code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
theBoard = {'7': ' ' , '8': ' ' , '9': ' ' ,
'4': ' ' , '5': ' ' , '6': ' ' ,
'1': ' ' , '2': ' ' , '3': ' ' }
def printBoard(board):
print(board['7'] + '|' + board['8'] + '|' + board['9'])
print('-+-+-')
print(board['4'] + '|' + board['5'] + '|' + board['6'])
print('-+-+-')
print(board['1'] + '|' + board['2'] + '|' + board['3'])
# -
printBoard(board)
def game():
turn = 'X'
count = 0
for i in range(10):
print("It's your turn," + turn + ".Move to which place?")
move = input()
if theBoard[move] == ' ':
theBoard[move] = turn
count += 1
else:
print("That place is already filled.\nMove to which place?")
continue
if count >= 5:
if theBoard['7'] == theBoard['8'] == theBoard['9'] != ' ': # across the top
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['4'] == theBoard['5'] == theBoard['6'] != ' ': # across the middle
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['2'] == theBoard['3'] != ' ': # across the bottom
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['4'] == theBoard['7'] != ' ': # down the left side
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['2'] == theBoard['5'] == theBoard['8'] != ' ': # down the middle
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['3'] == theBoard['6'] == theBoard['9'] != ' ': # down the right side
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['7'] == theBoard['5'] == theBoard['3'] != ' ': # diagonal
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['5'] == theBoard['9'] != ' ': # diagonal
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
# If neither X nor O wins and the board is full, we'll declare the result as 'tie'.
if count == 9:
print("\nGame Over.\n")
print("It's a Tie!!")
# we have to change the player after every move.
if turn =='X':
turn = 'O'
else:
turn = 'X'
printBoard(board):
# +
#Implementation of Two Player Tic-Tac-Toe game in Python.
''' We will make the board using dictionary
in which keys will be the location(i.e : top-left,mid-right,etc.)
and initialliy it's values will be empty space and then after every move
we will change the value according to player's choice of move. '''
theBoard = {'7': ' ' , '8': ' ' , '9': ' ' ,
'4': ' ' , '5': ' ' , '6': ' ' ,
'1': ' ' , '2': ' ' , '3': ' ' }
board_keys = []
for key in theBoard:
board_keys.append(key)
''' We will have to print the updated board after every move in the game and
thus we will make a function in which we'll define the printBoard function
so that we can easily print the board everytime by calling this function. '''
def printBoard(board):
print(board['7'] + '|' + board['8'] + '|' + board['9'])
print('-+-+-')
print(board['4'] + '|' + board['5'] + '|' + board['6'])
print('-+-+-')
print(board['1'] + '|' + board['2'] + '|' + board['3'])
# Now we'll write the main function which has all the gameplay functionality.
def game():
turn = 'X'
count = 0
for i in range(10):
printBoard(theBoard)
print("It's your turn," + turn + ".Move to which place?")
move = input()
if theBoard[move] == ' ':
theBoard[move] = turn
count += 1
else:
print("That place is already filled.\nMove to which place?")
continue
# Now we will check if player X or O has won,for every move after 5 moves.
if count >= 5:
if theBoard['7'] == theBoard['8'] == theBoard['9'] != ' ': # across the top
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['4'] == theBoard['5'] == theBoard['6'] != ' ': # across the middle
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['2'] == theBoard['3'] != ' ': # across the bottom
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['4'] == theBoard['7'] != ' ': # down the left side
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['2'] == theBoard['5'] == theBoard['8'] != ' ': # down the middle
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['3'] == theBoard['6'] == theBoard['9'] != ' ': # down the right side
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['7'] == theBoard['5'] == theBoard['3'] != ' ': # diagonal
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['5'] == theBoard['9'] != ' ': # diagonal
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
# If neither X nor O wins and the board is full, we'll declare the result as 'tie'.
if count == 9:
print("\nGame Over.\n")
print("It's a Tie!!")
# Now we have to change the player after every move.
if turn =='X':
turn = 'O'
else:
turn = 'X'
# Now we will ask if player wants to restart the game or not.
restart = input("Do want to play Again?(y/n)")
if restart == "y" or restart == "Y":
for key in board_keys:
theBoard[key] = " "
game()
if __name__ == "__main__":
game()
# -
# +
# Tic-Tac-Toe Program using
# random number in Python
# importing all necessary libraries
import numpy as np
import random
from time import sleep
# Creates an empty board
def create_board():
return(np.array([[0, 0, 0],
[0, 0, 0],
[0, 0, 0]]))
# Check for empty places on board
def possibilities(board):
l = []
for i in range(len(board)):
for j in range(len(board)):
if board[i][j] == 0:
l.append((i, j))
return(l)
# Select a random place for the player
def random_place(board, player):
selection = possibilities(board)
current_loc = random.choice(selection)
board[current_loc] = player
return(board)
# Checks whether the player has three
# of their marks in a horizontal row
def row_win(board, player):
for x in range(len(board)):
win = True
for y in range(len(board)):
if board[x, y] != player:
win = False
continue
if win == True:
return(win)
return(win)
# Checks whether the player has three
# of their marks in a vertical row
def col_win(board, player):
for x in range(len(board)):
win = True
for y in range(len(board)):
if board[y][x] != player:
win = False
continue
if win == True:
return(win)
return(win)
# Checks whether the player has three
# of their marks in a diagonal row
def diag_win(board, player):
win = True
y = 0
for x in range(len(board)):
if board[x, x] != player:
win = False
if win:
return win
win = True
if win:
for x in range(len(board)):
y = len(board) - 1 - x
if board[x, y] != player:
win = False
return win
# Evaluates whether there is
# a winner or a tie
def evaluate(board):
winner = 0
for player in [1, 2]:
if (row_win(board, player) or
col_win(board,player) or
diag_win(board,player)):
winner = player
if np.all(board != 0) and winner == 0:
winner = -1
return winner
# Main function to start the game
def play_game():
board, winner, counter = create_board(), 0, 1
print(board)
sleep(2)
while winner == 0:
for player in [1, 2]:
board = random_place(board, player)
print("Board after " + str(counter) + " move")
print(board)
sleep(2)
counter += 1
winner = evaluate(board)
if winner != 0:
break
return(winner)
# Driver Code
print("Winner is: " + str(play_game()))
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NjHnSAbHrInP"
# # 量子フーリエ変換
#
# 古典的なフーリエ変換は波や信号の解析において重要なツールであり、関数をそれぞれ異なる周波数を持つ成分に分解します。
#
# この離散的な対応である離散フーリエ変換は、$n$ 個の複素数 $x_0,\ldots,x_{N-1}$ に作用し、次式のように、別の $n$ 個の複素数列 $\tilde x_0,\ldots,\tilde x_{N-1}$ へと変換します。
#
# $$\tilde x_k = \sum_{y=0}^{N-1}e^{-\frac{2\pi ikn}N} \cdot x_k$$
#
# $n$ 個の量子ビットに対する量子フーリエ変換(一般にQFTと略します)は各基底状態 $x\in \{0,1\}^n$ に対して同様にはたらき、次式のように表されます(ただし、$N=2^n$)。
#
# $$\text{QFT}(\lvert x\rangle) = \frac 1{\sqrt N}\sum_{y=0}^{N-1}e^{\frac{2\pi ixy}N}\lvert y\rangle\qquad \qquad (1)$$
#
# ここでは、ビット列 $x\in\{0, 1\}^n$ を整数 $\sum_{j=0}^{n-1}2^{n-1-j}x_j$ として表記するものとします。
#
# QFTは事実上、計算基底から "フーリエ基底" に基底を変換したものと考えることができます。
# + [markdown] colab_type="text" id="CJhIv6UdASWQ"
# ## QFTのテンソル積分解
#
# いくつかの代数的な操作によって、式$(1)$の総和を純粋状態同士のテンソル積へと分解できることがわかります。
#
# $$\text{QFT}(\lvert x\rangle) = \frac 1{\sqrt N}\left(\vert 0\rangle + e^{\frac{2\pi ix}{2^1}}\vert 1\rangle\right)\otimes \left(\vert 0\rangle + e^{\frac{2\pi ix}{2^2}}\vert 1\rangle\right)\otimes \cdots \otimes \left(\vert 0\rangle + e^{\frac{2\pi ix}{2^n}}\vert 1\rangle\right).$$
#
# この定式化により、QFTを実現する量子回路をどのように実装すればよいかが明確になります。
#
# ## 制御位相回転
#
# 回路構成を見る前に、次のゲートを定義します。
#
# $$R_k = \begin{pmatrix}1 & 0\\ 0 & e^{\frac{2\pi i}{2^k}}\end{pmatrix},$$
#
# そしてこのゲートを制御ゲート化したものが、制御量子ビットの値に応じてターゲット量子ビットの位相を変えることを思い出しましょう。
# 言い換えると、$CR_k(\lvert 0\rangle|x\rangle) = |0\rangle \lvert x\rangle$ かつ
#
# \begin{aligned}CR_k(\lvert 1\rangle \lvert x\rangle) = \lvert 1\rangle\otimes R_k(\lvert x \rangle) &=\lvert 1\rangle\otimes R_k\left( \langle 0\lvert x\rangle \lvert 0\rangle + \langle 1 \lvert x\rangle \lvert 1\rangle\right)
# \\ & = \lvert 1\rangle\otimes \left( \langle 0 \lvert x\rangle \lvert 0\rangle + \langle 1 \lvert x\rangle e^{\frac{2\pi i}{2^k}} \lvert 1\rangle\right)\end{aligned}
#
# となります。
#
# より簡潔に、次のように表されます。
#
# $$CR_k(\lvert y\rangle \lvert x\rangle) = \lvert y\rangle \otimes \left( \langle 0 \vert x\rangle \lvert 0\rangle + \langle 1 \vert x\rangle e^{\frac{2\pi iy}{2^k}} \lvert 1\rangle\right).$$
#
# よって、
# $$CR_k = \begin{pmatrix}1 & 0 & 0 & 0\\ 0 & 1 & 0 & 0\\ 0& 0& 1 & 0\\ 0 & 0&0 &e^{\frac{2\pi i}{2^k}}\end{pmatrix} = \text{CPhase}\left(\frac{2\pi}{2^k}\right).$$
# + [markdown] colab_type="text" id="oXg3hf3yAkMS"
# ## 回路構造
#
# これで、QFTを実現する回路を構築するために必要な材料が揃いました。
# 以下は、$n=4$量子ビットの回路例です。
#
# 
#
# ここでは、$\lvert \varphi_1\rangle$から$\lvert \varphi_4\rangle$までの状態を見ていきます。
# + [markdown] colab_type="text" id="U1uxGVsCCi85"
# $\lvert \varphi_1\rangle$ において、先頭の量子ビットの状態を、基底状態 $x_0\in\{0,1\}$ 上で解析します。すると、$\lvert x_0\rangle$ は次のようになります(ここでは計算を簡単にするために、正規化は無視します)。
# # + $H$: $\lvert 0\rangle + e^{\pi i x_0}\lvert 1\rangle$,
# # + $R_2$: $\lvert 0\rangle + e^{\pi i x_0 + \frac{2\pi i x_1}{2^2}}\lvert 1\rangle$
# # + $R_3$: $\lvert 0\rangle + e^{\pi i x_0 + \frac{2\pi i x_1}{2^2} + \frac{2\pi i x_2}{2^3}}\lvert 1\rangle$
# # + $R_4$: $\lvert 0\rangle + e^{\pi i x_0 + \frac{2\pi i x_1}{2^2} + \frac{2\pi i x_2}{2^3} + \frac{2\pi i x_3}{2^4}}\lvert 1\rangle = \lvert 0\rangle + e^{\frac{2\pi i}{2^4} (8x_0 + 4x_1 + 2x_2 + x_3)}\lvert 1\rangle = |0\rangle + e^{\frac{2\pi i x}{2^4}}\lvert 1\rangle$.
#
# よって、$\lvert \varphi_1\rangle = \left(\lvert 0\rangle + e^{\frac{2\pi i x}{2^4}}\lvert 1\rangle\right)\otimes \lvert x_1x_2x_3\rangle$ となります。
#
# 帰納的に、$\lvert \varphi_2\rangle$ が量子ビット $\lvert x_1\rangle$ の状態を次のように変えることが簡単に確かめられます。
# # + $\lvert 0\rangle + e^{\frac{2\pi i}{2^3}(4x_1 + 2x_2 + x_3)}\lvert 1\rangle$.
#
# ただし、$e^{2\pi ix_0} = 1$ なので、$e^{\frac{2\pi i}{2^3}(4x_1 + 2x_2 + x_3)}=e^{\frac{2\pi i}{2^3}(8x_0 + 4x_1 + 2x_2 + x_3)}= e^{\frac{2\pi ix}{2^3}}$ とできます。
#
# よって $\lvert \varphi_2\rangle = \left(\lvert 0\rangle + e^{\frac{2\pi i x}{2^4}}\lvert 1\rangle\right)\otimes \left(\lvert 0\rangle + e^{\frac{2\pi i x}{2^3}}\lvert 1\rangle\right)\otimes \lvert x_2x_3\rangle$ となります。
#
# このように続けると、 $\lvert \varphi_4 \rangle$ は私達が求める量子ビットを逆の順序で与えることが確認できます。そのため2つのスワップ操作を行って、QFTの回路を完成させました。
#
# ## Blueqat によるQFTの実装
# -
# !pip install blueqat
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="3eRpgppa_1FI" outputId="d638cc92-9486-43bc-c94a-14a65c6705d7"
# モジュールをインポート
from blueqat import Circuit
import math
# -
# qftを作用させる関数
def apply_qft(circuit: Circuit(), qubits):
num_qubits = len(qubits)
for i in range(num_qubits):
circuit.h[qubits[i]]
for j in range(i+1, num_qubits):
circuit.cphase(math.pi/(2 ** (j-i)))[qubits[j],qubits[i]] # Apply gate CR_{j-i}(qubit j, qubit i)
# 最後に量子ビットの順序を逆にする
for i in range(int(num_qubits/2)):
circuit.swap(qubits[i],qubits[num_qubits-i-1])
# ### 回路の実行
n = 4 # QFTを行う量子ビット数
qc = Circuit()
qc.x[:] # 状態 |1111> を用意
apply_qft(qc, range(n))
qc.run()
# + [markdown] colab_type="text" id="JM9lexiT_034"
# QFTは個々の量子ビットの位相を変化させるだけなので、測定によって物理的に観測することはできません。以下は、$\text{QFT}(|1111\rangle)$から得られた上記の状態ベクトルを視覚的に表したものです。
#
# 
#
# -
# ## QFTの応用
#
# QFT has wide applications in Algorithms design, and is the building block for many influential quantum algorithms such as Quantum Phase Estimation [[1]] , Shor's Algorithm [[2]], and the Hidden Subgroup Problem [[3]].
