probability:
# $P(r,w,s)=P(w|r,s)P(r)P(s)$
#
# The conditional probability is then:
#
# $
# P(s|w)=\frac{\sum_{r} P(w|s,r)P(s) P(r)}{P(w)}=\frac{P(s) \sum_{r} P(w|s,r) P(r)}{P(w)}
# $
#
# Note that we are summing over all possible conditions for $r$ as we do not know if it was raining. Specifically, we want to know the probability of sprinklers having been on given the wet grass, $P(s=1|w=1)$:
#
# $
# P(s=1|w=1)=\frac{P(s = 1)( P(w = 1|s = 1, r = 1) P(r = 1)+ P(w = 1|s = 1,r = 0) P(r = 0))}{P(w = 1)}
# $
#
# where
#
# \begin{eqnarray}
# P(w=1)=P(s=1)( P(w=1|s=1,r=1 ) P(r=1) &+ P(w=1|s=1,r=0) P(r=0))\\
# +P(s=0)( P(w=1|s=0,r=1 ) P(r=1) &+ P(w=1|s=0,r=0) P(r=0))\\
# \end{eqnarray}
#
# This code has been written out below, you just need to insert the right numbers from the table.
# +
##############################################################################
# TODO for student: Write code to insert the correct conditional probabilities
# from the table; see the comments to match variable with table entry.
# Comment out the line below to test your solution
# raise NotImplementedError("Finish the simulation code first")
##############################################################################
Pw1r1s1 = 0.999 # the probability of wet grass given rain and sprinklers on
Pw1r1s0 = 0.99 # the probability of wet grass given rain and sprinklers off
Pw1r0s1 = 0.9 # the probability of wet grass given no rain and sprinklers on
Pw1r0s0 = 0.001 # the probability of wet grass given no rain and sprinklers off
Ps = 0.25 # the probability of the sprinkler being on
Pr = 0.1 # the probability of rain that day
# Uncomment once variables are assigned above
A= Ps * (Pw1r1s1 * Pr + (Pw1r0s1) * (1 - Pr))
B= (1 - Ps) * (Pw1r1s0 *Pr + (Pw1r0s0) * (1 - Pr))
print("Given that the grass is wet, the probability the sprinkler was on is: " +
str(A/(A + B)))
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W0D5_Statistics/solutions/W0D5_Tutorial2_Solution_204db048.py)
#
#
# -
# The probability you should get is about 0.7522.
#
# Your neighbour now tells you that it was indeed
# raining today, $P (r = 1) = 1$, so what is now the probability the sprinklers were on? Try changing the numbers above.
#
#
# ## Bonus Think!: Causality in the Brain
#
# In a causal stucture this is the correct way to calculate the probabilities. Do you think this is how the brain solves such problems? Would it be different for task involving novel stimuli (e.g. for someone with no previous exposure to sprinklers), as opposed to common stimuli?
#
# **Main course preview:** On W3D5 we will discuss causality further!
| W0/W0D5/tutorials/tutorial2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python3_7_6
# language: python
# name: py3_7_6
# ---
# # 직사각형 별찍기
# https://programmers.co.kr/learn/courses/30/lessons/12969
#
#
# ### 문제 설명
# - 이 문제에는 표준 입력으로 두 개의 정수 n과 m이 주어집니다.
# - 별(*) 문자를 이용해 가로의 길이가 n, 세로의 길이가 m인 직사각형 형태를 출력해보세요.
#
#
# ### 제한 조건
# - n과 m은 각각 1000 이하인 자연수입니다.
#
#
# ### 예시
# - 입력: 5 3
# - 출력:
# +
# *****
# *****
# *****
# -
# ### 나의 풀이:
a, b = map(int, input().strip().split(' '))
for i in range(b):
print('*'*int(a))
# ### 다른 사람 풀이:
a, b = map(int, input().strip().split(' '))
answer = ('*'*a +'\n')*b
print(answer)
| programmers.co.kr/level 1/[Algorithm] programmers_level 1_print star_2021.06.02.wed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cedeerwe/brutalna-akademia/blob/master/notebooks/03_spionske_hry.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="FWdf_UTa4CDd" colab_type="text"
# ## Inštrukcie
#
# Pred začiatkom vás poprosím spustiť príkaz z hornej lišty `Runtime -> Run All` alebo `Ctrl + F9`.
# Pre spustenie ľubovoľnej bunky kódu stlačte `Ctrl + Enter` alebo šípku vľavo od samotnej bunky.
# Toto prostredie obashuje veľa rôznych skratiek, ktoré sa spúšťajú stlačením kombinácie `Ctrl + M` a následne nejaké ďalšie tlačítko. Napríklad pomocníka na používanie tohto prostredia nájdete po stlačení `Ctrl + M` a následne `h`.
#
# Pri názvoch sekcií často nájdete nápis `1 cell hidden` alebo niečo podobné. Text pred nimi vám vždy prezradí, čo sa v nich nachádza. Rozbalte ich iba ak chcete pokračovať ďalej, sú zabalené napríklad preto, aby ste rovno nevideli riešenia a mali ste čas sa zamyslieť.
# + [markdown] id="pI82NOKd4a0g" colab_type="text"
# # Pomocný kód
#
#
# + id="20aLw8wcrQYB" colab_type="code" colab={}
import pandas as pd
table1 = pd.DataFrame({
"HH": {
"HH": 1 / 2,
"HZ": 1 / 2,
"ZH": 3 / 4,
"ZZ": 1 / 2
},
"HZ": {
"HH": 1 / 2,
"HZ": 1 / 2,
"ZH": 1 / 2,
"ZZ": 1 / 4
},
"ZH": {
"HH": 1 / 4,
"HZ": 1 / 2,
"ZH": 1 / 2,
"ZZ": 1 / 2
},
"ZZ": {
"HH": 1 / 2,
"HZ": 3 / 4,
"ZH": 1 / 2,
"ZZ": 1 / 2
},
}).round(3)
table2 = pd.DataFrame({
"HHH": {
"HHH": 1 / 2,
"HHZ": 1 / 2,
"HZH": 3 / 5,
"HZZ": 3 / 5,
"ZHH": 7 / 8,
"ZHZ": 7 / 12,
"ZZH": 7 / 10,
"ZZZ": 1 / 2,
},
"HHZ": {
"HHH": 1 / 2,
"HHZ": 1 / 2,
"HZH": 1 / 3,
"HZZ": 1 / 3,
"ZHH": 3 / 4,
"ZHZ": 3 / 8,
"ZZH": 1 / 2,
"ZZZ": 3 / 10,
},
"HZH": {
"HHH": 2 / 5,
"HHZ": 2 / 3,
"HZH": 1 / 2,
"HZZ": 1 / 2,
"ZHH": 1 / 2,
"ZHZ": 1 / 2,
"ZZH": 5 / 8,
"ZZZ": 5 / 12,
},
"HZZ": {
"HHH": 2 / 5,
"HHZ": 2 / 3,
"HZH": 1 / 2,
"HZZ": 1 / 2,
"ZHH": 1 / 2,
"ZHZ": 1 / 2,
"ZZH": 1 / 4,
"ZZZ": 1 / 8,
},
"ZHH": {
"HHH": 1 / 8,
"HHZ": 1 / 4,
"HZH": 1 / 2,
"HZZ": 1 / 2,
"ZHH": 1 / 2,
"ZHZ": 1 / 2,
"ZZH": 2 / 3,
"ZZZ": 2 / 5,
},
"ZHZ": {
"HHH": 5 / 12,
"HHZ": 5 / 8,
"HZH": 1 / 2,
"HZZ": 1 / 2,
"ZHH": 1 / 2,
"ZHZ": 1 / 2,
"ZZH": 2 / 3,
"ZZZ": 2 / 5,
},
"ZZH": {
"HHH": 3 / 10,
"HHZ": 1 / 2,
"HZH": 3 / 8,
"HZZ": 3 / 4,
"ZHH": 1 / 3,
"ZHZ": 1 / 3,
"ZZH": 1 / 2,
"ZZZ": 1 / 2,
},
"ZZZ": {
"HHH": 1 / 2,
"HHZ": 7 / 10,
"HZH": 7 / 12,
"HZZ": 7 / 8,
"ZHH": 3 / 5,
"ZHZ": 3 / 5,
"ZZH": 1 / 2,
"ZZZ": 1 / 2,
}
}).round(3)
table3 = pd.DataFrame({
"HHZ": {
"HHZ": 1 / 2,
"HZZ": 1 / 3,
"ZHH": 3 / 4,
"ZZH": 1 / 2,
},
"HZZ": {
"HHZ": 2 / 3,
"HZZ": 1 / 2,
"ZHH": 1 / 2,
"ZZH": 1 / 4,
},
"ZHH": {
"HHZ": 1 / 4,
"HZZ": 1 / 2,
"ZHH": 1 / 2,
"ZZH": 2 / 3,
},
"ZZH": {
"HHZ": 1 / 2,
"HZZ": 3 / 4,
"ZHH": 1 / 3,
"ZZH": 1 / 2,
}
}).round(3)
table4 = pd.DataFrame({
"HHZ": {
"HHZ": 0,
"HZZ": -2,
"ZHH": 3,
"ZZH": 0,
},
"HZZ": {
"HHZ": 2,
"HZZ": 0,
"ZHH": 0,
"ZZH": -3,
},
"ZHH": {
"HHZ": -3,
"HZZ": 0,
"ZHH": 0,
"ZZH": 2,
},
"ZZH": {
"HHZ": 0,
"HZZ": 3,
"ZHH": -2,
"ZZH": 0,
}
})
# + [markdown] id="RpkC7Jbg4eWE" colab_type="text"
# # Úloha č.1
#
# Poznáte ten pocit, idete si po ulici a zrazu máte dojem, že vás niekto špehuje. Že vás niekto nasleduje. Že nie ste v bezpečí ani vo vlastných myšlienkach, lebo vám do nich vidia. A vy máte strach, bojíte sa, lebo už zajtra sa idete zúčastniť majstrovstiev sveta v stratégiách na hru kameň-papier-nožnice. Ako máte vyhrať, keď presne vidia a vedia, akú stratégiu ste si prichystali?
#
# **Akú mám zvoliť stratégiu do zápasov v hre kameň-papier-nožnice, ak súperi dopredu poznajú moju stratégiu?**
#
#
# + [markdown] id="t-03nLJarXme" colab_type="text"
# ## Nápoveda
# + [markdown] id="z6G5wAtGraT7" colab_type="text"
# Ak budeme vedieť, čo zahráme, súper nás porazí. Vedeli by sme vymyslieť takú stratégiu, že nebudeme vedieť, čo zahráme?
# + [markdown] id="tXUYBWW0qzpr" colab_type="text"
# ## Riešenie
# + [markdown] id="roRdVZiJ6aPw" colab_type="text"
# Na tento typ otázky sa simulácia nehodí, pretože nie je jasné, čo by sme mali naprogramovať, aby nám to pomohlo.
#
# ### Odhad
#
# Ako nám napovedala nápoveda, potrebujeme zaručiť, aby sme nevedeli, čo zahráme. Na to existuje skvelý prostriedok: náhoda. Ak je naša stratégia náhodná, nevieme dopredu, čo zahráme a teda ani náš súper to nebude vedieť využiť. Ak záhrame rovnomerne náhodne cez všetky tri nástroje, náš súper nemôže mať šancu vyhrať vyššiu ako $\tfrac{1}{3}$. To by mohlo byť riešenie
#
# ### Matematika
#
# Vieme teda, že budeme chcieť hrať kameň, papier a nožnice s nejakými pravdepodobnosťami. Aké pravdepodobnosti by sme si ale mali vybrať? Rovnomerné dajú nášmu súperovi šancu na výhru $\tfrac{1}{3}$, vieme to zlepšiť? Skúsme.
#
# Môžme najprv zistiť, čo najlepšie by mohol spraviť špión a snažiť sa jeho šancu na výhru minimalizovať.
#
# **Pod-úloha**
#
# Predstavme si, že sme v úlohe špióna. Máme informáciu, že náš súper hrá kameň s pravdepodobnosťou $p_K$, papier s pravdepodobnosťou $p_P$ a nožnice s pravdepodobnosťou $p_N$. Čo máme spraviť, aby sme mali čo najväčšiu šancu vyhrať?
# + [markdown] id="QhYDGlTUtmkV" colab_type="text"
# ### Pokračovanie
# + [markdown] id="mKrNYCWTtkJH" colab_type="text"
#
#
# Riešenie je celkom intuitívne. Budeme hrať iba jeden nástroj a to ten, ktorý poráža najpravdepodobnejší nástroj nášho súpera. Ak by bolo napríklad $p_K = \tfrac{1}{2}$, $p_P = \tfrac{1}{3}$ a $p_N = \tfrac{1}{6}$, hrali by sme stále papier a vyhrali by sme s pravdepodobnosťou $\tfrac{1}{2}$. Ako ale dokážeme, že je to naozaj najviac, ako vieme dosiahnuť? Čo ak by sme sami hrali náhodnú stratégiu s pravdepodobnosťami $q_K, q_P$ a $q_N$? Nevieme s nejakým špeciálnym zvolením pravdepodobností dosiahnuť viac? Ak ešte stále nemáte rozmyslený dôkaz, teraz je posledná možnosť!
#
#
# + [markdown] id="jIFSu99B1008" colab_type="text"
# #### Pokračovanie
# + [markdown] id="8JDSn2an14fQ" colab_type="text"
# Môžme vyhrať troma rôznymi spôsobmi, kameň proti nožniciam, nožnice proti papieru alebo papier proti kameňu. Aby sa stala prvá možnosť, musíme si sami náhodne vybrať kameň s pravdepodobnosťou $q_K$ a súper si musí náhodne vybrať nožnice s pravdepodobnosťou $p_N$. Šanca na takúto konkrétnu výhru bude súčin týchto pravdepodobností, teda $q_K \cdot p_N$.
#
# Naša úplna šanca na výhru potom bude
#
# $$q_K \cdot p_N + q_P \cdot p_K + q_N \cdot p_P,$$
#
# kde navyše vieme, že
#
# $$q_K + q_P + q_N = 1 \qquad \text{a} \qquad p_K + q_P + q_N = 1.$$
#
# Ak vám nie je jasné, odkiaľ máme tieto vzorce, skúste si ich odvodiť. Označme si $p = \max(p_K, p_P, p_N)$. Naša šanca na výhru je potom zhora ohraničená ako
#
# $$q_K \cdot p_N + q_P \cdot p_K + q_N \cdot p_P \quad\leq\quad p \cdot (q_K + q_P + q_N) \quad=\quad p.$$
#
# Šancu na výhru $p$ vieme dosiahnuť presne tak, že budeme hrať iba nástroj porážajúci najpravdepodobnejší nástroj súpera, čo bolo treba dokázať. Ohraničili sme, že to môže byť najviac $p$ a ukázali sme, že toto $p$ vieme dosiahnuť.
#
# Ak má teda špión optimálnu stratégiu dosahujúcu maximum z našich pravdepodobností, aby mal čo najmenšiu šancu vyhrať, chceme minimalizovať maximum našich pravdepodobností. To dosiahneme tak, že všetky pravdepodobnosti spravíme rovné $\tfrac{1}{3}$.
#
# Riešenie je teda hrať kameň-papier-nožnice rovnomerne náhodne.
#
# Aby sme si ujasnili terminológiu do ďalších cvičení, odteraz budeme pojmom **stratégia** označovať priradenie pravdepodobností hraniu jednolivých možných **akcií**. V tomto prípade sme došli k optimálnym pravdepodobnostiam $\tfrac{1}{3}$ na hranie každej z akcií *kameň*, *papier* a *nožnice*.
# + [markdown] id="jCQplQon-k2j" colab_type="text"
# # Úloha č.2
#
# Ideme sa hrať hru z minulej hodiny $\rightarrow$ každý si vyberieme postupnosť hodov mincou, budeme hádzať spoločnou mincou a komu padne jeho postupnosť skôr, vyhral.
#
#
# V tjeto časti hráme turnaj v postupnostiach dĺžky 2. Náš súper má svojich špiónov. **Akú máme vybrať stratégiu, aby sme mali čo najväčšiu šancu na výhru?**
# + [markdown] id="OkcFg2xZqzM3" colab_type="text"
# ## Nápoveda 1
#
# + [markdown] id="kqQD-GWTq-kr" colab_type="text"
# Šance na výhry (z minulej hodiny) si zosumarizujeme do tabuľky. Ukazuje šancu na výhru riadka proti stĺpcu.
# + id="JZmiKhGyr9_n" colab_type="code" outputId="cd986b3c-f95d-4f27-b965-78e6df6b6522" colab={"base_uri": "https://localhost:8080/", "height": 173}
table1
# + [markdown] id="oSCfyDEZsFIs" colab_type="text"
# ## Nápoveda 2
# + [markdown] id="S30c3kG165GQ" colab_type="text"
# Vieme povedať o nejakej akcii, že ju určite nebudeme hrať?
# + [markdown] id="fGyMnAFT7CxN" colab_type="text"
# ## Riešenie
# + [markdown] id="4JrYUGdJuoBA" colab_type="text"
# ### Odhad
# + [markdown] id="HJK76HsP7YlT" colab_type="text"
# Všetky akcie, ktoré máme k dispozícií v tejto hre sú $HH$, $HZ$, $ZH$, $ZZ$. Šance výhry jednotlivých akcií nad ostatnými sú spísané v nasledovnej tabuľke.
# + id="QqlSuG788CuM" colab_type="code" outputId="f5d058f5-a5cf-4c02-a672-7a43af824d04" colab={"base_uri": "https://localhost:8080/", "height": 173}
table1
# + [markdown] id="Q9xBb-oGusIj" colab_type="text"
# Vidíme, že hrať $HH$ ani $ZZ$ sa nám neoplatí, lebo ich výsledky sú v každom prípade horšie ako hrať $HZ$ alebo $ZH$. Ostanú nám teda iba dve možnosti, ktoré majú proti sebe vyrovnané šance. Takže na prvý pohľad vyzerá, že riešenie je hrať $ZH$ alebo $HZ$.
# + [markdown] id="miTGmBLuAcjI" colab_type="text"
# ### Matematika
# + [markdown] id="PxzlAE2K-olv" colab_type="text"
# Z odhadu vyzerá jasne, čo by sme chceli dokázať a v tejto časti to skúsime spraviť poctivou a vydretou matematikou. Aj keď možno nie je potrebná spraviť takto detailne, aspoň si to nacvičíme na ťažšie úlohy.
#
# Označme si pravdepodobnosti hrania jednotlivých akcií v rámci našej stratégie ako $p_{HH}$, $p_{HZ}$, $p_{ZH}$ a $p_{ZZ}$ a pravdepodobnosti hrania akcií v súperovej stratégii ako $q_{HH}$, $q_{HZ}$, $q_{ZH}$, $q_{ZZ}$. Ak by sme poznali všetkých týchto 8 čísel, vedeli by sme si vypočítať našu šancu na výhru ako
#
# $$
# \tfrac{1}{2} p_{HH} \cdot q_{HH} + \tfrac{1}{2} p_{HH} \cdot q_{HZ} + \tfrac{1}{4} p_{HH} \cdot q_{ZH} + \tfrac{1}{2} p_{HH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{HZ} \cdot q_{HH} + \tfrac{1}{2} p_{HZ} \cdot q_{HZ} + \tfrac{1}{2} p_{HZ} \cdot q_{ZH} + \tfrac{3}{4} p_{HZ} \cdot q_{ZZ} +\\
# \tfrac{3}{4} p_{ZH} \cdot q_{HH} + \tfrac{1}{2} p_{ZH} \cdot q_{HZ} + \tfrac{1}{2} p_{ZH} \cdot q_{ZH} + \tfrac{1}{2} p_{ZH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{ZZ} \cdot q_{HH} + \tfrac{1}{4} p_{ZZ} \cdot q_{HZ} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZH} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZZ}.\\
# $$
#
# To vyzerá ako dosť veľký a strašideľný výraz. Na tento výraz sa chceme pozrieť z podobného uhla ako v minulej úlohe. To znamená, že pre našu fixnú stratégiu chceme zistiť, ako má naš súper reagovať svojou stratégiou, aby minimalizoval našu šancu na výhru. Naopak, my chceme zvoliť takú stratégiu, aby sme aj v tomto najhoršom prípade dosiahli čo najlepší výsledok.
# + [markdown] id="oYms6X1qDMeB" colab_type="text"
# **Matematicky povedané**, chceme vypočítať *maximum* cez našu voľbu stratégie *minima* cez súperov výber stratégie. **Matematicky zapísané**, chceme vypočítať
#
# $$
# \max_{p_{HH}, p_{HZ}, p_{ZH}, p_{ZZ}} \quad \min_{q_{HH}, q_{HZ}, q_{ZH}, q_{ZZ}} \left[ \\
# \tfrac{1}{2} p_{HH} \cdot q_{HH} + \tfrac{1}{2} p_{HH} \cdot q_{HZ} + \tfrac{1}{4} p_{HH} \cdot q_{ZH} + \tfrac{1}{2} p_{HH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{HZ} \cdot q_{HH} + \tfrac{1}{2} p_{HZ} \cdot q_{HZ} + \tfrac{1}{2} p_{HZ} \cdot q_{ZH} + \tfrac{3}{4} p_{HZ} \cdot q_{ZZ} +\\
# \tfrac{3}{4} p_{ZH} \cdot q_{HH} + \tfrac{1}{2} p_{ZH} \cdot q_{HZ} + \tfrac{1}{2} p_{ZH} \cdot q_{ZH} + \tfrac{1}{2} p_{ZH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{ZZ} \cdot q_{HH} + \tfrac{1}{4} p_{ZZ} \cdot q_{HZ} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZH} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZZ} \right].\\
# $$
#
# Venujte prosím chvíľu, aby ste si premysleli, že vypočítanie tohto výrazu je v skutočnosti cieľom úlohy č.2. Problém takéhoto typu sa vyskytuje veľmi často v teórii hier a nazýva sa aj min-max problémom, respektíve *sedlovou* (angl. *saddle point*) optimalizáciou (skúste prísť na to, prečo).
# + [markdown] id="VAUaNhFTEcaX" colab_type="text"
# Teraz skúsime v tomto výraze uvidieť postup z časti s odhadom. Chceme uvidieť, že pre dosiahnutie maxima musíme zvoliť $p_{HH} = 0$. Vieme to ukázať sporom. Uvažujme našu stratégiu, ktorá maximalizuje výraz vyššie a označme si $p_{HH} = a > 0$ a $p_{HZ} = b$. Potom stratégia v ktorej $p_{HH} = 0 $ a $p_{HZ} = a + b$ dáva priamym porovnaním väčšiu hodnotu:
#
# $$
# \tfrac{1}{2} \mathbf{a} \cdot q_{HH} + \tfrac{1}{2} \mathbf{a} \cdot q_{HZ} + \tfrac{1}{4} \mathbf{a} \cdot q_{ZH} + \tfrac{1}{2} \mathbf{a} \cdot q_{ZZ} +\\
# \tfrac{1}{2} \mathbf{b} \cdot q_{HH} + \tfrac{1}{2} \mathbf{b} \cdot q_{HZ} + \tfrac{1}{2} \mathbf{b} \cdot q_{ZH} + \tfrac{3}{4} \mathbf{b} \cdot q_{ZZ} +\\
# \tfrac{3}{4} p_{ZH} \cdot q_{HH} + \tfrac{1}{2} p_{ZH} \cdot q_{HZ} + \tfrac{1}{2} p_{ZH} \cdot q_{ZH} + \tfrac{1}{2} p_{ZH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{ZZ} \cdot q_{HH} + \tfrac{1}{4} p_{ZZ} \cdot q_{HZ} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZH} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZZ}.\\
# \leq \\
# \tfrac{1}{2} \mathbf{0} \cdot q_{HH} + \tfrac{1}{2} \mathbf{0} \cdot q_{HZ} + \tfrac{1}{4} \mathbf{0} \cdot q_{ZH} + \tfrac{1}{2} \mathbf{0} \cdot q_{ZZ} +\\
# \tfrac{1}{2}\mathbf{(a + b)} \cdot q_{HH} + \tfrac{1}{2}\mathbf{(a + b)} \cdot q_{HZ} + \tfrac{1}{2}\mathbf{(a + b)} \cdot q_{ZH} + \tfrac{3}{4}\mathbf{(a + b)} \cdot q_{ZZ} +\\
# \tfrac{3}{4} p_{ZH} \cdot q_{HH} + \tfrac{1}{2} p_{ZH} \cdot q_{HZ} + \tfrac{1}{2} p_{ZH} \cdot q_{ZH} + \tfrac{1}{2} p_{ZH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{ZZ} \cdot q_{HH} + \tfrac{1}{4} p_{ZZ} \cdot q_{HZ} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZH} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZZ}.\\
# $$
# Rovnosť môže nastať, ak by boli $q_{HH}$ a $q_{ZZ}$ nulové. V každom prípade ale z vyššieho vyplýva, že existuje maximálne riešenie, v ktorom je $p_{HH} = 0$ a budeme preto uvažovať iba takéto stratégie. Ak máme takýto vzťah medzi dvoma akciami $HH$ a $HZ$, hovorí sa, že akcia $HZ$ **majorizuje** akciu $HH$.
# + [markdown] id="CPQUEafgIYdo" colab_type="text"
# Obdobným spôsobom vieme ukázať aj to, že $p_{ZZ} = 0$ v maxime. Ostanú nám teda iba dve nenulové premenné, konkrétne $p_{HZ}$ a $p_{ZH}$. Vieme o nich, že ich súčet je 1, lebo sú to pravdepodobnosti. Využitím tohto vzťahu si označme $p_{HZ} = p$ a $p_{ZH} = 1 -p$. Naša pôvodná úloha sa teda pretransformovala na
#
# $$
# \max_{p} \quad \min_{q_{HH}, q_{HZ}, q_{ZH}, q_{ZZ}} \left[ \\
# \tfrac{1}{2} p \cdot q_{HH} + \tfrac{1}{2} p \cdot q_{HZ} + \tfrac{1}{2} p \cdot q_{ZH} + \tfrac{3}{4} p \cdot q_{ZZ} +\\
# \tfrac{3}{4} (1 - p) \cdot q_{HH} + \tfrac{1}{2} (1 - p) \cdot q_{HZ} + \tfrac{1}{2} (1 - p) \cdot q_{ZH} + \tfrac{1}{2} (1 - p) \cdot q_{ZZ} \right] \\
# = \\
# \max_{p} \quad \min_{q_{HH}, q_{HZ}, q_{ZH}, q_{ZZ}} \left[
# (\tfrac{3}{4} - \tfrac{1}{4}p) \cdot q_{HH} + \tfrac{1}{2} q_{HZ} + \tfrac{1}{2} q_{ZH} + (\tfrac{1}{2} + \tfrac{1}{4}p) \cdot q_{ZZ} \right] \\
# = \\
# \tfrac{1}{2} + \max_{p} \quad \min_{q_{HH}, q_{HZ}, q_{ZH}, q_{ZZ}} \left[
# \tfrac{1}{4}(1 - p) \cdot q_{HH} + \tfrac{1}{4}p \cdot q_{ZZ} \right],
# $$
# pričom v poslednom kroku sme využili, že súčet $q$ je 1. Oba členy, ktoré ostali v maximalizačno-minimalizačnej časti sú kladné. Minimum sa teda dosiahne pre hodnoty $q_{HH} = 0$ a $q_{ZZ} = 0$ nezávisle od voľby nášho $p$, čím dostaneme konečný výsledok $\tfrac{1}{2}$.
#
# V rámci dôkazu sme vyžadovali od našej stratégie iba aby platilo $p_{HH} = 0$ a $p_{ZZ} = 0$ a preto pravdepodobnosti zvolené pre zvyšné dve akcie môžu byť v skutočnosti ľubovoľné! Skúste sa o tom sami presvedčiť.
# + [markdown] id="nov66LuaMZnQ" colab_type="text"
# #### Into the matrix
#
#
# + [markdown] id="_6t3jApJMd4Z" colab_type="text"
# Ako ste si iste všimli, riešenie tejto úlohy bolo plné obrovských výrazov. Hneď na úvod sme na vás vybalili tento
#
# $$
# \tfrac{1}{2} p_{HH} \cdot q_{HH} + \tfrac{1}{2} p_{HH} \cdot q_{HZ} + \tfrac{1}{4} p_{HH} \cdot q_{ZH} + \tfrac{1}{2} p_{HH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{HZ} \cdot q_{HH} + \tfrac{1}{2} p_{HZ} \cdot q_{HZ} + \tfrac{1}{2} p_{HZ} \cdot q_{ZH} + \tfrac{3}{4} p_{HZ} \cdot q_{ZZ} +\\
# \tfrac{3}{4} p_{ZH} \cdot q_{HH} + \tfrac{1}{2} p_{ZH} \cdot q_{HZ} + \tfrac{1}{2} p_{ZH} \cdot q_{ZH} + \tfrac{1}{2} p_{ZH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{ZZ} \cdot q_{HH} + \tfrac{1}{4} p_{ZZ} \cdot q_{HZ} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZH} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZZ}.\\
# $$
#
# Hovorí niečo dôležité, ale je neefektívne zapísaný. Jednak zaberá veľa miesta a jednak každá z hodnôt $p$ a $q$ sa vyskytuje v tomto výraze až 4-krát, pričom $p$ sú rovnaké po riadkoch a $q$ po stĺpcoch. Je iba prirodzené, že matematici si tento vzor všimli a vymysleli jednoduchší spôsob zápisu tohto výrazu. Pokročilí matematici ho zapíšu takto:
#
# $$
# \begin{bmatrix}
# p_{HH} & p_{HZ} & p_{ZH} & p_{ZZ}
# \end{bmatrix}
# \begin{bmatrix}
# \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{1}{4} & \tfrac{1}{2} \\
# \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{3}{4} \\
# \tfrac{3}{4} & \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{1}{2} \\
# \tfrac{1}{2} & \tfrac{1}{4} & \tfrac{1}{2} & \tfrac{1}{2}
# \end{bmatrix} \begin{bmatrix}
# q_{HH} \\
# q_{HZ} \\
# q_{ZH} \\
# q_{ZZ}
# \end{bmatrix}
# $$
#
# Niektorí z vás sa už možno s podobným zápisom stretli, jedná sa o *matice* a *vektory* a násobenie medzi nimi. Skúste sa zamyslieť nad tým, ako asi takéto násobenie funguje, aby boli tieto dva výrazy rovnaké.
# + [markdown] id="_fSO4TN_QrZX" colab_type="text"
# Namiesto nášho max-min problému zapísaného ako
#
# $$
# \max_{p_{HH}, p_{HZ}, p_{ZH}, p_{ZZ}} \quad \min_{q_{HH}, q_{HZ}, q_{ZH}, q_{ZZ}} \left[ \\
# \tfrac{1}{2} p_{HH} \cdot q_{HH} + \tfrac{1}{2} p_{HH} \cdot q_{HZ} + \tfrac{1}{4} p_{HH} \cdot q_{ZH} + \tfrac{1}{2} p_{HH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{HZ} \cdot q_{HH} + \tfrac{1}{2} p_{HZ} \cdot q_{HZ} + \tfrac{1}{2} p_{HZ} \cdot q_{ZH} + \tfrac{3}{4} p_{HZ} \cdot q_{ZZ} +\\
# \tfrac{3}{4} p_{ZH} \cdot q_{HH} + \tfrac{1}{2} p_{ZH} \cdot q_{HZ} + \tfrac{1}{2} p_{ZH} \cdot q_{ZH} + \tfrac{1}{2} p_{ZH} \cdot q_{ZZ} +\\
# \tfrac{1}{2} p_{ZZ} \cdot q_{HH} + \tfrac{1}{4} p_{ZZ} \cdot q_{HZ} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZH} + \tfrac{1}{2} p_{ZZ} \cdot q_{ZZ}. \right]\\
# $$
#
# pokročilí matematici povedia, že: Definujme vektory
#
# $$\mathbf{p} = \begin{bmatrix}
# p_{HH} \\ p_{HZ} \\ p_{ZH} \\ p_{ZZ}
# \end{bmatrix}, \quad \mathbf{q} = \begin{bmatrix}
# q_{HH} \\ q_{HZ} \\ q_{ZH} \\ q_{ZZ}
# \end{bmatrix}$$
#
# a maticu
#
# $$
# \mathbf{A} = \begin{bmatrix}
# \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{1}{4} & \tfrac{1}{2} \\
# \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{3}{4} \\
# \tfrac{3}{4} & \tfrac{1}{2} & \tfrac{1}{2} & \tfrac{1}{2} \\
# \tfrac{1}{2} & \tfrac{1}{4} & \tfrac{1}{2} & \tfrac{1}{2}
# \end{bmatrix},
# $$
#
# a potom náš problém je jednoducho
#
# $$
# \max_{\mathbf{p}} ~\min_{\mathbf{q}} \quad \mathbf{p}^{\top} \mathbf{A} \mathbf{q}.
# $$
#
# Vyrázne jednoduchšie, čo poviete? Jednotlivé súčasti sú jasne oddelené a samotná úloha je už len priamočiarou kombináciou týchto výrazov.
#
# Matice a vektory sa teraz nebudeme učiť používať, ale do budúcnosti si z tohto pamätajte, že slúžia na zjednodušenie výpočtov a zápisov. Ak vás to zaujíma viac, pravidlá na ich používanie nájdete hocikde na internete.
# + [markdown] id="DTjSZboM_hOC" colab_type="text"
# # Úloha č.3
#
# Zopakujeme si predchádzajúcu úlohu s postupnosťami dĺžky 3. **Akú by sme mali zvoliť stratégiu?**
#
# Pozn.: Údaje v tabuľke sú zaokruhlené na 3 desatinné miesta, pôvodné čísla nájdete v sekcii pomocného kódu.
#
# + id="r50o8HC_b24M" colab_type="code" outputId="ea45403b-4426-47e6-f00c-4f200b08da8f" colab={"base_uri": "https://localhost:8080/", "height": 297}
table2
# + [markdown] id="QGhMQVn7cVhi" colab_type="text"
# ## Nápoveda
# + [markdown] id="rOZUuKNHclHq" colab_type="text"
# Nepotrebujete naviac nič nové oproti predchádzajúcej úlohe, akurát postup je komplikovanejší.
# + [markdown] id="JWlM-MxqcmxE" colab_type="text"
# ## Riešenie
# + [markdown] id="MLwkTrTFcrm1" colab_type="text"
# ### Odhad
# + [markdown] id="nXwxDPd0xb_c" colab_type="text"
# Po prvom prezretí tabuľky vidíme, že postupnosti $HHZ$ majorizuje $HHH$, $ZZH$ majorizuje $ZZZ$ a takisto aj $HZZ$ majorizuje $HZH$ a $ZHH$ majorizuje $ZHZ$. Ak vyradíme tieto štyri akcie, ostáva nám tabuľka
# + id="dlnervRifA0c" colab_type="code" outputId="3ace6b64-0a8e-4907-bad0-fb81e5eeb160" colab={"base_uri": "https://localhost:8080/", "height": 173}
table3
# + [markdown] id="R1qC14Cif5PY" colab_type="text"
# Po krátkom štúdiu tabuľky vidíme, že ku každej akcii vieme priradiť inú akciu, ktorá ju poráža. Máme teda niečo podobné ako kameň - papier - nožnice, akurát pravdepodobnosti sú odlišné. Môžme si ale všimnúť, že akcie $HZZ$ a $ZHH$ majú väčšie pravdepodobnosti výhry ako zvyšné dve a preto by sme očakávali, že ich budeme hrať s väčšiou pravdepodobnosťou.
# + [markdown] id="TgyGw8mmgh5b" colab_type="text"
# ### Matematika
# + [markdown] id="fYZfs4TLgjz9" colab_type="text"
# V sekcii s odhadom sme použili správny argument na majorizáciu a budeme pokračovať už len so zmenšenou tabuľkou.
# + id="QbuBwNzjhYet" colab_type="code" outputId="41da0a08-06fa-43bd-9b16-d07defe1492b" colab={"base_uri": "https://localhost:8080/", "height": 173}
table3
# + [markdown] id="sIcZCaHvhZdT" colab_type="text"
# #### Transformácia problému
#
# Pravdepodobnosti v tabuľke môžme interpretovať aj ako bodové zisky, ktoré sa snažíme maximalizovať. Z pohľadu stratégie je úplne jedno, či sú bodové zisky takéto, alebo 10 krát väčšie. Aby sme nemuseli pracovať so zlomkami, odpočítame si od celej tabuľky $\tfrac{1}{2}$ a vynásobíme ju číslom 12. Takto dostaneme celé čísla a vyrovnaná pravdepodobnosť bude zodpovedať bodovému zisku 0.
# + id="WPgYnaOiidHT" colab_type="code" outputId="bb70c50a-cc9a-426d-cd4b-7f549d4a51e6" colab={"base_uri": "https://localhost:8080/", "height": 173}
table4
# + [markdown] id="JRjdFldZifjo" colab_type="text"
# Premyslite si, že nájdenie optimálnej stratégie pre túto tabuľku je ekvivalentné s nájdením optimálnej stratégie pre pôvodnú tabuľku. Keďže je táto hra symetrická pre oboch hráčov a náš súper si môže vyberať stratégiu ako druhý, náš optimálny bodový zisk môže byť nanajvýš 0 (premyslite si). Ak dosiahneme bodový zisk $0$, bude to úspech.
# + [markdown] id="8zQEi5jZwqyt" colab_type="text"
# #### Kombinovaná majorizácia
# + [markdown] id="BXHBtj-m1Y6x" colab_type="text"
# Ako si môžme všimnúť, žiadna ďalšia akcia už nie je majorizovaná a ostáva otázkou, čo ďalej. Pre začiatok si matematicky napíšme náš sedlový problém:
#
# $$
# \max_{p_{HHZ}, p_{HZZ}, p_{ZHH}, p_{ZZH}} \quad \min_{q_{HHZ}, q_{HZZ}, q_{ZHH}, q_{ZZH}} \left[ \\
# 2 p_{HHZ} \cdot q_{HZZ} - 3 p_{HHZ} \cdot q_{ZHH} \\
# -2 p_{HZZ} \cdot q_{HHZ} + 3 p_{HZZ} \cdot q_{ZZH} \\
# 3 p_{ZHH} \cdot q_{HHZ} - 2 p_{ZHH} \cdot q_{ZZH} \\
# -3 p_{ZZH} \cdot q_{HZZ} + 2 p_{ZZH} \cdot q_{ZHH} \right] \\
# = \\
# \max_{p_{HHZ}, p_{HZZ}, p_{ZHH}, p_{ZZH}} \quad \min_{q_{HHZ}, q_{HZZ}, q_{ZHH}, q_{ZZH}} \left[ \\
# q_{HHZ} \cdot (3 p_{ZHH} - 2 p_{HZZ}) + \\
# q_{HZZ} \cdot (2 p_{HHZ} - 3 p_{ZZH}) + \\
# q_{ZHH} \cdot (2 p_{ZZH} - 3 p_{HHZ}) + \\
# q_{ZZH} \cdot (3 p_{HZZ} - 2 p_{ZHH}) \right]
# $$
#
# Po dlhších pohľadoch na tieto rovnice a na tabuľku si môžme všimnúť, že kombinácia akcií $HZZ$ a $ZHH$ majorizuje kombináciu akcií $HHZ$ a $ZZH$.
# + [markdown] id="hviHtfV-mgil" colab_type="text"
# Matematicky povedané, nech $$p_{HHZ} = p_{ZZH} = a, \qquad p_{HZZ} = b, \qquad p_{ZHH} = c$$ je naša stratégia. Potom platí, že stratégia $$p_{HHZ} = p_{ZZH} = 0, \qquad p_{HZZ} = (a + b), \qquad p_{ZHH} = (a + c)$$ dáva lepšie výsledky, pretože
#
# $$
# q_{HHZ} \cdot (3 c - 2 b) + q_{HZZ} \cdot (2 a - 3 a) + q_{ZHH} \cdot (2 a - 3 a) + q_{ZZH} \cdot (3 b - 2 c) \\
# < \\
# q_{HHZ} \cdot (3 (c + a) - 2 (b + a)) + q_{HZZ} \cdot 0 + q_{ZHH} \cdot 0 + q_{ZZH} \cdot (3 (b + a) - 2 (c + a)) \\
# = \\
# q_{HHZ} \cdot (3c - 2b + a)) + q_{ZZH} \cdot (3b - 2 c + a)
# $$
#
# Nerovnosť platí, lebo pri každom z členov $q$ máme zrazu číslo, ktoré je o $a$ väčšie a zároveň sú všetky členy $q$ nezáporné. Keďže $a$ je kladné, pravá (spodná) strana je nutne väčšia.
#
# Touto úvahou sme dokázali, že ak sú $p_{HHZ}$ a $p_{ZZH}$ rovnaké a nenulové, nemáme optimálne riešenie.
# + [markdown] id="tHzXv66CpVY3" colab_type="text"
# V skutočnosti ani nepotrebujeme podmienku, že musia byť rovnaké, pretože argument ako vyššie môžme použiť aj tak. Akonáhle vieme odčítať od oboch $p_{HHZ}$ a $p_{ZZH}$ rovnakú hodnotu a presunúť ich rovnomerne na pravdepodobnosti $p_{HZZ}$ a $p_{ZHH}$, vieme náš výraz zväčšiť (premyslite si, prípadne dokážte).
#
# Z toho vyplýva, že akonáhle sú obe pravdepodobnosti $p_{HHZ}$ a $p_{ZZH}$ nenulové, nemáme optimálne rozloženie pravdepodobností. Odteraz teda môžme uvažovať, že aspoň jedno z pravdepodobností $p_{HHZ}$ a $p_{ZZH}$ musí byť nula.
# + [markdown] id="HL1yJ6lorUlM" colab_type="text"
# #### Maximálne riešenie
#
# Nevieme síce zatiaľ ukázať, že aj druhá z pravdepodobností $p_{HHZ}$ a $p_{ZZH}$ musí byť nula, ale začína to tak vyzerať. Zamyslime sa teda nachvíľu, že ak by sme naozaj používali iba akcie $HZZ$ a $ZHH$, aká by bola optimálna stratégia. Majme teda stratégiu definovanú všeobecne ako
#
# $$
# p_{HHZ} = p_{ZZH} = 0, \qquad p_{HZZ} = p, \qquad p_{ZHH} = (1-p).
# $$
#
# Náš optimalizačný problém sa zmení na
#
# $$
# \max_{p} \quad \min_{q_{HHZ}, q_{HZZ}, q_{ZHH}, q_{ZZH}} \left[ q_{HHZ} \cdot (3 (1-p) - 2 p) + q_{ZZH} \cdot (3 p - 2 (1 - p)) \right] \\
# = \\
# \max_{p} \quad \min_{q_{HHZ}, q_{HZZ}, q_{ZHH}, q_{ZZH}} \left[ q_{HHZ} \cdot (3 - 5p) + q_{ZZH} \cdot (5p - 2) \right]
# $$
# Ak teda zvolíme ako $p$ ľubovoľné číslo medzi $\tfrac{2}{5}$ a $\tfrac{3}{5}$, náš bodový zisk budeme vedieť zdola ohraničiť nulou, čo bolo našim cieľom. Aby sme nemali kladný bodový zisk, bude musieť súper zvoliť stratégiu v ktorej $q_{HHZ} = q_{ZZH} = 0$. Takýmto spôsobom sa nám podarí hrať s vyrovnanými šancami na výhru.
# + [markdown] id="yAINoxT3tqbr" colab_type="text"
# #### Iba dve nenulové pravdepodobnosti
#
# Vráťme sa ešte naspäť k pôvodnému problému, lebo sme ešte nedokázali, že obe pravdepodobnosti $p_{HHZ}$ a $p_{ZZH}$ musia byť nutne nulové. Uvažujme teraz stratégiu danú ako
#
# $$
# p_{HHZ} = 0, \quad p_{ZZH} = a > 0 \qquad p_{HZZ} = b, \qquad p_{ZHH} = c
# $$
#
# Náš optimalizačný problém bude
# $$
# \max_{a, b, c} \quad \min_{q_{HHZ}, q_{HZZ}, q_{ZHH}, q_{ZZH}} \left[ q_{HHZ} \cdot (3 c - 2 b) + q_{HZZ} \cdot ( - 3 a) + q_{ZHH} \cdot (2 a) + q_{ZZH} \cdot (3 b - 2 c) \right]
# $$
# Môžme si všimnúť, že ak súper vždy zvolí akciu $HZZ$, bude náš zisk záporný, čo je horšie ako 0, ktorú sme vedeli dosiahnuť v predchádzajúcej časti. Toto riešenie teda nemôže byť optimálne. Obdobne ukážeme aj nutnú nulovosť pravdepodobnosti $p_{HHZ}$.
# + [markdown] id="pKJh9l1Av0Av" colab_type="text"
# #### Súhrn
#
# Aj keď to tak možno nevyzerá, práve sme skončili dôkaz. Ukázali sme, že:
# - Akcie $HHH$, $HZH$, $ZHZ$ a $ZZZ$ sú majorizované a nemusíme ich uvažovať.
# - Kombinácia akcií $HHZ$ a $ZZH$ je majorizovaná kombináciou $HZZ$ a $ZHH$, z čoho vyplýva, že $p_{HZZ}$ a $p_{ZHH}$ nemôžu byť zároveň nenulové.
# - Zvolením stratégie $$
# p_{HHZ} = p_{ZZH} = 0, \qquad p_{HZZ} = p, \qquad p_{ZHH} = (1-p).
# $$ pre $p \in [\tfrac{2}{5}, \tfrac{3}{5}]$ vieme dosiahnuť 50% šancu na výhru.
# - Zvolením stratégie, kde je práve jedna z pravdepodobností $p_{HHZ}$ a $p_{ZZH}$ nenulová vie náš súper pre nás zaručiť pravdepodobnosť na výhru menšiu ako 50%.
#
# Ak spojíme všetky tieto argumenty, dostaneme, že naša maximálna šanca na výhru je 50% a je dosiahnuteľná iba stratégiou opísanou v treťom bode.
#
# Prezentovali sme tu jedno z možných riešení, sada argumentov mohla byť aj odlišná. V úlohách tohto typu to tak často býva. Nezabudnite si porovnať svoj spôsob riešenia aj s ostatnými.
# + [markdown] id="DK5oco9hzxW0" colab_type="text"
# # Domáca úloha č. 1
#
# Pokračovanie úlohy č.1. Na ďalší zápas v kameň-papier-nožnice sme si tiež zamestnali svojich špiónov a prezradili nám, že náš súper nikdy nebude hrať kameň. **Akú zvolíme stratégiu?**
# + [markdown] id="UjjCJsL7nbN3" colab_type="text"
# # Domáca úloha č.2
#
# Pokračovanie úlohy č.3. Naši špióni nám prezradili, že náš súper si nikdy nevyberie postupnosť hlava-znak-znak. **Akú zvolíme stratégiu?**
# + [markdown] id="HLcm2ZjtJAAS" colab_type="text"
# # Úloha pre praktikov
#
# Nájdite optimálnu stratégiu pre nejakú situáciu, v ktorej ste sa v minulosti ocitli.
# + [markdown] id="KdVmXLffnQ9X" colab_type="text"
# # Úloha pre brutálnych
#
# Zopakujeme si úlohu 2 & 3 s postupnosťami dĺžky 4.
# + [markdown] id="RB0gclNG1o5o" colab_type="text"
# # Úloha pre akademikov
#
# Uvažujme mincovú hru o postupnostiach dĺžky $n$. Budú existovať akcie, ktoré budú vždy majorizované? Ktoré? Koľko ich je?
| notebooks/03_spionske_hry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import dask.dataframe as dd
from distributed import wait
from prefect import Flow, task
import prefect
df = dd.read_csv(
"s3://nyc-tlc/trip data/yellow_tripdata_2019-01.csv",
dtype={
"payment_type": "UInt8",
"VendorID": "UInt8",
"passenger_count": "UInt8",
"RatecodeID": "UInt8",
},
storage_options={"anon": True}
)
# -
df = df[['payment_type', 'fare_amount', 'trip_distance', 'passenger_count', 'tpep_pickup_datetime', 'tip_amount']]
# +
from dask_ml.xgboost import XGBRegressor
est = XGBRegressor()
est.fit(df.drop('tip_amount', axis=1), df['tip_amount'])
# -
| blogs/coiled-prefect-demo/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gfgullo/AVSpeechSynthesizer-Example/blob/master/CNN_%2B_LSTM_con_Keras.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="SDGEK3yNerdL" colab_type="text"
# # CNN + LSTM con Keras
#
# Importiamo i moduli che ci serviranno
# + id="Jz5zRrEteY_S" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Dense
from keras.datasets import imdb
from keras.preprocessing.sequence import pad_sequences
# + [markdown] id="uadT-PS2eyph" colab_type="text"
# ## Caricamento e preprocessing del dataset
# Carichiamo il dataset con Keras limitandolo alle 10.000 parole più comuni, poi tronchiamo/espandiamo le sequenze a 500 elementi con la funzione pad_sequences.
# + id="OYrwWo6iew4e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="61b350b1-5fe6-403a-ae8b-aecd698bd343"
num_words = 10000
maxlen = 500
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=num_words)
X_train = pad_sequences(X_train, maxlen = maxlen)
X_test = pad_sequences(X_test, maxlen = maxlen)
# + [markdown] id="eW4Hjx7lfEWo" colab_type="text"
# ## Creazione del modello
# + id="4S7eDkqhe3-g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="e52175b1-a309-4c62-d6dd-98c0ec95e96e"
from keras.layers import Embedding, LSTM
from keras.layers.convolutional import Conv1D, MaxPooling1D
model = Sequential()
model.add(Embedding(num_words, 50))
model.add(Conv1D(filters=32, kernel_size=3, padding="same", activation="relu"))
model.add(MaxPooling1D(pool_size=2))
model.add(LSTM(32, dropout=0.4))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + id="Ts4mnFOyfOlj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 311} outputId="7538e6e1-ab9a-482a-f3f1-6e40f2437b9e"
model.compile(loss='binary_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
model.fit(X_train, y_train, batch_size=512, validation_split=0.2, epochs=5)
model.evaluate(X_test, y_test)
# + [markdown] id="6QxhigwLf7dW" colab_type="text"
# Come vedi, questo approccio ibrido in cui abbiamo combinato uno strato convoluzionale con uno strato ricorrente ci ha portato al miglior risultato che abbiamo ottenuto fin'ora.
| CNN_+_LSTM_con_Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exothermic Batch Reactor
# ## Details
# <NAME> (18CHE160)
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from matplotlib import style
from scipy.integrate import odeint
from IPython.display import display, Math
# %config InlineBackend.figure_format = 'svg'
style.use("classic")
# -
# ## Data
#
# $\rho = 1000 \ \ (\frac{kg}{m^3})$
#
# ${C}_{p} = 0.96 \ \ (\frac{kcal}{kg-K})$
#
# k (at 298K) = 0.002 ${min}^{-1}$
#
# A = 15${m}^{2}$
#
# U = 600 $\frac{kcal}{hr - {m}^{2} - K}$
#
# Initial Conditions: at $t=0: {C}_{Ai} = 2 \ \ (\frac{kmol}{{m}^{3}})$
#
# ${T}_{i} = 298 K$
#
# ${V}_{reactor} = 10{m}^{3}$
#
#
# ## Reaction Details
#
# The reaction is exothermic
#
# A batch reactor is used to process this reactant A
#
# Reaction is of the form $A \ \ \rightarrow \ \ Products$
#
rho = 1000 # kg/m3
Cp = (0.96) # J/kmol/K
k = 0.002 #min^-1
Tc = 298 #K
V = 10 #m3
U = 600/60 #kcal/(min-m2-K)
A = 15 #m2
E_by_R = 10000 #K
ua_by_v = U*A/V # J/(sec-m3-K)
zeta = U*A/V/(rho*Cp)
ko = k/np.exp(-E_by_R/298)
ca_i = 2 #kmol/m3
Ti = 298 #K
#
# ## Equations
#
#
# The mass and energy balance equations yield
#
# $$\frac{{dX}_{A}}{dt} = k_o * exp(\frac{-E}{R \ \ * T_i + \Theta * {(\Delta T )}_{max}})* (1 - {X}_{A})$$
#
# and
#
# $$\frac{d \Theta}{dt} = k_o * exp(\frac{-E}{R \ \ * T_i + \Theta * {(\Delta T )}_{max}})* (1 - {X}_{A}) - (\frac{U*A}{V* \rho * C_p}) * (\Theta + \alpha)$$
#
# where $\Theta = (\frac{T - T_i}{{(\Delta T )}_{max}})$
#
# and $\alpha = (\frac{T_i - T_c}{{(\Delta T )}_{max}})$
#
# ## Defining the ODEs
def reactor_solver(del_h_r_given):
del_h_r = del_h_r_given*1000 # kcal/kmol
del_t_max = ca_i * (-del_h_r)/(rho*Cp)
alpha = (Ti-Tc)/del_t_max
#print("Delta T max: %.2f"% (del_t_max))
def balances(x, t):
# We define the ODEs here
[Xa,theta] = x
dXadt = ko*np.exp( -E_by_R/ (Ti + theta*del_t_max) )*(1-Xa)
dthetadt = ko*np.exp( -E_by_R/ (Ti + theta*del_t_max) )*(1-Xa) - zeta *(theta+alpha)
return [dXadt,dthetadt]
# Defining time arrays
t_0 = np.linspace(0,180, 5000)
# soln =[Xa,theta]
x0 = [0,0]
soln_0 = odeint(balances, x0, t_0)
return [t_0, soln_0,del_t_max]
def plot_sol(t, sol,del_h_r_given,del_t_max):
fig = plt.figure(facecolor="white")
plt.suptitle(r"$(\Delta H_r) = %.1f (\frac{kcal}{gmol}) \ \ , {(\Delta T)}_{max} = %.2f K$" % (del_h_r_given,del_t_max), fontsize=18)
plt.grid()
plt.plot(t, sol[:, 0], label=r'$X_A$', color='r', linewidth=1.5)
plt.plot(t, sol[:, 1], label=r'$\Theta $', color='b', linewidth=1.5)
plt.xlabel("Time (min)")
plt.ylabel(r"Conversion $(X_A)$ , Temperature $\Theta$")
plt.ylim([0,1])
plt.legend(loc="best")
# ## Solving ODEs and plotting for various cases
del_h_r_given = -15
[t_0, soln_0,del_t_max] =reactor_solver(del_h_r_given)
plot_sol(t_0, soln_0,del_h_r_given,del_t_max )
del_h_r_given = -20
[t_0, soln_0,del_t_max] =reactor_solver(del_h_r_given)
plot_sol(t_0, soln_0,del_h_r_given,del_t_max )
del_h_r_given = -30
[t_0, soln_0,del_t_max] =reactor_solver(del_h_r_given)
plot_sol(t_0, soln_0,del_h_r_given,del_t_max )
del_h_r_given = -35
[t_0, soln_0,del_t_max] =reactor_solver(del_h_r_given)
plot_sol(t_0, soln_0,del_h_r_given,del_t_max )
del_h_r_given = -40
[t_0, soln_0,del_t_max] =reactor_solver(del_h_r_given)
plot_sol(t_0, soln_0,del_h_r_given,del_t_max )
del_h_r_given = -45
[t_0, soln_0,del_t_max] =reactor_solver(del_h_r_given)
plot_sol(t_0, soln_0,del_h_r_given,del_t_max )
| PSL2 Assignment 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Packages
from pyomo.environ import *
import numpy as np
import random
import pandas as pd
from numpy import flatnonzero as find
from pypower.api import case9, ext2int, bustypes, makeBdc, rundcpf, ppoption, case39, rundcopf
from pypower.idx_bus import BUS_TYPE, REF, VA, PD, LAM_P, LAM_Q, MU_VMAX, MU_VMIN
from pypower.idx_gen import PG, MU_PMAX, MU_PMIN, MU_QMAX, MU_QMIN, PMAX, PMIN, GEN_BUS
from pypower.idx_brch import PF, PT, QF, QT, RATE_A, MU_SF, MU_ST
from pypower.idx_cost import COST
import matplotlib.pyplot as plt
# Run PyPower case to get initial condition
ppc = case39()
ppopt = ppoption(VERBOSE=0)
pp_dcpf = rundcopf(ppc, ppopt)
pp_bus = pp_dcpf["bus"]
pp_branch = pp_dcpf["branch"]
pp_gen = pp_dcpf["gen"]
# +
## convert to internal indexing
ppc = ext2int(ppc)
baseMVA, bus, gen, branch, gencost = \
ppc["baseMVA"], ppc["bus"], ppc["gen"], ppc["branch"], ppc["gencost"]
## get bus index lists of each type of bus
ref, pv, pq = bustypes(bus, gen)
pvpq = np.matrix(np.r_[pv, pq])
## generator info
gbus = gen[:, GEN_BUS].astype(int) ## what buses are they at?
