code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Modules
# +
import tensorflow as tf
print('tensorflow version:', tf.__version__)
import pandas as pd
print('pandas version:', pd.__version__)
import numpy as np
print('numpy version:', np.__version__)
import sklearn
print('sklearn version:', sklearn.__version__)
from sklearn import preprocessing
import matplotlib
print('matplotlib version:', matplotlib.__version__)
import matplotlib.pyplot as plt
# -
# # Configs
# +
# With numpy, when a value is printed display more values per line
# https://stackoverflow.com/questions/21971449/how-do-i-increase-the-cell-width-of-the-jupyter-ipython-notebook-in-my-browser
np.set_printoptions(linewidth=5000)
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# In Pandas, display more rows and columns
# https://stackoverflow.com/a/11711637/4375369
pd.set_option('display.max_rows', 100)
pd.set_option('display.max_columns', 100)
SEED_FOR_REPRODUCABILITY = 777
# -
# # Retrieve Dataset
# - Mnist digits
# +
(X_train_raw, y_train_raw), (X_test_raw, y_test_raw) = tf.keras.datasets.mnist.load_data()
X_train_raw, y_train_raw, X_test_raw, y_test_raw
# -
# # Prepare the datasets
# ## Shuffle the data sets
# - https://stackoverflow.com/questions/35076223/how-to-randomly-shuffle-data-and-target-in-python
# +
X_train_raw_shuffled, y_train_raw_shuffled = sklearn.utils.shuffle(X_train_raw, y_train_raw, random_state=SEED_FOR_REPRODUCABILITY)
X_test_raw_shuffled, y_test_raw_shuffled = sklearn.utils.shuffle(X_test_raw, y_test_raw, random_state=SEED_FOR_REPRODUCABILITY)
sample_index = 1
X_train_raw[sample_index], y_train_raw[sample_index], X_train_raw_shuffled[sample_index], y_train_raw_shuffled[sample_index],X_test_raw[sample_index], y_test_raw[sample_index], X_test_raw_shuffled[sample_index], y_test_raw_shuffled[sample_index]
# -
y_train_raw_shuffled[sample_index]
# ## Normalize the example data sets
# +
X_train_raw_shuffled_max_value = tf.math.reduce_max(X_train_raw_shuffled)
print("Maximim value:", X_train_raw_shuffled_max_value)
X_train_normalized = X_train_raw_shuffled / X_train_raw_shuffled_max_value
X_test_normalized = X_test_raw_shuffled / X_train_raw_shuffled_max_value
print("Before normalization (training):")
print(X_train_raw_shuffled[sample_index])
print("After normalization (training):")
print(X_train_normalized[sample_index].numpy())
print("Before normalization (test):")
print(X_test_raw_shuffled[sample_index])
print("After normalization (test):")
print(X_test_normalized[sample_index].numpy())
# -
# ## One-hot encode the label data
# - https://machinelearningmastery.com/how-to-one-hot-encode-sequence-data-in-python/
# +
y_train_one_hot_encoded = tf.keras.utils.to_categorical(y_train_raw_shuffled)
y_test_one_hot_encoded = tf.keras.utils.to_categorical(y_test_raw_shuffled)
print('train:')
print(y_train_one_hot_encoded[sample_index], y_train_raw_shuffled[sample_index])
print("test:")
print(y_test_one_hot_encoded[sample_index], y_test_raw_shuffled[sample_index])
# -
# # Summarize Accepted Data
# +
X_train_accepted = tf.expand_dims(X_train_normalized, axis=-1)
y_train_accepted = y_train_one_hot_encoded
X_test_accepted = tf.expand_dims(X_test_normalized, axis=-1)
y_test_accepted = y_test_one_hot_encoded
# -
# # Create the Model Architecture
# +
tf.random.set_seed(SEED_FOR_REPRODUCABILITY)
if model_001:
del(model_001)
model_001 = tf.keras.Sequential([
tf.keras.layers.Conv2D(64, (3, 3), activation=tf.keras.activations.relu, padding="same"),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation=tf.keras.activations.relu, padding="same"),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation=tf.keras.activations.relu, padding="same"),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation=tf.keras.activations.relu, padding="same"),
tf.keras.layers.MaxPool2D(pool_size=(2, 2)),
# tf.keras.layers.Conv2D(10, 3, activation=tf.keras.activations.relu),
# tf.keras.layers.MaxPool2D(
# pool_size=2,
# padding="valid"
# ),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(1000, activation=tf.keras.activations.relu),
tf.keras.layers.Dense(1000, activation=tf.keras.activations.relu),
tf.keras.layers.Dense(100, activation=tf.keras.activations.relu),
tf.keras.layers.Dense(100, activation=tf.keras.activations.relu),
tf.keras.layers.Dense(10, activation=tf.keras.activations.relu),
tf.keras.layers.Dense(10, activation=tf.keras.activations.softmax)
])
model_001.compile(
loss=tf.keras.losses.CategoricalCrossentropy(),
optimizer=tf.keras.optimizers.Adam(),
metrics=[ "accuracy" ]
)
model_001.build(input_shape=X_train_accepted.shape)
model_001.summary()
tf.keras.utils.plot_model(model_001, show_shapes=True)
# -
# # Fit the Architecture to the Data
# - Do the training
# +
def scheduler(epoch, lr):
print('learning rate', lr, 'epoch', epoch)
return 0.001
if epoch < 10:
return 0.001
else:
return 0.0001
# elif epoch < 10:
# return 0.0001
# elif epoch < 15:
# return 0.00001
# elif epoch < 20:
# return 0.000001
# else:
# return 0.0000001
learning_rate_scheduler_callback = tf.keras.callbacks.LearningRateScheduler(scheduler)
model_001_history = model_001.fit(
X_train_accepted,
y_train_accepted,
validation_data=(X_test_accepted, y_test_accepted),
epochs=25,
callbacks= [ learning_rate_scheduler_callback ]
)
# -
model_001.evaluate(X_test_accepted, y_test_accepted)
pd.DataFrame(model_001_history.history)
pd.DataFrame(model_001_history.history).plot()
plt.ylabel("loss/accuracy")
plt.xlabel("epochs")
pd.DataFrame(model_001_history.history['accuracy']).plot()
plt.ylabel("training accuracy")
plt.xlabel("epochs")
pd.DataFrame(model_001_history.history['val_accuracy']).plot()
plt.ylabel("validation accuracy")
plt.xlabel("epochs")
pd.DataFrame(model_001_history.history['val_loss']).plot()
plt.ylabel("validation loss")
plt.xlabel("epochs")
# # Summary
#
# - In this practice I found it challenging to get the above 99% that was achieved in the previous practice
# - I think this is because the test data was incorrect in the last practice. I had split the test data into test and validation data, without shuffling first. Today I just kept it as one and used the test for the validation as well.
# - I was still able to achieve >99% but after much tweaking of the architecture
# - In a future practice I would like to store model checkpoints as the training occurs to preserve the best found models
# - Additionally, it would be nice to try to generate more test data, by rotating the existing test data. Yesterday it seemed like some were not found because of rotation.
| practiceInstances/002MnistHandwritingRecognition/practice_002_mnist_handwriting_recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Refitting NumPyro models with ArviZ (and xarray)
#
# ArviZ is backend agnostic and therefore does not sample directly. In order to take advantage of algorithms that require refitting models several times, ArviZ uses `SamplingWrappers` to convert the API of the sampling backend to a common set of functions. Hence, functions like Leave Future Out Cross Validation can be used in ArviZ independently of the sampling backend used.
# Below there is an example of `SamplingWrapper` usage for [NumPyro](https://pyro.ai/numpyro/).
import arviz as az
import numpyro
import numpyro.distributions as dist
import jax.random as random
from numpyro.infer import MCMC, NUTS
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
import xarray as xr
numpyro.set_host_device_count(4)
# For this example, we will use a linear regression model.
# +
np.random.seed(26)
xdata = np.linspace(0, 50, 100)
b0, b1, sigma = -2, 1, 3
ydata = np.random.normal(loc=b1 * xdata + b0, scale=sigma)
# -
plt.plot(xdata, ydata)
# Now we will write the NumPyro Code:
def model(N, x, y=None):
b0 = numpyro.sample("b0", dist.Normal(0, 10))
b1 = numpyro.sample("b1", dist.Normal(0, 10))
sigma_e = numpyro.sample("sigma_e", dist.HalfNormal(10))
numpyro.sample("y", dist.Normal(b0 + b1 * x, sigma_e), obs=y)
data_dict = {
"N": len(ydata),
"y": ydata,
"x": xdata,
}
kernel = NUTS(model)
sample_kwargs = dict(
sampler=kernel,
num_warmup=1000,
num_samples=1000,
num_chains=4,
chain_method="parallel"
)
mcmc = MCMC(**sample_kwargs)
mcmc.run(random.PRNGKey(0), **data_dict)
# We have defined a dictionary `sample_kwargs` that will be passed to the `SamplingWrapper` in order to make sure that all
# refits use the same sampler parameters. We follow the same pattern with {func}`az.from_numpyro <arviz.from_numpyro>`.
dims = {"y": ["time"], "x": ["time"]}
idata_kwargs = {
"dims": dims,
"constant_data": {"x": xdata}
}
idata = az.from_numpyro(mcmc, **idata_kwargs)
del idata.log_likelihood
idata
# We are now missing the `log_likelihood` group because we have not used the `log_likelihood` argument in `idata_kwargs`. We are doing this to ease the job of the sampling wrapper. Instead of going out of our way to get Stan to calculate the pointwise log likelihood values for each refit and for the excluded observation at every refit, we will compromise and manually write a function to calculate the pointwise log likelihood.
#
# Even though it is not ideal to lose part of the straight out of the box capabilities of PyStan-ArviZ integration, this should generally not be a problem. We are basically moving the pointwise log likelihood calculation from the Stan Code to the Python code, in both cases, we need to manually write the function to calculate the pointwise log likelihood.
#
# Moreover, the Python computation could even be written to be compatible with [Dask](https://docs.dask.org/en/latest/). Thus it will work even in cases where the large number of observations makes it impossible to store pointwise log likelihood values (with shape `n_samples * n_observations`) in memory.
def calculate_log_lik(x, y, b0, b1, sigma_e):
mu = b0 + b1 * x
return stats.norm(mu, sigma_e).logpdf(y)
# This function should work for any shape of the input arrays as long as their shapes are compatible and can broadcast. There is no need to loop over each draw in order to calculate the pointwise log likelihood using scalars.
#
# Therefore, we can use {func}`xr.apply_ufunc <xarray.apply_ufunc>` to handle the broadcasting and preserve the dimension names:
log_lik = xr.apply_ufunc(
calculate_log_lik,
idata.constant_data["x"],
idata.observed_data["y"],
idata.posterior["b0"],
idata.posterior["b1"],
idata.posterior["sigma_e"],
)
idata.add_groups(log_likelihood=log_lik)
# The first argument is the function, followed by as many positional arguments as needed by the function, 5 in our case. As this case does not have many different dimensions nor combinations of these, we do not need to use any extra kwargs passed to `xr.apply_ufunc`.
#
# We are now passing the arguments to `calculate_log_lik` initially as `xr.DataArrays`. What is happening here behind the scenes is that `xr.apply_ufunc` is broadcasting and aligning the dimensions of all the DataArrays involved and afterwards passing NumPy arrays to `calculate_log_lik`. Everything works automagically.
#
# Now let's see what happens if we were to pass the arrays directly to `calculate_log_lik` instead:
calculate_log_lik(
idata.constant_data["x"].values,
idata.observed_data["y"].values,
idata.posterior["b0"].values,
idata.posterior["b1"].values,
idata.posterior["sigma_e"].values
)
# If you are still curious about the magic of xarray and `apply_ufunc`, you can also try to modify the `dims` used to generate the `InferenceData` a couple cells before:
#
# dims = {"y": ["time"], "x": ["time"]}
#
# What happens to the result if you use a different name for the dimension of `x`?
idata
# We will create a subclass of {class}`~arviz.SamplingWrapper`. Therefore, instead of having to implement all functions required by {func}`~arviz.reloo` we only have to implement {func}`~arviz.SamplingWrapper.sel_observations` (we are cloning {func}`~arviz.SamplingWrapper.sample` and {func}`~arviz.SamplingWrapper.get_inference_data` from the {class}`~arviz.SamplingWrapper` in order to use `apply_ufunc` instead of assuming the log likelihood is calculated within Stan).
#
# Let's check the 2 outputs of `sel_observations`.
# 1. `data__i` is a dictionary because it is an argument of `sample` which will pass it as is to `model.sampling`.
# 2. `data_ex` is a list because it is an argument to `log_likelihood__i` which will pass it as `*data_ex` to `apply_ufunc`.
#
# More on `data_ex` and `apply_ufunc` integration is given below.
# +
class NumPyroSamplingWrapper(az.SamplingWrapper):
def __init__(self, model, **kwargs):
self.rng_key = kwargs.pop("rng_key", random.PRNGKey(0))
super(NumPyroSamplingWrapper, self).__init__(model, **kwargs)
def sample(self, modified_observed_data):
self.rng_key, subkey = random.split(self.rng_key)
mcmc = MCMC(**self.sample_kwargs)
mcmc.run(subkey, **modified_observed_data)
return mcmc
def get_inference_data(self, fit):
# Cloned from PyStanSamplingWrapper.
idata = az.from_numpyro(mcmc, **self.idata_kwargs)
return idata
class LinRegWrapper(NumPyroSamplingWrapper):
def sel_observations(self, idx):
xdata = self.idata_orig.constant_data["x"]
ydata = self.idata_orig.observed_data["y"]
mask = np.isin(np.arange(len(xdata)), idx)
# data__i is passed to numpyro to sample on it -> dict of numpy array
# data_ex is passed to apply_ufunc -> list of DataArray
data__i = {"x": xdata[~mask].values, "y": ydata[~mask].values, "N": len(ydata[~mask])}
data_ex = [xdata[mask], ydata[mask]]
return data__i, data_ex
# -
loo_orig = az.loo(idata, pointwise=True)
loo_orig
# In this case, the Leave-One-Out Cross Validation (LOO-CV) approximation using [Pareto Smoothed Importance Sampling](https://arxiv.org/abs/1507.02646) (PSIS) works for all observations, so we will use modify `loo_orig` in order to make {func}`~arviz.reloo` believe that PSIS failed for some observations. This will also serve as a validation of our wrapper, as the PSIS LOO-CV already returned the correct value.
loo_orig.pareto_k[[13, 42, 56, 73]] = np.array([0.8, 1.2, 2.6, 0.9])
# We initialize our sampling wrapper. Let's stop and analyze each of the arguments.
#
# We use the `log_lik_fun` and `posterior_vars` argument to tell the wrapper how to call {func}`~xarray:xarray.apply_ufunc`. `log_lik_fun` is the function to be called, which is then called with the following positional arguments:
#
# log_lik_fun(*data_ex, *[idata__i.posterior[var_name] for var_name in posterior_vars]
#
# where `data_ex` is the second element returned by `sel_observations` and `idata__i` is the `InferenceData` object result of `get_inference_data` which contains the fit on the subsetted data. We have generated `data_ex` to be a tuple of DataArrays so it plays nicely with this call signature.
#
# We use `idata_orig` as a starting point, and mostly as a source of observed and constant data which is then subsetted in `sel_observations`.
#
# Finally, `sample_kwargs` and `idata_kwargs` are used to make sure all refits and corresponding `InferenceData` are generated with the same properties.
pystan_wrapper = LinRegWrapper(
mcmc,
rng_key=random.PRNGKey(7),
log_lik_fun=calculate_log_lik,
posterior_vars=("b0", "b1", "sigma_e"),
idata_orig=idata,
sample_kwargs=sample_kwargs,
idata_kwargs=idata_kwargs
)
# And eventually, we can use this wrapper to call {func}`~arviz.reloo`, and compare the results with the PSIS LOO-CV results.
loo_relooed = az.reloo(pystan_wrapper, loo_orig=loo_orig)
loo_relooed
loo_orig
| doc/source/user_guide/numpyro_refitting_xr_lik.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SVI Part II: Conditional Independence, Subsampling, and Amortization
#
# ## The Goal: Scaling SVI to Large Datasets
#
# For a model with $N$ observations, running the `model` and `guide` and constructing the ELBO involves evaluating log pdf's whose complexity scales badly with $N$. This is a problem if we want to scale to large datasets. Luckily, the ELBO objective naturally supports subsampling provided that our model/guide have some conditional independence structure that we can take advantage of. For example, in the case that the observations are conditionally independent given the latents, the log likelihood term in the ELBO can be approximated with
#
# $$ \sum_{i=1}^N \log p({\bf x}_i | {\bf z}) \approx \frac{N}{M}
# \sum_{i\in{\mathcal{I}_M}} \log p({\bf x}_i | {\bf z}) $$
#
# where $\mathcal{I}_M$ is a mini-batch of indices of size $M$ with $M<N$ (for a discussion please see references [1,2]). Great, problem solved! But how do we do this in Pyro?
#
# ## Marking Conditional Independence in Pyro
#
# If a user wants to do this sort of thing in Pyro, he or she first needs to make sure that the model and guide are written in such a way that Pyro can leverage the relevant conditional independencies. Let's see how this is done. Pyro provides two language primitives for marking conditional independencies: `plate` and `markov`. Let's start with the simpler of the two.
#
# ### sequential `plate`
#
# Let's return to the example we used in the [previous tutorial](svi_part_i.ipynb). For convenience let's replicate the main logic of `model` here:
#
# ```python
# def model(data):
# # sample f from the beta prior
# f = pyro.sample("latent_fairness", dist.Beta(alpha0, beta0))
# # loop over the observed data using pyro.sample with the obs keyword argument
# for i in range(len(data)):
# # observe datapoint i using the bernoulli likelihood
# pyro.sample("obs_{}".format(i), dist.Bernoulli(f), obs=data[i])
# ```
#
# For this model the observations are conditionally independent given the latent random variable `latent_fairness`. To explicitly mark this in Pyro we basically just need to replace the Python builtin `range` with the Pyro construct `plate`:
#
# ```python
# def model(data):
# # sample f from the beta prior
# f = pyro.sample("latent_fairness", dist.Beta(alpha0, beta0))
# # loop over the observed data [WE ONLY CHANGE THE NEXT LINE]
# for i in pyro.plate("data_loop", len(data)):
# # observe datapoint i using the bernoulli likelihood
# pyro.sample("obs_{}".format(i), dist.Bernoulli(f), obs=data[i])
# ```
#
# We see that `pyro.plate` is very similar to `range` with one main difference: each invocation of `plate` requires the user to provide a unique name. The second argument is an integer just like for `range`.
#
# So far so good. Pyro can now leverage the conditional independency of the observations given the latent random variable. But how does this actually work? Basically `pyro.plate` is implemented using a context manager. At every execution of the body of the `for` loop we enter a new (conditional) independence context which is then exited at the end of the `for` loop body. Let's be very explicit about this:
#
# - because each observed `pyro.sample` statement occurs within a different execution of the body of the `for` loop, Pyro marks each observation as independent
# - this independence is properly a _conditional_ independence _given_ `latent_fairness` because `latent_fairness` is sampled _outside_ of the context of `data_loop`.
#
# Before moving on, let's mention some gotchas to be avoided when using sequential `plate`. Consider the following variant of the above code snippet:
#
# ```python
# # WARNING do not do this!
# my_reified_list = list(pyro.plate("data_loop", len(data)))
# for i in my_reified_list:
# pyro.sample("obs_{}".format(i), dist.Bernoulli(f), obs=data[i])
# ```
#
# This will _not_ achieve the desired behavior, since `list()` will enter and exit the `data_loop` context completely before a single `pyro.sample` statement is called. Similarly, the user needs to take care not to leak mutable computations across the boundary of the context manager, as this may lead to subtle bugs. For example, `pyro.plate` is not appropriate for temporal models where each iteration of a loop depends on the previous iteration; in this case a `range` or `pyro.markov` should be used instead.
#
# ## vectorized `plate`
#
# Conceptually vectorized `plate` is the same as sequential `plate` except that it is a vectorized operation (as `torch.arange` is to `range`). As such it potentially enables large speed-ups compared to the explicit `for` loop that appears with sequential `plate`. Let's see how this looks for our running example. First we need `data` to be in the form of a tensor:
#
# ```python
# data = torch.zeros(10)
# data[0:6] = torch.ones(6) # 6 heads and 4 tails
# ```
#
# Then we have:
#
# ```python
# with plate('observe_data'):
# pyro.sample('obs', dist.Bernoulli(f), obs=data)
# ```
#
# Let's compare this to the analogous sequential `plate` usage point-by-point:
# - both patterns requires the user to specify a unique name.
# - note that this code snippet only introduces a single (observed) random variable (namely `obs`), since the entire tensor is considered at once.
# - since there is no need for an iterator in this case, there is no need to specify the length of the tensor(s) involved in the `plate` context
#
# Note that the gotchas mentioned in the case of sequential `plate` also apply to vectorized `plate`.
# ## Subsampling
#
# We now know how to mark conditional independence in Pyro. This is useful in and of itself (see the [dependency tracking section](svi_part_iii.ipynb) in SVI Part III), but we'd also like to do subsampling so that we can do SVI on large datasets. Depending on the structure of the model and guide, Pyro supports several ways of doing subsampling. Let's go through these one by one.
#
# ### Automatic subsampling with `plate`
#
# Let's look at the simplest case first, in which we get subsampling for free with one or two additional arguments to `plate`:
#
# ```python
# for i in pyro.plate("data_loop", len(data), subsample_size=5):
# pyro.sample("obs_{}".format(i), dist.Bernoulli(f), obs=data[i])
# ```
#
# That's all there is to it: we just use the argument `subsample_size`. Whenever we run `model()` we now only evaluate the log likelihood for 5 randomly chosen datapoints in `data`; in addition, the log likelihood will be automatically scaled by the appropriate factor of $\tfrac{10}{5} = 2$. What about vectorized `plate`? The incantantion is entirely analogous:
#
# ```python
# with plate('observe_data', size=10, subsample_size=5) as ind:
# pyro.sample('obs', dist.Bernoulli(f),
# obs=data.index_select(0, ind))
# ```
#
# Importantly, `plate` now returns a tensor of indices `ind`, which, in this case will be of length 5. Note that in addition to the argument `subsample_size` we also pass the argument `size` so that `plate` is aware of the full size of the tensor `data` so that it can compute the correct scaling factor. Just like for sequential `plate`, the user is responsible for selecting the correct datapoints using the indices provided by `plate`.
#
# Finally, note that the user must pass a `device` argument to `plate` if `data` is on the GPU.
#
# ### Custom subsampling strategies with `plate`
#
# Every time the above `model()` is run `plate` will sample new subsample indices. Since this subsampling is stateless, this can lead to some problems: basically for a sufficiently large dataset even after a large number of iterations there's a nonnegligible probability that some of the datapoints will have never been selected. To avoid this the user can take control of subsampling by making use of the `subsample` argument to `plate`. See [the docs](http://docs.pyro.ai/en/dev/primitives.html#pyro.plate) for details.
#
# ### Subsampling when there are only local random variables
#
# We have in mind a model with a joint probability density given by
#
# $$ p({\bf x}, {\bf z}) = \prod_{i=1}^N p({\bf x}_i | {\bf z}_i) p({\bf z}_i) $$
#
# For a model with this dependency structure the scale factor introduced by subsampling scales all the terms in the ELBO by the same amount. This is the case, for example, for a vanilla VAE. This explains why for the VAE it's permissible for the user to take complete control over subsampling and pass mini-batches directly to the model and guide; `plate` is still used, but `subsample_size` and `subsample` are not. To see how this looks in detail, see the [VAE tutorial](vae.ipynb).
#
#
# ### Subsampling when there are both global and local random variables
#
# In the coin flip examples above `plate` appeared in the model but not in the guide, since the only thing being subsampled was the observations. Let's look at a more complicated example where subsampling appears in both the model and guide. To make things simple let's keep the discussion somewhat abstract and avoid writing a complete model and guide.
#
# Consider the model specified by the following joint distribution:
#
# $$ p({\bf x}, {\bf z}, \beta) = p(\beta)
# \prod_{i=1}^N p({\bf x}_i | {\bf z}_i) p({\bf z}_i | \beta) $$
#
# There are $N$ observations $\{ {\bf x}_i \}$ and $N$ local latent random variables
# $\{ {\bf z}_i \}$. There is also a global latent random variable $\beta$. Our guide will be factorized as
#
# $$ q({\bf z}, \beta) = q(\beta) \prod_{i=1}^N q({\bf z}_i | \beta, \lambda_i) $$
#
# Here we've been explicit about introducing $N$ local variational parameters
# $\{\lambda_i \}$, while the other variational parameters are left implicit. Both the model and guide have conditional independencies. In particular, on the model side, given the $\{ {\bf z}_i \}$ the observations $\{ {\bf x}_i \}$ are independent. In addition, given $\beta$ the latent random variables $\{\bf {z}_i \}$ are independent. On the guide side, given the variational parameters $\{\lambda_i \}$ and $\beta$ the latent random variables $\{\bf {z}_i \}$ are independent. To mark these conditional independencies in Pyro and do subsampling we need to make use of `plate` in _both_ the model _and_ the guide. Let's sketch out the basic logic using sequential `plate` (a more complete piece of code would include `pyro.param` statements, etc.). First, the model:
#
# ```python
# def model(data):
# beta = pyro.sample("beta", ...) # sample the global RV
# for i in pyro.plate("locals", len(data)):
# z_i = pyro.sample("z_{}".format(i), ...)
# # compute the parameter used to define the observation
# # likelihood using the local random variable
# theta_i = compute_something(z_i)
# pyro.sample("obs_{}".format(i), dist.MyDist(theta_i), obs=data[i])
# ```
#
# Note that in contrast to our running coin flip example, here we have `pyro.sample` statements both inside and outside of the `plate` loop. Next the guide:
#
# ```python
# def guide(data):
# beta = pyro.sample("beta", ...) # sample the global RV
# for i in pyro.plate("locals", len(data), subsample_size=5):
# # sample the local RVs
# pyro.sample("z_{}".format(i), ..., lambda_i)
# ```
#
# Note that crucially the indices will only be subsampled once in the guide; the Pyro backend makes sure that the same set of indices are used during execution of the model. For this reason `subsample_size` only needs to be specified in the guide.
# ## Amortization
#
# Let's again consider a model with global and local latent random variables and local variational parameters:
#
# $$ p({\bf x}, {\bf z}, \beta) = p(\beta)
# \prod_{i=1}^N p({\bf x}_i | {\bf z}_i) p({\bf z}_i | \beta) \qquad \qquad
# q({\bf z}, \beta) = q(\beta) \prod_{i=1}^N q({\bf z}_i | \beta, \lambda_i) $$
#
# For small to medium-sized $N$ using local variational parameters like this can be a good approach. If $N$ is large, however, the fact that the space we're doing optimization over grows with $N$ can be a real probelm. One way to avoid this nasty growth with the size of the dataset is *amortization*.
#
# This works as follows. Instead of introducing local variational parameters, we're going to learn a single parametric function $f(\cdot)$ and work with a variational distribution that has the form
#
# $$q(\beta) \prod_{n=1}^N q({\bf z}_i | f({\bf x}_i))$$
#
# The function $f(\cdot)$—which basically maps a given observation to a set of variational parameters tailored to that datapoint—will need to be sufficiently rich to capture the posterior accurately, but now we can handle large datasets without having to introduce an obscene number of variational parameters.
# This approach has other benefits too: for example, during learning $f(\cdot)$ effectively allows us to share statistical power among different datapoints. Note that this is precisely the approach used in the [VAE](vae.ipynb).
#
# ## Tensor shapes and vectorized `plate`
#
# The usage of `pyro.plate` in this tutorial was limited to relatively simple cases. For example, none of the `plate`s were nested inside of other `plate`s. In order to make full use of `plate`, the user must be careful to use Pyro's tensor shape semantics. For a discussion see the [tensor shapes tutorial](tensor_shapes.ipynb).
# ## References
#
# [1] `Stochastic Variational Inference`,
# <br/>
# <NAME>, <NAME>, <NAME>, <NAME>
#
# [2] `Auto-Encoding Variational Bayes`,<br/>
# <NAME>, <NAME>
| tutorial/source/svi_part_ii.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:85% !important; }</style>"))
# %load_ext autoreload
# +
import os
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.python.client import device_lib
from tensorflow.keras import layers,models,utils
from tensorflow.keras import optimizers
from tensorflow.keras.applications import VGG16
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import matplotlib.pyplot as plt
# +
def get_available_gpus():
local_device_protos = device_lib.list_local_devices()
return [x.name for x in local_device_protos if x.device_type == 'GPU']
get_available_gpus()
# +
#import tensorflow as tf
with tf.device('/gpu:0'):
a = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a')
b = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b')
c = tf.matmul(a, b)
with tf.Session() as sess:
print (sess.run(c))
# -
conv_base = VGG16(weights='imagenet',include_top=False,input_shape=(150, 150, 3))
# ### Extracting features using the pretrained convolutional base
base_dir = './../data/cats_and_dogs_small/'
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
datagen = ImageDataGenerator(rescale=1./255)
batch_size = 20
def extract_features(directory, sample_count):
features = np.zeros(shape=(sample_count, 4, 4, 512))
labels = np.zeros(shape=(sample_count))
generator = datagen.flow_from_directory(directory, target_size=(150, 150), batch_size=batch_size, class_mode='binary')
i = 0
for inputs_batch, labels_batch in generator:
features_batch = conv_base.predict(inputs_batch)
features[i * batch_size : (i + 1) * batch_size] = features_batch
labels[i * batch_size : (i + 1) * batch_size] = labels_batch
i += 1
if i * batch_size >= sample_count:
break
return features, labels
train_features, train_labels = extract_features(train_dir, 2000)
validation_features, validation_labels = extract_features(validation_dir, 1000)
test_features, test_labels = extract_features(test_dir, 1000)
train_features = np.reshape(train_features, (2000, 4 * 4 * 512))
validation_features = np.reshape(validation_features, (1000, 4 * 4 * 512))
test_features = np.reshape(test_features, (1000, 4 * 4 * 512))
model = models.Sequential()
model.add(layers.Dense(256, activation='relu', input_dim=4 * 4 * 512))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer=optimizers.RMSprop(lr=2e-5), loss='binary_crossentropy',metrics=['acc'])
history = model.fit(train_features, train_labels, epochs=30, batch_size=20,
validation_data=(validation_features, validation_labels))
# +
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.figure(figsize=(15,7.5))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure(figsize=(15,7.5))
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
# ### Adding a densely connected classifier on top of the convolutional base
model = models.Sequential()
model.add(conv_base)
model.add(layers.Flatten())
model.add(layers.Dense(256, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.summary()
print('This is the number of trainable weights before freezing the conv base:', len(model.trainable_weights))
conv_base.trainable = False
print('This is the number of trainable weights after freezing the conv base:', len(model.trainable_weights))
train_datagen = ImageDataGenerator(rescale=1./255, rotation_range=40, width_shift_range=0.2, height_shift_range=0.2, shear_range=0.2,
zoom_range=0.2, horizontal_flip=True,)
test_datagen = ImageDataGenerator(rescale=1./255)
# +
train_generator = train_datagen.flow_from_directory(train_dir, target_size=(150, 150),
batch_size=20, class_mode='binary')
validation_generator = test_datagen.flow_from_directory(validation_dir,target_size=(150, 150),
batch_size=20,class_mode='binary')
# -
model.compile(optimizer=optimizers.RMSprop(lr=2e-5), loss='binary_crossentropy',metrics=['acc'])
history = model.fit_generator(train_generator, steps_per_epoch=100, epochs=30, validation_data=validation_generator, validation_steps=50)
# +
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.figure(figsize=(15,7.5))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure(figsize=(15,7.5))
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
# ### Fine tuning the model
conv_base.summary()
# +
conv_base.trainable = True
set_trainable = False
for layer in conv_base.layers:
if layer.name == 'block5_conv1':
set_trainable = True
if set_trainable:
layer.trainable = True
else:
layer.trainable = False
# -
model.compile(optimizer=optimizers.RMSprop(lr=2e-5), loss='binary_crossentropy',metrics=['acc'])
history = model.fit_generator(train_generator, steps_per_epoch=100, epochs=100, validation_data=validation_generator, validation_steps=50)
# +
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.figure(figsize=(15,7.5))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure(figsize=(15,7.5))
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# +
def smooth_curve(points, factor=0.8):
smoothed_points = []
for point in points:
if smoothed_points:
previous = smoothed_points[-1]
smoothed_points.append(previous * factor + point * (1 - factor))
else:
smoothed_points.append(point)
return smoothed_points
plt.figure(figsize=(15,7.5))
plt.plot(epochs, smooth_curve(acc), 'bo', label='Smoothed training acc')
plt.plot(epochs, smooth_curve(val_acc), 'b', label='Smoothed validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure(figsize=(15,7.5))
plt.plot(epochs, smooth_curve(loss), 'bo', label='Smoothed training loss')
plt.plot(epochs, smooth_curve(val_loss), 'b', label='Smoothed validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
test_generator = test_datagen.flow_from_directory(test_dir, target_size=(150, 150), batch_size=20, class_mode='binary')
test_loss, test_acc = model.evaluate_generator(test_generator, steps=50)
print('test acc:', test_acc)
| notebooks/dl-chollet/scripts/Dogs and Cats Classification with Pre-trained Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
def return_data(N, f, noise):
''' data generating function '''
X = np.random.uniform(0,6, N)
Y = f(X) + noise*np.random.rand(N)
return X, Y
def plot(X,Y):
plt.xlabel(r"$x$", fontsize=20); plt.ylabel(r"$y$", fontsize=20)
plt.xlim(0,6); plt.ylim(-1.5, 2.5)
plt.scatter(X, Y, alpha=0.5)
# -
# ## 1-2. 多項式フィッティング
# 早速ですが機械学習してみます。以下では、あらかじめ定義しておいたデータ生成関数をもちいて実際に10点ほどデータを取ってみたものです:
#
X, Y = return_data(10, np.sin, 1)
plot(X, Y); plt.show()
# これを多項式関数
#
# $$
# f_{M}(x) =
# \sum_{i=0}^M a_i x^i
# $$
#
# の $a_i$ をうまく調節してこのデータにフィッティングすることを考えます。 $M=\color{blue}1, \color{green}3, \color{red}{50}$ でやってみると,
f1 = np.poly1d(np.polyfit(X, Y, deg=1))
f3 = np.poly1d(np.polyfit(X, Y, deg=3))
f50 = np.poly1d(np.polyfit(X, Y, deg=50))
# どれくらいうまくフィッティングできているか見てみます:
plot(X, Y)
X_ = np.linspace(0, 6, 100)
for f in [f1, f3, f50]:
plt.plot(X_, f(X_), alpha=0.5, label=r"$f_{%d}$"%f.order)
plt.legend(fontsize=20); plt.show()
# のようになりました。$f_1$ は線形なので曲がった曲線が表現できず駄目に見えます。$f_3$ は結構いいように見えますが、より次数を上げた $f_{50}$ のほうが更に良く当てはめられているように思えます。
#
# ### 更にデータを加えてみる
# 一番データを上手くフィットしているモデル $f_{50}$ の性能を確かめるために、別のデータを取ってみます。
X_new, Y_new = return_data(30, np.sin, 1)
plot(X_new, Y_new)
X_ = np.linspace(0, 6, 100)
for f in [f1, f3, f50]:
plt.plot(X_, f(X_), alpha=0.5, label=r"$f_{%d}$"%f.order)
plt.legend(fontsize=20); plt.show()
# こうしてみると、むしろ $f_3$ のほうが良いことがわかります。
# $f_{50}$ はパラメータ数が多い分、与えられたデータへの当てはめは得意ですが、あまりにも過剰にパラメータがあると与えられたデータにのみ固執してしまい、未知のデータには使えなくなってしまうというわけです。これを **過剰適合** と呼びます。
#
# 一方で $f_1$ のようにパラメータ数が少なすぎて与えられたデータへの当てはめすらも上手くいかない状況を **過小適合** といいます。
#
# この例からわかるのは、過剰適合でも過小適合でもない、「いい塩梅のフィッティング」をしないと、与えられたデータの当てはめから知らないデータの振る舞いを上手く予想することはできないということです。この節ではここでの教訓が、機械学習のほとんどあらゆる場面で有効だということを理論的に説明することを目的とします。
| section1/1-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SBTi-Finance Tool - Quick Temperature Score Calculation
# This notebook provides a simple example of the SBTi-Finance Tool. It shows how to use it to calculate the temperature score for companies, aggregate them to a portfolio level to get the portfolio temperature score. It also shows you how to calculate the portfolio coverage.
#
# Please see the [methodology](https://sciencebasedtargets.org/wp-content/uploads/2020/09/Temperature-Rating-Methodology-V1.pdf), [guidance](https://sciencebasedtargets.org/wp-content/uploads/2020/10/Financial-Sector-Science-Based-Targets-Guidance-Pilot-Version.pdf) and the [technical documentation](https://sciencebasedtargets.github.io/SBTi-finance-tool/) for more details.
#
# See 1_analysis_example (on [Colab](https://colab.research.google.com/github/OFBDABV/SBTi/blob/master/examples/1_analysis_example.ipynb) or [Github](https://github.com/ScienceBasedTargets/SBTi-finance-tool/blob/master/examples/1_analysis_example.ipynb)) for more in depth example of how to work with Jupyter Notebooks in general and SBTi notebooks in particular.
# ### Install the SBTi Python module
# This is only required if you have not already installed the module.
# + pycharm={"is_executing": false}
# !pip install SBTi
# + pycharm={"is_executing": false}
import SBTi
from SBTi.data.excel import ExcelProvider
from SBTi.portfolio_aggregation import PortfolioAggregationMethod
from SBTi.portfolio_coverage_tvp import PortfolioCoverageTVP
from SBTi.temperature_score import TemperatureScore, Scenario
from SBTi.target_validation import TargetProtocol
from SBTi.interfaces import ETimeFrames, EScope
import pandas as pd
# -
# ## Download the dummy data provider
# We have prepared dummy data for you to be able to run the tool as it is to familiarise yourself with how it works. To use your own data; please check out to the [Data Requirements section](https://ofbdabv.github.io/SBTi/DataRequirements.html) of the technical documentation for more details on data requirements and formatting.
#
# *The dummy data may include some company names, but the data associated with those company names is completely random and any similarities with real world data is purely coincidental.
#
# + pycharm={"is_executing": false}
import urllib.request
import os
if not os.path.isdir("data"):
os.mkdir("data")
if not os.path.isfile("data/data_provider_example.xlsx"):
urllib.request.urlretrieve("https://github.com/ScienceBasedTargets/SBTi-finance-tool/raw/master/examples/data/data_provider_example.xlsx", "data/data_provider_example.xlsx")
if not os.path.isfile("data/example_portfolio.csv"):
urllib.request.urlretrieve("https://github.com/ScienceBasedTargets/SBTi-finance-tool/raw/master/examples/data/example_portfolio.csv", "data/example_portfolio.csv")
# -
# ##### Logging
# The SBTi module uses the Python standard library logging utilities to send log messages. The log level can be changed according to the user's needs.
# + pycharm={"is_executing": false}
import logging
root_logger = logging.getLogger()
root_logger.setLevel("INFO")
# -
# ## Create a data provider
# Data providers let you connect to the data source of your choice. In this case we are connecting to Excel as a data provider. For all available dataproviders check the implementation [here](https://github.com/ScienceBasedTargets/SBTi-finance-tool/tree/master/SBTi/data)
# + pycharm={"is_executing": false}
provider = ExcelProvider(path="data/data_provider_example.xlsx")
# -
# ## Load your portfolio
# In our case the portfolio is stored as a CSV file. The portfolio should at least have an "id" (the identifier of the company) and a "proportion" (the weight of the company in your portfolio e.g. the value of the shares you hold) column.
#
# Please see the technical documentation on [Data Legends](https://ofbdabv.github.io/SBTi/Legends.html#) for details on data requirements.
# + pycharm={"is_executing": false}
df_portfolio = pd.read_csv("data/example_portfolio.csv", encoding="iso-8859-1")
# + pycharm={"is_executing": false}
df_portfolio.head(5)
# -
# To load the data from the data provider, we have to pass a list of IPortfolioCompany instances. The module has a strict [data model](https://ofbdabv.github.io/SBTi/autoapi/SBTi/interfaces/index.html) to convert Pandas Dataframe to the right object types we supplied a utility function.
#
# + pycharm={"is_executing": false}
companies = SBTi.utils.dataframe_to_portfolio(df_portfolio)
# -
# ## Calculate the temperature scores
# In the amended portfolio you'll find your original portfolio, amended with the emissions and the temperature score.
# + pycharm={"is_executing": false}
temperature_score = TemperatureScore( # all available options:
time_frames=list(SBTi.interfaces.ETimeFrames), # ETimeFrames: SHORT MID and LONG
scopes=[EScope.S1S2, EScope.S3, EScope.S1S2S3], # EScopes: S3, S1S2 and S1S2S3
aggregation_method=PortfolioAggregationMethod.WATS # Options for the aggregation method are WATS, TETS, AOTS, MOTS, EOTS, ECOTS, and ROTS.
)
amended_portfolio = temperature_score.calculate(data_providers=[provider], portfolio=companies)
# -
# For every company the tool assigns a score for all the requested timeframe and scope combinations. In this example we used the full set resulting in 9 scores per company as displayed below:
# + pycharm={"is_executing": false}
amended_portfolio[['company_name', 'time_frame', 'scope', 'temperature_score']].head(9)
# -
# ## Calculate the aggregated temperature score
# Calculate an aggregated temperature score. This can be done using different aggregation methods. Here we'll use the "Weighted Average Temperature Score" (WATS) by initializing the TemperatureScore Object with PortfolioAggregationMethod.WATS. For more details, please refer to notebook 4 (on [Colab](https://colab.research.google.com/github/OFBDABV/SBTi/blob/master/examples/4_portfolio_aggregations.ipynb) or [GitHub](https://github.com/ScienceBasedTargets/SBTi-finance-tool/blob/master/examples/4_portfolio_aggregations.ipynb)) and the [methodology document](https://sciencebasedtargets.org/wp-content/uploads/2020/09/Temperature-Rating-Methodology-V1.pdf) sections 3.2. The temperature scores are calculated per time-frame/scope combination.
#
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
# Here we cast a ScoreAggregation object to a Pandas Dataframe for viewing the temp scores in a human readable way
pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
# ## Portfolio coverage
#
# The portfolio coverage provides insights in the proportion of the portfolio that has set SBTi-approved GHG emissions reduction targets. Only companies with SBTi-status "Approved" are included in the portfolio coverage.
#
# To calculate the portfolio coverage we use the same aggregation methods we use for the Portfolio Temperature Score. In this example we use the "Weighted Average Temperature Score" (WATS). For more details on aggregation methods and the portfolio coverage method, please refer to the [methodology document](https://sciencebasedtargets.org/wp-content/uploads/2020/09/Temperature-Rating-Methodology-V1.pdf) sections 3.2 and also turn to notebook 4 (on [Colab](https://colab.research.google.com/github/OFBDABV/SBTi/blob/master/examples/4_portfolio_aggregations.ipynb) or [GitHub](https://github.com/ScienceBasedTargets/SBTi-finance-tool/blob/master/examples/4_portfolio_aggregations.ipynb)) for more aggregation examples.
# + pycharm={"is_executing": false}
portfolio_coverage_tvp = PortfolioCoverageTVP()
coverage = portfolio_coverage_tvp.get_portfolio_coverage(amended_portfolio.copy(), PortfolioAggregationMethod.WATS)
print("Portfolio coverage is: {c:.2f}%".format(c=coverage))
| examples/2_quick_temp_score_calculation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Libraries
import re
# # Load article
#
# Download the text from [**here**](https://drive.google.com/file/d/1PEUMaDaPye5pxlA-SZTsnZ4k7WSzdLLe/view?usp=sharing) and load it into python using the following code snippet.
# Load lines into list
filename = 'data/Article.txt'
with open(filename, encoding='utf-8') as f:
lines = f.readlines()
# # Tasks
# ### Print the first 20 lines of article
print(lines[0:20])
for line in lines[0:20]:
print(line)
# ## Print out every line from the file that...
# ... that has 'q'
# +
regex = re.compile('.*', re.M)
for line in lines:
regex.split(line)
matches = re.findall(r'.*q.*', line)
for match in matches:
print(match)
# -
# ... that starts with 'H'
# +
regex = re.compile('.*', re.M)
for line in lines:
regex.split(line)
matches = re.findall(r'^H.*', line)
for match in matches:
print(match)
# -
# ... that has 'wh'
for line in lines:
regex.split(line)
matches = re.findall(r'.*wh.*', line)
for match in matches:
print(match)
# ... that has an 'q' or a 'Q'
for line in lines:
regex.split(line)
matches = re.findall(r'.*q.*', line, re.I)
for match in matches:
print(match)
# ... that has a '*' in it
for line in lines:
regex.split(line)
matches = re.findall(r'.*\*.*', line)
for match in matches:
print(match)
# ... that starts with an 'T' or an 't'
for line in lines:
regex.split(line)
matches = re.findall(r'^t.*', line, re.I)
for match in matches:
print(match)
# ... that starts with number
for line in lines:
regex.split(line)
matches = re.findall(r'^\d.*', line)
for match in matches:
print(match)
# ... that has both 'a' and 'e' and 'i' and 'o' and 'u' in it
for line in lines:
regex.split(line)
matches = re.findall(r'.*[a].*[e].*[i].*[o].*[u].*', line, re.I)
for match in matches:
print(match)
# ... that has an 'a' and somewhere later an 'e'
for line in lines:
regex.split(line)
matches = re.findall(r'.*a.*e.*', line)
for match in matches:
print(match)
# ... that does not have an 'i'
for line in lines:
regex.split(line)
matches = re.findall(r'^[^i]*$', line)
for match in matches:
if len(match)>1:
print(match)
# ... that does not have an 'i' nor 'z'
for line in lines:
matches = re.findall(r'^[^i|z]*$', line)
for match in matches:
if len(match)>1:
print(match)
# ... that has an 'x' but not 'y'
for line in lines:
regex.split(line)
matches = re.findall(r'^[^y]*$', line)
for match in matches:
if len(match)>1:
print(match)
# ... that has at least 2 consecutive vowels (a, e, i, o, u) like in the word "bear"
for line in lines:
regex.split(line)
matches = re.findall(r'.*[aeiou]{2}.*', line)
for match in matches:
print(match)
# ... that has at least 3 vowels
for line in lines:
regex.split(line)
matches = re.findall(r'.*[aeiou]{3}.*', line)
for match in matches:
print(match)
# ... that has at least 30 characters
for line in lines:
matches = re.findall(r".*", line)
for match in matches:
if len(match)>30:
print(match)
# ... has the same word appear twice in the same line
for line in lines:
matches = re.findall(r".*(?=(\b\w+\b).*(\1)).*", line)
for match in matches:
print(match)
# ## Print all the words
# Words with either 'Bar' or 'Baz' in them
for line in lines:
matches = re.findall(r'\b(?=\w*Bar|Baz)\w+\b', line)
for match in matches:
if len(match)>0:
print(match)
# Words with either 'Whe' or 'The' in them
for line in lines:
matches = re.findall(r'\b(?=\w*Whe|The)\w+\b', line)
for match in matches:
if len(match)>0:
print(match)
# Words containing a double character (e.g. 'oo')
for line in lines:
matches = re.findall(r'\b\w+(?=(\w)(\1))\w+\b', line)
for match in matches:
if len(match)>0:
print(match)
# ## Cleanup string codes so they contain only numbers
# * Remove slash and spaces
# +
codes = ['2373/ 8293',
' 8292342 / 8263',
'12/903820 ',
'8203184 / 02342 ']
result = []
for code in codes:
result.append(re.sub(r'\W', "", code))
result
# -
# ## Switch order of the numbers in string
# * Preserve slash /
# * Remove spaces
# +
codes = ['2373/ 8293',
' 8292342 / 8263',
'12/903820 ',
'8203184 / 02342 ']
result = []
for code in codes:
result.append(re.sub(r'\s', "", code))
result
| Regex/Regex Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py38)
# language: python
# name: py38
# ---
# +
import numpy as np
from matplotlib import pyplot as plt
import matplotlib as mpl
import datetime as dt
import pandas as pd
import f90nml
import matplotlib.dates as mdates
import netCDF4 as nc
mpl.rc('xtick', labelsize=8)
mpl.rc('ytick', labelsize=8)
mpl.rc('legend', fontsize=8)
mpl.rc('axes', titlesize=8)
mpl.rc('axes', labelsize=8)
mpl.rc('figure', titlesize=8)
mpl.rc('font', size=8)
mpl.rc('text', usetex=True)
mpl.rc('text.latex', preamble = r'''
\usepackage{txfonts}
\usepackage{lmodern}
''')
mpl.rc('font', family='sans-serif', weight='normal', style='normal')
# %matplotlib inline
# -
# ## Mesozooplankton
# REAL(wp), dimension (1:3) :: zz_rate_mesozoo_sumpeakval !uM N magnitude of mesozooplankton summer concentration peaks
# REAL(wp), dimension (1:3) :: zz_rate_mesozoo_sumpeakpos ! year-day times of mesozooplankton summer concentration peaks
# REAL(wp), dimension (1:3) :: zz_rate_mesozoo_sumpeakwid ! year-days widths of mesozooplankton summer concentration peaks
nml=f90nml.read('/data/eolson/results/MEOPAR/biomodelevalpaper/modParams/namelist_smelt_ref')
nmlold=f90nml.read('/data/eolson/results/MEOPAR/biomodelevalpaper/modParams/namelist_smelt_ref_old') # for now just load the same one
# +
#&nampismezo ! parameters for microzooplankton
winterconc = nml['nampismezo']['zz_rate_mesozoo_winterconc'] #uM N mesozooplankton background concentration
summerconc = 1.0 # uM N mesozooplankton relative summer concentration
sumpeakval = np.expand_dims(nml['nampismezo']['zz_rate_mesozoo_sumpeakval'],0) #uM N magnitude of mesozooplankton summer concentration peaks
sumpeakpos = np.expand_dims(nml['nampismezo']['zz_rate_mesozoo_sumpeakpos'],0) # year-day times of mesozooplankton summer concentration peaks
sumpeakwid = np.expand_dims(nml['nampismezo']['zz_rate_mesozoo_sumpeakwid'],0) # year-days widths of mesozooplankton summer concentration peaks,0)
#&nampismezo ! parameters for microzooplankton
winterconc_old = nmlold['nampismezo']['zz_rate_mesozoo_winterconc'] #uM N mesozooplankton background concentration
summerconc_old = 1.0 # uM N mesozooplankton relative summer concentration
sumpeakval_old = np.expand_dims(nmlold['nampismezo']['zz_rate_mesozoo_sumpeakval'],0) #uM N magnitude of mesozooplankton summer concentration peaks
sumpeakpos_old = np.expand_dims(nmlold['nampismezo']['zz_rate_mesozoo_sumpeakpos'],0) # year-day times of mesozooplankton summer concentration peaks
sumpeakwid_old = np.expand_dims(nmlold['nampismezo']['zz_rate_mesozoo_sumpeakwid'],0) # year-days widths of mesozooplankton summer concentration peaks,0)
# -
maxRate=nml['nampismezo']['zz_rate_mesozoo_r']
# check that grazing rate has been unchanged since original run:
if not maxRate==1.39e-5:
print('PROBLEM: GRAPH BELOW WILL BE INCORRECT BECAUSE GRAZING RATES ARE NOT CONSISTENT')
else:
print('rates ok')
# +
zz_day=np.arange(1,367) # in model, nday_year starts at 1 on jan 1
dts=[dt.datetime(2014,12,31)+dt.timedelta(days=float(ii)) for ii in zz_day]
zz_day=np.expand_dims(zz_day,1)
MesZoBar = winterconc + \
summerconc*(np.sum ( sumpeakval * np.exp(-(zz_day-sumpeakpos)**2/sumpeakwid**2),1) \
+ np.sum ( sumpeakval * np.exp( -(zz_day-sumpeakpos-365.25)**2/sumpeakwid**2),1) \
+ np.sum ( sumpeakval * np.exp( -(zz_day-sumpeakpos+365.25)**2/sumpeakwid**2),1) )
MesZoBar_old = winterconc_old + \
summerconc_old*(np.sum ( sumpeakval_old * np.exp(-(zz_day-sumpeakpos_old)**2/sumpeakwid_old**2),1) \
+ np.sum ( sumpeakval_old * np.exp( -(zz_day-sumpeakpos_old-365.25)**2/sumpeakwid_old**2),1) \
+ np.sum ( sumpeakval_old * np.exp( -(zz_day-sumpeakpos_old+365.25)**2/sumpeakwid_old**2),1) )
# -
fig,ax=plt.subplots(1,1,figsize=(4,3))
ax.plot(dts,MesZoBar_old*maxRate*24*3600,'--',color='grey')
ax.plot(dts,MesZoBar*maxRate*24*3600,'-',color='k')
ax.set_ylabel('$G_{max}^{MESZ}$ ($\muup$M N d$^{-1}$)')
#ax.set_xlabel('Date')
fig.autofmt_xdate(bottom=0.3, rotation=30, ha='right')
yearsFmt = mdates.DateFormatter('%b %d')
ax.xaxis.set_major_formatter(yearsFmt)
fig.savefig('/data/eolson/results/MEOPAR/biomodelevalpaper/figsMod/mesozoo.eps',dpi=400,transparent=True)
# +
##version with data
df=pd.read_excel('/ocean/eolson/MEOPAR/obs/MackasZoop/1-s2.0-S007966111300061X-mmc2.xls',skiprows=1)
df=df.dropna(0,how='any', subset=['Year','Month','Day']).dropna(1,how='any',thresh=200)
yd=[(dt.datetime(int(r.Year),int(r.Month),int(r.Day))-dt.datetime(int(r.Year)-1,12,31)).days for i,r in df.iterrows()]
df=df.assign(yd=yd).sort_values(by='yd')
dtsdf=[dt.datetime(2014,12,31)+dt.timedelta(days=int(ii)) for ii in df['yd'].values]
df['dts']=dtsdf
fig,ax=plt.subplots(1,1,figsize=(3,2))
p2,=ax.plot(dts,MesZoBar_old*maxRate*24*3600,'--',color='k',label='this study')
p1,=ax.plot(dts,MesZoBar*maxRate*24*3600,'-',color='k',label='Moore-Maley et al. (2016)')
ax.set_ylabel('$G_{max}^{MESZ}$ ($\muup$M N d$^{-1}$)')
fig.autofmt_xdate(bottom=0.3, rotation=30, ha='right')
yearsFmt = mdates.DateFormatter('%b %d')
ax.set_ylim(0,10)
ax.xaxis.set_major_formatter(yearsFmt)
ax2=ax.twinx()
p3,=ax2.plot(df.loc[:,['dts']],df.loc[:,['Total Biomass']],'.',color='silver', MarkerEdgeWidth=0,MarkerSize=4,label='Mackas et al. (2013)')
ax2.set_ylabel('Zooplankton Biomass (g m$^{-2}$)')
ax2.set_ylim()
ax.set_position(pos=(0.15,.16,.7,.8))
ax2.set_position(pos=(0.15,.16,.7,.8))
#fig.legend((p1,p2,p3),('this study','Moore-Maley et al. (2016)','Mackas et al. (2013)'),loc=2)
#fig.savefig('/data/eolson/results/MEOPAR/biomodelevalpaper/figsMod/mesozooWithMackas.eps',dpi=400,transparent=True)
# +
##version with data
df=pd.read_excel('/ocean/eolson/MEOPAR/obs/MackasZoop/1-s2.0-S007966111300061X-mmc2.xls',skiprows=1)
df=df.dropna(0,how='any', subset=['Year','Month','Day']).dropna(1,how='any',thresh=200)
yd=[(dt.datetime(int(r.Year),int(r.Month),int(r.Day))-dt.datetime(int(r.Year)-1,12,31)).days for i,r in df.iterrows()]
df=df.assign(yd=yd).sort_values(by='yd')
dtsdf=[dt.datetime(2014,12,31)+dt.timedelta(days=int(ii)) for ii in df['yd'].values]
df['dts']=dtsdf
fig,(ax,ax1,ax2)=plt.subplots(3,1,figsize=(3,2.2),sharex=True)
ax.yaxis.tick_left()
ax1.yaxis.tick_right()
ax1.yaxis.set_label_position('right')
ax2.yaxis.tick_right()
ax2.yaxis.set_label_position('right')
for iax in (ax,ax1,ax2):
iax.patch.set_alpha(0)
iax.set_xlim(dt.datetime(2015,1,1),dt.datetime(2015,12,31))
iax.tick_params(direction='in')
p2,=ax.plot(dts,MesZoBar_old*maxRate*24*3600,'--',color='k',label='this study')
p1,=ax.plot(dts,MesZoBar*maxRate*24*3600,'-',color='k',label='Moore-Maley et al. (2016)')
ax.set_ylabel('$G_{max}^{MESZ}$ ($\muup$M N d$^{-1}$)')
fig.autofmt_xdate(bottom=0.3, rotation=30, ha='right')
yearsFmt = mdates.DateFormatter('%b %d')
ax.set_ylim(0,5)
ax.xaxis.set_major_formatter(yearsFmt)
#ax2=ax.twinx()
p3,=ax1.plot(df.loc[:,['dts']],df.loc[:,['Total Biomass']],'.',color='silver', MarkerEdgeWidth=0,MarkerSize=4,label='Mackas et al. (2013)')
ax2.plot(df.loc[:,['dts']],df.loc[:,['Total Biomass']],'.',color='silver', MarkerEdgeWidth=0,MarkerSize=4,label='Mackas et al. (2013)')
ax2.set_ylabel('Zooplankton Biomass (g m$^{-2}$)')
#ax1.set_ylim()
ax.set_position(pos=(0.15,.15,.7,.8))
ax2.set_position(pos=(0.15,.15,.7,.63))
ax1.set_position(pos=(0.15,.8,.7,.15))
ax1.spines['bottom'].set_visible(False)
ax1.spines['left'].set_visible(False)
ax2.spines['top'].set_visible(False)
ax1.xaxis.tick_top()
ax1.tick_params(labeltop='off') # don't put tick labels at the top
ax2.xaxis.tick_bottom()
ax.xaxis.tick_bottom()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax1.set_ylim(90,160)
ax1.set_yticks((100,130,160))
ax2.set_ylim(0,83)
d = .015 # how big to make the diagonal lines in axes coordinates
# arguments to pass to plot, just so we don't keep repeating them
kwargs = dict(transform=ax1.transAxes, color='k', clip_on=False)
#ax1.plot((-d, +d), (-d, +d), **kwargs) # top-left diagonal
a=.6/.15
ax1.plot((1 - d, 1 + d), (-d*a, +d*a), **kwargs) # top-right diagonal
kwargs.update(transform=ax2.transAxes) # switch to the bottom axes
#ax2.plot((-d, +d), (1 - d, 1 + d), **kwargs) # bottom-left diagonal
ax2.plot((1 - d, 1 + d), (1 - d, 1 + d), **kwargs) # bottom-right diagonal
ax2.yaxis.set_label_coords(1.14,.63)
ax1.xaxis.set_ticklabels([]);
#fig.legend((p1,p2,p3),('this study','Moore-Maley et al. (2016)','Mackas et al. (2013)'),loc=2)
fig.savefig('/data/eolson/results/MEOPAR/biomodelevalpaper/figsMod/mesozooWithMackas.eps',dpi=400,transparent=True)
# +
## check axes equivalence
df=pd.read_excel('/ocean/eolson/MEOPAR/obs/MackasZoop/1-s2.0-S007966111300061X-mmc2.xls',skiprows=1)
df=df.dropna(0,how='any', subset=['Year','Month','Day']).dropna(1,how='any',thresh=200)
yd=[(dt.datetime(int(r.Year),int(r.Month),int(r.Day))-dt.datetime(int(r.Year)-1,12,31)).days for i,r in df.iterrows()]
df=df.assign(yd=yd).sort_values(by='yd')
dtsdf=[dt.datetime(2014,12,31)+dt.timedelta(days=int(ii)) for ii in df['yd'].values]
df['dts']=dtsdf
fig,(ax,ax1,ax2)=plt.subplots(3,1,figsize=(3,2.2),sharex=True)
ax.yaxis.tick_left()
ax1.yaxis.tick_right()
ax1.yaxis.set_label_position('right')
ax2.yaxis.tick_right()
ax2.yaxis.set_label_position('right')
for iax in (ax,ax1,ax2):
iax.patch.set_alpha(0)
iax.set_xlim(dt.datetime(2015,1,1),dt.datetime(2015,12,31))
iax.tick_params(direction='in')
p2,=ax.plot(dts,MesZoBar_old*maxRate*24*3600,'--',color='k',label='this study')
p1,=ax.plot(dts,MesZoBar*maxRate*24*3600,'-',color='k',label='Moore-Maley et al. (2016)')
ax.set_ylabel('$G_{max}^{MESZ}$ ($\muup$M N d$^{-1}$)')
fig.autofmt_xdate(bottom=0.3, rotation=30, ha='right')
yearsFmt = mdates.DateFormatter('%b %d')
ax.set_ylim(0,5)
ax.xaxis.set_major_formatter(yearsFmt)
#ax2=ax.twinx()
p3,=ax1.plot(df.loc[:,['dts']],df.loc[:,['Total Biomass']],'.',color='silver', MarkerEdgeWidth=0,MarkerSize=4,label='Mackas et al. (2013)')
ax2.plot(df.loc[:,['dts']],df.loc[:,['Total Biomass']],'.',color='silver', MarkerEdgeWidth=0,MarkerSize=4,label='Mackas et al. (2013)')
ax2.set_ylabel('Zooplankton Biomass (g m$^{-2}$)')
#ax1.set_ylim()
ax.set_position(pos=(0.15,.15,.7,.8))
ax2.set_position(pos=(0.15,.15,.7,.63))
ax1.set_position(pos=(0.15,.8,.7,.15))
ax1.spines['bottom'].set_visible(False)
ax1.spines['left'].set_visible(False)
ax2.spines['top'].set_visible(False)
ax1.xaxis.tick_top()
ax1.tick_params(labeltop='off') # don't put tick labels at the top
ax2.xaxis.tick_bottom()
ax.xaxis.tick_bottom()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax1.set_ylim(90,160)
ax1.set_yticks((100,130,160))
ax2.set_ylim(0,83)
d = .015 # how big to make the diagonal lines in axes coordinates
# arguments to pass to plot, just so we don't keep repeating them
kwargs = dict(transform=ax1.transAxes, color='k', clip_on=False)
#ax1.plot((-d, +d), (-d, +d), **kwargs) # top-left diagonal
a=.6/.15
ax1.plot((1 - d, 1 + d), (-d*a, +d*a), **kwargs) # top-right diagonal
kwargs.update(transform=ax2.transAxes) # switch to the bottom axes
#ax2.plot((-d, +d), (1 - d, 1 + d), **kwargs) # bottom-left diagonal
ax2.plot((1 - d, 1 + d), (1 - d, 1 + d), **kwargs) # bottom-right diagonal
ax2.yaxis.set_label_coords(1.14,.63)
ax1.xaxis.set_ticklabels([]);
ax.plot(dt.datetime(2015,12,20),3.79,'r.')
#fig.legend((p1,p2,p3),('this study','Moore-Maley et al. (2016)','Mackas et al. (2013)'),loc=2)
#fig.savefig('/data/eolson/results/MEOPAR/biomodelevalpaper/figsMod/mesozooWithMackas.eps',dpi=400,transparent=True)
# -
with nc.Dataset('/ocean/eolson/MEOPAR/NEMO-forcing/grid/mesh_mask201702_noLPE.nc') as fm:
tmask=np.copy(fm.variables['tmask'][0,0,:,:])
navlon=np.copy(fm.variables['nav_lon'][:,:])
navlat=np.copy(fm.variables['nav_lat'][:,:])
fig,ax=plt.subplots(1,1,figsize=(5,5))
ax.pcolormesh(navlon,navlat,tmask,cmap=plt.get_cmap('Greys_r'))
plt.plot(-1*df['Longitude (deg W)'],df['Latitude (deg N)'],'b.')
ax.set_xlim(-125.3,-123)
ax.set_ylim(48.7,50.1)
| notebooks/methods/mesozooplankton-paperFig.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pandas]
# language: python
# name: conda-env-pandas-py
# ---
# ## Understanding NumPy Array
# ### Getting help
help(np.array)
# +
# np.arange?
# -
# Creating an array
import numpy as np
a = np.array([2,4,6,8,10])
print(a)
# Creating an array using arange()
import numpy as np
a = np.arange(1,11)
print(a)
# +
import numpy as np
p = np.zeros((3,3)) # Create an array of all zeros
print(p)
q = np.ones((2,2)) # Create an array of all ones
print(q)
r = np.full((2,2), 4) # Create a constant array
print(r)
s = np.eye(4) # Create a 2x2 identity matrix
print(s)
t = np.random.random((3,3)) # Create an array filled with random values
print(t)
# -
# Creating an array using arange()
import numpy as np
a = np.arange(1,11)
print(type(a))
print(a.dtype)
# check shape pf Array
print(a.shape)
a = np.array([[5,6],[7,8]])
print(a)
print(a[0,0])
print(a[0,1])
print(a[1,0])
print(a[1,1])
# ## NumPy Array Numerical Data Types
print(np.float64(21))
print(np.int8(21.0))
print(np.bool(21))
print(np.bool(0))
print(np.bool(21.0))
print(np.float(True))
print(np.float(False))
# +
arr=np.arange(1,11, dtype= np.float32)
print(arr)
# -
np.int(42.0 + 1.j)
c= complex(42, 1)
print(c)
print(c.real,c.imag)
# +
# Creating an array
import numpy as np
a = np.array([2,4,6,8,10])
print(a.dtype)
# -
print(a.dtype.itemsize)
# +
# Create numpy array using arange() function
var1=np.arange(1,11, dtype='f')
print(var1)
# -
print(np.arange(1,6, dtype='D'))
print(np.dtype(float))
print(np.dtype('f'))
print(np.dtype('d'))
print(np.dtype('f8'))
# +
var2=np.array([1,2,3],dtype='float64')
print(var2.dtype.char)
# -
print(var2.dtype.type)
# ## Manipulating Shape of NumPy Array
# Create an array
arr = np.arange(12)
# +
# Reshape the array dimension
new_arr=arr.reshape(4,3)
print(new_arr)
# +
# Reshape the array dimension
new_arr2=arr.reshape(3,4)
print(new_arr2)
# -
# Create an array
arr=np.arange(1,10).reshape(3,3)
print(arr)
# flatten the array
print(arr.flatten())
# ravel() function
print(arr.ravel())
# Transpose the matrix
print(arr.transpose())
# resize the matrix
arr.resize(1,9)
print(arr)
# ## Stacking of Numpy arrays
arr1 = np.arange(1,10).reshape(3,3)
print(arr1)
arr2 = 2*arr1
print(arr2)
# +
arr3=np.hstack((arr1, arr2))
print(arr3)
# -
# Horizontal stacking using concatenate() function
arr4=np.concatenate((arr1, arr2), axis=1)
print(arr4)
arr5=np.vstack((arr1, arr2))
print(arr5)
arr6=np.concatenate((arr1, arr2), axis=0)
print(arr6)
arr7=np.dstack((arr1, arr2))
print(arr7)
# Create 1-D array
arr1 = np.arange(4,7)
print(arr1)
# Create 1-D array
arr2 = 2 * arr1
print(arr2)
# Create column stack
arr_col_stack = np.column_stack((arr1,arr2))
print(arr_col_stack)
# Create row stack
arr_row_stack = np.row_stack((arr1,arr2))
print(arr_row_stack)
# ## Partitioning Numpy Array
# Create an array
arr=np.arange(1,10).reshape(3,3)
print(arr)
# +
# Peroform horizontal splitting
arr_hor_split=np.hsplit(arr, 3)
print(arr_hor_split)
# +
# vertical split
arr_ver_split=np.vsplit(arr, 3)
print(arr_ver_split)
# +
# split with axis=0
arr_split=np.split(arr,3,axis=0)
print(arr_split)
# -
# split with axis=1
np.split(arr,3,axis=1)
# ## Changing Datatype of NumPy Arrays
# +
# Create an array
arr=np.arange(1,10).reshape(3,3)
print("Integer Array:",arr)
# Change datatype of array
arr=arr.astype(float)
# print array
print("Float Array:", arr)
# Check new data type of array
print("Changed Datatype:", arr.dtype)
# +
# Change datatype of array
arr=arr.astype(float)
# Check new data type of array
print(arr.dtype)
# +
# Create an array
arr=np.arange(1,10)
# Convert NumPy array to Python List
list1=arr.tolist()
print(list1)
# -
# ## Creating NumPy views and copies
# +
# Create NumPy Array
arr = np.arange(1,5).reshape(2,2)
print(arr)
# Create no copy only assignment
arr_no_copy=arr
# Create Deep Copy
arr_copy=arr.copy()
# Create shallow copy using View
arr_view=arr.view()
print("Original Array: ",id(arr))
print("Assignment: ",id(arr_no_copy))
print("Deep Copy: ",id(arr_copy))
print("Shallow Copy(View): ",id(arr_view))
# +
# Update the values of original array
arr[1]=[99,89]
# Check values of array view
print("View Array:\n", arr_view)
# Check values of array copy
print("Copied Array:\n", arr_copy)
# -
# ## Slicing NumPy Array
# Create NumPy Array
arr = np.arange(10)
print(arr)
print(arr[3:6])
print(arr[3:])
print(arr[-3:])
print(arr[2:7:2])
# ## Boolean and Fancy Indexing
# +
# Create NumPy Array
arr = np.arange(21,41,2)
print("Orignial Array:\n",arr)
# Boolean Indexing
print("After Boolean Condition:",arr[arr>30])
# +
# Create NumPy Array
arr = np.arange(1,21).reshape(5,4)
print("Orignial Array:\n",arr)
# Selecting 2nd and 3rd row
indices = [1,2]
print("Selected 1st and 2nd Row:\n", arr[indices])
# Selecting 3nd and 4th row
indices = [2,3]
print("Selected 3rd and 4th Row:\n", arr[indices])
# +
# Create row and column indices
row = np.array([1, 2])
col = np.array([2, 3])
print("Selected Sub-Array:", arr[row, col])
# -
# ## Broadcasting arrays
# Create NumPy Array
arr1 = np.arange(1,5).reshape(2,2)
print(arr1)
# Create another NumPy Array
arr2 = np.arange(5,9).reshape(2,2)
print(arr2)
# Add two matrices
print(arr1+arr2)
# Multiply two matrices
print(arr1*arr2)
# Add a scaler value
print(arr1 + 3)
# Multiply with a scalar value
print(arr1 * 3)
# ## Create DataFrame
# +
# Import pandas library
import pandas as pd
# Create empty DataFrame
df = pd.DataFrame()
# Header of dataframe.
df.head()
# -
df
# +
# Create dictionary of list
data = {'Name': ['Vijay', 'Sundar', 'Satyam', 'Indira'], 'Age': [23, 45, 46, 52 ]}
# Create the pandas DataFrame
df = pd.DataFrame(data)
# Header of dataframe.
df.head()
# -
# Pandas DataFrame by lists of dicts.
# Initialise data to lists.
data =[ {'Name': 'Vijay', 'Age': 23},{'Name': 'Sundar', 'Age': 25},{'Name': 'Shankar', 'Age': 26}]
# Creates DataFrame.
df = pd.DataFrame(data,columns=['Name','Age'])
# Print dataframe header
df.head()
# Creating DataFrame using list of tuples.
data = [('Vijay', 23),( 'Sundar', 45), ('Satyam', 46), ('Indira',52)]
# Create dataframe
df = pd.DataFrame(data, columns=['Name','Age'])
# Print dataframe header
df.head()
# ## Pandas Series
# Creating Pandas Series using Dictionary
dict1 = {0 : 'Ajay', 1 : 'Jay', 2 : 'Vijay'}
# Create Pandas Series
series = pd.Series(dict1)
# Show series
series
# load Pandas and NumPy
import pandas as pd
import numpy as np
# Create NumPy array
arr = np.array([51,65,48,59, 68])
# Create Pandas Series
series = pd.Series(arr)
series
# load Pandas and NumPy
import pandas as pd
import numpy as np
# Create Pandas Series
series = pd.Series(10, index=[0, 1, 2, 3, 4, 5])
series
# +
# Import pandas
import pandas as pd
# Load data using read_csv()
df = pd.read_csv("WHO_first9cols.csv")
# Show initial 5 records
df.head()
# -
# Show last 5 records
df.tail()
# Show the shape of DataFrame
print("Shape:", df.shape)
# Check the column list of DataFrame
print("List of Columns:", df.columns)
# Show the datatypes of columns
print("Data types:", df.dtypes)
# Select a series
country_series=df['Country']
# check datatype of series
type(country_series)
print(country_series.index)
# Convert Pandas Series into List
print(country_series.values)
# Country name
print(country_series.name)
# Pandas Series Slicing
country_series[-5:]
# Creating Pandas Series using Dictionary
dict1 = {0 : 'Ajay', 1 : 'Jay', 2 : 'Vijay'}
# Create Pandas Series
series = pd.Series(dict1)
# Show series
series
# load Pandas and NumPy
import pandas as pd
import numpy as np
# Create NumPy array
arr = np.array([51,65,48,59, 68])
# Create Pandas Series
series = pd.Series(arr)
series
# load Pandas and NumPy
import pandas as pd
import numpy as np
# Create Pandas Series
series = pd.Series(10, index=[0, 1, 2, 3, 4, 5])
series
# ## Querying Data
# !pip install quandl
# +
import quandl
sunspots = quandl.get("SIDC/SUNSPOTS_A")
sunspots.head()
# -
sunspots.head()
sunspots.tail()
sunspots.columns
# +
# Select columns
sunspots_filtered=sunspots[['Yearly Mean Total Sunspot Number','Definitive/Provisional Indicator']]
# Show top 5 records
sunspots_filtered.head()
# -
# Select rows using index
sunspots["20020101": "20131231"]
# +
# Boolean Filter
sunspots[sunspots['Yearly Mean Total Sunspot Number'] > sunspots['Yearly Mean Total Sunspot Number'].mean()]
# -
# ## Statistics
# +
# Import pandas
import pandas as pd
# Load data using read_csv()
df = pd.read_csv("WHO_first9cols.csv")
# Show initial 5 records
df.head()
# -
df.shape
# Describe the dataset
df.describe()
# Count number of observation
df.count()
# Compute median of all the columns
df.median()
# Compute minimum of all the columns
df.min()
# Compute maximum of all the columns
df.max()
# Compute standard deviation of all the columns
df.std()
# ## Grouping Pandas DataFrames
df.head()
# Group By Dataframe on the basis of Continent column
df.groupby('Continent').mean()
df.groupby('Continent').mean()['Adult literacy rate (%)']
# ## Joins
# +
# Import pandas
import pandas as pd
# Load data using read_csv()
dest = pd.read_csv("dest.csv")
# Show DataFrame
dest.head()
# +
# Load data using read_csv()
tips = pd.read_csv("tips.csv")
# Show DataFrame
tips.head()
# -
# Join DataFrames using Inner Join
df_inner= pd.merge(dest, tips, on='EmpNr', how='inner')
df_inner.head()
# Join DataFrames using Outer Join
df_outer= pd.merge(dest, tips, on='EmpNr', how='outer')
df_outer.head()
# Join DataFrames using Right Outer Join
df_right= pd.merge(dest, tips, on='EmpNr', how='right')
df_right
# Join DataFrames using Left Outer Join
df_left= pd.merge(dest, tips, on='EmpNr', how='left')
df_left
# ## Missing Values
# +
# Import pandas
import pandas as pd
# Load data using read_csv()
df = pd.read_csv("WHO_first9cols.csv")
# Show initial 5 records
df.head()
# -
# Count missing values in DataFrame
pd.isnull(df).sum()
# Count missing values in DataFrame
df.isnull().sum()
df.info()
# Drop all the missing values
df.dropna(inplace=True)
df.info()
# +
# Load data using read_csv()
df = pd.read_csv("WHO_first9cols.csv")
# Show initial 5 records
df.head()
# -
df.info()
# Fill missing values with 0
df.fillna(0,inplace=True)
df.info()
# ## Pivot Table
# +
# Import pandas
import pandas as pd
# Load data using read_csv()
purchase = pd.read_csv("purchase.csv")
# Show initial 10 records
purchase.head(10)
# -
# Summarise dataframe using pivot table
pd.pivot_table(purchase,values='Number', index=['Weather',],
columns=['Food'], aggfunc=np.sum)
# ## Dealing with dates
# Date range function
pd.date_range('01-01-2000', periods=45, freq='D')
# Convert argument to datetime
pd.to_datetime('1/1/1970')
# Convert argument to datetime in specified format
pd.to_datetime(['20200101', '20200102'], format='%Y%m%d')
# Value Error
pd.to_datetime(['20200101', 'not a date'])
# Handle value error
pd.to_datetime(['20200101', 'not a date'], errors='coerce')
| Chapter02/ch2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Data Science блог с помощью fastpages"
# > "Как запустить свой DS/ML/AI/Tech блог с минимумом сложностей связанных с хостингом и деплойем этого блога."
#
# - toc: true
# - badges: false
# - comments: true
# - categories: [tutorial]
# - image: images/favicon.png
# - sticky_rank: 1
# В конце февраля 2020 года ребята из `fast.ai` представили миру `fastpages` - платформу для ведения блога. Отмечу, что `fastpages` основан на `Jekyll`, о котором на Хабре есть множество постов.
#
# Примером блога на движке `fastpages` является [данный блог](https://maxbalashov.github.io/ml-notes/).
#
# Главное отличительная черта и преимущество `fastpages` состоит в поддерживаемых из коробки форматах постов:
# - Jupyter ноутбуки (расширение `.ipynb`);
# - Markdown файлы (расширение `.md`);
# - Word файлы (расширение `.docx`)
#
# Таким образом, автору блога необходимо сохранить пост в любом из перечисланных выше форматах в соответствующей директории:
# - './_notebooks/' для `.ipynb`;
# - './_posts/' для `.md`;
# - './_word/' для `.docx`.
#
# А все остальное сделает `fastpages`, как утверждают его авторы.
# `fastpages` использует Github Pages для хостинга и Github Actions для автоматизации публикации постов.
#
# Как я понимаю, `fastpages` является доработкой связки Github Pages + `Jekyll`, где можно сразу же из Jupyter ноутбука получить опубликованный пост.
# # Создание блога с помощью `fastpages` и `GitHub`
# Если хотите самостоятельно разобраться, то вот [официальная инструкция](https://github.com/fastai/fastpages#setup-instructions) по настройке в репозитории `fastpages`.
#
# Процесс настройки `fastpages`:
#
# 1. Создать собственную копию репозитория из шаблона `fastpages` по [ссылке](https://github.com/fastai/fastpages/generate)
# 
# 2. Далее автоматически откроется pull request (через ~ 30 секунд), который отвечает за настройку вашего блога, чтобы он мог начать работать.
# 
# 3. Вам нужно выполнить инструкции из полученного pull request'a и вы получите свою собственную уже работающую платформу для блога.
#
# ## Видео туториал
# > youtube: https://youtu.be/L0boq3zqazI
# # Настройка блога
#
# Есть возможность для персонализированной конфигурации вашего блога. Параметры конфигурации находятся в файле `./_config.yml`, некоторые из них приведены ниже:
# - `title` - название вашего блога, которое отображается в верхнем левом углу на каждой странице;
# - `description` - описание, которое будет отображаться в разных местах при предварительном просмотре вашего сайта (например, в социальных сетях);
# - `github_username` - позволяет вашему сайту отображать ссылку на вашу страницу GitHub в нижнем колонтитуле;
# - `github_repo` - позволяет вашему сайту отображать ссылки на ваш репозиторий для различных функций, таких как ссылки на GitHub, Google Colab и Binder для Jupyter ноутбуков;
# - `default_badges` - по умолчанию ссылки GitHub, Google Colab и Binder будут отображаться в постах созданных из Jupyter ноутбуков. Вы можете задать, какие из них будут отображаться по умолчанию, установив для соответствующего значения в `default_badges` значение `true` или `false`. Например, если вы хотите отключить ссылки на Binder, вы должны поправить `default_badges`:
# ```
# default_badges:
# github: true
# binder: false
# colab: true
# ```
# - `url` - это не нужно менять, если у вас нет собственного домена;
# - `baseurl` - см. комментарии в /_config.yml для получения инструкций ("Special Instructions for baseurl"). Если у вас нет настраиваемого домена, вы можете игнорировать эту опцию;
# - `twitter_username` - создает ссылку в нижнем колонтитуле на страницу Twitter;
# - `use_math` - установите значение `true`, чтобы получить поддержку математических формул `LaTeX`;
# - `show_description` - отображает на домашней странице описание под заголовком ваших постов в блоге. По умолчанию установлено значение `true`;
# - `google_analytics` - опционально можно использовать идентификатор Google Analytics;
# - `pagination` - максимальное количество постов, отображаемых на каждой странице вашей домашней страницы. Значение по умолчанию равно 15. Когда число постов превысит заданное значение, тогда произойдет разбивка на страницы, которая выглядит так:
# 
#
# - `show_tags` - включает отображение тегов внутри постов, которые выглядят следующим образом:
# 
#
# - `show_image` - при значении `true` включается возможность добавления изображений к постам на домашней странице. Выглядит следующим образом (первые 2 поста сопровождаются изображениями):
# 
# # Публикация постов из `.ipynb` с помощью `fastpages`
# 1. Сохраните исходный файл вашего поста (в одном из форматов: `.ipynb`, `.md` или `.docx`) в соответствующей папке репозитория (`./_notebooks`, `./_posts` или `./_word`). Пример имени для поста `2020-05-26-DS-fastpages-blog.ipynb`. Такое наименование является необходимым для отображения поста движком Jekyll ([больше деталей](https://jekyllrb.com/docs/posts/)).
#
# Важные аспекты наименования постов:
# - Вначале имени поста указывается дата в формате `YYYY-MM-DD-`;
# - Символ, следующий сразу за тире, должен быть буквой алфавита.
# 2. Сделайте [commit и push](https://help.github.com/en/github/managing-files-in-a-repository/adding-a-file-to-a-repository-using-the-command-line) ваших файлов на удаленный репозиторий GitHub в ветку `master`.
# 3. GitHub автоматически конвертирует ваши файлы в посты блога. Процесс конвертации займет ~5 минут. Можно перейти на вкладку «Actions» в репозитории на GitHub. Вы увидите три workflow, которые запускаются при каждом `push` в ветку `master`:
# - Check Configurations - процесс проверки ваших файлов (например, ссылок на изображения), перед обновлением контента в блоге;
# - CI - процесс непрерывного деплоя вашего блога;
# - GH Pages Status - процесс проверки доступа к блогу.
#
# Если эти процессы завершаются зеленой галочкой для последнего коммита, то сайт блога успешно обновился.
#
# 4. Для предварительного локального просмотра того, как ваш блог будет выглядеть, [см. этот раздел](https://github.com/fastai/fastpages#running-the-blog-on-your-local-machine).
#
# Ниже представлены различные возможности форматирования, которые `fastpages` поддерживает из коробки.
# ## Возможности форматирования постов
#
# Первая ячейка в вашем Jupyter ноутбуке (а также первые строки в Markdown файлах) содержит метаданные, которые могут включать/выключать опции связанные с постом.
#
# ```
# # "Title"
# > "Awesome summary"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME> & <NAME>
# - categories: [fastpages, jupyter]
# ```
#
# Для указания таких в Markdown файлах необходимо в начале файла задать опции как и в ноутбуке, только поместив эти метаданные между строк содержащих по три минуса, т.е. `---`.
#
# Выглядит это так:
#
# ```
# ---
# title: "Title"
# description: "Awesome description"
# toc: true
# layout: post
# categories: [markdown]
# ---
# ```
#
# > Note: Все, что определено в начале поста, должно соответствовать YAML разметке. Поэтому если вы хотите использовать двоеточие в заголовке, вы должны экранировать его двойными кавычками: `- title: "Deep learning: A tutorial"`
#
# Для большего понимания советую ознакомиться с [туториалом по YAML](https://rollout.io/blog/yaml-tutorial-everything-you-need-get-started/).
# Перечень управляющих конструкций для форматирования поста (взято [отсюда](https://github.com/fastai/fastpages#customizing-blog-posts-with-front-matter)):
# - `toc` - при значении `true` автоматически будет сгенерировано оглавление поста из заголовков, обозначенных Markdown разметкой;
# - `badges` \[notebooks only\] - при значении `true` отображаются ссылки `Google Colab`, `Binder` и `GitHub`, не работает при приватном репозитории;
# - `hide_github_badge` \[notebooks only\] - при значении `true` скроет ссылку на `GitHub`;
# - `hide_colab_badge` \[notebooks only\] - при значении `true` скроет ссылку на `Google Colab`;
# - `hide_binder_badge` \[notebooks only\] - при значении `true` скроет ссылку на `Binder`;
# - `branch` \[notebooks only\] - используется для дополнительной ссылки на ваш Jupyter ноутбук на Colab и GitHub. Значение по умолчанию: `master`;
# - `comments` - при значении `true` будут включены комментарии ([больше деталей](https://github.com/fastai/fastpages#enabling-comments));
# - `author` - при значении `true` отображаются имена авторов;
# - `categories` - позволяют группировать посты по тегам (на странице "Tags").
# - `image` - задает изображение для поста, которое будет отображаться на главной странице блога и в соц. сетях (Twitter) вместе с ссылкой на пост:
# - пример задания изображения к посту - `images/figure.png`;
# - изображение обязательно должно находиться внутри папке `/images` вашего репозитория;
# - `search_exclude` - позволяет скрывать пост в поиске блога (страница `Search`), стоит заменить, поиск работает только с латиницей;
# - `hide` - при значении `true` пост будет скрыт на главной странице блога, но будет доступен по прямой ссылке:
# - рекомендуется использовать [permalinks](https://jekyllrb.com/docs/permalinks/) для создания предсказуемых ссылок на сам пост;
# - если `search_exclude` будет иметь значение `true`, то пост можно будет найти через поиск блога (страница `Search`);
# - `sticky_rank` - позволяет закрепить пост на конкретной позиции, задав ему порядковый номер. Если двум постам задать одинаковый номер, то между собой они будут отсортированы по дате.
# ## Скрытие и сворачивание кода
#
# Приятной функциональностью этого движка для блога является возможность скрывать код и/или результаты его выполнения. Это позволяет не нагружать посты отображением простыни кода или огромного количество принтов (что бывает при обучении нейросетей по эпохам), скрывая эти большие по размеру элементы, но не выкидывая их из поста на совсем.
# Комментарий `#hide` в первой строке любой ячейки кода будет скрывать как ввод, так и вывод этой ячейки.
#
# Ниже есть ячейка, которая не отображается в посте, но присутствует в исходном jupyter ноутбуке. Можете проверить на [GitHub](https://github.com/MaxBalashov/ml-notes/blob/master/_notebooks/2020-05-26-DS-fastpages-blog.ipynb).
# +
#hide
print('Этой ячейки и результата ее запуска вы в блоге не увидите :)')
# -
# Комментарий `#hide_input` в первой строке любой ячейки кода будет скрывать только ввод этой ячейки.
# +
#hide_input
print('Вы увидите только результат запуска ячейки, а именно этот комментарий.')
# -
# Поместив флаг `#collapse-hide`, в первую строку любой ячейки, вы скроете код этой ячейки внутри поста. Но в замен появится кнопка, позволяющая показать эту ячейку.
#collapse-hide
import altair as alt
import pandas as pd
import numpy as np
# Флаг `#collapse-show` позволяет показать ячейку по умолчанию, но дает читателю возможность скрыть ее.
#collapse-show
np.random.seed(42)
source = pd.DataFrame(
np.cumsum(np.random.randn(100, 3), 0).round(2),
columns=['A', 'B', 'C'],
index=pd.RangeIndex(100, name='x')
)
source = source.reset_index().melt('x', var_name='category', value_name='y')
# ## Интерактивные графики с помощью [Altair](https://altair-viz.github.io/)
#
# Графики построенные с помощью библиотеки Altair внутри поста остаются интерактивными как в ноутбуке.
# +
#collapse-hide
# код отсюда: https://altair-viz.github.io/gallery/multiline_tooltip.html
# Create a selection that chooses the nearest point & selects based on x-value
nearest = alt.selection(type='single', nearest=True, on='mouseover',
fields=['x'], empty='none')
# The basic line
line = alt.Chart(source).mark_line(interpolate='basis').encode(
x='x:Q',
y='y:Q',
color='category:N'
)
# Transparent selectors across the chart. This is what tells us
# the x-value of the cursor
selectors = alt.Chart(source).mark_point().encode(
x='x:Q',
opacity=alt.value(0),
).add_selection(
nearest
)
# Draw points on the line, and highlight based on selection
points = line.mark_point().encode(
opacity=alt.condition(nearest, alt.value(1), alt.value(0))
)
# Draw text labels near the points, and highlight based on selection
text = line.mark_text(align='left', dx=5, dy=-5).encode(
text=alt.condition(nearest, 'y:Q', alt.value(' '))
)
# Draw a rule at the location of the selection
rules = alt.Chart(source).mark_rule(color='gray').encode(
x='x:Q',
).transform_filter(
nearest
)
# Put the five layers into a chart and bind the data
alt.layer(
line, selectors, points, rules, text
).properties(
width=600, height=300
)
# -
# ## Отображение таблиц
#
# Таблицы в опубликованных постах отображаются примерно как и в Jupyter ноутбуках.
source.head()
# ## Вставка изображений
# Вы можете добавлять изображения с подписями следующим образом:
#
# ``
#
# 
#
# Напомню, что подписи опциональны, и что изображения можно указывать как локально (в рамках репозитория блога), так и находящиеся в открытом доступе (имею в виду интернет).
# ## Анимированные гифки
#
# Гифки вставляются как изображения и полноценно отображаются в постах.
#
# 
# ## Видео Youtube
# Чтобы красиво вставить видео с Youtube достаточно использовать конструкцию:
#
# `> youtube: https://youtu.be/L0boq3zqazI`
#
# [Видео туториал](#Видео-туториал) по настройке блога прикреплен именно таким образом.
# ## Посты из Twitter
# Есть возможность отображать посты из Twitter.
#
# Например, ссылка на этот пост `> twitter: https://twitter.com/jakevdp/status/1204765621767901185?s=20` отобразит слудущее:
#
# > twitter: https://twitter.com/jakevdp/status/1204765621767901185?s=20
# ## `LaTeX` формулы
# Jupyter ноутбуки поддерживают синтаксис `LaTeX` формул. Чтобы формулы отображались в постах, нужно убедиться, что опция `use_math` включена внутри `_config.yml` (см. [Настройка блога](#Настройка-блога)).
#
# Следуюший `LaTeX` код:
# > `$$L(\theta) = \frac{1}{N} \sum_i^N{(y_i - \hat{y_i})^2} \rightarrow \min_{\theta}$$`
#
# будет отображен таким образом:
# $$L(\theta) = \frac{1}{N} \sum_i^N{(y_i - \hat{y_i})^2} \rightarrow \min_{\theta}$$
# ## Примечания
#
# Есть возможность отображать примечания различных типов.
#
# **Предупреждение:** `> Warning: There will be no second warning!`
#
# > Warning: There will be no second warning!
#
#
# **Важно:** `> Important: Pay attention! It's important.`
#
# > Important: Pay attention! It's important.
#
#
# **Подсказка:** `> Tip: This is my tip.`
#
# > Tip: This is my tip.
#
#
# **Заметка:** `> Note: Take note of this.`
#
# > Note: Take note of this.
#
#
# Если вставить в любое из примечаний ссылку, то она будет работать.
#
# Например,
#
# `> Note: A doc link to [an example website: fast.ai](https://www.fast.ai/) should also work fine.`
#
# отобразится так:
#
# > Note: A doc link to [an example website: fast.ai](https://www.fast.ai/) should also work fine.
# ## Отображение Emoji
# Если написать `Сейчас будет эмоджи :robot:.`, то получится
#
# Сейчас будет эмоджи :robot:.
#
# [Шпаргалка](https://www.webfx.com/tools/emoji-cheat-sheet/) по Emoji.
# ## Сноски
#
# В jupyter ноутбуках можно использовать сноски, однако синтаксис отличается от Markdown разметки. Это [руководство](https://github.com/fastai/fastpages/blob/master/_fastpages_docs/NOTEBOOK_FOOTNOTES.md) содержит более подробную информацию об этом синтаксисе, который выглядит следующим образом:
#
# ```
# {% raw %}For example, here is a footnote {% fn 1 %}.
# And another {% fn 2 %}
# {{ 'This is the footnote.' | fndetail: 1 }}
# {{ 'This is the other footnote. You can even have a [link](https://fastpages.fast.ai/jupyter/2020/02/20/test.html#Footnotes)' | fndetail: 2 }}{% endraw %}
# ```
#
# For example, here is a footnote {% fn 1 %}.
# And another {% fn 2 %}
# {{ 'This is the footnote.' | fndetail: 1 }}
# {{ 'This is the other footnote. You can even have a [link](https://fastpages.fast.ai/jupyter/2020/02/20/test.html#Footnotes)' | fndetail: 2 }}
# # Как `fastpages` конвертирует исходные файлы посты
#
# Для этого `fastpages` использует [nbdev](https://nbdev.fast.ai/index.html) для преобразования jupyter ноутбуков, word и `.md` файлов в посты блога. После того, как вы сохраните исходные файлы своих постов в папках `/_notebooks`, `/_word` или `/_posts`, то GitHub Actions c помощью nbdev автоматически преобразует их в конечный вид, в котором посты отображаются на сайте вашего блога.
# # `fast_template` - младший брат `fastpages`
# Стоит упомянуть, что ранее `fast.ai` выпустили аналогичный проект под названием [fast_template](https://github.com/fastai/fast_template/), который еще проще в настройке, но не поддерживает автоматическое создание постов из Word и Jupyter файлов, а также многие другие функции перечисленные выше. Поскольку `fastpages` более гибок и расширяем, его авторы рекомендуют использовать его там, где это возможно.
#
# Авторы предполагают, что `fast_template` может быть лучшим вариантом для тех, кто ведет не технические блоги. В этом случае посты можно создавать только с помощью встроенного онлайн-редактора Github, не заморачиваясь с использованием `git`.
# # Плюсы и минусы
#
# ## Что понравилось
# - простота создания и размещения блога и публикации контента;
# - возможность публиковать Jupyter ноутбуки в качестве постов + удобства оформления:
# - поддержка отображения интерактивных графиков;
# - скрытие/сворачивание кода;
# - поддержка отображения GIF-анимации;
# - интеграция видео с youtube и тд.
# - нет зависимости от сторонней платформы по типу Medium;
# - возможность разместить блог по собственному url;
# - параметр `badges` в метаинформации к посту позволяет прикрепить ссылки на `GitHub`, `Binder`, `Google Colab`, что позволяет сразу перейти от поста к коду и его исполнению;
# - комментарии для блога из коробки;
# - возможность прикрепить пост на конкретуню позицию на общей странице с помощью `sticky_rank`, смотреть [тут](#Возможности-форматирования-постов);
# - отсутствие сторонней рекламы;
#
# ## Что не понравилось или вызывало вопросы
# - непонятно, как сделать структурированный блог с вложенностью:
# - возможное решение [permalinks](https://jekyllrb.com/docs/permalinks/)];
# - структура нужна для объединения нескольких постов общей темой;
# - хочется структуру, чтобы в одной директорий хранить все, что связанно с постом (данные, изображения для ноутбуков) в одной папке, а не искать их в куче общих файлов и не городить какую-то структуру в этих общих для всех постов папках.
# - нет WYSIWYG (What You See Is What You Get):
# - `Jekyll` его и не подразумевает из коробки;
# - возможен [локальный запуск](https://github.com/fastai/fastpages#running-the-blog-on-your-local-machine) блога;
# - в `Jekyll` в заголовке и описании поста не поддерживаются обратные кавычки, квадратные скобки и тд.
# - `Jekyll` подразумевает использование `git` для публикации постов;
# - целесообразность хранения Jupyter ноутбуков в репозитории под вопросом;
# - непонятно, как привязать spell checker для Jupyter ноутбуков.
# # Резюме
# Команда fast.ai предложили DS сообществу интересный и достаточно функциональный инструмент для ведения блога, автору которого остается думать только о том, какой контент публиковать.
#
# Сложность использования практически минимальная, нужны базовые знания `git`, разметки Markdown и Jupyter Notebook. И никаких проблем с тем, как и где хостить и деплоить сам блог.
#
# Конечно, есть определенные пожелания по поводу функционала этого движка, для этого можно участвовать в развитии проекта, находя баги или предлогая те или иные улучшения. В последнем случае даже `pull request` не обязателен, порой хватает текстового описание тех или иных желаний пользователей.
#
# В заключение хочу сказать, что сам пользуюсь и всем советую.
# # DS/ML/AI блоги
# - [Пример блога](https://drscotthawley.github.io/devblog3/) на `fastpages` by <NAME>;
# - [Анализ малых данных](https://dyakonov.org/) блог Александра Дьяконова;
# - <NAME> [github.io](http://karpathy.github.io/), [medium](https://medium.com/@karpathy);
# - [Machine Learning Mastery](https://machinelearningmastery.com/blog/) by <NAME>;
#
# ## Блоги компаний
# - [Fast.ai](https://www.fast.ai/) + [fastpages blog](https://fastpages.fast.ai/);
# - [Airbnb](https://medium.com/airbnb-engineering/ai/home);
# - [Uber](https://eng.uber.com/category/articles/ai/);
# - [OpenAI](https://openai.com/blog/);
# - [DeepMind](https://deepmind.com/blog);
# - [Nvidia](https://blogs.nvidia.com/blog/category/deep-learning/) + [AI podcast](https://blogs.nvidia.com/ai-podcast/);
# - Microsoft [AI blog](https://blogs.microsoft.com/ai/) + [ML devblogs](https://devblogs.microsoft.com/cse/tag/,machine-learning-ml/)
# # Полезные ссылки
# - [Репозиторий](https://github.com/fastai/fastpages#setup-instructions) проекта `fastpages`;
# - [Introducing fastpages](https://fastpages.fast.ai/fastpages/jupyter/2020/02/21/introducing-fastpages.html);
# - [Туториал](https://fastpages.fast.ai/jupyter/2020/02/20/test.html) с примерами того, что можно сделать в `.ipynb` посте;
# - [Репозиторий](https://github.com/fastai/fast_template/) проекта `fast_template` + [статья от fast.ai](https://www.fast.ai/2020/01/16/fast_template/);
# - [Домашняя страница](https://nbdev.fast.ai/index.html) проекта `nbdev` + [репозиторий](https://github.com/fastai/nbdev) + [форум](https://forums.fast.ai/c/fastai-users/nbdev/);
# - Достаточно широкий [обзор](https://habr.com/ru/company/ruvds/blog/501012/) движков для блога;
# - Еще блоги \[[1](https://medium.com/datadriveninvestor/best-ai-ml-data-science-blogs-to-follow-in-2019-c3598032e3b8), [2](https://www.springboard.com/blog/machine-learning-blog/)\];
| _notebooks/2020-05-26-DS-fastpages-blog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="5CVwx4E09qJM" colab_type="code" outputId="54089c14-8afa-4744-dda3-deac961cf5ba" executionInfo={"status": "ok", "timestamp": 1559302496784, "user_tz": -330, "elapsed": 16584, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16620884610365813544"}} colab={"base_uri": "https://localhost:8080/", "height": 425}
# !apt-get install protobuf-compiler python-pil python-lxml python-tk
# !git clone https://github.com/tensorflow/models.git
# !cd models/research; protoc object_detection/protos/*.proto --python_out=.
# !cd models/research; export PYTHONPATH=$PYTHONPATH:`pwd`:`pwd`/slim; python object_detection/builders/model_builder_test.py
# + id="sCTFVe-6sU_n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1398} outputId="566b8c07-d8ab-4339-c036-b63ded6f2d26" executionInfo={"status": "ok", "timestamp": 1559303173317, "user_tz": -330, "elapsed": 12302, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16620884610365813544"}}
# !pip install object-detection
# + id="xIkwQ4vmn85F" colab_type="code" colab={}
import functools
import json
import os
import sys
import tensorflow as tf
# + id="_GYcbfk3GZ_6" colab_type="code" colab={}
sys.path.append('models')
sys.path.append('models/research')
sys.path.append('models/research/object_detection')
sys.path.append('models/research/slim')
# + id="X2MUtaeqn85M" colab_type="code" outputId="472b74a0-dd00-4338-9169-c1184e27b020" executionInfo={"status": "error", "timestamp": 1559303155376, "user_tz": -330, "elapsed": 757, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16620884610365813544"}} colab={"base_uri": "https://localhost:8080/", "height": 368}
from object_detection.builders import dataset_builder
from object_detection.builders import graph_rewriter_builder
from object_detection.builders import model_builder
from object_detection.legacy import trainer
from object_detection.utils import config_util
# + id="dp-DsTz_n85P" colab_type="code" colab={}
tf.logging.set_verbosity(tf.logging.INFO)
# + id="7ql-5mPTHbcC" colab_type="code" outputId="cb6b9a9c-f262-4c3c-be97-729f0ccd6dd2" executionInfo={"status": "ok", "timestamp": 1559293250570, "user_tz": -330, "elapsed": 40089, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16620884610365813544"}} colab={"base_uri": "https://localhost:8080/", "height": 122}
#Mount google drive to access the data from drive
from google.colab import drive
drive.mount('/content/drive')
# + id="vzUv5_BVn85T" colab_type="code" colab={}
train_dir = "/content/drive/My Drive/Programming/gun_detector/training/new"
num_clones = 1
clone_on_cpu = False
pipeline_config_path = "/content/drive/My Drive/Programming/gun_detector/training/ssd_inception_v2_coco_for_colab.config"
# + id="FF8uxwj3n85W" colab_type="code" colab={}
@tf.contrib.framework.deprecated(None, 'Use object_detection/model_main.py.')
def main():
assert train_dir, 'train_dir is misssing'
if pipeline_config_path:
configs = config_util.get_configs_from_pipeline_file(pipeline_config_path)
else:
None
model_config = configs['model']
train_config = configs['train_config']
input_config = configs['train_input_config']
model_fn = functools.partial(model_builder.build,
model_config=model_config,
is_training=True)
def get_next(config):
return dataset_builder.make_initializable_iterator(
dataset_builder.build(config)).get_next()
create_input_dict_fn = functools.partial(get_next, input_config)
env = json.loads(os.environ.get('TF_CONFIG', '{}'))
cluster_data = env.get('cluster', None)
cluster = tf.train.ClusterSpec(cluster_data) if cluster_data else None
task_data = env.get('task', None) or {'type': 'master', 'index': 0}
task_info = type('TaskSpec', (object,), task_data)
# Parameters for a single worker.
ps_tasks = 0
worker_replicas = 1
worker_job_name = 'lonely_worker'
task = 0
is_chief = True
master = ''
if cluster_data and 'worker' in cluster_data:
# Number of total worker replicas include "worker"s and the "master".
worker_replicas = len(cluster_data['worker']) + 1
if cluster_data and 'ps' in cluster_data:
ps_tasks = len(cluster_data['ps'])
if worker_replicas > 1 and ps_tasks < 1:
raise ValueError('At least 1 ps task is needed for distributed training.')
if worker_replicas >= 1 and ps_tasks > 0:
# Set up distributed training.
server = tf.train.Server(tf.train.ClusterSpec(cluster), protocol='grpc',
job_name=task_info.type,
task_index=task_info.index)
if task_info.type == 'ps':
server.join()
return
worker_job_name = '%s/task:%d' % (task_info.type, task_info.index)
task = task_info.index
is_chief = (task_info.type == 'master')
master = server.target
graph_rewriter_fn = None
if 'graph_rewriter_config' in configs:
graph_rewriter_fn = graph_rewriter_builder.build(
configs['graph_rewriter_config'], is_training=True)
trainer.train(create_input_dict_fn,
model_fn,
train_config,
master,
task,
num_clones,
worker_replicas,
clone_on_cpu,
ps_tasks,
worker_job_name,
is_chief,
train_dir,
graph_hook_fn=graph_rewriter_fn)
# + id="x-9giVwen85a" colab_type="code" outputId="3c4f5460-eb91-437e-d81c-d42b12401711" executionInfo={"status": "error", "timestamp": 1559296955813, "user_tz": -330, "elapsed": 45182, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16620884610365813544"}} colab={"base_uri": "https://localhost:8080/", "height": 6481}
main()
# + id="XibhagF3ItO-" colab_type="code" colab={}
| training/train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.5 64-bit
# name: python37564bit8409255418034dfeada81c2f6070bc9d
# ---
# + tags=[]
from cy_data_access.models.position import *
import pandas as pd
pd.set_option('expand_frame_repr', False) # 当列太多时不换行
connect_db_env(db_name=DB_POSITION)
selling = list(AIMSPositionSelling.objects.values())
df = pd.DataFrame(selling)
df.drop(['_cls', '_id'], axis=1, inplace=True)
print("""
{}
sum: {}
""".format(df, df['profit_amount'].sum()))
# -
| tests/db.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# !rm -rf gambiae.g?f.gz ag.db 2>/dev/null
# #!wget http://www.vectorbase.org/download/anopheles-gambiae-pestbasefeaturesagamp42gtfgz -O gambiae.gtf.gz
# !wget http://www.vectorbase.org/download/anopheles-gambiae-pestbasefeaturesagamp42gff3gz -O gambiae.gff.gz
import gffutils
import sqlite3
# !rm -f ag.db
try:
db = gffutils.create_db('gambiae.gff.gz', 'ag.db')
except sqlite3.OperationalError:
db = gffutils.FeatureDB('ag.db')
print(list(db.featuretypes()))
for feat_type in db.featuretypes():
print(feat_type, db.count_features_of_type(feat_type))
for contig in db.features_of_type('contig'):
print(contig)
from collections import defaultdict
num_mRNAs = defaultdict(int)
num_exons = defaultdict(int)
max_exons = 0
max_span = 0
for contig in db.features_of_type('contig'):
cnt = 0
for gene in db.region((contig.seqid, contig.start, contig.end), featuretype='gene'):
cnt += 1
span = abs(gene.start - gene.end) # strand
if span > max_span:
max_span = span
max_span_gene = gene
my_mRNAs = list(db.children(gene, featuretype='mRNA'))
num_mRNAs[len(my_mRNAs)] += 1
if len(my_mRNAs) == 0:
exon_check = [gene]
else:
exon_check = my_mRNAs
for check in exon_check:
num_exons[len(my_exons)] += 1
if len(my_exons) > max_exons:
max_exons = len(my_exons)
max_exons_gene = gene
print('contig %s, number of genes %d' % (contig.seqid, cnt))
print('Max number of exons: %s (%d)' % (max_exons_gene.id, max_exons))
print('Max span: %s (%d)' % (max_span_gene.id, max_span))
print(num_mRNAs)
print(num_exons)
| notebooks/02_Genomes/Annotations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fit the DDM on individual data
import rlssm
import pandas as pd
import os
# ## Import the data
# +
data = rlssm.load_example_dataset(hierarchical_levels = 1)
data.head()
# -
# ## Initialize the model
model = rlssm.DDModel(hierarchical_levels = 1)
# ## Fit
# sampling parameters
n_iter = 1000
n_chains = 2
n_thin = 1
model_fit = model.fit(
data,
thin = n_thin,
iter = n_iter,
chains = n_chains,
pointwise_waic=False,
verbose = False)
# ### get Rhat
model_fit.rhat
# ### get wAIC
model_fit.waic
# ## Posteriors
model_fit.samples.describe()
import seaborn as sns
sns.set(context = "talk",
style = "white",
palette = "husl",
rc={'figure.figsize':(15, 8)})
model_fit.plot_posteriors(height=5, show_intervals="HDI", alpha_intervals=.05);
# ## Posterior predictives
# ### Ungrouped
pp = model_fit.get_posterior_predictives_df(n_posterior_predictives=100)
pp
pp_summary = model_fit.get_posterior_predictives_summary(n_posterior_predictives=100)
pp_summary
model_fit.plot_mean_posterior_predictives(n_posterior_predictives=100, figsize=(20,8), show_intervals='HDI');
model_fit.plot_quantiles_posterior_predictives(n_posterior_predictives=100, kind='shades');
# ### Grouped
import numpy as np
# +
# Define new grouping variables, in this case, for the different choice pairs, but any grouping var can do
data['choice_pair'] = 'AB'
data.loc[(data.cor_option == 3) & (data.inc_option == 1), 'choice_pair'] = 'AC'
data.loc[(data.cor_option == 4) & (data.inc_option == 2), 'choice_pair'] = 'BD'
data.loc[(data.cor_option == 4) & (data.inc_option == 3), 'choice_pair'] = 'CD'
data['block_bins'] = pd.cut(data.trial_block, 8, labels=np.arange(1, 9))
# -
model_fit.get_grouped_posterior_predictives_summary(
grouping_vars=['block_label', 'choice_pair'],
quantiles=[.3, .5, .7],
n_posterior_predictives=100)
model_fit.get_grouped_posterior_predictives_summary(
grouping_vars=['block_bins'],
quantiles=[.3, .5, .7],
n_posterior_predictives=100)
model_fit.plot_mean_grouped_posterior_predictives(grouping_vars=['block_bins'],
n_posterior_predictives=100,
figsize=(20,8));
model_fit.plot_quantiles_grouped_posterior_predictives(
n_posterior_predictives=100,
grouping_var='choice_pair',
kind='shades',
quantiles=[.1, .3, .5, .7, .9]);
| docs/notebooks/DDM_fitting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kundajelab/dragonn/blob/master/paper_supplement/LowMemQC.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PgZwwRreD9P9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="cd3cecf0-42f6-4d03-b5ed-a5430d3a903e"
#uncomment the lines below if you are running this tutorial from Google Colab
# !pip install dragonn>=0.2.6
# + id="aK_u2unQD9QD" colab_type="code" colab={}
# Making sure our results are reproducible
from numpy.random import seed
seed(1234)
from tensorflow import set_random_seed
set_random_seed(1234)
# + id="CdnEx_EiD9QE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4a223c4f-5098-498e-948c-184a798c59b6"
#load dragonn tutorial utilities
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
from dragonn.tutorial_utils import *
# + [markdown] id="CIyS9aXkD9QI" colab_type="text"
# ## Input data <a name='1'>
# <a href=#outline>Home</a>
#
# Tutorials 1 - 3 have used simulated data generated with the simdna package. In this tutorial, we will examine how well CNN's are able to predict transcription factor binding for four TF's in vivo.