#
# [1]: https://en.wikipedia.org/wiki/Quantum_phase_estimation_algorithm
# [2]: https://en.wikipedia.org/wiki/Shor's_algorithm
# [3]: https://en.wikipedia.org/wiki/Hidden_subgroup_problem
| tutorial-ja/121_qft_ja.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working With Rotation Poles
#
# ## 1. Finite Rotation Poles
# A simple example to begin is to get the value of a finite rotation between two plates that should be in the rotation file (so we can check that it's right).
#
# Here is the line from the rotation file defining the relative position of Australia wrt Antarctica at 40.1 Ma:
#
# 801 40.100 17.1000 30.6000 -23.6800 802 ! AUS-ANT An18 Muller et.al 1997
#
# Here is the code to get this rotation using pygplates. Note that in the call to the function 'rotation_model.get_rotation', there are four input parameters
# 1. the time to reconstruct to (40.1 Ma)
# 2. the moving plate id (801 for Australia)
# 3. the time to reconstruct from (0 for present day)
# 4. the fixed plate id (802 for Antarctica)
#
# The code should print the Lat,Long,Angle of the finite pole as shown above.
# +
import pygplates
import numpy as np
input_rotation_filename = 'Data/Seton_etal_ESR2012_2012.1.rot'
rotation_model = pygplates.RotationModel(input_rotation_filename)
finite_rotation = rotation_model.get_rotation(40.1,801,0,802)
pole_lat,pole_lon,pole_angle = finite_rotation.get_lat_lon_euler_pole_and_angle_degrees()
print('Finite Pole Lat,Lon,Angle = %f,%f,%f ' % (pole_lat,pole_lon,pole_angle))
# -
# The finite pole can be calculated that describes the relative rotation of any plate relative to any other plate, at any time, so long as the reconstruction tree contains the right information to link the plates and interpolate rotations. For example, suppose we want a finite pole of rotation that describes the position of North America (plateid 101) relative to South America (plateid 201) at 102 Ma.
# +
finite_rotation = rotation_model.get_rotation(102,101,0,201)
pole_lat,pole_lon,pole_angle = finite_rotation.get_lat_lon_euler_pole_and_angle_degrees()
print('Finite Pole Lat,Lon,Angle = %f,%f,%f ' % (pole_lat,pole_lon,pole_angle))
# -
# ## 2. Stage Rotation Poles
#
# Suppose you want to get the stage pole of rotation for a plate pair over a certain time range. To do this, we ultimately use the 'pygplates.get_equivalent_stage_rotation' function. Intermediate steps involve getting rotation models for the begin and end times of the time period, from whichever rotation file we want to work with.
#
# Note that there are several other, similar functions for working with stage and finite rotations. Note also that many of the pygplates functions return more than just the 'Lat,Long,Angle' familiar from rotation files and desktop GPlates. So we have to use additional lines of code if we want to strip out certain values from the result.
# +
fixed_plate = 0
moving_plate = 802
to_time = 50
from_time = 55
stage_rotation = rotation_model.get_rotation(to_time,moving_plate,from_time,fixed_plate)
print(stage_rotation)
pole_lat,pole_lon,pole_angle = stage_rotation.get_lat_lon_euler_pole_and_angle_degrees()
print('Stage Pole Lat,Lon,Angle = %f,%f,%f ' % (pole_lat,pole_lon,pole_angle))
# -
# ### Stage Pole Sequences
# Suppose you wanted to do this for a series of successive time intervals. A loop to calculate stage poles for stages 10 Myr in duration, from 100 Ma to present, would look something like this:
# +
import numpy as np
time_step = 10
for time in np.arange(0,100,time_step):
to_time = time
from_time = time+time_step
stage_rotation = rotation_model.get_rotation(float(to_time),moving_plate,float(from_time),fixed_plate)
pole_lat,pole_lon,pole_angle = stage_rotation.get_lat_lon_euler_pole_and_angle_degrees()
print('Time interval = ',time,'-',time+time_step,', Stage Pole Lat,Lon,Angle = %f,%f,%f ' % (pole_lat,pole_lon,pole_angle))
# -
# ### A Practical Example - PAC-WANT stage poles in the Cenozoic
# One application of these values is to visualise the stability of stage poles describing the relative motion of two plates through time, and how this stability is influenced by noise in the reconstructions, as discussed by Iaffaldano et al (2012, Nat.Comm). Here we reproduce one of these examples, looking at relative motion between the Pacific and (West) Antarctic plates over the last ~45 Ma:
#
# +
fixed_plate = 901
moving_plate = 804
time_step = 1
Lats = []
Longs = []
Angles = []
for time in np.arange(0,42,time_step):
to_time = time
from_time = time+time_step
stage_rotation = rotation_model.get_rotation(float(to_time),moving_plate,float(from_time),fixed_plate)
pole_lat,pole_lon,pole_angle = stage_rotation.get_lat_lon_euler_pole_and_angle_degrees()
Lats.append(pole_lat)
Longs.append(pole_lon)
Angles.append(np.radians(pole_angle))
# These next lines are necessary becuase the answers come out in the northern hemisphere,
# need to check convention
Longs = np.add(Longs,180.)
Lats = np.multiply(Lats,-1)
# -
# To visualise how the stage pole migrates through time, we can plot the sequence on a map. In this case we do two subplots that show the same stage pole sequence, with colour mapped to time on the left, and magnitude of angular velocity on the right.
# +
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
import numpy as np
# %matplotlib inline
# Make the figure and a dummy orthographic map to get extents
fig = plt.figure(figsize=(12,6),dpi=300)
lat_0=-75. ; lon_0=130.
m1 = Basemap(projection='ortho',lon_0=lon_0,lat_0=lat_0,resolution=None)
# First subplot
ax = fig.add_axes([0.0,0.1,0.5,0.8], facecolor='k')
pmap = Basemap(projection='ortho',lon_0=lon_0,lat_0=lat_0,resolution='l',\
llcrnrx=m1.urcrnrx/-6.,llcrnry=m1.urcrnry/-6.,urcrnrx=m1.urcrnrx/6.,urcrnry=m1.urcrnry/6.)
clip_path = pmap.drawmapboundary(fill_color='white')
pmap.fillcontinents(color='grey', lake_color='white', zorder=0)
pmap.drawmeridians(np.arange(0, 360, 30))
pmap.drawparallels(np.arange(-90, 90, 30))
ax = plt.gca()
x,y = pmap(Longs,Lats)
pmap.plot(x, y, 'r', clip_path=clip_path,zorder=0)
l3=pmap.scatter(x, y, 200, c=np.arange(0,42,time_step),
cmap=plt.cm.jet_r,edgecolor='k',clip_path=clip_path,vmin=0,vmax=45)
cbar = pmap.colorbar(l3,location='right',pad="5%")
cbar.set_label('Time (Ma)',fontsize=12)
# Second subplot
ax = fig.add_axes([0.5,0.1,0.5,0.8],facecolor='k')
pmap = Basemap(projection='ortho',lon_0=lon_0,lat_0=lat_0,resolution='l',\
llcrnrx=m1.urcrnrx/-6.,llcrnry=m1.urcrnry/-6.,urcrnrx=m1.urcrnrx/6.,urcrnry=m1.urcrnry/6.)
clip_path = pmap.drawmapboundary(fill_color='white')
pmap.fillcontinents(color='grey', lake_color='white', zorder=0)
pmap.drawmeridians(np.arange(0, 360, 30))
pmap.drawparallels(np.arange(-90, 90, 30))
ax = plt.gca()
x,y = pmap(Longs,Lats)
pmap.plot(x, y, 'r', clip_path=clip_path,zorder=0)
l3=pmap.scatter(x, y, 200, c=np.degrees(Angles),
cmap=plt.cm.jet_r,edgecolor='k',clip_path=clip_path,vmin=0.5,vmax=1.2)
cbar = pmap.colorbar(l3,location='right',pad="5%")
cbar.set_label('Angular Rate (deg/Myr)',fontsize=12)
plt.show()
# -
# This plot is comparable to figures S2 and S6 of Iaffaldano et al (2012). Note that the stage pole sequence from the Seton et al (2012) rotation file contains PAC-WANT rotations based on Cande et al (1995) at ~10 Myr intervals, so the reconstruction are inevitably less noisy than rotations from a high resolution data set (e.g. the ~1 Myr intervals derived by Croon et al 2008). On the other hand, subtle changes in plate motion over shorter timescales than ~10 Ma investigated by Iaffaldano will not be fully represented in the Seton et al (2012) files.
| notebooks/pygplates-Working-with-Rotation-Poles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reset -f
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import sebaba.ml as sbbml
import sebaba.utils as utils
import matplotlib.pyplot as plt
import matplotlib.ticker as tkr
pd.options.display.float_format = "{:.4f}".format
data = pd.read_csv("data/ex1data.tsv", sep = "\t")
data.head()
# +
fig, ax = plt.subplots(figsize = (10.0, 6.5))
ax = sns.scatterplot(x = "population", y = "profit", data = data, color = "r")
ax.set_ylabel("Profit in $\$$10,000s", fontsize = 18)
ax.set_xlabel("Population in 10,000s", fontsize = 18)
ax.xaxis.set_major_locator(tkr.MaxNLocator(integer = True))
ax.margins(0.05)
ax.axis("tight")
ax.grid(True)
fig.tight_layout()
plt.xlim(0, 30)
plt.show()
# -
x = data[["population"]].values
y = data[["profit"]].values
x_train, x_test, y_train, y_test = utils.split_train_test(x, y, prop_train = 80)
model = sbbml.LinearRegression(alpha = 0.01, normalize = False)
model.fit(x_train, y_train)
model.theta
utils.plot_cost_function(cost = model.cost, width = 10.0, height = 6.5)
y_prime = model.predict(x_test)
utils.root_mean_squared_error(y_prime, y_test)
model = sbbml.LinearRegression(alpha = 0.01, normalize = True)
model.fit(x_train, y_train)
model.theta
utils.plot_cost_function(cost = model.cost, width = 10.0, height = 6.5)
y_prime = model.predict(x_test)
utils.root_mean_squared_error(y_prime, y_test)
x_prime = np.linspace(0, 30, 100).reshape(-1, 1)
y_prime = model.predict(x_prime)
# +
fig, ax = plt.subplots(figsize = (10.0, 6.5))
sns.scatterplot(data = data, x = "population", y = "profit", color = "r")
sns.lineplot(x = x_prime.flatten(), y = y_prime.flatten())
ax.set_ylabel("Profit in $\$$10,000s", fontsize = 18)
ax.set_xlabel("Population in 10,000s", fontsize = 18)
ax.xaxis.set_major_locator(tkr.MaxNLocator(integer = True))
ax.margins(0.05)
ax.axis("tight")
ax.grid(True)
fig.tight_layout()
plt.show()
# -
model = sbbml.LinearRegression(alpha = 0.01, normalize = True)
x_train_poly = utils.map_polynomial_features(x = x_train, degree = 3)
model.fit(x_train_poly, y_train)
utils.plot_cost_function(cost = model.cost, width = 10.0, height = 6.5)
x_test_poly = utils.map_polynomial_features(x = x_test, degree = 3)
y_prime = model.predict(x_test_poly)
utils.root_mean_squared_error(y_prime, y_test)
x_prime = np.linspace(0, 30, 100).reshape(-1, 1)
x_prime_poly = utils.map_polynomial_features(x = x_prime, degree = 3)
y_prime_poly = model.predict(x_prime_poly)
# +
fig, ax = plt.subplots(figsize = (10.0, 6.5))
ax = sns.scatterplot(data = data, x = "population", y = "profit", color = "red")
sns.lineplot(x = x_prime.flatten(), y = y_prime_poly.flatten())
ax.set_ylabel("Profit in $\$$10,000s", fontsize = 18)
ax.set_xlabel("Population in 10,000s", fontsize = 18)
ax.xaxis.set_major_locator(tkr.MaxNLocator(integer = True))
ax.margins(0.05)
ax.axis("tight")
ax.grid(True)
fig.tight_layout()
plt.xlim(0, 30)
plt.show()
# -
data = pd.read_csv("data/ex2data.tsv", sep = "\t"); data.head()
# +
fig, ax = plt.subplots(figsize = (10.0, 6.5))
ax = sns.scatterplot(data = data, x = "size", y = "number_bedrooms", s = 80, hue = "price", palette = "Set1")
ax.set_xlabel("House size", fontsize = 18)
ax.set_ylabel("Number of bedrooms", fontsize = 18)
ax.xaxis.set_major_locator(tkr.MaxNLocator(integer = True))
ax.margins(0.05)
ax.axis("tight")
ax.grid(True)
fig.tight_layout()
plt.show()
# -
x = data[["size", "number_bedrooms"]].values
y = data[["price"]].values
x_train, x_test, y_train, y_test = utils.split_train_test(x, y, prop_train = 80)
model = sbbml.LinearRegression(alpha = 0.01, normalize = True)
model.fit(x_train, y_train)
model.theta
utils.plot_cost_function(cost = model.cost, width = 10.0, height = 6.5)
y_prime = model.predict(x_test)
utils.root_mean_squared_error(y_prime, y_test)
model = sbbml.LinearRegression(alpha = 0.01, normalize = True)
x_train_poly = utils.map_polynomial_features(x = x_train, degree = 3)
model.fit(x_train_poly, y_train)
utils.plot_cost_function(cost = model.cost, width = 10.0, height = 6.5)
x_test_poly = utils.map_polynomial_features(x = x_test, degree = 3)
y_prime = model.predict(x_test_poly)
utils.root_mean_squared_error(y_prime, y_test)
model = sbbml.RidgeRegression(alpha = 0.01, gamma = 1.0, normalize = True)
x_train_poly = utils.map_polynomial_features(x = x_train, degree = 3)
model.fit(x_train_poly, y_train)
utils.plot_cost_function(cost = model.cost, width = 10.0, height = 6.5)
x_test_poly = utils.map_polynomial_features(x = x_test, degree = 3)
y_prime = model.predict(x_test_poly)
utils.root_mean_squared_error(y_prime, y_test)
| linear-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:py3_physeq]