refgen = find(gbus == ref)
## build B matrices and phase shift injections
B, Bf, _, _ = makeBdc(baseMVA, bus, branch)
B = B.todense()
Bf = Bf.todense()
# Problem dimensions
NG = gen.shape[0] # Number of generators
NB = bus.shape[0] # Number of buses
NBr = branch.shape[0] # Number of lines
NL = np.count_nonzero(pp_bus[:, PD]) # Number of loads
NW = 1 # Number of wind farms
# Map generator to bus
CG = np.zeros((NB,NG))
CG[gbus,range(NG)] = 1
# Map load to bus
CL = np.zeros((NB,NL))
load_bus = np.nonzero(pp_bus[:, PD])[0]
for i in range(len(load_bus)):
CL[load_bus[i], i] = 1
# Map wind farm to bus
CW = np.zeros((NB,1))
CW[5,0] = 1
# Generator capacity limit p.u.
Pmax = gen[:, PMAX]/baseMVA
Pmin = gen[:, PMIN]/baseMVA
# Line flow limit p.u.
Lmax = branch[:, RATE_A]/baseMVA
Lmin = - branch[:, RATE_A]/baseMVA
# Quadratic cost coefficients
# Convert to p.u.
CG2 = gencost[:, COST]*baseMVA**2
CG1 = gencost[:, COST+1]*baseMVA
CG0 = gencost[:, COST+2]
# Cost coefficient for wind power
CW1 = 100
CW0 = 0
# Cost coefficients for generation reserve
CRG_up = 10*np.ones((NG,1))*baseMVA
CRG_down = 10*np.ones((NG,1))*baseMVA
# Cost coefficients for load reserve
CRL_up = 9.8*np.ones((NL,1))*baseMVA
CRL_down = 9.8*np.ones((NL,1))*baseMVA
# Uncontrollable load
load = pp_bus[:,PD]
PL_total = load[load > 0]
PL = PL_total/baseMVA
# Generator and demand set points
# Power in p.u., Va in rad
Pg0 = pp_gen[:, PG]/baseMVA
Pd0 = pp_bus[:, PD]/baseMVA
Pinj0 = (np.matmul(CG, Pg0) - Pd0)
Va0 = pp_bus[:, VA]/180*np.pi
# -
# Read temperature data
weather_data = pd.read_csv("Weather_Data/NYC_Hourly_Normal_Temp.csv")
# Read wind power data
PW_data = pd.read_csv("spring_wind.csv",header=None).to_numpy()
PW_data = PW_data/500*300/baseMVA
# PW_data = PW_data[:, 0:4000]
Tamb = (weather_data.iloc[:,8] - 32)/1.8
Tamb_min = min(Tamb)
Tamb_10PCTL = (weather_data.iloc[:,6] - 32)/1.8
Tamb_90PCTL = (weather_data.iloc[:,7] - 32)/1.8
std_Tamb_10PCTL = (Tamb - Tamb_10PCTL)/1.282
std_Tamb_90PCTL = (Tamb_90PCTL - Tamb)/1.282
std_Tamb = (std_Tamb_10PCTL + std_Tamb_90PCTL)/10
def VS(dt,Tamb,num_TCL,Tset,delta,R,C,P_el,COP,baseMVA):
Tdown = np.zeros(num_TCL)
Tup = np.zeros(num_TCL)
T_ON = np.zeros(num_TCL)
T_OFF = np.zeros(num_TCL)
D = np.zeros(num_TCL)
B = 0
Pdown = 0
Pup = 0
Sdown = 0
Sup = 0
for i in range(0,num_TCL):
Tdown[i] = Tset[i] - delta/2
Tup[i] = Tset[i] + delta/2
T_ON[i] = C[i]*R[i]*np.log((Tdown[i] - Tamb - R[i]*P_el[i]*COP[i])/(Tup[i] - Tamb - R[i]*P_el[i]*COP[i]))
if ((Tdown[i] - Tamb - R[i]*P_el[i]*COP[i])/(Tup[i] - Tamb - R[i]*P_el[i]*COP[i]) < 0):
print(Tdown[i] - Tamb - R[i]*P_el[i]*COP[i])
print(Tup[i] - Tamb - R[i]*P_el[i]*COP[i])
T_OFF[i] = C[i]*R[i]*np.log((Tup[i] - Tamb)/(Tdown[i] - Tamb))
D[i] = T_ON[i]/(T_ON[i] + T_OFF[i])
if ( Tamb < Tdown[i] and Tamb + R[i]*P_el[i]*COP[i] > Tup[i] ):
B = B + P_el[i]*D[i]
Pup = Pup + P_el[i]
Sup = Sup + dt*P_el[i]*(1 - D[i])
return [B/1000/baseMVA,Pdown/1000/baseMVA,Pup/1000/baseMVA,Sdown,Sup]
# Optimization: Problem formulation
def optimizer(NG,NL,Tamb_sample1,Tamb_sample2,PW_sample,num_TCL,Tset,delta,R,C,P_el,COP,S0,PL,\
Pmax,Pmin,Lmax,Lmin,CG,CW,CL,CG0,CG1,CG2,CW0,CW1,CRG_up,CRG_down,CRL_up,CRL_down,nsample,PW_f):
model = ConcreteModel()
# Defining the variables
model.PG = Var(range(NG),within = NonNegativeReals, initialize=0)
model.PC = Var(range(NL),within = NonNegativeReals, initialize=0)
model.RG_down = Var(range(NG),within = NonNegativeReals, initialize=0)
model.RG_up = Var(range(NG),within = NonNegativeReals, initialize=0)
model.RG = Var(range(NG),range(nsample),within = Reals, initialize=0)
model.RL_down = Var(range(NL),within = NonNegativeReals, initialize=0)
model.RL_up = Var(range(NL),within = NonNegativeReals, initialize=0)
model.RL = Var(range(NL),range(nsample),within = Reals, initialize=0)
model.dG_down = Var(range(NG),within = NonNegativeReals, initialize=0)
model.dG_up = Var(range(NG),within = NonNegativeReals, initialize=0)
model.dL_down = Var(range(NL),within = NonNegativeReals, initialize=0)
model.dL_up = Var(range(NL),within = NonNegativeReals, initialize=0)
model.Va = Var(range(NB), within=Reals, initialize=0, bounds=(-1,1))
model.s1 = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.s2 = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.s3 = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.s4 = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.s5 = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.s6 = Var(range(nsample), within=NonNegativeReals, initialize=0)
# model.s7 = Var(range(nsample), within=NonNegativeReals, initialize=0)
# model.s8 = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.d = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.e = Var(range(nsample), within=NonNegativeReals, initialize=0)
model.dual = Suffix(direction=Suffix.IMPORT)
## Defining the deterministic constraints
model.c = ConstraintList()
# Power mismatch distribution
model.c.add(sum(model.dG_down[i] for i in range(NG)) + sum(model.dL_up[j] for j in range(NL)) == 1)
model.c.add(sum(model.dG_up[i] for i in range(NG)) + sum(model.dL_down[j] for j in range(NL)) == 1)
# Line flow limit constraints
for i in range(NBr):
model.c.add(sum(Bf[i,j]*model.Va[j] for j in range(NB)) <= Lmax[i])
model.c.add(sum(Bf[i,j]*model.Va[j] for j in range(NB)) >= Lmin[i])
PB_all = np.zeros((NL,nsample))
P_mis_wind_all = np.zeros(nsample)
identified_VS_all = np.zeros((NL,nsample,5))
# Defining the CC constraints
for i in range(nsample):
for j in range(NG):
# Generation capacity limit
model.c.add(model.PG[j] + model.RG[j,i] >= Pmin[j])
model.c.add(model.PG[j] + model.RG[j,i] <= Pmax[j])
# Reserve capacity limit
model.c.add(model.RG[j,i] >= -model.RG_down[j])
model.c.add(model.RG[j,i] <= model.RG_up[j])
# DC Power flow constraint
for j in range(NB):
model.c.add(sum(CG[j,k]*model.PG[k] + CG[j,k]*model.RG[k,i] for k in range(NG)) + CW[j]*PW_sample[i]\
- sum(CL[j,k]*PL[k] + CL[j,k]*model.PC[k] + CL[j,k]*model.RL[k,i] for k in range(NL)) >= sum(B[j,k]*model.Va[k] for k in range(NB)) - model.s1[i])
model.c.add(sum(CG[j,k]*model.PG[k] + CG[j,k]*model.RG[k,i] for k in range(NG)) + CW[j]*PW_sample[i]\
- sum(CL[j,k]*PL[k] + CL[j,k]*model.PC[k] + CL[j,k]*model.RL[k,i] for k in range(NL)) <= sum(B[j,k]*model.Va[k] for k in range(NB)) + model.s2[i])
# model.c.add(sum(CG[j,k]*model.PG[k] + CG[j,k]*model.RG[k,i] for k in range(NG)) + CW[j]*PW_sample[i]\
# - sum(CL[j,k]*PL[k] + CL[j,k]*model.PC[k] for k in range(NL)) >= sum(B[j,k]*model.Va[k] for k in range(NB)) - model.s1[i])
# model.c.add(sum(CG[j,k]*model.PG[k] + CG[j,k]*model.RG[k,i] for k in range(NG)) + CW[j]*PW_sample[i]\
# - sum(CL[j,k]*PL[k] + CL[j,k]*model.PC[k] for k in range(NL)) <= sum(B[j,k]*model.Va[k] for k in range(NB)) + model.s2[i])
P_mis_wind = PW_sample[i] - PW_f
P_mis_wind_all[i] = P_mis_wind
PB = np.zeros(NL)
# print(i)
# P_mis_load = 0
for j in range(NL):
identified_VS = VS(0.25,Tamb_sample1[i],num_TCL[j],Tset[0:num_TCL[j],j],delta,R[0:num_TCL[j],j],C[0:num_TCL[j],j],P_el[0:num_TCL[j],j],COP[0:num_TCL[j],j],baseMVA)
# print("VS0", j, identified_VS[0])
identified_VS_all[j,i,:] = identified_VS
PB[j] = identified_VS[0]
PB_all[:,i] = PB
model.c.add(model.e[i] >= P_mis_wind - sum(model.PC[j] for j in range(NL)) + sum(PB[j] for j in range(NL)))
model.c.add(model.d[i] >= - P_mis_wind + sum(model.PC[j] for j in range(NL)) - sum(PB[j] for j in range(NL)))
for j in range(NG):
model.c.add(model.RG[j,i] == model.dG_up[j]*model.d[i] - model.dG_down[j]*model.e[i])
for j in range(NL):
model.c.add(model.RL[j,i] == model.dL_up[j]*model.e[i] - model.dL_down[j]*model.d[i])
identified_VS1 = VS(0.25,Tamb_sample1[i],num_TCL[j],Tset[0:num_TCL[j],j],delta,R[0:num_TCL[j],j],C[0:num_TCL[j],j],P_el[0:num_TCL[j],j],COP[0:num_TCL[j],j],baseMVA)
# identified_VS2 = VS(0.25,Tamb_sample2[i],num_TCL,Tset[:,j],delta,R[:,j],C[:,j],P_el[:,j],COP[:,j],baseMVA)
model.c.add(model.PC[j] + model.RL[j,i] >= identified_VS1[1])
model.c.add(model.PC[j] + model.RL[j,i] <= identified_VS1[2])
model.c.add(model.RL[j,i] >= -model.RL_down[j])
model.c.add(model.RL[j,i] <= model.RL_up[j])
model.c.add(S0[j] + 0.25*(model.PC[j] + model.RL[j,i] - identified_VS1[0])*1000*baseMVA >= identified_VS1[3] - model.s3[i])
model.c.add(S0[j] + 0.25*(model.PC[j] + model.RL[j,i] - identified_VS1[0])*1000*baseMVA <= identified_VS1[4] + model.s4[i])
model.c.add(S0[j] + 0.75*(model.PC[j] - identified_VS1[0])*1000*baseMVA + 0.25*(model.PC[j] + model.RL[j,i]\
- identified_VS1[0])*1000*baseMVA >= identified_VS1[3] - model.s5[i])
model.c.add(S0[j] + 0.75*(model.PC[j] - identified_VS1[0])*1000*baseMVA + 0.25*(model.PC[j] + model.RL[j,i]\
- identified_VS1[0])*1000*baseMVA <= identified_VS1[4] + model.s6[i])
# model.c.add(S0[j] + 0.75*(model.PC[j] - identified_VS2[0])*1000*baseMVA + 0.25*(model.PC[j] + model.RL[j,i]\
# - identified_VS2[0])*1000*baseMVA >= identified_VS2[3] - model.s7[i])
# model.c.add(S0[j] + 0.75*(model.PC[j] - identified_VS2[0])*1000*baseMVA + 0.25*(model.PC[j] + model.RL[j,i]\
# - identified_VS2[0])*1000*baseMVA <= identified_VS2[4] + model.s8[i])
def cost(model, CG0, CG1, CG2, CRG_down, CRG_up, nsample, CRL_down, CRL_up):
cost = sum(CG0[i] for i in range(NG))\
+ sum(CG1[i]*model.PG[i] for i in range(NG))\
+ sum(CG2[i]*model.PG[i]**2 for i in range(NG))\
+ sum(CRG_up[i]*model.RG_up[i] for i in range(NG))\
+ sum(CRG_down[i]*model.RG_down[i] for i in range(NG))\
+ 1000*sum(model.s1[i] + model.s2[i] for i in range(nsample))\
+ 1e10*sum(model.s3[i] + model.s4[i] + model.s5[i] + model.s6[i] for i in range(nsample))\
+ 1000*sum(model.d[i] + model.e[i] for i in range(nsample))\
+ sum(CRL_up[i]*model.RL_up[i] for i in range(NL))\
+ sum(CRL_down[i]*model.RL_down[i] for i in range(NL))\
# + 1e10*sum(model.s7[i] + model.s8[i] for i in range(nsample))\
return cost
model.obj = Objective(expr=cost(model, CG0, CG1, CG2, CRG_down, CRG_up, nsample, CRL_down, CRL_up), sense=minimize)
return model, PB_all, P_mis_wind_all, identified_VS_all
# +
# Parameters related to the demand side
num_TCL = np.zeros(NL)
for k in range(NL):
num_TCL[k] = 50*np.ceil(PL[k]*baseMVA)
num_TCL = num_TCL.astype(int)
max_num_TCL = np.max(num_TCL).astype(int)
Tset = np.zeros((max_num_TCL, NL))
R = np.zeros((max_num_TCL, NL))
C = np.zeros((max_num_TCL, NL))
COP = np.zeros((max_num_TCL, NL))
P_el = np.zeros((max_num_TCL, NL))
Tset_set = [21,21.5,22,22.5,23]
for i in range(max_num_TCL):
for j in range(NL):
s = random.sample(Tset_set,1)
Tset[i,j] = s[0]
delta = 1
R_nom = 2
R_low = 0.8*R_nom
R_high = 1.2*R_nom
C_nom = 2
C_low = 0.8*C_nom
C_high = 1.2*C_nom
COP_nom = 2.5
COP_low = 0.8*COP_nom
COP_high = 1.2*COP_nom
np.random.seed(0)
for i in range(NL):
R[:,i] = np.random.uniform(R_low,R_high,max_num_TCL)
C[:,i] = np.random.uniform(C_low,C_high,max_num_TCL)
COP[:,i] = np.random.uniform(COP_low,COP_high,max_num_TCL)
for j in range(max_num_TCL):
P_el[j,i] = np.ceil((Tset[j,i] + delta/2 - Tamb_min)/(R[j,i]*COP[j,i])) + 3
# Sampling
eps = 0.1
beta = 0.1
num_decision_variables = 5*NG + 5*NL
num_samples = np.ceil((2/eps)*(np.log(1/beta) + num_decision_variables)).astype(int)
num_samples2 = np.ceil((num_decision_variables + 1 + np.log(1/beta) + np.sqrt(2*(num_decision_variables+1)*np.log(1/beta)))/eps).astype(int)
# np.random.seed(1)
# Tamb_sample = np.random.normal(mu_Tamb,sigma_Tamb,num_samples)
# PW_sample = np.random.choice(PW_data[i,:], size=num_samples)
P_B = np.zeros((NL,1))
Pdown = np.zeros((NL,1))
Pup = np.zeros((NL,1))
Sdown = np.zeros((NL,1))
Sup = np.zeros((NL,1))
S0 = np.zeros((NL,1))
for j in range(NL):
[P_B[j],Pdown[j],Pup[j],Sdown[j],Sup[j]] = \
VS(0.25,Tamb[0],num_TCL[j],Tset[0:num_TCL[j],j],delta,R[0:num_TCL[j],j],C[0:num_TCL[j],j],P_el[0:num_TCL[j],j],COP[0:num_TCL[j],j],baseMVA)
S0[j] = (Sdown[j] + Sup[j])/2
# -
# Sampling
eps = 0.2
beta = 0.2
num_decision_variables = 5*NG + 5*NL
num_samples = np.ceil((2/eps)*(np.log(1/beta) + num_decision_variables)).astype(int)
num_samples2 = np.ceil((num_decision_variables + 1 + np.log(1/beta) + np.sqrt(2*(num_decision_variables+1)*np.log(1/beta)))/eps).astype(int)
print(num_samples)
print(num_samples2)
# +
# Single period
# Define the model
# nsample= 50
# np.random.seed(1)
# Tamb_sample = np.random.normal(Tamb[0],std_Tamb[0],num_samples)
# PW_sample = np.random.choice(PW_data[0,:], size=num_samples)
# PW_f = np.median(PW_data[0,:])
# # PW_f = np.median(PW_data)
# model, PB_all, P_mis_wind, identified_VS_all = optimizer(NG,NL,
# Tamb_sample,PW_sample,num_TCL,Tset,delta,R,C,P_el,COP,S0,PL,\
# Pmax,Pmin,Lmax,Lmin,CG,CW,CL,CG0,CG1,CG2,CW0,CW1,CRG_up,CRG_down,CRL_up,CRL_down,nsample,PW_f)
# # Solve the problem
# solver = SolverFactory("bonmin")
# solver.solve(model)
# model.display()
# -
def cost(model,CG0,CG1,CG2,CRG_up,CRG_down,CRL_up,CRL_down,NG,NL):
r_pg = np.zeros(NG)
r_rg_up = np.zeros(NG)
r_rg_down = np.zeros(NG)
r_rl_up = np.zeros(NL)
r_rl_down = np.zeros(NL)
r_pc = np.zeros(NL)
gen_cost = np.zeros(NG)
gen_r_cost = np.zeros(NG)
load_r_cost = np.zeros(NL)
for i in range(NG):
r_pg[i] = model.PG[i]()
r_rg_up[i] = model.RG_up[i]()
r_rg_down[i] = model.RG_down[i]()
gen_cost[i] = CG0[i] + CG1[i]*r_pg[i] + CG2[i]*r_pg[i]**2
gen_r_cost[i] = CRG_up[i]*r_rg_up[i] + CRG_down[i]*r_rg_down[i]
for i in range(NL):
r_rl_up[i] = model.RL_up[i]()
r_rl_down[i] = model.RL_down[i]()
r_pc[i] = model.PC[i]()
load_r_cost[i] = CRL_up[i]*r_rl_up[i] + CRL_down[i]*r_rl_down[i]
total_cost = np.sum(gen_cost + gen_r_cost) + np.sum(load_r_cost)
return r_pg,r_rg_up,r_rg_down,r_rl_up,r_rl_down,r_pc,gen_cost,gen_r_cost,load_r_cost,total_cost
# +
# Multi-period
import time
S0_all = np.zeros((24,NL))
R_PC = np.zeros((24,NL))
models = []
Tamb_samples1 = []
Tamb_samples2 = []
PW_samples = []
PW_fs = []
PB_alls = []
P_mis_wind_alls = []
identified_VS_alls = []
P_B_test = []
Pdown_test = []
Pup_test = []
Sdown_test = []
Sup_test = []
# Initialize at hour 0
for j in range(NL):
[P_B[j],Pdown[j],Pup[j],Sdown[j],Sup[j]] = \
VS(0.25,Tamb[0],num_TCL[j],Tset[0:num_TCL[j],j],delta,R[0:num_TCL[j],j],C[0:num_TCL[j],j],P_el[0:num_TCL[j],j],COP[0:num_TCL[j],j],baseMVA)
S0_all[0,j] = (Sdown[j] + Sup[j])/2
nsample= 453
for h in range(24):
print("Hour ", h)
t_start = time.time()
print("S0: ", S0_all[h])
np.random.seed(1)
Tamb_sample1 = np.random.normal(Tamb[h],std_Tamb[h],nsample)
# Tamb_sample1 = Tamb[h]*np.ones(nsample)
Tamb_sample2 = np.random.normal(Tamb[h+1],std_Tamb[h+1],nsample)
PW_sample = np.random.choice(PW_data[h,:], size=nsample)
PW_f = np.median(PW_data[h, :])
# PW_sample = PW_f*np.ones(nsample)
# PW_sample[PW_sample > 0.15] = 0.1
# PW_f = np.median(PW_sample)
# Define the problem
model, PB_all, P_mis_wind_all, identified_VS_all = optimizer(NG,NL,Tamb_sample1,Tamb_sample2,PW_sample,num_TCL,Tset,delta,R,C,P_el,COP,S0_all[h,:],PL,\
Pmax,Pmin,Lmax,Lmin,CG,CW,CL,CG0,CG1,CG2,CW0,CW1,CRG_up,CRG_down,CRL_up,CRL_down,nsample,PW_f)
# print("S up: ", identified_VS_all[4])
# Solve the problem
solver = SolverFactory("bonmin")
solver.solve(model)
r_pc = model.PC[:]()
R_PC[h,:] = r_pc
models.append(model)
Tamb_samples1.append(Tamb_sample1)
# Tamb_samples2.append(Tamb_sample2)
PW_samples.append(PW_sample)
PW_fs.append(PW_f)
PB_alls.append(PB_all)
P_mis_wind_alls.append(P_mis_wind_all)
identified_VS_alls.append(identified_VS_all)
if h != 23:
for j in range(NL):
[P_B[j],Pdown[j],Pup[j],Sdown[j],Sup[j]] = \
VS(0.25,Tamb[h],num_TCL[j],Tset[0:num_TCL[j],j],delta,R[0:num_TCL[j],j],C[0:num_TCL[j],j],P_el[0:num_TCL[j],j],COP[0:num_TCL[j],j],baseMVA)
S0_all[h+1,j] = S0_all[h,j] + r_pc[j]*1000*baseMVA - P_B[j]*1000*baseMVA
print("S up %d: %f" %(j, Sup[j]))
P_B_test.append(P_B)
Pdown_test.append(Pdown)
Pup_test.append(Pup)
Sdown_test.append(Sdown)
Sup_test.append(Sup)
t_end = time.time()
print("Elapsed: ", t_end - t_start)
# +
import pickle
case_name = "case39_with_DR"
# Write models
file = open("Results/models_%s_%d" %(case_name,nsample), "wb")
pickle.dump(models, file)
file.close()
# Write S0_all
file = open("Results/S0all_%s_%d" %(case_name,nsample),"wb")
pickle.dump(S0_all, file)
file.close()
# Write R_PC
file = open("Results/R_PC_%s_%d" %(case_name,nsample), "wb")
pickle.dump(R_PC, file)
file.close()
# Write Tamb_samples1
file = open("Results/Tamb_samples1_%s_%d" %(case_name,nsample), "wb")
pickle.dump(Tamb_samples1, file)
file.close()
# Write Tamb_samples2
file = open("Results/Tamb_samples2_%s_%d" %(case_name,nsample), "wb")
pickle.dump(Tamb_samples2, file)
file.close()
# Write PW_samples
file = open("Results/PW_samples_%s_%d" %(case_name,nsample), "wb")
pickle.dump(PW_sample, file)
file.close()
# Write PW_fs
file = open("Results/PW_fs_%s_%d" %(case_name,nsample), "wb")
pickle.dump(PW_fs, file)
file.close()
# Write PB
file = open("Results/PB_alls_%s_%d" %(case_name,nsample), "wb")
pickle.dump(PB_alls, file)
file.close()
# Write identified_VS_alls
file = open("Results/VS_alls_%s_%d" %(case_name,nsample), "wb")
pickle.dump(identified_VS_alls, file)
file.close()
# Write Pmis_winds
file = open("Results/P_mis_winds_%s_%d" %(case_name,nsample), "wb")
pickle.dump(P_mis_wind_alls, file)
file.close()
# Write P_B_test
file = open("Results/P_B_test_%s_%d" %(case_name,nsample), "wb")
pickle.dump(P_B_test, file)
file.close()
# Write Pup_test
file = open("Results/Pup_test_%s_%d" %(case_name,nsample), "wb")
pickle.dump(Pup_test, file)
file.close()
# Write Pdown_test
file = open("Results/Pdown_test_%s_%d" %(case_name,nsample), "wb")
pickle.dump(Pdown_test, file)
file.close()
# Write Sup_test
file = open("Results/Sup_test_%s_%d" %(case_name,nsample), "wb")
pickle.dump(Sup_test, file)
file.close()
# Write Sdown_test
file = open("Results/Sdown_test_%s_%d" %(case_name,nsample), "wb")
pickle.dump(Sdown_test, file)
file.close()
| SOPF with DR_case39.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib
# ### Material utilizado em nosso laboratório:
#
# https://github.com/CSSEGISandData/COVID-19
#
# #### Casos Confirmados
# Importando a base .csv atualizada
casos = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv')
# Lista os últimos itens do dataframe
casos.tail()
# Quantitativo de Casos Covid-19 no Mundo
#
# Temos dados atualizados todos os dias.
#
# *Objetivo: Realizar resposta a seguinte pergunta: No mundo qual o total de casos confirmados?*
#
# ##### x.iloc[linhas,colunas]
# ##### : todas as linhas
casos.shape
# shape 0 = primeiro valor ou Linhas
# shape 1 = segundo valor ou colunas
casos.iloc[: , 4 : casos.shape[1]]
# ### Como somar estes valores?
# Vamos utilizar a numpy
np.sum(casos.iloc[: , 4 : casos.shape[1]])
# Ajustar esta visualização com o Pandas (ver últimos 10 dias)
# ### Pandas irá nos ajudar em "Casos" criando um novo dataframe
casos = np.sum(casos.iloc[: , 4 : casos.shape[1]])
casos.index = pd.DatetimeIndex(casos.index)
casos.tail(10)
casos.plot()
# <img src="https://www.fortenanoticia.com.br/wp-content/uploads/2020/01/China-registra-80-mortes-por-coronav%C3%ADrus-741x483.jpg" width=350 height=150>
# ### Vamos verificar as mortes - Mundo
#
mortes = pd.read_csv("https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv")
mortes.shape
mortes.tail()
mortes.iloc[: , 4 : mortes.shape[1]]
mortes = np.sum(mortes.iloc[ : , 4 : mortes.shape[1]])
mortes.index = pd.DatetimeIndex(mortes.index)
mortes.tail(10)
mortes.plot()
mortalidade = (casos/mortes)*100
mortalidade.plot()
# ### Recuperados
#
# Qual o número de recuperados no mundo
recuperados = pd.read_csv("https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv")
recuperados.shape
recuperados.tail()
recuperados = np.sum(recuperados.iloc[ : , 4 : recuperados.shape[1]])
recuperados.index = pd.DatetimeIndex(recuperados.index)
recuperados.tail(10)
recuperados.plot()
ativos = casos - (recuperados+mortes)
ativos.plot()
# ### Novo dataframe com todos estes dados que manipulamos
#
df = pd.concat([casos, ativos, recuperados, mortes], axis=1)
df.columns = (['casos', 'ativos', 'recuperados','mortes'])
df.tail()
df.plot()
| Python_fuzzy/Aula_IA_Covid_19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison of the accuracy of a cutting plane active learning procedure using the (i) analytic center; (ii) Chebyshev center; and (iii) random center on the diabetes data set
# # The set up
# +
import numpy as np
import pandas as pd
import active
import experiment
import logistic_regression as logr
from sklearn import datasets # The Iris dataset is imported from here.
from IPython.display import display
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 1
# %aimport active
# %aimport experiment
# %aimport logistic_regression
np.set_printoptions(precision=4)
# -
plt.rcParams['axes.labelsize'] = 15
plt.rcParams['axes.titlesize'] = 15
plt.rcParams['xtick.labelsize'] = 15
plt.rcParams['ytick.labelsize'] = 15
plt.rcParams['legend.fontsize'] = 15
plt.rcParams['figure.titlesize'] = 18
# # Importing and processing the diabetes data set
# In this experiment we work with a data set with 2 classes and 8 features where the classes are divided between whether or not a patient has diabetes and the 8 features correspond to 8 health measurements.
#
# The 2 classes in this data set are not linearly seperable and it is known that the data set has missing variables.
#
# We work with all features of the data set and randomly divide the data set into two halves, to be used for training and testing.
names = ['diabetes', 'num preg', 'plasma', 'bp', 'skin fold', 'insulin', 'bmi', 'pedigree', 'age']
data = pd.read_csv('diabetes_scale.csv', header=None, names=names)
data['ones'] = np.ones((data.shape[0], 1)) # Add a column of ones
data.head()
np.random.seed(1)
size = data.shape[0]
index = np.arange(size)
np.random.shuffle(index)
training_index = index[:int(size/2)]
testing_index = index[int(size/2):]
# # Experimental procedure
# See Section 7.5 of the report.
# # Logistic regression
# +
Y = data['diabetes']
X = data[['num preg', 'plasma', 'bp', 'skin fold', 'insulin', 'bmi', 'pedigree', 'age', 'ones']]
X = np.array(X)
Y = np.array(Y)
Y[Y==-1] = 0
X_diabetes_training = X[training_index]
Y_diabetes_training = Y[training_index]
X_diabetes_testing = X[testing_index]
Y_diabetes_testing = Y[testing_index]
# -
print(X_diabetes_testing)
print(Y_diabetes_testing)
print(X_diabetes_training)
print(Y_diabetes_training)
# # Using logistic regression as a benchmark
n = 10
iterations = 15
X_training = X_diabetes_training
Y_training = Y_diabetes_training
X_testing = X_diabetes_testing
Y_testing = Y_diabetes_testing
# Here we compute the average accuracy over 10 tests, where for each test the accuracy is computed by training logistic regression on 1 to 15 randomly selected patterns from the same fixed training set, which comprises a uniform sample of half the data set.
#
# We note here that the optimization process sometimes fails to converge.
average_accuracies_logr_15 = \
logr.experiment(n, iterations, X_testing, Y_testing, X_training, Y_training)
print(average_accuracies_logr_15)
# Here we compute the average accuracy over 10 tests, where for each test the accuracy is computed by training logistic regression on 1 to 30 randomly selected patterns from the same fixed training set, which comprises a uniform sample of half the data set.
#
# We compute the accuracy from training logistic regression on 50% and 100% of the training data set also.
#
# We note here that the optimization process sometimes fails to converge.
iterations = 30
average_accuracies_logr_30 = \
logr.experiment(n, iterations, X_testing, Y_testing, X_training, Y_training)
print(average_accuracies_logr_30)
# Here we train logistic regression on 50% of the training data set, selected random uniformly, and repeat this 10 times to compute the average accuracy.
n = 10
size = X_training.shape[0]
size_half = int(size/2)
accuracies = []
for i in range(n):
index_all = np.arange(size)
np.random.shuffle(index_all)
index_half = index_all[:size_half]
X_training_half = X_training[index_half]
Y_training_half = Y_training[index_half]
w_half = logr.train(X_training_half, Y_training_half)
predictions = logr.predict(w_half, X_training_half)
accuracy = logr.compute_accuracy(predictions, Y_training_half)
accuracies.append(accuracy)
accuracies = np.array(accuracies)
average_accuracy_training_half = np.sum(accuracies)/n
print('The average accuracy training on 50% of the training data set is',\
average_accuracy)
# Here we simply train logistic regression on half of the data and test it on the other half.
w_training = logr.train(X_training, Y_training)
predictions = logr.predict(w_training, X_testing)
accuracy_training_all = logr.compute_accuracy(predictions, Y_testing)
print("The accuracy training on the whole training data set is", accuracy)
# # Average accuracy of the cutting plane active learning procedure over 10 tests and 15 iterations using the (i) analytic center; (ii) Chebyshev center; and (iii) random center
# +
Y = data['diabetes']
X = data[['num preg', 'plasma', 'bp', 'skin fold', 'insulin', 'bmi', 'pedigree', 'age', 'ones']]
X = np.array(X)
Y = np.array(Y)
X_diabetes_training = X[training_index]
Y_diabetes_training = Y[training_index]
X_diabetes_testing = X[testing_index]
Y_diabetes_testing = Y[testing_index]
# -
n = 10
iterations = 15
X_testing = X_diabetes_testing
Y_testing = Y_diabetes_testing
X_training = X_diabetes_training
Y_training = Y_diabetes_training
average_accuracies_ac_15 = \
experiment.experiment(n, iterations, X_testing, Y_testing,
X_training, Y_training, center='ac',
sample=1, M=None)
average_accuracies_cc_15 = \
experiment.experiment(n, iterations, X_testing, Y_testing,
X_training, Y_training, center='cc',
sample=1, M=None)
average_accuracies_rand_15 = \
experiment.experiment(n, iterations, X_testing, Y_testing,
X_training, Y_training, center='random',
sample=1, M=None)
# +
plt.figure(figsize=(12,7))
queries = np.arange(1, iterations + 1)
plt.plot(queries, average_accuracies_logr_15, 'mx-', label='LR',
markevery=5,
lw=1.5, ms=10, markerfacecolor='none', markeredgewidth=1.5,
markeredgecolor = 'm')
plt.plot(queries, average_accuracies_ac_15, 'r^-', label='AC',
markevery=5,
lw=1.5, ms=10, markerfacecolor='none', markeredgewidth=1.5,
markeredgecolor = 'r')
plt.plot(queries, average_accuracies_cc_15, 'go-', label='CC',
markevery=5,
lw=1.5, ms=10, markerfacecolor='none', markeredgewidth=1.5,
markeredgecolor = 'g')
plt.plot(queries, average_accuracies_rand_15, 'bs-', label='Random',
markevery=5,
lw=1.5, ms=10, markerfacecolor='none', markeredgewidth=1.5,
markeredgecolor = 'b')
plt.plot(queries, [average_accuracy_training_half]*queries.shape[0], 'k--',
color = '0.4', label='LR - half',lw=1.5, ms=10)
plt.plot(queries, [accuracy_training_all]*queries.shape[0], 'k-',
color = '0.4', label='LR - all',lw=1.5, ms=10)
plt.xlabel('Number of iterations')
plt.ylabel('Accuracy averaged over %d tests' % n)
plt.title('Average accuracy of a cutting plane active learning procedure (diabetes data set)')
plt.legend(loc='best')
plt.savefig('diabetes_experiment_all_15.png', dpi=600, bbox_inches='tight', transparent=True)
plt.show()
| projects/david/lab/experiment_active_diabetes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''venv'': venv)'
# name: python382jvsc74a57bd0265455998716544771fe8f8e46d3bc336b7ce25fc2914091372dd288abc92a2d
# ---
import pandas as pd
log = pd.read_csv("./00_log.csv", header = None)
users = pd.read_csv("./00_users.csv", encoding="koi8-r", sep="\t")
log.columns = ["user_id","time","bet","win"]
# Посчитайте количество пропусков в столбце time. Метод isna() есть не только у DataFrame, но и у Series. Это значит, что применять его можно не только ко всей таблице, но и к каждому столбцу отдельно.
log.time.isna().value_counts()[True]
# Удалите все столбцы, где есть пропуски. Запишите в поле, сколько осталось столбцов в данных после этого.
tmp_log = log.copy()
tmp_log.dropna(axis=1,inplace=True)
tmp_log.info()
# Удалите все строки, где есть пропуски. Запишите в поле, сколько осталось строк в данных после этого.
tmp_log = log.copy()
tmp_log.dropna(axis=0, inplace=True)
tmp_log.info()
tmp_log = log.copy()
tmp_log.loc[10].isna()
# Удалите дубли среди столбцов user_id и time. Запишите в поле ниже, сколько осталось строк после удаления дублей.
#
tmp_log = log.copy()
tmp_log.drop_duplicates(subset=["user_id", "time"], inplace=True)
tmp_log.info()
log.time = log.time.apply(lambda s: s[1:] if type(s) == str else s)
log.head()
log.time = pd.to_datetime(log.time)
type(log.loc[3].time)
log.dropna(axis=0, inplace=True, subset=["time"])
log.info()
log.time.max()
log.time.apply(lambda x: x.minute).loc[13]
log = pd.read_csv("./00_log.csv", header = None)
log = log.dropna()
log.columns = ['user_id', 'time', 'bet', 'win']
log['time'] = log['time'].apply(lambda x: x[1:])
log['time'] = pd.to_datetime(log['time'])
log['time'] = log.time.apply(lambda x: x.minute)
log['time'].head()
| unit_3/python_12.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Evaluation metrics
# ---
#
# _You are currently looking at **version 1.0** of this notebook._
#
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
# -
df = pd.read_csv('../_data/fraud_data.csv')
# ### Label stats / distribution
df['Class'].describe()
np.bincount(df['Class'])
# ### Train-test split
X, y = df.iloc[:,:-1], df.iloc[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# ### Confusion matrix of dummy classifier
#
# Using `X_train`, `X_test`, `y_train`, and `y_test` (as defined above), train a dummy classifier that classifies everything as the majority class of the training data. What is the accuracy of this classifier? What is the recall?
#
# *This function should a return a tuple with two floats, i.e. `(accuracy score, recall score)`.*
from sklearn.dummy import DummyClassifier
from sklearn.metrics import confusion_matrix
# +
# Fit DummyClassifier
dummy_majority = DummyClassifier(strategy='most_frequent').fit(X_train, y_train)
# Predict
y_dummy_predictions = dummy_majority.predict(X_test)
# Get scores (manually)
cm = confusion_matrix(y_test, y_dummy_predictions)
cm
# +
### Accuracy and recall scores
# -
from sklearn.metrics import accuracy_score, recall_score, precision_score
from sklearn.svm import SVC
# +
TN, FP = cm[0, 0], cm[0, 1]
FN, TP = cm[1, 0], cm[1, 1]
# or
TN, FP, FN, TP = cm.ravel()
# Calculated
accuracy_sc_m = float((TN + TP) /(TN + FP + FN + TP))
recall_sc_m = float(TP /(TP + FN))
precision_sc_m = float(TP /(TP + FP + 10e-8)) # *avoid div/zero
# Function
accuracy_sc = accuracy_score(y_test, y_dummy_predictions)
recall_sc = recall_score(y_test, y_dummy_predictions)
precision_sc = precision_score(y_test, y_dummy_predictions)
accuracy_sc_m, recall_sc_m, precision_sc_m
accuracy_sc, recall_sc, precision_sc
# +
svm = SVC().fit(X_train, y_train)
y_predicted = svm.predict(X_test)
accuracy_sc = accuracy_score(y_test, y_predicted)
recall_sc = recall_score(y_test, y_predicted)
precision_sc = precision_score(y_test, y_predicted)
accuracy_sc, recall_sc, precision_sc
# -
# ### Confusion matrix using decision function with threshold
#
# - What is the confusion matrix when using a fraud threshold of -220?
# - Decision function returns prediction values for y, which can be converted to labels using a threshold
# - One can reduce the sensitivity(recall) and reduce the number of FN/falsely convicted/negative diagnosed
# +
svm = SVC(C=1e9, gamma=1e-07).fit(X_train, y_train)
fraud_threshold = np.linspace(100, 350, 6) * -1
for thres in fraud_threshold:
y_decision_scores = svm.decision_function(X_test) > thres
print('Fraud threshold: {}\n'.format(thres), confusion_matrix(y_test, y_decision_scores.astype('int')))
print('Sensitivity: {:.3f}\n'.format(recall_score(y_test, y_decision_scores)))
# -
# ### Question 5
#
# Train a logisitic regression classifier with default parameters using X_train and y_train.
#
# For the logisitic regression classifier, create a precision recall curve and a roc curve using y_test and the probability estimates for X_test (probability it is fraud).
#
# Looking at the precision recall curve, what is the recall when the precision is `0.75`?
#
# Looking at the roc curve, what is the true positive rate when the false positive rate is `0.16`?
#
# *This function should return a tuple with two floats, i.e. `(recall, true positive rate)`.*
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import roc_curve, auc, precision_recall_curve
# Fit lr, return probabilities
lr = LogisticRegression().fit(X_train, y_train)
y_proba = lr.predict_proba(X_test)[: ,1]
# +
plt.figure(figsize=(7, 7))
ax = plt.gca()
ax.set_xlim([0.0, 1.01]) # OOP
plt.ylim([0.0, 1.01]) # pyplot
# PR curve
precision, recall, _ = precision_recall_curve(y_test, y_proba)
ax.plot(precision, recall, label='Precision-Recall Curve')
ax.set_ylabel('Recall', fontsize=16)
ax.set_xlabel('Precision', fontsize=16)
# ROC curve
false_positive_rate, recall, _ = roc_curve(y_test, y_proba)
ax.plot(false_positive_rate, recall, label='ROC Curve')
ax.set_ylabel('True Positive Rate', fontsize=16)
ax.set_xlabel('False Positive Rate', fontsize=16)
ax.set_aspect('equal')
plt.show();
# -
# ### Cross validation by GridSearchCV
#
# - Perform a grid search over a selection of hyperparameters.
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
# +
lr = LogisticRegression(random_state=0)
C = [0.01, 0.1, 1, 10, 100]
penalty = ['l1', 'l2']
grid_values = {'C':C, 'penalty': penalty}
grid_lr_prec = GridSearchCV(lr,
param_grid=grid_values,
scoring='recall',
cv=5,
return_train_score=True)
grid_lr_prec.fit(X_train, y_train)
# -
# #### Mean test scores of each hyperparameter combination
df = pd.DataFrame(grid_lr_prec.cv_results_)
pivot = pd.pivot_table(df, values='mean_test_score', index=['param_C'], columns = ['param_penalty']).as_matrix()
pivot
# #### Plot mean scores
plt.figure()
sns.heatmap(pivot.reshape(5, 2), xticklabels=penalty, yticklabels=C, vmin=0.77, vmax=0.81)
plt.yticks(rotation=0);
| metrics/evaluation_metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ToCsmp1pc1wS"
# <table class="tfo-notebook-buttons" align="center">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/PracticalDL/Practical-Deep-Learning-Book/blob/master/code/chapter-5/1-develop-tool.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/PracticalDL/Practical-Deep-Learning-Book/blob/master/code/chapter-5/1-develop-tool.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="Pknp7a70h6IF"
# # Chapter 5 - From Novice to Master Predictor: Maximizing Convolutional Neural Network Accuracy
#
#
# We explore strategies to maximize the accuracy that our classifier can achieve, with the help of a range of tools including TensorBoard, What-If Tool, tf-explain, TensorFlow Datasets, AutoKeras, AutoAugment. Along the way, we conduct experiments to develop an intuition of what parameters might or might not work for your AI task. [Read online here.](https://learning.oreilly.com/library/view/practical-deep-learning/9781492034858/ch05.html)
#
# ## Tools
#
# In this file, we will develop a tool to experiment with various parameter settings of a model. One can choose amongst different kinds of augmentation techniques, use different datasets available in TensorFlow Datasets, choose to train either from scratch or use finetune from MobileNet or any model of your choice, all in the browser without any framework installs on your system.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="hJduD52zNu0Q" outputId="406835cb-d9a7-43f5-d522-47f7260b04ef"
# Perform all installations
# !pip install tensorflow-gpu==2.0.0
# !pip install tensorflow-datasets
# !pip install tensorwatch
# Get TensorBoard to run
# %load_ext tensorboard
# + colab={"base_uri": "https://localhost:8080/", "height": 471} colab_type="code" id="4UcsyIiLQr_9" outputId="407b6de2-003a-4987-a2b5-9d0f54d334b0"
# Import necessary packages
import tensorflow as tf
import tensorflow_datasets as tfds
# tfds makes a lot of progress bars and they take up a lot of screen space, so lets diable them
tfds.disable_progress_bar()
import math
import numpy as np
from keras.preprocessing import image
from tensorflow.keras.callbacks import CSVLogger
# + [markdown] colab_type="text" id="hZ7ZvJY-r2vY"
# To make experiments reproducible across runs, we control the amount of randomization possible. Randomization is introduced in initialization of weights of models, randomized shuffling of data.
#
# Random number generators can be made reproducible by initializing a seed and that’s exactly what we will do. Various frameworks have their own ways of setting a random seed, some of which are shown below:
# + colab={} colab_type="code" id="Kww2jNggsSm9"
tf.random.set_seed(1234)
np.random.seed(1234)
# + colab={} colab_type="code" id="QI_ZhgbpxcEE"
# Variables
BATCH_SIZE = 32
NUM_EPOCHS = 100
IMG_H = IMG_W = 224
IMG_SIZE = 224
LOG_DIR = './log'
DEGREES = 10 #for rotation
SHUFFLE_BUFFER_SIZE = 1024
IMG_CHANNELS = 3
# + [markdown] colab_type="text" id="Rv3IASONQeVt"
# ## Dataset
#
# Choose the dataset that we want to experiment on. We have tried to build this tool in such a way that it works with all the image datasets available in `TensorFlow Datasets`.
#
# To see all available datasets in `TensorFlow Datasets`, use the following `print` command.
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="eo8WzErmRz9C" outputId="7f2d7d9f-703c-41c3-a048-9d1b6fa2c32f"
# View all available datasets
print(tfds.list_builders())
# + colab={} colab_type="code" id="Jo_r3u6kQcZ6"
# Choose dataset
#dataset_name = "colorectal_histology"
#dataset_name = "caltech101"
#dataset_name = "oxford_flowers102"
dataset_name = "cats_vs_dogs"
# + [markdown] colab_type="text" id="n9pr4pVkSX2D"
# ### Dataset Preprocessing and Augmenting
#
# Let's define some preprocessing and augmentation functions.
#
# Note that bicubic resizing functionality is not available in TFDS yet
#
# All `tf.image` augmentations defined at https://www.tensorflow.org/api_docs/python/tf/image.
# + colab={} colab_type="code" id="fAp_67kDN_n0"
def preprocess(ds):
x = tf.image.resize_with_pad(ds['image'], IMG_SIZE, IMG_SIZE)
x = tf.cast(x, tf.float32)
x = (x / 127.5) - 1
return x, ds['label']
def augmentation(image, label):
#image = tf.image.resize_with_crop_or_pad(image, IMG_SIZE, IMG_SIZE)
# Random Crop: randomly crops an image and fits to given size
#image = tf.image.random_crop(image,[IMG_SIZE, IMG_SIZE, IMG_CHANNELS])
# Brightness: Adjust brightness by a given max_delta
image = tf.image.random_brightness(image, .1)
# Random Contrast: Add a random contrast to the image
image = tf.image.random_contrast(image, lower=0.0, upper=1.0)
# Flip: Left and right
image = tf.image.random_flip_left_right(image)
# Rotation: Only 90 degrees is currently supported.
# Not all images still look the same after a 90 degree rotation
# Most images are augmented by a 10-30 degree tilt
#image = tf.keras.preprocessing.image.random_rotation(image,10)
# Finally return the augmented image and label
return image, label
# + [markdown] colab_type="text" id="iahpaGwaSipp"
# ### Dataset Loading
#
# Develop handy functions to load training and validation data.
#
# Some of the datasets in `TensorFlow Datasets` do not have a `validation` split. For those datasets we take a small percentage of samples from the `training` set, and treat it as the `validation` set. Splitting the dataset using the `weighted_splits` takes care of randomizing and shuffling data between the splits.
# + colab={} colab_type="code" id="hsRP-IrrEPcS"
def get_dataset(dataset_name, *split_value):
# see all possible splits in dataset
_, info = tfds.load(dataset_name, with_info=True)
#print(info)
if "validation" in info.splits:
# then load train and validation
if len(split_value) == 1:
print(
"INFO: Splitting train dataset according to splits provided by user"
)
if "test" in info.splits:
print('INFO: Test dataset is available')
all_data = tfds.Split.TEST + tfds.Split.TRAIN
print("INFO: Added test data to train data")
train, info_train = tfds.load(dataset_name,
split=all_data,
with_info=True)
NUM_CLASSES = info_train.features['label'].num_classes
NUM_EXAMPLES = info_train.splits['train'].num_examples
else:
all_data = tfds.Split.TRAIN
split_train, _ = all_data.subsplit(
weighted=[split_value[0], 100 - split_value[0]])
# Load train with the new split
train, info_train = tfds.load(dataset_name,
split=split_train,
with_info=True)
else:
#load training dataset the without any user intervention way
print("INFO: Loading standard splits for training dataset")
train, info_train = tfds.load(dataset_name,
split=tfds.Split.TRAIN,
with_info=True)
# Load validation dataset as is standard
val, info_val = tfds.load(dataset_name,
split=tfds.Split.VALIDATION,
with_info=True)
NUM_CLASSES = info_train.features['label'].num_classes
NUM_EXAMPLES = info_train.splits['train'].num_examples
else:
# Validation not in default datasets
print(
"INFO: Defining a 90-10 split between training and validation as no default split exists."
)
# Here we have defined how to split the original train dataset into train and val
# Use 90% as train dataset and 10% as validation dataset
split_train, split_val = tfds.Split.TRAIN.subsplit(weighted=[9, 1])
train, info_train = tfds.load(dataset_name,
split=split_train,
with_info=True)
val, info_val = tfds.load(dataset_name,
split=split_val,
with_info=True)
NUM_CLASSES = info_train.features['label'].num_classes
# The total number of classes in training dataset should either be equal to or more than the total number of classes in validation dataset
assert NUM_CLASSES >= info_val.features['label'].num_classes
NUM_EXAMPLES = info_train.splits['train'].num_examples * 0.9
# Standard processing for training and validation set
IMG_H, IMG_W, IMG_CHANNELS = info_train.features['image'].shape
if IMG_H == None or IMG_H != IMG_SIZE:
IMG_H = IMG_SIZE
if IMG_W == None or IMG_W != IMG_SIZE:
IMG_W = IMG_SIZE
if IMG_CHANNELS == None:
IMG_CHANNELS = 3
# Training specific processing
train = train.map(preprocess).repeat().shuffle(SHUFFLE_BUFFER_SIZE).batch(
BATCH_SIZE)
train = train.map(augmentation)
train = train.prefetch(tf.data.experimental.AUTOTUNE)
# Validation specific processing
val = val.map(preprocess).repeat().batch(BATCH_SIZE)
val = val.prefetch(tf.data.experimental.AUTOTUNE)
return train, info_train, val, info_val, IMG_H, IMG_W, IMG_CHANNELS, NUM_CLASSES, NUM_EXAMPLES
# + [markdown] colab_type="text" id="rusRxOdWWZhZ"
# Now that we have defined all our helper functions, let's use them to get the dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="rpIYrRQwg1kx" outputId="1e005fa5-9ada-42d2-c3c0-5b2aeb141b6c"
train, info_train, val, info_val, IMG_H, IMG_W, IMG_CHANNELS, NUM_CLASSES, NUM_EXAMPLES = get_dataset(
dataset_name, 100)
print("\n\nIMG_H, IMG_W", IMG_H, IMG_W)
print("IMG_CHANNELS", IMG_CHANNELS)
print("NUM_CLASSES", NUM_CLASSES)
print("BATCH_SIZE", BATCH_SIZE)
print("NUM_EXAMPLES", NUM_EXAMPLES)
print("NUM_EPOCHS", NUM_EPOCHS)
# If you want to print even more information on both the splits, uncomment the lines below:
#print(info_train)
#print(info_val)
# + [markdown] colab_type="text" id="Chn8a67jWgn4"
# Great!
#
# ## Training
#
# Here we decide what kind of training to perform.
#
# We have defined a model from scratch as well as a model that performs transfer learning on MobileNet.
#
# Depending on what dataset we chose, and how different that dataset is from `ImageNet dataset`, we may learn a lot from the experiments on both kinds of trainings.
# + colab={} colab_type="code" id="HZ1lbZR3dy1i"
#choose "scratch" to train a new model from scratch
training_format = "scratch"
#choose transfer_learning to use finetuning on mobilenet
training_format = "transfer_learning"
# + colab={"base_uri": "https://localhost:8080/", "height": 86} colab_type="code" id="g0A6MFDBXyQr" outputId="e673f9a7-d0aa-448b-fbf5-80d52fb25ee7"
#Allow TensorBoard callbacks
tensorboard_callback = tf.keras.callbacks.TensorBoard(LOG_DIR,
histogram_freq=1,
write_graph=True,
write_grads=True,
batch_size=BATCH_SIZE,
write_images=True)
# -
# ### Model defination for training from scratch
# + colab={} colab_type="code" id="Mpp4mC6pX2uE"
def create_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, (3, 3),
activation='relu',
input_shape=(IMG_SIZE, IMG_SIZE, IMG_CHANNELS)),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
tf.keras.layers.Conv2D(32, (3, 3), activation='relu'),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
tf.keras.layers.Conv2D(32, (3, 3), activation='relu'),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
tf.keras.layers.Dropout(rate=0.3),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(rate=0.3),
tf.keras.layers.Dense(NUM_CLASSES, activation='softmax')
])
return model
def scratch(train, val, learning_rate):
model = create_model()
model.summary()
model.compile(
optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
earlystop_callback = tf.keras.callbacks.EarlyStopping(
monitor='val_accuracy', min_delta=0.0001, patience=5)
csv_logger = CSVLogger('colorectal-scratch-' + 'log.csv',
append=True,
separator=';')
model.fit(train,
epochs=NUM_EPOCHS,
steps_per_epoch=int(NUM_EXAMPLES / BATCH_SIZE),
validation_data=val,
validation_steps=1,
validation_freq=1,
callbacks=[tensorboard_callback, earlystop_callback, csv_logger])
return model
# -
# ### Model Defination for fine-tuning
#
# We will be using the MobileNet model for fine-tuning. We can decide how many layers of the model to train by un-freeze the top layers of the model as follows:
#
# ```
# #Our unfreeze_percentage variable helps us decide how many layers to unfreeze
# unfreeze_percentage = 0.75
#
# #Initially set all layers to fixed, i.e., not trainable
# mobile_net.trainable=False
#
# #Find total number of layers in base model
# num_layers = len(mobile_net.layers)
# print("Total number of layers in MobileNet: ", num_layers)
#
# #Set the last few layers to be trainable
# for layer_index in range(int(num_layers - unfreeze_percentage*num_layers), num_layers):
# print(layer_index, mobile_net.layers[layer_index])
#
# #set the layer to be trainable
# mobile_net.layers[layer_index].trainable = True
# ```
#
# All you need to do is unfreeze the last few layers in the `mobile_net` model or set them trainable. Then, you should recompile the model (necessary for these changes to take effect), and resume training.
# + colab={} colab_type="code" id="4WM_eV6RSyIW"
def transfer_learn(train, val, unfreeze_percentage, learning_rate):
mobile_net = tf.keras.applications.MobileNet(input_shape=(IMG_SIZE,
IMG_SIZE,
IMG_CHANNELS),
include_top=False)
# Use mobile_net.summary() to view the model
mobile_net.trainable = False
# Unfreeze some of the layers according to the dataset being used
num_layers = len(mobile_net.layers)
for layer_index in range(
int(num_layers - unfreeze_percentage * num_layers), num_layers):
mobile_net.layers[layer_index].trainable = True
model_with_transfer_learning = tf.keras.Sequential([
mobile_net,
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(NUM_CLASSES, activation='softmax')
], )
model_with_transfer_learning.compile(
optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate),
loss='sparse_categorical_crossentropy',
metrics=["accuracy"])
model_with_transfer_learning.summary()
earlystop_callback = tf.keras.callbacks.EarlyStopping(
monitor='val_accuracy', min_delta=0.0001, patience=10)
csv_logger = CSVLogger('colorectal-transferlearn-' + 'log.csv',
append=True,
separator=';')
model_with_transfer_learning.fit(
train,
epochs=NUM_EPOCHS,
steps_per_epoch=int(NUM_EXAMPLES / BATCH_SIZE),
validation_data=val,
validation_steps=1,
validation_freq=1,
callbacks=[tensorboard_callback, earlystop_callback, csv_logger])
return model_with_transfer_learning
# + [markdown] colab_type="text" id="OOg-rjnBugSj"
# Recall that `tf.keras.layers.BatchNormalization` and `tf.keras.layers.Dropout` are applicable only during training. They are turned off when calculating validation loss.
#
# Also, remember that training metrics report the average for an epoch, while validation metrics are evaluated after the epoch, so validation metrics see a model that has trained slightly longer.
#
# Run the following cells to start the training and then come back up to look at TensorBoard.
#
# ## TensorBoard
#
# Let's focus on what we can learn fom TensorBoard:
#
# 1. Visualize the training and validation accuracy and loss.
# 2. View the output from each layer by clicking on the `Images` tab. This takes some time to load so grab a coffee!
#
# We can edit the size of the results we are looking at, add additional contrast and brightness.
#
# 3. Visualize the graph of the network that we just trained.
# 4. The `Distributions` tab shows the weight distribution of the weight matrices of each of the layers. This is very useful when quantizing a model. And we will be learning more about this in the later chapters. We can view the histogram of this distribution in the `Histogram` tab.
#
#
#
#
#
# Note: You can ALT+Scroll in and out for zoom
# + colab={"base_uri": "https://localhost:8080/", "height": 836} colab_type="code" id="mWDKvkWARtQ2" outputId="b6fffb82-abfa-41a0-ecc1-815e56b7943a"
# Start TensorBoard
# %tensorboard --logdir ./log
# + colab={"base_uri": "https://localhost:8080/", "height": 714} colab_type="code" id="AU5zrBpccru3" outputId="18f4b088-b998-4e59-d3b5-0053a1c35724"
# select the percentage of layers to be trained while using the transfer learning
# technique. The selected layers will be close to the output/final layers.
unfreeze_percentage = 0
learning_rate = 0.001
if training_format == "scratch":
print("Training a model from scratch")
model = scratch(train, val, learning_rate)
elif training_format == "transfer_learning":
print("Fine Tuning the MobileNet model")
model = transfer_learn(train, val, unfreeze_percentage, learning_rate)
# +
# Save the model to load it in the What-If tool
tf.saved_model.save(model, "tmp/model/1/")
# -
# Load the saved model
loaded = tf.saved_model.load("/tmp/model/1/")
print(list(loaded.signatures.keys())) # ["serving_default"]
# Zip the directory so that we can download it
# !zip model.zip /tmp/model/*
# +
# If you are running this in Google Colab,
# Go to the content directory and download the trained model
# !pwd
# -
# ## Summary
#
# In our experiment, we were able to modify how many layers of the MobileNet V2 base model we wanted to train. Our training process nudged the weights from a more generic feature set to features associated specifically to the dataset at hand. This is relevant for datasets that are quite different from the ImageNet dataset, or are much smaller.
#
# As we learnt in this and the previous chapters, the higher up a layer is, the more specialized it is to the task at hand. The initial layers learn very simple features, like distinguishing an edge, a feature that is common to almost all images. Then, as we proceed to higher layers, features become more specific to the training dataset.
#
# Through fine-tuning, we attempted to nudge these higher-layer/specific features to work with a new dataset while still making use of the generic layers.
| code/chapter-5/1-develop-tool.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
input_str = [6,True]
# Here 1 represents the day of the week and Flase represents if on vacation or not, if on vacation then it will be true.
input_list =(input_str)
day_of_the_week = input_list[0] #first element is an integer denoting the day of the week
is_on_vacation = input_list[1] #this is a boolean denoting if its vacation or not
# write your code here
def alarm_time(day_of_the_week, is_on_vacation):
weekend = [6,7]
if is_on_vacation:
if day_of_the_week in weekend:
return 'off'
else:
return "10:00"
else:
if day_of_the_week in weekend:
return '10:00'
else:
return '7:00'
print(alarm_time(day_of_the_week, is_on_vacation))
# -
| Alarm Clock with Test Cases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Padding and Stride
# :label:`sec_padding`
#
#
# In the previous example of :numref:`fig_correlation`,
# our input had both a height and width of 3
# and our convolution kernel had both a height and width of 2,
# yielding an output representation with dimension $2\times2$.
# As we generalized in :numref:`sec_conv_layer`,
# assuming that
# the input shape is $n_h\times n_w$
# and the convolution kernel shape is $k_h\times k_w$,
# then the output shape will be
# $(n_h-k_h+1) \times (n_w-k_w+1)$.
# Therefore, the output shape of the convolutional layer
# is determined by the shape of the input
# and the shape of the convolution kernel.
#
# In several cases, we incorporate techniques,
# including padding and strided convolutions,
# that affect the size of the output.
# As motivation, note that since kernels generally
# have width and height greater than $1$,
# after applying many successive convolutions,
# we tend to wind up with outputs that are
# considerably smaller than our input.
# If we start with a $240 \times 240$ pixel image,
# $10$ layers of $5 \times 5$ convolutions
# reduce the image to $200 \times 200$ pixels,
# slicing off $30 \%$ of the image and with it
# obliterating any interesting information
# on the boundaries of the original image.
# *Padding* is the most popular tool for handling this issue.
#
# In other cases, we may want to reduce the dimensionality drastically,
# e.g., if we find the original input resolution to be unwieldy.
# *Strided convolutions* are a popular technique that can help in these instances.
#
# ## Padding
#
# As described above, one tricky issue when applying convolutional layers
# is that we tend to lose pixels on the perimeter of our image.
# Since we typically use small kernels,
# for any given convolution,
# we might only lose a few pixels,
# but this can add up as we apply
# many successive convolutional layers.
# One straightforward solution to this problem
# is to add extra pixels of filler around the boundary of our input image,
# thus increasing the effective size of the image.
# Typically, we set the values of the extra pixels to zero.
# In :numref:`img_conv_pad`, we pad a $3 \times 3$ input,
# increasing its size to $5 \times 5$.
# The corresponding output then increases to a $4 \times 4$ matrix.
# The shaded portions are the first output element as well as the input and kernel tensor elements used for the output computation: $0\times0+0\times1+0\times2+0\times3=0$.
#
# 
# :label:`img_conv_pad`
#
# In general, if we add a total of $p_h$ rows of padding
# (roughly half on top and half on bottom)
# and a total of $p_w$ columns of padding
# (roughly half on the left and half on the right),
# the output shape will be
#
# $$(n_h-k_h+p_h+1)\times(n_w-k_w+p_w+1).$$
#
# This means that the height and width of the output
# will increase by $p_h$ and $p_w$, respectively.
#
# In many cases, we will want to set $p_h=k_h-1$ and $p_w=k_w-1$
# to give the input and output the same height and width.
# This will make it easier to predict the output shape of each layer
# when constructing the network.
# Assuming that $k_h$ is odd here,
# we will pad $p_h/2$ rows on both sides of the height.
# If $k_h$ is even, one possibility is to
# pad $\lceil p_h/2\rceil$ rows on the top of the input
# and $\lfloor p_h/2\rfloor$ rows on the bottom.
# We will pad both sides of the width in the same way.
#
# CNNs commonly use convolution kernels
# with odd height and width values, such as 1, 3, 5, or 7.
# Choosing odd kernel sizes has the benefit
# that we can preserve the spatial dimensionality
# while padding with the same number of rows on top and bottom,
# and the same number of columns on left and right.
#
# Moreover, this practice of using odd kernels
# and padding to precisely preserve dimensionality
# offers a clerical benefit.
# For any two-dimensional tensor `X`,
# when the kernel's size is odd
# and the number of padding rows and columns
# on all sides are the same,
# producing an output with the same height and width as the input,
# we know that the output `Y[i, j]` is calculated
# by cross-correlation of the input and convolution kernel
# with the window centered on `X[i, j]`.
#
# In the following example, we create a two-dimensional convolutional layer
# with a height and width of 3
# and (**apply 1 pixel of padding on all sides.**)
# Given an input with a height and width of 8,
# we find that the height and width of the output is also 8.
#
# + origin_pos=2 tab=["pytorch"]
import torch
from torch import nn
# We define a convenience function to calculate the convolutional layer. This
# function initializes the convolutional layer weights and performs
# corresponding dimensionality elevations and reductions on the input and
# output
def comp_conv2d(conv2d, X):
# Here (1, 1) indicates that the batch size and the number of channels
# are both 1
X = X.reshape((1, 1) + X.shape)
Y = conv2d(X)
# Exclude the first two dimensions that do not interest us: examples and
# channels
return Y.reshape(Y.shape[2:])
# Note that here 1 row or column is padded on either side, so a total of 2
# rows or columns are added
conv2d = nn.Conv2d(1, 1, kernel_size=3, padding=1)
X = torch.rand(size=(8, 8))
comp_conv2d(conv2d, X).shape
# + [markdown] origin_pos=4
# When the height and width of the convolution kernel are different,
# we can make the output and input have the same height and width
# by [**setting different padding numbers for height and width.**]
#
# + origin_pos=6 tab=["pytorch"]
# Here, we use a convolution kernel with a height of 5 and a width of 3. The
# padding numbers on either side of the height and width are 2 and 1,
# respectively
conv2d = nn.Conv2d(1, 1, kernel_size=(5, 3), padding=(2, 1))
comp_conv2d(conv2d, X).shape
# + [markdown] origin_pos=8
# ## Stride
#
# When computing the cross-correlation,
# we start with the convolution window
# at the top-left corner of the input tensor,
# and then slide it over all locations both down and to the right.
# In previous examples, we default to sliding one element at a time.
# However, sometimes, either for computational efficiency
# or because we wish to downsample,
# we move our window more than one element at a time,
# skipping the intermediate locations.
#
# We refer to the number of rows and columns traversed per slide as the *stride*.
# So far, we have used strides of 1, both for height and width.
# Sometimes, we may want to use a larger stride.
# :numref:`img_conv_stride` shows a two-dimensional cross-correlation operation
# with a stride of 3 vertically and 2 horizontally.
# The shaded portions are the output elements as well as the input and kernel tensor elements used for the output computation: $0\times0+0\times1+1\times2+2\times3=8$, $0\times0+6\times1+0\times2+0\times3=6$.
# We can see that when the second element of the first column is outputted,
# the convolution window slides down three rows.
# The convolution window slides two columns to the right
# when the second element of the first row is outputted.
# When the convolution window continues to slide two columns to the right on the input,
# there is no output because the input element cannot fill the window
# (unless we add another column of padding).
#
# 
# :label:`img_conv_stride`
#
# In general, when the stride for the height is $s_h$
# and the stride for the width is $s_w$, the output shape is
#
# $$\lfloor(n_h-k_h+p_h+s_h)/s_h\rfloor \times \lfloor(n_w-k_w+p_w+s_w)/s_w\rfloor.$$
#
# If we set $p_h=k_h-1$ and $p_w=k_w-1$,
# then the output shape will be simplified to
# $\lfloor(n_h+s_h-1)/s_h\rfloor \times \lfloor(n_w+s_w-1)/s_w\rfloor$.
# Going a step further, if the input height and width
# are divisible by the strides on the height and width,
# then the output shape will be $(n_h/s_h) \times (n_w/s_w)$.
#
# Below, we [**set the strides on both the height and width to 2**],
# thus halving the input height and width.
#
# + origin_pos=10 tab=["pytorch"]
conv2d = nn.Conv2d(1, 1, kernel_size=3, padding=1, stride=2)
comp_conv2d(conv2d, X).shape
# + [markdown] origin_pos=12
# Next, we will look at (**a slightly more complicated example**).
#
# + origin_pos=14 tab=["pytorch"]
conv2d = nn.Conv2d(1, 1, kernel_size=(3, 5), padding=(0, 1), stride=(3, 4))
comp_conv2d(conv2d, X).shape
# + [markdown] origin_pos=16
# For the sake of brevity, when the padding number
# on both sides of the input height and width are $p_h$ and $p_w$ respectively, we call the padding $(p_h, p_w)$.
# Specifically, when $p_h = p_w = p$, the padding is $p$.
# When the strides on the height and width are $s_h$ and $s_w$, respectively,
# we call the stride $(s_h, s_w)$.
# Specifically, when $s_h = s_w = s$, the stride is $s$.
# By default, the padding is 0 and the stride is 1.
# In practice, we rarely use inhomogeneous strides or padding,
# i.e., we usually have $p_h = p_w$ and $s_h = s_w$.
#
# ## Summary
#
# * Padding can increase the height and width of the output. This is often used to give the output the same height and width as the input.