#
# We will learn to predict transcription factor binding for four transcription factors in the GM12878 cell line (one of the Tier 1 cell lines for the ENCODE project). First, we download the narrowPeak bed files for each of these transcription factors. You can skip the following code block if you already have the data downloaded.
# + id="joKrWPskD9QJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 210} outputId="d9c814ad-2728-43d7-88f1-e760f82f56de"
## SPI1, optimal IDR thresholded peaks, Myers lab, hg19
# https://www.encodeproject.org/experiments/ENCSR000BGQ/
# #!wget -O SPI1.narrowPeak.gz http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.narrowPeak.gz
## Download "ambiguous" peak sets -- these peaks are in the optimal overlap set across replicates, but are not
## found to be reproducible at a high confidence (p<0.05) by IDR
# #! wget -O SPI1.ambiguous.gz http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.ambiguous.gz
## Download the hg19 chromsizes file (We only use chroms 1 -22, X, Y for training)
# !wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.chrom.sizes
## Download the hg19 fasta reference genome (and corresponding .fai index)
# #!wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.genome.fa.gz
# #!wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.genome.fa.fai
# + [markdown] id="xPNOawx_D9QL" colab_type="text"
# ## Generating positive and negative bins for genome-wide training <a name='2'>
# <a href=#outline>Home</a>
# + [markdown] id="T5YXbDhrD9QM" colab_type="text"
# We will use the *genomewide_labels* function from the [seqdataloader](https://github.com/kundajelab/seqdataloader) package to generate positive and negative labels for the TF-ChIPseq peaks across the genome. We will treat each sample as a task for the model and compare the performance of the model on SPI1 task in the single-tasked and multi-tasked setting.
# + id="ty7OVrz2D9QO" colab_type="code" colab={}
from seqdataloader import *
# + id="P2MY4tuCD9QR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ecd61735-2357-49a1-e480-7338fa116d9a"
## seqdataloader accepts an input file, which we call tasks.tsv, with task names in column 1, the corresponding
## peak files in column 2, skip column 3 (which will be used for regression in Tutorial 5), and ambiguous peaks in
## column4
with open("tasks.tsv",'w') as f:
f.write("\t".join(["SPI1","SPI1.narrowPeak.gz","","SPI1.ambiguous.gz"])+'\n')
# ! cat tasks.tsv
# + [markdown] id="794ATwuBD9QV" colab_type="text"
# With the parameter configuration below, seqdataloader splits the genome into 1kb regions, with a stride of 50. Each 1kb region is centered at a 200 bp bin, with a left flank of 400 bases and a right flank of 400 bases.
#
# * Each 200 bp bin is labeled as positive if a narrowPeak summit overlaps with it.
#
# * The bin is labeled ambiguous (label = -1) and excluded from training if there is some overlap with the narrowPeak, but the peak summit does not lie in that overlap.
#
# * The bin is labeled negative if there is no overlap with the narrowPeak.
# + id="YNZ6kU5INojl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="89313292-1d6b-4098-d845-9732f774c404"
# !pwd
# + id="q5JWEtjXD9QW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 227} outputId="4c143a7a-e45f-4480-e0e3-56ce82cc2de9"
positives_train_set_params={
'store_positives_only':True,
'task_list':"tasks.tsv",
'outf':"positives.TF.train.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_exclude':['chr1','chr2','chr19'],
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':20,
'subthreads':2,
'allow_ambiguous':True,
'output_hdf5_low_mem':True,
'labeling_approach':'peak_summit_in_bin_classification'
}
genomewide_labels(positives_train_set_params)
# + id="aqbArOrhD9QY" colab_type="code" colab={}
positives_valid_set_params={
'store_positives_only':True,
'task_list':"tasks.tsv",
'outf':"positives.TF.valid.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_keep':'chr1',
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':20,
'subthreads':2,
'allow_ambiguous':True,
'output_hdf5_low_mem':True,
'labeling_approach':'peak_summit_in_bin_classification'
}
genomewide_labels(positives_valid_set_params)
# + id="xAOzk3rdD9Qb" colab_type="code" colab={}
positives_test_set_params={
'store_positives_only':True,
'task_list':"tasks.tsv",
'outf':"positives.TF.test.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_keep':['chr2','chr19'],
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':20,
'subthreads':2,
'allow_ambiguous':True,
'output_hdf5_low_mem':True,
'labeling_approach':'peak_summit_in_bin_classification'
}
genomewide_labels(positives_test_set_params)
# + id="XyDOKpsDD9Qf" colab_type="code" colab={}
# When provided with the --store-positives_only flag, the code generates all bins for each task that are labeled positive.
pd.read_hdf("SPI1.positives.TF.train.hdf5",start=0,stop=10)
# + id="eAjuqJM1D9Qj" colab_type="code" colab={}
from dragonn.generators import *
# + id="gCHAgATXD9Ql" colab_type="code" colab={}
#To prepare for model training, we import the necessary functions and submodules from keras
from keras.models import Sequential
from keras.layers.core import Dropout, Reshape, Dense, Activation, Flatten
from keras.layers.convolutional import Conv2D, MaxPooling2D
from keras.optimizers import Adadelta, SGD, RMSprop;
import keras.losses;
from keras.constraints import maxnorm;
from keras.layers.normalization import BatchNormalization
from keras.regularizers import l1, l2
from keras.callbacks import EarlyStopping, History, TensorBoard
from keras import backend as K
K.set_image_data_format('channels_last')
#we use a custom binary cross-entropy loss that can handle ambiguous labels (denoted with -1 ) and exclude them
# from the loss calculation
from dragonn.custom_losses import get_ambig_binary_crossentropy
# + id="MiPHtJcsD9Qn" colab_type="code" colab={}
from concise.metrics import tpr, tnr, fpr, fnr, precision, f1
from keras.constraints import max_norm
def initialize_model(ntasks=1):
#Define the model architecture in keras (regularized, 3-layer convolution model followed by 1 dense layer)
model=Sequential()
model.add(Conv2D(filters=50,kernel_size=(1,15),padding="same", kernel_constraint=max_norm(7.0,axis=-1),input_shape=(1,1000,4)))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Conv2D(filters=50,kernel_size=(1,15),padding="same"))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Conv2D(filters=50,kernel_size=(1,13),padding="same"))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(1,40)))
model.add(Flatten())
model.add(Dense(50))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Dense(ntasks))
model.add(Activation("sigmoid"))
#use the custom ambig_binary_crossentropy loss, indicating that a value of -1 indicates an ambiguous label
loss=get_ambig_binary_crossentropy(-1)
##compile the model, specifying the Adam optimizer, and binary cross-entropy loss.
model.compile(optimizer='adam', loss=loss,
metrics=[tpr,
tnr,
fpr,
fnr,
precision,
f1])
return model
# + id="uSbcq-9jD9Qq" colab_type="code" colab={}
#create the generators
from dragonn.generators import *
case1_spi1_train_gen=DataGenerator("SPI1.positives.TF.train.hdf5","hg19.genome.fa.gz",shuffled_ref_negatives=True,upsample=False,batch_size=256)
case1_spi1_valid_gen=DataGenerator("SPI1.positives.TF.valid.hdf5","hg19.genome.fa.gz",shuffled_ref_negatives=True,upsample=False,batch_size=256)
case1_ctcf_train_gen=DataGenerator("CTCF.positives.TF.train.hdf5","hg19.genome.fa.gz",shuffled_ref_negatives=True,upsample=False,batch_size=256)
case1_ctcf_valid_gen=DataGenerator("CTCF.positives.TF.valid.hdf5","hg19.genome.fa.gz",shuffled_ref_negatives=True,upsample=False,batch_size=256)
# + [markdown] id="Q4OsGDSAD9Qt" colab_type="text"
# We now follow the standard protocol we used in tutorials 1 - 3 to train a keras model, with the exception that we use the fit_generator function in keras, rather than the fit function.
# + id="HJVdXu_ED9Qu" colab_type="code" colab={}
callbacks=[EarlyStopping(patience=3,restore_best_weights=True),History()]
# + id="2x6by142D9Qz" colab_type="code" colab={}
#If you are running this notebook in google colab, uncomment the lines below to observe the model's training
# !mkdir logs
# %tensorboard --logdir logs
tensorboard_visualizer=TensorBoard(log_dir="logs", histogram_freq=0, batch_size=500, write_graph=True, write_grads=False, write_images=False, embeddings_freq=0, embeddings_layer_names=None, embeddings_metadata=None)
callbacks.append(tensorboard_visualizer)
# + id="Bt6twhsVD9Q4" colab_type="code" colab={} outputId="e5465ed2-5cd8-40a7-882c-5a2dd6f9d2e3"
#Train the SPI1 model
case1_spi1_model=initialize_model()
## use the keras fit_generator function to train the model with early stopping after 3 epochs
history_case1_spi1=case1_spi1_model.fit_generator(case1_spi1_train_gen,
validation_data=case1_spi1_valid_gen,
epochs=150,
verbose=1,
use_multiprocessing=True,
workers=40,
max_queue_size=100,
callbacks=callbacks)
# + id="EvgcnKtlD9Q9" colab_type="code" colab={}
## Plot the learning curves for SPI1
from dragonn.tutorial_utils import plot_learning_curve
plot_learning_curve(history_case1_spi1)
| tutorials/LowMemQC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ray Concepts - Task Parallelism (Part 2)
#
# The previous lesson explored Ray's core concepts and how they work. We learned how to define Ray _tasks_, run them, and retrieve the results. We also started learning about how Ray schedules tasks in a distributed environment.
#
# This lesson completes the discussion of Ray tasks by exploring how task dependencies are handled. We'll finish with a look under the hood at Ray's architecture and runtime behavior.
#
# > **Tip:** Recall that the [Ray Package Reference](https://ray.readthedocs.io/en/latest/package-ref.html) in the [Ray Docs](https://ray.readthedocs.io/en/latest/) is useful for exploring the API features we'll learn.
# +
# If you are running on Google Colab, uncomment and run the following linkes
# to install the necessary dependencies.
# print("Setting up colab environment")
# # !pip install -q ray
# # !pip install -q bokeh
# +
# Imports and initialize Ray. We're adding NumPy for the examples and the tutorial `util` library:
import ray, time, sys # New notebook, so new process
import numpy as np # Used for examples
# +
def pnd(n, duration, prefix=''):
"""Print an integer and a time duration, with an optional prefix."""
prefix2 = prefix if len(prefix) == 0 else prefix+' '
print('{:s}n: {:2d}, duration: {:6.3f} seconds'.format(prefix2, n, duration))
def pd(duration, prefix=''):
"""Print a time duration, with an optional prefix."""
prefix2 = prefix if len(prefix) == 0 else prefix+' '
print('{:s}duration: {:6.3f} seconds'.format(prefix2, duration))
# -
ray.init(ignore_reinit_error=True)
# Let's work with a new remote function. Previously, our `expensive` and `expensive_task` functions returned tuples that included time durations. Obviously the durations were useful for understanding how long the functions took to execute. Now, it will be more convenient to not return "metadata" like this, but just data values that we care about, because we are going to pass them to other functions.
#
# Hence, we'll define _dependency_ relationships between tasks. We'll learn how Ray handles these dependent, asynchronous computations.
#
# So, let's define a task to return a random NumPy array of some size `n`. As before, we'll add a sleep time, one tenth the size of `n`:
@ray.remote
def make_array(n):
time.sleep(n/10.0)
return np.random.standard_normal(n)
# Now define a task that can add two NumPy arrays together. The arrays need to be the same size, but we'll ignore any checking for this requirement.
@ray.remote
def add_arrays(a1, a2):
time.sleep(a1.size/10.0)
return np.add(a1, a2)
# Now lets use them!
start = time.time()
id1 = make_array.remote(20)
id2 = make_array.remote(20)
id3 = add_arrays.remote(id1, id2)
print(ray.get(id3))
pd(time.time() - start, prefix="Total time:")
# Something subtle and "magical" happened here; when we called `add_arrays`, we didn't need to call `ray.get()` first for `id1` and `id2`, since `add_arrays` expects NumPy arrays. Because `add_arrays` is a Ray task, Ray automatically does the extraction for us, so we can write code that looks more natural.
#
# Furthermore, note that the `add_arrays` task effectively depends on the outputs of the two `make_array` tasks. Ray won't run `add_arrays` until the other tasks are finished. Hence, _Ray handles task dependencies automatically for us._
#
# This is why the elapsed time is about 4 seconds. We used a size of 20, so we slept 2 seconds in each call to `make_array`, but those happened in parallel, _followed_ by a second sleep of 2 seconds in `add_arrays`.
# Recall from the previous lesson that we explored when to call `ray.get()` to avoid forcing tasks to become synchronous when they should be asynchronous. This additional example illustrates two key points:
#
# * _Don't ask for results you don't need._
# * _Don't ask for the results you need until you really need them._
#
# We don't need to see the objects for `id1` and `id2`. We only need the final array for `id3`.
# ## Using ray.wait() with ray.get()
#
# We've seen several examples of the best idiomatic way to use `ray.get()`. Here again is an example from the last lesson:
#
# ```python
# start = time.time()
# ids = [expensive_task.remote(n) for n in range(5)] # Fire off the asynchronous tasks
# for n2, duration in ray.get(ids): # Retrieve all the values from the list of futures
# p(n2, duration)
# pd(time.time() - start, prefix="Total time:")
# ```
#
# Let's try it again with our new methods:
start = time.time()
array_ids = [make_array.remote(n*10) for n in range(5)]
added_array_ids = [add_arrays.remote(id, id) for id in array_ids]
for array in ray.get(added_array_ids):
print(f'{array.size}: {array}')
pd(time.time() - start, prefix="Total time:")
# On my machine, I waited 8 seconds and then everything was printed at once.
# There are two fundamental problems with the way we've used `ray.get()` so far:
#
# 1. There's no timeout, in case something gets "hung".
# 2. We have to wait for _all_ the objects to be available before `ray.get()` returns.
#
# The ability to specify a timeout is essential in production code as a defensive measure. Many potential problems could happen in a real production system, any one of which could cause the task we're waiting on to take an abnormally long time to complete or never complete. Our application would be deadlocked waiting on this task. Hence, it's **strongly recommended** in production software to always use timeouts on blocking calls, so that the application can attempt some sort of recovery in situations like this, or at least report the error and "degrade gracefully".
#
# Actually, there _is_ a `timeout=<value>` option you can pass to `ray.get()` ([documentation](https://ray.readthedocs.io/en/latest/package-ref.html#ray.get)), but it will most likely be removed in a future release of Ray. Why remove it if timeouts are important? This change will simplify the implementation of `ray.get()` and encourage the use of `ray.wait()` for waiting ([documentation](https://ray.readthedocs.io/en/latest/package-ref.html#ray.wait)) instead, followed by using `ray.get()` to retrieve values for tasks that `ray.wait()` tells us are finished.
#
# Using `ray.wait()` is also the way to fix the second problem with using `ray.get()` by itself, that we have to wait for all tasks to finish before we get any values back. Some of those tasks might finish quickly, like our contrived examples that sleep for short durations compared to other invocations.
#
# When you have a list of asynchronous tasks, you want to process the results of them as soon they become available, even while others continue to run. Use `ray.wait()` for this purpose.
#
# Therefore, while `ray.get()` is simple and convenient, for _production code_, we recommend using `ray.wait()`, **with** timeouts, for blocking on running tasks. Then use `ray.get()` to retrieve values of completed tasks. Now we'll learn how to use these two together. For a longer discussion on `ray.wait()`, see [this blog post](https://medium.com/distributed-computing-with-ray/ray-tips-and-tricks-part-i-ray-wait-9ed7a0b9836d).
#
# Here is the previous example rewritten to use `ray.wait()`:
# +
start = time.time()
array_ids = [make_array.remote(n*10) for n in range(5)]
added_array_ids = [add_arrays.remote(id, id) for id in array_ids]
arrays = []
waiting_ids = list(added_array_ids) # Assign a working list to the full list of ids
while len(waiting_ids) > 0: # Loop until all tasks have completed
# Call ray.wait with:
# 1. the list of ids we're still waiting to complete,
# 2. tell it to return immediately as soon as one of them completes,
# 3. tell it wait up to 10 seconds before timing out.
ready_ids, remaining_ids = ray.wait(waiting_ids, num_returns=1, timeout=10.0)
print('Returned {:3d} completed tasks. (elapsed time: {:6.3f})'.format(len(ready_ids), time.time() - start))
new_arrays = ray.get(ready_ids)
arrays.extend(new_arrays)
for array in new_arrays:
print(f'{array.size}: {array}')
waiting_ids = remaining_ids # Reset this list; don't include the completed ids in the list again!
print(f"\nall arrays: {arrays}")
pd(time.time() - start, prefix="Total time:")
# -
# Now it still takes about 8 seconds to complete, 4 seconds for the longest invocation of `make_array` and 4 seconds for the invocation of `add_arrays`, but since the others complete more quickly, we see their results as soon as they become available, at 0, 2, 4, and 6 second intervals.
#
# > **Warning:** For each call to `ray.wait()` in a loop like this, it's important to remove the ids that have completed. Otherwise, `ray.wait()` will return immediately with the same list containg the first completed item, over and over again; you'll loop forever!! Resetting the list is easy, since the second list returned by `ray.wait()` is the rest of the items that are still running. So, that's what we use.
#
# Now let's try it with `num_returns = 2`:
# +
start = time.time()
array_ids = [make_array.remote(n*10) for n in range(5)]
added_array_ids = [add_arrays.remote(id, id) for id in array_ids]
arrays = []
waiting_ids = list(added_array_ids) # Assign a working list to the full list of ids
while len(waiting_ids) > 0: # Loop until all tasks have completed
# Call ray.wait with:
# 1. the list of ids we're still waiting to complete,
# 2. tell it to return immediately as soon as TWO of them complete,
# 3. tell it wait up to 10 seconds before timing out.
return_n = 2 if len(waiting_ids) > 1 else 1
ready_ids, remaining_ids = ray.wait(waiting_ids, num_returns=return_n, timeout=10.0)
print('Returned {:3d} completed tasks. (elapsed time: {:6.3f})'.format(len(ready_ids), time.time() - start))
new_arrays = ray.get(ready_ids)
arrays.extend(new_arrays)
for array in new_arrays:
print(f'{array.size}: {array}')
waiting_ids = remaining_ids # Reset this list; don't include the completed ids in the list again!
print(f"\nall arrays: {arrays}")
pd(time.time() - start, prefix="Total time:")
# -
# Now we get two at a time output. Note that we don't actually pass `num_returns=2` every time. If you ask for more items than the length of the input list, you get an error. So, we compute `num_returns`, using `2` except when there's only one task to wait on, in which case we use `1`. So, in fact, the output for `40` was a single task result, because we started with `5` and processed two at a time.
# ## Exercise 2
#
# The following cell is identical to the last one. Modify it to use a timeout of `2.5` seconds, shorter than our longest tasks. What happens now? Try using other times.
# +
start = time.time()
array_ids = [make_array.remote(n*10) for n in range(5)]
added_array_ids = [add_arrays.remote(id, id) for id in array_ids]
arrays = []
waiting_ids = list(added_array_ids) # Assign a working list to the full list of ids
while len(waiting_ids) > 0: # Loop until all tasks have completed
# Call ray.wait with:
# 1. the list of ids we're still waiting to complete,
# 2. tell it to return immediately as soon as TWO of them complete,
# 3. tell it wait up to 10 seconds before timing out.
return_n = 2 if len(waiting_ids) > 1 else 1
ready_ids, remaining_ids = ray.wait(waiting_ids, num_returns=return_n, timeout=2.5)
print('Returned {:3d} completed tasks. (elapsed time: {:6.3f})'.format(len(ready_ids), time.time() - start))
new_arrays = ray.get(ready_ids)
arrays.extend(new_arrays)
for array in new_arrays:
print(f'{array.size}: {array}')
waiting_ids = remaining_ids # Reset this list; don't include the completed ids in the list again!
print(f"\nall arrays: {arrays}")
pd(time.time() - start, prefix="Total time:")
# -
# In conclusion:
#
# > **Tips:**
# >
# > 1. Use `ray.wait()` with a timeout to wait for one or more running tasks. Then use `ray.get()` to retrieve the values for the finished tasks.
# > 2. Don't ask for results you don't need.
# > 3. Don't ask for the results you need until you really need them.
# ## Exercise 3
#
# Let's make sure you understand how to use `ray.wait()`. The definitions from Exercise 1 in the previous lesson are repeated in the next cell. Change the definitions to use Ray. In particular, use `ray.wait()` as we used it above. You can just use the default values for `num_returns` and `timeout` if you want. The second cell uses `assert` statements to check your work.
#
# > **Tip:** The solution is in the `solutions` folder.
# +
def slow_square(n):
time.sleep(n)
return n*n
@ray.remote
def fast_square(n):
return slow_square(n)
start = time.time()
square_ids = [fast_square.remote(n) for n in range(4)]
squares = []
waiting_ids = list(square_ids) # Assign a working list to the full list of ids
while len(waiting_ids) > 0: # Loop until all tasks have completed
# Call ray.wait with:
# 1. the list of ids we're still waiting to complete,
# 2. tell it to return immediately as soon as TWO of them complete,
# 3. tell it wait up to 10 seconds before timing out.
return_n = 2 if len(waiting_ids) > 1 else 1
ready_ids, remaining_ids = ray.wait(waiting_ids, num_returns=return_n, timeout=2.5)
new_squares = ray.get(ready_ids)
squares.extend(new_squares)
waiting_ids = remaining_ids # Reset this list; don't include the completed ids in the list again!
duration = time.time() - start
# -
assert squares == [0, 1, 4, 9], f'Did you use ray.get() to retrieve the values? squares = {squares}'
assert duration < 4.1, f'Did you use Ray to parallelize the work? duration = {duration}'
# ## What Is the Optimal Task Granularity
#
# How fine-grained should Ray tasks be? There's no fixed rule of thumb, but Ray clearly adds some overhead for task management and using object stores in a cluster. Therefore, it makes sense that tasks which are too small will perform poorly.
#
# We'll explore this topic over several more lessons, but for now, let's get a sense of the overhead while running in your setup.
#
# We'll continue to use NumPy arrays to create "load", but remove the `sleep` calls:
# +
def noop(n):
return n
def local_make_array(n):
return np.random.standard_normal(n)
@ray.remote
def remote_make_array(n):
return local_make_array(n)
# -
# Let's do `trials` runs for each experiment, to average out background noise:
trials=100
# First, let's use `noop` to baseline local function calls. Note that we call `print` for the duration, rathern than `pd`, because the overhead is so low the `pd` formatting will print `0.000`:
start = time.time()
[noop(t) for t in range(trials)]
print(f'{time.time() - start} seconds')
# Let's try the same run with `local_make_array(n)` for `n = 100000`:
start = time.time()
[local_make_array(100000) for _ in range(trials)]
print(f'{time.time() - start} seconds')
# So, we can safely ignore the "noop" overhead for now. For completeness, here's what happens with remote execution:
start = time.time()
ids = [remote_make_array.remote(100000) for _ in range(trials)]
ray.get(ids)
print(f'{time.time() - start} seconds')
# For arrays of 100000, using Ray is faster (at least on this test machine). The benefits of parallel computation, rather than synchronous, already outweight the Ray overhead.
#
# So, let's run some trials with increasingly large array sizes, to compare the performance with local vs. remote execution. First, we'll set up `matplotlib`:
local_durations = []
remote_durations = []
# These n values were determined by experimentation on this test machine.
# If you are using an old machine, and this cell takes a long time to execute,
# you could set the `trials` value above to a smaller number.
ns = [i*(10**j) for j in range(2,5) for i in [1,2,3,5,8]]
for n in ns:
start_local = time.time()
[local_make_array(n) for _ in range(trials)]
local_durations.append(time.time() - start_local)
start_remote = time.time()
ids = [remote_make_array.remote(n) for _ in range(trials)]
ray.get(ids)
remote_durations.append(time.time() - start_remote)
(ns, local_durations, remote_durations)
# +
import numpy as np
from bokeh.layouts import gridplot
from bokeh.plotting import figure, output_file, show
import bokeh.io
# The next two lines prevent Bokeh from opening the graph in a new window.
bokeh.io.reset_output()
bokeh.io.output_notebook()
tooltips = [
("name", "$name"),
("array size", "$x"),
("time", "$y")]
p1 = figure(x_axis_type="log", y_axis_type="log", title="Execution Times", tooltips=tooltips)
p1.grid.grid_line_alpha=0.3
p1.xaxis.axis_label = 'array size'
p1.yaxis.axis_label = 'time'
p1.line(ns, local_durations, color='#A6CEE3', legend_label='local', name='local')
p1.circle(ns, local_durations, color='darkgrey', size=4)
p1.line(ns, remote_durations, color='#B2DF8A', legend_label='remote', name='remote')
p1.square(ns, remote_durations, color='darkgrey', size=4)
p1.legend.location = "top_left"
show(gridplot([[p1]], plot_width=800, plot_height=400))
# -
# Here's a static image from a test run, in case the Bokeh plot isn't working. Your results may look a lot different!
# 
# Let's confirm what the graph shows as the crossing point:
i=0
while i < len(ns) and local_durations[i] < remote_durations[i]:
i=i+1
print('The Ray times are faster starting at n = {:d}, local = {:6.3f} vs. remote = {:6.3f}'.format(
ns[i], local_durations[i], remote_durations[i]))
# ## How Distributed Task Management Works
#
# > **Note:** If you just want to learn the Ray API, you can safely skip the rest of this lesson (notebook) for now. It continues the exploration of how Ray works internally, which we started in the previous lesson. However, you should come back to this material at some point, so you'll develop a better understanding of how Ray works.
# At the end of the last lesson, we examined Ray task scheduling at a high-level, by watching the Ray Dashboard and analyzing the performance times. Now we'll walk through some images that show the process Ray follows to place tasks around a cluster.
# Assume we will invoke the `make_array` task twice, then invoke `add_arrays` to sum the returned NumPy arrays. Graphically, it looks as follows:
#
# 
# How does this get scheduled in a cluster? Here we'll assume a three-node cluster that has resources for running two Ray worker tasks per node (under powered compared to what we learned using Ray Dashboard last lesson!).
# 
# First, assume that the driver program is running on Node1. So it will invoke the local scheduler to schedule the three tasks.
# 
# Immediately the ids for the task futures are returned. The _Global Control Store_ tracks where every task is running and every object is stored in the local _Object Stores_.
# 
# Suppose the local scheduler has available capacity in the first worker on the same node. It schedules the first `make_array` task there.
# 
# It decides to schedule the second `make_array` task in a worker on node 2.
# 
# When the two tasks finish, they place their result objects in their local object stores.
# 
# Now `add_array` can be scheduled, because the two tasks it depends on are done. Let's suppose it gets scheduled in the second worker on Node 1.
# 
# The first object it needs is already on the same node, in the object store, so the `add_arrays` task can _read it directly from shared memory_. No copying is required to the worker's process space.
# 
# However, the second object is on a different node, so Ray copies it to the local object store.
# 
# Now it can also be read from shared memory.
# 
# When `add_arrays` is finished, it writes its results to the local object store.
# 
# At this point, if the driver calls `ray.get(id3)`, it will return `obj3`.
# 
# Whew! Hopefully you have a better sense of what Ray does under the hood. Scheduling tasks on other nodes and copying objects between object stores is efficient, but incurs unavoidable network overhead.
| ray-core/03-TaskParallelism-Part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zj7ViHva58R_" colab_type="text"
# ### Package Preparation
# + id="QUz01gDzateB" colab_type="code" outputId="972f0211-adad-44ea-82f2-e6f2a4dfadb6" colab={"base_uri": "https://localhost:8080/", "height": 258}
# !pip install tf_sentencepiece
import tensorflow as tf
import numpy as np
import tensorflow_hub as hub
import tf_sentencepiece
from random import shuffle, choice
import re
import os
import datetime
from functools import reduce
from operator import itemgetter
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="KCsHRfC556Oo" colab_type="text"
# ### Configurations
# + id="i4zp8CgboUSO" colab_type="code" colab={}
xling_encoding_len = 512
max_seg = 10
level_class_cnt = 3
test_percentage = 0.1
validation_percentage = 0.1
dropout_rate = 0.5
eta = 1e-4
hidden_feature_dim = 100
attention_key_dim = 100
gru_feature_dim = 50
batch_size = 512
epochs = 8
label_re = re.compile('(\d+)\.\d+')
sentence_re = re.compile('(?:\.|!|\?)\s')
input_path = '/content/gdrive/My Drive/data_source/milnet/raw_text/gourmet.txt'
model_out_path = '/content/gdrive/My Drive/data_source/milnet/results/food_xling_c3.h5'
log_out_dir = '/content/gdrive/My Drive/data_source/milnet/log/'
sample_amount = 0
with open(input_path) as in_file:
sample_amount = len(in_file.read().split('\n\n')) - 1
sample_indices = [*range(sample_amount)]
shuffle(sample_indices)
train_samples = sample_indices[0:int(sample_amount * (1 - test_percentage - validation_percentage))]
validation_samples = sample_indices[int(sample_amount * (1 - test_percentage - validation_percentage)): int(sample_amount * (1 - test_percentage))]
test_samples = sample_indices[int(sample_amount * (1 - test_percentage)):]
# + [markdown] id="3lzuDqbj4oOn" colab_type="text"
# ### Data Preloading
# + id="kvM3Gi7P4m8t" colab_type="code" colab={}
g = tf.Graph()
with g.as_default():
text_input = tf.placeholder(dtype=tf.string, shape=[None])
en_de_embed = hub.Module("https://tfhub.dev/google/universal-sentence-encoder-xling/en-de/1")
embedded_text = en_de_embed(text_input)
init_op = tf.group([tf.global_variables_initializer(), tf.tables_initializer()])
g.finalize()
session = tf.Session(graph=g)
session.run(init_op)
# + id="0Amu2Opkazbg" colab_type="code" colab={}
def __pad_doc_encoding(doc_encoding, max_seg):
if doc_encoding.shape[0] > max_seg:
return doc_encoding[:max_seg]
elif doc_encoding.shape[0] < max_seg:
topad_len = max_seg - doc_encoding.shape[0]
pad_width = [(0, 0) if i != 0 else (0, topad_len) for i in range(len(doc_encoding.shape))]
return np.pad(doc_encoding, pad_width, 'constant', constant_values=0)
else:
return doc_encoding
def __label_map(raw_label):
if raw_label == 1 or raw_label == 2:
return 0
elif raw_label == 3:
return 1
else:
return 2
def __balance_data(feature_array, label_array):
to_balance_indices = np.concatenate([np.where(label_array == 2)[0], np.where(label_array == 4)[0]])
return np.delete(feature_array, to_balance_indices, axis=0), np.delete(label_array, to_balance_indices, axis=0)
def data_generator(sample_indices, input_path, segment_re, label_re,
batch_size=batch_size, max_seg=max_seg, xling_len=xling_encoding_len, epochs=epochs, use_balance=True):
global session, embedded_text, text_input
with open(input_path) as in_file:
file_content = [*itemgetter(*sample_indices)(in_file.read().split('\n\n'))]
for _ in range(epochs):
shuffle(file_content)
feature_cache, label_cache = [], []
batch_index = 0
for sample in file_content:
label_cache.append(sample.split('\n')[0])
feature_cache.append([*filter(lambda x: len(x) > 1, segment_re.split(' '.join(sample.split('\n')[1:])))])
batch_index += 1
if batch_index == batch_size:
len_lst = [*map(len, feature_cache)]
batch_features = session.run(embedded_text, feed_dict={text_input: reduce(lambda x, y: x + y, feature_cache)})
label_array = np.array([np.array([int(label_re.findall(l)[0])]) for l in label_cache])
feature_array = np.zeros((batch_size, max_seg, xling_len))
for index, length in enumerate(len_lst):
feature_array[index] = __pad_doc_encoding(np.array(batch_features[:length]), max_seg)
batch_features = batch_features[length:]
feature_array = np.array(feature_array)
if use_balance:
feature_array, label_array = __balance_data(feature_array, label_array)
yield feature_array, np.array([np.array([__label_map(l[0])]) for l in label_array])
feature_cache, label_cache = [], []
batch_index = 0
# + id="NJO9L0aAhUEG" colab_type="code" colab={}
shared_sublayer_cache = {}
def branch_execute(layer_in, sublayer, args={}):
instance_cnt = layer_in.shape[1]
sliced_inputs = [tf.keras.layers.Lambda(lambda x: x[:,i])(layer_in)
for i in range(instance_cnt)]
branch_layers = [sublayer(**{**{'layer_in': sliced_inputs[i]}, **args})
for i in range(instance_cnt)]
expand_layer = tf.keras.layers.Lambda(lambda x: tf.keras.backend.expand_dims(x, axis=1))
expanded_layers = [expand_layer(branch_layers[i]) for i in range(instance_cnt)]
concated_layer = tf.keras.layers.Concatenate(axis=1)(expanded_layers)
return concated_layer
def __seg_classifier_layer_share(layer_in, class_cnt, dropout_rate, eta):
global shared_sublayer_cache
if 'shared_seg_classifier_sublayers' not in shared_sublayer_cache:
shared_sublayer_cache['shared_seg_classifier_sublayers'] = {
'drop_out_layer': tf.keras.layers.Dropout(
dropout_rate
),
'dense_layer': tf.keras.layers.Dense(
units=class_cnt,
activation='softmax',
kernel_regularizer=tf.keras.regularizers.l2(eta),
bias_regularizer=tf.keras.regularizers.l2(eta)
)
}
shared_layers = shared_sublayer_cache['shared_seg_classifier_sublayers']
drop_out_layer = shared_layers['drop_out_layer'](layer_in)
dense_layer = shared_layers['dense_layer'](drop_out_layer)
return dense_layer
def __attention_layer_share(layer_in, attention_key_dim, dropout_rate, eta):
global shared_sublayer_cache
if 'shared_attention_sublayers' not in shared_sublayer_cache:
shared_sublayer_cache['shared_attention_sublayers'] = {
'drop_out_layer': tf.keras.layers.Dropout(
dropout_rate
),
'dense_layer': tf.keras.layers.Dense(
units=attention_key_dim,
activation='tanh',
kernel_regularizer=tf.keras.regularizers.l2(eta),
bias_regularizer=tf.keras.regularizers.l2(eta)
),
'nobias_dense_layer': tf.keras.layers.Dense(
units=1,
use_bias=False,
bias_regularizer=tf.keras.regularizers.l2(eta)
)
}
shared_layers = shared_sublayer_cache['shared_attention_sublayers']
drop_out_layer = shared_layers['drop_out_layer'](layer_in)
dense_layer = shared_layers['dense_layer'](drop_out_layer)
nobias_dense_layer = shared_layers['nobias_dense_layer'](dense_layer)
return nobias_dense_layer
def bidirectional_gru_layer(layer_in, gru_feature_dim):
bidirectional_layer = tf.keras.layers.Bidirectional(
tf.keras.layers.GRU(gru_feature_dim, return_sequences=True)
)(layer_in)
return bidirectional_layer
def merge_layer(layer_in, class_cnt, eta):
dot_layer = tf.keras.layers.Dot(axes=1)(layer_in)
flatten_layer = tf.keras.layers.Flatten()(dot_layer)
dense_layer = tf.keras.layers.Dense(
units=class_cnt,
activation='softmax',
kernel_regularizer=tf.keras.regularizers.l2(eta),
bias_regularizer=tf.keras.regularizers.l2(eta)
)(flatten_layer)
return dense_layer
def performance_judge(model, generator, class_cnt):
eps = np.finfo(float).eps
accuracy, precisions, recalls, f1s = [], [], [], []
for i, (features, labels) in enumerate(generator):
predicted = model.predict(features)
precisions.append([])
recalls.append([])
f1s.append([])
contingency_table = np.zeros((class_cnt, class_cnt))
for index in range(features.shape[0]):
contingency_table[int(labels[index][0])][np.argmax(predicted[index])] += 1
accuracy.append(np.trace(contingency_table) / features.shape[0])
for index in range(class_cnt):
precisions[i].append(contingency_table[index][index] / (np.sum(contingency_table[:, index]) + eps))
recalls[i].append(contingency_table[index][index] / (np.sum(contingency_table[index, :]) + eps))
f1s[i].append(2 * precisions[i][-1] * recalls[i][-1] / ((precisions[i][-1] + recalls[i][-1]) + eps))
precisions = [float(sum(l))/len(l) for l in zip(*precisions)]
recalls = [float(sum(l))/len(l) for l in zip(*recalls)]
f1s = [float(sum(l))/len(l) for l in zip(*f1s)]
print('Accuracy:', round(reduce(lambda x, y: x + y, accuracy) / len(accuracy), 3))
for index in range(class_cnt):
print('_____ Class', index, '_____')
print('Precision\t', round(precisions[index], 3))
print('Recall\t\t', round(recalls[index], 3))
print('F1 Score\t', round(f1s[index], 3))
# + id="5kSdKsCX9iwa" colab_type="code" outputId="4f62e343-fd70-4a4b-c49b-acc124b4d65c" colab={"base_uri": "https://localhost:8080/", "height": 1000}
print('Constructing Model ...', end='')
model_input = tf.keras.Input((max_seg, xling_encoding_len))
biglu_layer = bidirectional_gru_layer(
model_input,
gru_feature_dim=50
)
attention_layer = branch_execute(
biglu_layer,
sublayer=__attention_layer_share,
args={
'attention_key_dim': 100,
'dropout_rate': 0.5,
'eta': 1e-4
}
)
softmaxed_attention_layer = tf.keras.layers.Softmax(
axis=1
)(attention_layer)
classification_layer = branch_execute(
model_input,
sublayer=__seg_classifier_layer_share,
args={
'class_cnt': level_class_cnt,
'dropout_rate': 0.5,
'eta': 1e-4
}
)
merge_layer = merge_layer(
[softmaxed_attention_layer, classification_layer],
class_cnt=level_class_cnt,
eta=1e-4
)
model = tf.keras.Model(model_input, merge_layer)
print('\rModel Constructed. Compiling ...', end='')
model.compile(
optimizer=tf.keras.optimizers.Adam(clipvalue=0.5),
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=['accuracy']
)
print('\rModel Compiled.')
model.summary()
# + id="MaTGynpcAi61" colab_type="code" outputId="26d61b72-d480-419d-e9b6-7acec2ba20b5" colab={"base_uri": "https://localhost:8080/", "height": 887}
logdir = os.path.join(log_out_dir, datetime.datetime.now().strftime("%Y%m%d_%H%M%S"))
tensorboard_callback = tf.keras.callbacks.TensorBoard(logdir, histogram_freq=0)
model.fit_generator(
data_generator(train_samples, input_path, sentence_re, label_re, use_balance=True),
validation_data=data_generator(validation_samples, input_path, sentence_re, label_re, use_balance=True),
steps_per_epoch=(sample_amount * (1 - test_percentage - validation_percentage) // batch_size) - 1,
validation_steps=(sample_amount * (validation_percentage) // batch_size) - 1,
validation_freq=2,
epochs=epochs,
callbacks=[tensorboard_callback]
)
model.save(model_out_path)
print('########## Training Error ##########')
performance_judge(model, data_generator(train_samples, input_path, sentence_re, label_re, epochs=1, use_balance=True), level_class_cnt)
print('')
print('############ Test Error ############')
performance_judge(model, data_generator(train_samples, input_path, sentence_re, label_re, epochs=1, use_balance=True), level_class_cnt)
print(logdir)
# + id="u6mviP8_f_4p" colab_type="code" outputId="4b08b781-637b-46e9-f9be-f8162708678e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %load_ext tensorboard
# %tensorboard --logdir logs
# + id="GkM3V8yEVlTr" colab_type="code" colab={}
| milnet_xling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Anitápolis TFA multiple Inversions
# This notebook performs the inversion using Levenberg-Marquadt's algorithm of total field anomaly (TFA).
# +
import numpy as np
import matplotlib.pyplot as plt
import cPickle as pickle
import os
import pandas as pd
from fatiando.vis import mpl
# -
from datetime import datetime
today = datetime.today()
# dd/mm/YY/Hh/Mm
d4 = today.strftime("%d-%b-%Y-%Hh:%Mm")
d4
# ### Auxiliary functions
# +
import sys
sys.path.insert(0, '../../code')
import mag_polyprism_functions as mfun
# -
# # Input
# ### Importing model parameters
# output of inversion
inversion = dict()
data = pd.read_csv('anitapolis_mag.txt', skipinitialspace=True, delim_whitespace=True)
data['GPSALT'] = - data['GPSALT'] + 800
mask = (data['GPSALT'].get_values()<0.)
data = data[mask]
plt.figure(figsize=(10,7))
plt.title('Observed TFA', fontsize=20)
plt.tricontourf(data['Y'], data['X'], data['mag_res'], 20, cmap='RdBu_r').ax.tick_params(labelsize=12)
plt.plot(data['Y'], data['X'], 'k.')
plt.xlabel('$y$(km)', fontsize=18)
plt.ylabel('$x$(km)', fontsize=18)
clb = plt.colorbar(pad=0.025, aspect=40, shrink=1)
clb.ax.tick_params(labelsize=13)
#estimate = mpl.polygon(model0[0], '.-r', xy2ne=True)
#estimate.set_label('Initial estimate')
clb.ax.set_title('nT')
mpl.m2km()
plt.show()
xp = data['X'].get_values()
yp = data['Y'].get_values()
zp = data['GPSALT'].get_values()
alt = data['ALTURA'].get_values()
dobs = data['mag_res'].get_values()
plt.figure(figsize=(10,7))
plt.title('Observed TFA', fontsize=20)
plt.tricontourf(yp, xp, dobs, 20, cmap='RdBu_r').ax.tick_params(labelsize=12)
plt.xlabel('$y$(km)', fontsize=18)
plt.ylabel('$x$(km)', fontsize=18)
clb = plt.colorbar(pad=0.025, aspect=40, shrink=1)
clb.ax.tick_params(labelsize=13)
clb.ax.set_title('nT')
mpl.m2km()
plt.legend(loc=0, fontsize=12, shadow=bool, framealpha=1)
plt.show()
# ### Parameters of the initial model
# +
M = 20 # number of vertices per prism
L = 6 # number of prisms
P = L*(M+2) + 1 # number of parameters
#figura
incs = -21.
decs = -11.
int_min = 11.
int_max = 20.
intensity = np.linspace(int_min, int_max, 10)
# depth to the top, thickness and radius
z0_min = 0.
z0_max = 180.
z0 = np.linspace(z0_min, z0_max, 10)
dz = 900.
r = 700.
x0 = 6921000.
y0 = 688000.
# main field
inc, dec = [-37.05, -18.17]
# -
z0
intensity
# ### Limits
# +
# limits for parameters in meters
rmin = 10.
rmax = 1200.
y0min = 687000.
y0max = 699000.
x0min = 6916000.
x0max = 6923000.
dzmin = 10.
dzmax = 1000.
mmin, mmax = mfun.build_range_param(M, L, rmin, rmax, x0min, x0max, y0min, y0max, dzmin, dzmax)
# -
# ### Variation
# variation for derivatives
deltax = 0.01*np.max(100.)
deltay = 0.01*np.max(100.)
deltar = 0.01*np.max(100.)
deltaz = 0.01*np.max(100.)
# ### Outcropping parameters
# outcropping body parameters
m_out = np.zeros(M + 2)
#m_out = model['param_vec'][:M+2]
# ### Regularization parameters
# +
#lamb = th*0.01 # Marquadt's parameter
lamb = 10.0
dlamb = 10. # step for Marquadt's parameter
a1 = 1.0e-4 # adjacent radial distances within each prism
a2 = 1.0e-3 # vertically adjacent radial distances
a3 = 0. # outcropping cross-section
a4 = 0. # outcropping origin
a5 = 1.0e-4 # vertically adjacent origins
a6 = 1.0e-8 # zero order Tikhonov on adjacent radial distances
a7 = 1.0e-5 # zero order Tikhonov on thickness of each prism
# -
foldername = ''
delta = np.array([deltax, deltay, deltar, deltaz])
alpha = np.array([a1, a2, a3, a4, a5, a6, a7])
itmax = 30
itmax_marq = 10
tol = 1.0e-4 # stop criterion
# ### Inversion
inversion_results = []
for j, z in enumerate(z0):
for k, i in enumerate(intensity):
alpha = np.array([a1, a2, a3, a4, a5, a6, a7])
print 'inversion: %d top: %d intensity: %2.f' % (j*z0.size + k, z, i)
model0, m0 = mfun.initial_cylinder(M, L, x0, y0, z, dz, r, inc, dec, incs, decs, i)
d_fit, m_est, model_est, phi_list, model_list, res_list = mfun.levmarq_tf(
xp, yp, zp, m0, M, L, delta,
itmax, itmax_marq, lamb,
dlamb, tol, mmin, mmax,
m_out, dobs, inc, dec,
model0[0].props, alpha, z, dz
)
inversion_results.append([m_est, phi_list, model_list, dobs - d_fit])
# # Results
inversion['x'] = xp
inversion['y'] = yp
inversion['z'] = zp
inversion['observed_data'] = dobs
inversion['inc_dec'] = [incs, decs]
inversion['z0'] = z0
inversion['initial_dz'] = dz
inversion['intial_r'] = r
inversion['limits'] = [rmin, rmax, x0min, x0max, y0min, y0max, dzmin, dzmax]
inversion['regularization'] = np.array([a1, a2, a3, a4, a5, a6, a7])
inversion['tol'] = tol
inversion['main_field'] = [inc, dec]
inversion['intensity'] = intensity
inversion['results'] = inversion_results
# ### Folder to save the results
if foldername == '':
mypath = 'results/multiple-'+d4 #default folder name
if not os.path.isdir(mypath):
os.makedirs(mypath)
else:
mypath = 'results/multiple-'+foldername #defined folder name
if not os.path.isdir(mypath):
os.makedirs(mypath)
file_name = mypath+'/inversion.pickle'
with open(file_name, 'w') as f:
pickle.dump(inversion, f)
| code/anitapolis/multiple_inversion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# load in the libraries we will use
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import GridSearchCV
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.model_selection import ShuffleSplit
from sklearn.metrics import make_scorer
from sklearn.metrics import r2_score
import visuals as vs
# %matplotlib inline
train = pd.read_csv('houseTrain.csv')
test = pd.read_csv('houseTest.csv')
print len(train)
print len(test)
# First split the training data into the features and the target
features = train.drop(['Id', 'SalePrice'], axis=1)
target = train.SalePrice
# How many features are there?
print len(features.columns.values)
# Which columns are missing values?
nan_columns = features.columns[pd.isnull(features).any()].tolist()
for i in nan_columns:
print '{0}: {1} {2}'.format(i, features[i].isnull().sum(), features[i].dtypes)
# Let's turn the above into a function
def missing_data_info(df):
'''
Takes: a pandas dataframe which is checked for missing data in the columns. Uses any() method to check for
missing data.
Returns: prints out the name of columns with missing data, the number of missing values, and the dtype of the column
and returns a dictionary of the printed data in the form column_name: [#_missing_values, col_dtype]'''
nan_columns = df.columns[pd.isnull(df).any()].tolist()
nan_dict = {}
for i in nan_columns:
print '{0}: {1} {2}'.format(i, df[i].isnull().sum(), features[i].dtypes)
nan_dict[i] = [df[i].isnull().sum(), features[i].dtypes]
return nan_dict
# The Alley, FireplaceQu, PoolQC, Fence, MiscFeatures objects should be dropped
variables_to_drop = ['Alley', 'FireplaceQu', 'PoolQC', 'Fence', 'MiscFeature']
def graph_mean_on_scatter(df, x_name, y_name):
'''
Input:
df: the pandas data frame the data is contained in
x_name: the name of the df column to plot on the x-axis
y_name: the name of the df column to plot on the y-axis
Dependencies:
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
Returns:
A regplot which has any missing values plotted separately in order to evaluate the usefullness of
substituting in the mean for the missing data.'''
missing = pd.DataFrame(data=df[x_name][df[x_name].isnull()], columns=[x_name])
missing[y_name] = df[y_name][df[x_name].isnull()]
missing[x_name].fillna(value=np.nanmean(df[x_name]), inplace=True)
sns.regplot(x=x_name, y=y_name, data=df)
plt.plot(missing[x_name], missing[y_name], 's')
plt.show()
return
# Now need to decide how to fill the missing values from the remaining factors
# Let's start with LotFrontage
graph_mean_on_scatter(train, 'LotFrontage', 'SalePrice')
# +
# The mean LotFrontage values fit right in with the rest of the data. Let's fill in the missing data using these values
mean_LotFrontage = np.nanmean(features.LotFrontage)
features.LotFrontage.fillna(value=mean_LotFrontage, inplace=True)
# check that LotFrontage column isn't missing any daya
missing_data_info(features)
#success!
# -
# Let's try the same strategy above for MasVnrArea which is the Masonry Veneer Area
graph_mean_on_scatter(train, 'MasVnrArea', 'SalePrice')
# I'm going to use the mean value to fill the missing data
mean_MasVnrArea = np.nanmean(features.MasVnrArea)
features.MasVnrArea.fillna(value=mean_MasVnrArea, inplace=True)
# Look at how using the mean value works for GarageYrBlt
graph_mean_on_scatter(train, 'GarageYrBlt', 'SalePrice')
# I don't like that so much of imputed data is weighted down below the trend line.
# It might be that the value for GarageYrBlt doens't exists is because there is no garage!
# Looking back at the number of values of missing data for the garage features,
# there are an equal number of missing features. This would suggest that those homes don't have a garage.
# And in fact, the same pattern is true for the basement features.
# We can test this hypothesis by looking at the comparison between GarageYrBlt and YearBlt
graph_mean_on_scatter(train, 'GarageYrBlt', 'YearBuilt')
# Looks like most of the garages were built when the house was built, so this likely isn't going to add much information
# I'll add the garage features to the variables_to_drop list
variables_to_drop.extend(['GarageType', 'GarageYrBlt', 'GarageFinish', 'GarageQual', 'GarageCond'])
features.drop(variables_to_drop, axis=1, inplace=True)
missing_data_info(features)
# Let's look through the remaining columns that are missing data points. These are all categorical variables
# We'll start with MasVnrType
sns.boxplot(x='MasVnrType', y='SalePrice', data=train)
sns.stripplot(x="MasVnrType", y="SalePrice", data=train,
size=2, jitter=True, edgecolor="gray", alpha=0.5)
# The BrkFace veneer type looks like it is right in the middle. Since we used the mean values to for the
# MasVnrArea, I'll use BrkFace as the "mean" value to use as the replacement for this feaure
features.MasVnrType.fillna(value='BrkFace', inplace=True)
# ### Basement Condition
# The BsmtCond feature documentation says that NA values mean there is no basement. I want to see if it shows up as a factor
sns.boxplot(x='BsmtCond', y='SalePrice', data=train)
sns.stripplot(x='BsmtCond', y='SalePrice', data=train, size=2, jitter=True, edgecolor="gray", alpha=0.5)
# Since 'Na' string in the basement features doesn't count as a category, I'm going to replace the Na values with the
# string 'None'
for i in ['BsmtCond', 'BsmtExposure', 'BsmtFinType1', 'BsmtFinType2', 'BsmtQual']:
features.loc[:,i].fillna(value='NoBsmt', inplace=True)
missing_data_info(features)
# only 1 feature left with null values! This feature ranks the circuit breaker/fuse box quality of the house
# let's take a look at it
sns.boxplot(x='Electrical', y='SalePrice', data=train)
sns.stripplot(x='Electrical', y='SalePrice', data=train, size = 2, jitter=True, alpha=0.5)
train.SalePrice[train.Electrical.isnull()]
# +
# Since so many of the values are SBrkr and the price of the house missing the data is pretty close to the median of
# the SBrkr values, I'm going to assign the missing value to SBrkr
features.Electrical.fillna(value="SBrkr", inplace=True)
#and check that it worked
missing_data_info(features)
# +
# success! That was a good pratice of investigating and filling missing features. Now I will use just the numeric
# data to train a DecisionTreeRegressor model and make another submission
# <NAME>: "The best data scientists rapidly iterate."
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
numeric_features = features.select_dtypes(include=numerics)
# -
vs.ModelLearning(numeric_features, target)
vs.ModelComplexity(numeric_features, target)
# I'm going with a max depth of 7
reg = DecisionTreeRegressor(max_depth = 7)
reg.fit(numeric_features, target)
# prepare the test data
test.drop(variables_to_drop, axis=1, inplace=True)
numeric_test_features = test.select_dtypes(include=numerics)
numeric_test_features.drop('Id', axis=1, inplace=True)
# need to impute the missing data
the_missing_test_data = missing_data_info(numeric_test_features)
for k in the_missing_test_data.keys():
numeric_test_features[k].fillna(value=np.nanmean(numeric_test_features[k]), inplace=True)
missing_data_info(numeric_test_features)
# No missing data points in the test set so:
DTRegressor = reg.predict(numeric_test_features)
print len(DTRegressor)
test_id = test.Id
print len(test_id)
DTR_submission = pd.DataFrame(data=DTRegressor, index=test_id, columns=['SalePrice'])
print DTR_submission.head()
DTR_submission.to_csv('DTR_submission3.csv')
def performance_metric(y_true, y_predict):
""" Calculates and returns the performance score between
true and predicted values based on the metric chosen. """
# TODO: Calculate the performance score between 'y_true' and 'y_predict'
score = r2_score(y_true, y_predict)
# Return the score
return score
# this was worse than the first model!
# Let's do hyperparameter tuning
# split the data into training and testing
X_train, X_test, y_train, y_test = train_test_split(numeric_features, target, test_size = 0.2, random_state=0)
# make the scorer
r2_scorer = make_scorer(performance_metric)
parameters = {'max_depth':[1,3,6,10,20], 'min_samples_split':[2,5,10,15,18,20,23,25,28,30,40,50,100],
'min_samples_leaf':[1,5,10,12,15,17,20,40]}
reg = DecisionTreeRegressor()
clf = GridSearchCV(reg, parameters)
clf.fit(X_train, y_train)
clf.best_score_
clf.best_params_
GS_results = pd.DataFrame(clf.cv_results_).sort_values('mean_test_score', axis=0, ascending=False)
GS_results.head()
clf.score(X_test, y_test)
DTR = clf.predict(numeric_test_features)
print len(DTRegressor)
test_id = test.Id
print len(test_id)
DTR_submission = pd.DataFrame(data=DTR, index=test_id, columns=['SalePrice'])
print DTR_submission.head()
DTR_submission.to_csv('DTR_submission4.csv')
# +
# The decision tree with the hypertuned parameters increased my score by 188 positions (0.03 decrease in
# the root mean squared error of my predictions).
# -
categorical_variables = features.select_dtypes(include= ['object'])
categorical_variables.head()
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
le = LabelEncoder()
categorical_encoded = categorical_variables.apply(le.fit_transform)
cat_reg = DecisionTreeRegressor()
cat_reg.fit(categorical_encoded, target)
numeric_categorical_test = test.select_dtypes(['object'])
numeric_categorical_test = numeric_categorical_test.apply(le.fit_transform)
numeric_categorical_test.head()
DTR = cat_reg.predict(numeric_categorical_test)
print len(DTRegressor)
test_id = test.Id
print len(test_id)
DTR_submission = pd.DataFrame(data=DTR, index=test_id, columns=['SalePrice'])
print DTR_submission.head()
DTR_submission.to_csv('DTR_submission5.csv')
# submission score of 0.39 - pretty bad!
def make_submission(test_data, test_data_index, predictor, k):
'''
test_data = a pandas dataframe containing the test data
predictor = a trained classifier
k = the number of the entry. Used to keep older entries from being overwritten
output: writes a csv file to the current directory'''
predictions = predictor.predict(test_data)
test_id = test_data_index
submission = pd.DataFrame(data=predictions, index=test_id, columns=['SalePrice'])
print submission.head()
submission.to_csv('DTR_submission{}.csv'.format(k))
make_submission(numeric_categorical_test, test.Id, cat_reg, 6)
# concatenate the categorical_encoded and the numeric_features dataframes
all_features = pd.concat([categorical_encoded, numeric_features], axis=1)
all_features.head()
X_train, X_test, y_train, y_test = train_test_split(all_features, target, test_size = 0.2, random_state=0)
reg = DecisionTreeRegressor()
clf = GridSearchCV(reg, parameters)
clf.fit(X_train, y_train)
print clf.best_score_
print clf.best_params_
print clf.score(X_test, y_test)
all_features_test = pd.concat([numeric_categorical_test, numeric_test_features], axis=1)
make_submission(all_features_test, test.Id, clf, 6)
# these predictions are worse than the hypertuned submission using only the numeric features (submission 4)
# this is not what I was expecting because this model gave higher scores on the hold-out set duing the
# GridSearchCV and gave a better score on the held out test data
# +
# I think the next thing to do is clean up this notebook and then go through the categorical variables to determine
# which are ordinal and which are not, determine how to replace eadh variable type and then use one hot encoding on
# the non-ordinal categorical variables and label encoding on the ordinal ones
| HousePrices_DTRegressor2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="TqpAXjrxBizU"
import re
import json
import pandas as pd
# + id="J_lWEvabIHcs"
def _casefold(sentence):
'''
Args:
Input : raw sentence.
Output: lower case sentence.
'''
return str(sentence).lower()
# + id="OhvQ-dHsG02Q"
def _normalize(sentence):
'''
Turn sentence into its normal form, as long it's on the dictionary. :)
Args:
Input : lowercase sentence.
Output: normalized sentence.
'''
# replace duplicate chars, e.g okeee to oke
sentence = re.sub(r'(\w)\1{2,}', r'\1\1', sentence)
# remove \n and remove space
words = sentence.strip('\n').split()
with open('slangwords.json', 'r') as f:
file = json.load(f)
slang = {value:key for value, key in file.items()}
normal = [slang.get(word, word) for word in sentence.split()]
return ' '.join(normal)
# + id="p5AaZjx0aXgz"
def preprocess(sentence):
'''
Preprocessing the sentence.
Args:
Input : raw sentence.
Output: preprocessed sentence.
'''
sentence = _casefold(sentence)
sentence = _normalize(sentence)
return sentence
# + id="KcZvfsB3IiNt"
def to_DF1(df):
'''
Args:
Input : raw dataframe, results of lapor_scraping
query | report | institute | category
Output: dataframe, preprocessed string on `report` column
'''
reports = df.report.tolist()
preprocessed = [preprocess(report) for report in reports]
df['report'] = preprocessed
df.to_csv('data/df1.csv', index=False)
return df
# + id="Z8kbhS5K9spd"
def to_DF2(df):
'''
Args:
Input : DF1, preprocessed dataframe.
query | report | institute | category
Output: tokenized report from DF1.
index | report
--- | ---
1 | sentence1_token1
1 | sentence1_token2
2 | sentence2_token1
'''
df2 = df.copy()
df2.drop(['query', 'institute', 'category', 'label'], axis=1, inplace=True)
df2.insert(0, 'report_num', pd.factorize(df['report'])[0]+1)
df2.report = [r.split() for r in df.report]
df2 = df2.explode('report')
df2.to_csv('data/df2.csv', index=False)
return df2
# + id="Bi0HaTAK93v-"
def main():
df = pd.read_csv('lapor_scraping_results.csv')
df1 = to_DF1(df)
to_DF2(df1)
# + id="DWNHfK9X-L7c"
main()
| build_df/build_df.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## □ kNN 알고리즘이란 무엇인가?
# ```
# k Nearest Neighbor 의 약자로 k개의 최근접 이웃이라는 뜻
# ML 지도학습에서 분류에 해당하는 알고리즘
# ```
# ```
# 새로 들어온 데이터가 기존 데이터의 그룹에 어느 그룹에 속하는지 찾을 때 거리가 가까운 데이터의 그룹을 자기 그룹으로 선택하는 아주 간단한 알고리즘
# ```
#
# ## □ kNN 알고리즘의 장단점
# ```
# 장점
# 단순하고 효율적
# 모델을 훈련시키지 않음
# ```
# ```
# 단점
# 모델을 생성하지 않아서 특징과 클래스간의 관계를 이해하는 능력이 제약된다
# 적절한 k값을 모델 개발자가 직접 알아내야한다.
# ```
#
# ## □ kNN의 원리
# 새로 들어온 데이터가 기존 데이터 중에서 어느 데이터에 더 인접해 있는지 거리를 계산해서 가장 가까운 거리에 있는 데이터를 자기의 이웃으로 선택하는 것
# 거리를 계산할 때 사용하는 수학식 -> 유클리드 거리 계산식
#
# p.121
#
# ### ◆ 유클리드 거리 공식을 R로 구현하기
# 1. 두 점의 좌표를 지정한다
# ```R
# a=c(2,4)
# b=c(5,6)
# ```
# 2. 두 점 사이의 거리를 구한다
a=c(2,4)
b=c(5,6)
d=sqrt((b[1]-a[1])^2+(b[2]-a[2])^2)
print(d)
# ### ◆ 3차원에서 두 점 사이의 거리 구하기
a<-c(0,3,2)
b<-c(2,0,0)
sqrt(sum((a-b)^2))
# ### ※ 문제215. a 지점과 b 지점사이의 거리를 구하는 distance라는 함수를 생성하시오
dist<-function(a,b){
sqrt(sum((a-b)^2))
}
a<-c(0,3,2)
b<-c(2,0,0)
dist(a,b)
# ### ※ 문제216. 위에서 만든 distance 함수를 이용해서 여러개의 지점과 c(4,4) 지점과의 거리를 각각 비교하시오
x <- c(1,2,4,5,6,1)
y <- c(5,6,5,2,3,7)
dist<-function(a,b){
return (sqrt(sum((a-b)^2)))
}
temp <- c()
for (i in 1:length(x)){
temp <- append(temp,dist(c(x[i],y[i]),c(4,4)))
}
print(temp)
# ### ※ 문제217. 위의 결과에서 가장 작은 값만 출력하시오
print(min(temp))
# ### ※ 문제218. 토마토와 가장 가까운 거리에 있는 음식의 종류가 무엇인지 출력하시오
fruits <- data.frame(재료=c('사과','베이컨','당근','바나나','셀러리','치즈'),
단맛=c(10,1,10,7,3,1),
아삭한맛=c(9,4,1,10,10,1),
음식종류=c('과일','단백질','과일','채소','채소','단백질'))
토마토 <- c(6,4)
temp <- c()
dist<-function(a,b){
return (sqrt(sum((a-b)^2)))
}
for (i in 1:length(fruits$재료)){
temp <- append(temp,dist(c(fruits$단맛[i],fruits$아삭한맛[i]),토마토))
}
fruits$dist <- temp
min(fruits$dist)
# ### ※ 문제219. 위에서 만든 fruits의 파생변수인 dist를 이용해서 순위 파생변수를 추가하시오
library(dplyr)
fruits$rnk <- dense_rank(fruits$dist)
fruits
# ### ※ 문제220. 위의 결과에서 순위가 3위까지인것만 출력하시오
fruits[fruits$rnk<=3,'음식종류']
# ### ※ 문제221. 위의 결과에서 최빈값을 출력하시오
rs<-fruits[fruits$rnk<=3,'음식종류'] # rnk<=3 : k parameter
names(table(rs)[table(rs)==max(table(rs))])
| 10-2. ML ch.3-1 kNN algorithm (200623).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="n4fcdzt1L0fL"
# ## Mounting your google drive
#
# You can use google drive to store and access files e.g. storing and loading data from numpy or CSV files.
# Use the following command to mount your GDrive and access your files.
# + colab={"base_uri": "https://localhost:8080/"} id="ydOU6YpVLaow" executionInfo={"status": "ok", "timestamp": 1629392111866, "user_tz": -60, "elapsed": 28999, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="925720ba-6376-48c9-cb7a-cd46a9564935"
from google.colab import drive
drive.mount('/content/gdrive/')
# + colab={"base_uri": "https://localhost:8080/"} id="aSRYEjk782Cc" executionInfo={"status": "ok", "timestamp": 1629392135679, "user_tz": -60, "elapsed": 23817, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="037cad1f-3e8f-4824-b651-5050bd211547"
# !pip install ffmpeg
# !pip install vtk
# + id="lD9BrjrtYDPi" executionInfo={"status": "ok", "timestamp": 1629392162528, "user_tz": -60, "elapsed": 2787, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}}
import os
# change the current path. The user can adjust the path depend on the requirement
os.chdir("/content/gdrive/MyDrive/Cola-Notebooks/FYP/YF")
import vtktools
# + colab={"base_uri": "https://localhost:8080/"} id="zaGMonalKI3E" executionInfo={"status": "ok", "timestamp": 1629392163602, "user_tz": -60, "elapsed": 1077, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="87a9305c-3919-4ada-a265-38b59ce2fa6b"
# ! /opt/bin/nvidia-smi
# + id="k2FU1lqyFRva"
# # !unzip csv_data.zip
# + colab={"base_uri": "https://localhost:8080/"} id="sqsQSr0eyMDy" executionInfo={"status": "ok", "timestamp": 1629392187294, "user_tz": -60, "elapsed": 7417, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="c6d8c3b9-fa9b-4af0-8b9a-08c35904b683"
# %matplotlib inline
import numpy as np
import pandas as pd
import scipy
import numpy.linalg as la
import scipy.linalg as sl
import scipy.sparse.linalg as spl
import matplotlib.pyplot as plt
import torch.nn as nn # Neural network module
import scipy.sparse as sp
import scipy.optimize as sop
import progressbar
# making slopes
import torch
from torch.utils.data import TensorDataset
import torch.nn.functional as F
from matplotlib.pyplot import LinearLocator
import matplotlib as mpl
import matplotlib.colors as colors
# create an animation
from matplotlib import animation
from IPython.display import HTML
from matplotlib import animation
import math
import ffmpeg
# !pip install pycm livelossplot
# %pylab inline
from livelossplot import PlotLosses
from torch.utils.data import DataLoader
import torch.utils.data as Data
import time
import platform
print('python version', platform.python_version())
print('torch version', torch.__version__)
print('numpy version', np.version.version)
# + colab={"base_uri": "https://localhost:8080/"} id="Rk1Uza3iuS6d" executionInfo={"status": "ok", "timestamp": 1629392187294, "user_tz": -60, "elapsed": 7, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="37c9416a-19c3-41bb-c5e6-7f17a647846a"
def set_seed(seed):
"""
Use this to set ALL the random seeds to a fixed value and take out any randomness from cuda kernels
"""
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.backends.cudnn.benchmark = True ##uses the inbuilt cudnn auto-tuner to find the fastest convolution algorithms. -
torch.backends.cudnn.enabled = True
return True
device = 'cuda' # Set out device to GPU
print('Cuda installed, running on GPU!') # print sentence
# + [markdown] id="dGuU-LvdBV3_"
# # SFC-CAE
# + id="JQaaZNOlehqX" executionInfo={"status": "ok", "timestamp": 1629392222320, "user_tz": -60, "elapsed": 532, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}}
# These functions are saved in function.py and the note are also added to that file
def saveIndex(path_train, path_valid, path_test,train_index, valid_index, test_index):
# save training and validation loss
np.savetxt(path_train,train_index, delimiter=',')
np.savetxt(path_valid,valid_index, delimiter=',')
np.savetxt(path_test,test_index, delimiter=',')
def getIndex(path_train,path_valid,path_test):
train_index = np.loadtxt(path_train,delimiter=",")
valid_index = np.loadtxt(path_valid,delimiter=",")
test_index = np.loadtxt(path_test,delimiter=",")
return train_index,valid_index,test_index
def saveMode(path_train, path_valid, path_test,mode_train, mode_valid, mode_test):
# save training and validation loss
np.savetxt(path_train,mode_train.cpu().data.numpy(), delimiter=',')
np.savetxt(path_valid,mode_valid.cpu().data.numpy(), delimiter=',')
np.savetxt(path_test,mode_test.cpu().data.numpy(), delimiter=',')
def getMode(path_train,path_valid,path_test):
mode_train = np.loadtxt(path_train,delimiter=",")
mode_valid = np.loadtxt(path_valid,delimiter=",")
mode_test = np.loadtxt(path_test,delimiter=",")
return mode_train,mode_valid,mode_test
def saveCsv(pathcsv,EPOCH):
# save training and validation loss
losses_combined = np.zeros((EPOCH,3))
losses_combined[:,0] = np.asarray(epoch_list)
losses_combined[:,1] = np.asarray(loss_list)
losses_combined[:,2] = np.asarray(loss_valid)
np.savetxt(pathcsv, losses_combined , delimiter=',')
def PlotMSELoss(pathName,name):
epoch = pd.read_csv(pathName,usecols=[0]).values
train_loss = pd.read_csv(pathName,usecols=[1]).values
val_loss = pd.read_csv(pathName,usecols=[2]).values
fig = plt.figure(figsize=(10,7))
axe1 = plt.subplot(111)
axe1.semilogy(epoch,train_loss,label = "train")
axe1.plot(epoch,val_loss,label = "valid")
axe1.legend(loc = "best",fontsize=14)
axe1.set_xlabel("$epoch$",fontsize=14)
axe1.set_ylabel("$MSE loss$",fontsize=14)
axe1.set_title(name,fontsize=14)
def getTotal_decoded(training_decoded,valid_decoded,test_decoded,train_index,valid_index,test_index):
total_decoded = np.zeros((nTotal,nNodes,2))
for i in range(len(train_index)):
total_decoded[int(train_index[i]),:,0] = training_decoded.cpu().detach().numpy()[i,:,0]
total_decoded[int(train_index[i]),:,1] = training_decoded.cpu().detach().numpy()[i,:,1]
for i in range(len(valid_index)):
total_decoded[int(valid_index[i]),:,0] = valid_decoded.cpu().detach().numpy()[i,:,0]
total_decoded[int(valid_index[i]),:,1] = valid_decoded.cpu().detach().numpy()[i,:,1]
for i in range(len(test_index)):
total_decoded[int(test_index[i]),:,0] = test_decoded.cpu().detach().numpy()[i,:,0]
total_decoded[int(test_index[i]),:,1] = test_decoded.cpu().detach().numpy()[i,:,1]
return total_decoded
def getMSELoss(pathName):
epoch = pd.read_csv(pathName,usecols=[0]).values
train_loss = pd.read_csv(pathName,usecols=[1]).values
val_loss = pd.read_csv(pathName,usecols=[2]).values
return train_loss,val_loss,epoch
def index_split(train_ratio, valid_ratio, test_ratio, total_num):
if train_ratio + valid_ratio + test_ratio != 1:
raise ValueError("Three input ratio should sum to be 1!")
total_index = np.arange(total_num)
rng = np.random.default_rng()
total_index = rng.permutation(total_index)
knot_1 = int(total_num * train_ratio)
knot_2 = int(total_num * valid_ratio) + knot_1
train_index, valid_index, test_index = np.split(total_index, [knot_1, knot_2])
return train_index, valid_index, test_index
# + id="JhlQ5u1qCM4z" executionInfo={"status": "ok", "timestamp": 1629392226464, "user_tz": -60, "elapsed": 533, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}}
path_train = "/content/gdrive/MyDrive/Cola-Notebooks/FYP/YF/"+"new_FPC_train_index.csv"
path_valid = "/content/gdrive/MyDrive/Cola-Notebooks/FYP/YF/"+"new_FPC_valid_index.csv"
path_test = "/content/gdrive/MyDrive/Cola-Notebooks/FYP/YF/"+"new_FPC_test_index.csv"
# saveIndex(path_train, path_valid, path_test,train_index, valid_index, test_index)
# + colab={"base_uri": "https://localhost:8080/"} id="cSGMwBIACNyk" executionInfo={"status": "ok", "timestamp": 1629392280731, "user_tz": -60, "elapsed": 2615, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="4ae32151-7944-454c-d47c-2fdeaa3062d0"
# Load the train_index, valid_index and test_index
train_index,valid_index,test_index= getIndex(path_train,path_valid,path_test)
print(test_index)
# + [markdown] id="rMkDfCiOBcNa"
# ## load data
# + colab={"base_uri": "https://localhost:8080/"} id="PBNpFB2vC5gb" executionInfo={"status": "ok", "timestamp": 1629392293414, "user_tz": -60, "elapsed": 558, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="ac548529-7517-46ab-ae74-75f615a441ee"
os.chdir('/content/gdrive/MyDrive/Cola-Notebooks/FYP/YF')
print(os.getcwd())
# read in the data (1000 csv files)
nTrain = 1600
nValid = 200
nTest = 200
nTotal = nTrain + nValid + nTest
nNodes = 20550 # should really work this out
# The below method to load data is too slow. Therefore, we use load pt file
# [:, :, 2] is speed, [:, :, 3] is u, [:, :, 4] is v
# (speed not really needed)
# [:, :, 0] and [:, :, 1] are the SFC orderings
# training_data = np.zeros((nTrain,nNodes,5))
# for i in range(nTrain):
# data = np.loadtxt('csv_data/data_' +str(int(train_index[i]))+ '.csv', delimiter=',')
# training_data[i,:,:] = data
# training_data = np.array(training_data)
# print('size training data', training_data.shape)
# valid_data = np.zeros((nValid,nNodes,5))
# for i in range(nValid):
# data = np.loadtxt('csv_data/data_' +str(int(valid_index[i]))+ '.csv', delimiter=',')
# valid_data[i,:,:] = data
# valid_data = np.array(valid_data)
# print('size validation data', valid_data.shape)
# test_data = np.zeros((nTest,nNodes,5))
# for i in range(nTest):
# data = np.loadtxt('csv_data/data_' +str(int(test_index[i]))+ '.csv', delimiter=',')
# test_data[i,:,:] = data
# test_data = np.array(test_data)
# print('size test data', test_data.shape)
# total_data = np.zeros((nTotal,nNodes,5))
# for i in range(len(train_index)):
# total_data[int(train_index[i]),:,:] = training_data[i,:,:]
# for i in range(len(valid_index)):
# total_data[int(valid_index[i]),:,:] = valid_data[i,:,:]
# for i in range(len(test_index)):
# total_data[int(test_index[i]),:,:] = test_data[i,:,:]
# print('size total data', total_data.shape)
# + id="eGLp9fDhf8He"
# Before we save the pt file, we must load the data according to the above method
# torch.save(training_data, '/content/gdrive/MyDrive/FPC_new_random_train.pt')
# torch.save(valid_data, '/content/gdrive/MyDrive/FPC_new_random_valid.pt')
# torch.save(test_data, '/content/gdrive/MyDrive/FPC_new_random_test.pt')
# torch.save(total_data, '/content/gdrive/MyDrive/FPC_new_random_total.pt')
# + id="QdJ9D-6ykNjw" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629392389892, "user_tz": -60, "elapsed": 75271, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="4991ba34-8cd0-4dbb-ee87-fb393b86c652"
# load the data, this method save the time
training_data = torch.load('/content/gdrive/MyDrive/FPC_new_random_train.pt')
valid_data = torch.load('/content/gdrive/MyDrive/FPC_new_random_valid.pt')
test_data = torch.load('/content/gdrive/MyDrive/FPC_new_random_test.pt')
total_data = torch.load('/content/gdrive/MyDrive/FPC_new_random_total.pt')
print(training_data.shape)
print(valid_data.shape)
print(test_data.shape)
print(total_data.shape)
# + id="YXRsMOCwUxwU" executionInfo={"status": "ok", "timestamp": 1629392392966, "user_tz": -60, "elapsed": 3077, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}}
# rescale the data so that u and v data lies in the range [-1,1] (and speed in [0,1])
ma = np.max(training_data[:, :, 2])
mi = np.min(training_data[:, :, 2])
k = 1./(ma - mi)
b = 1 - k*ma
# this won't be used
training_data[:, :, 2] = k * training_data[:, :, 2] + b #- b
ma = np.max(training_data[:, :, 3])
mi = np.min(training_data[:, :, 3])
ku = 2./(ma - mi)
bu = 1 - ku*ma
training_data[:, :, 3] = ku * training_data[:, :, 3] + bu
valid_data[:, :, 3] = ku * valid_data[:, :, 3] + bu
test_data[:, :, 3] = ku * test_data[:, :, 3] + bu
total_data[:, :, 3] = ku * total_data[:, :, 3] + bu
ma = np.max(training_data[:, :, 4])
mi = np.min(training_data[:, :, 4])
kv = 2./(ma - mi)
bv = 1 - kv*ma
training_data[:, :, 4] = kv * training_data[:, :, 4] + bv
valid_data[:, :, 4] = kv * valid_data[:, :, 4] + bv
test_data[:, :, 4] = kv * test_data[:, :, 4] + bv
total_data[:, :, 4] = kv * total_data[:, :, 4] + bv
# + [markdown] id="VGN-qendDFff"
# ## Network architetcure
# + id="yrCzaq9PDJnx" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629392392967, "user_tz": -60, "elapsed": 5, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="d90a9530-8b80-4e28-ee83-85bb7060dbbc"
# SFC-CAE: one curve with nearest neighbour smoothing and compressing to 16 latent variables
print("compress to 16")
Latent_num = 16
torch.manual_seed(42)
# Hyper-parameters
EPOCH = 2001
BATCH_SIZE = 16
LR = 0.0001
k = nNodes # number of nodes - this has to match training_data.shape[0]
print(training_data.shape) # nTrain by number of nodes by 5
# Data Loader for easy mini-batch return in training
train_loader = Data.DataLoader(dataset = training_data, batch_size = BATCH_SIZE, shuffle = True)
# + id="imijLVOsDN1h" executionInfo={"status": "ok", "timestamp": 1629392393958, "user_tz": -60, "elapsed": 995, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}}
# Standard
class CNN_1(nn.Module):
def __init__(self):
super(CNN_1, self).__init__()
self.encoder_h1 = nn.Sequential(
# input shape (16,4,20550) # The first 16 is the batch size
nn.Tanh(),
nn.Conv1d(4, 8, 16, 4, 9),
# output shape (16, 8, 5139)
nn.Tanh(),
nn.Conv1d(8, 8, 16, 4, 9),
# output shape (16, 8,1286)
nn.Tanh(),
nn.Conv1d(8, 16, 16, 4, 9),
# output shape (16,16,323)
nn.Tanh(),
nn.Conv1d(16, 16, 16, 4, 9),
# output shape (16, 16, 82)
nn.Tanh(),
)
self.fc1 = nn.Sequential(
nn.Linear(16*82, 16),
nn.Tanh(),
)
self.fc2 = nn.Sequential(
nn.Linear(16, 16*82),
nn.Tanh(),
)
self.decoder_h1 = nn.Sequential(
# (16, 16, 82)
nn.Tanh(),
nn.ConvTranspose1d(16, 16, 17, 4, 9), # (16, 16, 323)
nn.Tanh(),
nn.ConvTranspose1d(16, 8, 16, 4, 9), # (16, 8, 1286)
nn.Tanh(),
nn.ConvTranspose1d(8, 8, 17, 4, 9), # (16, 8, 5139)
nn.Tanh(),
nn.ConvTranspose1d(8, 4, 16, 4, 9), # (16, 4, 20550)
nn.Tanh(),
)
# input sparse layers, initialize weight as 0.33, bias as 0
self.weight1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight1_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight1_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias1 = torch.nn.Parameter(torch.FloatTensor(torch.zeros(k)),requires_grad = True)
self.weight11 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight11_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight11_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias11 = torch.nn.Parameter(torch.FloatTensor(torch.zeros(k)),requires_grad = True)
self.weight2 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight2_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight2_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias2 = torch.nn.Parameter(torch.FloatTensor(torch.zeros(k)),requires_grad = True)
self.weight22 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight22_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight22_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias22 = torch.nn.Parameter(torch.FloatTensor(torch.zeros(k)),requires_grad = True)
self.weight3 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight3_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight3_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias3 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.zeros(k)),requires_grad = True)
self.weight33 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight33_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight33_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias33 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.zeros(k)),requires_grad = True)
self.weight4 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight4_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight4_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias4 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.zeros(k)),requires_grad = True)
self.weight44 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight44_0 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.weight44_1 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.ones(k)),requires_grad = True)
self.bias44 = torch.nn.Parameter(torch.FloatTensor(0.33 * torch.zeros(k)),requires_grad = True)
# output sparse layers, initialize weight as 0.083, bias as 0
self.weight_out1 = torch.nn.Parameter(torch.FloatTensor(0.083 *torch.ones(k)),requires_grad = True)
self.weight_out1_0 = torch.nn.Parameter(torch.FloatTensor(0.083* torch.ones(k)),requires_grad = True)
self.weight_out1_1 = torch.nn.Parameter(torch.FloatTensor(0.083* torch.ones(k)),requires_grad = True)
self.weight_out11 = torch.nn.Parameter(torch.FloatTensor(0.083 *torch.ones(k)),requires_grad = True)
self.weight_out11_0 = torch.nn.Parameter(torch.FloatTensor(0.083* torch.ones(k)),requires_grad = True)
self.weight_out11_1 = torch.nn.Parameter(torch.FloatTensor(0.083* torch.ones(k)),requires_grad = True)
self.weight_out2 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out2_0 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out2_1 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out22 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out22_0 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out22_1 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out3 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out3_0 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out3_1 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out33 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out33_0 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out33_1 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out4 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out4_0= torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out4_1 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out44 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out44_0= torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.weight_out44_1 = torch.nn.Parameter(torch.FloatTensor(0.083 * torch.ones(k)),requires_grad = True)
self.bias_out1 = torch.nn.Parameter(torch.FloatTensor(torch.zeros(k)),requires_grad = True)
self.bias_out2 = torch.nn.Parameter(torch.FloatTensor(torch.zeros(k)),requires_grad = True)
def forward(self, x):
# print("X_size",x.size())
# first curve
ToSFC1 = x[:, :, 0] # The first column is the first SFC ordering
ToSFC1Up = torch.zeros_like(ToSFC1)
ToSFC1Down = torch.zeros_like(ToSFC1)
ToSFC1Up[:-1] = ToSFC1[1:]
ToSFC1Up[-1] = ToSFC1[-1]
ToSFC1Down[1:] = ToSFC1[:-1]
ToSFC1Down[0] = ToSFC1[0]
batch_num = ToSFC1.shape[0]
x1 = x[:, :, 3:5] # The fourth column and fifth column are velocities u and v respectively
#print("x1", x1.shape) # # (16, 20550, 2)
x1_1d = torch.zeros((batch_num, 4, k)).to(device)
# first input sparse layer, then transform to sfc order1
for j in range(batch_num):
x1_1d[j, 0, :] = x1[j, :, 0][ToSFC1[j].long()] * self.weight1 + \
x1[j, :, 0][ToSFC1Up[j].long()] * self.weight1_0 + \
x1[j, :, 0][ToSFC1Down[j].long()] * self.weight1_1 + self.bias1
x1_1d[j, 1, :] = x1[j, :, 0][ToSFC1[j].long()] * self.weight11 + \
x1[j, :, 0][ToSFC1Up[j].long()] * self.weight11_0 + \
x1[j, :, 0][ToSFC1Down[j].long()] * self.weight11_1 + self.bias11
x1_1d[j, 2, :] = x1[j, :, 1][ToSFC1[j].long()] * self.weight2 + \
x1[j, :, 1][ToSFC1Up[j].long()] * self.weight2_0 + \
x1[j, :, 1][ToSFC1Down[j].long()] * self.weight2_1 + self.bias2
x1_1d[j, 3, :] = x1[j, :, 1][ToSFC1[j].long()] * self.weight22 + \
x1[j, :, 1][ToSFC1Up[j].long()] * self.weight22_0 + \
x1[j, :, 1][ToSFC1Down[j].long()] * self.weight22_1 + self.bias22
# first cnn encoder
encoded_1 = self.encoder_h1(x1_1d.view(-1, 4, k)) #(16,4,20550)
# print("encoded", encoded_1.shape)
# flatten and concatenate
encoded_3 = encoded_1.view(-1,16*82)
# print("Before FC", encoded_3.shape)
# fully connection
encoded = self.fc1(encoded_3) # (b,64)
# print("After encoder FC,the output of encoder",encoded.shape)
decoded_3 = self.decoder_h1(self.fc2(encoded).view(-1, 16, 82))
# print("The output of decoder: ", decoded_3.shape)
BackSFC1 = torch.argsort(ToSFC1)
BackSFC1Up = torch.argsort(ToSFC1Up)
BackSFC1Down = torch.argsort(ToSFC1Down)
decoded_sp = torch.zeros((batch_num, k, 2)).to(device)
# output sparse layer, resort according to sfc transform
for j in range(batch_num):
decoded_sp[j, :, 0] = decoded_3[j, 0, :][BackSFC1[j].long()]* self.weight_out1 + \
decoded_3[j, 0, :][BackSFC1Up[j].long()] * self.weight_out1_0 + \
decoded_3[j, 0, :][BackSFC1Down[j].long()] * self.weight_out1_1 + \
decoded_3[j, 1, :][BackSFC1[j].long()]* self.weight_out11 + \
decoded_3[j, 1, :][BackSFC1Up[j].long()] * self.weight_out11_0 + \
decoded_3[j, 1, :][BackSFC1Down[j].long()] * self.weight_out11_1 + self.bias_out1
decoded_sp[j, :, 1] = decoded_3[j, 2, :][BackSFC1[j].long()] * self.weight_out3 + \
decoded_3[j, 2, :][BackSFC1Up[j].long()] * self.weight_out3_0 + \
decoded_3[j, 2, :][BackSFC1Down[j].long()] * self.weight_out3_1 + \
decoded_3[j, 3, :][BackSFC1[j].long()] * self.weight_out33 + \
decoded_3[j, 3, :][BackSFC1Up[j].long()] * self.weight_out33_0 + \
decoded_3[j, 3, :][BackSFC1Down[j].long()] * self.weight_out33_1 + self.bias_out2
# resort 1D to 2D
decoded = F.tanh(decoded_sp) # both are BATCH_SIZE by nNodes by 2
return encoded, decoded
# + [markdown] id="SiZOP8Z3EqyY"
# ## Train
# + colab={"base_uri": "https://localhost:8080/"} id="7AK1uDOcEp2K" executionInfo={"status": "ok", "timestamp": 1627460693610, "user_tz": -60, "elapsed": 32653110, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "12928489296811262671"}} outputId="59e688af-7cb7-4553-8c74-898c113bbab7"
# train the autoencoder
t_train_0 = time.time()
autoencoder = CNN_1().to(device)
optimizer = torch.optim.Adam(autoencoder.parameters(), lr=LR)
loss_func = nn.MSELoss()
loss_list = []
loss_valid = []
epoch_list=[]
for epoch in range(EPOCH):
for step, x in enumerate(train_loader):
#print("x", x.shape)
b_y = x[:, :, 3:5].to(device)
b_x = x.to(device) # bx: False x: False
#print("b_y",b_y.shape)
encoded, decoded = autoencoder(b_x.float()) #decoded true by:False
loss = loss_func(decoded, b_y.float()) #Loss: True # mean square error
optimizer.zero_grad() # clear gradients for this training step
loss.backward() # backpropagation, compute gradients
optimizer.step() # apply gradients
loss_list.append(loss)
encoded, decoded = autoencoder(torch.tensor(valid_data).to(device))
error_autoencoder = (decoded.detach() - torch.tensor(valid_data[:,:, 3:5]).to(device))
MSE_valid = (error_autoencoder**2).mean()
loss_valid.append(MSE_valid)
epoch_list.append(epoch)
print('Epoch: ', epoch, '| train loss: %.6f' % loss.cpu().data.numpy(), '| valid loss: %.6f' % MSE_valid)
#save the weights every 500 epochs
if (epoch%500 == 0):
torch.save(autoencoder, "./SFC_CAE/pkl/II_Eran"+str(epoch) +"_LV"+str(Latent_num)+ "_B"+str(BATCH_SIZE)+"_n"+ str(nTrain)+"_L"+str(LR)+".pkl")
pathcsv= "./SFC_CAE/csv/II_Eran"+str(epoch)+"_LV"+str(Latent_num) + "_B"+str(BATCH_SIZE)+"_n"+ str(nTrain)+"_L"+str(LR)+".csv"
saveCsv(pathcsv,epoch+1)
t_train_1 = time.time()
# torch.save(autoencoder, path)
# + [markdown] id="7cgJFouPFHjB"
# ## Save and Plot loss
# + id="aliDL6J_HpaU" colab={"base_uri": "https://localhost:8080/", "height": 467} executionInfo={"status": "ok", "timestamp": 1629392537749, "user_tz": -60, "elapsed": 3613, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="490c66ca-1d64-4479-94e1-c6672199e971"
pathName = "./SFC_CAE/csv/II_Eran2000_LV16_B16_n1600_L0.0001.csv"
name = "SFC-CAE MSE loss of 16 compression variables"
PlotMSELoss(pathName,name)
# + id="lhLg1LThNLvv" executionInfo={"status": "ok", "timestamp": 1629392601925, "user_tz": -60, "elapsed": 16241, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}}
autoencoder = torch.load("./SFC_CAE/pkl/II_Eran2000_LV16_B16_n1600_L0.0001.pkl")
# + [markdown] id="heF699Yka3Nb"
# ## MSE
# + id="t8Hj5BVjFM5E" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629392832447, "user_tz": -60, "elapsed": 222866, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="048bf08e-c1a6-4fdd-8b28-86e8a0f4c6bd"
# pass training, validation and test data through the autoencoder
t_predict_0 = time.time()
mode_1train, training_decoded = autoencoder.to(device)(torch.tensor(training_data).to(device))
error_autoencoder = (training_decoded.cpu().detach().numpy() - training_data[:,:,3:5])
print("MSE_err of training data", (error_autoencoder**2).mean())
mode_1valid, valid_decoded = autoencoder.to(device)(torch.tensor(valid_data).to(device))
error_autoencoder = (valid_decoded.cpu().detach().numpy() - valid_data[:, :, 3:5])
print("Mse_err of validation data", (error_autoencoder**2).mean())
mode_1test, test_decoded = autoencoder.to(device)(torch.tensor(test_data).to(device))
error_autoencoder = (test_decoded.cpu().detach().numpy() - test_data[:, :, 3:5])
print("Mse_err of test data", (error_autoencoder**2).mean())
t_predict_1 = time.time()
total_decoded = getTotal_decoded(training_decoded,valid_decoded,test_decoded,train_index,valid_index,test_index)
error_autoencoder = (total_decoded - total_data[:, :, 3:5])
print("Mse_err of total data", (error_autoencoder**2).mean())
print(mode_1train.shape)
print(mode_1valid.shape)
print(mode_1test.shape)
print('Predict time:',t_predict_1-t_predict_0)
# + colab={"base_uri": "https://localhost:8080/"} id="Vnk8GoE9UXjJ" executionInfo={"status": "ok", "timestamp": 1629392894221, "user_tz": -60, "elapsed": 2735, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="7bd068e1-961b-4aff-a4cb-78e7677b8e81"
# Save the mode of the training, valid and test data. They will be used in hierarchical autoencoder
Latent_num = 16
torch.manual_seed(42)
BATCH_SIZE = 16
LR = 0.0001
nTrain = 1600
path_train = "./HAE/mode_new/II_mode1_LV"+str(Latent_num)+"_Eran"+str(2000) + "_B"+str(BATCH_SIZE)+"_n"+ str(nTrain)+"_L"+str(LR)+"_train.csv"
path_valid = "./HAE/mode_new/II_mode1_LV"+str(Latent_num)+"_Eran"+str(2000) + "_B"+str(BATCH_SIZE)+"_n"+ str(nTrain)+"_L"+str(LR)+"_valid.csv"
path_test = "./HAE/mode_new/II_mode1_LV"+str(Latent_num)+"_Eran"+str(2000) + "_B"+str(BATCH_SIZE)+"_n"+ str(nTrain)+"_L"+str(LR)+"_test.csv"
print(path_train)
saveMode(path_train,path_valid,path_test,mode_1train,mode_1valid,mode_1test)
# + colab={"base_uri": "https://localhost:8080/"} id="hOSvd19AU3Wg" executionInfo={"status": "ok", "timestamp": 1629392895410, "user_tz": -60, "elapsed": 574, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "08129041439789093783"}} outputId="3455df6b-f125-4f42-f064-a43bbb1d0d69"
mode_1train,mode_1valid,mode_1test = getMode(path_train,path_valid,path_test)
mode_1train = torch.from_numpy(mode_1train).to(device)
mode_1valid = torch.from_numpy(mode_1valid).to(device)
mode_1test = torch.from_numpy(mode_1test).to(device)
print(mode_1train.shape)
print(mode_1test.shape)
print(mode_1valid.shape)
print(mode_1valid)
# + [markdown] id="NkgzWnUHKz9k"
# ## Convert csv to vtu
# + id="M-UjkUT97pOM"
# Before convert csv file to vtu file, the range of data must be recovered
training_decoded[:, :, 0] = (training_decoded[:, :, 0] - bu)/ku
valid_decoded[:, :, 0] = (valid_decoded[:, :, 0] - bu)/ku
test_decoded[:, :, 0] = (test_decoded[:, :, 0] - bu)/ku
total_decoded[:, :, 0] = (total_decoded[:, :, 0] - bu)/ku
training_decoded[:, :, 1] = (training_decoded[:, :, 1] - bv)/kv
valid_decoded[:, :, 1] = (valid_decoded[:, :, 1] - bv)/kv
test_decoded[:, :, 1] = (test_decoded[:, :, 1] - bv)/kv
total_decoded[:, :, 1] = (total_decoded[:, :, 1] - bv)/kv
training_data[:, :, 3] = (training_data[:, :, 3] - bu)/ku
valid_data[:, :, 3] = (valid_data[:, :, 3] - bu)/ku
test_data[:, :, 3] = (test_data[:, :, 3] - bu)/ku
total_data[:, :, 3] = (total_data[:, :, 3] - bu)/ku
training_data[:, :, 4] = (training_data[:, :, 4] - bv)/kv
valid_data[:, :, 4] = (valid_data[:, :, 4] - bv)/kv
test_data[:, :, 4] = (test_data[:, :, 4] - bv)/kv
total_data[:, :, 4] = (total_data[:, :, 4] - bv)/kv
# + colab={"base_uri": "https://localhost:8080/"} id="LCMYEjFPK4iZ" executionInfo={"status": "ok", "timestamp": 1626611379200, "user_tz": -60, "elapsed": 43501, "user": {"displayName": "\u6768\u9492", "photoUrl": "", "userId": "12928489296811262671"}} outputId="5721f4d0-9d1d-43b2-aa5b-7bccd804619e"
# results = np.concatenate((training_decoded.cpu().data.numpy(), valid_decoded.cpu().data.numpy(), test_decoded.cpu().data.numpy()))
results = total_decoded
print('results shape', results.shape)
N = results.shape[1] * results.shape[2]
results = results.reshape((results.shape[0],N), order='F')
print('results shape', results.shape, type(results))
# The path can be defined by user depending on the requirements
path = "./SFC_CAE/CAE_II"+"_LV"+str(Latent_num) + "_B"+str(BATCH_SIZE)+'E_'+str(3000)+"_result.csv"
# write results to file
np.savetxt(path, results , delimiter=',')
| fpc_methods/SFC_CAE/SFC_CAE ipynbs/16_FPC_II16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Image denoising using dictionary learning
#
#
# An example comparing the effect of reconstructing noisy fragments
# of a raccoon face image using firstly online `DictionaryLearning` and
# various transform methods.
#
# The dictionary is fitted on the distorted left half of the image, and
# subsequently used to reconstruct the right half. Note that even better
# performance could be achieved by fitting to an undistorted (i.e.
# noiseless) image, but here we start from the assumption that it is not
# available.
#
# A common practice for evaluating the results of image denoising is by looking
# at the difference between the reconstruction and the original image. If the
# reconstruction is perfect this will look like Gaussian noise.
#
# It can be seen from the plots that the results of `omp` with two
# non-zero coefficients is a bit less biased than when keeping only one
# (the edges look less prominent). It is in addition closer from the ground
# truth in Frobenius norm.
#
# The result of `least_angle_regression` is much more strongly biased: the
# difference is reminiscent of the local intensity value of the original image.
#
# Thresholding is clearly not useful for denoising, but it is here to show that
# it can produce a suggestive output with very high speed, and thus be useful
# for other tasks such as object classification, where performance is not
# necessarily related to visualisation.
#
#
#
# +
print(__doc__)
from time import time
import matplotlib.pyplot as plt
import numpy as np
import scipy as sp
from sklearn.decomposition import MiniBatchDictionaryLearning
from sklearn.feature_extraction.image import extract_patches_2d
from sklearn.feature_extraction.image import reconstruct_from_patches_2d
try: # SciPy >= 0.16 have face in misc
from scipy.misc import face
face = face(gray=True)
except ImportError:
face = sp.face(gray=True)
# Convert from uint8 representation with values between 0 and 255 to
# a floating point representation with values between 0 and 1.
face = face / 255.
# downsample for higher speed
face = face[::4, ::4] + face[1::4, ::4] + face[::4, 1::4] + face[1::4, 1::4]
face /= 4.0
height, width = face.shape
# Distort the right half of the image
print('Distorting image...')
distorted = face.copy()
distorted[:, width // 2:] += 0.075 * np.random.randn(height, width // 2)
# Extract all reference patches from the left half of the image
print('Extracting reference patches...')
t0 = time()
patch_size = (7, 7)
data = extract_patches_2d(distorted[:, :width // 2], patch_size)
data = data.reshape(data.shape[0], -1)
data -= np.mean(data, axis=0)
data /= np.std(data, axis=0)
print('done in %.2fs.' % (time() - t0))
# #############################################################################
# Learn the dictionary from reference patches
print('Learning the dictionary...')
t0 = time()
dico = MiniBatchDictionaryLearning(n_components=100, alpha=1, n_iter=500)
V = dico.fit(data).components_
dt = time() - t0
print('done in %.2fs.' % dt)
plt.figure(figsize=(4.2, 4))
for i, comp in enumerate(V[:100]):
plt.subplot(10, 10, i + 1)
plt.imshow(comp.reshape(patch_size), cmap=plt.cm.gray_r,
interpolation='nearest')
plt.xticks(())
plt.yticks(())
plt.suptitle('Dictionary learned from face patches\n' +
'Train time %.1fs on %d patches' % (dt, len(data)),
fontsize=16)
plt.subplots_adjust(0.08, 0.02, 0.92, 0.85, 0.08, 0.23)
# #############################################################################
# Display the distorted image
def show_with_diff(image, reference, title):
"""Helper function to display denoising"""
plt.figure(figsize=(5, 3.3))
plt.subplot(1, 2, 1)
plt.title('Image')
plt.imshow(image, vmin=0, vmax=1, cmap=plt.cm.gray,
interpolation='nearest')
plt.xticks(())
plt.yticks(())
plt.subplot(1, 2, 2)
difference = image - reference
plt.title('Difference (norm: %.2f)' % np.sqrt(np.sum(difference ** 2)))
plt.imshow(difference, vmin=-0.5, vmax=0.5, cmap=plt.cm.PuOr,
interpolation='nearest')
plt.xticks(())
plt.yticks(())
plt.suptitle(title, size=16)
plt.subplots_adjust(0.02, 0.02, 0.98, 0.79, 0.02, 0.2)
show_with_diff(distorted, face, 'Distorted image')
# #############################################################################
# Extract noisy patches and reconstruct them using the dictionary
print('Extracting noisy patches... ')
t0 = time()
data = extract_patches_2d(distorted[:, width // 2:], patch_size)
data = data.reshape(data.shape[0], -1)
intercept = np.mean(data, axis=0)
data -= intercept
print('done in %.2fs.' % (time() - t0))
transform_algorithms = [
('Orthogonal Matching Pursuit\n1 atom', 'omp',
{'transform_n_nonzero_coefs': 1}),
('Orthogonal Matching Pursuit\n2 atoms', 'omp',
{'transform_n_nonzero_coefs': 2}),
('Least-angle regression\n5 atoms', 'lars',
{'transform_n_nonzero_coefs': 5}),
('Thresholding\n alpha=0.1', 'threshold', {'transform_alpha': .1})]
reconstructions = {}
for title, transform_algorithm, kwargs in transform_algorithms:
print(title + '...')
reconstructions[title] = face.copy()
t0 = time()
dico.set_params(transform_algorithm=transform_algorithm, **kwargs)
code = dico.transform(data)
patches = np.dot(code, V)
patches += intercept
patches = patches.reshape(len(data), *patch_size)
if transform_algorithm == 'threshold':
patches -= patches.min()
patches /= patches.max()
reconstructions[title][:, width // 2:] = reconstruct_from_patches_2d(
patches, (height, width // 2))
dt = time() - t0
print('done in %.2fs.' % dt)
show_with_diff(reconstructions[title], face,
title + ' (time: %.1fs)' % dt)
plt.show()
| 01 Machine Learning/scikit_examples_jupyter/decomposition/plot_image_denoising.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Mother:
def __init__(self):
self.name = "Manju"
def Print(self):
print("Print of Mother called")
class Father:
def __init__(self):
self.name = "Ajay"
def Print(self):
print("Print of Father called")
class Child(Father,Mother): # the order in which we have called , print will be in that order.. here father has been called first so father's print will be called first
# def __init__(self,name):
# self.name = name
def __init__(self):
super().__init__()
def PrintChild(self):
print("Name of child: ",self.name)
# -
# c = Child("Pramit")
c = Child()
c.PrintChild()
# +
# c = Child("pramit")
# c.Print()
# -
| 02.Data-Structures-and-Algorithms/05.OOPS-2/07.Multiple-inheritance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **iSSVD** is the algorithm developed for 'Robust Intergrative Biclustering for Multi-view Data'. It is based on sparse singular value decompostion (Lee et al. 2010) and stability selection (Meinshausen and Bühlmann, 2010 and Sill et al., 2011).
#
# This is a simple guide to show how to use the python package 'iSSVD'.
import pandas as pd
import numpy as np
from issvd_functions import issvd, issvd_diagnostics, gen_sim_vec, gen_tmp
from numpy.random import seed
# First generate example data. Here we generate one group of data that contains two views, each has dimension $200\times1,000$. We have four artificical biclusters in the data while each biclsuter contains $50$ samples and $100+100$ (100 for each view) variables. There is no overlaps between biclsuters. The noise level $\sigma$ is 0.1.
# +
seed(20)
data, rows, cols = gen_sim_vec(n=200,p=1000,D=2,rowsize=50, colsize=100,
numbers=1,sigma=0.1,nbicluster=4, orthonm=False)
df = data[0]
# -
# Then we run iSSVD.
res = issvd(X=df,standr=False,pointwise=True,steps=100,size=0.5,
vthr = 0.9,ssthr=[0.6,0.65],nbicluster=10,rows_nc=False,cols_nc=False,col_overlap=False
,row_overlap=False,pceru=0.1,pcerv=0.1,merr=0.0001,iters=100)
# We compare the biclusters with the original ones. The intermediate step is to convert the true biclsuter indices to the format identified by the diagnostic function.
# +
# Bicluster samples identified by iSSVD
Rows = res['Sample_index']
# Bicluster variables identified by iSSVD
Cols = res['Variable_index']
# True bicluster samples and variables
row_ind = rows[0]
col_ind = cols[0]
res1tmp, res2tmp = gen_tmp(Rows,Cols, row_ind, col_ind,n=200,p=1000,D=2)
rev, rel, f, fp, fn = issvd_diagnostics(res1tmp,res2tmp,row_ind,col_ind)
# -
print(f"Average Revovery is {rev.round(4)}.")
print(f"Average Relevance is {rel.round(4)}.")
print(f"F-score is {f.round(4)}.")
print(f"False positive rate is {fp.round(4)}.")
print(f"False negative rate is {fn.round(4)}.")