# language: R
# name: conda-env-py3_physeq-r
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Goal" data-toc-modified-id="Goal-1"><span class="toc-item-num">1 </span>Goal</a></span></li><li><span><a href="#Var" data-toc-modified-id="Var-2"><span class="toc-item-num">2 </span>Var</a></span></li><li><span><a href="#Init" data-toc-modified-id="Init-3"><span class="toc-item-num">3 </span>Init</a></span></li><li><span><a href="#LLMGAG" data-toc-modified-id="LLMGAG-4"><span class="toc-item-num">4 </span>LLMGAG</a></span><ul class="toc-item"><li><span><a href="#Setup" data-toc-modified-id="Setup-4.1"><span class="toc-item-num">4.1 </span>Setup</a></span><ul class="toc-item"><li><span><a href="#Run" data-toc-modified-id="Run-4.1.1"><span class="toc-item-num">4.1.1 </span>Run</a></span></li></ul></li></ul></li><li><span><a href="#Summary" data-toc-modified-id="Summary-5"><span class="toc-item-num">5 </span>Summary</a></span><ul class="toc-item"><li><span><a href="#Number-of-genes-assembled-&-clustered" data-toc-modified-id="Number-of-genes-assembled-&-clustered-5.1"><span class="toc-item-num">5.1 </span>Number of genes assembled & clustered</a></span></li><li><span><a href="#Taxonomy" data-toc-modified-id="Taxonomy-5.2"><span class="toc-item-num">5.2 </span>Taxonomy</a></span><ul class="toc-item"><li><span><a href="#Summary" data-toc-modified-id="Summary-5.2.1"><span class="toc-item-num">5.2.1 </span>Summary</a></span></li></ul></li><li><span><a href="#Annotations" data-toc-modified-id="Annotations-5.3"><span class="toc-item-num">5.3 </span>Annotations</a></span><ul class="toc-item"><li><span><a href="#COG-functional-categories" data-toc-modified-id="COG-functional-categories-5.3.1"><span class="toc-item-num">5.3.1 </span>COG functional categories</a></span></li><li><span><a href="#Grouped-by-taxonomy" data-toc-modified-id="Grouped-by-taxonomy-5.3.2"><span class="toc-item-num">5.3.2 </span>Grouped by taxonomy</a></span></li></ul></li><li><span><a href="#humann2-db-genes" data-toc-modified-id="humann2-db-genes-5.4"><span class="toc-item-num">5.4 </span>humann2 db genes</a></span><ul class="toc-item"><li><span><a href="#Summary" data-toc-modified-id="Summary-5.4.1"><span class="toc-item-num">5.4.1 </span>Summary</a></span><ul class="toc-item"><li><span><a href="#By-taxonomy" data-toc-modified-id="By-taxonomy-5.4.1.1"><span class="toc-item-num">5.4.1.1 </span>By taxonomy</a></span></li></ul></li></ul></li></ul></li><li><span><a href="#sessionInfo" data-toc-modified-id="sessionInfo-6"><span class="toc-item-num">6 </span>sessionInfo</a></span></li></ul></div>
# -
# # Goal
#
# * Run `LLMGAG` (metagenome assembly of genes) pipeline on animal gut microbiome metagenome study
# * study = PRJEB9357
# * host = House Cat
# # Var
# +
studyID = 'PRJEB9357'
base_dir = file.path('/ebio/abt3_projects/Georg_animal_feces/data/metagenome/multi-study/BioProjects/',
studyID)
tmp_out_dir = file.path('/ebio/abt3_projects/databases_no-backup/animal_gut_metagenomes/multi-study_MG-asmbl/',
studyID)
work_dir = file.path(base_dir, 'LLMGAG')
pipeline_dir = '/ebio/abt3_projects/methanogen_host_evo/bin/llmgag'
threads = 24
# -
# # Init
# +
library(dplyr)
library(tidyr)
library(ggplot2)
library(data.table)
set.seed(8304)
source('/ebio/abt3_projects/Georg_animal_feces/code/misc_r_functions/init.R')
# -
make_dir(base_dir)
make_dir(tmp_out_dir)
make_dir(work_dir)
# # LLMGAG
# ## Setup
cat_file(file.path(work_dir, 'config.yaml'))
# ### Run
# ```{bash}
# (snakemake_dev) @ rick:/ebio/abt3_projects/methanogen_host_evo/bin/llmgag
# $ screen -L -S llmgag-PRJEB9357 ./snakemake_sge.sh /ebio/abt3_projects/Georg_animal_feces/data/metagenome/multi-study/BioProjects/PRJEB9357/LLMGAG/config.yaml cluster.json /ebio/abt3_projects/Georg_animal_feces/data/metagenome/multi-study/BioProjects/PRJEB9357/LLMGAG/SGE_log 24
# ```
pipelineInfo(pipeline_dir)
# # Summary
# ## Number of genes assembled & clustered
F = file.path(work_dir, 'assembly', 'plass', 'genes.faa')
cmd = glue::glue('grep -c ">" {fasta}', fasta=F)
n_raw_seqs = system(cmd, intern=TRUE)
cat('Number of assembled sequences:', n_raw_seqs, '\n')
F = file.path(work_dir, 'cluster', 'linclust', 'clusters_rep-seqs.faa')
cmd = glue::glue('grep -c ">" {fasta}', fasta=F)
n_rep_seqs = system(cmd, intern=TRUE)
cat('Number of cluster rep sequences:', n_rep_seqs, '\n')
F = file.path(work_dir, 'humann2_db', 'clusters_rep-seqs.faa.gz')
cmd = glue::glue('gunzip -c {fasta} | grep -c ">"', fasta=F)
n_h2_seqs = system(cmd, intern=TRUE)
cat('Number of humann2_db-formatted seqs:', n_h2_seqs, '\n')
# ## Taxonomy
# reading in taxonomy table
## WARING: slow
F = file.path(work_dir, 'taxonomy', 'clusters_rep-seqs_tax_db.tsv.gz')
cmd = glue::glue('gunzip -c {file}', file=F)
coln = c('seqID', 'taxID', 'rank', 'spp', 'lineage')
levs = c('Domain', 'Phylum', 'Class', 'Order', 'Family', 'Genus', 'Species')
tax = fread(cmd, sep='\t', header=FALSE, col.names=coln, fill=TRUE) %>%
separate(lineage, levs, sep=':')
tax %>% dfhead
# number of sequences
tax$seqID %>% unique %>% length %>% print
# which ranks found?
tax$rank %>% table %>% print
# number of classifications per seqID
tax %>%
group_by(seqID) %>%
summarize(n = n()) %>%
ungroup() %>%
.$n %>% summary
# ### Summary
# +
# summarizing taxonomy
tax_s = tax %>%
filter(Domain != '',
Phylum != '') %>%
group_by(Domain, Phylum) %>%
summarize(n = seqID %>% unique %>% length) %>%
ungroup()
tax_s %>% dfhead
# +
# plotting by phylum
p = tax_s %>%
filter(n > 10) %>%
mutate(Phylum = Phylum %>% reorder(n)) %>%
ggplot(aes(Phylum, n, fill=Domain)) +
geom_bar(stat='identity', position='dodge') +
scale_y_log10() +
labs(y = 'No. of genes') +
coord_flip() +
theme_bw() +
theme(
axis.text.y = element_text(size=7)
)
dims(5,7)
plot(p)
# -
# top phyla
tax_s %>%
arrange(-n) %>%
head(n=30)
# +
# summarizing taxonomy
tax_s = tax %>%
filter(Domain != '',
Phylum != '',
Class != '') %>%
group_by(Domain, Phylum, Class) %>%
summarize(n = seqID %>% unique %>% length) %>%
ungroup()
tax_s %>% dfhead
# -
# top hits
tax_s %>%
arrange(-n) %>%
head(n=30)
# ## Annotations
# eggnog-mapper v2
cols = c(
"query_name",
"seed_eggNOG_ortholog",
"seed_ortholog_evalue",
"seed_ortholog_score",
"Predicted_taxonomic_group",
"Predicted_protein_name",
"Gene_Ontology_terms",
"EC_number",
"KEGG_ko",
"KEGG_Pathway",
"KEGG_Module",
"KEGG_Reaction",
"KEGG_rclass",
"BRITE",
"KEGG_TC",
"CAZy",
"BiGG_Reaction",
"tax_scope__eggNOG_taxonomic_level_used_for_annotation",
"eggNOG_OGs",
"bestOG",
"COG_Functional_Category",
"eggNOG_free_text_description"
)
F = file.path(work_dir, 'annotate', 'eggnog-mapper', 'clusters_rep-seqs.emapper.annotations.gz')
cmd = glue::glue('gunzip -c {file}', file=F, header=FALSE)
emap_annot = fread(cmd, sep='\t')
colnames(emap_annot) = cols
emap_annot = emap_annot %>%
dplyr::select(-Gene_Ontology_terms)
emap_annot %>% dfhead
# +
# adding taxonomy info
intersect(emap_annot$query_name, tax$seqID) %>% length %>% print
emap_annot = emap_annot %>%
left_join(tax, c('query_name'='seqID'))
emap_annot %>% dfhead
# -
n_annot_seqs = emap_annot$query_name %>% unique %>% length
cat('Number of rep seqs with eggnog-mapper annotations:', n_annot_seqs, '\n')
# ### COG functional categories
#
# * [wiki on categories](https://ecoliwiki.org/colipedia/index.php/Clusters_of_Orthologous_Groups_%28COGs%29)
# +
# summarizing by functional group
max_cat = emap_annot$COG_Functional_Category %>% unique %>% sapply(nchar) %>% max
emap_annot_s = emap_annot %>%
dplyr::select(query_name, COG_Functional_Category) %>%
separate(COG_Functional_Category, LETTERS[1:max_cat], sep='(?<=[A-Z])') %>%
gather(X, COG_func_cat, -query_name) %>%
filter(!is.na(COG_func_cat),
COG_func_cat != '') %>%
dplyr::select(-X)
emap_annot_s %>% dfhead
# +
# plotting summary
p = emap_annot_s %>%
ggplot(aes(COG_func_cat)) +
geom_bar() +
labs(x='COG functional category', y='No. of genes') +
theme_bw()
dims(9,3)
plot(p)
# +
# plotting summary
p = emap_annot_s %>%
group_by(COG_func_cat) %>%
summarize(perc_abund = n() / n_annot_seqs * 100) %>%
ungroup() %>%
ggplot(aes(COG_func_cat, perc_abund)) +
geom_bar(stat='identity') +
labs(x='COG functional category', y='% of all genes') +
theme_bw()
dims(9,3)
plot(p)
# -
# ### Grouped by taxonomy
# +
max_cat = emap_annot$COG_Functional_Category %>% unique %>% sapply(nchar) %>% max
emap_annot_s = emap_annot %>%
dplyr::select(query_name, COG_Functional_Category) %>%
separate(COG_Functional_Category, LETTERS[1:max_cat], sep='(?<=[A-Z])') %>%
gather(X, COG_func_cat, -query_name) %>%
left_join(tax, c('query_name'='seqID')) %>%
filter(!is.na(COG_func_cat),
COG_func_cat != '') %>%
dplyr::select(-X)
emap_annot_s %>% dfhead
# +
# plotting summary by domain
p = emap_annot_s %>%
ggplot(aes(COG_func_cat)) +
geom_bar() +
facet_wrap(~ Domain, scales='free_y') +
labs(x='COG functional category', y='No. of genes') +
theme_bw()
dims(9,5)
plot(p)
# +
# plotting summary by phylum
p = emap_annot_s %>%
group_by(Phylum) %>%
mutate(n = n()) %>%
ungroup() %>%
filter(n >= 1000) %>%
mutate(Phylum = Phylum %>% reorder(-n)) %>%
ggplot(aes(COG_func_cat, fill=Domain)) +
geom_bar() +
facet_wrap(~ Phylum, scales='free_y', ncol=3) +
labs(x='COG functional category', y='No. of genes') +
theme_bw()
dims(10,5)
plot(p)
# -
# ## humann2 db genes
# +
# gene IDs
F = file.path(work_dir, 'humann2_db', 'clusters_rep-seqs_annot-index.tsv')
hm2 = fread(F, sep='\t', header=TRUE) %>%
separate(new_name, c('UniRefID', 'Gene_length', 'Taxonomy'), sep='\\|') %>%
separate(Taxonomy, c('Genus', 'Species'), sep='\\.s__') %>%
separate(Species, c('Species', 'TaxID'), sep='__taxID') %>%
mutate(Genus = gsub('^g__', '', Genus))
hm2 %>% dfhead
# +
# adding taxonomy
intersect(hm2$original_name, tax$seqID) %>% length %>% print
hm2 = hm2 %>%
left_join(tax, c('original_name'='seqID'))
hm2 %>% dfhead
# -
# ### Summary
# number of unique UniRef IDs
hm2$UniRefID %>% unique %>% length
# duplicate UniRef IDs
hm2 %>%
group_by(UniRefID) %>%
summarize(n = n()) %>%
ungroup() %>%
filter(n > 1) %>%
arrange(-n) %>%
head(n=30)
# +
# number of genes with a taxID
hm2_f = hm2 %>%
filter(!is.na(TaxID))
hm2_f %>% nrow
# -
# #### By taxonomy
# +
# number of UniRefIDs
hm2_f_s = hm2_f %>%
group_by(Domain, Phylum) %>%
summarize(n = UniRefID %>% unique %>% length) %>%
ungroup()
p = hm2_f_s %>%
filter(n >= 10) %>%
mutate(Phylum = Phylum %>% reorder(n)) %>%
ggplot(aes(Phylum, n, fill=Domain)) +
geom_bar(stat='identity', position='dodge') +
scale_y_log10() +
coord_flip() +
labs(y='No. of UniRef IDs') +
theme_bw() +
theme(
axis.text.y = element_text(size=7)
)
dims(5,5)
plot(p)
# -
# # sessionInfo
sessionInfo()
| multi-study/06n_LLMGAG_PRJEB9357.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# _Lambda School Data Science — Tree Ensembles_
#
# # Model Interpretation
#
# ### Objectives
# - Partial Dependence Plots
# - Shapley Values
#
#
# ### Pre-reads
# 1. Kaggle / <NAME>: Machine Learning Explainability
# - https://www.kaggle.com/dansbecker/partial-plots
# - https://www.kaggle.com/dansbecker/shap-values
# 2. <NAME>: Interpretable Machine Learning
# - https://christophm.github.io/interpretable-ml-book/pdp.html
# - https://christophm.github.io/interpretable-ml-book/shapley.html
#
#
# ### Librarires
# - [PDPbox](https://github.com/SauceCat/PDPbox): `pip install pdpbox`
# - [shap](https://github.com/slundberg/shap): `conda install -c conda-forge shap` / `pip install shap`
#
#
# ### Types of explanations
#
# #### Global explanation: all features in relation to each other
# - Feature Importances (mean decrease impurity)
# - Permutation Importances
# - Drop-Column Importances
#
# #### Global explanation: individual feature in relation to target
# - Partial Dependence plots
#
# #### Individual prediction explanation
# - Shapley Values
#
# _Note that the coefficients from a linear model give you all three types of explanations!_
# # Titanic
# +
# %matplotlib inline
import pandas as pd
import seaborn as sns
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
def load_titanic():
df = sns.load_dataset('titanic')
df['age'] = df['age'].fillna(df['age'].mean())
df['class'] = df['class'].map({'First': 1, 'Second': 2, 'Third': 3})
df['female'] = df['sex'] == 'female'
X = df[['age', 'class', 'fare', 'female']]
y = df['survived']
return X, y
X, y = load_titanic()
# -
# #### Naive majority calss baseline
y.value_counts(normalize=True)
# #### Logistic Regression
lr = LogisticRegression(solver='lbfgs')
cross_val_score(lr, X, y, scoring='accuracy', cv=5, n_jobs=-1)
lr.fit(X, y)
pd.Series(lr.coef_[0], X.columns)
sns.regplot(x=X['age'], y=y, logistic=True, y_jitter=.05);
# #### Gradient Boosting
gb = GradientBoostingClassifier()
cross_val_score(gb, X, y, scoring='accuracy', cv=5, n_jobs=-1)
gb.fit(X, y)
pd.Series(gb.feature_importances_, X.columns)
# +
from pdpbox.pdp import pdp_isolate, pdp_plot
feature='age'
pdp_isolated = pdp_isolate(model=gb, dataset=X, model_features=X.columns, feature=feature)
pdp_plot(pdp_isolated, feature);
# -
# From [PDPbox documentation](https://pdpbox.readthedocs.io/en/latest/):
#
#
# >**The common headache**: When using black box machine learning algorithms like random forest and boosting, it is hard to understand the relations between predictors and model outcome. For example, in terms of random forest, all we get is the feature importance. Although we can know which feature is significantly influencing the outcome based on the importance calculation, it really sucks that we don’t know in which direction it is influencing. And in most of the real cases, the effect is non-monotonic. We need some powerful tools to help understanding the complex relations between predictors and model prediction.