# * The stride can reduce the resolution of the output, for example reducing the height and width of the output to only $1/n$ of the height and width of the input ($n$ is an integer greater than $1$).
# * Padding and stride can be used to adjust the dimensionality of the data effectively.
#
# ## Exercises
#
# 1. For the last example in this section, use mathematics to calculate the output shape to see if it is consistent with the experimental result.
# 1. Try other padding and stride combinations on the experiments in this section.
# 1. For audio signals, what does a stride of 2 correspond to?
# 1. What are the computational benefits of a stride larger than 1?
#
# + [markdown] origin_pos=18 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/68)
#
| scripts/d21-en/pytorch/chapter_convolutional-neural-networks/padding-and-strides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Machine Translation and the Dataset
# :label:`sec_machine_translation`
#
# We have used RNNs to design language models,
# which are key to natural language processing.
# Another flagship benchmark is *machine translation*,
# a central problem domain for *sequence transduction* models
# that transform input sequences into output sequences.
# Playing a crucial role in various modern AI applications,
# sequence transduction models will form the focus of the remainder of this chapter
# and :numref:`chap_attention`.
# To this end,
# this section introduces the machine translation problem
# and its dataset that will be used later.
#
#
# *Machine translation* refers to the
# automatic translation of a sequence
# from one language to another.
# In fact, this field
# may date back to 1940s
# soon after digital computers were invented,
# especially by considering the use of computers
# for cracking language codes in World War II.
# For decades,
# statistical approaches
# had been dominant in this field :cite:`Brown.Cocke.Della-Pietra.ea.1988,Brown.Cocke.Della-Pietra.ea.1990`
# before the rise
# of
# end-to-end learning using
# neural networks.
# The latter
# is often called
# *neural machine translation*
# to distinguish itself from
# *statistical machine translation*
# that involves statistical analysis
# in components such as
# the translation model and the language model.
#
#
# Emphasizing end-to-end learning,
# this book will focus on neural machine translation methods.
# Different from our language model problem
# in :numref:`sec_language_model`
# whose corpus is in one single language,
# machine translation datasets
# are composed of pairs of text sequences
# that are in
# the source language and the target language, respectively.
# Thus,
# instead of reusing the preprocessing routine
# for language modeling,
# we need a different way to preprocess
# machine translation datasets.
# In the following,
# we show how to
# load the preprocessed data
# into minibatches for training.
#
# + origin_pos=2 tab=["pytorch"]
import os
import torch
from d2l import torch as d2l
# + [markdown] origin_pos=4
# ## Downloading and Preprocessing the Dataset
#
# To begin with,
# we download an English-French dataset
# that consists of [bilingual sentence pairs from the Tatoeba Project](http://www.manythings.org/anki/).
# Each line in the dataset
# is a tab-delimited pair
# of an English text sequence
# and the translated French text sequence.
# Note that each text sequence
# can be just one sentence or a paragraph of multiple sentences.
# In this machine translation problem
# where English is translated into French,
# English is the *source language*
# and French is the *target language*.
#
# + origin_pos=5 tab=["pytorch"]
#@save
d2l.DATA_HUB['fra-eng'] = (d2l.DATA_URL + 'fra-eng.zip',
'94646ad1522d915e7b0f9296181140edcf86a4f5')
#@save
def read_data_nmt():
"""Load the English-French dataset."""
data_dir = d2l.download_extract('fra-eng')
with open(os.path.join(data_dir, 'fra.txt'), 'r') as f:
return f.read()
raw_text = read_data_nmt()
print(raw_text[:75])
# + [markdown] origin_pos=6
# After downloading the dataset,
# we proceed with several preprocessing steps
# for the raw text data.
# For instance,
# we replace non-breaking space with space,
# convert uppercase letters to lowercase ones,
# and insert space between words and punctuation marks.
#
# + origin_pos=7 tab=["pytorch"]
#@save
def preprocess_nmt(text):
"""Preprocess the English-French dataset."""
def no_space(char, prev_char):
return char in set(',.!?') and prev_char != ' '
# Replace non-breaking space with space, and convert uppercase letters to
# lowercase ones
text = text.replace('\u202f', ' ').replace('\xa0', ' ').lower()
# Insert space between words and punctuation marks
out = [
' ' + char if i > 0 and no_space(char, text[i - 1]) else char
for i, char in enumerate(text)]
return ''.join(out)
text = preprocess_nmt(raw_text)
print(text[:80])
# + [markdown] origin_pos=8
# ## Tokenization
#
# Different from character-level tokenization
# in :numref:`sec_language_model`,
# for machine translation
# we prefer word-level tokenization here
# (state-of-the-art models may use more advanced tokenization techniques).
# The following `tokenize_nmt` function
# tokenizes the the first `num_examples` text sequence pairs,
# where
# each token is either a word or a punctuation mark.
# This function returns
# two lists of token lists: `source` and `target`.
# Specifically,
# `source[i]` is a list of tokens from the
# $i^\mathrm{th}$ text sequence in the source language (English here) and `target[i]` is that in the target language (French here).
#
# + origin_pos=9 tab=["pytorch"]
#@save
def tokenize_nmt(text, num_examples=None):
"""Tokenize the English-French dataset."""
source, target = [], []
for i, line in enumerate(text.split('\n')):
if num_examples and i > num_examples:
break
parts = line.split('\t')
if len(parts) == 2:
source.append(parts[0].split(' '))
target.append(parts[1].split(' '))
return source, target
source, target = tokenize_nmt(text)
source[:6], target[:6]
# + [markdown] origin_pos=10
# Let us plot the histogram of the number of tokens per text sequence.
# In this simple English-French dataset,
# most of the text sequences have fewer than 20 tokens.
#
# + origin_pos=11 tab=["pytorch"]
d2l.set_figsize()
_, _, patches = d2l.plt.hist([[len(l)
for l in source], [len(l) for l in target]],
label=['source', 'target'])
for patch in patches[1].patches:
patch.set_hatch('/')
d2l.plt.legend(loc='upper right');
# + [markdown] origin_pos=12
# ## Vocabulary
#
# Since the machine translation dataset
# consists of pairs of languages,
# we can build two vocabularies for
# both the source language and
# the target language separately.
# With word-level tokenization,
# the vocabulary size will be significantly larger
# than that using character-level tokenization.
# To alleviate this,
# here we treat infrequent tokens
# that appear less than 2 times
# as the same unknown ("<unk>") token.
# Besides that,
# we specify additional special tokens
# such as for padding ("<pad>") sequences to the same length in minibatches,
# and for marking the beginning ("<bos>") or end ("<eos>") of sequences.
# Such special tokens are commonly used in
# natural language processing tasks.
#
# + origin_pos=13 tab=["pytorch"]
src_vocab = d2l.Vocab(source, min_freq=2,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
len(src_vocab)
# + [markdown] origin_pos=14
# ## Loading the Dataset
# :label:`subsec_mt_data_loading`
#
# Recall that in language modeling
# each sequence example,
# either a segment of one sentence
# or a span over multiple sentences,
# has a fixed length.
# This was specified by the `num_steps`
# (number of time steps or tokens) argument in :numref:`sec_language_model`.
# In machine translation, each example is
# a pair of source and target text sequences,
# where each text sequence may have different lengths.
#
# For computational efficiency,
# we can still process a minibatch of text sequences
# at one time by *truncation* and *padding*.
# Suppose that every sequence in the same minibatch
# should have the same length `num_steps`.
# If a text sequence has fewer than `num_steps` tokens,
# we will keep appending the special "<pad>" token
# to its end until its length reaches `num_steps`.
# Otherwise,
# we will truncate the text sequence
# by only taking its first `num_steps` tokens
# and discarding the remaining.
# In this way,
# every text sequence
# will have the same length
# to be loaded in minibatches of the same shape.
#
# The following `truncate_pad` function
# truncates or pads text sequences as described before.
#
# + origin_pos=15 tab=["pytorch"]
#@save
def truncate_pad(line, num_steps, padding_token):
"""Truncate or pad sequences."""
if len(line) > num_steps:
return line[:num_steps] # Truncate
return line + [padding_token] * (num_steps - len(line)) # Pad
truncate_pad(src_vocab[source[0]], 10, src_vocab['<pad>'])
# + [markdown] origin_pos=16
# Now we define a function to transform
# text sequences into minibatches for training.
# We append the special “<eos>” token
# to the end of every sequence to indicate the
# end of the sequence.
# When a model is predicting
# by
# generating a sequence token after token,
# the generation
# of the “<eos>” token
# can suggest that
# the output sequence is complete.
# Besides,
# we also record the length
# of each text sequence excluding the padding tokens.
# This information will be needed by
# some models that
# we will cover later.
#
# + origin_pos=17 tab=["pytorch"]
#@save
def build_array_nmt(lines, vocab, num_steps):
"""Transform text sequences of machine translation into minibatches."""
lines = [vocab[l] for l in lines]
lines = [l + [vocab['<eos>']] for l in lines]
array = torch.tensor([
truncate_pad(l, num_steps, vocab['<pad>']) for l in lines])
valid_len = (array != vocab['<pad>']).type(torch.int32).sum(1)
return array, valid_len
# + [markdown] origin_pos=18
# ## Putting All Things Together
#
# Finally, we define the `load_data_nmt` function
# to return the data iterator, together with
# the vocabularies for both the source language and the target language.
#
# + origin_pos=19 tab=["pytorch"]
#@save
def load_data_nmt(batch_size, num_steps, num_examples=600):
"""Return the iterator and the vocabularies of the translation dataset."""
text = preprocess_nmt(read_data_nmt())
source, target = tokenize_nmt(text, num_examples)
src_vocab = d2l.Vocab(source, min_freq=2,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
tgt_vocab = d2l.Vocab(target, min_freq=2,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
src_array, src_valid_len = build_array_nmt(source, src_vocab, num_steps)
tgt_array, tgt_valid_len = build_array_nmt(target, tgt_vocab, num_steps)
data_arrays = (src_array, src_valid_len, tgt_array, tgt_valid_len)
data_iter = d2l.load_array(data_arrays, batch_size)
return data_iter, src_vocab, tgt_vocab
# + [markdown] origin_pos=20
# Let us read the first minibatch from the English-French dataset.
#
# + origin_pos=21 tab=["pytorch"]
train_iter, src_vocab, tgt_vocab = load_data_nmt(batch_size=2, num_steps=8)
for X, X_valid_len, Y, Y_valid_len in train_iter:
print('X:', X.type(torch.int32))
print('valid lengths for X:', X_valid_len)
print('Y:', Y.type(torch.int32))
print('valid lengths for Y:', Y_valid_len)
break
# + [markdown] origin_pos=22
# ## Summary
#
# * Machine translation refers to the automatic translation of a sequence from one language to another.
# * Using word-level tokenization, the vocabulary size will be significantly larger than that using character-level tokenization. To alleviate this, we can treat infrequent tokens as the same unknown token.
# * We can truncate and pad text sequences so that all of them will have the same length to be loaded in minibatches.
#
#
# ## Exercises
#
# 1. Try different values of the `num_examples` argument in the `load_data_nmt` function. How does this affect the vocabulary sizes of the source language and the target language?
# 1. Text in some languages such as Chinese and Japanese does not have word boundary indicators (e.g., space). Is word-level tokenization still a good idea for such cases? Why or why not?
#
# + [markdown] origin_pos=24 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1060)
#
| scripts/d21-en/pytorch/chapter_recurrent-modern/machine-translation-and-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Benchmark-monolingual-model:-English-and-German" data-toc-modified-id="Benchmark-monolingual-model:-English-and-German-1"><span class="toc-item-num">1 </span>Benchmark monolingual model: English and German</a></span></li><li><span><a href="#Experiment-1:-Multilingual-eng-eng" data-toc-modified-id="Experiment-1:-Multilingual-eng-eng-2"><span class="toc-item-num">2 </span>Experiment 1: Multilingual eng-eng</a></span></li><li><span><a href="#Experiment-2:-Multilingual-de-de" data-toc-modified-id="Experiment-2:-Multilingual-de-de-3"><span class="toc-item-num">3 </span>Experiment 2: Multilingual de-de</a></span></li><li><span><a href="#Experiment-3:-Multilingual-eng-de" data-toc-modified-id="Experiment-3:-Multilingual-eng-de-4"><span class="toc-item-num">4 </span>Experiment 3: Multilingual eng-de</a></span></li><li><span><a href="#Experiment-4:-Multilingual-de-eng" data-toc-modified-id="Experiment-4:-Multilingual-de-eng-5"><span class="toc-item-num">5 </span>Experiment 4: Multilingual de-eng</a></span></li><li><span><a href="#Experiment-5:-Multilingual-mult-eng" data-toc-modified-id="Experiment-5:-Multilingual-mult-eng-6"><span class="toc-item-num">6 </span>Experiment 5: Multilingual mult-eng</a></span></li><li><span><a href="#Experiment-6:-Multilingual-mult-de" data-toc-modified-id="Experiment-6:-Multilingual-mult-de-7"><span class="toc-item-num">7 </span>Experiment 6: Multilingual mult-de</a></span></li><li><span><a href="#Comparisson" data-toc-modified-id="Comparisson-8"><span class="toc-item-num">8 </span>Comparisson</a></span></li><li><span><a href="#New-experiment-results" data-toc-modified-id="New-experiment-results-9"><span class="toc-item-num">9 </span>New experiment results</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Question-1:-Knowledge-transfer" data-toc-modified-id="Question-1:-Knowledge-transfer-9.0.1"><span class="toc-item-num">9.0.1 </span>Question 1: Knowledge transfer</a></span></li><li><span><a href="#Question-2" data-toc-modified-id="Question-2-9.0.2"><span class="toc-item-num">9.0.2 </span>Question 2</a></span></li></ul></li></ul></li></ul></div>
# -
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import pickle
# # Benchmark monolingual model: English and German
infile = open('./Results/benchmark-eng/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_b1 = pickle.load(infile)
infile.close()
infile = open('./Results/benchmark-eng/task_test_f1_per_epoch_0.pkl','rb')
test_f1_b1 = pickle.load(infile)
infile.close()
infile = open('./Results/benchmark-de/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_b2 = pickle.load(infile)
infile.close()
infile = open('./Results/benchmark-de/task_test_f1_per_epoch_0.pkl','rb')
test_f1_b2 = pickle.load(infile)
infile.close()
# +
plt.figure(figsize=(12,5))
# plot for training accuracy
plt.subplot(1, 2, 1)
plt.plot(train_acc_b1, linestyle='--', marker='o', label = 'English: SemEval2010-8')
plt.plot(train_acc_b2[0:11], linestyle='--', marker='o', label = 'German: SemEval2010-8-de')
plt.xlabel("epoch")
plt.ylabel("training accuracy")
plt.ylim(0, 0.9)
plt.legend()
# plot for training accuracy
plt.subplot(1, 2, 2)
plt.plot(test_f1_b1, linestyle='--', marker='o', label = 'English: SemEval2010-8')
plt.plot(test_f1_b2[0:11], linestyle='--', marker='o', label = 'German: SemEval2010-8-de')
plt.xlabel("epoch")
plt.ylabel("test F1-score")
plt.ylim(0, 0.9)
plt.legend()
plt.show()
# -
# # Experiment 1: Multilingual eng-eng
infile = open('./Results/eng-eng/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_1 = pickle.load(infile)
infile.close()
infile = open('./Results/eng-eng/task_test_f1_per_epoch_0.pkl','rb')
test_f1_1 = pickle.load(infile)
infile.close()
plt.plot(train_acc_1, linestyle='--', marker='o', label = 'train accuracy')
plt.plot(test_f1_1, linestyle='--', marker='o', label = 'test f1')
plt.legend()
plt.show()
# # Experiment 2: Multilingual de-de
infile = open('./Results/de-de/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_2 = pickle.load(infile)
infile.close()
infile = open('./Results/de-de/task_test_f1_per_epoch_0.pkl','rb')
test_f1_2 = pickle.load(infile)
infile.close()
plt.plot(train_acc_2, linestyle='--', marker='o', label = 'train accuracy')
plt.plot(test_f1_2, linestyle='--', marker='o', label = 'test f1')
plt.legend()
plt.show()
# # Experiment 3: Multilingual eng-de
infile = open('./Results/eng-de/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_3 = pickle.load(infile)
infile.close()
infile = open('./Results/eng-de/task_test_f1_per_epoch_0.pkl','rb')
test_f1_3 = pickle.load(infile)
infile.close()
plt.plot(train_acc_3, linestyle='--', marker='o', label = 'train accuracy')
plt.plot(test_f1_3, linestyle='--', marker='o', label = 'test f1')
plt.legend()
plt.show()
# # Experiment 4: Multilingual de-eng
infile = open('./Results/de-eng/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_4 = pickle.load(infile)
infile.close()
infile = open('./Results/de-eng/task_test_f1_per_epoch_0.pkl','rb')
test_f1_4 = pickle.load(infile)
infile.close()
plt.plot(train_acc_4, linestyle='--', marker='o', label = 'train accuracy')
plt.plot(test_f1_4, linestyle='--', marker='o', label = 'test f1')
plt.legend()
plt.show()
# # Experiment 5: Multilingual mult-eng
infile = open('./Results/mult_l-eng/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_5 = pickle.load(infile)
infile.close()
infile = open('./Results/mult_l-eng/task_test_f1_per_epoch_0.pkl','rb')
test_f1_5 = pickle.load(infile)
infile.close()
plt.plot(train_acc_5, linestyle='--', marker='o', label = 'train accuracy')
plt.plot(test_f1_5, linestyle='--', marker='o', label = 'test f1')
plt.legend()
plt.show()
# # Experiment 6: Multilingual mult-de
infile = open('./Results/mult_l-de/task_train_accuracy_per_epoch_0.pkl','rb')
train_acc_6 = pickle.load(infile)
infile.close()
infile = open('./Results/mult_l-de/task_test_f1_per_epoch_0.pkl','rb')
test_f1_6 = pickle.load(infile)
infile.close()
plt.plot(train_acc_6, linestyle='--', marker='o', label = 'train accuracy')
plt.plot(test_f1_6, linestyle='--', marker='o', label = 'test f1')
plt.legend()
plt.show()
# +
plt.plot(train_acc_1[0:11], linestyle='--', marker='o', label = 'English: SemEval2010-8')
plt.plot(train_acc_2, linestyle='--', marker='o', label = 'German: SemEval2010-8-de')
plt.plot(train_acc_5, linestyle='--', marker='o', label = 'Multilingual')
plt.legend()
plt.show()
# -
# # Comparisson
test_results = {"monolingual eng": test_f1_b1[-1],
"monolingual de": test_f1_b2[-1],
"1. multilingual eng-eng": test_f1_1[-1],
"3. multilingual eng-de": test_f1_3[-1],
"2. multilingual de-de": test_f1_2[-1],
"4. multilingual de-eng": test_f1_4[-1],
"5. multilingual mult-eng": test_f1_5[-1],
"6. multilingual mult-de": test_f1_6[-1]}
test_results
plt.barh(*zip(*test_results.items()), color = ['brown', 'brown', 'royalblue', 'royalblue', 'royalblue', 'royalblue', 'royalblue', 'royalblue'])
plt.xticks(rotation=90)
plt.show()
test_eng_results = {"monolingual eng": test_f1_b1[-1],
"1. multilingual eng-eng": test_f1_1[-1],
"3. multilingual eng-de": test_f1_3[-1]}
test_de_results = {"monolingual de": test_f1_b2[-1],
"2. multilingual de-de": test_f1_2[-1],
"4. multilingual de-eng": test_f1_4[-1]}
plt.barh(*zip(*test_eng_results.items()), color = ['brown', 'royalblue', 'royalblue'])
plt.xticks(rotation=90)
plt.show()
plt.barh(*zip(*test_de_results.items()), color = ['brown', 'royalblue', 'royalblue'])
plt.xticks(rotation=90)
plt.show()
test_results_eng = {"monolingual eng": test_f1_b1[-1],
"1. multilingual eng-eng": test_f1_1[-1],
"4. multilingual de-eng": test_f1_4[-1],
"5. multilingual mult-eng": test_f1_5[-1]
}
plt.bar(*zip(*test_results_eng.items()), color = ['brown', 'royalblue', 'royalblue', 'royalblue'])
plt.xticks(rotation=90)
plt.show()
test_results_de = {"monolingual de": test_f1_b2[-1],
"2. multilingual de-de": test_f1_2[-1],
"3. multilingual eng-de": test_f1_3[-1],
"6. multilingual mult-de": test_f1_6[-1]}
plt.bar(*zip(*test_results_de.items()), color = ['brown', 'royalblue', 'royalblue', 'royalblue'])
plt.xticks(rotation=90)
plt.show()
test_results_2 = {"monolingual eng": test_f1_b1[-1],
"1. multilingual eng-eng": test_f1_1[-1],
"monolingual de": test_f1_b2[-1],
"2. multilingual de-de": test_f1_2[-1]}
plt.bar(*zip(*test_results_2.items()), color = ['brown', 'royalblue', 'brown', 'royalblue'])
plt.xticks(rotation=90)
plt.show()
test_results_22 = {"1. multilingual eng-eng": test_f1_1[-1],
"5. multilingual mult-eng": test_f1_5[-1],
"2. multilingual de-de": test_f1_2[-1],
"6. multilingual mult-de": test_f1_6[-1]}
plt.barh(*zip(*test_results_22.items()), color = ['coral', 'coral', 'royalblue', 'royalblue'], align='center')
plt.xticks(rotation=0)
plt.show()
# # New experiment results
# + active=""
# ############################################################################################
# Model: BERT-base-multilingual
#
# TRAIN DATASET ('./data/Semeval_DE_NEW/TRAIN_FILE_DE_FINAL.TXT')
# F1 on train: 0.5750279955207166
# Accuracy on train: 0.6947828389830508
# Precision on train: 0.5831913685406019
# Recall on train: 0.5670900055218111
#
# TEST SMALL DE ('./data/Semeval_DE_NEW/TEST_SMALL_DE.TXT')
# F1 on test: 0.5090909090909091
# Accuracy on test: 0.5321691176470589
# Precision on test: 0.5714285714285714
# Recall on test: 0.45901639344262296
#
# TEST DE ('./data/Semeval_DE_NEW/TEST_FILE_DE_FINAL.TXT')
# F1 on test: 0.49531116794543906
# Accuracy on test: 0.616010581061693
# Precision on test: 0.501727115716753
# Recall on test: 0.4890572390572391
#
# TEST EN ('./data/Semeval/TEST_FILE_FULL.TXT')
# F1 on test: 0.3618657098923629
# Accuracy on test: 0.5685019045709703
# Precision on test: 0.5276532137518685
# Recall on test: 0.2753510140405616
#
# ############################################################################################
# Model: BERT-base-multilingual
#
# TRAIN DATASET ('./data/Semeval/TRAIN_FILE.TXT')
# F1 on train: 0.8047542735042735
# Accuracy on train: 0.875375
# Precision on train: 0.7996284501061571
# Recall on train: 0.8099462365591398
#
# TEST SMALL DE ('./data/Semeval_DE_NEW/TEST_SMALL_DE.TXT')
# F1 on test: 0.3364485981308411
# Accuracy on test: 0.3805147058823529
# Precision on test: 0.391304347826087
# Recall on test: 0.29508196721311475
#
# TEST DE ('./data/Semeval_DE_NEW/TEST_FILE_DE_FINAL.TXT')
# F1 on test: 0.3778529163144548
# Accuracy on test: 0.503474713055954
# Precision on test: 0.3794567062818336
# Recall on test: 0.37626262626262624
#
# TEST EN ('./data/Semeval/TEST_FILE_FULL.TXT')
# F1 on test: 0.7457892675283981
# Accuracy on test: 0.7988923416198878
# Precision on test: 0.7490165224232888
# Recall on test: 0.7425897035881436
#
# ############################################################################################
# Model: BERT-base-multilingual
#
# TRAIN DATASET ('./data/Semeval_ENDE_large/TRAIN_FILE.TXT')
# F1 on train: 0.7189965607930406
# Accuracy on train: 0.8090920781893004
# Precision on train: 0.719045049905584
# Recall on train: 0.7189480782198246
#
# TEST SMALL DE ('./data/Semeval_DE_NEW/TEST_SMALL_DE.TXT')
# F1 on test: 0.5137614678899083
# Accuracy on test: 0.546875
# Precision on test: 0.5833333333333334
# Recall on test: 0.45901639344262296
#
# TEST DE ('./data/Semeval_DE_NEW/TEST_FILE_DE_FINAL.TXT')
# F1 on test: 0.48817567567567566
# Accuracy on test: 0.6312544835007174
# Precision on test: 0.48983050847457626
# Recall on test: 0.48653198653198654
#
# TEST EN ('./data/Semeval/TEST_FILE_FULL.TXT')
# F1 on test: 0.7442405310425614
# Accuracy on test: 0.7981655974338413
# Precision on test: 0.745113369820172
# Recall on test: 0.7433697347893916
# +
# benchmarks
b_en = 0.7670
b_de = test_f1_b2[-1]
# results of the experiments (F1_score)
de_train = 0.5750279955207166
de_de = 0.51531116794543906
de_de_s = 0.5090909090909091
de_en = 0.3618657098923629
en_train = 0.8047542735042735
en_de = 0.3778529163144548
en_de_s = 0.3364485981308411
en_en = 0.7457892675283981
mult_train = 0.7189965607930406
mult_de = 0.49817567567567566
mult_de_s = 0.5137614678899083
mult_en = 0.7442405310425614
# -
test_results = {"monolingual eng": b_en,
"monolingual de": b_de,
"1. multilingual eng-eng": en_en,
"2. multilingual eng-de": en_de,
"3. multilingual eng-de-small": en_de_s,
"4. multilingual de-eng": de_en,
"5. multilingual de-de": de_de,
"6. multilingual de-de-small": de_de_s,
"7. multilingual mult-eng": mult_en,
"8. multilingual mult-de": mult_de,
"9. multilingual mult-de-small": mult_de_s,}
test_results = {"monolingual eng": b_en,
"monolingual de": b_de,
"train: en, test: en": en_en,
"train: en, test: de": en_de,
"train: en, test: de100": en_de_s,
"train: de, test: eng": de_en,
"train: de, test: de": de_de,
"train: de, test: de100": de_de_s,
"train: mult, test: eng": mult_en,
"train: mult, test: de": mult_de,
"train: mult, test: de100": mult_de_s,}
test_results
plt.barh(*zip(*test_results.items()), color = ['coral']*2+['royalblue']*9, align='center')
plt.xticks(rotation=0)
plt.xlabel("F1-score")
plt.show()
test_results_MTB = {"BERT fine-tuning": en_en,
"BERT MTB + fine-tuning": 0.67}
# +
plt.bar(*zip(*test_results_MTB.items()), color = ['coral', 'royalblue'])
plt.xticks(rotation=0)
test_results_MTB = {"BERT fine-tuning": test_f1_b1[-1],
"BERT MTB + fine-tuning": 0.67}
plt.xlabel("Approach")
plt.ylabel("F1-score on test sample")
for i, v in enumerate(test_results_MTB.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
plt.show()
# -
# ### Question 1: Knowledge transfer
# +
q2_en_pretrain = {"en": en_en,
"de": en_de,
"de100": en_de_s
}
q2_de_pretrain = {"de": de_de,
"de100": de_de_s,
"en": de_en
}
plt.figure(figsize=(12,5))
# plot for English
plt.subplot(1, 2, 1)
plt.bar(*zip(*q2_en_pretrain.items()), color = ['coral']*0+['royalblue']*1+['seagreen']*2, align='center')
plt.xticks(rotation=0)
plt.xlabel("Test Dataset")
plt.ylabel("F1-score on Test Dataset")
plt.title('Train dataset: SemEval2010-8')
for i, v in enumerate(q2_en_pretrain.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
# plot for German
plt.subplot(1, 2, 2)
plt.bar(*zip(*q2_de_pretrain.items()), color = ['coral']*0+['royalblue']*2+['seagreen']*1, align='center')
plt.xticks(rotation=0)
plt.xlabel("Test Dataset")
plt.title('Train dataset: SemEval2010-8-de')
plt.ylim(0, 0.78)
for i, v in enumerate(q2_de_pretrain.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
plt.show()
# +
# Multilingual results
q2_de_pretrain = {"de": mult_de,
"de100": mult_de_s,
"en": mult_en
}
# plot for multilingual
plt.bar(*zip(*q2_de_pretrain.items()), color = ['coral']*0+['royalblue']*3+['seagreen']*0, align='center')
plt.xticks(rotation=0)
plt.xlabel("Test Dataset")
plt.ylabel("F1-score")
plt.title('Train dataset: Multilingual')
plt.ylim(0, 0.78)
for i, v in enumerate(q2_de_pretrain.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
plt.show()
# -
# ### Question 2
# +
q2_en_pretrain = {"Monolingual": b_en,
"Multilingual": en_en
}
q2_de_pretrain = {"Monolingual": b_de,
"Multilingual": de_de
}
plt.figure(figsize=(12,5))
# plot for English
plt.subplot(1, 2, 1)
plt.bar(*zip(*q2_en_pretrain.items()), color = ['coral']*1+['royalblue']*1+['seagreen']*0, align='center')
plt.xticks(rotation=0)
plt.xlabel("Model")
plt.ylabel("F1-score on Test Dataset")
plt.title('Train: SemEval2010-8 Train \n Test: SemEval2010-8 Test')
plt.ylim(0, 0.8)
for i, v in enumerate(q2_en_pretrain.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
# plot for German
plt.subplot(1, 2, 2)
plt.bar(*zip(*q2_de_pretrain.items()), color = ['coral']*1+['royalblue']*2+['seagreen']*0, align='center')
plt.xticks(rotation=0)
plt.xlabel("Model")
plt.title('Train: SemEval2010-8-de Train \n Test: SemEval2010-8-de Test')
plt.ylim(0, 0.8)
for i, v in enumerate(q2_de_pretrain.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
plt.show()
# +
q2_2_en = {"train: en": en_en,
"train: mult": mult_en}
q2_2_de = {"train: de": de_de,
"train: mult": mult_de}
# +
plt.figure(figsize=(12,5))
# plot for English
plt.subplot(1, 2, 1)
plt.bar(*zip(*q2_2_en.items()), color = ['coral']*1+['royalblue']*1+['seagreen']*0, align='center')
plt.xticks(rotation=0)
plt.xlabel("Train Dataset")
plt.ylabel("F1-score on Test Dataset")
plt.title('Test: SemEval2010-8 Test')
plt.ylim(0, 0.8)
for i, v in enumerate(q2_2_en.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
# plot for German
plt.subplot(1, 2, 2)
plt.bar(*zip(*q2_2_de.items()), color = ['coral']*1+['royalblue']*2+['seagreen']*0, align='center')
plt.xticks(rotation=0)
plt.xlabel("Train Dataset")
plt.title('Test: SemEval2010-8-de Test')
plt.ylim(0, 0.8)
for i, v in enumerate(q2_2_de.values()):
plt.text(i-.25, v - 0.05 , str(round(v*100,2))+'%', color='white', fontweight='bold')
plt.show()
# -
plt.barh(*zip(*q2_2_en.items()), color = ['coral']*0+['royalblue']*2, align='center')
plt.xticks(rotation=0)
#plt.ylabel("Experiment setting")
plt.xlabel("F1-score on test sample")
plt.xlim(0, 0.8)
plt.show()
plt.barh(*zip(*q2_2_de.items()), color = ['coral']*0+['royalblue']*2, align='center')
plt.xticks(rotation=0)
#plt.ylabel("Experiment setting")
plt.xlabel("F1-score on test sample")
plt.xlim(0, 0.8)
plt.show()
| Jupyter Notebooks/Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
file = "D:\\My Personal Documents\\Learnings\\Data Science\\Data Sets\\Twiter Sentiment Analysis\\train_E6oV3lV.csv"
test = 'D:\\My Personal Documents\\Learnings\\Data Science\\Data Sets\\Twiter Sentiment Analysis\\test_tweets_anuFYb8.csv'
tweets = pd.read_csv(file)
test_tweets=pd.read_csv(test)
tweets1=tweets[tweets.label==1]
tweets=tweets.append(tweets1)
tweets=tweets.append(tweets1)
tweets=tweets.append(tweets1)
label=tweets.label
tweets[tweets.label==1].count()
tweets=tweets.append(test_tweets)
tweets.groupby(tweets.label).tweet.count()
cleaned_tweet = tweets.tweet.str.replace("[^a-zA-Z]", " ")
cleaned_tweet=cleaned_tweet.apply(lambda x :" ".join(w for w in x.split() if w != 'user' ))
from nltk.stem.porter import PorterStemmer
#from nltk.stem.snowball import SnowballStemmer
stem=PorterStemmer()
cleaned_tweet = cleaned_tweet.apply(lambda x : " ".join( stem.stem(w) for w in x.split()))
cleaned_tweet = cleaned_tweet.apply(lambda x : " ".join( w for w in x.split()))
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer(max_features=9300,stop_words='english')
bow=tfidf.fit_transform(cleaned_tweet)
train_bow = bow[:38688,:]
test_bow = bow[38688:,:]
from sklearn.model_selection import train_test_split
x_train,x_test,y_train, y_test = train_test_split(train_bow,label,test_size=0.2,random_state=3, stratify=label)
print(x_train.shape)
print(y_train.shape)
print(x_test.shape)
print(y_test.shape)
x_train[0][0]
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.optimizers import SGD
from keras.datasets import mnist
from keras.callbacks import ModelCheckpoint
# +
#n_class=2
#y_train = keras.utils.to_categorical(y_train,n_class)
#y_test = keras.utils.to_categorical(y_test,n_class)
# -
import os
output_dir = 'model_tweet_output1/dense'
modelcheckpoint = ModelCheckpoint(filepath=output_dir+"/weights.{epoch:02d}.hdf5")
if not os.path.exists(output_dir):
os.makedirs(output_dir)
model = Sequential()
model.add(Dense(3000,activation='relu', input_shape=(9300,)))
model.add(Dropout(0.2))
#model.add(Dense(2000,activation='relu', input_shape=(9300,)))
#model.add(Dropout(0.3))
model.add(Dense(1,activation='sigmoid'))
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
model.fit(x_train,y_train,epochs=5,verbose=1, validation_data=(x_test,y_test),callbacks=[modelcheckpoint])
model.load_weights(output_dir+'/weights.03.hdf5')
y_hat=model.predict(test_bow)
y_hat = pd.DataFrame(y_hat, columns=['Prob']).Prob.apply(lambda x : 1 if x >=0.6 else 0)
y_hat.to_csv('D:\\My Personal Documents\\Learnings\\Data Science\\Data Sets\\Twiter Sentiment Analysis\\submit_TFIDF_MLP.csv')
| notebooks/NN - Twitten Sentiment Analysis Challenge - MLP - TFIDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lista Prodotti DOP - Unione Europea
#
# La lista dei prodotti DOP nell'Unione Europea è aggiornata dal Dipartimento per l'Agricoltura e lo Sviluppo Rurale.
# http://ec.europa.eu/agriculture/quality/door/list.html?locale=it
#
# http://nbviewer.jupyter.org/gist/jtbaker/57a37a14b90feeab7c67a687c398142c?flush_cache=true
#
# Data Visualization Folium
#
# http://apps.socib.es/Leaflet.TimeDimension/examples/example12.html
#
# https://github.com/python-visualization/folium/blob/master/examples/Colormaps.ipynb
#
import pandas as pd
import geopandas as gpd
import numpy as np
import folium #https://nbviewer.jupyter.org/github/python-visualization/folium/blob/master/examples
import json
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# %pylab inline
url = 'http://ec.europa.eu/agriculture/quality/door/Denominations.xls?locale'
df = pd.read_excel(url, skiprows = 3, header=0)
df.head(2)
print (df.columns.tolist())
def desc_ita(x):
if x[' Type ']=='PDO':
return 'DOP'
elif x[' Type ']=='PGI':
return 'IGP'
elif x[' Type ']=='TSG':
return 'STG'
df['Type IT'] = df.apply(lambda x: desc_ita(x), axis=1)
df.head(2)
# Filtro solo i registrati e aggrego per Paese
df=df[(df[' Status ']=='Registered')]
df_agg = df.groupby([' ISO '],as_index=False).count()
df_eu = df_agg[[' ISO ',' Dossier Number ']].sort_values([' Dossier Number '], ascending=False)
df_eu.head(2)
# # Data Visualization
# Utilizzo GeoPandas
gdf = gpd.read_file("euro2.geojson")
gdf1=gdf.merge(df_eu, left_on='wb_a2', right_on=' ISO ', how='left').fillna(0)
gdf1.head(2)
# +
# mappa
icon_url = 'http://www.ildatomancante.it/external/Parmigiano4.png'
centroid=gdf1.geometry.centroid
m=folium.Map(location=[centroid.y.mean(), centroid.x.mean()], zoom_start=4)
# layer 1
folium.GeoJson(gdf1[['geometry',' ISO ',' Dossier Number ','sovereignt']],
name="Produttori BIO in Europa",
style_function=lambda x: {
"weight":2,
'color':'black',
'fillColor':'green' if x['properties'][' Dossier Number '] else 'white',
'fillOpacity':0.5},
highlight_function=lambda x: {'weight':3, 'color':'black'},
smooth_factor=2.0,
).add_to(m)
max_size = max(gdf1[' Dossier Number '])
# I can add marker one by one on the map
for i, v in gdf1.iterrows():
centroid=v.geometry.centroid
size = v[' Dossier Number ']/max_size * 100
icon = folium.features.CustomIcon(icon_url,icon_size=(size,size))
popup = '<strong>Paese:</strong> ' + v.sovereignt + '<br>' + '<strong>Prodotti: </strong>'+str(v[' Dossier Number '])
folium.Marker([centroid.y, centroid.x], icon=icon, popup=popup).add_to(m)
m.save('FoodPorn_Prodotti_DOP.html')
m
| Lista_Prodotti_DOP_EU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.5
# language: julia
# name: julia-1.6
# ---
# # Kmer Graph
DATE_TASK = "2022-03-06-ecoli-phapecoctavirus-core-genome"
DIR = mkpath("$(homedir())/workspace/$DATE_TASK")
cd(DIR)
DATE, TASK = match(r"^(\d{4}-\d{2}-\d{2})-(.*)$", DATE_TASK).captures
# +
import Pkg
Pkg.update()
pkgs = [
"JSON",
"HTTP",
"Dates",
"uCSV",
"DelimitedFiles",
"DataFrames",
"ProgressMeter",
"BioSequences",
"FASTX",
"Distances",
"StatsPlots",
"StatsBase",
"Statistics",
"MultivariateStats",
"Random",
"Primes",
"SparseArrays",
"SHA",
"GenomicAnnotations",
"Combinatorics",
"OrderedCollections",
"Downloads",
"Clustering",
"Revise",
"Mmap",
"Graphs",
"MetaGraphs",
"FileIO"
]
for pkg in pkgs
try
eval(Meta.parse("import $pkg"))
catch
Pkg.add(pkg)
eval(Meta.parse("import $pkg"))
end
end
# works but can't update locally, need to push and restart kernel to activate changes
# "https://github.com/cjprybol/Mycelia.git#master",
# didn't work
# "$(homedir())/git/Mycelia#master",
pkg_path = "$(homedir())/git/Mycelia"
try
eval(Meta.parse("import $(basename(pkg_path))"))
catch
# Pkg.add(url=pkg)
Pkg.develop(path=pkg_path)
# pkg = replace(basename(pkg), ".git#master" => "")
# pkg = replace(basename(pkg), "#master" => "")
eval(Meta.parse("import $(basename(pkg_path))"))
end
# +
function save_graph(graph::Graphs.AbstractGraph, outfile::String)
if !occursin(r"\.jld2$", outfile)
outfile *= ".jld2"
end
FileIO.save(outfile, Dict("graph" => graph))
return outfile
end
function load_graph(file::String)
return FileIO.load(file)["graph"]
end
# -
# https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?&id=$(tax_id)
# https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?lvl=0&id=2733124
root_tax_id = 2733124
child_tax_ids = vcat(Mycelia.taxonomic_id_to_children(root_tax_id), root_tax_id)
# child_tax_ids = vcat(child_tax_ids, root_tax_id)
# +
# TODO
# here is where we should apply a filter where host == Escherichia
# need to load host information into neo4j taxonomy
# +
# # refseq_metadata = Mycelia.load_refseq_metadata()
# ncbi_metadata = Mycelia.load_genbank_metadata()
# +
# show(ncbi_metadata[1:1, :], allcols=true)
# +
# tax_id_filter = map(taxid -> taxid in child_tax_ids, ncbi_metadata[!, "taxid"])
# is_right_host = map(x -> occursin(r"Escherichia"i, x), ncbi_metadata[!, "organism_name"])
# not_excluded = ncbi_metadata[!, "excluded_from_refseq"] .== ""
# is_full = ncbi_metadata[!, "genome_rep"] .== "Full"
# # assembly_levels = ["Complete Genome"]
# assembly_levels = ["Complete Genome", "Chromosome"]
# # assembly_levels = ["Complete Genome", "Chromosome", "Scaffold"]
# # assembly_levels = ["Complete Genome", "Chromosome", "Scaffold", "Contig"]
# assembly_level_filter = map(x -> x in assembly_levels, ncbi_metadata[!, "assembly_level"])
# full_filter = is_full .& not_excluded .& assembly_level_filter .& tax_id_filter .& is_right_host
# count(full_filter)
# +
# TODO
# here is another place we could enforce host == escherichia
# we'll use a manual filter as a temporary solution
# +
# ncbi_metadata_of_interest = ncbi_metadata[full_filter, :]
# +
# https://www.ncbi.nlm.nih.gov/sviewer/viewer.cgi?db=nuccore&report=genbank&id=GCA_021354775
# +
# for col in names(ncbi_metadata_of_interest)
# @show col, ncbi_metadata_of_interest[1, col]
# end
# +
# GCA_002956955.1
# +
# # can I also get genbank record?????
# # for extension in ["genomic.fna.gz", "protein.faa.gz"]
# for extension in ["genomic.fna.gz", "protein.faa.gz", "genomic.gbff.gz"]
# outdir = mkpath(joinpath(DIR, extension))
# ProgressMeter.@showprogress for row in DataFrames.eachrow(ncbi_metadata_of_interest)
# url = Mycelia.ncbi_ftp_path_to_url(row["ftp_path"], extension)
# outfile = joinpath(outdir, basename(url))
# if !isfile(outfile)
# try
# Downloads.download(url, outfile)
# catch e
# # @show e
# showerror(stdout, e)
# # @assert extension == "protein.faa.gz"
# # here is where we should call prodigal to fill in protein annotations if we don't otherwise see them
# end
# end
# end
# end
# -
extension = "genomic.fna.gz"
outdir = mkpath(joinpath(DIR, extension))
fastx_files = filter(x -> !occursin(".ipynb_checkpoints", x), readdir(outdir, join=true))
kmer_size = Mycelia.assess_kmer_saturation(fastx_files)
graph = Mycelia.fastx_to_kmer_graph(BioSequences.BigDNAMer{kmer_size}, fastx_files)
graph_outfile = "$DIR/root-tax-id_$(root_tax_id).k_$(kmer_size).genome-graph"
# Mycelia.save_graph(graph, graph_outfile)
save_graph(graph, graph_outfile * ".jld2")
Mycelia.graph_to_gfa(graph, graph_outfile * ".gfa")
function graph_to_neo4j_tables(graph, base_directory)
base_directory = graph_outfile * ".neo4j-tables"
mkpath(base_directory)
node_table = joinpath(base_directory, "nodes.tsv")
edge_table = joinpath(base_directory, "edges.tsv")
# +
# write out node tsv
# -
open(node_table, "w") do io
println(io, join(["kmer", "count"], '\t'))
for vertex in Graphs.vertices(graph)
if haskey(graph.vprops[vertex], :kmer)
sequence = graph.vprops[vertex][:kmer]
else
sequence = graph.vprops[vertex][:sequence]
end
depth = graph.vprops[vertex][:count]
fields = ["$sequence", "$(depth)"]
line = join(fields, '\t')
println(io, line)
end
end
# +
# write out edge tsv
# -
open(edge_table, "w") do io
println(io, join(["source", "source_orientation", "destination", "destination_orientation"], '\t'))
for edge in Graphs.edges(graph)
for o in graph.eprops[edge][:orientations]
link = [edge.src,
o.source_orientation,
edge.dst,
o.destination_orientation]
line = join(link, '\t')
println(io, line)
end
end
end
NEO4J_BIN_DIR = "/home/jupyter-cjprybol/software/neo4j-community-4.4.3/bin"
if !occursin(NEO4J_BIN_DIR, ENV["PATH"])
ENV["PATH"] = "$(NEO4J_BIN_DIR):" * ENV["PATH"]
end
DOMAIN = "e9d58dd0.databases.neo4j.io"
USERNAME="neo4j"
PASSWORD=readline(joinpath(homedir(), ".config", "neo4j", "public-phapecoctavirus.password.txt"));
ADDRESS="neo4j+s://$(DOMAIN):7687"
DATABASE = "neo4j"
cmd = "CREATE CONSTRAINT ON (k:Kmer) ASSERT k.sequence IS UNIQUE"
@time cypher(address = ADDRESS, username = USERNAME, password = PASSWORD, database = DATABASE, cmd = cmd)
# ```
# gcloud iam service-accounts keys create ~/.config/gcloud/url-signer-key.json --iam-account="928365250020-compute@developer.gserviceaccount.com"
# # gsutil -m signurl -d 24h -m GET "~/.config/gcloud/url-signer-key.json" "gs://neo4j-upload/public-phapecoctavirus/edges.tsv"
# gsutil signurl -d 24h ~/.config/gcloud/url-signer-key.json gs://neo4j-upload/public-phapecoctavirus/nodes.tsv
# gsutil signurl -d 24h ~/.config/gcloud/url-signer-key.json gs://neo4j-upload/public-phapecoctavirus/edges.tsv
# ```
# +
# println(read(`gsutil signurl -d 24h '~/.config/gcloud/url-signer-key.json' 'gs://neo4j-upload/public-phapecoctavirus/nodes.tsv'`))
# -
NODES_FILE = "https://storage.googleapis.com/neo4j-upload/public-phapecoctavirus/nodes.tsv?x-goog-signature=1af3780c722389bd6c32463e61759196ef2397d47a07ce0eab4cbc4f4d9fc08a9c977bc6084189fc0cb2367a33223510227274ce7d358f510c66d76583dd21d8e700e62798dc1ebe51114f59e50ef07d23db8c9662c848425dbe9edf97085cbc3c113c2b533bf87150712530c53a52276d6d04588b97456a3c8982dae75e2a72b84c5b10717b556ffe1f0dda275d33f43d295ee97381a89dccdd90b953a6dd20d7fc7a92ea2ee4f0a43c153e36df4e589a48dbe722b11f722ec5be0d3cfa2429b5300a59e320c2f13da6da128930a0baafcce406d3d1b227d6298d9ec31bc67826c75e2a36232dcfe6df79cc2fbbb636af5cc634412e94df6b3082d76064f175&x-goog-algorithm=GOOG4-RSA-SHA256&x-goog-credential=928365250020-compute%40developer.gserviceaccount.com%2F20220310%2Fus-east1%2Fstorage%2Fgoog4_request&x-goog-date=20220310T145905Z&x-goog-expires=86400&x-goog-signedheaders=host"
# EDGES_FILE = "https://storage.googleapis.com/neo4j-upload/public-phapecoctavirus/edges.tsv?x-goog-signature=38c14f7944a00742abcc6684adbf32b0457e5fde690d8db0d341a716cd0097410668351a56eb28dacfcf705dd20e10377887555e9ba5e64a5d6716338aa452638badc51409846d0b84b0a67e39b44c72164aff9aa0b377d4e1aecc8a64cc326cb1317b6c651c6c17a3c039d5bab3189130d9a5090788905ed13414b023c7d067631df246fb8676cdc49c07b5b38361f00a484b022e1d6ac212917abb1871ed9be0f566deb7cafad3bfd5c0a5e334aec29d0ae13bf3690bc737b96b8a49afb13dafa5a494c8fbb9b817ac080ae379ea7b40f67160a338e301a8a471e93698f17fb9cfc39e279b00d9a6a584fa44e0efc2ea8d20f7c3a6286151a450222d712cbf&x-goog-algorithm=GOOG4-RSA-SHA256&x-goog-credential=928365250020-compute%40developer.gserviceaccount.com%2F20220309%2Fus-east1%2Fstorage%2Fgoog4_request&x-goog-date=20220309T184444Z&x-goog-expires=86400&x-goog-signedheaders=host"
# cmd =
# """
# FIELDTERMINATOR '\t' LOAD CSV WITH HEADERS FROM '$(NODES_FILE)' AS row
# MERGE (k:Kmer {kmer: row.kmer})
# """
# cmd = rstrip(replace(cmd, '\n' => ' '))
cmd =
"""
CALL apoc.periodic.iterate(
"LOAD CSV WITH HEADERS FROM '$(NODES_FILE)' AS row FIELDTERMINATOR '\t' RETURN row",
"MERGE (k:Kmer {kmer: row.kmer})",
{batchSize:1000})
"""
cmd = rstrip(replace(cmd, '\n' => ' '))
cypher_cmd = Mycelia.cypher(address = ADDRESS, username = USERNAME, password = PASSWORD, database = DATABASE, cmd = cmd)
@time run(cypher_cmd)
# +
# write a batching script that will load the CSV, write temporary files to GCP, grab the signed url, upload with cypher, delete the temp file, and then repeat until complete
| _research/2022-03-06-ecoli-phapecoctavirus-core-genome-graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
# # AirBnb Feature Importance Project
# ## Motivation
#
# Traveling to a nice touristic cosmopolitan city is different to travel to a small city or to a not so turistic one. Thats clear in our minds and there is no doubt about it. But could we tell the most important characteristics of an AirBnb acomodation that differ between the most visited cities and the not so visited ones. <br>
#
# With that said, the main motivation of the project is translated into the following research questions:
#
# ### Research Questions
# 1. Which are the 10 most important characteristics to influence the overall review of an acomodation?
# 2. What are the general differences between a top rated acomodation and a almost top one?
# 3. Which are the main review aspects that influences the overall review score?
# 4. How would be the most likely description of a acomodation in a small city like Salem, Oregon based on the findings of this study?
#
# ## Importing libraries
# +
import pandas as pd
import numpy as np
import gzip
from io import BytesIO
import re
from decimal import Decimal
import seaborn as sns
import matplotlib.pyplot as plt
import dataframe_image as dfi
import nltk
nltk.download('vader_lexicon')
from nltk.sentiment.vader import SentimentIntensityAnalyzer
from tqdm import tqdm
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.metrics import r2_score, mean_squared_error
import lightgbm as lgb
from xgboost import XGBRegressor
# -
# ## Toolkit functions
def remove_nonnumeric_chars(s):
'''
INPUT:
s - numeric string with non-numeric charachters
OUTPUT:
np.float16 - numeric value before the dot
'''
try:
return np.float64(Decimal(re.sub(r'[^\d.]', '', s)))
except:
return np.nan
def create_dummy_df(df, cat_cols, dummy_na, drop_first=True):
'''
INPUT:
df - pandas dataframe with categorical variables you want to dummy
cat_cols - list of strings that are associated with names of the categorical columns
dummy_na - Bool holding whether you want to dummy NA vals of categorical columns or not
OUTPUT:
df - a new dataframe that has the following characteristics:
1. contains all columns that were not specified as categorical
2. removes all the original columns in cat_cols
3. dummy columns for each of the categorical columns in cat_cols
4. if dummy_na is True - it also contains dummy columns for the NaN values
5. Use a prefix of the column name with an underscore (_) for separating
'''
cat_df_out = pd.get_dummies(df[cat_cols], prefix=cat_cols, prefix_sep='_', dummy_na=dummy_na, drop_first=drop_first)
df_out = cat_df_out.join(df.drop(cat_cols, axis=1))
return df_out
def get_sequences(vector, split_val):
''''
INPUT:
vector - raw serie (pd.Series)
split_val - reference str to split the series
OUTPUT:
median, mean, max, min values of the sequences found in the serie
'''
sequences = ''.join([str(val) for val in vector]).split(split_val)
sequences_count = [len(val) for val in sequences]
sequences_cleansed = [val for val in sequences_count if val>0]
if len(sequences_cleansed) > 0:
return np.median(sequences_cleansed), np.mean(sequences_cleansed), np.max(sequences_cleansed), np.min(sequences_cleansed)
else:
return 0,0,0,0
def plot_importance(data, feature, title=None ,rows=10):
'''
INPUT:
data - dataframe based on which the plot will be done
feature - feature to plot
title - plot title, default=None
rows - number of rows to show
'''
corr_abs = data.iloc[:rows]
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(5,7), dpi=200)#, sharey=True)
gs = fig.add_gridspec(1, 3)
sns.heatmap(corr_abs, annot=True, cmap='magma_r', cbar=False,ax=ax[0])
sns.barplot(x=feature, y=corr_abs.index, data=corr_abs, palette="magma")
ax[0].axes.get_xaxis().set_visible(False)
ax[1].axes.get_xaxis().set_visible(False)
ax[1].axes.get_yaxis().set_visible(False)
fig.suptitle(title, fontsize=20)
sns.despine(bottom=True)
# ## ETL
#
# ### tourism data
# read and load tourism data
tourism = pd.read_csv('datasets/raw/city_tourism.csv',sep=';')
# making sure to cleanse the names
for col in ['city','city_name','state','region','country']:
tourism[col] = tourism[col].apply(lambda x: x.strip())
# preparing the cities names to iterate through the cities
cities = tourism['city'].values
# ### calendar data
# +
trusted_calendar_dataset = pd.DataFrame()
for city in tqdm(cities):
# defining path to read data
calendar_path = f'datasets/raw/{city}/calendar.csv.gz'
# extracting data
with gzip.open(calendar_path,'r') as f:
df_calendar = pd.read_csv(BytesIO(f.read()))
# dropping NaN values (not much missing values)
df_calendar.dropna(inplace=True)
# object to numeric transformation
toNumeric_columns = ['price','adjusted_price']
for col in toNumeric_columns:
df_calendar[col] = df_calendar[col].apply(remove_nonnumeric_chars)
# str to datetime transformation
df_calendar['date'] = pd.to_datetime(df_calendar['date'])
# dealing with categorical values
cat_cols_lst = ['available']
df_calendar = create_dummy_df(df_calendar, cat_cols_lst, dummy_na=False)
# Aggregating by listing_id
df_calendar_agg = df_calendar.groupby('listing_id').agg({'date':['min','max'],
'price':['min', 'max', 'mean'],
'minimum_nights':['min','max','mean'],
'maximum_nights':['min','max'],
'available_t':'sum'
})
# Renaming aggregated columns
# concat agg column names
col_names = []
for cols in df_calendar_agg.columns:
new_col = '_'.join(cols)
col_names.append(new_col)
# rename columns
df_calendar_agg.columns = col_names
# Feature engineering
# number of days online
df_calendar_agg['online_days'] = (df_calendar_agg['date_max']-df_calendar_agg['date_min']).apply(lambda x: x.days)+1
df_calendar_agg.drop('date_max', axis=1, inplace=True)
df_calendar_agg.drop('date_min', axis=1, inplace=True)
# occupied days
df_calendar_agg['ocupied_days'] = df_calendar_agg['online_days'] - df_calendar_agg['available_t_sum']
# total revenue generated by the allocation
df_calendar_agg['revenue'] = df_calendar_agg['ocupied_days'] * df_calendar_agg['price_mean']
# ocupation rate
df_calendar_agg['ocupation_rate'] = df_calendar_agg['ocupied_days'] / df_calendar_agg['online_days']
# geographical and tourism data
df_calendar_agg['city'] = city
df_calendar_agg = df_calendar_agg.join(tourism.set_index('city'), on='city', how='left')
df_calendar_agg['foreign_visitors'] = df_calendar_agg['foreign_visitors'].astype('str').apply(lambda x: re.sub('\.', '', x))
df_calendar_agg['visitors_per_capita'] = df_calendar_agg['visitors_per_capita'].apply(lambda x: re.sub(',','.',x)).astype(np.float64)
# calculating relative price based on each city
df_calendar_agg['price_mean_rel'] = df_calendar_agg['price_mean'] / df_calendar_agg['price_mean'].mean()
df_calendar_agg['price_min_rel'] = df_calendar_agg['price_min'] / df_calendar_agg['price_min'].mean()
df_calendar_agg['price_max_rel'] = df_calendar_agg['price_max'] / df_calendar_agg['price_max'].mean()
# calculating ocupation statistics
d = {'listing_id':[],
'ocupation_duration_median':[],
'ocupation_duration_mean':[],
'ocupation_duration_max':[],
'ocupation_duration_min':[]
}
# loop through each acomodation to impute the statistics
for listing_id in df_calendar_agg.index:
vector = df_calendar.loc[df_calendar['listing_id']==listing_id]['available_t']
median_val, mean_val, max_val, min_val = get_sequences(vector, '1')
d['listing_id'].append(listing_id)
d['ocupation_duration_median'].append(median_val)
d['ocupation_duration_mean'].append(mean_val)
d['ocupation_duration_max'].append(max_val)
d['ocupation_duration_min'].append(min_val)
# join with calculated statistics
ocupation_duration_stats = pd.DataFrame(d)
df_calendar_agg = df_calendar_agg.join(ocupation_duration_stats.set_index('listing_id'))
# concat data to the calendar dataset
trusted_calendar_dataset = pd.concat([trusted_calendar_dataset, df_calendar_agg])
# loading data to the trusted directory
trusted_calendar_dataset.to_csv('datasets/trusted/calendar_dataset.csv')
# -
# ### listing data
# +
# creating empty dataset for listings data
trusted_listing_dataset = pd.DataFrame()
for city in tqdm(cities):
# defining path to read data
listing_path = f'datasets/raw/{city}/listings.csv.gz'
with gzip.open(listing_path,'r') as f:
df_listing = pd.read_csv(BytesIO(f.read()))
missing_cols_100 = df_listing.columns[df_listing.isna().mean()==1]
df_listing.drop(missing_cols_100, axis=1, inplace=True)
# for the ones below 5%, dropping the rows
to_drop_rows = df_listing.columns[(df_listing.isna().mean()<0.1)]
df_listing.dropna(subset=to_drop_rows, inplace=True)
object_cols = df_listing.columns[df_listing.dtypes=='object']
# cleansing host data
to_drop = ['host_url', 'host_name', 'host_location', 'host_about', 'host_thumbnail_url', 'host_picture_url', 'host_neighbourhood', 'host_verifications']
df_listing.drop(to_drop, axis=1, inplace=True)
# dropping not needed columns
to_drop = ['listing_url', 'name', 'description', 'picture_url', 'neighbourhood', 'amenities', 'price', 'has_availability',
'neighborhood_overview','calendar_last_scraped','last_review',
'bathrooms_text']
df_listing.drop(to_drop, axis=1, inplace=True)
# dropping not needed columns
to_drop = ['scrape_id', 'host_id', 'host_listings_count', 'neighbourhood_cleansed', 'latitude', 'longitude', 'minimum_nights',
'maximum_nights','minimum_minimum_nights','maximum_minimum_nights','minimum_maximum_nights','maximum_maximum_nights','minimum_nights_avg_ntm','maximum_nights_avg_ntm',
'availability_30','availability_60','availability_365','availability_90','number_of_reviews_ltm','calculated_host_listings_count','calculated_host_listings_count_entire_homes',
'calculated_host_listings_count_shared_rooms','calculated_host_listings_count_private_rooms', 'last_scraped']
df_listing.drop(to_drop, axis=1, inplace=True)
# object to numeric transformation
toNumeric_columns = ['host_response_rate','host_acceptance_rate']
for col in toNumeric_columns:
df_listing[col] = df_listing[col].apply(remove_nonnumeric_chars)
# dealing with categorical columns
categorical_cols = ['host_response_time', 'host_is_superhost', 'host_has_profile_pic', 'host_identity_verified', 'room_type', 'instant_bookable']
df_listing = create_dummy_df(df_listing, categorical_cols, dummy_na=True)
# list of date columns
date_cols = ['host_since','first_review']
# str to datetime transformation
for col in date_cols:
df_listing[col] = pd.to_datetime(df_listing[col])
# transforming the date into the number of days until the last date
for col in date_cols:
df_listing[col] = (df_listing[col].max()-df_listing[col]).apply(lambda x: x.days)
# imputing the median value to the missing values
# The method chosen for NaN values Imputing was the median values, because they are succeptible to the outliars
# and influences less the whole distribution of the variables
df_listing['first_review'] = df_listing['first_review'].fillna(np.median(df_listing['first_review']))
# imputing the median value to the missing values
df_listing = df_listing.fillna(df_listing.median())
# concat datasets
trusted_listing_dataset = pd.concat([trusted_listing_dataset, df_listing])
# drop features that are not always found in axis
for col in ['license','neighbourhood_group_cleansed', 'property_type']:
try:
trusted_listing_dataset.drop(col, axis=1, inplace=True)
except:
print(f'{col} not found in axis')
# making sure that no NaN column is encountered in the dataset
trusted_listing_dataset = trusted_listing_dataset.fillna(trusted_listing_dataset.median())
# loading data to the trusted directory
trusted_listing_dataset.to_csv('datasets/trusted/listing_dataset.csv')
# -
# ### review data
# +
# creating empty dataset for the review data
trusted_reviews_dataset = pd.DataFrame()
for city in tqdm(cities):
# defining path to read data
reviews_path = f'datasets/raw/{city}/reviews.csv.gz'
with gzip.open(reviews_path,'r') as f:
df_reviews = pd.read_csv(BytesIO(f.read()))
# dropping nan values
df_reviews.dropna(inplace=True)
# dropping useless columns
df_reviews.drop(['id', 'reviewer_id', 'reviewer_name'], axis=1, inplace=True)
# performing sentiment analysis on the comments given
sid = SentimentIntensityAnalyzer()
df_reviews["review_comment_score"] = df_reviews['comments'].apply(sid.polarity_scores)
df_reviews["review_comment_score"] = df_reviews["review_comment_score"].apply(lambda review: review['compound'])
# transforming the date into the number of days until the last date
df_reviews['date'] = pd.to_datetime(df_reviews['date'])
df_reviews['review_date'] = (df_reviews['date'].max()-df_reviews['date']).apply(lambda x: x.days)
# dropping the transformed columns
df_reviews.drop(['date','comments'], axis=1, inplace=True)
# Grouping by listing_id
df_reviews_agg = df_reviews.groupby('listing_id').agg({'review_comment_score':['min','max','mean','median'],
'review_date':['min', 'max', 'mean','median'],
})
# concat agg column names
col_names = []
for cols in df_reviews_agg.columns:
new_col = '_'.join(cols)
col_names.append(new_col)
# rename columns
df_reviews_agg.columns = col_names
# concat data to the reviews dataset
trusted_reviews_dataset = pd.concat([trusted_reviews_dataset, df_reviews_agg])
# loading data to the trusted directory
trusted_reviews_dataset.to_csv('datasets/trusted/reviews_dataset.csv')
# -
# ## Data Preparation
# Reading the datasets
# +
# extracting the data
calendar_path = 'datasets/trusted/calendar_dataset.csv'
listing_path = 'datasets/trusted/listing_dataset.csv'
reviews_path = 'datasets/trusted/reviews_dataset.csv'
calendar_trusted = pd.read_csv(calendar_path)
listing_trusted = pd.read_csv(listing_path)
reviews_trusted = pd.read_csv(reviews_path)
# -
# Joining all the raw datasets
df_refined = reviews_trusted.set_index('listing_id')\
.join(listing_trusted.set_index('id'), how='left') \
.join(calendar_trusted.set_index('listing_id'), how='left')
# drop nan values before persisting dataset
df_refined.dropna(inplace=True)
# Reducing the outliars
# +
min_val = 50
df_refined['host_acceptance_rate'] = df_refined['host_acceptance_rate'].apply(lambda x: x if x>=min_val else min_val)
max_val = 3
df_refined['price_max_rel'] = df_refined['price_max_rel'].apply(lambda x: x if x<=max_val else max_val)
max_val = 1000
df_refined['price_max'] = df_refined['price_max'].apply(lambda x: x if x<=max_val else max_val)
max_val = 100
df_refined['minimum_nights_min'] = df_refined['minimum_nights_min'].apply(lambda x: x if x<=max_val else max_val)
max_val = 3
df_refined['price_min_rel'] = df_refined['price_min_rel'].apply(lambda x: x if x<=max_val else max_val)
max_val = 30
df_refined['host_total_listings_count'] = df_refined['host_total_listings_count'].apply(lambda x: x if x<=max_val else max_val)
max_val = 10
df_refined['reviews_per_month'] = df_refined['reviews_per_month'].apply(lambda x: x if x<=max_val else max_val)
# -
# loading the data
df_refined.to_csv('datasets/refined/refined_dataset.csv')
# ## Modelling
# Loading the refined dataset
df_refined = pd.read_csv('datasets/refined/refined_dataset.csv')
# Loading the most important features for the XGBoost model
most_important_XGBoost = pd.read_csv('datasets/refined/most_important_XGBoost.csv', index_col=0)
# #### Using the most important features without relation to the review scores
# +
# listing the most important features plus the target feature
most_important_features = list(most_important_XGBoost.iloc[:10].index) + ['review_scores_rating']
# creating a copy of the dataset in memory to be used on the trainig process
df_refined_feed_model = df_refined[most_important_features].copy()
# using the standard scaler to standardize the data
scaler = StandardScaler()
# separating the train and target features
X = df_refined_feed_model[most_important_features].drop('review_scores_rating', axis=1)
y = df_refined_feed_model['review_scores_rating']
# train and test datasets split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=42)
# standardization of the dataset
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# check the shape of the train and test datasets
print(f'features X_train: {len(X_train[1])}\nfeatures X_test: {len(X_test[1])}')
# Learning - XGBoost
xgbc = XGBRegressor()
xgbc.fit(X_train, y_train)
xgbc_pred_train=xgbc.predict(X_train)
xgbc_pred_test=xgbc.predict(X_test)
# performing crossvalidation test
xgscore = np.mean(cross_val_score(estimator=xgbc,
X=X_train, y=y_train, cv=5))
# printing cross validation results
print(f'XGBoost cross val score: {xgscore}')
# analysing train and test data results
train_score_xgbc = r2_score(y_train, xgbc_pred_train)
test_score_xgbc = r2_score(y_test, xgbc_pred_test)
# printing r2 scores of training and test
print(f'test score: {test_score_xgbc} \ntrain score: {train_score_xgbc}')
# getting the most important features
dict_importance = xgbc.get_booster().get_score(importance_type="total_gain")
d={}
# cleanse the data
for index, value in dict_importance.items():
d[X.columns[int(index[1:])]] = value
# transforming data to a dataframe
most_important_XGBoost = pd.DataFrame(d, index=['Total gain']).T
most_important_XGBoost = most_important_XGBoost[['Total gain']].sort_values(ascending=False, by='Total gain')
# plot most important features
plot_importance(most_important_XGBoost, 'Total gain', 'XGBoost Model [Total gain]')
# -
# #### Using the review scores
# +
# listing the features to be used on the training model
most_important_features = [col for col in df_refined.columns if ('review_scores' in col)
and('accuracy' not in col)
and('value' not in col) ]
# creating a copy of the dataset in memory to be used on the trainig process
df_refined_feed_model = df_refined[most_important_features[:10]].copy()
# using the standard scaler to standardize the data
scaler = StandardScaler()
# separating the train and target features
X = df_refined_feed_model[most_important_features].drop('review_scores_rating', axis=1)
y = df_refined_feed_model['review_scores_rating']
# train and test datasets split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=42)
# standardization of the dataset
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# check the shape of the train and test datasets
print(f'features X_train: {len(X_train[1])}\nfeatures X_test: {len(X_test[1])}')
# Learning - XGBoost
xgbc = XGBRegressor()
xgbc.fit(X_train, y_train)
xgbc_pred_train=xgbc.predict(X_train)
xgbc_pred_test=xgbc.predict(X_test)
# performing crossvalidation test
xgscore = np.mean(cross_val_score(estimator=xgbc,
X=X_train, y=y_train, cv=5))
# printing cross validation results
print(f'XGBoost cross val score: {xgscore}')
# analysing train and test data results
train_score_xgbc = r2_score(y_train, xgbc_pred_train)
test_score_xgbc = r2_score(y_test, xgbc_pred_test)
# printing r2 scores of training and test
print(f'test score: {test_score_xgbc} \ntrain score: {train_score_xgbc}')
# getting the most important features
dict_importance = xgbc.get_booster().get_score(importance_type="total_gain")
d={}
# cleanse the data
for index, value in dict_importance.items():
d[X.columns[int(index[1:])]] = value
# transforming data to a dataframe
most_important_XGBoost_review = pd.DataFrame(d, index=['Total gain']).T
most_important_XGBoost_review = most_important_XGBoost_review[['Total gain']].sort_values(ascending=False, by='Total gain')
# plot most important features
plot_importance(most_important_XGBoost_review, 'Total gain', 'XGBoost Model [Total gain]')
# -
# ## Results
#
# ### Question 1
# Which are the 10 most important characteristics to influence the overall review of an acomodation?
most_important_XGBoost = pd.read_csv('datasets/refined/most_important_XGBoost.csv', index_col=0)
most_important_XGBoost.iloc[:10]
# ### Question 2
# What are the general differences between a top rated acomodation and a almost top one?
# Creating the feature score_group to further answer the questions
df_refined['score_group'] = pd.qcut(df_refined['review_scores_rating'], q=2, labels=['almost top', 'top'])
df_comparison = df_refined.groupby('score_group')[list(most_important_XGBoost.iloc[:10].index)+['review_scores_rating']].median().copy()
# The analysis is made based on the median values of each one, so that the outliars do not influence the observations so much.