# Next, we visualize the biclusters.
import seaborn as sns
import matplotlib.pyplot as plt
# +
# Before clustered
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(15,3))
sns.heatmap(df[0],ax=axes[0])
sns.heatmap(df[0],ax=axes[1])
# +
# After Clustered
new_df = []
for d in range(2):
cs = []
d=0
col = np.array([],dtype=int)
for i in range(4):
r1 = df[d][Rows[i],:]
col = np.append(col,Cols[i][d])
c1 = r1[:,col]
c2 = np.delete(r1, col, axis=1)
c3 = np.concatenate([c1,c2], axis=1)
cs.append(c3)
new_df.append(np.vstack(cs))
fign, axesn = plt.subplots(nrows=1, ncols=2, figsize=(15,3))
sns.heatmap(new_df[0],ax=axesn[0])
sns.heatmap(new_df[0],ax=axesn[1])
# -
# ! jupyter nbconvert --to html Guide.ipynb
| iSSVD/Guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Overview
# This notebook guides you through process of testing if the Jupyter Notebook server is authorized to access the Earth Engine servers, and provides a way to authorize the server, if needed.
# # Testing if the Jupyter Notebook server is authorized to access Earth Engine
#
# To begin, first verify that you can import the Earth Engine Python API package by running the following cell.
import ee
# Next, try to initialize the `ee` Python package.
try:
ee.Initialize()
print('The Earth Engine package initialized successfully!')
except ee.EEException as e:
print('The Earth Engine package failed to initialize!')
except:
print("Unexpected error:", sys.exc_info()[0])
raise
# If the initialization succeeded, you can stop here. Congratulations! If not, continue on below...
# # Authenticating to the Earth Engine servers
# If the initialization process failed, you will need to authenticate the Jupyter Notebook server so that it can communicate with the Earth Engine servers. You can initiate the authentication process by running the following command.
#
# *Note that `earthengine authenticate` is a system command (rather than a Python command), and the cell uses the [%%bash cell magic](http://ipython.readthedocs.io/en/stable/interactive/magics.html#cellmagic-bash) in the first line of the cell to indicate that the cell contents should be executed using a bash shell.*
# + language="bash"
# earthengine authenticate --quiet
# -
# Once you have obtained an authorization code from the previous step, paste the code into the following cell and run it.
# + language="bash"
# earthengine authenticate --authorization-code=PLACE_AUTH_CODE_HERE
# -
# # Removing authentication credentials
# Authentication credentials are stored as a file in the user's configuration directory. If you need to remove the authentication credentials, run the following cell.
# + language="bash"
# rm ~/.config/earthengine
| python/examples/ipynb/authorize_notebook_server.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A brief introduction to Species Distribution Models in Python
#
# *Predicting spatial distributions for ecological species leveraging Python's ever-strengthening machine learning stack.*
#
# ---
#
# **Author : [<NAME>](mailto:<EMAIL>)** | November 2020
#
# ---
#
# ***Species Distributions Models (SDMs)*** are an important and widely used tool for ecologists, agriculture scientists, conservation biologists, and many other geospatial science enthusiasts. While dozens of tutorials leverage the traditional R stack for SDMs, with packages such as Raster, implementations of SDMs in Python are, ***surprisingly***, rather limited. To bridge this gap, we explore a SDM workflow powered by Python's machine learning capabilities. The methods employed barely scratch the surface of available techniques, and hopefully this introduction can serve as a springboard to further exploration.
#
# If you are completely new to SDMs, it may be prudent to start <a target="_blank" rel="noopener noreferrer" href="https://www.google.com/search?q=species+distribution+modeling&oq=species+distribution+modeling&aqs=chrome..69i57j35i39j0l3j69i61l2j69i60.6284j0j1&sourceid=chrome&ie=UTF-8">here</a>. SDMs associate presence locations of a species to climate variables, giving you the power to predict species suitability across an entire landscape. First, environmental variables are sampled from presence coordinates. Second, a statistical model (here, SK-Learn classifiers) defines a species-environment relationship. Third, the species–environment relationship is mapped across the study space, denoting a potential distribution of the species (referred to as interpolation). Projecting to future/past climates or to novel geographic areas is referred to as extrapolation. A typical workflow is as follows:
#
# `conceptualization` -> `data pre-processing` -> `model training/assessment` ->
# `interpolate/extrapolate` -> `iterate`
#
# ## Tutorial Objectives
# ---
#
# 1. Create a SDM workspace with a Python codebase.
# 2. Run a suite of SDMs with your ML classifiers of choice.
# 3. Visualize model predictions with climate features (1970-2000).
# ## Section 1 | Set up
# ---
#
# ### 1.1 | Workspace
#
# The first step is to create `inputs`/`outputs` folders in our working file directory. It is best practices to keep the data and results separated, as `outputs` folder should be completely reproducible.
import os
os.mkdir("inputs")
os.mkdir("outputs")
# We now install the additional dependencies we will need for our SDMs, with four primary libraries:
#
# * ***scikit-learn***: De-facto Python machine learning
# * ***pyimpute***: spatial classification
# * ***rasterio***: reads and writes geospatial rasters
# * ***geopandas***: spatial operations in Python made easy
#
# These can be installed at the terminal using `pip install LIBRARY`, but you may find it cleaner to create a conda virtual environment from `requirements-py.txt` (see <a target="_blank" rel="noopener noreferrer" href="https://github.com/daniel-furman/py-sdms-tutorial">Git repo</a>).
# ### 1.2 | Data Processing
#
# We first need to download a geodatabase (here we use a `.shp` file) denoting presence/absence coordinates, which can be directly loaded into Python as a GeoPandas `GeoDataFrame` (a tabular data structure for geometric data types). Here, the `CLASS` column is a binary indication of the presence/absence of the species. For this tutorial, we are using Joshua trees (*Yucca brevifolia*) as the example species:
#
# <img src="jtree.jpg" width = 500/>
#
# To follow along chunk by chunk, clone the <a target="_blank" rel="noopener noreferrer" href="https://github.com/daniel-furman/py-sdms-tutorial">Git repo</a> and open `Intro-to-SDMs-Py.ipynb` in your working directory of choice.
import geopandas as gpd
import shutil
import glob
# grab jtree data after cloning Git repo
for f in sorted(glob.glob('data/jtree*')):
shutil.copy(f,'inputs/')
# or grab your data of choice and move to 'inputs/'
pa = gpd.GeoDataFrame.from_file("inputs/jtree.shp")
pa.sample(5) # GeoDataFrame for the species
# We now check that there are no duplicate or `NaN` coordinates, as well as inspect the shapefile's attributes.
print("number of duplicates: ", pa.duplicated(subset='geometry', keep='first').sum())
print("number of NA's: ", pa['geometry'].isna().sum())
print("Coordinate reference system is: {}".format(pa.crs))
print("{} observations with {} columns".format(*pa.shape))
# We can map the species presences (`pa==1`).
pa[pa.CLASS == 1].plot(marker='*', color='green', markersize=8)
# And we can map the background points (`pa == 0`).
pa[pa.CLASS == 0].plot(marker='+', color='black', markersize=4)
# However, if you don't have a geospatial database with presences/absence coordinates, there are some easy steps to create one for ***virtually any species*** of interest! You can start by searching the open-data Global Biodiversity Information Facility (<a target="_blank" rel="noopener noreferrer" href="https://www.gbif.org">GBIF</a>), downloading a species database to `.csv`, and migrating to R to pipe the database to `.shp` (e.g. see `data-pre-processing.R` in the <a target="_blank" rel="noopener noreferrer" href="https://github.com/daniel-furman/py-sdms-tutorial">Git repo</a> or the additional information section below).
# ### Section 2 | Mapping species suitability
# ---
#
# In this section we will train our machine learning classifiers and make spatial predictions of the species distribution over current conditions (1970-2000).
#
# First, we load 19 bioclimatic features (here we use 2.5 arc-minute resolution) from the publicly available [WorldClim database](https://www.worldclim.org) (v. 2.1, Fick & Hijmans, 2017).
# grab climate features - cropped to joshua tree study area
for f in sorted(glob.glob('data/bioclim/bclim*.asc')):
shutil.copy(f,'inputs/')
raster_features = sorted(glob.glob(
'inputs/bclim*.asc'))
# check number of features
print('\nThere are', len(raster_features), 'raster features.')
# We are now ready to use `pyimpute` to generate the raster maps of suitability. We first prep the pyimpute workflow:
from pyimpute import load_training_vector
from pyimpute import load_targets
train_xs, train_y = load_training_vector(pa, raster_features, response_field='CLASS')
target_xs, raster_info = load_targets(raster_features)
train_xs.shape, train_y.shape # check shape, does it match the size above of the observations?
# and we implemement several `scikit-learn` classifiers:
# +
# import machine learning classifiers
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from xgboost import XGBClassifier
from lightgbm import LGBMClassifier
CLASS_MAP = {
'rf': (RandomForestClassifier()),
'et': (ExtraTreesClassifier()),
'xgb': (XGBClassifier()),
'lgbm': (LGBMClassifier())
}
from pyimpute import impute
from sklearn import model_selection
# model fitting and spatial range prediction
for name, (model) in CLASS_MAP.items():
# cross validation for accuracy scores (displayed as a percentage)
k = 5 # k-fold
kf = model_selection.KFold(n_splits=k)
accuracy_scores = model_selection.cross_val_score(model, train_xs, train_y, cv=kf, scoring='accuracy')
print(name + " %d-fold Cross Validation Accuracy: %0.2f (+/- %0.2f)"
% (k, accuracy_scores.mean() * 100, accuracy_scores.std() * 200))
# spatial prediction
model.fit(train_xs, train_y)
os.mkdir('outputs/' + name + '-images')
impute(target_xs, model, raster_info, outdir='outputs/' + name + '-images',
class_prob=True, certainty=True)
# -
# All done! We have a `responses.tif` raster which is the predicted class (0 or 1) and `probability_1.tif` with a continuous suitability scale. Let's average the continuous output for the four models and plot our map.
# +
from pylab import plt
# define spatial plotter
def plotit(x, title, cmap="Blues"):
plt.imshow(x, cmap=cmap, interpolation='nearest')
plt.colorbar()
plt.title(title, fontweight = 'bold')
import rasterio
distr_rf = rasterio.open("outputs/rf-images/probability_1.0.tif").read(1)
distr_et = rasterio.open("outputs/et-images/probability_1.0.tif").read(1)
distr_xgb = rasterio.open("outputs/xgb-images/probability_1.0.tif").read(1)
distr_lgbm = rasterio.open("outputs/lgbm-images/probability_1.0.tif").read(1)
distr_averaged = (distr_rf + distr_et + distr_xgb + distr_lgbm)/4
plotit(distr_averaged, "Joshua Tree Range, averaged", cmap="Greens")
# -
# Lastly, let's zoom in to Joshua Tree National Park and inspect the suitability there.
plotit(distr_averaged[100:150, 100:150], "Joshua Tree National Park Suitability", cmap="Greens")
# ### Additional resources
# ---
# 1. <a target="_blank" rel="noopener noreferrer" href="https://cran.r-project.org/web/packages/dismo/vignettes/sdm.pdf">Species distribution modeling with R</a> (Hijmans and Elith, 2017)
# 2. Pyimpute's <a target="_blank" rel="noopener noreferrer" href="https://github.com/perrygeo/pyimpute/blob/master/README.md">README.md</a>
# 3. A study <a target="_blank" rel="noopener noreferrer" href="https://www.researchgate.net/publication/229149956_Selecting_Pseudo-Absences_for_Species_Distribution_Models_How_Where_and_How_Many">on generating pseudo absence points</a> (Barbet-Massin et al., 2012)
# 4. A study <a target="_blank" rel="noopener noreferrer" href="https://www.nature.com/articles/s41598-018-25437-1
# ">on SDM transferability and pixel size</a> (Manzoor et al., 2018)
# 5. A study <a target="_blank" rel="noopener noreferrer" href="https://onlinelibrary.wiley.com/doi/full/10.1111/ddi.13161">on SDMs for invasive species</a> (Lake et al., 2020)
# 6. A book <a target="_blank" rel="noopener noreferrer" href="https://www.amazon.com/Mapping-Species-Distributions-Biodiversity-Conservation/dp/0521700027
# ">on mapping SDMs </a> (Franklin, 2009)
# 7. A more modern <a target="_blank" rel="noopener noreferrer" href="https://damariszurell.github.io/SDM-Intro/"> SDMs tutorial</a> (Zurell, 2020)
# 8. A study <a target="_blank" rel="noopener noreferrer" href="https://onlinelibrary.wiley.com/doi/full/10.1111/j.1600-0587.2012.07348.x">on collinearity among model variables</a> (<NAME> et al., 2012)
#
#
#
#
# ### Data Citations
# ---
#
# 1. GBIF.org (01 November 2020) GBIF Occurrence' Download https://doi.org/10.15468/dl.g6swrm
# 2. <NAME>. and <NAME>, 2017. WorldClim 2: new 1km spatial resolution climate surfaces for global land areas. International Journal of Climatology 37 (12): 4302-4315.
| Python-sdm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # pygohome
#
# ## Python, Let's Go Home. Quickly.
#
# *pygohome* is a 100% personal route optimizer in a known environment based on experience.
#
# *You* walk/ride/drive frequently between known locations (home, work, school, shops, family, friends, …) using different routes, but would like to know the optimal route, that should take you the less time possible? *pygohome* uses your recorded GPX tracks to build a route network of *your* world with estimation on how long *you* need to get from A to B using the mean of transport of your choice.
# ## How it works
#
# 1. **Choose your mean of transport.** Bicycle works great, walking should too. Motorized vehicles may be served better by real-time online services.
#
# 2. **Track all your trips** using that mean of transport. Start the tracking before you leave, stop the tracking after you arrive. Ride/walk/drive normally, stop at lights, don't speed. OsmAnd works great (list of other apps needed). 1 or 2 seconds tracking interval is ok.
#
# 3. **Add POIs (and intersections) as waypoints.** Transfer all GPX files to your computer. Load all of them into some GPX editor (JOSM works great).
#
# 4. **Create a new GPX file with waypoints** and add all points of interest (home, work, or any place where you started, deliberately paused or ended a trip) with an attribute `name=…`. Then add unnamed waypoints on each intersection where your tracks cross, split, or join. This is the biggest future plan in *pygohome* to offer a good editor with checks for consistency. Now you have to do it manually.
#
# 5. **Run the cell below** and an empty map will appear. All the action happens here from now on.
#
# 6. **Load your GPX files** with the tracks AND the file with waypoints. The current limit is 10MB for one upload, so if you have more data, do it in more batches.
#
# 7. **Find the fastest route** where `src` and `dst` are the names of the POIs and `Quantile` is the a float number between `0.0` (take into account your best time on each road segment) and `1.0` (worst time), while `0.8` is considered a safe value.
from pygohome.jupygohome import gohome
gohome()
| notebooks/pygohome.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Ellipsoidal nested rejection sampling
#
# This example demonstrates how to use ellipsoidal nested rejection sampling [1] to sample from the posterior distribution for a logistic model fitted to model-simulated data. For a tutorial on nested sampling, see: [INSERT LINK to NESTED SAMPLING TUTORIAL NOTEBOOK].
#
# [1] "A nested sampling algorithm for cosmological model selection", <NAME>, <NAME> and <NAME>, [arXiv:astro-ph/0508461v2](https://arxiv.org/abs/astro-ph/0508461).
# First create fake data.
# +
from __future__ import print_function
import pints
import pints.toy as toy
import numpy as np
import matplotlib.pyplot as plt
# Load a forward model
model = toy.LogisticModel()
# Create some toy data
r = 0.015
k = 500
real_parameters = [r, k]
times = np.linspace(0, 1000, 100)
signal_values = model.simulate(real_parameters, times)
# Add independent Gaussian noise
sigma = 10
observed_values = signal_values + pints.noise.independent(sigma, signal_values.shape)
# Plot
plt.plot(times,signal_values,label = 'signal')
plt.plot(times,observed_values,label = 'observed')
plt.xlabel('Time')
plt.ylabel('Values')
plt.legend()
plt.show()
# -
# Create the nested sampler that will be used to sample from the posterior.
# +
# Create an object with links to the model and time series
problem = pints.SingleOutputProblem(model, times, observed_values)
# Create a log-likelihood function (adds an extra parameter!)
log_likelihood = pints.GaussianLogLikelihood(problem)
# Create a uniform prior over both the parameters and the new noise variable
log_prior = pints.UniformLogPrior(
[0.01, 400, sigma * 0.5],
[0.02, 600, sigma * 1.5])
# Create a nested ellipsoidal rejectection sampler
sampler = pints.NestedEllipsoidSampler(log_likelihood, log_prior)
# Set number of iterations
sampler.set_iterations(4000)
# Set enlargement factor (a value of 1 means uniformly sample from the minimum volume
# ellipsoid around sampled points, whereas > 1 widens the sampling volume)
sampler.set_enlargement_factor(1.25)
# Set number of initial rejection samples (before ellipsoidal sampling begins)
sampler.set_rejection_samples(200)
# Set the number of posterior samples to generate
sampler.set_posterior_samples(400)
# Set gaps between updating ellipsoid
sampler.set_ellipsoid_update_gap(100)
# Set the number of active points
sampler.set_active_points_rate(400)
sampler.set_log_to_screen(False)
# -
# Run the sampler!
samples, marginal_log_likelihood = sampler.run()
print('Done!')
# ## Plot posterior samples versus true parameter values (dashed lines)
# +
# Plot output
import pints.plot
pints.plot.histogram([samples], ref_parameters=[r, k, sigma])
plt.show()
# +
vTheta = samples[0]
pints.plot.pairwise(samples, kde=True)
plt.show()
# -
# ## Plot posterior predictive simulations versus the observed data
pints.plot.series(samples[:100], problem)
plt.show()
# ## Marginal likelihood estimate
print('marginal log-likelihood = ' + str(marginal_log_likelihood))
| examples/sampling-ellipsoidal-nested-rejection-sampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['MKL_NUM_THREADS'] = '1'
os.environ['OPENBLAS_NUM_THREADS'] = '1'
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import pymc3 as pm
import arviz as az
import matplotlib.pyplot as plt
import warnings
warnings.simplefilter('ignore')
# -
maize_data = pd.read_csv('../data/maize.csv')
maize_data.plot.scatter(x='TTsum_c', y='CropBiomassWt', s=50);
x, y = maize_data[['TTsum_c', 'CropBiomassWt']].values.T
x
maize_data.TTsum_c.values.reshape(-1,1)
# ## Gaussian Process
with pm.Model() as gp_maize_model:
# Lengthscale
ρ = pm.HalfCauchy('ρ', 5)
η = pm.HalfCauchy('η', 5)
M = pm.gp.mean.Linear(coeffs=(maize_data.CropBiomassWt/maize_data.TTsum_c).mean())
K = (η**2) * pm.gp.cov.ExpQuad(1, ρ)
Xu = pm.gp.util.kmeans_inducing_points(10, x.reshape(-1,1))
σ = pm.HalfNormal('σ', 50)
yield_gp = pm.gp.MarginalSparse(mean_func=M, cov_func=K, approx="FITC")
yield_gp.marginal_likelihood('CropBiomassWt', X=x.reshape(-1,1), Xu=Xu, y=y, noise=σ)
with gp_maize_model:
gp_trace = pm.sample(1000, tune=2000, cores=1, random_seed=42)
az.plot_trace(gp_trace, var_names=['ρ', 'η', 'σ']);
# +
X_pred = np.linspace(250, 2000, 100)
with gp_maize_model:
maize_pred = recruit_gp.conditional("maize_pred", X_pred.reshape(-1, 1))
gp_maize_samples = pm.sample_posterior_predictive(gp_trace, vars=[maize_pred], samples=3, random_seed=42)
# -
ax = maize_data.plot.scatter(x='TTsum_c', y='CropBiomassWt', c='k', s=50)
ax.set_ylim(0, None)
for x in gp_maize_samples['maize_pred']:
ax.plot(X_pred, x);
with gp_maize_model:
gp_maize_samples = pm.sample_posterior_predictive(gp_trace, vars=[maize_pred], samples=100, random_seed=42)
from pymc3.gp.util import plot_gp_dist
fig, ax = plt.subplots(figsize=(8,6))
plot_gp_dist(ax, gp_maize_samples['maize_pred'], X_pred)
maize_data.plot.scatter(x='TTsum_c', y='CropBiomassWt', c='k', s=50, ax=ax)
ax.set_ylim(0, 350);
with gp_maize_model:
maize_pred_noise = recruit_gp.conditional("maize_pred_noise", X_pred.reshape(-1,1), pred_noise=True)
gp_maize_samples = pm.sample_posterior_predictive(gp_trace, vars=[maize_pred_noise], samples=500, random_seed=42)
from pymc3.gp.util import plot_gp_dist
fig, ax = plt.subplots(figsize=(8,6))
plot_gp_dist(ax, gp_maize_samples['maize_pred_noise'], X_pred)
maize_data.plot.scatter(x='TTsum_c', y='CropBiomassWt', c='k', s=50, ax=ax)
ax.set_ylim(0, 350);
# ### Exercise
#
# We might be interested in what may happen if the population gets very large -- say, 600 or 800 spawners. We can predict this, though it goes well outside the range of data that we have observed. Generate predictions from the posterior predictive distribution that covers this range of spawners.
#
# *Hint: you need to add a new `conditional` variable.*
# +
# Write answer here
| notebooks/maize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="1NF27HFLxZMK" papermill={"duration": 0.017078, "end_time": "2021-10-11T21:33:48.361316", "exception": false, "start_time": "2021-10-11T21:33:48.344238", "status": "completed"} tags=[]
# ## Library Imports
# + id="ZaibE8jTxTRV" papermill={"duration": 0.033182, "end_time": "2021-10-11T21:33:48.410697", "exception": false, "start_time": "2021-10-11T21:33:48.377515", "status": "completed"} tags=[]
from time import time
notebook_start_time = time()
# + id="tCefPeQFxwtS" papermill={"duration": 5.659781, "end_time": "2021-10-11T21:33:54.086492", "exception": false, "start_time": "2021-10-11T21:33:48.426711", "status": "completed"} tags=[]
import os
import re
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import torch
from torch import nn, optim
from torch.utils.data import Dataset
from torch.utils.data import DataLoader as DL
from torch.nn.utils import weight_norm as WN
from torchvision import models, transforms
from sklearn.model_selection import KFold
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
# + [markdown] id="iWlt7AdSxxWm" papermill={"duration": 0.014926, "end_time": "2021-10-11T21:33:54.116919", "exception": false, "start_time": "2021-10-11T21:33:54.101993", "status": "completed"} tags=[]
# ## Constants and Utilities
# + id="ZpMGLKhKx0EX" papermill={"duration": 0.074246, "end_time": "2021-10-11T21:33:54.206623", "exception": false, "start_time": "2021-10-11T21:33:54.132377", "status": "completed"} tags=[]
SEED = 49
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
DATA_PATH = "../input/petfinder-pawpularity-score"
FEATURE_PATH = "../input/petfinder-pf-nc-ua-all-dataset"
MODEL_NAME = "densenet169"
DEBUG = False
verbose = False
sc_y = StandardScaler()
# + id="yIR_gIEPyZrc" papermill={"duration": 0.02819, "end_time": "2021-10-11T21:33:54.249938", "exception": false, "start_time": "2021-10-11T21:33:54.221748", "status": "completed"} tags=[]
def breaker(num=50, char="*") -> None:
print("\n" + num*char + "\n")
def get_targets(path: str) -> np.ndarray:
df = pd.read_csv(os.path.join(path, "train.csv"), engine="python")
targets = df["Pawpularity"].copy().values
return targets.reshape(-1, 1)
def show_graphs(L: list, title=None) -> None:
TL, VL = [], []
for i in range(len(L)):
TL.append(L[i]["train"])
VL.append(L[i]["valid"])
x_Axis = np.arange(1, len(L) + 1)
plt.figure()
plt.plot(x_Axis, TL, "r", label="train")
plt.plot(x_Axis, VL, "b", label="valid")
plt.grid()
plt.legend()
if title:
plt.title("{} Loss".format(title))
else:
plt.title("Loss")
plt.show()
# + [markdown] id="7y0fd6rWy9C7" papermill={"duration": 0.015974, "end_time": "2021-10-11T21:33:54.280755", "exception": false, "start_time": "2021-10-11T21:33:54.264781", "status": "completed"} tags=[]
# ## Dataset Template and Build Dataloader
# + id="TtCaj5UAy7ar" papermill={"duration": 0.028455, "end_time": "2021-10-11T21:33:54.323761", "exception": false, "start_time": "2021-10-11T21:33:54.295306", "status": "completed"} tags=[]
class DS(Dataset):
def __init__(self, features=None, targets=None):
self.features = features
self.targets = targets
def __len__(self):
return self.features.shape[0]
def __getitem__(self, idx):
return torch.FloatTensor(self.features[idx]), torch.FloatTensor(self.targets[idx])
def build_dataloaders(tr_features: np.ndarray, va_features: np.ndarray,
tr_targets: np.ndarray, va_targets: np.ndarray,
batch_size: int, seed: int):
if verbose:
breaker()
print("Building Train and Validation DataLoaders ...")
tr_data_setup = DS(features=tr_features, targets=tr_targets)
va_data_setup = DS(features=va_features, targets=va_targets)
dataloaders = {
"train" : DL(tr_data_setup, batch_size=batch_size, shuffle=True, generator=torch.manual_seed(seed)),
"valid" : DL(va_data_setup, batch_size=batch_size, shuffle=False)
}
return dataloaders
# + [markdown] id="8wmsPRi6zjyn" papermill={"duration": 0.014622, "end_time": "2021-10-11T21:33:54.353917", "exception": false, "start_time": "2021-10-11T21:33:54.339295", "status": "completed"} tags=[]
# ## Build Model
# + id="2GYZ4teczjIm" papermill={"duration": 0.029085, "end_time": "2021-10-11T21:33:54.398988", "exception": false, "start_time": "2021-10-11T21:33:54.369903", "status": "completed"} tags=[]
def build_model(IL: int, seed: int):
class ANN(nn.Module):
def __init__(self, IL=None):
super(ANN, self).__init__()
self.predictor = nn.Sequential()
self.predictor.add_module("BN", nn.BatchNorm1d(num_features=IL, eps=1e-5))
self.predictor.add_module("FC", WN(nn.Linear(in_features=IL, out_features=1)))
def get_optimizer(self, lr=1e-3, wd=0):
params = [p for p in self.parameters() if p.requires_grad]
return optim.Adam(params, lr=lr, weight_decay=wd)
def get_plateau_scheduler(self, optimizer=None, patience=5, eps=1e-8):
return optim.lr_scheduler.ReduceLROnPlateau(optimizer=optimizer, patience=patience, eps=eps, verbose=True)
def forward(self, x1, x2=None):
if x2 is not None:
return self.predictor(x1), self.predictor(x2)
else:
return self.predictor(x1)
if verbose:
breaker()
print("Building Model ...")
print("\n{} -> 1".format(IL))
torch.manual_seed(seed)
model = ANN(IL=IL)
return model
# + [markdown] id="uuMvELK2zqh0" papermill={"duration": 0.015047, "end_time": "2021-10-11T21:33:54.428811", "exception": false, "start_time": "2021-10-11T21:33:54.413764", "status": "completed"} tags=[]
# ## Fit and Predict Helpers
# + id="8DQ9EUrIzpmn" papermill={"duration": 0.042856, "end_time": "2021-10-11T21:33:54.486610", "exception": false, "start_time": "2021-10-11T21:33:54.443754", "status": "completed"} tags=[]
def fit(model=None, optimizer=None, scheduler=None,
epochs=None, early_stopping_patience=None,
dataloaders=None, fold=None, lr=None, wd=None, verbose=False) -> tuple:
name = "./LR_{}_WD_{}_Fold_{}_state.pt".format(lr, wd, fold)
if verbose:
breaker()
print("Training Fold {}...".format(fold))
breaker()
# else:
# print("Training Fold {}...".format(fold))
Losses = []
bestLoss = {"train" : np.inf, "valid" : np.inf}
start_time = time()
for e in range(epochs):
e_st = time()
epochLoss = {"train" : np.inf, "valid" : np.inf}
for phase in ["train", "valid"]:
if phase == "train":
model.train()
else:
model.eval()
lossPerPass = []
for X, y in dataloaders[phase]:
X, y = X.to(DEVICE), y.to(DEVICE)
optimizer.zero_grad()
with torch.set_grad_enabled(phase == "train"):
output = model(X)
loss = torch.nn.MSELoss()(output, y)
if phase == "train":
loss.backward()
optimizer.step()
lossPerPass.append(loss.item())
epochLoss[phase] = np.mean(np.array(lossPerPass))
Losses.append(epochLoss)
if early_stopping_patience:
if epochLoss["valid"] < bestLoss["valid"]:
bestLoss = epochLoss
BLE = e + 1
torch.save({"model_state_dict": model.state_dict(),
"optim_state_dict": optimizer.state_dict()},
name)
early_stopping_step = 0
else:
early_stopping_step += 1
if early_stopping_step > early_stopping_patience:
if verbose:
print("\nEarly Stopping at Epoch {}".format(e))
break
if epochLoss["valid"] < bestLoss["valid"]:
bestLoss = epochLoss
BLE = e + 1
torch.save({"model_state_dict": model.state_dict(),
"optim_state_dict": optimizer.state_dict()},
name)
if scheduler:
scheduler.step(epochLoss["valid"])
if verbose:
print("Epoch: {} | Train Loss: {:.5f} | Valid Loss: {:.5f} | Time: {:.2f} seconds".format(e+1, epochLoss["train"], epochLoss["valid"], time()-e_st))
if verbose:
breaker()
print("Best Validation Loss at Epoch {}".format(BLE))
breaker()
print("Time Taken [{} Epochs] : {:.2f} minutes".format(len(Losses), (time()-start_time)/60))
breaker()
print("Training Completed")
breaker()
return Losses, BLE, name
#####################################################################################################
def predict_batch(model=None, dataloader=None, mode="test", path=None) -> np.ndarray:
model.load_state_dict(torch.load(path, map_location=DEVICE)["model_state_dict"])
model.to(DEVICE)
model.eval()
y_pred = torch.zeros(1, 1).to(DEVICE)
if re.match(r"valid", mode, re.IGNORECASE):
for X, _ in dataloader:
X = X.to(DEVICE)
with torch.no_grad():
output = model(X)
y_pred = torch.cat((y_pred, output.view(-1, 1)), dim=0)
elif re.match(r"test", mode, re.IGNORECASE):
for X in dataloader:
X = X.to(DEVICE)
with torch.no_grad():
output = model(X)
y_pred = torch.cat((y_pred, output.view(-1, 1)), dim=0)
return y_pred[1:].detach().cpu().numpy()
# + [markdown] id="YQP876MS0OrX" papermill={"duration": 0.015738, "end_time": "2021-10-11T21:33:54.517505", "exception": false, "start_time": "2021-10-11T21:33:54.501767", "status": "completed"} tags=[]
# ## Train
# + id="OrGICYXg0QEM" papermill={"duration": 0.033581, "end_time": "2021-10-11T21:33:54.566357", "exception": false, "start_time": "2021-10-11T21:33:54.532776", "status": "completed"} tags=[]
def train(features: np.ndarray, targets: np.ndarray,
n_splits: int, batch_size: int, lr: float, wd: float,
epochs: int, early_stopping: int,
patience=None, eps=None) -> list:
metrics = []
KFold_start_time = time()
if verbose:
breaker()
print("\tLR : {}, WD: {}".format(lr, wd))
breaker()
print("Performing {} Fold CV ...".format(n_splits))
fold = 1
for tr_idx, va_idx in KFold(n_splits=n_splits, shuffle=True, random_state=SEED).split(features):
tr_features, va_features = features[tr_idx], features[va_idx]
tr_targets, va_targets = targets[tr_idx], targets[va_idx]
tr_targets = sc_y.fit_transform(tr_targets)
va_targets = sc_y.transform(va_targets)
dataloaders = build_dataloaders(tr_features, va_features,
tr_targets, va_targets,
batch_size, SEED)
model = build_model(IL=tr_features.shape[1], seed=SEED).to(DEVICE)
optimizer = model.get_optimizer(lr=lr, wd=wd)
scheduler = None
if isinstance(patience, int) and isinstance(eps, float):
scheduler = model.get_plateau_scheduler(optimizer, patience, eps)
L, _, name = fit(model=model, optimizer=optimizer, scheduler=scheduler,
epochs=epochs, early_stopping_patience=early_stopping,
dataloaders=dataloaders, fold=fold, lr=lr, wd=wd, verbose=verbose)
y_pred = predict_batch(model=model, dataloader=dataloaders["valid"], mode="valid", path=name)
RMSE = np.sqrt(mean_squared_error(sc_y.inverse_transform(y_pred), sc_y.inverse_transform(va_targets)))
if verbose:
print("\nValidation RMSE [Fold {}]: {:.5f}".format(fold, RMSE))
breaker()
show_graphs(L)
metrics_dict = {"Fold" : fold, "LR" : lr, "WD" : wd, "RMSE" : RMSE}
metrics.append(metrics_dict)
fold += 1
if verbose:
breaker()
print("Total Time to {} Fold CV : {:.2f} minutes".format(n_splits, (time() - KFold_start_time)/60))
return metrics, (time() - KFold_start_time)/60
# + [markdown] id="UGUqaAzy1C33" papermill={"duration": 0.014556, "end_time": "2021-10-11T21:33:54.595796", "exception": false, "start_time": "2021-10-11T21:33:54.581240", "status": "completed"} tags=[]
# ## Main
# + id="_0yK3IwR1CEs" papermill={"duration": 0.03379, "end_time": "2021-10-11T21:33:54.644995", "exception": false, "start_time": "2021-10-11T21:33:54.611205", "status": "completed"} tags=[]
def main():
DEBUG = False
########### Params ###########
if DEBUG:
n_splits = 10
patience, eps = 5, 1e-8
epochs, early_stopping = 5, 5
batch_size = 128
lrs = [1e-2, 1e-3]
wds = [0.0, 1e-1]
else:
n_splits = 10
patience, eps = 5, 1e-8
epochs, early_stopping = 100, 8
batch_size = 128
lrs = [1e-3, 5e-4, 1e-4]
wds = [0.0, 1e-1, 1e-2, 1e-3, 1e-4, 1e-5, 1e-6]
##############################
complete_metrics = []
if verbose:
breaker()
print("Loading Data ...")
else:
breaker()
features = np.load(os.path.join(FEATURE_PATH, "{}_features.npy".format(MODEL_NAME)))
targets = get_targets(DATA_PATH)
for lr in lrs:
for wd in wds:
# Without Scheduler
metrics, time_taken = train(features, targets, n_splits, batch_size, lr, wd, epochs, early_stopping, patience=None, eps=None)
# # With Plateau Scheduler
# metrics = train(features, targets, n_splits, batch_size, lr, wd, epochs, early_stopping, patience=patience, eps=eps)
complete_metrics.append(metrics)
if not verbose:
print("LR : {}, WD: {} -> {:.2f} minutes".format(lr, wd, time_taken))
if verbose:
breaker()
for i in range(len(complete_metrics)):
for j in range(len(complete_metrics[i])):
print(complete_metrics[i][j])
rmse = []
for i in range(len(complete_metrics)):
for j in range(len(complete_metrics[i])):
rmse.append(complete_metrics[i][j]["RMSE"])
best_index = rmse.index(min(rmse))
best_index_1 = best_index // n_splits
best_index_2 = best_index % n_splits
breaker()
print("Best RMSE: {:.5f} using LR: {} and WD: {}".format(complete_metrics[best_index_1][best_index_2]["RMSE"],
complete_metrics[best_index_1][best_index_2]["LR"],
complete_metrics[best_index_1][best_index_2]["WD"]))
breaker()
with open("complete_metrics.pkl", "wb") as fp:
pickle.dump(complete_metrics, fp)
# + id="9Vf0T4VN2Z1Q" outputId="d8f15<PASSWORD>-4<PASSWORD>" papermill={"duration": 2161.000679, "end_time": "2021-10-11T22:09:55.660909", "exception": false, "start_time": "2021-10-11T21:33:54.660230", "status": "completed"} tags=[]
main()
# + papermill={"duration": 0.036502, "end_time": "2021-10-11T22:09:55.720425", "exception": false, "start_time": "2021-10-11T22:09:55.683923", "status": "completed"} tags=[]
if not verbose:
with open("complete_metrics.pkl", "rb") as fp:
params = pickle.load(fp)
rmse = []
for i in range(len(params)):
for j in range(len(params[i])):
rmse.append(params[i][j]["RMSE"])
best_index = rmse.index(min(rmse))
if DEBUG:
best_index_1 = best_index // 3
best_index_2 = best_index % 3
else:
best_index_1 = best_index // 10
best_index_2 = best_index % 10
breaker()
print("Params: {}".format(params[best_index_1][best_index_2]))
breaker()
# + id="YmNqGwy98SyF" papermill={"duration": 0.033019, "end_time": "2021-10-11T22:09:55.776473", "exception": false, "start_time": "2021-10-11T22:09:55.743454", "status": "completed"} tags=[]
breaker()
print("Notebook Rumtime : {:.2f} minutes".format((time() - notebook_start_time)/60))
breaker()
| PF-2/Notebooks/Analysis/D169 (LR,WD) (NC) (BN,WN).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simuation Final Project v3
# ## DATA604 - Summer 2020
# ### Completed by: <NAME>
# ### Date: July 16, 2020
#
# The simulation models an NBA game for a single team in addition to simulating a matchup between two teams. The matchup simulation can be extrapolated as a playoff series.
#
# At the heart of the game model is the model of a single team possession. The possession is considered a discrete event in this model. The outcome of a single possession is zero to four points.
#
# The model is based on overall team statistics for the NBA season 2019-20 which is currently on hiatus due to the Covid-19 pandemic. The statistics were gathered from two sources: stats.nba.com and basketball-reference.com.
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
from pandas import *
import numpy as np
import math
import time
# -
# ## Retrieve Statistics
#
# The team statistics for the 30 NBA teams are stored in a CSV file in the same directory as this Jupyter notebook. The file is read from the current directory and then the columns are renamed for clarity. Then, the numeric columns are type-casted to numeric data types as initially all data is read in as strings. The statistics are then output to the file for visual inspection.
# +
# Read in statistics for all 30 NBA teams
filename = 'Teams_Stats_Sim_v3.csv'
all_teams = read_csv(filename, header=0, decimal='M')
# Rename the columns
all_teams.columns = ['team', 'turnover_pct_per_play', 'fga_pct_per_play', 'off_reb_per_play', 'two_pt_fg_attempts_pct',
'two_pt_fg_shooting_pct', 'three_pt_fg_shooting_pct', 'ft_shooting_pct', 'shooting_foul_drawn_pct',
'pace', 'off_rating', 'def_rating']
# Cast the appropriate columns to numeric instead of string
cols = ['turnover_pct_per_play', 'fga_pct_per_play', 'off_reb_per_play', 'two_pt_fg_attempts_pct',
'two_pt_fg_shooting_pct', 'three_pt_fg_shooting_pct', 'ft_shooting_pct', 'shooting_foul_drawn_pct',
'pace', 'off_rating', 'def_rating']
all_teams[cols] = all_teams[cols].apply(pd.to_numeric, errors='coerce')
# Output the complete input file just for visual validation
all_teams
# -
# ## Approach
#
# - The model is constructed using many concise functions to replicate elements of the possession.
#
# - The season statistics for each team retrieved from the CSV file are defined as a system object.
#
# - The game statistics for each team are defined and maintained as a state object.
# ### Statistics used
#
# - Turnover percentage per play
#
# - Field Goal Attempt percentage per play
#
# - Offensive Rebound percentage per play
#
# - Two-point Field Goal Attempt percentage per play
#
# - Two-point Field Goal Shooting percentage
#
# - Three-point Field Goal Shooting percentage
#
# - Free-Throw Shooting percentage
#
# - Shooting Foul Drawn percentage per play
#
# - Overal Team Pace (possessions per game)
#
# - Overall Team Offensive Rating
#
# - Overall Team Defensive Rating
# Following function maps in the input data (season statistics) from the primary dataframe to a system object for one team based on the input parameter.
def map_team_stats_to_system(team):
"""
Given the team 3-character identifier, create a system object based on the input file of team statistics
"""
team_stats = all_teams[all_teams['team']== team]
team_sys = System(turnover_pct_per_play=team_stats.iloc[0]['turnover_pct_per_play'],
fga_pct_per_play =team_stats.iloc[0]['fga_pct_per_play'],
off_reb_per_play =team_stats.iloc[0]['off_reb_per_play'],
two_pt_fg_attempts_pct =team_stats.iloc[0]['two_pt_fg_attempts_pct'],
two_pt_fg_shooting_pct =team_stats.iloc[0]['two_pt_fg_shooting_pct'],
three_pt_fg_shooting_pct =team_stats.iloc[0]['three_pt_fg_shooting_pct'],
ft_shooting_pct =team_stats.iloc[0]['ft_shooting_pct'],
shooting_foul_drawn_pct =team_stats.iloc[0]['shooting_foul_drawn_pct'],
pace =team_stats.iloc[0]['pace'],
off_rating =team_stats.iloc[0]['off_rating'],
def_rating =team_stats.iloc[0]['def_rating'])
return team_sys
# Test map_team_stats_to_system
atl = map_team_stats_to_system('ATL')
# Function used to create and initialize the game statistics State object for a game simulation
def initialize_game_stats_state():
"""
Initialize the game statistics to zero and return State object
"""
game_stats_state = State(turnovers=0,
two_point_field_goal_attempts=0,
two_point_field_goal_makes=0,
three_point_field_goal_attempts=0,
three_point_field_goal_makes=0,
free_throw_attempts=0,
free_throw_makes=0,
shooting_fouls_drawn=0,
offensive_rebounds=0)
return game_stats_state
# initialize_game_stats_state
stats_state = initialize_game_stats_state()
# Function to build the System object and State object for a team as input to a game simulation.
#
# Display of the composite object shows the two attributes are the game statistics state and overall statistics system.
def create_team_game_object(team_name):
"""
Create a team object as a composite of the game stats State object and the season statistics System object
"""
team = State(game_stats_state=initialize_game_stats_state(),
overall_stats_sys=map_team_stats_to_system(team_name))
return team
# +
# Test create_team_game_object
spurs = create_team_game_object('SAS')
spurs
# -
# Create a system object to represent a team
# team_A object used throughout the Jupyter notebook to individually test functions
team_A = map_team_stats_to_system('MIL')
# Utility function to convert a percentage between 0.0 and 1.0 to an integer from zero to one thousand.
def convert_pct_to_int(pct):
"""
Utility function to convert any percentage to an integer of range 0-1000
"""
return int(pct * 1000)
# Test convert_pct_to_int function
convert_pct_to_int(.47)
# Following function determines if a turnover occurs based on the team percentage
def did_turnover_occur(team):
"""
Determine if a possession results in a turnover
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.turnover_pct_per_play)
if (rand_num <= pct):
result = True
return result
# Test did_turnover_occur
did_turnover_occur(team_A)
# Following function determines if a field goal attempt occurs based on the team percentage
def did_field_goal_attempt_occur(team):
"""
Determine if a possession results in a field goal attempt
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.fga_pct_per_play)
if (rand_num <= pct):
result = True
return result
# Test did_field_goal_attempt_occur
did_field_goal_attempt_occur(team_A)
# Following function determines if an offensive rebound occurs based on the team percentage
def did_offensive_rebound_occur(team):
"""
Determine if a play results in a offensive rebound
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.off_reb_per_play)
# Setting offensive rebounding percentage to 20%
pct = 200
if (rand_num <= pct):
result = True
return result
# Test did_offensive_rebound_occur
did_offensive_rebound_occur(team_A)
# Following function determines if a two-point FG attempt occurs as opposed to a three-point FG attempt based on the team percentage
def is_2P_FGA(team):
"""
Determine if a two point field goal attempt occurs
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.two_pt_fg_attempts_pct)
if (rand_num <= pct):
result = True
return result
# Test is_2P_FGA
is_2P_FGA(team_A)
# Following function determines if a two-point FG was successful based on the team percentage
def is_2P_FGA_made(team):
"""
Determine if a two-point field goal attempt was successful
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.two_pt_fg_shooting_pct)
if (rand_num <= pct):
result = True
return result
# Test is_2P_FGA_made
is_2P_FGA_made(team_A)
# Following function determines if a three-point FG was successful based on the team percentage
def is_3P_FGA_made(team):
"""
Determine if a three-point field goal attempt was successful
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.three_pt_fg_shooting_pct)
if (rand_num <= pct):
result = True
return result
# Test is_3P_FGA_made
is_3P_FGA_made(team_A)
# Following function determines if a free-throw attempt was successful based on the team percentage
def is_FTA_made(team):
"""
Determine if a free-throw attempt was successful
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.ft_shooting_pct)
if (rand_num <= pct):
result = True
return result
# Test is_FTA_made
is_FTA_made(team_A)
# Following function determines if a shooting fould occurs based on the team percentage
#
# - Note: The percentage is static for all team's at a very low percentage.
def did_shooting_foul_occur(team):
"""
Determine if a shooting foul occurred
"""
result = False
# Retrieve random number
rand_num = np.random.uniform(1, 1000)
# Convert percent to int
pct = convert_pct_to_int(team.shooting_foul_drawn_pct)
# Hardcoding 50 for test purposes
pct = 50
if (rand_num <= pct):
result = True
return result
# Test did_shooting_foul_occur
did_shooting_foul_occur(team_A)
# Following function simulates a free-throw attempt based on the team percentage
def simulate_FT_attempt(team):
"""
Simulate a free throw attempt based on the team's free-throw percentage
"""
# result represents the points scored
result_pts = 0
if (is_FTA_made(team)):
result_pts += 1
return result_pts
# Test simulate_FT_attempt
simulate_FT_attempt(team_A)
# Following function simulates multiple free-throw attempts based on the input parameter 'attempts'
def simulate_FT_attempts(team, attempts):
"""
Simulate free throw attempts
"""
# result represents the points scored
result_pts = 0
# Loop through the number of attempts
for a in range(1, attempts+1):
result_pts += simulate_FT_attempt(team)
return result_pts
# Test simulate_FT_attempts
simulate_FT_attempts(team_A, 2)
# In order to determine if a team grabs an offensive rebound, the model must know if the final free throw attempt was a made basket or a miss. The below function will return the number of points scored for the given count of attempts along with a flag indicating the state of the final attempt.
def simulate_FT_attempts_with_final_FT_result(team, attempts):
"""
Simulate free throw attempts
"""
# result represents the points scored
result_pts = 0
result_final_FT_miss = False
# Loop through the number of attempts
for a in range(1, attempts+1):
result = simulate_FT_attempt(team)
result_pts += result
# For final shot, set final FT miss flag
if (a == attempts):
if (result == 0):
result_final_FT_miss = True
return result_pts, result_final_FT_miss
# Test simulate_FT_attempts_with_final_FT_result
simulate_FT_attempts_with_final_FT_result(team_A, 2)
# To accurately reflect a true basketball possession, the model takes into account the possibility of the "and-one" shooting foul on a made field goal.
def simulate_2P_FG_attempt_with_shooting_foul_possible(team):
"""
Simulate a two-point field goal attempt
"""
# result represents the points scored
result_pts = 0
result_miss_fg = False
result_miss_ft = False
result_2P_FGA = is_2P_FGA_made(team)
if (result_2P_FGA):
# FG attempt was made, add two points
result_pts += 2
is_shooting_foul_drawn = did_shooting_foul_occur(team)
if (is_shooting_foul_drawn):
# simulate free throw attempt
pts, is_final_FT_missed = simulate_FT_attempts_with_final_FT_result(team_A, 1)
result_pts += pts
result_miss_ft = is_final_FT_missed
else:
result_pts = 0
result_miss_fg = True
return result_pts, result_miss_fg, result_miss_ft
# Test simulate_2P_FG_attempt_with_shooting_foul_possible
simulate_2P_FG_attempt_with_shooting_foul_possible(team_A)
# Following function simulates a two-point FG attempt
def simulate_2P_FG_attempt(team):
"""
Simulate a two-point field goal attempt
"""
# result represents the points scored
result_pts = 0
if (is_2P_FGA_made(team)):
result_pts += 2
return result_pts
# Test simulate_2P_FG_attempt
simulate_2P_FG_attempt(team_A)
# Following function simulates a three-point FG attempt
def simulate_3P_FG_attempt(team):
"""
Simulate a three-point field goal attempt
"""
# result represents the points scored
result_pts = 0
if (is_3P_FGA_made(team)):
result_pts += 3
return result_pts
# Test simulate_3P_FG_attempt
simulate_3P_FG_attempt(team_A)
# Again, to accurately reflect a true basketball possession, the model takes into account the possibility of the "and-one" shooting foul on a made field goal.
def simulate_3P_FG_attempt_with_shooting_foul_possible(team):
"""
Simulate a three-point field goal attempt
"""
# result represents the points scored
result_pts = 0
result_miss_fg = False
result_miss_ft = False
result_3P_FGA = is_3P_FGA_made(team)
if (result_3P_FGA):
# FG attempt was made, add three points
result_pts += 3
is_shooting_foul_drawn = did_shooting_foul_occur(team)
if (is_shooting_foul_drawn):
# simulate free throw attempt
pts, is_final_FT_missed = simulate_FT_attempts_with_final_FT_result(team_A, 1)
result_pts += pts
result_miss_ft = is_final_FT_missed
else:
result_pts = 0
result_miss_fg = True
return result_pts, result_miss_fg, result_miss_ft
# Test simulate_3P_FG_attempt_with_shooting_foul_possible
simulate_3P_FG_attempt_with_shooting_foul_possible(team_A)
# This function will modify a team's field-goal percentage based on the difference between the team's offensive rating and the opponent's defensive rating. Please note, a better defensive rating is a lower rating. The linear formula used to determine the amount to change is based on all the games of the 2019-20 NBA season to date.
#
# A good defensive team will cause a decrease in the field-goal shooting percentage.
#
# This function, along with the following two, are used to better simulate the matchup between two teams. Simply running the simulation for two teams independent of impact on one another ends up rewarding strong offensive teams. By creating this functions to account for opponent's defensive rating, the percentages of a team are adjusted according to the specific opponent.
def calculate_fg_pct_based_on_opp(team_off_rating, opp_def_rating, team_fg_pct):
"""
Calculate a shooting percentage based on the overall team offensive rating and opponent's defensive rating
"""
# 0.00355*x + -0.00505
# Based on linear regression formula
return (0.00355 * (opp_def_rating - team_off_rating) - 0.00505) + team_fg_pct
calculate_fg_pct_based_on_opp(111.3, 101.6, 0.47)
# This function will modify a team's three-point field-goal percentage based on the difference between the team's offensive rating and the opponent's defensive rating. Please note, a better defensive rating is a lower rating. The linear formula used to determine the amount to change is based on all the games of the 2019-20 NBA season to date.
#
# A good defensive team will cause a decrease in the three-point field-goal shooting percentage.
def calculate_3p_fg_pct_based_on_opp(team_off_rating, opp_def_rating, team_3p_fg_pct):
"""
Calculate a shooting percentage based on the overall team offensive rating and opponent's defensive rating
"""
# 0.00463*x + -0.00847
# Based on linear regression formula
return (0.00463 * (opp_def_rating - team_off_rating) - 0.00847) + team_3p_fg_pct
calculate_3p_fg_pct_based_on_opp(111.3, 101.6, 0.383)
# This function will modify a team's turnover percentage based on the difference between the team's offensive rating and the opponent's defensive rating. Please note, a better defensive rating is a lower rating. The linear formula used to determine the amount to change is based on all the games of the 2019-20 NBA season to date.
#
# A good defensive team will cause an increase in the turnover percentage.
def calculate_turnover_pct_based_on_opp(team_off_rating, opp_def_rating, turnover_pct):
"""
Calculate a turnover percentage based on the overall team offensive rating and opponent's defensive rating
"""
# -0.000993*x + -0.00478
# Based on linear regression formula
return (-0.000993 * (opp_def_rating - team_off_rating) - 0.00478) + turnover_pct
calculate_turnover_pct_based_on_opp(111.3, 101.6, 0.136)
calculate_turnover_pct_based_on_opp(112.3, 114.6, 0.136)
# Utility function used to keep a running total of the game statistics from each possession simulation.
def combine_poss_stats(poss_stats_running, poss_stats_instance):
poss_stats_running.turnovers += poss_stats_instance.turnovers
poss_stats_running.two_point_field_goal_attempts += poss_stats_instance.two_point_field_goal_attempts
poss_stats_running.two_point_field_goal_makes += poss_stats_instance.two_point_field_goal_makes
poss_stats_running.three_point_field_goal_attempts += poss_stats_instance.three_point_field_goal_attempts
poss_stats_running.three_point_field_goal_makes += poss_stats_instance.three_point_field_goal_makes
poss_stats_running.free_throw_attempts += poss_stats_instance.free_throw_attempts
poss_stats_running.free_throw_makes += poss_stats_instance.free_throw_makes
poss_stats_running.shooting_fouls_drawn += poss_stats_instance.shooting_fouls_drawn
poss_stats_running.offensive_rebounds += poss_stats_instance.offensive_rebounds
return poss_stats_running
# +
# Test combine_poss_stats
running = State(turnovers=1,
two_point_field_goal_attempts=3,
two_point_field_goal_makes=4,
three_point_field_goal_attempts=2,
three_point_field_goal_makes=7,
free_throw_attempts=10,
free_throw_makes=8,
shooting_fouls_drawn=2,
offensive_rebounds=5)
instanc = State(turnovers=6,
two_point_field_goal_attempts=5,
two_point_field_goal_makes=2,
three_point_field_goal_attempts=2,
three_point_field_goal_makes=8,
free_throw_attempts=4,
free_throw_makes=9,
shooting_fouls_drawn=2,
offensive_rebounds=1)
running = combine_poss_stats(running, instanc)
running
# -
# This function, simulate_possession, is the heart of the NBA game model. This is the primary function in which a possession is simulated with the result being the number of points scored in the possession along with the game statistics of the possession itself.
def simulate_possession(team):
"""
Simulate a possession for the given team
"""
# Track stats for possession
poss_stats = initialize_game_stats_state()
result_pts = 0
is_turnover = did_turnover_occur(team)
if (is_turnover):
# Turnover occured, return 0 points
# Line not necessary, but leaving in for clarity
poss_stats.turnovers += 1
result_pts = 0
else:
# Check if FGA occurred vs. FT attempts
is_FGA = did_field_goal_attempt_occur(team)
if (is_FGA):
# FG attempted: 2P or 3P
result_2P_FGA = is_2P_FGA(team)
if (result_2P_FGA):
# 2P Field Goal attempted
pts, miss_fg, miss_ft = simulate_2P_FG_attempt_with_shooting_foul_possible(team)
result_pts += pts
# Count shot attempt
poss_stats.two_point_field_goal_attempts += 1
if (pts == 2):
poss_stats.two_point_field_goal_makes += 1
if (miss_ft):
poss_stats.free_throw_attempts += 1
poss_stats.shooting_fouls_drawn += 1
elif (pts == 3):
poss_stats.two_point_field_goal_makes += 1
poss_stats.free_throw_attempts += 1
poss_stats.free_throw_makes += 1
poss_stats.shooting_fouls_drawn += 1
# Opportunity for offensive rebound
if (miss_fg or miss_ft):
offensive_rebound = did_offensive_rebound_occur(team)
if (offensive_rebound):
# Possession starts over
poss_stats.offensive_rebounds += 1
# Recursively call simulate_possession
poss_pts_recur, poss_stats_recur = simulate_possession(team)
result_pts += poss_pts_recur
poss_stats = combine_poss_stats(poss_stats, poss_stats_recur)
# else: Offensive rebound did not occur; Possession over
# else: Nothing here, as opposing team will inbound ball; Possession over
else:
# 3P Field Goal attempted
pts, miss_fg, miss_ft = simulate_3P_FG_attempt_with_shooting_foul_possible(team)
result_pts += pts
# Count shot attempt
poss_stats.three_point_field_goal_attempts += 1
if (pts == 3):
poss_stats.three_point_field_goal_makes += 1
if (miss_ft):
poss_stats.free_throw_attempts += 1
poss_stats.shooting_fouls_drawn += 1
elif (pts == 4):
poss_stats.three_point_field_goal_makes += 1
poss_stats.free_throw_attempts += 1
poss_stats.free_throw_makes += 1
poss_stats.shooting_fouls_drawn += 1
# Opportunity for offensive rebound
if (miss_fg or miss_ft):
offensive_rebound = did_offensive_rebound_occur(team)
if (offensive_rebound):
# Possession starts over
poss_stats.offensive_rebounds += 1
# Recursively call simulate_possession
poss_pts_recur, poss_stats_recur = simulate_possession(team)
result_pts += poss_pts_recur
poss_stats = combine_poss_stats(poss_stats, poss_stats_recur)
# else: Offensive rebound did not occur; Possession over
# else: Nothing here, as opposing team will inbound ball; Possession over
else:
# Free throw attempts
pts, miss = simulate_FT_attempts_with_final_FT_result(team_A, 2)
result_pts += pts
poss_stats.free_throw_attempts += 2
poss_stats.shooting_fouls_drawn += 1
if (pts == 1):
poss_stats.free_throw_makes += 1
elif (pts == 2):
poss_stats.free_throw_makes += 2
# Opportunity for offensive rebound
if (miss):
offensive_rebound = did_offensive_rebound_occur(team)
if (offensive_rebound):
# Possession starts over
poss_stats.offensive_rebounds += 1
# Recursively call simulate_possession
poss_pts_recur, poss_stats_recur = simulate_possession(team)
result_pts += poss_pts_recur
poss_stats = combine_poss_stats(poss_stats, poss_stats_recur)
# else: Offensive rebound did not occur; Possession over
# else: Nothing here, as opposing team will inbound ball; Possession over
return result_pts, poss_stats
# Test simulate_possession
simulate_possession(team_A)
# simulate_game function simply calls the simulate_possession function a total number of times based on the team's pace of play.
def simulate_game(team):
"""
Simulate a game by summing results of all possessions
"""
score = 0
score_at_poss = TimeSeries()
# Track stats for game
game_stats = initialize_game_stats_state()
score_at_poss[0] = score
for p in range(1, math.ceil(team.pace)+1):
poss_pts, poss_stats = simulate_possession(team)
score += poss_pts
score_at_poss[p] = score
game_stats = combine_poss_stats(game_stats, poss_stats)
return score_at_poss, game_stats
# simuate_game function is verified along with a plot to illustrate the team's score increasing along with the possessions.
# +
# Test simulate_game
game_score, game_stats = simulate_game(team_A)
plot(game_score, label='Points')
decorate(title='Running Game Score',
xlabel='Possession',
ylabel='Score')
game_score.max()
# -
# As a means of validation, confirming the model fits the real-world system expectations, the final game statistics State object is output to screen to allow for comparison with the team's real per game totals.
game_stats
# +
# Let's find the average score for a team (Spurs) ... should be about 113
# Validation of the simulate_game model
spurs = map_team_stats_to_system('SAS')
results = TimeSeries()
turnovers = TimeSeries()
two_point_field_goal_attempts = TimeSeries()
two_point_field_goal_makes = TimeSeries()
three_point_field_goal_attempts = TimeSeries()
three_point_field_goal_makes = TimeSeries()
free_throw_attempts = TimeSeries()
free_throw_makes = TimeSeries()
shooting_fouls_drawn = TimeSeries()
offensive_rebounds = TimeSeries()
for g in range(1, 11):
game_score, game_stats = simulate_game(spurs)
results[g] = game_score.max()
turnovers[g] = game_stats.turnovers
two_point_field_goal_attempts[g] = game_stats.two_point_field_goal_attempts
two_point_field_goal_makes[g] = game_stats.two_point_field_goal_makes
three_point_field_goal_attempts[g] = game_stats.three_point_field_goal_attempts
three_point_field_goal_makes[g] = game_stats.three_point_field_goal_makes
free_throw_attempts[g] = game_stats.free_throw_attempts
free_throw_makes[g] = game_stats.free_throw_makes
shooting_fouls_drawn[g] = game_stats.shooting_fouls_drawn
offensive_rebounds[g] = game_stats.offensive_rebounds
plot(game_score)
decorate(title='Running Game Score',
xlabel='Possession',
ylabel='Score')
print("Avg Score :", results.mean())
print("Avg TOs :", turnovers.mean())
print("Avg 2P FGAs:", two_point_field_goal_attempts.mean())
print("Avg 2P FGMs:", two_point_field_goal_makes.mean())
print("Avg 3P FGAs:", three_point_field_goal_attempts.mean())
print("Avg 3P FGMs:", three_point_field_goal_makes.mean())
print("Avg FTAs :", free_throw_attempts.mean())
print("Avg FTMs :", free_throw_makes.mean())
print("Avg SFs :", shooting_fouls_drawn.mean())
print("Avg OffRebs:", offensive_rebounds.mean())
# -
# For documentation purposes: I ran the above simulation 100 times for the Spurs at 11:28am CT 7/12/20. Here are the results:
#
# - Avg Score : 113.26
#
# - Avg TOs : 11.33
#
# - Avg 2P FGAs: 59.18
#
# - Avg 2P FGMs: 31.03
#
# - Avg 3P FGAs: 29.18
#
# - Avg 3P FGMs: 11.2
#
# - Avg FTAs : 23.74
#
# - Avg FTMs : 17.6
#
# - Avg SFs : 12.91
#
# - Avg OffRebs: 9.52
#
# Spurs 2019-20 season per-game averages according to https://stats.nba.com/teams/traditional/
#
# - Points : 113.2
#
# - Turnovers : 12.3
#
# - 2P FGAs : 60.8 (89.5 - 28.7)
#
# - 2P FGMs : 31.3 (42.0 - 10.7)
#
# - 3P FGAs : 28.7
#
# - 3P FGMs : 10.7
#
# - FTAs : 22.8
#
# - FTMs : 18.4
#
# - SFs : --
#
# - OffRebs : 8.8
# Below plot shows the distribution of final game score for each of the simulations run.
# +
game_totals_ser = pandas.Series(results)
game_totals_ser.plot.hist(bins=20, alpha=0.5, legend=False)
decorate(title='Distribution of Scores',
xlabel='Score',
ylabel='Frequency',
legend=None)
# -
# In order to ensure the same pace for two teams, the average of the two teams' paces are calculated.
def calculate_pace_of_game(team_one_pace, team_two_pace):
"""
Calculate pace of a game between two teams
"""
# Calculate average of the pace given the two teams
return (team_one_pace + team_two_pace) / 2
pace = calculate_pace_of_game(100.6, 101.3)
# The below function, simulate_matchup_between_two_teams, is the heart of the model that simulates a game between two teams.
def simulate_matchup_between_two_teams(team_1, team_2, games):
"""
Simulate a matchup between two teams.
High level: The simulate_game function is called for each team, and then the results (final scores) are compared
Algorithm does identify tie games and runs a simulation for overtime until there is a winner
games: games per matchup simulation
"""
m_start_time = time.time()
# Map statistics from input file
team_one = map_team_stats_to_system(team_1)
team_two = map_team_stats_to_system(team_2)
# Calculate pace of game based on the two teams
matchup_pace = calculate_pace_of_game(team_one.pace, team_two.pace)
# Modify shooting percentages based on offensive and defensive team ratings
team_one.two_pt_fg_shooting_pct = calculate_fg_pct_based_on_opp(team_one.off_rating, team_two.def_rating, team_one.two_pt_fg_shooting_pct)
team_one.three_pt_fg_shooting_pct = calculate_3p_fg_pct_based_on_opp(team_one.off_rating, team_two.def_rating, team_one.three_pt_fg_shooting_pct)
team_two.two_pt_fg_shooting_pct = calculate_fg_pct_based_on_opp(team_two.off_rating, team_one.def_rating, team_two.two_pt_fg_shooting_pct)
team_two.three_pt_fg_shooting_pct = calculate_3p_fg_pct_based_on_opp(team_two.off_rating, team_one.def_rating, team_two.three_pt_fg_shooting_pct)
# Modify turnover percentages based on offensive and defensive team ratings
team_one.turnover_pct_per_play = calculate_turnover_pct_based_on_opp(team_one.off_rating, team_two.def_rating, team_one.turnover_pct_per_play)
team_two.turnover_pct_per_play = calculate_turnover_pct_based_on_opp(team_two.off_rating, team_one.def_rating, team_two.turnover_pct_per_play)
# Initialize matchup state
matchup = State(team_one_wins=0,
team_two_wins=0,
overtime_games=0,
games=0,
team_one_score=TimeSeries(),
team_two_score=TimeSeries(),
team_one=team_1,
team_two=team_2)
# Create TimeSeries object to track the game result difference
game_result_diff = TimeSeries()
# Loop through a game X number of times based on input parameter
for g in range(1, games+1):
matchup.games += 1
# Always re-initialize pace
team_one.pace = matchup_pace
team_two.pace = matchup_pace
gm_team_one, gm_team_one_stats = simulate_game(team_one)
gm_team_two, gm_team_two_stats = simulate_game(team_two)
if (gm_team_one.max() > gm_team_two.max()):
matchup.team_one_wins += 1
matchup.team_one_score[g] = gm_team_one.max()
matchup.team_two_score[g] = gm_team_two.max()
game_result_diff[g] = gm_team_one.max() - gm_team_two.max()
elif (gm_team_two.max() > gm_team_one.max()):
matchup.team_two_wins += 1
matchup.team_one_score[g] = gm_team_one.max()
matchup.team_two_score[g] = gm_team_two.max()
game_result_diff[g] = gm_team_one.max() - gm_team_two.max()
else:
# Tie: Send game to overtime
matchup.overtime_games += 1
score_tied = True
# Modify pace for 5 minutes
ot_pace = (matchup_pace / 48) * 5
team_one.pace = ot_pace
team_two.pace = ot_pace
while (score_tied):
gm_team_one, gm_team_one_stats = (simulate_game(team_one))
gm_team_two, gm_team_two_stats = (simulate_game(team_two))
if (gm_team_one.max() > gm_team_two.max()):
matchup.team_one_wins += 1
matchup.team_one_score[g] = gm_team_one.max()
matchup.team_two_score[g] = gm_team_two.max()
game_result_diff[g] = gm_team_one.max() - gm_team_two.max()
score_tied = False
elif (gm_team_two.max() > gm_team_one.max()):
matchup.team_two_wins += 1
matchup.team_one_score[g] = gm_team_one.max()
matchup.team_two_score[g] = gm_team_two.max()
game_result_diff[g] = gm_team_one.max() - gm_team_two.max()
score_tied = False
print("Matchup run time: %s seconds ---" % (time.time() - m_start_time))
return matchup, game_result_diff
# +
# Test map_team_stats_to_system
team_A = map_team_stats_to_system('DAL')
team_A
# +
# Test simulate_matchup_between_two_teams
matchup_results, pt_diffs = simulate_matchup_between_two_teams('MIL', 'LAL', 5)
print(matchup_results.team_one, "wins:", matchup_results.team_one_wins)
print(matchup_results.team_two, "wins:", matchup_results.team_two_wins)
print("Overtime games:", matchup_results.overtime_games)
print("Total Games:", matchup_results.games)
# -
# A plot of the score difference for each game of the two-team matchup simulation.
# +
pt_diffs_ser = pandas.Series(pt_diffs)
pt_diffs_ser.plot.hist(bins=20, alpha=0.5, label='MIL')
decorate(title='Distribution of Scores Difference',
xlabel='Difference',
ylabel='Frequency',
legend='MIL')
# -
pt_diffs.mean()
# Plot of the point distribution for both teams identified above.
# +
from matplotlib import pyplot
team_one_scores = pandas.Series(matchup_results.team_one_score)
team_two_scores = pandas.Series(matchup_results.team_two_score)
pyplot.hist(team_one_scores, bins=20, alpha=0.5, label=matchup_results.team_one)
pyplot.hist(team_two_scores, bins=20, alpha=0.5, label=matchup_results.team_two)
pyplot.legend(loc='upper right')
decorate(title='Distribution of Scores',
xlabel='Score',
ylabel='Frequency',
legend=None)
pyplot.show()
# -
# Plot of the point totals for both teams for all simulations
plot(matchup_results.team_one_score)
plot(matchup_results.team_two_score)
decorate(title='Score by Game Simulation',
xlabel='Game',
ylabel='Final Score',
legend=None)
# Following function determines the series winner based on the win totals
def determine_series_winner(team_one, team_two, results):
"""
Determine the series winner based on the results object from a matchup
"""
winner = team_two
if (results.team_one_wins > results.team_two_wins):
winner = team_one
return winner
# Utility function to output the results of a series outcome to show progress in the simulate_playoffs function along with the results of each series.
def output_playoff_series_result(team_one, team_two, results):
"""
Display to screen the playoff series result
"""
print("===== Series Result =====")
print(team_one, "wins:", results.team_one_wins)
print(team_two, "wins:", results.team_two_wins)
if (results.team_one_wins > results.team_two_wins):
print ("Series Winner:", team_one)
else:
print ("Series Winner:", team_two)
# simulate_playoffs function runs a tournament simulation for sixteen teams.
def simulate_playoffs(eastern_conf, western_conf, games_per_series_sim):
"""
Simulate NBA playoffs
"""
# Set number of game simulations per matchup
games = games_per_series_sim
# ===== Simulate first round =====
# Eastern conference
first_rd_east_series_one_results, first_rd_east_series_one_pt_diffs = simulate_matchup_between_two_teams(eastern_conf[0], eastern_conf[7], games)
first_rd_east_series_one_winner = determine_series_winner(eastern_conf[0], eastern_conf[7], first_rd_east_series_one_results)
output_playoff_series_result(eastern_conf[0], eastern_conf[7], first_rd_east_series_one_results)
first_rd_east_series_two_results, first_rd_east_series_two_pt_diffs = simulate_matchup_between_two_teams(eastern_conf[3], eastern_conf[4], games)
first_rd_east_series_two_winner = determine_series_winner(eastern_conf[3], eastern_conf[4], first_rd_east_series_two_results)
output_playoff_series_result(eastern_conf[3], eastern_conf[4], first_rd_east_series_two_results)
first_rd_east_series_three_results, first_rd_east_series_three_pt_diffs = simulate_matchup_between_two_teams(eastern_conf[1], eastern_conf[6], games)
first_rd_east_series_three_winner = determine_series_winner(eastern_conf[1], eastern_conf[6], first_rd_east_series_three_results)
output_playoff_series_result(eastern_conf[1], eastern_conf[6], first_rd_east_series_three_results)
first_rd_east_series_four_results, first_rd_east_series_four_pt_diffs = simulate_matchup_between_two_teams(eastern_conf[2], eastern_conf[5], games)
first_rd_east_series_four_winner = determine_series_winner(eastern_conf[2], eastern_conf[5], first_rd_east_series_four_results)
output_playoff_series_result(eastern_conf[2], eastern_conf[5], first_rd_east_series_four_results)
# Western conference
first_rd_west_series_one_results, first_rd_west_series_one_pt_diffs = simulate_matchup_between_two_teams(western_conf[0], western_conf[7], games)
first_rd_west_series_one_winner = determine_series_winner(western_conf[0], western_conf[7], first_rd_west_series_one_results)
output_playoff_series_result(western_conf[0], western_conf[7], first_rd_west_series_one_results)
first_rd_west_series_two_results, first_rd_west_series_two_pt_diffs = simulate_matchup_between_two_teams(western_conf[3], western_conf[4], games)
first_rd_west_series_two_winner = determine_series_winner(western_conf[3], western_conf[4], first_rd_west_series_two_results)
output_playoff_series_result(western_conf[3], western_conf[4], first_rd_west_series_two_results)
first_rd_west_series_three_results, first_rd_west_series_three_pt_diffs = simulate_matchup_between_two_teams(western_conf[1], western_conf[6], games)
first_rd_west_series_three_winner = determine_series_winner(western_conf[1], western_conf[6], first_rd_west_series_three_results)
output_playoff_series_result(western_conf[1], western_conf[6], first_rd_west_series_three_results)
first_rd_west_series_four_results, first_rd_west_series_four_pt_diffs = simulate_matchup_between_two_teams(western_conf[2], western_conf[5], games)
first_rd_west_series_four_winner = determine_series_winner(western_conf[2], western_conf[5], first_rd_west_series_four_results)
output_playoff_series_result(western_conf[2], western_conf[5], first_rd_west_series_four_results)
# ===== Simulate second round =====
# Eastern Conference
second_rd_east_series_one_results, second_rd_east_series_one_pt_diffs = simulate_matchup_between_two_teams(first_rd_east_series_one_winner, first_rd_east_series_two_winner, games)
second_rd_east_series_one_winner = determine_series_winner(first_rd_east_series_one_winner, first_rd_east_series_two_winner, second_rd_east_series_one_results)
output_playoff_series_result(first_rd_east_series_one_winner, first_rd_east_series_two_winner, second_rd_east_series_one_results)
second_rd_east_series_two_results, second_rd_east_series_two_pt_diffs = simulate_matchup_between_two_teams(first_rd_east_series_three_winner, first_rd_east_series_four_winner, games)
second_rd_east_series_two_winner = determine_series_winner(first_rd_east_series_three_winner, first_rd_east_series_four_winner, second_rd_east_series_two_results)
output_playoff_series_result(first_rd_east_series_three_winner, first_rd_east_series_four_winner, second_rd_east_series_two_results)
# Western Conference
second_rd_west_series_one_results, second_rd_west_series_one_pt_diffs = simulate_matchup_between_two_teams(first_rd_west_series_one_winner, first_rd_west_series_two_winner, games)
second_rd_west_series_one_winner = determine_series_winner(first_rd_west_series_one_winner, first_rd_west_series_two_winner, second_rd_west_series_one_results)
output_playoff_series_result(first_rd_west_series_one_winner, first_rd_west_series_two_winner, second_rd_west_series_one_results)
second_rd_west_series_two_results, second_rd_west_series_two_pt_diffs = simulate_matchup_between_two_teams(first_rd_west_series_three_winner, first_rd_west_series_four_winner, games)
second_rd_west_series_two_winner = determine_series_winner(first_rd_west_series_three_winner, first_rd_west_series_four_winner, second_rd_west_series_two_results)
output_playoff_series_result(first_rd_west_series_three_winner, first_rd_west_series_four_winner, second_rd_west_series_two_results)
# ===== Simulate conference finals =====
# Eastern Conference
third_rd_east_series_one_results, third_rd_east_series_one_pt_diffs = simulate_matchup_between_two_teams(second_rd_east_series_one_winner, second_rd_east_series_two_winner, games)
third_rd_east_series_one_winner = determine_series_winner(second_rd_east_series_one_winner, second_rd_east_series_two_winner, third_rd_east_series_one_results)
output_playoff_series_result(second_rd_east_series_one_winner, second_rd_east_series_two_winner, third_rd_east_series_one_results)
# Western Conference
third_rd_west_series_one_results, third_rd_west_series_one_pt_diffs = simulate_matchup_between_two_teams(second_rd_west_series_one_winner, second_rd_west_series_two_winner, games)
third_rd_west_series_one_winner = determine_series_winner(second_rd_west_series_one_winner, second_rd_west_series_two_winner, third_rd_west_series_one_results)
output_playoff_series_result(second_rd_west_series_one_winner, second_rd_west_series_two_winner, third_rd_west_series_one_results)
# ===== Simulate NBA finals =====
nba_finals_results, nba_finals_pt_diffs = simulate_matchup_between_two_teams(third_rd_east_series_one_winner, third_rd_west_series_one_winner, games)
nba_finals_winner = determine_series_winner(third_rd_east_series_one_winner, third_rd_west_series_one_winner, nba_finals_results)
output_playoff_series_result(third_rd_east_series_one_winner, third_rd_west_series_one_winner, nba_finals_results)
return nba_finals_winner
# Test output_playoff_series_result
output_playoff_series_result('DAL', 'NYK', matchup_results)
# ## Playoff simulation
#
# Set the variable 'simulations_per_series' appropriately. 101 should be sufficient. Remember, the higher the number, the longer the playoff simulation will take to complete.
# +
# Standings as of July 11, 2020
east = ['MIL', 'TOR', 'BOS', 'MIA', 'IND', 'PHI', 'BKN', 'ORL']
west = ['LAL', 'LAC', 'DEN', 'UTA', 'OKC', 'HOU', 'DAL', 'MEM']
simulations_per_series = 5
# Test simulate_playoffs
start_time = time.time()
simulate_playoffs(east, west, simulations_per_series)
print("--- %s seconds ---" % (time.time() - start_time))
| FinalProject/FinalProject_v3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from mikeio.dfs0 import Dfs0
from mikeio.eum import TimeStep, ItemInfo, EUMType, EUMUnit
from datetime import datetime, timedelta
# # Create a timeseries
# +
data = []
nt = 10
d1 = np.zeros(nt)
data.append(d1)
d2 = np.ones(nt)
data.append(d2)
items = [ItemInfo("Zeros", EUMType.Water_Level, EUMUnit.meter),
ItemInfo("Ones", EUMType.Discharge, EUMUnit.meter_pow_3_per_sec)]
dfs = Dfs0()
dfs.create(filename="test.dfs0", data=data, items=items, title="Zeros and ones",
start_time=datetime(2000,1,1),
timeseries_unit=TimeStep.DAY, dt=7)
# -
# ## From comma separated file
# +
import pandas as pd
df = pd.read_csv("../tests/testdata/co2-mm-mlo.csv", parse_dates=True,index_col='Date',na_values=-99.99)
df.head()
# -
# Remove missing values
df = df.dropna()
df = df[["Average","Trend"]]
df.plot()
# A dataframe with a datetimeindex can be used to create a dfs 0 with a non-equidistant time axis.
df.to_dfs0("mauna_loa_co2.dfs0")
# To get a equidistant time axis first interpolate to hourly values.
df_h = df.resample('h').interpolate()
df_h.to_dfs0("mauna_loa_co2_eq_1hr.dfs0")
# # Read a timeseries
res = dfs.read("test.dfs0")
res.items
res.time
res.data
# ## Or as a Pandas dataframe
# +
dfs0file = r"../tests/testdata/da_diagnostic.dfs0"
dfs = Dfs0()
df = dfs.to_dataframe(dfs0file)
df.head()
# -
dfs._dfs.FileInfo.TimeAxis.TimeAxisType
# +
dfs0file = r"../tests/testdata/random.dfs0"
dfs = Dfs0()
df = dfs.to_dataframe(dfs0file)
df.head()
# -
dfs._dfs.FileInfo.TimeAxis.TimeStep
# ## Create a timeseries with non-equidistant data
# +
data = []
d1 = np.random.uniform(low=0.0, high=5.0, size=5)
data.append(d1)
datetimes = [
datetime(2000, 1, 1, 0, 0),
datetime(2000, 1, 8, 0, 0),
datetime(2000, 1, 10, 0, 0),
datetime(2000, 2, 22, 0, 0),
datetime(2000, 11, 29, 0, 0)
]
dfs = Dfs0()
dfs.create(filename="neq.dfs0",
datetimes=datetimes,
data=data,
title="Non equidistant"
)
# -
# ## Create a timeseries with accumulated timestep
# ## Find correct eum units
EUMType.search("prec")
EUMType.Precipitation_Rate.units
# +
from DHI.Generic.MikeZero.DFS import DataValueType
n= 1000
d1 = np.random.random([n])
d2 = np.random.random([n])
data = []
data.append(d1)
data.append(d2)
start_time = datetime(2017, 1, 1)
time_vector = []
t = start_time
random_dt = np.random.choice([1.0,2.0],size=n)
for i in range(n):
t = t + timedelta(hours=random_dt[i])
time_vector.append(t)
title = 'Hello Test'
items = [ItemInfo("Water level",EUMType.Water_Level), # use default units
ItemInfo("Precipitation", EUMType.Precipitation_Rate)]
data_value_type = [DataValueType.Instantaneous, DataValueType.Accumulated]
dfs = Dfs0()
dfs.create(filename='accumulated.dfs0', data=data,
datetimes=time_vector,
items=items, title=title,
data_value_type=data_value_type)
# -
ds = dfs.read("accumulated.dfs0")
ds.items
# # Modify an existing timeseries
#
# The `write` method allows to modify the data without the need of specifying names, units etc. If you need to add variables, rename, change units, you must use `create` instead.
res = dfs.read("test.dfs0")
res
res['Ones']
# Modify the data in some way...
data = res.data
data[1] = data[1]*np.pi
data[1]
from shutil import copyfile
copyfile("test.dfs0","modified.dfs0")
dfs.write("modified.dfs0", data)
res = dfs.read("modified.dfs0")
res['Ones']
# The second item is modified.
res['Zeros']
# ## Convert units
#
# Read a file with waterlevel i meters.
# +
dfs = Dfs0()
filename = r"C:\Program Files (x86)\DHI\2020\MIKE Zero\Examples\MIKE_21\FlowModel_FM\HD\Oresund\Data\1993\Boundary_Conditions\waterlevel_viken.dfs0"
ds = dfs.read(filename)
ds.items
# -
import matplotlib.pyplot as plt
plt.plot(ds.time,ds.data[0])
plt.ylabel(ds.items[0].name);
# The aim is to convert this timeseries to feet (1m = 3.3 ft)
data = ds.data
data[0] = data[0]*3.3
# Which units are acceptable?
ds.items[0].type.units
# +
dfs = Dfs0()
items = ds.items
items[0].name = "Viken"
items[0].unit = EUMUnit.feet
dfs.create(filename='wl_feet.dfs0', data=ds.data,
start_time=ds.time[0],
dt = 1800,
items=items)
# -
# 
# ## Clean up
# +
import os
os.remove("test.dfs0")
os.remove("modified.dfs0")
os.remove("neq.dfs0")
os.remove("accumulated.dfs0")
os.remove("wl_feet.dfs0")
os.remove("mauna_loa_co2_eq_1hr.dfs0")
os.remove("mauna_loa_co2.dfs0")
# -
| notebooks/Dfs0 - Timeseries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
tf.__version__
tensor_a = tf.constant( "Hello, Tensorflow" )
tensor_a
tensor_a_str = tensor_a.numpy().decode( 'utf-8' )
tensor_a_str
from PIL import Image
| src/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Falkner-Skan
#
# As an example, let's solve the Falkner-Skan boundary layer equation - a familiar friend (or foe) to any battle-hardened aerodynamicist. The actual equation looks like this, as written in *Aerodynamics of Viscous Fluids* by <NAME> (internal MIT draft), Eq. 3.32:
#
# -----
#
# With $F, U, S$ as functions of $\eta$, and with $()'$ denoting a derivative w.r.t. $\eta$:
#
# $ F' = U $
#
# $ U' = S $
#
# $ S' = -\frac{1+a}{2} F S - a (1 - U^2) $
#
# with the following three boundary conditions:
#
# $ F(0) = 0 $
#
# $ U(0) = 0 $
#
# $ U(\infty) = 1 $, which we will approximate as $ U(10) = 1 $
#
# And here, $a$ is some parameter that we know describing the edge velocity profile $u_e \propto x^a$ , typically in the range $-0.0904 < a < 2$ or so.
#
# -----
#
# For the non-aerodynamicists reading this, don't worry if this doesn't mean much to you - the actual equation is fairly inconsequential. This serves as a good example of an ODE for several reasons:
#
# * It is nonlinear
# * It is higher-order (third-order, specifically)
# * It is a boundary value problem rather than an initial value problem, so explicit solution is difficult. (Note: explicit solution is not impossible; see the shooting method. However, for this problem, it is horribly unstable and the implicit solution that we will demonstrate is far superior.)
#
# If you haven't already, read the [notes on ODE solving in this folder's README](README.md).
#
# ## Solving the ODE
#
# Let's solve the Falkner-Skan ODE, for the value of our constant $a = 0.1$.
#
# First, we'll do boilerplate stuff that you all know by now, having read earlier tutorials:
# + pycharm={"name": "#%%\n"}
import aerosandbox as asb
import aerosandbox.numpy as np
opti = asb.Opti() # Initialize an optimization/analysis environment
a = opti.parameter(value=0.1) # Initialize constants
# -
# Now, let's define some derivatives:
# + pycharm={"name": "#%%\n"}
n_points = 100 # Number of discretization points
eta = np.linspace(0, 10, n_points) # Discretize eta from 0 to 10.
F = opti.variable( # Create a variable F that represents some function F(eta)
init_guess=eta + 10 / 3 * (1 - eta / 10) ** 3 # We get this by integrating our guess for U.
)
U = opti.derivative_of( # Create a variable U that represents a derivative of F(eta) with respect to eta.
variable=F,
with_respect_to=eta,
derivative_init_guess=1 - (1 - eta / 10) ** 2 # We guess that the velocity profile is quadratic.
)
S = opti.derivative_of( # Create a variable S that represents a derviative of U(eta) with respect to eta.
variable=U,
with_respect_to=eta,
derivative_init_guess=0.2 * (1 - eta / 10) # We get this by differentiating our guess for U.
)
# -
# Define the governing equations: (Note: derivatives between $F$, $U$, and $S$ were already constrained when they were initialized with the `opti.derivative_of()` syntax, so we just need to implement the last equation.)
# + pycharm={"name": "#%%\n"}
opti.constrain_derivative(
variable=S,
with_respect_to=eta,
derivative=-(1 + a) / 2 * F * S - a * (1 - U ** 2)
)
# -
# Then, define the boundary conditions:
# + pycharm={"name": "#%%\n"}
opti.subject_to([
F[0] == 0,
U[0] == 0,
U[-1] == 1,
])
# -
# And solve (flagging `verbose=False` to keep things tidy):
# + pycharm={"name": "#%%\n"}
sol = opti.solve(verbose=False)
# -
# Let's print a few of the results, and plot them:
# + pycharm={"name": "#%%\n"}
print("U(eta) at each of our values of eta in `eta`:")
print(
sol.value(U)
)
# + pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(palette=sns.color_palette("husl", 3))
fig, ax = plt.subplots(1, 1, figsize=(6.4, 4.8), dpi=200)
for var in ["F", "U", "S"]:
plt.plot(
sol.value(eval(var)),
eta,
label=var
)
plt.xlim(-0.1, 1.1)
plt.xlabel(r"Boundary Layer Parameter")
plt.ylabel(r"$\eta$")
plt.title(rf"Falkner-Skan Solution for $a = {sol.value(a)}$")
plt.tight_layout()
plt.legend()
plt.show()
# -
# Nice!
#
# ## Plotting Families of the ODE
#
# For kicks, we can also plot $U(\eta)$ for a bunch of different values of $a$:
# + pycharm={"name": "#%%\n"}
a_values = [-0.09044] + list(np.arange(0, 2.1, 0.1))
fig, ax = plt.subplots(1, 1, figsize=(6.4, 4.8), dpi=200)
colors = plt.cm.rainbow(np.linspace(1, 0, len(a_values)))
for i, value in enumerate(a_values):
opti.set_value(a, value)
sol = opti.solve(verbose=False)
plt.plot(
sol.value(U),
eta,
label=rf"$a = {sol.value(a):.1f}$" if i != 0 else rf"$a = {sol.value(a):.4f}$",
color=colors[i],
zorder=3 + len(a_values) - i
)
plt.xlim(-0.1, 1.1)
plt.xlabel(r"$U(\eta)$")
plt.ylabel(r"$\eta$")
plt.title(r"Falkner-Skan Solutions for Various $a$ Values")
plt.tight_layout()
plt.legend(ncol=3)
plt.show()
# -
# Cool. So far, this has all been pretty normal. But, let's change it up a bit.
#
# ## Doing Inverse Analysis
#
# Take a look at our solution above for $a \approx -0.0904$. Something interesting is happening here - this is an incipient separation flow. We can tell because the nondimensional shear $S(\eta) = dU/d\eta$ at the wall ($\eta = 0$) is going to zero.
#
# But our value of $-0.0904$ was just a guess, found by trial and error. Suppose now we want to ask the following question: at what exact value of $a$ does the flow separate, as indicated by the wall shear $S(0)$ going exactly to zero?
#
# If we had solved this ODE with traditional methods, this would be a big pain - we would need to add $a$ as an unknown and define another residual equation after differentiating with respect to $a$ before resolving.
#
# One might suggest trial and error instead - why not guess a series of values of $a$ and pick the best one, or do something clever by post-processing the solve? The problem is that for $a < -0.0904$ no solution exists, so trial and error is really tedious and difficult. This is common - often the forward problem is well-posed but the inverse problem is not. So, we need to get a bit clever:
#
# Here, we can solve the inverse problem super easily just by changing $a$ from a parameter to a variable and adding "zero wall shear" as an additional constraint:
# + pycharm={"name": "#%%\n"}
opti = asb.Opti() # Initialize an optimization/analysis environment
a = opti.variable(init_guess=0) # `a` is a variable now, initialized to a guess of 0 (the Blasius solution)
eta = np.linspace(0, 10, n_points) # Discretize eta from 0 to 10.
F = opti.variable( # Create a variable F that represents some function F(eta)
init_guess=eta + 10 / 3 * (1 - eta / 10) ** 3
)
U = opti.derivative_of( # Create a variable U that represents a derivative of F(eta) with respect to eta.
variable=F,
with_respect_to=eta,
derivative_init_guess=1 - (1 - eta / 10) ** 2
)
S = opti.derivative_of( # Create a variable S that represents a derviative of U(eta) with respect to eta.
variable=U,
with_respect_to=eta,
derivative_init_guess=0.2 * (1 - eta / 10)
)
opti.constrain_derivative(
variable=S,
with_respect_to=eta,
derivative=-(1 + a) / 2 * F * S - a * (1 - U ** 2)
)
opti.subject_to([
F[0] == 0,
U[0] == 0,
U[-1] == 1,
S[0] == 0, # We simply constrain the wall shear to be exactly 0.
])
# -
# Then, we solve again and print our answer for $a$:
# + pycharm={"name": "#%%\n"}
sol = opti.solve(verbose=False)
print(f"Value of `a` at incipient separation: {sol.value(a)}")
# + [markdown] pycharm={"name": "#%% md\n"}
# So, we can get the value of $a$ at incipient separation without any kind of manual fussing with residual Jacobians - it's all abstracted for us.
#
# This type of solve is called the *inverse solution*, and it comes up **all the time** in engineering.
#
# Another example of the inverse problem: we might analyze an airfoil at an angle of attack of 5 degrees and find that it has a lift coefficient of 0.6. Suppose then that we want to find the angle of attack that corresponds to a lift coefficient of 0.5 - in essense, *go backwards* from lift coefficient to angle of attack.
#
# Solving the inverse problem is often quite tedious without the abstracted approach used here, but it's easily implemented in AeroSandbox.
| tutorial/03 - Trajectory Optimization and Optimal Control/01 - Solving ODEs with AeroSandbox/01 - Falkner-Skan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
from itertools import cycle
import torch
import torch.nn as nn
import torch.optim as optim
from torchvision import datasets
from torch.autograd import Variable
from tensorboardX import SummaryWriter
from utils import weights_init
import matplotlib.pyplot as plt
from data_loader import MNIST_Paired
from torch.utils.data import DataLoader
from networks import Encoder, Decoder, Discriminator
from utils import imshow_grid, mse_loss, reparameterize, transform_config
import utils
# +
import argparse
parser = argparse.ArgumentParser()
# add arguments
parser.add_argument('--cuda', type=bool, default=True, help="run the following code on a GPU")
parser.add_argument('--batch_size', type=int, default=64, help="batch size for training")
parser.add_argument('--image_size', type=int, default=28, help="height and width of the image")
parser.add_argument('--num_channels', type=int, default=1, help="number of channels in the image")
parser.add_argument('--initial_learning_rate', type=float, default=0.0001, help="starting learning rate")
parser.add_argument('--style_dim', type=int, default=16, help="dimension of style latent space")
parser.add_argument('--class_dim', type=int, default=16, help="dimension of class latent space")
parser.add_argument('--num_classes', type=int, default=10, help="number of classes on which the data set trained")
# arguments to control per iteration training of architecture
parser.add_argument('--generator_times', type=int, default=2, help="number of times the generator is run")
parser.add_argument('--discriminator_times', type=int, default=1, help="number of times the discriminator is run")
parser.add_argument(
'--discriminator_limiting_accuracy', type=float, default=0.8, help="acc. at which discriminator is stopped training"
)
parser.add_argument('--beta_1', type=float, default=0.5, help="default beta_1 val for adam")
parser.add_argument('--beta_2', type=float, default=0.999, help="default beta_2 val for adam")
# paths to save models
parser.add_argument('--encoder_save', type=str, default='encoder', help="model save for encoder")
parser.add_argument('--decoder_save', type=str, default='decoder', help="model save for decoder")
parser.add_argument('--discriminator_save', type=str, default='discriminator', help="model save for discriminator")
parser.add_argument('--log_file', type=str, default='log.txt', help="text file to save training logs")
parser.add_argument('--load_saved', type=bool, default=False, help="flag to indicate if a saved model will be loaded")
parser.add_argument('--start_epoch', type=int, default=0, help="flag to set the starting epoch for training")
parser.add_argument('--end_epoch', type=int, default=50, help="flag to indicate the final epoch of training")
FLAGS = parser.parse_known_args()[0]
# +
encoder = Encoder(style_dim=FLAGS.style_dim, class_dim=FLAGS.class_dim)
encoder.apply(weights_init)
decoder = Decoder(style_dim=FLAGS.style_dim, class_dim=FLAGS.class_dim)
decoder.apply(weights_init)
# load saved models
encoder.load_state_dict(torch.load(os.path.join('checkpoints', FLAGS.encoder_save), map_location=torch.device('cpu')))
decoder.load_state_dict(torch.load(os.path.join('checkpoints', FLAGS.decoder_save), map_location=torch.device('cpu')))
# +
X_1 = torch.FloatTensor(FLAGS.batch_size, FLAGS.num_channels, FLAGS.image_size, FLAGS.image_size)
X_2 = torch.FloatTensor(FLAGS.batch_size, FLAGS.num_channels, FLAGS.image_size, FLAGS.image_size)
paired_mnist = MNIST_Paired(root='mnist', download=True, train=True, transform=transform_config)
loader = cycle(DataLoader(paired_mnist, batch_size=FLAGS.batch_size, shuffle=True, num_workers=0, drop_last=True))
image_batch_1, image_batch_2, _ = next(loader)
# shuffle batch1
image_batch_1 = image_batch_1[torch.randperm(FLAGS.batch_size)]
# +
X_1.copy_(image_batch_1)
X_2.copy_(image_batch_2)
style_mu_1, style_logvar_1, class_1 = encoder(Variable(X_1))
style_1 = reparameterize(training=False, mu=style_mu_1, logvar=style_logvar_1)
_, __, class_2 = encoder(Variable(X_2))
reconstructed_X_1 = decoder(style_1, class_1)
reconstructed_X_2 = decoder(style_1, class_2)
# +
def get_image_list(batch):
return [batch[i, 0, :, :].detach().numpy() for i in range(batch.shape[0])]
re_x1_im = get_image_list(reconstructed_X_1)
re_x2_im = get_image_list(reconstructed_X_2)
x1_im = get_image_list(X_1)
x2_im = get_image_list(X_2)
# -
utils.imshow_grid(x1_im)
utils.imshow_grid(x2_im)
utils.imshow_grid(re_x1_im)
utils.imshow_grid(re_x2_im)
| check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from keras.models import Model
from keras.layers import Input
from keras.layers.pooling import MaxPooling3D
from keras import backend as K
import json
from collections import OrderedDict
def format_decimal(arr, places=6):
return [round(x * 10**places) / 10**places for x in arr]
DATA = OrderedDict()
# ### MaxPooling3D
# **[pooling.MaxPooling3D.0] input 4x4x4x2, pool_size=(2, 2, 2), strides=None, padding='valid', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=None, padding='valid', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(290)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.0'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.1] input 4x4x4x2, pool_size=(2, 2, 2), strides=(1, 1, 1), padding='valid', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=(1, 1, 1), padding='valid', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(291)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.1'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.2] input 4x5x2x3, pool_size=(2, 2, 2), strides=(2, 1, 1), padding='valid', data_format='channels_last'**
# +
data_in_shape = (4, 5, 2, 3)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=(2, 1, 1), padding='valid', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(282)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.2'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.3] input 4x4x4x2, pool_size=(3, 3, 3), strides=None, padding='valid', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(3, 3, 3), strides=None, padding='valid', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(283)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.3'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.4] input 4x4x4x2, pool_size=(3, 3, 3), strides=(3, 3, 3), padding='valid', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(3, 3, 3), strides=(3, 3, 3), padding='valid', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(284)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.4'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.5] input 4x4x4x2, pool_size=(2, 2, 2), strides=None, padding='same', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=None, padding='same', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(285)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.5'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.6] input 4x4x4x2, pool_size=(2, 2, 2), strides=(1, 1, 1), padding='same', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=(1, 1, 1), padding='same', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(286)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.6'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.7] input 4x5x4x2, pool_size=(2, 2, 2), strides=(1, 2, 1), padding='same', data_format='channels_last'**
# +
data_in_shape = (4, 5, 4, 2)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=(1, 2, 1), padding='same', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(287)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.7'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.8] input 4x4x4x2, pool_size=(3, 3, 3), strides=None, padding='same', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(3, 3, 3), strides=None, padding='same', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(288)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.8'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.9] input 4x4x4x2, pool_size=(3, 3, 3), strides=(3, 3, 3), padding='same', data_format='channels_last'**
# +
data_in_shape = (4, 4, 4, 2)
L = MaxPooling3D(pool_size=(3, 3, 3), strides=(3, 3, 3), padding='same', data_format='channels_last')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(289)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.9'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.10] input 2x3x3x4, pool_size=(3, 3, 3), strides=(2, 2, 2), padding='valid', data_format='channels_first'**
# +
data_in_shape = (2, 3, 3, 4)
L = MaxPooling3D(pool_size=(3, 3, 3), strides=(2, 2, 2), padding='valid', data_format='channels_first')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(290)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.10'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.11] input 2x3x3x4, pool_size=(3, 3, 3), strides=(1, 1, 1), padding='same', data_format='channels_first'**
# +
data_in_shape = (2, 3, 3, 4)
L = MaxPooling3D(pool_size=(3, 3, 3), strides=(1, 1, 1), padding='same', data_format='channels_first')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(291)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.11'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[pooling.MaxPooling3D.12] input 3x4x4x3, pool_size=(2, 2, 2), strides=None, padding='valid', data_format='channels_first'**
# +
data_in_shape = (3, 4, 4, 3)
L = MaxPooling3D(pool_size=(2, 2, 2), strides=None, padding='valid', data_format='channels_first')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(292)
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['pooling.MaxPooling3D.12'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# ### export for Keras.js tests
# +
import os
filename = '../../../test/data/layers/pooling/MaxPooling3D.json'
if not os.path.exists(os.path.dirname(filename)):
os.makedirs(os.path.dirname(filename))
with open(filename, 'w') as f:
json.dump(DATA, f)
# -
print(json.dumps(DATA))
| notebooks/layers/pooling/MaxPooling3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: kgtk-env
# language: python
# name: kgtk-env
# ---
# # Generating Useful Wikidata Files
#
# This notebook generates files that contain derived data that is useful in many applications. The input to the notebook is the full Wikidata or a subset of Wikidata. It also works for arbutrary KGs as long as they follow the representation requirements of Wikidata:
#
# - the *instance of* relation is represented using the `P31` property
# - the *subclass of* relation is represented using the `P279` property
# - all properties declare a datatype, and the data types must be one of the datatypes in Wikidata.
#
# Inputs:
#
# - `claims_file`: contains all statements, which consist of edges `node1/label/node2` where `label` is a property in Wikidata (e.g., sitelinks, labels, aliases and description are not in the claims file.
# - `item_file`: the subset of the `claims_file` consistin of edges for property of data type `wikibase-item`
# - `label_file`, `alias_file` and `description_file` containing labels, aliases and descriptions. It is assume that these files contain the labels, aliases and descriptions of all nodes appearing in the claims file. Users may provide these files for specific languages only.
#
# Outputs:
#
# - **Instance of (P31):** `derived.P31.tsv.gz` contains all the `instance of (P31)` edges present in the claims file.
# - **Subclass of (P279):** `derived.P279.tsv.gz` contains all the `subclass of (P279)` edges present in the claims file.
# - **Is A (isa):** `derived.isa.tsv.gz` contains edges `node`isa/node2` where either `node1/P31/node2` or `node1/P279/node2`
# - **Closure of subclass of (P279star):** `derived.P279star.tsv.gz` contains edges `node1/P279star/node2` where `node2` is reachable from `node1` via zero or more hops using the `P279` property. Note that for example, `Q44/P279star/Q44`. An example when this file is useful is when you want to find all the instance of a class, including instances of subclasses of the given class.
# - **In/out degrees:** `metadata.out_degree.tsv.gz` contains the out degree of every node, and `metadata.in_degree.tsv.gz` contains the in degree of every node.
# - **Pagerank:** outputs page rank on the directed graph in `metadata.pagerank.directed.tsv.gz` and page rank of the directed graph in `metadata.pagerank.undirected.tsv.gz`.
# ### Batch Invocation
# Example batch command. The second argument is a notebook where the output will be stored. You can load it to see progress.