# [Animation by <NAME>](https://twitter.com/ChristophMolnar/status/1066398522608635904), author of [_Interpretable Machine Learning_](https://christophm.github.io/interpretable-ml-book/)
#
# > Partial dependence plots show how a feature affects predictions of a Machine Learning model on average.
# > 1. Define grid along feature
# > 2. Model predictions at grid points
# > 3. Line per data instance -> ICE (Individual Conditional Expectation) curve
# > 4. Average curves to get a PDP (Partial Dependence Plot)
# #### Compare Predictions
# +
from sklearn.model_selection import cross_val_predict
y_pred_lr = cross_val_predict(lr, X, y, cv=5, n_jobs=-1)
y_pred_gb = cross_val_predict(gb, X, y, cv=5, n_jobs=-1)
preds = pd.DataFrame({'true': y, 'lr': y_pred_lr, 'gb': y_pred_gb})
gb_right = preds['gb'] == preds['true']
lr_wrong = preds['lr'] != preds['true']
len(preds[gb_right & lr_wrong]) / len(preds)
# -
preds[gb_right & lr_wrong].head()
# +
data_for_prediction = X.loc[27]
data_for_prediction
# -
# #### Explain individual prediction
#
# https://www.kaggle.com/dansbecker/shap-values
# +
import shap
# Create object that can calculate shap values
explainer = shap.TreeExplainer(gb)
# Calculate Shap values
shap_values = explainer.shap_values(data_for_prediction)
shap.initjs()
shap.force_plot(explainer.expected_value, shap_values, data_for_prediction)
# -
# # Lending Club
# +
import category_encoders as ce
import pandas as pd
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import cross_val_score
from sklearn.pipeline import make_pipeline
# Load data from https://www.kaggle.com/c/ds1-tree-ensembles/data
X_train = pd.read_csv('train_features.csv')
X_test = pd.read_csv('test_features.csv')
y_train = pd.read_csv('train_labels.csv')['charged_off']
sample_submission = pd.read_csv('sample_submission.csv')
def wrangle(X):
X = X.copy()
# Drop some columns
X = X.drop(columns='id') # id is random
X = X.drop(columns=['member_id', 'url', 'desc']) # All null
X = X.drop(columns='title') # Duplicative of purpose
X = X.drop(columns='grade') # Duplicative of sub_grade
# Transform sub_grade from "A1" - "G5" to 1.1 - 7.5
def wrangle_sub_grade(x):
first_digit = ord(x[0]) - 64
second_digit = int(x[1])
return first_digit + second_digit/10
X['sub_grade'] = X['sub_grade'].apply(wrangle_sub_grade)
# Convert percentages from strings to floats
X['int_rate'] = X['int_rate'].str.strip('%').astype(float)
X['revol_util'] = X['revol_util'].str.strip('%').astype(float)
# Transform earliest_cr_line to an integer: how many days it's been open
X['earliest_cr_line'] = pd.to_datetime(X['earliest_cr_line'], infer_datetime_format=True)
X['earliest_cr_line'] = pd.Timestamp.today() - X['earliest_cr_line']
X['earliest_cr_line'] = X['earliest_cr_line'].dt.days
# Create features for three employee titles: teacher, manager, owner
X['emp_title'] = X['emp_title'].str.lower()
X['emp_title_teacher'] = X['emp_title'].str.contains('teacher', na=False)
X['emp_title_manager'] = X['emp_title'].str.contains('manager', na=False)
X['emp_title_owner'] = X['emp_title'].str.contains('owner', na=False)
# Drop categoricals with high cardinality
X = X.drop(columns=['emp_title', 'zip_code'])
# Transform features with many nulls to binary flags
many_nulls = ['sec_app_mths_since_last_major_derog',
'sec_app_revol_util',
'sec_app_earliest_cr_line',
'sec_app_mort_acc',
'dti_joint',
'sec_app_collections_12_mths_ex_med',
'sec_app_chargeoff_within_12_mths',
'sec_app_num_rev_accts',
'sec_app_open_act_il',
'sec_app_open_acc',
'revol_bal_joint',
'annual_inc_joint',
'sec_app_inq_last_6mths',
'mths_since_last_record',
'mths_since_recent_bc_dlq',
'mths_since_last_major_derog',
'mths_since_recent_revol_delinq',
'mths_since_last_delinq',
'il_util',
'emp_length',
'mths_since_recent_inq',
'mo_sin_old_il_acct',
'mths_since_rcnt_il',
'num_tl_120dpd_2m',
'bc_util',
'percent_bc_gt_75',
'bc_open_to_buy',
'mths_since_recent_bc']
for col in many_nulls:
X[col] = X[col].isnull()
# For features with few nulls, do mean imputation
for col in X:
if X[col].isnull().sum() > 0:
X[col] = X[col].fillna(X[col].mean())
# Return the wrangled dataframe
return X
# Wrangle train and test in the same way
X_train = wrangle(X_train)
X_test = wrangle(X_test)
# +
# %%time
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=0.2, stratify=y_train, random_state=42)
encoder = ce.OrdinalEncoder()
X_train = encoder.fit_transform(X_train)
X_val = encoder.transform(X_val)
gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_pred_proba = gb.predict_proba(X_val)[:,1]
print('Validation ROC AUC:', roc_auc_score(y_val, y_pred_proba))
# -
# ### Partial Dependence Plot
# +
from pdpbox.pdp import pdp_isolate, pdp_plot
feature='int_rate'
pdp_isolated = pdp_isolate(model=gb, dataset=X_val, model_features=X_val.columns, feature=feature)
pdp_plot(pdp_isolated, feature);
# -
# ### Individual predictions
import numpy as np
y_pred = (y_pred_proba >= 0.5).astype(int)
confidence = np.abs(y_pred_proba - 0.5)
preds = pd.DataFrame({'y_val': y_val, 'y_pred': y_pred, 'y_pred_proba': y_pred_proba, 'confidence': confidence})
# True positives, with high confidence
preds[(y_val==1) & (y_pred==1)].sort_values(by='confidence', ascending=False).head()
data_for_prediction = X_val.loc[17575]
explainer = shap.TreeExplainer(gb)
shap_values = explainer.shap_values(data_for_prediction)
shap.force_plot(explainer.expected_value, shap_values, data_for_prediction)
# True negatives, with high confidence
preds[(y_val==0) & (y_pred==0)].sort_values(by='confidence', ascending=False).head()
data_for_prediction = X_val.loc[1778]
shap_values = explainer.shap_values(data_for_prediction)
shap.force_plot(explainer.expected_value, shap_values, data_for_prediction)
# False positives, with high (mistaken) confidence
preds[(y_val==0) & (y_pred==1)].sort_values(by='confidence', ascending=False).head()
data_for_prediction = X_val.loc[33542]
shap_values = explainer.shap_values(data_for_prediction)
shap.force_plot(explainer.expected_value, shap_values, data_for_prediction)
# False negatives, with high (mistaken) confidence
preds[(y_val==1) & (y_pred==0)].sort_values(by='confidence', ascending=False).head()
data_for_prediction = X_val.loc[30492]
shap_values = explainer.shap_values(data_for_prediction)
shap.force_plot(explainer.expected_value, shap_values, data_for_prediction)
# Most uncertain predictions (least confidence)
preds.sort_values(by='confidence', ascending=True).head()
data_for_prediction = X_val.loc[22527]
shap_values = explainer.shap_values(data_for_prediction)
shap.force_plot(explainer.expected_value, shap_values, data_for_prediction)
| module3-model-interpretation/LS_DS_413_Model_Interpretation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Salary prediction, episode II: make it actually work (4 points)
#
# Your main task is to use some of the tricks you've learned on the network and analyze if you can improve __validation MAE__. Try __at least 3 options__ from the list below for a passing grade. Write a short report about what you have tried. More ideas = more bonus points.
#
# __Please be serious:__ " plot learning curves in MAE/epoch, compare models based on optimal performance, test one change at a time. You know the drill :)
#
# You can use either pure __tensorflow__ or __keras__. Feel free to adapt the seminar code for your needs.
#
# +
# < A whole lot of your code > - models, charts, analysis
# -
# ### A short report
#
# Please tell us what you did and how did it work.
#
# `<YOUR_TEXT_HERE>`, i guess...
# ## Recommended options
#
# #### A) CNN architecture
#
# All the tricks you know about dense and convolutional neural networks apply here as well.
# * Dropout. Nuff said.
# * Batch Norm. This time it's `L.BatchNormalization`
# * Parallel convolution layers. The idea is that you apply several nn.Conv1d to the same embeddings and concatenate output channels.
# * More layers, more neurons, ya know...
#
#
# #### B) Play with pooling
#
# There's more than one way to perform pooling:
# * Max over time - our `L.GlobalMaxPool1D`
# * Average over time (excluding PAD)
# * Softmax-pooling:
# $$ out_{i, t} = \sum_t {h_{i,t} \cdot {{e ^ {h_{i, t}}} \over \sum_\tau e ^ {h_{j, \tau}} } }$$
#
# * Attentive pooling
# $$ out_{i, t} = \sum_t {h_{i,t} \cdot Attn(h_t)}$$
#
# , where $$ Attn(h_t) = {{e ^ {NN_{attn}(h_t)}} \over \sum_\tau e ^ {NN_{attn}(h_\tau)}} $$
# and $NN_{attn}$ is a dense layer.
#
# The optimal score is usually achieved by concatenating several different poolings, including several attentive pooling with different $NN_{attn}$ (aka multi-headed attention).
#
# The catch is that keras layers do not inlude those toys. You will have to [write your own keras layer](https://keras.io/layers/writing-your-own-keras-layers/). Or use pure tensorflow, it might even be easier :)
#
# #### C) Fun with words
#
# It's not always a good idea to train embeddings from scratch. Here's a few tricks:
#
# * Use a pre-trained embeddings from `gensim.downloader.load`. See last lecture.
# * Start with pre-trained embeddings, then fine-tune them with gradient descent. You may or may not want to use __`.get_keras_embedding()`__ method for word2vec
# * Use the same embedding matrix in title and desc vectorizer
#
#
# #### D) Going recurrent
#
# We've already learned that recurrent networks can do cool stuff in sequence modelling. Turns out, they're not useless for classification as well. With some tricks of course..
#
# * Like convolutional layers, LSTM should be pooled into a fixed-size vector with some of the poolings.
# * Since you know all the text in advance, use bidirectional RNN
# * Run one LSTM from left to right
# * Run another in parallel from right to left
# * Concatenate their output sequences along unit axis (dim=-1)
#
# * It might be good idea to mix convolutions and recurrent layers differently for title and description
#
#
# #### E) Optimizing seriously
#
# * You don't necessarily need 100 epochs. Use early stopping. If you've never done this before, take a look at [early stopping callback](https://keras.io/callbacks/#earlystopping).
# * In short, train until you notice that validation
# * Maintain the best-on-validation snapshot via `model.save(file_name)`
# * Plotting learning curves is usually a good idea
#
# Good luck! And may the force be with you!
# +
import gensim.downloader
embeddings = gensim.downloader.load("fasttext-wiki-news-subwords-300")
# If you're low on RAM or download speed, use "glove-wiki-gigaword-100" instead. Ignore all further asserts.
# -
| week02_classification/homework_part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" src="media/Assets&ArchHeader.jpg">
# # Managing the Db2 Data Management Console
#
# This Jupyter Notebook contains examples of how to setup and manage the Db2 Data Management Console. It covers how to add additional users using database authentication, how to explore and manage connections and setup and manage monitoring profiles.
#
# The Db2 Data Management Console is more than a graphical user interface. It is a set of microservices that you can use to script your use of Db2.
#
# <img align="left" src="media/DMC.png">
# ### Launching the Db2 Console
# You can launch the Db2 Console using the browser running in the Virtual desktop or from your own desktop.
#
# Inside the virtual machine desktop you can follow the link: http://localhost:11080.
#
# To use the Db2 Console from your own browser, look for the link in your welcome note. It will start with **services-uscentral.skytap.com**
#
# You can log into the console using:
# * userid: db2inst1
# * password: <PASSWORD>
# ### Db2 Console Open API
# This Jupyter Notebook contains examples of how to use the Open APIs and the composable interface that are available in the Db2 Data Management Console. Everything in the User Interface is also available through an open and fully documented RESTful Services API. The full set of APIs are documented as part of the Db2 Data Management Console user interface. In this hands on lab you can connect to the documentation directly through this link: [Db2 Data Management Console RESTful APIs](http://localhost:11080/dbapi/api/index_enterprise.html).
# ### Where to find this sample online
# You can find a copy of this notebook at https://github.com/Db2-DTE-POC/db2dmc.
# ### First we will import a few helper classes
# We need to pull in a few standard Python libraries so that we can work with REST, JSON and a library called Pandas. Pandas lets us work with DataFrames, which are a very powerful way to work with tabular data in Python.
# Import the class libraries
import requests
import ssl
import json
from pprint import pprint
from requests import Response
import pandas as pd
import time
from requests.packages.urllib3.exceptions import InsecureRequestWarning
requests.packages.urllib3.disable_warnings(InsecureRequestWarning)
from IPython.display import IFrame
from IPython.display import display, HTML
from pandas import json_normalize
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
# ### The Db2 Class
# Next we will create a Db2 helper class that will encapsulate the Rest API calls that we can use to directly access the Db2 Data Management Console service without having to use the user interface.
#
# To access the service we need to first authenticate with the service and create a reusable token that we can use for each call to the service. This ensures that we don't have to provide a userID and password each time we run a command. The token makes sure this is secure.
#
# Each request is constructed of several parts. First, the URL and the API identify how to connect to the service. Second the REST service request that identifies the request and the options. For example '/metrics/applications/connections/current/list'. And finally some complex requests also include a JSON payload. For example running SQL includes a JSON object that identifies the script, statement delimiters, the maximum number of rows in the results set as well as what do if a statement fails.