# 1. **reviews_per_month** - the top acomodatios receives generally less reviews per month than the not so top ones.
# 2. **review_date_min** - there are no huge difference between them.
# 3. **host_since** - the hosts of the top acomodations are in generall hosts for a longer period in AirBnb.
# 4. **review_date_max** - the higher ranked acomodations are generally newer than their counterparts.
# 5. **number_of_reviews** - the higher ranked acomodations have less reviews.
# 6. **host_total_listings_count** - the hosts owns less acomodations for the higher ranked listings.
# 7. **revenue** - The revenue related to the top acomodations is higher than the others.
# 8. **price_min** - The acomodation price of the top ranked acomodations is higher than the others.
# 9. **host_acceptance_rate** - no difference observed
# 10. **host_response_rate** - no difference obeserved
# 11. **price_max_rel** - The acomodation price of the top ranked acomodations is higher than the others.
# 12. **host_is_superhost_t** - Being a superhost is a sign of the most well ranked acomodations.
#
df_comparison_styled = df_comparison.T.style.background_gradient(cmap='magma',axis=1) #adding a gradient based on values in cell
df_comparison_styled
dfi.export(df_comparison_styled,"images/comparison.png")
# ### Question 3
# Which are the main review aspects that influences the overall review score?
most_important_XGBoost_review
| ProductionNotebook_AirBnb_feature_importance_prd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 7.3
# language: ''
# name: sagemath
# ---
# <h3>FACTORES</h3>
factor(2^127+1)
log(2^127+1,base=10)+1
(log(2^127+1,base=10)+1).n()
floor((log(2^127+1,base=10)+1).n())
def factorizar(N1,N2):
for k in srange(N1,N2):
print k,2^(10*k)+1,floor(log(2^(10*k)+1,base=10).n())+1,factor(2^(10*k)+1)
factorizar(1,5)
time factorizar(1,20)
time factorizar(1,30)
# Tarda bastante en ejecutar $factorizar(1,40)$, y cuando se interrumpe el cálculo se ve que está parado en $2^{35}+1$. Calculamos ese caso concreto:
time factorizar(35,36)
n = 268501*7416361*47392381*2430065924693517198550322751963101*1038213793447841940908293355871461401
time factor(n)
n2 = 2430065924693517198550322751963101*1038213793447841940908293355871461401
time factor(n2)
# Vemos que tarda lo mismo en factorizar $2^{35}+1$ que en factorizar el producto $n$ de sus factores mayores que $10000$, o el producto $n2$ de sus dos factores más grandes.
# <h3>POTENCIAS</h3>
# <p>¿Es mayor $n^{n+1}$ o $(n+1)^n$?</p>
50^(51)>51^(50)
[n^(n+1)>(n+1)^n for n in srange(2,100)]
time L = [n^(n+1)>(n+1)^n for n in srange(3,1000)]
all(L)
L2 = [n^(n+1)-(n+1)^n>(n-3)*n^n for n in srange(3,10)]
L2
time L3 = [n^(n+1)-(n+1)^n>(n-3)*n^n for n in srange(3,1000)]
all(L3)
# <h3>CUADRADOS</h3>
# <p>¿Es cierto que si tenemos $3$ enteros, $x,y,z$, tales que $x^2+y^2=z^2$ NO es posible que ambos $x$ e $y$ sean impares?¿Es cierto que siempre el producto $x\cdot y\cdot z$ de tales enteros es múltiplo de $60$?</p>
[(x,y) for x in srange(1,100) for y in srange(1,100) if is_square(x^2+y^2) and (x%2==1 and y%2==1)]
[(x,y) for x in srange(1,1000) for y in srange(1,1000) if is_square(x^2+y^2) and (x%2==1 and y%2==1)]
# <h3>Otros ejercicios</h3>
#
# <h3>OTROS EJERCICIOS</h3>
# <ol>
# <li>¿Qué enteros primos $p$ se pueden representar como la suma de dos cuadrados de enteros? </li>
# <li>¿Cuántos ceros hay al final, por la derecha, en $100!$ (factorial de cien)?¿Y en $365!$?</li>
# <li> ¿Es cierto que la suma $$1!+2!+3!+\dots+n!$$ no puede ser un cuadrado si $n\ge 4$?</li>
# <li>¿Existen grupos de $k$ enteros positivos consecutivos cuya suma sea una potencia de $2$?¿Y si permitimos que sean enteros positivos o negativos?</li>
# </ol>
fact
| 2_Curso/Laboratorio/SAGE-noteb/IPYNB/INTRO/primera-clase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation,Flatten
from keras.layers import Embedding
from keras.layers import Conv1D, GlobalMaxPooling1D
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from keras.preprocessing.text import Tokenizer
from keras.preprocessing import sequence
from keras import metrics
from sklearn.preprocessing import LabelEncoder
from sklearn.utils import shuffle
from keras import metrics
from keras.layers.merge import concatenate
import sys
sys.path.append('../')
from Utilities.model_visualization import model_to_png
from PIL import Image # used for loading images
import numpy as np
from numpy import asarray
import matplotlib.pyplot as plt
import os # used for navigating to image path
from keras.layers import Input
import cv2
from keras.layers import concatenate
from keras.models import Model
from keras.applications import VGG16
import pandas as pd
from glob import glob
import tensorflow as tf
from keras.callbacks import ModelCheckpoint
from sklearn.model_selection import StratifiedKFold
from keras.preprocessing.image import ImageDataGenerator, img_to_array, load_img
from keras.callbacks import EarlyStopping
import pickle
from scipy import misc
from keras import optimizers
from keras.utils import plot_model
from numpy import array
from numpy import argmax
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
import time
import sys
# -
# <h1>Loading Data</h1>
df = pd.read_pickle('processed_df.pkl')
print(df.iloc[0])
split_pos = int(df['text'].count()*0.8)
train = df[:split_pos]
test = df[split_pos:]
# <h1>CNN Text Model</h1>
# +
dataColumn = 'text'
labelColumn = 'text_info'
tags = train[labelColumn]
texts = train[dataColumn]
tags_Y = test[labelColumn]
texts_Y = test[dataColumn]
# -
print(train.shape)
# +
num_max = 1000
# preprocess
le = LabelEncoder()
tags = le.fit_transform(tags.astype(str))
tok = Tokenizer(num_words=num_max)
tok.fit_on_texts(texts)
mat_texts = tok.texts_to_matrix(texts,mode='count')
print(tags[:5])
print(mat_texts[:5])
print(tags.shape,mat_texts.shape)
# For testing data
le_Y = LabelEncoder()
tags_Y = le_Y.fit_transform(tags_Y.astype(str))
tok_Y = Tokenizer(num_words=num_max)
tok_Y.fit_on_texts(texts_Y)
mat_texts_Y = tok.texts_to_matrix(texts_Y,mode='count')
# +
# for cnn preproces
max_len = 100
cnn_texts_seq = tok.texts_to_sequences(texts)
cnn_texts_mat = sequence.pad_sequences(cnn_texts_seq,maxlen=max_len)
# For testing data
cnn_texts_seq_Y = tok.texts_to_sequences(texts_Y)
cnn_texts_mat_Y = sequence.pad_sequences(cnn_texts_seq_Y,maxlen=max_len)
# -
filepath = "text_weights.h5"
checkpoint = ModelCheckpoint(filepath, monitor='val_acc', verbose=1, save_best_only=True, mode='max', period=1)
early_stopping = EarlyStopping(monitor='val_acc', min_delta=0, patience=4, verbose=1, mode='max')
callbacks_list = [checkpoint, early_stopping]
def get_hybrid_model(): # Pre Trained Embeddings
# load the whole embedding into memory
embeddings_index = dict()
f = open('Embeddings/glove.6B.100d.txt', encoding="utf8")
for line in f:
values = line.split()
word = values[0]
coefs = asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
# create a weight matrix for words in training docs
embedding_matrix = np.zeros((len(tok.word_index) + 1, 100))
for word, i in tok.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
#text classifier
inputs = Input(shape=(100,))
e = Embedding(len(tok.word_index) + 1,
100,
weights=[embedding_matrix],
input_length=max_len,
trainable=False)(inputs)
x = Dropout(0.2)(e)
x = Conv1D(128,
3,
padding='valid',
activation='relu',
strides=1)(x)
x = GlobalMaxPooling1D()(x)
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
hybrid_link = Dense(32, activation='relu', name='hybrid_link')(x)
x = Dense(1, activation='sigmoid', name='Text_Classifier')(hybrid_link)
text_classifier = x
#image classifier
IMAGE_SIZE = [224, 224] # we will keep the image size as (64,64). You can increase the size for better results.
vgg = VGG16(input_shape = (224, 224, 3), weights = None, include_top = True) # input_shape = (64,64,3) as required by VGG
x = (vgg.layers[-2].output)
image_model = Dense(3, activation = 'softmax',name='Hybrid_Classifier')(x) # adding the output layer with softmax function as this is a multi label classification problem.
#hybrid model
concatenate_layer = concatenate([image_model, hybrid_link])
hybrid = Dense(4, activation='softmax')(concatenate_layer)
model = Model(inputs=[vgg.input, inputs], outputs=[hybrid,text_classifier])
return model
# +
model = get_hybrid_model()
model.compile(loss='binary_crossentropy',
optimizer= optimizers.adam(lr=0.00008),
metrics=['acc',metrics.binary_accuracy])
model.summary()
plot_model(model, to_file='multiple_inputs_outputs.png')
# -
# <h1>CNN Image</h1>
# +
IMG_SIZE =224
dataset_dir = 'H:/FYP DATASETS/FYP DATASETS/Crisis/'
def load_img(img):
path = os.path.join(dataset_dir, img)
rows=224
columns=224
img= cv2.resize(cv2.imread(path,cv2.IMREAD_COLOR),(rows,columns),interpolation=cv2.INTER_CUBIC)
return img
# -
for index, row in train.iterrows():
train.at[index,'image_path'] = load_img(row['image_path'])
def encode_label(damage):
# integer encode
damage = np.array(damage)
label_encoder = LabelEncoder()
integer_encoded = label_encoder.fit_transform(damage)
# binary encode
onehot_encoder = OneHotEncoder(sparse=False)
integer_encoded = integer_encoded.reshape(len(integer_encoded), 1)
onehot_encoded = onehot_encoder.fit_transform(integer_encoded)
return onehot_encoded
y = encode_label(train.iloc[:]['damage'])
print(train.iloc[0])
print(train.damage.unique())
train_images = train['image_path'].tolist()
# no need to convert y to list as it is 1 dim encoding takes care of it
train_images = np.array(train_images)
train_text = np.array(train['text'].tolist())
print(cnn_texts_mat.shape)
# +
history = model.fit(x=[train_images,cnn_texts_mat], y=[y,tags],
epochs=40,
batch_size=25,
validation_split=0.2,
shuffle=True,
verbose=1)
# -
model.save_weights('hybrid_only.h5')
| Version 7 (Hybrid)/Basic/Hybrid_only.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using dicom2stl.py to extract an iso-surface from a volume
# This notebook gives a basic introduction to using the `'dicom2stl.py'` script to extract an iso-surface from a volume image.
# +
import os, sys
# download dicom2stl if it's not here already
if not os.path.isdir('dicom2stl'):
# !{'git clone https://github.com/dave3d/dicom2stl.git'}
# -
# ## Create a test volume that is 4 Gaussian blobs arranged in a tetrahedron
from dicom2stl.tests import create_data
tetra = create_data.make_tetra()
# ## Display the tetra volume using [ITK Widgets](https://github.com/InsightSoftwareConsortium/itkwidgets)
try:
import itkwidgets
except:
# !{sys.executable} -m pip install itkwidgets
import itkwidgets
itkwidgets.view(tetra, cmap='Grayscale', vmin=100)
# ## Write the tetra volume to a file
# +
try:
import SimpleITK as sitk
except:
# !{sys.executable} -m pip install SimpleITK
import SimpleITK as sitk
sitk.WriteImage(tetra, "tetra.nii.gz")
# -
# ## Show the command line options for dicom2stl.py
# !{'./dicom2stl/dicom2stl.py -h'}
# ## Extract an iso-surface from the tetra volume
# The `'-i'` flag tells the script the intensity value to use for the iso-surface, `150` in this case. The `'-o'` flag specifies the output file, `tetra.stl`. The script can output STL, VTK or PLY files. And `tetra.nii.gz` is input volume.
# !{'./dicom2stl/dicom2stl.py -i 150 -o tetra.stl tetra.nii.gz'}
# ## Load the mesh
from dicom2stl.utils import vtkutils
mesh = vtkutils.readMesh('tetra.stl')
# ## Display the mesh with the volume
itkwidgets.view(tetra, cmap='Grayscale', geometries=[mesh], vmin=100)
| server/back/dicom2stl/examples/Isosurface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Evaluating a Classification Model
#
# In this exercise, you will create a pipeline for a classification model, and then apply commonly used metrics to evaluate the resulting classifier.
#
# ### Prepare the Data
#
# First, import the libraries you will need and prepare the training and test data:
# +
from pyspark.sql.types import *
from pyspark.sql.functions import *
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.feature import VectorAssembler, StringIndexer, MinMaxScaler
from pyspark.ml.evaluation import BinaryClassificationEvaluator
# Load the source data
flightSchema = StructType([
StructField("DayofMonth", IntegerType(), False),
StructField("DayOfWeek", IntegerType(), False),
StructField("Carrier", StringType(), False),
StructField("OriginAirportID", StringType(), False),
StructField("DestAirportID", StringType(), False),
StructField("DepDelay", IntegerType(), False),
StructField("ArrDelay", IntegerType(), False),
StructField("Late", IntegerType(), False),
])
data = spark.read.csv('wasb://spark@<YOUR_ACCOUNT>.blob.core.windows.net/data/flights.csv', schema=flightSchema, header=True)
# Split the data
splits = data.randomSplit([0.7, 0.3])
train = splits[0]
test = splits[1]
# -
# ### Define the Pipeline and Train the Model
# Now define a pipeline that creates a feature vector and trains a classification model
monthdayIndexer = StringIndexer(inputCol="DayofMonth", outputCol="DayofMonthIdx")
weekdayIndexer = StringIndexer(inputCol="DayOfWeek", outputCol="DayOfWeekIdx")
carrierIndexer = StringIndexer(inputCol="Carrier", outputCol="CarrierIdx")
originIndexer = StringIndexer(inputCol="OriginAirportID", outputCol="OriginAirportIdx")
destIndexer = StringIndexer(inputCol="DestAirportID", outputCol="DestAirportIdx")
numVect = VectorAssembler(inputCols = ["DepDelay"], outputCol="numFeatures")
minMax = MinMaxScaler(inputCol = numVect.getOutputCol(), outputCol="normNums")
featVect = VectorAssembler(inputCols=["DayofMonthIdx", "DayOfWeekIdx", "CarrierIdx", "OriginAirportIdx", "DestAirportIdx", "normNums"], outputCol="features")
lr = LogisticRegression(labelCol="Late", featuresCol="features")
pipeline = Pipeline(stages=[monthdayIndexer, weekdayIndexer, carrierIndexer, originIndexer, destIndexer, numVect, minMax, featVect, lr])
model = pipeline.fit(train)
# ### Test the Model
# Now you're ready to apply the model to the test data.
prediction = model.transform(test)
predicted = prediction.select("features", col("prediction").cast("Int"), col("Late").alias("trueLabel"))
predicted.show(100, truncate=False)
# ### Compute Confusion Matrix Metrics
# Classifiers are typically evaluated by creating a *confusion matrix*, which indicates the number of:
# - True Positives
# - True Negatives
# - False Positives
# - False Negatives
#
# From these core measures, other evaluation metrics such as *precision* and *recall* can be calculated.
tp = float(predicted.filter("prediction == 1.0 AND truelabel == 1").count())
fp = float(predicted.filter("prediction == 1.0 AND truelabel == 0").count())
tn = float(predicted.filter("prediction == 0.0 AND truelabel == 0").count())
fn = float(predicted.filter("prediction == 0.0 AND truelabel == 1").count())
metrics = spark.createDataFrame([
("TP", tp),
("FP", fp),
("TN", tn),
("FN", fn),
("Precision", tp / (tp + fp)),
("Recall", tp / (tp + fn))],["metric", "value"])
metrics.show()
# ### View the Raw Prediction and Probability
# The prediction is based on a raw prediction score that describes a labelled point in a logistic function. This raw prediction is then converted to a predicted label of 0 or 1 based on a probability vector that indicates the confidence for each possible label value (in this case, 0 and 1). The value with the highest confidence is selected as the prediction.
prediction.select("rawPrediction", "probability", "prediction", col("Late").alias("trueLabel")).show(100, truncate=False)
# Note that the results include rows where the probability for 0 (the first value in the **probability** vector) is only slightly higher than the probability for 1 (the second value in the **probability** vector). The default *discrimination threshold* (the boundary that decides whether a probability is predicted as a 1 or a 0) is set to 0.5; so the prediction with the highest probability is always used, no matter how close to the threshold.
# ### Review the Area Under ROC
# Another way to assess the performance of a classification model is to measure the area under a *received operator characteristic (ROC) curve* for the model. The **spark.ml** library includes a **BinaryClassificationEvaluator** class that you can use to compute this. A ROC curve plots the True Positive and False Positive rates for varying threshold values (the probability value over which a class label is predicted). The area under this curve gives an overall indication of the models accuracy as a value between 0 and 1. A value under 0.5 means that a binary classification model (which predicts one of two possible labels) is no better at predicting the right class than a random 50/50 guess.
evaluator = BinaryClassificationEvaluator(labelCol="Late", rawPredictionCol="rawPrediction", metricName="areaUnderROC")
auc = evaluator.evaluate(prediction)
print ("AUC = ", auc)
| Lab03/Classification Evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/satyajitghana/TSAI-DeepNLP-END2.0/blob/main/05_NLP_Augment/SST_Dataset_Augmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="mNwTHtsDKavA"
# # Stanford Sentiment TreeBank Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="t7udJjj-Ly-i" outputId="359259d3-487a-45ed-85c5-ab7539568daf"
# ! pip install pytorch-lightning --quiet
# ! pip install nlpaug --quiet
# ! pip install google-trans-new --quiet
# ! pip install swifter --quiet
# + [markdown] id="JDoFLfACKYSC"
# ## Get to know RAW Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="pBGGh-9H6c2l" outputId="b71a62f1-f2fc-437d-f8dc-9f54b241f379"
# ! wget http://nlp.stanford.edu/~socherr/stanfordSentimentTreebank.zip
# + colab={"base_uri": "https://localhost:8080/"} id="vr-4d0VL6m1Z" outputId="796627da-b567-4b40-956b-b1e853fec2d0"
# ! unzip stanfordSentimentTreebank.zip
# + id="hykrz_eo6jEP"
import os
import pandas as pd
from tqdm.auto import tqdm
import swifter
tqdm.pandas()
# + id="K6KYG1MZ7RDT"
sst_dir = 'stanfordSentimentTreebank'
# + id="jsA_Tbi-6wnl"
sentiment_labels = pd.read_csv(os.path.join(sst_dir, "sentiment_labels.txt"), names=['phrase_ids', 'sentiment_values'], sep="|", header=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="31iuctus7XuA" outputId="b9b3ff51-87df-469f-cb09-30b9834f7d70"
sentiment_labels.head()
# + id="2_TxJUzH_n-s"
def discretize_label(label):
if label <= 0.2: return 0
if label <= 0.4: return 1
if label <= 0.6: return 2
if label <= 0.8: return 3
return 4
# + id="xUPYflCD_tK0"
sentiment_labels['sentiment_values'] = sentiment_labels['sentiment_values'].apply(discretize_label)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="cBWMAPxY_50P" outputId="873774fc-19ff-464b-8177-51d04eb2d738"
sentiment_labels.head()
# + id="bXNcIsSj7cwH"
sentence_ids = pd.read_csv(os.path.join(sst_dir, "datasetSentences.txt"), sep="\t")
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="v2crGNOx7d2E" outputId="aa2a6235-67b2-4b47-f83a-1bb2eb738bbf"
sentence_ids.head()
# + id="zjOqAFYp7hiT"
dictionary = pd.read_csv(os.path.join(sst_dir, "dictionary.txt"), sep="|", names=['phrase', 'phrase_ids'])
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="2A9yaLg87ifW" outputId="bd08a33d-5995-4389-de34-718d4f595ea8"
dictionary.head()
# + id="7eyow7Jp7k-C"
train_test_split = pd.read_csv(os.path.join(sst_dir, "datasetSplit.txt"))
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="sg2TEkZi7l_h" outputId="589b448f-c918-4910-d72d-6834470f8a37"
train_test_split.head()
# + id="MeNZoW0p70bL"
sentence_phrase_merge = pd.merge(sentence_ids, dictionary, left_on='sentence', right_on='phrase')
sentence_phrase_split = pd.merge(sentence_phrase_merge, train_test_split, on='sentence_index')
dataset = pd.merge(sentence_phrase_split, sentiment_labels, on='phrase_ids')
# + id="xTWYox_XXHjd"
dataset['phrase_cleaned'] = dataset['sentence'].str.replace(r"\s('s|'d|'re|'ll|'m|'ve|n't)\b", lambda m: m.group(1))
# + colab={"base_uri": "https://localhost:8080/", "height": 262} id="CypFM4Pf6vCB" outputId="f731d79d-dde5-4e03-a830-39cf20b239eb"
dataset.head()
# + colab={"base_uri": "https://localhost:8080/"} id="d-SFEbgW75P0" outputId="7fcb6e0c-b2e8-467c-ffa3-56f16e20d110"
dataset.iloc[100]
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="tMixwrUy7_hm" outputId="dd48fa25-11a8-404f-de65-816f73c56adb"
dataset.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="VtB2CSpNBHGO" outputId="3fefb05f-6551-4ac3-a4ea-6cd46ca12654"
dataset.info()
# + id="bKs8WE_SYdFr"
dataset.to_csv('sst_dataset_cleaned.csv')
# + [markdown] id="OVveexEZL3uE"
# ## Augmentation Time
# + id="IgPnYxWQtwaC"
import pandas as pd
import numpy as np
from tqdm.auto import tqdm
import swifter
tqdm.pandas()
# + id="Z_nZx-W3hr98"
import nlpaug.augmenter.char as nac
import nlpaug.augmenter.word as naw
import nlpaug.augmenter.sentence as nas
import nlpaug.flow as nafc
from nlpaug.util import Action
# + id="sWJcqMK6LPpJ"
# ! gdown https://drive.google.com/uc?id=1p-e89cyFD2_U1Wx8r9iFmjqSCTnLUt7y
# + id="yetCFPWkuNUL"
dataset = pd.read_csv('sst_dataset_cleaned.csv', index_col=0)
# + [markdown] id="KZcBXExqB_--"
# ## BackTranslate Module
# + id="iI3difHWB_Pu"
from nlpaug.augmenter.word import WordAugmenter
import google_trans_new
from google_trans_new import google_translator
import random
class BackTranslateAug(WordAugmenter):
def __init__(self, name='BackTranslateAug', aug_min=1, aug_max=10,
aug_p=0.3, stopwords=None, tokenizer=None, reverse_tokenizer=None,
device='cpu', verbose=0, stopwords_regex=None):
super(BackTranslateAug, self).__init__(
action=Action.SUBSTITUTE, name=name, aug_min=aug_min, aug_max=aug_max,
aug_p=aug_p, stopwords=stopwords, tokenizer=tokenizer, reverse_tokenizer=reverse_tokenizer,
device=device, verbose=0, stopwords_regex=stopwords_regex)
self.translator = google_translator()
def substitute(self, data):
if not data:
return data
if self.prob() < self.aug_p:
trans_lang = random.choice(list(google_trans_new.LANGUAGES.keys()))
trans_text = self.translator.translate(data, lang_src='en', lang_tgt=trans_lang)
en_text = self.translator.translate(trans_text, lang_src=trans_lang, lang_tgt='en')
return en_text
return data
# + [markdown] id="otgqDIOU1JBr"
# Random Deletion
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="rxt6g-7aiG8M" outputId="db3731f1-c563-4642-f56c-f157d75dfd91"
text = dataset['sentence'].iloc[0]
text
# + colab={"base_uri": "https://localhost:8080/"} id="6VYPIAyc2O8n" outputId="32cc8957-5bb6-473d-cb57-ccd81bb74d24"
aug = naw.RandomWordAug(aug_max=3)
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + [markdown] id="4xVqr-J81LPQ"
# Random Swap
# + colab={"base_uri": "https://localhost:8080/"} id="DXlN8Vso0Zkc" outputId="c47c723f-b84e-475c-d46c-3da5a131df80"
aug = naw.RandomWordAug(action="swap", aug_max=3)
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + [markdown] id="DZ4I_lNQ1lDz"
# Back Translator
# + colab={"base_uri": "https://localhost:8080/"} id="PDQweEVj1wnU" outputId="28c1a65e-c8bc-4056-a5ba-01c53d122180"
aug = BackTranslateAug(aug_max=3, aug_p=1)
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + colab={"base_uri": "https://localhost:8080/"} id="uadCzPLXuseh" outputId="0b25b5c0-61b3-4439-d4a3-71c6c86074cb"
aug = naw.SynonymAug(aug_src='wordnet')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + [markdown] id="rJlaXn4ZiZak"
# Use the backtranslator model to augment the entire dataset, call this backtranslated column of the dataset
# + id="Q9XlfyRIh_Ir"
dataset_aug = dataset.copy()
# + [markdown] id="0YuuRWCI6uGO"
# ## Synonym Augmentor
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["4cb90b1116314ec196c9a805d7c49230", "<KEY>", "7682466a0fbe4b72916e31d8fd97cb9a", "b33794b44b0e4a579872585bd690ac92", "e158db9e591244e691f43e8e6faacc10", "5eaf8b392dca450c9be5edac2141e354", "818c2e2bfec74072afdba10bdf094712", "2c4dde80f973431bad98cdc23b6a3b73"]} id="SkGdzUIyjF2_" outputId="9aad8876-53c4-447c-f49b-689ad0debdde"
aug = naw.SynonymAug(aug_src='wordnet')
synonym_sentences = dataset_aug['sentence'].progress_apply(aug.augment)
# + colab={"base_uri": "https://localhost:8080/"} id="wDVK4N1GuUx0" outputId="87d09062-9865-478c-e108-410fc1cad0e1"
synonym_sentences.head()
# + id="E5WMQGVf05ET"
dataset_aug['synonym_sentences'] = synonym_sentences
# + id="WWVpilwR0-tV"
dataset_aug.to_csv('sst_dataset_synonym.csv')
# + [markdown] id="7wdNj5OuJwNU"
# ## Back Translate
# + [markdown] id="xElNTVSjK_dh"
# See https://docs.google.com/spreadsheets/d/e/2PACX-1vQ5G4wKHEXkseaSy_8khXdmUqfx2jVUK4T-ITSeq8AMB1QWJoyZrpzelCf8Sb70mhs0knjqCEdZguWT/pubhtml for how it was done
# + colab={"base_uri": "https://localhost:8080/"} id="VRo9-vT9Jvcq" outputId="f0b8b4fd-9668-4f06-e09d-57416232b956"
# ! gdown https://drive.google.com/uc?id=1eD_yJb4avApTCET1Q-eNco89FT-QF46g
# + id="RwKwtGMSJ5lK"
translated_dset = pd.read_csv('/content/sst_dataset_translated.csv', index_col=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 501} id="aWsl2DMt03-1" outputId="6d3bf24f-1cd7-44dc-e918-b17f16008d06"
translated_dset.head()
# + id="FYHWNRb2KJIO"
dataset_aug['backtranslated'] = translated_dset['sentence_trans_aug'].copy()
# + [markdown] id="G-HJchoBLFpK"
# ## Final Augmented Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 414} id="BheFuJYoKSrm" outputId="f0425ded-9517-4fd4-bc87-ff9907d41a5b"
dataset_aug.head()
# + id="0HLQGnYiKTmC"
dataset_aug.to_csv('sst_dataset_augmented.csv')
# + id="PEH16XPQKc9u"
| 05_NLP_Augment/SST_Dataset_Augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Part III: Flows and Dependencies
#
# In this part, we show how to follow where specific (faulty) values come from, and why they came to be.
#
# * [Tracking Failure Origins](Slicer.ipynb) shows how to instrument programs such that _data flows_ as well as _control flows_ and the associated dependencies are automatically tracked, allowing to easily find out where a value came from.
| docs/notebooks/03_Dependencies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Fern Fractals
import numpy as np
import matplotlib.pyplot as plt
# The Barnsley Fern is a fractal that resembles the Black Spleenwort species of fern. It is constructed by plotting a sequence of points in the $(x,y)$ plane, starting at $(0,0)$, generated by following the affine transformations $f_1$, $f_2$, $f_2$, and $f_4$ where each transformation is applied to the previous point and chosen at random with probabilities $p_1 = 0.01$, $p_2 = 0.85$, $p_3 = 0.07$, and $p_4 = 0.07$
# $$ f_1(x,y) =
# \begin{bmatrix}
# 0 & 0 \\
# 0 & 0.16
# \end{bmatrix}
# \begin{bmatrix}
# x \\
# y
# \end{bmatrix}
# $$
# $$ f_2(x,y) =
# \begin{bmatrix}
# 0.85 & 0.04 \\
# -0.04 & 0.85
# \end{bmatrix}
# \begin{bmatrix}
# x \\
# y
# \end{bmatrix}
# +
# \begin{bmatrix}
# 0 \\
# 1.6
# \end{bmatrix}
# $$
# $$ f_3(x,y) =
# \begin{bmatrix}
# 0.2 & -.26 \\
# 0.23 & 0.22
# \end{bmatrix}
# \begin{bmatrix}
# x \\
# y
# \end{bmatrix}
# +
# \begin{bmatrix}
# 0 \\
# 1.6
# \end{bmatrix}
# $$
# $$ f_4(x,y) =
# \begin{bmatrix}
# -0.15 & 0.28 \\
# 0.26 & 0.24
# \end{bmatrix}
# \begin{bmatrix}
# x \\
# y
# \end{bmatrix}
# +
# \begin{bmatrix}
# 0 \\
# 0.44
# \end{bmatrix}
# $$
# ### Part 0: Introduction to vector tranformations
# The functions listed above are known as vector transformations. They are extremely similar to functions you're used to, but instead of operating on scalars, they operate on vectors. Here is an extremely brief introduction to what vector functions are and how we'll use them in this assignment:
#
# The functions we're used to seeing in math take a number input and produce some other number.
# $$ f(x) = y$$
#
# Similarily, vector functions take a vector input and produce some other vector.
# $$ f(\ \vec v\ ) = \vec u $$
# While programming, you've probably already dealt with something extremely similar. For example this following function is a vector transformation. This function takes a vector as an argument ($\vec{v} = [x, y]$) and returns some other vector ($\vec u = [a,b]$).
#
# def my_program(x, y):
# a = 3 * x
# b = 5 * y
# return a, b
#
# In the functions defined above, the input vector [x, y] is multiplied by a matrix and added to another vector. Here's an example of how matrix multiplication works.
# $$
# \begin{bmatrix}
# a & b \\
# c & d
# \end{bmatrix}
# \begin{bmatrix}
# x \\
# y
# \end{bmatrix} =
# \begin{bmatrix}
# ax + by \\
# cx + dy
# \end{bmatrix}
# $$
# The programming function that correspond to this would look like:
#
# def function(x, y):
# new_x = a*x + b*y
# new_y = c*x + d*y
# return new_x, new_y
# ### Part 1: Defining the transformation functions
# Let's start by creating functions for the transformations defined above.
# Define f1
def f1(x, y):
# TO DO: fill me in
return(0, 0.16*y)
pass
assert(abs (f1(1,1)[0] - (0)) < 0.0001)
assert(abs (f1(1,1)[1] - (0.16)) < 0.0001)
# TO DO: create an additional assert statement
assert(abs(f1(1,2)[1] - (0.32)) < 0.0001)
# Define f2
def f2(x, y):
# TO DO: fill me in
x_new = 0.85*x + 0.04*y
y_new = -0.04*x + 0.85*y + 1.6
return(x_new, y_new)
pass
assert(abs (f2(1,1)[0] - (0.89)) < 0.0001)
assert(abs (f2(1,1)[1] - (2.41)) < 0.0001)
# TO DO: create an additional assert statement
assert(abs(f2(2,2)[1] - (3.22)) < 0.001)
# Define f3
def f3(x, y):
# TO DO: fill me in
x_new = 0.2*x + -0.26*y
y_new = 0.23*x + 0.22*y + 1.6
return(x_new, y_new)
pass
assert(abs (f3(1,1)[0] - (-0.06)) < 0.0001)
assert(abs (f3(1,1)[1] - (2.05)) < 0.0001)
# TO DO: create an additional assert statement
assert(abs(f3(1,2)[1] - (2.27)) < 0.0001)
# Define f4
def f4(x, y):
# TO DO: fill me in
x_new = -0.15*x + 0.28*y
y_new = 0.26*x + 0.24*y + 0.44
return(x_new, y_new)
pass
assert(abs (f4(1,1)[0] - (0.13)) < 0.0001)
assert(abs (f4(1,1)[1] - (0.94)) < 0.0001)
# TO DO: create an additional assert statement
assert(abs((f4(1,2)[1]) - (1.18)) < 0.0001)
# ### Part 2: Applying the transformations
# Now that we have our transformations defined, let's apply them! Defining this function is not hard, but it's also not intuitive. Below I've outlined one way you can create your `generate_fractal` function.
# To give you some intuition into what this function is doing, you can think of it as creating a "picture". We start with an `x` and `y` value and apply one of the transformations above. The tranformation gives us a new `x` and `y` value. We plot this `x` and `y` and then we repeat this process for some number of iterations (N).
#
# Here's a walk through of one way you can implement the function below:
# 1. Initialize `x` and `y` to 0.
# 2. Create two arrays (I'll refer to them as `XA` and `YA`) both with length N. As you iterate and apply your transformation functions, you'll store your resulting `x` and `y` values in these arrays.
# 3. Create a list of the functions you defined above
# 4. Make a `for` loop that iterates N times over the following:
# 1. Randomly select one of your functions above using the probabilities listed in the introduction, I recommend `np.random.choice(...)`. Make sure your list of probabilities is in the same order as your list of functions.
# 2. Get your new `x`, `y` values from calling the function selected in the previous step with your existing `x`, `y` values
# 3. Set the `ith` index of `XA` and `YA` to the new `x` and `y` values respectively. (The `ith` index in this case is whatever number loop you are on)
# 5. Create a scatter plot of your X and Y value arrays
# 1. Use `plt.scatter(....)` and include the arguments `s = 1` and `marker = "o"` to get the cleanest image
# 2. Additionally, add color! To do this, create an additional array of length N (I'll refer to this as `color_array`). The numbers in this array are up to you, but should not be all zero).
# 3. In your `plt.scatter(...)` call, add arguments `c = color_array` and `cmap = "Greens"`. Once this is working for you, you're welcome to look up and use other colormaps.
#
# <b> I recommend increasing your figure size so your plot is easier to see. Do this by adding `plt.figure(figsize=(10,10))` before you call `plt.scatter(...)`</b>
# When your testing your function, use at least N = 5000, or it'll be extremely hard to tell from the plot if you've done it correctly.
def generate_fractal(N):
# TO DO: Fill me in!
x = 0
y = 0
XA = np.empty(N)
YA = np.empty(N)
funcs = f1, f2, f3, f4
p = [0.01, 0.85, 0.07, 0.07]
for i in range(N):
x, y = np.random.choice(funcs, p=p)(x, y)
XA[i] = x
YA[i] = y
plt.scatter(XA, YA, s=0.05, c='#4F7942')
pass
# ### Part 3: Run your code with N = 100,000.
# +
# Call your function with N = 100,000.
# NOTE: This should only take a few seconds to run.
# If your code takes longer, see if there's anything you can change that'll
# improve performance (is there anything in the loop that doesn't need to be?)
N = 100000
generate_fractal(N)
# -
| Assignment6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. Click `PCA-maSigFun` at the menu
#
# 2. select your data, if you didn't found your data, please click the blue words `expression` or `convariates` to `Upload Data` page for upload data.
#
# **Quantitative factor**: name of the numerical variable of the experimental design, normally the time.
#
# **Qualitative factor**: name of the categorical variable of the experimental design.
#
# **Control group**: name of the reference series in the regression model (in the example is Ctr).
#
# Here we set example data. all options as following this picture.
#
# 
#
# 3. Select database. SEA supposts 11 organism. But our example data not belongs to any database. So we choose our annotion data.
#
# 
#
# 4. About `options`, here we set as defaults.
#
# **Polynomial degree**: degree for the regression model. The maximum allowed degree is #time_points – 1.
#
# **Alpha**: significant level for gene selection.
#
# **R-Squared cut-off**: required level of the goodness of fit of the regression model. This parameter is between 0 and 1. Higher values indicate well fitted models.Recommend values are between [0.4,0.8].
#
# **Cut-off**: Variability level to select Principal Components in each category.
#
# **Selection factor**: criterion to select components can be:
#
# Proportion of acumulated variability. Posible cut-off values are in (0,1).
#
# Proportion of variability of each PC. Posible cut-off values are in (0,1).
#
# Average: components are selected that explain more than “cut-off” times the average component variability. The recommended “cut-off” values are in [1,1.5].
#
#
# 5. You must give a name `job name` to your PCA-maSigFun analysis to easy find out which is this analysis.
#
#
# 6. At the end, click `Run` to analysis.
# # Results
# 
# # maSigVisualization
# Click `Send to maSig visualization`, we get the interface of `maSigVisualization` and had fill all options.
# 
# **Clustering method**: available methods are:
#
# 'hclust': hierarchical clustering
# 'kmeans': k-means
#
# **Number of clusters**: groups to split gene selection to show results.
#
#
# **PCA parameters**: Threshold for significant gene contribution for the PCA model. This threshold allows the identification of the genes that most contribute to the selected components. It can be computed by applying several procedures:
#
#
# **Resampling**: where a null Leverage distribution is created by permuting columns of expression data and genes are selected at the “alpha” percentile of the null distribution.
#
# **minAS**: where a density function is calculated on the data and genes are selected on a local minimum basis [7].
#
# **Gamma**: where a gamma distribution is adjusted to the distributions of the gene loadings, and genes are selected at the “alpha” percentile of the gamma distribution [7].
#
# **Custom**: where the user can decide the threshold.
#
# 
# 
# 
| time-course/workflows/SEA/PCA-maSigFun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd # DateFrame, Series
import numpy as np # Scientific Computing package - Array
from sklearn import tree
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn.model_selection import train_test_split
from matplotlib import pyplot as plt
import seaborn as sns
import graphviz
import pydotplus
import io
from scipy import misc
# %matplotlib inline
# -
# # Spotify Song Attributes EDA
# - Import Dataset
# - EDA to visualize data and observe structure
# - Train a classifier (Decision Tree)
# - Predict target using the trained classifier
data = pd.read_csv('data.csv')
data.head()
data.describe()
train, test = train_test_split(data, test_size=0.15)
print('Training size: {}; Test size: {}'.format(len(train), len(test)))
# Custom Color Palette
red_blue = ['#19B5FE', '#EF4836']
palette = sns.color_palette(red_blue)
sns.set_palette(palette)
sns.set_style('white')
pos_tempo = data[data['target'] == 1]['tempo']
neg_tempo = data[data['target'] == 0]['tempo']
pos_dance= data[data['target'] == 1]['danceability']
neg_dance = data[data['target'] == 0]['danceability']
pos_duration = data[data['target'] == 1]['duration_ms']
neg_duration = data[data['target'] == 0]['duration_ms']
pos_loudness = data[data['target'] == 1]['loudness']
neg_loudness = data[data['target'] == 0]['loudness']
pos_speech = data[data['target'] == 1]['speechiness']
neg_speech = data[data['target'] == 0]['speechiness']
pos_valence = data[data['target'] == 1]['valence']
neg_valence = data[data['target'] == 0]['valence']
pos_energy = data[data['target'] == 1]['energy']
neg_energy = data[data['target'] == 0]['energy']
pos_acoustic = data[data['target'] == 1]['acousticness']
neg_acoustic = data[data['target'] == 0]['acousticness']
pos_key = data[data['target'] == 1]['key']
neg_key = data[data['target'] == 0]['key']
pos_instrumental = data[data['target'] == 1]['instrumentalness']
neg_instrumental = data[data['target'] == 0]['instrumentalness']
fig = plt.figure(figsize=(12, 8))
plt.title('Song Tempo Like / Dislike Distribution')
pos_tempo.hist(alpha=0.7, bins=30, label='positive')
neg_tempo.hist(alpha=0.7, bins=30, label='negative')
plt.legend(loc='upper right')
plt.show()
# +
fig2 = plt.figure(figsize=(15, 15))
# Danceability
ax3 = fig2.add_subplot(331)
ax3.set_xlabel('Danceability')
ax3.set_ylabel('Count')
ax3.set_title('Song Danceability Like / Dislike Distribution')
pos_dance.hist(alpha=0.5, bins=30)
ax4 = fig2.add_subplot(331)
neg_dance.hist(alpha=0.5, bins=30)
# Duration
ax5 = fig2.add_subplot(332)
ax5.set_xlabel('Duration')
ax5.set_ylabel('Count')
ax5.set_title('Song Duration Like / Dislike Distribution')
pos_duration.hist(alpha=0.5, bins=30)
ax6 = fig2.add_subplot(332)
neg_duration.hist(alpha=0.5, bins=30)
# Loudness
ax7 = fig2.add_subplot(333)
ax7.set_xlabel('Loudness')
ax7.set_ylabel('Count')
ax7.set_title('Song Loudness Like / Dislike Distribution')
pos_loudness.hist(alpha=0.5, bins=30)
ax8 = fig2.add_subplot(333)
neg_loudness.hist(alpha=0.5, bins=30)
# Speechiness
ax9 = fig2.add_subplot(334)
ax9.set_xlabel('Speechiness')
ax9.set_ylabel('Count')
ax9.set_title('Song Speechiness Like / Dislike Distribution')
pos_speech.hist(alpha=0.5, bins=30)
ax10 = fig2.add_subplot(334)
neg_speech.hist(alpha=0.5, bins=30)
# Valence
ax11 = fig2.add_subplot(335)
ax11.set_xlabel('Valence')
ax11.set_ylabel('Count')
ax11.set_title('Song Valence Like / Dislike Distribution')
pos_valence.hist(alpha=0.5, bins=30)
ax12 = fig2.add_subplot(335)
neg_valence.hist(alpha=0.5, bins=30)
# Energy
ax13 = fig2.add_subplot(336)
ax13.set_xlabel('Energy')
ax13.set_ylabel('Count')
ax13.set_title('Song Energy Like / Dislike Distribution')
pos_energy.hist(alpha=0.5, bins=30)
ax14 = fig2.add_subplot(336)
neg_energy.hist(alpha=0.5, bins=30)
# Acoustic
ax15 = fig2.add_subplot(337)
ax15.set_xlabel('Acousticness')
ax15.set_ylabel('Count')
ax15.set_title('Song Acousticness Like / Dislike Distribution')
pos_acoustic.hist(alpha=0.5, bins=30)
ax16 = fig2.add_subplot(337)
neg_acoustic.hist(alpha=0.5, bins=30)
# Key
ax17 = fig2.add_subplot(338)
ax17.set_xlabel('Key')
ax17.set_ylabel('Count')
ax17.set_title('Song Key Like / Dislike Distribution')
pos_key.hist(alpha=0.5, bins=30)
ax18 = fig2.add_subplot(338)
neg_key.hist(alpha=0.5, bins=30)
# Instrumentalness
ax19 = fig2.add_subplot(339)
ax19.set_xlabel('Instrumentalness')
ax19.set_ylabel('Count')
ax19.set_title('Song Instrumental Like / Dislike Distribution')
pos_instrumental.hist(alpha=0.5, bins=30)
ax20 = fig2.add_subplot(339)
neg_instrumental.hist(alpha=0.5, bins=30)
# -
c = DecisionTreeClassifier(min_samples_split=100)
features = ['danceability', 'duration_ms', 'loudness', 'speechiness', 'valence', 'energy', 'acousticness', 'key', 'instrumentalness']
# +
X_train = train[features]
y_train = train['target']
X_test = test[features]
y_test = test['target']
# -
dt = c.fit(X_train, y_train)
def show_tree(tree, features, path):
f = io.StringIO()
export_graphviz(tree, out_file=f, feature_names=features)
pydotplus.graph_from_dot_data(f.getvalue()).write_png(path)
img = misc.imread(path)
plt.rcParams['figure.figsize'] = (20,20)
plt.imshow(img)
show_tree(dt, features, 'decision_tree_1')
y_pred = c.predict(X_test)
y_pred
# +
from sklearn.metrics import accuracy_score
score = accuracy_score(y_test, y_pred) * 100
# -
print("Accuracy using Decision Tree: " )
| Spotify-EDA-DecisionTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
# %matplotlib inline
hpi_url = "https://raw.githubusercontent.com/TrainingByPackt/Interactive-Data-Visualization-with-Python/master/datasets/hpi_data_countries.tsv"
# Once downloaded, read it into a DataFrame using pandas
hpi_df = pd.read_csv(hpi_url, sep='\t')
# +
import seaborn as sns
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_subplot(111)
ax = sns.scatterplot(x='Wellbeing (0-10)', y='Happy Planet Index', hue='Region', data=hpi_df)
plt.show()
# -
import altair as alt
alt.Chart(hpi_df).mark_circle().encode(
x='Wellbeing (0-10):Q',
y='Happy Planet Index:Q',
color='Region:N',
).interactive()
| Lesson04/Exercise27.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/https-deeplearning-ai/tensorflow-1-public/blob/master/C3/W3/ungraded_labs/C3_W3_Lab_5_sarcasm_with_bi_LSTM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Q2MY4-M1zuhV"
# # Ungraded Lab: Training a Sarcasm Detection Model using Bidirectional LSTMs
#
# In this lab, you will revisit the [News Headlines Dataset for Sarcasm Detection](https://www.kaggle.com/rmisra/news-headlines-dataset-for-sarcasm-detection/home) dataset and use it to train a Bi-LSTM Model.
#
# + [markdown] id="S-AgItE6z80t"
# ## Download the Dataset
#
# First, you will download the JSON file and extract the contents into lists.
# + id="k_Wlz9i10Dmn"
# Download the dataset
# !wget https://storage.googleapis.com/tensorflow-1-public/course3/sarcasm.json
# + id="Pr4R0I240GOh"
import json
# Load the JSON file
with open("./sarcasm.json", 'r') as f:
datastore = json.load(f)
# Initialize the lists
sentences = []
labels = []
# Collect sentences and labels into the lists
for item in datastore:
sentences.append(item['headline'])
labels.append(item['is_sarcastic'])
# + [markdown] id="zN9-ojV55UCR"
# ## Split the Dataset
#
# You will then split the lists into train and test sets.
# + id="50H0ZrJf035i"
training_size = 20000
# Split the sentences
training_sentences = sentences[0:training_size]
testing_sentences = sentences[training_size:]
# Split the labels
training_labels = labels[0:training_size]
testing_labels = labels[training_size:]
# + [markdown] id="MYVNY4tE5YbN"
# ## Data preprocessing
#
# Next, you will generate the vocabulary and padded sequences.
# + id="hodsUZib1Ce7"
import numpy as np
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
vocab_size = 10000
max_length = 120
trunc_type='post'
padding_type='post'
oov_tok = "<OOV>"
# Initialize the Tokenizer class
tokenizer = Tokenizer(num_words=vocab_size, oov_token=oov_tok)
# Generate the word index dictionary
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
# Generate and pad the training sequences
training_sequences = tokenizer.texts_to_sequences(training_sentences)
training_padded = pad_sequences(training_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
# Generate and pad the testing sequences
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
# Convert the labels lists into numpy arrays
training_labels = np.array(training_labels)
testing_labels = np.array(testing_labels)
# + [markdown] id="o23gJhj95el5"
# ## Build and Compile the Model
#
# The architecture here is almost identical to the one you used in the previous lab with the IMDB Reviews. Try to tweak the parameters and see how it affects the training time and accuracy (both training and validation).
# + id="jGwXGIXvFhXW"
import tensorflow as tf
# Parameters
embedding_dim = 16
lstm_dim = 32
dense_dim = 24
# Model Definition with LSTM
model_lstm = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(lstm_dim)),
tf.keras.layers.Dense(dense_dim, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
# Set the training parameters
model_lstm.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# Print the model summary
model_lstm.summary()
# + [markdown] id="krcQGm7B5g9A"
# ## Train the Model
# + colab={"background_save": true} id="nEKV8EMj11BW"
NUM_EPOCHS = 10
# Train the model
history_lstm = model_lstm.fit(training_padded, training_labels, epochs=NUM_EPOCHS, validation_data=(testing_padded, testing_labels))
# + id="g9DC6dmLF8DC"
import matplotlib.pyplot as plt
# Plot Utility
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
# Plot the accuracy and loss history
plot_graphs(history_lstm, 'accuracy')
plot_graphs(history_lstm, 'loss')
| C3/W3/ungraded_labs/C3_W3_Lab_5_sarcasm_with_bi_LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hrishipoola/Customer_RFMT_Metrics_Segmentation/blob/main/Customer_RFMT_Metrics_%26_Segmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9tzncua3zv05"
# # Customer RFMT Metrics & Segmentation
# + [markdown] id="0u-agRJ6z2T_"
# ## Table of Contents
#
# 1. Introduction
# 2. Install & Import Packages
# 3. Load & Process Data
# <br> Orders
# <br> Payments
# <br> Outliers
# <br> Merge
# 4. Create RFMT Metrics
# 5. Manually Segment & Score
# 6. K-means Clustering
# <br> Box Cox Transformation
# <br> Scaling
# <br> Optimal K
# <br> Fit
# 7. Segments
# <br> Snake Plot
# <br> Relative Importants of Metrics
# 8. References
#
#
# + [markdown] id="1799BiwSvGzl"
# ## 1. Introduction
#
# Today, we'll construct recency, frequency, monetary value, and tenure (RFMT) metrics and segments using Brazilian ecommerce marketplace [Olist's sales transactions data](https://www.kaggle.com/olistbr/brazilian-ecommerce?select=olist_orders_dataset.csv) dating from October 2016 to October 2018.
#
# RFMT metrics can be used to segment customers in order to identify which customers are responsive to marketing, engaged, contribute to churn, high spenders vs. low-value purchasers, or have upselling or cross-selling potential. Understanding segments can help us better tailor product, sales, and marketing activities and investments. For example, at-risk customers may have high monetary value and frequency, but weak recency and could be targeted with promotions and renewals. In our case, we'll define our metrics as:
#
# - Recency: days since last transaction (delivery)
# - Frequency: number of transactions during time period
# - Monetary value: total spend during time period
# - Tenure: days since first purchase order
#
# To construct RFMT metrics, we'll need order id, purchase history, order status, delivery dates, and spend details by unique customer id. Olist's data schema shows that the orders data set contains unique customer ids, order status, and delivery dates, while the payments data set contains spend. Let's merge these two data sets together on order id to get what we need.
#
# We'll use RFMT metrics to segment customers, first manually by building RFMT scores along with arbitrary cutoffs and then using K-means clustering to uncover segemnts in the data (an alternative to K-means would be non-zero matrix factorization (NMF)). We'll then compare our 4 resulting segments and relative importance of segment metrics.