#
# ```
# papermill Wikidata\ Useful\ Files.ipynb useful-files.out.ipynb \
# -p claims_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/all.tsv.gz \
# -p label_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/part.label.en.tsv.gz \
# -p item_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/part.wikibase-item.tsv.gz \
# -p property_item_file = /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/part.property.wikibase-item.tsv.gz \
# -p output_path <local folder> \
# -p output_folder useful_files_v4 \
# -p temp_folder temp.useful_files_v4 \
# -p delete_database no
# -p languages es,ru,zh-cn
# ```
# + tags=["parameters"]
# Parameters
# Folder on local machine where to create the output and temporary folders
output_path = "/Users/pedroszekely/Downloads/kypher"
# The names of the output and temporary folders
output_folder = "useful_wikidata_files_v4"
temp_folder = "temp.useful_wikidata_files_v4"
# The location of input files
wiki_root_folder = "/Volumes/GoogleDrive/Shared drives/KGTK/datasets/wikidata-20200803-v4/"
claims_file = "claims.tsv.gz"
label_file = "labels.en.tsv.gz"
alias_file = "aliases.en.tsv.gz"
description_file = "descriptions.en.tsv.gz"
item_file = "claims.wikibase-item.tsv.gz"
label_all = "labels.tsv.gz"
alias_all = "aliases.tsv.gz"
description_all = "descriptions.tsv.gz"
# Location of the cache database for kypher
cache_path = "/Users/pedroszekely/Downloads/kypher/temp.useful_wikidata_files_v4"
# Whether to delete the cache database
delete_database = False
# Whether to compute pagerank as it may not run on the laptop
compute_pagerank = False
languages = ''
# -
languages = languages.split(',')
# +
import io
import os
import subprocess
import sys
import numpy as np
import pandas as pd
import altair as alt
# -
# ## Set up environment and folders to store the files
#
# - `OUT` folder where the output files go
# - `TEMP` folder to keep temporary files , including the database
# - `kgtk` shortcut to invoke the kgtk software
# - `kypher` shortcut to invoke `kgtk query with the cache database
# - `CLAIMS` the `all.tsv` file of wikidata that contains all edges except label/alias/description
# - `LABELS` the file with the English labels
# - `ITEMS` the wikibase-item file (currently does not include node1 that are properties so for now we need the net file
# - `STORE` location of the cache file
if cache_path:
os.environ['STORE'] = "{}/wikidata.sqlite3.db".format(cache_path)
else:
os.environ['STORE'] = "{}/{}/wikidata.sqlite3.db".format(output_path, temp_folder)
os.environ['OUT'] = "{}/{}".format(output_path, output_folder)
os.environ['TEMP'] = "{}/{}".format(output_path, temp_folder)
os.environ['kgtk'] = "kgtk"
os.environ['kgtk'] = "time kgtk --debug"
os.environ['kypher'] = "time kgtk --debug query --graph-cache " + os.environ['STORE']
os.environ['CLAIMS'] = wiki_root_folder + claims_file
os.environ['LABELS'] = wiki_root_folder + label_file
os.environ['ALIASES'] = wiki_root_folder + alias_file
os.environ['DESCRIPTIONS'] = wiki_root_folder + description_file
os.environ['ITEMS'] = wiki_root_folder + item_file
# Echo the variables to see if they are all set correctly
# !echo $OUT
# !echo $TEMP
# !echo $kgtk
# !echo $kypher
# !echo $CLAIMS
# !echo $LABELS
# !echo $ALIASES
# !echo $LABELS
# !echo $DESCRIPTIONS
# !echo $STORE
# !alias col="column -t -s $'\t' "
# Go to the output directory and create the subfolders for the output files and the temporary files
# cd $output_path
# !mkdir -p $OUT
# !mkdir -p $TEMP
# Clean up the output and temp folders before we start
# +
# # !rm $OUT/*.tsv $OUT/*.tsv.gz
# # !rm $TEMP/*.tsv $TEMP/*.tsv.gz
# -
if delete_database:
print("Deleteddatabase")
# !rm $STORE
# !ls -l $OUT
# !ls $TEMP
# !ls -l "$CLAIMS"
# !ls -l "$LABELS"
# !ls -l "$ALIASES"
# !ls -l "$LABELS"
# !ls -l "$DESCRIPTIONS"
# !ls $STORE
# !zcat < "$CLAIMS" | head | col
# ### Preview the input files
# It is always a good practice to peek a the files to make sure the column headings are what we expect
# !$kypher -i "$CLAIMS" --limit 10 | col
# Force creation of the index on the label column
# !$kypher -i "$CLAIMS" -o - \
# --match '(i)-[:P31]->(c)' \
# --limit 5 \
# | column -t -s $'\t'
# Force creation of the index on the node2 column
# !$kypher -i "$CLAIMS" -o - \
# --match '(i)-[r]->(:Q5)' \
# --limit 5 \
# | column -t -s $'\t'
# ### Count the number of edges
# Counting takes a long time
# !$kypher -i "$CLAIMS" \
# --match '()-[r]->()' \
# --return 'count(r) as count' \
# --limit 10
# ### Get labels, aliases and descriptions for other languages
for lang in languages:
cmd = f"kgtk --debug query --graph-cache {os.environ['STORE']} -i {wiki_root_folder}{label_all} -o {output_path}/{output_folder}/labels.{lang}.tsv.gz --match '(n1)-[l:label]->(n2)' --where 'n2.kgtk_lqstring_lang_suffix = \"{lang}\"' --return 'n1, l.label, n2, l.id' "
# !{cmd}
for lang in languages:
cmd = f"kgtk --debug query --graph-cache {os.environ['STORE']} -i {wiki_root_folder}{alias_all} -o {output_path}/{output_folder}/aliases.{lang}.tsv.gz --match '(n1)-[l:alias]->(n2)' --where 'n2.kgtk_lqstring_lang_suffix = \"{lang}\"' --return 'n1, l.label, n2, l.id' "
# !{cmd}
for lang in languages:
cmd = f"kgtk --debug query --graph-cache {os.environ['STORE']} -i {wiki_root_folder}{description_all} -o {output_path}/{output_folder}/descriptions.{lang}.tsv.gz --match '(n1)-[l:description]->(n2)' --where 'n2.kgtk_lqstring_lang_suffix = \"{lang}\"' --return 'n1, l.label, n2, l.id' "
# !{cmd}
# ### Create the P31 and P279 files
# Create the `P31` file
# !$kypher -i "$CLAIMS" -o $OUT/derived.P31.tsv.gz \
# --match '(n1)-[l:P31]->(n2)' \
# --return 'l, n1, l.label, n2'
# Create the P279 file
# !gzcat $OUT/derived.P31.tsv.gz | head | col
# !$kypher -i "$CLAIMS" -o $OUT/derived.P279.tsv.gz \
# --match '(n1)-[l:P279]->(n2)' \
# --return 'l, n1, l.label, n2'
# ### Create the file that contains all nodes reachable via P279 starting from a node2 in P31 or a node1 in P279
# First compute the roots
# !$kypher -i $OUT/derived.P279.tsv.gz -o $TEMP/P279.n1.tsv.gz \
# --match '(n1)-[l]->()' \
# --return 'n1 as id'
# !$kypher -i $OUT/derived.P31.tsv.gz -o $TEMP/P31.n2.tsv.gz \
# --match '()-[l]->(n2)' \
# --return 'n2 as id'
# !$kgtk cat --mode NONE -i $TEMP/P31.n2.tsv.gz $TEMP/P279.n1.tsv.gz \
# | gzip > $TEMP/P279.roots.1.tsv.gz
# !$kgtk sort2 --mode NONE --column id -i $TEMP/P279.roots.1.tsv.gz \
# | gzip > $TEMP/P279.roots.2.tsv.gz
# We have lots of duplicates
# !zcat < $TEMP/P279.roots.2.tsv.gz | head
# !$kgtk compact -i $TEMP/P279.roots.2.tsv.gz --mode NONE \
# --presorted \
# --columns id \
# > $TEMP/P279.roots.tsv
# Now we can invoke the reachable-nodes command
# !$kgtk reachable-nodes \
# --rootfile $TEMP/P279.roots.tsv \
# --selflink \
# -i $OUT/derived.P279.tsv.gz \
# | gzip > $TEMP/P279.reachable.tsv.gz
# !zcat < $TEMP/P279.reachable.tsv.gz | head | col
# The reachable-nodes command produces edges labeled `reachable`, so we need one command to rename them.
# !$kypher -i $TEMP/P279.reachable.tsv.gz -o $TEMP/P279star.1.tsv.gz \
# --match '(n1)-[]->(n2)' \
# --return 'n1, "P279star" as label, n2 as node2'
# Now we can concatenate these files to produce the final output
# !$kgtk sort2 -i $TEMP/P279star.1.tsv.gz -o $TEMP/P279star.2.tsv.gz
# Make sure there are no duplicates
# !$kgtk compact --presorted -i $TEMP/P279star.2.tsv.gz -o $TEMP/P279star.3.tsv.gz
# Add ids
# !$kgtk add-id --id-style node1-label-node2-num -i $TEMP/P279star.3.tsv.gz -o $OUT/derived.P279star.tsv.gz
# !zcat < $OUT/derived.P279star.tsv.gz | head | col
# This is how we would do the typical `?item P31/P279* ?class` in Kypher.
# The example shows how to get all the counts of instances of subclasses of city (Q515).
# !$kypher -i $OUT/derived.P31.tsv.gz -i $OUT/derived.P279star.tsv.gz -i "$LABELS" \
# --match 'P31: (n1)-[:P31]->(c), P279star: (c)-[]->(:Q515), label: (n1)-[:label]->(label), label: (c)-[:label]->(c_label)' \
# --return 'distinct c as class, count(c) as count, c_label as `class name`, n1 as instance, label as `label`' \
# --order-by 'count(c) desc, c, n1' \
# --limit 10 \
# | col
# Illustrate that it is indeed `P279*`
# !$kypher -i $OUT/derived.P31.tsv.gz -i $OUT/derived.P279star.tsv.gz -i "$LABELS" \
# --match 'P31: (n1)-[:P31]->(c), P279star: (c)-[]->(:Q63440326), label: (n1)-[:label]->(label), label: (c)-[:label]->(c_label)' \
# --return 'distinct c as class, c_label as `class name`, n1 as instance, label as `label`' \
# --order-by 'c, n1' \
# --limit 10 \
# | col
# Test that `P279star` is indeed star
# !$kypher -i $OUT/derived.P279star.tsv.gz \
# --match '(n1:Q44)-[:P279star]->(n2:Q44)'
# ### Create a file to do generalized Is-A queries
# The idea is that `(n1)-[:isa]->(n2)` when `(n1)-[:P31]->(n2)` or `(n1)-[:P279]->(n2)`
#
# We do this by concatenating the files and renaming the relation
# !$kgtk cat -i $OUT/derived.P31.tsv.gz $OUT/derived.P279.tsv.gz \
# | gzip > $TEMP/isa.1.tsv.gz
# !$kypher -i $TEMP/isa.1.tsv.gz -o $OUT/derived.isa.tsv.gz \
# --match '(n1)-[]->(n2)' \
# --return 'n1, "isa" as label, n2'
# Example of how to use the `isa` relation
# !$kypher -i $OUT/derived.isa.tsv.gz -i $OUT/derived.P279star.tsv.gz -i "$LABELS" -o - \
# --match 'isa: (n1)-[l:isa]->(c), P279star: (c)-[]->(:Q44), label: (n1)-[:label]->(label)' \
# --return 'distinct n1, l.label, "Q44" as node2, label as n1_label' \
# --limit 10 \
# | col
# ### Create files with `isa/P279* and P31/P279*`
# This file is useful to find all nodes that are below a q-node via P279 or isa.
#
# > These files are very large and take many hours to compute
# !$kypher -i "$CLAIMS" -i "$P279STAR" -i "$ISA" \
# --match '\
# isa: (n1)-[]->(n2), \
# P279star: (n2)-[]->(n3)' \
# --return 'distinct n1 as node1, "isa_star" as label, n3 as node2' \
# -o "$TEMP"/derived.isastar_1.tsv.gz
# Now add ids and sort it
# !$kgtk add-id --id-style wikidata -i "$TEMP"/derived.isastar_1.tsv.gz \
# / sort2 -o "$OUT"/derived.isastar.tsv.gz
# It is very big
# !zcat < "$OUT"/derived.isastar.tsv.gz | wc
# Also calculate the same file by for P31/P279*
# !$kypher -i "$CLAIMS" -i "$P279STAR" \
# --match '\
# claims: (n1)-[:P31]->(n2), \
# P279star: (n2)-[]->(n3)' \
# --return 'distinct n1 as node1, "P31P279star" as label, n3 as node2' \
# -o "$TEMP"/derived.P31P279star.gz
# Add ids and sort it
# !$kgtk add-id --id-style wikidata -i "$TEMP"/derived.P31P279star.gz \
# / sort2 -o "$OUT"/derived.P31P279star.tsv.gz
# It is also very big
# !zcat < "$OUT"/derived.P31P279star.tsv.gz | wc
# ## Compute pagerank
# Now compute pagerank. These commands will exceed 16GB memory for graphs containing over 25 million nodes.
if compute_pagerank:
# !$kgtk graph-statistics -i "$ITEMS" -o $OUT/metadata.pagerank.directed.tsv.gz \
# --page-rank-property directed_pagerank \
# --pagerank --statistics-only \
# --log $TEMP/metadata.pagerank.directed.summary.txt
if compute_pagerank:
# !cat $TEMP/metadata.pagerank.directed.summary.txt
if compute_pagerank:
# !$kgtk graph-statistics -i "$ITEMS" -o $OUT/metadata.pagerank.undirected.tsv.gz \
# --page-rank-property undirected_pagerank \
# --pagerank --statistics-only --undirected \
# --log $TEMP/metadata.pagerank.undirected.summary.txt
if compute_pagerank:
# !cat $TEMP/metadata.pagerank.undirected.summary.txt
# ## Compute Degrees
# Kypher can compute the out degree by counting the node2s for each node1
# !$kypher -i "$CLAIMS" -o $TEMP/metadata.out_degree.tsv.gz \
# --match '(n1)-[l]->()' \
# --return 'distinct n1 as node1, count(distinct l) as node2, "out_degree" as label'
# !$kgtk add-id --id-style node1-label-node2-num -i $TEMP/metadata.out_degree.tsv.gz \
# / sort2 -o $OUT/metadata.out_degree.tsv.gz
# To count the in-degree we only care when the node2 is a wikibase-item
# BUG in kypher, sometimes the following command will not work, as in we'll see multilple rows for a Qnode, which is
# fixable by deleting cache
# !$kypher -i "$CLAIMS" -o $TEMP/metadata.in_degree.tsv.gz \
# --match '()-[l]->(n2 {`wikidatatype`:"wikibase-item"})' \
# --return 'distinct n2 as node1, count(distinct l) as node2, "in_degree" as label' \
# --order-by 'n2'
# +
# rename columns before adding ids, KGTK will complain
# df = pd.read_csv('{}/metadata.in_degree.tsv.gz'.format(os.environ['TEMP']), sep='\t')
# df = df.rename(columns={"node2": "node1"})
# df = df.rename(columns={'count(DISTINCT graph_1_c1."id")': "node2"})
# df.to_csv('{}/metadata.in_degree.1.tsv.gz'.format(os.environ['TEMP']), sep='\t', index=False)
# -
# !$kgtk add-id --id-style node1-label-node2-num -i $TEMP/metadata.in_degree.tsv.gz \
# / sort2 -o $OUT/metadata.in_degree.tsv.gz
# !zcat < $OUT/metadata.in_degree.tsv.gz | head | col
# Calculate the distribution so we can make a nice chart
# !$kypher -i $OUT/metadata.in_degree.tsv.gz -o $OUT/statistics.in_degree.distribution.tsv \
# --match '(n1)-[]->(n2)' \
# --return 'distinct n2 as in_degree, count(distinct n1) as count, "count" as label' \
# --order-by 'cast(n2, integer)'
# !head $OUT/metadata.in_degree.distribution.tsv | col
# !$kypher -i $OUT/metadata.out_degree.tsv.gz -o $OUT/statistics.out_degree.distribution.tsv \
# --match '(n1)-[]->(n2)' \
# --return 'distinct n2 as out_degree, count(distinct n1) as count, "count" as label' \
# --order-by 'cast(n2, integer)'
# Draw some charts
# +
data = pd.read_csv(
os.environ["OUT"] + "/statistics.in_degree.distribution.tsv", sep="\t"
)
alt.Chart(data).mark_circle(size=60).encode(
x=alt.X("in_degree", scale=alt.Scale(type="log")),
y=alt.Y("count", scale=alt.Scale(type="log"), title="count of nodes"),
tooltip=["in_degree", "count"],
).interactive().properties(title="Distribution of In Degree")
# +
data = pd.read_csv(
os.environ["OUT"] + "/statistics.out_degree.distribution.tsv", sep="\t"
)
alt.Chart(data).mark_circle(size=60).encode(
x=alt.X("out_degree", scale=alt.Scale(type="log")),
y=alt.Y("count", scale=alt.Scale(type="log"), title="count of nodes"),
tooltip=["out_degree", "count"],
).interactive().properties(title="Distribution of Out Degree")
# -
# ## Summary of results
# !ls -lh $OUT/*
# Highest page rank
if compute_pagerank:
# !$kypher -i $OUT/metadata.pagerank.undirected.tsv.gz -i "$LABELS" -o - \
# --match 'pagerank: (n1)-[:undirected_pagerank]->(page_rank), label: (n1)-[:label]->(label)' \
# --return 'distinct n1, label as label, page_rank as `undirected page rank' \
# --order-by 'page_rank desc' \
# --limit 10 \
# | col
| use-cases/Wikidata Useful Files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D3_ModelFitting/W1D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Tutorial 1: Linear regression with MSE
# **Week 1, Day 3: Model Fitting**
#
# **By Neuromatch Academy**
#
# **Content creators**: <NAME>, <NAME>, <NAME> with help from <NAME>
#
# **Content reviewers**: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
#
#
#
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ___
# # Tutorial Objectives
#
# *Estimated timing of tutorial: 30 minutes*
#
# This is Tutorial 1 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
#
# In this tutorial, we will learn how to fit simple linear models to data.
# - Learn how to calculate the mean-squared error (MSE)
# - Explore how model parameters (slope) influence the MSE
# - Learn how to find the optimal model parameter using least-squares optimization
#
# ---
#
# **acknowledgements:**
# - we thank <NAME>, much of today's tutorials are inspired by exercises asigned in his mathtools class.
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/2mkq4/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# ---
# # Setup
# + cellView="both"
import numpy as np
import matplotlib.pyplot as plt
# + cellView="form"
#@title Figure Settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form"
#@title Plotting Functions
def plot_observed_vs_predicted(x, y, y_hat, theta_hat):
""" Plot observed vs predicted data
Args:
x (ndarray): observed x values
y (ndarray): observed y values
y_hat (ndarray): predicted y values
theta_hat (ndarray):
"""
fig, ax = plt.subplots()
ax.scatter(x, y, label='Observed') # our data scatter plot
ax.plot(x, y_hat, color='r', label='Fit') # our estimated model
# plot residuals
ymin = np.minimum(y, y_hat)
ymax = np.maximum(y, y_hat)
ax.vlines(x, ymin, ymax, 'g', alpha=0.5, label='Residuals')
ax.set(
title=fr"$\hat{{\theta}}$ = {theta_hat:0.2f}, MSE = {mse(x, y, theta_hat):.2f}",
xlabel='x',
ylabel='y'
)
ax.legend()
# -
# ---
# # Section 1: Mean Squared Error (MSE)
# + cellView="form"
# @title Video 1: Linear Regression & Mean Squared Error
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1tA411e7NW", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="HumajfjJ37E", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# This video covers a 1D linear regression and mean squared error.
#
# <details>
# <summary> <font color='blue'>Click here for text recap of video </font></summary>
#
# **Linear least squares regression** is an old but gold optimization procedure that we are going to use for data fitting. Least squares (LS) optimization problems are those in which the objective function is a quadratic function of the
# parameter(s) being optimized.
#
# Suppose you have a set of measurements: for each data point or measurement, you have $y_{i}$ (the "dependent" variable) obtained for a different input value, $x_{i}$ (the "independent" or "explanatory" variable). Suppose we believe the measurements are proportional to the input values, but are corrupted by some (random) measurement errors, $\epsilon_{i}$, that is:
#
# \begin{align}
# y_{i}= \theta x_{i}+\epsilon_{i}
# \end{align}
#
# for some unknown slope parameter $\theta.$ The least squares regression problem uses **mean squared error (MSE)** as its objective function, it aims to find the value of the parameter $\theta$ by minimizing the average of squared errors:
#
# \begin{align}
# \min _{\theta} \frac{1}{N}\sum_{i=1}^{N}\left(y_{i}-\theta x_{i}\right)^{2}
# \end{align}
# We will now explore how MSE is used in fitting a linear regression model to data. For illustrative purposes, we will create a simple synthetic dataset where we know the true underlying model. This will allow us to see how our estimation efforts compare in uncovering the real model (though in practice we rarely have this luxury).
#
# First we will generate some noisy samples $x$ from [0, 10) along the line $y = 1.2x$ as our dataset we wish to fit a model to.
# + cellView="form"
# @title
# @markdown Execute this cell to generate some simulated data
# setting a fixed seed to our random number generator ensures we will always
# get the same psuedorandom number sequence
np.random.seed(121)
# Let's set some parameters
theta = 1.2
n_samples = 30
# Draw x and then calculate y
x = 10 * np.random.rand(n_samples) # sample from a uniform distribution over [0,10)
noise = np.random.randn(n_samples) # sample from a standard normal distribution
y = theta * x + noise
# Plot the results
fig, ax = plt.subplots()
ax.scatter(x, y) # produces a scatter plot
ax.set(xlabel='x', ylabel='y');
# -
# Now that we have our suitably noisy dataset, we can start trying to estimate the underlying model that produced it. We use MSE to evaluate how successful a particular slope estimate $\hat{\theta}$ is for explaining the data, with the closer to 0 the MSE is, the better our estimate fits the data.
# ## Coding Exercise 1: Compute MSE
#
# In this exercise you will implement a method to compute the mean squared error for a set of inputs $\mathbf{x}$, measurements $\mathbf{y}$, and slope estimate $\hat{\theta}$. Here, $\mathbf{x}$ and $\mathbf{y}$ are vectors of data points. We will then compute and print the mean squared error for 3 different choices of theta.
#
# As a reminder, the equation for computing the estimated y for a single data point is:
#
# $$\hat{y}_{i}= \theta x_{i}$$
#
# and for mean squared error is:
#
# \begin{align}
# \min _{\theta} \frac{1}{N}\sum_{i=1}^{N}\left(y_{i}-\hat{y}_i\right)^{2}
# \end{align}
# +
def mse(x, y, theta_hat):
"""Compute the mean squared error
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
theta_hat (float): An estimate of the slope parameter
Returns:
float: The mean squared error of the data with the estimated parameter.
"""
####################################################
## TODO for students: compute the mean squared error
# Fill out function and remove
raise NotImplementedError("Student exercise: compute the mean squared error")
####################################################
# Compute the estimated y
y_hat = ...
# Compute mean squared error
mse = ...
return mse
theta_hats = [0.75, 1.0, 1.5]
for theta_hat in theta_hats:
print(f"theta_hat of {theta_hat} has an MSE of {mse(x, y, theta_hat):.2f}")
# + cellView="both"
# to_remove solution
def mse(x, y, theta_hat):
"""Compute the mean squared error
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
theta_hat (float): An estimate of the slope parameter
Returns:
float: The mean squared error of the data with the estimated parameter.
"""
# Compute the estimated y
y_hat = theta_hat * x
# Compute mean squared error
mse = np.mean((y - y_hat)**2)
return mse
theta_hats = [0.75, 1.0, 1.5]
for theta_hat in theta_hats:
print(f"theta_hat of {theta_hat} has an MSE of {mse(x, y, theta_hat):.2f}")
# -
# The result should be:
#
# theta_hat of 0.75 has an MSE of 9.08\
# theta_hat of 1.0 has an MSE of 3.0\
# theta_hat of 1.5 has an MSE of 4.52
#
#
#
#
# We see that $\hat{\theta} = 1.0$ is our best estimate from the three we tried. Looking just at the raw numbers, however, isn't always satisfying, so let's visualize what our estimated model looks like over the data.
#
#
# + cellView="form"
#@title
#@markdown Execute this cell to visualize estimated models
fig, axes = plt.subplots(ncols=3, figsize=(18, 4))
for theta_hat, ax in zip(theta_hats, axes):
# True data
ax.scatter(x, y, label='Observed') # our data scatter plot
# Compute and plot predictions
y_hat = theta_hat * x
ax.plot(x, y_hat, color='r', label='Fit') # our estimated model
ax.set(
title= fr'$\hat{{\theta}}$= {theta_hat}, MSE = {mse(x, y, theta_hat):.2f}',
xlabel='x',
ylabel='y'
);
axes[0].legend()
# -
# ## Interactive Demo 1: MSE Explorer
#
# Using an interactive widget, we can easily see how changing our slope estimate changes our model fit. We display the **residuals**, the differences between observed and predicted data, as line segments between the data point (observed response) and the corresponding predicted response on the model fit line.
#
# - What value of $\hat{\theta}$ results in the lowest MSE?
# - Is this a good way of estimating $\theta$?
#
# + cellView="form"
#@title
#@markdown Make sure you execute this cell to enable the widget!
@widgets.interact(theta_hat=widgets.FloatSlider(1.0, min=0.0, max=2.0))
def plot_data_estimate(theta_hat):
y_hat = theta_hat * x
plot_observed_vs_predicted(x, y, y_hat, theta_hat)
# +
# to_remove explanation
"""
1. Theta_hat = 1.2 results in the minimum MSE
2. No, this is really inefficient - we do not want to fit models by changing parameters
on a slider!
"""
# -
# While visually exploring several estimates can be instructive, it's not the most efficient for finding the best estimate to fit our data. Another technique we can use is choose a reasonable range of parameter values and compute the MSE at several values in that interval. This allows us to plot the error against the parameter value (this is also called an **error landscape**, especially when we deal with more than one parameter). We can select the final $\hat{\theta}$ ($\hat{\theta}_\textrm{MSE}$) as the one which results in the lowest error.
# + cellView="form"
# @title
# @markdown Execute this cell to loop over theta_hats, compute MSE, and plot results
# Loop over different thetas, compute MSE for each
theta_hat_grid = np.linspace(-2.0, 4.0)
errors = np.zeros(len(theta_hat_grid))
for i, theta_hat in enumerate(theta_hat_grid):
errors[i] = mse(x, y, theta_hat)
# Find theta that results in lowest error
best_error = np.min(errors)
theta_hat = theta_hat_grid[np.argmin(errors)]
# Plot results
fig, ax = plt.subplots()
ax.plot(theta_hat_grid, errors, '-o', label='MSE', c='C1')
ax.axvline(theta, color='g', ls='--', label=r"$\theta_{True}$")
ax.axvline(theta_hat, color='r', ls='-', label=r"$\hat{{\theta}}_{MSE}$")
ax.set(
title=fr"Best fit: $\hat{{\theta}}$ = {theta_hat:.2f}, MSE = {best_error:.2f}",
xlabel=r"$\hat{{\theta}}$",
ylabel='MSE')
ax.legend();
# -
# We can see that our best fit is $\hat{\theta}=1.18$ with an MSE of 1.45. This is quite close to the original true value $\theta=1.2$!
#
# ---
# # Section 2: Least-squares optimization
#
# *Estimated timing to here from start of tutorial: 20 min*
#
# While the approach detailed above (computing MSE at various values of $\hat\theta$) quickly got us to a good estimate, it still relied on evaluating the MSE value across a grid of hand-specified values. If we didn't pick a good range to begin with, or with enough granularity, we might miss the best possible estimator. Let's go one step further, and instead of finding the minimum MSE from a set of candidate estimates, let's solve for it analytically.
#
# We can do this by minimizing the cost function. Mean squared error is a convex objective function, therefore we can compute its minimum using calculus. Please see video or Bonus Section 1 for this derivation! After computing the minimum, we find that:
#
# \begin{align}
# \hat\theta = \frac{\mathbf{x}^\top \mathbf{y}}{\mathbf{x}^\top \mathbf{x}}
# \end{align}
# where $\mathbf{x}$ and $\mathbf{y}$ are vectors of data points.
#
# This is known as solving the normal equations. For different ways of obtaining the solution, see the notes on [Least Squares Optimization](https://www.cns.nyu.edu/~eero/NOTES/leastSquares.pdf) by <NAME>.
# ## Coding Exercise 2: Solve for the Optimal Estimator
#
# In this exercise, you will write a function that finds the optimal $\hat{\theta}$ value using the least squares optimization approach (the equation above) to solve MSE minimization. It shoud take arguments $x$ and $y$ and return the solution $\hat{\theta}$.
#
# We will then use your function to compute $\hat{\theta}$ and plot the resulting prediction on top of the data.
# +
def solve_normal_eqn(x, y):
"""Solve the normal equations to produce the value of theta_hat that minimizes
MSE.
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
float: the value for theta_hat arrived from minimizing MSE
"""
################################################################################
## TODO for students: solve for the best parameter using least squares
# Fill out function and remove
raise NotImplementedError("Student exercise: solve for theta_hat using least squares")
################################################################################
# Compute theta_hat analytically
theta_hat = ...
return theta_hat
theta_hat = solve_normal_eqn(x, y)
y_hat = theta_hat * x
plot_observed_vs_predicted(x, y, y_hat, theta_hat)
# + cellView="both"
# to_remove solution
def solve_normal_eqn(x, y):
"""Solve the normal equations to produce the value of theta_hat that minimizes
MSE.
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
float: the value for theta_hat arrived from minimizing MSE
"""
# Compute theta_hat analytically
theta_hat = (x.T @ y) / (x.T @ x)
return theta_hat
theta_hat = solve_normal_eqn(x, y)
y_hat = theta_hat * x
with plt.xkcd():
plot_observed_vs_predicted(x, y, y_hat, theta_hat)
# -
# We see that the analytic solution produces an even better result than our grid search from before, producing $\hat{\theta} = 1.21$ with MSE = 1.43!
# ---
# # Summary
#
# *Estimated timing of tutorial: 30 minutes*
#
# Linear least squares regression is an optimization procedure that can be used for data fitting:
#
# - Task: predict a value for $y_i$ given $x_i$
# - Performance measure: $\textrm{MSE}$
# - Procedure: minimize $\textrm{MSE}$ by solving the normal equations
#
# **Key point**: We fit the model by defining an *objective function* and minimizing it.
#
# **Note**: In this case, there is an *analytical* solution to the minimization problem and in practice, this solution can be computed using *linear algebra*. This is *extremely* powerful and forms the basis for much of numerical computation throughout the sciences.
# # Notation
# \begin{align}
# x_{i} &\quad \text{input, independent variable}\\
# y_{i} &\quad \text{measurement, dependent variable}\\
# \mathbf{x} &\quad \text{vector of input values}\\
# \mathbf{y} &\quad \text{vector of measurements}\\
# \hat{y}_{i} &\quad \text{estimate of dependent variable}\\
# \epsilon_{i} &\quad \text{measurement error}\\
# \theta &\quad \text{slope parameter}\\
# \hat{\theta} &\quad \text{estimated slope parameter}\\
# \hat{\theta}_\text{MSE} &\quad \text{slope parameter estimated via the mean squared error}\\
# \textrm{MSE} &\quad \text{mean squared error}\\
# \end{align}
# ---
# # Bonus
# ---
# ## Bonus Section 1: Least Squares Optimization Derivation
#
# We will outline here the derivation of the least squares solution.
#
# We first set the derivative of the error expression with respect to $\theta$ equal to zero,
#
# \begin{align}
# \frac{d}{d\theta}\frac{1}{N}\sum_{i=1}^N(y_i - \theta x_i)^2 = 0 \\
# \frac{1}{N}\sum_{i=1}^N-2x_i(y_i - \theta x_i) = 0
# \end{align}
#
# where we used the chain rule. Now solving for $\theta$, we obtain an optimal value of:
#
# \begin{align}
# \hat\theta = \frac{\sum_{i=1}^N x_i y_i}{\sum_{i=1}^N x_i^2}
# \end{align}
#
# Which we can write in vector notation as:
#
# \begin{align}
# \hat\theta = \frac{\mathbf{x}^\top \mathbf{y}}{\mathbf{x}^\top \mathbf{x}}
# \end{align}
#
#
# This is known as solving the *normal equations*. For different ways of obtaining the solution, see the notes on [Least Squares Optimization](https://www.cns.nyu.edu/~eero/NOTES/leastSquares.pdf) by <NAME>.
| tutorials/W1D3_ModelFitting/W1D3_Tutorial1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Titanic Project
#
# ## In this project we will discuss the data analysis of Titanic crash and will try to look deeper into the information to get the answers of different questions that are raised while looking into the collected data by kaggle.
#
# ## The Questions that I think should be clearlly answerable from the provided dataset
#
# ### 1) Is it possible that pclass(Socio-economic class) has any interference with number of lives saved?
# #### hypothesis 1) pclass had no effect on the lives saved
# ### 2) Did gender played any role in saving lives of the Passanger?
# #### hypothesis 2) gender didnot play any role in saving lives.
#
#
# From the Titanic data, we can see the various features present for each passenger on the ship:
#
# Survived: Outcome of survival (0 = No; 1 = Yes)
# Pclass: Socio-economic class (1 = Upper class; 2 = Middle class; 3 = Lower class)
# Name: Name of passenger
# Sex: Sex of the passenger
# Age: Age of the passenger (Some entries contain NaN)
# SibSp: Number of siblings and spouses of the passenger aboard
# Parch: Number of parents and children of the passenger aboard
# Ticket: Ticket number of the passenger
# Fare: Fare paid by the passenger
# Cabin Cabin number of the passenger (Some entries contain NaN)
# Embarked: Port of embarkation of the passenger (C = Cherbourg; Q = Queenstown; S = Southampton)
#
# This information is available on kaggle's website
#
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# #### First thing first taking a look at dataset to create a idea to answer the raised questions.
data = pd.read_csv('titanic-data.csv')
print(data.head())
print(data.tail())
# #### After looking into the data and proposed questions, I think following coloums dosent play much role in the dataset
#
# Name
# Ticket
# Cabin
# Fare
# Embarked
# Parch
# SibSp
#
# So the above listed coloums will be removed.
# And it will be followed by the data cleaning to take care of missing values
clean_data = data.drop(['Name','Ticket','Cabin','Fare','Embarked', 'Parch', 'SibSp'], axis=1)
print(clean_data.head())
# #### Now Dealing with the missing values as they may affect the outcome
# #### Therefore we will calculate the mean value and then will replace it from NAN
age_mean = np.mean(clean_data['Age'])
print age_mean
# #### So to Handle the missing values, replace null by mean
clean_data['Age'] = clean_data['Age'].replace(np.nan, age_mean)
print clean_data.head()
print clean_data.tail()
# #### So from above display we now no longer need the age coloum as now we have a updated age where every person is having a age value.
# #### Now calculating the standerd statistics for the dataset to get a genral idea of distribution
#
clean_data.describe()
# #### The above chart provides the various values for the dataset such as total count, mean, standerd deviation, Maximum and minimum
# #### list of survived and dead which will provide us the final stage to answer our questions
survived = clean_data[clean_data.Survived==True]
dead = clean_data[clean_data.Survived==False]
print survived.head()
print'---------------'
print dead.head()
# #### As we can see from the above display that the list of survived and dead is created and now we can start our analysis from this list
# #### Now in order to answer our first question we will group the data as per there Socio-economic class and then we will plot them to get our answers
class_of_survived = survived.groupby(['Pclass']).size()
class_of_dead = dead.groupby(['Pclass']).size()
# ### # 1) Is it possible that pclass(Socio-economic class) has any interference with number of lives saved?
#
print class_of_survived
print'-----------------'
print class_of_dead
survived.groupby(['Pclass']).size().plot(kind='bar',stacked=True)
plt.ylabel('Count')
plt.xlabel('Socio-economic class (1 = Upper class; 2 = Middle class; 3 = Lower class)')
plt.title('Survived',fontsize=14)
# Pclass: Socio-economic class (1 = Upper class; 2 = Middle class; 3 = Lower class)
#
# From the above graph we can see that lots of lives were saved
dead.groupby(['Pclass']).size().plot(kind='bar',stacked=True)
plt.ylabel('Count')
plt.xlabel('Socio-economic class (1 = Upper class; 2 = Middle class; 3 = Lower class)')
plt.title('NOT-Survived',fontsize=14)
# Pclass: Socio-economic class (1 = Upper class; 2 = Middle class; 3 = Lower class)
#
# The above plot shows the distribution
# +
data_socio = pd.pivot_table(data=clean_data,
values='PassengerId', index='Pclass', columns='Survived', aggfunc='count')
print data_socio
# +
upper = data_socio.loc[1]
middle = data_socio.loc[2]
lower = data_socio.loc[3]
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(8,4))
pie_1 = axes[0].pie(upper, labels=['Not survived','Survived'],
autopct='%1.1f%%', colors=['gold', 'lightskyblue'])
axes[0].set_title('upper')
axes[0].axis('equal')
pie_2 = axes[1].pie(middle, labels=['Not survived','Survived'],
autopct='%1.1f%%', startangle=90, colors=['gold', 'lightskyblue'])
axes[1].set_title('middle')
axes[1].axis('equal')
pie_3 = axes[2].pie(lower, labels=['Not survived','Survived'],
autopct='%1.1f%%', startangle=90, colors=['gold', 'lightskyblue'])
axes[2].set_title('lower')
axes[2].axis('equal')
plt.subplots_adjust(wspace=1)
# -
# #### To get a more detailed look, I have again ploted the socio-economic class but this time using pie chart and it gives us even more information compared to bar plot
# #### Lets put some thought on what could have happened when titanic went down in the sea, from the above charts and displayed value one thing is clear that lots of passenger did survived the accident. But when we look closer we come to our first question that is " Does Socio-economic class has any effect of saving the number of lifes ?"
#
# #### So to answer this we first divided the dataset into survived and dead and then they were further grouped on there Socio-economic classes, this step has reveled that the passenger class has a remarkable effect on the number of lives saved. if we take a look on the chart titled as Not-Survived we can clearly see that coloum representing the lower class is very high as compare to other coloums.
#
# #### So for our first hypothesis that pclass had no effect on the lives saved.
# #### null is rejected as from the chart it is visible in conclusion we will discuss what could be the reason behind this.
# ### #2) Did gender played any role in saving lives of the Passanger?
# #### Now to answer our next question we will regroup the data as per there gender
gender_based_survived = pd.crosstab(survived['Survived'], survived['Sex'])
gender_based_dead = pd.crosstab(dead['Survived'], dead['Sex'])
print gender_based_survived
print gender_based_dead
pd.crosstab(survived['Survived'], survived['Sex']).plot(kind = 'bar', stacked = False)
plt.title('Survived')
plt.ylabel('Count')
plt.xlabel('gender (1 = survived)')
pd.crosstab(dead['Survived'], dead['Sex']).plot(kind = 'bar',stacked = False)
plt.title('Not - Survived')
plt.ylabel('Count')
plt.xlabel('gender (0 = didnot survive)')
# +
data_gender = pd.pivot_table(data=clean_data,
values='PassengerId', index='Sex', columns='Survived', aggfunc='count')
print data_gender
# +
female = data_gender.loc['female']
male = data_gender.loc['male']
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(8,4))
pie_1 = axes[0].pie(female, labels=['Not survived','Survived'],
autopct='%1.1f%%', colors=['gold', 'lightskyblue'])
axes[0].set_title('Female')
axes[0].axis('equal')
pie_2 = axes[1].pie(male, labels=['Not survived','Survived'],
autopct='%1.1f%%', startangle=90, colors=['gold', 'lightskyblue'])
axes[1].set_title('Male')
plt.axis('equal')
# -
# #### Now using the pie chart to compare the data of survived and Not-survived for male and female passenger
# #### For the Second question "Did gender played any role in saving lives ?"
# #### our hypothesis says that gender had nothing to do with lives saved.
# #### but if we take a look at charts its very clear that female passengers were saved as the priority.
# #### So based on this we reject the null hypothesis.
# ### Conclusion:
# #### From the above two questions and there hypothesis analysis we must make some conclusion about what could have have happened. And as per my analysis and understanding about the build of titanic 1)Socio-economic group did played a rol when it came to passenger survived as, passengers from upper class had there cabins towards the upper deck, which made it very easy for them to access the life saving boats. As compared to lower class whoes cabins were situated at the lower decks as a result when the accident happened they didnt had much of the time to react and resulted in tremandous loss of life. 2) Also about the gender being a one of the major player in saving lives, I think as it was the most and obvious decision to save females first as a part of first and most basic instinct of a human.
#
# #### And as a part of further exploration i will definatlly look into the possiblity of age factor and one thing I think I will enjoy solving is all those who travled with family did maneged to get atleast one of there family member to survive.
| Titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Semi-supervised Image Classification and Generation with PCA and K-Means
# ### By <NAME> and <NAME>
# ----------------------------------------------------------------------------------------------------------------------
# **Abstract**: Image classification and image generation are long-standing problems in the fields of artificial intelligence and computer science research. While many modern approaches to these tasks make use of deep neural networks like Generative Adversarial Networks and Convolutional Neural Networks, we demonstrate that simple image recognition and generation may be accomplished with classical, unsupervised methods in linear algebra, including Principal Component Analysis and K-means clustering.
# ----------------------------------------------------------------------------------------------------------------------
# ### Load in packages that we will use
using MLDatasets, LinearAlgebra, Clustering, ImageCore, Images, MultivariateStats, StatsBase, Noise, PyPlot, Plots #, Plots
# ### Load in dataset
# +
# load full training set
train_x, train_y = MNIST.traindata()
# load full test set
test_x, test_y = MNIST.testdata()
println("Training set size: " * string(size(train_x)))
println("Testing set size: " * string(size(test_x)))
p0 = Plots.plot(title = "0", Gray.(transpose(MNIST.traintensor(indexin(0, train_y)[1]))))
p1 = Plots.plot(title = "1", Gray.(transpose(MNIST.traintensor(indexin(1, train_y)[1]))))
p2 = Plots.plot(title = "2", Gray.(transpose(MNIST.traintensor(indexin(2, train_y)[1]))))
p3 = Plots.plot(title = "3", Gray.(transpose(MNIST.traintensor(indexin(3, train_y)[1]))))
p4 = Plots.plot(title = "4", Gray.(transpose(MNIST.traintensor(indexin(4, train_y)[1]))))
p5 = Plots.plot(title = "5", Gray.(transpose(MNIST.traintensor(indexin(5, train_y)[1]))))
p6 = Plots.plot(title = "6", Gray.(transpose(MNIST.traintensor(indexin(6, train_y)[1]))))
p7 = Plots.plot(title = "7", Gray.(transpose(MNIST.traintensor(indexin(7, train_y)[1]))))
p8 = Plots.plot(title = "8", Gray.(transpose(MNIST.traintensor(indexin(8, train_y)[1]))))
p9 = Plots.plot(title = "9", Gray.(transpose(MNIST.traintensor(indexin(9, train_y)[1]))))
#Plots.plot(p0, p1, p2, p3, p4, p5, p6, p7, p8, p9, layout=(2, 5), axis=([], false))
PyPlot.imshow(
transpose(MNIST.traintensor(30)), cmap="gray"
)
# -
# ### Reformat training data into a matrix of flattened vectors where each column is a single image.
# ### Then plot an image to ensure it was done correctly
# +
reshaped_train_x = reshape(train_x, (28*28, size(train_x)[3]))
reshaped_test_x = reshape(test_x, (28*28, size(test_x)[3]))
println("Training set size: " * string(size(reshaped_train_x)))
println("Testing set size: " * string(size(reshaped_test_x)))
# make sure we've correctly maintained the data
function reshape_flattened_vector(flat_vector)
n_rows = 28
n_cols = 28
# reshape flat vector to 2d image, for some reason img gets
# flipped along diagonal when reshaping so we transpose it
return transpose(reshape(flat_vector, (n_rows, n_cols)))
end
first_ex = reshaped_train_x[:,3]
first_ex_reshaped = reshape_flattened_vector(first_ex)
# show a few of the images to be sure
# Plots.plot(Gray.(first_ex_reshaped))
# -
# ### This method will give us all the images for a given digit
# ### For example, this is useful when we want to pull all the images containing 4's, etc.
# get images of a certain class
function get_mnist_digitclass(digitclass, data, labels)
digitclass_vec = Array{Float64}(undef, 0, size(data)[1])
# print(size(digitclass_vec))
for i = 1:size(data)[2] # number of samples
if(labels[i] == digitclass)
digitclass_vec = [ digitclass_vec; transpose(data[:,i]) ]
end
end
return transpose(digitclass_vec)
end
fours = get_mnist_digitclass(4, reshaped_train_x, train_y)
# +
fig = figure(figsize=(10, 4))
subplot(211)
PyPlot.imshow(
show_sample_img(fours, 17), cmap="gray"
)
axis("off")
subplot(212)
PyPlot.imshow(
show_sample_img(fours, 2), cmap="gray"
)
axis("off")
suptitle("Four Styles",size=24)
# -
# ### Use SVD to reduce a random image in our training set
test_svd = svd(MNIST.traintensor(3))
println("Singular values: ", test_svd.S)
# ### Plot the SVD representation of our digit
reconstructed_test_svd = transpose((test_svd.U * Diagonal(test_svd.S) * test_svd.Vt))
println(MNIST.trainlabels(3))
Plots.plot(Gray.(reconstructed_test_svd))
# ### Using "low rank" approximation for images
# ### We grab the first 3 singular elements from our SVD to show that the digit is still recognizable even with significantly less data
# rank 3 approximation for test_svd
test_svd_3 = transpose(test_svd.U[:, 1:3] * Diagonal(test_svd.S[1:3]) * test_svd.Vt[1:3, :])
Plots.plot(Gray.(test_svd_3))
# ### Build principal components of MNIST
# #### Helper functions that perform the actual PCA training, displaying sample images and displaying the 3D scatter plots of the projections on to the low dimensional subspace
# +
function pca_m(data, n_reduced_dims)
data = Float64.(data)
pca_model = fit(PCA, data; maxoutdim=n_reduced_dims)
princ_vecs = transform(pca_model, data)
proj_mat = projection(pca_model)
reconstructed = reconstruct(pca_model, princ_vecs)
return (pca_model, princ_vecs, proj_mat)
end
function show_sample_img(data, sample_no)
data_sample = data[:,sample_no]
return data_sample_reshaped = reshape_flattened_vector(data_sample)
# return Plots.plot(Gray.(data_sample_reshaped), axis=([], false))
end
function display_proj(proj_coords, r_start, r_end, color, label)
(x1, y1, z1) = (proj_coords[r_start:r_end, 1],
proj_coords[r_start:r_end, 2],
proj_coords[r_start:r_end, 3])
return scatter3D(x1, y1, z1, color = color, label = label, s = 60)
end
function get_digit_projections(data, labels, digit_class)
for i = 1:size(labels)[1]
if labels[i] == digit_class
println(labels[i])
end
end
end
# -
# ### Train the PCA and then project down to subspace, plot in 3D the projections of the fours images and fives images
# +
function least_squares(A, b)
return inv(transpose(A) * A) * transpose(A) * b
end
function kdim_subspace_proj(basis_matrix, samples)
# project samples on to k-dimension subspace of best fit spanned by
# the vectors in the columns of the basis matrix
proj_matrix = basis_matrix * inv(transpose(basis_matrix)*basis_matrix) * transpose(basis_matrix)
# project columns of samples onto columnspace of basis matrix
proj_samples = proj_matrix * samples
return proj_samples
end
function kdim_proj_coords(basis_matrix, projected_samples)
# get coordinates of projected samples within new kdim subspace, expect a k-dim vector out
return least_squares(basis_matrix, projected_samples)
end
# +
# determine embedding dimension, train PCA on mnist samples
embedding_dim = 16
pca_model, princ_vecs, pca_proj_mat = pca_m(reshaped_test_x, embedding_dim)
println("Proj Matrix: " * string(size(pca_proj_mat)))
# get projection coordinates of samples onto k-dim subspace of best fit
kdim_proj_vecs = transpose(reshaped_test_x) * pca_proj_mat
kdim_proj_vecs = transpose(kdim_proj_vecs)
println("Projection coordinates: " * string(size(kdim_proj_vecs)))
# +
p0 = get_mnist_digitclass(0, kdim_proj_vecs, test_y)
p1 = get_mnist_digitclass(1, kdim_proj_vecs, test_y)
p2 = get_mnist_digitclass(2, kdim_proj_vecs, test_y)
p3 = get_mnist_digitclass(3, kdim_proj_vecs, test_y)
p4 = get_mnist_digitclass(4, kdim_proj_vecs, test_y)
p5 = get_mnist_digitclass(5, kdim_proj_vecs, test_y)
p6 = get_mnist_digitclass(6, kdim_proj_vecs, test_y)
p7 = get_mnist_digitclass(7, kdim_proj_vecs, test_y)
p8 = get_mnist_digitclass(8, kdim_proj_vecs, test_y)
p9 = get_mnist_digitclass(9, kdim_proj_vecs, test_y)
fig = figure(figsize=(10,10))
p0_plot = scatter3D(p0[1, :], p0[2, :], p0[3, :], label = "Zero", s = 20)
p1_plot = scatter3D(p1[1, :], p1[2, :], p1[3, :], label = "One", s = 20)
# p2_plot = scatter3D(p2[1, :], p2[2, :], p2[3, :], label = "Two", s = 20)
# p3_plot = scatter3D(p3[1, :], p3[2, :], p3[3, :], label = "Three", s = 20)
# p4_plot = scatter3D(p4[1, :], p4[2, :], p4[3, :], label = "Four", s = 20)
# p5_plot = scatter3D(p5[1, :], p5[2, :], p5[3, :], label = "Five", s = 20)
# p6_plot = scatter3D(p6[1, :], p6[2, :], p6[3, :], label = "Six", s = 20)
# p7_plot = scatter3D(p7[1, :], p7[2, :], p7[3, :], label = "Seven", s = 20)
# p8_plot = scatter3D(p8[1, :], p8[2, :], p8[3, :], label = "Eight", s = 20)
# p9_plot = scatter3D(p9[1, :], p9[2, :], p9[3, :], label = "Nine", s = 20)
title("3D Scatter of Digit Projections")
legend(loc="upper right")
PyPlot.savefig("3d-scatter-4-9.png")
# -
# ### K-means clustering
# +
function acc_score(pred, truth)
score = 0
for i = 1:length(pred)
if(pred[i] == truth[i])
score += 1
end
end
return score / length(pred)
end
function get_corr_labels(arr, index)
indexed_arr = Array{Float64}(undef, 0)
for i = 1:length(arr)
if index[i] == 1
indexed_arr = [indexed_arr; arr[i]]
end
end
return indexed_arr
end
# gets most probable ground truth label corresponding to each bucket
# so that we know what cluster corresponds to what number
function get_cluster_assoc(cluster_labels, truth_labels)
reference_labels = Dict()
unique_labels = countmap(cluster_labels)
for i = 1:length(unique_labels)
index = [ifelse(x == i, 1, 0) for x in cluster_labels]
indexed_truth_labels = get_corr_labels(truth_labels, index)
counted_indexed_truth_labels = countmap(indexed_truth_labels)
reference_labels[i] = Int(findmax(counted_indexed_truth_labels)[2])
end
return reference_labels
end
# -
# ### The code below performs the k-means clustering and prints out the dimensions of the projection matrix, the associated value with each clustering bin from our k-means, and the total accuracy our k-means clustering algorithm had
# +
# perform actual k-means clustering
R = kmeans(kdim_proj_newcoords, 75; maxiter=20, init=:kmpp)
a = assignments(R)
c = counts(R)
# projection matrix dims
println("Projections: " * string(size(kdim_proj_newcoords)))
# mappings between clustering bins and which number they represent
assoc_vals = get_cluster_assoc(a, test_y)
predicted_labels = [Int(assoc_vals[bin]) for bin in a]
println("Accuracy: " * string(acc_score(predicted_labels, test_y)))
for x in sort(collect(zip(values(assoc_vals),keys(assoc_vals)))) println(x) end
# +
# perform k-means with different bin sizes and plot accuracy as bin size increases
acc_bins = Array{Float64}(undef, 0)
bin_size = Array{Float64}(undef, 0)
for i = 1:4:256
R_i = kmeans(kdim_proj_newcoords, i; maxiter=20, init=:kmpp)
a_i = assignments(R_i)
# mappings between clustering bins and which number they represent
assoc_vals = get_cluster_assoc(a_i, test_y)
predicted_labels = [Int(assoc_vals[bin]) for bin in a_i]
acc_bins = [acc_bins; acc_score(predicted_labels, test_y)]
bin_size = [bin_size;i]
end
# +
fig = figure(figsize=(10,5))
xlabel("Cluster Count", size=18)
ylabel("K-means Model Accuracy",size=18)
PyPlot.scatter(bin_size, acc_bins)
PyPlot.plot(bin_size, acc_bins)
title("K-means Accuracy by Cluster Count", size = 24)
PyPlot.grid("on")
ylim(0, 1)
PyPlot.savefig("kmeans-performance.png")
# -
# ### Using our clustering bins from above, we can take the center of each bin and use its value to "reconstruct" what a generic digit from that bin looks like!
# +
# transform k-means centers back to global coordinate frame
println("Kmeans Centers: " * string(size(R.centers)))
kmeans_center_images = reconstruct(pca_model, R.centers)
PyPlot.imshow(
show_sample_img(kmeans_center_images, 27), cmap="gray"
)
PyPlot.savefig("kmeans-performance.png")
# -
# ### Here we add some random noise to the vector and show what our model outputs
# +
print(size(R.centers)) # embedding_dim dimensional vector
noisy_centers = mult_gauss(R.centers, 0.4)
kmeans_noisy_centers = reconstruct(pca_model, noisy_centers)
fig = figure(figsize=(10, 4))
subplot(141)
PyPlot.imshow(
show_sample_img(kmeans_noisy_centers, 27), cmap="gray"
)
title("7", size=24)
axis("off")
subplot(142)
PyPlot.imshow(
show_sample_img(kmeans_noisy_centers, 22), cmap="gray"
)
title("5", size=24)
axis("off")
subplot(143)
PyPlot.imshow(
show_sample_img(kmeans_noisy_centers, 26), cmap="gray"
)
title("3", size=24)
axis("off")
subplot(144)
PyPlot.imshow(
show_sample_img(kmeans_noisy_centers, 34), cmap="gray"
)
title("9", size=24)
axis("off")
suptitle("Noisy Embedded Digit Reconstruction",size=24)
PyPlot.savefig("noisy-digits.png")
# -
# This analyzes the dot product similarity between the centers of two clusters.
# We use the centers of the bins representing the digits 9 and 4.
# The two digits look somewhat similar, so we expect the centers to be somewhat similar
# +
function euclidian_similarity(x, y)
return sqrt(sum((x - y) .^ 2))
end
function cosine_similarity(x, y)
return dot(x, y) / (norm(x) * norm(y))
end
# now look at dot product similarity of cluster centers
println("Similarity of 9 and 4 center (cosine similarity): ")
c1 = R.centers[:, 8]
c2 = R.centers[:, 22]
println("Cosine Similarity: " * string(cosine_similarity(c1, c2)))
println("Euc. Similarity: " * string(euclidian_similarity(c1, c2)))
# -
function digit_similarity(digit_one, digit_two)
digit_one_bins = [i.first for i in assoc_vals if i.second == digit_one]
digit_two_bins = [i.first for i in assoc_vals if i.second == digit_two]
sum = 0
for bin_one in digit_one_bins
for bin_two in digit_two_bins
sum += cosine_similarity(R.centers[:, bin_one], R.centers[:, bin_two])
end
end
return sum/(size(digit_one_bins, 1) * size(digit_two_bins, 1))
end
# +
similarity_matrix = Array{Float64}(undef, 10, 10)
for i = 1:10
for j = 1:10
similarity_matrix[i, j] = digit_similarity(i-1, j-1)
end
end
rounded_matrix = [round(i, digits=3) for i in similarity_matrix]
println(rounded_matrix)
# -
# # THANKS FOR READING!
# For any questions: please email <EMAIL> or <EMAIL>
| .ipynb_checkpoints/Final-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DAT210x - Programming with Python for DS
# ## Module6- Lab5
import pandas as pd
# Useful information about the dataset used in this assignment can be [found here](https://archive.ics.uci.edu/ml/machine-learning-databases/mushroom/agaricus-lepiota.names).
# Load up the mushroom dataset into dataframe `X` and verify you did it properly, and that you have not included any features that clearly shouldn't be part of the dataset.
#
# You should not have any doubled indices. You can check out information about the headers present in the dataset using the link we provided above. Also make sure you've properly captured any NA values.
# +
#name_lst = ['classes', 'cap-shape', 'cap-surface', 'cap-color', 'bruises?', 'odor', 'gill-attachment', 'gill-spacing', 'gill-size', 'gill-color', 'stalk-shape', 'stalk-root', 'stalk-surface-above-ring', 'stalk-surface-below-ring', 'stalk-color-above-ring', 'stalk-color-below-ring', 'veil-type', 'veil-color', 'ring-number', 'ring-type', 'spore-print-color', 'population', 'habitat']
name_lst = ['cat','cap-shape', 'cap-surface', 'cap-color', 'bruises?', 'odor', 'gill-attachment', 'gill-spacing', 'gill-size', 'gill-color', 'stalk-shape', 'stalk-root', 'stalk-surface-above-ring', 'stalk-surface-below-ring', 'stalk-color-above-ring', 'stalk-color-below-ring', 'veil-type', 'veil-color', 'ring-number', 'ring-type', 'spore-print-color', 'population', 'habitat']
X = pd.read_csv('Datasets/agaricus-lepiota.data',header = None,names = name_lst, na_values='?')
# -
# An easy way to show which rows have nans in them:
X[pd.isnull(X).any(axis=1)]
# For this simple assignment, just drop any row with a nan in it, and then print out your dataset's shape:
X = X.dropna(axis = 0)
print("After dropping all rows with any NaNs, shape of X is:", X.shape)
# Copy the labels out of the dataframe into variable `y`, then remove them from `X`.
#
# Encode the labels, using the `.map()` trick we presented you in Module 5, using `canadian:0`, `kama:1`, and `rosa:2`.
y = X.cat.map({'p':0,'e':1})
X = X.drop(labels=['cat'], axis = 1)
# Encode the entire dataframe using dummies:
# Split your data into `test` and `train` sets. Your `test` size should be 30% with `random_state` 7.
#
# Please use variable names: `X_train`, `X_test`, `y_train`, and `y_test`:
X = pd.get_dummies(X)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state = 7)
# Create an DT classifier. No need to set any parameters:
from sklearn import tree
model = tree.DecisionTreeClassifier()
# Train the classifier on the `training` data and labels; then, score the classifier on the `testing` data and labels:
model.fit(X_train, y_train)
score = model.score(X_test, y_test)
print("High-Dimensionality Score: ", round((score*100), 3))
# Use the code on the course's SciKit-Learn page to output a .DOT file, then render the .DOT to .PNGs.
#
# You will need graphviz installed to do this. On macOS, you can `brew install graphviz`. On Windows 10, graphviz installs via a .msi installer that you can download from the graphviz website. Also, a graph editor, gvedit.exe can be used to view the tree directly from the exported tree.dot file without having to issue a call. On other systems, use analogous commands.
#
# If you encounter issues installing graphviz or don't have the rights to, you can always visualize your .dot file on the website: http://webgraphviz.com/.
tree.export_graphviz(model.tree_, out_file = 'tree.dot', feature_names = X.columns)
from subprocess import call
call(['dot', '-T', 'png', 'tree.dot', '-o', 'tree.png'])
| Module6/.ipynb_checkpoints/Module6 - Lab5-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **밑바닥부터 시작하는 데이터과학**
# Data Science from Scratch
# - https://github.com/joelgrus/data-science-from-scratch/blob/master/first-edition/code-python3/decision_trees.py
# # **17장 의사결정나무**
# 다양한 **의사결정경로** 와 **결과** 를 나타냅니다
# 1. **분류나무 (Classification Tree) :** 범주형 결과를 반환 합니다
# 1. **회귀나무 (Regression Tree) :** 숫자형 결과를 반환 합니다
# ## **1 엔트로피**
# **얼마만큼의 정보를 담고 있는지** 를 확인하는 방법으로 **데이터 불확실성 (uncertainty)** 을 나타 냅니다
# 1. 특정 $ c_i $ 클래스에 속할 확률 $ p_i $ 을 엔트로피로 계산하면 $ H(S) = -p_1 \log_2 p_1 - ... - p_n \log_2 p_n $$
# 1. **데이터가 단일 클래스에 속한** 경우 : **불확실성은 없고 엔트로피는 낮다** (그룹이 안정적)
# 1. **데이터가 클래스에 고르게 분포된** 경우 : **불확실성과 엔트로피가 높다** (파벌이 많다)
# +
from collections import Counter, defaultdict
from functools import partial
import math, random
# 클래스에 속할 확률을 입력하면 엔트로피를 계산 합니다
def entropy(class_probabilities):
return sum(-p * math.log(p, 2) for p in class_probabilities if p)
def class_probabilities(labels):
total_count = len(labels)
return [count / total_count
for count in Counter(labels).values()]
def data_entropy(labeled_data):
labels = [label for _, label in labeled_data]
probabilities = class_probabilities(labels)
return entropy(probabilities)
# -
# ## **2 파티션 엔트로피와 의사결정나무**
# 1. 의사결정나무는 **여러개의 파티션** 으로 분할된 **여러 데이터셋** 을 비교합니다
# 1. **여러개의 파티션** 을 계산하더라도, **전체에 대한 데이터셋 엔트로피 계산이** 필요 합니다
# 1. 데이터 $ S $ 를 $ q_1, ... q_m $ 의 비율을 찾는 파티션 $ S_i, ... S_m $ 을 나누는 공식은 다음과 같습니다
#
# $$ H = q_1 H(S_1) + ... + q_m H(S_m) $$
# subsets는 레이블이있는 데이터의 list 입니다
def partition_entropy(subsets):
total_count = sum(len(subset) for subset in subsets)
return sum( data_entropy(subset) * len(subset) / total_count
for subset in subsets )
# ## **3 의사결정나무 만들기**
# 의사결정나무는 **결정노드 (Decison Node)** 와 **잎 노드 (leaf node)** 로 구성됩니다
# 1. **결정노드 (Decison Node) :** 질문의 답에 따른 경로를 안내 합니다
# 1. **잎 노드 (leaf node) :** 예측값이 무엇인지를 알려 줍니다
import json
with open('./data/data.json', 'r') as f:
inputs = json.load(f)
inputs = inputs['inputs']
len(inputs), inputs[:2]
# +
# attribute 에 따라 inputs 의 파티션을 나눕니다
def group_by(items, key_fn):
groups = defaultdict(list)
for item in items:
key = key_fn(item)
groups[key].append(item)
return groups
def partition_by(inputs, attribute):
return group_by(inputs, lambda x: x[0][attribute])
# 주어진 파티션에 대응되는 엔트로피 계산하기
def partition_entropy_by(inputs,attribute):
partitions = partition_by(inputs, attribute)
return partition_entropy(partitions.values())
# 각 데이터에 엔트로피를 최소로 하는 파티션을 찾습니다
for key in ['level','lang','tweets','phd']:
print(key, ":", partition_entropy_by(inputs, key))
# +
# 직급(level) 을 기준으로 나눌 때 최소 엔트로피를 갖습니다
# 이를 기준으로 직급을 기준으로 SubTree 를 생성합니다.
senior_inputs = [(input, label)
for input, label in inputs if input["level"] == "Senior"]
# 위 분석결과 tweets 값이 0으로, 다음 분류기준이 됩니다
for key in ['lang', 'tweets', 'phd']:
print(key, partition_entropy_by(senior_inputs, key))
# -
# ## **5 종합하기**
# 보다 일반화된 방법을 적용해 보겠습니다
# +
# 실제 Tree 구축용 함수
def build_tree_id3(inputs, split_candidates=None):
# partition 첫단계 실행
if split_candidates is None:
split_candidates = inputs[0][0].keys()
num_inputs = len(inputs) # 입력 데이터에서 True, False 갯수를 확인
num_trues = len([label for item, label in inputs if label])
num_falses = num_inputs - num_trues
if num_trues == 0: return False # True 가 없을때 False 반환
if num_falses == 0: return True # False 가 없을때 True 반환
if not split_candidates: # 파티션 기준사용할 변수가 없을 때
return num_trues >= num_falses # 다수결로 결과를 출력
best_attribute = min(split_candidates, # 적합한 변수로 파티션을 나눈다
key=partial(partition_entropy_by, inputs))
partitions = partition_by(inputs, best_attribute)
new_candidates = [a for a in split_candidates if a != best_attribute]
# 재귀적 방법으로 서브Tree 를 구축합니다
subtrees = { attribute : build_tree_id3(subset, new_candidates)
for attribute, subset in partitions.items() }
subtrees[None] = num_trues > num_falses # 기본값
return (best_attribute, subtrees)
print("building the tree")
tree = build_tree_id3(inputs)
tree
# +
# 의사 결정나무 Tree 로 주어진 입력값 inputs 을 분류 합니다
def classify(tree, input):
# leaf 노드일 때 값을 반환합니다
if tree in [True, False]:
return tree
# 위에 비해당시 Partition 을 나눕니다
attribute, subtree_dict = tree
subtree_key = input.get(attribute)
# subtree 가 존재하지 않을 때 None 서브트리를 추가
if subtree_key not in subtree_dict:
subtree_key = None
# 적절한 SubTree 를 선택합니다
subtree = subtree_dict[subtree_key]
return classify(subtree, input)
print("Junior / Java / tweets / no phd", classify(tree,
{ "level" : "Junior",
"lang" : "Java",
"tweets" : "yes",
"phd" : "no"} ))
# -
print("Junior / Java / tweets / phd", classify(tree,
{ "level" : "Junior",
"lang" : "Java",
"tweets" : "yes",
"phd" : "yes"} ))
print("Intern :", classify(tree, { "level" : "Intern" } ))
print("Senior :", classify(tree, { "level" : "Senior" } ))
# ## **6 랜덤 포레스트**
# 결정나무의 한계인 **OverFitting**을 방지하는 방법으로 **여러개의 결정나무를 만든 뒤, 다수결과 결과를 결정** 하는 방법 입니다.
# 1. 랜덤한 모델을 찾기위해서 **bootstrap** 을 활용 합니다
# 1. 즉 inputs 이 아닌, **bootstrap_sample(input)** 로 여러모델을 학습한 뒤 결정을 합니다
# 1. 이러한 방식을 **bootstrap aggregation** 또는 **bagging(배깅)** 이라고 합니다
def forest_classify(trees, input):
votes = [classify(tree, input) for tree in trees]
vote_counts = Counter(votes)
return vote_counts.most_common(1)[0][0]
| 16.DecisionTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Enterprise Deep Learning with TensorFlow: openSAP
# ## SAP Innovation Center Network
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ## Load the necessary modules
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tensorflow as tf
import numpy as np
from functools import reduce
import os
import shutil
# -
# ## Setting up our environment
tf.logging.set_verbosity(tf.logging.INFO)
TRAINING_POS_PATH = "data/sentiment/rt-polarity-pos.txt"
TRAINING_NEG_PATH = "data/sentiment/rt-polarity-neg.txt"
NUM_WORDS = 200
TRAIN_RATIO = .8
NUM_EPOCHS = 50
BATCH_SIZE = 512
LEARNING_RATE = .001
NUM_LAYERS = 1
NUM_UNITS = 10
DROPOUT_PROB = .8
EMBEDDING_SIZE = 10
MODEL_PATH = "dump/"
EVERY_N_ITER = 500
CLEAN = True
# Clean
if CLEAN:
if os.path.exists(MODEL_PATH):
shutil.rmtree(MODEL_PATH)
os.mkdir(MODEL_PATH)
# ## Creating our data set
# +
# create data/sentiment folder, if it doesn't exist
import os
if not os.path.exists("data/sentiment"):
os.makedirs("data/sentiment")
#Download sentence polarity movie review dataset
# https://www.cs.cornell.edu/people/pabo/movie-review-data/
# !wget https://raw.githubusercontent.com/abromberg/sentiment_analysis/master/polarityData/rt-polaritydata/rt-polarity-pos.txt --directory-prefix=./data/sentiment/
# !wget https://raw.githubusercontent.com/abromberg/sentiment_analysis/master/polarityData/rt-polaritydata/rt-polarity-neg.txt --directory-prefix=./data/sentiment/
# -
# Load datasets
def clean_corpus(fname):
sentences = []
for line in open(fname, encoding="utf-8", errors="ignore"):
# Remove leading/trailing whitespace at the boundaries
line = line.strip()
# Don't include empty lines
if not line:
continue
# Remove non-alphanumeric characters (excl. spaces)
line = "".join([char for char in line if char.isalnum() or char == " "])
# Remove leading/trailing spaces between words
line = " ".join([word for word in line.split(" ") if word.strip()])
# Split into words
words = line.split(" ")
# Add to collection
sentences.append(words)
return sentences
# Define input
sentences_pos = clean_corpus(fname=TRAINING_POS_PATH)
sentences_neg = clean_corpus(fname=TRAINING_NEG_PATH)
# Truncate after word threshold
sentences_pos = [sentence[:NUM_WORDS] for sentence in sentences_pos]
sentences_neg = [sentence[:NUM_WORDS] for sentence in sentences_neg]
# Determine dictionary (0 is used for padding)
sentences = sentences_pos + sentences_neg
dictionary = [word for sentence in sentences for word in sentence]
dictionary = list(set(dictionary))
dictionary = dict(zip(dictionary, range(1, len(dictionary) + 1)))
# Convert sentences to sequences of integers
sentences_pos = [[dictionary[word] for word in sentence] for sentence in sentences_pos]
sentences_neg = [[dictionary[word] for word in sentence] for sentence in sentences_neg]
# Check integrity
dictionary_inv = {b: a for a, b in dictionary.items()}
print("POS: " + " ".join([dictionary_inv[index] for index in sentences_pos[0]]))
print("NEG: " + " ".join([dictionary_inv[index] for index in sentences_neg[0]]))
# Pad sentences to same length from the left side (with zeros)
def pad_zeros(sentence):
if len(sentence) == NUM_WORDS:
return sentence
else:
return [0] * (NUM_WORDS - len(sentence)) + sentence
sentences_pos = [pad_zeros(sentence) for sentence in sentences_pos]
sentences_neg = [pad_zeros(sentence) for sentence in sentences_neg]
# +
# Create data set
data_pos = np.array(sentences_pos, dtype=np.int32)
data_pos_labels = np.ones(shape=[len(sentences_pos)], dtype=np.int32)
data_neg = np.array(sentences_neg, dtype=np.int32)
data_neg_labels = np.zeros(shape=[len(sentences_neg)], dtype=np.int32)
data = np.vstack((data_pos, data_neg))
data_labels = np.concatenate((data_pos_labels, data_neg_labels))
# +
# Split into training/test set
np.random.shuffle(data)
num_rows = data.shape[0]
split_train = int(num_rows * TRAIN_RATIO)
train, train_labels = data[:split_train, :], data_labels[:split_train]
test, test_labels = data[split_train:, :], data_labels[split_train:]
# -
# Create input function
def get_input_fn(x, y=None, batch_size=128, num_epochs=1, shuffle=False):
return tf.estimator.inputs.numpy_input_fn(x={"x": x},
y=y,
batch_size=batch_size,
num_epochs=num_epochs,
shuffle=shuffle)
# Set model params
model_params = {"learning_rate": LEARNING_RATE,
"num_layers": NUM_LAYERS,
"num_units": NUM_UNITS,
"embedding_size": EMBEDDING_SIZE,
"dropout_prob": DROPOUT_PROB,
"vocabulary_size": len(dictionary) + 1}
# Log loss
loss_hook = tf.train.LoggingTensorHook(["loss"], every_n_iter=EVERY_N_ITER)
# ## Define our model
# Define LSTM model function
def lstm_model_fn(features, labels, mode, params):
# Define input layer
input_layer = features["x"]
# Embedding layer
word_embeddings = tf.get_variable(name="word_embeddings",
shape=[params["vocabulary_size"], params["embedding_size"]],
initializer=tf.random_normal_initializer())
input_layer = tf.nn.embedding_lookup(word_embeddings, input_layer)
# LSTM (with dropout)
basic_lstm_cells = [tf.contrib.rnn.BasicLSTMCell(num_units=params["num_units"],
activation=tf.nn.tanh)
for _ in range(params["num_layers"])]
dropout_lstm_cells = [tf.nn.rnn_cell.DropoutWrapper(basic_lstm_cell, output_keep_prob=params["dropout_prob"])
for basic_lstm_cell in basic_lstm_cells]
multi_lstm_cells = tf.nn.rnn_cell.MultiRNNCell(dropout_lstm_cells)
outputs, states = tf.nn.dynamic_rnn(multi_lstm_cells, input_layer, dtype=tf.float32)
# Extract final state (last hidden state of sequence of topmost layer)
final_state = states[-1].h
# Fully connected layer (with linear activation)
logits = tf.squeeze(tf.layers.dense(inputs=final_state, units=1, activation=None))
# Define output
sentiment = tf.sigmoid(logits)
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(
mode=mode,
predictions={"sentiment": sentiment})
# Cast labels
labels = tf.cast(labels, dtype=tf.float32)
# Define loss
loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(labels=labels, logits=logits), name="loss")
with tf.name_scope("summaries"):
tf.summary.scalar("cross_entropy", loss)
# Optimizer
optimizer = tf.train.RMSPropOptimizer(learning_rate=params["learning_rate"])
train_op = optimizer.minimize(loss=loss, global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(
mode=mode,
loss=loss,
train_op=train_op)
# ## Instantiate our estimator
# Instantiate Estimator
nn = tf.estimator.Estimator(model_fn=lstm_model_fn,
params=model_params,
model_dir=MODEL_PATH)
# ## Start Training
# Train
nn.train(input_fn=get_input_fn(x=train,
y=train_labels,
batch_size=BATCH_SIZE,
num_epochs=NUM_EPOCHS,
shuffle=True),
hooks=[loss_hook])
# +
# Test
eval_dict = nn.evaluate(input_fn=get_input_fn(x=test,
y=test_labels,
batch_size=test.shape[0]))
print("Cross entropy (test set): {0:.2f}".format(eval_dict["loss"]))
# -
# ## Start Prediction
# Predict
prediction = nn.predict(input_fn=get_input_fn(x=test,
y=test_labels,
batch_size=test.shape[0]))
sentiments = np.array([p["sentiment"] for p in prediction])
# Find indices that would sort array (ascending order)
idx = np.argsort(sentiments)
idx_lo = idx[:5]
idx_hi = idx[-5:]
# ## What is the sentiment of some example sentences?
# Map word indices back to strings
dictionary_inv[0] = ""
map2str = np.vectorize(dictionary_inv.__getitem__)
test_str = map2str(test)
test_str = np.apply_along_axis(lambda row: reduce(lambda a, b: a.strip() + " " + b.strip(), row), axis=1, arr=test_str)
# Most negative
print("NEGATIVE:")
for i in idx_lo:
print("\t{} ::: {:.3f}".format(test_str[i], sentiments[i]))
print()
# Most positive
print("POSITIVE:")
for i in idx_hi[::-1]:
print("\t{} ::: {:.3f}".format(test_str[i], sentiments[i]))
| Week_03_Unit_05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# # Example Use of USB stream and Sockets
# > Example usage see link
# https://www.pythonforthelab.com/blog/using-pyzmq-for-inter-process-communication-part-1/
#
# ## Send and receive socket wih FLIR cam
# +
from time import sleep
from imutils.video import FPS
import imutils
import cv2
from FLIRCam.USB_video_stream import *
import zmq
print('Starting1')
context = zmq.Context()
pubsocket = context.socket(zmq.PUB)
pubsocket.bind("tcp://*:5555")
subsocket = context.socket(zmq.SUB)
subsocket.connect("tcp://localhost:5555")
subsocket.setsockopt(zmq.SUBSCRIBE, b'camera_frame')
width = 1000
height = 750
fvs = USBVideoStream() # .start()
fvs.start()
sleep(0.5)
topic = 'camera_frame'
i=0
while True:
try:
i += 1
while not fvs.more():
pass
snd_frame = fvs.read()
snd_frame = imutils.resize(snd_frame['image'], width=width, height=height)
pubsocket.send_string(topic, zmq.SNDMORE)
pubsocket.send_pyobj(snd_frame)
rec_topic = subsocket.recv_string()
rec_frame = subsocket.recv_pyobj()
cv2.putText(rec_frame, f'Received frame number {i}, {rec_topic}, {rec_frame.shape}',
(10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
cv2.imshow("Cam1", rec_frame)
if cv2.waitKey(100) == 27:
break # esc to quit
except KeyboardInterrupt:
print("W: interrupt received, stopping…")
break
fvs.stop()
del fvs
subsocket.close()
pubsocket.close() # clean up
cv2.destroyAllWindows()
print('Finished')
# -
# ## Send and receive socket wih Webcam
# +
from time import sleep
import zmq
import cv2
print('Starting1')
context = zmq.Context()
pubsocket = context.socket(zmq.PUB)
pubsocket.bind("tcp://*:5555")
subsocket = context.socket(zmq.SUB)
subsocket.connect("tcp://localhost:5555")
subsocket.setsockopt(zmq.SUBSCRIBE, b'camera_frame')
print('Starting2')
cam = cv2.VideoCapture(0)
sleep(0.5)
topic = 'camera_frame'
print('Starting3')
i=0
while True:
try:
i += 1
ret, snd_frame = cam.read()
pubsocket.send_string(topic, zmq.SNDMORE)
pubsocket.send_pyobj(snd_frame)
rec_topic = subsocket.recv_string()
rec_frame = subsocket.recv_pyobj()
cv2.putText(rec_frame, f'Received frame number {i}, {rec_topic}, {rec_frame.shape}',
(10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
cv2.imshow("Cam1", rec_frame)
if cv2.waitKey(100) == 27:
break # esc to quit
except KeyboardInterrupt:
print("W: interrupt received, stopping…")
break
cam.release()
subsocket.close()
pubsocket.close() # clean up
cv2.destroyAllWindows()
print('Finished')
# -
# ## Show FLIR camera
# +
def show_FLIRcam(mirror=False, width=1000, height=750):
fvs = USBVideoStream().start()
while True:
# while not fvs.more():
# pass
img = fvs.read_wait()
if img is not None:
img = imutils.resize(img['image'], width=width, height=height)
if mirror:
img = cv2.flip(img, 1)
cv2.imshow('FLIR cam', img)
if cv2.waitKey(1) == 27:
break # esc to quit
fvs.stop()
del fvs
cv2.destroyAllWindows()
show_FLIRcam(mirror=True)
# -
# ## Show OpenCV web camera
# +
def show_webcam(mirror=False):
cam = cv2.VideoCapture(0)
while True:
ret_val, img = cam.read()
if mirror:
img = cv2.flip(img, 1)
cv2.imshow('my webcam', img)
if cv2.waitKey(1) == 27:
break # esc to quit
cv2.destroyAllWindows()
cam.release()
show_webcam(mirror=True)
| nbs/10_Example_Sockets_and_USB_Streaming.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <p><font size="6"><b>Visualization - Matplotlib</b></font></p>
#
# > *© 2021, <NAME> and <NAME>. Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
# # Matplotlib
# [Matplotlib](http://matplotlib.org/) is a Python package used widely throughout the scientific Python community to produce high quality 2D publication graphics. It transparently supports a wide range of output formats including PNG (and other raster formats), PostScript/EPS, PDF and SVG and has interfaces for all of the major desktop GUI (graphical user interface) toolkits. It is a great package with lots of options.
#
# However, matplotlib is...
#
# > The 800-pound gorilla — and like most 800-pound gorillas, this one should probably be avoided unless you genuinely need its power, e.g., to make a **custom plot** or produce a **publication-ready** graphic.
#
# > (As we’ll see, when it comes to statistical visualization, the preferred tack might be: “do as much as you easily can in your convenience layer of choice [nvdr e.g. directly from Pandas, or with seaborn], and then use matplotlib for the rest.”)
#
# (quote used from [this](https://dansaber.wordpress.com/2016/10/02/a-dramatic-tour-through-pythons-data-visualization-landscape-including-ggplot-and-altair/) blogpost)
#
# And that's we mostly did, just use the `.plot` function of Pandas. So, why do we learn matplotlib? Well, for the *...then use matplotlib for the rest.*; at some point, somehow!
#
# Matplotlib comes with a convenience sub-package called ``pyplot`` which, for consistency with the wider matplotlib community, should always be imported as ``plt``:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ## - dry stuff - The matplotlib `Figure`, `Axes` and `Axis`
#
# At the heart of **every** plot is the figure object. The "Figure" object is the top level concept which can be drawn to one of the many output formats, or simply just to screen. Any object which can be drawn in this way is known as an "Artist" in matplotlib.
#
# Lets create our first artist using pyplot, and then show it:
fig = plt.figure()
plt.show()
# On its own, drawing the figure artist is uninteresting and will result in an empty piece of paper (that's why we didn't see anything above).
#
# By far the most useful artist in matplotlib is the **Axes** artist. The Axes artist represents the "data space" of a typical plot, a rectangular axes (the most common, but not always the case, e.g. polar plots) will have 2 (confusingly named) **Axis** artists with tick labels and tick marks.
#
# 
#
# There is no limit on the number of Axes artists which can exist on a Figure artist. Let's go ahead and create a figure with a single Axes artist, and show it using pyplot:
ax = plt.axes()
# Matplotlib's ``pyplot`` module makes the process of creating graphics easier by allowing us to skip some of the tedious Artist construction. For example, we did not need to manually create the Figure artist with ``plt.figure`` because it was implicit that we needed a figure when we created the Axes artist.
#
# Under the hood matplotlib still had to create a Figure artist, its just we didn't need to capture it into a variable.
# ## - essential stuff - `pyplot` versus Object based
# Some example data:
x = np.linspace(0, 5, 10)
y = x ** 2
# Observe the following difference:
# **1. pyplot style: plt.** (you will see this a lot for code online!)
ax = plt.plot(x, y, '-')
# **2. object oriented**
from matplotlib import ticker
# +
x = np.linspace(0, 5, 10)
y = x ** 10
fig, ax = plt.subplots()
ax.plot(x, y, '-')
ax.set_title("My data")
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter("%.1f"))
# -
# Although a little bit more code is involved, the advantage is that we now have **full control** of where the plot axes are placed, and we can easily add more than one axis to the figure:
# +
fig, ax1 = plt.subplots()
ax1.plot(x, y, '-')
ax1.set_ylabel('y')
ax2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
ax2.set_xlabel('x')
ax2.plot(x, y*2, 'r-')
# -
# And also Matplotlib advices the object oriented style:
#
# 
# <div class="alert alert-info" style="font-size:18px">
#
# <b>REMEMBER</b>:
#
# <ul>
# <li>Use the <b>object oriented</b> power of Matplotlib</li>
# <li>Get yourself used to writing <code>fig, ax = plt.subplots()</code></li>
# </ul>
# </div>
fig, ax = plt.subplots()
ax.plot(x, y, '-')
# ...
# ## An small cheat-sheet reference for some common elements
# +
x = np.linspace(-1, 0, 100)
fig, ax = plt.subplots(figsize=(10, 7))
# Adjust the created axes so that its topmost extent is 0.8 of the figure.
fig.subplots_adjust(top=0.9)
ax.plot(x, x**2, color='0.4', label='power 2')
ax.plot(x, x**3, color='0.8', linestyle='--', label='power 3')
ax.vlines(x=-0.75, ymin=0., ymax=0.8, color='0.4', linestyle='-.')
ax.fill_between(x=x, y1=x**2, y2=1.1*x**2, color='0.85')
ax.axhline(y=0.1, color='0.4', linestyle='-.')
ax.axhspan(ymin=0.65, ymax=0.75, color='0.95')
fig.suptitle('Figure title', fontsize=18,
fontweight='bold')
ax.set_title('Axes title', fontsize=16)
ax.set_xlabel('The X axis')
ax.set_ylabel('The Y axis $y=f(x)$', fontsize=16)
ax.set_xlim(-1.0, 1.1)
ax.set_ylim(-0.1, 1.)
ax.text(0.5, 0.2, 'Text centered at (0.5, 0.2)\nin data coordinates.',
horizontalalignment='center', fontsize=14)
ax.text(0.5, 0.5, 'Text centered at (0.5, 0.5)\nin relative Axes coordinates.',
horizontalalignment='center', fontsize=14,
transform=ax.transAxes, color='grey')
ax.annotate('Text pointing at (0.0, 0.75)', xy=(0.0, 0.75), xycoords="data",
xytext=(20, 40), textcoords="offset points",
horizontalalignment='left', fontsize=14,
arrowprops=dict(facecolor='black', shrink=0.05, width=1))
ax.legend(loc='lower right', frameon=True, ncol=2, fontsize=14)
# -
# Adjusting specific parts of a plot is a matter of accessing the correct element of the plot:
#
# 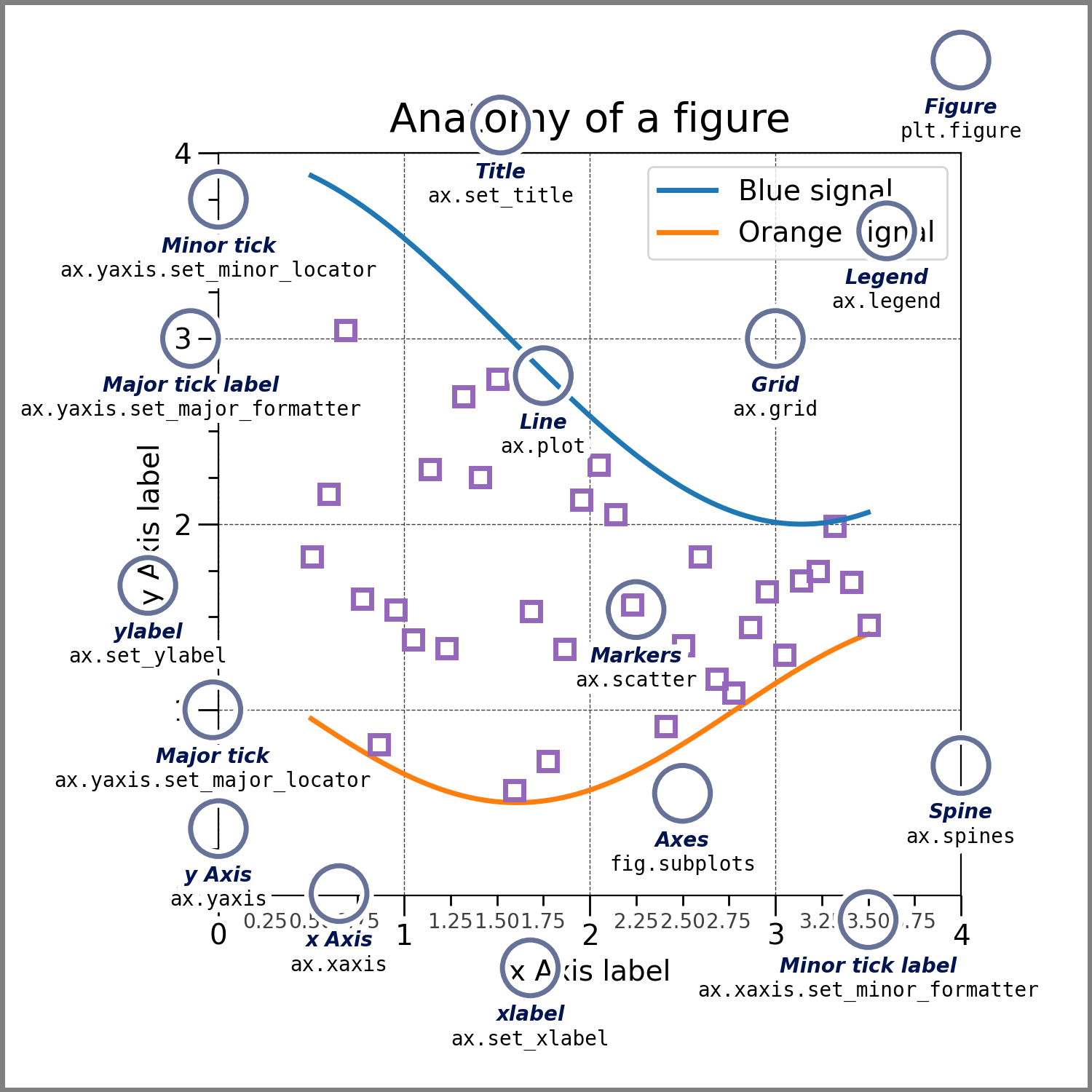
# For more information on legend positioning, check [this post](http://stackoverflow.com/questions/4700614/how-to-put-the-legend-out-of-the-plot) on stackoverflow!
# ## Exercises
# For these exercises we will use some random generated example data (as a Numpy array), representing daily measured values:
data = np.random.randint(-2, 3, 100).cumsum()
data
# <div class="alert alert-success">
#
# **EXERCISE 1**
#
# Make a line chart of the `data` using Matplotlib. The figure should be 12 (width) by 4 (height) in inches. Make the line color 'darkgrey' and provide an x-label ('days since start') and a y-label ('measured value').
#
# Use the object oriented approach to create the chart.
#
# <details><summary>Hints</summary>
#
# - When Matplotlib only receives a single input variable, it will interpret this as the variable for the y-axis
# - Check the cheat sheet above for the functions.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/visualization_01_matplotlib1.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 2**
#
# The data represents each a day starting from Jan 1st 2021. Create an array (variable name `dates`) of the same length as the original data (length 100) with the corresponding dates ('2021-01-01', '2021-01-02',...). Create the same chart as in the previous exercise, but use the `dates` values for the x-axis data.
#
# Mark the region inside `[-5, 5]` with a green color to show that these values are within an acceptable range.
#
# <details><summary>Hints</summary>
#
# - As seen in notebook `pandas_04_time_series_data`, Pandas provides a useful function `pd.date_range` to create a set of datetime values. In this case 100 values with `freq="D"`.
# - Make sure to understand the difference between `axhspan` and `fill_between`, which one do you need?
# - When adding regions, adding an `alpha` level is mostly a good idea.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/visualization_01_matplotlib2.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 3**
#
# Compare the __last ten days__ ('2021-04-01' till '2021-04-10') in a bar chart using darkgrey color. For the data on '2021-04-01', use an orange bar to highlight the measurement on this day.
#
# <details><summary>Hints</summary>
#
# - Select the last 10 days from the `data` and `dates` variable, i.e. slice [-10:].
# - Similar to a `plot` method, Matplotlib provides a `bar` method.
# - By plotting a single orange bar on top of the grey bars with a second bar chart, that one is highlithed.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/visualization_01_matplotlib3.py
# -
# ## I do not like the style...
# **...understandable**
# Matplotlib had a bad reputation in terms of its default styling as figures created with earlier versions of Matplotlib were very Matlab-lookalike and mostly not really catchy.
#
# Since Matplotlib 2.0, this has changed: https://matplotlib.org/users/dflt_style_changes.html!
#
# However...
# > *Des goûts et des couleurs, on ne discute pas...*
#
# (check [this link](https://fr.wiktionary.org/wiki/des_go%C3%BBts_et_des_couleurs,_on_ne_discute_pas) if you're not french-speaking)
#
# To account different tastes, Matplotlib provides a number of styles that can be used to quickly change a number of settings:
plt.style.available
# +
x = np.linspace(0, 10)
with plt.style.context('seaborn-whitegrid'): # 'seaborn', ggplot', 'bmh', 'grayscale', 'seaborn-whitegrid', 'seaborn-muted'
fig, ax = plt.subplots()
ax.plot(x, np.sin(x) + x + np.random.randn(50))
ax.plot(x, np.sin(x) + 0.5 * x + np.random.randn(50))
ax.plot(x, np.sin(x) + 2 * x + np.random.randn(50))
# -
# We should not start discussing about colors and styles, just pick **your favorite style**!
plt.style.use('seaborn')
# or go all the way and define your own custom style, see the [official documentation](https://matplotlib.org/3.1.1/tutorials/introductory/customizing.html) or [this tutorial](https://colcarroll.github.io/yourplotlib/#/).
# <div class="alert alert-info">
#
# <b>REMEMBER</b>:
#
# * If you just want **quickly a good-looking plot**, use one of the available styles (`plt.style.use('...')`)
# * Otherwise, creating `Figure` and `Axes` objects makes it possible to change everything!
#
# </div>
# ## Advanced subplot configuration
# The function to setup a Matplotlib Figure we have seen up to now, `fig, ax = plt.subplots()`, supports creating both a single plot and multiple subplots with a regular number of rows/columns:
fig, ax = plt.subplots(2, 3, figsize=(5, 5))
# A typical issue when plotting multiple elements in the same Figure is the overlap of the subplots. A straight-forward approach is using a larger Figure size, but this is not always possible and does not make the content independent from the Figure size. Matplotlib provides the usage of a [__constrained-layout__](https://matplotlib.org/stable/tutorials/intermediate/constrainedlayout_guide.html) to fit plots within your Figure cleanly.
fig, ax = plt.subplots(2, 3, figsize=(5, 5), constrained_layout=True)
# When more advanced layout configurations are required, the usage of the [gridspec](https://matplotlib.org/stable/api/gridspec_api.html#module-matplotlib.gridspec) module is a good reference. See [gridspec demo](https://matplotlib.org/stable/gallery/userdemo/demo_gridspec03.html#sphx-glr-gallery-userdemo-demo-gridspec03-py) for more information. A useful shortcut to know about is the [__string-shorthand__](https://matplotlib.org/stable/tutorials/provisional/mosaic.html#string-short-hand) to setup subplot layouts in a more intuitive way, e.g.
axd = plt.figure(constrained_layout=True).subplot_mosaic(
"""
ABD
CCD
"""
)
axd;
# ## Interaction with Pandas
# What we have been doing while plotting with Pandas:
import pandas as pd
flowdata = pd.read_csv('data/vmm_flowdata.csv',
index_col='Time',
parse_dates=True)
flowdata.plot.line() # remark default plot() is a line plot
# Under the hood, it creates an Matplotlib Figure with an Axes object.
# ### Pandas versus matplotlib
# #### Comparison 1: single plot
flowdata.plot(figsize=(16, 6), ylabel="Discharge m3/s") # SHIFT + TAB this!
# Making this with matplotlib...
fig, ax = plt.subplots(figsize=(16, 6))
ax.plot(flowdata)
ax.legend(["L06_347", "LS06_347", "LS06_348"])
# is still ok!
# #### Comparison 2: with subplots
axs = flowdata.plot(subplots=True, sharex=True,
figsize=(16, 8), colormap='viridis', # Dark2
fontsize=15, rot=0)
axs[0].set_title("EXAMPLE");
# Mimicking this in matplotlib (just as a reference, it is basically what Pandas is doing under the hood):
# +
from matplotlib import cm
import matplotlib.dates as mdates
colors = [cm.viridis(x) for x in np.linspace(0.0, 1.0, len(flowdata.columns))] # list comprehension to set up the colors
fig, axs = plt.subplots(3, 1, figsize=(16, 8))
for ax, col, station in zip(axs, colors, flowdata.columns):
ax.plot(flowdata.index, flowdata[station], label=station, color=col)
ax.legend()
if not ax.get_subplotspec().is_last_row():
ax.xaxis.set_ticklabels([])
ax.xaxis.set_major_locator(mdates.YearLocator())
else:
ax.xaxis.set_major_locator(mdates.YearLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y'))
ax.set_xlabel('Time')
ax.tick_params(labelsize=15)
# -
# Is already a bit harder ;-). Pandas provides as set of default configurations on top of Matplotlib.
# ### Best of both worlds...
# +
fig, (ax0, ax1) = plt.subplots(2, 1) #prepare a Matplotlib figure
flowdata.plot(ax=ax0) # use Pandas for the plotting
# +
fig, ax = plt.subplots(figsize=(15, 5)) #prepare a matplotlib figure
flowdata.plot(ax=ax) # use pandas for the plotting
# Provide further adaptations with matplotlib:
ax.set_xlabel("")
ax.grid(which="major", linewidth='0.5', color='0.8')
fig.suptitle('Flow station time series', fontsize=15)
# +
fig, (ax0, ax1) = plt.subplots(2, 1, figsize=(16, 6)) #provide with matplotlib 2 axis
flowdata[["L06_347", "LS06_347"]].plot(ax=ax0) # plot the two timeseries of the same location on the first plot
flowdata["LS06_348"].plot(ax=ax1, color='0.7') # plot the other station on the second plot
# further adapt with matplotlib
ax0.set_ylabel("L06_347")
ax1.set_ylabel("LS06_348")
ax1.legend()
# -
# <div class="alert alert-info">
#
# <b>Remember</b>:
#
# * You can do anything with matplotlib, but at a cost... <a href="http://stackoverflow.com/questions/tagged/matplotlib">stackoverflow</a>
# * The preformatting of Pandas provides mostly enough flexibility for quick analysis and draft reporting. It is not for paper-proof figures or customization
#
# If you take the time to make your perfect/spot-on/greatest-ever matplotlib-figure: Make it a <b>reusable function</b>!
#
# `fig.savefig()` to save your Figure object!
#
# </div>
# ## Exercise
flowdata = pd.read_csv('data/vmm_flowdata.csv',
index_col='Time',
parse_dates=True)
flowdata.head()
# <div class="alert alert-success">
#
# **EXERCISE 4**
#
# Pandas supports different types of charts besides line plots, all available from `.plot.xxx`, e.g. `.plot.scatter`, `.plot.bar`,... Make a bar chart to compare the mean discharge in the three measurement stations L06_347, LS06_347, LS06_348. Add a y-label 'mean discharge'. To do so, prepare a Figure and Axes with Matplotlib and add the chart to the created Axes.
#
# <details><summary>Hints</summary>
#
# * You can either use Pandas `ylabel` parameter to set the label or add it with Matploltib `ax.set_ylabel()`
# * To link an Axes object with Pandas output, pass the Axes created by `fig, ax = plt.subplots()` as parameter to the Pandas plot function.
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/visualization_01_matplotlib4.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 5**
#
# To compare the stations data, make two subplots next to each other:
#
# - In the left subplot, make a bar chart of the minimal measured value for each of the station.
# - In the right subplot, make a bar chart of the maximal measured value for each of the station.
#
# Add a title to the Figure containing 'Minimal and maximal discharge from 2009-01-01 till 2013-01-02'. Extract these dates from the data itself instead of hardcoding it.
#
# <details><summary>Hints</summary>
#
# - One can directly unpack the result of multiple axes, e.g. `fig, (ax0, ax1) = plt.subplots(1, 2,..` and link each of them to a Pands plot function.
# - Remember the remark about `constrained_layout=True` to overcome overlap with subplots?
# - A Figure title is called `suptitle` (which is different from an Axes title)
# - f-strings ([_formatted string literals_](https://docs.python.org/3/tutorial/inputoutput.html#formatted-string-literals)) is a powerful Python feature (since Python 3.6) to use variables inside a string, e.g. `f"some text with a {variable:HOWTOFORMAT}"` (with the format being optional).
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/visualization_01_matplotlib5.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 6**
#
# Make a line plot of the discharge measurements in station `LS06_347`.
#
# The main event on November 13th caused a flood event. To support the reader in the interpretation of the graph, add the following elements:
#
# - Add an horizontal red line at 20 m3/s to define the alarm level.
# - Add the text 'Alarm level' in red just above the alarm levl line.
# - Add an arrow pointing to the main peak in the data (event on November 13th) with the text 'Flood event on 2020-11-13'
#
# Check the Matplotlib documentation on [annotations](https://matplotlib.org/stable/gallery/text_labels_and_annotations/annotation_demo.html#annotating-plots) for the text annotation
#
# <details><summary>Hints</summary>
#
# - The horizontal line is explained in the cheat sheet in this notebook.
# - Whereas `ax.text` would work as well for the 'alarm level' text, the `annotate` method provides easier options to shift the text slightly relative to a data point.
# - Extract the main peak event by filtering the data on the maximum value. Different approaches are possible, but the `max()` and `idxmax()` methods are a convenient option in this case.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/visualization_01_matplotlib6.py
# -
# # Need more matplotlib inspiration?
# For more in-depth material:
# * http://www.labri.fr/perso/nrougier/teaching/matplotlib/
# * notebooks in matplotlib section: http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/Index.ipynb#4.-Visualization-with-Matplotlib
# * main reference: [matplotlib homepage](http://matplotlib.org/)
# <div class="alert alert-info" style="font-size:18px">
#
# **Galleries!**
#
# Galleries are great to get inspiration, see the plot you want, and check the code how it is created:
#
# * [matplotlib gallery](https://matplotlib.org/stable/gallery/index.html)
# * [seaborn gallery](https://seaborn.pydata.org/examples/index.html)
# * [python Graph Gallery](https://python-graph-gallery.com/)
#
# </div>
| notebooks/visualization_01_matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rt Live Model
#
# Based on [Rt.live](https://Rt.live) model, original source code on [Github](https://github.com/rtcovidlive/covid-model).
#
# Adapted for South Africa - <NAME>
#
# Last update - 19 Apr 2021
# +
# %load_ext autoreload
# %autoreload 2
from pip._internal import main
try:
import pymc3 as pm
except:
from pip._internal import main
main(['install', 'pymc3'])
import pymc3 as pm
import pandas as pd
import numpy as np
import arviz as az
from matplotlib import pyplot as plt
from covid.models.generative import GenerativeModel
from covid.data import summarize_inference_data
# %config InlineBackend.figure_format = 'retina'
from covid.data import get_and_process_covidtracking_data, summarize_inference_data
# -
# ## Setup
# While testing, run US model first to baseline the latest model files. Set `us_run` to **True** For the latest stable model, checkout commit from 2 July 2020.
#
# For automated calculation run `rtlive-model-za.py`. Github Action `rtlive-model-za.yaml` will run automatically in on Github.
us_run = False
# ## Troubleshooting
#
# 1. Ensure that all modules are installed with `conda` from `requirements.txt`
# 1. How to solve Theano library warnings on some OS platforms: [SO](https://stackoverflow.com/questions/53423610/how-to-update-scan-cython-code-in-theano)
# # Baseline model with US data
#
# For the latest stable model, checkout commit from 2 July 2020.
# ## Fetch data and select the state's data
if us_run:
df = get_and_process_covidtracking_data(run_date=pd.Timestamp.today()-pd.Timedelta(days=1))
if us_run:
region = "OR"
model_data = df.loc[region]
model_data.tail()
# ## Create the model instance and sample
if us_run:
gm = GenerativeModel(region, model_data)
gm.sample()
# ## Summarize Model Output
result = pd.DataFrame()
if us_run:
result = summarize_inference_data(gm.inference_data)
result.tail(10)
# ## Plot Model Output
# +
def plot_results():
fig, ax = plt.subplots(figsize=(10,5))
result.test_adjusted_positive.plot(c="g", label="Test-adjusted")
result.test_adjusted_positive_raw.plot(c="g", alpha=.5, label="Test-adjusted (raw)", style="--")
result.infections.plot(c="b", label="Infections")
gm.observed.positive.plot(c='r', alpha=.7, label="Reported Positives")
fig.set_facecolor('w')
ax.legend();
if us_run:
plot_results()
# +
def plot_rt():
fig, ax = plt.subplots(figsize=(10,5))
ax.set_title(f"{region} $R_t$")
samples = gm.trace['r_t']
x=result.index
cmap = plt.get_cmap("Reds")
percs = np.linspace(51, 99, 40)
colors = (percs - np.min(percs)) / (np.max(percs) - np.min(percs))
samples = samples.T
result["median"].plot(c="k", ls='-')
for i, p in enumerate(percs[::-1]):
upper = np.percentile(samples, p, axis=1)
lower = np.percentile(samples, 100-p, axis=1)
color_val = colors[i]
ax.fill_between(x, upper, lower, color=cmap(color_val), alpha=.8)
ax.axhline(1.0, c="k", lw=1, linestyle="--")
fig.set_facecolor('w')
if us_run:
plot_rt()
# -
# # South African Results
# ## Download data
url = 'https://raw.githubusercontent.com/dsfsi/covid19za/master/data/covid19za_provincial_cumulative_timeline_confirmed.csv'
states_cases = pd.read_csv(url, parse_dates=['date'], dayfirst=True, index_col=0)
states_cases.tail()
url = 'https://raw.githubusercontent.com/dsfsi/covid19za/master/data/covid19za_timeline_testing.csv'
states_tests = pd.read_csv(url, parse_dates=['date'], dayfirst=True, index_col=0)
states_tests.tail()
# ## Cleanup data
cases = pd.Series(states_cases['total'], index=states_cases.index, name='cases')
cases
casezero = states_cases.index[0]
caselast = states_cases.index[-1]
casezero, caselast
idx = pd.date_range(casezero, caselast)
tests_all = pd.Series(states_tests['cumulative_tests'], index=states_tests.index, name='tests')
tests_all
tests = tests_all.loc[casezero:caselast]
tests
combined_model = pd.concat([cases, tests], axis=1)
combined_model
# Assume previous day testing result for missing data point
combined_model.loc[casezero,'tests'] = 163
# Reindex complete date range
filled_model = combined_model.reindex(idx, method='ffill')
filled_model
# Do a final clean up, assume previous day results for missing data points
final_filled_model = filled_model.ffill(axis=0)
final_filled_model
# Add delta columns for models
final_filled_model['positive'] = final_filled_model['cases'].diff()
final_filled_model['total'] = final_filled_model['tests'].diff()
final_filled_model
df_model = final_filled_model.iloc[1:]
df_model
# ## Calculate Rt
region = 'Total RSA'
gm = GenerativeModel(region, df_model)
gm.sample()
# ## Show results
result = summarize_inference_data(gm.inference_data)
result.tail(10)
plot_results()
plot_rt()
export_results = result[['median','upper_80','lower_80','infections','test_adjusted_positive']]
export_results = export_results.rename(columns={'median':'Median','upper_80':'High_80','lower_80':'Low_80','infections':'Infections','test_adjusted_positive':'Adjusted_Postive'})
export_results.tail()
# ## Export results
export_results.to_csv('../../data/calc/calculated_rt_sa_mcmc.csv', float_format='%.3f')
| notebooks/covid-model/rtlive-model-za.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import requests
import numpy as np
import matplotlib.pyplot as plt
import json
import re
import torch
from torch import nn
from torch.utils.data import DataLoader, Dataset, SequentialSampler, RandomSampler
from torch.utils.data.distributed import DistributedSampler
import torch.nn.functional as F
with open('Key') as f:
key = f.read()
symbol = 'AMD'
interval = '1min'
# api_call = f'https://www.alphavantage.co/query?function=TIME_SERIES_INTRADAY&symbol={symbol}&interval={interval}&outputsize=full&apikey={key}'
api_call = f'https://www.alphavantage.co/query?function=TIME_SERIES_DAILY&symbol={symbol}&outputsize=full&apikey={key}'
req = requests.get(api_call)
# +
amd_ts = json.loads(req.text)
meta = amd_ts['Meta Data']
data = amd_ts['Time Series (Daily)']
amd_df = pd.DataFrame(data).T
# +
col_name = {'1. open': 'Open', '2. high': 'High', '3. low':'Low', '4. close': 'Close', '5. volume': 'Volume', 'index': 'Time'}
amd_df = amd_df.reset_index()
amd_df = amd_df.rename(columns=col_name)
amd_open = amd_df[['Time', 'Open']]
amd_open = amd_open.rename(columns={'Time':'ds', 'Open': 'y'})
# -
class regressor(torch.nn.Module):
def __init__(self):
super(regressor, self).__init__()
self.in_layer = nn.Linear(10, 32)
self.layer_1 = nn.Linear(32, 16)
self.layer_2 = nn.Linear(16, 4)
self.output = nn.Linear(4, 1)
self.ReLU = nn.ReLU()
def forward(self, x):
x = self.in_layer(x)
x = self.ReLU(x)
x = self.layer_1(x)
x = self.ReLU(x)
x = self.layer_2(x)
x = self.ReLU(x)
x = self.output(x)
return x
reg_net = regressor()
reg_net = reg_net.float()
optimizer = torch.optim.Adam(reg_net.parameters(), lr=.0001)
criterion = torch.nn.MSELoss()
# Create autoregressive dataset
dataset = pd.concat([amd_open[['y']].rename(columns={'y': f'x_{i}'}).shift(i) for i in range(1, 11)], axis=1)
dataset.head()
x = dataset.to_numpy()[10:].astype(np.double)
y = amd_open['y'].to_numpy()[10:].astype(np.double)
num_epochs = 1
for epoch in range(num_epochs):
for xi, yi in zip(x, y):
xt = torch.tensor(xi)
yt = torch.tensor(yi)
optimizer.zero_grad()
estimate = reg_net(xt.float())
loss = criterion(estimate, yt)
loss.backward()
optimizer.step()
prediction = reg_net(torch.tensor(x).float())
pred = prediction.detach().numpy()
idx = list(amd_open['ds'])
plt.plot(range(len(pred)), pred, 'ro')
plt.plot(range(len(y)), y, 'b')
plt.show()
| DataCollect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="qN8P0AnTnAhh"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="p8SrVqkmnDQv"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="AftvNA5VMemJ"
# # Federated Reconstruction for Matrix Factorization
# + [markdown] id="coAumH42q9nz"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/federated/tutorials/federated_reconstruction_for_matrix_factorization"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/federated/blob/master/docs/tutorials/federated_reconstruction_for_matrix_factorization.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/federated/blob/master/docs/tutorials/federated_reconstruction_for_matrix_factorization.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/federated/docs/tutorials/federated_reconstruction_for_matrix_factorization.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="mxV9o4VmWNti"
# This tutorial explores *partially local federated learning*, where some client parameters are never aggregated on the server. This is useful for models with user-specific parameters (e.g. matrix factorization models) and for training in communication-limited settings. We build on concepts introduced in the [Federated Learning for Image Classification](https://www.tensorflow.org/federated/tutorials/federated_learning_for_image_classification) tutorial; as in that tutorial, we introduce high-level APIs in `tff.learning` for federated training and evaluation.
#
# We begin by motivating partially local federated learning for [matrix factorization](https://en.wikipedia.org/wiki/Matrix_factorization_(recommender_systems)). We describe [Federated Reconstruction](https://arxiv.org/abs/2102.03448), a practical algorithm for partially local federated learning at scale. We prepare the MovieLens 1M dataset, build a partially local model, and train and evaluate it.
# + id="I8pu6-dckG_u"
#@test {"skip": true}
# !pip install --quiet --upgrade tensorflow-federated-nightly
# !pip install --quiet --upgrade nest-asyncio
import nest_asyncio
nest_asyncio.apply()
# + id="2txfde-th95B"
import collections
import functools
import io
import os
import requests
import zipfile
from typing import List, Optional, Tuple
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
import tensorflow_federated as tff
np.random.seed(42)
# + [markdown] id="229PrhyXaw_Y"
# ## Background: Matrix Factorization
#
# [Matrix factorization](https://en.wikipedia.org/wiki/Matrix_factorization_(recommender_systems)) has been a historically popular technique for learning recommendations and embedding representations for items based on user interactions. The canonical example is movie recommendation, where there are $n$ users and $m$ movies, and users have rated some movies. Given a user, we use their rating history and the ratings of similar users to predict the user's ratings for movies they haven't seen. If we have a model that can predict ratings, it's easy to recommend users new movies that they'll enjoy.
#
# For this task, it's useful to represent users' ratings as an $n \times m$ matrix $R$:
#
# 
#
# This matrix is generally sparse, since users typically only see a small fraction of the movies in the dataset. The output of matrix factorization is two matrices: an $n \times k$ matrix $U$ representing $k$-dimensional user embeddings for each user, and an $m \times k$ matrix $I$ representing $k$-dimensional item embeddings for each item. The simplest training objective is to ensure that the dot product of user and item embeddings are predictive of observed ratings $O$:
#
# $$argmin_{U,I} \sum_{(u, i) \in O} (R_{ui} - U_u I_i^T)^2$$
#
# This is equivalent to minimizing the mean squared error between observed ratings and ratings predicted by taking the dot product of the corresponding user and item embeddings. Another way to interpret this is that this ensures that $R \approx UI^T$ for known ratings, hence "matrix factorization". If this is confusing, don't worry–we won't need to know the details of matrix factorization for the rest of the tutorial.
# + [markdown] id="7O37nOQRvAjw"
# ## Exploring MovieLens Data
#
# Let's start by loading the [MovieLens 1M](https://grouplens.org/datasets/movielens/1m/) data, which consists of 1,000,209 movie ratings from 6040 users on 3706 movies.
# + id="DwxoBLaWneOE"
def download_movielens_data(dataset_path):
"""Downloads and copies MovieLens data to local /tmp directory."""
if dataset_path.startswith('http'):
r = requests.get(dataset_path)
z = zipfile.ZipFile(io.BytesIO(r.content))
z.extractall(path='/tmp')
else:
tf.io.gfile.makedirs('/tmp/ml-1m/')
for filename in ['ratings.dat', 'movies.dat', 'users.dat']:
tf.io.gfile.copy(
os.path.join(dataset_path, filename),
os.path.join('/tmp/ml-1m/', filename),
overwrite=True)
download_movielens_data('http://files.grouplens.org/datasets/movielens/ml-1m.zip')
# + id="Y6_bskRUniqB"
def load_movielens_data(
data_directory: str = "/tmp",
) -> Tuple[pd.DataFrame, pd.DataFrame]:
"""Loads pandas DataFrames for ratings, movies, users from data directory."""