#
# The full set of APIs are documents as part of the Db2 Data Management Console user interface. In this hands on lab you can connect to that directly through this link: [Db2 Data Management Console RESTful APIs](http://localhost:11080/dbapi/api/index_enterprise.html).
# Run the Db2 Class library
# Used to construct and reuse an Autentication Key
# Used to construct RESTAPI URLs and JSON payloads
class Db2Console():
def __init__(self, url, verify = False, proxies=None, ):
self.url = url
self.proxies = proxies
self.verify = verify
def authenticate(self, userid, password, profile=""):
credentials = {'userid':userid, 'password':password}
r = requests.post(self.url+'/auth/tokens', verify=self.verify, json=credentials, proxies=self.proxies)
if (r.status_code == 200):
bearerToken = r.json()['token']
if profile == "":
self.headers = {'Authorization': 'Bearer'+ ' '+bearerToken}
return True;
else:
self.headers = {'Authorization': 'Bearer'+ ' '+bearerToken, 'X-DB-Profile': profile}
return True;
else:
print ('Unable to authenticate, no bearer token obtained')
return False;
def printResponse(self, r, code):
if (r.status_code == code):
pprint(r.json())
else:
print (r.status_code)
print (r.content)
def getRequest(self, api, json=None):
return requests.get(self.url+api, verify = self.verify, headers=self.headers, proxies = self.proxies, json=json)
def postRequest(self, api, json=None):
return requests.post(self.url+api, verify = self.verify, headers=self.headers, proxies = self.proxies, json=json)
def deleteRequest(self, api, json=None):
return requests.delete(self.url+api, verify = self.verify, headers=self.headers, proxies = self.proxies, json=json)
def putRequest(self, api, json=None):
return requests.put(self.url+api, verify = self.verify, headers=self.headers, proxies = self.proxies, json=json)
def getStatusCode(self, response):
return (response.status_code)
def getJSON(self, response):
return (response.json())
def runSQL(self, script, limit=10, separator=';', stopOnError=False):
sqlJob = {'commands': script, 'limit':limit, 'separator':separator, 'stop_on_error':str(stopOnError)}
return self.postRequest('/sql_jobs',sqlJob)
def getSQLJobResult(self, jobid):
return self.getRequest('/sql_jobs/'+jobid)
def getUserPriviledges(self, profile=''):
if profile == '' :
return self.getRequest('/userProfilePrivileges')
else :
return self.getRequest('/userProfilePrivileges/'+profile)
def assignUserPrivileges(self, profile, user):
json = [{'profileName': profile, 'USER':[user], 'OWNER':[]}]
return self.postRequest('/userProfilePrivileges?action=assign', json)
def assignOwnerPrivileges(self, profile, owner):
json = [{'profileName': profile, 'USER':[], 'OWNER':[owner]}]
return self.postRequest('/userProfilePrivileges?action=assign', json)
def revokeProfilePrivileges(self, profile, user):
json = [{'profileName': profile, 'USER':[user]}]
return self.postRequest('/userProfilePrivileges?action=revoke', json)
def getConnectionProfile(self,profile):
return self.getRequest('/dbprofiles/'+profile)
def getMonitorStatus(self):
return self.getRequest('/monitor')
def getConnectionProfiles(self):
return self.getRequest('/dbprofiles')
def getConsoleRepository(self):
return self.getRequest('/repository')
def getExportConnectionProfiles(self):
return self.getRequest('/dbprofiles/transfer/export?exportCred=true')
def postConnectionProfile(self, connectionName, dbName, port, host, userid, password, comment):
json = {"name":connectionName,"location":"","databaseName":dbName,"dataServerType":"DB2LUW","port":port,"host":host,"URL":"jdbc:db2://"+host+":"+port+"/"+dbName+":retrieveMessagesFromServerOnGetMessage=true;","sslConnection":"false","disableDataCollection":"false","collectionCred":{"securityMechanism":"3","user":userid,"password":password},"operationCred":{"securityMechanism":"3","user":userid,"password":password,"saveOperationCred":"true"},"comment":comment}
return self.postRequest('/dbprofiles', json)
def putConnectionProfileUpdate(self, connectionName, dbName, port, host, userid, password, comment):
json = {"name":connectionName,"location":"","databaseName":dbName,"dataServerType":"DB2LUW","port":port,"host":host,"URL":"jdbc:db2://"+host+":"+port+"/"+dbName+":retrieveMessagesFromServerOnGetMessage=true;","sslConnection":"false","disableDataCollection":"false","collectionCred":{"securityMechanism":"3","user":userid,"password":password},"operationCred":{"securityMechanism":"3","user":userid,"password":password,"saveOperationCred":"true"},"comment":comment}
return self.putRequest('/dbprofiles/'+connectionName, json)
def postTestConnection(self, dbName, port, host, userid, password):
json = {"name":"","location":"","databaseName":dbName,"dataServerType":"DB2LUW","port":port,"host":host,"URL":"jdbc:db2://"+host+":"+port+"/"+dbName+":retrieveMessagesFromServerOnGetMessage=true;","sslConnection":"false","disableDataCollection":"false","operationCred":{"securityMechanism":"3","user":userid,"password":password}}
return self.postRequest('/dbprofiles/testConnection', json)
def deleteConnectionProfile(self, connectionName):
return self.deleteRequest('/dbprofiles/'+connectionName)
def getMonitoringProfiles(self):
return self.getRequest('/monitorprofile/front')
def getMonitoringProfile(self, profileID):
return self.getRequest('/monitorprofile/front/'+profileID)
def putMonitoringProfile(self, profileID, json):
return self.putRequest('/monitorprofile/front/'+profileID, json)
def getProfileIndex(self, profileName):
r = self.getMonitoringProfiles()
if (self.getStatusCode(r)==200):
json = self.getJSON(r)
profileList = pd.DataFrame(json_normalize(json['resources']))[['name','id']]
display(profileList)
profileList.set_index('name',inplace=True)
try:
profileIndex = profileList.loc[profileName][0]
except KeyError:
profileIndex = 0
print(profileName + " not found")
return profileIndex
else:
print(self.getStatusCode(r))
# ### Db2 Data Management Console Connection
# To connect to the Db2 Data Management Console service you need to provide the URL, the service name (v4) and profile the console user name and password as well as the name of the connection profile used in the console to connect to the database you want to work with. For this lab we are assuming that the following values are used for the connection:
# * Userid: db2inst1
# * Password: <PASSWORD>
# * Connection: sample
#
# **Note:** If the Db2 Data Management Console has not completed initialization, the connection below will fail. Wait for a few moments and then try it again.
# +
# Connect to the Db2 Data Management Console service
Console = 'http://localhost:11080'
user = 'peterconsole'
password = '<PASSWORD>'
# Set up the required connection
databaseAPI = Db2Console(Console+'/dbapi/v4')
if databaseAPI.authenticate(user, password) :
print("Token Created")
else :
print("Token Creation Failed")
database = Console
# -
# ### Confirm the connection
# To confirm that your connection is working you can the Console connection profiles.
# Get Console Connection Profiles
r = databaseAPI.getConnectionProfiles()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)))
else:
print(databaseAPI.getStatusCode(r))
# ### Get Repository Configuration
# You can also get details on the repository configuration. You can see that we are using a local database name HISTORY to store all the monitoring data collected by the console.
# Get Console Repository Configuration
r = databaseAPI.getConsoleRepository()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)[['databaseName','status','host','port','collectionCred.user']]).transpose())
else:
print(databaseAPI.getStatusCode(r))
# ### Running SQL Routines
# We need to run some SQL scripts later in the lab. So run the next two cells. They define routines that run SQL scripts and display the results of those scripts. If you want to learn more about how these routines were developed, check out the [Analyzing SQL Workloads notebook](http://localhost:8888/notebooks/Db2_Data_Management_Console_SQL.ipynb).
def runSQL(profile,user, password, sqlText):
if databaseAPI.authenticate(user, password, profile) :
runID = databaseAPI.getJSON(databaseAPI.runSQL(sqlText))['id']
json = databaseAPI.getJSON(databaseAPI.getSQLJobResult(runID))
while 'results' not in json :
json = databaseAPI.getJSON(databaseAPI.getSQLJobResult(runID))
fulljson = json
while json['results'] != [] or (json['status'] != "completed" and json['status'] != "failed") :
json = databaseAPI.getJSON(databaseAPI.getSQLJobResult(runID))
while 'results' not in json :
json = databaseAPI.getJSON(databaseAPI.getSQLJobResult(runID))
for results in json['results'] :
fulljson['results'].append(results)
time.sleep(1)
return fulljson
else :
print('Could not authenticate')
print('runSQL routine defined')
def displayResults(json):
for results in json['results']:
print('Statement: '+str(results['index'])+': '+results['command'])
if 'error' in results :
print(results['error'])
elif 'rows' in results :
df = pd.DataFrame(results['rows'],columns=results['columns'])
print(df)
else :
print('No errors. Row Affected: '+str(results['rows_affected']))
print()
print('displayResults routine defined')
# ## Accessing the Db2 Console through custom URLs
# This lab provides direct links to specific pages in the Db2 Console to make it easy to navigate during the lab. Since you can use this lab from a browser running in the virtual desktop or from your own browser on your own desktop, we just need to save the location of the console you are using.
#
# By default it is setup to use links that will work from the virtual machine desktop. To run from your own browser on your own desktop, enter the port location of the Db2 Console provided in your welcome note and run the cell below.
externalConsole = "http://services-uscentral.skytap.com:18995"
database = externalConsole
# ### Testing the Custom URL
# Run the cell below.
#
# It will generate a link to the console that works for your selected environment. Notice that it brings up the whole console including the full navigation menu.
#
# Right click on the link and select **Open Link in New Tab** or **Open Link in New Window**
print(database+'/console')
# You can also navigate to a specific page in the console by specifying the full URL or the page. For example, run the following cell and click on the link to navigate to the page that displays the tables in the Sample database:
print(database+'/console/#explore/table?profile=Sample')
# You can also choose to create a URL that just links a page without the full navigation menu. Simply add **?mode=compact** to the URL as an option. For example:
print(database+'/console/?mode=compact#explore/table?profile=Sample')
# ## Setting up Repository Authentication
# ### Reviewing current console authentication and switching to repository authentication
# Let's start by looking at the current authentication settings in the console. To see the authentication settings you can either:
#
# Use the full console to switch to Repository Database Authentication:
# 1. Open the Console
# 2. Click the gear icon at the top right of the page
# 3. Select **Authentication Setting**
#
# Use the embedded page from the console to switch to Repostory Database Authentication.
# 1. Run the cell below.
# 2. Click the link in the cell.
print(database+'/console/?mode=compact#settings/authentication')
# To support multiple users you need a way to authenticate those other users. There are two choices.
# 1. **Repository Database Authentication**. This delegates the authentication of console users to the Db2 repository database used by the console to store historical monitoring data.
# 2. **LDAP**. This delegates the authentication to an external LDAP service.
# In this example we use the Repository Database to authenticate console users. You associate console users and administrators to Db2 Authorities, Groups, UDFs or Role.
#
# In this lab we are going to setup the first option, Repository Database Authentication.
#
# Before switching to Repository Database Authentication lets create some new users in the operating system that Db2 can recognize.
# ### Adding Operating System Users
# The repository database uses standard Db2 authentiation. The simplest way to set this up is to use the userid authentication that is built into the operating system that your Db2 database is running on.
#
# To add new users to the console you need to add new users to your operating system.
#
# In this hands-on lab you can create those new users through the Linux settings console.
#
# 1. Click **Applications** at the upper-left of the screen
# 1. Click **Sundry** in the menu
# 1. Click **Users and Groups** in the menu
# 1. Enter the db2inst1 password **<PASSWORD>**
# 1. Click **Add User** at the upper-left of the page
# 1. Type **peterconsole** into the **Username** field.
# 1. Type **Peter** into the **Full Name** field
# 1. Enter **DataConsole** into the **password fields**
# 1. Click **OK**
#
# Repeat steps 5 to 10 for the following new users:
# 1. **paulconsole** with full name **Paul**
# 1. **maryconsole** with full name **Mary**
# ### Creating Roles for Authentication
# Now that you have three new users we need to create two database roles to classify the access allowd for each id. Either an Admin or User role (CONSOLE_ADM, CONSOLE_USR).
# +
user = 'DB2INST1'
password = '<PASSWORD>'
profile = 'Repository'
script = 'CREATE ROLE CONSOLE_ADM; CREATE ROLE CONSOLE_USR'
print('Adding Roles')
displayResults(runSQL(profile, user, password, script))
print('done')
# -
# ### Adding Users to the Console using Repository Authentication
# Now that you have three new users lets add them grant them database authorities and CONSOLE roles. The following cell will grant a user to either an Admin or User role (CONSOLE_ADM, CONSOLE_USR).
# +
userList = {'newUser':['peterconsole', 'paulconsole', 'maryconsole'], 'Type':['Admin', 'User', 'User']}
userListDF = pd.DataFrame(userList)
display(userListDF)
user = 'db2inst1'
password = '<PASSWORD>'
profile = 'Repository'
for row in range(0, len(userListDF)):
Type = userListDF['Type'].iloc[row]
newUser = userListDF['newUser'].iloc[row]
script = 'GRANT DBADM, CREATETAB, BINDADD, CONNECT, CREATE_NOT_FENCED, IMPLICIT_SCHEMA, LOAD ON DATABASE TO USER '+newUser+';'
if userListDF['Type'].iloc[row] == 'Admin' :
script = script + 'GRANT ROLE "CONSOLE_ADM" TO USER '+newUser+' WITH ADMIN OPTION;'
else :
script = script + 'GRANT ROLE "CONSOLE_USR" TO USER '+newUser+';'
print('Adding User: '+newUser)
displayResults(runSQL(profile, user, password, script))
print('done')
# -
# ### Switch Authentication Method in the Console
# Next that we can users that are assigned to the correct role, we can switch from using the single default user to using Repository Database authentication for multiple users. You can do this through the Db2 Data Management Console and navigating to the **Repository Setting** page or by running the next cell below. Pick one of the two methods below to get to the Repository Settings page. Then follow the 10 steps to change the authentication method.