#
# Future areas to explore:
#
# - Tailor metrics to product categories. For example, we could weight R and F higher and M lower for FMCG (e.g., cosmetics, headphones), while weighting M higher and R and F lower for durable goods (e.g., washing machines)
# - Merge [marketing funnel data set](https://www.kaggle.com/olistbr/marketing-funnel-olist/home) to understand and model how customer journey shapes purchasing behavior
# + [markdown] id="bhEyM7QHF036"
# ## 2. Install & Import Packages
# + id="kbLgqVNosvzL"
import pandas as pd
import numpy as np
from datetime import date, timedelta
from scipy import stats
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import KMeans
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib import style
sns.set_style("darkgrid")
style.use('fivethirtyeight')
import io
from io import BytesIO
from zipfile import ZipFile
# + colab={"base_uri": "https://localhost:8080/"} id="FkY_S_w9wWZk" outputId="30e720ea-ca29-4fe0-8c96-325327487563"
# Mount google drive
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="OkScCXUJGRj5"
# ## 3. Load & Process Data
# + [markdown] id="T3c5tPUSSjO4"
# ### Orders
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 72} id="m-NcyURn6CmR" outputId="e399f882-f6fc-40bd-bbe0-00e00e78d02c"
from google.colab import files
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="yD3wDUetws6l" outputId="fc927c35-3730-49d4-c19c-d7b46f283d08"
# Read in csv zip file
zipfile = ZipFile(BytesIO(uploaded['olist_orders_dataset.csv.zip']))
orders = pd.read_csv(zipfile.open('olist_orders_dataset.csv'))
orders.head()
# + colab={"base_uri": "https://localhost:8080/"} id="W0RK59ZYRZ5-" outputId="d3e4987a-b80f-40a0-a5c8-88d016c50127"
orders.info()
# + colab={"base_uri": "https://localhost:8080/"} id="RuAGj8P3W_4x" outputId="03fa14e8-ae0c-48cb-a9b7-3a46a6a1c6ca"
# Double-check for duplicate orders - there are no duplicates
orders.duplicated().value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="QzR6ageARitE" outputId="11516a9b-49ca-4cef-c574-a57f790c3a2c"
# Convert date columns to datetime format
dates = ['order_purchase_timestamp',
'order_approved_at',
'order_delivered_carrier_date',
'order_delivered_customer_date',
'order_estimated_delivery_date']
for col in dates:
orders[col] = pd.to_datetime(orders[col])
orders.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="HoRFkvTsR2oy" outputId="6ae6f93a-9ae2-4ece-e49e-b50dde7986bb"
orders['order_status'].value_counts()
# + id="v04RtRHCR-lA"
# Filter orders
delivered = orders.query('order_status == "delivered"')
# + colab={"base_uri": "https://localhost:8080/"} id="f69Fcof0SLFs" outputId="936d3bd3-dd87-44ad-96f2-1167577a4e71"
delivered.isnull().sum()
# + [markdown] id="R5itq3qPSn2E"
# ### Payments
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 72} id="dZUmBa35Sngf" outputId="103785f5-fe70-4b62-def0-5bd7a790479f"
from google.colab import files
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="bhnUFq3LAJ-q" outputId="c9bce4d2-d841-4710-ad55-9e4b683f14e1"
# Read in csv zip file
zipfile = ZipFile(BytesIO(uploaded['olist_order_payments_dataset.csv.zip']))
payments = pd.read_csv(zipfile.open('olist_order_payments_dataset.csv'))
payments.head()
# + colab={"base_uri": "https://localhost:8080/"} id="JmVRszbCS3q6" outputId="d2a16a42-fc99-4ff8-8dc6-68c06be65751"
payments.info()
# + colab={"base_uri": "https://localhost:8080/"} id="xqkt4zs7ZMll" outputId="963537a9-f65e-4fe3-ccf0-2c1faba77f8b"
# Double-check for duplicate orders - there are no duplicates
payments.duplicated().value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="rSkpXMWXlRkL" outputId="75b0cd24-e3dc-4d62-a047-ecf1c7648322"
# Keep only non-zero payment values
payments = payments.query('payment_value > 0')
payments.describe()
# + [markdown] id="DBllaKiGUGhB"
# ### Outliers
#
# Let's take a look at payment value outliers (beyond 3 standard deviations).
# + colab={"base_uri": "https://localhost:8080/", "height": 189} id="SC3HfD7uTFsg" outputId="c6f941ec-628d-4875-faad-fce711fd70fb"
# Box plot: distribution is skewed with higher end outliers
square = dict(markerfacecolor='salmon', markeredgecolor='salmon', marker='.')
payments.payment_value.plot(kind='box', vert=False, flierprops=square, figsize=(18,1.5))
plt.xlabel('R$')
plt.title('Distribution of Payment Values')
# + colab={"base_uri": "https://localhost:8080/"} id="T-Rhzn8yYc6z" outputId="abed10ea-c2d3-4d0b-8db8-350b5393fecc"
# Remove outliers (beyond 3 standard deviations)
z = np.abs(stats.zscore(payments['payment_value']))
payments_filtered = payments[(z < 3)]
outliers = payments[(z > 3)] # Keep the outliers as a separate dataframe in case we want to explore them
print('There are {} outlier payment values.'.format(outliers.shape[0]))
# + colab={"base_uri": "https://localhost:8080/"} id="wHv9Vy_3V3Zy" outputId="eb9fef00-92b1-4d1c-d31d-b5c9b9473b0c"
# Payments with outliers removed is less skewed
payments_filtered.payment_value.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 189} id="CkYrjpDLTjph" outputId="ce818c3e-88d7-4d76-aedb-2f8a2c311e18"
# Box plot: distribution is skewed with higher end outliers
square = dict(markerfacecolor='salmon', markeredgecolor='salmon', marker='.')
payments_filtered.payment_value.plot(kind='box', vert=False, flierprops=square, figsize=(18,1.5))
plt.xlabel('R$')
plt.title('Distribution of Payment Values Ex-Outliers')
# + colab={"base_uri": "https://localhost:8080/", "height": 399} id="Po_8aga1XoJv" outputId="2b683b03-8b05-4b7d-ddfb-cd3b4b7ac321"
# Plot distribution
sns.displot(payments_filtered['payment_value'],bins=30, kde=True, color='turquoise')
plt.title('Distribution of Payment Values (Ex-Outliers)')
plt.xlabel('R$')
# + colab={"base_uri": "https://localhost:8080/", "height": 399} id="4tRwXZhqjHXA" outputId="d52720cc-797c-4c43-e955-8bb1a552f143"
# Plot distribution
sns.displot(outliers['payment_value'],bins=50, color='salmon')
plt.title('Distribution of Payment Value Outliers')
plt.xlabel('R$')
# + [markdown] id="mPUHCOPHi3AY"
# Let's actually keep the outliers in as they could represent an important group of customers to understand and segment.
#
# In the future, we could remove the outliers and run a further analysis and segmentation focusing only on core customers. We could also dive into the outlier group itself.
# + [markdown] id="r8oI-vALWGFs"
# ### Merge
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="46yuBkU6AX8C" outputId="b4cdc06f-2fdc-4636-9e1d-7e354519086d"
# Merge delivered and payments
df = delivered.merge(payments, on ='order_id',how = 'outer')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="52NMDSGIF6jo" outputId="a735bea3-482e-41b7-8ca4-a1c356614946"
df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="_bNuHdsXWEp5" outputId="71e64fab-1953-4044-fd82-ae979ea53c9f"
print('Delivery timestamps range from {} to {}.'.format(min(df.order_delivered_customer_date), max(df.order_delivered_customer_date)))
# + [markdown] id="6DrXlL6BVT40"
# ## 4. Create RFMT Metrics
#
#
#
# + id="oXu_C8cOUuud"
# RFMT metrics
snapshot_date = df['order_delivered_customer_date'].max() + timedelta(days=1)
rfmt = df.groupby('customer_id') \
.agg({'order_delivered_customer_date': lambda x: (snapshot_date - x.max()).days, # recency
'order_id': 'count', # frequency
'payment_value': 'sum', # monetary value
'order_purchase_timestamp': lambda x: (snapshot_date - x.min()).days # tenure
})
rfmt.dropna(inplace=True)
rfmt = rfmt.query('payment_value > 0') # keep only non-zero monetary values, box cox transformation requires positive values
rfmt.columns = ['recency','frequency','monetary_value','tenure']
# + [markdown] id="xyNApFdlDfHW"
# ## 5. Manually Segment & Score
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="eJf5t6MJVDWR" outputId="1ddfa290-bfb0-466a-f23e-9b67f2c90153"
# Recency
recency_labels = range(3,0,-1) # low recency is better than high recency
recency_group = pd.qcut(rfmt['recency'], 3, labels=recency_labels, duplicates='drop')
# Frequency
frequency_labels = range(1,2)
frequency_group = pd.qcut(rfmt['frequency'], 1, labels=frequency_labels, duplicates='drop')
# Monetary value
monetary_labels = range(1,5)
monetary_group = pd.qcut(rfmt['monetary_value'], 4, labels=monetary_labels, duplicates='drop')
# Tenure
tenure_labels = range(1,5)
tenure_group = pd.qcut(rfmt['tenure'], 4, labels=tenure_labels, duplicates='drop')
rfmt = rfmt.assign(R=recency_group.values, F=frequency_group.values, M=monetary_group.values, T=tenure_group.values)
rfmt['RFMT_Segment'] = rfmt['R'].astype(str) + rfmt['F'].astype(str) + rfmt['M'].astype(str) + rfmt['T'].astype(str)
rfmt['RFMT_Score'] = rfmt[['R','F','M','T']].sum(axis=1)
rfmt
# + colab={"base_uri": "https://localhost:8080/"} id="X7WGv6t-Voys" outputId="594df54f-865a-49d2-f350-955c2903942f"
# 10 largest RFMT segments
rfmt.groupby('RFMT_Segment').size().sort_values(ascending=False)[:10]
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="DGaOQ1XdVuXw" outputId="b4f36bcf-67ab-49f0-b7c7-1ce98eea2546"
rfmt.groupby('RFMT_Score').agg({'recency':'mean',
'frequency':'mean',
'monetary_value':['mean','count'],
'tenure': 'mean'
}).round(1)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="QvFKynyQWB7Q" outputId="9f96bf3a-c1ae-481b-c361-d41f606566e4"
def segment_name(df):
if df['RFMT_Score'] >= 10:
return 'Gold'
elif df['RFMT_Score'] >=8 and df['RFMT_Score'] <=10:
return 'Silver'
else:
return 'Bronze'
rfmt['Segment_Name'] = rfmt.apply(segment_name, axis=1)
rfmt.groupby('Segment_Name').agg({'recency':'mean',
'frequency':'mean',
'monetary_value':['mean','count'],
'tenure':'mean'
}).round(1)
# + [markdown] id="wsZTVG0NWiQi"
# ## 6. K-means clustering
#
# Instead of manually segmenting and scoring using arbitrary cutoffs, let's apply K-means clustering to uncover segments with distinct purchasing behavior.
#
#
# + [markdown] id="dNjdgdKfIAuM"
# ### Box Cox Transformation
#
# K-means requires that variables have symmetric (non-skewed) distribution, the same mean, and the same variance. Let's unskew our data using Box Cox transformation and scale it to normalize our RFMT metrics with mean around 0 and st dev around 1.
# + id="eTISTUSLWqz1"
# RFMT metrics
snapshot_date = df['order_delivered_customer_date'].max() + timedelta(days=1)
rfmt = df.groupby('customer_id') \
.agg({'order_delivered_customer_date': lambda x: (snapshot_date - x.max()).days, # recency
'order_id': 'count', # frequency
'payment_value': 'sum', # monetary value
'order_purchase_timestamp': lambda x: (snapshot_date - x.min()).days # tenure
})
rfmt.dropna(inplace=True)
rfmt = rfmt.query('payment_value > 0') # keep only non-zero monetary values
rfmt.columns = ['recency','frequency','monetary_value','tenure']
# + colab={"base_uri": "https://localhost:8080/", "height": 321} id="3eJvj0gLXhhG" outputId="7d1a0ec8-e2b0-4496-feb6-611175ca8a40"
fig, axes = plt.subplots(1, 4, figsize=(16,4))
rfmt.hist('recency', bins=10, color='turquoise', ax=axes[0])
rfmt.hist('frequency', bins=3, color='salmon', ax=axes[1])
rfmt.hist('monetary_value', bins=100, color='gold',ax=axes[2])
rfmt.hist('tenure', bins=10, color='slategray',ax=axes[3])
axes[0].set_title('Recency')
axes[1].set_title('Frequency')
axes[2].set_title('Monetary Value')
axes[3].set_title('Tenure')
# + [markdown] id="IbRuFPKju7w3"
# RFM variables are skewed, particularly frequency and monetary value.
# + id="_QWlKwyqX4ei"
def boxcox_df(x):
# Since stats.boxcox returns 2 objects, this function can be applied to a dataframe and returns 1 object
x_boxcox, _ = stats.boxcox(x) # we don't care about second object
return x_boxcox
rfmt_boxcox = rfmt.apply(boxcox_df, axis=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 321} id="99ff9A_kYC0F" outputId="e05643c9-7b3b-4840-96da-1010e35ebed5"
fig, axes = plt.subplots(1, 4, figsize=(16,4))
rfmt_boxcox.hist('recency', bins=10, color='turquoise', ax=axes[0])
rfmt_boxcox.hist('frequency', bins=3, color='salmon', ax=axes[1])
rfmt_boxcox.hist('monetary_value', bins=40, color='gold',ax=axes[2])
rfmt_boxcox.hist('tenure', bins=10, color='slategray',ax=axes[3])
axes[0].set_title('Box Cox Recency')
axes[1].set_title('Box Cox Frequency')
axes[2].set_title('Box Cox Monetary Value')
axes[3].set_title('Box Cox Tenure')
# + [markdown] id="Ys6gRh0Mvfkd"
# Box cox-transformed variables are less skewed, though frequency remains skewed because most values are 1.
# + [markdown] id="hY4CcOoBI4za"
# ### Scaling
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="MtcxWJInY3ef" outputId="e42c405e-9354-4d18-9f12-352bf9d738cb"
scaler = StandardScaler()
scaler.fit(rfmt_boxcox)
rfmt_scaled = scaler.transform(rfmt_boxcox) # stored as numpy array
rfmt_scaled_df = pd.DataFrame(data=rfmt_scaled,
index=rfmt_boxcox.index,
columns=rfmt_boxcox.columns)
rfmt_scaled_df.agg(['mean','std']).round()
# + [markdown] id="eto8yAhuxLaA"
# As we expect, all means are close to 0 and all standard deviations are close to 1.
# + colab={"base_uri": "https://localhost:8080/", "height": 321} id="cRPmkBWeZMR0" outputId="5b58842e-135f-4538-d676-79bb3c5d713a"
fig, axes = plt.subplots(1, 4, figsize=(16,4))
rfmt_scaled_df.hist('recency', bins=10, color='turquoise', ax=axes[0])
rfmt_scaled_df.hist('frequency', bins=3, color='salmon', ax=axes[1])
rfmt_scaled_df.hist('monetary_value', bins=40, color='gold',ax=axes[2])
rfmt_scaled_df.hist('tenure', bins=10, color='slategray',ax=axes[3])
axes[0].set_title('Normalized Recency')
axes[1].set_title('Normalized Frequency')
axes[2].set_title('Normalized Monetary Value')
axes[3].set_title('Normalized Tenure')
# + [markdown] id="nNX3Ynr02Gbn"
# ### Optimal K
# + colab={"base_uri": "https://localhost:8080/", "height": 343} id="G5qq2_dnZgzv" outputId="6a57c3cb-70ea-49bb-a26c-e978e134b2d0"
sse={}
for k in range(1,11):
kmeans = KMeans(n_clusters=k, random_state=1111)
kmeans.fit(rfmt_scaled_df)
sse[k] = kmeans.inertia_
sns.lineplot(x=list(sse.keys()), y=list(sse.values()))
plt.title('Elbow Plot')
plt.xlabel('Number of Clusters')
plt.ylabel('Squared Standard Error')
# + [markdown] id="LBWGOt47Zzyc"
# Using the elbow method, the optimal number of clusters is 3 or 4, after which we have diminishing returns to SSE. Let's try both and compare.
# + [markdown] id="ZAZRgC1mJ-4c"
# ### Fit
# + [markdown] id="Pzi6YCsRBtVH"
# #### 3 Clusters
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="g2jT01v6Z7O-" outputId="bf03f99d-eeed-4f42-86fb-715d3f9fce5c"
kmeans = KMeans(n_clusters=3, random_state=1111)
kmeans.fit(rfmt_scaled_df)
cluster_labels = kmeans.labels_
rfmt_3_clusters = rfmt.assign(cluster = cluster_labels) # add cluster column to original rfm dataframe
rfmt_3_clusters.groupby(['cluster']) \
.agg({
'recency':'mean',
'frequency':'mean',
'monetary_value':['mean','count'],
'tenure':'mean'
}).round(0)
# + [markdown] id="X5FzKpf5Bphw"
# #### 4 Clusters
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="oPb2GfJJaP-W" outputId="58e38041-34a4-46c0-8e3b-428912691876"
kmeans = KMeans(n_clusters=4, random_state=1111)
kmeans.fit(rfmt_scaled_df)
cluster_labels = kmeans.labels_
rfmt_4_clusters = rfmt.assign(cluster = cluster_labels) # add cluster column to original rfm dataframe
rfmt_4_clusters.groupby(['cluster']) \
.agg({
'recency':'mean',
'frequency':'mean',
'monetary_value':['mean','count'],
'tenure':'mean'
}).round(0)
# + [markdown] id="S6iqFKjECsU-"
# While the 3 cluster segementation is simpler, 4 clusters provides more insight, better separating monetary values on the low and high ends. Cluster 2 stands out in size as a small group of higher frequency buyers.
#
# - Cluster 0: weak recency, highest spend, longest tenure customers
#
# - Cluster 1: strongest recency, moderate spend customers
#
# - Cluster 2: small group of higher frequency, moderate spend customers
#
# - Cluster 3: weak recency, lowest spend customers
# + [markdown] id="1GXbmwaFKTVV"
# ## 7. Segments
# + [markdown] id="KufAUWNuEYjK"
# ### Snake Plot
# + colab={"base_uri": "https://localhost:8080/", "height": 401} id="5jivuZL1auxa" outputId="57a186e4-11d3-4d71-9de1-6ec1df621b3d"
rfmt_scaled_df['cluster'] = rfmt_4_clusters['cluster']
# Melt to long format for easier plotting
rfmt_melted = pd.melt(rfmt_scaled_df.reset_index(),
id_vars = ['customer_id', 'cluster'],
value_vars = ['recency','frequency','monetary_value','tenure'],
var_name = 'Metric',
value_name = 'Value')
plt.figure(figsize=(12,5))
sns.lineplot(x='Metric', y='Value', hue='cluster', data=rfmt_melted)
plt.title('Snake Plot of Standardized RFMT')
# + [markdown] id="cbUszIIrLZif"
# The snake plot visualizes the differences between the clusters. For example, we can more easily see higher frequency Cluster 2, low monetary value Cluster 3, and high monetary value Cluster 0.
# + [markdown] id="bLnYvQQmK7-r"
# ### Relative Importance of Segment Metrics
# + [markdown] id="4lHp2Q_MMm9a"
# To understand the importance of each metric to each segment relative to the population, let's calculate relative importance. The farther the ratio is from 0, the higher the metric's importance for that segment compared to the population average.
# + colab={"base_uri": "https://localhost:8080/", "height": 263} id="8wUGFJjnbXRK" outputId="def0d6ef-55d5-419b-8145-bfe08a483ebd"
cluster_avg = rfmt_4_clusters.groupby(['cluster']).mean()
population_avg = rfmt.mean()
relative_importance = cluster_avg / population_avg - 1
plt.figure(figsize=(10,3))
sns.heatmap(data=relative_importance, annot=True, fmt='.2f', cmap='magma_r')
plt.title('Relative Importance of Metric')
# + [markdown] id="BoIv4YiePq-q"
# Recency is particulary important for Cluster 1, frequency for Cluster 2, monetary value for Cluster 0 and Cluster 3, and tenure for Cluster 1.
#
# Based on our understanding customer segment purchasing behavior, we can better tailor product, sales, and marketing activities and investments.
# + [markdown] id="9XWxEGtrKbhv"
# ## 8. References
# + [markdown] id="C3fibwUgFx51"
# https://learn.datacamp.com/courses/customer-segmentation-in-python
#
# https://www.geeksforgeeks.org/box-cox-transformation-using-python/
#
# https://en.wikipedia.org/wiki/Power_transform#Box%E2%80%93Cox_transformation
#
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.boxcox.html
| Customer_RFMT_Metrics_&_Segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Read and Write file
# !echo "Hello! Welcome to demofile.txt\nThis file is for testing purposes.\nGood Luck!\n" >> demofile.txt
f = open("demofile.txt", "r")
print(f.read())
f.close()
f = open("demofile.txt", "r")
print(f.read(5))
f.close()
f = open("demofile.txt", "r")
print(f.readline())
print(f.readline())
f.close()
f = open("demofile.txt", "r")
my_readline = f.__iter__()
next(my_readline)
f.readline()
id(f.readline())
id(my_readline)
dir(f.readline())
f = open("demofile.txt", "r")
for line in f:
print(line)
print("-------")
f.close()
# # Writing a file
# +
f = open("demofile.txt", "a")
f.write("Now the file has more content!\n")
f.close()
#open and read the file after the appending:
f = open("demofile.txt", "r")
print(f.read())
# +
f = open("demofile.txt", "w")
f.write("Woops! I have deleted the content!")
f.close()
#open and read the file after the appending:
f = open("demofile.txt", "r")
print(f.read())
# -
# # os
import os
os.getcwd()
os.listdir(os.getcwd())
os.path.abspath("./")
os.path.abspath("demotext.txt")
os.path.dirname(os.path.abspath("./demotext.txt"))
os.path.join(os.getcwd(),"demotext.txt")
# # subprocess
# !ls
import subprocess
my_process = subprocess.run("ls", capture_output=True)
my_process.stdout
# !ls -a
my_process = subprocess.run(["ls", "-a"], capture_output=True)
my_process.stdout
| 2020-09-06-python-os-subprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manual Neural Network
#
# In this notebook we will manually build out a neural network that mimics the TensorFlow API. This will greatly help your understanding when working with the real TensorFlow!
# ____
# ### Quick Note on Super() and OOP
class SimpleClass():
def __init__(self,str_input):
print("SIMPLE"+str_input)
class ExtendedClass(SimpleClass):
def __init__(self):
print('EXTENDED')
s = ExtendedClass()
class ExtendedClass(SimpleClass):
def __init__(self):
super().__init__(" My String")
print('EXTENDED')
s = ExtendedClass()
# ________
# ## Operation
class Operation():
"""
An Operation is a node in a "Graph". TensorFlow will also use this concept of a Graph.
This Operation class will be inherited by other classes that actually compute the specific
operation, such as adding or matrix multiplication.
"""
def __init__(self, input_nodes = []):
"""
Intialize an Operation
"""
self.input_nodes = input_nodes # The list of input nodes
self.output_nodes = [] # List of nodes consuming this node's output
# For every node in the input, we append this operation (self) to the list of
# the consumers of the input nodes
for node in input_nodes:
node.output_nodes.append(self)
# There will be a global default graph (TensorFlow works this way)
# We will then append this particular operation
# Append this operation to the list of operations in the currently active default graph
_default_graph.operations.append(self)
def compute(self):
"""
This is a placeholder function. It will be overwritten by the actual specific operation
that inherits from this class.
"""
pass
# ## Example Operations
#
# ### Addition
class add(Operation):
def __init__(self, x, y):
super().__init__([x, y])
def compute(self, x_var, y_var):
self.inputs = [x_var, y_var]
return x_var + y_var
# ### Multiplication
class multiply(Operation):
def __init__(self, a, b):
super().__init__([a, b])
def compute(self, a_var, b_var):
self.inputs = [a_var, b_var]
return a_var * b_var
# ### Matrix Multiplication
class matmul(Operation):
def __init__(self, a, b):
super().__init__([a, b])
def compute(self, a_mat, b_mat):
self.inputs = [a_mat, b_mat]
return a_mat.dot(b_mat)
# ## Placeholders
class Placeholder():
"""
A placeholder is a node that needs to be provided a value for computing the output in the Graph.
"""
def __init__(self):
self.output_nodes = []
_default_graph.placeholders.append(self)
# ## Variables
class Variable():
"""
This variable is a changeable parameter of the Graph.
"""
def __init__(self, initial_value = None):
self.value = initial_value
self.output_nodes = []
_default_graph.variables.append(self)
# ## Graph
class Graph():
def __init__(self):
self.operations = []
self.placeholders = []
self.variables = []
def set_as_default(self):
"""
Sets this Graph instance as the Global Default Graph
"""
global _default_graph
_default_graph = self
# ## A Basic Graph
#
# $$ z = Ax + b $$
#
# With A=10 and b=1
#
# $$ z = 10x + 1 $$
#
# Just need a placeholder for x and then once x is filled in we can solve it!
g = Graph()
g.set_as_default()
A = Variable(10)
b = Variable(1)
# Will be filled out later
x = Placeholder()
y = multiply(A,x)
z = add(y,b)
# ## Session
import numpy as np
# ### Traversing Operation Nodes
def traverse_postorder(operation):
"""
PostOrder Traversal of Nodes. Basically makes sure computations are done in
the correct order (Ax first , then Ax + b). Feel free to copy and paste this code.
It is not super important for understanding the basic fundamentals of deep learning.
"""
nodes_postorder = []
def recurse(node):
if isinstance(node, Operation):
for input_node in node.input_nodes:
recurse(input_node)
nodes_postorder.append(node)
recurse(operation)
return nodes_postorder
# +
class Session:
def run(self, operation, feed_dict = {}):
"""
operation: The operation to compute
feed_dict: Dictionary mapping placeholders to input values (the data)
"""
# Puts nodes in correct order
nodes_postorder = traverse_postorder(operation)
for node in nodes_postorder:
if type(node) == Placeholder:
node.output = feed_dict[node]
elif type(node) == Variable:
node.output = node.value
else: # Operation
node.inputs = [input_node.output for input_node in node.input_nodes]
node.output = node.compute(*node.inputs)
# Convert lists to numpy arrays
if type(node.output) == list:
node.output = np.array(node.output)
# Return the requested node value
return operation.output
# -
sess = Session()
result = sess.run(operation=z,feed_dict={x:10})
result
10*10 + 1
# ** Looks like we did it! **
# +
g = Graph()
g.set_as_default()
A = Variable([[10,20],[30,40]])
b = Variable([1,1])
x = Placeholder()
y = matmul(A,x)
z = add(y,b)
# -
sess = Session()
result = sess.run(operation=z,feed_dict={x:10})
result
# ## Activation Function
import matplotlib.pyplot as plt
# %matplotlib inline
def sigmoid(z):
return 1/(1+np.exp(-z))
sample_z = np.linspace(-10,10,100)
sample_a = sigmoid(sample_z)
plt.plot(sample_z,sample_a)
# #### Sigmoid as an Operation
class Sigmoid(Operation):
def __init__(self, z):
# a is the input node
super().__init__([z])
def compute(self, z_val):
return 1/(1+np.exp(-z_val))
# ## Classification Example
from sklearn.datasets import make_blobs
data = make_blobs(n_samples = 50,n_features=2,centers=2,random_state=75)
data
features = data[0]
plt.scatter(features[:,0],features[:,1])
labels = data[1]
plt.scatter(features[:,0],features[:,1],c=labels,cmap='coolwarm')
# DRAW A LINE THAT SEPERATES CLASSES
x = np.linspace(0,11,10)
y = -x + 5
plt.scatter(features[:,0],features[:,1],c=labels,cmap='coolwarm')
plt.plot(x,y)
# ## Defining the Perceptron
#
# $$ y = mx + b $$
#
# $$ y = -x + 5 $$
#
# $$ f1 = mf2 + b , m=1$$
#
# $$ f1 = -f2 + 5 $$
#
# $$ f1 + f2 - 5 = 0 $$
#
# ### Convert to a Matrix Representation of Features
# $$ w^Tx + b = 0 $$
#
# $$ \Big(1, 1\Big)f - 5 = 0 $$
#
# Then if the result is > 0 its label 1, if it is less than 0, it is label=0
#
#
# ### Example Point
#
# Let's say we have the point f1=2 , f2=2 otherwise stated as (8,10). Then we have:
#
# $$
# \begin{pmatrix}
# 1 , 1
# \end{pmatrix}
# \begin{pmatrix}
# 8 \\
# 10
# \end{pmatrix} + 5 = $$
np.array([1, 1]).dot(np.array([[8],[10]])) - 5
# Or if we have (4,-10)
np.array([1,1]).dot(np.array([[4],[-10]])) - 5
# ### Using an Example Session Graph
g = Graph()
g.set_as_default()
x = Placeholder()
w = Variable([1,1])
b = Variable(-5)
z = add(matmul(w,x),b)
a = Sigmoid(z)
sess = Session()
sess.run(operation=a,feed_dict={x:[8,10]})
sess.run(operation=a,feed_dict={x:[0,-10]})
# # Great Job!
| study_python/tensorflow/Tensorflow-Bootcamp-master/01-Neural-Network-Basics/Manual Neural Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Imports
from matplotlib import pyplot as plt
import cv2 as cv
import numpy as np
import os
import pickle
# Open and convert a input
# image from BGR to GRAYSCALE
image = cv.imread(filename = 'Figures/input.png',
flags = cv.IMREAD_GRAYSCALE)
# +
# FREAK is just a feature descriptor
# Initiate STAR detector
STAR = cv.xfeatures2d.StarDetector_create()
# Initiate FREAK descriptor
FREAK = cv.xfeatures2d.FREAK_create()
# +
# Find the keypoints with FREAK
keypoints = STAR.detect(image, None)
# Print number of keypoints detected
print("Number of keypoints Detected:", len(keypoints), "\n")
# +
# Save Keypoints to a file
index = []
for point in keypoints:
temp = (point.pt,
point.size,
point.angle,
point.response,
point.octave,
point.class_id)
index.append(temp)
# File name
filename = "Outputs/FREAK-keypoints.txt"
# Delete a file if it exists
if os.path.exists(filename):
os.remove(filename)
# Open a file
file = open(filename, "wb")
# Write
file.write(pickle.dumps(index))
# Close a file
file.close()
# +
# Compute the descriptors with FREAK
keypoints, descriptors = FREAK.compute(image, keypoints)
# Print the descriptor size in bytes
print("Size of Descriptor:", FREAK.descriptorSize(), "\n")
# Print the descriptor type
print("Type of Descriptor:", FREAK.descriptorType(), "\n")
# Print the default norm type
print("Default Norm Type:", FREAK.defaultNorm(), "\n")
# Print shape of descriptor
print("Shape of Descriptor:", descriptors.shape, "\n")
# -
# Draw only 50 keypoints on input image
image = cv.drawKeypoints(image = image,
keypoints = keypoints[:50],
outImage = None,
flags = cv.DRAW_MATCHES_FLAGS_DRAW_RICH_KEYPOINTS)
# +
# Plot input image
# Turn interactive plotting off
plt.ioff()
# Create a new figure
plt.figure()
plt.axis('off')
plt.imshow(image)
plt.show()
plt.imsave(fname = 'Figures/feature-detection-FREAK.png',
arr = image,
dpi = 300)
# Close it
plt.close()
| feature-detection-FREAK.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Recognizing hand-written digits
#
#
# An example showing how the scikit-learn can be used to recognize images of
# hand-written digits.
#
# This example is commented in the
# `tutorial section of the user manual <introduction>`.
#
# +
print(__doc__)
# Author: <NAME> <gael dot varoquaux at normalesup dot org>
# License: BSD 3 clause
# Standard scientific Python imports
import matplotlib.pyplot as plt
# Import datasets, classifiers and performance metrics
from sklearn import datasets, svm, metrics
from sklearn.model_selection import train_test_split
# The digits dataset
digits = datasets.load_digits()
# The data that we are interested in is made of 8x8 images of digits, let's
# have a look at the first 4 images, stored in the `images` attribute of the
# dataset. If we were working from image files, we could load them using
# matplotlib.pyplot.imread. Note that each image must have the same size. For these
# images, we know which digit they represent: it is given in the 'target' of
# the dataset.
_, axes = plt.subplots(2, 4)
images_and_labels = list(zip(digits.images, digits.target))
for ax, (image, label) in zip(axes[0, :], images_and_labels[:4]):
ax.set_axis_off()
ax.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
ax.set_title('Training: %i' % label)
# To apply a classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.images)
data = digits.images.reshape((n_samples, -1))
# Create a classifier: a support vector classifier
classifier = svm.SVC(gamma=0.001)
# Split data into train and test subsets
X_train, X_test, y_train, y_test = train_test_split(
data, digits.target, test_size=0.5, shuffle=False)
# We learn the digits on the first half of the digits
classifier.fit(X_train, y_train)
# Now predict the value of the digit on the second half:
predicted = classifier.predict(X_test)
images_and_predictions = list(zip(digits.images[n_samples // 2:], predicted))
for ax, (image, prediction) in zip(axes[1, :], images_and_predictions[:4]):
ax.set_axis_off()
ax.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
ax.set_title('Prediction: %i' % prediction)
print("Classification report for classifier %s:\n%s\n"
% (classifier, metrics.classification_report(y_test, predicted)))
disp = metrics.plot_confusion_matrix(classifier, X_test, y_test)
disp.figure_.suptitle("Confusion Matrix")
print("Confusion matrix:\n%s" % disp.confusion_matrix)
plt.show()
| sklearn/sklearn learning/demonstration/auto_examples_jupyter/classification/plot_digits_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Coding begins now!!!
# 1) Define the environment representation in the following code block
# + active=""
# #PEAS Description of the task environment here :
# Performance : minimizing travelling distance(Optimal), gettig to the correct destination(Complete), Time and Memory it use to find the solution, minimizing the total cost of search
# Environment : Roads, Cities, Other traffic
# Actuators : Steering, Accelerator, Signals, Horn, Display
# Sensors : Camera, Sonar, Speedometer, Keyboard, Engine Sensors
# -
#Environment representation goes here
#Fully observable :Agent know the complete state of the environment at each point in time
#Single agent :Single agent is solving the problem
#Deterministic :Each action has exactly one outcome
#Sequential :Each action could affect all the futute decisions
#Static :Environment is not changing while agent is deliberating
#Discrete :At any given state there are only finitely many actions to choose from
#Known :Knows which states are reached by each actions
# Define the haversine formula and what it does
# + active=""
# #haversine formula representation
# The Haversine formula calculates the shortest distance between two points on a sphere using their latitudes and longitudes measured along the surface. Haversine formula will be used to find the heuristic function of the problem.
# -
# 2) Define a function which calculates the heuristic distance from each city to the destination city in the following code block
#Function for calculating distance from each node/city to Destination
class Heuristics(object):
def __init__(self, dest):
self.data_dict = {
'Panji': [15.4909, 73.8278],
'Raichur': [16.2076, 77.3463],
'Mangalore': [12.9141, 74.8560],
'Bellari': [15.1394, 76.9214],
'Tirupati': [13.6288, 79.4192],
'Kurnool': [15.8281, 78.0373],
'Kozhikode': [11.2588, 75.7804],
'Bangalore': [12.9716, 77.5946],
'Nellore': [14.4426, 79.9865],
'Chennai': [13.0827, 80.2707]
}
self.dest = dest
def h(self, n):
LatLong1 = self.data_dict[n]
LatLong2 = self.data_dict[self.dest]
import math
if (n == self.dest):
distance = 0
else:
# distance between latitudes
# and longitudes
dLat = (LatLong2[0] - LatLong1[0]) * math.pi / 180.0
dLon = (LatLong2[1] - LatLong1[1]) * math.pi / 180.0
# convert to radians
lat1 = (LatLong1[0]) * math.pi / 180.0
lat2 = (LatLong2[0]) * math.pi / 180.0
# apply formulae
a = (pow(math.sin(dLat / 2), 2) +
pow(math.sin(dLon / 2), 2) *
math.cos(lat1) * math.cos(lat2));
rad = 6371
c = 2 * math.asin(math.sqrt(a))
# distance
distance = rad * c
return distance
# +
#Code Block 2 ..
# -
# 3) Implementation of A* Algorithm . Feel free to add code blocks for each methods needed starting here.
# Please modularize the implementation of A* and write each of them in a code block.
#Code Block 1
class Graph(object):
def __init__(self, adjacency_list):
self.adjacency_list = adjacency_list
def get_neighbors(self, v):
return self.adjacency_list[v]
#Code Block 2 ..
class AStar(object):
def __init__(self, start_node, stop_node, graph):
self.start_node = start_node
self.stop_node = stop_node
self.graph = graph
self.h_func = Heuristics(stop_node)
self.visited_node_count = 0
def a_star_algorithm(self):
open_list = set([self.start_node])
closed_list = set([])
# g contains current distances from start_node to all other nodes
# the default value (if it's not found in the map) is +infinity
g = {}
g[self.start_node] = 0
# parents contains an adjacency map of all nodes
parents = {}
parents[self.start_node] = self.start_node
while len(open_list) > 0:
n = None
# find a node with the lowest value of f() - evaluation function
for v in open_list:
if n == None or g[v] + self.h_func.h(v) < g[n] + self.h_func.h(n):
n = v;
if n == None:
print('Path does not exist!')
return None
# if the current node is the stop_node
# then we begin reconstructin the path from it to the start_node
if n == self.stop_node:
reconst_path = []
self.n_cost = g[n]
while parents[n] != n:
reconst_path.append(n)
n = parents[n]
reconst_path.append(self.start_node)
reconst_path.reverse()
self.best_path = reconst_path
return None
# for all neighbors of the current node do
for (m, weight) in self.graph.get_neighbors(n):
# if the current node isn't in both open_list and closed_list
# add it to open_list and note n as it's parent
self.visited_node_count +=1
if m not in open_list and m not in closed_list:
open_list.add(m)
parents[m] = n
g[m] = g[n] + weight
# otherwise, check if it's quicker to first visit n, then m
# and if it is, update parent data and g data
# and if the node was in the closed_list, move it to open_list
else:
if g[m] > g[n] + weight:
g[m] = g[n] + weight
parents[m] = n
if m in closed_list:
closed_list.remove(m)
open_list.add(m)
# remove n from the open_list, and add it to closed_list
# because all of his neighbors were inspected
open_list.remove(n)
closed_list.add(n)
print('Path does not exist!')
return None
# Call your main function/algorithm block in the next code block with appropriate input representation
#Computation call
adjacency_list = {
'Panji': [('Mangalore', 365), ('Bellari', 409), ('Raichur', 457)],
'Raichur': [('Panji', 457), ('Tirupati', 453), ('Kurnool', 100)],
'Mangalore': [('Panji', 365), ('Kozhikode', 233), ('Bangalore', 352)],
'Bellari': [('Panji', 409), ('Tirupati', 379), ('Bangalore', 311)],
'Tirupati': [('Raichur', 453), ('Kurnool', 340), ('Nellore', 136), ('Chennai', 153), ('Bellari', 379)],
'Kurnool': [('Raichur', 100), ('Tirupati', 340), ('Nellore', 325)],
'Kozhikode': [('Mangalore', 233), ('Bangalore', 356)],
'Bangalore': [('Mangalore', 352), ('Kozhikode', 356), ('Bellari', 311), ('Chennai', 346)],
'Nellore': [('Chennai', 175), ('Tirupati', 136), ('Kurnool', 325)],
'Chennai': [('Nellore', 175), ('Tirupati', 153), ('Bangalore', 346)]
}
graph = Graph(adjacency_list)
a_star = AStar('Panji', 'Chennai', graph)
a_star.a_star_algorithm()
# (3.1) Path taken to reach destination from Panaji
# Execute statement to retrieve the path taken here
print(a_star.best_path)
# (3.2) Cost of the path
# Execute statement to retrieve the cost of the path here
print(a_star.n_cost)
# (3.3) Total Number of nodes vistied to get this state
# Execute statement to retrieve the total number of nodes visited to get this state here
print(a_star.visited_node_count)
# <center>All the best!! Happy Coding!!Let human intelligence prevail</center>
# <center>**********************************************************</center>
| A-star/a-star.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Video Actor Synchroncy and Causality (VASC)
# ## RAEng: Measuring Responsive Caregiving Project
# ### <NAME>, 2020
# ### https://github.com/infantlab/VASC
#
# # Step 3: Analyse the data using scipy statsmodels
#
# This script correlates and compares the timeseries of wireframes for the two figures in the video `["parent", "infant"]`
#
# We start by reloading the saved parquet file containing the multi-index numpy array of all [OpenPose](https://github.com/CMU-Perceptual-Computing-Lab/openpose) data from all pairs of individuals.
#
#
# +
import sys
import os
import json
import math
import numpy as np
import pandas as pd
import pyarrow.parquet as pq
import matplotlib.pyplot as plt
# %matplotlib inline
import logging
import ipywidgets as widgets #let's us add buttons and sliders to this page.
from ipycanvas import Canvas
import vasc #a module of our own functions (found in vasc.py in this folder)
#turn on debugging
logger = logging.getLogger()
logger.setLevel(logging.INFO)
# %pdb on
# +
jupwd = os.getcwd() + "\\"
# where's the project data folder?
projectpath = "C:\\Users\\cas\\OneDrive - Goldsmiths College\\Projects\\Measuring Responsive Caregiving\\VASCTutorial"
#where are your video files?
videos_in = "C:\\Users\\cas\\OneDrive - Goldsmiths College\\Projects\\Measuring Responsive Caregiving\\VASCTutorial\\demovideos"
# locations of videos and output
videos_out = projectpath + "\\out"
videos_out_openpose = videos_out + "\\openpose"
videos_out_timeseries = videos_out + "\\timeseries"
videos_out_analyses = videos_out + "\\analyses"
# -
# ### 3.1 Load the clean data as a DataFrame
#
# Reload the clean data file created in step 2.
#retrieve the list of base names of processed videos.
try:
with open(videos_out + '\\clean.json') as json_file:
videos = json.load(json_file)
print("Existing clean.json found..")
except:
print("File clean.json not found.")
# +
print('reading parquet file:')
df = pq.read_table(videos_out_timeseries + '\\cleandata.parquet').to_pandas()
#sort the column names as this helps with indexing
df = df.sort_index(axis = 1)
print(df.head())
# -
# ## 3.2 Process the data
#
# Next we set all 0 values to as missing value `np.nan` to enable interpolation.
# Then use numpy's built in `interpolate` method.
# +
df = df.replace(0.0, np.nan)
#are we going to use all the data or a subset?
first = 0
last = 8500
df = df.truncate(before = first, after = last)
# -
df = df.interpolate()
#take a quick look
print(df.head())
df.shape
# ### 3.2.1 Mean movements
# We create a dictionary of the subsets of OpenPose coordinates we want to average and then call `mean` on the Pandas dataframe. e.g.
#
# ```
# meanpoints = {
# "headx" : [0, 3, 45, 48, 51, 54],
# "heady" : [1, 4, 46, 49, 52, 55],
# "allx" : [0, 3, 6, 9, ...],
# "ally" : [1, 4, 7, 10, ...]
# }
# ```
#
# Then we call the `vasc.averageCoordinateTimeSeries` function to average across sets of coordinates. For a given set of videos and people. For example
#
# In:
# ```
# videos = "All"
# people = "Both"
# df2 = vasc.averageCoordinateTimeSeries(df,meanpoints,videos,people)
# df2.head
# ```
#
# Out:
# ```
# person infant parent
# avgs headx heady xs ys headx
# 501 565.996600 369.840600 534.895615 398.482538 471.686200
# 502 567.231800 369.887600 534.354198 398.706552 471.849400
# 503 567.228600 370.159600 534.444328 398.678133 471.711600
# 504 566.912600 369.857000 535.369536 398.551636 472.309400
# ... ... ... ... ... ...
# ```
#
# +
meanpoints = {"head" : vasc.headxys,
"headx": vasc.headx,
"heady": vasc.heady,
"arms" : vasc.armsxys,
"armsx": vasc.armsx,
"armsy": vasc.armsy,
"all" : vasc.xys,
"allx" : vasc.xs,
"ally" : vasc.ys}
vids = "All"
people = ["infant","parent"]
#average across the points in each group (all points of head etc. )
avgdf = vasc.averageCoordinateTimeSeries(df,meanpoints,vids,people)
# -
avgdf.head
# ### 3.2.2 Rolling window of movements
#
# One thing we'd like to know is if mothers move in response to infants. The raw time series are probably too noisy to tell us this so instead we can look at few alternatives
#
# 1. **Smoothed** - if we average the signal over a short rolling window we smooth out any high-frequency jitter.
# 2. **Variance** - the variance of movement over a short rolling window. First we apply 2 second long (50 frame) rolling window to each coordinate of the body and use the stddev or variance function `std()` or `var()` . Then we take averages as in the step above. However, this time we combine x and y coordinates as this is now a movement index.
#
#
#
# +
win = 50 #2 seconds
halfwin = math.floor(win/2)
smoothdf = df.rolling(window = 5).mean()
smoothdf = smoothdf.truncate(before = first, after = last)
vardf = df.rolling(window = win, min_periods = halfwin).var()
vardf = vardf.truncate(before = first + 50, after = last) # cut out the empty bits at the start
smoothdf = vasc.averageCoordinateTimeSeries(smoothdf,meanpoints,vids,people)
vardf = vasc.averageCoordinateTimeSeries(vardf,meanpoints,vids,people)
# -
# Let's create a widget to plot some graphs of the data
# +
vidlist = [] #used to fill dropdown options
for vid in videos:
vidlist.append(vid)
pickvid = widgets.Dropdown(
options= vidlist,
value= vidlist[0],
description='Subject:'
)
features = []
for f in meanpoints:
features.append(f)
pickfeature = widgets.Dropdown(
options= features,
value= features[0],
description='Feature:'
)
linetypes = ["Mean point", "Smoothed Mean (5 frames)","Variance over 2 secs"]
picktype = widgets.Dropdown(
options= linetypes,
value= linetypes[0],
description='Line type:'
)
def pickvid_change(change):
if change['name'] == 'value' and (change['new'] != change['old']):
updateAll(True)
def pickfeature_change(change):
if change['name'] == 'value' and (change['new'] != change['old']):
updateAll(True)
def picktype_change(change):
if change['name'] == 'value' and (change['new'] != change['old']):
updateAll(True)
pickvid.observe(pickvid_change, 'value')
pickfeature.observe(pickfeature_change, 'value')
picktype.observe(picktype_change, 'value')
button_update = widgets.Button(description="Redraw")
output = widgets.Output()
def drawGraphs(vid, feature, linetype):
"""Plot input signals"""
plt.ion()
f,ax=plt.subplots(4,1,figsize=(14,10),sharex=True)
ax[0].set_title('Infant')
ax[1].set_title('Parent')
ax[1].set_xlabel('Frames')
who = ["infant","parent"]
if linetype == linetypes[0]:
usedf = avgdf
elif linetype == linetypes[1]:
usedf = smoothdf
else:
usedf = vardf
#to select a single column..
infant = usedf[(vid, people[0], feature)].to_frame()
parent = usedf[(vid, people[1], feature)].to_frame()
n = np.arange(usedf.shape[0])
#selecting multiple columns slightly messier
#infant = df3.loc[50:,(vid, part[0], ('head','arms', 'all'))]
#parent = df3.loc[50:,(vid, part[1], ('head','arms', 'all'))]
ax[0].plot(n,infant)
ax[1].plot(n,parent, color='b')
#calculate the correlations in a shorter rolling window
r_window_size = 120
rolling_r = usedf[(vid, who[0], feature)].rolling(window=r_window_size, center=True).corr(vardf[(vid, who[1], feature)])
usedf.loc[:,(vid, slice(None), feature)].plot(ax=ax[2])
ax[2].set(xlabel='Frame',ylabel='Movement index for parent and infant')
rolling_r.plot(ax=ax[3])
ax[3].set(xlabel='Frame',ylabel='Pearson r')
ax[3].set_title("Local correlation with rolling window size " + str(r_window_size))
plt.show()
def updateAll(forceUpdate = False):
output.clear_output(wait = True)
if forceUpdate:
logging.debug('forceUpdate')
#slider.value = 0
#slider.max = videos[pickvid.value][pickcam.value]["end"]
with output:
display(pickvid,pickfeature,picktype,button_update)
drawGraphs(pickvid.value,pickfeature.value,picktype.value)
#draw everything for first time
updateAll(True)
output
# -
# ### 3.3 Movement analysis
#
# First we run some simple correlations between the mother and infant.
infant = vardf[(vid, people[0], 'head')].to_frame()
infant.head
print(type(infant))
#vid = "SS003"
vardf[(vid, people[0], 'head')].corr(vardf[(vid, people[1], 'head')])
who = ["infant","parent"]
parts = ["head","arms","all"]
results = pd.DataFrame(columns = ("corrHead","lagHead","corrArms","lagArms","corrAll","lagAll","DyadSynScore"),
index = videos)
#loop through colculate for each pair
for vid in videos:
thisrow = []
for part in parts:
#to select a single column..
pearson = vardf[(vid, people[0], part)].corr(vardf[(vid, people[1], part)])
thisrow.append(pearson) #this is for correlation
thisrow.append(None) #this is for maximum lag
thisrow.append(None) #don't have DyadSynScore yet
results.loc[vid] = thisrow
#take a quick look
results
# ## 3.4 Comparing to human coding.
#
# We have a spreadsheet of syhnchrony scores for each parent infant dyad. Here we see if we can find a measure that correlates with the human scores.
#
# First, load up the spreadsheet..
# +
excelpath = projectpath + "\\SS_CARE.xlsx"
filename, file_format = os.path.splitext(excelpath)
if file_format and file_format == 'xls':
# use default reader
videolist = pd.read_excel(excelpath)
else:
#since dec 2020 read_excel no longer supports xlsx (!?) so need to use openpyxl like so..
videolist = pd.read_excel(excelpath, engine = "openpyxl")
videolist = videolist.set_index("subject")
# -
#take a quick look
videolist
# #copy the dyad syncrhony and maternal sensitivity scores into our data frame.
results["DyadSynScore"] = videolist["DyadSyn"]
results["MatSensScore"] = videolist["MatSens"]
#take a quick look
results
#scatter plots of these results.
plt.scatter(results["DyadSynScore"],results["corrArms"], )
plt.title("Correlation between expert rated synchrony and time series correlations")
plt.xlabel("Dyad Synchroncy Score")
plt.ylabel("Dyad Correlation")
plt.show()
#
rolling_r.mean()
# So
# +
d1 = vardf[(vid, who[0], parts[0])]
d2 = vardf[(vid, who[1], parts[0])]
seconds = 5
fps = 25
wholeads = who[0] + 'leads <> ' + who[1] + ' leads'
rs = [vasc.crosscorr(d1,d2, lag) for lag in range(-int(seconds*fps-1),int(seconds*fps))]
offset = np.ceil(len(rs)/2)-np.argmax(rs)
f,ax=plt.subplots(figsize=(14,3))
ax.plot(rs)
ax.axvline(np.ceil(len(rs)/2),color='k',linestyle='--',label='Center')
ax.axvline(np.argmax(rs),color='r',linestyle='--',label='Peak synchrony')
ax.set(title=f'Offset = {offset} frames\n' + wholeads,ylim=[.0,1],xlim=[0,300], xlabel='Offset',ylabel='Pearson r')
ax.set_xticklabels([int(item-150) for item in ax.get_xticks()]);
plt.legend()
# -
# ## 3.4 Granger Causality
#
# The next thing to look at is if the movements of the infant predict the movements of the parent. This would suggest parent is responding to the infant.
#
# +
https://towardsdatascience.com/granger-causality-and-vector-auto-regressive-model-for-time-series-forecasting-3226a64889a6
https://www.machinelearningplus.com/time-series/time-series-analysis-python/
https://towardsdatascience.com/four-ways-to-quantify-synchrony-between-time-series-data-b99136c4a9c9
# -
| Step3.AnalyseData.scipy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/MIT-LCP/bidmc-datathon/blob/master/02_severity_of_illness.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="y4AOVdliM8gm"
# # eICU Collaborative Research Database
#
# # Notebook 2: Severity of illness
#
# This notebook introduces high level admission details relating to a single patient stay, using the following tables:
#
# - patient
# - admissiondx
# - apacheapsvar
# - apachepredvar
# - apachepatientresult
#
#
# + [markdown] colab_type="text" id="e0lUnIkYOyv4"
# ## Load libraries and connect to the database
# + colab={} colab_type="code" id="SJ6l1i3fOL4j"
# Import libraries
import numpy as np
import os
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.path as path
# Make pandas dataframes prettier
from IPython.display import display, HTML
# Access data using Google BigQuery.
from google.colab import auth
from google.cloud import bigquery
# + colab={} colab_type="code" id="TE4JYS8aO-69"
# authenticate
auth.authenticate_user()
# + colab={} colab_type="code" id="oVavf-ujPOAv"
# Set up environment variables
project_id='bidmc-datathon'
os.environ["GOOGLE_CLOUD_PROJECT"]=project_id
# + [markdown] colab_type="text" id="a1CAI3GjQYE0"
# ## Selecting a single patient stay¶
#
# As we have seen, the patient table includes general information about the patient admissions (for example, demographics, admission and discharge details). See: http://eicu-crd.mit.edu/eicutables/patient/
#
# ## Questions
#
# Use your knowledge from the previous notebook and the online documentation (http://eicu-crd.mit.edu/) to answer the following questions:
#
# - Which column in the patient table is distinct for each stay in the ICU (similar to `icustay_id` in MIMIC-III)?
# - Which column is unique for each patient (similar to `subject_id` in MIMIC-III)?
# + colab={} colab_type="code" id="R6huFICkSQAd"
# view distinct ids
# %%bigquery
SELECT DISTINCT(patientunitstayid)
FROM `physionet-data.eicu_crd_demo.patient`
# + colab={} colab_type="code" id="yEBIFRBqRo4y"
# set the where clause to select the stay of interest
# %%bigquery patient
SELECT *
FROM `physionet-data.eicu_crd_demo.patient`
WHERE patientunitstayid = <your_id_here>
# + colab={} colab_type="code" id="LjIL2XR6TAyp"
patient
# + [markdown] colab_type="text" id="QSbKYqF0TQ1n"
# ## Questions
#
# - Which type of unit was the patient admitted to? Hint: Try `patient['unittype']` or `patient.unittype`
# - What year was the patient discharged from the ICU? Hint: You can view the table columns with `patient.columns`
# - What was the status of the patient upon discharge from the unit?
# + [markdown] colab_type="text" id="izaH0XwwUxDD"
# ## The admissiondx table
#
# The `admissiondx` table contains the primary diagnosis for admission to the ICU according to the APACHE scoring criteria. For more detail, see: http://eicu-crd.mit.edu/eicutables/admissiondx/
# + colab={} colab_type="code" id="dlj3UCDTTEjj"
# set the where clause to select the stay of interest
# %%bigquery admissiondx
SELECT *
FROM `physionet-data.eicu_crd_demo.admissiondx`
WHERE patientunitstayid = <your_id_here>
# + colab={} colab_type="code" id="3wdEHFLJVMKm"
# View the columns in this data
admissiondx.columns
# + colab={} colab_type="code" id="tbOA44lAVNLr"
# View the data
admissiondx.head()
# + colab={} colab_type="code" id="Hc0y4ueOVWOk"
# Set the display options to avoid truncating the text
pd.set_option('display.max_colwidth', -1)
admissiondx.admitdxpath
# + [markdown] colab_type="text" id="mSb_BrgvWDdD"
# ## Questions
#
# - What was the primary reason for admission?
# - How soon after admission to the ICU was the diagnoses recorded in eCareManager? Hint: The `offset` columns indicate the time in minutes after admission to the ICU.
# + [markdown] colab_type="text" id="rd3Tw6_kWwlS"
# ## The apacheapsvar table
#
# The apacheapsvar table contains the variables used to calculate the Acute Physiology Score (APS) III for patients. APS-III is an established method of summarizing patient severity of illness on admission to the ICU, taking the "worst" observations for a patient in a 24 hour period.
#
# The score is part of the Acute Physiology Age Chronic Health Evaluation (APACHE) system of equations for predicting outcomes for ICU patients. See: http://eicu-crd.mit.edu/eicutables/apacheApsVar/
# + colab={} colab_type="code" id="fXOzR5XWVdNa"
# set the where clause to select the stay of interest
# %%bigquery apacheapsvar
SELECT *
FROM `physionet-data.eicu_crd_demo.apacheapsvar`
WHERE patientunitstayid = <your_id_here>
# + colab={} colab_type="code" id="mL_lVORdXDIg"
apacheapsvar.head()
# + [markdown] colab_type="text" id="8x_Z8q4jXH7D"
# ## Questions
#
# - What was the 'worst' heart rate recorded for the patient during the scoring period?
# - Was the patient oriented and able to converse normally on the day of admission? (hint: the verbal element refers to the Glasgow Coma Scale).
# + [markdown] colab_type="text" id="XplJvhIYX432"
# # apachepredvar table
#
# The apachepredvar table provides variables underlying the APACHE predictions. Acute Physiology Age Chronic Health Evaluation (APACHE) consists of a groups of equations used for predicting outcomes in critically ill patients. See: http://eicu-crd.mit.edu/eicutables/apachePredVar/
# + colab={} colab_type="code" id="iAIFESy9XFhC"
# set the where clause to select the stay of interest
# %%bigquery apachepredvar
SELECT *
FROM `physionet-data.eicu_crd_demo.apachepredvar`
WHERE patientunitstayid = <your_id_here>
# + colab={} colab_type="code" id="LAu7G72cYEY1"
apachepredvar.columns
# + [markdown] colab_type="text" id="IEaS6L9OY0vJ"
# ## Questions
#
# - Was the patient ventilated during (APACHE) day 1 of their stay?
# - Is the patient recorded as having diabetes?
# + [markdown] colab_type="text" id="nrTEkjxqZD2l"
# # `apachepatientresult` table
#
# The `apachepatientresult` table provides predictions made by the APACHE score (versions IV and IVa), including probability of mortality, length of stay, and ventilation days. See: http://eicu-crd.mit.edu/eicutables/apachePatientResult/
# + colab={} colab_type="code" id="M2RCJNBgZOJ2"
# set the where clause to select the stay of interest
# %%bigquery apachepatientresult
SELECT *
FROM `physionet-data.eicu_crd_demo.apachepatientresult`
WHERE patientunitstayid = <your_id_here>
# + colab={} colab_type="code" id="4whVaOP1Za8f"
apachepatientresult
# + [markdown] colab_type="text" id="5YO_GQcNZUWR"
# ## Questions
#
# - What versions of the APACHE score are computed?
# - How many days during the stay was the patient ventilated?
# - How long was the patient predicted to stay in hospital?
# - Was this prediction close to the truth?
| 02_severity_of_illness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# < [Energies and the PyMOL Mover](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/03.03-Energies-and-the-PyMOLMover.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Basic Folding Algorithm](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/04.01-Basic-Folding-Algorithm.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/04.00-Introduction-to-Folding.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # Introduction to Folding
# Any folding algorithm requires…
#
#
# - …a search strategy, an algorithm to generate many candidate structures (or decoys) and…
#
#
# - …a scoring function to discriminate near-native structures from all the others.
#
#
# In this workshop you will write your own Monte Carlo protein folding algorithm from scratch, and we will explore a couple of the tricks used by Simons et al. (1997, 1999) to speed up the folding search.
#
#
# ## Suggested Readings
# 1. <NAME> et al., “Assembly of Protein Structures from Fragments,” *J. Mol. Biol.*
# 268, 209-225 (1997).
# 2. <NAME> et al., “Improved recognition of protein structures,” *Proteins* 34, 82-95
# (1999).
# 3. Chapter 4 (Monte Carlo methods) of <NAME> & <NAME>, *Computer
# Simulation of Liquids*, Oxford University Press, 1989.
# **Chapter contributors:**
#
# - <NAME> (Johns Hopkins University); this chapter was adapted from the [PyRosetta book](https://www.amazon.com/PyRosetta-Interactive-Platform-Structure-Prediction-ebook/dp/B01N21DRY8) (<NAME>, <NAME>, <NAME>, <NAME>).