# Load pandas DataFrames from data directory. Assuming data is formatted as
# specified in http://files.grouplens.org/datasets/movielens/ml-1m-README.txt.
ratings_df = pd.read_csv(
os.path.join(data_directory, "ml-1m", "ratings.dat"),
sep="::",
names=["UserID", "MovieID", "Rating", "Timestamp"], engine="python")
movies_df = pd.read_csv(
os.path.join(data_directory, "ml-1m", "movies.dat"),
sep="::",
names=["MovieID", "Title", "Genres"], engine="python")
# Create dictionaries mapping from old IDs to new (remapped) IDs for both
# MovieID and UserID. Use the movies and users present in ratings_df to
# determine the mapping, since movies and users without ratings are unneeded.
movie_mapping = {
old_movie: new_movie for new_movie, old_movie in enumerate(
ratings_df.MovieID.astype("category").cat.categories)
}
user_mapping = {
old_user: new_user for new_user, old_user in enumerate(
ratings_df.UserID.astype("category").cat.categories)
}
# Map each DataFrame consistently using the now-fixed mapping.
ratings_df.MovieID = ratings_df.MovieID.map(movie_mapping)
ratings_df.UserID = ratings_df.UserID.map(user_mapping)
movies_df.MovieID = movies_df.MovieID.map(movie_mapping)
# Remove nulls resulting from some movies being in movies_df but not
# ratings_df.
movies_df = movies_df[pd.notnull(movies_df.MovieID)]
return ratings_df, movies_df
# + [markdown] id="nqVrh1o9t1cZ"
# Let's load and explore a couple Pandas DataFrames containing the rating and movie data.
# + id="OkAh5nt_n4ll"
ratings_df, movies_df = load_movielens_data()
# + [markdown] id="6aNtIwvNuP7v"
# We can see that each rating example has a rating from 1-5, a corresponding UserID, a corresponding MovieID, and a timestamp.
# + id="G4qap4n-C83I"
ratings_df.head()
# + [markdown] id="5og9HO-ZubIJ"
# Each movie has a title and potentially multiple genres.
# + id="5TyN-30NC91Z"
movies_df.head()
# + [markdown] id="YWsip1k5ue5B"
# It's always a good idea to understand basic statistics of the dataset:
# + id="8I1jgmDOCqt4"
print('Num users:', len(set(ratings_df.UserID)))
print('Num movies:', len(set(ratings_df.MovieID)))
# + id="1aO07Lg21Joa"
ratings = ratings_df.Rating.tolist()
plt.hist(ratings, bins=5)
plt.xticks([1, 2, 3, 4, 5])
plt.ylabel('Count')
plt.xlabel('Rating')
plt.show()
print('Average rating:', np.mean(ratings))
print('Median rating:', np.median(ratings))
# + [markdown] id="poMbHDQguqPA"
# We can also plot the most popular movie genres.
# + id="1gYdfRoOw04z"
movie_genres_list = movies_df.Genres.tolist()
# Count the number of times each genre describes a movie.
genre_count = collections.defaultdict(int)
for genres in movie_genres_list:
curr_genres_list = genres.split('|')
for genre in curr_genres_list:
genre_count[genre] += 1
genre_name_list, genre_count_list = zip(*genre_count.items())
plt.figure(figsize=(11, 11))
plt.pie(genre_count_list, labels=genre_name_list)
plt.title('MovieLens Movie Genres')
plt.show()
# + [markdown] id="evWb8hg8vk-P"
# This data is naturally partitioned into ratings from different users, so we'd expect some heterogeneity in data between clients. Below we display the most commonly rated movie genres for different users. We can observe significant differences between users.
# + id="EfAeZ7f0GlSo"
def print_top_genres_for_user(ratings_df, movies_df, user_id):
"""Prints top movie genres for user with ID user_id."""
user_ratings_df = ratings_df[ratings_df.UserID == user_id]
movie_ids = user_ratings_df.MovieID
genre_count = collections.Counter()
for movie_id in movie_ids:
genres_string = movies_df[movies_df.MovieID == movie_id].Genres.tolist()[0]
for genre in genres_string.split('|'):
genre_count[genre] += 1
print(f'\nFor user {user_id}:')
for (genre, freq) in genre_count.most_common(5):
print(f'{genre} was rated {freq} times')
print_top_genres_for_user(ratings_df, movies_df, user_id=0)
print_top_genres_for_user(ratings_df, movies_df, user_id=10)
print_top_genres_for_user(ratings_df, movies_df, user_id=19)
# + [markdown] id="-p88NsfPwTOP"
# ## Preprocessing MovieLens Data
#
# We'll now prepare the MovieLens dataset as a list of `tf.data.Dataset`s representing each user's data for use with TFF.
#
# We implement two functions:
# * `create_tf_datasets`: takes our ratings DataFrame and produces a list of user-split `tf.data.Dataset`s.
# * `split_tf_datasets`: takes a list of datasets and splits them into train/val/test by *user*, so the val/test sets contain only ratings from users **unseen** during training. Typically in standard centralized matrix factorization we actually split so that the val/test sets contain held-out ratings from **seen** users, since unseen users don't have user embeddings. In our case, we'll see later that the approach we use to enable matrix factorization in FL also enables quickly reconstructing user embeddings for unseen users.
# + id="DHwb2AsvtIwO"
def create_tf_datasets(ratings_df: pd.DataFrame,
batch_size: int = 1,
max_examples_per_user: Optional[int] = None,
max_clients: Optional[int] = None) -> List[tf.data.Dataset]:
"""Creates TF Datasets containing the movies and ratings for all users."""
num_users = len(set(ratings_df.UserID))
# Optionally limit to `max_clients` to speed up data loading.
if max_clients is not None:
num_users = min(num_users, max_clients)
def rating_batch_map_fn(rating_batch):
"""Maps a rating batch to an OrderedDict with tensor values."""
# Each example looks like: {x: movie_id, y: rating}.
# We won't need the UserID since each client will only look at their own
# data.
return collections.OrderedDict([
("x", tf.cast(rating_batch[:, 1:2], tf.int64)),
("y", tf.cast(rating_batch[:, 2:3], tf.float32))
])
tf_datasets = []
for user_id in range(num_users):
# Get subset of ratings_df belonging to a particular user.
user_ratings_df = ratings_df[ratings_df.UserID == user_id]
tf_dataset = tf.data.Dataset.from_tensor_slices(user_ratings_df)
# Define preprocessing operations.
tf_dataset = tf_dataset.take(max_examples_per_user).shuffle(
buffer_size=max_examples_per_user, seed=42).batch(batch_size).map(
rating_batch_map_fn,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
tf_datasets.append(tf_dataset)
return tf_datasets
def split_tf_datasets(
tf_datasets: List[tf.data.Dataset],
train_fraction: float = 0.8,
val_fraction: float = 0.1,
) -> Tuple[List[tf.data.Dataset], List[tf.data.Dataset], List[tf.data.Dataset]]:
"""Splits a list of user TF datasets into train/val/test by user.
"""
np.random.seed(42)
np.random.shuffle(tf_datasets)
train_idx = int(len(tf_datasets) * train_fraction)
val_idx = int(len(tf_datasets) * (train_fraction + val_fraction))
# Note that the val and test data contains completely different users, not
# just unseen ratings from train users.
return (tf_datasets[:train_idx], tf_datasets[train_idx:val_idx],
tf_datasets[val_idx:])
# + id="T6pJVpHfns9q"
# We limit the number of clients to speed up dataset creation. Feel free to pass
# max_clients=None to load all clients' data.
tf_datasets = create_tf_datasets(
ratings_df=ratings_df,
batch_size=5,
max_examples_per_user=300,
max_clients=2000)
# Split the ratings into training/val/test by client.
tf_train_datasets, tf_val_datasets, tf_test_datasets = split_tf_datasets(
tf_datasets,
train_fraction=0.8,
val_fraction=0.1)
# + [markdown] id="T2SdGARZ0-cm"
# As a quick check, we can print a batch of training data. We can see that each individual example contains a MovieID under the "x" key and a rating under the "y" key. Note that we won't need the UserID since each user only sees their own data.
# + id="9D2rCgcwFP4E"
print(next(iter(tf_train_datasets[0])))
# + [markdown] id="VOaSLuFK18G7"
# We can plot a histogram showing the number of ratings per user.
# + id="98VwSFBe1GPM"
def count_examples(curr_count, batch):
return curr_count + tf.size(batch['x'])
num_examples_list = []
# Compute number of examples for every other user.
for i in range(0, len(tf_train_datasets), 2):
num_examples = tf_train_datasets[i].reduce(tf.constant(0), count_examples).numpy()
num_examples_list.append(num_examples)
plt.hist(num_examples_list, bins=10)
plt.ylabel('Count')
plt.xlabel('Number of Examples')
plt.show()
# + [markdown] id="eqz6oRm22FWM"
# Now that we've loaded and explored the data, we'll discuss how to bring matrix factorization to federated learning. Along the way, we'll motivate partially local federated learning.
# + [markdown] id="PMZLj5WprMJM"
# ## Bringing Matrix Factorization to FL
#
# While matrix factorization has been traditionally used in centralized settings, it's especially relevant in federated learning: user ratings may live on separate client devices, and we may want to learn embeddings and recommendations for users and items without centralizing the data. Since each user has a corresponding user embedding, it's natural to have each client store their user embedding–this scales much better than a central server storing all the user embeddings.
#
# One proposal for bringing matrix factorization to FL goes as follows:
# 1. The server stores and sends the item matrix $I$ to sampled clients each round
# 2. Clients update the item matrix and their personal user embedding $U_u$ using SGD on the above objective
# 3. Updates to $I$ are aggregated on the server, updating the server copy of $I$ for the next round
#
# This approach is *partially local*–that is, some client parameters are never aggregated by the server. Though this approach is appealing, it requires clients to maintain state across rounds, namely their user embeddings. Stateful federated algorithms are less appropriate for cross-device FL settings: in these settings the population size is often much larger than the number of clients that participate in each round, and a client usually participates at most once during the training process. Besides relying on state that may not be initialized, stateful algorithms can result in performance degradation in cross-device settings due to state getting *stale* when clients are infrequently sampled. Importantly, in the matrix factorization setting, a stateful algorithm leads to all unseen clients missing trained user embeddings, and in large-scale training the majority of users may be unseen. For more on the motivation for stateless algorithms in cross-device FL, see [Wang et al. 2021 Sec. 3.1.1](https://arxiv.org/pdf/2107.06917.pdf) and [Reddi et al. 2020 Sec. 5.1](https://arxiv.org/abs/2003.00295).
#
# Federated Reconstruction ([Singhal et al. 2021](https://arxiv.org/abs/2102.03448)) is a stateless alternative to the aforementioned approach. The key idea is that instead of storing user embeddings across rounds, clients reconstruct user embeddings when needed. When FedRecon is applied to matrix factorization, training proceeds as follows:
# 1. The server stores and sends the item matrix $I$ to sampled clients each round
# 2. Each client freezes $I$ and trains their user embedding $U_u$ using one or more steps of SGD (reconstruction)
# 3. Each client freezes $U_u$ and trains $I$ using one or more steps of SGD
# 4. Updates to $I$ are aggregated across users, updating the server copy of $I$ for the next round
#
# This approach does not require clients to maintain state across rounds. The authors also show in the paper that this method leads to fast reconstruction of user embeddings for unseen clients (Sec. 4.2, Fig. 3, and Table 1), allowing the majority of clients who do not participate in training to have a trained model, enabling recommendations for these clients.
# + [markdown] id="imwLf1zksCjN"
# ## Defining the Model
#
# We'll next define the local matrix factorization model to be trained on client devices. This model will include the full item matrix $I$ and a single user embedding $U_u$ for client $u$. Note that clients will not need to store the full user matrix $U$.
#
# We'll define the following:
# - `UserEmbedding`: a simple Keras layer representing a single `num_latent_factors`-dimensional user embedding.
# - `get_matrix_factorization_model`: a function that returns a [`tff.learning.reconstruction.Model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/Model) containing the model logic, including which layers are globally aggregated on the server and which layers remain local. We need this additional information to initialize the Federated Reconstruction training process. Here we produce the `tff.learning.reconstruction.Model` from a Keras model using [`tff.learning.reconstruction.from_keras_model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/from_keras_model). Similar to `tff.learning.Model`, we can also implement a custom `tff.learning.reconstruction.Model` by implementing the class interface.
# + id="nSLMxPDP3D72"
class UserEmbedding(tf.keras.layers.Layer):
"""Keras layer representing an embedding for a single user, used below."""
def __init__(self, num_latent_factors, **kwargs):
super().__init__(**kwargs)
self.num_latent_factors = num_latent_factors
def build(self, input_shape):
self.embedding = self.add_weight(
shape=(1, self.num_latent_factors),
initializer='uniform',
dtype=tf.float32,
name='UserEmbeddingKernel')
super().build(input_shape)
def call(self, inputs):
return self.embedding
def compute_output_shape(self):
return (1, self.num_latent_factors)
def get_matrix_factorization_model(
num_items: int,
num_latent_factors: int) -> tff.learning.reconstruction.Model:
"""Defines a Keras matrix factorization model."""
# Layers with variables will be partitioned into global and local layers.
# We'll pass this to `tff.learning.reconstruction.from_keras_model`.
global_layers = []
local_layers = []
# Extract the item embedding.
item_input = tf.keras.layers.Input(shape=[1], name='Item')
item_embedding_layer = tf.keras.layers.Embedding(
num_items,
num_latent_factors,
name='ItemEmbedding')
global_layers.append(item_embedding_layer)
flat_item_vec = tf.keras.layers.Flatten(name='FlattenItems')(
item_embedding_layer(item_input))
# Extract the user embedding.
user_embedding_layer = UserEmbedding(
num_latent_factors,
name='UserEmbedding')
local_layers.append(user_embedding_layer)
# The item_input never gets used by the user embedding layer,
# but this allows the model to directly use the user embedding.
flat_user_vec = user_embedding_layer(item_input)
# Compute the dot product between the user embedding, and the item one.
pred = tf.keras.layers.Dot(
1, normalize=False, name='Dot')([flat_user_vec, flat_item_vec])
input_spec = collections.OrderedDict(
x=tf.TensorSpec(shape=[None, 1], dtype=tf.int64),
y=tf.TensorSpec(shape=[None, 1], dtype=tf.float32))
model = tf.keras.Model(inputs=item_input, outputs=pred)
return tff.learning.reconstruction.from_keras_model(
keras_model=model,
global_layers=global_layers,
local_layers=local_layers,
input_spec=input_spec)
# + [markdown] id="-B3FPaRiwY3n"
# Analagous to the interface for Federated Averaging, the interface for Federated Reconstruction expects a `model_fn` with no arguments that returns a `tff.learning.reconstruction.Model`.
# + id="vNBRQW9EwneZ"
# This will be used to produce our training process.
# User and item embeddings will be 50-dimensional.
model_fn = functools.partial(
get_matrix_factorization_model,
num_items=3706,
num_latent_factors=50)
# + [markdown] id="fQVpVIfnwvPg"
# We'll next define `loss_fn` and `metrics_fn`, where `loss_fn` is a no-argument function returning a Keras loss to use to train the model, and `metrics_fn` is a no-argument function returning a list of Keras metrics for evaluation. These are needed to build the training and evaluation computations.
#
# We'll use Mean Squared Error as the loss, as mentioned above. For evaluation we'll use rating accuracy (when the model's predicted dot product is rounded to the nearest whole number, how often does it match the label rating?).
# + id="FDJUfeSNwxIL"
class RatingAccuracy(tf.keras.metrics.Mean):
"""Keras metric computing accuracy of reconstructed ratings."""
def __init__(self,
name: str = 'rating_accuracy',
**kwargs):
super().__init__(name=name, **kwargs)
def update_state(self,
y_true: tf.Tensor,
y_pred: tf.Tensor,
sample_weight: Optional[tf.Tensor] = None):
absolute_diffs = tf.abs(y_true - y_pred)
# A [batch_size, 1] tf.bool tensor indicating correctness within the
# threshold for each example in a batch. A 0.5 threshold corresponds
# to correctness when predictions are rounded to the nearest whole
# number.
example_accuracies = tf.less_equal(absolute_diffs, 0.5)
super().update_state(example_accuracies, sample_weight=sample_weight)
loss_fn = lambda: tf.keras.losses.MeanSquaredError()
metrics_fn = lambda: [RatingAccuracy()]
# + [markdown] id="ecM_vru8xg2j"
# ## Training and Evaluation
#
# Now we have everything we need to define the training process. One important difference from the [interface for Federated Averaging](https://www.tensorflow.org/federated/api_docs/python/tff/learning/build_federated_averaging_process) is that we now pass in a `reconstruction_optimizer_fn`, which will be used when reconstructing local parameters (in our case, user embeddings). It's generally reasonable to use `SGD` here, with a similar or slightly lower learning rate than the client optimizer learning rate. We provide a working configuration below. This hasn't been carefully tuned, so feel free to play around with different values.
#
# Check out the [documentation](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/build_training_process) for more details and options.
# + id="YQsX0FgtwsoE"
# We'll use this by doing:
# state = training_process.initialize()
# state, metrics = training_process.next(state, federated_train_data)
training_process = tff.learning.reconstruction.build_training_process(
model_fn=model_fn,
loss_fn=loss_fn,
metrics_fn=metrics_fn,
server_optimizer_fn=lambda: tf.keras.optimizers.SGD(1.0),
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(0.5),
reconstruction_optimizer_fn=lambda: tf.keras.optimizers.SGD(0.1))
# + [markdown] id="ssbbds4vzhXJ"
# We can also define a computation for evaluating our trained global model.
# + id="KHi7J330PtxO"
# We'll use this by doing:
# eval_metrics = evaluation_computation(state.model, tf_val_datasets)
# where `state` is the state from the training process above.
evaluation_computation = tff.learning.reconstruction.build_federated_evaluation(
model_fn,
loss_fn=loss_fn,
metrics_fn=metrics_fn,
reconstruction_optimizer_fn=functools.partial(
tf.keras.optimizers.SGD, 0.1))
# + [markdown] id="h_V_ZwlE0DSl"
# We can initialize the training process state and examine it. Most importantly, we can see that this server state only stores item variables (currently randomly initialized) and not any user embeddings.
# + id="I_kOjFVKQoNX"
state = training_process.initialize()
print(state.model)
print('Item variables shape:', state.model.trainable[0].shape)
# + [markdown] id="yPFqgTV21lJO"
# We can also try to evaluate our randomly initialized model on validation clients. Federated Reconstruction evaluation here involves the following:
#
# 1. The server sends the item matrix $I$ to sampled evaluation clients
# 2. Each client freezes $I$ and trains their user embedding $U_u$ using one or more steps of SGD (reconstruction)
# 3. Each client calculates loss and metrics using the server $I$ and reconstructed $U_u$ on an unseen portion of their local data
# 4. Losses and metrics are averaged across users to calculate overall loss and metrics
#
# Note that steps 1 and 2 are the same as for training. This connection is important, since training the same way we evaluate leads to a form of *meta-learning*, or learning how to learn. In this case, the model is learning how to learn global variables (item matrix) that lead to performant reconstruction of local variables (user embeddings). For more on this, see [Sec. 4.2](https://arxiv.org/abs/2102.03448) of the paper.
#
# It's also important for steps 2 and 3 to be performed using disjoint portions of clients' local data, to ensure fair evaluation. By default, both the training process and evaluation computation use every other example for reconstruction and use the other half post-reconstruction. This behavior can be customized using the `dataset_split_fn` argument (we'll explore this further later).
# + id="JiBOGFsWWBiU"
# We shouldn't expect good evaluation results here, since we haven't trained
# yet!
eval_metrics = evaluation_computation(state.model, tf_val_datasets)
print('Initial Eval:', eval_metrics['eval'])
# + [markdown] id="aZUZwjWp4iJu"
# We can next try running a round of training. To make things more realistic, we'll sample 50 clients per round randomly without replacement. We should still expect train metrics to be poor, since we're only doing one round of training.
# + id="lOTfqrVcVfJf"
federated_train_data = np.random.choice(tf_train_datasets, size=50, replace=False).tolist()
state, metrics = training_process.next(state, federated_train_data)
print(f'Train metrics:', metrics['train'])
# + [markdown] id="Rr3ZS9jz5Mj0"
# Now let's set up a training loop to train over multiple rounds.
# + id="VJBzOPNYwp9q"
NUM_ROUNDS = 20
train_losses = []
train_accs = []
state = training_process.initialize()
# This may take a couple minutes to run.
for i in range(NUM_ROUNDS):
federated_train_data = np.random.choice(tf_train_datasets, size=50, replace=False).tolist()
state, metrics = training_process.next(state, federated_train_data)
print(f'Train round {i}:', metrics['train'])
train_losses.append(metrics['train']['loss'])
train_accs.append(metrics['train']['rating_accuracy'])
eval_metrics = evaluation_computation(state.model, tf_val_datasets)
print('Final Eval:', eval_metrics['eval'])
# + [markdown] id="yM-jAGNm5di7"
# We can plot training loss and accuracy over rounds. The hyperparameters in this notebook have not been carefully tuned, so feel free to try different clients per round, learning rates, number of rounds, and total number of clients to improve these results.
# + id="h6w702JmR-3V"
plt.plot(range(NUM_ROUNDS), train_losses)
plt.ylabel('Train Loss')
plt.xlabel('Round')
plt.title('Train Loss')
plt.show()
plt.plot(range(NUM_ROUNDS), train_accs)
plt.ylabel('Train Accuracy')
plt.xlabel('Round')
plt.title('Train Accuracy')
plt.show()
# + [markdown] id="ZTzKkT-a5kgX"
# Finally, we can calculate metrics on an unseen test set when we're finished tuning.
# + id="Iq0UxEBBJcR-"
eval_metrics = evaluation_computation(state.model, tf_test_datasets)
print('Final Test:', eval_metrics['eval'])
# + [markdown] id="Mr2fRxic6Lfi"
# ## Further Explorations
#
# Nice work on completing this notebook. We suggest the following exercises to explore partially local federated learning further, roughly ordered by increasing difficulty:
#
# * Typical implementations of Federated Averaging take multiple local passes (epochs) over the data (in addition to taking one pass over the data across multiple batches). For Federated Reconstruction we may want to control the number of steps separately for reconstruction and post-reconstruction training. Passing the `dataset_split_fn` argument to the training and evaluation computation builders enables control of the number of steps and epochs over both reconstruction and post-reconstruction datasets. As an exercise, try performing 3 local epochs of reconstruction training, capped at 50 steps and 1 local epoch of post-reconstruction training, capped at 50 steps. Hint: you'll find [`tff.learning.reconstruction.build_dataset_split_fn`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/build_dataset_split_fn) helpful. Once you've done this, try tuning these hyperparameters and other related ones like learning rates and batch size to get better results.
#
# * The default behavior of Federated Reconstruction training and evaluation is to split clients' local data in half for each of reconstruction and post-reconstruction. In cases where clients have very little local data, it can be reasonable to reuse data for reconstruction and post-reconstruction for the training process only (not for evaluation, this will lead to unfair evaluation). Try making this change for the training process, ensuring the `dataset_split_fn` for evaluation still keeps reconstruction and post-reconstruction data disjoint. Hint: [`tff.learning.reconstruction.simple_dataset_split_fn`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/simple_dataset_split_fn) might be useful.
#
# * Above, we produced a `tff.learning.Model` from a Keras model using `tff.learning.reconstruction.from_keras_model`. We can also implement a custom model using pure TensorFlow 2.0 by [implementing the model interface](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/Model). Try modifying `get_matrix_factorization_model` to build and return a class that extends `tff.learning.reconstruction.Model`, implementing its methods. Hint: the source code of [`tff.learning.reconstruction.from_keras_model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/reconstruction/from_keras_model) provides an example of extending the `tff.learning.reconstruction.Model` class. Refer also to the [custom model implementation in the EMNIST image classification tutorial](https://www.tensorflow.org/federated/tutorials/federated_learning_for_image_classification#customizing_the_model_implementation) for a similar exercise in extending a `tff.learning.Model`.
#
# * In this tutorial, we've motivated partially local federated learning in the context of matrix factorization, where sending user embeddings to the server would trivially leak user preferences. We can also apply Federated Reconstruction in other settings as a way to train more personal models (since part of the model is completely local to each user) while reducing communication (since local parameters are not sent to the server). In general, using the interface presented here we can take any federated model that would typically be trained fully globally and instead partition its variables into global variables and local variables. The example explored in the [Federated Reconstruction paper](https://arxiv.org/abs/2102.03448) is personal next word prediction: here, each user has their own local set of word embeddings for out-of-vocabulary words, enabling the model to capture users' slang and achieve personalization without additional communication. As an exercise, try implementing (as either a Keras model or a custom TensorFlow 2.0 model) a different model for use with Federated Reconstruction. A suggestion: implement an EMNIST classification model with a personal user embedding, where the personal user embedding is concatenated to the CNN image features before the last Dense layer of the model. You can reuse much of the code from this tutorial (e.g. the `UserEmbedding` class) and the [image classification tutorial](https://www.tensorflow.org/federated/tutorials/federated_learning_for_image_classification).
#
# \
# If you're still looking for more on partially local federated learning, check out the [Federated Reconstruction paper](https://arxiv.org/abs/2102.03448) and [open-source experiment code](https://github.com/google-research/federated/tree/master/reconstruction).
| site/en-snapshot/federated/tutorials/federated_reconstruction_for_matrix_factorization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
tsv_file='clinical.tsv'
csv_table=pd.read_table(tsv_file,sep='\t')
csv_table.to_csv('clinical.csv',index=False)
df = pd.DataFrame(csv_table)
df.to_csv('Users\crystalrubalcava\Desktop\final_project\clinical.csv', index=False)
| Scripts/python_tsv_to_csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/amindazad/DS-Unit-2-Linear-Models/blob/master/DS-Unit-2-Linear-Models/module4-logistic-regression/Amin_Azad_LS_DS_214_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1dThjul9bJZ9" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# ## Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [X] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [X] Begin with baselines for classification.
# - [X] Use scikit-learn for logistic regression.
# - [X] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [X] Get your model's test accuracy. (One time, at the end.)
# - [X] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Make exploratory visualizations.
# - [ ] Do one-hot encoding.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Get and plot your coefficients.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="kkd3jjUnbJaE" colab_type="code" colab={}
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# + id="khc4GfQRbJaH" colab_type="code" colab={}
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# + id="II_sOEgxbJaK" colab_type="code" colab={}
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# + id="A0rPb2btbJaN" colab_type="code" colab={}
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# + id="-kVPJ8embJaQ" colab_type="code" colab={}
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
# + id="vcKxrZ7YbJaT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="f499ae4d-5d35-4f60-ce27-83e51f906004"
# Scanning the dataset ...
df
# + id="Avdpk6uWbd1C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="64c96d74-bd93-44cc-ca6a-edcc56364473"
# Checking data types
df.dtypes
# + id="cilGEMiKbvbe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="eae87b1c-8c2f-4aaa-e37b-2f17b9b20a5b"
# Convert the date column to pandas date data format
import pandas as pd
df['Date'] = pd.to_datetime(df['Date'], infer_datetime_format=True)
df['Date']
# + id="fGtMyZNeckIZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="41da6c17-2476-4b42-c520-63ce20d137bb"
# train/validate/test split, Train = 2016 & earlier. Validate = 2017. Test = 2018 & later.
train = df[(df['Date']>='2016-01-01')&(df['Date']<'2017-01-01')]
val = df[(df['Date']>='2017-01-01')&(df['Date']<'2018-01-01')]
test = df[df['Date']>='2018-01-01']
train.shape, val.shape ,test.shape
# + id="u9qjV58Ufazn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="6cdcb979-e69e-47fc-cf6e-0834c8468a60"
# Calculate baseline for classification
# Lets start with defining a target
target = 'Great'
y_train = train[target]
y_train.value_counts(normalize=True)
# + id="JVZ-RZDDk4mz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a552cf68-1018-46c6-a46e-51cbcbc69ff9"
from sklearn.metrics import accuracy_score
#Define the baseline by assuming if we guessed how many times would have been right?
majority_class=y_train.mode()[0]
y_pred=[majority_class]*len(y_train)
#Calculate the accuracy score
accuracy_score(y_train, y_pred)
# + id="AiDUecaQyGXi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 322} outputId="3764c0d1-2fe0-4162-eff9-f19e29a5682e"
df.head()
# + id="yEuhUbDbCQ5L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2b642eac-dbe1-4665-ea03-602971e07ac6"
#OneHotCoding the features
#checking the non numeric values and sort them by cordinality
print(train.shape)
train.describe(exclude='number').T.sort_values(by='unique')
# + id="RN3GyBOhDWrB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 425} outputId="364078fe-dfb0-4df1-87d3-1aced3cf7d87"
import category_encoders as ce
encoder = ce.OneHotEncoder(use_cat_names=True)
X_train_encoded = encoder.fit_transform(X_train)
X_train_encoded
# + id="Oqx0i1Fexb0H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="7c292696-2f43-4546-db2b-6c2b7f5913a3"
#Logistic regression
import category_encoders as ce
encoder = ce.OneHotEncoder(use_cat_names=True)
# Select features and get rid of Nan values
features = ['Yelp', 'Google','Cost','Hunger', 'Tortilla', 'Temp', 'Meat', 'Wrap', 'Burrito']
#Estimator API 5 steps
#1. Import estimator class
from sklearn.linear_model import LogisticRegression
#2. Instantiate the model
log_reg = LogisticRegression(solver='lbfgs')
#3. Arrange X feature matrices and y vectors and clean Nans meanwhile
from sklearn.impute import SimpleImputer
X_train = train[features]
X_train_encoded = encoder.fit_transform(X_train)
X_train_imputed = imputer.fit_transform(X_train_encoded)
y_train = train[target]
X_val = val[features]
X_val_encoded = encoder.transform(X_val)
X_val_imputed = imputer.transform(X_val_encoded)
y_val = val[target]
#4.Fit the model
log_reg.fit(X_train_imputed, y_train)
#5. Apply the model to new data
log_reg.predict(X_val_imputed)
log_reg.score(X_val_imputed, y_val)
# + id="kf0uFLAI07Wb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="df434758-616a-4875-f2ec-4ec8d1f4f19a"
#Get the test accuracy
X_test = test[features]
X_test_encoded = encoder.transform(X_test)
X_test_imputed = imputer.transform(X_test_encoded)
y_pred = log_reg.predict(X_test_imputed)
y_test = test[target]
log_reg.score(X_test_imputed, y_test)
# + id="Tt9n9YQuA_5p" colab_type="code" colab={}
| DS-Unit-2-Linear-Models/module4-logistic-regression/Amin_Azad_LS_DS_214_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from functools import reduce
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
# %config InlineBackend.figure_format = 'retina'
# %matplotlib inline
plt.style.use('seaborn')
# -
# # Import __[Global Ecological Footprint 2016](https://www.kaggle.com/footprintnetwork/ecological-footprint)__ dataset
ecological = pd.read_csv("./Data/countries.csv")
ecological
# # Import __[World Happiness Report 2016](https://www.kaggle.com/unsdsn/world-happiness)__ dataset
happiness = pd.read_csv("./Data/2016.csv")
happiness
# # Remove countries that aren't on both datasets
#
# Note that I had to name Vietnam to be the same in the csv files
data = reduce(lambda left, right: pd.merge(left, right, on='Country'), [ecological, happiness])
data
# # Remove countries with NaNs as any element
print(data.isnull().values.any()) # Check if any contain NaNs
data[data.isna().any(axis=1)] # Display which ones contain NaN
data = data.dropna(how='any')
data
# # Output CSV
# +
data['GDP per Capita'] = data['GDP per Capita'].replace('[\$,]', '', regex=True).astype(float)
data
data.to_csv('./Data/data.csv', index = False)
# -
# # Import CSV
#
data = pd.read_csv("./Data/data.csv")
data
data['GDP per Capita'] = data['GDP per Capita'].replace('[\$,]', '', regex=True).astype(float)
data
# +
plt.style.use('seaborn')
plt.figure()
plt.scatter(data['GDP per Capita'], np.log(data['HDI']), marker='o', color='r')
plt.scatter(data['GDP per Capita'], np.log(data['Happiness Score']), marker='o', color='b')
plt.scatter(data['GDP per Capita'], np.log(data['Earths Required']), marker='o', color='g')
plt.title('HDI vs GDP per Capita')
plt.xlabel('GDP per Capita')
plt.show()
# +
data = data.sort_values('HDI')
plt.figure(figsize=(20, 10))
region_data = {k : [] for k in set(data['Region_x'])}
for k, d in data.groupby('Region_x'):
region_data[k].append(np.mean(d.HDI))
region_data[k].append(np.var(d.HDI))
region_data[k].append(np.max(d.HDI) - np.min(d.HDI))
region_data[k].append(np.mean(d['GDP per Capita']))
region_data[k].append(np.var(d['GDP per Capita']))
region_data[k].append(np.max(d['GDP per Capita']) - np.min(d['GDP per Capita']))
plt.bar(d.Country, d.HDI, width=.4, align='edge')
plt.xticks(rotation=90)
plt.show()
df = pd.DataFrame(region_data)
df.index = ['HDI: Mean', 'HDI: Variance', 'HDI: Range', 'GDP: Mean','GDP: Variance', 'GDP: Range']
df.T.plot.bar()
plt.figure(figsize=(20, 10))
plt.scatter(df.T['HDI: Mean'], df.T['GDP: Mean'])
# -
#plt.figure(figsize=(20, 10))
plt.figure(figsize=(20, 10))
labels = []
for k, d in data.groupby('Region_x'):
labels.append(str(k))
plt.plot(d.HDI, d['GDP per Capita'])
plt.legend(labels)
plt.show()
| .ipynb_checkpoints/Data cleanup-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# data['rate'] = data.gains_uvxy - data.gains_spy
# 对rate 进行正态判断,大于1个std,进行操作
# -
import yfinance as yf
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import statsmodels
from statsmodels.tsa.stattools import coint
import pandas as pd
import scipy.stats as st
import os
import math
# +
reload = True
file_name = 'uvxy_300d'
if not reload and os.path.exists(file_name):
hist = pd.read_csv(file_name)
else:
tmp = yf.Ticker("UVXY")
hist = tmp.history(period="300d")
hist.to_csv(file_name)
file_name = 'spy_300d'
if not reload and os.path.exists(file_name):
spy_hist = pd.read_csv(file_name)
else:
tmp = yf.Ticker("SPY")
spy_hist = tmp.history(period="300d")
spy_hist.to_csv(file_name)
print(hist.shape)
print(spy_hist.shape)
# -
hist.shape
# +
# hist.head(10)
# -
hist['gains'] = (hist.Close - hist.Close.shift(1))/hist.Close.shift(1) * 100
spy_hist['gains'] = (spy_hist.Close - spy_hist.Close.shift(1))/spy_hist.Close.shift(1) * 100
hist['gains'].describe()
spy_hist['gains'].describe()
sns.distplot(hist.gains)
# +
# st.norm.cdf(-15, hist['gains'].mean(), hist['gains'].std())
# -
sns.distplot(spy_hist.gains)
st.norm.cdf(-2, spy_hist['gains'].mean(), spy_hist['gains'].std())
spy_hist.Close.plot(figsize=(10,10))
hist.Close.plot()
spy_hist.gains.fillna(0, inplace=True)
hist.gains.fillna(0, inplace=True)
# +
# hist.gains.cumsum().plot()
# spy_hist.gains.cumsum().plot()
# +
# 差分
hist['close_diff'] = hist.Close - hist.Close.shift(1) + 1000
spy_hist['close_diff'] = spy_hist.Close - spy_hist.Close.shift(1) + 1000
hist.dropna(inplace=True)
spy_hist.dropna(inplace=True)
hist.head()
# -
# 对数
hist['close_log'] = hist['Close'].apply(math.log)
spy_hist['close_log'] = spy_hist['Close'].apply(math.log)
spy_hist.head()
# +
score, pvalue, _ = coint(hist.gains, spy_hist.gains)
print(score, pvalue)
score, pvalue, _ = coint(hist.Close, spy_hist.Close)
print(score, pvalue)
score, pvalue, _ = coint(hist.close_diff, spy_hist.close_diff)
print(score, pvalue)
score, pvalue, _ = coint(hist.close_log, spy_hist.close_log)
print(score, pvalue)
# -
print(np.corrcoef(hist.Close, spy_hist.Close))
print(np.corrcoef(hist.close_diff, spy_hist.close_diff))
print(np.corrcoef(spy_hist.gains, hist.gains))
hist.reset_index(inplace=True)
spy_hist.reset_index(inplace=True)
col = ['Date','gains','Close','close_diff']
data = pd.merge(hist[col], spy_hist[col],suffixes=['_uvxy','_spy'], on='Date')
data.set_index(keys='Date')
# data['rate'] = (data.gains_uvxy+1) / (data.gains_spy+1)
# data['rate'] = data.gains_uvxy - data.gains_spy # 这个收益是 150
data['rate'] = data.close_diff_uvxy / data.close_diff_spy # 差分+1000,这个收益是 188.78997135162348
data
data.rate.describe()
sns.distplot(data.rate)
data.fillna(0,inplace=True)
data.isna().sum()
print((data.rate>50).sum())
# data = data[abs(data['rate'])<50]
(data.rate>50).sum()
# data.rate.plot()
# plt.axhline(data.rate.mean(), color='red', linestyle='--')
# +
def zscore(series):
print(series.mean(), series.std())
return (series - series.mean()) / np.std(series)
z_score = zscore(data.rate)
z_score.plot(figsize=(10,10))
plt.axhline(z_score.mean())
plt.axhline(1.0, color='red')
plt.axhline(-1.0, color='green')
plt.show()
# -
# # 开始预测
# +
print(z_score.shape)
train = z_score[:200]
test = z_score[200:]
print(train.shape, test.shape)
plt.figure(figsize=(15,7))
train.plot()
buy = train.copy()
sell = train.copy()
buy[train>-1] = 0
sell[train<1] = 0
# buy[~((data['gains_uvxy']>-1) & (data['gains_spy']>0))] = 0
# sell[~((data['gains_uvxy']<1) & (data['gains_spy']<0))] = 0
buy.plot(color='g', linestyle='None', marker='^')
sell.plot(color='r', linestyle='None', marker='^')
x1,x2,y1,y2 = plt.axis()
# plt.axis((x1,x2,data.rate.min(),data.rate.max()))
plt.legend(['Ratio', 'Buy Signal', 'Sell Signal'])
plt.show()
# +
# Plot the prices and buy and sell signals from z score
plt.figure(figsize=(10,10))
S1 = hist.iloc[:200].Close
S2 = spy_hist.iloc[:200].Close
S1.plot(color='b')
S2.plot(color='c')
buyR = 0*S1.copy()
sellR = 0*S1.copy()
# When buying the ratio, buy S1 and sell S2
buyR[buy!=0] = S1[buy!=0]
sellR[buy!=0] = S2[buy!=0]
# When selling the ratio, sell S1 and buy S2
buyR[sell!=0] = S2[sell!=0]
sellR[sell!=0] = S1[sell!=0]
buyR.plot(color='g', linestyle='None', marker='^')
sellR.plot(color='r', linestyle='None', marker='^')
tmp = 150+ 10*z_score[0:200]
tmp.plot()
x1,x2,y1,y2 = plt.axis()
plt.axis((x1,x2,min(S1.min(),S2.min()),max(S1.max(),S2.max())))
plt.legend(['UVXY','SPY', 'Buy Signal', 'Sell Signal'])
plt.show()
# +
# Trade using a simple strategy
def trade(data_, z_score_, window1, window2):
data_.reset_index(inplace=True,drop=True)
z_score_.reset_index(inplace=True,drop=True)
S1 = data_['gains_uvxy']
S2 = data_['gains_spy']
close = data_['Close_uvxy']
date = data_['Date']
# If window length is 0, algorithm doesn't make sense, so exit
if (window1 == 0) or (window2 == 0):
return 0
# Compute rolling mean and rolling standard deviation
# Simulate trading
# Start with no money and no positions
money = 0
countS1 = 0
countS2 = 0
for i in range(len(S1)):
# buy
if z_score_[i]<-7:
if countS1<0:
money -= close[i]*abs(countS1)
countS1 = 0
money -= close[i]
countS1 += 1
print('Buying volatility %s at %f %f %f %s %s'%(date[i],z_score_[i], close[i], money+countS1*close[i], countS1,countS2))
elif z_score_[i]>0:
if countS1>0:
money += close[i] * countS1
countS1 = 0
money += close[i]
countS1 -= 1
print('Selling volatility %s at %f %f %f %s %s'%(date[i], z_score_[i], close[i], money+countS1*close[i], countS1,countS2))
return money+countS1*close.iloc[-1]
trade(data.iloc[:200],z_score.iloc[:200], 5, 60)
# -
data[hist['Date'] == "2020-04-01"]
data[hist['Date'] == "2020-03-30"]
trade(data.iloc[200:],z_score[200:], 5, 60)
data.iloc[1]
z_score.reset_index(inplace=True,drop=True)
z_score
| uvxy_base_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp scraper
# -
# # Scraper
#
# > This module is used to scrape https://www.point2homes.com for all residential in Scottsdale Arizona
#hide
from nbdev.showdoc import *
from lxml.etree import HTML
from selenium import webdriver
from selenium.webdriver.common.by import By
import xmltodict as xd
from time import sleep
from lxml import etree
import pandas as pd
def get_text(xpath):
return re.sub('<[^<]+?>', '', etree.tostring(xpath, method='html', with_tail=False).decode('ascii')).replace('\n', '').replace(' ', '')
def extract_properties(land):
data = xd.parse(etree.tostring(land))['div']['div']
try:
address = data[0]['@data-address']
except:
print("Missing Address")
address = ''
try:
price = data[1]['@data-price']
try:
lot_size = data[2]['ul']['li'][0]['strong']
except:
# print("Missing lot_size")
lot_size = ''
try:
beds = data[2]['ul']['li'][0]['strong']
except:
# print("missing beds")
beds = ''
try:
baths = data[2]['ul']['li'][1]['strong']
except:
# print("missing baths")
baths = ''
try:
sqft = data[2]['ul']['li'][2]['strong']
except:
# print("missing sqft")
sqft = ''
try:
property_type = data[2]['ul']['li'][3]['#text']
except:
# print("Missing property_type")
property_type = ''
link = data[4]['a']['@href']
properties = {
'address': address,
'price':price,
'lot_size':lot_size,
'beds':beds,
'baths':baths,
'sqft':sqft,
'property_type':property_type,
'link':link
}
return properties
except Exception as e:
raise e
print(f'Failed on {address}')
def read_land_column(lands):
land_list = []
for land in lands:
land_list.append(extract_properties(land))
return land_list
url = "https://www.point2homes.com/CA/Real-Estate-Maps/BC/Vancouver-Island.html"
browser = webdriver.Firefox()
browser.get(url)
# +
page_source = HTML(browser.page_source)
lands = page_source.xpath("//div[@class='item_information']")
# -
land_data = []
# + jupyter={"outputs_hidden": true}
while(1):
# Read Page
page_source = HTML(browser.page_source)
# Extract Land Column
lands = page_source.xpath("//div[@class='item_information']")
# Append data
land_data = land_data + read_land_column(lands)
# Report in.
print(f"Successfully Scraped: {len(lands)}")
# Go to next page
next_button = browser.find_element(By.XPATH, "//a[starts-with(@class,'pager-next')]")
if 'disabled' in next_button.get_attribute('class'):
break
else:
print('Next Page')
next_button.click()
sleep(8)
# -
df = pd.DataFrame(land_data)
df
import re
def extract_numbers(s):
return float(''.join(re.findall("[-+]?\d*\.\d+|\d+", s)) or 0)
df['price_float'] = df['price'].apply(extract_numbers)
df['sqft_float'] = df['sqft'].apply(extract_numbers)
df['Dollar per Sqft'] = df['price_float'] / df['sqft_float']
df = df.sort_values('Dollar per Sqft')
df['link'] = 'https://www.point2homes.com' + df['link']
df['Dollar per Sqft'] = df['Dollar per Sqft'].apply(lambda x: str(round(x, 2)))
df = df.drop_duplicates('address')
df.to_csv('vanland_homes.csv', index=False)
df[df['baths'] >= 3]
df.to_csv("ScottsdaleAZ_raw.csv", index=False)
df.head(100).to_csv("ScottsdaleAZTop100.csv", index=False)
len(pages)extract_numbers
df = pd.DataFrame(land_list)
df.sort_values('price_float').to_csv('VancouverIsland-Cheapest.csv',index=False)
df.iloc[92].values
# +
next_page = browser.find_element(By.XPATH, "//input[starts-with(@name,'username')]")
password = browser.find_element(By.XPATH, "//input[starts-with(@name,'password')]")
username.send_keys("<EMAIL>")
password.send_keys("<PASSWORD>")
browser.find_element(By.XPATH, "//input[starts-with(@name,'Login')]").click()
# -
pager-next
# +
# price, ac, price/ac, datetime, id, area
# -
1038 / 100
item = {
'address':
'price':
'characteristics':
'labels':
'detail_link'
}
import pandas as pd
from lxml import etree
etree.tostring(tree)
etree.tostringlist(tree)
pd.read_html(tree)
tree = lands[0]
price = tree.xpath("//div[@class='price']")
price
e = price[0]
e.text
address = tree.xpath("//div[@class='item_address']")
address[0].attrib['data-address']
data-address
descriptions = tree.xpath("//div[@class='info member-info']")
links = tree.xpath("//a[@class='learn-more']")
# + jupyter={"outputs_hidden": true}
lands
# -
| 03_ScottsdaleAZ.ipynb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Preprocessing: translating MoMA files to Pandas
import os, re, glob
import numpy as np
import pandas as pd
from colicycle.MoMAobj import Momaobj
import colicycle.time_mat_operations as tmo
import colicycle.momaprocess as mp
folder_prefix = '../PreProcessed/'
# +
#say where data are located and how many channelss are present in each experiment
data_folder = [[folder_prefix+'20180709_GW296_glucose8aa37_1_MMStack/',2],
[folder_prefix+'20180711_GW296_glucose37_1_MMStack/',2],
[folder_prefix+'20180706_GW296_glycerol37_1_MMStack/',2],
[folder_prefix+'20170327_GW339_temp/',3]
]
tosave_folder = folder_prefix
# -
name = [os.path.basename(os.path.normpath(d[0])) for d in data_folder]
for ind, dataset in enumerate(data_folder):
mom = Momaobj(data_folder=dataset[0],col_nb=dataset[1], no_raw=True)
for i in range(len(mom.pos_list)):
mom.pos = mom.pos_list[i]
mom.gl = mom.gl_list[i]
time_mat_pd = mp.parse_exported(mom.get_momapath())
time_mat_pd['length'] = time_mat_pd.pixlim.apply(lambda x: x[:,1]-x[:,0])
time_mat_pd = tmo.essential_props(time_mat_pd)
#time_mat_pd = mp.moma_cleanup(time_mat_pd)
time_mat_pd = pd.concat([time_mat_pd, time_mat_pd.apply(lambda row: tmo.exponential_fit(row), axis=1)], axis=1)
time_mat_pd['pix_max']=time_mat_pd.pixlim.apply(lambda x: x[:,1])
time_mat_pd['pix_min']=time_mat_pd.pixlim.apply(lambda x: x[:,0])
time_mat_pd['full_cellcycle'] = time_mat_pd.exit_type.apply(lambda x: True if x=='DIVISION' else False)
time_mat_pd['full_cellcycle'] = time_mat_pd.exit_type.apply(lambda x: True if x=='DIVISION' else False)
time_mat_pd[time_mat_pd.born == -1]['full_cellcycle'] = False
time_mat_pd['mother_id'] = time_mat_pd['mother_id'].fillna(-1).astype(int)
filename = tosave_folder+'/'+name[ind]+'/step1/step1_time_mat'+'_pos'+mom.pos+'_GL'+mom.gl+'.pkl'
os.makedirs(os.path.dirname(filename), exist_ok=True)
time_mat_pd['Ti'] = np.nan
time_mat_pd['Li'] = np.nan
time_mat_pd['Li_fit'] = np.nan
time_mat_pd.to_pickle(filename)
| DataProcessing/step1_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Well, I have done it. It's layered cluster
# Unfortunately, it's not any faster....
from PIL import Image
import pickle
import vocabulary
import os
from pylab import *
from numpy import *
from scipy.cluster.vq import *
# -
import sift
import imagesearch3
imagesearch3 = reload(imagesearch3)
import mysparse
mysparse = reload(mysparse)
with open('caltech_imlist.pkl', 'rb') as f:
imlist = pickle.load(f)
featlist = pickle.load(f)
nbr_features = len(featlist)
descr = []
feature = sift.read_features_from_file(featlist[0])[1]
descr.append(feature[4:])
descriptors = descr[0] # stack all features for k-means
for i in arange(1, nbr_features):
feature = sift.read_features_from_file(featlist[i])[1]
descr.append(feature[4:])
descriptors = vstack((descriptors, descr[i]))
descriptors_short = descriptors[::100]
print descriptors_short.shape
cnt=0
word_list=[]
def divide_branch_with_center(data, branch, k, min_size, depth):
global cnt
# print depth, ':', len(branch)
div = min(k, len(branch))
if (div<=1) or (len(branch)<=min_size):
return []
centroids, distortion = kmeans(data[branch], k)
code, distance = vq(data[branch], centroids)
new_branch = []
for i in range(k):
ind = where(code==i)[0]
if len(ind)==0:
continue
else:
cnt += 1
word_list.append(centroids[i])
new_branch.append((centroids[i], distance[i], divide_branch_with_center(data, branch[ind], k, min_size, depth+1), cnt-1))
return new_branch
import sys
sys.setrecursionlimit(10000)
min_size = min(10, int(descriptors_short.shape[0]/100))
tree = array([i for i in range(descriptors_short.shape[0])])
cnt=0
word_list=[]
branches = ([0]*descriptors_short.shape[1], 0, divide_branch_with_center(descriptors, tree, 4, min_size, 0))
feature = sift.read_features_from_file(featlist[0])[1]
loc, desc = feature[:4], feature[4:]
print desc
def get_distance(l1, l2):
try:
return sqrt(sum([(i1-i2)**2 for i1, i2 in zip(l1, l2)]))
except:
print l1, l2
raise
def get_word(node, d):
if len(node[2])==0:
return node[3]
min_distance = get_distance(node[2][0][0], d)
next_node = 0
for i in arange(1, len(node)):
distance = get_distance(node[2][i][0], d)
if (distance<min_distance):
min_distance = distance
next_node = i
return get_word(node[2][next_node], d)
node = branches
print get_word(node, desc[0])
print word_list[927]
class vocabulary4:
def __init__(self, node, word_list, name):
self.node = node
self.word_list = word_list
self.name = name
def project(self, d):
nbr_words = len(self.word_list)
imhist = zeros((nbr_words))
for d0 in d:
w = get_word(self.node, d0)
imhist[w]+=1
return imhist
def get_word(self, node, d):
if len(node[2])==0:
return node[3]
min_distance = get_distance(node[2][0][0], d)
next_node = 0
for i in arange(1, len(node)):
distance = get_distance(node[2][i][0], d)
if (distance<min_distance):
min_distance = distance
next_node = i
return get_word(node[2][next_node], d)
voc = vocabulary4(branches, word_list, "clustered vocabulary")
h = voc.project(desc)
import imagesearch
imagesearch = reload(imagesearch)
os.remove('test4.db')
indx = imagesearch.Indexer('test4.db', voc)
indx.create_tables()
def average_color(imn):
im = array(Image.open(imn))
if size(im.shape)>2:
col = [int(average(im[:, :, c])+0.5) for c in [0, 1, 2]]
else:
col = [128, 128, 128]
return col
for i in range(nbr_images):
locs, descr = sift.read_features_from_file(featlist[i])
indx.add_to_index(imlist[i], descr)
indx.add_to_index_color(imlist[i], average_color(imlist[i]))
indx.db_commit()
from sqlite3 import dbapi2 as sqlite
con = sqlite.connect('test4.db')
print con.execute('select count (filename) from imlist').fetchone()
print con.execute('select * from imlist').fetchone()
print con.execute('select col0, col1, col2 from imcolor where imid=1000').fetchone()
imid = 1525
figure()
imshow(Image.open(imlist[imid-1]))
axis('off')
show()
nbr_results = 10
imagesearch = reload(imagesearch)
src = imagesearch.Searcher('test4.db', voc)
res = [w[1] for w in src.query(imlist[imid-1])[:nbr_results]]
imagesearch.plot_results(src, res, figsize=(16, 8))
| Chapter-7/CV Book Ch 7 Exercise 7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finding and Visualizing Time Series Motifs of All Lengths using the Matrix Profile
# ## Import Some Packages
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import cm
import ipywidgets as widgets
from ipywidgets import interact, Layout
import stumpy
plt.style.use('https://raw.githubusercontent.com/TDAmeritrade/stumpy/main/docs/stumpy.mplstyle')
# -
# ## EOG Example
#
# See [Figure 1](https://www.cs.ucr.edu/~eamonn/PAN_SKIMP%20%28Matrix%20Profile%20XX%29.pdf)
eog_df = pd.read_csv("https://zenodo.org/record/4733142/files/eog.csv?download=1")
m_250 = 250
m_500 = 500
mp_250 = stumpy.stump(eog_df["EOG"], m=m_250)
mp_500 = stumpy.stump(eog_df["EOG"], m=m_500)
motif_idx_250 = np.argmin(mp_250[:, 0])
motif_idx_500 = np.argmin(mp_500[:, 0])
nn_idx_250 = mp_250[motif_idx_250, 1]
nn_idx_500 = mp_500[motif_idx_500, 1]
fig, axs = plt.subplots(3)
axs[0].plot(eog_df["EOG"].values)
axs[1].plot(np.arange(m_250), eog_df.iloc[motif_idx_250 : motif_idx_250 + m_250])
axs[1].plot(np.arange(m_250), eog_df.iloc[nn_idx_250 : nn_idx_250 + m_250])
axs[2].plot(np.arange(m_500), eog_df.iloc[motif_idx_500 : motif_idx_500 + m_500])
axs[2].plot(np.arange(m_500), eog_df.iloc[nn_idx_500 : nn_idx_500 + m_500])
plt.show()
# ## Compute the Pan Matrix Profile using STIMP
#
# Essentially, `stumpy.stimp` implements [Table 2](https://www.cs.ucr.edu/~eamonn/PAN_SKIMP%20%28Matrix%20Profile%20XX%29.pdf).
min_m, max_m = 100, 1000
eog = stumpy.stimp(eog_df["EOG"].values, min_m=min_m, max_m=max_m, percentage=0.01) # This percentage controls the extent of `stumpy.scrump` completion
percent_m = 0.01 # The percentage of windows to compute
n = np.ceil((max_m - min_m) * percent_m).astype(int)
for _ in range(n):
eog.update()
# Above, we select a range of window sizes from `min_m = 3` to `max_m = 1000` and we arrange the windows in this range according to a breadth first order (this is done automatically by `stumpy.stimp` and see [slide 32](https://drive.google.com/file/d/1eT9oHOAKoi4oGkUX26V9aZIopov0Pxt5/view)) and/or [Section C](https://www.cs.ucr.edu/~eamonn/PAN_SKIMP%20%28Matrix%20Profile%20XX%29.pdf). The order of the window sizes to be processed can be found in `.M_` attribute:
eog.M_[:n]
# Notice that we don't compute the matrix profiles (approximated using `stumpy.scrump` at 1% and with `pre-scrump` turned on) for all of the window sizes and, instead, we select only 1% of the window sizes (`percent_m = 0.01`) for this task. So, only a total of `n` matrix profiles were computed. Now, let's plot our pan matrix profile along with the locations of our motif pairs (vertical red lines):
# +
fig = plt.figure()
fig.canvas.toolbar_visible = False
fig.canvas.header_visible = False
fig.canvas.footer_visible = False
lines = [motif_idx_250, motif_idx_500, nn_idx_250, nn_idx_500]
color_map = cm.get_cmap("Greys_r", 256)
im = plt.imshow(eog.PAN_, cmap=color_map, origin="lower", interpolation="none", aspect="auto")
plt.xlabel("Time", fontsize="20")
plt.ylabel("m", fontsize="20")
plt.clim(0.0, 1.0)
plt.colorbar()
plt.tight_layout()
# Draw some vertical lines where each motif and nearest neighbor are located
if lines is not None:
for line in lines:
plt.axvline(x=line, color='red')
plt.show()
# -
# Now, we'll compute 2% more matrix profiles (for a total of 1% + 2% = 3%) for additional windows:
for _ in range(2 * n):
eog.update()
# +
fig = plt.figure()
fig.canvas.toolbar_visible = False
fig.canvas.header_visible = False
fig.canvas.footer_visible = False
lines = [motif_idx_250, motif_idx_500, nn_idx_250, nn_idx_500]
color_map = cm.get_cmap("Greys_r", 256)
im = plt.imshow(eog.PAN_, cmap=color_map, origin="lower", interpolation="none", aspect="auto")
plt.xlabel("Time", fontsize="20")
plt.ylabel("m", fontsize="20")
plt.clim(0.0, 1.0)
plt.colorbar()
plt.tight_layout()
# Draw some vertical lines where each motif and nearest neighbor are located
if lines is not None:
for line in lines:
plt.axvline(x=line, color='red')
plt.show()
# -
# Notice how the pan matrix profile has become a bit clearer and a less "blocky"?
# ## Steamgen Example
steam_df = pd.read_csv("https://zenodo.org/record/4273921/files/STUMPY_Basics_steamgen.csv?download=1")
m = 640
mp = stumpy.stump(steam_df["steam flow"], m=m)
motif_idx = np.argmin(mp[:, 0])
nn_idx = mp[motif_idx, 1]
fig, axs = plt.subplots(2)
axs[0].plot(steam_df["steam flow"].values)
axs[1].plot(np.arange(m), steam_df["steam flow"].iloc[motif_idx : motif_idx + m])
axs[1].plot(np.arange(m), steam_df["steam flow"].iloc[nn_idx : nn_idx + m])
plt.show()
# ### Compute the Pan Matrix Profile using STIMP
min_m, max_m = 100, 3000
steam = stumpy.stimp(steam_df['steam flow'], min_m=min_m, max_m=max_m, percentage=0.01) # This percentage controls the extent of `stumpy.scrump` completion
percent_m = 0.01 # The percentage of windows to compute
n = np.ceil((max_m - min_m) * percent_m).astype(int)
for _ in range(n):
steam.update()
# +
fig = plt.figure()
fig.canvas.toolbar_visible = False
fig.canvas.header_visible = False
fig.canvas.footer_visible = False
lines = [motif_idx, nn_idx]
color_map = cm.get_cmap("Greys_r", 256)
im = plt.imshow(steam.PAN_, cmap=color_map, origin="lower", interpolation="none", aspect="auto")
plt.xlabel("Time", fontsize="20")
plt.ylabel("m", fontsize="20")
plt.clim(0.0, 1.0)
plt.colorbar()
plt.tight_layout()
if lines is not None:
for line in lines:
plt.axvline(x=line, color='red')
plt.show()
# -
for _ in range(2 * n):
steam.update()
# + tags=[]
fig = plt.figure()
fig.canvas.toolbar_visible = False
fig.canvas.header_visible = False
fig.canvas.footer_visible = False
lines = [motif_idx, nn_idx]
color_map = cm.get_cmap("Greys_r", 256)
im = plt.imshow(steam.PAN_, cmap=color_map, origin="lower", interpolation="none", aspect="auto")
plt.xlabel("Time", fontsize="20")
plt.ylabel("m", fontsize="20")
plt.clim(0.0, 1.0)
plt.colorbar()
plt.tight_layout()
if lines is not None:
for line in lines:
plt.axvline(x=line, color='red')
plt.show()
# -
# ## Bonus Section
# +
# %matplotlib widget
plt.style.use('https://raw.githubusercontent.com/TDAmeritrade/stumpy/main/docs/stumpy.mplstyle')
plt.ioff()
fig = plt.figure()
fig.canvas.toolbar_visible = False
fig.canvas.header_visible = False
plt.ion()
ax = plt.gca()
ax.format_coord = lambda x, y: f'Time = {x:.0f}, m = {y:.0f}'
lines = [motif_idx, nn_idx]
color_map = cm.get_cmap("Greys_r", 256)
im = plt.imshow(steam.PAN_, cmap=color_map, origin="lower", interpolation="none", aspect="auto")
plt.xlabel("Time", fontsize="20")
plt.ylabel("m", fontsize="20")
plt.clim(0.0, 1.0)
plt.colorbar()
plt.tight_layout()
if lines is not None:
for line in lines:
plt.axvline(x=line, color='red')
def update_slider(change):
PAN = steam.pan(threshold=change['new'])
im.set_data(PAN)
fig.canvas.draw_idle()
threshold = 0.2
slider = widgets.FloatSlider(value=threshold, min=0.0, max=1.0, step=0.01, readout_format='.2f', layout=Layout(width='80%'), description='Threshold:')
slider.observe(update_slider, names='value')
widgets.VBox([fig.canvas, slider])
# -
| docs/Tutorial_Pan_Matrix_Profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# **Step 1: Loading scikit learn version of Diabetes data set**
from sklearn.datasets import load_diabetes
diabetes = load_diabetes()
print(diabetes['DESCR'])
# **Step 2: Splitting the data into train set and test set**
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(diabetes.data,diabetes.target, random_state = 2810)
# **Step 3: Calculating train and test R-squared**
from sklearn.linear_model import Lasso
lasso = Lasso()
lasso.fit(X_train,y_train)
print(lasso.score(X_train,y_train))
print(lasso.score(X_test,y_test))
print(lasso.coef_)
print(diabetes.feature_names)
# **There are only two features and their names are 'bmi' and 's5' respectively**
# **Step 4: Loading original diabetes dataset**
unscaled_diabetes = np.genfromtxt('/home/smith/Downloads/diabetes.data.txt', delimiter = '\t', skip_header=1)
# **Step 5: Splitting the data into train set and test set**
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(unscaled_diabetes[:,0:10],unscaled_diabetes[:,10], random_state = 2810)
# **Step 6: Calculating train and test R-squared on the original dataset**
lasso = Lasso()
lasso.fit(X_train,y_train)
print(lasso.score(X_train,y_train))
print(lasso.score(X_test,y_test))
print(lasso.coef_)
# **Using unscaled version only made one feature coefficient 0, namely 's4', whilst using the previous one had only 2 non-zero coefficient. Using Unscaled version has improved training R-squared over the previous one**
# **Step 7: Preprocessing the data whilst avoiding Data Snooping**
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# **Step 8: Repeating step 3 for the current training and test data**
from sklearn.linear_model import Lasso
lasso = Lasso()
lasso.fit(X_train_scaled,y_train)
print(lasso.score(X_train_scaled,y_train))
print(lasso.score(X_test_scaled,y_test))
print(lasso.coef_)
# **using scaled version causes all coefficient to be non-zero, which is different from previous 2 cases. The train and test R-squared has shown a minor improvement over unscaled test set.**
# **The scaled version's results are closer to unscaled version rather than the scikit learn version due to the presence of data snooping in the scikit learn dataset.**
# **Step 9: Plot the test R-squared vs the number of features used**
alphas = [0.1,0.3,0.5,0.8,1,2,3,5,10,15,20,25,30,35,40,45,50]
number_coeff = np.zeros(len(alphas))
r_test = np.zeros(len(alphas))
for i in range(len(alphas)):
lasso1 = Lasso(alpha = alphas[i])
lasso1.fit(X_train_scaled,y_train)
r_test[i] = lasso1.score(X_test_scaled,y_test)
number_coeff[i]= np.sum(lasso1.coef_!=0)
plt.plot(number_coeff,r_test)
plt.xlabel('Number of coefficients')
plt.ylabel('Test R-squared')
plt.title('Test R-squared vs Number of features')
plt.grid()
print(number_coeff)
# **I will choose the point at which number of coefficient is less and the test R-squared is high, which is at number_coeff =4. Doing this will also provide good interpretability**
# **Step 10: Choose the regularization parameter for the Lasso using cross-validation on the training set.**
from sklearn.model_selection import cross_val_score
best_score = 0
for i in [0.1,0.3,0.5,0.8,1,1.5,2,2.53,5,10,15,20,25,30,35,40,45,50]:
lasso2 = Lasso(alpha = i)
scores = cross_val_score(lasso2, X_train_scaled , y_train, cv=5)
score = np.mean(scores)
if score > best_score:
best_score = score
best_alpha = i
lasso3 = Lasso(alpha = best_alpha)
lasso3.fit(X_train_scaled,y_train)
print(lasso3.score(X_train_scaled,y_train))
print(lasso3.score(X_test_scaled,y_test))
print(best_alpha)
print(best_score)
print(lasso3.coef_)
# **Step 11: Implement an inductive conformal predictor**
# **(a) Split the training set**
X_proper,X_cal,y_proper,y_cal = train_test_split(X_train,y_train,test_size = 99, random_state = 2810)
# **(b) Preprocess the training set proper, calibration set, and test set**
scaler1 = StandardScaler()
scaler1.fit(X_proper)
X_proper_scaled = scaler1.transform(X_proper)
X_cal_scaled = scaler1.transform(X_cal)
X_tes_scaled = scaler1.transform(X_test)
# **(c) Using the nonconformity measure α = |y − yˆ|**
lasso_c = Lasso(alpha = best_alpha)
lasso_c.fit(X_proper_scaled,y_proper)
y_pred_cal = lasso_c.predict(X_cal_scaled)
non_conf = abs(y_cal - y_pred_cal)
sorted_ncs = np.sort(non_conf)
#calculating for significance level 5%
e = 0.05
aug_len = len(y_cal)+1
k = np.ceil((1-e)*aug_len)
c1 = sorted_ncs[int(k)-1]
#calculating for significance level 20%
e = 0.2
aug_len = len(y_cal)+1
k = np.ceil((1-e)*aug_len)
c2 = sorted_ncs[int(k)-1]
[c1, c2]
y_pred_test = lasso_c.predict(X_tes_scaled)
pred_set_c1 = np.array([y_pred_test - c1, y_pred_test+c1]).T
pred_set_c2 = np.array([y_pred_test - c2, y_pred_test+c2]).T
#test error rate for 20% significance level
print(np.mean((y_test >= pred_set_c2[:,0])&(pred_set_c2[:,1]>=y_test)))
#test error rate for 5% significance level
print(np.mean((y_test >= pred_set_c1[:,0])&(pred_set_c1[:,1]>=y_test)))
print(c1-c2)
#Prediction interval
# **Step 12:Results**
# (a) The training R-squared is 0.3784148518466054
#
# The test R-squared is 0.32473224605708073
#
# Number of Featured used = 2
# (b) The training R-squared is 0.5465817497702268
#
# The test R-squared is 0.34893320451969656
#
# Number of Featured used = 9
# (c) The training R-squared is 0.5520822070008307
#
# The test R-squared is 0.38035967571695484
#
# Number of Featured used = 10
# (d) The training R-squared is 0.5473130230215644
#
# The test R-squared is 0.3878215277221756
#
# Number of Featured used = 7
#
# Best chosen Alpha = 2
#
# (e) Length of prediction intervals = 34.594119089867135
#
# Test- error rate for 5% significance level = 0.9009009009009009
#
# Test- error rate for 20% significance level = 0.7027027027027027
#
# **1) Interesting observation about Lasso: In our dataset, the number of coefficients made zero by Lasso is highly dependent on the random state of the split. Lasso seems to be very sensitive to Data Snooping**
# **2) Tie-breaking: We could add a tie-breaking random variable in our calculations to take ties into the account.**
# **3) Validity of Conformal predictors can be tested by using Cross-validation**
| Inductive Conformal Predictors and Lasso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:hackathon]
# language: python
# name: conda-env-hackathon-py
# ---
import os
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib import rcParams
from scipy.signal import convolve2d
from scipy.ndimage import gaussian_filter, convolve1d
from skimage.io import imread, imsave
from skimage.filters import threshold_otsu
from skimage.segmentation import clear_border
from skimage.measure import label, regionprops
from skimage.morphology import closing, square, erosion, dilation
from skimage.color import label2rgb
from skimage import img_as_uint
from skimage.feature import blob_dog
from mpl_toolkits.mplot3d import Axes3D
import glob
# ## We will be using image processing techniques and the skimage package to determine whether or not an ionic liquid - solvent mixture is phase separated
# ## The first step will be to convolve the image. This process smooths out some of the shadowing from the VMD-rendered image
def _convolveImage(image, kernel):
def scaleIt(cvld):
cvld[cvld > 255.0] = 255.0
cvld[cvld < 0.0] = 0.0
return cvld
convolved = np.ones(image.shape)
for i in range(convolved.shape[-1]):
cvld = convolve2d(image[:,:,i], kernel, boundary='fill', mode='same',
fillvalue=0.)
convolved[:,:,i] = scaleIt(cvld)
#plt.imshow(convolved.astype(int))
return convolved.astype(int)
# ## Next we will threshold a gray-scale image via Otsu's method
def _apply_otsu(gray):
thresh_otsu = threshold_otsu(gray)
im_bw = gray < thresh_otsu
return im_bw
# ### In some image processing tutorial, the borders are cleared so that objects near the border aren't counted. I am skipping this step so that in theory I can still count these objects.
# ### To clean up the image, I am going to play around with a cutoff to delete objects that may be either noise or a single atom, which we don't want to count towards total objects. A filtered image as well as the image properties will be passed into `_cutoff_particles`
def _cutoff_particles(image, image_props, cutoff=300):
im_bw_filt = image > 1
# Loop through image properties and delete small objects
n_regions = 0
for prop in im_props:
if prop.area < cutoff:
im_bw_filt[image==prop.label] == False
else:
n_regions += 1
print('Number of individual regions = {}'.format(n_regions))
return n_regions
# ### Now we will loop through our images and see how our image processing workflow performs
"""hetero_list = list()
sigma = 8
unsharp_strength = 0.8
kernel_size = 10
kernel = np.ones((kernel_size, kernel_size)) / kernel_size
kernel[0,:]
for filepath in glob.iglob('/Users/raymatsumoto/science/keras-phase-sep/data/train-images/hetero/*.png'):
image = imread(filepath)
blurred = gaussian_filter(image, sigma=0.8)
convolved = _convolveImage(image - unsharp_strength * blurred, kernel)
gray = convolved[:,:,0]
im_bw = _apply_otsu(gray)
im_labeled, n_labels = label(im_bw, background=0, return_num=True)
im_labeled += 1
im_props = regionprops(im_labeled)
n_regions = _cutoff_particles(im_labeled, im_props, cutoff=150)
hetero_list.append(n_regions)"""
"""homo_list = list()
sigma = 8
unsharp_strength = 0.8
kernel_size = 10
kernel = np.ones((kernel_size, kernel_size)) / kernel_size
kernel[0,:]
for filepath in glob.iglob('/Users/raymatsumoto/science/keras-phase-sep/data-otsu/train/homo/*.png'):
print(filepath)
image = imread(filepath)
im_labeled, n_labels = label(image, background=0, return_num=True)
im_labeled += 1
im_props = regionprops(im_labeled)
n_regions = _cutoff_particles(im_labeled, im_props, cutoff=150)
homo_list.append(n_regions)"""
# ## Experiment with k-means clustering for colors
from sklearn.cluster import KMeans
from sklearn.metrics import pairwise_distances_argmin
from sklearn.utils import shuffle
# +
n_colors = 3
image = np.array(image, dtype=np.float64) / 255
w, h, d = original_shape = tuple(image.shape)
assert d == 3
image_array = np.reshape(image_array, (w * h, d))
image_array_sample = shuffle(image_array, random_state=0)[:1000]
kmeans = KMeans(n_clusters=n_colors, random_state=0).fit(image_array_sample)
# -
labels=kmeans.predict(image_array)
img = 'data/train/homo/0f19f16fc5cbd07025c80a51a30683e6-1.png'
image = cv2.imread(img)
len(image)
# +
import cv2
from sklearn.cluster import KMeans
from skimage.transform import resize
class DominantColors:
CLUSTERS = None
IMAGE = None
COLORS = None
LABELS = None
def __init__(self, image, clusters, filename):
self.CLUSTERS = clusters
self.IMAGE = image
self.FILE = filename
def dominantColors(self):
#read image
img = cv2.imread(self.IMAGE)
img = cv2.resize(img, dsize=(50, 50), interpolation=cv2.INTER_CUBIC)
#convert to rgb from bgr
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
#reshaping to a list of pixels
img = img.reshape((img.shape[0] * img.shape[1], 3))
#save image after operations
self.IMAGE = img
#using k-means to cluster pixels
kmeans = KMeans(n_clusters = self.CLUSTERS)
kmeans.fit(img)
#the cluster centers are our dominant colors.
self.COLORS = kmeans.cluster_centers_
#save labels
self.LABELS = kmeans.labels_
#returning after converting to integer from float
return self.COLORS.astype(int)
def rgb_to_hex(self, rgb):
return '#%02x%02x%02x' % (int(rgb[0]), int(rgb[1]), int(rgb[2]))
def plotClusters(self):
#plotting
fig = plt.figure(figsize=(10,8))
ax = Axes3D(fig)
for label, pix in zip(self.LABELS, self.IMAGE):
ax.scatter(pix[0], pix[1], pix[2], color = self.rgb_to_hex(self.COLORS[label]))
ax.set_xlabel('Red', fontsize=18, labelpad=13)
ax.set_ylabel('Green', fontsize=18, labelpad=13)
ax.set_zlabel('Blue', fontsize=18, labelpad=16)
ax.tick_params(axis = 'both', which = 'major', labelsize = 18)
plt.tight_layout()
plt.savefig(self.FILE)
plt.show()
img = 'data/train/homo/0f19f16fc5cbd07025c80a51a30683e6-1.png'
#img = 'data/train/homo/007364fd56b31f36321ad4c0e64281bb-1.png'
clusters = 1
dc = DominantColors(img, clusters, 'red.pdf')
colors = dc.dominantColors()
dc.plotClusters()
print(colors)
# -
img = 'data/train/homo/007364fd56b31f36321ad4c0e64281bb-1.png'
clusters = 1
dc = DominantColors(img, clusters, 'blue.pdf')
colors = dc.dominantColors()
dc.plotClusters()
print(colors)
list(colors[0]).index(max(colors[0]))
image = imread('data/train-images/homo/0f19f16fc5cbd07025c80a51a30683e6-1.png')
#image = imread('data/train-images/homo/24ba6ce31eb7b9c0191f31cbd0c8c942-5.tga')
plt.imshow(image)
sigma = 8
unsharp_strength = 0.8
kernel_size = 10
kernel = np.ones((kernel_size, kernel_size)) / kernel_size
blurred = gaussian_filter(image, sigma=0.8)
convolved = _convolveImage(image - unsharp_strength * blurred, kernel)
plt.imshow(convolved)
gray = convolved[:,:,0]
im_bw = _apply_otsu(gray)
(im_bw == True).any()
plt.imshow(im_bw)
im_bw[:,:90] = 0
im_bw[:,-90:] = 0
im_bw[:90,:] = 0
im_bw[-90:,:] = 0
test_clear = clear_border(im_bw)
imsave('test.png', img_as_uint(test_clear))
plt.imshow(test_clear)
image = imread('data-otsu/train/homo/156ab47c6ef918c17616d304b2b588b9-1homo-0.png')
plt.imshow(image)
np.max([region.area for region in regionprops(label(test_clear))])
def label_regions(image):
label_image = label(image)
image_label_overlay = label2rgb(label_image, image=image)
fig, ax = plt.subplots(figsize=(10, 6))
ax.imshow(image_label_overlay)
for region in regionprops(label_image):
# take regions with large enough areas
if region.area >= 100:
# draw rectangle around segmented coins
minr, minc, maxr, maxc = region.bbox
rect = mpatches.Rectangle((minc, minr), maxc - minc, maxr - minr,
fill=False, edgecolor='red', linewidth=2)
ax.add_patch(rect)
ax.set_axis_off()
plt.tight_layout()
plt.show()
return regionprops(label_image)
# +
sigma = 8
unsharp_strength = 0.8
kernel_size = 9
kernel = np.ones((kernel_size, kernel_size)) / kernel_size
kernel[0,:]
image = imread('/Users/raymatsumoto/science/keras-phase-sep/data/test-images/homo/5e541ed97dbbd7387dffd5fc29eccd63-3.png')
blurred = gaussian_filter(image, sigma=0.8)
convolved = _convolveImage(image - unsharp_strength * blurred, kernel)
close = closing(convolved, square(3))
gray = convolved[:,:,0]
im_bw = _apply_otsu(gray)
im_labeled, n_labels = label(im_bw, background=0, return_num=True)
im_labeled += 1
im_props = regionprops(im_labeled)
plt.imshow(im_labeled,cmap=plt.cm.gray)
n_regions = _cutoff_particles(im_labeled, im_props, cutoff=50)
# -
fig, ax = plt.subplots(figsize=(10,10))
ax.imshow(convolved, cmap=plt.cm.gray)
| examples/image_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot random init
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as sts
import glob
paths_gpomdp = glob.glob("data/random-init-nn/gpomdp/progress*.csv")
paths_remps = glob.glob("data/random-init-nn/remps/progress*.csv")
paths
# -
dfs = [pd.read_csv(p) for p in paths_gpomdp]
om = np.array([df.Omega[0] for df in dfs])
ret = np.array([np.max(df.ReturnsMean) for df in dfs])
om
dfs_remps = [pd.read_csv(p) for p in paths_remps]
om_remps = np.array([df.Omega[0] for df in dfs_remps])
ret_remps = np.array([np.max(df.ReturnsMean) for df in dfs_remps])
om_remps
# +
ind = np.argsort(om)
ind_remps = np.argsort(om_remps)
plt.plot(om[ind], ret[ind], om_remps[ind_remps], ret_remps[ind_remps])
plt.savefig("random-init.png")
# +
file_name="random-init.csv"
to_write = np.concatenate((np.reshape(om, (-1,1)), np.reshape(om_remps, (-1,1)), np.reshape(ret,(-1,1)), np.reshape(ret_remps,(-1,1))), axis=1)
np.savetxt(file_name, to_write, delimiter=',', header="Omega_gpomdp, Omega_remps, Return_gpomdp, Return_remps", comments='')
| thesis_presentation/plots/cartpole/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 平均の種類
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import numpy as np
import math as m
import warnings
warnings.filterwarnings('ignore')
# +
fig =plt.figure(figsize=(6,6))
N = 101 # 公差を5刻みにするように番兵をつける
# グラフ領域
corr = 1.0 # 原点0.0から始めるための補正値
g_x = np.linspace(-1.0, 1.0, N)
g_y = np.linspace(-1.0, 1.0, N)
# 円
r_x = np.cos(g_x * np.pi)
r_y = np.sin(g_y * np.pi)
plt.plot(r_x + corr, r_y + corr, color="black")
print(f"C: [({np.min(r_x)}, {np.min(r_y)}), ({np.max(r_x)}, {np.max(r_y)})]")
# 線a
def a(x):
return x * 0
a_min = np.min(r_x)
a_max = a_min + 0.75 * 2 # length
a_d = abs(a_max - a_min)
a_x = np.linspace(a_min, a_max, N)
a_y = a(a_x)
print(f"A: [({np.min(a_x)}, {np.min(a_y)}), ({np.max(a_x)}, {np.max(a_y)})]")
plt.plot(a_x + corr, a_y + corr, color="red")
# 線b
def b(x):
return x * 0
b_min, b_max = a_max, np.max(r_x)
b_d = abs(b_max - b_min)
b_x = np.linspace(b_min, b_max, N)
b_y = b(b_x)
print(f"B: [({np.min(b_x)}, {np.min(b_y)}), ({np.max(b_x)}, {np.max(b_y)})]")
plt.plot(b_x + corr, b_y + corr, color="blue")
# 相加平均/算術平均 Arithmetic Mean
radius = (a_d + b_d) / 2 # 相加平均は半径
am_x = radius - corr
am_y = m.sqrt(1 - am_x**2)
print(f"AM: ({am_x}, {am_y})")
plt.vlines(x=am_x + corr, ymin=0 + corr, ymax=am_y + corr, color="green")
# 相乗平均/幾何平均 Geometric Mean
gm_x = a_max
gm_y = m.sqrt(1 - gm_x**2)
print(f"GM: ({gm_x}, {gm_y})")
plt.vlines(x=gm_x+ corr, ymin=0 + corr, ymax=gm_y + corr, color="purple")
# 調和平均 Harmonic Mean
hm_x = np.linspace(am_x, gm_x, N)
hm_y = np.linspace(0, gm_y, N)
plt.plot(hm_x + corr, hm_y + corr, color="orange")
# 描画
plt.grid()
plt.show()
| math/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # AssocPlace
# # Set up Analysis
# ### Initial Imports
import sys; sys.prefix
# +
import pandas as pd
import moss
from scipy import stats
import scipy as sp
import seaborn as sns
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import os.path as op
# Gather project info & functions
from ap_setup_project import *
# for plotting
sns.set(style='ticks', context='poster', font_scale=1.3)
# %matplotlib inline
# R for stats
# # %load_ext rpy2.ipython
# # %R require(lme4)
# # %R require(lmerTest)
# -
sns.__path__
# ### Gather experiment info
dirs = dict()
dirs['basedir'] = op.join(op.expanduser('~'), 'Experiments/AssocPlace')
dirs['datadir'] = op.join(dirs['basedir'], 'data')
dirs['analydir'] = op.join(dirs['basedir'], 'analysis')
dirs['subj_info_file'] = op.join(dirs['datadir'], 'group_info.csv')
exp = gather_experiment_info(exp_name='AP', dirs=dirs)
subj_info = pd.read_csv(dirs['subj_info_file'])
subj_info.head()
# ### Subj info
subj_info = subj_info[pd.isnull(subj_info.remove)]
subj_info
# ### Set up filepaths & load in data
# +
ds = pd.DataFrame() # study
dt = pd.DataFrame() # test
study = True
test = True
questionnaires_shock = True
questionnaires_post = True
for subid in subj_info.subid:
print subid
if study:
# add study file
study_file = op.join(dirs['datadir'], subid, subid + '_behav_study.csv')
d = pd.read_csv(study_file)
d['subid'] = subid
ds = ds.append(d, ignore_index=True)
if test:
# add test file
test_file = op.join(dirs['datadir'], subid, subid + '_behav_freeresp.csv')
d2 = pd.read_csv(test_file)
# print d2.head()
d2['subid'] = subid
dt = dt.append(d2, ignore_index=True)
# Compiled group data
if questionnaires_shock:
q_file = op.join(dirs['basedir'], 'data/Quest/Questionnaires_shockblock_group.csv')
dq_shock = pd.read_csv(q_file, index_col=0)
if questionnaires_post:
q_file = op.join(dirs['basedir'], 'data/Quest/Questionnaires_group.csv')
dq_post = pd.read_csv(q_file, index_col=0)
# -
# #### Remove runs for some subjects
dt.shape
subj106_run1 = (dt.subid == 'ap106') & (dt.run == 1) # this subj said they likely reversed responses for run 1, so remove
dt = dt.loc[np.invert(subj106_run1),:]
11789 - 11747
dt.shape
# #### Remove some trials for beach repeats (48 trials total, 2 for 24 subs)
data = dt.loc[dt.associate == 'beach'].groupby(['subid']).count().reset_index()
beach_counts = dt.loc[dt.associate == 'beach'].groupby(['subid']).count().reset_index()
sub_list = beach_counts[beach_counts.trial == 2].subid
trials_remove = (dt.subid.isin(sub_list)) & (dt.associate == 'beach')
dt = dt.loc[np.invert(trials_remove),:]
dt.shape
11747 - 11699
# #### Remove a couple trials from ap151 from when talking to him re: squeezeball
# Note that two preceding trials are removed for shock and shock + 1
trials_remove = (dt.subid == 'ap151') & (dt.target.isin(['OLIVE', 'CRAB']))
dt = dt.loc[np.invert(trials_remove),:]
dt.shape
# #### Remove beach from ds too.
ds.head()
print ds.shape
sub_list = beach_counts[beach_counts.trial == 2].subid
trials_remove = (ds.subid.isin(sub_list)) & (ds.pic == 'beach')
ds = ds.loc[np.invert(trials_remove),:]
print ds.shape
23646 - 23502
# ### Assign subid to group (pilot, control, stress) and gender (male, female)
ds = ds.merge(subj_info, on='subid', how='outer')
dt = dt.merge(subj_info, on='subid', how='outer')
# ### Number subjects per group
ds.groupby(['subid', 'group']).mean().reset_index().groupby('group').count().subid
dt.groupby(['subid', 'group']).mean().reset_index().groupby('group').count().subid
# #### Male counts
dt[dt.gender=='male'].groupby(['subid', 'group']).mean().reset_index().groupby('group').count().subid
# # Preprocessing
# ## Study
ds.head()
# ##### Deal with no responses (NR): Set ISI resp as resp, if ISI was within a second after stim-offset
sns.distplot(ds.respRT[ds.resp != 'NR'])
sns.distplot(ds.ISIrespRT[ds.resp == 'NR'])
ds.respRT[(ds.resp == 'NR') & (ds.ISIrespRT < 4)] = ds.ISIrespRT[(ds.resp == 'NR') & (ds.ISIrespRT < 4)]
ds.resp[(ds.resp == 'NR') & (ds.ISIrespRT < 4)] = ds.ISIresp[(ds.resp == 'NR') & (ds.ISIrespRT < 4)]
sns.distplot(ds.respRT[ds.resp != 'NR'])
# +
ds = ds.replace(to_replace='WI', value='indoor')
ds = ds.replace(to_replace='WO', value='outdoor')
ds = ds.replace(to_replace='UR', value='unrelated')
ds = ds.replace(to_replace='R', value='related')
ds = ds.replace(to_replace='NR', value='no response')
ds.head()
# -
# ## Test
dt.head()
# #### Distribution of RTs during ITI
sns.distplot(dt.ISIrespRT[(dt.ISIresp != 'NR')] + 4)
# +
# drop this column so join function doesn't get confused
dt = dt.drop('index',1)
# Split cond to condition and reps
dt['cond_orig'] = dt['cond']
dt = dt.drop('cond', 1)
dt.cond_orig[dt.cond_orig == 'F'] = 'F_0'
dt = dt.join(pd.DataFrame(dt.cond_orig.str.split('_').tolist(), columns= ['cond', 'reps']))
#Replace no responses with ISI responses, update RT (change resp last), but only if ISI within a sec after offset
dt.respRT[(dt.resp == 'NR') & (dt.ISIrespRT < 1)] = dt.ISIrespRT[(dt.resp == 'NR') & (dt.ISIrespRT < 1)] + 4
dt.acc[(dt.resp == 'NR') & (dt.ISIrespRT < 1)] = dt.ISIacc[(dt.resp == 'NR') & (dt.ISIrespRT < 1)]
dt.accSpec[(dt.resp == 'NR') & (dt.ISIrespRT < 1)] = dt.ISIaccSpec[(dt.resp == 'NR') & (dt.ISIrespRT < 1)]
dt.resp[(dt.resp == 'NR') & (dt.ISIrespRT < 1)] = dt.ISIresp[(dt.resp == 'NR') & (dt.ISIrespRT < 1)]
drop_cols = ['ISIrespRT', 'ISIresp', 'ISIacc', 'ISIaccSpec']
for drop_col in drop_cols:
dt = dt.drop(drop_col,1)
# Remove shock and post shock trials
shockTrials = pd.Series(dt.shockTrial)
lagged = shockTrials.shift(1) # shift forward one
combined = lagged + shockTrials
dt['shock_and_post'] = combined # merged; shock and post shock = 1
dt.shock_and_post[0] = dt.shockTrial[0] # first trial = first trial since no prev
dt.ix[dt.group == 'control-fmri', 'shock_and_post'] = 0 # set controls to 0
dt.ix[dt.shockCond == 'safe', 'shock_and_post'] = 0 # set safe cond to 0
dt = dt.query('shock_and_post < 1') #remove trials
print set(dt.shockTrial) # confirm that it worked; shockTrial = 0 only (1s for controls/safe)
dt = dt.drop('shockTrial',1)
dt = dt.drop('shock_and_post',1)
# reset index post removing shock trials
dt = dt.reset_index()
dt = dt.drop('index',1)
dt.head()
# -
sns.distplot(dt.respRT[(dt.resp != 'NR')])
dt = dt.replace(to_replace='TI', value='indoor')
dt = dt.replace(to_replace='TO', value='outdoor')
dt = dt.replace(to_replace='F', value='foil')
dt = dt.replace(to_replace='NR', value='no response')
# +
# Convert accSpec to acc, split to col for Confidence
dt['accSpec_tosplit'] = dt['accSpec']
dt.accSpec_tosplit[dt.accSpec_tosplit == 'CR'] = 'CR_N'
dt.accSpec_tosplit[dt.accSpec_tosplit == 'MI'] = 'MI_N'
dt.accSpec_tosplit[dt.accSpec_tosplit == 'no response'] = 'no response_N'
dt = dt.join(pd.DataFrame(dt.accSpec_tosplit.str.split('_').tolist(),
columns= ['accSpec2', 'conf']))
# clear out the unneeded cols
dt = dt.drop('accSpec2',1)
dt = dt.drop('accSpec_tosplit',1)
dt.head()
# -
dt.group2 = dt.group
dt.group2[dt.group2 == 'control'] = 'control-behav'
dt.group2[dt.group2 == 'stress'] = 'stress-behav'
dt = dt.join(pd.DataFrame(dt.group2.str.split('-').tolist(), columns= ['stress_group', 'modality']))
dt.head()
# ### Save out for stats
ds.to_csv('/Volumes/group/awagner/sgagnon/AP/data/behav/df_study.csv')
dt.to_csv('/Volumes/group/awagner/sgagnon/AP/data/behav/df_test.csv')
| AP/analysis/AssocPlace_generate_mergedbehav.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Pytorch (base env)
# language: python
# name: base
# ---
import torch.utils.data as utils
# +
import torch
import torch.nn as nn
import torchvision.transforms as transforms
import torchvision.datasets as dsets
from torch.autograd import Variable
from torch.optim import lr_scheduler
from sklearn.metrics import balanced_accuracy_score
from torch.utils.data import Dataset, DataLoader
import pandas as pd
import pickle
import numpy as np
from barbar import Bar
import time
# %matplotlib inline
import matplotlib.pyplot as plt
# -
torch.manual_seed(27)
def plot_acc(history):
fig, axarr = plt.subplots(figsize=(12,6), ncols=2)
axarr[0].plot(range(0, len(history)), history['acc'], label='train score')
axarr[0].plot(range(0, len(history)), history['val_acc'], label='test score')
axarr[0].set_xlabel('Number of Epochs', fontsize=18)
axarr[0].set_ylabel('Accuracy', fontsize=18)
axarr[0].set_ylim([0,1])
axarr[1].plot(range(0, len(history)), history['acc'], label='train score')
axarr[1].plot(range(0, len(history)), history['val_acc'], label='test score')
axarr[1].set_xlabel('Number of Epochs', fontsize=18)
axarr[1].set_ylabel('Accuracy', fontsize=18)
axarr[1].set_ylim([0.7,1])
plt.legend()
plt.show()
DATAPATH = 'data/features/'
MODELPATH = 'output/models/'
# ### STEP 2: LOADING DATASET
class CattleSoundDataset(Dataset):
""" FreeSound dataset."""
# Initialize your data, download, etc.
def __init__(self, X, y):
self.len = X.shape[0]
self.x_data = torch.from_numpy(X)
self.y_data = torch.from_numpy(y)
def __getitem__(self, index):
return (self.x_data[index], self.y_data[index])
def __len__(self):
return self.len
X_train = np.load('data/train_test/X_mel_train.npy')
X_test = np.load('data/train_test/X_mel_test.npy')
y_train = np.load('data/train_test/y_mel_train.npy')
y_test = np.load('data/train_test/y_mel_test.npy')
# +
print('X_train:', X_train.shape)
print('X_test:', X_test.shape)
print('y_train:', y_train.shape)
print('y_test:', y_test.shape)
# -
train_dataset = CattleSoundDataset(X_train, y_train)
test_dataset = CattleSoundDataset(X_test, y_test)
# ### STEP 2: MAKING DATASET ITERABLE
batch_size = 32
n_iters = 1000
num_epochs = n_iters / (len(train_dataset) / batch_size)
num_epochs = int(num_epochs)
num_epochs
transformations = transforms.Compose([transforms.ToTensor()])
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size= batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
dataset_sizes = {'train':len(train_loader.dataset),'valid':len(test_loader.dataset)}
dataloaders = {'train':train_loader,'valid':test_loader}
# ### STEP 3: CREATE MODEL CLASS
class ConvBlock(nn.Module):
def __init__(self, in_channels, out_channels):
super().__init__()
self.conv1 = nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel_size=3, stride=1, padding=(1,1)),
nn.BatchNorm2d(out_channels),
nn.ReLU(),
)
self.conv2 = nn.Sequential(
nn.Conv2d(out_channels, out_channels, kernel_size=3, stride=1, padding=(1,1)),
nn.BatchNorm2d(out_channels),
nn.ReLU(),
)
self.max_pool = nn.MaxPool2d(2)
self._init_weights()
def _init_weights(self):
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal_(m.weight)
if m.bias is not None:
nn.init.zeros_(m.bias)
elif isinstance(m, nn.BatchNorm2d):
nn.init.constant_(m.weight, 1)
nn.init.zeros_(m.bias)
def forward(self, x):
x = self.conv1(x)
x = self.conv2(x)
x = self.max_pool(x)
return x
class CNNModel(nn.Module):
def __init__(self, num_classes):
super().__init__()
self.conv = nn.Sequential(
ConvBlock(in_channels=1, out_channels=32),
ConvBlock(in_channels=32, out_channels=64),
ConvBlock(in_channels=64, out_channels=64),
ConvBlock(in_channels=64, out_channels=32)
)
self.fc = nn.Sequential(
nn.Dropout(0.5),
nn.Linear(640, 320),
nn.PReLU(),
nn.BatchNorm1d(320),
nn.Dropout(0.1),
nn.Linear(320, num_classes),
)
def forward(self, x):
out = self.conv(x)
out = out.view(out.size(0), -1)
#x = torch.mean(x, dim=3)
#x, _ = torch.max(x, dim=2)
out = self.fc(out)
return out
# ### STEP 4: INSTANTIATE MODEL CLASS
model = CNNModel(num_classes=3)
# +
#######################
# USE GPU FOR MODEL #
#######################
if torch.cuda.is_available():
model.cuda()
# -
# ### STEP 5: INSTANTIATE LOSS CLASS
criterion = nn.CrossEntropyLoss().cuda()
# ### STEP 6: INSTANTIATE OPTIMIZER CLASS
# +
learning_rate = 0.001
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
exp_lr_scheduler = lr_scheduler.StepLR(optimizer, step_size=7, gamma=0.1)
# -
# ### STEP 7: TRAIN THE MODEL
def train_model(model, criterion, optimizer, scheduler, num_epochs=num_epochs, graph=False):
since = time.time()
best_model_wts = model.state_dict()
best_acc = 0.0
history = pd.DataFrame()
train_acc = []
val_acc = []
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'valid']:
if phase == 'train':
scheduler.step()
model.train(True) # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Calculate Accuracy
y_pred = []
y_true = []
# Iterate over data.
for data in dataloaders[phase]:
# get the inputs
inputs, labels = data
# wrap them in Variable
if torch.cuda.is_available():
inputs = Variable(inputs.unsqueeze(1).cuda())
labels = Variable(labels.cuda())
else:
inputs, labels = Variable(inputs.unsqueeze(1)), Variable(labels)
# zero the parameter gradients
optimizer.zero_grad()
# forward
outputs = model(inputs.to(dtype=torch.float))
_, preds = torch.max(outputs.data, 1)
loss = criterion(outputs, labels)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.data
running_corrects += torch.sum(preds == labels.data)
if torch.cuda.is_available():
y_pred += preds.cpu().numpy().tolist()
y_true += labels.cpu().numpy().tolist()
else:
y_pred += preds.numpy()
y_true += labels.numpy()
epoch_loss = running_loss / dataset_sizes[phase]
#epoch_acc = running_corrects / dataset_sizes[phase]
epoch_acc = balanced_accuracy_score(y_true, y_pred)
if phase == 'train':
train_acc.append(epoch_acc)
else:
val_acc.append(epoch_acc)
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
phase, epoch_loss, epoch_acc))
# deep copy the model
if phase == 'valid' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = model.state_dict()
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(
time_elapsed // 60, time_elapsed % 60))
print('Best val Acc: {:4f}'.format(best_acc))
# load best model weights
model.load_state_dict(best_model_wts)
history['val_acc'] = val_acc
history['acc'] = train_acc
if graph:
print()
print('==========' * 10)
plot_acc(history)
return model
# +
model = train_model(model, criterion, optimizer, exp_lr_scheduler,
num_epochs=num_epochs, graph=True)
# + active=""
# # Print model's state_dict
# print("Model's state_dict:")
# for param_tensor in model.state_dict():
# print(param_tensor, "\t", model.state_dict()[param_tensor].size())
#
# # Print optimizer's state_dict
# print("Optimizer's state_dicat:")
# for var_name in optimizer.state_dict():
# print(var_name, "\t", optimizer.state_dict()[var_name])
# -
# ### STEP 8: SAVING THE MODEL
state = {
'epoch': num_epochs,
'state_dict': model.state_dict(),
'optimizer': optimizer.state_dict()
}
torch.save(state, MODELPATH+'cnn_pretrained.model')
inputs, labels = next(iter(dataloaders['train']))
inputs.shape
conv = nn.Sequential(
ConvBlock(in_channels=1, out_channels=32),
ConvBlock(in_channels=32, out_channels=64),
ConvBlock(in_channels=64, out_channels=64),
ConvBlock(in_channels=64, out_channels=32)
)
inputs, labels = Variable(inputs.unsqueeze(1)), Variable(labels)
output = conv(inputs.to(dtype=torch.float))
output.shape
output = output.view(output.size(0), -1)
output.shape
X
| notebooks/cnn/train_cnn_lenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of the input data
# - What is typical customer/fraudster behavior?
# - Which type of aggregated information could be useful for the simulator?
# - Where are structural differences between fraud/non-fraud?
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
from datetime import datetime, timedelta
import utils_data
from os.path import join
from IPython.display import display
dates_2016 = [datetime(2016, 1, 1) + timedelta(days=i) for i in range(366)]
# ##### Read in dataset and split into fraud/non-fraud
# +
dataset01, dataset0, dataset1 = utils_data.get_real_dataset()
datasets = [dataset0, dataset1]
out_folder = utils_data.FOLDER_REAL_DATA_ANALYSIS
# -
# ##### Print some basic info about the dataset
print(dataset01.head())
data_stats = utils_data.get_real_data_stats()
data_stats.to_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'aggregated_data.csv'))
display(data_stats)
# Percentage of fraudulent cards also in genuine transactions:
most_used_card = dataset0['CardID'].value_counts().index[0]
print("Card (ID) with most transactions: ", most_used_card)
# ## 1. TIME of TRANSACTION:
# Here we analyse number of transactions regarding time.
# ### 1.1 Activity per day:
plt.figure(figsize=(15, 5))
plt_idx = 1
for d in datasets:
plt.subplot(1, 2, plt_idx)
trans_dates = d["Global_Date"].apply(lambda date: date.date())
all_trans = trans_dates.value_counts().sort_index()
date_num = matplotlib.dates.date2num(all_trans.index)
plt.plot(date_num, all_trans.values, 'k.', label='num trans.')
plt.plot(date_num, np.zeros(len(date_num))+np.sum(all_trans)/366, 'g--',label='average')
plt_idx += 1
plt.title(d.name, size=20)
plt.xlabel('days (1.1.16 - 31.12.16)', size=15)
plt.xticks([])
plt.xlim(matplotlib.dates.date2num([datetime(2016,1,1), datetime(2016,12,31)]))
if plt_idx == 2:
plt.ylabel('num transactions', size=15)
plt.legend(fontsize=15)
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'time_day-in-year'))
plt.show()
# Analysis:
# - It's interesting that there seems to be some kind of structure in the fraudster behavior. I.e., there are many days on which the number of frauds is exactly the same. This must either be due to some peculiarity in the data (are these days where fraud was investigated more?) or because the fraudsters do coordinated attacks
# ### 1.2 Activity per day in a month:
# +
monthdays_2016 = np.unique([dates_2016[i].day for i in range(366)], return_counts=True)
monthdays_2016 = monthdays_2016[1][monthdays_2016[0]-1]
plt.figure(figsize=(12, 5))
plt_idx = 1
monthday_frac = np.zeros((31, 2))
idx = 0
for d in datasets:
# get the average number of transactions per day in a month
monthday = d["Local_Date"].apply(lambda date: date.day).value_counts().sort_index()
monthday /= monthdays_2016
if idx > -1:
monthday_frac[:, idx] = monthday.values / np.sum(monthday.values, axis=0)
idx += 1
plt.subplot(1, 2, plt_idx)
plt.plot(monthday.index, monthday.values, 'ko')
plt.plot(monthday.index, monthday.values, 'k-', markersize=0.1)
plt.plot(monthday.index, np.zeros(31)+np.sum(monthday)/31, 'g--', label='average')
plt.title(d.name, size=20)
plt.xlabel('day in month', size=15)
if plt_idx == 1:
plt.ylabel('avg. num transactions', size=15)
plt_idx += 1
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'time_day-in-month'))
plt.show()
# save the resulting data
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT, 'monthday_frac'), monthday_frac)
# -
# Analysis:
# - the amount of transactions does not depend on the day in a month in a utilisable way
# ### 1.3 Activity per weekday:
# +
weekdays_2016 = np.unique([dates_2016[i].weekday() for i in range(366)], return_counts=True)
weekdays_2016 = weekdays_2016[1][weekdays_2016[0]]
plt.figure(figsize=(12, 5))
plt_idx = 1
weekday_frac = np.zeros((7, 2))
idx = 0
for d in datasets:
weekday = d["Local_Date"].apply(lambda date: date.weekday()).value_counts().sort_index()
weekday /= weekdays_2016
if idx > -1:
weekday_frac[:, idx] = weekday.values / np.sum(weekday.values, axis=0)
idx += 1
plt.subplot(1, 2, plt_idx)
plt.plot(weekday.index, weekday.values, 'ko')
plt.plot(weekday.index, weekday.values, 'k-', markersize=0.1)
plt.plot(weekday.index, np.zeros(7)+np.sum(weekday)/7, 'g--', label='average')
plt.title(d.name, size=20)
plt.xlabel('weekday', size=15)
plt.xticks(range(7), ['Mo', 'Tu', 'We', 'Th', 'Fr', 'Sa', 'Su'])
if plt_idx == 1:
plt.ylabel('avg. num transactions', size=15)
plt_idx += 1
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'time_day-in-week'))
plt.show()
# save the resulting data
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT, 'weekday_frac'), weekday_frac)
# -
# Analysis:
# - the amount of transactions does not depend on the day in a week in a utilisable way
# ### 1.4 Activity per month in a year:
# +
monthdays = np.array([31, 29, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31])
plt.figure(figsize=(12, 5))
plt_idx = 1
month_frac = np.zeros((12, 2))
idx = 0
for d in datasets:
month = d["Local_Date"].apply(lambda date: date.month).value_counts().sort_index()
# correct for different number of days in a month
month = month / monthdays[month.index.values-1] * np.mean(monthdays[month.index.values-1])
if idx > -1:
month_frac[month.index-1, idx] = month.values / np.sum(month.values, axis=0)
idx += 1
plt.subplot(1, 2, plt_idx)
plt.plot(month.index, month.values, 'ko')
plt.plot(month.index, month.values, 'k-', markersize=0.1)
plt.plot(range(1,13), np.zeros(12)+np.sum(month)/12, 'g--', label='average')
plt.title(d.name, size=20)
plt.xlabel('month', size=15)
plt.xticks(range(1, 13), ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec'])
if plt_idx == 1:
plt.ylabel('num transactions', size=15)
plt_idx += 1
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'time_month-in-year'))
plt.show()
# save the resulting data
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT, 'month_frac'), month_frac)
# -
# Analysis:
# - people buy more in summer than in winter
# ### 1.5 Activity per hour of day:
# +
plt.figure(figsize=(12, 5))
plt_idx = 1
hour_frac = np.zeros((24, 2))
idx = 0
for d in datasets:
hours = d["Local_Date"].apply(lambda date: date.hour).value_counts().sort_index()
hours /= 366
if idx > -1:
hour_frac[hours.index.values, idx] = hours.values / np.sum(hours.values, axis=0)
idx += 1
plt.subplot(1, 2, plt_idx)
plt.plot(hours.index, hours.values, 'ko')
plt.plot(hours.index, hours.values, 'k-', markersize=0.1, label='transactions')
plt.plot(range(24), np.zeros(24)+np.sum(hours)/24, 'g--', label='average')
plt.title(d.name, size=20)
plt.xlabel('hour', size=15)
# plt.xticks([])
if plt_idx == 1:
plt.ylabel('avg. num transactions', size=15)
plt_idx += 1
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'time_hour-in-day'))
plt.show()
# save the resulting data
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT, 'hour_frac'), hour_frac)
# -
# Analysis:
# - the hour of day is very important: people spend most in the evening and least during the night; fraud is usually committed in the night
# +
# extract only hours
date_hour_counts = dataset0["Local_Date"].apply(lambda d: d.replace(minute=0, second=0)).value_counts(sort=False)
hours = np.array(list(map(lambda d: d.hour, list(date_hour_counts.index))))
counts = date_hour_counts.values
hour_mean = np.zeros(24)
hour_min = np.zeros(24)
hour_max = np.zeros(24)
hour_std = np.zeros(24)
for h in range(24):
hour_mean[h] = np.mean(counts[hours==h])
hour_min[h] = np.min(counts[hours==h])
hour_max[h] = np.max(counts[hours==h])
hour_std[h] = np.std(counts[hours==h])
print(np.vstack((range(24), hour_min, hour_max, hour_mean, hour_std)).T)
# -
# ### 1.6 TEST: Do the above calculated fractions lead to the correct amount of transactions?
# +
# total number of transactions we want in one year
aggregated_data = pd.read_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'aggregated_data.csv'), index_col=0)
trans_per_year = np.array(aggregated_data.loc['transactions'].values, dtype=np.float)[1:]
# transactions per day in a month
frac_monthday = np.load(join(utils_data.FOLDER_SIMULATOR_INPUT, 'monthday_frac.npy'))
# transactions per day in a week
frac_weekday = np.load(join(utils_data.FOLDER_SIMULATOR_INPUT, 'weekday_frac.npy'))
# transactions per month in a year
frac_month = np.load(join(utils_data.FOLDER_SIMULATOR_INPUT, 'month_frac.npy'))
# transactions hour in a day
frac_hour = np.load(join(utils_data.FOLDER_SIMULATOR_INPUT, 'hour_frac.npy'))
cust_idx = 0
std_transactions = 1000
num_customers = 200
# get the probability of a transaction in a given hour
curr_date = datetime(2016, 1, 1)
num_trans = 0
for i in range(366*24):
new_trans = float(trans_per_year[cust_idx])
new_trans *= frac_month[curr_date.month-1, cust_idx]
new_trans *= frac_monthday[curr_date.day-1, cust_idx]
new_trans *= 7 * frac_weekday[curr_date.weekday(), cust_idx]
new_trans *= frac_hour[curr_date.hour, cust_idx]
num_trans += new_trans
curr_date += timedelta(hours=1)
print(curr_date)
print(trans_per_year[cust_idx])
print(num_trans)
print("")
# the difference happens because some months have longer/shorter days.
# We did not want to scale up the transactions on day 31 because that's unrealistic.
curr_date = datetime(2016, 1, 1)
num_trans = 0
for i in range(366*24):
for c in range(num_customers):
# num_trans is the number of transactions the customer will make in this hour
# we assume that we have enough customers to model that each customer can make max 1 transaction per hour
cust_trans = float(trans_per_year[cust_idx])
cust_trans += np.random.normal(0, std_transactions, 1)[0]
cust_trans /= num_customers
cust_trans *= frac_month[curr_date.month-1, cust_idx]
cust_trans *= frac_monthday[curr_date.day-1, cust_idx]
cust_trans *= 7 * frac_weekday[curr_date.weekday(), cust_idx]
cust_trans *= frac_hour[curr_date.hour, cust_idx]
cust_trans += np.random.normal(0, 0.01, 1)[0]
if cust_trans > np.random.uniform(0, 1, 1)[0]:
num_trans += 1
curr_date += timedelta(hours=1)
print(curr_date)
print(trans_per_year[cust_idx])
print(num_trans)
print("")
# -
# ## 2. COUNTRY
# ### 2.1 Country per transaction:
# +
country_counts = pd.concat([d['Country'].value_counts() for d in datasets], axis=1)
country_counts.fillna(0, inplace=True)
country_counts.columns = ['non-fraud', 'fraud']
country_counts[['non-fraud', 'fraud']] /= country_counts.sum(axis=0)
# save the resulting data
country_counts.to_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'country_frac.csv'))
countries_large = []
for c in ['non-fraud', 'fraud']:
countries_large.extend(country_counts.loc[country_counts[c] > 0.05].index)
countries_large = np.unique(countries_large)
countries_large_counts = []
for c in countries_large:
countries_large_counts.append(country_counts.loc[c, 'non-fraud'])
countries_large = [countries_large[np.argsort(countries_large_counts)[::-1][i]] for i in range(len(countries_large))]
plt.figure(figsize=(10,5))
bottoms = np.zeros(3)
for i in range(len(countries_large)):
c = countries_large[i]
plt.bar((0, 1, 2), np.concatenate((country_counts.loc[c], [0])), label=c, bottom=bottoms)
bottoms += np.concatenate((country_counts.loc[c], [0]))
# fill up the rest
plt.bar((0, 1), 1-bottoms[:-1], bottom=bottoms[:-1], label='rest')
plt.legend(fontsize=20)
plt.xticks([0, 1], ['non-fraud', 'fraud'], size=15)
plt.ylabel('fraction transactions made', size=15)
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'country_distribution'))
plt.show()
# -
# ## 3. CURRENCY
# ### 3.1 Currency per Transaction
# +
currency_counts = pd.concat([d['Currency'].value_counts() for d in datasets], axis=1)
currency_counts.fillna(0, inplace=True)
currency_counts.columns = ['non-fraud', 'fraud']
currency_counts[['non-fraud', 'fraud']] /= currency_counts.sum(axis=0)
currencies_large = []
for c in ['non-fraud', 'fraud']:
currencies_large.extend(currency_counts.loc[currency_counts[c] > 0].index)
currencies_large = np.unique(currencies_large)
currencies_large_counts = []
for c in currencies_large:
currencies_large_counts.append(currency_counts.loc[c, 'non-fraud'])
currencies_large = [currencies_large[np.argsort(currencies_large_counts)[::-1][i]] for i in range(len(currencies_large))]
plt.figure(figsize=(10,5))
bottoms = np.zeros(3)
for i in range(len(currencies_large)):
c = currencies_large[i]
plt.bar((0, 1, 2), np.concatenate((currency_counts.loc[c], [0])), label=c, bottom=bottoms)
bottoms += np.concatenate((currency_counts.loc[c], [0]))
plt.legend(fontsize=20)
plt.xticks([0, 1], ['non-fraud', 'fraud'], size=15)
plt.ylabel('fraction of total transactions made', size=15)
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'currency_distribution'))
plt.show()
# -
# ### 3.1 Currency per country
# Check how many cards make purchases in several currencies:
curr_per_cust = dataset0[['CardID', 'Currency']].groupby('CardID')['Currency'].value_counts().index.get_level_values(0)
print(len(curr_per_cust))
print(len(curr_per_cust.unique()))
print(len(curr_per_cust) - len(curr_per_cust.unique()))
# CONCLUSION: Only 243 cards out of 54,000 puchased things in several currencies.
# Estimate the probability of selection a currency, given a country:
# +
curr_per_country0 = dataset0.groupby(['Country'])['Currency'].value_counts(normalize=True)
curr_per_country1 = dataset1.groupby(['Country'])['Currency'].value_counts(normalize=True)
curr_per_country0.to_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'currency_per_country0.csv'))
curr_per_country1.to_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'currency_per_country1.csv'))
# -
# ## 4. Merchants
# ### 4.1: Merchants per Currency
plt.figure(figsize=(7,5))
currencies = dataset01['Currency'].unique()
merchants = dataset01['MerchantID'].unique()
for curr_idx in range(len(currencies)):
for merch_idx in range(len(merchants)):
plt.plot(range(len(currencies)), np.zeros(len(currencies))+merch_idx, 'r-', linewidth=0.2)
if currencies[curr_idx] in dataset01.loc[dataset01['MerchantID'] == merch_idx, 'Currency'].values:
plt.plot(curr_idx, merch_idx, 'ko')
plt.xticks(range(len(currencies)), currencies)
plt.ylabel('Merchant ID', size=15)
plt.xlabel('Currency', size=15)
plt.tight_layout()
plt.show()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'currency_per_merchant'))
# We conclude from this that most merchants only sell things in one currenyc; thus, we will let each customer select the merchant given the currency that the customer has (which is unique).