#
# Use the full console to switch to Repository Database Authentication:
# 1. Click the link http://localhost:11080/console
# 2. Click the gear icon at the top right of the page
# 3. Select **Authentication**
#
# Use the embedded page from the console to switch to Repostory Database Authentication
# 1. Run the cell below
URL = database+'/console/?mode=compact#settings/authentication'
print(URL)
# Now configure the console to use the Repository database to authenticate users.
# 1. Select **Repository** from the list of available Authentiation Types.
# 2. Scroll down and select **Db2 Roles** from the list of User role matting methods
# 3. Enter CONSOLE_ADM in the Admin roles field
# 4. Enter CONSOLE_USR in the User roles field
# 5. Click Next
# 6. Enter **Peter** in the Test user ID field
# 7. Enter **<PASSWORD>** in the Test user password field
# 8. Click **Test**. You should see confirmation that the test succeeded
# 9. Select **Save** at the bottom right of the page. A confirmation dialog appears.
# 10. Click **Yes**. You should now see that Repository authentication is set and enabled successfully
#
# ### Test a New Console User
# To test the new user and authentication method you can log out and into the console and you can run the cell below. After signing in you run SQL for the first time. This prompts a new user to enter a userid and password to authenticate against the Db2 database. You can save this authentication pair if you want to run SQL through the service later.
#
# 1. Open the console http://localhost:11080/console
# 2. Click the user icon at the very top right of the screen
# 3. Select **sign out** a confirmation dialog appears
# 4. Select **Yes**
# 5. Enter **Peter** in the user field
# 6. Enter **DataConsole** in the password field
# 7. Click **History** Connection name in the Databases homepage
# 8. Click the **Run SQL** icon at the left of the page
# 9. Enter **select TABSCHEMA, TABNAME from syscat.tables**
# 9. Click **Run All**
# 9. Enter **Peter** and **DataConsole** in the userid and password field
# 10. IMPORTANT - Click **Save Credentials to repository**
# 11. Click **Test Connection**
# 12. Click **Save**
# This next cell runs a simple query through the console service using one of the new users. This only works if you chose to save Peter's credentials to the repository.
sql = 'select TABSCHEMA, TABNAME from syscat.tables'
displayResults(runSQL('Repository', 'peterconsole', 'DataConsole', sql))
# ### Grant Authority to DBINST1 to Access the Console
# When you added **Peter** to the CONSOLE_ADM role you also included the **WITH ADMIN OPTION** that means that **Peter** can now grant other users access to the **CONSOLE_ADM** group. Run the SQL in the next cell to grant access to DB2INST to the Console as an Administrator.
sql = 'GRANT ROLE "CONSOLE_ADM" TO USER "DB2INST1"'
displayResults(runSQL('Repository', 'peterconsole', 'DataConsole', sql))
# Now lets try running the previous select statement using the **DB2INST1** user id.
sql = 'select TABSCHEMA, TABNAME from syscat.tables'
displayResults(runSQL('Repository', 'db2inst1', 'db2inst1', sql))
# ## Manage Console User Priviledges
# In the console you can control which users can access specific database connections. These are divided into Owners, who can grant privleges, and users who can only use the connection.
# ### Get User Connection Profile Privileges
# The next cell lets you control exactly which users can access specific databases. You can also access this through the console **Users and Privileges** settings page in the main console.
URL = database+'/console/?mode=compact#settings/users'
print(URL)
# If you have a lot to users and databases to manage it may be useful to script these changes. The next cell includes an example of how to retrieve the current privileges through the console APIs.
# Get User Connection Profile Privleges
r = databaseAPI.getUserPriviledges()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)))
else:
print(databaseAPI.getStatusCode(r))
# ### Change User Connection Profile Privileges
# If you have a lot of changes to make at once, the following cells are examples of how to script changes to multiple user and database connection priviledges. The first two cells in this section define reusable routines that let you add and revoke privileges with a single call.
# +
# Change User Connection Profile Privileges
def addProfilePrivileges(profile, name, userType):
if userType == 'user':
r = databaseAPI.assignUserPrivileges(profile, name)
else:
r = databaseAPI.assignOwnerPrivileges(profile, name)
if (databaseAPI.getStatusCode(r)==201):
print(name+' added to: '+profile+" as a new "+userType+".")
else:
print(databaseAPI.getStatusCode(r))
print('Created routine to add profile privileges')
# +
# Revoke User Connection Profile Privileges
def revokeProfilePrivileges(profile, name) :
r = databaseAPI.revokeProfilePrivileges(profile, name)
if (databaseAPI.getStatusCode(r)==201):
print(name+' privilege revoked from: '+profile)
else:
print(databaseAPI.getStatusCode(r))
print('Created routine to revoke profile privileges')
# -
addProfilePrivileges('Repository', 'paulconsole', 'owner')
revokeProfilePrivileges('Repository', 'paulconsole')
# These next two examples, show how to process an entire list of changes through a simple loop. Both to grant and revoke user privileges.
# +
userList = {'userName':['peterconsole', 'paulconsole', 'maryconsole', 'peterconsole', 'paulconsole', 'maryconsole'], 'Profile':['Repository', 'Repository', 'Repository', 'Sample', 'Sample','Sample'],'Type':['owner', 'user', 'user', 'owner', 'user', 'user']}
userListDF = pd.DataFrame(userList)
display(userListDF)
for row in range(0, len(userListDF)):
profile = userListDF['Profile'].iloc[row]
userType = userListDF['Type'].iloc[row]
name = userListDF['userName'].iloc[row]
addProfilePrivileges(profile, name, userType)
print('done')
# +
userList = {'userName':['peterconsole', 'paulconsole', 'maryconsole', 'peterconsole', 'paulconsole', 'maryconsole'], 'Profile':['Repository', 'Repository', 'Repository', 'Sample', 'Sample','Sample']}
userListDF = pd.DataFrame(userList)
display(userListDF)
for row in range(0, len(userListDF)):
profile = userListDF['Profile'].iloc[row]
name = userListDF['userName'].iloc[row]
revokeProfilePrivileges(profile, name)
print('done')
# -
# ## Managing Database Connection Profiles
# To monitor and manage more Db2 databases through the console, you need to add a new connection profile for each database. The connection profile includes information on how to connect to the database (host, port, and database name) as well as log in information for monitoring and management.
# ### Adding Connection Profiles
# Let's start by checking the list of current database profiles available in the console.
#
# Run the next two cells.
#
# The first cell opens the Connection Profile setting page. You can access this through the full console (http://localhost:11080/console) by selecting **Connection Profile** from the settings menu at the top right of the screen.
#
# The second cell lists the connection information by accessing the console API.
print(database+'/console/?mode=compact#connection/manager')
# Get Console Connection Profiles
r = databaseAPI.getConnectionProfiles()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)))
else:
print(databaseAPI.getStatusCode(r))
# ### Export Connection Profile List
# To copy information from one instance of the Db2 Data Management Console to another you can export the current connection profile by selecting the **up arrow** icon at the top right of the Connection Profile page. You can then import the same file back into another console to move connection information from on console to another.
#
# You can also export the connection profile list through the console API. The example cells below create a csv file with the full connection profile information. You can then read back the csv file to see the contents of the csv file.
# +
# Export Console Connection Profiles
r = databaseAPI.getExportConnectionProfiles()
if (databaseAPI.getStatusCode(r)==200):
with open('connectionProfile.csv','wb') as file:
file.write(r.content)
print('Created connectionProfile.csv')
else:
print(databaseAPI.getStatusCode(r))
# -
# Import Console Connection Profile
df = pd.read_csv("connectionProfile.csv")
df = df.drop(df.index[1]) #Drop the second row
df = df.iloc[:, :-1] # Drop the last column
display(df)
# Scroll to the right and find the he **collectionCred** column.
#
# This is an encrypted version of the userid and password used to connect to the database.
# Anytime that the Db2 Data Management Console stores a userid and password, it encrypts it using a routine included with the console. If you export and import a connection profile you can choose to export with our without credentials. If you choose to include the credentials you can use the same userid and passwords from the original connection profile list when you import into a new console.
# ### Create a new Db2 Database
# In this next section you will create a new and add it to the Db2 Data Management Console.
# To create a new Db2 database you need to access the Db2 Terminal:
#
# 1. Click the **Terminal** icon at the bottom left of the screen
# 2. Switch to the DB2INST1 USER by running the following command
# ```
# su - db2inst1
# ```
# 3. Enter the password **<PASSWORD>**
# 4. Create a database with the following command where **dbone** is the name of the new database
# ```
# db2 create database dbone
# ```
# ### Create a new Database Connection Profile through the User Interface
# You can add the database through the Connection Profile page or though the console API. Let's try with the console first. Either open the Connection Profile page through the full console (http://localhost:11080/console) and select **Connection Profile** from the settings menu at the top right of the screen, or run the cell below.
print(database+'/console/?mode=compact#connection/manager/add')
# To create a new connection:
# 1. Enter **databaseOne** in the **Connection name** field
# 2. Enter **localhost** in the **Host** field
# 3. Enter **50001** in the **Port** field
# 4. Enter **dbone** in the **Database** field
# 5. Enter **db2inst1** in the **User name** field
# 6. Enter **<PASSWORD>** in the **Password** field
# 7. Select **Enable operation**
# 8. Enter **db2inst1** in the **User name** field
# 9. Enter **<PASSWORD>** in the **Password** field
# 10. Select **Save credentials to repository**
# 11. Click **Save**
# Now, lets delete the connection you just created so we can recreate it using the console API.
# 1. Click the checkbox beside **databaseone**
# 2. Click **delete**
# 3. Click **Yes** to confirm that you want to delete the databaseone connection. This does not drop the actual database.
# ### List the current Connection Profiles
# Now we can perform the same actions using the console API. Let's start by listing the current database connections. Run the next cell.
# Get Console Connection Profiles
r = databaseAPI.getConnectionProfiles()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)))
else:
print(databaseAPI.getStatusCode(r))
# ### Check Connection Information without Creating a new Profile
# Just like the user interface has a **Test connection** button you can test that your userid and password connects to the database before you create a connection profile. The next cell will check that the following connection information is valid.
dbName = 'dbone'
port = '50001'
host = 'localhost'
userid = 'db2inst1'
password = '<PASSWORD>'
r = databaseAPI.postTestConnection(dbName, port, host, userid, password)
if (databaseAPI.getStatusCode(r)==200):
print(databaseAPI.getJSON(r))
else:
print(databaseAPI.getStatusCode(r))
# ### Create a new Database Connection Profile
# The next cell adds a connection to the **dbone** database.
connectionName = 'databaseOne'
dbName = 'dbone'
port = '50001'
host = 'localhost'
userid = 'db2inst1'
password = '<PASSWORD>'
comment = 'New connection profile test'
r = databaseAPI.postConnectionProfile(connectionName, dbName, port, host, userid, password, comment)
if (databaseAPI.getStatusCode(r)==201):
print("Created connection profile "+connectionName)
else:
print(databaseAPI.getStatusCode(r))
# ### List the current Connection Profiles
# Next run the routine to list the connection profiles to confirm that the connection to dbone is now available.
# Get Console Connection Profiles
r = databaseAPI.getConnectionProfiles()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)))
else:
print(databaseAPI.getStatusCode(r))
# ### Update an Existing Database Connection Profile
# You can also update an exsiting connection profile. The next cell is an example of updating the comment field. You can use the same routine to update the userid and password, host or portnumber as well.
connectionName = 'databaseOne'
dbName = 'dbone'
port = '50001'
host = 'localhost'
userid = 'db2inst1'
password = '<PASSWORD>'
comment = 'Updated Comment'
r = databaseAPI.putConnectionProfileUpdate(connectionName, dbName, port, host, userid, password, comment)
if (databaseAPI.getStatusCode(r)==200):
print("Updated connection profile "+connectionName)
else:
print(databaseAPI.getStatusCode(r))
# Check that the comment field has been updated with the new comment. Run the cell below and check the comment field near the bottom of the list.
# +
# Check Connection Profile
connectionName = 'databaseOne'
r = databaseAPI.getConnectionProfile(connectionName)
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
display(pd.DataFrame(json_normalize(json)).transpose())
else:
print(databaseAPI.getStatusCode(r))
# -
# ### Delete a connection profile
# Finally you can delete existing database connection profiles from the console using a single API call.
# +
connectionName = 'databaseOne'
r = databaseAPI.deleteConnectionProfile(connectionName)
if (databaseAPI.getStatusCode(r)==200):
print("Deleted connection profile "+connectionName)
else:
print(databaseAPI.getStatusCode(r))
# -
# ## Manage Monitoring Profiles
# Each monitored database is associated with a monitoring profile. The profile defines what kind of data is collected, when it is collected and how long it is retained in the repository database. The monitoring profile also configures which alerts are enabled.
#
# Multiple database can be associated with each profile. There is also one default profile that any new database connection profile is added to when it is created.
# ### List Monitor Profiles
# Let's start by listing the current monitoring profiles.
#
# You can access this through the full console (http://localhost:11080/console) by selecting **Monitoring Profile** from the settings menu at the top right of the screen. You can also go directly to the monitoring profiles page by running the next cell.
#
# 1. Click the elipses (...) in the **Actions** column at the right side of the list and click **Edit**
# 2. Browse through the monitoring profile page to see the default settings for this database.
# 3. Click **Cancel** to return to the monitoring profile list.
print(database+'/console/?mode=compact#monitorprofile')
# ### Creating a New Monitoring Profile
# Use the user interface to create a new profile.
# 1. Click **New profile** at the top right of the screen
# 2. Enter **Test Profile** in the **Profile name (New)** field
# 3. Click **Select database list**
# 4. Click the checkbox beside **SAMPLE**
# 5. Click **Save** to add asign the SAMPLE database to the new monitoring profile
# 6. Click **Save** to create the new profile
# ### Using the API to access Monitoring Profiles
# You can access the same information throug the console API. The next cell lists all the available monitoring profiles.
r = databaseAPI.getMonitoringProfiles()
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
profileList = pd.DataFrame(json_normalize(json['resources']))
display(profileList)
else:
print(databaseAPI.getStatusCode(r))
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Get a Monitor Profile
# To get the details of a single monitoring profile, run the cell below. The complete JSON string that is returned can be long. So we only display some key information. However you can uncomment the line that prints the entire JSON string if you want to see everything.
# +
profileName = "Test"
# Find the identifier used to access this profile
profileID = str(databaseAPI.getProfileIndex(profileName))
# Retrieve the monitoring profile information for the identifier
r = databaseAPI.getMonitoringProfile(profileID)
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
# print(json)
# Print out the base information of the monitoring profile
print(json['base_info'])
# Print the common monitoring settings
print(json['monitor_config']['common_settings'])
else:
print(databaseAPI.getStatusCode(r))
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Change a Monitoring Profile
# Now that we have the complete monitoring profile as a JSON string, we can update the string and put it back to update the monitoring profile settings. In this example you change the database included in the profile and change the data collection period.