# <!--NAVIGATION-->
# < [Energies and the PyMOL Mover](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/03.03-Energies-and-the-PyMOLMover.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Basic Folding Algorithm](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/04.01-Basic-Folding-Algorithm.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/04.00-Introduction-to-Folding.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
| notebooks/04.00-Introduction-to-Folding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import packages
import pandas as pd
import numpy as np
#to plot within notebook
import matplotlib.pyplot as plt
# %matplotlib inline
#setting figure size
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 20,10
#for normalizing data
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=(0, 1))
#read the file
df = pd.read_csv('msft.csv')
df.columns = ['Date', 'Open', 'High', 'Low', 'Close', "Volume"]
#print the head
df.head()
# +
#setting index as date
df['Date'] = pd.to_datetime(df.Date,format='%Y-%m-%d')
df.index = df['Date']
#plot
plt.figure(figsize=(16,8))
plt.plot(df['Close'], label='Close Price history')
# +
#creating dataframe with date and the target variable
data = df.sort_index(ascending=True, axis=0)
new_data = pd.DataFrame(index=range(0,len(df)),columns=['Date', 'Close'])
for i in range(0,len(data)):
new_data['Date'][i] = data['Date'][i]
new_data['Close'][i] = data['Close'][i]
# +
train = new_data[:-1000]
valid = new_data[-1000:]
train.shape
# +
preds = []
for i in range(0,valid.shape[0]):
a = train['Close'][len(train)-1000+i:].sum() + sum(preds)
b = a/1000
preds.append(b)
# checking the results (RMSE value)
rms=np.sqrt(np.mean(np.power((np.array(valid['Close'])-preds),2)))
print('\n RMSE value on validation set:')
print(rms)
# -
#creating a separate dataset
new_data = pd.DataFrame(index=range(0,len(df)),columns=['Date', 'Close', 'Year', 'Month', 'Day',
'WeekOfYr', 'DayOfYr', 'DayOfWk',
'StartOfYr', 'EndOfYr', 'StartOfQtr', 'EndOfQtr',
'StartOfMth', 'EndOfMth', 'StartOfWk', 'EndOfWk'])
for i in range(0,len(data)):
new_data['Date'][i] = data['Date'][i]
new_data['Close'][i] = data['Close'][i]
# +
import calendar
import datetime
def is_month_start_end(date):
last_day_of_mth = calendar.monthrange(date.year, date.month)[1]
return (date.day == 1, date.day == last_day_of_mth)
def is_quarter_start_end(date):
start_qtr_mths = [1, 4, 7, 10]
end_qtr_mths = [3, 6, 9, 13]
# Check for start of quarter
qtr_start = (date.month in start_qtr_mths) and (date.day == 1)
# Check for end of quarter
qtr_end = (date.month in end_qtr_mths) and (date.day == calendar.monthrange(date.year, date.month)[1])
return (qtr_start, qtr_end)
def is_year_start_end(date):
yr_start = (date.day == 1) and (date.month == 1)
yr_end = (date.day == 31) and (date.month == 12)
return (yr_start, yr_end)
def get_date_features(date):
year = date.year
month = date.month
day = date.day
(year, wkOfYr, dayOfWk) = datetime.date(year, month, day).isocalendar()
dayOfYr = datetime.date(year, month, day).timetuple().tm_yday
return year, month, day, wkOfYr, dayOfYr, dayOfWk, is_year_start_end(date), is_quarter_start_end(date), is_month_start_end(date), ((dayOfWk == 1), (dayOfWk == 5))
# -
get_date_features(datetime.date(2020, 6, 3))
# +
import datetime
for i in range(0,len(new_data)):
features = get_date_features(new_data['Date'][i])
new_data['Year'][i] = features[0]
new_data['Month'][i] = features[1]
new_data['Day'][i] = features[2]
new_data['WeekOfYr'][i] = features[3]
new_data['DayOfYr'][i] = features[4]
new_data['DayOfWk'][i] = features[5]
new_data['StartOfYr'][i] = features[6][0]
new_data['EndOfYr'][i] = features[6][1]
new_data['StartOfQtr'][i] = features[7][0]
new_data['EndOfQtr'][i] = features[7][1]
new_data['StartOfMth'][i] = features[8][0]
new_data['EndOfMth'][i] = features[8][1]
new_data['StartOfWk'][i] = features[9][0]
new_data['EndOfWk'][i] = features[9][1]
# -
new_data.head(10)
# +
new_data = new_data.drop('Date', axis=1)
#split into train and validation
train = new_data[:-1000]
valid = new_data[-1000:]
x_train = train.drop('Close', axis=1)
y_train = train['Close']
x_valid = valid.drop('Close', axis=1)
y_valid = valid['Close']
# +
#implement linear regression
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(x_train,y_train)
preds = model.predict(x_valid)
rms=np.sqrt(np.mean(np.power((np.array(y_valid)-np.array(preds)),2)))
rms
# +
#plot
valid['Predictions'] = 0
valid['Predictions'] = preds
valid.index = new_data[-1000:].index
train.index = new_data[:-1000].index
plt.plot(train['Close'])
plt.plot(valid[['Close', 'Predictions']])
# +
#importing libraries
from sklearn import neighbors
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=(0, 1))
new_data.drop('Date', axis=1, inplace=True)
#split into train and validation
train = new_data[:-1000]
valid = new_data[-1000:]
# +
x_train = train.drop('Close', axis=1)
y_train = train['Close']
x_valid = valid.drop('Close', axis=1)
y_valid = valid['Close']
# -
train
# +
#scaling data
x_train_scaled = scaler.fit_transform(x_train)
x_train = pd.DataFrame(x_train_scaled)
x_valid_scaled = scaler.fit_transform(x_valid)
x_valid = pd.DataFrame(x_valid_scaled)
# -
x_valid.shape
# +
#using gridsearch to find the best parameter
params = {'n_neighbors':[2,3,4,5,6,7,8,9]}
knn = neighbors.KNeighborsRegressor()
model = GridSearchCV(knn, params, cv=5)
#fit the model and make predictions
model.fit(x_train,y_train)
preds = model.predict(x_valid)
# -
#rmse
rms=np.sqrt(np.mean(np.power((np.array(y_valid)-np.array(preds)),2)))
rms
#plot
valid['Predictions'] = 0
valid['Predictions'] = preds
plt.plot(valid[['Close', 'Predictions']])
plt.plot(train['Close'])
# +
from pmdarima.arima import auto_arima
data = df.sort_index(ascending=True, axis=0)
train = data[:-1000]
valid = data[-1000:]
training = train['Close']
validation = valid['Close']
model = auto_arima(training, start_p=1, start_q=1,max_p=3, max_q=3, m=12,start_P=0, seasonal=True,d=1, D=1, trace=True,error_action='ignore',suppress_warnings=True)
model.fit(training)
forecast = model.predict(n_periods=1000)
forecast = pd.DataFrame(forecast,index = valid.index,columns=['Prediction'])
# -
rms=np.sqrt(np.mean(np.power((np.array(valid['Close'])-np.array(forecast['Prediction'])),2)))
rms
| jupyter/timeseries-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37
# language: python
# name: py37
# ---
import sys
# TODO: clean this up
sys.path.insert(0, "/Users/danieldubovski/projects/deep_query_optimization/")
# sys.path
from dqo.datasets import QueriesDataset, ExtendedQueriesDataset
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
ds = ExtendedQueriesDataset("imdb:extended")
db = ds.schema()
df = ds.load()
df = ds.df.query('runtime > 0.01')
df.runtime.describe()
len(df['query'].unique()), len(df)
df = df.drop_duplicates('query')
# ---
df = ds.groom()
# ----
len(df.query('runtime > 0.01 and runtime < 500')), len(df)
df['bucket'] = df.runtime.apply(np.log2).apply(np.round).apply(lambda x: max(0, x)).apply(lambda x: min(x,8)).astype(int)
plt.hist(df['bucket'])
df.groupby('bucket').count()
mean_count = int(df.groupby('bucket').count().mean().head(1)[0])
mean_count
# cap to limit variance
df = df.groupby('bucket').head(mean_count * 4)
ds.df = df
df = ds.groom()
len(ds.df), len(df)
ds.input_path
df.query('nodes > 125').count()
df = df.query('nodes < 125')
ds.df = df
df.columns
ds.save(split=True)
df.groupby('nodes').count()
plt.hist(ds.df['nodes'])
# ___
# # augment
# ---
bkts = sorted(list(ds.df[ds.df['bucket'] > 0].groupby('bucket').count().runtime))
min_bucket = bkts[1]
min_bucket
sample = df[df['bucket'] > 0].groupby('bucket').head(min_bucket)
buckets = sample.runtime.apply(np.log2).apply(lambda x: min(x, 8)).apply(lambda x: max(0, x))
plt.hist(buckets)
import os
batch_size = 1000
_df = sample
for i in range(0,len(_df), batch_size):
aug_df = ds.augment(_df[i:i+batch_size])
f_name = os.path.join(ds.input_path, f'aug_{batch_size}_{i}:{i + len(aug_df)}')
print(i, len(aug_df), f_name)
aug_df.to_csv(f_name, header=False, index=False, columns=['query', 'runtime'])
len(aug_df)
ds.df.to_csv(os.path.join(ds.input_path, 'clean.csv'), header=False, index=False, columns=['query', 'runtime'])
buckets = aug_df.runtime.apply(np.log2).apply(lambda x: min(x, 8)).apply(lambda x: max(0, x))
plt.hist(buckets)
ds.save()
df['partition'] = df.runtime.apply(np.log2).apply(lambda x: min(x, 8)).apply(lambda x: max(0, x))
slow_df = df[df['partition'] > 4]
len(slow_df)
import os
os.getcwd()
with open('/Users/danieldubovski/projects/deep_query_optimization/dqo/localhost_imdb.qcp', 'w+') as dcp_file:
for idx, row in slow_df.iterrows():
dcp_file.write(f"{row['query']}\n")
from dqo.relational import SQLParser
from tqdm import tqdm
# +
queries = []
for idx, row in tqdm(ds.df.iterrows(), total=ds.df.shape[0]):
queries.append(SQLParser.to_query(row['query']))
# -
db
# +
from collections import defaultdict, Counter
tables = defaultdict(int)
for query in queries:
c = Counter([r.name for r in query._relations])
for k, v in c.items():
tables[k] += v
tables_df = pd.DataFrame(tables.items(), columns=['table', 'count'])
tables_df
# -
# # SAVE
ds.df = df
ds.save(split=True)
len(df)
| dqo/datasets/groom.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href='http://www.holoviews.org'><img src="assets/hv+bk.png" alt="HV+BK logos" width="40%;" align="left"/></a>
# <div style="float:right;"><h2>08. Deploying Bokeh Apps</h2></div>
# In the previous sections we discovered how to use a ``HoloMap`` to build a Jupyter notebook with interactive visualizations that can be exported to a standalone HTML file, as well as how to use ``DynamicMap`` and ``Streams`` to set up dynamic interactivity backed by the Jupyter Python kernel. However, frequently we want to package our visualization or dashboard for wider distribution, backed by Python but run outside of the notebook environment. Bokeh Server provides a flexible and scalable architecture to deploy complex interactive visualizations and dashboards, integrating seamlessly with Bokeh and with HoloViews.
#
# For a detailed background on Bokeh Server see [the bokeh user guide](http://bokeh.pydata.org/en/latest/docs/user_guide/server.html). In this tutorial we will discover how to deploy the visualizations we have created so far as a standalone bokeh server app, and how to flexibly combine HoloViews and Bokeh APIs to build highly customized apps. We will also reuse a lot of what we have learned so far---loading large, tabular datasets, applying datashader operations to them, and adding linked streams to our app.
# ## A simple bokeh app
#
# The preceding sections of this tutorial focused solely on the Jupyter notebook, but now let's look at a bare Python script that can be deployed using Bokeh Server:
with open('./apps/server_app.py', 'r') as f:
print(f.read())
# Of the three parts of this app, part 2 should be very familiar by now -- load some taxi dropoff locations, declare a Points object, datashade them, and set some plot options.
#
# Step 1 is new: Instead of loading the bokeh extension using ``hv.extension('bokeh')``, we get a direct handle on a bokeh renderer using the ``hv.renderer`` function. This has to be done at the top of the script, to be sure that options declared are passed to the Bokeh renderer.
#
# Step 3 is also new: instead of typing ``app`` to see the visualization as we would in the notebook, here we create a Bokeh document from it by passing the HoloViews object to the ``renderer.server_doc`` method.
#
# Steps 1 and 3 are essentially boilerplate, so you can now use this simple skeleton to turn any HoloViews object into a fully functional, deployable Bokeh app!
#
# ## Deploying the app
#
# Assuming that you have a terminal window open with the ``hvtutorial`` environment activated, in the ``notebooks/`` directory, you can launch this app using Bokeh Server:
#
# ```
# bokeh serve --show apps/server_app.py
# ```
#
# If you don't already have a favorite way to get a terminal, one way is to [open it from within Jupyter](../terminals/1), then make sure you are in the ``notebooks`` directory, and activate the environment using ``source activate hvtutorial`` (or ``activate tutorial`` on Windows). You can also [open the app script file](../edit/apps/server_app.py) in the inbuilt text editor, or you can use your own preferred editor.
# Exercise: Modify the app to display the pickup locations and add a tilesource, then run the app with bokeh serve
# Tip: Refer to the previous notebook
# ## Iteratively building a bokeh app in the notebook
#
# The above app script can be built entirely without using Jupyter, though we displayed it here using Jupyter for convenience in the tutorial. Jupyter notebooks are also often helpful when initially developing such apps, allowing you to quickly iterate over visualizations in the notebook, deploying it as a standalone app only once we are happy with it.
#
# To illustrate this process, let's quickly go through such a workflow. As before we will set up our imports, load the extension, and load the taxi dataset:
# +
import holoviews as hv
import geoviews as gv
import dask.dataframe as dd
from holoviews.operation.datashader import datashade, aggregate, shade
from bokeh.models import WMTSTileSource
hv.extension('bokeh', logo=False)
usecols = ['tpep_pickup_datetime', 'dropoff_x', 'dropoff_y']
ddf = dd.read_csv('../data/nyc_taxi.csv', parse_dates=['tpep_pickup_datetime'], usecols=usecols)
ddf['hour'] = ddf.tpep_pickup_datetime.dt.hour
ddf = ddf.persist()
# -
# Next we define a ``Counter`` stream which we will use to select taxi trips by hour.
# +
stream = hv.streams.Counter()
points = hv.Points(ddf, kdims=['dropoff_x', 'dropoff_y'])
dmap = hv.DynamicMap(lambda counter: points.select(hour=counter%24).relabel('Hour: %s' % (counter % 24)),
streams=[stream])
shaded = datashade(dmap)
hv.opts('RGB [width=800, height=600, xaxis=None, yaxis=None]')
url = 'https://server.arcgisonline.com/ArcGIS/rest/services/World_Imagery/MapServer/tile/{Z}/{Y}/{X}.jpg'
wmts = gv.WMTS(WMTSTileSource(url=url))
overlay = wmts * shaded
# -
# Up to this point, we have a normal HoloViews notebook that we could display using Jupyter's rich display of ``overlay``, as we would with an any notebook. But having come up with the objects we want interactively in this way, we can now display the result as a Bokeh app, without leaving the notebook. To do that, first edit the following cell to change "8888" to whatever port your jupyter session is using, in case your URL bar doesn't say "localhost:8888/".
#
# Then run this cell to launch the Bokeh app within this notebook:
renderer = hv.renderer('bokeh')
server = renderer.app(overlay, show=True, websocket_origin='localhost:8888')
# We could stop here, having launched an app, but so far the app will work just the same as in the normal Jupyter notebook, responding to user inputs as they occur. Having defined a ``Counter`` stream above, let's go one step further and add a series of periodic events that will let the visualization play on its own even without any user input:
dmap.periodic(1)
# You can stop this ongoing process by clearing the cell displaying the app.
#
# Now let's open the [text editor](../edit/apps/periodic_app.py) again and make this edit to a separate app, which we can then launch using Bokeh Server separately from this notebook.
# Exercise: Copy the example above into periodic_app.py and modify it so it can be run with bokeh serve
# Hint: Use hv.renderer and renderer.server_doc
# Note that you have to run periodic **after** creating the bokeh document
# ## Combining HoloViews with bokeh models
# Now for a last hurrah let's put everything we have learned to good use and create a bokeh app with it. This time we will go straight to a [Python script containing the app](../edit/apps/player_app.py). If you run the app with ``bokeh serve --show ./apps/player_app.py`` from [your terminal](../terminals/1) you should see something like this:
#
# <img src="./assets/tutorial_app.gif"></img>
# This more complex app consists of several components:
#
# 1. A datashaded plot of points for the indicated hour of the daty (in the slider widget)
# 2. A linked ``PointerX`` stream, to compute a cross-section
# 3. A set of custom bokeh widgets linked to the hour-of-day stream
#
# We have already covered 1. and 2. so we will focus on 3., which shows how easily we can combine a HoloViews plot with custom Bokeh models. We will not look at the precise widgets in too much detail, instead let's have a quick look at the callback defined for slider widget updates:
#
# ```python
# def slider_update(attrname, old, new):
# stream.event(hour=new)
# ```
#
# Whenever the slider value changes this will trigger a stream event updating our plots. The second part is how we combine HoloViews objects and Bokeh models into a single layout we can display. Once again we can use the renderer to convert the HoloViews object into something we can display with Bokeh:
#
# ```python
# renderer = hv.renderer('bokeh')
# plot = renderer.get_plot(hvobj, doc=curdoc())
# ```
#
# The ``plot`` instance here has a ``state`` attribute that represents the actual Bokeh model, which means we can combine it into a Bokeh layout just like any other Bokeh model:
#
# ```python
# layout = layout([[plot.state], [slider, button]], sizing_mode='fixed')
# curdoc().add_root(layout)
# ```
# Advanced Exercise: Add a histogram to the bokeh layout next to the datashaded plot
# Hint: Declare the histogram like this: hv.operation.histogram(aggregated, bin_range=(0, 20))
# then use renderer.get_plot and hist_plot.state and add it to the layout
# # Onwards
#
# Although the code above is more complex than in previous sections, it's actually providing a huge range of custom types of interactivity, which if implemented in Bokeh alone would have required far more than a notebook cell of code. Hopefully it is clear that arbitrarily complex collections of visualizations and interactive controls can be built from the components provided by HoloViews, allowing you to make simple analyses very easily and making it practical to make even quite complex apps when needed. The [user guide](http://holoviews.org/user_guide), [gallery](http://holoviews.org/gallery/index.html), and [reference gallery](http://holoviews.org/reference) should have all the information you need to get started with all this power on your own datasets and tasks. Good luck!
| notebooks/08-deploying-bokeh-apps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy import signal
from scipy import fftpack
# from matplotlib import pyplot as plt
# from crawlab_toolbox import plotting
import pandas as pd
import os
import sys
# insert at 1, 0 is the script path (or '' in REPL)
sys.path.insert(1, '../dependencies/')
from plotting import *
# -
sklearn = pd.read_csv('data/Scikit_learn_citations.csv')
tensorflow = pd.read_csv('data/Tensorflow_citations.csv')
# +
bar_plot(sklearn.Year.values,
sklearn.Citations.values,
'Year','Citations',
title='Scikit-Learn Citations (Google Scholar)',
showplot=True,
save_plot=True,
folder='figures',
filename='ScikitLearnCitations',
template='wide',
file_type='svg',
transparent=True)
bar_plot(tensorflow.Year.values,
tensorflow.Citations.values,
'Year','Citations',
title='Tensorflow Citations (Google Scholar)',
showplot=True,
save_plot=True,
folder='figures',
filename='TensorflowCitations',
template='wide',
file_type='svg',
transparent=True)
# -
| Dissertation-Notebooks/Chapter-4/Citation_Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Fvj2rC--SRfj"
# # The MIT License
# + [markdown] id="mmr3mcc7SRfp"
# Copyright 2020 <NAME>, <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# + [markdown] id="hkMuqKjGLXMH"
# # Attention
#
# Before running this file, please confirm **API keys (Face API, remove.bg, TSUNA)** are prepared.
# + id="LCuZJ99i38MT"
# API Keys
faceAPI_key = input("API Key of Face: ")
removebg_key = input("API Key of remove.bg: ")
tsuna_key = input("API Key of TSUNA: ")
# + id="TMdCldbS38MP"
import cv2
import time
import glob
import requests
import argparse, os
import bs4, shutil, ssl
import numpy as np
import matplotlib.pyplot as plt
import http.client, urllib.request, urllib.parse, json
from bs4 import BeautifulSoup
from PIL import Image, ImageDraw, ImageFont
# + id="KDsL7qZm-Vwl"
video_path = input("Video Filename (e.g. ****.mp4): ") #Filename
summary_text = input("Video Title (Japanese only): ")
search_word = input("Video Key Phrase (This will be searched on Google): ")
# + [markdown] id="h2m0McX7LLku"
# # Step 1: Frame Sampling
# Sampled images are saved in the "processing/step1" folder.
# + id="uUAS5L1_38MW"
def movie_to_image():
output_path = "processing/step1/"
os.makedirs(output_path, exist_ok=True)
#load the video
capture = cv2.VideoCapture(video_path)
img_count = 0 #number of sampling images
frame_count = 0 #number of frames
num_cut = int(capture.get(cv2.CAP_PROP_FRAME_COUNT)/300) #sample approximately 300 images
while(capture.isOpened()):
ret, frame = capture.read()
if ret == False:
break
if frame_count % num_cut == 0:
img_file_name = output_path + str(img_count) + ".jpg"
cv2.imwrite(img_file_name, frame)
img_count += 1
frame_count += 1
capture.release()
return img_count
img_sum = movie_to_image()
print("Frame-Sampling Finished.")
# + [markdown] id="DFAmgsD4M0uz"
# # Step 2: Emotion Recognition
#
# Free version of Face API recognizes 20 images/min.
#
# Two images with the highest probability of happiness and surprise are saved as the "happiness.jpg" file and the "surprise.jpg" file in the "processing/step2" folder.
# + id="3x08nlxM38MY"
file = "processing/step1/"
output_path = "processing/step2/"
os.makedirs(output_path, exist_ok=True)
# extract the image with the highest emotion for each emotion
max_emotion = {"anger":["", 0.0], "contempt":["", 0.0], "disgust":["", 0.0], "fear":["", 0.0],
"happiness":["", 0.0], "neutral":["", 0.0], "sadness":["", 0.0], "surprise": ["", 0.0]}
emo_1 = ["anger","contempt","disgust","fear","happiness","neutral","sadness","surprise"]
img_name = ""
cnt = 0
def RaadJson(datas):
emotion = []
for data in datas:
if data == "error":
return 0
f = data["faceAttributes"]
d = f["emotion"]
for name in emo_1:
emotion.append(d[name])
return emotion
def Recognize(emotion):
data = np.array(emotion)
if data.size == 0:
return
for num in range(8):
if data[num] > max_emotion[emo_1[num]][1]:
max_emotion[emo_1[num]][0] = img_name
max_emotion[emo_1[num]][1] = data[num]
headers = {
#Request headers
"Content-Type" : "application/octet-stream",
"Ocp-Apim-Subscription-Key" : faceAPI_key,
}
params = urllib.parse.urlencode({
#Request parameters
"returnFaceId" : "false",
'returnFaceLandmarks': 'false',
'returnFaceAttributes': 'emotion'
})
try:
conn = http.client.HTTPSConnection('westcentralus.api.cognitive.microsoft.com')
for i in range(0, img_sum):
img_name = file + str(i) + ".jpg"
if cnt > 20:
time.sleep(60)
cnt = 0
f = open(img_name, "rb")
conn.request("POST", "/face/v1.0/detect?%s" % params, f, headers)
response = conn.getresponse()
data = response.read()
data = json.loads(data)
if RaadJson(data) == 0:
cnt += 1
continue
emotion = RaadJson(data)
Recognize(emotion)
cnt += 1
conn.close()
for key, value in max_emotion.items():
if key == "happiness" or key == "surprise":
if value[0] == "":
continue
pic = cv2.imread(value[0])
cv2.imwrite(output_path + key + ".jpg", pic)
except Exception as e:
print("[Errno {0}] {1}".format(e.errno, e.strerror))
print("Emotion-Recognition Finished.")
# + [markdown] id="a1tK6s95-Vwp"
# # Step 3: Image Insertion
#
# Images dowloaded from Google are saved in the "processing/step3/search_deta" folder.
#
# Representative object image which background is trimmed is saved as the "paste_img.png" file in the "processing/step3" folder.
# + id="ku_1aARl38Mc"
output_path = "processing/step3/"
os.makedirs(output_path, exist_ok=True)
def detect_face(img):
face_img = img.copy()
face_rects = face_cascade.detectMultiScale(face_img)
return face_rects
def img_add_msg(img, message, location, emotion, j):
font_path = "system/NotoSansCJKjp-Bold.otf"
font_size = 140
font = ImageFont.truetype(font_path, font_size)
img = Image.fromarray(img)
draw = ImageDraw.Draw(img)
size=draw.textsize(message, font=font)
largex = size[0]
largey = size[1]
#determine where the string must be placed
#priority: bottom → upper left → upper right
length = len(location)
checklu = np.zeros(length)
checkru = np.zeros(length)
checkd = np.zeros(length)
for i in range(length):
if location[i][1] + location[i][3] + largey + 60 < img.height:
checkd[i] = 1
if all(checkd):
textloc = ((img.width-largex)/2, img.height-largey-60) #bottom
else:
font_size = 60
font = ImageFont.truetype(font_path, font_size)
size2=draw.textsize(message, font=font)
x = size2[0]
y = size2[1]
for i in range(length):
if x + 100 < location[i][0] or y + 60 < location[i][1]:
checklu[i] = 1
if all(checklu):
textloc = (100, 60) #upper left
else:
textloc = (img.width-x-50, 30) #upper right
pos = np.array(textloc)
#insert the text
bw = 1
draw.text(pos-(-bw, -bw), message, font=font, fill='black')
draw.text(pos-(-bw, +bw), message, font=font, fill='black')
draw.text(pos-(+bw, -bw), message, font=font, fill='black')
draw.text(pos-(+bw, +bw), message, font=font, fill='black')
draw.text(pos, message, font=font, fill=(0, 0, 255, 0))
img = np.array(img) # convert PIL into cv2(NumPy)
cv2.imwrite("result/recommend_" + emotion + "/000" + str(j) + ".jpg", img)
# + id="K4pHhxf438Me"
data_dir = "processing/step3/search_data/"
os.makedirs(data_dir, exist_ok=True)
ssl._create_default_https_context = ssl._create_unverified_context
def image(search_word, num):
src_list = []
Res = requests.get("https://www.google.com/search?hl=jp&q=" + search_word + "&btnG=Google+Search&tbs=0&safe=off&tbm=isch")
Html = Res.text
Soup = bs4.BeautifulSoup(Html,'lxml')
links = Soup.find_all("img")
i = 0
cnt = 0
while cnt < num:
src = links[i].get("src")
if src[len(src)-3:] == "gif":
i += 1
continue
else:
src_list.append(src)
i += 1
cnt += 1
return src_list
def download_img(url, file_name):
r = requests.get(url, stream=True)
if r.status_code == 200:
with open(file_name +".jpg", 'wb') as f:
r.raw.decode_content = True
shutil.copyfileobj(r.raw, f)
num = 5
srcs = image(search_word, num)
for i in range(num):
file_name = data_dir + str(i)
download_img(srcs[i], file_name)
print("Image-Search Finished.")
# + id="3L7iWpRi38Mh"
# calculation of the similarity
def average_hash(target_file, size):
img = Image.open(target_file)
img = img.convert('RGB')
img = img.resize((size, size), Image.ANTIALIAS)
px = np.array(img.getdata()).reshape((size, size, 3))
avg = px.mean()
px = 1 * (px > avg)
return px
def hamming_dist(a, b):
a = a.reshape(1, -1)
b = b.reshape(1, -1)
dist = (a != b).sum()
return dist
size = 64
images = glob.glob(os.path.join(data_dir, "*.jpg"))
rate = 20.0
result = []
diffmin = 10000
for j, targetf in enumerate(images):
target_dist = average_hash(targetf, size)
for i, fname in enumerate(images, j+3):
dist = average_hash(fname, size)
diff = hamming_dist(target_dist, dist)/256
result.append([diff, targetf, fname])
if diff < diffmin and diff != 0:
diffmin = diff
choice1 = targetf
choice2 = fname
print("Successful Image Selected.")
# + id="Ol1MBEYJ38Mj"
# Inserting a specified image on an image in OpenCV format
class CvOverlayImage(object):
def __init__(self):
pass
@classmethod
def overlay(
cls,
cv_background_image,
cv_overlay_image,
point,
):
overlay_height, overlay_width = cv_overlay_image.shape[:2]
# background image
cv_rgb_bg_image = cv2.cvtColor(cv_background_image, cv2.COLOR_BGR2RGB)
pil_rgb_bg_image = Image.fromarray(cv_rgb_bg_image)
pil_rgba_bg_image = pil_rgb_bg_image.convert('RGBA')
# foreground image
cv_rgb_ol_image = cv2.cvtColor(cv_overlay_image, cv2.COLOR_BGRA2RGBA)
pil_rgb_ol_image = Image.fromarray(cv_rgb_ol_image)
pil_rgba_ol_image = pil_rgb_ol_image.convert('RGBA')
pil_rgba_bg_temp = Image.new('RGBA', pil_rgba_bg_image.size,
(255, 255, 255, 0))
pil_rgba_bg_temp.paste(pil_rgba_ol_image, point, pil_rgba_ol_image)
result_image = \
Image.alpha_composite(pil_rgba_bg_image, pil_rgba_bg_temp)
cv_bgr_result_image = cv2.cvtColor(
np.asarray(result_image), cv2.COLOR_RGBA2BGRA)
return cv_bgr_result_image
# + id="t0k6NIhA38Mm"
response = requests.post(
'https://api.remove.bg/v1.0/removebg',
files={'image_file': open(choice2, 'rb')},
data={'size': 'auto'},
headers={'X-Api-Key': removebg_key},
)
if response.status_code == requests.codes.ok:
with open("processing/step3/paste_img.png", 'wb') as out:
out.write(response.content)
print("Successful Cutout")
else:
print("Error:", response.status_code, response.text)
# + id="7viX0ftF38Mo"
cutimage = Image.open("processing/step3/paste_img.png")
crop = cutimage.split()[-1].getbbox()
newimage = cutimage.crop(crop)
newimage.save("processing/step3/paste_img.png", quality=95)
# + id="ETxR4EFQ38Mq"
# paste the foreground image
def img_paste(background, location):
foreground = cv2.imread("processing/step3/paste_img.png", cv2.IMREAD_UNCHANGED)
original_h, original_w = foreground.shape[:2]
if original_h/original_w>1.5:
foreground = cv2.resize(foreground, (int(background.shape[0]*0.7*foreground.shape[1]/foreground.shape[0]),int(background.shape[0]*0.7)))
elif original_h/original_w>1:
foreground = cv2.resize(foreground, (int(background.shape[0]*0.55*foreground.shape[1]/foreground.shape[0]),int(background.shape[0]*0.55)))
else:
foreground = cv2.resize(foreground, (int(background.shape[0]*0.4*foreground.shape[1]/foreground.shape[0]),int(background.shape[0]*0.4)))
fore_h, fore_w = foreground.shape[:2]
length = len(location)
checkl = np.zeros(length)
for i in range(length):
if fore_w + 100 < location[i][0]:
checkl[i] = 1
if all(checkl):
point = (100, int((background.shape[0] - fore_h)/2)) #left
else:
point = (background.shape[1]-fore_w-100, int((background.shape[0] - fore_h)/2)) #right
image = CvOverlayImage.overlay(background, foreground,point)
return image
# + [markdown] id="jAvqm4MH-Vwu"
# # Step 4: Text Insertion
#
# The thumbnails which are recommended to users are saved in the following folder:
#
# - **happiness**: the "result/recommend_happiness" folder
#
# - **surprise**: the "result/recommend_surprise" folder
# + id="bJsawnt338Ma"
# summarize the title
def summary():
headers = {"x-api-key": tsuna_key}
url = "https://clapi.asahi.com/headline-generation"
query = {"text" : summary_text, "types" : "paper", "length" : "8", "n_head": 5}
r = requests.post(url, headers=headers, data=query)
data = json.loads(r.text)
return data["headline"]
# + id="wd3v5ZHe38Ms"
title_data = summary()
# + id="9ii8hNSE38Mu"
#original image
happiness = 'processing/step2/happiness.jpg'
surprise = 'processing/step2/surprise.jpg'
face_cascade = cv2.CascadeClassifier('system/haarcascade_frontalface_default.xml')
# happiness
happiness_img = cv2.imread(happiness, cv2.IMREAD_UNCHANGED)
location_happiness = detect_face(happiness_img)
os.makedirs("result/recommend_happiness", exist_ok=True)
h_image = img_paste(happiness_img, location_happiness)
for i in range(0, 5):
img_add_msg(h_image, title_data[i], location_happiness, "happiness", i)
#surprise
surprise_img = cv2.imread(surprise, cv2.IMREAD_UNCHANGED)
location_surprise = detect_face(surprise_img)
os.makedirs("result/recommend_surprise", exist_ok=True)
s_image = img_paste(surprise_img, location_surprise)
for i in range(0, 5):
img_add_msg(s_image, title_data[i], location_surprise, "surprise", i)
print("Output Finished.")
| sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# # GLM Analysis (Simulated)
#
# In this example we simulate a block design
# functional near-infrared spectroscopy (fNIRS)
# experiment and analyse
# the simulated signal. We investigate the effect additive noise and
# measurement length has on response amplitude estimates.
# :depth: 2
#
# +
# sphinx_gallery_thumbnail_number = 3
# Authors: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import mne
import mne_nirs
import matplotlib.pylab as plt
import numpy as np
from mne_nirs.experimental_design import make_first_level_design_matrix
from mne_nirs.statistics import run_glm
from nilearn.plotting import plot_design_matrix
np.random.seed(1)
# -
# ## Simulate noise free NIRS data
#
# First we simulate some noise free data. We simulate 5 minutes of data with a
# block design. The inter stimulus interval of the stimuli is uniformly
# selected between 15 and 45 seconds.
# The amplitude of the simulated signal is 4 uMol and the sample rate is 3 Hz.
# The simulated signal is plotted below.
#
#
# +
sfreq = 3.
amp = 4.
raw = mne_nirs.simulation.simulate_nirs_raw(
sfreq=sfreq, sig_dur=60 * 5, amplitude=amp, isi_min=15., isi_max=45.)
raw.plot(duration=300, show_scrollbars=False)
# -
# ## Create design matrix
#
# Next we create a design matrix based on the annotation times in the simulated
# data. We use the nilearn plotting function to visualise the design matrix.
# For more details on this procedure see `tut-fnirs-hrf`.
#
#
design_matrix = make_first_level_design_matrix(raw, stim_dur=5.0,
drift_order=1,
drift_model='polynomial')
fig, ax1 = plt.subplots(figsize=(10, 6), nrows=1, ncols=1)
fig = plot_design_matrix(design_matrix, ax=ax1)
# ## Estimate response on clean data
#
# Here we run the GLM analysis on the clean data.
# The design matrix had three columns, so we get an estimate for our simulated
# event, the first order drift, and the constant.
# We see that the estimate of the first component is 4e-6 (4 uM),
# which was the amplitude we used in the simulation.
# We also see that the mean square error of the model fit is close to zero.
#
#
# +
glm_est = run_glm(raw, design_matrix)
def print_results(glm_est, truth):
"""Function to print the results of GLM estimate"""
print("Estimate:", glm_est.theta()[0][0],
" MSE:", glm_est.MSE()[0],
" Error (uM):", 1e6*(glm_est.theta()[0][0] - truth * 1e-6))
print_results(glm_est, amp)
# -
# ## Simulate noisy NIRS data (white)
#
# Real data has noise. Here we add white noise, this noise is not realistic
# but serves as a reference point for evaluating the estimation process.
# We run the GLM analysis exactly as in the previous section
# and plot the noisy data and the GLM fitted model.
# We print the response estimate and see that is close, but not exactly correct,
# we observe the mean square error is similar to the added noise.
# Note that the clean data plot is so similar to the GLM estimate that it is hard to see unless zoomed in.
#
#
# +
# First take a copy of noise free data for comparison
raw_noise_free = raw.copy()
raw._data += np.random.normal(0, np.sqrt(1e-11), raw._data.shape)
glm_est = run_glm(raw, design_matrix)
plt.plot(raw.times, raw_noise_free.get_data().T * 1e6)
plt.plot(raw.times, raw.get_data().T * 1e6, alpha=0.3)
plt.plot(raw.times, glm_est.theta()[0][0] * design_matrix["A"].values * 1e6)
plt.xlabel("Time (s)")
plt.ylabel("Haemoglobin (uM)")
plt.legend(["Clean Data", "Noisy Data", "GLM Estimate"])
print_results(glm_est, amp)
# -
# ## Simulate noisy NIRS data (colored)
#
# Here we add colored noise which better matches what is seen with real data.
# Again, the same GLM procedure is run.
# The estimate is reported below, and even though the signal was difficult to
# observe in the raw data, the GLM analysis has extracted an accurate estimate.
# However, the error is greater for the colored than white noise.
#
#
# +
raw = raw_noise_free.copy()
cov = mne.Covariance(np.ones(1) * 1e-11, raw.ch_names,
raw.info['bads'], raw.info['projs'], nfree=0)
raw = mne.simulation.add_noise(raw, cov,
iir_filter=[1., -0.58853134, -0.29575669,
-0.52246482, 0.38735476,
0.02428681])
design_matrix = make_first_level_design_matrix(raw, stim_dur=5.0,
drift_order=1,
drift_model='polynomial')
glm_est = run_glm(raw, design_matrix)
plt.plot(raw.times, raw_noise_free.get_data().T * 1e6)
plt.plot(raw.times, raw.get_data().T * 1e6, alpha=0.3)
plt.plot(raw.times, glm_est.theta()[0][0] * design_matrix["A"].values * 1e6)
plt.xlabel("Time (s)")
plt.ylabel("Haemoglobin (uM)")
plt.legend(["Clean Data", "Noisy Data", "GLM Estimate"])
print_results(glm_est, amp)
# -
# ## How does increasing the measurement length affect estimation accuracy?
#
# The easiest way to reduce error in your response estimate is to collect more
# data. Here we simulated increasing the recording time to 30 minutes.
# We run the same analysis and observe that the error is reduced from
# approximately 0.6 uM for 5 minutes of data to 0.25 uM for 30 minutes of data.
#
#
# +
raw = mne_nirs.simulation.simulate_nirs_raw(
sfreq=sfreq, sig_dur=60 * 30, amplitude=amp, isi_min=15., isi_max=45.)
cov = mne.Covariance(np.ones(1) * 1e-11, raw.ch_names,
raw.info['bads'], raw.info['projs'], nfree=0)
raw = mne.simulation.add_noise(raw, cov,
iir_filter=[1., -0.58853134, -0.29575669,
-0.52246482, 0.38735476,
0.02428681])
design_matrix = make_first_level_design_matrix(raw, stim_dur=5.0,
drift_order=1,
drift_model='polynomial')
glm_est = run_glm(raw, design_matrix)
plt.plot(raw.times, raw.get_data().T * 1e6, alpha=0.3)
plt.plot(raw.times, glm_est.theta()[0][0] * design_matrix["A"].values * 1e6)
plt.xlabel("Time (s)")
plt.ylabel("Haemoglobin (uM)")
plt.legend(["Noisy Data", "GLM Estimate"])
print_results(glm_est, amp)
# -
# ## Using autoregressive models in the GLM to account for noise structure
#
# An auto regressive noise model can be used account for temporal structure
# in the noise. To account for the noise properties in the example above,
# a fifth order auto regressive model is used below. Given this
# is a simulation, we can verify if the correct estimate of the noise
# properties was extracted from the data and if this
# improved the response estimate.
#
#
# +
glm_est = run_glm(raw, design_matrix, noise_model='ar5')
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(15, 6))
plt.plot([-0.58853134, -0.29575669, -0.52246482, 0.38735476, 0.02428681],
axes=axes) # actual values from model above
plt.plot(glm_est.model()[0].rho * -1.0, axes=axes) # estimates
plt.legend(["Simulation AR coefficients", "Estimated AR coefficients"])
plt.xlabel("Coefficient")
# -
# We can see that the estimates from the GLM AR model are quite accurate,
# but how does this affect the accuracy of the response estimate?
#
#
print_results(glm_est, amp)
# The response estimate using the AR(5) model is more accurate than the
# AR(1) model (error of 0.25 vs 2.8 uM).
#
#
# ## Conclusion?
#
# In this example we have generated a noise free signal containing simulated
# haemodynamic responses. We were then able to accurately estimate the amplitude
# of the simulated signal. We then added noise and illustrated that the
# estimate provided by the GLM was correct, but contained some error. We
# observed that as the measurement time was increased, the estimated
# error decreased.
# We also observed in this idealised example that including an appropriate
# model of the noise can improve the accuracy of the response estimate.
#
#
| examples/mne-nirs-website-examples/auto_examples/general/plot_10_hrf_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: all,-execution,-papermill,-trusted
# formats: ipynb,py//py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown] tags=[]
# # Description
# %% [markdown] tags=[]
# This notebook computes predicted expression correlations between all genes in the MultiPLIER models.
#
# It also has a parameter set for papermill to run on a single chromosome to run in parallel (see under `Settings` below).
# %% [markdown] tags=[]
# # Modules
# %% tags=[]
# %load_ext autoreload
# %autoreload 2
# %% tags=[]
import numpy as np
from scipy.spatial.distance import squareform
import pandas as pd
from tqdm import tqdm
import conf
from entity import Gene
# %% [markdown] tags=[]
# # Settings
# %% tags=["parameters"]
# specifies a single chromosome value
# by default, run on all chromosomes
chromosome = "all"
# %% tags=["injected-parameters"]
# Parameters
chromosome = 20
# %% tags=[]
if chromosome == "all":
from time import sleep
message = """
WARNING: you are going to compute correlations of gene predicted expression across all chromosomes without parallelism.
It is recommended that you look at the README.md file in this subfolder (nbs/08_gsa_gls/README.md) to know how to do that.
It will continue in 20 seconds.
"""
print(message)
sleep(20)
# %% [markdown] tags=[]
# # Load data
# %% [markdown] tags=[]
# ## MultiPLIER Z
# %% tags=[]
multiplier_z_genes = pd.read_pickle(
conf.MULTIPLIER["MODEL_Z_MATRIX_FILE"]
).index.tolist()
# %% tags=[]
len(multiplier_z_genes)
# %% tags=[]
multiplier_z_genes[:10]
# %% [markdown] tags=[]
# ## Get gene objects
# %% tags=[]
multiplier_gene_obj = {
gene_name: Gene(name=gene_name)
for gene_name in multiplier_z_genes
if gene_name in Gene.GENE_NAME_TO_ID_MAP
}
# %% tags=[]
len(multiplier_gene_obj)
# %% tags=[]
multiplier_gene_obj["GAS6"].ensembl_id
# %% tags=[]
_gene_obj = list(multiplier_gene_obj.values())
genes_info = pd.DataFrame(
{
"name": [g.name for g in _gene_obj],
"id": [g.ensembl_id for g in _gene_obj],
"chr": [g.chromosome for g in _gene_obj],
}
)
# %% tags=[]
genes_info.shape
# %% tags=[]
genes_info.head()
# %% [markdown] tags=[]
# ## Get tissues names
# %% tags=[]
db_files = list(conf.PHENOMEXCAN["PREDICTION_MODELS"]["MASHR"].glob("*.db"))
# %% tags=[]
assert len(db_files) == 49
# %% tags=[]
tissues = [str(f).split("mashr_")[1].split(".db")[0] for f in db_files]
# %% tags=[]
tissues[:5]
# %% [markdown] tags=[]
# # Test
# %% tags=[]
genes_info[genes_info["chr"] == "13"]
# %% tags=[]
_gene_list = [
Gene("ENSG00000134871"),
Gene("ENSG00000187498"),
Gene("ENSG00000183087"),
Gene("ENSG00000073910"),
Gene("ENSG00000133101"),
Gene("ENSG00000122025"),
Gene("ENSG00000120659"),
Gene("ENSG00000133116"),
]
tissue = "Whole_Blood"
# %% tags=[]
# %%timeit
for gene_idx1 in range(0, len(_gene_list) - 1):
gene_obj1 = _gene_list[gene_idx1]
for gene_idx2 in range(gene_idx1 + 1, len(_gene_list)):
gene_obj2 = _gene_list[gene_idx2]
gene_obj1.get_expression_correlation(
gene_obj2,
tissue,
)
# %% [markdown] tags=[]
# # Compute correlation per chromosome
# %% tags=[]
all_chrs = genes_info["chr"].dropna().unique()
assert all_chrs.shape[0] == 22
if chromosome != "all":
chromosome = str(chromosome)
assert chromosome in all_chrs
# run only on the chromosome specified
all_chrs = [chromosome]
# # For testing purposes
# all_chrs = ["13"]
# tissues = ["Whole_Blood"]
# genes_info = genes_info[genes_info["id"].isin(["ENSG00000134871", "ENSG00000187498", "ENSG00000183087", "ENSG00000073910"])]
for chr_num in all_chrs:
print(f"Chromosome {chr_num}", flush=True)
genes_chr = genes_info[genes_info["chr"] == chr_num]
print(f"Genes in chromosome{genes_chr.shape}", flush=True)
gene_chr_objs = [Gene(ensembl_id=gene_id) for gene_id in genes_chr["id"]]
gene_chr_ids = [g.ensembl_id for g in gene_chr_objs]
n = len(gene_chr_objs)
n_comb = int(n * (n - 1) / 2.0)
print(f"Number of gene combinations: {n_comb}", flush=True)
for tissue in tissues:
print(f"Tissue {tissue}", flush=True)
# check if results exist
output_dir = conf.PHENOMEXCAN["LD_BLOCKS"]["BASE_DIR"] / "gene_corrs" / tissue
output_file = output_dir / f"gene_corrs-{tissue}-chr{chr_num}.pkl"
if output_file.exists():
_tmp_data = pd.read_pickle(output_file)
if _tmp_data.shape[0] > 0:
print("Already run, stopping.")
continue
gene_corrs = []
pbar = tqdm(ncols=100, total=n_comb)
i = 0
for gene_idx1 in range(0, len(gene_chr_objs) - 1):
gene_obj1 = gene_chr_objs[gene_idx1]
for gene_idx2 in range(gene_idx1 + 1, len(gene_chr_objs)):
gene_obj2 = gene_chr_objs[gene_idx2]
gene_corrs.append(
gene_obj1.get_expression_correlation(gene_obj2, tissue)
)
pbar.update(1)
pbar.close()
# testing
gene_corrs_flat = pd.Series(gene_corrs)
print(f"Min/max values: {gene_corrs_flat.min()} / {gene_corrs_flat.max()}")
assert gene_corrs_flat.min() >= -1.001
assert gene_corrs_flat.max() <= 1.001
# save
gene_corrs_data = squareform(np.array(gene_corrs, dtype=np.float32))
np.fill_diagonal(gene_corrs_data, 1.0)
gene_corrs_df = pd.DataFrame(
data=gene_corrs_data,
index=gene_chr_ids,
columns=gene_chr_ids,
)
output_dir.mkdir(exist_ok=True, parents=True)
display(output_file)
gene_corrs_df.to_pickle(output_file)
# %% [markdown] tags=[]
# # Testing
# %% tags=[]
# data = pd.read_pickle(
# conf.PHENOMEXCAN["LD_BLOCKS"]["BASE_DIR"] / "gene_corrs" / "Whole_Blood" / "gene_corrs-Whole_Blood-chr13.pkl"
# )
# %% tags=[]
# assert data.loc["ENSG00000134871", "ENSG00000187498"] > 0.97
# %% tags=[]
| nbs/15_gsa_gls/gene_corrs/10-gene_expr_correlations-chr20.run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Shubham0Rajput/Feature-Detection-with-AKAZE/blob/master/AKAZE_code.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pKtBjHdbpdZQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="37d8f437-9b5b-4282-9a87-e6bb730d058c"
#IMPORT FILES
import matplotlib.pyplot as plt
import cv2
#matplotlib inline
#MOUNTIING DRIVE
from google.colab import drive
drive.mount('/content/drive')
# + id="k6YDoxkjqrU3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 383} outputId="169192bb-bf02-49a5-c5c4-a1ebf96afe3d"
from __future__ import print_function
import cv2 as cv
import numpy as np
import argparse
from math import sqrt
import matplotlib.pyplot as plt
imge1 = cv.imread('/content/drive/My Drive/e2.jpg')
img1 = cv.cvtColor(imge1, cv.COLOR_BGR2GRAY) # queryImage
imge2 = cv.imread('/content/drive/My Drive/e1.jpg')
img2 = cv.cvtColor(imge2, cv.COLOR_BGR2GRAY) # trainImage
if img1 is None or img2 is None:
print('Could not open or find the images!')
exit(0)
fs = cv.FileStorage('/content/drive/My Drive/H1to3p.xml', cv.FILE_STORAGE_READ)
homography = fs.getFirstTopLevelNode().mat()
## [AKAZE]
akaze = cv.AKAZE_create()
kpts1, desc1 = akaze.detectAndCompute(img1, None)
kpts2, desc2 = akaze.detectAndCompute(img2, None)
## [AKAZE]
## [2-nn matching]
matcher = cv.DescriptorMatcher_create(cv.DescriptorMatcher_BRUTEFORCE_HAMMING)
nn_matches = matcher.knnMatch(desc1, desc2, 2)
## [2-nn matching]
## [ratio test filtering]
matched1 = []
matched2 = []
nn_match_ratio = 0.8 # Nearest neighbor matching ratio
for m, n in nn_matches:
if m.distance < nn_match_ratio * n.distance:
matched1.append(kpts1[m.queryIdx])
matched2.append(kpts2[m.trainIdx])
## [homography check]
inliers1 = []
inliers2 = []
good_matches = []
inlier_threshold = 2.5 # Distance threshold to identify inliers with homography check
for i, m in enumerate(matched1):
col = np.ones((3,1), dtype=np.float64)
col[0:2,0] = m.pt
col = np.dot(homography, col)
col /= col[2,0]
dist = sqrt(pow(col[0,0] - matched2[i].pt[0], 2) +\
pow(col[1,0] - matched2[i].pt[1], 2))
if dist > inlier_threshold:
good_matches.append(cv.DMatch(len(inliers1), len(inliers2), 0))
inliers1.append(matched1[i])
inliers2.append(matched2[i])
## [homography check]
## [draw final matches]
res = np.empty((max(img1.shape[0], img2.shape[0]), img1.shape[1]+img2.shape[1], 3), dtype=np.uint8)
img0 = cv.drawMatches(img1, inliers1, img2, inliers2, good_matches, res)
#img0 = cv.drawMatchesKnn(img1,inliers1,img2,inliers2,res,None,flags=cv.DrawMatchesFlags_NOT_DRAW_SINGLE_POINTS)
cv.imwrite("akaze_result.png", res)
inlier_ratio = len(inliers1) / float(len(matched1))
print('A-KAZE Matching Results')
print('*******************************')
print('# Keypoints 1: \t', len(kpts1))
print('# Keypoints 2: \t', len(kpts2))
print('# Matches: \t', len(matched1))
print('# Inliers: \t', len(inliers1))
print('# Inliers Ratio: \t', inlier_ratio)
print('# Dist: \t', dist)
plt.imshow(img0),plt.show()
## [draw final matches]
# + id="jPYXTRj5oHk1" colab_type="code" colab={}
| AKAZE_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup code:
import urllib.parse as urlparse
from aiidalab_widgets_base import AiiDACodeSetup, valid_aiidacode_args
parsed_url = urlparse.parse_qs(urlparse.urlsplit(jupyter_notebook_url).query)
args = valid_aiidacode_args(parsed_url)
code = AiiDACodeSetup(**args)
display(code)
| setup_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
# +
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import time
import itertools as it
import helpers_07
# %matplotlib inline
# -
# # Transfer Learning
# Our goal this week is to apply a *previously trained network* (that was trained on the ILSVRC dataset), modify it slightly, and with only minimal retraining on a new dataset (called HamsterHare), use it to predict on the new task. Let's go.
# # A Different Dataset
# As per usual, we've hidden the complexity of grabbing the data. Because the dataset is a bit bigger, for now, we only return paths to files (and not full images as a NumPy array). We'll deal with that more in a bit.
helpers_07.download_hh("data/hh")
# note: label=1 means hare; label=0 means hamster
readable_hh_labels = {1:"hare", 0:"hamster"}
(train_files, train_labels,
test_files, test_labels) = helpers_07.train_test_split_hh_filenames(test_pct=.3)
print("\n".join(str(t) for t in train_files[:5]))
print("\n".join(str(t) for t in train_labels[:5]))
# Let's see what we're dealing with:
from scipy.misc import imread
img = imread(train_files[0], mode="RGB")[:,:,:3]
plt.imshow(img)
plt.title(readable_hh_labels[train_labels[0]])
plt.axis('off');
# # (Re-)Loading a Model and Accessing Named Model Elements
# ##### Named Elements
# In a few minutes, we are going to reload the AlexNet we built last week. When we do so, we will have agrat network, but we will have no variables referencing the Tensors and Operations inside of it. We need to deal with that, so at minimum we can feed it data. We'll even go beyond that: we'll add Operations to the network so we can retrain portions of it.
#
# To get started, let's look at a simpler example.
test_graph = tf.Graph()
with test_graph.as_default():
a = tf.constant(3, name="constant_a")
b = tf.constant(4, name="constant_b")
c = tf.multiply(a,b, name="mul_for_c")
# We can access operations. A few points:
# * ops are the "nodes" in the graph
# * they are operations that perform computation
# * op name is same as we passed to name=
print("Getting an Operation")
print(test_graph.get_operation_by_name('mul_for_c'))
# And we can access tensors. A few points:
# * tensors are the data or edges in the Graph
# * they often are the result of an Operation
# * tensor name is the `op:<number>`
# * many ops have only one output (the data we want), so the number is often 0
# * some ops have multiple outputs, to the numbers go 0,1,2, for diff output tensors
print("Getting a Tensor:")
new_c_ref = test_graph.get_tensor_by_name('mul_for_c:0')
print(c)
print(new_c_ref)
print(c is new_c_ref) # aka, refer to -same- object
# # Loading a Graph
# Now, let's bring back in the AlexNet we (possibly struggled) to create last week.
# create a placeholder graph to "rehydrate" our freeze-dried AlexNet
old_alex_graph = tf.Graph()
with old_alex_graph.as_default():
# importing the graph will populate new_alex_graph
saver = tf.train.import_meta_graph("../week_06/saved_models/alexnet.meta")
# And we can use some `get_*_by_name` methods to extract out tensors and operations. Note, we have to know these ahead of time (but, see below!).
# note the form is scope/name= ; for output tensor, we can tack on :0
print(old_alex_graph.get_tensor_by_name('inputs/images:0'), "\n",
old_alex_graph.get_operation_by_name('inputs/images'), sep="\n")
# Now, we have sort of an idealized scenario: we made the graph last week. So, we know (or we could go back to our old code an lookup) the names of the tensors and operations. But, what if (1) someone else gave us the graph, (2) we lost our old code, (3) we needed to programmatically get the names of tensors/operations. What would we do? Fortunately, this is a solved problem. We use `get_operations` as in `old_alex_graph.get_operations`.
print("\n".join(str(op.name) for op in old_alex_graph.get_operations()[:5]))
# The way we'll actually use this is to grab the specific tensors we need to work with in our new/transfered use of our AlexNet. Namely, we need the inputs and we need whatever "end point" we are going to cut off the old AlexNet and move to our new AlexNet. We're just going to cut out the last layer (`fc8` from last time). So, we'll pipe `fc7` somewhere else. More to come on that shortly.
# get references (Python variables) that refer
# to the named elements we need to access
inputs = old_alex_graph.get_tensor_by_name('inputs/images:0')
fc7 = old_alex_graph.get_tensor_by_name("alexnet/fully_connected_1/fc7:0")
# ## Exercise
# See if you can pickout the operations in the `fully_connected` name scope from the old graph.
# ### Solution
# ## A Retraining Model
# Since our original AlexNet model didn't have any training component (we populated its weights directly from NumPy arrays), we have some work to do, if we want to add training capabilities. Here's a template from our older trainable models:
# ## Exercise
# Look inside the following code and try to build up a model that resuses as much as possible from the prior model. Here's was out outline of steps from the slides:
#
# 1. Get handle to output from second-to-last layer
# 2. Create a new fully connected layer
# * number of neurons equal to the number of output classes)
# 3. Create new softmax cross-entropy loss
# 4. Create a training op to minimize the new loss
# * Set var_list parameter to be just the new layer variables
# 5. Train with new data!
class TransferedAlexNet:
def __init__(s, init_graph):
with init_graph.as_default():
#
# The original AlexNet from last week didn't have:
# labels, loss, training
# Also, it's prediction was structured for 1000 output classes
#
# Since we passed in init_graph above, we are working *with* that
# old AlexNet. But you can add to it. For example: here we are *adding*
# a labels placeholder to the original model:
with tf.name_scope('inputs'):
s.labels = tf.placeholder(tf.int32, shape=[None], name='labels')
#
# revisit one of your older models, and add in the remaining pieces:
# learning_rate, loss, global_step, training, a new prediction system, etc.
#
# you'll also need a new final layer to replace the fc8 layer from last time
# you can use helper_07.fully_connected_xavier_relu_layer
# to replace it ...
#
#
# FILL ME IN
#
init = tf.global_variables_initializer()
s.session = tf.Session(graph=init_graph)
s.session.run(init)
def fit(s, train_dict):
tr_loss, step, tr_acc, _ = s.session.run([s.loss, s.inc_step, s.pred_accuracy, s.train],
feed_dict=train_dict)
return tr_loss, step, tr_acc
def predict(s, test_dict):
ct_correct, preds = s.session.run([s.pred_correct, s.prediction],
feed_dict=test_dict)
return ct_correct, preds
# ### Solution
# The following code adds one nice twist. If you look at lines 31 and 37 (use Control-m followed by l - little 'ell' in the cell to get line numbers), you'll see that we setup a way to only optimize on the selected variables: in this case, the variables from our new end layer. This saves a ton of time (fewer parameters to work with) -and- it prevents us from losing the work done in the prior (very long/large) training steps (i.e., the work done before we even got our old AlexNet weights).
class TransferedAlexNet:
def __init__(s, init_graph, num_tgt_classes):
with init_graph.as_default():
with tf.name_scope('inputs'):
# have input placeholder from original graph
s.labels = tf.placeholder(tf.int32, shape=[None], name='labels')
with tf.name_scope('hyperparams'):
s.learning_rate = tf.placeholder(tf.float32, name='learning_rate')
s.one_hot_labels = tf.one_hot(s.labels, 2) # , dtype=tf.float32)
#
# we're going to rewire the outputs from the old fc7 to our new layer
#
orig_fc7 = init_graph.get_tensor_by_name("alexnet/fully_connected_1/fc7:0")
with tf.name_scope('new_top_layer'):
# the old fc8 (which we are replacing) had 1000 nodes for 1000 classes
s.logits = helpers_07.fully_connected_xavier_relu_layer(orig_fc7, num_tgt_classes)
with tf.name_scope('loss'):
smce = tf.nn.softmax_cross_entropy_with_logits
s.loss = tf.reduce_mean(smce(logits=s.logits, labels=s.one_hot_labels),
name="loss")
with tf.name_scope('global_step'):
global_step = tf.Variable(0, trainable=False, name='global_step')
s.inc_step = tf.assign_add(global_step, 1, name='inc_step')
# use to_train_vars = None to train on all trainable (including those from original)
to_train_vars = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, "new_top_layer")
with tf.name_scope('train'):
decayed_rate = tf.train.exponential_decay(s.learning_rate, global_step,
600, 0.998, True)
momopt = tf.train.MomentumOptimizer
s.train = momopt(decayed_rate, 0.9).minimize(s.loss, var_list=to_train_vars)
# there is a prediction namescope in the original model
# note, that the variable assignments here are attributes of this class
# and refer to distinct operations compared to the original model
with tf.name_scope('new_prediction'):
s.softmax = tf.nn.softmax(s.logits, name="softmax")
s.prediction = tf.cast(tf.arg_max(s.softmax, 1), tf.int32)
s.pred_correct = tf.equal(s.labels, s.prediction)
s.pred_accuracy = tf.reduce_mean(tf.cast(s.pred_correct, tf.float32))
init = tf.global_variables_initializer()
s.session = tf.Session(graph=init_graph)
s.session.run(init)
def fit(s, train_dict):
tr_loss, step, tr_acc, _ = s.session.run([s.loss, s.inc_step, s.pred_accuracy, s.train],
feed_dict=train_dict)
return tr_loss, step, tr_acc
def predict(s, test_dict):
ct_correct, preds = s.session.run([s.pred_correct, s.prediction],
feed_dict=test_dict)
return ct_correct, preds
# # Retraining
# Now, let's put this all together and retrain with some new data. I'm going to show a few options that demonstrate different techniques you might need. One issue we pushed under the hood, was that of the input shape. In TensorFlow, single batches of images must all be the same size. Even if the AlexNet we built can rescale images to 227 by 227, the inputs all need to be of one, common size. Even worse, our Hamster Hare images are all varying sizes. To show off an alternative use of TensorFlow, here is a set of helpers to rescale images (one image at a time).
# +
class TF_ReadAndScale:
'rescale images to a common size'
def __init__(self, img_size=[227, 227]):
self.img_size = img_size
ras_graph = tf.Graph()
with ras_graph.as_default():
self.img_path = tf.placeholder(tf.string)
raw_data = tf.read_file(self.img_path)
jpg_image = tf.image.decode_jpeg(raw_data, channels=3)
self.scaled_img = tf.image.resize_images(jpg_image, img_size)
self.session = tf.Session(graph=ras_graph)
def scale(self, img_path):
return self.session.run(self.scaled_img, feed_dict={self.img_path:img_path})
def image_files_into_array(img_file_lst, dtype=np.uint8, limit=None):
'take a list of filenames; return an array of images'
scaler = TF_ReadAndScale()
num_images = len(img_file_lst)
img_array_shape = [num_images] + scaler.img_size + [3]
img_array = np.empty(img_array_shape, dtype=dtype)
for tf, img_home in it.islice(zip(img_file_lst, img_array), limit):
img_home[:] = scaler.scale(tf)
return img_array
# -
# With that out of the way, let's build our model and get set to feed it new data.
# danger, rerunning will modify a modified model
# b/c the reruns share the old_alex_graph
# and it is updating through the reference
new_alex = TransferedAlexNet(old_alex_graph, num_tgt_classes=2)
inputs = old_alex_graph.get_tensor_by_name('inputs/images:0')
# We're going to save ourselves a lot of processing time (at the cost of some memory usage) but rescaling all of the images before we pass them for training. See below for an alternative.
# +
# you can set a limit here to either use all the data (limit = None)
# or, set to a small integer if you want to debug/time test (not for production)
limit = None # use all of a batch
# limit = 10 # use just 10 images per batch (completely ignore remainder)
batch_size = 32 # you can scale this up, if you want more images through your net at once
# way above, we did:
# (train_files, train_labels,
# test_files, test_labels) = helpers_07.train_test_split_hh_filenames(test_pct=.3)
# this load all of the images into memory; could be a problem on some machines
img_array = image_files_into_array(train_files, dtype=np.float32)
lbl_array = np.array(train_labels)
for epoch in range(5):
start = time.time()
batcher = helpers_07.array_batches(img_array, lbl_array, batch_size)
for image_batch, label_batch in it.islice(batcher, limit):
train_dict = {inputs : image_batch,
new_alex.labels : label_batch,
new_alex.learning_rate : 0.05}
tr_loss, step, tr_acc = new_alex.fit(train_dict)
end = time.time()
info_update = "Epoch: {:2d} Step: {:5d} Loss: {:8.2f} Acc: {:5.2f} Time: {:5.2f}"
print(info_update.format(epoch, step, tr_loss, tr_acc, (end - start) / 60.0))
# -
# The following is an alternative that shows off what you might have to do if you have large data and/or small physical memory size. Instead of loading all the images into memory at once, we simply load them "on demand" as needed by the batches. Note, the specific loop below also means that we rescale the images every time they are opened. We could rescale and save and then reopen the saved version, if we wanted.
def do_it():
scaler = TF_ReadAndScale()
for epoch in range(5):
start = time.time()
batcher = helpers_07.list_batches(train_files, train_labels, 32)
for file_batch, label_batch in it.islice(batcher, 10):
image_batch = [scaler.scale(a_file) for a_file in file_batch]
train_dict = {inputs : image_batch,
new_alex.labels : label_batch,
new_alex.learning_rate : 0.05}
tr_loss, step, tr_acc = new_alex.fit(train_dict)
end = time.time()
info_update = "Epoch: {:2d} Step: {:5d} Loss: {:8.2f} Acc: {:5.2f} Time: {:5.2f}"
print(info_update.format(epoch, step, tr_loss, tr_acc, (end - start) / 60.0))
# do_it() # disabled
# You may want to run this once to see the difference in running times compared to the above "scale them all once" method.