# Estimate the probability of selection a merchat, given the currency:
# +
merch_per_curr0 = dataset0.groupby(['Currency'])['MerchantID'].value_counts(normalize=True)
merch_per_curr1 = dataset1.groupby(['Currency'])['MerchantID'].value_counts(normalize=True)
merch_per_curr0.to_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'merchant_per_currency0.csv'))
merch_per_curr1.to_csv(join(utils_data.FOLDER_SIMULATOR_INPUT, 'merchant_per_currency1.csv'))
# -
# ### 4.2 Number transactions per merchant
# +
merchant_count0 = dataset0['MerchantID'].value_counts().sort_index()
merchant_count1 = dataset1['MerchantID'].value_counts().sort_index()
plt.figure(figsize=(15,10))
ax = plt.subplot(2, 1, 1)
ax.bar(merchant_count0.index.values, merchant_count0.values)
rects = ax.patches
for rect, label in zip(rects, merchant_count0.values):
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width()/2, height, label, ha='center', va='bottom')
plt.ylabel('num transactions')
plt.xticks([])
plt.xlim([-0.5, data_stats.loc['num merchants', 'all']+0.5])
ax = plt.subplot(2, 1, 2)
ax.bar(merchant_count1.index.values, merchant_count1.values)
rects = ax.patches
for rect, label in zip(rects, merchant_count1.values):
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width()/2, height, label, ha='center', va='bottom')
plt.ylabel('num transactions')
plt.xlabel('Merchant ID')
plt.xlim([-0.5, data_stats.loc['num merchants', 'all']+0.5])
plt.tight_layout()
plt.show()
# -
# ## 5. Transaction Amount
# ### 5.1 Amount over time
plt.figure(figsize=(12, 10))
plt_idx = 1
for d in datasets:
plt.subplot(2, 1, plt_idx)
plt.plot(range(d.shape[0]), d['Amount'], 'k.')
# plt.plot(date_num, amount, 'k.', label='num trans.')
# plt.plot(date_num, np.zeros(len(date_num))+np.mean(all_trans), 'g',label='average')
plt_idx += 1
# plt.title(d.name, size=20)
plt.xlabel('transactions', size=15)
plt.xticks([])
if plt_idx == 2:
plt.ylabel('amount', size=15)
plt.legend(fontsize=15)
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'amount_day-in-year'))
plt.show()
print(dataset0.loc[dataset0['Amount'] == 5472.53,['Local_Date', 'CardID', 'MerchantID', 'Amount', 'Currency', 'Country']])
# ### 5.2 Amount distribution
plt.figure(figsize=(10,5))
bins = [0, 5, 25, 50, 100, 1000, 11000]
plt_idx = 1
for d in datasets:
amount_counts, loc = np.histogram(d["Amount"], bins=bins)
amount_counts = np.array(amount_counts, dtype=np.float)
amount_counts /= np.sum(amount_counts)
plt.subplot(1, 2, plt_idx)
am_bot = 0
for i in range(len(amount_counts)):
plt.bar(plt_idx, amount_counts[i], bottom=am_bot, label='{}-{}'.format(bins[i], bins[i+1]))
am_bot += amount_counts[i]
plt_idx += 1
plt.ylim([0, 1.01])
plt.legend()
# plt.title("Amount distribution")
plt_idx += 1
plt.show()
plt.figure(figsize=(12, 10))
plt_idx = 1
for d in datasets:
plt.subplot(2, 1, plt_idx)
min_amount = min(d['Amount'])
max_amount = max(d['Amount'])
plt.plot(range(d.shape[0]), np.sort(d['Amount']), 'k.', label='transaction')
# plt.plot(date_num, amount, 'k.', label='num trans.')
plt.plot(np.linspace(0, d.shape[0], 100), np.zeros(100)+np.mean(d['Amount']), 'g--',label='average')
plt_idx += 1
plt.title(d.name, size=20)
plt.ylabel('amount', size=15)
if plt_idx == 3:
plt.xlabel('transactions', size=15)
else:
plt.legend(fontsize=15)
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'amount_day-in-year'))
plt.show()
# For each merchant, we will have a probability distribution over the amount spent
# +
from scipy.optimize import curve_fit
def sigmoid(x, x0, k):
y = 1 / (1 + np.exp(-k * (x - x0)))
return y
num_merchants = data_stats.loc['num merchants', 'all']
num_bins = 20
merchant_amount_distr = np.zeros((2, num_merchants, 2*num_bins+1))
plt.figure(figsize=(15, 5))
plt_idx = 1
for dataset in [dataset0, dataset1]:
for m in dataset0['MerchantID'].unique():
# get all transactions from this merchant
trans_merch = dataset.loc[dataset['MerchantID']==m]
num_transactions = trans_merch.shape[0]
if num_transactions > 0:
# get the amounts paid for the transactions with this merchant
amounts = trans_merch['Amount']
bins_height, bins_edges = np.histogram(amounts, bins=num_bins)
bins_height = np.array(bins_height, dtype=np.float)
bins_height /= np.sum(bins_height)
merchant_amount_distr[int(plt_idx > 7), (plt_idx-1)%7, :] = np.concatenate((bins_height, bins_edges))
plt.subplot(2, num_merchants, plt_idx)
plt.hist(amounts, bins=num_bins)
plt_idx += 1
plt.tight_layout()
plt.show()
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT,'merchant_amount_distr'), merchant_amount_distr)
# +
from scipy.optimize import curve_fit
def sigmoid(x, x0, k):
y = 1 / (1 + np.exp(-k * (x - x0)))
return y
num_merchants = data_stats.loc['num merchants', 'all']
merchant_amount_parameters = np.zeros((2, num_merchants, 4))
plt.figure(figsize=(15, 5))
plt_idx = 1
for dataset in [dataset0, dataset1]:
for m in dataset0['MerchantID'].unique():
# get all transactions from this merchant
trans_merch = dataset.loc[dataset['MerchantID']==m]
num_transactions = trans_merch.shape[0]
if num_transactions > 0:
# get the amounts paid for the transactions with this merchant
amounts = np.sort(trans_merch['Amount'])
min_amount = min(amounts)
max_amount = max(amounts)
amounts_normalised = (amounts - min_amount) / (max_amount - min_amount)
plt.subplot(2, num_merchants, plt_idx)
plt.plot(np.linspace(0, 1, num_transactions), amounts, '.')
# fit sigmoid
x_vals = np.linspace(0, 1, 100)
try:
p_sigmoid, _ = curve_fit(sigmoid, np.linspace(0, 1, num_transactions), amounts_normalised)
amounts_predict = sigmoid(x_vals, *p_sigmoid)
amounts_predict_denormalised = amounts_predict * (max_amount - min_amount) + min_amount
plt.plot(x_vals, amounts_predict_denormalised)
except:
# fit polynomial
p_poly = np.polyfit(np.linspace(0, 1, num_transactions), amounts_normalised, 2)
amounts_predict = np.polyval(p_poly, x_vals)
p_sigmoid, _ = curve_fit(sigmoid, x_vals, amounts_predict)
amounts_predict = sigmoid(x_vals, *p_sigmoid)
amounts_predict_denormalised = amounts_predict * (max_amount - min_amount) + min_amount
plt.plot(x_vals, amounts_predict_denormalised)
merchant_amount_parameters[int(plt_idx > 7), (plt_idx-1)%7] = [min_amount, max_amount, p_sigmoid[0], p_sigmoid[1]]
plt_idx += 1
plt.tight_layout()
plt.show()
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT,'merchant_amount_parameters'), merchant_amount_parameters)
print(merchant_amount_parameters)
# -
# We conclude that the normal customers and fraudsters follow roughly the same distribution, so we will only have one per merchant; irrespective of whether a genuine or fraudulent customer is making the transaction.
# +
from scipy.optimize import curve_fit
def sigmoid(x, x0, k):
y = 1 / (1 + np.exp(-k * (x - x0)))
return y
num_merchants = data_stats.loc['num merchants', 'all']
merchant_amount_parameters = np.zeros((2, num_merchants, 4))
plt.figure(figsize=(6, 3))
plt_idx = 1
dataset = dataset0
m = dataset0['MerchantID'].unique()[0]
# get all transactions from this merchant
trans_merch = dataset.loc[dataset['MerchantID']==m]
num_transactions = trans_merch.shape[0]
# get the amounts paid for the transactions with this merchant
amounts = np.sort(trans_merch['Amount'])
min_amount = min(amounts)
max_amount = max(amounts)
amounts_normalised = (amounts - min_amount) / (max_amount - min_amount)
plt.plot(range(num_transactions), amounts, 'k-', linewidth=2, label='real')
# fit sigmoid
x_vals = np.linspace(0, 1, 100)
x = np.linspace(0, 1, num_transactions)
p_sigmoid, _ = curve_fit(sigmoid, np.linspace(0, 1, num_transactions), amounts_normalised)
amounts_predict = sigmoid(x_vals, *p_sigmoid)
amounts_predict_denormalised = amounts_predict * (max_amount - min_amount) + min_amount
plt.plot(np.linspace(0, num_transactions, 100), amounts_predict_denormalised, 'm--', linewidth=3, label='approx')
merchant_amount_parameters[int(plt_idx > 7), (plt_idx-1)%7] = [min_amount, max_amount, p_sigmoid[0], p_sigmoid[1]]
plt.xlabel('transaction count', fontsize=20)
plt.ylabel('price', fontsize=20)
plt.legend(fontsize=15)
plt.tight_layout()
plt.savefig(join(utils_data.FOLDER_REAL_DATA_ANALYSIS, 'merchant_price_sigmoid_fit'))
plt.show()
# -
# ## Customers
# Here we want to find out how long customers/fraudsters return, i.e., how often the same credit card is used over time.
plt.figure(figsize=(15, 30))
plt_idx = 1
dist_transactions = [[], []]
for d in datasets:
# d = d.loc[d['Date'].apply(lambda date: date.month) < 7]
# d = d.loc[d['Date'].apply(lambda date: date.month) > 3]
plt.subplot(1, 2, plt_idx)
trans_idx = 0
for card in dataset01['CardID'].unique():
card_times = d.loc[d['CardID'] == card, 'Global_Date']
dist_transactions[plt_idx-1].extend([(card_times.iloc[i+1] - card_times.iloc[i]).days for i in range(len(card_times)-1)])
if plt_idx == 2:
num_c = 2
else:
num_c = 10
if len(card_times) > num_c:
card_times = card_times.apply(lambda date: date.date())
card_times = matplotlib.dates.date2num(card_times)
plt.plot(card_times, np.zeros(len(card_times)) + trans_idx, 'k.', markersize=1)
plt.plot(card_times, np.zeros(len(card_times)) + trans_idx, 'k-', linewidth=0.2)
trans_idx += 1
min_date = matplotlib.dates.date2num(min(dataset01['Global_Date']).date())
max_date = matplotlib.dates.date2num(max(dataset01['Global_Date']).date())
# plt.xlim([min_date, max_date])
plt.xticks([])
for m in range(1,13):
datenum = matplotlib.dates.date2num(datetime(2016, m, 1))
plt.plot(np.zeros(2)+datenum, [-1, 1000], 'r-', linewidth=0.5)
if plt_idx == 1:
plt.ylim([0,300])
else:
plt.ylim([0, 50])
plt_idx += 1
plt.show()
# average distance between two transactions with the same card
print(np.mean(dist_transactions[0]))
print(np.mean(dist_transactions[1]))
# At a given transaction, estimate the probability of doing another transaction with the same card.
# +
prob_stay = np.zeros(2)
for k in range(2):
dataset = [dataset0, dataset1][k]
creditcards = dataset.loc[dataset['Global_Date'].apply(lambda d: d.month) > 3]
creditcards = creditcards.loc[creditcards['Global_Date'].apply(lambda d: d.month) < 6]
creditcard_counts = creditcards['CardID'].value_counts()
creditcardIDs = creditcards['CardID']
data = dataset.loc[dataset['Global_Date'].apply(lambda d: d.month) > 3]
single = 0
multi = 0
for i in range(len(creditcards)):
cc = creditcards.iloc[i]['CardID']
dd = creditcards.iloc[i]['Global_Date']
cond1 = data['CardID'] == cc
cond2 = data['Global_Date'] > dd
if len(data.loc[np.logical_and(cond1, cond2)]) == 0:
single += 1
else:
multi += 1
prob_stay[k] = multi/(single+multi)
print('probability of doing another transaction:', prob_stay[k], '{}'.format(['non-fraud', 'fraud'][k]))
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT, 'prob_stay'), prob_stay)
# -
# ## Fraud behaviour
# +
cards0 = dataset0['CardID'].unique()
cards1 = dataset1['CardID'].unique()
print('cards total:', len(np.union1d(cards0, cards1)))
print('fraud cards:', len(cards1))
print('intersection:', len(np.intersect1d(cards0, cards1)))
# go through the cards that were in both sets
cards0_1 = []
cards1_0 = []
cards010 = []
for cib in np.intersect1d(cards0, cards1):
date0 = dataset0.loc[dataset0['CardID']==cib].iloc[0]['Global_Date']
date1 = dataset1.loc[dataset1['CardID']==cib].iloc[0]['Global_Date']
if date0 < date1:
cards0_1.append(cib)
# genuine purchases after fraud
dates00 = dataset0.loc[dataset0['CardID']==cib].iloc[1:]['Global_Date']
if len(dates00)>0:
if sum(dates00>date1)>0:
cards010.append(cib)
else:
cards1_0.append(cib)
print('first genuine then fraud: ', len(cards0_1))
print('first fraud then genuine: ', len(cards1_0))
print('genuine again after fraud: ', len(cards010))
prob_stay_after_fraud = len(cards010)/len(cards0_1)
print('prob of purchase after fraud: ', prob_stay_after_fraud)
np.save(join(utils_data.FOLDER_SIMULATOR_INPUT, 'prob_stay_after_fraud'), prob_stay_after_fraud )
# +
plt.figure(figsize=(10, 25))
dist_transactions = []
trans_idx = 0
data_compromised = dataset01.loc[dataset01['CardID'].apply(lambda cid: cid in np.intersect1d(cards0, cards1))]
no_trans_after_fraud = 0
trans_after_fraud = 0
for card in data_compromised['CardID'].unique():
cards_used = data_compromised.loc[data_compromised['CardID'] == card, ['Global_Date', 'Target']]
dist_transactions.extend([(cards_used.iloc[i+1, 0] - cards_used.iloc[i, 0]).days for i in range(len(cards_used)-1)])
card_times = cards_used['Global_Date'].apply(lambda date: date.date())
card_times = matplotlib.dates.date2num(card_times)
plt.plot(card_times, np.zeros(len(card_times)) + trans_idx, 'k-', linewidth=0.9)
cond0 = cards_used['Target'] == 0
plt.plot(card_times[cond0], np.zeros(len(card_times[cond0])) + trans_idx, 'g.', markersize=5)
cond1 = cards_used['Target'] == 1
plt.plot(card_times[cond1], np.zeros(len(card_times[cond1])) + trans_idx, 'r.', markersize=5)
if max(cards_used.loc[cards_used['Target']==0, 'Global_Date']) > max(cards_used.loc[cards_used['Target']==1, 'Global_Date']):
trans_after_fraud += 1
else:
no_trans_after_fraud += 1
trans_idx += 1
min_date = matplotlib.dates.date2num(min(dataset01['Global_Date']).date())
max_date = matplotlib.dates.date2num(max(dataset01['Global_Date']).date())
plt.xticks([])
plt.ylim([0, trans_idx])
# print lines for months
for m in range(1,13):
datenum = matplotlib.dates.date2num(datetime(2016, m, 1))
plt.plot(np.zeros(2)+datenum, [-1, 1000], 'r-', linewidth=0.5)
plt_idx += 1
plt.show()
print("genuine transactions after fraud: ", trans_after_fraud)
print("fraud is the last transaction: ", no_trans_after_fraud)
# -
# when a fraudster uses an existing card, are country and currency always the same?
# +
plt.figure(figsize=(10, 25))
dist_transactions = []
trans_idx = 0
for card in data_compromised['CardID'].unique():
cards_used = data_compromised.loc[data_compromised['CardID'] == card, ['Global_Date', 'Target', 'Country', 'Currency']]
if len(cards_used['Country'].unique()) > 1 or len(cards_used['Currency'].unique()) > 1:
print(cards_used)
print("")
# -
| data/analyse_data.ipynb |
% -*- coding: utf-8 -*-
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% some housekeeping stuff
register_graphics_toolkit ("gnuplot");
available_graphics_toolkits ();
graphics_toolkit ("gnuplot")
clear
% end of housekeeping
% # Introduction to Matrices
%
% A matrix is an ordered set of numbers listed rectangular
% form
%
% $$ \mathbf{A} = \begin{bmatrix}
% 2 & 5 & 7 & 8 \\
% 5 & 6 & 8 & 9 \\
% 3 & 9 & 0 & 1
% \end{bmatrix} $$
%
% by convention matrices are generally assigned bold
% symbols in mathematical notation
% and the entries don’t have to be integers (or even real)
% The matrix A above has three rows and four columns. It
% is a 3 × 4 matrix
% we denote the element of the second row and the fourth
% column $a_{2,4} = 9$
% +
%plot -s 600,600 -f 'svg'
%A=[2 5 7 8
%5 6 8 9
%3 9 0 1]
%size(A)
%[n,m]=size(A)
%a=A(2,4)
% -
% A **square matrix** has $n$ rows and $n$ columns
% $$ \mathbf{B} = \begin{bmatrix}
% 2 & 5 & 7 \\
% 5 & 6 & 8 \\
% 3 & 9 & 0
% \end{bmatrix}
% $$
% A **diagonal matrix** is a square matrix with all non-diagonal elements equal to zero
% $$ \mathbf{C} = \begin{bmatrix}
% 2 & 0 & 0 \\
% 0 & 6 & 0 \\
% 0 & 0 & 2
% \end{bmatrix}
% $$
% **diagonal elements** are denoted $a_{i,i}$ with $i=1,2,3, ...$
% +
%plot -s 600,600 -f 'svg'
%input the two matrices.
%make the diagonal one using D = diag(v)
%returns a square diagonal matrix with the elements of vector v on the main diagonal.
% -
% a **vector** is a matrix with only one row or column
%
% a **row matrix** is a matrix with one row (also called a row vector)
%
% a **column matrix** is a matrix with one column (also called a column vector)
%
% **transpose of a matrix** is indicated as $\mathbf{A}^\prime$ or $\mathbf{A}^{\mathrm{T}}$ and results from $a_{i,j}=a_{j,i}^\prime$
%
% $$ \mathbf{D} = \begin{bmatrix}
% 2 & 8 & 0 & 6 \\
% 1 & 6 & 0 & 2\\
% 3 & 0 & 2 & 1
% \end{bmatrix}
% \qquad
% \mathbf{D}^\prime = \begin{bmatrix}
% 2 & 1 & 3 \\
% 8 & 6 & 0\\
% 0 & 0 & 2\\
% 6 & 2 & 1
% \end{bmatrix}
% $$
% +
%plot -s 600,600 -f 'svg'
%input D and take the transpose. D=D'
% -
% **identity matrix** is a diagonal matrix with all diagonal elements=1
%
% **sum of matrices** if two matrices are of the same size ($n \times m$) then to add them we just sum the corresponding entries
%
% $$ \begin{bmatrix} 2 & 8\\
% 1 & 6\\
% 3 & 0
% \end{bmatrix}
% +
% \begin{bmatrix} 2 & 1\\
% 8 & 6\\
% 0 & 0
% \end{bmatrix}
% =
% \begin{bmatrix}
% 4 & 9 \\
% 9 & 12\\
% 3 & 0
% \end{bmatrix}
% $$
% +
%plot -s 600,600 -f 'svg'
%perform that matrix addition.
% -
% multiplication by a scaler (a scaler is just a number) $c\mathbf{A} = [c a_{i,j}]$
%
% multiply two matrices $\mathbf{A}$ and $\mathbf{B}$ is only possible if the second dimension of $\mathbf{A}$ is the same as the first dimension of $\mathbf{B}$
%
% so for $\mathbf{A}$ with dimension $m \times n$ and $\mathbf{B}$ with dimension $n \times p$ multiplication is defined as $\mathbf{AB}=\mathbf{C}$ where
%
% $$ c_{i,j}=\sum_{k=1}^{n} a_{i,k}b_{k,j} $$
%
% the dimension of $\mathbf{C}$ is $m \times p$
%
% **EXAMPLE** ...
%
% $$ \mathbf{A}=\begin{bmatrix} 1 & 3 \\ 2 & 4 \end{bmatrix} \qquad \text{and} \qquad \mathbf{B}=\begin{bmatrix} 2 & 5 & 1 \\ 4 & 12 & 6 \end{bmatrix} $$
%
% $\mathbf{A}$ is $2 \times 2$ and $\mathbf{B}$ is $2 \times 3$
%
% then the product is $$ \mathbf{C}=[c_{i,j}]=\sum_{k=1}^{2} a_{i,k}b_{k,j} $$
%
% $$ \begin{bmatrix} (1)(2)+(3)(4) & (1)(5)+(3)(12) & (1)(1)+(3)(6)\\ (2)(2)+(4)(4) & (2)(5)+(4)(12)&(2)(1)+(4)(6) \end{bmatrix}
% =\begin{bmatrix} 14 & 41 & 19 \\ 20 & 58 & 26 \end{bmatrix}
% $$
% +
%plot -s 600,600 -f 'svg'
%CONFIRM the matrix multiliplication. and the final matrix dimension.
% -
% a $n \times n$ matrix $\mathbf{A}$ is said to be invertible if there exists an $n \times n$ matrix $\mathbf{B}$ such that $\mathbf{AB}=\mathbf{BA}=\mathbf{I}$ where $\mathbf{I}$ is an $n \times n$ identity matrix
%
% note: in general $\mathbf{AB}$ is not necessarily equal to $\mathbf{BA}$
%
% the inverse of a matrix is usually written $\mathbf{A}^{-1}$
% +
%plot -s 600,600 -f 'svg'
%pick any nxn matrix to take the inverse of.
%Ainv=inv(A);
%and verify that you get the inverse
%use who command to see what matricess you have in your workspace.
% -
% # SAMPLING STATISTICS
%
% most statistics you are probably familiar with are called sampling statistics
%
% when measuring data we think of taking a finite number of samples from infinitly many possible measurements
%
% so we have this idea of the "true" value and our task is to figure out how good an estimate of that true value we have
%
% if there was no such thing as error we'd always measure the true value
%
% there are two types of error, random and systematic
%
% sampling statitics assume random error
%
% if errors are random then they are distributed as a normal distribution about the mean
%
% normal distribution (or Gaussian distribution)
% $$ f(x) = \dfrac{1}{\sqrt{2\pi\sigma^2}}e^{\frac{-(x-\mu)^2}{2\sigma^2}} $$
%
% so for various values of $x$ we can calculate $f(x)$ dependent on the true mean ($\mu$) and the standard deviation ($\sigma$)
%
% the mean determines the center value and $\sigma$ determines the spread
%
% for infinitely many samples this distribution gives the frequency of each occurance
%
% integrating from $-\infty$ to $\infty$ we'll get a value of 1
%
% # EXERCISE 1
%
% let's write a function to return $f(x)$ for a given $x$, $\mu$ and $\sigma$
%
% then let's write a script to determine and plot distributions with mean 1.5 and $\sigma$ 0.1, 0.25 and 0.5 on the same graph
%
% finally, let's add an integration step to the script to show the area is always one
%
% +
%plot -s 600,600 -f 'svg'
% put all these in an mfile a script to sequentially execute the commands (and
% store all the variables in the same workspace)
%anonymous function
f = @(x,mu,sigma) (1/sqrt(2*pi*sigma^2))*exp(-((x-mu).^2)./2*sigma^2);
%note use of "." for array-wise instead of matrix operation
%define row vector and mu and sigma
x=-6:0.1:9; mu=1.5; sigma=0.5;
% call the function, make sure variables are in the right order
y=f(x,mu,sigma);
% plot it
plot(x,y)
% make plot look nicer. add labels. thicker lines. also add the other two lines
% -
% # EXERCISE 2
%
% octave/matlab can generate random numbers with a normal distribution
%
% let's write a script to generate 2, 3, 10, 100, 1000 random numbers with a mean of 1.5 and a standard deviation of 0.25
%
% calculate the sample mean and standard deviation for these sets
%
% plot histograms for each data set and the theoretical distributions on the sample plot
| .ipynb_checkpoints/LECTURE1_matrices_statistics-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import
# +
# Basic
import pandas as pd
import numpy as np
# ML Toolkit
from robusta.crossval import *
# %load_ext memory_profiler
# -
# # Binary Classification
# ## Data
# +
from catboost.datasets import amazon
X_train, X_test = amazon()
y_train = X_train['ACTION']
X_train.drop(columns='ACTION', inplace=True)
X_train.index.name = 'id'
X_test.set_index('id', inplace=True)
X_train
# -
# ## Task
scoring = 'roc_auc'
cv = 5
# ## Model
# +
from lightgbm import LGBMClassifier
model = LGBMClassifier()
# -
# ## Predict Probability
# ### Averaging (used by default)
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, method='predict_proba',
verbose=2, n_jobs=-1)
y_pred
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, method='predict_proba',
avg_type='auto', verbose=2, n_jobs=-1)
y_pred # the same
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, method='predict_proba',
avg_type='mean', verbose=2, n_jobs=-1)
y_pred # the same
# -
# ### Rank Averaging
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, method='predict_proba',
avg_type='rank', verbose=2, n_jobs=-1)
y_pred
# -
# ## Predict Classes
# ### Soft Vote (used by default)
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, verbose=2, n_jobs=-1)
y_pred.value_counts()
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, avg_type='soft',
verbose=2, n_jobs=-1)
y_pred.value_counts() # the same
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, avg_type='auto',
verbose=2, n_jobs=-1)
y_pred.value_counts() # the same
# -
# ### Hard Vote
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, avg_type='hard',
verbose=2, n_jobs=-1)
y_pred.value_counts() # differs
# -
# # Regression
# ## Task
scoring = 'r2'
cv = 5
# ## Model
# +
from lightgbm import LGBMRegressor
model = LGBMRegressor()
# +
_, y_pred = crossval_predict(model, cv, X_train, y_train, X_new=X_test,
scoring=scoring, verbose=2, n_jobs=-1)
y_pred
| examples/crossval-predict-task-type.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>EnergyUsagePrediction.model_def.asum.v1_0_7.ipynb</b>
# <br/>For my use case "Energy usage prediction based on historical weather and energy usage data.". The original dataset can be downloaded from <a href="https://www.kaggle.com/taranvee/smart-home-dataset-with-weather-information">kaggle</a>
# <br/>The dataset used in this step (feature engineering) has already been transformed in the ETL step.
# <br/>Data exploration is described/performed in "EnergyUsagePrediction.data_exp.asum.1_0_5.Ipynb"
# <br/>ETL is described/performed in "EnergyUsagePrediction.etl.asum.1_0_8.Ipynb"
# <br/>Feature engineering is described/performed in "EnergyUsagePrediction.feature_eng.asum.1_0_8.Ipynb"
# <br/>
# <br/>This task defines the machine learning or deep learning model.
# <br/>
# <br/>Load <i>smart-home-dataset-with-weather-information_post_feature_eng.csv</i> file into pandas dataframe
#
# +
import types
import numpy as np
import pandas as pd
from botocore.client import Config
import ibm_boto3
def __iter__(self): return 0
# @hidden_cell
# The following code accesses a file in your IBM Cloud Object Storage. It includes your credentials.
# You might want to remove those credentials before you share the notebook.
client_x = ibm_boto3.client(service_name='s3',
ibm_api_key_id='[credentials]',
ibm_auth_endpoint="https://iam.cloud.ibm.com/oidc/token",
config=Config(signature_version='oauth'),
endpoint_url='https://s3.eu-geo.objectstorage.service.networklayer.com')
body = client_x.get_object(Bucket='default-donotdelete-pr-dczw8ajohz6wjh',Key='smart-home-dataset-with-weather-information_post_feature_eng.csv')['Body']
# add missing __iter__ method, so pandas accepts body as file-like object
if not hasattr(body, "__iter__"): body.__iter__ = types.MethodType( __iter__, body )
df = pd.read_csv(body)
df.head()
# -
# import the necessary packages
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Dense
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.layers.experimental import preprocessing
#from keras import backend as K
# %matplotlib inline
import matplotlib.pyplot as plt
# For usability we define constants for the labels
#
# +
lbTimestamp = 'Timestamp'
lbTotalEneryUsage = 'TotalUsage_kW'
lbTemperature = 'Temperature_F'
lbTemperatureNormalized = 'Temperature_F_normalized'
lbHumidity = 'Humidity'
lbHumidityNormalized = 'Humidity_normalized'
lbPressure = 'Pressure_hPa'
lbPressureNormalized = 'Pressure_hPa_normalized'
lbWindSpeed = 'WindSpeed'
lbWindSpeedNormalized = 'WindSpeed_normalized'
lbCloudCover = 'cloudCover'
lbCloudCoverNormalized = 'cloudCover_normalized'
lbWindBearing = 'WindBearing'
lbWindBearingNormalized = 'WindBearing_normalized'
lbPrecipIntensity = 'PrecipIntensity'
lbPrecipIntensityNormalized = 'PrecipIntensity_normalized'
lbDewPoint = 'dewPoint_F'
lbDewPointNormalized = 'dewPoint_F_normalized'
lbDayOfYear='dayOfYear'
lbDayOfYearNormalized='dayOfYear_normalized'
lbHourOfDay='hourOfDay'
lbHourOfDayNormalized='hourOfDay_normalized'
lbMinuteOfDay='minuteOfDay'
lbMinuteOfDayNormalized='minuteOfDay_normalized'
lbWeatherIndicatorClearDay = 'weatherIndicator_clear-day'
lbWeatherIndicatorClearNight = 'weatherIndicator_clear-night'
lbWeatherIndicatorCloudy = 'weatherIndicator_cloudy'
lbWeatherIndicatorFog = 'weatherIndicator_fog'
lbWeatherIndicatorPartlyCloudyDay = 'weatherIndicator_partly-cloudy-day'
lbWeatherIndicatorPartlyCloudyNight = 'weatherIndicator_partly-cloudy-night'
lbWeatherIndicatorRain = 'weatherIndicator_rain'
lbWeatherIndicatorSnow = 'weatherIndicator_snow'
lbWeatherIndicatorWind = 'weatherIndicator_wind'
# -
df.info()
# We take 80% of the dataset for training, 20 procent for testing.
# <br/>We use a seed to build deterministic training and test data.
inputColumns = [lbTemperatureNormalized,
lbHumidityNormalized,
lbWindSpeedNormalized,
lbWindBearingNormalized,
lbDewPointNormalized,
lbPrecipIntensityNormalized,
lbDayOfYearNormalized,
lbHourOfDayNormalized,
lbMinuteOfDayNormalized,
lbWeatherIndicatorClearDay,
lbWeatherIndicatorClearNight,
lbWeatherIndicatorCloudy,
lbWeatherIndicatorFog,
lbWeatherIndicatorPartlyCloudyDay,
lbWeatherIndicatorPartlyCloudyNight,
lbWeatherIndicatorRain,
lbWeatherIndicatorSnow,
lbWeatherIndicatorWind]
outputColumns = [lbTotalEneryUsage]
train=df.sample(frac=0.8,random_state=42) #random state is a seed value
test=df.drop(train.index)
train_x=train[inputColumns]
test_x=test[inputColumns]
train_x.info()
train_y=train[outputColumns]
test_y=test[outputColumns]
train_y.info()
# First we will use a traditional machine learning algorithm: LinearRegression
from sklearn import linear_model
# +
# with sklearn
regr = linear_model.LinearRegression()
regr.fit(train_x, train_y)
# -
print('Intercept: \n', regr.intercept_)
print('Coefficients: \n', regr.coef_)
predicted = regr.predict(test_x)
import sklearn.metrics as sm
print("Mean absolute error =", round(sm.mean_absolute_error(test_y, predicted), 2))
print("Mean squared error =", round(sm.mean_squared_error(test_y, predicted), 2))
print("Median absolute error =", round(sm.median_absolute_error(test_y, predicted), 2))
print("Explain variance score =", round(sm.explained_variance_score(test_y, predicted), 2))
print("R2 score =", round(sm.r2_score(test_y, predicted), 2))
from sklearn.metrics import mean_squared_error
mean_squared_error(test_y, predicted)
samples = test.sample(200,random_state=42)
samples_x=samples[inputColumns]
samples_y=samples[outputColumns]
predictedSamples = regr.predict(samples_x)
figure=plt.figure(figsize=(12,12))
samples_y = samples.reset_index()
plt.plot(samples_y[lbTotalEneryUsage], figure=figure)
plt.xlabel("x")
plt.ylabel("actual+predicted")
plt.plot(predictedSamples, figure=figure)
plt.show()
# Now let's start with a Deep Learning approach using Keras Sequential Model
# +
# Create Keras model
model = Sequential()
batch_size = 32
input_dim=18
model.add(Dense(batch_size*input_dim, kernel_initializer = "uniform",input_dim=input_dim, name="input"))
model.add(Dense(256, activation="relu", name="hiddenlayer1"))
model.add(Dense(1, name="output"))
# Gradient descent algorithm
adam = Adam(0.001)
model.compile(loss='mse', optimizer=adam)
history = model.fit(train_x, train_y, epochs=15, batch_size=batch_size)
# -
plt.plot(history.history['loss'])
plt.xlabel("No. of Iterations")
plt.ylabel("J(Theta1 Theta0)/Cost")
plt.show()
model.evaluate(test_x, test_y)
predicted = model.predict(test_x)
print("Mean absolute error =", round(sm.mean_absolute_error(test_y, predicted), 2))
print("Mean squared error =", round(sm.mean_squared_error(test_y, predicted), 2))
print("Median absolute error =", round(sm.median_absolute_error(test_y, predicted), 2))
print("Explain variance score =", round(sm.explained_variance_score(test_y, predicted), 2))
print("R2 score =", round(sm.r2_score(test_y, predicted), 2))
def printSamplePredictedVsActual(myModel, testData):
test_sample= testData.sample(200,random_state=42)
test_sample = test_sample.reset_index()
test_sample_x = test_sample[inputColumns]
test_sample_y = test_sample[outputColumns]
test_predicted = myModel.predict(test_sample_x)
figure=plt.figure(figsize=(12,12))
plt.plot(test_sample[outputColumns], figure=figure)
plt.xlabel("x")
plt.ylabel("actual")
plt.plot(test_predicted, figure=figure)
plt.show()
printSamplePredictedVsActual(model, test)
print(str(predicted.min()) +','+str(predicted.max())+str(predicted.mean()))
# +
# Create Keras model2
model2 = Sequential()
batch_size = 32
input_dim=18
model2.add(Dense(batch_size*input_dim, kernel_initializer = "uniform",input_dim=input_dim, name="input"))
model2.add(Dropout(0.2))
model2.add(Dense(256, activation="relu", name="hiddenlayer1"))
model2.add(Dense(256, activation="relu", name="hiddenlayer2"))
model2.add(Dense(256, activation="relu", name="hiddenlayer3"))
model2.add(Dense(256, activation="relu", name="hiddenlayer4"))
model2.add(Dense(256, activation="relu", name="hiddenlayer5"))
model2.add(Dense(1, name="output"))
# Gradient descent algorithm
#adam = Adam(0.1)
adam = Adam(0.00001)
model2.compile(loss='mse', optimizer=adam)
history = model2.fit(train_x, train_y, epochs=25, batch_size=batch_size)
# -
plt.plot(history.history['loss'])
plt.xlabel("No. of Iterations")
plt.ylabel("J(Theta1 Theta0)/Cost")
plt.show()
test_x=test[inputColumns]
test_y=test[outputColumns]
model2.evaluate(test_x, test_y)
predicted = model2.predict(test_x)
print("Mean absolute error =", round(sm.mean_absolute_error(test_y, predicted), 2))
print("Mean squared error =", round(sm.mean_squared_error(test_y, predicted), 2))
print("Median absolute error =", round(sm.median_absolute_error(test_y, predicted), 2))
print("Explain variance score =", round(sm.explained_variance_score(test_y, predicted), 2))
print("R2 score =", round(sm.r2_score(test_y, predicted), 2))
printSamplePredictedVsActual(model2, test)
# So far, the Dense Neural network performs best.
| EnergyUsagePrediction.model_def.asum.v1_0_7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Tutorial 2: Profiles
# ====================
#
# In this example, we'll create a `Grid` of Cartesian $(y,x)$ coordinates and pass it to the `light_profiles` module to
# create images on this `Grid` and the `mass_profiles` module to create deflection-angle maps on this grid.
# +
# %matplotlib inline
from pyprojroot import here
workspace_path = str(here())
# %cd $workspace_path
print(f"Working Directory has been set to `{workspace_path}`")
import autolens as al
import autolens.plot as aplt
# -
# Lets use the same `Grid` as the previous tutorial (if you skipped that tutorial, I recommend you go back to it!)
grid = al.Grid.uniform(shape_2d=(100, 100), pixel_scales=0.05, sub_size=2)
# Next, lets create a `LightProfile` using the `light_profiles` module, which in **PyAutoLens** is imported as `lp` for
# conciseness. we'll use an `EllipticalSersic` function, which is an analytic function often use to depict galaxies.
#
# (If you are unsure what the `elliptical_comps` are, I'll give a description of them at the end of the tutorial.)
sersic_light_profile = al.lp.EllipticalSersic(
centre=(0.0, 0.0),
elliptical_comps=(0.0, 0.111111),
intensity=1.0,
effective_radius=1.0,
sersic_index=2.5,
)
# We can print a `Profile` to confirm its parameters.
print(sersic_light_profile)
# We can pass a `Grid` to a `LightProfile` to compute its intensity at every `Grid` coordinate, using a `_from_grid`
# method
light_image = sersic_light_profile.image_from_grid(grid=grid)
# Much like the `Grid` objects in the previous tutorials, these functions return **PyAutoLens** `Array` objects which are
# accessible in both 2D and 1D.
print(light_image.shape_2d)
print(light_image.shape_1d)
print(light_image.in_2d[0, 0])
print(light_image.in_1d[0])
print(light_image.in_2d)
print(light_image.in_1d)
# The values computed (e.g. the image) are calculated on the sub-grid and the returned values are stored on the sub-grid,
# which in this case is a 200 x 200 grid.
print(light_image.sub_shape_2d)
print(light_image.sub_shape_1d)
print(light_image.in_2d[0, 0])
print(light_image[0])
# The benefit of storing all the values on the sub-grid, is that we can now use these values to bin-up the regular grid`s
# shape by taking the mean of each intensity value computed on the sub-grid. This ensures that aliasing effects due to
# computing intensities at only one pixel coordinate inside a full pixel does not degrade the image we create.
print("intensity of top-left `Grid` pixel:")
print(light_image.in_2d_binned[0, 0])
print(light_image.in_1d_binned[0])
# If you find these 2D and 1D `Array`'s confusing - I wouldn't worry about it. From here on, we'll pretty much just use
# these `Array`'s as they returned to us from functions and not think about if they should be in 2D or 1D. Nevertheless,
# its important that you understand **PyAutoLens** offers these 2D and 1D representations - as it`ll help us later when we
# cover fititng lens data!
#
# We can use a `Profile` `Plotter`.to plot this image.
aplt.LightProfile.image(light_profile=sersic_light_profile, grid=grid)
# To perform ray-tracing, we need to create a `MassProfile` from the `mass_profiles` module, which we import as `mp` for
# conciseness.
#
# A `MassProfile` is an analytic function that describes the distribution of mass in a galaxy, and therefore
# can be used to derive its surface-density, gravitational potential and most importantly, its deflection angles. For
# those unfamiliar with lensing, the deflection angles describe how light is bent by the `MassProfile` due to the
# curvature of space-time.
# +
sis_mass_profile = al.mp.SphericalIsothermal(centre=(0.0, 0.0), einstein_radius=1.6)
print(sis_mass_profile)
# -
# Just like above, we can pass a `Grid` to a `MassProfile` to compute its deflection angles. These are returned as the
# `Grid``s we used in the previous tutorials, so have full access to the 2D / 1D methods and mappings. And, just like
# the image above, they are computed on the sub-grid, so that we can bin up their values to compute more accurate
# deflection angles.
#
# (If you are new to gravitiational lensing, and are unclear on what a `deflection-angle` means or what it is used for,
# then I'll explain all in tutorial 4 of this chapter. For now, just look at the pretty pictures they make, and worry
# about what they mean in tutorial 4!).
# +
mass_profile_deflections = sis_mass_profile.deflections_from_grid(grid=grid)
print("deflection-angles of `Grid` sub-pixel 0:")
print(mass_profile_deflections.in_2d[0, 0])
print("deflection-angles of `Grid` sub-pixel 1:")
print(mass_profile_deflections.in_2d[0, 1])
print()
print("deflection-angles of `Grid` pixel 0:")
print(mass_profile_deflections.in_2d_binned[0, 1])
print()
print("deflection-angles of central `Grid` pixels:")
print(mass_profile_deflections.in_2d_binned[49, 49])
print(mass_profile_deflections.in_2d_binned[49, 50])
print(mass_profile_deflections.in_2d_binned[50, 49])
print(mass_profile_deflections.in_2d_binned[50, 50])
# -
# A `Profile` `Plotter`.can plot these deflection angles.
#
# (The black and red lines are the `critical curve` and `caustic` of the `MassProfile`. we'll cover what these are in
# a later tutorial.)
aplt.MassProfile.deflections_y(mass_profile=sis_mass_profile, grid=grid)
aplt.MassProfile.deflections_x(mass_profile=sis_mass_profile, grid=grid)
# `MassProfile`'s have a range of other properties that are used for lensing calculations, a couple of which we've plotted
# images of below:
#
# - Convergence: The surface mass density of the `MassProfile` in dimensionless units which are convenient for
# lensing calcuations.
# - Potential: The gravitational of the `MassProfile` again in convenient dimensionless units.
# - Magnification: Describes how much brighter each image-pixel appears due to focusing of light rays by the `MassProfile`.
#
# Extracting `Array`'s of these quantities from **PyAutoLens** is exactly the same as for the image and deflection angles above.
# +
mass_profile_convergence = sis_mass_profile.convergence_from_grid(grid=grid)
mass_profile_potential = sis_mass_profile.potential_from_grid(grid=grid)
mass_profile_magnification = sis_mass_profile.magnification_from_grid(grid=grid)
# -
# Plotting them is equally straight forward.
# +
aplt.MassProfile.convergence(mass_profile=sis_mass_profile, grid=grid)
aplt.MassProfile.potential(mass_profile=sis_mass_profile, grid=grid)
aplt.MassProfile.magnification(mass_profile=sis_mass_profile, grid=grid)
# -
# Congratulations, you`ve completed your second **PyAutoLens** tutorial! Before moving on to the next one, experiment with
# **PyAutoLens** by doing the following:
#
# 1) Change the `LightProfile`'s effective radius and Sersic index - how does the image's appearance change?
# 2) Change the `MassProfile`'s einstein radius - what happens to the deflection angles, potential and convergence?
# 3) Experiment with different `LightProfile`'s and `MassProfile`'s in the light_profiles and mass_profiles modules.
# In particular, use the `EllipticalIsothermal` `Profile`.to introduce ellipticity into a `MassProfile`.
# ___Elliptical Components___
#
# The `elliptical_comps` describe the ellipticity of the geometry of the light and mass profiles. You may be more
# familiar with a coordinate system where the ellipse is defined in terms of:
#
# - axis_ratio = semi-major axis / semi-minor axis = b/a
# - position angle phi, where phi is in degrees.
#
# We can use the **PyAutoLens** `convert` module to determine the elliptical components from the axis-ratio and phi,
# noting that the position angle phi is defined counter-clockwise from the positive x-axis.
# +
elliptical_comps = al.convert.elliptical_comps_from(axis_ratio=0.5, phi=45.0)
print(elliptical_comps)
# -
# The elliptical components are related to the axis-ratio and position angle phi as follows:
#
# fac = (1 - axis_ratio) / (1 + axis_ratio)
#
# elliptical_comp[0] = elliptical_comp_y = fac * np.sin(2 * phi)
# elliptical_comp[1] = elliptical_comp_x = fac * np.cos(2 * phi)
#
# The reason we use the elliptical components, instead of the axis-ratio and phi, to define a `Profile` geometry is that it
# improves the lens modeling process. What is lens modeling? You'll find out in chapter 2!
| howtolens/chapter_1_introduction/tutorial_2_profiles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import mglearn
import matplotlib.pyplot as plt
# %matplotlib inline
# -
ram_prices = pd.read_csv(os.path.join(mglearn.datasets.DATA_PATH, "ram_price.csv"))
ram_prices.head()
ram_prices.describe()
plt.semilogy(ram_prices.date, ram_prices.price)
plt.xlabel('Year')
plt.ylabel('Price in $/Mbyte')
from sklearn.tree import DecisionTreeRegressor
from sklearn.linear_model import LinearRegression
# +
# use the data to forecast prices after the year 2000
data_train = ram_prices[ram_prices.date < 2000]
data_test = ram_prices[ram_prices.date >= 2000]
# +
# predict prices based on date
X_train = data_train.date[:, np.newaxis]
X_test = data_test.date[:, np.newaxis]
# we use a log transform to get a simpler relationship of data to target
y_train = np.log(data_train.price)
y_test = np.log(data_test.price)
# -
tree = DecisionTreeRegressor()
tree.fit(X_train, y_train)
linear_reg = LinearRegression()
linear_reg.fit(X_train, y_train)
# +
# predit on ALL data
X_all = ram_prices.date[:, np.newaxis]
# -
X_all
pred_tree = tree.predict(X_all)
pred_tree
pred_lr = linear_reg.predict(X_all)
pred_lr
# +
# undo log-transform
price_tree = np.exp(pred_tree)
price_tree
# -
price_lr = np.exp(pred_lr)
price_lr
plt.semilogy(data_train.date, data_train.price, label='Training data')
plt.semilogy(data_test.date, data_test.price, label='Test data')
plt.semilogy(ram_prices.date, price_tree, label='Tree prediction')
plt.semilogy(ram_prices.date, price_lr, label='Linear prediction')
plt.legend()
| Coding_exercices/Decision Tree Regressor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shor's Algorithm
#
# Prerequisites: Quantum Fourier Transformation, Quantum Phase Estimation
#
# Shor's algorithm is an algorithm used to factor integers in polynomial time. To do so, we first find a solution for the problem of period finding for $a^x\ mod\ N$. If we can do so, we can factor integers in polynomial time too, as it is possible to convert the factoring problem to the period finding problem in polyomial time.
#
# # Period Finding.
# ### Problem
# Find the period of the function : $a^x\ mod\ N$
# Where $a,\ N \in \mathbb{N},\ a\ <\ N,\ gcd(a,\ N)\ =\ 1$
# The period of the function, $r$, is defined as the smallest non-zero integer such that
# $a^r\ mod\ N\ =\ 1$
#
#
# For example, for a = 3, and N = 35, the period r = 12.
#
# ### Solution
# Shor's solution used quantum phase estimation on the unitary operator : $U|y\rangle \equiv |ay \bmod N \rangle$
# To see how this helps us, lets say we start in state |1⟩. Now suppose that each successive application of U will multiply the state of our register by a (mod N), then after r applications we will arrive at state |1⟩ again. Example with a = 3, and N = 35.
# $\begin{aligned}
# U|1\rangle &= |3\rangle & \\
# U^2|1\rangle &= |9\rangle \\
# U^3|1\rangle &= |27\rangle \\
# & \vdots \\
# U^{(r-1)}|1\rangle &= |12\rangle \\
# U^r|1\rangle &= |1\rangle
# \end{aligned}$
#
# So a superposition of the states in this cycle ( |u0⟩ ) would be an eigenstate of U :
#
# $|u_0\rangle = \tfrac{1}{\sqrt{r}}\sum_{k=0}^{r-1}{|a^k \bmod N\rangle}$
#
# For a = 3, N = 35, it would be:
# $\begin{aligned}
# |u_0\rangle &= \tfrac{1}{\sqrt{12}}(|1\rangle + |3\rangle + |9\rangle \dots + |4\rangle + |12\rangle) \\[10pt]
# U|u_0\rangle &= \tfrac{1}{\sqrt{12}}(U|1\rangle + U|3\rangle + U|9\rangle \dots + U|4\rangle + U|12\rangle) \\[10pt]
# &= \tfrac{1}{\sqrt{12}}(|3\rangle + |9\rangle + |27\rangle \dots + |12\rangle + |1\rangle) \\[10pt]
# &= |u_0\rangle
# \end{aligned}$
#
# The eigenstate has an eigen value of 1, but this is not that useful. A more useful case is when the phase of the kth state is proportional to k:
# $\begin{aligned}
# |u_1\rangle &= \tfrac{1}{\sqrt{r}}\sum_{k=0}^{r-1}{e^{-\tfrac{2\pi i k}{r}}|a^k \bmod N\rangle}\\[10pt]
# U|u_1\rangle &= e^{\tfrac{2\pi i}{r}}|u_1\rangle
# \end{aligned}$
#
# For a = 3, N = 35, it would be:
# $\begin{aligned}
# |u_1\rangle &= \tfrac{1}{\sqrt{12}}(|1\rangle + e^{-\tfrac{2\pi i}{12}}|3\rangle + e^{-\tfrac{4\pi i}{12}}|9\rangle \dots + e^{-\tfrac{20\pi i}{12}}|4\rangle + e^{-\tfrac{22\pi i}{12}}|12\rangle) \\[10pt]
# U|u_1\rangle &= \tfrac{1}{\sqrt{12}}(|3\rangle + e^{-\tfrac{2\pi i}{12}}|9\rangle + e^{-\tfrac{4\pi i}{12}}|27\rangle \dots + e^{-\tfrac{20\pi i}{12}}|12\rangle + e^{-\tfrac{22\pi i}{12}}|1\rangle) \\[10pt]
# U|u_1\rangle &= e^{\tfrac{2\pi i}{12}}\cdot\tfrac{1}{\sqrt{12}}(e^{\tfrac{-2\pi i}{12}}|3\rangle + e^{-\tfrac{4\pi i}{12}}|9\rangle + e^{-\tfrac{6\pi i}{12}}|27\rangle \dots + e^{-\tfrac{22\pi i}{12}}|12\rangle + e^{-\tfrac{24\pi i}{12}}|1\rangle) \\[10pt]
# U|u_1\rangle &= e^{\tfrac{2\pi i}{12}}|u_1\rangle
# \end{aligned}$
#
# This is extremely useful as the eigenvalue contains r. The r has to be included to make sure the phase differences between the r computational basis states are equal. To generalise it further, we can multiply an integer s to this phase difference, and it will still show up in our eigenvalue:
# $\begin{aligned}
# |u_s\rangle &= \tfrac{1}{\sqrt{r}}\sum_{k=0}^{r-1}{e^{-\tfrac{2\pi i s k}{r}}|a^k \bmod N\rangle}\\[10pt]
# U|u_s\rangle &= e^{\tfrac{2\pi i s}{r}}|u_s\rangle
# \end{aligned}$
#
# Example for a = 3, N = 35:
# $\begin{aligned}
# |u_s\rangle &= \tfrac{1}{\sqrt{12}}(|1\rangle + e^{-\tfrac{2\pi i s}{12}}|3\rangle + e^{-\tfrac{4\pi i s}{12}}|9\rangle \dots + e^{-\tfrac{20\pi i s}{12}}|4\rangle + e^{-\tfrac{22\pi i s}{12}}|12\rangle) \\[10pt]
# U|u_s\rangle &= \tfrac{1}{\sqrt{12}}(|3\rangle + e^{-\tfrac{2\pi i s}{12}}|9\rangle + e^{-\tfrac{4\pi i s}{12}}|27\rangle \dots + e^{-\tfrac{20\pi i s}{12}}|12\rangle + e^{-\tfrac{22\pi i s}{12}}|1\rangle) \\[10pt]
# U|u_s\rangle &= e^{\tfrac{2\pi i s}{12}}\cdot\tfrac{1}{\sqrt{12}}(e^{-\tfrac{2\pi i s}{12}}|3\rangle + e^{-\tfrac{4\pi i s}{12}}|9\rangle + e^{-\tfrac{6\pi i s}{12}}|27\rangle \dots + e^{-\tfrac{22\pi i s}{12}}|12\rangle + e^{-\tfrac{24\pi i s}{12}}|1\rangle) \\[10pt]
# U|u_s\rangle &= e^{\tfrac{2\pi i s}{12}}|u_s\rangle
# \end{aligned}$
#
# We now have a unique eigenstate for each integer value of s where $0 \leq s \leq r-1$. If we sum up all these eigenstates, the different phases cancel out all computational bias states except $|1\rangle$:
# $\tfrac{1}{\sqrt{r}}\sum_{s=0}^{r-1} |u_s\rangle = |1\rangle$
#
# Example with a = 7 and N = 15 (smaller r, r = 4):
# $\begin{aligned}
# \tfrac{1}{2}(\quad|u_0\rangle &= \tfrac{1}{2}(|1\rangle \hphantom{e^{-\tfrac{2\pi i}{12}}}+ |7\rangle \hphantom{e^{-\tfrac{12\pi i}{12}}} + |4\rangle \hphantom{e^{-\tfrac{12\pi i}{12}}} + |13\rangle)\dots \\[10pt]
# + |u_1\rangle &= \tfrac{1}{2}(|1\rangle + e^{-\tfrac{2\pi i}{4}}|7\rangle + e^{-\tfrac{\hphantom{1}4\pi i}{4}}|4\rangle + e^{-\tfrac{\hphantom{1}6\pi i}{4}}|13\rangle)\dots \\[10pt]
# + |u_2\rangle &= \tfrac{1}{2}(|1\rangle + e^{-\tfrac{4\pi i}{4}}|7\rangle + e^{-\tfrac{\hphantom{1}8\pi i}{4}}|4\rangle + e^{-\tfrac{12\pi i}{4}}|13\rangle)\dots \\[10pt]
# + |u_3\rangle &= \tfrac{1}{2}(|1\rangle + e^{-\tfrac{6\pi i}{4}}|7\rangle + e^{-\tfrac{12\pi i}{4}}|4\rangle + e^{-\tfrac{18\pi i}{4}}|13\rangle)\quad) = |1\rangle \\[10pt]
# \end{aligned}$
#
# Since the computational basis state |1⟩ is a superposition of these eigenstates, which means if we do QPE on U using the state |1⟩ , we will measure a phase $\phi = \frac{s}{r}$, where s is a random integer between 0 and r−1 . We finally use the continued fractions algorithm on ϕ to find r .
#
# # Factoring:
#
# Now we have r , we might be able to use this to find a factor of N . Since:
#
# $a^r \bmod N = 1$
#
# then:
#
# $(a^r - 1) \bmod N = 0$
#
# which mean N must divide $a^r−1$ . And if r is also even, then we can write:
#
# $a^r -1 = (a^{r/2}-1)(a^{r/2}+1)$
#
# (if r is not even, we cannot go further and must try again with a different value for a ). There is then a high probability that the greatest common divisor of either $a^{r/2}−1$ , or $a^{r/2}+1$ is a factor of N
| Sheets/Shor's Algorithm/Shor's Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow-GPU
# language: python
# name: tf-gpu
# ---
# ## Facebook API Access
# +
# https://developers.facebook.com/tools/access_token/
ACCESS_TOKEN = '<KEY>'
# -
# ## Example 1. Making Graph API requests over HTTP
# +
import requests
import json
base_url = 'https://graph.facebook.com/me'
# Specify which fields to retrieve
fields = 'id,name,likes.limit(10){about}'
url = '{0}?fields={1}&access_token={2}'.format(base_url, fields, ACCESS_TOKEN)
# This API is HTTP-based and could be requested in the browser,
# with a command line utlity like curl, or using just about
# any programming language by making a request to the URL.
# Click the hyperlink that appears in your notebook output
# when you execute this code cell to see for yourself...
print(url)
# Interpret the response as JSON and convert back
# to Python data structures
content = requests.get(url).json()
# Pretty-print the JSON and display it
print(json.dumps(content, indent=1))
# -
# ## Example 2. Querying the Graph API with Python
# +
import facebook # pip install facebook-sdk
import json
# A helper function to pretty-print Python objects as JSON
def pp(o):
print(json.dumps(o, indent=1))
# Create a connection to the Graph API with your access token
g = facebook.GraphAPI(ACCESS_TOKEN, version='2.8')
# Execute a few example queries:
# Get my ID
pp(g.get_object('me'))
# -
# Get the connections to an ID
# Example connection names: 'feed', 'likes', 'groups', 'posts'
pp(g.get_connections(id='me', connection_name='likes'))
# Search for a location, may require approved app
pp(g.request("search", {'type': 'place', 'center': '40.749444, -73.968056', 'fields': 'name, location'}))
| Facebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import math
from random import sample, shuffle
import matplotlib.pyplot as plt
from sklearn.metrics.pairwise import pairwise_distances
from matplotlib.animation import FuncAnimation
from itertools import islice
# -
def visualize_data(X, file_name):
plt.figure(figsize=(6,6))
plt.scatter(X[:,0], X[:,1],c='r',s=10)
plt.xticks(np.arange(min(X[:,0]), max(X[:,0])+1, 1))
plt.yticks(np.arange(min(X[:,1]), max(X[:,1])+1, 2))
plt.plot(X[:,0], X[:,1], "b-")
plt.xlabel('$x_1$')
plt.ylabel('$x_2$')
plt.xticks(np.arange(min(X[:,0]), max(X[:,0])+1, 1))
plt.yticks(np.arange(min(X[:,1]), max(X[:,1])+1, 2))
plt.title("Travelling Salesman Problem")
plt.savefig(file_name)
plt.show()
# +
import argparse
import numpy as np
import math
from random import sample, shuffle
import matplotlib.pyplot as plt
from sklearn.metrics.pairwise import pairwise_distances
from itertools import islice
class GeneticAlgo():
def __init__(self, genes) -> None:
'''
Parameters
genes: cities
Returns
'''
self.n_genes = genes.shape[0]
self.distances = pairwise_distances(genes, metric='euclidean')
def init_population(self, n_chromosomes: int) -> None:
"""
Generate a population of n_chromosomes consisting of different permutations of n_genes.
Parameters
n_chromosomes: number of chromosomes/ population size
Returns
"""
self.n_chromosomes = n_chromosomes
population = []
n = 0
while n < self.n_chromosomes:
chromosome = np.random.permutation(self.n_genes)
if not any([np.array_equal(self.n_chromosomes, x) for x in population]):
population.append(chromosome)
n += 1
self.population = population
def fitness(self, chromosome: list) -> int:
"""
Sum distances between cities by the order of genes in THAT chromosome.
Parameters
chromosome: a permutation of genes
Returns
fitness: distance
"""
fitness = 0
for i in range(len(chromosome) - 1):
fitness += self.distances[chromosome[i], chromosome[i + 1]]
fitness += self.distances[chromosome[0], chromosome[len(chromosome) - 1]]
return fitness
def tournament_selection(self, selection_factor, k=2, p=0):
"""
Select k random chromosomes as parents, keep the best one (highest fitness) and iterate for n=selection_factor times.
If p != 0 and k == 2 then stochastic tournament selection is applied with probability p of getting the best chromosome.
Parameters
selection_factor: iteration times
k: the number of chromosomes to select
p: the probability to add the best one
Returns
"""
parents = []
if p != 0 and k == 2:
for i in range(selection_factor):
s = sample(self.population, k)
if np.random.rand() < p:
parents.append(min([[x, fitness(x)] for x in s], key=lambda x: x[1])[0])
else:
parents.append(max([[x, fitness(x)] for x in s], key=lambda x: x[1])[0])
else:
for i in range(selection_factor):
s = sample(self.population, k)
parents.append(min([[x, self.fitness(x)] for x in s], key=lambda x: x[1])[0])
return parents
def mutation(self, chromosome: list):
"""
Switch two random genes of the chromosome.
Parameters
chromosome: list of genes
Return
"""
mutated = np.array(chromosome)
gene1, gene2 = sample(range(len(chromosome)), 2)
mutated[gene1] = chromosome[gene2]
mutated[gene2] = chromosome[gene1]
return mutated
def _two_point_crossover(self, parent1, parent2):
"""
Produces offsprings of parent1 and parent2 by using two points crossover.
Parameters:
parent1: 1st chromsome
parent2: 2nd chromsome
Returns
"""
n_genes = len(parent1)
cross_point1, cross_point2 = np.random.choice(n_genes, 2)
sample(range(n_genes), 2)
if (cross_point1 > cross_point2):
tmp = cross_point1
cross_point1 = cross_point2
cross_point2 = tmp
child1 = np.array(parent1)
child2 = np.array(parent2)
j1 = cross_point1
j2 = cross_point2
for i in range(cross_point1, cross_point2):
while parent2[j1] not in parent1[cross_point1:cross_point2]:
j1 = (j1 + 1) % n_genes
child1[i] = parent2[j1]
j1 = (j1 + 1) % n_genes
while parent1[j2] not in parent2[cross_point1:cross_point2]:
j2 = (j2 + 1) % n_genes
child2[i] = parent1[j2]
j2 = (j2 + 1) % n_genes
return child1, child2
def _UOX_crossover(self, parent1, parent2):
'''
Parameter
parent1:
parent2:
Return
'''
mask1 = np.random.randint(2, size=len(parent1))
mask2 = 1 - mask1
child1 = parent1 * mask1 + parent2 * mask2
child2 = parent2 * mask1 + parent1 * mask2
return child1, child2
def crossover(self, parent1, parent2, mode):
'''
Parameter
parent1: 1st chromsome
parent2: 2nd chromsome
mode: type of crossover
Return
'''
if mode == 'UOX':
return self._UOX_crossover(np.array(parent1), np.array(parent2))
elif mode == 'two_point':
return self._two_point_crossover(np.array(parent1), np.array(parent2))
def create_offspring(self, parents, mode, p_crossover=1, p_mutation=0.1):
"""
Apply crossover and mutation to the parents.
Parameters
parents: all the chromsomes to reproduce children
mode: crossover method
p_crossover: the probability to have crossover.
p_mutation: the probability to have mutations.
Return
"""
offsprings = []
for i in range(len(parents)):
if np.random.rand() <= p_crossover:
p1, p2 = sample(parents, 2)
c1, c2 = self.crossover(p1, p2, mode)
if (not any([np.array_equal(c1, x) for x in parents]) and
not any([np.array_equal(c1, x) for x in offsprings])):
offsprings.append(c1)
if (not any([np.array_equal(c2, x) for x in parents]) and
not any([np.array_equal(c2, x) for x in offsprings])):
offsprings.append(c2)
for parent in parents:
if np.random.rand() <= p_mutation:
c = self.mutation(parent)
if (not any([np.array_equal(c, parent) for parent in parents]) and
not any([np.array_equal(c, x) for x in offsprings])):
offsprings.append(c)
return offsprings
def elitism_replacement(self, offspring, n_elite):
"""
Keep the n_elite best parents and substitute the others with the best offsprings
without changing the number of chromosomes.
Parameters
offspring: all the offspring chromsomes
n_elite: the number of parents to keep
Return
"""
self.population.sort(key=lambda x: self.fitness(x))
new_population = self.population[:n_elite]
offspring.sort(key=lambda x: self.fitness(x))
new_population.extend(offspring[:(len(self.population) - n_elite)])
self.population = new_population
# +
from easydict import EasyDict
cfg = EasyDict({
'crossover':'two_point',
'elite':0.1,
'file':'ulysses22.tsp',
'n_chromosomes':200,
'n_iter':400,
'p_crossover':0.6,
'p_mutation':0.2,
'selection_factor':90})
print(cfg)
if cfg.file == 'eil51.tsp':
start_line = 6
with open('eil51.tsp') as f:
lines = []
for line in islice(f, start_line, 57):
_, x1, x2 = line.split()
lines.append([x1, x2])
cities = np.asarray(lines, dtype=np.float32)
elif cfg.file == 'ulysses22.tsp':
start_line = 7
with open('ulysses22.tsp') as f:
lines = []
for line in islice(f, start_line, 29):
_, x1, x2 = line.split()
lines.append([x1, x2])
cities = np.asarray(lines, dtype=np.float32)
ga = GeneticAlgo(genes=cities)
ga.init_population(n_chromosomes=cfg.n_chromosomes)
plot_data = []
for i in np.arange(cfg.selection_factor):
parents = ga.tournament_selection(selection_factor=cfg.selection_factor)
offspring = ga.create_offspring(parents=parents, mode=cfg.crossover, p_crossover=1, p_mutation=0.1)
ga.elitism_replacement(offspring=offspring, n_elite= int(cfg.n_chromosomes*cfg.elite))
best_chromosome = min(ga.population, key=lambda x: ga.fitness(x))
plot_data.append([i,ga.fitness(best_chromosome)])
result={}
best_chromosomes = min(ga.population, key=lambda x: ga.fitness(x))
fitness = ga.fitness(best_chromosomes)
result = {'best_chromosome': best_chromosomes,
'fitness': fitness}
print(result)
# -
plot = np.array(plot_data)
def visualize_(X):
plt.plot(X[:,0],X[:,1],c='b')
plt.xlabel('$iteration$')
plt.ylabel('$fitness$')
plt.figure(figsize=(4,4))
plt.show()
visualize_(plot)
visualize_data(cities[best_chromosomes], 'best.png')
| Genatic Algorithm/TSP.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # Перечислимые типы (enums)
// ## 1. Базовые возможности
enum Color
{
White, // 0
Red, // 1
Green, // 2
Blue, // 3
Orange, // 4
}
// +
Color white = Color.White;
Console.WriteLine(white); // White
Color red = (Color)1; // Так можно приводить к типу перечисления
Console.WriteLine(red); // Red
Color unknown = (Color)42; // Нет ошибки!
Console.WriteLine(unknown); // 42
// -
Color green = Enum.Parse<Color>("Green");
green.ToString()
Enum.TryParse<Color>("Blue", out Color blue);
blue.ToString()
// Посмотрим, какими типами можно задавать перечисления
enum Dummy : object {}
// ## 2. Приведение перечислимых типов
enum Fruit
{
Melon, // 0
Tomato, // 1
Apple, // 2
Blueberry, // 3
Orange, // 4
}
Fruit orange = Color.Orange; // Безопасность типов -> ошибка
Fruit tomato = (Fruit)Color.Red; // А вот так уже можно
Console.WriteLine(tomato);
Color unknownColor = (Color)42;
Fruit unknownFruit = (Fruit)unknownColor;
Console.WriteLine(unknownFruit);
// +
// Любой enum имеет следующую цепочку наследования: MyEnum <- System.Enum <- System.ValueType <- System.Object
Enum enumEnum = Color.Blue;
ValueType enumValueType = Color.Blue;
object enumObj = Color.Blue; // BOXING
Console.WriteLine($"{enumEnum}, {enumValueType}, {enumObj}");
// -
// ## 3. Использование одного целочисленного значения для нескольких enum значений
public enum Subject
{
Programming = 0,
DiscreteMath = 1,
Algebra = 2,
Calculus = 3,
Economics = 4,
MostDifficultSubject = Algebra,
MostUsefulSubject = Programming,
// MostHatefulSubject = Programming
}
// +
Console.WriteLine(Subject.Programming);
Console.WriteLine(Subject.MostUsefulSubject);
Console.WriteLine((Subject)0);
Console.WriteLine(Subject.Programming == Subject.MostUsefulSubject)
// +
Console.WriteLine(Subject.Algebra);
Console.WriteLine(Subject.MostDifficultSubject);
Console.WriteLine((Subject)2);
Console.WriteLine(Subject.Algebra == Subject.MostDifficultSubject)
// -
// ## 4. Рефлексия перечислимых типов
// Статический метод Enum.GetUnderlyingType возвращает целочисленный тип для енама
Enum.GetUnderlyingType(typeof(Subject))
Enum.GetUnderlyingType(typeof(Subject))
// В типе System.Type также есть метод GetEnumUnderlyingType
typeof(Subject).GetEnumUnderlyingType()
// Который работает только с объектами-типами енамов
typeof(short).GetEnumUnderlyingType()
// Можно получить все значения енама c помощью Enum.GetValues(Type)
var enumValues = Enum.GetValues(typeof(Subject)); // Аналог: typeof(Subject).GetEnumValues();
foreach(var value in enumValues){
Console.WriteLine(value);
}
Enum.GetNames(typeof(Subject)) // Аналог: typeof(Subject).GetEnumNames()
// Проверка, есть ли в енаме соответствующее значение.
Enum.IsDefined(typeof(Subject), 3)
Enum.IsDefined(typeof(Subject), 42)
// ## 5. Битовые флаги
[Flags]
enum FilePermission : byte
{
None = 0b00000000,
Read = 0b00000001,
Write = 0b00000010,
Execute = 0b00000100,
Rename = 0b00001000,
Move = 0b00010000,
Delete = 0b00100000,
User = Read | Execute,
ReadWrite = Read | Write,
Admin = Read | Write | Execute | Rename | Move | Delete
}
// [Про FlagsAttribute](https://docs.microsoft.com/ru-ru/dotnet/api/system.flagsattribute?view=net-5.0)
FilePermission permission = FilePermission.User;
permission.HasFlag(FilePermission.Read)
// Пример использования:
// ```
// void RenameFile(File file, User user)
// {
// if (!user.Permission.HasFlag(FilePermission.Rename)) {
// throw new SomeException("you can't.")
// }
// ...
// }
// ```
for (int i = 0; i <= 16; ++i) {
FilePermission fp = (FilePermission)i;
Console.WriteLine(fp.ToString("G"));
}
// Пример из стандартной библиотеки: System.AttributeTargets
[Flags, Serializable]
public enum AttributeTargets {
Assembly = 0x0001,
Module = 0x0002,
Class = 0x0004,
Struct = 0x0008,
Enum = 0x0010,
Constructor = 0x0020,
Method = 0x0040,
Property = 0x0080,
Field = 0x0100,
Event = 0x0200,
Interface = 0x0400,
Parameter = 0x0800,
Delegate = 0x1000,
ReturnValue = 0x2000,
GenericParameter = 0x4000,
All = Assembly | Module | Class | Struct | Enum |
Constructor | Method | Property | Field | Event |
Interface | Parameter | Delegate | ReturnValue |
GenericParameter
}
// ## 6. Методы расширения для enum
// Перечислениям можно "добавлять функциональность" с помощью методов расширения
//public static class EnumExtentions
//{
public static int GetMark(this Subject subject)
{
return subject switch
{
Subject.Programming => 8,
Subject.DiscreteMath => 10,
Subject.Algebra => 5,
Subject.Calculus => 7,
Subject.Economics => 6,
_ => 0,
};
}
//}
Subject prog = Subject.Programming;
prog.GetMark()
| Materials/Enums.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Save and Restore
# In this post we are going to talk about how to save the parameters into the disk and restore the saved parameters from the disk. The savable/restorable paramters of the network are __Variables__ (i.e. weights and biases).
# ## TLDR:
#
# To save and restore your variables, all you need to do is to call the `tf.train.Saver()` at the end of you graph.
#
# ```python
# # create the graph
# X = tf.placeholder(..)
# Y = tf.placeholder(..)
# w = tf.get_variable(..)
# b = tf.get_variable(..)
# ...
# loss = tf.losses.mean_squared_error(..)
# optimizer = tf.train.AdamOptimizer(..).minimize(loss)
# ...
#
# saver = tf.train.Saver()
# ```
#
#
# __In the train mode__, in the session we will initialize the variables and run our network. At the end of training, we will save the variables using `saver.save()`:
#
# ```python
# # TRAIN
# with tf.Session() as sess:
# sess.run(tf.global_variables_initializer())
# # train our model
# for step in range(steps):
# sess.run(optimizer)
# ...
# saved_path = saver.save(sess, './my-model', global_step=step)
# ```
#
# This will create 3 files (`data`, `index`, `meta`) with a suffix of the step you saved your model.
#
# __In the test mode__, in the session we will restore the variables using `saver.restore()` and validate or test our model.
#
# ```python
# # TEST
# with tf.Session() as sess:
# saver.restore(sess, './my-model')
# ...
# ```
#
# ## 0. Import the required libraries:
#
# We will start with importing the required Python libraries.
#imports
import tensorflow as tf
import os
# ## 1. Save and Restore Two Variables:
# ### 1.1 Save:
# We will start with saving and restoring two variables in TensorFlow. We will create a graph with two variables. Let's create two variables `a = [3 3]` and `b = [5 5 5]`:
# create variables a and b
a = tf.get_variable("A", initializer=tf.constant(3, shape=[2]))
b = tf.get_variable("B", initializer=tf.constant(5, shape=[3]))
# Notice the __lower__case letter as python name and __UPPER__case letter as TensorFlow name. It will be important when we want to import the graph in restoring the data.
# __Recall from the [Tensor Types Tutorial](https://github.com/easy-tensorflow/easy-tensorflow/blob/master/1_TensorFlow_Basics/Tutorials/2_Tensor_Types.ipynb):__ Variables need to be initialized before being used. To do so, we have to invoke a __variable initializer operation__ and run the operation on the session. This is the easiest way to initialize variables which initializes all variables at once.
# initialize all of the variables
init_op = tf.global_variables_initializer()
# Now, on the session, we can initialize the variables and run the to see the values:
# run the session
with tf.Session() as sess:
# initialize all of the variables in the session
sess.run(init_op)
# run the session to get the value of the variable
a_out, b_out = sess.run([a, b])
print('a = ', a_out)
print('b = ', b_out)
# __Important Note:__ All of the variables exist in the scope of the session. So, after the session is closed, we will loose the variable.
#
# In order to save the variable, we will call the saver function using `tf.train.Saver()` in our graph. This function will find all the variables in the graph. We can see the list of all variables in `_var_list`. Let's create a `saver` object and take a look at the `_var_list` in the object:
# create saver object
saver = tf.train.Saver()
for i, var in enumerate(saver._var_list):
print('Var {}: {}'.format(i, var))
# So, our graph consists of two variables that listed above.
#
# __Important Note__: Notice the `:0` at the end of the variable name. For more about tensor naming check [here](https://stackoverflow.com/questions/36150834/how-does-tensorflow-name-tensors).
#
# Now that the saver object is created in the graph, in the session, we can call the `saver.save()` function to save the variables in the disk. We have to pass the created session (`sess`) and the path to the file that we want to save the variables:
# run the session
with tf.Session() as sess:
# initialize all of the variables in the session
sess.run(init_op)
# save the variable in the disk
saved_path = saver.save(sess, './saved_variable')
print('model saved in {}'.format(saved_path))
# If you check your working directory, you will notice that 3 new files have been created with the name `saved_variable` in them.
for file in os.listdir('.'):
if 'saved_variable' in file:
print(file)
# __.data:__ Contains variable values
#
# __.meta:__ Contains graph structure
#
# __.index:__ Identifies checkpoints
# ### 1.2. Restore:
# Now that all the things that you need is saved in the disk, you can load your saved variables in the session using `saver.restore()`:
# run the session
with tf.Session() as sess:
# restore the saved vairable
saver.restore(sess, './saved_variable')
# print the loaded variable
a_out, b_out = sess.run([a, b])
print('a = ', a_out)
print('b = ', b_out)
# Notice that this time we did not initialize the variables in our session. Instead, we restored them from the disk.
# __Important Note:__ In order to restore the parameters, the graph should be defined. Since we defined the graph in top, we didn't have a problem restoring the parameters. But what happens if we have not loaded the graph?
# delete the current graph
tf.reset_default_graph()
try:
with tf.Session() as sess:
# restore the saved vairable
saver.restore(sess, './saved_variable')
# print the loaded variable
a_out, b_out = sess.run([a, b])
print('a = ', a_out)
print('b = ', b_out)
except Exception as e:
print(str(e))
# We can define the graph in two ways.
# #### 1.2.1. Define the graph from scratch and then run the session:
# This way is simple if you have your graph. So, what you can do is to create the graph and then restore your variables:
# +
# delete the current graph
tf.reset_default_graph()
# create a new graph
# create variables a and b
a = tf.get_variable("A", initializer=tf.constant(3, shape=[2]))
b = tf.get_variable("B", initializer=tf.constant(5, shape=[3]))
# initialize all of the variables
init_op = tf.global_variables_initializer()
# create saver object
saver = tf.train.Saver()
# run the session
with tf.Session() as sess:
# restore the saved vairable
saver.restore(sess, './saved_variable')
# print the loaded variable
a_out, b_out = sess.run([a, b])
print('a = ', a_out)
print('b = ', b_out)
# -
# Keep in mind that the graph should be exactly like the one that you saved. ButwWhat if we do not know the exact graph and we are using someone else's pre-trained model?
# #### 1.2.2. Restore the graph from `.meta` file.
#
# When we save the variables, it creates a `.meta` file. This file contains the graph structure. Therefore, we can import the meta graph using `tf.train.import_meta_graph()` and restore the values of the graph. Let's import the graph and see all tensors in the graph:
# +
# delete the current graph
tf.reset_default_graph()
# import the graph from the file
imported_graph = tf.train.import_meta_graph('saved_variable.meta')
# list all the tensors in the graph
for tensor in tf.get_default_graph().get_operations():
print (tensor.name)
# -
# If you recall from section 1.1, we defined the python names with __lower__case letters and in TensorFlow names with __UPPER__case letters. You can see that what we have here are the __UPPER__case letter variables. It means that `tf.train.Saver()` saves the variables with the TensorFlow name. Now that we have the imported graph, and we know that we are interested in `A` and `B` tensors, we can restore the parameters:
# run the session
with tf.Session() as sess:
# restore the saved vairable
imported_graph.restore(sess, './saved_variable')
# print the loaded variable
a_out, b_out = sess.run(['A:0','B:0'])
print('a = ', a_out)
print('b = ', b_out)
# __Important Note:__ Notice that in `sess.run()` we provided the TensorFlow name of the tensors `'A:0'` and `'B:0'` instead of `a` and `b`.
# ## 2. Save and Restore Variables of a Sample Linear Model:
# Now that we have learnt how to save and restore parameters, we can write a simple model and try to save and restore the __weights__ and __biases__ in this network.
#
# We will build a simple linear model. If you do not know about the linear model, check our [Linear Classifier Tutorial](https://github.com/easy-tensorflow/easy-tensorflow/tree/master/2_Linear_Classifier).
# +
# delete the current graph
tf.reset_default_graph()
# Data Dimensions
img_h = img_w = 28 # MNIST images are 28x28
img_size_flat = img_h * img_w # 28x28=784, the total number of pixels
n_classes = 10 # Number of classes, one class per digit
# Load MNIST data
from tensorflow.examples.tutorials.mnist import input_data
data = input_data.read_data_sets("MNIST/", one_hot=True)
# Hyper-parameters
learning_rate = 0.001 # The optimization initial learning rate
batch_size = 100 # Training batch size
num_steps = 1000 # Total number of training steps
# Placeholders for inputs (x), outputs(y)
x = tf.placeholder(tf.float32, shape=[None, img_size_flat], name='X')
y = tf.placeholder(tf.float32, shape=[None, n_classes], name='Y')
W = tf.get_variable('W',
dtype=tf.float32,
shape=[img_size_flat, n_classes],
initializer=tf.truncated_normal_initializer(stddev=0.01))
b = tf.get_variable('b',
dtype=tf.float32,
initializer=tf.constant(0., shape=[n_classes], dtype=tf.float32))
# Calculate the output logits as: output_logits = W*x + b
output_logits = tf.matmul(x, W) + b
# Convert logits to probabilities
y_pred = tf.nn.softmax(output_logits)
# Define the loss function, optimizer, and accuracy
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(labels=y, logits=output_logits), name='loss')
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate, name='Adam-op').minimize(loss)
correct_prediction = tf.equal(tf.argmax(output_logits, 1), tf.argmax(y, 1), name='correct_pred')
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32), name='accuracy')
# -
# At the end of graph, will call the `tf.train.Saver()` to save all the variables.
# create saver object
saver = tf.train.Saver()
# Now we can run the model and save the variables.
# run the session
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for i in range(num_steps):
# Get a batch of training examples and their corresponding labels.
x_batch, y_true_batch = data.train.next_batch(batch_size)
# Put the batch into a dict to be fed into the placeholders
feed_dict_train = {x: x_batch, y: y_true_batch}
sess.run(optimizer, feed_dict=feed_dict_train)
feed_dict_valid = {x: data.validation.images, y: data.validation.labels}
loss_test, acc_test = sess.run([loss, accuracy], feed_dict=feed_dict_valid)
print('---------------------------------------------------------')
print("Validation loss: {0:.2f}, Validation accuracy: {1:.01%}".format(loss_test, acc_test))
print('---------------------------------------------------------')
# save the variable in the disk
saved_path = saver.save(sess, './linear_model')
print('model saved in {}'.format(saved_path))
# We can check that the model is saved in `./linear_model`.
for file in os.listdir('.'):
if 'linear_model' in file:
print(file)
# Let's restore the model and pull out the trained variables. at this time, the garph still exists in the memory. So, we can restore it and evaluate the network on the test set:
# +
# Test the network after training
# run the session
with tf.Session() as sess:
# restore the saved vairable
saver.restore(sess, './linear_model')
# Accuracy
feed_dict_test = {x: data.test.images, y: data.test.labels}
loss_test, acc_test = sess.run([loss, accuracy], feed_dict=feed_dict_test)
print('---------------------------------------------------------')
print("Test loss: {0:.2f}, test accuracy: {1:.01%}".format(loss_test, acc_test))
print('---------------------------------------------------------')
print()
# print the loaded variable
weight, bias = sess.run(['W:0','b:0'])
print('W = ', weight)
print('b = ', bias)
# -
# Recall from __Section 1.2__, if we do not have the graph, we can restore the values of the graph using `tf.train.import_meta_graph()`:
# +
# delete the current graph
tf.reset_default_graph()
# import the graph from the file
imported_graph = tf.train.import_meta_graph('linear_model.meta')
# list all the tensors in the graph
for tensor in tf.get_default_graph().get_operations():
print (tensor.name)
# -
# Let's say that I am interested in `loss` and `accuracy` of my model. We can easily get the values of corresponding tensors, providing the correct placeholders:
# run the session
with tf.Session() as sess:
# restore the saved vairable
imported_graph.restore(sess, './linear_model')
# Accuracy
feed_dict_test = {'X:0': data.test.images, 'Y:0': data.test.labels}
loss_test, acc_test = sess.run(['loss:0', 'accuracy:0'], feed_dict=feed_dict_test)
print('---------------------------------------------------------')
print("Test loss: {0:.2f}, test accuracy: {1:.01%}".format(loss_test, acc_test))
print('---------------------------------------------------------')
print()
# Thanks for reading! If you have any question or doubt, feel free to leave a comment in our [website](http://easy-tensorflow.com/).
| 1_TensorFlow_Basics/Tutorials/4_Save_and_Restore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: advbox
# language: python
# name: advbox
# ---
# 介绍如何在pytorch环境下,使用FGSM算法攻击基于MNIST数据集预训练的CNN/MLP模型。运行该文件前,需要先运行指定文件生成对应的模型:
#
# cd tutorials
# python mnist_model_pytorch.py
#
# Jupyter notebook中使用Anaconda中的环境需要单独配置,默认情况下使用的是系统默认的Python环境,以使用advbox环境为例。
# 首先在默认系统环境下执行以下命令,安装ipykernel。
#
# conda install ipykernel
# conda install -n advbox ipykernel
#
# 在advbox环境下激活,这样启动后就可以在界面上看到advbox了。
#
# python -m ipykernel install --user --name advbox --display-name advbox
#
#调试开关
import logging
#logging.basicConfig(level=logging.INFO,format="%(filename)s[line:%(lineno)d] %(levelname)s %(message)s")
#logger=logging.getLogger(__name__)
import sys
import torch
import torchvision
from torchvision import datasets, transforms
from torch.autograd import Variable
import torch.utils.data.dataloader as Data
from advbox.adversary import Adversary
from advbox.attacks.gradient_method import FGSM
from advbox.models.pytorch import PytorchModel
from tutorials.mnist_model_pytorch import Net
# +
TOTAL_NUM = 500
pretrained_model="tutorials/mnist-pytorch/net.pth"
loss_func = torch.nn.CrossEntropyLoss()
#使用MNIST测试数据集 随机挑选TOTAL_NUM个
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('tutorials/mnist-pytorch/data', train=False, download=True, transform=transforms.Compose([
transforms.ToTensor(),
])),
batch_size=1, shuffle=True)
# Define what device we are using
logging.info("CUDA Available: {}".format(torch.cuda.is_available()))
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Initialize the network
model = Net().to(device)
# Load the pretrained model
model.load_state_dict(torch.load(pretrained_model, map_location='cpu'))
# Set the model in evaluation mode. In this case this is for the Dropout layers
model.eval()
# advbox demo
m = PytorchModel(
model, loss_func,(0, 1),
channel_axis=1)
#实例化FGSM
attack = FGSM(m)
#设置攻击步长为0.1
attack_config = {"epsilons": 0.1}
# use test data to generate adversarial examples
total_count = 0
fooling_count = 0
for i, data in enumerate(test_loader):
inputs, labels = data
inputs, labels=inputs.numpy(),labels.numpy()
total_count += 1
adversary = Adversary(inputs, labels[0])
# FGSM non-targeted attack
adversary = attack(adversary, **attack_config)
if adversary.is_successful():
fooling_count += 1
print(
'attack success, original_label=%d, adversarial_label=%d, count=%d'
% (labels, adversary.adversarial_label, total_count))
else:
print('attack failed, original_label=%d, count=%d' %
(labels, total_count))
if total_count >= TOTAL_NUM:
print(
"[TEST_DATASET]: fooling_count=%d, total_count=%d, fooling_rate=%f"
% (fooling_count, total_count,
float(fooling_count) / total_count))
break
print("fgsm attack done")
# -
| ebook/ebook_mnist_fgsm_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# # Old Faithful and Clustering
faithful = pd.read_csv("faithful.csv")
display(faithful.head())
display(faithful.describe())
# +
import seaborn as sns
plt.figure(figsize=(10,5))
plt.scatter(faithful["eruptions"], faithful["waiting"])
plt.xlabel("eruptions")
plt.ylabel("waiting")
plt.xlim(0,6)
plt.ylim(30,100)
plt.figure(figsize=(10,5))
sns.kdeplot(faithful["eruptions"], faithful["waiting"])
plt.scatter(faithful["eruptions"], faithful["waiting"])
plt.xlim(0,6)
plt.show(30,100)
# -
# There are two distinct modes to the data: one with eruption values (voulmes?) of 1 to 3 and low waiting times, and a second cluster with larger eruptions and longer waiting times. Notably, there are very few eruptions in the middle.
# ## Review: PCA
# First, we import data on different types of crime in each US state
USArrests = pd.read_csv("USArrests.csv")
USArrests['StateAbbrv'] = ["AL", "AK", "AZ", "AR", "CA", "CO", "CT","DE", "FL", "GA", "HI", "ID", "IL", "IN", "IA", "KS", "KY", "LA", "ME", "MD", "MA", "MI", "MN", "MS", "MO", "MT", "NE", "NV", "NH", "NJ","NM", "NY", "NC", "ND", "OH", "OK", "OR", "PA", "RI", "SC","SD", "TN", "TX", "UT", "VT", "VA", "WA", "WV", "WI", "WY"]
display(USArrests.head())
display(USArrests.describe())
# The data has more dimensinons than we can easily visualize, so we use PCA to condense it. As usual, we scale the data before applying PCA. (Note that we scale everything, rather than fitting on train and carrying that scaling to future data-- we won't be using a test set here, so it's correct to use all the data to scale).
# +
from sklearn import preprocessing
df = USArrests[['Murder','Assault','UrbanPop','Rape']]
scaled_df = pd.DataFrame(preprocessing.scale(df), index=USArrests['State'], columns = df.columns)
fitted_pca = PCA().fit(scaled_df)
USArrests_pca = fitted_pca.transform(scaled_df)
# -
# The biplot function plots the first two PCA components, and provides some helpful annotations
# +
def biplot(scaled_data, fitted_pca, original_dim_labels, point_labels):
pca_results = fitted_pca.transform(scaled_data)
pca1_scores = pca_results[:,0]
pca2_scores = pca_results[:,1]
# plot each point in 2D post-PCA space
plt.scatter(pca1_scores,pca2_scores)
# label each point
for i in range(len(pca1_scores)):
plt.text(pca1_scores[i],pca2_scores[i], point_labels[i])
#for each original dimension, plot what an increase of 1 in that dimension means in this space
for i in range(fitted_pca.components_.shape[1]):
raw_dims_delta_on_pca1 = fitted_pca.components_[0,i]
raw_dims_delta_on_pca2 = fitted_pca.components_[1,i]
plt.arrow(0, 0, raw_dims_delta_on_pca1, raw_dims_delta_on_pca2 ,color = 'r',alpha = 1)
plt.text(raw_dims_delta_on_pca1*1.1, raw_dims_delta_on_pca2*1.1, original_dim_labels[i], color = 'g', ha = 'center', va = 'center')
plt.figure(figsize=(8.5,8.5))
plt.xlim(-3.5,3.5)
plt.ylim(-3.5,3.5)
plt.xlabel("PC{}".format(1))
plt.ylabel("PC{}".format(2))
plt.grid()
biplot(scaled_df, fitted_pca,
original_dim_labels=scaled_df.columns,
point_labels=USArrests['State'])
# -
# The red arrows and green text give us a sense of direction. If any state had 'murder' increase by one (scaled) unit, it would move in the direction of the 'murder' line by that amount. An increase by one (scaled) unit of both 'murder' and 'Urban Pop' would apply both moves.
#
# We can also make inferrences about what combination of crimes and population puts California at its observed point.
# ## Extra: Variance Captured
# As usual, we want to know how what proportion of the variance each PC captures
# +
plt.figure(figsize=(11,8.5))
plt.plot(range(1,5),fitted_pca.explained_variance_ratio_,"-o")
plt.xlabel("Principal Component")
plt.ylabel("Proportion of Variance Explained")
plt.ylim(0,1)
plt.show()
print("Proportion of variance explained by each PC:")
print(fitted_pca.explained_variance_ratio_)
# -
# Even more usefully, we can plot how much of the total variation we'd capture by using N PCs. The PCA-2 plot above has 86.7% of the total variance.
# +
plt.figure(figsize=(11,8.5))
plt.plot(range(1,5),np.cumsum(fitted_pca.explained_variance_ratio_),"-o")
plt.xlabel("Principal Component")
plt.ylabel("Cumulative Proportion of Variance Explained")
plt.ylim(0,1.1)
plt.show()
print("Total variance capturted when using N PCA components:")
print(np.cumsum(fitted_pca.explained_variance_ratio_))
# -
# # Scaling and Distances
# Returning to the arrest/crime data, we again inspect the data and its PCA plot
np.random.seed(123)
arrests_sample = USArrests.sample(6)
arrests_sample
np.random.seed(123)
np.round(scaled_df.sample(6),2)
plt.figure(figsize=(10,5))
biplot(scaled_df, fitted_pca,
original_dim_labels=scaled_df.columns,
point_labels=USArrests['State'])
# ## Distances
# One of the key ideas in clustering is the distance or disimilarity between points. Euclidean distance is common, though one is free to define domain-specific measures of how similar/distant two observations are.
from scipy.spatial.distance import pdist
from scipy.spatial.distance import squareform
# The `pdist` function computes the distances between all pairs of data points (which can be quite expensive for large data). `squareform` turns the result into a numpy array (the raw format avoids storing redundant values)
#
# The distances between a handful of states are shown below. Hawaii and Indiana are relatively similar on these variables, while Maine and New Mexico are relatively different.
# +
dist_eucl = pdist(scaled_df,metric="euclidean")
distances = pd.DataFrame(squareform(dist_eucl), index=USArrests["State"].values, columns=USArrests["State"].values)
sample_distances = distances.loc[arrests_sample["State"], arrests_sample["State"]]
sample_distances
# -
# For visualization, we can make a heatmap of the sample state's distances
plt.figure(figsize=(11,8.5))
sns.heatmap(sample_distances,cmap="mako")
plt.show()
# We can likewise heatmap all the states.
# +
import seaborn as sns
plt.figure(figsize=(11,8.5))
sns.heatmap(distances)
plt.show()
# -
# # Kmeans
# Kmeans is a classical, workhorse clustering algorithm, and a common place to start. It assumes there are K centers and, starting from random guesses, algorithmically improves its guess about where the centers must be.
# +
from sklearn.cluster import KMeans
#random_state parameter sets seed for random number generation
arrests_km = KMeans(n_clusters=3,n_init=25,random_state=123).fit(scaled_df)
arrests_km.cluster_centers_
# -
# We can read off where the 3 cluster centers are. (The value 3 is chosen arbitratially- soon we'll see how to tell what number of clusters seems to work best)
pd.DataFrame(arrests_km.cluster_centers_,columns=['Murder','Assault','UrbanPop','Rape'])
# The .lables_ tell us which cluster each point was assigned to
scaled_df_cluster = scaled_df.copy()
scaled_df_cluster['Cluster'] = arrests_km.labels_
scaled_df_cluster.head()
# The mean of the points in each cluster is the cluster center found by K-means
scaled_df_cluster.groupby('Cluster').mean()
# ## Silhouette Plots
# Silhouette plots give rich information on the quality of a clustering
# +
from sklearn.metrics import silhouette_samples, silhouette_score
import matplotlib.cm as cm
#modified code from http://scikit-learn.org/stable/auto_examples/cluster/plot_kmeans_silhouette_analysis.html
def silplot(X, clusterer, pointlabels=None):
cluster_labels = clusterer.labels_
n_clusters = clusterer.n_clusters
# Create a subplot with 1 row and 2 columns
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.set_size_inches(11,8.5)
# The 1st subplot is the silhouette plot
# The silhouette coefficient can range from -1, 1 but in this example all
# lie within [-0.1, 1]
ax1.set_xlim([-0.1, 1])
# The (n_clusters+1)*10 is for inserting blank space between silhouette
# plots of individual clusters, to demarcate them clearly.
ax1.set_ylim([0, len(X) + (n_clusters + 1) * 10])
# The silhouette_score gives the average value for all the samples.
# This gives a perspective into the density and separation of the formed
# clusters
silhouette_avg = silhouette_score(X, cluster_labels)
print("For n_clusters = ", n_clusters,
", the average silhouette_score is ", silhouette_avg,".",sep="")
# Compute the silhouette scores for each sample
sample_silhouette_values = silhouette_samples(X, cluster_labels)
y_lower = 10
for i in range(0,n_clusters+1):
# Aggregate the silhouette scores for samples belonging to
# cluster i, and sort them
ith_cluster_silhouette_values = \
sample_silhouette_values[cluster_labels == i]
ith_cluster_silhouette_values.sort()
size_cluster_i = ith_cluster_silhouette_values.shape[0]
y_upper = y_lower + size_cluster_i
color = cm.nipy_spectral(float(i) / n_clusters)
ax1.fill_betweenx(np.arange(y_lower, y_upper),
0, ith_cluster_silhouette_values,
facecolor=color, edgecolor=color, alpha=0.7)
# Label the silhouette plots with their cluster numbers at the middle
ax1.text(-0.05, y_lower + 0.5 * size_cluster_i, str(i))
# Compute the new y_lower for next plot
y_lower = y_upper + 10 # 10 for the 0 samples
ax1.set_title("The silhouette plot for the various clusters.")
ax1.set_xlabel("The silhouette coefficient values")
ax1.set_ylabel("Cluster label")
# The vertical line for average silhouette score of all the values
ax1.axvline(x=silhouette_avg, color="red", linestyle="--")
ax1.set_yticks([]) # Clear the yaxis labels / ticks
ax1.set_xticks([-0.1, 0, 0.2, 0.4, 0.6, 0.8, 1])
# 2nd Plot showing the actual clusters formed
colors = cm.nipy_spectral(cluster_labels.astype(float) / n_clusters)
ax2.scatter(X[:, 0], X[:, 1], marker='.', s=200, lw=0, alpha=0.7,
c=colors, edgecolor='k')
xs = X[:, 0]
ys = X[:, 1]
if pointlabels is not None:
for i in range(len(xs)):
plt.text(xs[i],ys[i],pointlabels[i])
# Labeling the clusters
centers = clusterer.cluster_centers_
# Draw white circles at cluster centers
ax2.scatter(centers[:, 0], centers[:, 1], marker='o',
c="white", alpha=1, s=200, edgecolor='k')
for i, c in enumerate(centers):
ax2.scatter(c[0], c[1], marker='$%d$' % int(i), alpha=1,
s=50, edgecolor='k')
ax2.set_title("The visualization of the clustered data.")
ax2.set_xlabel("Feature space for the 1st feature")
ax2.set_ylabel("Feature space for the 2nd feature")
plt.suptitle(("Silhouette analysis for KMeans clustering on sample data "
"with n_clusters = %d" % n_clusters),
fontsize=14, fontweight='bold')
# +
fitted_km = KMeans(n_clusters=4,n_init=25,random_state=123).fit(scaled_df)
silplot(scaled_df.values, fitted_km)
# +
# Objects with negative silhouette
sil = silhouette_samples(scaled_df, fitted_km.labels_)
USArrests.loc[sil<=0,:]
# -
# ## Elbow plots
# +
wss = []
for i in range(1,11):
fitx = KMeans(n_clusters=i, init='random', n_init=5, random_state=109).fit(scaled_df)
wss.append(fitx.inertia_)
plt.figure(figsize=(11,8.5))
plt.plot(range(1,11), wss, 'bx-')
plt.xlabel('Number of clusters $k$')
plt.ylabel('Inertia')
plt.title('The Elbow Method showing the optimal $k$')
plt.show()
# -
# # Silhouette Score
# +
from sklearn.metrics import silhouette_score
scores = [0]
for i in range(2,11):
fitx = KMeans(n_clusters=i, init='random', n_init=5, random_state=109).fit(scaled_df)
score = silhouette_score(scaled_df, fitx.labels_)
scores.append(score)
plt.figure(figsize=(11,8.5))
plt.plot(range(1,11), np.array(scores), 'bx-')
plt.xlabel('Number of clusters $k$')
plt.ylabel('Average Silhouette')
plt.title('The Silhouette Method showing the optimal $k$')
plt.show()
# -
# # Gap Statistic
# +
from gap_statistic import OptimalK
from sklearn.datasets.samples_generator import make_blobs
gs_obj = OptimalK()
n_clusters = gs_obj(scaled_df.values, n_refs=50, cluster_array=np.arange(1, 15))
print('Optimal clusters: ', n_clusters)
# -
gs_obj.gap_df.head()
gs_obj.plot_results()
# # Hierarchical Clustering
# K-means is a very 'hard' clustering: points belong to exactly one cluster, no matter what. A hierarchical clustering creates a nesting of clusters as existing clusters are merged or split.
#
# Dendograms (literally: branch graphs) can show the pattern of splits/merges.
# +
import scipy.cluster.hierarchy as hac
from scipy.spatial.distance import pdist
plt.figure(figsize=(11,8.5))
dist_mat = pdist(scaled_df, metric="euclidean")
ward_data = hac.ward(dist_mat)
hac.dendrogram(ward_data, labels=USArrests["State"].values);
plt.show()
# -
# # DBSCAN
# DBSCAN is a more modern clustering approach that allows points to not be part of any cluster, and determines the number of clusters by itself.
# First, let's look at out data
multishapes = pd.read_csv("multishapes.csv")
ms = multishapes[['x','y']]
msplot = ms.plot.scatter(x='x',y='y',c='Black',title="Multishapes data",figsize=(11,8.5))
msplot.set_xlabel("X")
msplot.set_ylabel("Y")
plt.show()
# To the eye, there's a pretty clear structure to the data
# However, K-means struggles to find a good clustering
shape_km = KMeans(n_clusters=5,n_init=25,random_state=123).fit(ms)
plt.figure(figsize=(10,10))
plt.scatter(ms['x'],ms['y'], c=shape_km.labels_);
plt.scatter(shape_km.cluster_centers_[:,0],shape_km.cluster_centers_[:,1], c='r', marker='h', s=100);
#todo: labels? different markers?
# DB Scan uses a handful of parameters, including the number of neighbors a point must have to be considered 'core' (`min_samples`) and the distance within which neighbors must fall (`epsilon`). Most reasonable values of min_samples yeild the same results, but tuning epsilon is important.
#
# The function below implement's the authors suggestion for setting epsilon: look at the nearest-neighbor distances and find a level where they begin to grow rapidly.
# +
from sklearn.neighbors import NearestNeighbors
def plot_epsilon(df, min_samples):
fitted_neigbors = NearestNeighbors(n_neighbors=min_samples).fit(df)
distances, indices = fitted_neigbors.kneighbors(df)
dist_to_nth_nearest_neighbor = distances[:,-1]
plt.plot(np.sort(dist_to_nth_nearest_neighbor))
plt.xlabel("Index\n(sorted by increasing distances)")
plt.ylabel("{}-NN Distance (epsilon)".format(min_samples-1))
plt.tick_params(right=True, labelright=True)
# -
plot_epsilon(ms, 3)
# The major slope occurs around eps=0.15 when min_samples set to 3.
# +
from sklearn.cluster import DBSCAN
fitted_dbscan = DBSCAN(eps=0.15).fit(ms)
plt.figure(figsize=(10,10))
plt.scatter(ms['x'],ms['y'], c=fitted_dbscan.labels_);
# -
# We see good results with the suggested epsilon. A lower epsilon (0.12) won't quite merge all the clustersWe
# ## DBSCAN on crime data
#
# Returning to the crime data, let's tune epsilon and see what clusters are returned
plot_epsilon(scaled_df, 5)
# The optimal value is either around 1.67 or 1.4
fitted_dbscan = DBSCAN(eps=1.4).fit(scaled_df)
fitted_dbscan.labels_
# At this `epsilon` and `min_samples`, all but one state are included in cluster 0. The remaining point (Alaska) is not part of any cluster
| content/lectures/lecture04/notebook/cs109b_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# default_exp linkage
# -
# # Linkage analysis pipeline
# This pipeline is using paramlink2 to do linkage analysis. The R code is bridged to python through rpy2. It run linkage analysis from batch to batch. Its input is the intermediate result of seqlink.