# +
profileName = "Test"
# Get the existing json for the monitoring profile
profileID = str(databaseAPI.getProfileIndex(profileName))
r = databaseAPI.getMonitoringProfile(profileID)
if (databaseAPI.getStatusCode(r)==200):
json = databaseAPI.getJSON(r)
else:
print(databaseAPI.getStatusCode(r))
# Change the json that describes the list of databases assigned to this monitoring profile
json['base_info']['assign_database_list'] = "HISTORY"
json['monitor_config']['common_settings']['collect_data_every'] = 10
# Put the updated profile back
r = databaseAPI.putMonitoringProfile(profileID, json)
if (databaseAPI.getStatusCode(r)==200):
print(profileName+' Monitoring Profile Updated')
print(json['base_info'])
print(json['monitor_config']['common_settings'])
else:
print(databaseAPI.getStatusCode(r))
# -
# ## Cleaning up
# ### Re-enable the original setup administrator account
# The setup administrator account is automatically disabled when the administrator account is configured the authentication settings. However, during emergency situations (for example, when the configured LDAP server is down, or the entries of administrator accounts are removed or renamed) when none of the administrator account is available to log into the console, it becomes necessary to re-enable the setup administrator account.
#
# To re-enable setup administrator account:
# 1. Click the **Files** icon at the bottom left of the screen
# 2. Select the **Home** directory
# 3. Select the **dmc** directory
# 3. Select the **Config** directory
# 4. Select the **dewebserver_override.properties** file
# 5. Hit the **Enter** key
# 6. In the file editor change the property value named **auth** to **superAdmin** (auth=superAdmin, no spaces before or after)
# 7. Click **Save** at the top right of the editor screen
# 8. If necessary log out of the IBM Db2 Data Management Console and log in with the setup administrator account.
# Userid: db2inst1
# Password: <PASSWORD>
# 9. Reload the Db2 Console browser page
# 10. Navigate to Authentication Setting page and configure the authentication settings. You should see that the settings have returned to the default configuration.
# #### Credits: IBM 2019-2021, <NAME> [<EMAIL>]
| console/Db2_Data_Management_Console_Management.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 开源爬虫工具OpenDataTools及其使用样例
# ## 项目介绍
#
# + OpenDataTools是一个开源爬虫工具,通过爬虫将各种数据接口简化,方便用户使用. 由QuantOS团队开发.
#
# + 目前的版本是:0.0.5
#
# + 目前只支持 空气质量AQI 的数据获取.
#
# + 项目地址: https://github.com/PKUJohnson/OpenData, 感兴趣的同学可以去研究代码.
# ## 样例介绍
#
# 空气质量数据AQI, 数据来源于 环保部网站. http://datacenter.mep.gov.cn/
# ## 准备工作
#
# 安装opendatatools(开源的数据爬虫工具)
# + pip install opendatatools
#
# 安装pyecharts
# + pip install pyecharts
# + pip install echarts-countries-pypkg
# + pip install echarts-china-provinces-pypkg
# + pip install echarts-china-cities-pypkg
# ## Case 1: API介绍(OpenDataTools获取空气质量数据)
from opendatatools import aqi
# 获取历史某日全国各大城市的AQI数据
# 返回DataFrame
df_aqi = aqi.get_daily_aqi('2018-05-27')
df_aqi
# +
# 获取实时全国各大城市的AQI数据
#df_aqi = aqi.get_hour_aqi('2018-05-28 11:00:00')
# 如果不指定时间点,会尝试获取最近的数据
df_aqi = aqi.get_hour_aqi()
df_aqi
# -
# 获取单个城市的AQI历史数据
aqi.get_daily_aqi_onecity('北京市')
#获取单个城市某日的AQI小时数据
aqi_hour = aqi.get_hour_aqi_onecity('北京市', '2018-05-26')
aqi_hour.set_index('time', inplace=True)
aqi_hour
# ## Case 2 : 获取实时全国AQI数据并画地图展示
# +
# encoding: utf-8
from opendatatools import aqi
from pyecharts import Geo
import pandas as pd
def draw_realtime_aqi_map(time = None):
from opendatatools import aqi
df_aqi = aqi.get_hour_aqi(time)
# some city cannot by process by echart
echart_unsupported_city = [
"菏泽市", "襄阳市", "恩施州", "湘西州","阿坝州", "延边州",
"甘孜州", "凉山州", "黔西南州", "黔东南州", "黔南州", "普洱市", "楚雄州", "红河州",
"文山州", "西双版纳州", "大理州", "德宏州", "怒江州", "迪庆州", "昌都市", "山南市",
"林芝市", "临夏州", "甘南州", "海北州", "黄南州", "海南州", "果洛州", "玉树州", "海西州",
"昌吉州", "博州", "克州", "伊犁哈萨克州"]
if time is None and len(df_aqi) > 0:
time = df_aqi['time'][0]
data = []
for index, row in df_aqi.iterrows():
city = row['city']
aqi = row['aqi']
if city in echart_unsupported_city:
continue
data.append( (city, aqi) )
geo = Geo("全国最新主要城市空气质量(AQI) - %s" % time , "数据来源于环保部网站",
title_color="#fff",
title_pos="center", width=1000,
height=600, background_color='#404a59')
attr, value = geo.cast(data)
geo.add("", attr, value, visual_range=[0, 150],
maptype='china',visual_text_color="#fff",
symbol_size=10, is_visualmap=True,
label_formatter='{b}', # 指定 label 只显示城市名
tooltip_formatter='{c}', # 格式:经度、纬度、值
label_emphasis_textsize=15, # 指定标签选中高亮时字体大小
label_emphasis_pos='right' # 指定标签选中高亮时字体位置
)
return geo
# -
draw_realtime_aqi_map()
# ## Case 3: 获取历史某日全国AQI数据并画地图展示
# +
# encoding: utf-8
from opendatatools import aqi
from pyecharts import Geo
import pandas as pd
def draw_his_aqi_map(date):
from opendatatools import aqi
df_aqi = aqi.get_daily_aqi(date)
#df_aqi.to_csv("aqi_daily.csv")
# some city cannot by process by echart
echart_unsupported_city = [
"菏泽市", "襄阳市", "恩施州", "湘西州","阿坝州", "延边州",
"甘孜州", "凉山州", "黔西南州", "黔东南州", "黔南州", "普洱市", "楚雄州", "红河州",
"文山州", "西双版纳州", "大理州", "德宏州", "怒江州", "迪庆州", "昌都市", "山南市",
"林芝市", "临夏州", "甘南州", "海北州", "黄南州", "海南州", "果洛州", "玉树州", "海西州",
"昌吉州", "博州", "克州", "伊犁哈萨克州"]
data = []
for index, row in df_aqi.iterrows():
city = row['city']
aqi = row['aqi']
if city in echart_unsupported_city:
continue
data.append( (city, aqi) )
geo = Geo("全国主要城市空气质量(AQI) - %s" % date , "数据来源于环保部网站",
title_color="#fff",
title_pos="center", width=1000,
height=600, background_color='#404a59')
attr, value = geo.cast(data)
geo.add("", attr, value, visual_range=[0, 150],
maptype='china',visual_text_color="#fff",
symbol_size=10, is_visualmap=True,
label_formatter='{b}', # 指定 label 只显示城市名
tooltip_formatter='{c}', # 格式:经度、纬度、值
label_emphasis_textsize=15, # 指定标签选中高亮时字体大小
label_emphasis_pos='right' # 指定标签选中高亮时字体位置
)
return geo
# -
draw_his_aqi_map('2018-05-27')
draw_his_aqi_map('2017-05-27')
draw_his_aqi_map('2016-05-27')
# ## Case 4 : 看某几个城市历史一段时间的走势图
# +
# encoding: utf-8
from pyecharts import Line
import pandas as pd
def draw_city_aqi(cities, start_date = None, end_date = None):
from opendatatools import aqi
line = Line("城市AQI趋势图")
data_dict = {}
for city in cities:
print("getting data for %s" % city)
df_aqi = aqi.get_daily_aqi_onecity(city)
df_aqi.set_index('date', inplace=True)
df_aqi.sort_index(ascending=True, inplace=True)
if start_date is not None:
df_aqi = df_aqi[df_aqi.index >= start_date]
if end_date is not None:
df_aqi = df_aqi[df_aqi.index <= end_date]
data_dict[city] = df_aqi
axis_x = df_aqi.index
axis_y = df_aqi['aqi']
line.add("aqi curve for %s" % (city), axis_x, axis_y, mark_point=["average"])
return line
# -
draw_city_aqi(['北京市','上海市'], start_date = '2018-01-01', end_date = '2018-05-31')
# ## Case 5 : 看某个城市日内小时走势图
# +
from pyecharts import Line
import pandas as pd
def draw_city_aqi_hour(cities, date):
from opendatatools import aqi
line = Line("城市AQI小时趋势图")
data_dict = {}
for city in cities:
print("getting data for %s" % city)
df_aqi = aqi.get_hour_aqi_onecity(city, date)
df_aqi.set_index('time', inplace=True)
df_aqi.sort_index(ascending=True, inplace=True)
data_dict[city] = df_aqi
axis_x = df_aqi.index
axis_y = df_aqi['aqi']
line.add("aqi curve for %s" % (city), axis_x, axis_y, mark_point=["average"])
return line
# -
draw_city_aqi_hour(['北京市', '上海市'], '2018-05-28')
#
| notebook/opendatatools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ds17-unit-1
# language: python
# name: ds17-unit-1
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/IBMikeNichols/Daily-Warm-Ups/blob/master/Unit-1/basics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="eaCRXrCTg1GI" colab_type="text"
# # Basics
# + [markdown] id="_E_4A16jg1GJ" colab_type="text"
# ## load a csv from the following url:
#
# #### https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/
#
#
# + id="da2hZDXgg1GK" colab_type="code" colab={}
# A good practice is to keep your import statements to the top of your
# notebooks
import pandas as pd
# + id="L1p1dNNUg1GO" colab_type="code" colab={}
# One thing you should begin practicing early is commenting your code!
# Another good practice is to use descriptive variable names
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data"
column_names=['mpg', 'cylinders', 'displacement', 'horsepower', 'weight', 'acceleration', 'model year', 'origin', 'car name']
# looks like this file is tab delimited?
df = pd.read_csv(url, header=None, names=column_names, delim_whitespace=True)
# + id="YtQrEpCUg1GS" colab_type="code" outputId="10fba73e-ff16-441c-bdc5-cbd4b52ea725" colab={"base_uri": "https://localhost:8080/", "height": 204}
# now print the first 5 rows of the data you just loaded
df.head()
# + id="Kjmwor3zg1GW" colab_type="code" outputId="00acb6f1-c1c6-44ab-dd55-520e8097d8d9" colab={"base_uri": "https://localhost:8080/", "height": 266}
# and the last 7
df.tail(7)
# + id="mEjsxF2pg1Ga" colab_type="code" outputId="365efc00-fa37-4ef6-c96c-7ccdf4bb342e" colab={"base_uri": "https://localhost:8080/", "height": 297}
# how many rows are in the dataset?
df.describe()
# + id="ty_r3e9Fg1Ge" colab_type="code" outputId="ce1ed52c-e24a-4cb0-bb76-709af59fbd6f" colab={}
# how many columns
# + id="n5VChHsfg1Gh" colab_type="code" outputId="b18f3b94-154d-4e23-c0d2-18185d759027" colab={}
# what is the shape of the dataset
# + [markdown] id="Ho8gLGlGg1Gl" colab_type="text"
# ### From the data directory load the iris dataset
# + id="fzbSHSXwg1Gm" colab_type="code" outputId="f4904dbd-2255-47f6-900e-6f5fe9d6736f" colab={}
column_headers = [
"sepal-length", "sepal-width", "petal-length", "petal_width", "class"
]
df =
# and print out the first 5 rows
df.head()
# + [markdown] id="mAMilAWUhPDE" colab_type="text"
# ## Make a basic graph using either of the above data sources when you're finished post the graph in the ds17 channel on slack
# + id="LDI-Lv4mg1Gs" colab_type="code" colab={}
# + [markdown] id="CeKc8dQyhdhf" colab_type="text"
# ## Save a copy of your notebook to GitHub and submit a pull request to this repository
#
#
| Unit-1/basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementing FIR filters
# <div align="right"><a href="https://people.epfl.ch/paolo.prandoni"><NAME></a>, <a href="https://www.epfl.ch/labs/lcav/">LCAV, EPFL</a></div>
# <br />
# Digital filters are fully described by their constant-coefficient difference equation (CCDE) and a CCDE can be easily translated into a few lines of code to obtain a simple filter implementation.
#
# In the case of FIR filters the coefficients in the CCDE coincide with the values of the impulse response, so that FIR filters can also be implemented as a convolution (aka an inner product) or by performing a multiplication in the frequency domain, as we will see shortly.
#
# When applying an FIR filter to a finite-length input, we also need to be mindful of the so called "border effects", that is, of the output samples that involve data indices outside of the input range.
#
# In this notebook we will look at different FIR implementations and at the associated caveats.
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
# ## 1. FIR implementation for real-time use
#
# The classic way to implement a causal filter that works in real time is the _"one-in one-out"_ approach. This will require the algorithm to keep track of past input samples and we can achieve that by implementing a persistent delay line. In Python we can either define a class or use function attributes; classes are tidier and reusable:
class FIR_loop():
def __init__(self, h):
self.h = h
self.ix = 0
self.M = len(h)
self.buf = np.zeros(self.M)
def filter(self, x):
y = 0
self.buf[self.ix] = x
for n in range(0, self.M):
y += self.h[n] * self.buf[(self.ix+self.M-n) % self.M]
self.ix = (self.ix + 1) % self.M
return y
# +
# simple moving average:
h = np.ones(5)/5
# instantiate a moving average FIR and use it
f = FIR_loop(h)
for n in range(0, 10):
print(round(f.filter(n), 3), end=' ')
# -
# While there's nothing wrong with the above implementation, when the data to be filtered is known in advance, it makes no sense to explicitly iterate over each data point and it's better to use an optimized built-in function to perform a convolution; in Numpy, such a command is `convolve`. When we do so, however, we need to take border effects into account.
# ## 2. Offline implementations and border effects
#
# When filtering a finite-length data vector, we need to decide how to deal with the "out of range" indexes that appear in the convolution. Recall that the $n$-th output sample is defined as
#
# $$
# y[n] = \sum_{k=-\infty}^{\infty} h[k]x[n-k]
# $$
#
# If the filter is an $M$-tap FIR, $h[n]$ is nonzero only fo $0 \leq n \leq M-1$; we can therefore write
#
# $$
# y[n] = \sum_{k=0}^{M-1} h[k]x[n-k]
# $$
#
# Now assume that the input is a length-$N$ signal, namely, $x[n]$ is defined only for $0 \leq n \le N$; also, we can safely consider $N > M$, otherwise exchange the roles of $x$ and $h$. Then $y[n]$ can be computed only only for $M - 1 \le n \le N-1$ since, for any other value of $n$, the sum will contain a term $x[n-k]$ where $n-k$ is outside of the valid range of indexes for the input.
#
# This means that, if we start with an $N$-point input, we can fully compute only $N-M+1$ output samples. While this may not be a problem in some applications, especially if $N \gg M$, it certainly is troublesome if repeated filtering operations end up "chipping away" at the signal little by little.