# # Evaluation
# ## Exercise
# Using the `.predict` method we gave `TransferedAlexNet`, write some code to evaluate it on a test set. Remember, we need to scale the test images before they are fed to our model. Here's a template (that needs several modification to be used) from the Week 04 notebook:
# +
#total_correct = 0
#for batch_data, batch_labels in batches(test_data, test_labels, 200):
# test_dict = {}
# correctness, curr_preds = model.predict(test_dict)
# total_correct += correctness.sum()
#print(total_correct / len(test_data))
# -
# ### Solution
| Intel-TF101-Class7/Week7_Transfer_Learning_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 425} id="iCl1TNdNyQq_" executionInfo={"status": "error", "timestamp": 1629215083907, "user_tz": -540, "elapsed": 3189, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="c8a12e6a-0a37-4db6-9eff-c8728e908791"
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
import pandas as pd
# 데이터 입력
df = pd.read_csv('sample_data/iris.csv', names = ["sepal_length", "sepal_width", "petal_length", "petal_width", "species"])
# + id="tku6uAt-ySbs" executionInfo={"status": "aborted", "timestamp": 1629215083902, "user_tz": -540, "elapsed": 7, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}}
# 데이터 분류
dataset=df.copy()
# 데이터 분류
Y_obj=dataset.pop("species")
X=dataset.copy()
# 문자열을 숫자로 변환
Y_encoded=pd.get_dummies(Y_obj)
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="QkRArrcYyUAC" executionInfo={"status": "error", "timestamp": 1629215093766, "user_tz": -540, "elapsed": 292, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="900a4337-a14e-4984-e702-07dabd62d0c4"
# 전체 데이터에서 학습 데이터와 테스트 데이터(0.1)로 구분
X_train1, X_test, Y_train1, Y_test = train_test_split(X, Y_encoded, test_size=0.1,shuffle=True) ## shuffle=True로 하면 데이터를 섞어서 나눔
## 학습 셋에서 학습과 검증 데이터(0.2)로 구분
X_train, X_valid, Y_train, Y_valid = train_test_split(X_train1, Y_train1, test_size=0.2, shuffle=True) ## shuffle=True로 하면 데이터를 섞어서 나눔
# + colab={"base_uri": "https://localhost:8080/", "height": 372} id="411dV4ozyXEj" executionInfo={"status": "error", "timestamp": 1629215110867, "user_tz": -540, "elapsed": 282, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="5bc466c1-df07-425f-8b7b-77c4219ade5c"
## input layer / hidden layer / output layer 각 perceptron 갯수
## 모델의 파라미터( w, b) 값들이 불러와짐.
model=tf.keras.models.load_model('iris_multi_model.h5')
model.summary()
# + id="pCe2_PRmybP0" executionInfo={"status": "error", "timestamp": 1629215124090, "user_tz": -540, "elapsed": 282, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="78c4aa1e-fadb-422a-9fc8-7781266f7364" colab={"base_uri": "https://localhost:8080/", "height": 241}
#
# 모델 컴파일
loss=tf.keras.losses.categorical_crossentropy
optimizer = tf.keras.optimizers.SGD(learning_rate=0.01)
model.compile(loss=loss,
optimizer=optimizer,
metrics=[tf.keras.metrics.categorical_accuracy])
## model fit은 histoy를 반환한다. 훈련중의 발생하는 모든 정보를 담고 있는 딕셔너리.
result=model.fit(X_train, Y_train, epochs=200, batch_size=50, validation_data=(X_valid,Y_valid)) # validation_data=(X_valid,Y_valid)을 추가하여 학습시 검증을 해줌.
model.save('iris_multi_model2.h5')
## histoy는 딕셔너리이므로 keys()를 통해 출력의 key(카테고리)를 확인하여 무엇을 받고 있는지 확인.
print(result.history.keys())
# + id="5kJkNcaEyefC"
### history에서 loss와 val_loss의 key를 가지는 값들만 추출
loss = result.history['loss']
val_loss = result.history['val_loss']
### loss와 val_loss를 그래프화
epochs = range(1, len(loss) + 1)
plt.subplot(211) ## 2x1 개의 그래프 중에 1번째
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
### history에서 binary_accuracy와 val_binary_accuracy key를 가지는 값들만 추출
acc = result.history['categorical_accuracy']
val_acc = result.history['val_categorical_accuracy']
### binary_accuracy와 val_binary_accuracy key를 그래프화
plt.subplot(212) ## 2x1 개의 그래프 중에 2번째
plt.plot(epochs, acc, 'ro', label='Training acc')
plt.plot(epochs, val_acc, 'r', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
# model.evalueate를 통해 테스트 데이터로 정확도 확인하기.
## model.evaluate(X_test, Y_test)의 리턴값은 [loss, binary_acuuracy ] -> 위 model.compile에서 metrics=[ keras.metrics.binary_accuracy]옵션을 주어서 binary acuuracy 출력됨.
print("\n Test Accuracy: %.4f" % (model.evaluate(X_test, Y_test)[1]))
model.save('iris_multi_model.h5')
## 그래프 띄우기
plt.show()
| tensorflow/day3/practice/P_03_05_iris_multi_classification_retrain.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# + [markdown] colab_type="text" id="EVPW4kLCmaSc"
# ## Day 20 作業:將 missing 值替代為其他值
#
# 由於現實世界中的資料可能有缺漏,因此資料中就會有 missing 值存在,而在資料科學及機器學習中,缺漏值的處理關係到分析或預測的結果。
#
# 在今天的作業中,請將資料集中的年齡缺漏值,依性別分別替換為資料集中男性或女性的年齡平均值。
#
# 本次作業使用的資料集:Kaggle Titanic: Machine Learning from Disaster 的 train 資料集
#
# 競賽主頁:[https://www.kaggle.com/c/titanic/overview](https://www.kaggle.com/c/titanic/overview)
#
# 請自行下載資料集 **train.csv**:[https://www.kaggle.com/c/titanic/data](https://www.kaggle.com/c/titanic/data)
# + colab={} colab_type="code" id="eeWlfZJSmaSe"
using DataFrames, CSV
# + colab={} colab_type="code" id="IkfIcUkjmaTS" outputId="7c455008-ef6c-410c-a8ba-1ad7c6edbb2d"
df = CSV.read("../data/train.csv")
# + colab={} colab_type="code" id="otCGP_AtmaTm" outputId="3ff0bb8d-1103-47d8-fe43-47f18622f8c5"
size(df)
# + [markdown] colab_type="text" id="3JwkiUhYmaT5"
# 呼叫 `describe()` 函式時,加上 `:nmissing` 參數,顯示各 column 的 missing 值數目。
#
# 可以看到 Age 共有 177 個缺漏值。
# + colab={} colab_type="code" id="QWT4tm45maT7" outputId="d0c6a1ae-b249-4a32-9284-c3ffd4910b96"
describe(df, :nmissing)
# + [markdown] colab_type="text" id="feAkzNA-maUC"
# 計算男性及女性的平均年齡。
#
# 【提示】使用 `skipmissing()` 去掉缺漏值後,再計算平均值。
# + colab={} colab_type="code" id="wRiM0Gq_maUD"
using Statistics
# -
male = df[df.Sex.=="male", :Age]
female = df[df.Sex.=="female", :Age]
# + colab={} colab_type="code" id="54yYKzADmaUM"
male_age_avg = round(mean(skipmissing(male)))
# + colab={} colab_type="code" id="4w92N00-maUT"
female_age_avg = round(mean(skipmissing(female)))
# + [markdown] colab_type="text" id="-O9fseP7maUa"
# 將 Age column 中的 missing 值,依性別替換為上面計算出來的平均年齡值。
#
# 【提示】可產生新的 column 來存放新的結果,而不取代掉原先的 Age column。
# -
function is_male(df, index)
if df[index, :Sex] == "male"
return true
else
return false
end
end
# + colab={} colab_type="code" id="UEtMiJrOmaUb"
age_col = []
# 請輸入程式碼
for (index, val) in enumerate(df.Age)
if is_male(df, index)
push!(age_col, coalesce(val, male_age_avg))
else
push!(age_col, coalesce(val, female_age_avg))
end
end
age_col
# -
df[:, :NewAge] = age_col
# + colab={} colab_type="code" id="szWNK43vmaUh"
describe(df)
| homework/julia_020_hw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# visual confirmation of ICs
# get 30 ICs
import nipype.interfaces.io as nio
import os
PD_ICA_file='/templateflow/PD_ICAs';
ICs_list=list(range(30))
ICs_list=["{:0>4d}".format(x) for x in ICs_list]
# file import
ds_ic = nio.DataGrabber(infields=['IC_id'])
ds_ic.inputs.base_directory = PD_ICA_file # database
ds_ic.inputs.template = 'melodic_IC_%4s.nii.gz' # from cwd
ds_ic.inputs.sort_filelist = True
ds_ic.inputs.IC_id = ICs_list
res_ic = ds_ic.run()
ic_list=res_ic.outputs.outfiles
atlas_09_masked='/templateflow/tpl-MNI152NLin2009cAsym/tpl-MNI152NLin2009cAsym_res-02_T1w.nii.gz'
GROUPS=['PD','ET','NC']
OUT_DIR='/output/PD_ICA/'
SUB_LIST=[]; AGE_LIST=[]; JCOB_LIST=[];
for group_name in GROUPS:
current_group=group_name
current_sub_list_file = '/codes/devel/PD_Marker/'+current_group+'_info_ICA.list'
# create dir for output
current_OUT_DIR=OUT_DIR+current_group+'/'
if not os.path.exists(current_OUT_DIR):
os.makedirs(current_OUT_DIR)
#read sub list
with open(current_sub_list_file, 'r') as f_sub:
sub_list_raw= f_sub.readlines()
sub_list = [x[0:-1].split('\t')[0] for x in sub_list_raw] # remove
age_list = [int(x[0:-1].split('\t')[1]) for x in sub_list_raw]
SUB_LIST.append(sub_list); AGE_LIST.append(age_list);
N_sub=len(sub_list)
print(group_name, ': ', N_sub)
# grab group Jacobians
ds_jacobian = nio.DataGrabber(infields=['sub_id'])
ds_jacobian.inputs.base_directory = current_OUT_DIR # database
ds_jacobian.inputs.template = '%s_desc-preproc_T1w_space-MNI2009c_Warp_Jacobian.nii.gz' # from cwd
ds_jacobian.inputs.sort_filelist = True
ds_jacobian.inputs.sub_id = sub_list
res_jacobian = ds_jacobian.run()
jacobian_list=res_jacobian.outputs.outfiles
JCOB_LIST.append(jacobian_list)
pd_sub_list = SUB_LIST[0]; et_sub_list = SUB_LIST[1]; nc_sub_list = SUB_LIST[2];
pd_age_list = AGE_LIST[0]; et_age_list = AGE_LIST[1]; nc_age_list = AGE_LIST[2];
pd_jaco_list=JCOB_LIST[0]; et_jaco_list=JCOB_LIST[1]; nc_jaco_list=JCOB_LIST[2];
# +
# corr IC * subj
import nibabel as nib
import time
ICx_OUT_DIR = '/output/PD_ICA/IC_Ximg/'
N_IC=len(ic_list); N_PD=len(pd_sub_list); N_ET=len(et_sub_list) ;N_NC=len(nc_sub_list);
nc_jaco_x_list=[]
i_PD_IC=0
PD_ICA_img=nib.load(ic_list[i_PD_IC]);
def grab_group_corr_ICA_all_sv(ic_list, img_list, id_list, out_dir):
import nibabel as nib
from nilearn.image import resample_to_img
from nilearn.image import math_img
from scipy import stats
import numpy as np
x_list=[];
N_ic=len(ic_list);
N_sub=len(img_list);
#print('gourp++')
for j in range(N_ic):
y_list=[]
for i in range(N_sub):
# cal img * img
ic_img=nib.load(ic_list[j])
sub_img=nib.load(img_list[i])
sub_img_re = resample_to_img(sub_img, ic_img)
#sub_img_x = math_img("img1 * img2", img1=ic_img, img2=sub_img_re)
#nib.save(sub_img_x, out_dir+id_list[i]+'-xIC'+str(j)+'.nii.gz');
# cal naive corr
ic_data = ic_img.get_fdata().reshape(-1);
nz_pos=np.flatnonzero(ic_data)
ic_val=list(ic_data.ravel()[nz_pos])
sub_dat = sub_img_re.get_fdata().reshape(-1);
sub_val=list(stats.zscore(sub_dat.ravel()[nz_pos]))
#_list.append([sub_img_x, np.corrcoef(sub_val, ic_val)[0,1]])
y_list.append(np.corrcoef(sub_val, ic_val)[0,1])
x_list.append(y_list)
return x_list
##
# just look at ic7
ic7_list=[ic_list[i_PD_IC]]
N_IC=len(ic_list); N_PD=len(pd_sub_list); N_ET=len(et_sub_list) ;N_NC=len(nc_sub_list);
t0=time.time()
nc_jaco_corr_list=grab_group_corr_ICA_all_sv(ic7_list, nc_jaco_list, nc_sub_list, ICx_OUT_DIR)
print('NC group corr IC takes: ', str(time.time()-t0))
pd_jaco_corr_list=grab_group_corr_ICA_all_sv(ic7_list, pd_jaco_list, pd_sub_list, ICx_OUT_DIR)
print('NC+PD group corr IC takes: ', str(time.time()-t0))
et_jaco_corr_list=grab_group_corr_ICA_all_sv(ic7_list, et_jaco_list, et_sub_list, ICx_OUT_DIR)
print('all group corr IC takes: ', str(time.time()-t0))
import pandas as pd
ic7_corr=pd_jaco_corr_list[0]+et_jaco_corr_list[0]+nc_jaco_corr_list[0]
ic_corr_lable=['PD']*N_PD+['ET']*N_ET+['NC']*N_NC
group_corr=pd.DataFrame({'IC Correlation': ic7_corr, 'Study Group': ic_corr_lable})
group_corr.to_csv('group_ic-sub_corr_ic0.csv')
print(group_corr)
# -
# plot and sv Dagher 30 ICs
from nilearn import plotting
GROUPS=['PD','ET','NC']
OUT_DIR='/output/PD_ICA/figs'
atlas_09_masked='/templateflow/tpl-MNI152NLin2009cAsym/tpl-MNI152NLin2009cAsym_res-02_T1w.nii.gz'
N_ic = len(ic_list)
i_ic=7
VMIN= 3; VMAX=16; # vmin=VMIN;
#plotting.plot_glass_brain(ic_list[i_ic], title='no th '+str(i_ic)+', with TH=0', display_mode='lyrz',
# black_bg=True, colorbar=True, plot_abs=False, vmin=VMIN, vmax=VMAX, threshold=5)
plotting.plot_stat_map(ic_list[i_ic], bg_img=atlas_09_masked, display_mode='z', threshold=3, vmax=VMAX, \
title="PD-ICA Axial", draw_cross=False, cut_coords=[-12, -7, 20], \
output_file=OUT_DIR+'/PD-ICA_Axial.png')
plotting.plot_stat_map(ic_list[i_ic], bg_img=atlas_09_masked, display_mode='x', threshold=3, vmax=VMAX, \
title="PD-ICA Sagittal", draw_cross=False, cut_coords=[-12, -7, 20], \
output_file=OUT_DIR+'/PD-ICA_Sagittal.png')
plotting.plot_stat_map(ic_list[i_ic], bg_img=atlas_09_masked, display_mode='y', threshold=3, vmax=VMAX, \
title="PD-ICA Coronal", draw_cross=False, cut_coords=[-12, -7, 20], \
output_file=OUT_DIR+'/PD-ICA_Coronal.png')
#plot group corr
import seaborn as sns
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
group_corr = pd.read_csv('group_ic-sub_corr_ic0.csv', sep=',', header=0, index_col=0)
#group_corr=pd.DataFrame({'IC Correlation': ic7_corr, 'Study Group': ic_corr_lable})
plt.figure(1, figsize=(8, 6))
ax = sns.violinplot(x="Study Group", y="IC Correlation", data=group_corr, inner='point')
plt.ylim(-0.35, 0.35)
plt.savefig('group_corr_ic0.png')
# +
import matplotlib.pyplot as plt
import numpy as np
plt.style.use({'figure.figsize':(12, 8)})
vmin1=-.2; vmax1 = .2
df=[df_1, df_2, df_3];
# Fixing random state for reproducibility
#np.random.seed(19680801)
fig, axs = plt.subplots(1, 3)
cm = ['RdBu_r', 'viridis']
ax2 = axs[1]
pcm2 = ax2.imshow(df_1, cmap=cm[0], vmin= vmin1, vmax = vmax1)
ax2.set_title("y=IC, x=NC subjects")
#fig.colorbar(pcm1, ax=ax1)
ax1 = axs[0]
pcm1 = ax1.imshow(df_2, cmap=cm[0], vmin= vmin1, vmax = vmax1)
ax1.set_title("y=IC, x=PD subjects")
ax3 = axs[2]
pcm3 = ax3.imshow(df_3, cmap=cm[0], vmin= vmin1, vmax = vmax1)
ax3.set_title("y=IC, x=ET subjects")
fig.colorbar(pcm3, ax=ax3)
plt.show()
| codes/devel/PD_Marker/fig1_ADIC_in_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: qic
# language: python
# name: qic
# ---
# <img src="https://s3-ap-southeast-1.amazonaws.com/he-public-data/wordmark_black65ee464.png" width="700">
# # Classical Support Vector Machines
#
# This notebook will serve as a summary of some of the resources below and is not meant to be used as a stand-alone reading material for Classical Support Vector Machines. We encourage you to complete reading the resources below before going forward with the notebook.
#
# ### Resources:
# 1. MIT Open Courseware lecture: https://youtu.be/_PwhiWxHK8o
# 3. MIT lecture slides: http://web.mit.edu/6.034/wwwbob/svm-notes-long-08.pdf
# 4. SVM Wikipedia page : https://en.wikipedia.org/wiki/Support_vector_machine
# 2. SVM tutorial using sklearn: https://jakevdp.github.io/PythonDataScienceHandbook/05.07-support-vector-machines.html
# ## Contents
#
# 1. [Introduction](#intro)
# 2. [SVMs as Linear Classifiers](#linear)
# 3. [Lagrange Multipliers and the Primal and Dual form](#primal)
# 4. [Class Prediction For a New Datapoint](#pred)
# 5. [Classifying Linearly Separable Data](#class-linear)
# 6. [Dealing With Non-Linearly Separable Data](#non-linear)
# 7. [Feature Map and Kernel](#kernel)
# 8. [Additional Resources](#add)
# ## Introduction <a id="intro"></a>
# +
# installing a few dependencies
# !pip install --upgrade seaborn==0.10.1
# !pip install --upgrade scipy==1.4.1
# !pip install --upgrade scikit-learn==0.23.1
# !pip install --upgrade matplotlib==3.2.0
# the output will be cleared after installation
from IPython.display import clear_output
clear_output()
# -
# Suppose you are a Botanist trying to distinguish which one of the **three species** a flower belongs to just by looking at **four features** of a flower - the length and the width of the sepals and petals. As part of your research you create a **dataset** of these features for a set of flowers for which the **species is already known**, where each **datapoint** of this dataset corresponds to a single flower. Now, your colleague brings in a new flower and asks you which species it belongs to. You could go into the lab and do the necessary tests to figure out what species it is, however, the lab is under renovation. So, left with no other choice you pull up the dataset that you created earlier and after a few minutes of trying to find a pattern you realise that this new flower has a petal width and sepal length similar to all the flowers of species 1. Thus, you **predict** this new flower to be of the species 1. This process of assigning a new datapoint to one of the known **classes** (flower species) is called **classfication**. And, as we used a dataset where we knew the classes corresponding to the datapoints before-hand, thus, this classification procedure comes under the umbrella of [**supervised learning**](https://en.wikipedia.org/wiki/Supervised_learning).
#
#
# Support Vector Machines (SVMs) are **supervised learning models** that are mainly used for **classification** and **regression** tasks. In the context of classification, which is the topic of discussion, we use SVMs to find a **linear decision boundary with maximum width** splitting the space such that datapoints belonging to different classes are on either side of the boundary. Classification takes place based on which side of the decision boundary a new datapoint lands.
#
#
# Before we try to understand how SVMs work, let's take a look at the [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set) which was the dataset mentioned in the first paragraph.
# +
# importing the iris dataset
from sklearn.datasets import load_iris
import numpy as np
iris = load_iris()
print("Number of datapoints: {}".format(iris['data'].shape[0]))
print("Number of features: {}".format(iris['data'].shape[1]))
print("Sample of the dataset:")
print(iris['data'][:5])
print("Unique species : {}".format(np.unique(iris['target'])))
# -
# Looking at the first 5 datapoints of the Iris dataset we realize that each datapoint is an array with four features. The number of features in a dataset is called the **dimension of the dataset**. Further, there are three unique species, which implies, three **classes** in the dataset. It's important to note that SVMs are natively binary classification algorithms, i.e, can classify between only 2 classes. However, there are methods to convert a binary classifier to a multi-class classifier, mentioned [here](https://datascience.stackexchange.com/questions/46514/how-to-convert-binary-classifier-to-multiclass-classifier). Let us now dig deeper into the mathematics of how SVMs work.
#
# **Reminder:** Read the resources provided above to understand the next section with greater degree of clarity.
# ## SVMs as Linear Classifiers <a id="linear"></a>
# 
# Source: [wikipedia](https://en.wikipedia.org/wiki/Support_vector_machine)
# Our input dataset is of the form $(\vec{x}_{1}, y_{1}), ..., (\vec{x}_{n}, y_{n})$,
# where, $\vec{x}$ is a $d$ dimensional vector where $d$ is the number of features and $y_{i}$'s are the labels $ y_{i} \in {-1, +1}$ as it is a binary classification problem.
#
# $\vec{w}$ is a vector perpendicular to the **decision boundary** (hyperplane that cuts the space into two parts and is the result of classification). Let $\vec{u}$ be a vector representing a point on our feature space. Now, to understand whether a point is on the +ve side or the -ve side we'll have to project the point $\vec{u}$ onto $\vec{w}$, which will give us a scaled version of $\vec{u}$'s projection in the direction perpendicular to the decision boundary. Depending on the value of this quantity we'll have the point either on the +ve side or the -ve side. This can be represented mathematically as
#
#
# $$\begin{equation} \vec{w}\cdot\vec{x}_{+} + b \geq 1 \label{eq:vector_ray} \tag{1}\end{equation}$$
# $$\begin{equation} \vec{w}\cdot\vec{x}_{-} + b \leq -1 \tag{2}\end{equation}$$
#
# where, $\vec{x}_{+}$ is a datapoint with label $y_{i} = +1$,<br>
# $\vec{x}_{-}$ is a datapoint with label $y_{i} = -1$ and<br>
# b is parameter that has to be learnt
#
# These two lines are separated by a distance of $\frac{2}{||{\vec{w}}||}$. The line in the middle of both of these, i.e,
#
# $$\begin{equation} \vec{w}\cdot\vec{u} + b = 0 \tag{3}\end{equation}$$
#
# is the equation of the hyperplane denoting our decision boundary. Together, the space between (1) and (2) forms what is usually know as the **street** or the **gutter**.
#
# equation (1) and (2) can be conveniently combined to give
#
# $$y_{i}(\vec{w}\cdot\vec{x}_{i} + b) \geq 1\tag{4}$$
#
# And the limiting case would be
#
# $$y_{i}(\vec{w}\cdot\vec{x}_{i} + b) -1 = 0 \tag{5}$$
#
# Which is attained when the points lie on the edges of the street, i.e, on (1) or (2). These points are responsible for the change in the width of the street and are called **support vectors**. Once the support vectors are calculated in the training phase we only need these vectors to classify new datapoints during the prediction phase, hence reducing the computational load significantly. Equation (4) is a constraint in the optimization process of maximizing the street width $\frac{2}{||{\vec{w}}||}$. In the next section let us see how we can combine the optimization problem and the constraints together into a single optimization equation using the concept of lagrange multipliers.
# ## Lagrange Multipliers and the Primal and Dual form <a id="primal"></a>
# Support Vector Machines are trying to solve the optimization problem of maximizing the street width $\frac{2}{||{\vec{w}}||}$ (which is equivalent to minimizing $\frac{||w||^2}{2}$) with the contraint $y_{i}(\vec{w}\cdot\vec{x}_{i} + b) \geq 1$. This can be elegantly written in a single equation with the help of [lagrange multipliers](https://en.wikipedia.org/wiki/Lagrange_multiplier). The resulting equation to be minimized is called the **primal form** (6).
#
# **Primal form:** $$ L_{p} = \frac{||w||^2}{2} - \sum_{i}{\alpha_{i}[y_{i}(\vec{w}\cdot\vec{x}_{i} + b) -1]}\tag{6}$$
#
# $$\frac{{\partial L}}{\partial \vec{w}} = \vec{w} - \sum_{i}{\alpha_{i}y_{i}\vec{x_{i}}}$$
#
# equating $\frac{{\partial L}}{\partial \vec{w}}$ to 0 we get,
#
# $$ \vec{w} = \sum_{i}{\alpha_{i}y_{i}\vec{x_{i}}}\tag{7}$$
#
# $$\frac{{\partial L}}{\partial \vec{b}} = \sum_{i}{\alpha_{i}y_{i}}$$
#
# and equating $\frac{{\partial L}}{\partial \vec{b}}$ to 0 we convert the primal form to the dual form,
#
# $$\sum_{i}{\alpha_{i}y_{i}} = 0\tag{8}$$
#
# $$L = \frac{1}{2}(\sum_{i}{\alpha_{i}y_{i}\vec{x_{i}}})(\sum_{j}{\alpha_{j}y_{j}\vec{x_{j}}}) - (\sum_{i}{\alpha_{i}y_{i}\vec{x_{i}}})(\sum_{j}{\alpha_{j}y_{j}\vec{x_{j}}}) - \sum_{i}{\alpha_{i}y_{i}b} + \sum_{i}{\alpha_{i}}$$
#
# **Dual form:** $$L_{d} = \sum_{i}{\alpha_{i}} - \frac{1}{2}\sum_{i}\sum_{j}\alpha_{i}\alpha_{j}y_{i}y_{j}(\vec{x}_{i}\cdot\vec{x}_{j})\tag{9}$$
# subject to: $$\sum_{i}{\alpha_{i}y_{i}} = 0$$
# Taking a closer look at the dual form $L_{d}$ we can see that it is a function quadratic in the lagrange multipler terms which can be solved efficiently on a classical computer using [quadratic programming](https://en.wikipedia.org/wiki/Quadratic_programming) techniques. However, Note that finding the dot product $\vec{x}_{i}\cdot\vec{x}_{j}$ becomes computationally expensive as the dimension of our data increases. In the days to come we'll learn how a quantum computer could be used to classify a classical dataset using an algorithm called the Variational Quantum Classifier (VQC) Algorithm as given in [this paper](https://arxiv.org/abs/1804.11326). Understanding of Classical SVM may not be required, however, some of the concepts such as kernels and feature maps will be crucial in understanding the VQC algorithm.
# ## Class Prediction for a New Datapoint <a id="pred"></a>
# The output of the training step are values of lagrange multipliers. Now, when a new datapoint $\vec z$ is given lets see how we can find the classification result corresponding to it:
#
# * Step 1: Use the obtained values of lagrange multipliers to calculate the value of $\vec{w}$ using $(7)$.
# * Step 2: Substitute the value of $\vec{w}$ in equation $(5)$ and substitute a support vector in the place of $\vec{x}_{i}$ to find the value of $b$.
# * Step 3: Find the value of $\vec{w}\cdot\vec{z} + b$. If it $>0$ then assign $\vec{z}$ a label $y_{z} = 1$ and $y_{z} = -1$ if the obtained value is $< 0$.
# ## Classifying Linearly Separable Data <a id="class-linear"></a>
# Lets switch gears and look at how we can use scikit-learn's Support Vector Classifier method to draw a decision boundary on a linearly separable dataset. This section of the notebook is a recap of resource \[4\] and thus we recommend reading it before going forward. The code used in this section is from the corresponding Github [repo](https://github.com/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/05.07-Support-Vector-Machines.ipynb).
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
# use seaborn plotting defaults
import seaborn as sns; sns.set()
# -
# ### Importing the dataset
# we are importing the make_blobs dataset as it can be clearly seen to be linearly separable
from sklearn.datasets.samples_generator import make_blobs
X, y = make_blobs(n_samples=50, centers=2,
random_state=0, cluster_std=0.60)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='autumn');
from sklearn.svm import SVC # "Support vector classifier"
model = SVC(kernel='linear', C=1E10)
model.fit(X, y)
# helper plotting function
def plot_svc_decision_function(model, ax=None, plot_support=True):
if ax is None:
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
x = np.linspace(xlim[0], xlim[1], 30)
y = np.linspace(ylim[0], ylim[1], 30)
Y, X = np.meshgrid(y, x)
xy = np.vstack([X.ravel(), Y.ravel()]).T
P = model.decision_function(xy).reshape(X.shape)
# plot decision boundary and margins
ax.contour(X, Y, P, colors='k',
levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
if plot_support:
ax.scatter(model.support_vectors_[:, 0],
model.support_vectors_[:, 1],
s=300, linewidth=1, facecolors='none', edgecolors='b');
ax.set_xlim(xlim)
ax.set_ylim(ylim)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='autumn')
plot_svc_decision_function(model);
# As we can see SVM works quite well when it deals with linearly separable datasets. The points which lie on the dotted lines denoted by $y_{i}(\vec{w}\cdot\vec{x}_{i} + b) = \pm1$ are the **support vectors**. Part of the reason why SVMs are popular are because, during the classification step only support vectors are used to classify a new point. This reduces the computational load significantly. The reason for this is because values of the lagrange multipliers turn out to be zero for vectors which are not support vectors.
model.support_vectors_
# ## Dealing With Non-Linearly Separable Data <a id="non-linear"></a>
# In the previous example we've seen how we can find a model to classify linearly separable data. Lets look at an example and see if SVMs can find a solution when the data is non-linear.
# +
from sklearn.datasets.samples_generator import make_circles
X, y = make_circles(100, factor=.1, noise=.1)
clf = SVC(kernel='linear').fit(X, y)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='autumn')
plot_svc_decision_function(clf, plot_support=False);
# -
# When the data is circular, like in the example above, SVM fails to find a satisfactory linear classification model. However, if we cleverly introduce a new parameter $r$ such that $r = e^{-x^{2}}$ and using that as our third parameter construct a new dataset (see picture below), we'll observe that a plane can be drawn horizontally passing through, say, $r=0.7$ to classify the dataset! This method in which we are mapping our dataset into a higher dimension to be able to find a linear boundary in the higher dimension is called a **feature map**.
r = np.exp(-(X ** 2).sum(1))
# +
from mpl_toolkits import mplot3d
# from ipywidgets import interact, fixed
def plot_3D(elev=30, azim=30, X=X, y=y):
ax = plt.subplot(projection='3d')
ax.scatter3D(X[:, 0], X[:, 1], r, c=y, s=50, cmap='autumn')
ax.view_init(elev=elev, azim=azim)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('r')
plot_3D()
# interact(plot_3D, elev=[-90, 90], azip=(-180, 180),
# X=fixed(X), y=fixed(y));
# -
# ## Feature Map and Kernel <a id="kernel"></a>
# As we have seen earlier a **feature map** maps our (non-linearly separable) input data to a higher dimensional **feature space** where our data is now linearly separable. This helps circumvent the problem of dealing with non-linearly separable data, however, a new problem arises. As we keep increasing the dimension of our data, computing the coordinates of our data and the dot product $\phi(\vec{x}_{i})\cdot\phi(\vec{x}_{j})$ in this higher dimentional feature space becomes computationally expensive. This is where the idea of the [Kernel functions](https://en.wikipedia.org/wiki/Kernel_method) comes in.
#
# Kernel functions allow us to deal with our data in the higher dimensional feature space (where our data is linearly separable) without ever having to compute the dot product in that space.
#
# if $\phi(\vec{x})$ is the feature map, then the corresponding kernel function is the dot product $\phi(\vec{x}_{i})\cdot\phi(\vec{x}_{j})$, therefore, the kernel function $k$ is
#
# $$k(x_{i},x_{j}) = \phi(\vec{x}_{i})\cdot\phi(\vec{x}_{j})$$
#
# Therefore, the corresponding transformed optimization problem can we written as,
#
# **Primal form:** $$ L_{p} = \frac{||w||^2}{2} - \sum_{i}{\alpha_{i}[y_{i}(\vec{w}\cdot\phi(\vec{x}_{i}) + b) -1]}\tag{6}$$
#
# **Dual form:** $$L_{d} = \sum_{i}{\alpha_{i}} - \frac{1}{2}\sum_{i}\sum_{j}\alpha_{i}\alpha_{j}y_{i}y_{j}(\phi(\vec{x}_{i})\cdot\phi(\vec{x}_{j}))$$
# or $$L_{d} = \sum_{i}{\alpha_{i}} - \frac{1}{2}\sum_{i}\sum_{j}\alpha_{i}\alpha_{j}y_{i}y_{j}k(x_{i},x_{j})$$
# subject to: $$\sum_{i}{\alpha_{i}y_{i}} = 0$$
# where $$ \vec{w} = \sum_{i}{\alpha_{i}y_{i}\phi(\vec{x_{i}})}$$
# To understand why Kernel functions are useful lets look at an example using the Radial Basis Function (rbf) Kernel.
#
# the rbf kernel is written as,
#
# $$k(x_{i},x_{j}) = exp(-||x_{i} - x_{j}||^{2}/2\sigma^{2}) $$
#
# where $\sigma$ is a tunable parameter
#
# What we should understand here is that the rbf kernel projects our data into an infinite dimensional feature space, however, the computational power required to compute the kernel function's value is quite negligible! As you see, we don't have to compute the dot product of the infinite dimensional vectors. This is how kernels help SVMs tackle non-linearly separable data.
#
# Rbf kernel in action:
clf = SVC(kernel='rbf', C=1E6)
clf.fit(X, y)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='autumn')
plot_svc_decision_function(clf)
plt.scatter(clf.support_vectors_[:, 0], clf.support_vectors_[:, 1],
s=300, lw=1, facecolors='none');
# In the next notebook we will learn how to use Quantum Computers to do the same task of classification and why it may be advantageous in the future.
# ## Additional Resources <a id="add"></a>
# 1. Andrew NG notes: http://cs229.stanford.edu/notes/cs229-notes3.pdf
# 2. Andrew NG lecture: https://youtu.be/lDwow4aOrtg
| Day 5/Classical Support Vector Machines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Let's start by importing the libraries
#Importing NumPY and Pandas Library
import numpy as np
import pandas as pd
#Importing Data Visualization Libraries
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ### Read the dataset
result = pd.read_csv("StudentsPerformance.csv")
# # Let's start exploring the data
result.head()
# ### Lets change the column name according to our convenience
result.columns = map(str.upper, result.columns)
result.head()
# ### Checking the type of data in each column
result.info()
# ### Let's check for null values in each column
result.isna().sum()
# ### Statistical Analysis on the data
result.describe()
# #### The lowest marks in Math is 0, in Reading is 17, and writing is 10 !
# #### The highest in all the 3 subjects is 100 !
# ### Lets analyse the marks of the students in Math, Reading and Writing
sns.pairplot(result, hue = 'GENDER', palette = 'coolwarm')
# ### Heatmap for our data
# Matrix form for correlation data
result.corr()
sns.heatmap(result.corr(), cmap = 'PuRd', annot = True)
# #### We can interpret that reading and writing score are highly correlated, while the writing score and math score is the least, compared to others!
# ### Lets add an "AVERAGE" column to the dataset
result["AVERAGE"] = (result["MATH SCORE"] + result["READING SCORE"] + result["WRITING SCORE"])/3
result.head()
# ### Lets find out the relation between Reading and Writing Score
sns.lmplot(x='READING SCORE',y='WRITING SCORE',data=result,hue='GENDER')
# ### Let's find out how Gender affects Ethnicity and Math Scores
result.pivot_table(values='MATH SCORE',index='GENDER',columns='RACE/ETHNICITY')
pvresult = result.pivot_table(values='MATH SCORE',index='GENDER',columns='RACE/ETHNICITY')
sns.heatmap(pvresult, annot = True)
# #### We can see that Group E Males have the highest average Math Scores, while Group A Females have the least!
# ### Similarly lets analyse for Reading Scores
result.pivot_table(values='READING SCORE',index='GENDER',columns='RACE/ETHNICITY')
pvresult = result.pivot_table(values='READING SCORE',index='GENDER',columns='RACE/ETHNICITY')
sns.heatmap(pvresult,cmap='YlOrRd',linecolor='white',linewidths=1, annot = True)
# #### We can see that Group E females on an average, fair better than others in Reading!
# ### Now, for Writing Scores
result.pivot_table(values='WRITING SCORE',index='GENDER',columns='RACE/ETHNICITY')
pvresult = result.pivot_table(values='WRITING SCORE',index='GENDER',columns='RACE/ETHNICITY')
sns.heatmap(pvresult, cmap = 'YlGnBu',linecolor='black',linewidths=1, annot = True)
# #### We can interpret that Group A Males have the least writing skills while Group E females have the most!
# ### Lets analyse the average scores now
result.pivot_table(values='AVERAGE',index='GENDER',columns='RACE/ETHNICITY')
pvresult = result.pivot_table(values='AVERAGE',index='GENDER',columns='RACE/ETHNICITY')
sns.heatmap(pvresult, cmap = 'Reds', annot = True)
# #### On an average, we can say that Group E Females score the best, while Group A males, least!
# ### Lets see the percentage distribution of Males and Females in the Dataset
(result.GENDER.value_counts()/len(result)) * 100
gender = result['GENDER'].value_counts()
labels = result.GENDER.unique()
plt.pie(gender,labels=labels,autopct="%1.1f%%",shadow=True,explode=(0.04,0.04),startangle=90)
plt.title('GENDER DISTRIBUTION',fontsize=15)
plt.show()
result.GENDER.value_counts()
sns.countplot(x='GENDER', data=result, palette = 'magma')
# #### We can see that the gender distribution is almost 50-50 !
gender = result.groupby("GENDER")
gender.mean()
gender.describe().transpose()
# #### Therefore, it is safe to assume that Males are slightly better than Females in Math, while Females outscore Males in Reading and Writing !
# ### Finding out the percentage of students who have taken Test Preparation Course Prior to taking Tests
(result['TEST PREPARATION COURSE'].value_counts()/len(result)) * 100
test = result['TEST PREPARATION COURSE'].value_counts()
labels = result["TEST PREPARATION COURSE"].unique()
plt.pie(test,labels=labels,autopct="%1.1f%%",shadow=True,explode=(0.04,0.04),startangle=90)
plt.title('TEST PREPARATION COURSE',fontsize=15)
plt.show()
tpc = result.groupby("TEST PREPARATION COURSE")
tpc.mean()
# #### We can say that Test Preparation Course has definitely improved the scores of students!
# ### Now, lets see how Test Preparation Course has helped students in improving their Test Scores, Gender wise
fig, ax = plt.subplots(1, 3, figsize=(16,4))
sns.violinplot(x="TEST PREPARATION COURSE", y='MATH SCORE', data=result,hue='GENDER',split=True,palette='PuRd', ax = ax[0])
sns.violinplot(x="TEST PREPARATION COURSE", y='READING SCORE', data=result,hue='GENDER',split = True,
palette='Purples', ax = ax[1])
sns.violinplot(x="TEST PREPARATION COURSE", y='WRITING SCORE', data=result,hue='GENDER',split = True,
palette='RdPu', ax = ax[2])
# ### Lets see how Test Preparation Course has helped to improve the average marks of the students
sns.boxplot(x="TEST PREPARATION COURSE", y="AVERAGE", hue = "GENDER", data = result)
# #### We can see that definitely, Test Preparation Course has helped improve their scores!
# ### Does a Parent's Level of Education influence the student's performance? Lets find out!
p_edu = result.groupby("PARENTAL LEVEL OF EDUCATION")
p_edu.mean()
# +
fig, ax = plt.subplots(3, 1, figsize=(16,16))
sns.boxplot(x = 'PARENTAL LEVEL OF EDUCATION', y = 'MATH SCORE', data = result, ax = ax[0], palette = "magma")
sns.boxplot(x = 'PARENTAL LEVEL OF EDUCATION', y = 'READING SCORE', data = result, ax = ax[1], palette = "plasma")
sns.boxplot(x = 'PARENTAL LEVEL OF EDUCATION', y = 'WRITING SCORE', data = result, ax = ax[2], palette = "inferno")
# -
# ### Now, lets see how Parental Level of Education has affected the average scores
sns.boxplot(x="TEST PREPARATION COURSE", y='AVERAGE', data=result,hue='GENDER', palette='inferno')
# #### Yeah! Parental Level of Education does improve the scores of students!
# ### Lets find the count of students belonging to a particular Race/Ethnicity
# Lets find the percentage distribution
(result["RACE/ETHNICITY"].value_counts()/len(result)) * 100
sns.countplot(x='RACE/ETHNICITY', data=result, palette = 'Reds')
sns.despine()
# #### A majority of the students belong to Group C, while Group A has the least number of students!
sns.boxplot(x = 'RACE/ETHNICITY', y = 'AVERAGE', data = result, palette = "magma")
# #### Therefore, we can see that Group E students have a higher average than others!
# ### Lets see how the distribution Parental Level Of Education varies with Race/Ethnicity
plt.figure(figsize = (16,5))
sns.countplot(x="PARENTAL LEVEL OF EDUCATION", hue="RACE/ETHNICITY", data=result, palette='viridis')
# ### Lets find out the percentage of students who receive standard and reduced Lunch
(result["LUNCH"].value_counts()/len(result)) * 100
lunch = result['LUNCH'].value_counts()
labels = result["LUNCH"].unique()
plt.pie(test,labels=labels,autopct="%1.1f%%",shadow=True,explode=(0.04,0.04),startangle=90)
plt.title('LUNCH DISTRIBUTION',fontsize=15)
plt.show()
# Plotting the figures
fig, ax = plt.subplots(3, 1, figsize=(16,16))
sns.swarmplot(x="RACE/ETHNICITY", y='MATH SCORE', data=result,hue='LUNCH',palette='Purples', ax = ax[0])
sns.swarmplot(x="RACE/ETHNICITY", y='READING SCORE', data=result,hue='LUNCH', palette='Blues', ax = ax[1])
sns.swarmplot(x="RACE/ETHNICITY", y='WRITING SCORE', data=result,hue='LUNCH', palette='Greens', ax = ax[2])
# ### Lets see if Lunch affects the scores of students
p_edu = result.groupby("LUNCH")
p_edu.mean()
# #### Students with Standard Lunch seem to score better than those with Free/Reduced Lunch !
# ### Lets see how type of Lunch differs due to Race/Ethnicity
sns.countplot(x="RACE/ETHNICITY", hue="LUNCH", data=result, palette='Oranges')
# #### Group C receives the majority of free/reduced Lunches while Group A receives the least
# ### Is Free/Reduced Lunch Gender Biased? Lets find out!
sns.countplot(x="LUNCH", data=result,hue = 'GENDER', palette='YlGnBu')
# #### The number of females receiving Standard or Free/Reduced Lunch is higher in both the cases!
| Shreya_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] lc_cell_meme={"current": "a9c2e890-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2ea3e-495a-11e8-8185-0242ac130002", "previous": null}
# # About: GCE - Set! Go! (Google Compute Engine)
#
# ---
#
# Get and set a virtual machine, aka compute instance of "Google Compute Engine" with specified machine type and OS image. *[Here are just set and go becasue GCE is always ready for you..]*
#
# Google Compute Engine上で仮想マシンの確保する。
# + [markdown] lc_cell_meme={"current": "a9c2ea3e-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2ebba-495a-11e8-8185-0242ac130002", "previous": "a9c2e890-495a-11e8-8185-0242ac130002"}
# ## *Operation Note*
#
# *This is a cell for your own recording. ここに経緯を記述*
# + [markdown] lc_cell_meme={"current": "a9c2ebba-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2ed22-495a-11e8-8185-0242ac130002", "previous": "a9c2ea3e-495a-11e8-8185-0242ac130002"}
# # 動作環境の確認
#
# このNotebookは、 [Google Python Client Library](https://github.com/google/google-api-python-client) を使ってマシンの確保を行います。
#
# このNotebook環境にGoogle Python Client Libraryがインストールされている必要があります。インストールされていない場合は、以下のセル実行に失敗し、 `ImportError` となります。
# + lc_cell_meme={"current": "a9c2ed22-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2ee8a-495a-11e8-8185-0242ac130002", "previous": "a9c2ebba-495a-11e8-8185-0242ac130002"}
from googleapiclient import discovery
# + [markdown] lc_cell_meme={"current": "a9c2ee8a-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2eff2-495a-11e8-8185-0242ac130002", "previous": "a9c2ed22-495a-11e8-8185-0242ac130002"}
# もし、インストールされていない場合は、[Using the Python Client Library](https://cloud.google.com/compute/docs/tutorials/python-guide)を参考にインストールしてください。
# + lc_cell_meme={"current": "a9c2eff2-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f15a-495a-11e8-8185-0242ac130002", "previous": "a9c2ee8a-495a-11e8-8185-0242ac130002"}
# #!pip2 install --upgrade google-api-python-client
#from googleapiclient import discovery
# + [markdown] lc_cell_meme={"current": "a9c2f15a-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f2b8-495a-11e8-8185-0242ac130002", "previous": "a9c2eff2-495a-11e8-8185-0242ac130002"}
# # 設定の定義
#
# どのようなマシンを確保するか?を定義していく。
# + [markdown] lc_cell_meme={"current": "a9c2f2b8-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f420-495a-11e8-8185-0242ac130002", "previous": "a9c2f15a-495a-11e8-8185-0242ac130002"}
# ## Credentialの指定
#
# Google Compute EngineにアクセスするためのCredentialを指示する。
#
# - JSONフォーマットのService Account情報
# - プロジェクトID
#
# を用意しておく。
# + lc_cell_meme={"current": "a9c2f420-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f588-495a-11e8-8185-0242ac130002", "previous": "a9c2f2b8-495a-11e8-8185-0242ac130002"}
creds = <KEY>'
target_project_id = 'my_project_id'
# + [markdown] lc_cell_meme={"current": "a9c2f588-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f6e6-495a-11e8-8185-0242ac130002", "previous": "a9c2f420-495a-11e8-8185-0242ac130002"}
# computeサービスを取得しておく。
# + lc_cell_meme={"current": "a9c2f6e6-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f83a-495a-11e8-8185-0242ac130002", "previous": "a9c2f588-495a-11e8-8185-0242ac130002"}
import os
from oauth2client.client import GoogleCredentials
credentials = GoogleCredentials.from_stream(os.path.expanduser(creds))
compute = discovery.build('compute', 'v1', credentials=credentials)
compute
# + [markdown] lc_cell_meme={"current": "a9c2f83a-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2f9a2-495a-11e8-8185-0242ac130002", "previous": "a9c2f6e6-495a-11e8-8185-0242ac130002"}
# ## ゾーンの設定
#
# どのZoneにインスタンスを確保するかを定義しておく。
# + lc_cell_meme={"current": "a9c2f9a2-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2fb00-495a-11e8-8185-0242ac130002", "previous": "a9c2f83a-495a-11e8-8185-0242ac130002"}
target_zone = 'us-central1-f'
# + [markdown] lc_cell_meme={"current": "a9c2fb00-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2fc5e-495a-11e8-8185-0242ac130002", "previous": "a9c2f9a2-495a-11e8-8185-0242ac130002"}
# ## マシン名の決定
#
# マシン名を決める。まず、現在のインスタンス名の一覧を確認する。
# + lc_cell_meme={"current": "a9c2fc5e-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2fdbc-495a-11e8-8185-0242ac130002", "previous": "a9c2fb00-495a-11e8-8185-0242ac130002"}
instances = compute.instances().list(zone=target_zone, project=target_project_id).execute()
map(lambda i: i['name'], instances['items'] if 'items' in instances else [])
# + [markdown] lc_cell_meme={"current": "a9c2fdbc-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c2ff24-495a-11e8-8185-0242ac130002", "previous": "a9c2fc5e-495a-11e8-8185-0242ac130002"}
# すでにあるインスタンスとは重複しないような名前を設定する。
# + lc_cell_meme={"current": "a9c2ff24-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c30082-495a-11e8-8185-0242ac130002", "previous": "a9c2fdbc-495a-11e8-8185-0242ac130002"}
machine_name = 'test-gce'
# + [markdown] lc_cell_meme={"current": "a9c30082-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c301e0-495a-11e8-8185-0242ac130002", "previous": "a9c2ff24-495a-11e8-8185-0242ac130002"}
# ## マシンタイプの指定
#
# まず、このZoneで利用可能なMachine Typeを取得する。
# + lc_cell_meme={"current": "a9c301e0-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3033e-495a-11e8-8185-0242ac130002", "previous": "a9c30082-495a-11e8-8185-0242ac130002"}
machineTypes = compute.machineTypes().list(project=target_project_id, zone=target_zone).execute()['items']
map(lambda t: (t['name'], t['description']), machineTypes)
# + [markdown] lc_cell_meme={"current": "a9c3033e-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3049c-495a-11e8-8185-0242ac130002", "previous": "a9c301e0-495a-11e8-8185-0242ac130002"}
# 利用したいマシンタイプ名を設定する。
# + lc_cell_meme={"current": "a9c3049c-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3062c-495a-11e8-8185-0242ac130002", "previous": "a9c3033e-495a-11e8-8185-0242ac130002"}
machine_type = 'f1-micro'
# + [markdown] lc_cell_meme={"current": "a9c3062c-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3078a-495a-11e8-8185-0242ac130002", "previous": "a9c3049c-495a-11e8-8185-0242ac130002"}
# ## イメージの指定
#
# イメージの一覧を確認する。`project`には、利用したいイメージの所属プロジェクトを指定する。
# + lc_cell_meme={"current": "a9c3078a-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c308e8-495a-11e8-8185-0242ac130002", "previous": "a9c3062c-495a-11e8-8185-0242ac130002"}
images = compute.images().list(project='ubuntu-os-cloud').execute()['items']
images = filter(lambda i: i['status'] == 'READY' and 'deprecated' not in i, images)
map(lambda i: (i['name'], i['selfLink']), images)
# + [markdown] lc_cell_meme={"current": "a9c308e8-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c30a46-495a-11e8-8185-0242ac130002", "previous": "a9c3078a-495a-11e8-8185-0242ac130002"}
# 利用したいイメージのURLを設定する。
# + lc_cell_meme={"current": "a9c30a46-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c30ba4-495a-11e8-8185-0242ac130002", "previous": "a9c308e8-495a-11e8-8185-0242ac130002"}
source_disk_image = 'https://www.googleapis.com/compute/v1/projects/ubuntu-os-cloud/global/images/ubuntu-1404-trusty-v20160610'
# + [markdown] lc_cell_meme={"current": "a9c30ba4-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c30d02-495a-11e8-8185-0242ac130002", "previous": "a9c30a46-495a-11e8-8185-0242ac130002"}
# ## キーペアの設定
#
# 現在のSSHキーの一覧を取得する。
# + lc_cell_meme={"current": "a9c30d02-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c30e56-495a-11e8-8185-0242ac130002", "previous": "a9c30ba4-495a-11e8-8185-0242ac130002"}
projectMetadata = compute.projects().get(project=target_project_id).execute()
currentSSHKeys = filter(lambda metadata: metadata['key'] == 'sshKeys', projectMetadata['commonInstanceMetadata']['items']) \
if 'commonInstanceMetadata' in projectMetadata and 'items' in projectMetadata['commonInstanceMetadata'] else []
currentSSHKeys = currentSSHKeys[0]['value'].split('\n') if currentSSHKeys else []
currentSSHKeys
# + [markdown] lc_cell_meme={"current": "a9c30e56-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c30fb4-495a-11e8-8185-0242ac130002", "previous": "a9c30d02-495a-11e8-8185-0242ac130002"}
# SSHのキー一覧にこのNotebook環境のキーがなければ、追加する。
# + lc_cell_meme={"current": "a9c30fb4-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31112-495a-11e8-8185-0242ac130002", "previous": "a9c30e56-495a-11e8-8185-0242ac130002"}
pub_key = None
with open(os.path.expanduser('~/.ssh/ansible_id_rsa.pub'), 'r') as f:
pub_key = f.readlines()[0].strip()
if not filter(lambda k: k.endswith(pub_key), currentSSHKeys):
currentSSHKeys.append('ansible:' + pub_key)
currentSSHKeys
# + [markdown] lc_cell_meme={"current": "a9c31112-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31266-495a-11e8-8185-0242ac130002", "previous": "a9c30fb4-495a-11e8-8185-0242ac130002"}
# Metadataに反映する。
# + lc_cell_meme={"current": "a9c31266-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c313c4-495a-11e8-8185-0242ac130002", "previous": "a9c31112-495a-11e8-8185-0242ac130002"}
compute.projects().setCommonInstanceMetadata(project=target_project_id,
body={'items': [{'key': 'sshKeys',
'value': '\n'.join(currentSSHKeys)}]}).execute()
# + [markdown] lc_cell_meme={"current": "a9c313c4-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31518-495a-11e8-8185-0242ac130002", "previous": "a9c31266-495a-11e8-8185-0242ac130002"}
# # マシンの確保
# + [markdown] lc_cell_meme={"current": "a9c31518-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31676-495a-11e8-8185-0242ac130002", "previous": "a9c313c4-495a-11e8-8185-0242ac130002"}
# ## インスタンスの起動
#
# 設定した情報を用いてマシンを確保する。
# + lc_cell_meme={"current": "a9c31676-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c317ca-495a-11e8-8185-0242ac130002", "previous": "a9c31518-495a-11e8-8185-0242ac130002"}
config = {
'name': machine_name,
'machineType': "zones/{}/machineTypes/{}".format(target_zone, machine_type),
'disks': [
{
'boot': True,
'autoDelete': True,
'initializeParams': {
'sourceImage': source_disk_image,
}
}
],
'networkInterfaces': [{
'network': 'global/networks/default',
'accessConfigs': [
{'type': 'ONE_TO_ONE_NAT', 'name': 'External NAT'}
]
}],
'serviceAccounts': [{
'email': 'default',
'scopes': [
'https://www.googleapis.com/auth/devstorage.read_write',
'https://www.googleapis.com/auth/logging.write'
]
}],
'metadata': {
'items': []
}
}
new_vm = compute.instances().insert(project=target_project_id, zone=target_zone, body=config).execute()
new_vm
# + [markdown] lc_cell_meme={"current": "a9c317ca-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31928-495a-11e8-8185-0242ac130002", "previous": "a9c31676-495a-11e8-8185-0242ac130002"}
# インスタンスがRunningになるまで待つ。
# + lc_cell_meme={"current": "a9c31928-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31a7c-495a-11e8-8185-0242ac130002", "previous": "a9c317ca-495a-11e8-8185-0242ac130002"}
import time
status = compute.instances().get(project=target_project_id, zone=target_zone,
instance=machine_name).execute()['status']
while status == 'PROVISIONING' or status == 'STAGING':
time.sleep(30)
status = compute.instances().get(project=target_project_id, zone=target_zone,
instance=machine_name).execute()['status']
assert(status == 'RUNNING')
# + [markdown] lc_cell_meme={"current": "a9c31a7c-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31bda-495a-11e8-8185-0242ac130002", "previous": "a9c31928-495a-11e8-8185-0242ac130002"}
# ## IPアドレスの取得
# + lc_cell_meme={"current": "a9c31bda-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31d38-495a-11e8-8185-0242ac130002", "previous": "a9c31a7c-495a-11e8-8185-0242ac130002"}
vm_desc = compute.instances().get(project=target_project_id, zone=target_zone, instance=machine_name).execute()
ip_address = vm_desc['networkInterfaces'][0]['accessConfigs'][0]['natIP']
ip_address
# + [markdown] lc_cell_meme={"current": "a9c31d38-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31e96-495a-11e8-8185-0242ac130002", "previous": "a9c31bda-495a-11e8-8185-0242ac130002"}
# pingが通ることを確認。
# + lc_cell_meme={"current": "a9c31e96-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c31ff4-495a-11e8-8185-0242ac130002", "previous": "a9c31d38-495a-11e8-8185-0242ac130002"}
# !ping -c 4 {ip_address}
# + [markdown] lc_cell_meme={"current": "a9c31ff4-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32148-495a-11e8-8185-0242ac130002", "previous": "a9c31e96-495a-11e8-8185-0242ac130002"}
# ## Ansibleから操作可能であることを確認
#
# [キーペアの設定](#キーペアの設定-3.6)でansibleユーザとしてこの環境の公開鍵をInjectionしている。
# + lc_cell_meme={"current": "a9c32148-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3229c-495a-11e8-8185-0242ac130002", "previous": "a9c31ff4-495a-11e8-8185-0242ac130002"}
import tempfile
work_dir = tempfile.mkdtemp()
work_dir
# + lc_cell_meme={"current": "a9c3229c-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c323fa-495a-11e8-8185-0242ac130002", "previous": "a9c32148-495a-11e8-8185-0242ac130002"}
import os
snapshot_hosts = os.path.join(work_dir, 'hosts')
with open(snapshot_hosts, 'w') as f:
f.write('{address}\n'.format(address=ip_address))
# !cat { snapshot_hosts }
# + [markdown] lc_cell_meme={"current": "a9c323fa-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3254e-495a-11e8-8185-0242ac130002", "previous": "a9c3229c-495a-11e8-8185-0242ac130002"}
# Ansible経由でansibleユーザとしてSSHできることを確認する。
#
# > インスタンス起動直後の場合、UNREACHABLEとなるおそれがある。その場合は再度実行すること。
# + lc_cell_meme={"current": "a9c3254e-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c326a2-495a-11e8-8185-0242ac130002", "previous": "a9c323fa-495a-11e8-8185-0242ac130002"}
# !ansible -a 'whoami' -i { snapshot_hosts } all
# + [markdown] lc_cell_meme={"current": "a9c326a2-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32800-495a-11e8-8185-0242ac130002", "previous": "a9c3254e-495a-11e8-8185-0242ac130002"}
# # Inventoryの更新
#
# Inventoryに、作成したマシンのIPアドレスを追加する。変更する前に、現在の内容をコピーしておく。
# + lc_cell_meme={"current": "a9c32800-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32954-495a-11e8-8185-0242ac130002", "previous": "a9c326a2-495a-11e8-8185-0242ac130002"}
# !cp inventory {work_dir}/inventory-old
# + [markdown] lc_cell_meme={"current": "a9c32954-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32ab2-495a-11e8-8185-0242ac130002", "previous": "a9c32800-495a-11e8-8185-0242ac130002"}
# [Inventory](../edit/inventory) を修正する。
# + lc_cell_meme={"current": "a9c32ab2-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32c06-495a-11e8-8185-0242ac130002", "previous": "a9c32954-495a-11e8-8185-0242ac130002"}
# !diff -ur {work_dir}/inventory-old inventory
# + [markdown] lc_cell_meme={"current": "a9c32c06-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32d64-495a-11e8-8185-0242ac130002", "previous": "a9c32ab2-495a-11e8-8185-0242ac130002"}
# 追加したグループ名で、ansibleのpingモジュールが実行可能かどうかを確認する。
# + lc_cell_meme={"current": "a9c32d64-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c32eb8-495a-11e8-8185-0242ac130002", "previous": "a9c32c06-495a-11e8-8185-0242ac130002"}
target_group = 'test-gce'
# + lc_cell_meme={"current": "a9c32eb8-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c33016-495a-11e8-8185-0242ac130002", "previous": "a9c32d64-495a-11e8-8185-0242ac130002"}
# !ansible -m ping {target_group}
# + [markdown] lc_cell_meme={"current": "a9c33016-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c33174-495a-11e8-8185-0242ac130002", "previous": "a9c32eb8-495a-11e8-8185-0242ac130002"}
# 特にエラーとならなければOK。これで完了とする。
# + [markdown] lc_cell_meme={"current": "a9c33174-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c332c8-495a-11e8-8185-0242ac130002", "previous": "a9c33016-495a-11e8-8185-0242ac130002"}
# # 後始末
#
# 一時ディレクトリを削除する。
# + lc_cell_meme={"current": "a9c332c8-495a-11e8-8185-0242ac130002", "history": [], "next": "a9c3341c-495a-11e8-8185-0242ac130002", "previous": "a9c33174-495a-11e8-8185-0242ac130002"}
# !rm -fr {work_dir}
# + lc_cell_meme={"current": "a9c3341c-495a-11e8-8185-0242ac130002", "history": [], "next": null, "previous": "a9c332c8-495a-11e8-8185-0242ac130002"}
| D01_GCE - Set! Go! (Google Compute Engine).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeightWatcher GPT
#
#
#
from IPython.display import Image
# ## Summary of Results
#
#
# Suppress the powerlaw package warnings
# "powerlaw.py:700: RuntimeWarning: divide by zero encountered in true_divide"
# "powerlaw.py:700: RuntimeWarning: invalid value encountered in true_divide"
import warnings
warnings.simplefilter(action='ignore', category=RuntimeWarning)
# +
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext watermark
# %watermark
# -
# ### Import WeightWatcher
#
# set custom Logging at WARN Level
# +
import logging
logging.basicConfig(level=logging.INFO)
logger = logging.getLogger(__name__)
logger.setLevel(logging.WARN)
import torch
import weightwatcher as ww
import pytorch_transformers as ppb
print("Torch ",torch.__version__)
print("WeightWatcher ",ww.__version__)
print("PyTorch Transformers",ppb.__version__)
# -
# ### Create all models now
device = torch.device("cpu")
model = ppb.BertModel.from_pretrained(init_checkpoint_pt)
model.to(device)
series_name = 'GPT'
all_names = [ 'densenet121', 'densenet169', 'densenet201', 'densenet161']
colors = ['blue', 'red']
all_models = []
all_models.append(models.densenet121(pretrained=True))
all_models.append(models.densenet161(pretrained=True))
all_models.append(models.densenet169(pretrained=True))
all_models.append(models.densenet201(pretrained=True))
# ### Get reported accuracies from pytorch website
#
# https://pytorch.org/docs/stable/torchvision/models.html
#
#
# <pre>
# <table class="docutils align-default">
# <colgroup>
# <col style="width: 55%" />
# <col style="width: 22%" />
# <col style="width: 22%" />
# </colgroup>
# <thead>
# <thead>
# <tr class="row-odd"><th class="head"><p>Network</p></th>
# <th class="head"><p>Top-1 error</p></th>
# <th class="head"><p>Top-5 error</p></th>
# </tr>
# </thead>
# <tbody>
#
# <tr class="row-even"><td><p>Densenet-121</p></td>
# <td><p>25.35</p></td>
# <td><p>7.83</p></td>
# </tr>
# <tr class="row-odd"><td><p>Densenet-169</p></td>
# <td><p>24.00</p></td>
# <td><p>7.00</p></td>
# </tr>
# <tr class="row-even"><td><p>Densenet-201</p></td>
# <td><p>22.80</p></td>
# <td><p>6.43</p></td>
# </tr>
# <tr class="row-odd"><td><p>Densenet-161</p></td>
# <td><p>22.35</p></td>
# <td><p>6.20</p></td>
# </tr>
#
# </tbody>
# </table>
# </pre>
# +
top1_errors= {
"densenet121": 25.35,
"densenet169": 24.00,
"densenet201": 22.80,
"densenet161": 22.35
}
# +
top5_errors= {
"densenet121": 7.83,
"densenet169": 7.00,
"densenet201": 6.43,
"densenet161": 6.20
}
# -
# ## Run WeightWatcher, collect summary and details (as dataframes) for all models
all_details = []
all_summaries = []
for im, name in enumerate(all_names):
watcher = ww.WeightWatcher(model=all_models[im], logger=logger)
results = watcher.analyze(alphas=True, softranks=True, spectralnorms=True, mp_fit=True)
summary = watcher.get_summary()
all_summaries.append(summary)
details = watcher.get_details(results=results)
details.drop(columns=['slice', 'slice_count'], inplace=True)
details.dropna(inplace=True)
details['NxM'] = pd.to_numeric(details.N * details.M)
all_details.append(details)
plt.rcParams.update({'font.size': 16})
from pylab import rcParams
rcParams['figure.figsize'] = 10,10
# +
from sklearn.linear_model import LinearRegression
from sklearn import metrics
def plot_test_accuracy(metric, xlabel, title):
"""Create plot of Metric vs Reported Test Accuracy, and run Linear Regression"""
num = len(all_names)
xs, ys = np.empty(num), np.empty(num)
for im, modelname in enumerate(all_names):
summary = all_summaries[im]
x = summary[metric]
xs[im] = x
error = top1_errors[modelname]
y = 100.0-error
ys[im] = y
label = modelname
plt.scatter(x, y, label=label)
xs = xs.reshape(-1,1)
ys = ys.reshape(-1,1)
regr = LinearRegression()
regr.fit(xs, ys)
y_pred = regr.predict(xs)
plt.plot(xs, y_pred, color='red', linewidth=1)
rmse = np.sqrt(metrics.mean_squared_error(ys, y_pred))
r2 = metrics.r2_score(ys, y_pred)
title2 = "RMSE: {0:.2} R2: {0:.2}".format(rmse, r2)
plt.legend()
plt.title(r"Test Accuracy vs "+title+"\n"+title2)
plt.ylabel(r"Test Accuracy")
plt.xlabel(xlabel);
plt.show()
# -
def plot_metrics_histogram(metric, xlabel, title, log=False, valid_ids = []):
transparency = 1.0
if len(valid_ids) == 0:
valid_ids = range(len(all_details)-1)
for im, details in enumerate(all_details):
if im in valid_ids:
vals = details[metric].to_numpy()
if log:
vals = np.log10(np.array(vals+0.000001, dtype=np.float))
plt.hist(vals, bins=100, label=all_names[im], alpha=transparency, color=colors[im], density=True)
transparency -= 0.15
plt.legend()
plt.title(title)
plt.xlabel(xlabel)
plt.show()
def plot_metrics_depth(metric, ylabel, title, log=False, valid_ids = []):
transparency = 1.0
if len(valid_ids) == 0:
valid_ids = range(len(all_details)-1)
for im, details in enumerate(all_details):
if im in valid_ids:
details = all_details[im]
name = all_names[im]
x = details.index.to_numpy()
y = details[metric].to_numpy()
if log:
y = np.log10(np.array(y+0.000001, dtype=np.float))
plt.scatter(x,y, label=name, color=colors[im])
plt.legend()
plt.title(title)
plt.xlabel("Layer id")
plt.ylabel(xlabel)
plt.show()
# ## Metrics vs Test Accuracy
# +
metric = "lognorm"
xlabel = r"$\langle\log\Vert W\Vert\rangle_{F}$"
title = "Average Log Frobenius Norm "+xlabel
plot_test_accuracy(metric, xlabel, title)
metric = "alpha"
xlabel = r"$\alpha$"
title = "Average Alpha "+xlabel
plot_test_accuracy(metric, xlabel, title)
metric = "alpha_weighted"
xlabel = r"$\hat{\alpha}$"
title = "Average Weighted Alpha "+xlabel
plot_test_accuracy(metric, xlabel, title)
metric = "spectralnorm"
xlabel = r"$\langle\Vert\mathbf{W}\Vert_{2}\rangle$"
title = "Average Log Spectral Norm "+xlabel
plot_test_accuracy(metric, xlabel, title)
metric = "softranklog"
xlabel = r"$\langle\mathcal{R}_{s}\rangle$"
title = "Average Log Stable Rank "+xlabel
plot_test_accuracy(metric, xlabel, title)
metric = "logpnorm"
xlabel = r"$\langle\Vert\mathbf{W}\Vert^{\alpha}\rangle$"
title = "Average Log pNorm "+xlabel
plot_test_accuracy(metric, xlabel, title)
# -
# ## Histogram of metrics for all layers
first_n_last_ids = [0, len(all_details)-1]
# +
metric = "lognorm"
xlabel = r"Log Frobenius Norm $\langle\log\Vert W\Vert\rangle_{F}$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title)
plot_metrics_histogram(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "alpha"
xlabel = r"Alpha $\alpha$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title)
plot_metrics_histogram(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "alpha_weighted"
xlabel = r"Weighted Alpha $\hat{\alpha}$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title)
plot_metrics_histogram(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "softranklog"
xlabel = r"Log Stable Rank $\mathcal{R}_{s}$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title)
plot_metrics_histogram(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "spectralnorm"
xlabel = r"Log Spectral Norm $\Vert\mathbf{W}\Vert_{2}$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title, log=True)
plot_metrics_histogram(metric, xlabel, title, log=True, valid_ids = first_n_last_ids)
metric = "softrank_mp"
xlabel = r"Log MP Soft Rank $\mathcal{R}_{mp}$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title)
plot_metrics_histogram(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "logpnorm"
xlabel = r"Log p-Norm $\Vert\mathbf{W}\Vert^{\alpha}$"
title = series_name+" "+xlabel
plot_metrics_histogram(metric, xlabel, title)
plot_metrics_histogram(metric, xlabel, title, valid_ids = first_n_last_ids)
# -
# ## Metrics as a function of depth
# +
metric = "lognorm"
xlabel = r"Log Frobenius Norm $\langle\log\Vert W\Vert\rangle_{F}$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title)
plot_metrics_depth(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "alpha"
xlabel = r"Alpha $\alpha$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title)
plot_metrics_depth(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "alpha_weighted"
xlabel = r"Weighted Alpha $\hat{\alpha}$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title)
plot_metrics_depth(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "softranklog"
xlabel = r"Log Stable Rank $\mathcal{R}_{s}$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title)
plot_metrics_depth(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "spectralnorm"
xlabel = r"Log Spectral Norm $\Vert\mathbf{W}\Vert_{2}$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title, log=True)
plot_metrics_depth(metric, xlabel, title, log=True, valid_ids = first_n_last_ids)
metric = "softrank_mp"
xlabel = r"Log MP Soft Rank $\mathcal{R}_{mp}$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title)
plot_metrics_depth(metric, xlabel, title, valid_ids = first_n_last_ids)
metric = "logpnorm"
xlabel = r"Log p-Norm $\Vert\mathbf{W}\Vert^{\alpha}$"
title = series_name+" "+xlabel
plot_metrics_depth(metric, xlabel, title)
plot_metrics_depth(metric, xlabel, title, valid_ids = first_n_last_ids)
# -
| WeightWatcher-GPT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from bs4 import BeautifulSoup as bs
import requests
url="https://www.radiojavan.com/mp3s/browse/popular/month"
pattern="https://host2.rjmusicmedia.com/media/mp3/mp3-256/Erfan-Sepehr-Khalse-Shabaye-Tehran-(Ft-Farshid).mp3"
myList=[]
html=requests.get(url)
soup=bs(html.content , "lxml")
mydivs = soup.findAll("div", {"class": "block_container"})
for divs in mydivs:
myList.append(divs.find('a')['href'])
# -
print(myList)
myName=[]
for item in myList:
myName.append(item[10:])
print(myName)
a="https://host2.rjmusicmedia.com/media/mp3/mp3-256/"
b=".mp3"
myUrl=[]
for item in myName:
myUrl.append(a+item+b)
print(myUrl)
file = open("Exported.txt", "w")
for item in myUrl:
file.write(item + '\n')
file.close()
| AGRadioJavan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mean Flow Frequency Curve
#
# __Description__: Calculates the mean flow frequency curve using [Bulletin 17C](https://pubs.usgs.gov/tm/04/b05/tm4b5.pdf) confidence limits calculated in [HEC-SSP](https://www.hec.usace.army.mil/software/hec-ssp/).