# Next, I will make it to a sos pipeline. Run all the chromosomes in parallel.
# +
#export
import numpy as np
import pandas as pd
import pickle
from itertools import repeat
import numbers
#Import necessary packages
import rpy2.robjects as robjects
from rpy2.robjects.packages import importr
from rpy2.robjects import pandas2ri
base = importr('base')
base.options(expressions = 5e5)
#Must be activated
pandas2ri.activate()
paramlink2=importr('paramlink2')
pedprobr=importr('pedprobr')
pedtools = importr('pedtools')
import time
from concurrent.futures import ProcessPoolExecutor,ThreadPoolExecutor
# -
# ## Functions to deal with haplotypes
# +
#export
def get_allele(s):
a = s[1] if s[0].isupper() else s[0]
return 0 if a=='?' else int(a)
def name_haps(snps):
name = []
for i in snps:
name += [i+'_A0',i+'_A1']
return name
def get_fam_hap(haps,variants,vcf=None):
new_haps,new_iid = [],[]
iid = haps[:,1]
haps = haps[:,2:]
for i in range(0,haps.shape[0],2):
cur_iid=iid[i]
new_iid.append(cur_iid)
if vcf is None or vcf[cur_iid]:#have vcf
hap_a01 = []
for a0,a1 in zip(haps[i],haps[i+1]): #loop through variants
hap_a01 += [get_allele(a0),get_allele(a1)]
else:
hap_a01 = [0,0]*haps.shape[1] #set missing vcf to 0
new_haps.append(hap_a01)
new_haps = pd.DataFrame(new_haps)
new_haps.index = new_iid
new_haps.columns = name_haps(variants)
#remove variants with only 1 or 2 as alleles, return None
idx=[]
for i in range(0,new_haps.shape[1],2):
v=set(new_haps.iloc[:,i]).union(set(new_haps.iloc[:,i+1]))
if 1 not in v or 2 not in v:
idx.append(False)
else:
idx.append(True)
if sum(idx)==0:
return None
return new_haps.loc[:,np.repeat(np.array(idx),2)],idx
def get_fam_geno(haps,variants,vcf=None):
new_haps,new_iid = [],[]
iid = haps[:,1]
haps = haps[:,5:]
for i in range(haps.shape[0]):
cur_iid=iid[i]
new_iid.append(cur_iid)
if vcf is None or vcf[cur_iid]:#have vcf
hap_a01 = []
for a01 in haps[i]: #loop through variants
hap_a01 += [int(a) for a in a01]
else:
hap_a01 = [0,0]*haps.shape[1] #set missing vcf to 0
new_haps.append(hap_a01)
new_haps = pd.DataFrame(new_haps)
new_haps.index = new_iid
new_haps.columns = name_haps(variants)
#remove variants with only 1 or 2 as alleles, return None
idx=[]
for i in range(0,new_haps.shape[1],2):
v=set(new_haps.iloc[:,i]).union(set(new_haps.iloc[:,i+1]))
if 1 not in v or 2 not in v:
idx.append(False)
else:
idx.append(True)
if sum(idx)==0:
return None
return new_haps.loc[:,np.repeat(np.array(idx),2)],idx
# -
# ## All genes from haps to peds
# #### compare multithread and multiprocess
# +
#export
def format_haps_bunch(dhaps,fam,vcfs=None,cutoff=None,haplotype=True):
gene_variants = {}
gene_haps = {}
for g in dhaps.keys():
haps = dhaps[g]['predata']
with ProcessPoolExecutor(max_workers = 10) as executor:
if haplotype:
results = executor.map(get_fam_hap,[haps[k][2] for k in haps.keys()],[haps[k][0] for k in haps.keys()],[vcfs[k] if vcfs else None for k in haps.keys()])
else:
results = executor.map(get_fam_geno,[haps[k][2] for k in haps.keys()],[haps[k][0] for k in haps.keys()],[vcfs[k] if vcfs else None for k in haps.keys()])
for f,hap in zip(haps.keys(),results):
if hap is None: #remove only have 1 or 2 variants
continue
if f not in gene_variants.keys():
gene_variants[f] = {'genes':[],'variants':[],'freqs':[]}
gene_haps[f] = hap[0]
else:
gene_haps[f] = pd.concat([gene_haps[f],hap[0]],axis=1)
idx=hap[1] #False for variants only have 1 or 2.
gene_variants[f]['genes'] += [g]*sum(idx)
gene_variants[f]['variants'] += list(haps[f][0][idx])
gene_variants[f]['freqs'] += list(haps[f][1][idx])
for i,j in gene_variants.items():
j=pd.DataFrame(j)
if cutoff is not None:
frq_idx=np.array(j['freqs'])>cutoff
j=j.loc[frq_idx,:]
gene_haps[i]=gene_haps[i].loc[:,np.repeat(frq_idx,2)]
redup_idx = ~gene_haps[i].columns.duplicated()
gene_haps[i] = pd.concat([fam[i],gene_haps[i].iloc[:,redup_idx]],axis=1)
j['uniq'] = list(redup_idx[range(0,len(redup_idx),2)])
gene_variants[i] = j
return gene_variants,gene_haps
def calculate_ped_lod(ped,afreq=None,rho=0,model = "AD",chrom = "AUTOSOMAL",penetrances = [0.01,0.9,0.9],dfreq=0.001):
def _calculate_ped_lod(mped, aff, model,rho):
res = paramlink2.lod(mped, aff, model,rho)
try:
res = pd.DataFrame(res)[['MARKER','LOD']]
except:
res = pd.DataFrame([[ped.columns[6],res[0]]],columns=['MARKER','LOD'])
return res
aff=ped.iloc[:,5]
mped = pedtools.as_ped(ped.drop(ped.columns[5], axis=1),famid_col = 1,id_col = 2,fid_col = 3,mid_col = 4,sex_col = 5)
if afreq is not None:
mped = pedtools.setLocusAttributes(mped,locusAttributes=[base.list(afreq=base.c(1-i,i)) for i in afreq])
modAD = paramlink2.diseaseModel(model,chrom,pd.Series(penetrances),dfreq)
if isinstance(rho,numbers.Number):
res = _calculate_ped_lod(mped, aff = aff, model = modAD,rho=rho)
else:
res=None
for r in rho:
tmp = _calculate_ped_lod(mped, aff = aff, model = modAD,rho=r)
if res is None:
res=tmp
res.columns = ['MARKER','LOD'+str(round(r,2))]
else:
res['LOD'+str(round(r,2))]=tmp.LOD
res.index=list(res.MARKER)
res=res.iloc[:,1:]
return res
def parallel_lods(haps,afreqs=None,rho=0):
start = time.perf_counter()
if afreqs is None:
with ProcessPoolExecutor(max_workers = 10) as executor:
results = executor.map(calculate_ped_lod,haps.values(),repeat(rho))
else:
with ProcessPoolExecutor(max_workers = 10) as executor:
results = executor.map(calculate_ped_lod,haps.values(),afreqs,repeat(rho))
print(time.perf_counter()-start)
return {k:res for k,res in zip(haps.keys(),results)}
def sum_variant_lods(lods):
variants = {}
for lod in lods:
for m,l in zip(lod['MARKER'],lod['LOD']):
if m in variants.keys():
variants[m] += l
else:
variants[m] = l
var_lst = []
for var,lod in variants.items():
snp = var[:-3]
var_lst.append(snp.split(':')+[snp,lod])
variants=pd.DataFrame(var_lst,columns=['CHR','POS','A0','A1','SNP','LOD'])
variants.POS = variants.POS.astype(int)
variants.sort_values('POS')
return variants
# -
# ## Testing
import pandas as pd
import numpy as np
import pickle
from SEQLinkage.linkage import *
# ### Read fam
fam17 = pd.read_csv('../data/new_trim_ped_famless17_no:xx.fam',delim_whitespace=True,header=None,names=['fid','iid','fathid','mothid','sex','ad'])
fam17.index = list(fam17.iid)
fam17.ad[fam17.ad==-9]=0
fam17_d = {}
for i in fam17.fid.unique():
fam17_d[i] = fam17[fam17.fid==i]
# ## Read haplotypes
import glob
glob.glob('../data/wg20220316/chr9test/tmp/CACHE/chr9test*.pickle')[-3:]
for i in glob.glob('../data/wg20220316/chr9test/tmp/CACHE/chr9test*.pickle')[-3:]:
print(i)
run_gene_lods(i[:-7],fam17_d,cutoff=0.05)
for i in glob.glob('../data/wg20220316/chr10test/tmp/CACHE/chr10test*.pickle'):
print(i)
run_gene_lods(i[:-7],fam17_d)
def run_gene_lods(file,fam,rho=0,cutoff=None):
with open(file+'.pickle', 'rb') as handle:
genes = pickle.load(handle)
gene_variants,gene_fam_haps = format_haps_bunch(genes,fam)
if cutoff is not None:
for f,variants in gene_variants.items():
gene_fam_haps[f]=gene_fam_haps[f].loc[:,[True]*6+list(np.repeat((variants.freqs>cutoff)[variants.uniq],2))]
res = parallel_lods(gene_fam_haps.values(),rho)
smy_res = sum_variant_lods(res)
with open(file+'cutoff'+str(cutoff)+'_rho'+str(rho)+'.result','wb') as handle:
pickle.dump(smy_res, handle, protocol=pickle.HIGHEST_PROTOCOL)
def format_haps_by_genes(file,fam,cutoff=None):
with open(file+'.pickle', 'rb') as handle:
genes = pickle.load(handle)
gene_variants,gene_fam_haps = format_haps_bunch(genes,fam)
if cutoff is not None:
for f,variants in gene_variants.items():
gene_fam_haps[f]=gene_fam_haps[f].loc[:,[True]*6+list(np.repeat((variants.freqs>cutoff)[variants.uniq],2))]
with open(file+'cutoff'+str(cutoff)+'.input','wb') as handle:
pickle.dump([gene_variants,gene_fam_haps], handle, protocol=pickle.HIGHEST_PROTOCOL)
file='../data/wg20220316/chr22test/tmp/CACHE/chr22test24cutoff0.05.input'
with open(file, 'rb') as handle:
gene_variants,gene_fam_haps = pickle.load(handle)
res = parallel_lods(gene_fam_haps.values(),np.arange(0,0.5,0.05))
with open(file[:-6]+'.lods','wb') as handle:
pickle.dump(res, handle, protocol=pickle.HIGHEST_PROTOCOL)
sos run nbs/seqlink_sos.ipynb lods --cwd data/wg20220316 --fam_path data/new_trim_ped_famless17_no:xx.fam --chrom 22 -j 1
format_haps_by_genes('../data/wg20220311/chr19test/CACHE/chr19test43',fam17_d,cutoff=0.05)
run_gene_lods('../data/wg20220311/chr19test/CACHE/chr19test44',fam17_d,rho=0.05,cutoff=0.05)
with open('../data/wg20220316/fam17_vcf.pickle', 'rb') as handle:
fam17_vcf = pickle.load(handle)
with open('../data/wg20220421/chr22test/tmp/CACHE/chr22test0.pickle', 'rb') as handle:
genes = pickle.load(handle)
gene_variants,gene_fam_haps = format_haps_bunch(genes,fam17_d,fam17_vcf,cutoff=0.01,haplotype=False)
with open('../data/wg20220316/chr22test/tmp/CACHE/chr22test24cutoff0.05.input', 'rb') as handle:
gene_variants,gene_fam_haps = pickle.load(handle)
tmp=gene_fam_haps['1007']
for x,y in zip(gene_fam_haps.values(),[gene_variants[k] for k in gene_fam_haps.keys()]):
if (x.shape[1]-6)/2!=sum(y.uniq):
print(x.fid[0])
afreqs = []
for k in gene_fam_haps.keys():
variants= gene_variants[k]
variants=variants.freqs[variants.uniq]
#variants=variants[variants>0.05]
afreqs.append(list(variants.round(decimals=3)))
tmp=[]
for i in range(10):
try:
hap=gene_fam_haps[list(gene_fam_haps.keys())[i]]
frq=afreqs[i]
tmp.append(calculate_ped_lod(hap,frq,np.arange(0,0.5,0.05)))
except:
print(i)
res = parallel_lods(gene_fam_haps,afreqs,np.arange(0,0.5,0.05))
len(res)
0<1<2
genes['APOE']['predata']['1007'][2]
# ### Functions of heterogeneity
with open('../data/wg20220316/chr22test/tmp/CACHE/chr22test24cutoff0.05.lods', 'rb') as handle:
res = pickle.load(handle)
res[0]
res=list(res)
variants = sorted(list(set().union(*[i.index for i in res])))
def format_fam_lods(res):
new_res,variants=[],[]
for i in res:
new_res.append(i)
variants.append(i.index)
variants = list(set().union(*variants))
cutoff = len(new_res)//10
var_res={}
for v in variants:
varlods = [r.loc[v] for r in res if v in r.index]
if len(varlods)>cutoff:
var_res[v]=pd.concat(varlods,axis=1)
return var_res
start = time.perf_counter()
var_res=format_fam_lods(res)
print(time.perf_counter()-start)
list(var_res.keys())[:10]
def hlod_fun(Li, sign=1):
def _fun(alpha):
return sign * sum(np.log10(alpha*np.power(10, Li) + 1 - alpha))
return _fun
start = time.perf_counter()
var_sovs=[]
for var,res in var_res.items():
for theta in res.index:
sov = minimize_scalar(hlod_fun(list(res.loc[theta]), -1), bounds=(0,1), method='bounded', options={'xatol':1e-8})
var_sovs.append([var,theta,sov.x,-sov.fun])
print(time.perf_counter()-start)
def min_hlod_func(res):
var_sovs=[]
for theta in res.index:
sov = minimize_scalar(hlod_fun(list(res.loc[theta]), -1), bounds=(0,1), method='bounded', options={'xatol':1e-8})
var_sovs.append([var,theta,sov.x,-sov.fun])
return var,theta,sov.x,-sov.fun
start = time.perf_counter()
results1=[]
with ProcessPoolExecutor(max_workers = 10) as executor:
results = executor.map(min_hlod_func,var_res.values())
#for i in results:
# results1.append(i)
print(time.perf_counter()-start)
for i in results:
pass
results=list(results)
# ### Pipeline of heterogeneity
from scipy.optimize import minimize_scalar
lod_files=glob.glob('../data/wg20220316/chr22test/tmp/CACHE/chr22test*cutoff0.05.lods')
for file in lod_files:
print(file[:-5])
with open(file, 'rb') as handle:
res = pickle.load(handle)
var_res=format_fam_lods(res)
start = time.perf_counter()
var_sovs,best_sovs=[],[]
for var,res in var_res.items():
best_sov=[var,'LOD0.5',0,0]
for theta in res.index:
sov = minimize_scalar(hlod_fun(list(res.loc[theta]), -1), bounds=(0,1), method='bounded', options={'xatol':1e-8})
var_sov=[var,theta,sov.x,-sov.fun]
var_sovs.append(var_sov)
if best_sov[3]<var_sov[3]:
best_sov=var_sov
best_sovs.append(best_sov)
print(time.perf_counter()-start)
var_sovs=pd.DataFrame(var_sovs)
best_sovs=pd.DataFrame(best_sovs)
with open(file[:-5]+'.hlods','wb') as handle:
pickle.dump(var_sovs, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open(file[:-5]+'.besthlod','wb') as handle:
pickle.dump(best_sovs, handle, protocol=pickle.HIGHEST_PROTOCOL)
lod_files=glob.glob('../data/wg20220316/chr22test/tmp/CACHE/chr22test*cutoff0.05unimputed.lods')
for file in lod_files:
print(file[:-5])
with open(file, 'rb') as handle:
res = pickle.load(handle)
var_res=format_fam_lods(res)
start = time.perf_counter()
var_sovs,best_sovs=[],[]
for var,res in var_res.items():
best_sov=[var,'LOD0.5',0,0]
for theta in res.index:
sov = minimize_scalar(hlod_fun(list(res.loc[theta]), -1), bounds=(0,1), method='bounded', options={'xatol':1e-8})
var_sov=[var,theta,sov.x,-sov.fun]
var_sovs.append(var_sov)
if best_sov[3]<var_sov[3]:
best_sov=var_sov
best_sovs.append(best_sov)
print(time.perf_counter()-start)
var_sovs=pd.DataFrame(var_sovs)
best_sovs=pd.DataFrame(best_sovs)
with open(file[:-5]+'.hlods','wb') as handle:
pickle.dump(var_sovs, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open(file[:-5]+'.besthlod','wb') as handle:
pickle.dump(best_sovs, handle, protocol=pickle.HIGHEST_PROTOCOL)
lod_files=glob.glob('../data/wg20220425genes/chr22test/tmp/CACHE/chr22test*cutoff0.05unimputed.lods')
for file in lod_files:
print(file[:-5])
with open(file, 'rb') as handle:
res = pickle.load(handle)
var_res=format_fam_lods(res.values())
start = time.perf_counter()
var_sovs,best_sovs=[],[]
for var,res in var_res.items():
best_sov=[var,'LOD0.5',0,0]
for theta in res.index:
sov = minimize_scalar(hlod_fun(list(res.loc[theta]), -1), bounds=(0,1), method='bounded', options={'xatol':1e-8})
var_sov=[var,theta,sov.x,-sov.fun]
var_sovs.append(var_sov)
if best_sov[3]<var_sov[3]:
best_sov=var_sov
best_sovs.append(best_sov)
print(time.perf_counter()-start)
var_sovs=pd.DataFrame(var_sovs)
best_sovs=pd.DataFrame(best_sovs)
with open(file[:-5]+'.hlods','wb') as handle:
pickle.dump(var_sovs, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open(file[:-5]+'.besthlod','wb') as handle:
pickle.dump(best_sovs, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('../data/wg20220316/chr22test/tmp/CACHE/chr22test24cutoff0.05'+'.besthlod','rb') as handle:
best_sovs=pickle.load(handle)
best_sovs
# ### Pipeline of linkage analysis (without haplotype imputation)
ped_vcf=pd.read_csv('../data/new_trim_ped.csv')
ped_vcf.index=list(ped_vcf.iid)
fam17_vcf={}
for k,v in fam17_d.items():
fam17_vcf[k]=ped_vcf.vcf[v.index]
with open('../data/wg20220316/fam17_vcf.pickle','wb') as handle:
pickle.dump(fam17_vcf, handle, protocol=pickle.HIGHEST_PROTOCOL)
file='../data/wg20220316/chr22test/tmp/CACHE/chr22test24cutoff0.05.input'
with open(file, 'rb') as handle:
gene_variants,gene_fam_haps = pickle.load(handle)
res = parallel_lods(gene_fam_haps.values(),np.arange(0,0.5,0.05))
with open(file[:-6]+'.lods','wb') as handle:
pickle.dump(res, handle, protocol=pickle.HIGHEST_PROTOCOL)
hap=gene_fam_haps['1007']
def unimput_haps():
pass
hap.shape
for k,hap in gene_fam_haps.items():
hap.loc[~fam17_vcf[k],[False]*6+[True]*(hap.shape[1]-6)]=0
ped_vcf.vcf[gene_fam_haps['1007'].iid]
# pseudomarker -p test_f10.ped -m test_f10.map --dom
# # Merlin to linkage
cmap=pd.read_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22.map',sep='\t')
new_map=cmap.iloc[:,[0,2,1]]
new_map.columns = ['Chromosome','Haldane','Name']
new_map.to_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22_new.map',header=True,index=False,sep='\t')
new_map
cped = pd.read_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22.ped',sep='\t',header=None)
cped.shape
for i in range(0,cped.shape[1]-6,2):
tmp0 = cped.iloc[:,6+i]
tmp1 = cped.iloc[:,7+i]
ind = (tmp0==0) | (tmp1==0)
tmp0[ind]=0
tmp1[ind]=0
tmp0[tmp0.astype(int)>2]=2
tmp1[tmp1.astype(int)>2]=2
cped[5]=cped[5].replace(-9,0)
cped.index = list(cped[1])
cped=cped.sort_index()
cped.to_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22_new.ped',header=False,index=False,sep='\t')
cped.iloc[:,:26].to_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22_new_f10.ped',header=False, index=False,sep='\t')
new_map[:10].to_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22_new_f10.map',header=True,index=False,sep='\t')
cped
# ## Run paramlink2 on CHP markers
cped = pd.read_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22.ped',sep='\t',header=None)
cped=cped.replace('?',0)
cped = pd.concat([cped.iloc[:,:4].astype(str),cped.iloc[:,4:].astype(int)],axis=1)
cped.index = list(cped[1])
cped=cped.sort_index()
cped[5]=cped[5].replace(-9,0)
tmp = cped.iloc[:,6:]
tmp[tmp>2]=2
cped = pd.concat([cped.iloc[:,:6],tmp],axis=1)
cped_d={}
for i in cped[0].unique():
cped_d[i]=cped[cped[0]==i]
calculate_ped_lod(cped_d['1137'])
cped_res = parallel_lods(cped_d.values())
cmap
variants = {}
for lod in cped_res:
for m,l in zip(lod['MARKER'],lod['LOD']):
if pd.isna(l):
continue
if m in variants.keys():
variants[m] += l
else:
variants[m] = l
#variants=pd.DataFrame(variants)
variants
cped_d['1007'].to_csv('../data/wg20220316/chr22test/MERLIN/chr22test.chr22_new_1007.ped',header=False,index=False,sep='\t')
lod_files=glob.glob('../data/wg20220425genes/chr22test/tmp/CACHE/chr22test*cutoff0.05unimputed.lods')
for file in lod_files:
print(file[:-5])
with open(file, 'rb') as handle:
res = pickle.load(handle)
tmp=list(res.values())[0]
if 'chr22:32532265:G:A_A0' in tmp.index:
var_res=format_fam_lods(res.values())
var_res['chr22:32532265:G:A_A0'].transpose()
var_res['chr22:32532265:G:A_A0'].sum(axis=1)
var_res['chr22:32532265:G:A_A0'].transpose().sort_values('LOD0.0')
tmp=var_res['chr22:32532265:G:A_A0'].transpose().loc[:,'LOD0.0']
sum(tmp>0)
tmp[tmp>0].sum()
tmp[tmp<0].sum()
| nbs/12_linkage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# language: python
# name: python373jvsc74a57bd0210f9608a45c0278a93c9e0b10db32a427986ab48cfc0d20c139811eb78c4bbc
# ---
from data_loading.data_loader import load_data
data = load_data()
import torch
import numpy as np
np.random.shuffle(data)
X = []
y = []
for d in data:
X.append(d[0])
y.append(d[1])
VAL_SPLIT = 0.25
VAL_SPLIT = len(X) * VAL_SPLIT
VAL_SPLIT = int(VAL_SPLIT)
X_train = X[:-VAL_SPLIT]
y_train = y[:-VAL_SPLIT]
X_test = X[-VAL_SPLIT:]
y_test = y[-VAL_SPLIT:]
print(len(X_train))
print(len(y_train))
print(len(X_test))
print(len(y_test))
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
print(len(X_train))
print(len(y_train))
print(len(X_test))
print(len(y_test))
# ## Modelling
from torchvision import models
device = torch.device('cuda')
import torch.nn as nn
# model = models.resnet18(pretrained=True).to(device)
# inf = model.fc.in_features
# model.fc = nn.Linear(inf,2)
from models.baseline_model import BaseLine_Model
model = BaseLine_Model().to(device)
model = model.to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
BATCH_SIZE = 32
EPOCHS = 100
import wandb
PROJECT_NAME = 'Face-Mask-Detection'
from tqdm import tqdm
wandb.init(project=PROJECT_NAME,name='test')
for _ in tqdm(range(EPOCHS),leave=False):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
preds = model(X_batch)
preds.to(device)
print(preds.shape)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item()})
wandb.finish()
for index in range(len(preds)):
print(torch.argmax(torch.round(preds)[index]))
print(y_batch[index])
print('\n')
| wandb/run-20210521_221444-5y0aquoo/tmp/code/train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CMS simulaatio avoimella datalla
#
# [](https://mybinder.org/v2/gh/cms-opendata-education/cms-jupyter-materials-finnish/master?filepath=%2FOppimiskokonaisuudet%2FAvoin-data-hiukkasfysiikassa%2Fcms-simulaatio.ipynb)
#
#
# Maailman isoin hiukkaskiihdytin LHC (Large Hadron Collider) kiihdyttää CERN:issä protoneita joiden törmäyksistä CMS (Compact Muon Solenoid) kerää dataa. LHC törmäyttää protoni ryppäitä noin 40 miljoonaa kertaa sekunnissa [2], mutta vain osa tapahtumista johtaa suurienergisiin törmäyksiin. Osa törmäyksen energiasta muuttuu massaksi ($E=mc^2$) ja muodostuu uusia hiukkasia [4]. Raskaat ja lyhytikäiset hiukkaset hajoavat edelleen kevyempiin hiukkasiin, joita hiukkasilmaisin havaitsee. Tutkijoiden tehtäväksi jää selvittää mitä tapahtui törmäyksen ja mitatun datan välissä. Esimerkiksi Higgsin hiukkasta ilmaisin ei havaitse, mutta sen hajoamisesta syntyviä kevyempiä hiukkasia voidaan mitata.
#
#
# Tässä tehtävässä pääset tarkastelemaan aitoa dataa joka on kerätty CMS ilmaisimella. Käytetään visuaalista työkalulla joka löytyy CERN:in avoimen datan portaalista:
# [http://opendata.cern.ch/visualise/events/CMS](http://opendata.cern.ch/visualise/events/CMS).
#Vielä nopea katsaus LHC hiukkaskiihdyttimeen ja CMS hiukkasilmaisimeen, videon (2.51min).
from IPython.display import HTML
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/pQhbhpU9Wrg" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>')
#www.youtube.com/embed/pQhbhpU9Wrg
# ****
# ## CMS ilmaisimen rakenteen tutkiskelu
# CMS hiukkasilmaisin on rakenteeltaan kuin sipuli. Jokaisella kerroksella on tärkeä tehtävä törmäyksestä sinkoilevien hiukkasten ratojen määrittämisessä. Avataan visualisointi työkalu [CMS event display](http://opendata.cern.ch/visualise/events/CMS) ja ruvetaan tutkimaan CMS:n rakennetta lisäämällä kerroksia vasemman reunan "Detector" valikosta. Parhaan näkymän saa vaihtamalla "Ortographic projecton" asetukseen ja tarkastelemalla ilmaisinlieriötä pohjasta.
# <br>
# <img src="../../Kuvat/EventDisplay_aloitus.PNG" align="center" width="600px">
# <br>
# ### Kerrosten tehtävät
# Eri hiukkaset vaikuttavat aineen kanssa eri tavoin, eli tarvitaan erilaisia kerroksia mittamaan hiukkasten energioita ja ratoja. Tässä lyhyt tiivistelmä [kattavammasta esityksestä.](https://cms-docdb.cern.ch/cgi-bin/PublicDocDB/RetrieveFile?docid=12398&filename=SetOfPostersFN_HQ_small_16092014.pdf)
# - **Jälki-ilmaisin**
# Jälki-ilmaisin koostuu kahden tyyppisisitä pii osista. Jälki-ilmaisin havaitsee varattujen hiukkasten reitit, kun ne vuorovaikuttavat elekromagneettisesti ilmaisimen kanssa. Tarkan paikkadatan avulla voidaan määrittää protonien törmäyskohdat ja syntyneiden raskaampien ydinten hajoamispaikat. Myös hiukkasen radan kaarevuussäteen avulla voidaan laskea sen momentti.
# - _Pixel_ -ilmaisin on rakennettu pienistä piisoluista (65 miljoonaa kappaletta), jotka mittaavat varattujen hiukkasten radat hyvin tarkasti.
# - _Tracker_ -kerros on valmistettu piiliuskoista jotka niin ikään mittaavat hiukkasten ratoja.
# - **Sähkömagneettinen kalorimetri (ECAL)**
# Elektronien ja fotonien energiat saadaan mitattua hyvin tarkasti sillä niiden törmäyks ECAL-kerrokseen aiheuttaa sähkömagneettisen ryöpyn joka mitataan tuikeilmaisimilla. Törmänneen elektronin tai fotonin energia on suoraan verrannollinen tuikeilmaisimien havaitsemaan valon määrään.
# - **Hadronikalorimetri (HCAL)**
# Hadronikalorimetri pysäyttää hadroneiksi kutsutut hiukkaset kuten protonit ja netronit. HCAL-kerrokseen saapuvat hadronit menettävät liike-energiaansa hiukkasryöppyihin joiden synnyttämien tuikevalojen avulla saadaan lasketuksi hadronin alkuperäinen energia.
# - **Myonijärjestelmä**
# CMS ilmaisin on nimensä (Compact Muon Solenoid) mukaan suunniteltu erityisesti havaitsemaan myoneita. Myonit ovat vaikeasti havaittavia ja ne kulkevatkin ECAL ja HCAL kerrosten läpi pysähtymättä. Kuitenkin positiivisina hiukkasina niiden kulkiessa kaasulla täytettyjen kammioiden läpi (*drift tubes*) kaasu ionisoituu ja vapautuneet electronit sekä positiiviset ionit kulkeutuvat (*Cathode Strip Chambers*) anodeille ja katodeile. Signaalin avulla voidaan laskea myonin paikka tietyllä ajanhetkellä. *Resistive Plate Chambers* ovat myös osa myonijärjestelmää luoden signaalia, joka voidaan siirtää eteenpäin käsiteltäväksi.
#
#
#
# Neutriinot havaitaan epäsuorasti liikemäärän säilymisen kautta.
#
# <br>
# <img src="../../Kuvat/CMS_Slice.gif" align="center" width="700px">
# <br>
#
# (_Superconductin_ _Solenoid_ on todella tehokas solenoidimagneetti joka aiheuttaa CMS:n siäsosiin noin 100,000 kertaan maan magneettikenttää vahvemman magneettikentän. Vahva kenttä saa varattujen hiukkasten liikeradat kaartumaan ja mahdollistaa niiden luokittelun ja liikemäärän määrittämisen.)
#
# Tarkat kuvaukset eri osien toiminnasta englanniksi: http://cms.web.cern.ch/news/detector-overview
#
# ****
# ## Törmäysten visualisointi
# Event Displayn avulla voi tarkastella CMS-mittausaseman todellisia tuloksia. Tässä tehtävässä käytetään protonitörmäyksistä kerättyä dataa, joka sittemmin johti Higgsin hiukkasen löytämiseen vuonna 2012.
#
# #### Aloitetaan törmäyksien tutkiminen
#
# 1. Avaa [CMS event display](http://opendata.cern.ch/visualise/events/CMS)
# 1. Paina vasemmasta yläkulmasta $\color{green}{\text{Open file}}$
# 1. Valitse $\color{green}{\text{Open file(s)}}$
# 1. Valitse $\color{green}{\text{HiggsCandidates/}}$
# 1. Valitse toinen mahdollisista Higgsin bosonin hajoamistavoista
# > $\color{green}{\text{4lepton.ig}}$ nimisissä tiedostoissa syntyy neljä leptonia. Niissä saattaa esiintyä Higgsin hajoaminen kahdeksi Z bosoniksi, jotka edelleen hajoavat joko elektroni/positroni ($e^- e^+$) tai myoni/antimyoni ($\mu^- \mu^+$) pareiksi. Syntyy siis yhteensä neljä leptonia.
#
# <img src="../../Kuvat/higgs-4l.png" align="center" width="200px">
# > $\color{green}{\text{diphoton.ig}}$ tiedostoissa törmäyksessä syntyi 2 fotonia. Ne saattavat olla peräisin tapahtumasarjasta jossa Higgs hajoaa kahdeksi fotoniksi.
#
# <img src="../../Kuvat/higgs-2photon.png" align="center" width="200px">
# 1. Valitse mikä tahansa ajo ja paina $\color{green}{\text{load}}$
# 1. Törmäyksen pitäisi nyt näkyä event displayssa.
# 1. Poista vasemman reunan valikosta, $\color{orange}{\text{Tracking}}$ otsikon alta, valinta $\color{orange}{\text{Tracks (reco.)}}$
# > Mikäli valitsit $\color{green}{\text{4lepton.ig}}$ valitse $\color{orange}{\text{Physics}}$ otsikon alta $\color{orange}{\text{Electron Tracks (GSF)}}$ ja $\color{orange}{\text{Tracker Muons (Reco)}}$ näkyviksi. Huom: Valinta on näkyvissä vain jos kyseinen hiukkanen on läsnä tapahtumissa.
# - Löydätkö kaikki neljä leptonia? Montako elektronia ja montako myonia valitsemassasi ajossa syntyi?
# - Voisivatko leptonit olla peräisin Higgsin hajoamisesta?
# - Tutki mihin ilmaisimen kerrokseen eri leptonien liike pysähtyy. (Matching muon chambers valinta paljastaa ne myonijärjestelmän kammiot joiden läpi myoni kulkee.) Vastaako tulos kerrosten kuvauksia?
#
# > Jos taas valitsit $\color{green}{\text{diphoton.ig}}$ tiedoston, valitse $\color{orange}{\text{Physics}}$ otsikon alta $\color{orange}{\text{Photons}}$ näkyviksi. Valinta on mahdollinen vain mikäli kyseisellä ajolla CMS todella havaitsi fotoneita.
# - Löydätkö kaksi fotonia? Voisivatko ne olla peräisin Higgsin hajoamisesta?
# - Tutki mihin ilmaisimen kerrokseen fotonien liike pysähtyy. Vastaako tulos kerrosten kuvauksia?
#
# Kokeile eri datalla ja vertaa tuloksia.
#
#
# ****
# Vuonna 2012 Atlaksen ja CMS:n tutkiat julkistivat löytäneensä Higsin bosonin. Teoreettisesti sen olemassaolo oli jo ennustettu, muttaa kokeellisesti se havaittiin ensimmäisen kerran LHC kiihdyttimen monen vuoden datan avulla. Higgsin bosoni täydentää standardimallia, muttei tee siitä ns. "kaiken teoriaa". CMS hiukkasilmaisimen tuottamaa dataa tutkitaan tauotta CERN:issä ja yliopistoissa ympäri maailmaa. Dataa tutkimalla tutkijat pyrkivät ymmärtämään paremmin aineen rakennetta ja esimerkiksi pimeän aineen ja antimateriaalin luonnetta.
# +
from IPython.lib.display import YouTubeVideo
YouTubeVideo('0QPTzogoJ-Q')
#https://youtu.be/0QPTzogoJ-Q
# -
# ****
# ## Lähteet
# [1] CMS koeasema, Diat suomeksi, Luettu 2019.
# Url: [https://cms-docdb.cern.ch/cgi-bin/PublicDocDB/RetrieveFile?docid=12398&filename=SetOfPostersFN_HQ_small_16092014.pdf](https://cms-docdb.cern.ch/cgi-bin/PublicDocDB/RetrieveFile?docid=12398&filename=SetOfPostersFN_HQ_small_16092014.pdf).
#
# [2] Detector overview. © Copyright CERN (2008-2014)
# Url: [http://cms.web.cern.ch/news/detector-overview](http://cms.web.cern.ch/news/detector-overview)
#
# [3] CMS Guide to education use of CMS Open Data
# Url: [http://opendata.cern.ch/docs/cms-guide-for-education](http://opendata.cern.ch/docs/cms-guide-for-education)
#
# [4] 12 steps - From idea to discovery, luettu 2019
# Url: [https://home.cern/science/physics/12-steps-idea-discovery](https://home.cern/science/physics/12-steps-idea-discovery)
#
| Opetusmateriaalit/AvoinDataHiukkasfysiikassa/2_CMS-simulaatio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import math
import pickle
from scipy import stats
import scipy.io
from scipy.spatial.distance import pdist
from scipy.linalg import cholesky
from scipy.io import loadmat
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.metrics import classification_report,roc_auc_score,recall_score,precision_score
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.decomposition import PCA
from pyearth import Earth
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
from sklearn.model_selection import StratifiedKFold
from src import SMOTE
from src import CFS
from src import metrices_V2 as metrices
import platform
from os import listdir
from os.path import isfile, join
from glob import glob
from pathlib import Path
import sys
import os
import copy
import traceback
from pathlib import Path
import matplotlib.pyplot as plt
# +
def apply_smote(df):
df.reset_index(drop=True,inplace=True)
cols = df.columns
smt = SMOTE.smote(df)
df = smt.run()
df.columns = cols
return df
def apply_cfs(df):
y = df.Bugs.values
X = df.drop(labels = ['Bugs'],axis = 1)
X = X.values
selected_cols = CFS.cfs(X,y)
cols = df.columns[[selected_cols]].tolist()
cols.append('Bugs')
return df[cols],cols
# -
def load_product_data(project):
# Processing Product files
understand_path = 'data/understand_files_all/' + project + '_understand.csv'
understand_df = pd.read_csv(understand_path)
understand_df = understand_df.dropna(axis = 1,how='all')
cols_list = understand_df.columns.values.tolist()
for item in ['Kind', 'Name','commit_hash', 'Bugs']:
if item in cols_list:
cols_list.remove(item)
cols_list.insert(0,item)
understand_df = understand_df[cols_list]
cols = understand_df.columns.tolist()
understand_df = understand_df.drop_duplicates(cols[4:len(cols)])
df = understand_df
cols = df.columns.tolist()
cols.remove('Bugs')
cols.append('Bugs')
_df = df
df = df[cols]
for item in ['Kind', 'Name','commit_hash']:
if item in cols:
df = df.drop(labels = [item],axis=1)
df.dropna(inplace=True)
df.reset_index(drop=True, inplace=True)
y = df.Bugs
X = df.drop('Bugs',axis = 1)
cols = X.columns
scaler = MinMaxScaler()
X = scaler.fit_transform(X)
X = pd.DataFrame(X,columns = cols)
return X,y,_df
def load_process_data(project):
understand_path = 'data/understand_files_all/' + project + '_understand.csv'
understand_df = pd.read_csv(understand_path)
commits = understand_df.commit_hash.unique()
commit_guru_file_level_path = 'data/commit_guru_file_level/' + project + '_file.csv'
commit_guru_path = 'data/commit_guru/' + project + '.csv'
commit_guru_file_level_df = pd.read_csv(commit_guru_file_level_path)
commit_guru_file_level_df['commit_hash'] = commit_guru_file_level_df.commit_hash.str.strip('"')
commit_guru_df = pd.read_csv(commit_guru_path)
commit_guru_df = commit_guru_df[['commit_hash','contains_bug']]
values = {'contains_bug': False}
commit_guru_df = commit_guru_df.fillna(values)
# commit_guru_df = commit_guru_df[commit_guru_df.commit_hash.isin(commits)]
df = commit_guru_file_level_df.merge(commit_guru_df,on='commit_hash')
df.rename(columns={"contains_bug": "Bugs"},inplace=True)
df = df[df['file_name'].str.contains('.java')]
df = df.drop(['commit_hash','file_name'],axis = 1)
df.dropna(inplace=True)
df.reset_index(drop=True, inplace=True)
df = df.drop_duplicates()
y = df.Bugs
X = df.drop('Bugs',axis = 1)
cols = X.columns
scaler = MinMaxScaler()
X = scaler.fit_transform(X)
X = pd.DataFrame(X,columns = cols)
return X,y
def load_both_data(project,metric):
understand_path = 'data/understand_files_all/' + project + '_understand.csv'
understand_df = pd.read_csv(understand_path)
understand_df = understand_df.dropna(axis = 1,how='all')
cols_list = understand_df.columns.values.tolist()
for item in ['Kind', 'Name','commit_hash', 'Bugs']:
if item in cols_list:
cols_list.remove(item)
cols_list.insert(0,item)
understand_df = understand_df[cols_list]
cols = understand_df.columns.tolist()
understand_df = understand_df.drop_duplicates(cols[4:len(cols)])
understand_df['Name'] = understand_df.Name.str.rsplit('.',1).str[1]
commit_guru_file_level_path = 'data/commit_guru_file_level/' + project + '_file.csv'
commit_guru_file_level_df = pd.read_csv(commit_guru_file_level_path)
commit_guru_file_level_df['commit_hash'] = commit_guru_file_level_df.commit_hash.str.strip('"')
commit_guru_file_level_df = commit_guru_file_level_df[commit_guru_file_level_df['file_name'].str.contains('.java')]
commit_guru_file_level_df['Name'] = commit_guru_file_level_df.file_name.str.rsplit('/',1).str[1].str.split('.').str[0].str.replace('/','.')
commit_guru_file_level_df = commit_guru_file_level_df.drop('file_name',axis = 1)
df = understand_df.merge(commit_guru_file_level_df,how='left',on=['commit_hash','Name'])
cols = df.columns.tolist()
cols.remove('Bugs')
cols.append('Bugs')
df = df[cols]
for item in ['Kind', 'Name','commit_hash']:
if item in cols:
df = df.drop(labels = [item],axis=1)
# df.dropna(inplace=True)
df = df.drop_duplicates()
df.reset_index(drop=True, inplace=True)
y = df.Bugs
X = df.drop('Bugs',axis = 1)
cols = X.columns
scaler = MinMaxScaler()
X = scaler.fit_transform(X)
X = pd.DataFrame(X,columns = cols)
imp_mean = IterativeImputer(random_state=0)
X = imp_mean.fit_transform(X)
X = pd.DataFrame(X,columns = cols)
if metric == 'process':
X = X[['la', 'ld', 'lt', 'age', 'ndev', 'nuc', 'ns', 'exp', 'sexp', 'rexp', 'nd']]
elif metric == 'product':
X = X.drop(['la', 'ld', 'lt', 'age', 'ndev', 'nuc', 'ns', 'exp', 'sexp', 'rexp', 'nd'],axis = 1)
else:
X = X
return X,y
def run_self(project,metric):
X,y = load_both_data(project,metric)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.40, random_state=18)
if metric == 'process':
loc = X_test['la'] + X_test['lt']
elif metric == 'product':
loc = X_test.CountLineCode
else:
loc = X_test['la'] + X_test['lt']
df_smote = pd.concat([X_train,y_train],axis = 1)
df_smote = apply_smote(df_smote)
y_train = df_smote.Bugs
X_train = df_smote.drop('Bugs',axis = 1)
clf = RandomForestClassifier()
clf.fit(X_train,y_train)
importance = clf.feature_importances_
print(len(importance))
predicted = clf.predict(X_test)
abcd = metrices.measures(y_test,predicted,loc)
pf = abcd.get_pf()
recall = abcd.calculate_recall()
precision = abcd.calculate_precision()
f1 = abcd.calculate_f1_score()
g_score = abcd.get_g_score()
pci_20 = abcd.get_pci_20()
ifa = abcd.get_ifa()
try:
auc = roc_auc_score(y_test, predicted)
except:
auc = 0
print(classification_report(y_test, predicted))
return recall,precision,pf,f1,g_score,auc,pci_20,ifa,importance
def run_self_k(project,metric):
precision = []
recall = []
pf = []
f1 = []
g_score = []
auc = []
pci_20 = []
ifa = []
importance = []
X,y = load_both_data(project,metric)
for _ in range(5):
skf = StratifiedKFold(n_splits=5)
for train_index, test_index in skf.split(X, y):
X_train, X_test = X.loc[train_index], X.loc[test_index]
y_train, y_test = y.loc[train_index], y.loc[test_index]
if metric == 'process':
loc = X_test['la'] + X_test['lt']
elif metric == 'product':
loc = X_test.CountLineCode
else:
loc = X_test['la'] + X_test['lt']
df_smote = pd.concat([X_train,y_train],axis = 1)
df_smote = apply_smote(df_smote)
y_train = df_smote.Bugs
X_train = df_smote.drop('Bugs',axis = 1)
clf = RandomForestClassifier()
clf.fit(X_train,y_train)
importance = clf.feature_importances_
predicted = clf.predict(X_test)
abcd = metrices.measures(y_test,predicted,loc)
pf.append(abcd.get_pf())
recall.append(abcd.calculate_recall())
precision.append(abcd.calculate_precision())
f1.append(abcd.calculate_f1_score())
g_score.append(abcd.get_g_score())
pci_20.append(abcd.get_pci_20())
ifa.append(abcd.get_ifa())
try:
auc.append(roc_auc_score(y_test, predicted))
except:
auc.append(0)
# print(classification_report(y_test, predicted))
return recall,precision,pf,f1,g_score,auc,pci_20,ifa,importance
proj_df = pd.read_csv('projects.csv')
projects = proj_df.repo_name.tolist()
precision_list = {}
recall_list = {}
pf_list = {}
f1_list = {}
g_list = {}
auc_list = {}
pci_20_list = {}
ifa_list = {}
featue_importance = {}
for project in projects[150:]:
try:
if project == '.DS_Store':
continue
print("+++++++++++++++++ " + project + " +++++++++++++++++")
recall,precision,pf,f1,g_score,auc,pci_20,ifa,importance = run_self_k(project,'product')
recall_list[project] = recall
precision_list[project] = precision
pf_list[project] = pf
f1_list[project] = f1
g_list[project] = g_score
auc_list[project] = auc
pci_20_list[project] = pci_20
ifa_list[project] = ifa
featue_importance[project] = importance
except Exception as e:
print(e)
continue
final_result = {}
final_result['precision'] = precision_list
final_result['recall'] = recall_list
final_result['pf'] = pf_list
final_result['f1'] = f1_list
final_result['g'] = g_list
final_result['auc'] = auc_list
final_result['pci_20'] = pci_20_list
final_result['ifa'] = ifa_list
final_result['featue_importance'] = featue_importance
with open('results/Performance/commit_guru_file_specific/product_700_rf_5_fold_5_repeat.pkl', 'wb') as handle:
pickle.dump(final_result, handle, protocol=pickle.HIGHEST_PROTOCOL)
| random_notebook_Experiments/File-level analysis-product.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import xml.etree.cElementTree as ET
from xml.etree.ElementTree import ElementTree,Element
import os
ET.register_namespace('',"urn:mpeg:dash:schema:mpd:2011")
def read_xml(in_path):
'''读取并解析xml文件
in_path: xml路径
return: ElementTree'''
tree = ElementTree()
tree.parse(in_path)
return tree
def write_xml(tree, out_path):
'''将xml文件写出
tree: xml树
out_path: 写出路径'''
tree.write(out_path,encoding="utf-8",xml_declaration=True)
def if_match(node, kv_map):
'''判断某个节点是否包含所有传入参数属性
node: 节点
kv_map: 属性及属性值组成的map'''
for key in kv_map:
if node.get(key) != kv_map.get(key):
return False
return True
#---------------search -----
def find_nodes(tree, path):
'''查找某个路径匹配的所有节点
tree: xml树
path: 节点路径'''
return tree.findall(path)
def get_node_by_keyvalue(nodelist, kv_map):
'''根据属性及属性值定位符合的节点,返回节点
nodelist: 节点列表
kv_map: 匹配属性及属性值map'''
i = 0
result_nodes = []
for node in nodelist:
i=i+1
if if_match(node, kv_map):
#if (i == 2):
result_nodes.append(node)
return result_nodes
#---------------change -----
def change_node_properties(num, nodelist, kv_map, is_delete=False):
'''修改/增加 /删除 节点的属性及属性值
nodelist: 节点列表
kv_map:属性及属性值map'''
i = 0
for node in nodelist:
i=i+1
for key in kv_map:
if is_delete:
if key in node.attrib:
del node.attrib[key]
else:
if (i==num):
node.set(key, kv_map.get(key))
def change_node_text(nodelist, text, is_add=False, is_delete=False):
'''改变/增加/删除一个节点的文本
nodelist:节点列表
text : 更新后的文本'''
for node in nodelist:
if is_add:
node.text += text
elif is_delete:
node.text = ""
else:
node.text = text
def create_node(tag, property_map, content):
'''新造一个节点
tag:节点标签
property_map:属性及属性值map
content: 节点闭合标签里的文本内容
return 新节点'''
element = Element(tag, property_map)
element.text = content
return element
def add_child_node(nodelist, element):
'''给一个节点添加子节点
nodelist: 节点列表
element: 子节点'''
for node in nodelist:
node.append(element)
def del_node_by_tagkeyvalue(nodelist, tag, kv_map):
'''同过属性及属性值定位一个节点,并删除之
nodelist: 父节点列表
tag:子节点标签
kv_map: 属性及属性值列表'''
for parent_node in nodelist:
children = parent_node.getchildren()
for child in children:
if child.tag == tag and if_match(child, kv_map):
parent_node.remove(child)
def get_tag_name(xml_element):
""" Module to remove the xmlns tag from the name
eg: '{urn:mpeg:dash:schema:mpd:2011}SegmentTemplate'
Return: SegmentTemplate
"""
# try:
tag_name = xml_element[xml_element.find('}')+1:]
# except TypeError:
# config_dash.LOG.error("Unable to retrieve the tag. ")
# return None
return tag_name
# +
tree = read_xml('dash_coaster_10x10_qp28.mpd')
root = tree.getroot()
child_period = root[1];
dir_path = "coaster_10x10/"
i=0
for adaptation_set in child_period:
adaptation_set.remove(adaptation_set[0])
# print(adaptation_set[0])
if i==0:
spec=adaptation_set[1][0].attrib['media'].split("_")
for k in range(1,61):
name = "_".join(spec[0:-1])+"_"+str(k)+".m4s"
file_size = os.path.getsize(dir_path+name)*8/1024.0
new_ele = create_node("SegmentSize", {"id":name,"size":str(file_size), "scale":"Kbits"}, "")
adaptation_set[1].append(new_ele)
else:
spec=adaptation_set[0][0].attrib['media'].split("_")
for k in range(1,61):
name = "_".join(spec[0:-1])+"_"+str(k)+".m4s"
file_size = os.path.getsize(dir_path+name)*8/1024.0
new_ele = create_node("SegmentSize", {"id":name,"size":str(file_size), "scale":"Kbits"}, "")
adaptation_set[0].append(new_ele)
i=i+1
write_xml(tree, "./dash_coaster_10x10_qp28.mpd")
# -
| dist/parse_mpd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using the Python SDK for autoRIFT
#
# HyP3's Python SDK `hyp3_sdk` provides a convenience wrapper around the HyP3 API and HyP3 jobs.
#
#
# The HyP3 SDK can be installed using [Anaconda/Miniconda](https://docs.conda.io/projects/conda/en/latest/user-guide/install/download.html#anaconda-or-miniconda)
# (recommended) via [`conda`](https://anaconda.org/conda-forge/hyp3_sdk):
#
# ```
# conda install -c conda-forge hyp3_sdk
# ```
#
# Or using [`pip`](https://pypi.org/project/hyp3-sdk/):
#
# ```
# python -m pip install hyp3_sdk
# ```
#
# Full documentation of the SDK can be found in the [HyP3 documentation](https://hyp3-docs.asf.alaska.edu/using/sdk/)
# +
# initial setup
import hyp3_sdk as sdk
AUTORIFT_API = 'https://hyp3-autorift.asf.alaska.edu/'
# -
# ## Authenticating to the API
#
# The SDK will attempt to pull your [NASA Earthdata Login](https://urs.earthdata.nasa.gov/) credentials out of `~/.netrc`
# by default, or you can pass your credentials in directly
# .netrc
hyp3 = sdk.HyP3(AUTORIFT_API)
# or enter your credentials
from getpass import getpass
username = 'MY_EDL_USERNAME'
password = getpass() # will prompt for a password
hyp3 = sdk.HyP3(AUTORIFT_API, username=username, password=password)
# ## Submitting jobs
#
# AutoRIFT jobs can be submitted using the `hyp3.submit_autorift_job()` method.
#
# ### Sentinel-1
#
# Sentinel-1 jobs are submitted using the [ESA granule ID](https://sentinel.esa.int/web/sentinel/user-guides/sentinel-1-sar/naming-conventions)
# +
s1_pairs = [
('S1A_IW_SLC__1SSH_20170221T204710_20170221T204737_015387_0193F6_AB07',
'S1B_IW_SLC__1SSH_20170227T204628_20170227T204655_004491_007D11_6654'),
('S1B_IW_SLC__1SDH_20180821T204618_20180821T204645_012366_016CC2_59A7',
'S1B_IW_SLC__1SDH_20180809T204617_20180809T204644_012191_01674F_9345'),
('S1A_IW_SLC__1SSH_20151214T080202_20151214T080229_009035_00CF68_780C',
'S1A_IW_SLC__1SSH_20151120T080202_20151120T080229_008685_00C5A7_105E'),
('S1A_IW_SLC__1SSH_20150909T162413_20150909T162443_007640_00A97D_922B',
'S1A_IW_SLC__1SSH_20150828T162412_20150828T162431_007465_00A4AF_DC3E'),
]
s1_jobs = sdk.Batch()
for g1, g2 in s1_pairs:
s1_jobs += hyp3.submit_autorift_job(g1, g2, name='s1-example')
# -
# Here we've given each job the name `s1-example`, which we can use later to search for these jobs.
#
# ### Sentinel-2
#
# Seninel-2 jobs can be submitted using either the [ESA granule ID](https://sentinel.esa.int/web/sentinel/user-guides/sentinel-2-msi/naming-convention)
# or the [Element 84 Earth Search ID](https://registry.opendata.aws/sentinel-2/)
# +
s2_pairs = [
# Can be either ESA granule IDs
('S2B_MSIL1C_20200612T150759_N0209_R025_T22WEB_20200612T184700',
'S2A_MSIL1C_20200627T150921_N0209_R025_T22WEB_20200627T170912'),
# or Element 84 Earth Search ID
('S2B_22WEB_20200612_0_L1C', 'S2A_22WEB_20200627_0_L1C'),
]
s2_jobs = sdk.Batch()
for g1, g2 in s2_pairs:
s2_jobs += hyp3.submit_autorift_job(g1, g2, name='s2-example')
# -
# ### Landsat-8 Collection 2
#
# Landsat-8 Collection 2 jobs are submitted using the [USGS scene ID](https://www.usgs.gov/faqs/what-naming-convention-landsat-collection-2-level-1-and-level-2-scenes?qt-news_science_products=0#qt-news_science_products)
# +
l8_pairs = [
('LC08_L1TP_009011_20200703_20200913_02_T1',
'LC08_L1TP_009011_20200820_20200905_02_T1'),
]
l8_jobs = sdk.Batch()
for g1, g2 in l8_pairs:
l8_jobs += hyp3.submit_autorift_job(g1, g2, name='l8-example')
# -
# ## Monitoring jobs
#
# One jobs are submitted, you can either watch the jobs until they finish
s1_jobs = hyp3.watch(s1_jobs)
# which will require you to keep the cell/terminal running, or you can come back later and search for jobs
s1_jobs = hyp3.find_jobs(name='s1-example')
s1_jobs = hyp3.watch(s1_jobs)
# ### Downloading files
#
# Batches are collections of jobs. They provide a snapshot of the job status when the job was created or last
# refreshed. To get updated information on a batch
s1_jobs = hyp3.refresh(s1_jobs)
# `hyp3.watch()` will return a refreshed batch once the batch has completed.
#
# Batches can be added together
print(f'Number of Jobs:\n S1:{len(s1_jobs)}\n S2:{len(s2_jobs)}\n L8:{len(l8_jobs)}')
all_jobs = s1_jobs + s2_jobs + l8_jobs
print(f'Total number of Jobs: {len(all_jobs)}')
# You can check if every job is complete and if every job was successful
all_jobs.complete()
all_jobs.succeeded()
# and filter jobs by status
succeeded_jobs = all_jobs.filter_jobs(succeeded=True, running=False, failed=False)
failed_jobs = all_jobs.filter_jobs(succeeded=False, running=False, failed=True)
# You can download the files for all successful jobs
file_list = succeeded_jobs.download_files('./')
# *Note: only succeeded jobs will have files to download.*
| docs/sdk_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import matplotlib.pyplot as plt
# Set up device and manual seed
torch.manual_seed(1)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# Create data
x = torch.arange(500, out=torch.FloatTensor()).view((-1, 1))
inverted_index = torch.randperm(1000, out=torch.FloatTensor())[:500].view(-1, 1, 1)
print(inverted_index.shape)
print(inverted_index[:2,:])
inverted_index = torch.sort(inverted_index, dim=0)
inverted_index = inverted_index[0]
max_document = inverted_index.max()
inverted_index_scaled = inverted_index / max_document
print(inverted_index_scaled.shape)
inverted_index[:2,:]
from sklearn.preprocessing import StandardScaler
y = StandardScaler().fit_transform(inverted_index.reshape(-1, 1).numpy())
y = torch.from_numpy(y)
inverted_index_scaled.shape
y.shape
x[:2,:]
class LII_LSTM(nn.Module):
def __init__(self, input_size=1, hidden_size=10, num_layers=1):
super(LII_LSTM, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.lstm = nn.LSTM(self.input_size, self.hidden_size)
self.linear = nn.Linear(self.hidden_size, 1) # output size is 1
def forward(self, input_seq):
lstm_out, _ = self.lstm(input_seq.view(input_seq.size()[0], 1, 1))
index_out = self.linear(lstm_out)
return index_out
def train(model, optimizer, index, target, scheduler=None, epochs=2000):
index.to(device)
target.to(device)
model.train()
for e in range(1, epochs+1):
# Zero out the grad
optimizer.zero_grad()
# Get output
prediction = model(index)
loss = F.mse_loss(prediction, target)
# Take gradient step
loss.backward()
optimizer.step()
# Take scheduler step
if scheduler:
scheduler.step(loss)
# Print loss
if e % 50 == 0:
#if e % 200 == 0:
with torch.no_grad():
plot_pred = prediction.detach().numpy().reshape(-1)
plot_index = index.numpy().reshape(-1)
plot_target = target.numpy().reshape(-1)
error = target - prediction
max_error = abs(error.max().item())
title = "Train Epoch {}: Loss - {}; Max error - {}".format(e, loss.item(), max_error)
plt.plot(plot_index, plot_target, label="True")
plt.plot(plot_index, plot_pred, label="Prediction")
plt.title(title)
plt.legend(loc="best")
plt.show()
return prediction
# +
# # Create model and setup optimizer
# lii_lstm = LII_LSTM()
# lii_lstm.to(device)
# optimizer = optim.Adam(params=lii_lstm.parameters(), lr=0.01)
# +
# Train model on unscaled inverted indices
# prediction = train(lii_lstm, optimizer, x, inverted_index)
# +
# Compare prediction and ground truth
# for i in range(prediction.size()[0]):
# print("True value: {} vs. Predicted: {}".format(int(inverted_index[i].item()),
# int(round(prediction[i].item()))))
# -
# Train model on scaled inverted indices
lii_lstm = LII_LSTM()
lii_lstm.to(device)
optimizer = optim.Adam(params=lii_lstm.parameters(), lr=0.01)
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer, patience=20, verbose=True, threshold=10e-6)
prediction_scaled = train(lii_lstm, optimizer, x, inverted_index_scaled, scheduler, epochs=500)
# Compare prediction and ground truth
for i in range(prediction_scaled.size()[0]):
print("True value: {} vs. Predicted: {}".format(int((inverted_index_scaled[i].item()*max_document).round()),
int((prediction_scaled[i].item()*max_document).round())))
| notebooks/.ipynb_checkpoints/lii_lstm_yg-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming with Python
#
# ## Episode 5 - Making Choices
#
# Teaching: 30 min,
# Exercises: 30 min
#
# ## Objectives
# - Write conditional statements including if, elif, and else branches.
# - Correctly evaluate expressions containing and and or.
# ### How can my programs do different things based on data values?
#
# In our last lesson, we discovered something suspicious was going on in our inflammation data by drawing some plots.
# How can we use Python to automatically recognise the different features we saw, and take a different action for each?
#
# In this lesson, we'll learn how to write code that runs only when certain conditions are true.
# ### Conditionals
# We can ask Python to take different actions, depending on a condition, with an `if` statement:
# ```
# num = 37
# if num > 100:
# print('greater')
# else:
# print('not greater')
# print('done')
# ```
# +
num = input()
num = int(num)
if num > 50:
print("great!")
else:
print("not great at all!")
# -
# The second line of this code uses the keyword `if` to tell Python that we want to make a choice.
#
# If the test (condition) that follows the `if` statement is *true*, the body of the `if` (i.e., the lines indented underneath it) are executed. If the test is *false*, the body of the `else` is executed instead.
#
# Only one or the other is ever executed.
# ### Executing a Conditional
#
# `if` statements don't have to include an `else` if it is not required. If there isn't one, Python simply does nothing if the test is *false*:
# ```
# num = 53
# print('before conditional...')
# if num > 100:
# print(num,' is greater than 100')
# print('...after conditional')
# ```
num = 600
print('before conditional...')
if num > 100:
print(num,' is greater than 100')
print('...after conditional')
# We can also chain several tests together using `elif`, which is short for "else if".
#
# The following Python code uses `elif` to print the sign (positive or negative) of a number.
#
# ```
# num = -3
#
# if num > 0:
# print(num, 'is positive')
# elif num == 0:
# print(num, 'is zero')
# else:
# print(num, 'is negative')
# ```
# Note that to test for *equality* we use a double equals sign ``==`` rather than a single equals sign `=` which is used to assign values.
# ### `and`'s and `or`'s
#
# We can also combine tests using `and` and `or`.
#
# `and` is only *true* if both parts are *true*:
#
# ```
# if (1 > 0) and (-1 > 0):
# print('both parts are true')
# else:
# print('at least one part is false')
# ```
if (1 > 0) and (-1 > 0):
print('both parts are true')
else:
print('at least one part is false')
# while `or` is true if at least one part is true:
# ```
# if (1 < 0) or (-1 < 0):
# print('at least one test is true')
# ```
if (1 < 0) or (-1 < 0):
print('at least one test is true')
# ### `True` and `False`
#
# `True` and `False` are special words (keywords) in Python called *booleans*, which represent truth values. A statement such as `1 < 0` returns the value `False`, while `-1 < 0` returns the value `True`. The can be used in place of conditional expressions:
# ```
# print(True of False
# ```
# +
# Multiple conditions
num = input()
num = int(num)
if num > 50 and num < 75 and num % 2 == 0:
print("number is between 50 and 75 and even")
elif num > 50 and num < 75:
print("number is between 50 and 75 and either even or not")
else:
print("number is not between 50 and 75")
# -
# ### Checking our Data
# Now that we've seen how conditionals work, we can use them to check for the suspicious features we saw in our inflammation data.
#
# Let's get back to where we left off in a previous episode:
#
# ```
# import numpy
# import matplotlib.pyplot
#
# data = numpy.loadtxt(fname='data/inflammation-03.csv', delimiter=',')
# min_plot = matplotlib.pyplot.plot(numpy.max(data, axis=0))
# matplotlib.pyplot.show()
# ```
# +
import numpy
import matplotlib.pyplot
data = numpy.loadtxt(fname='data/inflammation-01.csv', delimiter=',')
min_plot = matplotlib.pyplot.plot(numpy.max(data, axis=0))
matplotlib.pyplot.show()
# -
# From the plots, we see that maximum daily inflammation exhibits a strange behaviour and raises one unit a day. Wouldn't it be a good idea to detect such behaviour and report it as suspicious? Let's do that!
#
# However, instead of checking every single day of the study, let's merely check if maximum inflammation in the beginning (day 0) and in the middle (day 20) of the study are equal to the corresponding day numbers.
#
# First we'll get the max's for day 0 and day 20:
# ```
# max_inflammation_0 = numpy.max(data, axis=0)[0]
# max_inflammation_20 = numpy.max(data, axis=0)[20]
# print(max_inflammation_0)
# print(max_inflammation_20)
# ```
max_inflammation_0 = numpy.max(data, axis=0)[0]
max_inflammation_20 = numpy.max(data, axis=0)[20]
print(max_inflammation_0)
print(max_inflammation_20)
# and then we'll check them for suspicious values:
# ```
# if max_inflammation_0 == 0 and max_inflammation_20 == 20:
# print('Suspicious looking maxima!')
# ```
#
# Note, we can add parentheses `( )` to improve clarity (and order of evaluation)
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')
# We also saw a different problem in the third dataset; the minima per day were all zero (looks like a healthy person snuck into our study).
# ```
# data = numpy.loadtxt(fname='data/inflammation-03.csv', delimiter=',')
# min_plot = matplotlib.pyplot.plot(numpy.min(data, axis=0))
# matplotlib.pyplot.show()
# ```
# Let's have quick look at the all the `inflammation-03.csv` data set to confirm our suspicions
# ```
# matplotlib.pyplot.imshow(data)
# ```
# and look closely at the last row.
data = numpy.loadtxt(fname='data/inflammation-03.csv', delimiter=',')
min_plot = matplotlib.pyplot.plot(numpy.min(data, axis=0))
matplotlib.pyplot.show()
matplotlib.pyplot.imshow(data)
# We can also get python to check for this condition by summing all the minima and testing the result:
# ```
# if numpy.sum(numpy.min(data, axis=0)) == 0:
# print('Minima add up to zero!')
# ```
#
# this will become an `elif` in our script.
if numpy.sum(numpy.min(data, axis=0)) == 0:
print('Minima add up to zero!')
# And if neither of these conditions are true, we can use an `else` statement to give the all-clear
#
# So let's combine all these tests into a single script:
# ```
# data = numpy.loadtxt(fname='data/inflammation-01.csv', delimiter=',')
#
# max_inflammation_0 = numpy.max(data, axis=0)[0]
# max_inflammation_20 = numpy.max(data, axis=0)[20]
#
# if max_inflammation_0 == 0 and max_inflammation_20 == 20:
# print('Suspicious looking maxima!')
# elif numpy.sum(numpy.min(data, axis=0)) == 0:
# print('Minima add up to zero!')
# else:
# print('Seems OK!')
# ```
#
# give it a go with a selection of data files (eg, `inflammation-01.csv`, `inflammation-03.csv`)
# +
data = numpy.loadtxt(fname='data/inflammation-01.csv', delimiter=',')
max_inflammation_0 = numpy.max(data, axis=0)[0]
max_inflammation_20 = numpy.max(data, axis=0)[20]
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')
elif numpy.sum(numpy.min(data, axis=0)) == 0:
print('Minima add up to zero!')
else:
print('Seems OK!')
# +
data = numpy.loadtxt(fname='data/inflammation-03.csv', delimiter=',')
max_inflammation_0 = numpy.max(data, axis=0)[0]
max_inflammation_20 = numpy.max(data, axis=0)[20]
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')
elif numpy.sum(numpy.min(data, axis=0)) == 0:
print('Minima add up to zero!')
else:
print('Seems OK!')
# -
# ### Minima add up to zero!
# In this script, we have asked Python to do something different depending on the condition of our data. Here we printed messages in all cases, but we could also imagine not using the else catch-all so that messages are only printed when something is wrong, freeing us from having to manually examine every plot for features we've seen before.
# ## Exercises
# #### How Many Paths?
# Consider this code:
# ```
# if 4 > 5:
# print('A')
# elif 4 == 5:
# print('B')
# elif 4 < 5:
# print('C')
# ```
# Which of the following would be printed if you were to run this code? Why did you pick this answer?
# ```
# A
# B
# C
# B and C
# ```
if 4 > 5:
print('A')
elif 4 == 5:
print('B')
elif 4 < 5:
print('C')
# Solution: C
# ### What Is Truth?
#
# `True` and `False` booleans are not the only values in Python that are *true* and *false*.
#
# In fact, any value can be used in an `if` or `elif`. After reading and running the code below, explain what the rule is for which values are considered true and which are considered false.
#
# ```
# if True:
# print('True is true - not very surprising')
# if '':
# print('empty string is true')
# if 'word':
# print('word is true')
# if []:
# print('empty list is true')
# if [1, 2, 3]:
# print('non-empty list is true')
# if 0:
# print('zero is true')
# if 1:
# print('one is true')
# ```
if True:
print('True is true - not very surprising')
if '':
print('empty string is true')
if 'word':
print('word is true')
if []:
print('empty list is true')
if [1, 2, 3]:
print('non-empty list is true')
if 0:
print('zero is true')
if 1:
print('one is true')
# Solution:
# ### That's Not Not What I Meant
# Sometimes it is useful to check whether some condition is *not true*. The boolean operator `not` can do this explicitly.
#
# After reading and running the code below, write some `if` statements that use `not` to test the rule that you formulated in the previous challenge.
# ```
# if not '':
# print('empty string is not true')
# if not 'word':
# print('word is not true')
# if not not True:
# print('not not True is true')
# ```
if not '':
print('empty string is not true')
if not 'word':
print('word is not true')
if not not True:
print('not not True is true')
if not []:
print('empty list is not true')
if not 0:
print('zero is not true')
# Solution:
# ### Close Enough
# Write some conditions that print True if the variable a is within 10% of the variable b and False otherwise.
# Compare your implementation with your partner's: do you get the same answer for all possible pairs of numbers?
# +
a = input()
a = int(a)
b = 10
if abs(a-b)/(a-b) < 0.1:
print("True")
else:
print("False")
# -
# ### In-Place Operators
# Python (and most other languages in the C family) provides in-place operators that work like this:
# ```
# x = 1 # original value
# x += 1 # add one to x, assigning result back to x
# x *= 3 # multiply x by 3
# print(x)
# ```
#
# Write some code that sums the positive and negative numbers in a list separately, using in-place operators. Do you think the result is more or less readable than writing the same without in-place operators?
# +
positive_sum = 0
negative_sum = 0
numbers = [1,-2,3,4,-5,0, 6]
for i in numbers:
print(i)
print('Sum of Positive numbers:', positive_sum)
# -
# ### Sorting a List Into Buckets
#
# In our data folder, large data sets are stored in files whose names start with "inflammation-" and small data sets – in files whose names start with "small-". We also have some other files that we do not care about at this point. We'd like to break all these files into three lists called `large_files`, `small_file`, and `other_files`, respectively.
#
# Add code to the template below to do this. Note that the string method `startswith` returns `True` if and only if the string it is called on starts with the string passed as an argument, e.g.:
# ```
# print("String".startswith("Str"))
# print("String".startswith("somethingelse"))
# ```
#
# note that `startswith` is case sensitive.
# Use the following Python code as your starting point:
files = ['inflammation-01.csv',
'myscript.py',
'inflammation-02.csv',
'small-01.csv',
'small-02.csv']
large_files = []
small_files = []
other_files = []
# Your solution should:
#
# - loop over the names of the files
# - figure out which group each filename belongs
# - append the filename to that list
# - In the end the three lists should be:
# ```
# large_files = ['inflammation-01.csv', 'inflammation-02.csv']
# small_files = ['small-01.csv', 'small-02.csv']
# other_files = ['myscript.py']
# ```
# ### Counting Vowels
# Write a loop that counts the number of vowels in a character string.
#
# Test it on a few individual words and full sentences.
#
# Once you are done, compare your solution to your neighbour's. Did you make the same decisions about how to handle the letter 'y' (which some people think is a vowel, and some do not)?
sentence = "Python is Fun!"
for c in sentence:
print(c)
# ## Key Points
#
# - Use the `if` keyword to start a conditional statement, the `elif` keyword to provide additional tests, and the `else` keyword to provide a default.
#
# - The bodies of the branches of conditional statements must be indented.
#
# - Use `==` to test for equality.
#
# - X `and` Y is only true if both X and Y are true.
#
# - X `or` Y is true if either X or Y, or both, are true.
#
# - Zero `0`, the empty string `""`, and the empty list `[]` are considered false; all other numbers, strings, and lists are considered true.
#
# - `True` and `False` represent truth values.
# ### Save, and version control your changes
#
# - save your work: `File -> Save`
# - add all your changes to your local repository: `Terminal -> git add .`
# - commit your updates a new Git version: `Terminal -> git commit -m "End of Episode 5"`
# - push your latest commits to GitHub: `Terminal -> git push`
| lessons/python/ep5-conditionals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_pytorch_p36
# language: python
# name: conda_pytorch_p36
# ---
# # Moon Data Classification
#
# In this notebook, you'll be tasked with building and deploying a **custom model** in SageMaker. Specifically, you'll define and train a custom, PyTorch neural network to create a binary classifier for data that is separated into two classes; the data looks like two moon shapes when it is displayed, and is often referred to as **moon data**.
#
# The notebook will be broken down into a few steps:
# * Generating the moon data
# * Loading it into an S3 bucket
# * Defining a PyTorch binary classifier
# * Completing a training script
# * Training and deploying the custom model
# * Evaluating its performance
#
# Being able to train and deploy custom models is a really useful skill to have. Especially in applications that may not be easily solved by traditional algorithms like a LinearLearner.
#
# ---
# Load in required libraries, below.
# +
# data
import pandas as pd
import numpy as np
from sklearn.datasets import make_moons
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Generating Moon Data
#
# Below, I have written code to generate some moon data, using sklearn's [make_moons](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_moons.html) and [train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html).
#
# I'm specifying the number of data points and a noise parameter to use for generation. Then, displaying the resulting data.
# +
# set data params
np.random.seed(0)
num_pts = 600
noise_val = 0.25
# generate data
# X = 2D points, Y = class labels (0 or 1)
X, Y = make_moons(num_pts, noise=noise_val)
# Split into test and training data
X_train, X_test, Y_train, Y_test = train_test_split(X, Y,
test_size=0.25, random_state=1)
# -
# plot
# points are colored by class, Y_train
# 0 labels = purple, 1 = yellow
plt.figure(figsize=(8,5))
plt.scatter(X_train[:,0], X_train[:,1], c=Y_train)
plt.title('Moon Data')
plt.show()
# ## SageMaker Resources
#
# The below cell stores the SageMaker session and role (for creating estimators and models), and creates a default S3 bucket. After creating this bucket, you can upload any locally stored data to S3.
# sagemaker
import boto3
import sagemaker
from sagemaker import get_execution_role
# +
# SageMaker session and role
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
# default S3 bucket
bucket = sagemaker_session.default_bucket()
# -
# ### EXERCISE: Create csv files
#
# Define a function that takes in x (features) and y (labels) and saves them to one `.csv` file at the path `data_dir/filename`. SageMaker expects `.csv` files to be in a certain format, according to the [documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/cdf-training.html):
# > Amazon SageMaker requires that a CSV file doesn't have a header record and that the target variable is in the first column.
#
# It may be useful to use pandas to merge your features and labels into one DataFrame and then convert that into a `.csv` file. When you create a `.csv` file, make sure to set `header=False`, and `index=False` so you don't include anything extraneous, like column names, in the `.csv` file.
# +
import os
def make_csv(x, y, filename, data_dir):
'''Merges features and labels and converts them into one csv file with labels in the first column.
:param x: Data features
:param y: Data labels
:param file_name: Name of csv file, ex. 'train.csv'
:param data_dir: The directory where files will be saved
'''
# make data dir, if it does not exist
if not os.path.exists(data_dir):
os.makedirs(data_dir)
output_path = os.path.join(data_dir,filename)
# your code here
pd.concat([pd.DataFrame(y), pd.DataFrame(x)], axis=1).to_csv(output_path, header=False, index=False)
# nothing is returned, but a print statement indicates that the function has run
print('Path created: '+str(data_dir)+'/'+str(filename))
# -
# The next cell runs the above function to create a `train.csv` file in a specified directory.
# +
data_dir = 'data_moon' # the folder we will use for storing data
name = 'train.csv'
# create 'train.csv'
make_csv(X_train, Y_train, name, data_dir)
# -
# ### Upload Data to S3
#
# Upload locally-stored `train.csv` file to S3 by using `sagemaker_session.upload_data`. This function needs to know: where the data is saved locally, and where to upload in S3 (a bucket and prefix).
# +
# specify where to upload in S3
prefix = 'sagemaker/moon-data'
# upload to S3
input_data = sagemaker_session.upload_data(path=data_dir, bucket=bucket, key_prefix=prefix)
print(input_data)
# -
# Check that you've uploaded the data, by printing the contents of the default bucket.
# iterate through S3 objects and print contents
for obj in boto3.resource('s3').Bucket(bucket).objects.all():
print(obj.key)
# ---
# # Modeling
#
# Now that you've uploaded your training data, it's time to define and train a model!
#
# In this notebook, you'll define and train a **custom PyTorch model**; a neural network that performs binary classification.
#
# ### EXERCISE: Define a model in `model.py`
#
# To implement a custom classifier, the first thing you'll do is define a neural network. You've been give some starting code in the directory `source`, where you can find the file, `model.py`. You'll need to complete the class `SimpleNet`; specifying the layers of the neural network and its feedforward behavior. It may be helpful to review the [code for a 3-layer MLP](https://github.com/udacity/deep-learning-v2-pytorch/blob/master/convolutional-neural-networks/mnist-mlp/mnist_mlp_solution.ipynb).
#
# This model should be designed to:
# * Accept a number of `input_dim` features
# * Create some linear, hidden layers of a desired size
# * Return **a single output value** that indicates the class score
#
# The returned output value should be a [sigmoid-activated](https://pytorch.org/docs/stable/nn.html#sigmoid) class score; a value between 0-1 that can be rounded to get a predicted, class label.
#
# Below, you can use !pygmentize to display the code in the `model.py` file. Read through the code; all of your tasks are marked with TODO comments. You should navigate to the file, and complete the tasks to define a `SimpleNet`.
# !pygmentize source/model.py
# ## Training Script
#
# To implement a custom classifier, you'll also need to complete a `train.py` script. You can find this in the `source` directory.
#
# A typical training script:
#
# * Loads training data from a specified directory
# * Parses any training & model hyperparameters (ex. nodes in a neural network, training epochs, etc.)
# * Instantiates a model of your design, with any specified hyperparams
# * Trains that model
# * Finally, saves the model so that it can be hosted/deployed, later
#
# ### EXERCISE: Complete the `train.py` script
#
# Much of the training script code is provided for you. Almost all of your work will be done in the if __name__ == '__main__': section. To complete the `train.py` file, you will:
#
# * Define any additional model training hyperparameters using `parser.add_argument`
# * Define a model in the if __name__ == '__main__': section
# * Train the model in that same section
#
# Below, you can use !pygmentize to display an existing train.py file. Read through the code; all of your tasks are marked with TODO comments.
# !pygmentize source/train.py
# ### EXERCISE: Create a PyTorch Estimator
#
# You've had some practice instantiating built-in models in SageMaker. All estimators require some constructor arguments to be passed in. When a custom model is constructed in SageMaker, an **entry point** must be specified. The entry_point is the training script that will be executed when the model is trained; the `train.py` function you specified above!
#
# See if you can complete this task, instantiating a PyTorch estimator, using only the [PyTorch estimator documentation](https://sagemaker.readthedocs.io/en/stable/sagemaker.pytorch.html) as a resource. It is suggested that you use the **latest version** of PyTorch as the optional `framework_version` parameter.
#
# #### Instance Types
#
# It is suggested that you use instances that are available in the free tier of usage: `'ml.c4.xlarge'` for training and `'ml.t2.medium'` for deployment.
# +
# import a PyTorch wrapper
from sagemaker.pytorch import PyTorch
# specify an output path
output_path='s3://{}/{}/'.format(bucket, prefix)
# instantiate a pytorch estimator
estimator = PyTorch(role=role,
train_instance_count=1,
train_instance_type='ml.c4.xlarge',
output_path=output_path, # specified, above
framework_version='1.4.0',
entry_point='train.py',
source_dir='source',
sagemaker_session=sagemaker_session,
hyperparameters={
'input_dim':2,
'hidden_dim':20,
'output_dim':1,
'epochs':80
})
# -
# ## Train the Estimator
#
# After instantiating your estimator, train it with a call to `.fit()`. The `train.py` file explicitly loads in `.csv` data, so you do not need to convert the input data to any other format.
# %%time
# train the estimator on S3 training data
estimator.fit({'train': input_data})
# ## Create a Trained Model
#
# PyTorch models do not automatically come with `.predict()` functions attached (as many Scikit-learn models do, for example) and you may have noticed that you've been give a `predict.py` file. This file is responsible for loading a trained model and applying it to passed in, numpy data. When you created a PyTorch estimator, you specified where the training script, `train.py` was located.
#
# > How can we tell a PyTorch model where the `predict.py` file is?
#
# Before you can deploy this custom PyTorch model, you have to take one more step: creating a `PyTorchModel`. In earlier exercises you could see that a call to `.deploy()` created both a **model** and an **endpoint**, but for PyTorch models, these steps have to be separate.
#
# ### EXERCISE: Instantiate a `PyTorchModel`
#
# You can create a `PyTorchModel` (different that a PyTorch estimator) from your trained, estimator attributes. This model is responsible for knowing how to execute a specific `predict.py` script. And this model is what you'll deploy to create an endpoint.
#
# #### Model Parameters
#
# To instantiate a `PyTorchModel`, ([documentation, here](https://sagemaker.readthedocs.io/en/stable/sagemaker.pytorch.html#sagemaker.pytorch.model.PyTorchModel)) you pass in the same arguments as your PyTorch estimator, with a few additions/modifications:
# * **model_data**: The trained `model.tar.gz` file created by your estimator, which can be accessed as `estimator.model_data`.
# * **entry_point**: This time, this is the path to the Python script SageMaker runs for **prediction** rather than training, `predict.py`.
#
# +
# %%time
# importing PyTorchModel
from sagemaker.pytorch import PyTorchModel
# Create a model from the trained estimator data
# And point to the prediction script
model = PyTorchModel(role=role,
framework_version='1.4.0',
entry_point='predict.py',
source_dir='source',
model_data=estimator.model_data
)
# -
# ### EXERCISE: Deploy the trained model
#
# Deploy your model to create a predictor. We'll use this to make predictions on our test data and evaluate the model.
# %%time
# deploy and create a predictor
predictor = model.deploy(initial_instance_count =1, instance_type='ml.t2.medium')
# ---
# ## Evaluating Your Model
#
# Once your model is deployed, you can see how it performs when applied to the test data.
#
# The provided function below, takes in a deployed predictor, some test features and labels, and returns a dictionary of metrics; calculating false negatives and positives as well as recall, precision, and accuracy.
# +
# code to evaluate the endpoint on test data
# returns a variety of model metrics
def evaluate(predictor, test_features, test_labels, verbose=True):
"""
Evaluate a model on a test set given the prediction endpoint.
Return binary classification metrics.
:param predictor: A prediction endpoint
:param test_features: Test features
:param test_labels: Class labels for test data
:param verbose: If True, prints a table of all performance metrics
:return: A dictionary of performance metrics.
"""
# rounding and squeezing array
test_preds = np.squeeze(np.round(predictor.predict(test_features)))
# calculate true positives, false positives, true negatives, false negatives
tp = np.logical_and(test_labels, test_preds).sum()
fp = np.logical_and(1-test_labels, test_preds).sum()
tn = np.logical_and(1-test_labels, 1-test_preds).sum()
fn = np.logical_and(test_labels, 1-test_preds).sum()
# calculate binary classification metrics
recall = tp / (tp + fn)
precision = tp / (tp + fp)
accuracy = (tp + tn) / (tp + fp + tn + fn)
# print metrics
if verbose:
print(pd.crosstab(test_labels, test_preds, rownames=['actuals'], colnames=['predictions']))
print("\n{:<11} {:.3f}".format('Recall:', recall))
print("{:<11} {:.3f}".format('Precision:', precision))
print("{:<11} {:.3f}".format('Accuracy:', accuracy))
print()
return {'TP': tp, 'FP': fp, 'FN': fn, 'TN': tn,
'Precision': precision, 'Recall': recall, 'Accuracy': accuracy}
# -
# ### Test Results
#
# The cell below runs the `evaluate` function.
#
# The code assumes that you have a defined `predictor` and `X_test` and `Y_test` from previously-run cells.
# get metrics for custom predictor
metrics = evaluate(predictor, X_test, Y_test, True)
# ## Delete the Endpoint
#
# Finally, I've add a convenience function to delete prediction endpoints after we're done with them. And if you're done evaluating the model, you should delete your model endpoint!
# Accepts a predictor endpoint as input
# And deletes the endpoint by name
def delete_endpoint(predictor):
try:
boto3.client('sagemaker').delete_endpoint(EndpointName=predictor.endpoint)
print('Deleted {}'.format(predictor.endpoint))
except:
print('Already deleted: {}'.format(predictor.endpoint))
# delete the predictor endpoint
delete_endpoint(predictor)
# ## Final Cleanup!
#
# * Double check that you have deleted all your endpoints.
# * I'd also suggest manually deleting your S3 bucket, models, and endpoint configurations directly from your AWS console.
#
# You can find thorough cleanup instructions, [in the documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/ex1-cleanup.html).
# ---
# # Conclusion
#
# In this notebook, you saw how to train and deploy a custom, PyTorch model in SageMaker. SageMaker has many built-in models that are useful for common clustering and classification tasks, but it is useful to know how to create custom, deep learning models that are flexible enough to learn from a variety of data.
| Moon_Data/Moon_Classification_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PSF Generation Validation
#
# This validation process involves the following:
#
# - Determine a set of PSF configurations to test
# - Create a PSF for each of those configurations using both Flowdec and PSFGenerator
# - Store any more detailed visualizations/statistics in a separate notebook for each configuration
# - Aggregate top-line statistics like SSIM and PSNR to visualize across all configurations
import papermill as pm
import os
import os.path as osp
import pandas as pd
from flowdec import psf as fd_psf
nb_dir = 'results'
# Show default arguments and use these to derive test cases
fd_psf.GibsonLanni().get_arg_parser().print_help()
# +
# Build test case argument sets
args = [
{
'name': 'particle-pos-p1um',
# "num_samples" is increased here from default of 1000
# because approximations are significantly further off otherwise
'args': {'psf_args': '{"pz": 1.0, "num_samples": 2000}'}
}, {
'name': 'particle-pos-n1um',
'args': {'psf_args': '{"pz": -1.0, "num_samples": 2000}'}
}, {
'name': 'red-light',
'args': {'psf_args': '{"pz": 0.0, "wavelength": 0.7}'}
}, {
'name': 'blue-light',
# "min_wavelength" is lowered here from default of .436 microns
# because approximations are significantly further off otherwise
'args': {'psf_args': '{"pz": 0.0, "wavelength": 0.45, "min_wavelength": 0.35}'}
}, {
'name': 'green-light',
'args': {'psf_args': '{"pz": 0.0, "wavelength": 0.55, "min_wavelength": 0.35}'}
}, {
'name': 'low-zres',
'args': {'psf_args': '{"pz": 0.0, "res_axial": 1.5, "num_samples": 2000}'}
}, {
'name': 'hi-zres',
'args': {'psf_args': '{"pz": 0.0, "res_axial": 0.1}'}
}, {
'name': 'large-xy',
'args': {'psf_args': '{"pz": 0.0, "size_x": 512, "size_y": 400, "size_z": 16}'}
}, {
'name': 'large-z',
'args': {'psf_args': '{"pz": 0.0, "size_x": 32, "size_y": 64, "size_z": 128}'}
},
]
# -
# Run a comparison process for each configuration defined above
os.makedirs(nb_dir, exist_ok=True)
for arg in args:
print('Executing comparisons for configuration "{}"'.format(arg['name']))
nb_template = 'template.ipynb'
out_path = osp.join(nb_dir, arg['name'] + '.ipynb')
pm.execute_notebook(nb_template, out_path, parameters=arg['args'])
# Aggregate results across all configurations
df = pm.read_notebooks(nb_dir).dataframe
df['key'] = df['key'].apply(lambda v: v.replace('.ipynb', ''))
df.head()
def extract_series(df, prop):
df = df[df['name'] == prop].set_index('key')
return pd.concat([pd.Series(r['value']).rename('value').to_frame().assign(key=k) for k, r in df.iterrows()])
df_meas = extract_series(df, 'measures')
df_meas.head()
# ## Scores
#
# This table shows SSIM and PSNR scores (between Flowdec and PSFGenerator) on the original PSF image scale as well as a logarithmic scale:
# **SSIM**
pd.concat([
df_meas.loc['ssim_log'].set_index('key')['value'].rename('ssim_log_scale'),
df_meas.loc['ssim_original'].set_index('key')['value'].rename('ssim_original_scale')
], axis=1)
# **PSNR**
pd.concat([
df_meas.loc['psnr_log'].set_index('key')['value'].rename('psnr_log_scale'),
df_meas.loc['psnr_original'].set_index('key')['value'].rename('psnr_original_scale')
], axis=1)
# ## Maximum Differences
#
# This section shows the maximum difference for any single voxel value between Flowdec and PSFGenerator PSFs versus the range of the original PSF images:
# +
def extract_df(df, prop):
df = df[df['name'] == prop].set_index('key')
return pd.concat([pd.DataFrame(r['value']).assign(key=k) for k, r in df.iterrows()])
df_stat = extract_df(df, 'df_original')
# -
df_extrema = pd.concat([
df_stat.loc['max'].set_index('key')['Diff'].rename('max_diff'),
df_stat.loc['min'].set_index('key')['Diff'].rename('min_diff'),
df_stat.loc['max'].set_index('key')['Flowdec'].rename('Fowdec PSF Max Value'),
df_stat.loc['min'].set_index('key')['Flowdec'].rename('flowdec PSF Min Value'),
df_stat.loc['max'].set_index('key')['PSFGenerator'].rename('PSFGenerator PSF Max Value'),
df_stat.loc['min'].set_index('key')['PSFGenerator'].rename('PSFGenerator PSF Max Value'),
], axis=1)
df_extrema['Absolute Max Difference'] = df_extrema[['min_diff', 'max_diff']].abs().max(axis=1)
df_extrema = df_extrema.drop(['min_diff', 'max_diff'], axis=1)
df_extrema
ax = df_extrema.sort_values('Absolute Max Difference').plot(figsize=(18, 6), kind='bar', legend='side')
ax.set_yscale('log')
ax.set_xlabel('Test Case')
ax.set_ylabel('Voxel Value')
ax.set_title('Dynamic Range of PSFs vs Largest Difference by Test Case')
| python/validation/psfgeneration/validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NAXofsM3bmti"
# # Notebook
# *by <NAME> and <NAME>*
#
# Local helioseismology techniques allow the detection of active regions in the non-visible solar hemisphere (far-side) by analyzing the oscillations in the visible side of the Sun. However, this identification is challenged by the low signal-to-noise of the seismic data, and only strong active regions can be reliably detected.
#
# In this notebook, we will show a method to improve the detection of active regions in far-side seismic maps using a machine learning algorithm.
#
# This work is published in [Felipe & <NAME>, 2019, A&A, 632, 82](https://www.aanda.org/articles/aa/abs/2019/12/aa36838-19/aa36838-19.html)
#
#
# 
# Detection of a far-side active region.
#
#
# + [markdown] colab_type="text" id="a-tx1jmDe_SJ"
# ## Introduction
# One of the most remarkable applications of local helioseismology is the
# detection of active regions in the non-visible hemisphere of the Sun (on the far side).
# This was first achieved using the technique of helioseismic holography
# ([Lindsey & Braun 2000, Science, 287, 1799](https://science.sciencemag.org/content/287/5459/1799.full), [Braun & Lindsey 2001, ApJ, 560, 189](https://iopscience.iop.org/article/10.1086/324323)).
#
# Helioseismic holography uses the wavefield measured in a region of the solar surface (called "pupil") to determine the wave field at a focus point that is located at the surface or at a certain depth. This inference is
# performed assuming that the observed wave field at the pupil (e.g., the line-of-sight Doppler velocity) is produced by waves converging toward the focus point or waves diverging from that point. Far-side helioseismic holography is a particular application of this method, where the pupil is located at the near-side hemisphere and the focus points are located at the surface in the far-side hemisphere (see [Lindsey & Braun 2017, Space Weather, 15, 761](https://ui.adsabs.harvard.edu/abs/2017SpWea..15..761L/abstract)).
#
# The identification of active regions is founded
# on the fact that they introduce a phase shift between ingoing and outgoing waves. This phase shift (which can be characterized as a travel-time shift) is mainly due to the depression of the photosphere in
# magnetized regions, which causes the upcoming waves to reach the upper turning point a few seconds earlier in active regions than in quiet-Sun regions ([Felipe et al. 2017, A&A, 604, 126](https://ui.adsabs.harvard.edu/link_gateway/2017A%26A...604A.126F/PUB_HTML)). In this way, when an active region is located at the focus point, a negative phase shift (reduction in the travel
# time) is found.
#
# + [markdown] colab_type="text" id="j7QQkI664yAI"
# ## Why using a neural network approach?
#
# One of the main limitations of far-side helioseismology is the reduced signal-to-noise ratio. The signature of an active region detected on the far side has a
# signal-to-noise ratio of around 10, which means that only large and strong active regions can be reliably detected in far-side phase-shift maps (about several hundered active regions per solar cycle).
#
# Our aim in this work is to apply convolutional neural networks to learn a very fast and robust mapping between consecutive maps of estimated seismic maps and the probability map of the presence of an active region on the far side. The recent success of machine learning is no doubt a consequence of our ability to train very deep neural networks (DNNs). DNNs can be seen as a very flexible and differentiable parametric mapping between an input space and an output space. These highly parameterized
# DNNs are then tuned by optimizing a loss function that measures the ability of the DNN to map the input space onto the output space over a predefined training set. The combination of loss function and specific architecture has to be chosen to solve the specific problem at hand.
#
# Arguably the largest number of applications of DNNs has been in computer vision. Problems belonging to the realm of machine vision can hardly be solved using classical methods, be they based on machine learning or on rule-based methods. Only now, with the application of very DNNs, have we been able to produce real advances. Applications in science, and specifically in astrophysics and solar physics, have leveraged the results of machine vision to solve problems that were difficult or impossible to deal with in the past with classical techniques.
# + [markdown] colab_type="text" id="yBuqxmQRDl39"
# ## Description of the neural network
# In this notebook, we present a description of the neural network developed for the detection of far-side active regions. We have included a running example of the application of the network and the tools employed for the interpretation of the results.
#
# We have omitted the materials employed for the training set. They are publicly available and their locations are indicated. We have described the transformations applied to the original data to convert them into the data fed to the neural network for the training.
# + [markdown] colab_type="text" id="wzqcG6esWc12"
# ### Training set
# We have designed a neural network that can identify the presence of active
# regions on the far side. As input, the network uses far-side phase-shift maps
# computed using helioseismic holography. As a proxy for the presence of active
# regions, we employed Helioseismic and Magnetic Imager (HMI) magnetograms measured on the near side (facing Earth). The details of the data are discussed in the following sections.
# The training set that we describe in this section was used to supervise the parameter tuning of the neural network with the aim of generalizing this to
# new data.
# + [markdown] colab_type="text" id="eJbD8wSyEujP"
# #### HMI magnetograms
# The HMI magnetograms are one of the data products from the Solar Dynamics Observatory available through the Joint Science Operations Center (JSOC). In order to facilitate the comparison with the far-side seismic maps (next section), we are interested in magnetograms that are remapped onto a Carrington coordinate grid. We used data from the JSOC series *hmi.Mldailysynframe\_720s*. This data series contains synoptic maps constructed of HMI magnetograms collected over a 27-day solar rotation, where the first 120 degrees in longitude are replaced by data within 60 degrees of the central meridian of the visible hemisphere observed approximately at one time. These
# maps are produced daily at 12 UT. We only employed the 120 degrees in longitude
# including the magnetogram visible on the disk at one time. Magnetograms between
# 2010 June 1 (the first date available for the *hmi.Mldailysynframe\_720s*
# data) and 2018 October 26 were extracted. Because one magnetogram is taken per day, this means a total of 3066 magnetograms.
#
# Because new active regions emerge and old regions decay,
# magnetograms obtained on the near side are an inaccurate characterization of the
# active regions on the far side half a rotation earlier or later. We have
# partially corrected this problem. The far-side maps are associated with the
# magnetogram that is obtained when the seismically probed region has fully rotated to the
# Earth side, that is, 13.5 days after the measurement of the far-side map. We
# removed the active regions that emerge on the near side because they were absent when the far-side seismic data were taken. In order to identify the
# emerging active regions, we have employed the Solar Region Summary (SRS)
# files (available at [ftp.swpc.noaa.gov/pub/warehouse/](ftp://ftp.swpc.noaa.gov/pub/warehouse/), where the NOAA registered active regions are listed. All the active regions that appear for the first time at a longitude greater than $-60^{\circ}$ (where 0 corresponds to the central meridian of the visible hemisphere and the minus sign indicates the eastern hemisphere) were masked in the magnetograms. The value of the magnetogram was set to zero in an area 15 degrees wide in longitude and 12 degrees wide in latitude, centered in the location of the active region reported in the SRS file of that date (after correcting for the longitude because we employed magnetograms retrieved at 12 UT and in the SRS files the location of the active regions are reported for 24 UT on the previous day). The active regions that emerge in the visible hemisphere too close to an active region that had appeared on the eastern limb due to the solar rotation were not masked. Of the 1652 active regions labeled by NOAA during the temporal period employed for the training set, 967 were masked because they emerged in the visible hemisphere.
#
# The neural network is trained with binary maps, where the zeros correspond to quiet regions and the ones to active regions. This binary mask is built from the corrected magnetograms as follows. A Gaussian smoothing with a standard deviation of 3 degrees was applied to the corrected magnetograms. This smoothing removed all small-scale activity in the map and facilitated the segmentation of active regions of importance in the magnetogram.
# Then, regions with a magnetic flux higher than 30 Mx cm$^2$ were identified as active regions (and set to 1), and regions with lower magnetic flux were set to 0. The middle panel in the bottom row from Fig. 1 shows the magnetogram after the active regions that emerged in the visible solar hemisphere were removed and after Gaussian smoothing was applied. The active region visible in the original magnetogram (bottom left panel in Fig. 1) at a longitude $-30^{\circ}$ and a latitude $-5^{\circ}$ emerged on the near side and was therefore masked. The bottom right panel of Fig. 1 shows the binary map in which the location of the remaining active regions is indicated, those whose magnetic flux is above the selected threshold. Their positions match that of some regions with strong negative travel times in the seismic maps from about half a rotation earlier (case "t-13.0" in the top row of Fig. 1).
# 
# **Fig. 1.** Example of one of the elements from the training set. Panels in the top row show 11 far-side seismic maps, each of them obtained from the analysis of 24 h of HMI Doppler data. The horizontal axis is the longitude (a total of 120°) and the vertical axis is the latitude (between −72° and 72°). The label above the panels indicates the number of days prior to the time t when the corresponding magnetogram was acquired (in this example, t is 2015 December 10 at 12:00 UT). Bottom row: magnetograms we used as a proxy for the presence of active regions. Left panel: original magnetogram in heliospheric coordinates, middle panel: magnetogram after active regions that emerged in the near side are removed and after a Gaussian smoothing was applied, and right panel: binary map in which a value of 1 indicates the presence of an active region in the locations whose magnetic flux in the smoothed magnetogram is above the selected threshold. Red contours in the bottom left panel delimit the regions where the binary map is 1. The neural network is trained by associating the 11 far-side seismic maps (top row) with the binary map.
#
#
#
# + [markdown] colab_type="text" id="jiGXTckVWnvL"
# #### Far-side phase-shift maps
#
# Phase-shift maps of the far-side region of the Sun are available through JSOC. They are computed from
# HMI Doppler data using temporal series of one or five days. The processing of
# series of five days is a novel approach since 2014, introduced to improve the
# signal-to-noise ratio of the phase-shift maps. They are provided in Carrington
# heliographic coordinates with a cadence of 12 hours (maps are obtained at 0 and
# 12 UT). In this work, we focus on the far-side maps computed from 24 hours
# of Doppler data. We employed far-side maps between 2010 May 18 and 2018 October 12. For each map, we selected a $120^{\circ}$ region in longitude centered at the Carrington longitude of the central meridian of the visible hemisphere 13.5 days after the date of the far-side map. In this way, corrected magnetograms from which
# the new active regions are removed are associated with far-side maps that sample the same region in longitude. The training employed 11 consecutive far-side maps for each corrected magnetogram, which improved the seismic signal. These 11 consecutive far-side maps correspond to six days of data. The latitude span of the maps is
# between $-72^{\circ}$ and $72^{\circ}$. We chose a sampling of $1^{\circ}$ in both latitude and longitude.
# + [markdown] colab_type="text" id="nzIk_0FORyPd"
#
# ##Architecture
# The neural network of choice in
# this work is a U-net ([Ronneberger et al. 2015, ArXiv](https://arxiv.org/abs/1505.04597)), a fully
# convolutional architecture that has been used extensively for dense segmentation of images and displayed in Fig. 2. The U-net
# is an encoder-decoder
# network, in which the input is
# successively reduced in size through contracting layers and is finally increased in size through
# expanding layers. This encoder-decoder architecture has three main advantages, all
# of them a consequence of the contracting and expanding layers. The first
# advantage is that the contracting layers reduce the size of the images at each step.
# This makes the network faster because convolutions have to be carried out
# over smaller images. The second advantage is that this contraction couples
# together pixels in the input image that were far apart, so that smaller kernels
# can be used in convolutional layers (we used $3 \times 3$ kernels) and the network
# is able to better exploit multiscale information. The final
# advantage is a consequence of the skip connections (gray
# arrows), which facilitates training by explicitly
# propagating multiscale information from the contracting layers to the
# expanding layers.
#
# As shown in Fig. 2, the specific U-net architecture
# we used in this work is a combination of several
# differentiable operations. The first operation, indicated with blue arrows, is
# the consecutive application of convolutions with 3$\times$3 kernels,
# batch normalization (BN), which normalizes the input so that its mean
# is close to zero and its variance close to unity (which is known to
# be an optimal range of values for neural networks to work best) and
# a rectified linear unit (ReLU) activation function, given
# by $\sigma(x)=\max(0,x)$. This combination
# Conv+BN+ReLU was repeated twice as indicated in
# the legend of Fig. 2. Red arrows refer to
# max-pooling [(Goodfellow et al. 2016, Deep Learning, MIT Press)](http://www.deeplearningbook.org/), which reduces the resolution
# of the images by a factor 2 by computing the maximum of all non-overlapping
# $2 \times 2$ patches in the image. The expanding layers again increase the size of the images
# through bilinear interpolation (green arrows) followed by convolutional layers.
# Additionally, the layers in the encoding part transfer information to the
# decoding part through skip connections (gray arrows), which greatly
# improves the ability and stability of the network.
# Finally, because the output is a probability map, we forced it to be in the $[0,1]$ range
# through a sigmoid activation function that was applied in the last layer after a final
# $1 \times 1$ convolution that we used to reduce the number of channels from 16 to 1.
#
# The neural
# network was trained by minimizing the binary cross entropy between the output of
# the network per pixel ($p_i$) and the binarized magnetograms ($y_i$), summed
# over all pixels in the output magnetogram ($N$),
# \begin{equation}
# \ell = -\frac{1}{N} \sum_{i=1}^{N} y_{i} \cdot \log p_i+
# \left(1-y_{i}\right) \cdot \log \left(1-p_i\right)
# .\end{equation}
# To optimize the previous loss function, we employed the Adam optimizer [(Kingma & Ba 2014, ArXiv)](https://arxiv.org/abs/1412.6980) with a
# constant learning rate of 3$\times$10$^{-4}$ during 300 epochs and a batch
# size of 30.
#
# The neural network can be downloaded from the repository [https://github.com/aasensio/farside](https://github.com/aasensio/farside).
#
# Here we show the model.
#
# 
# **Fig 2.** U-net architecture. The vertical extent of the blocks indicates the size of the image, and the numbers above each block shows the number of channels.
#
# + [markdown] colab_type="text" id="DRJ_bGcsHD2o"
# ### Model
# + colab={} colab_type="code" id="wDqQzSV5XpQq"
#We first import the necessary modules
import numpy as np
import torch
import torch.nn as nn
import torch.utils.data
import torch.nn.functional as F
# + colab={} colab_type="code" id="s0RTGTVBXyug"
class double_conv(nn.Module):
'''(conv => BN => ReLU) * 2'''
def __init__(self, in_ch, out_ch):
super(double_conv, self).__init__()
self.conv = nn.Sequential(
nn.Conv2d(in_ch, out_ch, 3, padding=1),
nn.BatchNorm2d(out_ch),
nn.ReLU(inplace=True),
nn.Conv2d(out_ch, out_ch, 3, padding=1),
nn.BatchNorm2d(out_ch),
nn.ReLU(inplace=True)
)
def forward(self, x):
x = self.conv(x)
return x
class inconv(nn.Module):
def __init__(self, in_ch, out_ch):
super(inconv, self).__init__()
self.conv = double_conv(in_ch, out_ch)
def forward(self, x):
x = self.conv(x)
return x
class down(nn.Module):
def __init__(self, in_ch, out_ch):
super(down, self).__init__()
self.mpconv = nn.Sequential(
nn.MaxPool2d(2),
double_conv(in_ch, out_ch)
)
def forward(self, x):
x = self.mpconv(x)
return x
class up(nn.Module):
def __init__(self, in_ch, out_ch, bilinear=True):
super(up, self).__init__()
self.bilinear = bilinear
# would be a nice idea if the upsampling could be learned too,
if not bilinear:
self.up = nn.ConvTranspose2d(in_ch//2, in_ch//2, 2, stride=2)
self.conv = double_conv(in_ch, out_ch)
def forward(self, x1, x2):
if (self.bilinear):
x1 = torch.nn.functional.interpolate(x1, scale_factor=2)
else:
x1 = self.up(x1)
# input is CHW
diffY = x2.size()[2] - x1.size()[2]
diffX = x2.size()[3] - x1.size()[3]
x1 = F.pad(x1, (diffX // 2, diffX - diffX//2,
diffY // 2, diffY - diffY//2))
# for padding issues, see
# https://github.com/HaiyongJiang/U-Net-Pytorch-Unstructured-Buggy/commit/0e854509c2cea854e247a9c615f175f76fbb2e3a
# https://github.com/xiaopeng-liao/Pytorch-UNet/commit/8ebac70e633bac59fc22bb5195e513d5832fb3bd
x = torch.cat([x2, x1], dim=1)
x = self.conv(x)
return x
class outconv(nn.Module):
def __init__(self, in_ch, out_ch):
super(outconv, self).__init__()
self.conv = nn.Conv2d(in_ch, out_ch, 1)
def forward(self, x):
x = self.conv(x)
return x
class UNet(nn.Module):
def __init__(self, n_channels, n_classes, n_hidden=64):
super(UNet, self).__init__()
self.inc = inconv(n_channels, n_hidden)
self.down1 = down(n_hidden, 2*n_hidden)
self.down2 = down(2*n_hidden, 4*n_hidden)
self.down3 = down(4*n_hidden, 8*n_hidden)
self.down4 = down(8*n_hidden, 8*n_hidden)
self.up1 = up(16*n_hidden, 4*n_hidden)
self.up2 = up(8*n_hidden, 2*n_hidden)
self.up3 = up(4*n_hidden, n_hidden)
self.up4 = up(2*n_hidden, n_hidden)
self.outc = outconv(n_hidden, n_classes)
def forward(self, x):
x1 = self.inc(x)
x2 = self.down1(x1)
x3 = self.down2(x2)
x4 = self.down3(x3)
x5 = self.down4(x4)
x = self.up1(x5, x4)
x = self.up2(x, x3)
x = self.up3(x, x2)
x = self.up4(x, x1)
x = self.outc(x)
return torch.sigmoid(x)
# + [markdown] colab_type="text" id="K4_TAFawb__k"
# ### Forward model
# + colab={} colab_type="code" id="zJTupgjcYTsg"
class deep_farside(object):
def __init__(self, maxbatch):
self.cuda = torch.cuda.is_available()
self.device = torch.device("cuda" if self.cuda else "cpu")
torch.backends.cudnn.benchmark = True
self.max_batch = maxbatch
def init_model(self, checkpoint=None, n_hidden=16):
self.checkpoint = checkpoint
self.model = UNet(n_channels=11, n_classes=1, n_hidden=n_hidden).to(self.device)
if (self.cuda):
checkpoint = torch.load('{0}.pth'.format(self.checkpoint))
else:
checkpoint = torch.load('{0}.pth'.format(self.checkpoint), map_location=lambda storage, loc: storage)
self.model.load_state_dict(checkpoint['state_dict'])
def forward(self, phase):
n_cases, n_phases, nx, ny = phase.shape
assert (n_phases == 11), "n. phases is not 11"
print("Normalizing data...")
phase = np.nan_to_num(phase)
phase -= np.mean(phase)
phase /= np.std(phase)
phase[phase>0] = 0.0
self.model.eval()
n_batches = n_cases // self.max_batch
n_remaining = n_cases % self.max_batch
print(" - Total number of maps : {0}".format(n_cases))
print(" - Total number of batches/remainder : {0}/{1}".format(n_batches, n_remaining))
magnetograms = np.zeros((n_cases,nx,ny))
left = 0
print("Predicting magnetograms...")
with torch.no_grad():
for i in range(n_batches):
right = left + self.max_batch
phases = torch.from_numpy(phase[left:right,:,:,:].astype('float32')).to(self.device)
output = self.model(phases)
magnetograms[left:right,:,:] = output.cpu().numpy()[:,0,:,:]
left += self.max_batch
if (n_remaining != 0):
right = left + n_remaining
phases = torch.from_numpy(phase[left:right,:,:,:].astype('float32')).to(self.device)
output = self.model(phases)
magnetograms[left:right,:,:] = output.cpu().numpy()[:,0,:,:]
return magnetograms
# + [markdown] colab_type="text" id="jX1K-XmkB-yQ"
# #Interpretation of the results
# The neural network returns a probability $P$ map with values in the range $[0,1]$. An active region is then identified by examining these probability maps, instead of directly evaluating the travel times of the far-side seismic maps. We defined an integrated probability $P_{\rm i}$, computed
# as the integral of the probability $P$ in a continuous feature. The concept of ``integrated probability'' is equivalent to the ``seismic strength'' defined by the traditional method. Rather than simply search for continuous regions with strong negative travel times, an approach that is hindered by the usual strong noise of the seismic data, the neural network provides a cleaner picture of the locations where an active region is most probable. However, the probability maps usually exhibit some significant values in regions with negative travel time as a result of noise.