#
# __Input__: [HEC-SSP](https://www.hec.usace.army.mil/software/hec-ssp/) .rpt files containing flow frequency data for a specific USGS Stream Gage calculated at a range of confidence limits.
#
# __Output__: The mean flow frequency curve table (annual exceedance probability verses discharge).
#
# *Acknowledgement*: This notebook was adapated from the Excel Workbook titled, "Calculating Mean (Expected) Flow Frequency Cruve Using Bulletin 17C Conf...," which was provided by <EMAIL> from the USACE.
#
# ---
# ## Load Libraries, Parameters, and Data:
# ### Libraries:
import sys
sys.path.append('../../core')
from meanffc import*
# ### Parameters:
# #### Site specific (papermill):
# + active=""
# ## Filenames and paths:
# gage_ID = '01134500' # The USGS Station ID whose mean flow frequency curve will be calculated using this notebook.
# inputs_dir = pl.Path(os.getcwd())/'Inputs' # The directory/path where the SSP files can be found within the root directory
# outputs_dir = pl.Path(os.getcwd())/'Outputs' # The directory/path to save the outputs
#
# ## Options
# version = '2_2' # The version of HEC-SSP used to calculate the .rpt files. Either 2.1 or 2.2, specified as 2_1 or 2_2 here, respectively.
# max_cl = 0.99 # Specify the upper confidence limit to restrict the confidence limit range, i.e. set the upper limit to 0.99 to restrict the range to 0.01 to 0.99.
# adj_flows = 1.0 # [cfs]; amount to adjust the discharge for each annual exceedance probability if the discharge does not increase with decreasing probability.
# round_decimals = 1 # Number of decimal places to include in the final mean curve table.
# verbose = True # Option for displaying print statements.
# Extrapolate = True # True indicates that you want to extrapolate while False indicates that you want to only interpolate within the bounds of the minimum and maximum discharge for a specific CL.
# exclude_tails = True # If True than the integration to calculate the mean curve only includes the area between the lower and upper confidence limits.
# -
# ##### Make directory if it does not exist:
make_directories([outputs_dir], verbose)
# #### Project specific:
standard_AEP = [0.9, 0.5, 0.2, 0.1, 0.05, 0.02, 0.01, 0.005, 0.002, 0.001, 5E-04, 2E-04, 1E-04, 5E-05, 2E-05, 1E-05, 5E-06, 2E-06, 1E-06] # Selected standard reporting AEP values
# ### Data:
# #### Identify the HEC-SSP results files:
# +
inputs_dir = pl.Path(inputs_dir)
ssp_results = list_ssp_files(inputs_dir, max_cl, verbose)
# -
# ##### Construct a summary table:
df = make_ssp_table(ssp_results, version)
df.head(2)
# ##### Check that discharge increases with decreasing AEP:
df = monotonic_test(df, adj_flows)
# ##### Extract the confidence limits and annual exceedance probabilities:
# +
CL = list(map(float, df.columns))
AEP = df.index
# -
# ---
# ## Calculate the Mean Flow Frequency Curve:
# ### Transform the HEC-SSP Results:
# +
data = np.log10(df)
CLz = zvar(CL)
AEPz = zvar(AEP)
# -
# #### Plot the transformed data to visually check for errors:
plot_ssp_transform(data, AEPz, CLz)
# ### Determine the AEP for Each Flow and CL:
# #### Bin the discharge and calculate the annual exceedance probability:
# +
Q = binq(data)
res = interp_aep(data, Q, CLz, AEPz, Extrapolate)
# -
# ##### Plot the results:
plot_ssp_interp(res)
# #### Take the inverse of the standard normal z variate of the AEP:
restrans = zvar_inv(res, CL)
# ##### Plot the results:
plot_ssp_interptrans(restrans)
# #### Calculate the Mean (Expected Value) of the AEP:
AEPm = mean_aep(restrans, exclude_tails)
# ##### Transform the mean AEP:
AEPmz = zvar(AEPm)
# ##### Plot the results:
plot_ssp_meanmed(AEPz, data, AEPmz, Q)
# ---
# ## Construct a Summary Table and Save:
# ### Transform the Standard AEPs and Initialize a Summary Table:
# +
standard_AEPz = zvar(standard_AEP)
table = ffc_summary(standard_AEP, standard_AEPz)
# -
# ### Calculate the Mean and Median Flow:
# +
table['Q_Mean_cfs'] = np.round(10**table['AEPz'].apply(interp_q(AEPmz, Q)), round_decimals)
table['Q_Median_cfs'] = np.round(10**table['AEPz'].apply(interp_q(AEPz, np.array(data['0.5']))), round_decimals)
# -
# #### Save the Results:
# +
mean_curve_table = table.copy().drop(columns=['AEPz'])
outputs_dir = pl.Path(outputs_dir)
mean_curve_table.to_csv(outputs_dir/'MeanCurve_{}.csv'.format(gage_ID))
sb.glue('mean_curve', mean_curve_table.to_dict())
mean_curve_table.head(2)
# -
# #### Plot the Results (Static):
plot_ssp_meanmedffc(table, gage_ID)
# #### Plot the Results (Interactive):
plotly_ssp_meanffc(table, gage_ID)
# ---
# ## End
| notebooks/fluvial/SSP_to_Mean_Curve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''.venv'': venv)'
# name: python3
# ---
import re
from nltk.tokenize.casual import TweetTokenizer, casual_tokenize
from nltk.tokenize import RegexpTokenizer
from gatenlp import Document
from gatenlp.processing.tokenizer import NLTKTokenizer
from gatenlp.processing.tokenizer import SplitPatternTokenizer
# +
txt = " This is a 💩 document. It has two sentences and 14 tokens. "
doc = Document(txt)
doc
# -
tok = NLTKTokenizer(nltk_tokenizer=RegexpTokenizer(r'\w+|\$[\d\.]+|\S+'), space_token_type="SpaceToken")
doc = tok(Document(txt))
doc
tok = NLTKTokenizer(nltk_tokenizer=casual_tokenize, space_token_type="SpaceToken")
doc = tok(Document(txt))
doc
tok = SplitPatternTokenizer(split_pattern=re.compile(r'\s+'), space_token_type="SpaceToken")
doc = tok(Document(txt))
doc
tok = SplitPatternTokenizer(split_pattern=re.compile(r'\s+'),
token_pattern=re.compile(r'[a-zA-Z]'),
space_token_type="SpaceToken")
doc = tok(Document(txt))
doc
tok = SplitPatternTokenizer(split_pattern=" ",
token_pattern="o",
space_token_type="SpaceToken")
doc = tok(Document(txt))
doc
# +
import os
from gatenlp import logger, Document
import spacy
from gatenlp.lib_spacy import spacy2gatenlp, apply_spacy
nlp = spacy.load("en_core_web_sm")
txt = "<NAME> was born in Hawaii. He was elected president in 2008. "
doc = Document(txt)
annset = doc.annset()
ann = annset.add(0,32 ,"Sentence",{})
ann = annset.add(33,67,"Sentence",{})
anns = doc.annset()
sents = anns.with_type("Sentence")
assert len(sents) == 2
tokens = anns.with_type("Token")
assert len(tokens) == 0
gdoc=apply_spacy(nlp,doc , setname="spacy", containing_anns=anns)
annsOut = gdoc.annset("spacy")
sents = annsOut.with_type("Sentence")
assert len(sents) == 2
tokens = annsOut.with_type("Token")
assert len(tokens) == 14
doc
| debug/debug-spacy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Shahid1993/colab-notebooks/blob/master/predictive_keyboard_using_RNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-XyxJMP9j4Bb" colab_type="text"
# # [Making a Predictive Keyboard using Recurrent Neural Networks — TensorFlow for Hackers (Part V)](https://medium.com/@curiousily/making-a-predictive-keyboard-using-recurrent-neural-networks-tensorflow-for-hackers-part-v-3f238d824218)
# + [markdown] id="43qiGz1ikK4t" colab_type="text"
# This time we will build a model that predicts the next word (a character actually) based on a few of the previous. We will extend it a bit by asking it for 5 suggestions instead of only 1.
#
# # Recurrent Neural Networks
#
# Our weapon of choice for this task will be Recurrent Neural Networks (RNNs). But why? What’s wrong with the type of networks we’ve used so far? Nothing! Yet, they lack something that proves to be quite useful in practice — `memory`!
#
# In short, `RNN` models provide a way to not only examine the current input but the one that was provided one step back, as well. If we turn that around, we can say that the decision reached at time step t-1 directly affects the future at step t.
#
# 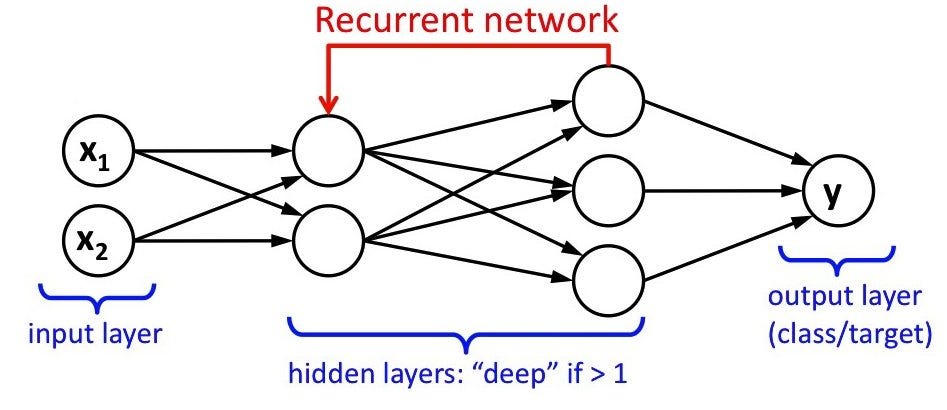
#
# It seems like a waste to throw out the memory of what you’ve seen so far and start from scratch every time. That’s what other types of Neural Networks do.
#
# # Definition
#
# `RNNs` define a recurrence relation over time steps which is given by:
#
# 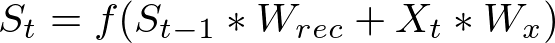
#
# Where St is the state at time step t, Xt an exogenous input at time t, Wrec and Wx are weights parameters. The feedback loops gives memory to the model because it can remember information between time steps.
#
# `RNNs` can compute the current state St from the current input Xt and previous state St−1 or predict the next state from St+1 from the current St and current input Xt. Concretely, we will pass a sequence of 40 characters and ask the model to predict the next one. We will append the new character and drop the first one and predict again. This will continue until we complete a whole word.
#
# # LSTMs
#
# Two major problems torment the `RNNs` — vanishing and exploding gradients. In traditional `RNNs` the gradient signal can be multiplied a large number of times by the weight matrix. Thus, the magnitude of the weights of the transition matrix can play an important role.
#
# If the weights in the matrix are small, the gradient signal becomes smaller at every training step, thus making learning very slow or completely stops it. This is called vanishing gradient. Let’s have a look at applying the sigmoid function multiple times, thus simulating the effect of vanishing gradient:
#
# 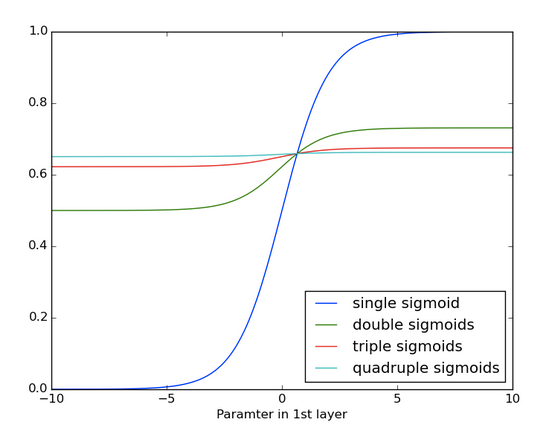
#
# Conversely, the exploding gradient refers to the weights in this matrix being so large that it can cause learning to diverge.
#
# `LSTM` model is a special kind of `RNN` that learns long-term dependencies. It introduces new structure — the memory cell that is composed of four elements: input, forget and output gates and a neuron that connects to itself:
#
# 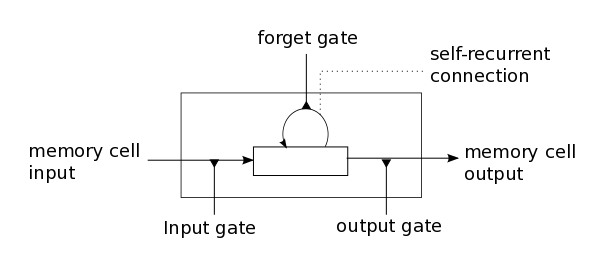
#
# `LSTMs` fight the gradient vanishing problem by preserving the error that can be backpropagated through time and layers. By maintaining a more constant error, they allow for learning long-term dependencies. On another hand, exploding is controlled with gradient clipping, that is the gradient is not allowed to go above some predefined value.
#
# + [markdown] id="86dHl5yOlPvC" colab_type="text"
# # Setup
#
# Let’s properly seed our random number generator and import all required modules:
# + id="FyG245CbEyOj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2f2187df-c688-45e5-fe69-4e65b91ca69d"
import numpy as np
np.random.seed(42)
import tensorflow as tf
tf.set_random_seed(42)
from keras.models import Sequential, load_model
from keras.layers import Dense, Activation
from keras.layers import LSTM, Dropout
from keras.layers import TimeDistributed
from keras.layers.core import Dense, Activation, Dropout, RepeatVector
from keras.optimizers import RMSprop
import matplotlib.pyplot as plt
import pickle
import sys
import heapq
import seaborn as sns
from pylab import rcParams
# %matplotlib inline
sns.set(style='whitegrid', palette='muted', font_scale=1.5)
rcParams['figure.figsize'] = 12, 5
# + [markdown] id="q7WepqMeP8Im" colab_type="text"
# # Loading the data
#
# We will use <NAME>’s Beyond Good and Evil as a training corpus for our model. The text is not that large and our model can be trained relatively fast using a modest GPU.
# + id="i6uTIz7yIdqP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3cd5e35e-2bd6-45f0-9da8-a82ac3bd6c23"
path = './sample_data/nietzsche.txt'
text = open(path).read().lower()
print('corpus length:', len(text))
# + [markdown] id="K2mMQUz6QIMW" colab_type="text"
# # Preprocessing
#
# Let’s find all unique chars in the corpus and create char to index and index to char maps:
# + id="o7MXMyBPPbcu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f1887907-0abc-42ac-b78a-6931c1e312c2"
chars = sorted(list(set(text)))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
print(f'unique chars: {len(chars)}')
# + [markdown] id="5FwDdojDQglR" colab_type="text"
# Next, let’s cut the corpus into chunks of 40 characters, spacing the sequences by 3 characters. Additionally, we will store the next character (the one we need to predict) for every sequence:
# + id="17sfI9SJQL_n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="cc834ee8-d955-4692-b6c2-92a2a6ffe178"
SEQUENCE_LENGTH = 40
step = 3
sentences = []
next_chars = []
for i in range(0, len(text) - SEQUENCE_LENGTH, step):
sentences.append(text[i: i + SEQUENCE_LENGTH])
next_chars.append(text[i + SEQUENCE_LENGTH])
print(f'num training examples: {len(sentences)}')
# + [markdown] id="ZEVtGZfEQwmq" colab_type="text"
# It is time for generating our features and labels. We will use the previously generated sequences and characters that need to be predicted to create one-hot encoded vectors using the `char_indices` map:
# + id="bpGyjuN_Q2sr" colab_type="code" colab={}
X = np.zeros((len(sentences), SEQUENCE_LENGTH, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
X[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
# + [markdown] id="0wFNfNWiQ-kg" colab_type="text"
# Let’s have a look at a single training sequence:
# + id="u5v7uuslQ59c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6eaabbe5-2ada-43d6-eed9-70d4e9b74d40"
sentences[100]
# + [markdown] id="x2SJ9Ko-RFL0" colab_type="text"
# The character that needs to be predicted for it is:
# + id="ho9TltNFRBCe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bee61a74-0536-4941-bcdd-03f5616d5e19"
next_chars[100]
# + [markdown] id="aydxJMHjRKog" colab_type="text"
# The encoded (one-hot) data looks like this:
# + id="9GWKVSnYRHI4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="999f66ee-73bd-4ec3-d483-400c02f4338c"
X[0][0]
# + id="GJX3M2C0RNXk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="b3b0a701-3c39-4ec8-9ce2-b2159c590619"
y[0]
# + [markdown] id="WR5slh2cRSdU" colab_type="text"
# And for the dimensions:
# + id="3Bq2v41uRPt9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="57a1d723-5526-4b77-9cf2-a39eb59347a2"
X.shape
# + id="evOzqrpKRUOE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f7f8bf3d-107d-4825-9070-a77acd6cdfb4"
y.shape
# + [markdown] id="ylvSVPOBRbxR" colab_type="text"
# We have 200288 training examples, each sequence has length of 40 with 59 unique chars.
# + [markdown] id="8nhrR1xnRuTU" colab_type="text"
# # Building the model
#
# The model we’re going to train is pretty straight forward. Single `LSTM` layer with `128` neurons which accepts input of shape (`40` — the length of a sequence, `57` — the number of unique characters in our dataset). A fully connected layer (for our output) is added after that. It has `57` neurons and softmax for activation function:
# + id="lSdgR19fRXt0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="cccd17f0-121a-4d21-d5ac-bcfaa1ad4a59"
model = Sequential()
model.add(LSTM(128, input_shape=(SEQUENCE_LENGTH, len(chars))))
model.add(Dense(len(chars)))
model.add(Activation('softmax'))
# + [markdown] id="BXtC_FywSSMw" colab_type="text"
# # Training
#
# Our model is trained for `20` epochs using `RMSProp` optimizer and uses `5%`` of the data for validation:
# + id="APLCGJaCSFlr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 937} outputId="dfa84c72-7b5a-426e-ca2c-ec82b5abef3b"
optimizer = RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
history = model.fit(X, y, validation_split=0.05, batch_size=128, epochs=20, shuffle=True).history
# + [markdown] id="TiFOP3MaapiA" colab_type="text"
# # Saving
#
# It took a lot of time to train our model. Let’s save our progress:
# + id="fU1E-_JmSZdm" colab_type="code" colab={}
model.save('keras_model.h5')
pickle.dump(history, open("history.p", "wb"))
# + [markdown] id="OE3KSLw9azAg" colab_type="text"
# And load it back, just to make sure it works:
# + id="S1r_uhXtatai" colab_type="code" colab={}
model = load_model('keras_model.h5')
history = pickle.load(open("history.p", "rb"))
# + [markdown] id="DSaKat0sa5f0" colab_type="text"
# # Evaluation
#
# Let’s have a look at how our accuracy and loss change over training epochs:
# + id="WucBUr4Ta1Mm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 372} outputId="6a518c95-66bb-4526-d34d-6beb3d80dd3f"
plt.plot(history['acc'])
plt.plot(history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left');
# + id="cmmS8HPqa8vB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 372} outputId="2f72ac26-3ced-457b-aa9c-a9fb31aea2f5"
plt.plot(history['loss'])
plt.plot(history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left');
# + [markdown] id="BjfU6rn5bIFX" colab_type="text"
# # Let’s put our model to the test
#
# Finally, it is time to predict some word completions using our model! First, we need some helper functions. Let’s start by preparing our input text:
# + id="kzx509pvbA-5" colab_type="code" colab={}
def prepare_input(text):
x = np.zeros((1, SEQUENCE_LENGTH, len(chars)))
for t, char in enumerate(text):
x[0, t, char_indices[char]] = 1.
return x
# + [markdown] id="Sg28ULFIbbj8" colab_type="text"
# Remember that our sequences must be `40` characters long. So we make a tensor with shape `(1, 40, 57)`, initialized with zeros. Then, a value of `1` is placed for each character in the passed text. We must not forget to use the lowercase version of the text:
# + id="D_AZTnbKbMS7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="0ad67771-7367-4014-8d01-a9486a7c7a17"
prepare_input("This is an example of input for our LSTM".lower())
# + [markdown] id="mLRi77rXgLY0" colab_type="text"
# Next up, the sample function:
# + id="ObRnqsZVgIiD" colab_type="code" colab={}
def sample(preds, top_n=3):
preds = np.asarray(preds).astype('float64')
preds = np.log(preds)
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
return heapq.nlargest(top_n, range(len(preds)), preds.take)
# + [markdown] id="7wJfgdm7gTZk" colab_type="text"
# This function allows us to ask our model what are the next n most probable characters. Isn’t that heap just cool?
#
#
# Now for the prediction functions themselves:
# + id="fb2Dfh2rgOwK" colab_type="code" colab={}
def predict_completion(text):
original_text = text
generated = text
completion = ''
while True:
x = prepare_input(text)
preds = model.predict(x, verbose=0)[0]
next_index = sample(preds, top_n=1)[0]
next_char = indices_char[next_index]
text = text[1:] + next_char
completion += next_char
if len(original_text + completion) + 2 > len(original_text) and next_char == ' ':
return completion
# + [markdown] id="H4mVUqJkgj6s" colab_type="text"
# This function predicts next character until space is predicted (you can extend that to punctuation symbols, right?). It does so by repeatedly preparing input, asking our model for predictions and sampling from them.
#
#
#
# <br>
#
# The final piece of the puzzle — `predict_completions` wraps everything and allow us to predict multiple completions:
# + id="jf3XSX_wgYdq" colab_type="code" colab={}
def predict_completions(text, n=3):
x = prepare_input(text)
preds = model.predict(x, verbose=0)[0]
next_indices = sample(preds, n)
return [indices_char[idx] + predict_completion(text[1:] + indices_char[idx]) for idx in next_indices]
# + [markdown] id="t0wTfqxHhSe8" colab_type="text"
# Let’s use sequences of `40` characters that we will use as seed for our completions. All of these are quotes from <NAME> himself:
# + id="p-7Qt-ycgx_F" colab_type="code" colab={}
quotes = [
"It is not a lack of love, but a lack of friendship that makes unhappy marriages.",
"That which does not kill us makes us stronger.",
"I'm not upset that you lied to me, I'm upset that from now on I can't believe you.",
"And those who were seen dancing were thought to be insane by those who could not hear the music.",
"It is hard enough to remember my opinions, without also remembering my reasons for them!"
]
# + id="6Hejr06lhXhq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="447a3cd0-8816-41a7-82f5-3285c1f60bdc"
for q in quotes:
seq = q[:40].lower()
print(seq)
print(predict_completions(seq, 5))
print()
# + [markdown] id="lwwedD_Hhrzk" colab_type="text"
# Apart from the fact that the completions look like proper words (remember, we are training our model on characters, not words), they look pretty reasonable as well! Perhaps better model and/or more training will provide even better results?
#
# <br>
#
# # Conclusion
#
# We’ve built a model using just a few lines of code in Keras that performs reasonably well after just 20 training epochs. Can you try it with your own text? Why not predict whole sentences? Will it work that well in other languages?
# + [markdown] id="mTb2Rh2IjMlv" colab_type="text"
# # References
#
# - [Recurrent Nets in TensorFlow](https://danijar.com/introduction-to-recurrent-networks-in-tensorflow/)
# - [The Unreasonable Effectiveness of Recurrent Neural Networks](https://karpathy.github.io/2015/05/21/rnn-effectiveness/)
# - [Understanding LSTM Networks](https://colah.github.io/posts/2015-08-Understanding-LSTMs/)
# - [How to implement RNN in Python](https://peterroelants.github.io/posts/rnn_implementation_part01/)
# - [LSTM Networks for Sentiment Analysis](http://deeplearning.net/tutorial/lstm.html)
# - [cs231n — Recurrent Neural Networks](http://cs231n.stanford.edu/slides/2016/winter1516_lecture10.pdf)
# + id="zEcCG4g2hcQF" colab_type="code" colab={}
| predictive_keyboard_using_RNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
import sys, os
import csv
import datetime
import numpy as np
import netCDF4
import ast
#import scipy.stats
#import json
sys.path.append("../../larda")
import pyLARDA
import pyLARDA.helpers as h
import pyLARDA.Transformations as pLTransf
import matplotlib
#matplotlib.use('Agg')
import matplotlib.pyplot as plt
from scipy import stats
import selection_criteria as s_c
# -
# %matplotlib inline
# +
def gen_dt_list(start, end, delta=24):
l = []
current = start
while current <= end:
l.append(current)
current = current + datetime.timedelta(hours=delta)
return l
def print_cloud_props(cloud):
dt_begin = ts_to_dt(float(cloud['Begin_Date_Unix']))
dt_end = ts_to_dt(float(cloud['End_Date_Unix']))
iwc_n = float(cloud['IWC_TOP_N'])
n_prof = float(cloud['N_Profiles'])
print(dt_begin.strftime("%Y%m%d-%H%M"), '-', dt_end.strftime("%Y%m%d-%H%M"), ':',
"{:>7.1f}".format(float(cloud['CBH'])), "{:>7.1f}".format(float(cloud['CTH'])),
"{:>7.2f}".format(float(cloud['CTT'])-273.15),
" {:.2e}".format(float(cloud['IWC_TOP_MED'])),
" {:.2f}".format(iwc_n/n_prof),
cloud['Cloud_Run'], cloud['Cloud_Type'])
def load_data(filename):
data = []
with open(filename, 'r') as f:
#with open('cloud_collection_LEIPZIG_all.csv', 'r') as f:
reader = csv.reader(f, delimiter=';')
header = next(reader)
for row in reader:
#data.append(row)
comp = row[1].replace('_', '')
if row[0][:14] == comp:
data.append({k:v for k, v in zip(header, row)})
else:
print('corrupted row ', row)
break
return data
# +
filename = {
"Pun_larda3": '../cloud_collections/cloud_collection_lacros_dacapo_all.csv',
"Lim_larda3": '../cloud_collections/cloud_collection_lacros_cycare_all_w_dl.csv',
"Lei_larda3": '../cloud_collections/cloud_collection_lacros_leipzig_all.csv',
}
clouds_new = load_data(filename['Pun_larda3'])
# +
clouds_new_filtered = []
for i, cloud in enumerate(clouds_new[:]):
ffcloud = lambda s: float(cloud[s])
dt_begin = h.ts_to_dt(ffcloud('Begin_Date_Unix'))
dt_end = h.ts_to_dt(ffcloud('End_Date_Unix'))
duration = dt_end-dt_begin
#conds = s_c.conditions_ice_wo_CTH(cloud)
conds = s_c.standard(cloud)
if all(conds[::2]):
clouds_new_filtered.append(cloud)
print(len(clouds_new_filtered))
# +
start = datetime.datetime(2018,11,28)
end = datetime.datetime(2020,12,31)
def filterfunc(dt, c):
ffcloud = lambda s: float(c[s])
dt_begin = h.ts_to_dt(ffcloud('Begin_Date_Unix'))
return dt < dt_begin < dt+datetime.timedelta(minutes=23*60+59)
sorted_day = {}
for dt in gen_dt_list(start, end):
new_today = [c for c in clouds_new_filtered if filterfunc(dt, c)]
if len(new_today) > 0:
print(dt.strftime("%Y%m%d"), f" clouds {len(new_today):2.0f} ")
#print(new_today)
sorted_day[dt.strftime("%Y%m%d")] = {'clouds': new_today}
# +
larda_rsd2 = pyLARDA.LARDA('remote', uri="http://larda3.tropos.de")
larda_rsd2.connect('lacros_dacapo')
# +
dt = datetime.datetime(2018,11,28)
# dt = datetime.datetime(2019,1,2)
# dt = datetime.datetime(2019,10,22)
# dt = datetime.datetime(2019,10,5)
#[dt, dt+datetime.timedelta(minutes=23*60+59)]
cases = {
"20181128": {'time_interval': [datetime.datetime(2018,11,28, 0, 20),
datetime.datetime(2018,11,28, 9, 0)],
'range_interval': [2100, 5200]}
}
case = cases["20181128"]
cloudnet_class = larda_rsd2.read("CLOUDNET", "CLASS", case['time_interval'], case['range_interval'])
cloudnet_Z = larda_rsd2.read("CLOUDNET", "Z", case['time_interval'], case['range_interval'])
cloudnet_beta = larda_rsd2.read("CLOUDNET", "beta", case['time_interval'], case['range_interval'])
cloudnet_VEL = larda_rsd2.read("CLOUDNET", "VEL", case['time_interval'], case['range_interval'])
shaun_VEL = larda_rsd2.read("SHAUN", "VEL", case['time_interval'], case['range_interval'])
T = larda_rsd2.read("CLOUDNET","T", case['time_interval'], case['range_interval'])
def toC(datalist):
return datalist[0]['var']-273.15, datalist[0]['mask']
T = pyLARDA.Transformations.combine(toC, [T], {'var_unit': "C"})
# +
def set_interval(ax):
ax.xaxis.set_major_locator(matplotlib.dates.HourLocator(interval=2))
ax.xaxis.set_minor_locator(matplotlib.dates.MinuteLocator(byminute=np.arange(0,60,20)))
return ax
def format_class(ax):
ax.tick_params(axis='both', which='major', labelsize=13.5)
ax.xaxis.label.set_size(13.5)
ax.yaxis.label.set_size(13.5)
ax.images[0].colorbar.ax.tick_params(axis='both', which='major', labelsize=11)
ax.images[0].colorbar.ax.yaxis.label.set_size(13.5)
return ax
def format_plt(ax):
ax.tick_params(axis='both', which='major', labelsize=13.5)
ax.xaxis.label.set_size(13.5)
ax.yaxis.label.set_size(13.5)
ax.images[0].colorbar.ax.tick_params(axis='both', which='major', labelsize=13.5)
ax.images[0].colorbar.ax.yaxis.label.set_size(13.5)
return ax
def add_custom_contour(ax, T):
dt = [h.ts_to_dt(t) for t in T['ts']]
rg = T['rg']/1000
ax.contour(
dt, rg, T['var'].T,
levels=[-20, -15,-10,-5],
linestyles=['-', ':', '--', '-.'],
colors='black'
)
return ax
fig_ar = 8/4.5
xsize = 7
pltfmt = dict(figsize=[xsize, xsize/fig_ar],
rg_converter=True)
rg_interval_km = [h/1000 for h in case['range_interval']]
fig, ax = pyLARDA.Transformations.plot_timeheight2(
cloudnet_class,
range_interval=rg_interval_km, **pltfmt)
#ax.yaxis.set_major_locator(matplotlib.ticker.MultipleLocator(1000))
from matplotlib import patches
for c in sorted_day[dt.strftime("%Y%m%d")]['clouds']:
dt_begin = h.ts_to_dt(float(c['Begin_Date_Unix']))
dt_end = h.ts_to_dt(float(c['End_Date_Unix']))
cbh = float(c['CBH'])/1000
cth = float(c['CTH'])/1000
duration = dt_end - dt_begin
height = cth-cbh
print(duration, height)
rect = patches.Rectangle(
(dt_begin,cbh),duration,height,linewidth=2,
edgecolor='none',facecolor='grey', alpha=0.15,
)
ax.add_patch(rect)
rect = patches.Rectangle(
(dt_begin,cbh),duration,height,linewidth=2,
edgecolor='darkred',facecolor='none'
)
ax.add_patch(rect)
ax = set_interval(ax)
ax = format_class(ax)
ax.images[0].colorbar.ax.set_ylabel('')
savename = "../plots/case/{}_cloudnet_class.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250)
contour = {'data': T, 'levels': np.arange(-37,11,4)}
cloudnet_Z['var_lims'] = [-40, 5]
cloudnet_Z['colormap'] = 'jet'
fig, ax = pyLARDA.Transformations.plot_timeheight2(
cloudnet_Z, range_interval=rg_interval_km,
#contour=contour,
z_converter="lin2z", **pltfmt)
ax = add_custom_contour(ax, T)
ax = set_interval(ax)
ax = format_plt(ax)
ax.images[0].colorbar.ax.set_ylabel('Cloudnet reflectivity [dBZ]')
savename = "../plots/case/{}_cloudnet_Z.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250)
fig, ax = pyLARDA.Transformations.plot_timeheight2(
cloudnet_VEL, range_interval=rg_interval_km,
**pltfmt)
ax = set_interval(ax)
ax = format_plt(ax)
ax.images[0].colorbar.ax.set_ylabel('Cloudnet velocity [m s$^{-1}$]')
savename = "../plots/case/{}_cloudnet_VEL.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250)
cloudnet_beta['colormap'] = 'jet'
fig, ax = pyLARDA.Transformations.plot_timeheight2(
cloudnet_beta, range_interval=rg_interval_km,
z_converter="log", **pltfmt)
ax = set_interval(ax)
ax = format_plt(ax)
ax.images[0].colorbar.ax.set_ylabel('Cloudnet beta [sr$^{-1}$ m$^{-1}$]')
savename = "../plots/case/{}_cloudnet_beta.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250)
fig, ax = pyLARDA.Transformations.plot_timeheight2(
shaun_VEL, range_interval=rg_interval_km,
**pltfmt)
ax = set_interval(ax)
ax = format_plt(ax)
ax.images[0].colorbar.ax.set_ylabel('Doppler lidar velocity [m s$^{-1}$]')
savename = "../plots/case/{}_shaun_VEL.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250)
# +
ctt = np.array([float(d['CTT']) for d in sorted_day[dt.strftime("%Y%m%d")]['clouds']])
ctt[ctt == 0.0] = np.nan
frac_prof_ice = np.array([float(d['IWC_TOP_N'])/float(d['N_Profiles']) for d in sorted_day[dt.strftime("%Y%m%d")]['clouds']])
ilcr = np.array([float(d['ILCR_MED']) for d in sorted_day[dt.strftime("%Y%m%d")]['clouds']])
ilcr[ilcr == 0.0] = 1e-50
fig, ax = plt.subplots(figsize=(6, 5))
# ax.set_facecolor('lightgrey')
# sc = ax.scatter(ctt-273.15, frac_prof_ice, s=12,
# #label=trace_geo_names[int(elem)],
# norm=matplotlib.colors.LogNorm(vmin=5e-5,vmax=2e-1),
# #vmin=-0.5, vmax=0.5,
# c=ilcr, cmap='plasma_r'
# )
sc = ax.scatter(ctt-273.15, frac_prof_ice,
marker='s', facecolors='none', edgecolors='r',
s=50, linewidth=2,
)
sc.cmap.set_under('grey')
cbar = fig.colorbar(sc, extend='min')
cbar.ax.set_ylabel('ILCR []', fontsize=14)
cbar.ax.tick_params(axis='y', which='major', direction='in',
right=True,
width=1.5, length=5, labelsize=12)
ax.set_xlabel('CTT [°C]', fontsize=14)
ax.set_ylabel('Fraction prof. containing ice [%]', fontsize=14)
ax.set_ylim([-0.05, 1.05])
ax.set_xlim([-43, 3])
print('no cases ', len(sorted_day[dt.strftime("%Y%m%d")]['clouds']), ctt[~np.isnan(ctt)].shape[0])
print(np.sum(frac_prof_ice > 1e-50))
ax.set_title('layered_all')
ax.text(0.97, 0.94, 'n={}'.format(ctt[~np.isnan(ctt)].shape[0]), horizontalalignment='right',
#verticalalignment='bottom',
transform=ax.transAxes, color='k', fontsize=13)
ax.xaxis.set_minor_locator(matplotlib.ticker.AutoMinorLocator())
ax.yaxis.set_minor_locator(matplotlib.ticker.AutoMinorLocator())
ax.tick_params(axis='both', which='major', direction='in',
top=True, right=True,
width=1.5, length=5, labelsize=12)
ax.tick_params(axis='both', which='minor', direction='in',
top=True, right=True,
width=1.5, length=2.5)
#ax.legend(fontsize=11)
plt.tight_layout()
# fname = '../plots/{}_frac_profiles_ilcr_{}.png'.format(camp, 'layered_all')
# plt.savefig(fname, dpi=250)
savename = "../plots/case/{}_ctt_frac_location.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250, transparent=True)
# +
hvel_corr = True
toarray = lambda s: np.array(ast.literal_eval(s))
for cloud in sorted_day[dt.strftime("%Y%m%d")]['clouds']:
fig, ax = plt.subplots(figsize=[2.5, 2.1])
ctt = float(cloud['CTT'])-273.15
if hvel_corr:
vel = max(float(cloud['VEL']),0.1)
else:
vel = 1
autocor_time = toarray(cloud['v_dl_autocor_time'])
autocor_coeff = toarray(cloud['v_dl_autocor_coeff'])
autocorr_lt_thres = np.where(autocor_coeff > 0.8)[0]
i_above_thres = autocorr_lt_thres[-1] if len(autocorr_lt_thres) > 0 else 0
autocorr_at_thres = autocor_time[i_above_thres]*vel if len(autocor_time) > 0 else 0
cth_std = float(cloud['CTH_STD'])
thick_med = float(cloud['Cloud_Thickness_MED'])
thick_std = float(cloud['Cloud_Thickness_STD'])
ct_ac = float(cloud['CTH_autocorr_08_time'])
print(f"{cloud['A_Unique_Identifier']:>18} {ctt:6.2f} {vel:6.2f} |\
{cth_std:6.1f} {thick_med:6.1f} {thick_std:6.1f} {ct_ac:6.1f} |\
{i_above_thres:3} {autocorr_at_thres:7.2f}")
ax.plot(autocor_time*vel, autocor_coeff,
linewidth=1.5)
ax.axhline(0.8, linewidth=1.2, color='dimgrey', linestyle='--')
ax.set_xscale('log')
if hvel_corr:
ax.set_xlim([1e0, 1e5])
ax.set_xlabel('Shift [m]', fontsize=12)
else:
ax.set_xlim([1e0, 3e3])
ax.set_xlabel('Shift [s]', fontsize=12)
ax.set_ylabel('DL vel. autocorr.', fontsize=12)
ax.set_ylim([-0.1, 1.05])
#ax.yaxis.set_minor_locator(matplotlib.ticker.MultipleLocator(0.5))
ax.yaxis.set_minor_locator(matplotlib.ticker.AutoMinorLocator())
ax.tick_params(axis='both', which='major', top=True, right=True,
width=1.5, length=4, labelsize=11)
ax.tick_params(axis='both', which='minor', top=True, right=True,
width=1.5, length=2.5)
#ax.legend(fontsize=12)
ax.xaxis.set_minor_locator(matplotlib.ticker.LogLocator(base=10.0,subs=(0.2,0.4,0.6,0.8),numticks=12))
ax.tick_params(axis='both', which='both', right=True, top=True)
ax.tick_params(axis='both', which='major', labelsize=12,
width=2, length=5.5)
ax.tick_params(axis='both', which='minor', width=1.3, length=3)
fig.tight_layout()
savename = "../plots/case/{}_shaun_autocorr_VEL.png".format(cloud['A_Unique_Identifier'])
fig.savefig(savename, dpi=250)
fig, ax = plt.subplots(figsize=[2.5, 2.1])
colors = ['darkorange', 'lightseagreen', 'royalblue']
colors = ['#FF5126', '#37DC94', '#162C9B']
for i, cloud in enumerate(sorted_day[dt.strftime("%Y%m%d")]['clouds']):
ctt = float(cloud['CTT'])-273.15
if hvel_corr:
vel = max(float(cloud['VEL']),0.1)
else:
vel = 1
autocor_time = toarray(cloud['v_dl_autocor_time'])
autocor_coeff = toarray(cloud['v_dl_autocor_coeff'])
autocorr_lt_thres = np.where(autocor_coeff > 0.8)[0]
i_above_thres = autocorr_lt_thres[-1] if len(autocorr_lt_thres) > 0 else 0
autocorr_at_thres = autocor_time[i_above_thres]*vel if len(autocor_time) > 0 else 0
cth_std = float(cloud['CTH_STD'])
thick_med = float(cloud['Cloud_Thickness_MED'])
thick_std = float(cloud['Cloud_Thickness_STD'])
ct_ac = float(cloud['CTH_autocorr_08_time'])
print(f"{cloud['A_Unique_Identifier']:>18} {ctt:6.2f} {vel:6.2f} |\
{cth_std:6.1f} {thick_med:6.1f} {thick_std:6.1f} {ct_ac:6.1f} |\
{i_above_thres:3} {autocorr_at_thres:7.2f}")
ax.plot(autocor_time*vel, autocor_coeff,
linewidth=1.5, color=colors[i])
ax.plot([1e-1,7e3], [0.8, 0.8],
linewidth=1.2, color='dimgrey', linestyle='--')
ax.set_xscale('log')
if hvel_corr:
ax.set_xlim([1e0, 1e5])
ax.set_xlabel('Shift [m]', fontsize=12)
else:
ax.set_xlim([1e0, 3e3])
ax.set_xlabel('Shift [s]', fontsize=12)
ax.set_ylabel('Vel. autocorr.', fontsize=12)
ax.set_ylim([-0.1, 1.05])
#ax.yaxis.set_minor_locator(matplotlib.ticker.MultipleLocator(0.5))
ax.yaxis.set_minor_locator(matplotlib.ticker.AutoMinorLocator())
ax.tick_params(axis='both', which='major', top=True, right=True,
width=1.5, length=4, labelsize=11)
ax.tick_params(axis='both', which='minor', top=True, right=True,
width=1.5, length=2.5)
#ax.legend(fontsize=12)
ax.xaxis.set_minor_locator(matplotlib.ticker.LogLocator(base=10.0,subs=(0.5,1), numticks=100))
ax.xaxis.set_minor_formatter(matplotlib.ticker.NullFormatter())
ax.xaxis.set_major_locator(matplotlib.ticker.LogLocator(base=100.0, subs=(1.0,), numticks=100))
ax.tick_params(axis='both', which='both', right=True, top=True)
ax.tick_params(axis='both', which='major', labelsize=12,
width=2, length=5.5)
ax.tick_params(axis='both', which='minor', width=1.3, length=3)
# hide top and right borders
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
fig.tight_layout()
savename = "../plots/case/{}_DL_combined_autocorr.png".format(dt.strftime('%Y%m%d'))
fig.savefig(savename, dpi=250, transparent=True)
# -
| analysis_code/2021-02-19_plot_case_study_paper.pub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow import keras
from tensorflow.keras import *
from tensorflow.keras.models import *
from tensorflow.keras.layers import *
from tensorflow.keras.regularizers import l2#正则化L2
import tensorflow as tf
import numpy as np
import pandas as pd
normal_all = pd.read_csv(r'F:\张老师课题学习内容\code\数据集\试验数据(包括压力脉动和振动)\2013.9.12-未发生缠绕前\2013-9-12压力脉动\1250-txt\normal_350_通道7.txt')
chanrao_all = pd.read_csv(r'F:\张老师课题学习内容\code\数据集\试验数据(包括压力脉动和振动)\2013.9.17-发生缠绕后\压力脉动\1250-txt\chanrao_350_通道7.txt')
normal=normal_all[["通道7"]]
chanrao=chanrao_all[["通道7"]]
chanrao=chanrao[0:120000]
#水泵的两种故障类型信号normal正常,chanrao故障
normal=normal.values.reshape(-1, 800)#(120000,1)-(150, 800)150条长度为800
chanrao=chanrao.values.reshape(-1, 800)
print(normal_all.shape,chanrao_all.shape)
print(normal.shape,chanrao.shape)
# +
import numpy as np
def yuchuli(data,label):#(7:1)(616:88)
#打乱数据顺序
np.random.shuffle(data)
train = data[0:120,:]
test = data[120:150,:]
label_train = np.array([label for i in range(0,120)])
label_test =np.array([label for i in range(0,30)])
return train,test ,label_train ,label_test
def stackkk(a,b,c,d,e,f,g,h):
aa = np.vstack((a, e))
bb = np.vstack((b, f))
cc = np.hstack((c, g))
dd = np.hstack((d, h))
return aa,bb,cc,dd
x_tra0,x_tes0,y_tra0,y_tes0 = yuchuli(normal,0)
x_tra1,x_tes1,y_tra1,y_tes1 = yuchuli(chanrao,1)
tr1,te1,yr1,ye1=stackkk(x_tra0,x_tes0,y_tra0,y_tes0 ,x_tra1,x_tes1,y_tra1,y_tes1)
x_train=tr1
x_test=te1
y_train = yr1
y_test = ye1
#打乱数据
state = np.random.get_state()
np.random.shuffle(x_train)
np.random.set_state(state)
np.random.shuffle(y_train)
state = np.random.get_state()
np.random.shuffle(x_test)
np.random.set_state(state)
np.random.shuffle(y_test)
#对训练集和测试集标准化
def ZscoreNormalization(x):
"""Z-score normaliaztion"""
x = (x - np.mean(x)) / np.std(x)
return x
x_train=ZscoreNormalization(x_train)
x_test=ZscoreNormalization(x_test)
# print(x_test[0])
#转化为一维序列
x_train = x_train.reshape(-1,800,1)
x_test = x_test.reshape(-1,800,1)
print(x_train.shape,x_test.shape)
def to_one_hot(labels,dimension=2):
results = np.zeros((len(labels),dimension))
for i,label in enumerate(labels):
results[i,label] = 1
return results
one_hot_train_labels = to_one_hot(y_train)
one_hot_test_labels = to_one_hot(y_test)
# -
x = layers.Input(shape=[800,1,1])
#普通卷积层
conv1 = layers.Conv2D(filters=16, kernel_size=(2, 1), activation='relu',padding='valid',name='conv1')(x)
#池化层
POOL1 = MaxPooling2D((2,1))(conv1)
#普通卷积层
conv2 = layers.Conv2D(filters=32, kernel_size=(2, 1), activation='relu',padding='valid',name='conv2')(POOL1)
#池化层
POOL2 = MaxPooling2D((2,1))(conv2)
#Dropout层
Dropout=layers.Dropout(0.1)(POOL2 )
Flatten=layers.Flatten()(Dropout)
#全连接层
Dense1=layers.Dense(100, activation='relu')(Flatten)
Dense2=layers.Dense(2, activation='softmax')(Dense1)
model = keras.Model(x, Dense2)
model.summary()
# +
#定义优化
model.compile(loss='categorical_crossentropy',
optimizer='adam',metrics=['accuracy'])
# -
import time
time_begin = time.time()
history = model.fit(x_train,one_hot_train_labels,
validation_split=0.1,
epochs=50,batch_size=10,
shuffle=True)
time_end = time.time()
time = time_end - time_begin
print('time:', time)
# +
import time
time_begin = time.time()
score = model.evaluate(x_test,one_hot_test_labels, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
time_end = time.time()
time = time_end - time_begin
print('time:', time)
# +
#绘制acc-loss曲线
import matplotlib.pyplot as plt
plt.plot(history.history['loss'],color='r')
plt.plot(history.history['val_loss'],color='g')
plt.plot(history.history['accuracy'],color='b')
plt.plot(history.history['val_accuracy'],color='k')
plt.title('model loss and acc')
plt.ylabel('Accuracy')
plt.xlabel('epoch')
plt.legend(['train_loss', 'test_loss','train_acc', 'test_acc'], loc='center right')
# plt.legend(['train_loss','train_acc'], loc='upper left')
#plt.savefig('1.png')
plt.show()
# +
import matplotlib.pyplot as plt
plt.plot(history.history['loss'],color='r')
plt.plot(history.history['accuracy'],color='b')
plt.title('model loss and sccuracy ')
plt.ylabel('loss/sccuracy')
plt.xlabel('epoch')
plt.legend(['train_loss', 'train_sccuracy'], loc='center right')
plt.show()
# -
| BCNcode/1_pressure pulsation signal/1250/TRANSMITTER NO.2/CNN/CNN_1250-Q=350.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
s = '<EMAIL>, <EMAIL>, <EMAIL>'
result = re.findall(r'[a-z]+@[a-z]+\.[a-z]+', s)
print(result)
print(len(result))
result = re.findall(r'([a-z]+)@([a-z]+)\.([a-z]+)', s)
print(result)
result = re.findall(r'(([a-z]+)@([a-z]+)\.([a-z]+))', s)
print(result)
result = re.findall('[0-9]+', s)
print(result)
s = '<EMAIL>, <EMAIL>, <EMAIL>'
result = re.finditer(r'[a-z]+@[a-z]+\.[a-z]+', s)
print(result)
print(type(result))
for m in result:
print(m)
l = list(re.finditer(r'[a-z]+@[a-z]+\.[a-z]+', s))
print(l)
print(l[0])
print(type(l[0]))
print(l[0].span())
print([m.span() for m in re.finditer(r'[a-z]+@[a-z]+\.[a-z]+', s)])
result = re.finditer(r'[a-z]+@[a-z]+\.[a-z]+', s)
for m in result:
print(m)
print(list(result))
| notebook/re_findall_finditer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Heading
print("O<NAME>")
a=5
b=2
a**b
a//b
a/float(b)
a%b
min(a,b)
a != b
a += 3
a = [1, "hello", 5.5]
a
len(a)
a[0] # First element index is 0
a.append("how are you?")
a.pop(1)
a
for x in a:
print(x)
print(a)
for i,x in enumerate(a):
print("element {}: {}".format(i, x))
a[0] = 10
a
b = (-1, "ola", 2.5) # Python Tuple
b[1] = "Goodbye"
x,y,z = b
y
a = {"name":"Mary", "age":23, "sign":"capricorn"} # Python Dictionary
a["name"]
a["job"] = "student"
a
# ## Python Functions
def func2(a, b=4, c=5):
if ( a > 2 and b < 10):
return a
elif c == 5:
return b
else:
return a + b +c
func2(4,c=6, b = 11)
# Numpy Module
import numpy as np
a = np.array([0, 2, 4, 6, 8, 10, 12,14,16])
a
a.shape
a[::-1]
a=np.array([[0, 1, 2, 3], [4,5,6,7],[8, 9, 10, 11]])
a
a.shape
a[:,0:2]
a.T
np.mean(a)
a.mean(axis=1)
import matplotlib.pyplot as plt
# %matplotlib inline
x=np.linspace(-5,5, 50)
y = np.sin(x)
y2 = y ** 2
y3 = -x / 5.0
plt.figure()
plt.plot(x,y,label='sin')
plt.plot(x,y2, '.', label='$\sin^{2}$')
plt.plot(x,y3,linewidth=3)
plt.xlabel("X axis")
plt.ylabel("Y axis")
plt.legend()
plt.show()
fig, ax = plt.subplots(2, sharex = True)
ax[0].plot(x,y)
ax[1].plot(x,y2)
ax[1].set_ylabel('y axis')
plt.show()
y,x = np.mgrid[0:20, 0:30]
z = (x-4)**2 + y **2
plt.figure()
plt.pcolormesh(x,y,z)
plt.show()
# SciPy
from scipy.optimize import curve_fit
def f(x,a,b,c):
return a * np.exp(-b * x ) +c
n=60
x = np.linspace(0,5, n)
y = f(x, 5, 2, 0.5) + 2 * np.random.rand(n)
popt, pcov = curve_fit (f, x, y)
perr = np.sqrt(np.diag(pcov))
y_fit = f(x, *popt)
msd = np.sum((y-y_fit) ** 2)/n
pnames = ['a', 'b, c']
result = ''
for na
| PythonBasicCourse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="YHWCIX5PnBk0" colab_type="code" colab={}
# !pip install datadotworld
# !pip install datadotworld[pandas]
# + id="_SkAGgZU0N9H" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="72tR4vT0sTc9" colab_type="code" colab={}
from google.colab import drive
# + id="7pLLOisoqUrZ" colab_type="code" colab={}
# !dw configure
# + id="9mKDoPrqqnbk" colab_type="code" colab={}
import datadotworld as dw
# + id="YKotQGLUrQtV" colab_type="code" colab={}
drive.mount("/content/drive")
# + id="uQnciXExshgD" colab_type="code" colab={}
# cd "drive/My Drive/Colab Notebooks/First_task"
# + id="Jo9a0pi9utvk" colab_type="code" colab={}
# !git clone {GITHUB_URL}
# + id="BB95W7H4vhLg" colab_type="code" colab={}
# cd dw_matix/
# + id="K1paYvBjwdKn" colab_type="code" colab={}
data = dw.load_dataset('datafiniti/womens-shoe-prices')
# + id="dgMc3BtBwubg" colab_type="code" colab={}
df = data.dataframes["7003_1"]
df.shape
# + colab_type="code" id="f8iOQbHTLl_n" colab={}
df.columns
# + colab_type="code" id="7IsPhkKALlSK" colab={}
df.prices_currency.unique()
# + id="6WPa4l_pxYUK" colab_type="code" colab={}
df.prices_currency.value_counts()
# + id="gKIC-CJFxknX" colab_type="code" colab={}
df_usd = df[df.prices_currency == "USD"].copy()
# + id="8lVODkrhx6WQ" colab_type="code" colab={}
df.prices_amountmin.unique()
# + id="tW7gCg3HzU1G" colab_type="code" colab={}
df_usd['prices_amountmin'] = df_usd.prices_amountmin.astype(np.float)
price_min = df_usd['prices_amountmin']
# + id="BiHul9Nl1URf" colab_type="code" outputId="6ecabef4-8990-4f77-b72e-969362c4794c" executionInfo={"status": "ok", "timestamp": 1581620370103, "user_tz": -60, "elapsed": 869, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07118407884086592178"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
filter_max = np.percentile(price_min, 99)
filter_max
# + id="5C9ZvoF82CHT" colab_type="code" colab={}
price_min < filter_max
df_usd_filter =df_usd[price_min < filter_max]
# + id="dgalCDKD3Vok" colab_type="code" outputId="bec9bdc4-b3f7-42f6-f011-e0e0beb13ded" executionInfo={"status": "ok", "timestamp": 1581620377840, "user_tz": -60, "elapsed": 982, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07118407884086592178"}} colab={"base_uri": "https://localhost:8080/", "height": 285}
df_usd_filter.prices_amountmin.hist(bins=200)
# + id="LXZkvRZ869Fl" colab_type="code" colab={}
df_usd_filter.to_csv('data/women_shoes.csv', index=False)
| transformacja_1/Day3.ipynb |