#
# It is necessary to define an unequivocal
# criterion to decide whether a region with increased probability is claimed as an active region. We chose to define a threshold in the integrated probability as the minimum value for the detection of seismic sources, in the same way as the traditional method establishes a threshold in the seismic strength. The selection of the threshold was based on the evaluation of the artificial set of far-side maps for which we know the exact location of the seismic sources (see [Felipe & <NAME>, 2019, A&A, 632, 82](https://www.aanda.org/articles/aa/abs/2019/12/aa36838-19/aa36838-19.html)). A value of $P_{\rm i}=100$ proved to be a good compromise between the success in detecting the seismic sources and avoiding the claim of false positives. We note that when the network is applied to real data, false positives can be easily dealt with by discarding the cases where the detection does no appear consistently in successive dates at the same location.
# + [markdown] colab_type="text" id="K0evx1_1bvw0"
# ## Examples
#
# In this section, we apply the network to actual far-side seismic maps obtained from HMI.
# First, we need to install photutils and an appropriate version of astropy, since some of their routines will be employed for the interpretation of the network output.
# + colab={"base_uri": "https://localhost:8080/", "height": 233} colab_type="code" id="dv5eyevuXjeh" outputId="95688219-0c9c-40ee-8b2e-dc353ce92ee0"
# !pip install photutils astropy==3.2.3
# + colab={} colab_type="code" id="WXRDNJkPC0Qw"
#import some modules
import h5py
import matplotlib.pyplot as plt
from astropy.convolution import Gaussian2DKernel
from astropy.stats import gaussian_fwhm_to_sigma
from photutils import detect_sources
from photutils import detect_threshold
import scipy.io
# %matplotlib inline
# + [markdown] colab_type="text" id="ocFON_TAb3Ot"
# Next, we download the data needed for these examples. We require the trained model (2019-04-02-11:27:48_hid_16_lr_0.0003_wd_0.0.pth) and some observed far-side maps. Each of the files farside_NN2019_003_dlong140.sav and test.h5 contains a set of consecutive far-side HMI seismic maps. The individual seismic maps have 140 points in longitude, with a resolution of 1 degree and centered at the central meridian of the non-visible solar hemisphere. The latitude coverage spans from -72 to 71 degrees, with the same resolution of 1 degree.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 611} colab_type="code" id="aKXG6p1UZ0bm" outputId="77ecadc4-3a0d-4521-eccb-94fb2c8ad173"
# !wget -O 2019-04-02-11:27:48_hid_16_lr_0.0003_wd_0.0.pth https://owncloud.iac.es/index.php/s/2xJpktVzVSx4YGy/download
# !wget -O farside_NN2019_003_dlong140.sav https://owncloud.iac.es/index.php/s/Xtxn7OJ1fliUdw1/download
# !wget -O test.h5 https://owncloud.iac.es/index.php/s/iax6sNFf9UYTtxR/download
# + [markdown] colab_type="text" id="HYf4MC3YcDqP"
# ### Example 1
# The file test.h5 includes a series of HMI far-side seismic maps, with the latitude and longitude coverage and resolution described above. First, we read the seismic maps.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="qWS4kCTUYW6w" outputId="f7b8e028-263a-47b6-8ea8-dae3a4c1fdb9"
f = h5py.File('test.h5', 'r')
f.keys()
# + [markdown] colab_type="text" id="eYi5yfqSGVl1"
# Next, we plot a random selection of those maps. Each panel shows a seismic map computed from 24 hours of Doppler velocity temporal series measured with HMI. The figure shows the general appearance of the far-side seismic maps. The horizontal axes are the longitude, and the vertical axes correspond to the latitude. The maps exhibit a distribution of positive (yellow) and negative (black) travel-time shifts. Negative travel-time shifts may correspond to far-side active regions but, as illustrated in these examples, these maps are very noisy and must be carefully interpreted.
# + colab={"base_uri": "https://localhost:8080/", "height": 581} colab_type="code" id="7KlZgL5za6cx" outputId="c8c15cf8-b887-48f7-dd29-fe9a328bb27e"
fig, ax = plt.subplots(nrows=3, ncols=4, figsize=(10,10))
for i in range(3):
for j in range(4):
ax[i,j].imshow(f['phases'][i,j,:,:])
# + [markdown] colab_type="text" id="mox4gnmBHctu"
# We compute the probability maps applying the neural network to continuous series of 11 farside maps.
# + colab={} colab_type="code" id="CIa2PS37a-hu"
deep_farside_network = deep_farside(maxbatch=20)
deep_farside_network.init_model(checkpoint='2019-04-02-11:27:48_hid_16_lr_0.0003_wd_0.0', n_hidden=16)
# + colab={"base_uri": "https://localhost:8080/", "height": 89} colab_type="code" id="U1bzFPu9bFqe" outputId="9bd1e373-057a-4ecd-9d7b-12479f849250"
prob = deep_farside_network.forward(f['phases'][:])
# + [markdown] colab_type="text" id="-5tI90KzHsZr"
# We can plot the probability maps obtained for a few randomly selected cases. These examples show a few small patches with increased probability. However, we need to evaluate each of these features to check if the can be claim as active regions.
# + colab={"base_uri": "https://localhost:8080/", "height": 595} colab_type="code" id="ddAWw2HFbHxO" outputId="f0726a1c-989a-498a-b806-6cd7bf73d555"
fig, ax = plt.subplots(nrows=2, ncols=2, figsize=(10,10))
ax = ax.flatten()
for i in range(4):
ax[i].imshow(prob[i,:,:])
# + [markdown] colab_type="text" id="PhL7svM3Ijkn"
# We employ the following routines to select features present in an specific map. In this example, we identify the feature found in the bottom left panel of the previous figure.
# + colab={} colab_type="code" id="9yV1JZE9bKo9"
sigma = 3.0 * gaussian_fwhm_to_sigma # FWHM = 3.
kernel = Gaussian2DKernel(sigma, x_size=3, y_size=3)
kernel.normalize()
segm = detect_sources(prob[2,:,:], 0.01, npixels=5, filter_kernel=kernel)
# + colab={"base_uri": "https://localhost:8080/", "height": 287} colab_type="code" id="DSho4YT7bNa9" outputId="752034f7-db8e-46d5-8b1d-73903757740b"
plt.imshow(segm)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="3shND5-4JWEt" outputId="b31be737-2951-4bb2-d64a-985bbc3ca7c6"
tmp = prob[2,:,:]
(tmp[segm.data==1]).sum()
# + [markdown] colab_type="text" id="gpPojzDLJc_r"
# In this case, we obtain an integrated probability $P_i$=32. This value is below the threshold indicated in the previous section ($P_i$=100) and, thus, this feature cannot be claim as a far-side active region.
# + [markdown] colab_type="text" id="T9EfOVWI-DZv"
# ##Example 2
#
# The file farside_NN2019_003_dlong140.sav contains 11 consecutive far-side HMI seismic maps. They were employed for the detection of the far-side active region labeled NN-2019-003 in [Felipe & Asensio Ramos, 2019, A&A, 632, 82](https://www.aanda.org/articles/aa/abs/2019/12/aa36838-19/aa36838-19.html) as illustrated in the second row of Fig. 6 from that paper. These seismic maps were measured between 1 February 2019 at 00:00 UT and 6 February 2019 at 00:00 UT, with a cadence of 12 hours.
# Similarly to the previous example, we start by reading the data and applying the forward model to the set of seismic maps.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="7WARHm4tbPMO" outputId="b41663a3-a2ee-4563-909f-6eb3f4ef0e02"
tmp = scipy.io.readsav('farside_NN2019_003_dlong140.sav')
tmp['data_out'].shape
# + colab={"base_uri": "https://localhost:8080/", "height": 89} colab_type="code" id="rr9Jo7WPbT0t" outputId="c4f06b6b-0add-4e31-e0e3-8414c5c8c830"
prob = deep_farside_network.forward(tmp['data_out'][None,:,:,:])
# + [markdown] colab_type="text" id="l8zl53z5_q_o"
# The forward model returns the following probability map:
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="w1eZBJr9bWKd" outputId="fee198a0-73c5-435a-8cbe-37e580509aab"
plt.imshow(prob[0,:,:], origin='lower')
# + [markdown] colab_type="text" id="32AfAkOo_0bO"
# We identify the individual continuous regions with a certain probability for the presence of active regions. In this example, there are two independent features.
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="UjCwC9QsbYZN" outputId="cfffd564-d681-4708-a038-983bcddcb4f6"
sigma = 3.0 * gaussian_fwhm_to_sigma # FWHM = 3.
kernel = Gaussian2DKernel(sigma, x_size=3, y_size=3)
kernel.normalize()
segm = detect_sources(prob[0,:,:], 0.01, npixels=5, filter_kernel=kernel)
plt.imshow(segm, origin='lower')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="kb8Kzo8-bbZ9" outputId="2c77aac8-8f5c-4009-d7cf-873e8778e1cc"
tmp = prob[0,:,:]
(tmp[segm.data==2]).sum()
# + [markdown] colab_type="text" id="Ss_VqyNHd7kl"
# The big feature exhibits an integrated probability of $P_i$=205. This is above the threshold selected to claim a region with increased probability as an active region ($P_i$=100). We note that the value computed here is slightly different from the value reported in the publication. This discrepancy is due to the use of a different method for identifying the features in the probability map, but it does not affect the interpretation of the results.
# With regard to the small feature found in the previous figure:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="j96eFo3sKeWl" outputId="c73b4076-d716-4009-9fcd-cc50e9f03f06"
tmp = prob[0,:,:]
(tmp[segm.data==1]).sum()
# + [markdown] colab_type="text" id="sZJMhOc5KuRD"
# Its integrated probability is $P_i$=36 and, thus, our approach cannot guarantee its association to an actual far-side active region.
| book/_build/jupyter_execute/07/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#import matplot library
import matplotlib.pyplot as plt
#import seaborn library
import seaborn as sns
# %matplotlib inline
#load flights data from sns dataset (built in)
flight_data = sns.load_dataset('flights')
#view top 5 records
flight_data.head()
#use pivot method to re-arrange the dataset
flight_data = flight_data.pivot('month','year','passengers')
#view dataset
flight_data
#use heatmap method to generate the heatmap of the flights data
sns.heatmap(flight_data)
| Matplot_Heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced Altair: Multiple Coordinated Views
import altair as alt
import pandas as pd
import numpy as np
# +
flu = pd.read_csv('flunet2010_11countries.csv', header=[0,1])
cols = flu.columns.tolist()
normed = pd.melt(flu, id_vars=[cols[0]], value_vars=cols[1:], var_name=['continent','country'])
normed = normed.rename(columns={normed.columns[0]: 'week'})
normed.head()
# setup renderer for Jupyter Notebooks (not needed for Juptyer Lab)
alt.renderers.enable('notebook')
# -
# ## Visualization 1
#
# #### Create Linked Plots Showing Flu Cases per Country and Total Flu Cases per Week
# #### Selections:
# * Click to select individual countries.
# * Hold shift and click to select multiple countries.
# * Brush barchart to narrow top view.
# ## Visualization 2
#
# #### Create an Overview+Detail Plot Showing Flu Cases per Country
# ## Visualization 3
# #### Create Linked Plots Showing Flu Cases per Country per Week and Total Flu Cases per Country
# For this visualization we create two linked plots. One that shows flu cases per country per week and a second on that show the total of all flu cases per country.
| notebooks/flu_data_scaffold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 10. Align double ended measurements
# The cable length was initially configured during the DTS measurement. For double ended
# measurements it is important to enter the correct length so that the forward channel and the
# backward channel are aligned.
#
# This notebook shows how to better align the forward and the backward measurements. Do this before the calibration steps.
import os
from dtscalibration import read_silixa_files
from dtscalibration.datastore_utils import suggest_cable_shift_double_ended, shift_double_ended
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# suggest_cable_shift_double_ended?
# +
filepath = os.path.join('..', '..', 'tests', 'data', 'double_ended2')
ds_aligned = read_silixa_files(
directory=filepath,
timezone_netcdf='UTC',
file_ext='*.xml') # this one is already correctly aligned
# -
# Because our loaded files were already nicely aligned, we are purposely offsetting the forward and backward channel by 3 `spacial indices'.
ds_notaligned = shift_double_ended(ds_aligned, 3)
# The device-calibrated temperature doesnot have a valid meaning anymore and is dropped
suggested_shift = suggest_cable_shift_double_ended(
ds_notaligned,
np.arange(-5, 5),
plot_result=True,
figsize=(12,8))
# The two approaches suggest a shift of -3 and -4. It is up to the user which suggestion to follow. Usually the two suggested shift are close
ds_restored = shift_double_ended(ds_notaligned, suggested_shift[0])
print(ds_aligned.x, 3*'\n', ds_restored.x)
# Note that our fiber has become shorter by 2*3 spatial indices
| examples/notebooks/10Align_double_ended_measurements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# ## AutoGluon-Tabularでのカスタムモデルを用いた機械学習モデルの開発
# このノートブックでは、AutoGluon-Tabular を用いて、独自のモデルの持ち込みと組み合わせた高精度な機械学習モデル構築の自動化をご体感頂きます。AutoGluon-Tabular の `AbstractModel` クラスを活用することで、お好みのアルゴリズムの学習やアンサンブルモデルの構築を(半)自動化することができます。
# ## 準備
# 必要となるライブラリーをインストールします。
# !pip install --upgrade mxnet
# !pip install autogluon
# ライブラリをインポートし、必要な設定を行います。
# +
import time
import autogluon as ag
from autogluon import TabularPrediction as task
from autogluon.task.tabular_prediction.hyperparameter_configs import get_hyperparameter_config
from autogluon.utils.tabular.data.label_cleaner import LabelCleaner
from autogluon.utils.tabular.ml.models.abstract.abstract_model import AbstractModel
from autogluon.utils.tabular.ml.utils import infer_problem_type
# 予測対象となるカラム名を指定します。
label_column = 'class'
# 学習したモデルを保存するディレクトリを指定します。
savedir = 'ag_models/'
# -
# ### データの取得
# このサンプルでは、ある人の年収が50Kを超えるかどうかを二値分類する機械学習モデルを構築します。そのためのデータをダウンロードし、学習用データを準備します。
# +
train_data = task.Dataset(file_path='https://autogluon.s3.amazonaws.com/datasets/Inc/train.csv')
test_data = task.Dataset(file_path='https://autogluon.s3.amazonaws.com/datasets/Inc/test.csv')
train_data = train_data.head(500)
train_data.head()
# -
# 正解ラベルと特徴量を分離します。
# +
X_train = train_data.drop(columns=[label_column])
y_train = train_data[label_column]
problem_type = infer_problem_type(y=y_train)
# -
# 正解ラベルを学習や推論で活用するデータ型へ変換するために、`LabelCleaner` を作成します。str 型だったものが、int 型となっています。これを活用するとデータ型を戻すこともできます。
label_cleaner = LabelCleaner.construct(problem_type=problem_type, y=y_train)
y_train_clean = label_cleaner.transform(y_train)
# テストデータも同様に準備します。
X_test = test_data.drop(columns=[label_column])
y_test = test_data[label_column]
y_test_clean = label_cleaner.transform(y_test)
# ### 独自モデルの準備
# AutoGluon の組み込みアルゴリズム以外を活用する場合には、`AbstractModel` でラップします。今回の例では、 `scikit-learn` の `Naive Bayse` アルゴリズムを活用します。
class NaiveBayesModel(AbstractModel):
def preprocess(self, X):
cat_columns = X.select_dtypes(['category', 'object']).columns
X = X.drop(cat_columns, axis=1)
return super().preprocess(X).fillna(0)
def _fit(self, X_train, y_train, **kwargs):
from sklearn.naive_bayes import GaussianNB
X_train = self.preprocess(X_train)
self.model = GaussianNB(**self.params)
self.model.fit(X_train, y_train)
# ## 独自モデルのみの学習
# まずは、先程準備した `NaiveBayesModel` のみを学習させてみましょう。
naive_bayes_model = NaiveBayesModel(path='AutogluonModels/', name='CustomNaiveBayes', problem_type=problem_type)
naive_bayes_model.fit(X_train=X_train, y_train=y_train_clean)
# 学習済モデルは保存することができ、使用する場合には、`predictor = task.load(savedir)` のように保存用ディレクトリからロードすることができます。
# 今回学習させたモデルについて、その精度を評価してみましょう。
# +
y_pred = naive_bayes_model.predict(X_test)
y_pred_orig = label_cleaner.inverse_transform(y_pred)
score = naive_bayes_model.score(X_test, y_test_clean)
print(f'test score ({naive_bayes_model.eval_metric.name}) = {score}')
# -
print("Summary of class variable: \n", train_data[label_column].describe())
# ## AutoGluon の `task` クラスを使った `NaiveBayesModel` の学習
#
# ### 学習
# 今度は、先程と違い Autogluon の `task` クラスを使って学習を行います。これにより、異なるハイパーパラメーターでの学習や、その評価がより簡単に行えます。独自モデルを `task` クラスで活用するために、`custome_hyperparameters` 変数の中で `NaiveBayesModel` を指定しています。今回は `var_smoothing` に異なる3つの値を設定して学習を行い、評価をします。
# +
custom_hyperparameters = {NaiveBayesModel: [{},{'var_smoothing': 0.00001}, {'var_smoothing': 0.000002}]}
predictor = task.fit(train_data=train_data, label=label_column, hyperparameters=custom_hyperparameters)
# -
# ### 推論
#
# `leaderboard` メソッドを使うと学習の過程で生成されたそれぞれのモデルについて、テストデータ、バリデーションデータでの性能、時間などが表示されます。
predictor.leaderboard(test_data)
# 今回学習させたモデルについて、その精度を評価してみましょう。
# +
y_pred = predictor.predict(test_data)
score = naive_bayes_model.score(X_test, y_test_clean)
print(f'test score ({naive_bayes_model.eval_metric.name}) = {score}')
# -
# ## AutoGluon の `task` クラスを使い、`NaiveBayesModel` と他のモデルの学習
# 次に、`NaiveBayesModel` に加えて、AutoGluon の組み込みアルゴリズムも学習させます。
# ハイパーパラメーターの探索を行う場合には、`hp_tune=True` を選択します。また、調整したいハイパーパラメーターの探索領域を辞書形式で渡すことができます。その他、探索試行の回数や、時間制限の目安、探索戦略などを指定することができます。`fit` メソッドの引数に`auto_stack=True` を指定することで、複数層スタッキングを行う事ができます。
custom_hyperparameters
# ### 学習
# +
custom_hyperparameters.update(get_hyperparameter_config('default'))
predictor = task.fit(train_data=train_data, label=label_column,
auto_stack=True, hyperparameters=custom_hyperparameters)
# -
# ### 推論
#
# `leaderborad` メソッドや `evaluate_predictions` メソッドを使ってモデルの評価を見てみましょう。
# +
predictor.leaderboard(test_data)
y_pred = predictor.predict(test_data)
perf = predictor.evaluate_predictions(y_true=y_test, y_pred=y_pred, auxiliary_metrics=True)
# -
| brazil_ecommerce/working/jp-AutoGluon-Tabular-Custom.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # VV10 Non-local correlation kernel
# One of the largest deficiencies with semilocal functionals is the lack of long-range correlation effects. This most notable expresses itself as the lack of disperssion in the interactions between molecules. VV10 was expressly created to bridge the gap between the expensive of true non-local correlation and a computational tractable form. We will begin by writing the overall expression:
#
# $$E_c^{\rm{nl}} = \frac{1}{2}\int \int d{\bf r}d{\bf r'}\rho({\bf r})\Phi({\bf r},{\bf r'})\rho({\bf r'})$$
#
# Where the two densities are tied together through the $\Phi$ operator.
#
# For VV10 we have:
# $$
# \begin{align}
# \Phi &= -\frac{3}{2gg'(g + g')}\\
# g &= \omega_0({\rm r}) R^2 + \kappa({\rm r)}\\
# g' &= \omega_0({\rm r}) R^2 + \kappa({\rm r')}
# \end{align}
# $$
#
# Where $w_{0}$:
#
# $$
# \begin{align}
# \omega_{0}(r) &= \sqrt{\omega_{g}^2(r) + \frac{\omega_p^2(r)}{3}} \\
# \omega_g^2(r) &= C \left | \frac{\nabla \rho({\bf r})}{\rho({\bf r})} \right |^4 \\
# \omega_p^2(r) &= 4 \pi \rho({\bf r})
# \end{align}
# $$
#
# and finally:
#
# $$\kappa({\bf r}) = b * \frac{3 \pi}{2} \left [ \frac{\rho({\bf r})}{9\pi} \right ]^\frac{1}{6}$$
#
# While there are several expression, this is actually quite easy to compute. First let us examine how the VV10 energy is reintegrated:
#
# $$E_c^{\rm{VV10}} = \int d{\bf r} \rho{\bf r} \left [ \beta + \frac{1}{2}\int d{\bf r'} \rho{\bf r'} \Phi({\bf r},{\bf r'}) \right]$$
#
#
# +
import psi4
import numpy as np
import ks_helper as ks
mol = psi4.geometry("""
He 0 0 -5
He 0 0 5
symmetry c1
""")
options = {'BASIS': 'aug-cc-pVDZ',
'DFT_SPHERICAL_POINTS': 110,
'DFT_RADIAL_POINTS': 20}
# -
# ## VV10 coefficients
# First let us build set and build a few static coefficients:
coef_C = 0.0093
coef_B = 5.9
coef_beta = 1.0 / 32.0 * (3.0 / (coef_B ** 2.0)) ** (3.0 / 4.0)
# ## VV10 kernel
# First let us construct a function that compute $\omega_0$ and $\kappa$ quantities. To make one piece simpler let us first examine a piece of $\omega_g$:
# $$\left |\frac{\nabla \rho({\bf r})}{\rho({\bf r})} \right|^4$$
#
# quantity. Recall that
#
# $$\gamma({\bf r}) = \nabla\rho({\bf r})\cdot\nabla\rho({\bf r})$$
#
# therefore, we can simplify the above to:
#
# $$\frac{\nabla \rho({\bf r})}{\rho({\bf r})} = \left | \frac{\gamma({\bf r})}{\rho({\bf r})\cdot({\bf r})} \right | ^2 $$
def compute_vv10_kernel(rho, gamma):
kappa_pref = coef_B * (1.5 * np.pi) / ((9.0 * np.pi) ** (1.0 / 6.0))
# Compute R quantities
Wp = (4.0 / 3.0) * np.pi * rho
Wg = coef_C * ((gamma / (rho * rho)) ** 2.0)
W0 = np.sqrt(Wg + Wp)
kappa = rho ** (1.0 / 6.0) * kappa_pref
return W0, kappa
# ## VV10 energy and gradient evaluation
#
# Yup so just integrate it out. Pretty easy.
# +
def compute_vv10(D, Vpot):
nbf = D.shape[0]
Varr = np.zeros((nbf, nbf))
total_e = 0.0
tD = 2.0 * np.array(D)
points_func = Vpot.properties()[0]
superfunc = Vpot.functional()
xc_e = 0.0
vv10_e = 0.0
# First loop over the outer set of blocks
for l_block in range(Vpot.nblocks()):
# Obtain general grid information
l_grid = Vpot.get_block(l_block)
l_w = np.array(l_grid.w())
l_x = np.array(l_grid.x())
l_y = np.array(l_grid.y())
l_z = np.array(l_grid.z())
l_npoints = l_w.shape[0]
points_func.compute_points(l_grid)
# Compute the functional itself
ret = superfunc.compute_functional(points_func.point_values(), -1)
xc_e += np.vdot(l_w, np.array(ret["V"])[:l_npoints])
v_rho = np.array(ret["V_RHO_A"])[:l_npoints]
v_gamma = np.array(ret["V_GAMMA_AA"])[:l_npoints]
# Begin VV10 information
l_rho = np.array(points_func.point_values()["RHO_A"])[:l_npoints]
l_gamma = np.array(points_func.point_values()["GAMMA_AA"])[:l_npoints]
l_W0, l_kappa = compute_vv10_kernel(l_rho, l_gamma)
phi_kernel = np.zeros_like(l_rho)
phi_U = np.zeros_like(l_rho)
phi_W = np.zeros_like(l_rho)
# Loop over the inner set of blocks
for r_block in range(Vpot.nblocks()):
# Repeat as for the left blocks
r_grid = Vpot.get_block(r_block)
r_w = np.array(r_grid.w())
r_x = np.array(r_grid.x())
r_y = np.array(r_grid.y())
r_z = np.array(r_grid.z())
r_npoints = r_w.shape[0]
points_func.compute_points(r_grid)
r_rho = np.array(points_func.point_values()["RHO_A"])[:r_npoints]
r_gamma = np.array(points_func.point_values()["GAMMA_AA"])[:r_npoints]
r_W0, r_kappa = compute_vv10_kernel(r_rho, r_gamma)
# Build the distnace matrix
R2 = (l_x[:, None] - r_x) ** 2
R2 += (l_y[:, None] - r_y) ** 2
R2 += (l_z[:, None] - r_z) ** 2
# Build g
g = l_W0[:, None] * R2 + l_kappa[:, None]
gp = r_W0 * R2 + r_kappa
#
F_kernal = -1.5 * r_w * r_rho / (g * gp * (g + gp))
F_U = F_kernal * ((1.0 / g) + (1.0 / (g + gp)))
F_W = F_U * R2
phi_kernel += np.sum(F_kernal, axis=1)
phi_U += -np.sum(F_U, axis=1)
phi_W += -np.sum(F_W, axis=1)
# Compute those derivatives
kappa_dn = l_kappa / (6.0 * l_rho)
w0_dgamma = coef_C * l_gamma / (l_W0 * l_rho ** 4.0)
w0_drho = 2.0 / l_W0 * (np.pi/3.0 - coef_C * np.power(l_gamma, 2.0) / (l_rho ** 5.0))
# Sum up the energy
vv10_e += np.sum(l_w * l_rho * (coef_beta + 0.5 * phi_kernel))
# Perturb the derivative quantities
v_rho += coef_beta + phi_kernel + l_rho * (kappa_dn * phi_U + w0_drho * phi_W)
v_rho *= 0.5
v_gamma += l_rho * w0_dgamma * phi_W
# Recompute to l_grid
lpos = np.array(l_grid.functions_local_to_global())
points_func.compute_points(l_grid)
nfunctions = lpos.shape[0]
# Integrate the LDA and GGA quantities
phi = np.array(points_func.basis_values()["PHI"])[:l_npoints, :nfunctions]
phi_x = np.array(points_func.basis_values()["PHI_X"])[:l_npoints, :nfunctions]
phi_y = np.array(points_func.basis_values()["PHI_Y"])[:l_npoints, :nfunctions]
phi_z = np.array(points_func.basis_values()["PHI_Z"])[:l_npoints, :nfunctions]
# LDA
Vtmp = np.einsum('pb,p,p,pa->ab', phi, v_rho, l_w, phi)
# GGA
l_rho_x = np.array(points_func.point_values()["RHO_AX"])[:l_npoints]
l_rho_y = np.array(points_func.point_values()["RHO_AY"])[:l_npoints]
l_rho_z = np.array(points_func.point_values()["RHO_AZ"])[:l_npoints]
tmp_grid = 2.0 * l_w * v_gamma
Vtmp += np.einsum('pb,p,p,pa->ab', phi, tmp_grid, l_rho_x, phi_x)
Vtmp += np.einsum('pb,p,p,pa->ab', phi, tmp_grid, l_rho_y, phi_y)
Vtmp += np.einsum('pb,p,p,pa->ab', phi, tmp_grid, l_rho_z, phi_z)
# Sum back to the correct place
Varr[(lpos[:, None], lpos)] += Vtmp + Vtmp.T
print(" VV10 NL energy: %16.8f" % vv10_e)
xc_e += vv10_e
return xc_e, Varr
ks.ks_solver("VV10", mol, options, compute_vv10)
# -
# Refs:
# - <NAME>.; <NAME>., *J. Chem. Phys.*, **2010**, *133*, 244103
| Tutorials/04_Density_Functional_Theory/4d_VV10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/suyash091/EEG-MNIST-Analysis/blob/master/Training_EEG_Lasso.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="0zeYcI6pz2VU" colab_type="code" colab={}
import pandas as pd
import gc
import numpy as np
# + id="bMy3RrZf2Ho5" colab_type="code" outputId="3e4e9adb-a4e5-4678-91bf-a9715b18d6de" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + id="ln-56IUl6iVk" colab_type="code" colab={}
Df1=pd.read_csv('/content/drive/My Drive/BCI MNIST/xvar1.csv')
Df2=pd.read_csv('/content/drive/My Drive/BCI MNIST/xvar2.csv')
Df3=pd.read_csv('/content/drive/My Drive/BCI MNIST/xvar3.csv')
# + id="VAcEgmdp_qQB" colab_type="code" colab={}
bigdata = pd.concat([Df1, Df2, Df3], ignore_index=True, sort =False)
del Df1
del Df2
del Df3
# + id="72od6WwFIi2k" colab_type="code" outputId="e412c5e3-892c-4db9-acfd-c69b3626d509" colab={"base_uri": "https://localhost:8080/", "height": 253}
bigdata.drop(bigdata.columns[bigdata.columns.str.contains('unnamed',case = False)],axis = 1, inplace = True)
bigdata.head()
# + id="LljaJcVxIslc" colab_type="code" colab={}
pdf1=pd.read_csv('/content/drive/My Drive/BCI MNIST/yvar1.csv')
pdf2=pd.read_csv('/content/drive/My Drive/BCI MNIST/yvar2.csv')
pdf3=pd.read_csv('/content/drive/My Drive/BCI MNIST/yvar3.csv')
# + id="qypUiETeLX-D" colab_type="code" colab={}
bigpred = pd.concat([pdf1, pdf2, pdf3], ignore_index=True, sort =False)
# + id="3Oj7j-elRvcu" colab_type="code" colab={}
bigpred=bigpred['0'].map({'-1-1-1-1-1-1-1-1-1-1-1-1-1-1':10,'88888888888888':8,'66666666666666':6,'00000000000000': 0,'99999999999999':9,'55555555555555':5,'22222222222222':2,'11111111111111':1,'77777777777777':7,'33333333333333':3,'44444444444444':4,88888888888888:8,66666666666666:6,00000000000000: 0,99999999999999:9,55555555555555:5,22222222222222:2,11111111111111:1,77777777777777:7,33333333333333:3,44444444444444:4})
# + id="VMBZZz2lLeBE" colab_type="code" outputId="5a493900-c4f3-4db8-ac1c-cab08c01ad86" colab={"base_uri": "https://localhost:8080/", "height": 34}
del pdf1
del pdf2
del pdf3
gc.collect()
# + id="_YINLtuoOba5" colab_type="code" outputId="e8b5f106-100c-4558-c5d4-fd22ce4dd425" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(bigpred.dropna())==len(bigpred)
# + id="5HK5YlvtSgFZ" colab_type="code" outputId="9f303475-66be-4c73-8cbe-b73fdd501b33" colab={"base_uri": "https://localhost:8080/", "height": 119}
bigpred.head()
# + id="xBPc3Cf4Nbd7" colab_type="code" outputId="a6b9653b-d357-46f9-c2cc-de1e1fb70173" colab={"base_uri": "https://localhost:8080/", "height": 320}
bigpred.drop(bigpred.columns[bigpred.columns.str.contains('unnamed',case = False)],axis = 1, inplace = True)
#bigpred.head(100)
# + id="9OXsffciLlhd" colab_type="code" outputId="858e444f-3365-4fbd-d72c-673d88b9dfb5" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(bigdata)==len(bigpred)
# + id="GpD9aJXGLqdu" colab_type="code" colab={}
#Import models from scikit learn module:
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import KFold #For K-fold cross validation
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn import metrics
import numpy as np
from sklearn.metrics import roc_curve, auc, precision_score, confusion_matrix, explained_variance_score, max_error, mean_absolute_error, mean_squared_error
from sklearn.linear_model import LogisticRegression, LinearRegression, Ridge, Lasso
#Generic function for making a classification model and accessing performance:
def classification_model(model, predictors, outcome):
history=model.fit(predictors,outcome.values.ravel())
predictions = model.predict(predictors)
accuracy = metrics.accuracy_score(predictions,outcome)
print('Accuracy : %s' % '{0:.3%}'.format(accuracy))
kf = KFold(n_splits=2)
error = []
for train, test in kf.split(predictors):
train_predictors = (predictors.iloc[train,:])
train_target = outcome.iloc[train]
model.fit(train_predictors, train_target.values.ravel())
error.append(model.score(predictors.iloc[test,:], outcome.iloc[test]))
print('Cross-Validation Score : %s' % '{0:.3%}'.format(np.mean(error)))
return history
# + id="IctR9mDZNUo8" colab_type="code" outputId="25fc1087-da9c-4799-e160-b037822098b4" colab={"base_uri": "https://localhost:8080/", "height": 204}
idx = np.random.permutation(df1.index)
# + id="6D0OwDLYwOui" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f70c190d-a7c8-4e9a-8e41-d2cd27c8fe51"
from sklearn.model_selection import train_test_split
df=bigdata
bigdata=(df-df.values.min())/(df.values.max()-df.values.min())
print('start')
X_train, X_test, y_train, y_test = train_test_split(bigdata, bigpred, test_size=0.20, random_state=42)
model = Lasso(alpha=0.1, copy_X=True, fit_intercept=True, max_iter=1000,normalize=False, positive=False, precompute=False, random_state=None,selection='cyclic', tol=0.0001, warm_start=False)
model.fit(X_train,y_train)
print('stop')
# + id="Zhfnto0BMbcn" colab_type="code" colab={}
#scaler = StandardScaler()
idx = np.random.permutation(X_test.index)
X_test=X_test.reindex(idx)
y_test=y_test.reindex(idx)
predictions = model.predict(X_test)
#prc=precision_score(predictions,y_test, average=None)
#cfm=confusion_matrix(predictions,y_test)
#accuracy = metrics.accuracy_score(predictions,y_test)
#false_positive_rate, true_positive_rate, thresholds = roc_curve(y_test, predictions)
#auc = auc(false_positive_rate, true_positive_rate)
#print(accuracy)
#print(prc)
#print(cfm)
#history=classification_model(model,X_test,y_test)
# + id="GxBhYzrVK4jz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="016a976f-9b3a-4aee-f527-27270cc7ff14"
print(explained_variance_score(predictions,y_test))
print(max_error(predictions,y_test))
print(mean_absolute_error(predictions, y_test, multioutput='raw_values'))
print(mean_squared_error(predictions, y_test, multioutput='raw_values'))
| Training_EEG_Lasso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The problem statement I am interested in is: **predicting the publisher's name from a given title**. For approaching this problem, first I am going to need a dataset consisting of article/post titles with their sources mentioned.
#
# The dataset I am going to use is already there as a BigQuery public dataset ([link](https://console.cloud.google.com/bigquery?p=bigquery-public-data&d=hacker_news&page=dataset)). But the dataset needs to be shaped a bit aligning to my needs.
#
# This dataset contains all stories and comments from Hacker News from its launch in 2006 to present. Each story contains a story ID, the author that made the post, when it was written, and the number of points the story received.
# The problem can be modeled as a text classification problem essentially. The steps I would be following:
# - Derive a small subset of the dataset from BiqQuery as the original dataset is moderately large (402 MB)
# - Preprocess the dataset initially
# - Get the data ready for feeding to a sequence model
# - Tokenize the titles
# - Pad the sequences
# - Create embedding matrix w.r.t pretrained embeddings
# - Build, train and evaluate the model
#
# Let's begin.
#
# **Note**: It is advisable to use Google Colab. To be able to use BigQuery you need to get some setup. Follow the instructions [here](https://colab.research.google.com/notebooks/bigquery.ipynb).
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="aXqlI97G24Xx" outputId="78888093-fa79-4aa4-96c1-5f36924cd932"
# Authenticate yourself with the Google creds
from google.colab import auth
auth.authenticate_user()
print('Authenticated')
# + colab={} colab_type="code" id="TiFL6K5U3ZCq"
# Set your Project ID
import os
PROJECT = 'fast-ai-exploration'
os.environ['PROJECT'] = PROJECT
# -
# I can now run a BQ query to get you a sense of the dataset.
# + colab={} colab_type="code" id="lsrAecORbhQ3"
# %%bigquery --project $PROJECT data_preview
SELECT
url, title, score
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
LENGTH(title) > 10
AND score > 10
AND LENGTH(url) > 0
LIMIT 10
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="_IOISh6Ibym2" outputId="e822251d-17bc-4f3b-a962-a9d68e4ccb11"
data_preview.head()
# -
# The problem in the current data is in place of `url` I need the source of the URL. For example, `https://github.com/Groundworkstech/Submicron` should appear as `github`. I would also want to rename the `url` column to `source`. But first, let me figure out the distribution in the titles belonging to several sources.
# + colab={} colab_type="code" id="bd-bzWav3J1H"
# %%bigquery --project $PROJECT source_num_articles
SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
COUNT(title) AS num_articles
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
GROUP BY
source
ORDER BY num_articles DESC
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="WXxlCxZa3rE1" outputId="da844436-5e3c-48df-d4dd-02f2bfe20be0"
source_num_articles.head()
# -
# Not major, but a slight class imbalance is there. I am going to exclude the titles belonging to `youtube`.
# + colab={} colab_type="code" id="gzscWESV36Uu"
# %%bigquery --project $PROJECT full_data
SELECT source, LOWER(REGEXP_REPLACE(title, '[^a-zA-Z0-9 $.-]', ' ')) AS title FROM
(SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
title
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
)
WHERE (source = 'github' OR source = 'nytimes' OR source = 'techcrunch' or source = 'blogspot')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="xOQKxkwt4u6m" outputId="444bcf17-c2e5-4c5b-fbc2-52d2dadaa352"
full_data.head()
# -
# The data is gathered now and as the next step, I would like perform a basic round of EDA. I would start by running a `.shape`.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Vv6hPlEs5cGB" outputId="5e74995f-5843-4458-bb9a-00519cabe90e"
# Number of (rows, coulumns)
full_data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="UF1Kay5Z4138" outputId="b6a4ae3f-3f30-4f3e-8f7d-f46d3e52c187"
# Class distribution
full_data.source.value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="pF1tEPij5MPq" outputId="fd00511f-b1dd-4f79-8c99-957a36dc2202"
# Missing value inspection
full_data.isna().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="Lnvx6hf05T7y" outputId="d4cec7f0-ce9b-46af-8ff4-8c902ba211b6"
# How is the length of the titles distributed?
full_data['title'].apply(len).describe()
# + colab={} colab_type="code" id="krs_FTVE45_C"
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('ggplot')
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="QCZh1CeK5JUT" outputId="9b8779a3-6554-4c4a-d9a4-475c58b5728d"
text_lens = full_data['title'].apply(len).values
plt.figure(figsize=(10,8))
sns.set()
g = sns.distplot(text_lens, kde=False, hist_kws={'rwidth':1})
g.set_xlabel('Title length')
g.set_ylabel('Count')
plt.show()
# -
# I would now like to perform some manual inspections so to figure out
# - how many titles fall above the minimum title length (11)
# - how many titles have the maximum length (138)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="bmVrxUfe5u15" outputId="999f1fd9-f36e-4481-8639-5e24e7359138"
(text_lens <= 11).sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="elTQh8XI9bcz" outputId="3ef89ed3-ced8-44e3-8612-6db41e8491b7"
(text_lens == 138).sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="Vn1aCuXr9l6s" outputId="20b71a9d-3149-4385-ceb0-a3203a64f944"
full_data[text_lens <= 11]['title'].head()
# + colab={"base_uri": "https://localhost:8080/", "height": 80} colab_type="code" id="dx7zMNwE95IL" outputId="15172a3d-4fe9-4f28-9f4c-2b1d1c977e55"
full_data[text_lens == 138][['title', 'source']]
# -
# Since, there is only one title with the title length of 138, I would remove it to avoid any skew.
# + colab={} colab_type="code" id="aQoLKSxg9_V2"
full_data = full_data[text_lens < 138].reset_index(drop=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ubuR__Ge_o1v" outputId="bba4e9f2-2921-40cd-db3e-0b0f2646ecda"
full_data.shape
# -
# ### Data splits with `pandas` `sample()`
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="L85exTkF-Z0b" outputId="5f443949-4eaa-4d28-fd38-499b29edecc3"
# 80% for train
train = full_data.sample(frac=0.8)
full_data.drop(train.index, axis=0, inplace=True)
# 10% for validation
valid = full_data.sample(frac=0.5)
full_data.drop(valid.index, axis=0, inplace=True)
# 10% for test
test = full_data
train.shape, valid.shape, test.shape
# -
# ### Check the class distributions in the splits
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="MqPWSFaCAfqv" outputId="d5dbc6d8-f68d-4ed6-c30a-e92ca0717144"
train.source.value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="BDJW9-gQAj1b" outputId="7bebb721-09a5-47e9-d90d-dec6509ec11d"
valid.source.value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="uxFd_Hw_AoNO" outputId="2346273d-2fca-4923-dd9a-e408345eba9b"
test.source.value_counts()
# + colab={} colab_type="code" id="6_wdbM5xABVj"
# !mkdir data
# -
# ### Serialize the datasets for later use
# + colab={} colab_type="code" id="cQwbtp_Z_VmJ"
train.to_csv('data/train.csv')
valid.to_csv('data/valid.csv')
test.to_csv('data/test.csv')
| Predicting publisher's name from an article with TF 2.0/Data_Collection_and_basic_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: Data Analysis
#
# This is the follow up to [Tutorial: Data Collection](./Readout-Data-Collection.ipynb). We have measured bitstrings for the single-qubit circuit $R_y(\theta)$ for various `theta`s. In this analysis, we compute $\langle Z \rangle (\theta)$, compare to the anayltically expected true value, and fit to a depolarizing noise model with T1 decay during readout.
# ## Loading data
#
# We can use utilities in ReCirq to query the filesystem and load in a dataset. Please recall that all tasks have an associated `EXPERIMENT_NAME` and a `dataset_id` which define the top two hierarchies in the filesystem. We import these values from the data collection script to ensure consistency.
# +
import cirq
import recirq
from recirq.readout_scan.tasks import EXPERIMENT_NAME, DEFAULT_BASE_DIR
# -
# `recirq.iterload_records` uses these two bits of information to iterate over records saved using `recirq.save` (in the data collection script.
#
# This also gives you a chance to do post-processing on the data. In general, you should do some massaging of the data and put the results into a pandas DataFrame. DataFrames are great for doing statistics and visualizations across tabular data.
# +
import numpy as np
import pandas as pd
records = []
# Load all data, do some light processing
for record in recirq.iterload_records(dataset_id='2020-02-tutorial', base_dir=DEFAULT_BASE_DIR):
# Expand task dataclass into columns
recirq.flatten_dataclass_into_record(record, 'task')
# Unwrap BitArray into np.ndarray
all_bitstrings = [ba.bits for ba in record['all_bitstrings']]
# Compute <Z>
record['z_vals'] = [np.mean((-1)**bitstrings, axis=0).item() for bitstrings in all_bitstrings]
# Don't need to carry around the full array of bits anymore
del record['all_bitstrings']
records.append(record)
df = pd.DataFrame(records)
print(len(df))
df.head()
# -
# ## Plot the data
#
# A good first step.
# +
# %matplotlib inline
from matplotlib import pyplot as plt
entry = df.iloc[0] # Pick the first qubit
plt.plot([], []) # advance color cycle in anticipation of future analysis
plt.plot(entry['thetas'], entry['z_vals'], 'o-')
plt.xlabel('Theta', fontsize=14)
plt.ylabel(r'$\langle Z \rangle$', fontsize=14)
plt.title("Qubit {}".format(entry['qubit']), fontsize=14)
plt.tight_layout()
# -
# ## How does it compare to analytical results?
#
# You could imagine setting up a separate task for computing and saving analytic results. For this single qubit example, we'll just compute it on the fly.
# +
qubit = cirq.LineQubit(0)
thetas = df.iloc[0]['thetas']
class _DummyMeasurementGate(cirq.IdentityGate):
"""A dummy measurement used to trick simulators into applying
readout error when using PauliString.expectation_from_xxx."""
def _measurement_key_(self):
return 'dummy!'
def __repr__(self):
if self.num_qubits() == 1:
return '_DummyMeasurementGate'
return '_DummyMeasurementGate({!r})'.format(self.num_qubits())
def __str__(self):
if (self.num_qubits() == 1):
return 'dummyM'
else:
return 'dummyM({})'.format(self.num_qubits())
def _circuit_diagram_info_(self, args):
from cirq import protocols
return protocols.CircuitDiagramInfo(
wire_symbols=('dM',) * self.num_qubits(), connected=True)
def dummy_measure(qubits):
return _DummyMeasurementGate(num_qubits=len(qubits)).on(*qubits)
def get_circuit(theta):
return cirq.Circuit([
cirq.ry(theta).on(qubit),
dummy_measure([qubit])
])
true_z_vals = []
for theta in thetas:
wf = cirq.final_wavefunction(get_circuit(theta))
op = cirq.Z(qubit) * 1.
true_z_val = op.expectation_from_wavefunction(wf, qubit_map={qubit:0}, check_preconditions=False)
true_z_vals.append(np.real_if_close(true_z_val).item())
true_z_vals = np.array(true_z_vals)
true_z_vals
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(11, 4))
ax1.plot(thetas, true_z_vals, '-', label='True')
ax1.plot(entry['thetas'], entry['z_vals'], 'o-', label='Data')
ax2.plot([], []) # advance color cycle
ax2.plot(entry['thetas'], np.abs(true_z_vals - entry['z_vals']), 'o-', label='|Data - True|')
ax1.legend(loc='best', frameon=False)
ax2.legend(loc='best', frameon=False)
ax1.set_xlabel('Theta', fontsize=14)
ax2.set_xlabel('Theta', fontsize=14)
fig.tight_layout()
# -
# ## Learn a model
#
# Our experimental data has some wiggles in it, but it also has a clear pattern of deviation from the true values. We can hypothesize a (parameterized) noise model and then use function minimization to fit the noise model parameters.
# +
import scipy.optimize
import cirq.contrib.noise_models as ccn
def get_obj_func(data_expectations):
all_results = []
def obj_func(x):
depol_prob, decay_prob, readout_prob = x
if depol_prob < 0 or decay_prob < 0 or readout_prob < 0:
# emulate constraints by returning a high cost if we
# stray into invalid territory
return 1000
sim = cirq.DensityMatrixSimulator(
noise=ccn.DepolarizingWithDampedReadoutNoiseModel(
depol_prob=depol_prob, decay_prob=decay_prob, bitflip_prob=readout_prob))
results = []
for theta in thetas:
density_result = sim.simulate(get_circuit(theta))
op = cirq.Z(qubit) * 1.
true_z_val = op.expectation_from_wavefunction(density_result.final_density_matrix, qubit_map=density_result.qubit_map, check_preconditions=False)
results.append(np.real_if_close(true_z_val).item())
results = np.array(results)
all_results.append(results)
cost = np.sum(np.abs(results - data_expectations))
return cost
return obj_func, all_results
# -
def print_result(x):
depol_prob, decay_prob, readout_prob = x
print(f'depol = {depol_prob:.2%}')
print(f'decay = {decay_prob:.2%}')
print(f'readout = {readout_prob:.2%}')
dfb = df
dfb = dfb.head(5) # Remove this to do all qubits
len(dfb)
# +
# Initial values
depol_prob = 0.01
decay_prob = 0.01
readout_prob = 0.01
opt_results = []
for i, entry in dfb.iterrows():
ofunc, results = get_obj_func(entry['z_vals'])
opt_result = scipy.optimize.minimize(ofunc,
[depol_prob, decay_prob, readout_prob],
method='nelder-mead',
options={'disp': True})
label = f"{entry['qubit'].row}, {entry['qubit'].col}"
print("Qubit", label)
print_result(opt_result.x)
opt_results.append(opt_result)
data_expectations = entry['z_vals']
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(11, 4))
ax1.plot(thetas, true_z_vals, label='True')
ax1.plot(thetas, data_expectations, 'o-', label=f'{label} Data')
ax1.plot(thetas, results[-1], '.-', label='Fit')
ax2.plot([], []) # advance color cycle
ax2.plot(thetas, np.abs(true_z_vals - data_expectations), 'o-', label='|Data - True|')
ax2.plot(thetas, np.abs(true_z_vals - results[-1]), '-', label='|Fit - True|')
ax1.legend(loc='best')
ax2.legend(loc='best')
fig.tight_layout()
plt.show()
| docs/Readout-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Counting Rings in Cropped Images - Real Data
#
# Mod of [<NAME>'s WWF 01_Custom.ipynb](https://walkwithfastai.com/Custom).
#
#
# Here we'll take cropped images of antinodes and try to count the rings, by fashioning a regression model out of a one-class classification model and scaling the output sigmoid (via fastai's y_range parameter) so that our fitted values stay within the linear regime of the sigmoid.
#
# And we also want to "clamp" our output between a min of about 0.2 rings and a max of 11 rings, because that's how the dataset was created; so sigmoid makes a good choice for this "clamping" too.
# +
#all_slow
# -
# ## Installs & Imports
#hide
#Run once per session
# !pip install fastai espiownage -q --upgrade
import espiownage
from espiownage.core import *
sysinfo()
print(f"espiownage version {espiownage.__version__}")
# And import our libraries
from fastai.vision.all import *
from espiownage.core import *
# Below you will find the exact imports for everything we use today
# +
from fastcore.foundation import L
from fastcore.xtras import Path # @patch'd properties to the Pathlib module
from fastai.callback.fp16 import to_fp16
from fastai.callback.schedule import fit_one_cycle, lr_find
from fastai.data.external import untar_data, URLs
from fastai.data.block import RegressionBlock, DataBlock
from fastai.data.transforms import get_image_files, Normalize, RandomSplitter, parent_label
from fastai.interpret import ClassificationInterpretation
from fastai.learner import Learner # imports @patch'd properties to Learner including `save`, `load`, `freeze`, and `unfreeze`
from fastai.optimizer import ranger
from fastai.vision.augment import aug_transforms, RandomResizedCrop, Resize
from fastai.vision.core import imagenet_stats
from fastai.vision.data import ImageBlock
from fastai.vision.learner import cnn_learner
from fastai.vision.utils import download_images, verify_images
# -
# # Run parameters
dataset_name = 'cyclegan' # choose from:
# - cleaner (*real* data that's clean-er than "preclean"),
# - preclean (unedited aggregates of 15-or-more volunteers)
# - spnet (original SPNet Real dataset)
# - cyclegan (original SPNet CGSmall dataset)
# - fake (newer than SPNet fake, this includes non-int ring #s)
use_wandb = False # WandB.ai logging
project = 'count_in_crops' # project name for wandb
# Optional: WandB tracking
if use_wandb:
# !pip install wandb -qqq
import wandb
from fastai.callback.wandb import *
from fastai.callback.tracker import SaveModelCallback
wandb.login()
# # Download and prepare data
#skip
path = get_data(dataset_name) / 'crops'; path
fnames = get_image_files(path)
print(f"{len(fnames)} total cropped images")
ind = 1 # pick one cropped image
fnames[ind]
# For labels, we want the ring count which is the number between the last '_' and the '.png'
# +
def label_func(x):
return round(float(x.stem.split('_')[-1]),2)
print(label_func(fnames[ind]))
# -
cropsize = (300,300) # pixels
croppedrings = DataBlock(blocks=(ImageBlock, RegressionBlock(n_out=1)),
get_items=get_image_files,
splitter=RandomSplitter(),
get_y=label_func,
item_tfms=Resize(cropsize, ResizeMethod.Squish),
batch_tfms=[*aug_transforms(size=cropsize, flip_vert=True, max_rotate=360.0),
Normalize.from_stats(*imagenet_stats)])
dls = croppedrings.dataloaders(path, bs=32)
dls.show_batch(max_n=9)
# ## Train model
opt = ranger
y_range=(0.2,13) # balance between "clamping" to range of real data vs too much "compression" from sigmoid nonlinearity
learn = cnn_learner(dls, resnet34, n_out=1, y_range=y_range, metrics=[mae, acc_reg05,acc_reg1,acc_reg15,acc_reg2], loss_func=MSELossFlat(), opt_func=opt)
learn.lr_find()
#learn.fine_tune(10, 1e-2)
lr = 5e-3
learn.fine_tune(30, lr, freeze_epochs=2) # accidentally ran this twice in a row :-O
learn.save(f'crop-rings-{dataset_name}')
# ## Interpretation
learn.load(f'crop-rings-{dataset_name}');
preds, targs, losses = learn.get_preds(with_loss=True) # validation set only
len(preds)
# I'll define a method to show a single prediction
def showpred(ind, preds, targs, losses, dls): # show prediction at this index
print(f"preds[{ind}] = {preds[ind]}, targs[{ind}] = {targs[ind]}, loss = {losses[ind]}")
print(f"file = {os.path.basename(dls.valid.items[ind])}")
print("Image:")
dls.valid.dataset[ind][0].show()
showpred(0, preds, targs, losses, dls)
# And now we'll run through predictions for the whole validation set:
# +
#preds, targs, losses = learn.get_preds(with_loss=True)
results = []
for i in range(len(preds)):
line_list = [dls.valid.items[i].stem]+[round(targs[i].cpu().numpy().item(),2), round(preds[i][0].cpu().numpy().item(),2), losses[i].cpu().numpy(), i]
results.append(line_list)
# store as pandas dataframe
res_df = pd.DataFrame(results, columns=['filename', 'target', 'prediction', 'loss','i'])
# -
# We can do our own version of printing top_losses:
res_df = res_df.sort_values('loss', ascending=False)
res_df.head()
# +
def show_top_losses(res_df, preds, targs, losses, dls, n=5):
for j in range(n):
showpred(res_df.iloc[j]['i'], preds, targs, losses, dls)
show_top_losses(res_df, preds, targs, losses, dls)
# -
# So then we can these results output into a CSV file, and use it to direct our data-cleaning efforts, i.e. look at the top-loss images first!
res_df.to_csv(f'ring_count_top_losses_{dataset_name}.csv', index=False)
# ## When in doubt, look at the data...
# Let's take a look at plots of this data
df2 = res_df.reset_index(drop=True)
plt.plot(df2["target"],'o',label='target')
plt.plot(df2["prediction"],'s', label='prediction')
plt.xlabel('Top-loss order (left=worse, right=better)')
plt.legend(loc='lower right')
plt.ylabel('Ring count')
plt.plot(df2["target"],df2["prediction"],'o')
plt.xlabel('Target ring count')
plt.ylabel('Predicted ring count')
plt.axis('square')
print(f"Target ring count range: ({df2['target'].min()}, {df2['target'].max()})")
print(f"Predicted ring count range: ({df2['prediction'].min()}, {df2['prediction'].max()})")
| nbs/count_in_crops_cyclegan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from parsePython import *
from code2vec import *
from UseData import *
import numpy as np
# +
def move(l1, l2):
result = []
flag = False
for i in l2:
for j in l1:
if i==j:
result.append(j)
flag = True
break
if (flag == False):
result.append(0)
else:
flag = False
for k in l1:
if k not in result:
result.append(k)
return result
def pad(x, size):
if size <= len(x):
return x[:size]
out = [0]*size
out[:len(x)] = x
return out
def get_vec(java_data, py_data):
_c2v = Code2Vec()
_ud = UseData()
_m, _a, _t = _ud.load_use_data()
Xs = []
Xt = []
for i in java_data:
py_token = cv.getToken(py_data[i])
py_api = cv.getAPISequence(py_data[i])
py_meth = cv.getMethodName(py_data[i])
py_token_vec = _c2v.convert_tokens(py_token)
py_apiseq_vec = _c2v.convert_apiseq(py_api)
py_meth_vec= _c2v.convert_methname(py_meth)
java_token_vec = _t[i-1]
java_apiseq_vec = _a[i-1]
java_meth_vec = _m[i-1]
py_token_vec = move(py_token_vec, java_token_vec)
py_apiseq_vec = move(py_apiseq_vec, java_apiseq_vec)
py_meth_vec = move(py_meth_vec, java_meth_vec)
padded_java_token_vec = pad(java_token_vec, 50)
padded_java_apiseq_vec = pad(java_apiseq_vec, 30)
padded_java_meth_vec = pad(java_meth_vec, 6)
padded_py_token_vec = pad(py_token_vec, 50)
padded_py_apiseq_vec = pad(py_apiseq_vec, 30)
padded_py_meth_vec = pad(py_meth_vec, 6)
temp = np.concatenate((padded_java_meth_vec, padded_java_apiseq_vec, padded_java_token_vec), axis=0)
Xs.append(temp)
temp = np.concatenate((padded_py_meth_vec, padded_py_apiseq_vec, padded_py_token_vec), axis=0)
Xt.append(temp)
return Xs, Xt
# -
cv = CodeVisitor()
c2v = Code2Vec()
ud = UseData()
m, a, t = ud.load_use_data()
# +
java_data = {17:"public static String getNextTag ( String xmlData , int position ) { String nextTag = null ; if ( xmlData != null && ! xmlData . isEmpty ( ) && position < xmlData . length ( ) && xmlData . substring ( position ) . contains ( "<" ) ) { while ( xmlData . charAt ( position ) != '<' ) { position ++ ; } int startIndex = position ; if ( xmlData . substring ( position ) . contains ( ">" ) ) { while ( xmlData . charAt ( position ) != '>' ) { position ++ ; } nextTag = xmlData . substring ( startIndex , position + 1 ) ; } } return nextTag ; } ",
7:"public float bottom ( float margin ) { return pageSize . bottom ( marginBottom + margin ) ; } ",
12:"public RGBColor toGreyScale ( ) { return new RGBColor ( 0.30 * getRed ( ) + 0.59 * getGreen ( ) + 0.11 * getBlue ( ) ) ; } ",
21:"public void addClientSocketListener ( ClientSocketListener listener ) { if ( listener == null ) throw new NullPointerException ( ) ; listeners . add ( listener ) ; }",
23:"final void addProcessToGcListLocked ( ProcessRecord proc ) { boolean added = false ; for ( int i = mProcessesToGc . size ( ) - 1 ; i >= 0 ; i -- ) { if ( mProcessesToGc . get ( i ) . lastRequestedGc < proc . lastRequestedGc ) { added = true ; mProcessesToGc . add ( i + 1 , proc ) ; break ; } } if ( ! added ) { mProcessesToGc . add ( 0 , proc ) ; } } ",
34:"public boolean isFocused ( ) { if ( m_Control == null ) return false ; return m_Control . isFocusControl ( ) ; } ",
36:"public static long getFreeDiskSpace ( boolean checkInternal ) { String status = Environment . getExternalStorageState ( ) ; long freeSpace = 0 ; if ( status . equals ( Environment . MEDIA_MOUNTED ) ) { freeSpace = freeSpaceCalculation ( Environment . getExternalStorageDirectory ( ) . getPath ( ) ) ; } else if ( checkInternal ) { freeSpace = freeSpaceCalculation ( \"/\" ) ; } else { return - 1 ; } return freeSpace ; }",
38:"public void removePropertyChangeListener ( String name , PropertyChangeListener pcl ) { m_bcSupport . removePropertyChangeListener ( name , pcl ) ; } ",
40:"public void incDfsUsed ( long value ) { used . addAndGet ( value ) ; } ",
49:"public void removeElements ( List elements ) { if ( elements . size ( ) > 0 ) { fElements . removeAll ( elements ) ; if ( fTable != null ) { fTable . remove ( elements . toArray ( ) ) ; } dialogFieldChanged ( ) ; } } "
}
py_data = {17:"def getNextTag(xmlData, position):\n\tnextTag = None\n\tif xmlData!=None and xmlData!='' and position < len(xmlData) and '<' in xmlData[position:]:\n\t\twhile xmlData[position]!='<':\n\t\t\tposition+=1\n\t\tstartIndex = position\n\t\tif '>' in xmlData[position:]:\n\t\t\twhile xmlData[position] !='>':\n\t\t\t\tposition+=1\n\t\t\tnextTag = xmlData[startIndex:position+1]\n\treturn nextTag",
7:"def bottom(margin):\n\treturn pageSize.bottom(marginBottom+margin)",
12:"def toGreyScale():\n\treturn RGBColor(0.30*getRed() + 0.59*getGreen() + 0.11*getBlue())",
21:"def addClientSocketListenser(listener):\n\tif listener == None:\n\t\traise NullPointerException()\n\tlisteners.add(listener)",
23:"def addProcessToGcListLocked(proc):\n\tadded = False\n\tfor i in range(mProcessesToGc.size()-1, 0, -1):\n\t\tif mProcessesToGc.get(i).lastRequestedGc < proc.lastRequestedGc:\n\t\t\tadded = True\n\t\t\tmProcessesToGc.add(i+1, proc)\n\t\t\tbreak\n\tif not added:\n\t\tmProcessesToGc.add(0, proc)",
34:"def isFocused():\n\tif m_Control==None:\n\t\treturn False\n\treturn m_Control.isFocusControl()",
36:"def getFreeDiskSpace(checkInternal):\n\tstatus=Environment.getExternalStorageState()\n\tfreeSpace = 0\n\tif status.equals(Environment.MEDIA_MOUNTED):\n\t\tfreeSpace = freeSpaceCalculation(Environment.getExternalStorageDirectory().getPath())\n\telif checkInternal:\n\t\tfreeSpace = freeSpaceCalculation(\"/\")\n\telse:\n\t\treturn -1\n\treturn freeSpace",
38:"def removePropertyChangeListener(name, pcl):\n\tm_bcSupport.removePropertyChangeListener(name, pcl)",
40:"def incDfsUsed(value):\n\tused.addAndGet(value)",
49:"def removeElements(elements):\n\tif len(elements)>0:\n\t\tfElements.removeAll(elements)\n\t\tif fTable!=None:\n\t\t\tfTable.remove(elements.toArray())\n\t\tdialogFieldChange()"
}
# -
i=17
token = cv.getToken(py_data[i])
api = cv.getAPISequence(py_data[i])
meth = cv.getMethodName(py_data[i])
print(token)
print(api)
print(meth)
py_token_vec = c2v.convert_tokens(token)
py_apiseq_vec = c2v.convert_apiseq(api)
py_meth_vec= c2v.convert_methname(meth)
print(py_token_vec)
print(py_apiseq_vec)
print(py_meth_vec)
java_token_vec = t[i-1]
java_apiseq_vec = a[i-1]
java_meth_vec = m[i-1]
print(java_token_vec)
print(java_apiseq_vec)
print(java_meth_vec)
py_token_vec = move(py_token_vec, java_token_vec)
py_apiseq_vec = move(py_apiseq_vec, java_apiseq_vec)
py_meth_vec = move(py_meth_vec, java_meth_vec)
print(py_token_vec)
print(py_apiseq_vec)
print(py_meth_vec)
Xs, Xt = get_vec(java_data, py_data)
labels = []
for i in range(len(Xs)):
labels.append(i)
for i in range(len(Xs)):
labels.append(i)
data = np.concatenate((Xs, Xt))
cv.printAST(pydata[i])
a={1:2,2:3,3:4}
a.get(1)
it = a.iteritems()
next(it)
| keras/Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Exploration - My data exploration lab notes
# # Code Like a Pythonista
# ### Iterating over list/set/dict
# Iterating over set
animals = {'cat', 'dog', 'fish', 'monkey'}
for animal in animals:
print('{}'.format(animal))
# Iterating over set (and generating an index)
animals = {'cat', 'dog', 'fish', 'monkey'}
for idx, animal in enumerate(animals):
print('{}: {}'.format(idx,animal))
# ### List/Set/Dict Comprehensions
#
# Python supports list comprehensions, that can be used to construct lists in a very natural way, similar to mathematic construction.
#
#
# [ output_expression() for(set of values to iterate) if(conditional filtering) ]
#
# +
#List comprehension
list1 = [x**2 for x in range(10)]
print(list1)
list2 = [x for x in list1 if x % 2 == 0]
print(list2)
#Set comprehension
nums = {x for x in range(10)}
print(nums) # Prints "{0, 1, 2, 3, 4, 5}"
#Dict comprehension
mcase = {'a':10, 'b': 34, 'A': 7, 'Z':3}
mcase_frequency = { k.lower() : mcase.get(k.lower(), 0) + mcase.get(k.upper(), 0) for k in mcase.keys() }
print(mcase_frequency)
#Set comprehension from list
names = [ 'Bob', 'JOHN', 'alice', 'bob', 'ALICE', 'J', 'Bob' ]
names_set = { name[0].upper() + name[1:].lower() for name in names if len(name) > 1 }
print(names_set)
#Nested list comprehension
matrix = [ [ 1 if item_idx == row_idx else 0 for item_idx in range(0, 3) ] for row_idx in range(0, 3) ]
print(matrix)
# -
# ### The power of comprehension
# +
numbers = range(20)
numbers_doubled_odds = []
for n in numbers:
if n%2 == 1:
numbers_doubled_odds.append(n*2)
print(numbers_doubled_odds)
#vs
numbers_doubled_odds = [n*2 for n in numbers if n%2==1]
print(numbers_doubled_odds)
# -
# Calculating prime numbers
noprimes = [j for i in range(2, 8) for j in range(i*2, 100, i)]
primes = [x for x in range(2, 100) if x not in noprimes]
print(primes)
# # Basic Numpy data structures
# +
import numpy as np
array1D = np.array([1,2,3,4,5,6,7,8, 9, 10])
#Standard print
print('Data in arr1D:\n', array1D)
#The last line is evaluated
array1D
# +
array2D = np.array([[1,2,3,4],[5,6,7,8]])
print('Data in arr2D:\n', array2D)
array2D
# +
# Slicing works the same as in standard Python
array2D = np.array([[1,2,3,4],[5,6,7,8]])
print(array2D)
mini_array2D = array2D[:2, 1:3]
print(mini_array2D)
# -
# ## Creating arrays
# +
array2D = np.zeros((2,4))
print(array2D)
array2D = np.ones((2,4))
print(array2D)
array2D = np.full((2,4),0.8)
print(array2D)
array2D = np.random.random((2,4))
print(array2D)
print(type(array2D))
print(array2D.shape)
# -
# ## Changing the shape of arrays
# +
array1D = np.arange(12)
print(array1D,'\n')
array2D = array1D.reshape(2,6)
print(array2D,'\n')
array2D = array1D.reshape(6,2)
print(array2D,'\n')
array3D = array1D.reshape(2,2,3)
print(array3D,'\n')
array3D = array1D.reshape(3,2,2)
print(array3D,'\n')
# -
# ## Statistics
# +
print('\n Numpy 2-dim array')
tab10n5 = np.random.randn(10,5)
print(tab10n5)
print('\n Standard deviation of array')
print(np.std(tab10n5))
# +
#TODO Statistics functions
# -
# ## Stacking together different arrays
#
# Take a quick look at [tutorial](https://docs.scipy.org/doc/numpy-dev/user/quickstart.html)
# and fill the next cell
#
# +
#TODO stacking arrays
# -
# ## Plotting
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 3*np.pi, 50)
plt.plot(x, np.sin(x**2))
plt.title('A simple chirp');
# -
# ## Importing data
# +
import csv
import numpy as np
data = np.genfromtxt('country-of-birth-london-min.csv', skip_header=1, delimiter=';')
print(data[:5,:])
# -
raw_data = np.genfromtxt('country-of-birth-london-min.csv', delimiter=';', dtype=None)
names = raw_data[:5,:1].astype(str)
print(names)
# +
#TODO - Calculate summary statistics
# +
#TODO - Visualize summary using matplotlib charts
# -
# # Miniproject - data exploration
#
# [Global Terrorism Database](https://www.kaggle.com/START-UMD/gtd)
# # Instructions:
#
# 1. Download data set, Global Terrorism Database, from https://www.kaggle.com/START-UMD/gtd
# 2. Take a quick look at the data set. Check what's inside, how the data is structured, and where the data is corrupted (missing values, bad structure, etc).
# 3. Think and create 5 questions to the data. Try to ask yourself what's really interesting in the data set. What's not so obvious. E.g. some trends, patterns, correlations.
# 4. Create a jupyter notebook and use python, numpy, pandas, matplotlib (at least) to provide all the answers to your questions.
# 5. Create a new github repository, and put your jupyter notebook there.
# 6. Create readme.md file as well in your github root directory with all necessary instructions (what is in the repo, what libs are necessary to run the code, where to find data set and where to save it - this is necessary because the dataset is too big for github repo).
# 7. Provide the necessary documentation and introduction in your notebook using markdown language, at least: data source description, data structure, importing process, data processing process.
# 8. Put some data visualization in your notebook. Sometimes it's much easier to present the answer using a chart rather than numbers
# 9. Check if your notebook run smoothly - use 'Reset & Run All' command from the menu. Save it.
# 10. Export the notebook as HTML as well, and save the file in the repo.
# 11. Do not forget to commit/push all the changes to your repo on hithub.
# 12. Smile :) You did a good job!
#
# FAQ:
# 1. Can I take a look at different solution provided at kaggle? Yes, you can. But check more than one solution. Try to understand what the authors are trying to solve, and how could it be used in your project. Try to find really good examples - easy to understand and not so complicated. Remember - you create the notebook as an instruction to someone else! Try to not complicate the process.
# 2. Jupyter notebook provide R kernel, so can I use R instead? No. Even if you love R, try to solve the project using Python.
#
| dataexp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Research plan**
#
# [Part 0. Mobile Price Classification](#mpc) <br>
# [Part 1. Feature and data explanation](#part1) <br>
# [Part 2. Primary data analysis](#EDA) <br>
# [Part 3. Primary visual data analysis](#part3) <br>
# [Part 4. Insights and found dependencies](#part4) <br>
# [Part 5. Metrics selection](#part5) <br>
# [Part 6. Model selection](#part6) <br>
# [Part 7. Data preprocessing](#part7) <br>
# [Part 8. Cross-validation and adjustment of model hyperparameters](#part8) <br>
# [Part 9. Creation of new features and description of this process](#part9) <br>
# [Part 10. Plotting training and validation curves](#part10) <br>
# [Part 11. Prediction for test or hold-out samples](#part11)<br>
# [Part 12. Conclusions](#part12)<br>
# [Bonus Part. Clustering](#bonus)<br>
# # <center> Mobile Price Classification <a id='mpc'></a>
# <center> Автор: <NAME>
# Oldi zdes'?
# <img src="https://habrastorage.org/webt/xp/br/v9/xpbrv9am-knejmbih1h9b7ndk20.jpeg" />
# # Part 1. Feature and data explanation <a id='part1'></a>
# Bob has started his own mobile company. He wants to give tough fight to big companies like Apple,Samsung etc.
#
# He does not know how to estimate price of mobiles his company creates. In this competitive mobile phone market you cannot simply assume things. To solve this problem he collects sales data of mobile phones of various companies.
#
# Bob wants to find out some relation between features of a mobile phone(eg:- RAM,Internal Memory etc) and its selling price.
#
# In this project we do have to predict price range indicating how high the price is.
# Download dataset from [Kaggle page](https://www.kaggle.com/iabhishekofficial/mobile-price-classification)
# <br>
# Dataset contain train (with target variable) and test (without target variable) samples.
# <br>
# For the <span style="color:red">train</span> sample, we will solve the <span style="color:red">multiclass classification</span> problem with 4 class, and for the <span style="color:blue">test</span> sample we will solve the <span style="color:blue">clustering</span> problem.
# ### The dataset has the following features (copied from Kaggle):
# Every object - it is a unique mobile phone.
# - **battery_power** - Total energy a battery can store in one time measured in mAh (quantitative);
# - **blue** - Has bluetooth or not (binary);
# - **clock_speed** - speed at which microprocessor executes instructions (quantitative);
# - **dual_sim** - Has dual sim support or not (binary);
# - **fc** - Front Camera mega pixels (categorical);
# - **four_g** - Has 4G or not (binary);
# - **int_memory** - Internal Memory in Gigabytes (quantitative);
# - **m_dep** - Mobile Depth in cm (categorical);
# - **mobile_wt** - Weight of mobile phone (quantitative);
# - **n_cores** - Number of cores of processor (categorical);
# - **pc** - Primary Camera mega pixels (categorical);
# - **px_height** - Pixel Resolution Heigh (quantitative);
# - **px_width** - Pixel Resolution Width (quantitative);
# - **ram** - Random Access Memory in Megabytes (quantitative);
# - **sc_h** - Screen Height of mobile in cm (categorical);
# - **sc_w** - Screen Width of mobile in cm (categorical);
# - **talk_time** - longest time that a single battery charge will last when you are (quantitative);
# - **three_g** - Has 3G or not (binary);
# - **touch_screen** - Has touch screen or not (binary);
# - **wifi** - Has wifi or not (binary);
# <br>
#
# - **price_range** - This is the `target variable` with value of 0(low cost), 1(medium cost), 2(high cost) and 3(very high cost). Contain only the in train sample
# # Part 2. Primary data analysis <a id='EDA'></a>
# Importing libraries:
# +
import numpy as np
import pandas as pd
import seaborn as sns
from pylab import rcParams
rcParams['figure.figsize'] = 10, 8
# #%config InlineBackend.figure_format = 'svg'
import warnings
warnings.simplefilter('ignore')
from sklearn.decomposition import PCA
from matplotlib import pyplot as plt
from sklearn.manifold import TSNE
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.multiclass import OneVsRestClassifier, OneVsOneClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_predict, StratifiedKFold, validation_curve
from sklearn.metrics import roc_auc_score, accuracy_score, precision_score, recall_score,\
f1_score, make_scorer, classification_report, confusion_matrix
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
pd.set_option('display.max_rows', 20)
pd.set_option('display.max_columns', 21)
from sklearn import metrics
from sklearn.cluster import KMeans, AgglomerativeClustering, AffinityPropagation, SpectralClustering
from tqdm import tqdm_notebook
from sklearn.metrics.cluster import adjusted_rand_score
from scipy.cluster import hierarchy
from scipy.spatial.distance import pdist
from sklearn.model_selection import learning_curve
from sklearn.model_selection import ShuffleSplit
# -
# Let`s look at data:
data_train = pd.read_csv('../data/mobile/train.csv')
data_test = pd.read_csv('../data/mobile/test.csv')
data_test.drop(columns='id', inplace=True)
data_train.head()
data_test.head()
# In our samples we have quantitative features, categorical and binary features
#
# <br>
# And our samples haven't missing items in the data:
data_train.info()
data_test.info()
# Look at the distribution of target feature:
data_train.groupby('price_range')[['price_range']].count().rename(columns={'price_range': 'count'}).T
# Ok, it is a toy dataset..)We see that the target variable is uniform distributed
# # Part 3. Primary visual data analysis <a id='part3'></a>
# Let's draw plot of correlation matrix (before this, drop a boolean variables):
corr_matrix = data_train.drop(['blue', 'dual_sim', 'four_g', 'three_g', 'touch_screen', 'wifi'], axis=1).corr()
fig, ax = plt.subplots(figsize=(16,12))
sns.heatmap(corr_matrix,annot=True,fmt='.1f',linewidths=0.5);
# Ok, we see that there is a correlation between the `target` variable and four features: `battery_power`, `px_height`, `px_width` and `ram`.
#
#
# And some variables are correlated with each other: `pc` and `fc` (photo modules), `sc_w` and `sc_h` (screen width and heght), `px_width` and `px_height` (pixel resolution heigh and width).
# Draw plot of distribution of target variable:
data_train['price_range'].value_counts().plot(kind='bar',figsize=(14,6))
plt.title('Distribution of target variable');
# Ok, we again see that the target variable is uniform distributed
# Look at the distribution of quantitative features:
features = list(data_train.drop(['price_range', 'blue', 'dual_sim',\
'four_g', 'fc', 'm_dep', 'n_cores',\
'pc', 'sc_h', 'sc_w', 'three_g', 'wifi', 'touch_screen'], axis=1).columns)
data_train[features].hist(figsize=(20,12));
# Let's look at the interaction of different features among themselves with `sns.pairplot`:
sns.pairplot(data_train[features + ['price_range']], hue='price_range');
# We see that the `ram` feature of a good separates our objects by different price categories.
# Construct the `sns.boxplot`, describe the distribution statistics of quantitative traits:
# +
fig, axes = plt.subplots(nrows=2, ncols=4, figsize=(20, 12))
for idx, feat in enumerate(features):
sns.boxplot(x='price_range', y=feat, data=data_train, ax=axes[int(idx / 4), idx % 4])
axes[int(idx / 4), idx % 4].set_xlabel('price_range')
axes[int(idx / 4), idx % 4].set_ylabel(feat);
# -
# We see that it is better to difference our price categories the following features: `battery_power`, `px_height`, `px_width` и `ram`. As well as the plot of the correlation matrix.
# Ok, let's plot the distribution for `sc_w` - categorical feature:
fig, ax = plt.subplots(figsize=(16,10))
sns.countplot(x='sc_w', hue='price_range', data=data_train);
# Wee see that count of our object decreases with increasing width
# plot the distribution for `sc_w` - categorical feature:
fig, ax = plt.subplots(figsize=(16,10))
sns.countplot(x='sc_h', hue='price_range', data=data_train);
# Now let's look at the connection of binary features of `blue`, `dual_sim`, `four_g` and `three_g` with our target `price_range`.
# +
_, axes = plt.subplots(1, 4, sharey=True, figsize=(16,6))
sns.countplot(x='blue', hue='price_range', data=data_train, ax=axes[0]);
sns.countplot(x='dual_sim', hue='price_range', data=data_train, ax=axes[1]);
sns.countplot(x='four_g', hue='price_range', data=data_train, ax=axes[2]);
sns.countplot(x='three_g', hue='price_range', data=data_train, ax=axes[3]);
# -
# All about the same, but count objects with 3G more than without.
# Now, let's build a t-SNE representation:
X = data_train.drop('price_range', axis=1)
y = data_train.price_range
# %%time
tsne = TSNE(random_state=17)
tsne_representation = tsne.fit_transform(X)
fig, ax = plt.subplots(figsize=(16,10))
cmap = sns.cubehelix_palette(dark=.1, light=.8, as_cmap=True)
sns.scatterplot(tsne_representation[:, 0], tsne_representation[:, 1],\
s=100, hue=data_train['price_range'], palette="Accent");
plt.title('t-SNE projection');
# We see that the object is well distinguished.
# Let's look at another representation of the `scaled data` colored by binary features:
# %%time
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
tsne2 = TSNE(random_state=17)
tsne_representation2 = tsne2.fit_transform(X_scaled)
# +
_, axes = plt.subplots(2, 2, sharey=True, figsize=(16,10))
axes[0][0].scatter(tsne_representation2[:, 0], tsne_representation2[:, 1],
c=data_train['three_g'].map({0: 'blue', 1: 'orange'}));
axes[0][1].scatter(tsne_representation2[:, 0], tsne_representation2[:, 1],
c=data_train['four_g'].map({0: 'blue', 1: 'orange'}));
axes[1][0].scatter(tsne_representation2[:, 0], tsne_representation2[:, 1],
c=data_train['blue'].map({0: 'blue', 1: 'orange'}));
axes[1][1].scatter(tsne_representation2[:, 0], tsne_representation2[:, 1],
c=data_train['dual_sim'].map({0: 'blue', 1: 'orange'}));
# -
# Ok, we see that the binary features are a bunch).
# # Part 4. Insights and found dependencies <a id='part4'></a>
# Combining the observation from the previous paragraphs, the following is to be denoted:<ul>
# <li>The dataset under analysis don`t contain omissions. That's no wonder: we have a toy dataset.</li>
# <li>The distribution of the target feature, <b>price_range</b> is uniform distribution, so this is again because we have toy data.</li>
# <li>The <b>pc</b> and <b>fc</b> columns to be interconnected features, as well as <b>sc_w</b> and <b>sc_h</b>, and <b>px_width</b> and <b>px_height</b> interconnected to. This is understandable, because the screen must be of a certain shape, basically the phone screens are similar to each other.</li>
# <li>The some features have a good correlation with the target variable <b>price_range</b>, such as <b>battery_power</b>, <b>px_height</b>, <b>px_width</b>, <b>ram</b>.</li>
# <li>On the sns.pairplot <b>ram</b> feature and the <b>t-SNE</b> representation we see that the objects are well separated from each other, and separated linearly, respectively, assume that models that work well with linearly separable objects, and those models that take into account distance, will be very useful here </li>
# <li>For the next part of creating features, there are some baselines that we can come up with.</li>
# </ul>
# # Part 5. Metrics selection <a id='part5'></a>
# We have a problem of multi-class classification. It is necessary to predict the class itself, not the probability of belonging to the class, so we use the metrics from the classification problem, namely `accuracy`, `precision`, `recall `, `f1`. The basic metric we will have is `accuracy` but we will use `classification_report` to estimate other metrics.
#
# We can use `accuracy`, because we have uniform distribution of target variable.
#
# $$\mathcal accuracy = \dfrac{1}{l}\sum_{i=1}^l [a(x_{i})=y_{i}]$$
#
# We will also consider the `confusion matrix`, columns `i` - true class label, line `j` - assessment of class membership from our algorithm, where $q_{ij}$:
#
# $$\mathcal q_{ij} = \sum_{m=1}^l [a(x_{m})=i][y_{m}=j]$$
# # Part 6. Model selection <a id='part6'></a>
# So, we have a problem of multi-class classification, and as we already know our task linearly separable.
# That's why we can use `LogisticRegression`. Well, we have four classes, and to solve this problem is well suited `OneVsOneClassifier` - a model that trains K(K-1) models for each pair of classes.
#
# With a problem of multi-class classification the following models also work well by default:
#
# - KNeighborsClassifier
# - RandomForestClassifier
# - SVC
# # Part 7. Data preprocessing <a id='part7'></a>
# We divide our sample into a matrix of features and a vector of answers:
X = data_train.drop('price_range', axis=1)
y = data_train.price_range
# Let's make a split into a train sample and hold-out sample:
X_train_part, X_valid, y_train, y_valid = train_test_split(X, y, test_size=0.3, stratify=y, random_state=17)
# Some models should not be scaled, but for others it is necessary:
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
X_train_part_scaled, X_valid_scaled, y_train, y_valid = train_test_split(X_scaled, y,\
test_size=0.3, stratify=y, random_state=17)
# # Part 8. Cross-validation and adjustment of model hyperparameters <a id='part8'></a>
# ## `LogisticRegression` with scaled features:
lr = LogisticRegression(random_state=17)
lr.fit(X_train_part_scaled, y_train);
print(accuracy_score(y_valid, lr.predict(X_valid_scaled)))
print(classification_report(y_valid, lr.predict(X_valid_scaled)))
# #### Confusion matrix for `LogisticRegression`:
tab = pd.crosstab(y_valid, lr.predict(X_valid_scaled), margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = tab.index
tab
# For `GridSearchCV` we choose the parameters of `LogisticRegression`: <b>C</b> - Inverse of regularization strength, smaller values specify stronger regularization. <b>solver</b> - Algorithm to use in the optimization problem. <b>class_weight</b> - Weights associated with classes in the form {class_label: weight}.
# +
params = {'C': np.logspace(-5, 5, 11),
'solver' : ['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'],
'class_weight' : ['balanced', None]}
lr_grid = GridSearchCV(lr, params, n_jobs=-1, cv=5, scoring='accuracy', verbose=1)
lr_grid.fit(X_train_part_scaled, y_train);
# -
print(accuracy_score(y_valid, lr_grid.predict(X_valid_scaled)))
print(classification_report(y_valid, lr_grid.predict(X_valid_scaled)))
lr_grid.best_params_, lr_grid.best_score_
# Nice, after `GridSearchCV` we see that score increase.
# #### Confusion matrix for `LogisticRegression` after `GridSearchCV`:
tab = pd.crosstab(y_valid, lr_grid.predict(X_valid_scaled), margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = tab.index
tab
# ## `KNeighborsClassifier` with unscaled features:
# +
kneigh = KNeighborsClassifier()
kneigh.fit(X_train_part, y_train)
print(accuracy_score(y_valid, kneigh.predict(X_valid)))
print(classification_report(y_valid, kneigh.predict(X_valid)))
# -
# #### Confusion matrix for `KNeighborsClassifier`:
tab = pd.crosstab(y_valid, kneigh.predict(X_valid), margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = tab.index
tab
# ## `OneVsOneClassifier` with scaled features:
# +
clf = OneVsOneClassifier(LogisticRegression(random_state=17))
clf.fit(X_train_part_scaled, y_train);
print(accuracy_score(y_valid, clf.predict(X_valid_scaled)))
print(classification_report(y_valid, clf.predict(X_valid_scaled)))
# -
# #### Doing `GridSearchCV` for `OneVsOneClassifier` with `LogisticRegression`:
# +
params = {'estimator__C': np.logspace(-5, 5, 11),
'estimator__solver' : ['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'],
'estimator__class_weight' : ['balanced', None]}
clf_grid = GridSearchCV(clf, params, n_jobs=-1, cv=5, scoring='accuracy', verbose=1)
clf_grid.fit(X_train_part_scaled, y_train);
# -
print(accuracy_score(y_valid, clf_grid.predict(X_valid_scaled)))
print(classification_report(y_valid, clf_grid.predict(X_valid_scaled)))
clf_grid.best_params_, clf_grid.best_score_
# #### Confusion matrix for `OneVsOneClassifier` after `GridSearchCV`:
tab = pd.crosstab(y_valid, clf_grid.predict(X_valid_scaled), margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = tab.index
tab
# For this task `OneVsOneClassifier` very good classifier!
# ## `RandomForestClassifier` with unscaled features:
rf_clf = RandomForestClassifier(random_state=17)
rf_clf.fit(X_train_part, y_train)
print(accuracy_score(y_valid, rf_clf.predict(X_valid)))
print(classification_report(y_valid, rf_clf.predict(X_valid)))
#print(confusion_matrix(y_valid, rf_clf.predict(X_valid)))
# #### Let's see `feature_importances_ ` for `RandomForestClassifier`:
pd.DataFrame({'feat': X_train_part.columns,
'coef': np.abs(rf_clf.feature_importances_).flatten().tolist()}).\
sort_values(by='coef', ascending=False).head()
# #### No wonder the correlation matrix told us that already.
# #### Confusion matrix for `RandomForestClassifier`:
tab = pd.crosstab(y_valid, rf_clf.predict(X_valid), margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = tab.index
tab
# ## `SVC` with unscaled features:
# +
svc = SVC(kernel='linear', probability=True, random_state=17)
svc.fit(X_train_part, y_train);
print(accuracy_score(y_valid, svc.predict(X_valid)))
print(classification_report(y_valid, svc.predict(X_valid)))
# +
svc = SVC(kernel='linear', probability=True, random_state=17)
svc.fit(X_train_part, y_train);
print(accuracy_score(y_valid, svc.predict(X_valid)))
print(classification_report(y_valid, svc.predict(X_valid)))
# -
# #### Doing `GridSearchCV` for `SVC`:
# +
# %%time
params_svc = {'C': np.logspace(-1, 1, 3),
'decision_function_shape': ['ovr', 'ovo'],
'class_weight' : ['balanced', None]}
svc_grid = GridSearchCV(svc, params_svc, n_jobs=-1, cv=3, scoring='accuracy', verbose=1)
svc_grid.fit(X_train_part, y_train);
# -
print(accuracy_score(y_valid, svc_grid.predict(X_valid)))
print(classification_report(y_valid, svc_grid.predict(X_valid)))
svc_grid.best_params_, svc_grid.best_score_
# #### Confusion matrix for `SVC` after `GridSearchCV`:
tab = pd.crosstab(y_valid, svc_grid.predict(X_valid), margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = tab.index
tab
# ### We have 2 models with amazing score - `OneVsOneClassifier` with `LogisticRegression` (scaled features), and `SVC` (unscaled features), with `accuracy = 0.9766` and `accuracy = 0.98` after `GridSearchCV` respectively!
# # Part 9. Creation of new features and description of this process <a id='part9'></a>
# The `inch` (abbreviation: in or ″) is a unit of length in the (British) imperial and United States customary systems of measurement. It is equal to 1⁄36 yard or 1⁄12 of a foot. Derived from the Roman uncia ("twelfth"), the word inch is also sometimes used to translate similar units in other measurement systems, usually understood as deriving from the width of the human thumb. Standards for the exact length of an inch have varied in the past, but since the adoption of the international yard during the 1950s and 1960s it has been based on the metric system and defined as exactly <b>2.54 cm</b>.
# Pixels per inch (`ppi`) or pixels per centimeter (ppcm) are measurements of the pixel density (resolution) of an electronic image device, such as a computer monitor or television display, or image digitizing device such as a camera or image scanner. Horizontal and vertical density are usually the same, as most devices have square pixels, but differ on devices that have non-square pixels.
# So, i think `ppi` it is a good feature, because, than the larger the value, the sharper the image.
#
# Let's check this.
data_train2 = data_train.copy()
data_train2['inch'] = (np.sqrt(data_train2['sc_h']**2 + data_train2['sc_w']**2)/2.54).astype('int')
data_train2['ppi'] = np.sqrt(data_train2['px_width']**2 + data_train2['px_height']**2)/data_train2['inch']
# Also make a feature that is based on the current modern phones:
data_train2['top'] = ((data_train2['touch_screen'] ==1)|\
(data_train2['ppi'] >=500)&\
(data_train2['inch'] >=5)&\
(data_train2['four_g'] ==1)|\
(data_train2['blue'] ==1)|\
(data_train2['int_memory'] >=36)|\
(data_train2['ram'] >=2600)).astype('int64')
data_train2['top'].value_counts()
# Ok, let's check these features on our models:
# For `SVC` unscaled matrix features:
X_train2, y2 = data_train2.drop(['price_range','inch'], axis=1), data_train2['price_range']
X_train_part2, X_valid2, y_train2, y_valid2 = train_test_split\
(X_train2, y2, test_size=.3, stratify=y2, random_state=17)
# +
svc2 = SVC(kernel='linear', probability=True, random_state=17)
svc2.fit(X_train_part2, y_train2);
print(accuracy_score(y_valid2, svc2.predict(X_valid2)))
print(classification_report(y_valid2, svc2.predict(X_valid2)))
# +
# %%time
params_svc2 = {'C': np.logspace(-1, 1, 3),
'decision_function_shape': ['ovr', 'ovo'],
'class_weight' : ['balanced', None]}
svc_grid2 = GridSearchCV(svc2, params_svc2, n_jobs=-1, cv=3, scoring='accuracy', verbose=1)
svc_grid2.fit(X_train_part2, y_train2);
# -
print(accuracy_score(y_valid2, svc_grid2.predict(X_valid2)))
print(classification_report(y_valid2, svc_grid2.predict(X_valid2)))
svc_grid2.best_params_, svc_grid2.best_score_
# For `OneVsOneClassifier` with `LogisticRegression` unscaled matrix features:
X2 = data_train2.drop(['price_range','inch'], axis=1)
scaler2 = StandardScaler()
X_scaled2, y2 = scaler2.fit_transform(X2), data_train2['price_range']
X_train_part_scaled2, X_valid_scaled2, y_train2, y_valid2 = train_test_split\
(X_scaled2, y2, test_size=.3, stratify=y2, random_state=17)
# +
clf2 = OneVsOneClassifier(LogisticRegression(random_state=17))
clf2.fit(X_train_part_scaled2, y_train2);
print(accuracy_score(y_valid2, clf2.predict(X_valid_scaled2)))
print(classification_report(y_valid2, clf2.predict(X_valid_scaled2)))
# +
params2 = {'estimator__C': np.logspace(-5, 5, 11),
'estimator__solver' : ['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'],
'estimator__class_weight' : ['balanced', None]}
clf_grid2 = GridSearchCV(clf2, params2, n_jobs=-1, cv=5, scoring='accuracy', verbose=1)
clf_grid2.fit(X_train_part_scaled2, y_train2);
# -
print(accuracy_score(y_valid2, clf_grid2.predict(X_valid_scaled2)))
print(classification_report(y_valid2, clf_grid2.predict(X_valid_scaled2)))
clf_grid2.best_params_, clf_grid2.best_score_
# Ok, with new features we observe the following situation: `OneVsOneClassifier` with `LogisticRegression` by comparison with a default train sample after `GridSearchCV` increase score and now - `accuracy = 0.98`.
# `SVC` with new features and without using `GridSearchCV` increase score : there was `accuracy = 0.9766` it became like this `accuracy = 0.9783`. But `GridSearchCV` is not increased the result.
# # Part 10. Plotting training and validation curves <a id='part10'></a>
def plot_learning_curve(estimator, title, X, y, ylim=None, cv=None,
n_jobs=None, train_sizes=np.linspace(.1, 1.0, 5)):
plt.figure()
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
return plt
# Plotting training and validation curves for grid model with new features `SVC`:
svc3 = SVC(C=0.1, kernel='linear', probability=True, class_weight='balanced', random_state=17)
title = "Learning Curves (SVM, Linear kernel, C=0.1)"
plot_learning_curve(svc3, title, X_train_part2, y_train2, (0.7, 1.01), cv=20, n_jobs=4)
plt.show()
# Plotting training and validation curves for grid model with new features `OneVsOneClassifier` with `LogisticRegression`:
clf3 = OneVsOneClassifier(LogisticRegression(C=100,\
class_weight='balanced', solver='newton-cg', random_state=17))
title = "Learning Curves (OneVsOneClassifier, LogisticRegression base model, C=100)"
plot_learning_curve(clf3, title, X_train_part_scaled2, y_train2, (0.7, 1.01), cv=20, n_jobs=4)
plt.show()
# We see that the curves practically converge, this indicates a high quality of the forecast and if we continue to move to the right (add data to the model), we can still improve the quality of the validation.
# # Part 11. Prediction for test or hold-out samples <a id='part11'></a>
# Was discussed in Part 8 and Part 9
# # Part 12. Conclusions <a id='part12'></a>
# We had a problem with multi-class classification, and we saw that the following methods do a better job: `OneVsOneClassifier` with `LogisticRegression` and `SVC`.
# We got very good score.
#
# Now Bob knows how to evaluate phones of his own production!
#
# Further ways to improve the solution:
#
# - To collect additional characteristics about the components of the phone (Manufacturer, type, brand);
# - Collect data about other phones;
# - Make more new features;
# - Combine multiple predictive models;
#
# # Bonus Part. Clustering <a id='bonus'></a>
# ## Сonsider the train sample:
# Reduce the dimension while preserving the variance:
pca = PCA(n_components=0.9, random_state=17).fit(X2)
X_pca = pca.transform(X2)
# Projections our data for the first two dimension:
plt.figure(figsize=(16,12))
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y2, s=100, cmap=plt.cm.get_cmap('nipy_spectral', 4));
plt.colorbar();
# t-SNE representation our data for the first two dimension:
# +
# %%time
tsne3 = TSNE(random_state=17)
X_tsne = tsne3.fit_transform(X2)
plt.figure(figsize=(16,10))
plt.scatter(X_tsne[:, 0], X_tsne[:, 1], c=y2,
edgecolor='none', alpha=0.7, s=200,
cmap=plt.cm.get_cmap('viridis', 4))
plt.colorbar()
plt.title('t-SNE projection')
# -
# K-MEANS Clustering:
kmeans = KMeans(n_clusters=4,random_state=17, n_jobs=1)
kmeans.fit(X_pca)
kmeans_labels = kmeans.labels_+1
plt.figure(figsize=(16,12))
plt.scatter(X_pca[:, 0], X_pca[:, 1],\
c=kmeans_labels, s=20,\
cmap=plt.cm.get_cmap('nipy_spectral', 4));
plt.colorbar();
# Confusion matrix are very bad:
tab = pd.crosstab(y2, kmeans_labels, margins=True)
tab.index = ['low cost', 'medium cost', 'high cost', 'very high cost', 'all']
tab.columns = ['cluster' + str(i + 1) for i in range(4)] + ['all']
tab
pd.Series(tab.iloc[:-1,:-1].max(axis=1).values /
tab.iloc[:-1,-1].values, index=tab.index[:-1])
inertia = []
for k in tqdm_notebook(range(1, 12)):
kmeans = KMeans(n_clusters=k, random_state=17).fit(X2)
inertia.append(np.sqrt(kmeans.inertia_))
plt.plot(range(1, 12), inertia, marker='s');
plt.xlabel('$k$')
plt.ylabel('$J(C_k)$');
# Agglomerative Clustering:
ag = AgglomerativeClustering(n_clusters=4,
linkage='ward').fit(X_pca)
ag_labels = ag.labels_+1
plt.figure(figsize=(16,12))
plt.scatter(X_pca[:, 0], X_pca[:, 1],\
c=ag_labels, s=20,\
cmap=plt.cm.get_cmap('nipy_spectral', 4));#cmap='viridis');
plt.colorbar();
# Score ARI for K-MEANS and Agglomerative Clustering:
adjusted_rand_score(y2, ag.labels_)
adjusted_rand_score(y2, kmeans.labels_)
# Dendrogram:
# +
distance_mat = pdist(X2) # pdist calculates the upper triangle of the distance matrix
Z = hierarchy.linkage(distance_mat, 'single') # linkage is agglomerative clustering algorithm
plt.figure(figsize=(10, 5))
dn = hierarchy.dendrogram(Z, color_threshold=0.5)
# -
# ### A summary of the score on the train sample:
# +
algorithms = []
algorithms.append(KMeans(n_clusters=4, random_state=17))
algorithms.append(AffinityPropagation())
algorithms.append(SpectralClustering(n_clusters=4, random_state=17,
affinity='nearest_neighbors'))
algorithms.append(AgglomerativeClustering(n_clusters=4))
data = []
for algo in algorithms:
algo.fit(X_pca)
data.append(({
'ARI': metrics.adjusted_rand_score(y2, algo.labels_),
'AMI': metrics.adjusted_mutual_info_score(y2, algo.labels_),
'Homogenity': metrics.homogeneity_score(y2, algo.labels_),
'Completeness': metrics.completeness_score(y2, algo.labels_),
'V-measure': metrics.v_measure_score(y2, algo.labels_),
'Silhouette': metrics.silhouette_score(X_pca, algo.labels_)}))
results = pd.DataFrame(data=data, columns=['ARI', 'AMI', 'Homogenity',
'Completeness', 'V-measure',
'Silhouette'],
index=['K-means', 'Affinity',
'Spectral', 'Agglomerative'])
results
# -
# ## Сonsider the test sample:
X3 = data_test
pca3 = PCA(n_components=0.9, random_state=17).fit(X3)
X_pca3 = pca3.transform(X3)
kmeans = KMeans(n_clusters=4,random_state=17, n_jobs=1)
kmeans.fit(X_pca3)
kmeans_labels = kmeans.labels_+1
plt.figure(figsize=(16,12))
plt.scatter(X_pca3[:, 0], X_pca3[:, 1],\
c=kmeans_labels, s=20,\
cmap=plt.cm.get_cmap('nipy_spectral', 4));#cmap='viridis');
plt.colorbar();
ag = AgglomerativeClustering(n_clusters=4,
linkage='ward').fit(X_pca3)
ag_labels = ag.labels_+1
plt.figure(figsize=(16,12))
plt.scatter(X_pca3[:, 0], X_pca3[:, 1],\
c=ag_labels, s=20,\
cmap=plt.cm.get_cmap('nipy_spectral', 4));#cmap='viridis');
plt.colorbar();
# ### We can only evaluate with silhouette:
metrics.silhouette_score(X_pca3, ag_labels)
metrics.silhouette_score(X_pca3, kmeans_labels)
| jupyter_english/projects_indiv/mobile_price_classification_trefilov_andrew.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
medium_df=pd.read_csv('/Volumes/other/college/h2/medium_data.csv')
medium_df
medium_df.info()
medium_df.columns
required_columns = ['title','subtitle', 'image', 'claps', 'responses',
'reading_time', 'publication', 'date']
medium_df_eda = medium_df[required_columns].copy()
medium_df.info()
medium_df_eda.info()
medium_df_eda.date = pd.to_datetime(medium_df_eda.date)
medium_df_eda.info()
print(medium_df_eda.date[0])
print(medium_df_eda.date[0].day)
print(medium_df_eda.date[0].month)
print(medium_df_eda.date[0].year)
medium_df_eda.image = medium_df_eda.image.str.replace('[0-9.]','',regex=True)
medium_df_eda.image.value_counts()
#what? we have 2 rows with no image. let's remove these two rows to make this dataset cleaner.
medium_df_eda.image.replace('',np.nan, inplace=True)
medium_df_eda.dropna(subset=['image'], inplace=True)
medium_df_eda.image.value_counts()
# +
#And its removed :) But we also have some upper extensions in upper case let's turn them into lower case
# -
medium_df_eda.image = medium_df_eda.image.str.lower()
medium_df_eda.image.value_counts()
#Exploratory Analysis and Visualization¶
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
publication_articles_count = medium_df_eda.publication.value_counts().rename_axis('publications').reset_index(name='counts')
publication_articles_count
plt.figure(figsize=(12,6))
plt.xlabel("number of articles")
plt.title("Number of articles by publications")
sns.barplot(y=publication_articles_count.publications,x=publication_articles_count.counts);
articles_df = medium_df_eda.date.value_counts().rename_axis('dates').reset_index(name='counts')
articles_df.info()
articles_df['month'] = articles_df.dates.dt.month_name()
articles_df['day_of_week'] = articles_df.dates.dt.day_name()
articles_df
rest_data = articles_df.pivot_table(index='month', columns='day_of_week', values='counts', aggfunc='sum', fill_value=0)
rest_data = rest_data[['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday','Sunday']]
# +
#The second line of the code is used to get weeks in order i.e. mon, tue, etc. Otherwise it will be sorted alphabetically.
# -
rest_data
plt.figure(figsize=(12,6))
plt.title("when were the blogs posted?")
sns.heatmap(rest_data, cmap="Greens", linewidths=.5)
# +
#Now that we understood what is the best day to post an article, let's try to get understand how long should the article be.
#we are going to plot the graph with number of claps vs reading time of the article. Assumtion is that a person will give a clap to article only when he reads it completly and finds it useful/entertaining.
# -
plt.figure(figsize=(12,6))
plt.xlabel("Reading time in minutes")
plt.ylabel("number of claps")
sns.scatterplot(medium_df_eda.reading_time,medium_df_eda.claps)
plt.figure(figsize=(19,6))
plt.xlabel("Reading time in minutes")
plt.ylabel("number of claps")
sns.scatterplot(medium_df_eda.responses, medium_df_eda.reading_time)
# +
#so, lets check what image extentions are uses in thes 60k articles dataframe. Pie chart with percentages will be useful for this analysis so lets plot that by using value_counts method
# -
medium_df_eda['image'].value_counts().plot(kind='pie', figsize=(10, 9), autopct='%1.1f%%')
plt.legend(medium_df_eda.image.unique())
plt.title("Type of images used")
medium_df_eda
#Q: what percent of articles have subtitle after heading, is it necessary?¶
percentage = medium_df_eda.subtitle.count()/len(medium_df_eda) * 100
print("About {} percent of articles have subtitles".format(percentage))
#Q: The dataset that we are using is in which time span
print("The data of blogs we have is from {} to {}".format(medium_df_eda.date.min(), medium_df_eda.date.max()))
# +
#So we have data for 2019 starting from 26th january to 30th December
# +
#Q: what is the avarage reading time of articles according to the publications
# -
avg_reading_time_df = medium_df_eda[["publication","reading_time"]].groupby("publication").mean()
print(avg_reading_time_df)
print("--------------------------------------")
print(medium_df_eda.publication.value_counts())
# +
#so from this we can see that most of the articles are written by The Startup publication and their avarage reading time is around 6 minutes and this also backs up our analysis i.e. ideal reading time is 5 - 10 minutes from the scatterplots.
# +
#Q: Which articles among the publications have maximum number of claps?
# -
indexes = medium_df_eda.groupby(['publication'], sort=False)['claps'].transform(max) == medium_df_eda['claps']
# +
#The code above will give us indexes of the rows with maximum number of claps and then we can select those rows from out dataset
# -
best_articles = medium_df_eda[indexes]
best_articles
# +
#So, these are the articles from specfic publications with maximum number of claps
# +
#Q: What is the reading time for the article with highest claps among the publications
# -
for index, row in best_articles.iterrows():
print("The article '{}' from '{}' publication has highest number of claps i.e. '{}'".format(row.title, row.publication, row.claps))
print("\n")
for index, row in best_articles.iterrows():
print("The best article from {} has reading time of {} minutes".format(row.publication, row.reading_time))
print("\n")
# +
#Q: What is the avarage reading time for the article with highest claps among the publications
# -
best_articles.reading_time.mean()
# +
#This number again backs up two of our analysis which we did previously i.e. 5 - 10 minutes. Hence we can say that reading time plays an effective role in success of the article.
# +
#Q: what percent of best blogs have subtitles?
# -
percentage = best_articles.subtitle.count()/len(best_articles) * 100
print("About {} percent of articles have subtitles".format(percentage))
| EDA-on-Medium-Blogs.ipynb |