#
# The solution is to "embed" the finite-length input data signal into an infinite-length sequence and, as always, the result will depend on the embedding that we choose, that is, finite support extension or periodization. Incidentally, please note that an FIR impulse response is by definition infinite sequence, albeit a finite-support one, since it is the response of the filter to the sequence $\delta[n]$.
#
# Independently of the method, the embedding will always create "artificial" data points that are said to suffer from **border effects**. We will now explore these effects using a moving average filter and a finite-length input with a simple shape.
# +
# let's use a simple moving average:
M = 5
h = np.ones(M)/float(M)
# let's build a signal with a ramp and a plateau
x = np.concatenate((np.arange(1, 9), np.ones(5) * 8, np.arange(8,0,-1)))
plt.stem(x, use_line_collection=True);
print(f'signal length: {len(x)}')
# -
# ### 2.1. No border effects (output shorter than input)
#
# We may choose to accept the loss of data points and use only the $N-M+1$ output samples that correspond to a full overlap between the input data and the impulse response. This can be achieved by selecting `mode='valid'` in `correlate`:
y = np.convolve(x, h, mode='valid')
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
# ### 2.2. finite-support extension
#
# #### 2.2.1. Full convolution
#
# By embedding the input into a finite-support signal, the convolution sum is now well defined for all values of $n$, which now creates a new problem: the output will be potentially nonzero for all values of $n$ for which $x[n-k]$ is nonzero, that is for $0 \le n \le N+M-1$: we end up with a *longer* support for the output sequence. This is the default in `correlate` and corresponds to `mode='full'`:
y = np.convolve(x, h, mode='full')
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
# #### 2.2.2 Truncated convolution
#
# If we want to preserve the same length for input and output, we need to truncate the result. You can keep the *first* $N$ samples and discard the tail:
y = np.convolve(x, h, mode='full')[:len(x)]
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
# Note that this corresponds exactly to the online implementation of the FIR filter, stopped after $N$ input points
f = FIR_loop(h)
y1 = np.zeros(len(x))
for n, xn in enumerate(x):
y1[n] = f.filter(xn)
plt.stem(y, use_line_collection=True);
# Alternatively, you can discard half the extra samples from the beginning and half from the end of the output and distribute the border effect evenly; this is achieved in `correlate` by setting `mode='same'`:
y = np.convolve(x, h, mode='same')
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
# ### 2.3. Periodic extension
#
# The other way in which we can embed a length-$N$ signal into a sequence is to build an $N$-periodic extension $\tilde{x}[n] = x[n \mod N]$. The convolution in this case will return an $N$-periodic output:
#
# $$
# \tilde{y}[n] = \sum_{k=0}^{M-1} h[k]\tilde{x}[n-k]
# $$
#
# #### 2.3.1. Circular convolution
#
# We can easily implement a circular convolution using `convolve`: since the overlap between time-reversed impulse response and input is already good for the last $N-M$ points of the output, we just need to consider two periods of the input to compute the first $M$:
def cconv(x, h):
# as before, we assume len(h) < len(x)
N = len(x)
xp = np.concatenate((x, x))
# full convolution
y = np.convolve(xp, h)
return y[N:2*N]
y = cconv(x, h)
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
# OK, clearly the result is not necessarily what we expected; note however that in both circular and "normal" convolution, you still have $M-1$ output samples "touched" by border effects, it's just that the border effects act differently in the two cases.
#
# #### 2.3.2. Normal convolution via circular convolution
#
# Interestingly, if you zero-pad the input signal with $M-1$ zeros, you can still obtain a "normal" convolution using a circular convolution:
# +
y = cconv(np.concatenate((x, np.zeros(M-1))), h)
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
# plot in red the *difference* with the standard conv
plt.stem(y - np.convolve(x, h, mode='full'), markerfmt='ro', use_line_collection=True);
# -
# This seems to be a completely redundant way of convolving two signals but it is interesting? because circular convolution can be efficiently implemented in the frequency-domain, as we will now show.
# ## 3. Offline FIR implementations using the FFT
#
# The convolution theorem states that, for infinite sequences, convolution in the time domain corresponds to multiplication in the frequency domain. With this, we could compute the output of a filter using the inverse Fourier transform like so:
#
# $$
# (x\ast y)[n] = \mbox{IDTFT}\{X(e^{j\omega})Y(e^{j\omega})\}[n]
# $$
#
# Can we apply this result to the finite-length case? In other words, what is the inverse DFT of the product of two DFTs? Let's see:
#
# \begin{align}
# \sum_{k=0}^{N-1}X[k]Y[k]e^{j\frac{2\pi}{N}nk} &= \sum_{k=0}^{N-1}\sum_{p=0}^{N-1}x[p]e^{-j\frac{2\pi}{N}pk}\sum_{q=0}^{N-1}y[q]e^{-j\frac{2\pi}{N}qk} \,e^{j\frac{2\pi}{N}nk} \\
# &= \sum_{p=0}^{N-1}\sum_{q=0}^{N-1}x[p]y[q]\sum_{k=0}^{N-1}e^{j\frac{2\pi}{N}(n-p-q)k} \\
# &= N\sum_{p=0}^{N-1}x[p]y[(n-p) \mod N]
# \end{align}
#
# The results follows from the fact that $\sum_{k=0}^{N-1}e^{j\frac{2\pi}{N}(n-p-q)k}$ is nonzero only for $n-p-q$ multiple of $N$; as $p$ varies from $0$ to $N-1$, the corresponding value of $q$ between $0$ and $N$ that makes $n-p-q$ multiple of $N$ is $(n-p) \mod N$.
#
# So the fundamental result is: **the inverse DFT of the product of two DFTs is the circular convolution of the underlying time-domain sequences!**
#
#
# To apply this result to FIR filtering, the first step is to choose the space for the DFTs. In our case we have a finite-length data vector of length $N$ and a finite-support impulse response of length $M$ with $M<N$ so let's operate in $\mathbb{C}^N$ by zero-padding the impulse response to size $N$. Also, we most likely want the normal convolution, so let's zero-pad both signals by an additional $M-1$ samples as explained in Section 2.3.2 above.
def DFTconv(x, h, mode='full'):
# we want the compute the full convolution
N = len(x)
M = len(h)
X = np.fft.fft(x, n=N+M-1)
H = np.fft.fft(h, n=N+M-1)
# we're using real-valued signals, so drop the imaginary part
y = np.real(np.fft.ifft(X * H))
if mode == 'valid':
# only N-M+1 points, starting at M-1
return y[M-1:N]
elif mode == 'same':
return y[int((M-1)/2):int((M-1)/2+N)]
else:
return y
# Let's verify that the results are the same
# +
y = np.convolve(x, h, mode='valid')
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
y = DFTconv(x, h, mode='valid')
print(f'signal length: {len(y)}')
plt.stem(y, markerfmt='ro', use_line_collection=True);
# +
y = np.convolve(x, h, mode='same')
print(f'signal length: {len(y)}')
plt.stem(y, use_line_collection=True);
y = DFTconv(x, h, mode='same')
print(f'signal length: {len(y)}')
plt.stem(y, markerfmt='ro', use_line_collection=True);
# -
# ### 3.1. Computational requirements
#
# Of course the question at this point is: why go through the trouble of taking DFTs if all we want is the standard convolution? The answer is: **computational efficiency.**
#
# If you look at the convolution sum, each output sample requires $M$ multiplications (and $M-1$ additions but let's just consider multiplications). In order to filter an $N$-point signal we will need $NM$ multiplications. Assume $N \approx M$ and you can see that the computational requirements are on the order of $M^2$. If we go the DFT route using an efficient FFT implementation we have approximately:
#
# * $M\log_2 M$ multiplication to compute $H[k]$
# * $M\log_2 M$ multiplication to compute $X[k]$
# * $M\log_2 M$ multiplication to compute $X[k]H[k]$
# * $M\log_2 M$ multiplication to compute the inverse DFT
#
# Even considering that we now have to use complex multiplications (which will cost twice as much), we can estimate the cost of the DFT based convolution at around $8M\log_2M$, which is smaller than $M^2$ as soon as $M>44$.
# In practice, the data vector is much longer than the impulse response so that filtering via standard convolution requires on the order of $MN$ operations. Two techniques, called [Overlap Add](https://en.wikipedia.org/wiki/Overlap%E2%80%93add_method) and [Overlap Save](https://en.wikipedia.org/wiki/Overlap%E2%80%93save_method)
# can be used to divide the convolution into $N/M$ independent convolutions between $h[n]$ and an $M$-sized piece of $x[n]$; FFT-based convolution can then be used on each piece. While the exact cost per sample of each technique is a bit complicated to estimate, as a rule of thumb **as soon as the impulse response is longer than 50 samples, it's more convenient to use DFT-based filtering.**
# # ---------------------------------
#
#
#
#
# ### Did you like this Notebook?
# Yes, no, maybe? Why don't you give us some feedback using the completely anonymous form below? Thank you!
from IPython.display import IFrame
IFrame('https://www.surveymonkey.com/r/NOTOSURVEY?notebook_set=COM303¬ebook_id=FIRImplementation', 600, 800)
| FIRimplementation/FIRImplementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## How likely is it for people in the same room to share the same birth date?
# P(same) = 1 - P(no-overlap)
#
# P(no overlap) = 365/365 * 364/365 * 363/365 etc
#Import libraries
import pandas as pd
import numpy as np
import seaborn as sns
import os
import matplotlib.pyplot as plt
# %matplotlib inline
# **Create function to calculate the probability of overlap**
# +
#Function to determine probability of being born on the same day
def prob_overlap(room = 20):
return 1 - np.product((365 - np.arange(room)) / 365)
#Plot the probabilities when the room size increases
plt.figure(figsize=(16,8))
plt.plot([prob_overlap(r) for r in range(1, 35)]);
# -
# **Create function to simulate and then caculate the probability of overlap**
#Simulation function to create room with random birthdays and verify how ofter we have people sharing the same birth data
#1)Create empty array to keep track of results
#2)Run simulation and verify if the length of unique values equals the lenght of all random birthdays in the room
#3)Return the probability of being born on the same date together with the array that keeps track
def simulate_room(simulations=1000, room=20):
track_array = np.empty((0,1), bool)
for simul in range(simulations):
track_array = np.append(track_array,
len(np.unique(np.random.randint(1, 365, room))) == room
)
probability = (simulations - np.sum(track_array)) / simulations
return probability
# **Visualize calculation versus simulation**
plt.figure(figsize=(16,8))
plt.plot([prob_overlap(room=r) for r in range(1, 35)], label='calculate');
plt.plot([simulate_room(room=r) for r in range(1, 35)], label='simulate');
plt.legend();
# **Use dataset to validate**
#Set working directory
os.chdir('C:\\Users\\<NAME>\\DiP\\Python klasje\\data')
df = pd.read_csv('birthdays.csv')
#Function to calculate the probability of being born on a specific date based on the dataset
def prob_born(dataf):
prob_born_df = (dataf
.assign(date = lambda d: pd.to_datetime(d['date']))
.assign(day_of_week = lambda d: d['date'].dt.dayofyear)
.groupby('day_of_week')
.agg(n_births = ('births', 'sum'))
.assign(prob = lambda d: d['n_births'] / np.sum(d['n_births']))
)
return prob_born_df
#Create dataframe containing probabilities for the different days
birth_prob_df = df.pipe(prob_born)
#Plot the probability to be born for the different days
plt.figure(figsize=(16,8))
birth_prob_df['prob'].plot()
#Simulation function to create room with random birthdays based on the USA birthday probability distribution
def simulate_room_true(dataf, simulations=1000, room=20):
track_array = np.empty((0,1), bool)
for simul in range(simulations):
track_array = np.append(track_array,
len(np.unique(np.random.choice(dataf.index, p=dataf['prob'], size=room))) == room
)
probability = (simulations - np.sum(track_array)) / simulations
return probability
# **Visualize calculation versus true simulation**
plt.figure(figsize=(16,8))
plt.plot([prob_overlap(room=r) for r in range(1, 35)], label='calculate');
plt.plot([simulate_room_true(dataf=birth_prob_df, room=r) for r in range(1, 35)], label='simulate_true')
plt.legend();
| notebooks/birthday_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Part 1: Training Tensorflow 2.0 Model on Azure Machine Learning Service
#
# ## Overview of the part 1
# This notebook is Part 1 (Preparing Data and Model Training) of a four part workshop that demonstrates an end-to-end workflow using Tensorflow 2.0 on Azure Machine Learning service. The different components of the workshop are as follows:
#
# - Part 1: [Preparing Data and Model Training](https://github.com/microsoft/bert-stack-overflow/blob/master/1-Training/AzureServiceClassifier_Training.ipynb)
# - Part 2: [Inferencing and Deploying a Model](https://github.com/microsoft/bert-stack-overflow/blob/master/2-Inferencing/AzureServiceClassifier_Inferencing.ipynb)
# - Part 3: [Setting Up a Pipeline Using MLOps](https://github.com/microsoft/bert-stack-overflow/tree/master/3-ML-Ops)
# - Part 4: [Explaining Your Model Interpretability](https://github.com/microsoft/bert-stack-overflow/blob/master/4-Interpretibility/IBMEmployeeAttritionClassifier_Interpretability.ipynb)
#
# **This notebook will cover the following topics:**
#
# - Stackoverflow question tagging problem
# - Introduction to Transformer and BERT deep learning models
# - Introduction to Azure Machine Learning service
# - Preparing raw data for training using Apache Spark
# - Registering cleanup training data as a Dataset
# - Debugging the model in Tensorflow 2.0 Eager Mode
# - Training the model on GPU cluster
# - Monitoring training progress with built-in Tensorboard dashboard
# - Automated search of best hyper-parameters of the model
# - Registering the trained model for future deployment
# ## Prerequisites
# This notebook is designed to be run in Azure ML Notebook VM. See [readme](https://github.com/microsoft/bert-stack-overflow/blob/master/README.md) file for instructions on how to create Notebook VM and open this notebook in it.
# ### Check Azure Machine Learning Python SDK version
#
# This tutorial requires version 1.0.69 or higher. Let's check the version of the SDK:
# +
import azureml.core
print("Azure Machine Learning Python SDK version:", azureml.core.VERSION)
# -
# ## Stackoverflow Question Tagging Problem
# In this workshop we will use powerful language understanding model to automatically route Stackoverflow questions to the appropriate support team on the example of Azure services.
#
# One of the key tasks to ensuring long term success of any Azure service is actively responding to related posts in online forums such as Stackoverflow. In order to keep track of these posts, Microsoft relies on the associated tags to direct questions to the appropriate support team. While Stackoverflow has different tags for each Azure service (azure-web-app-service, azure-virtual-machine-service, etc), people often use the generic **azure** tag. This makes it hard for specific teams to track down issues related to their product and as a result, many questions get left unanswered.
#
# **In order to solve this problem, we will build a model to classify posts on Stackoverflow with the appropriate Azure service tag.**
#
# We will be using a BERT (Bidirectional Encoder Representations from Transformers) model which was published by researchers at Google AI Reasearch. Unlike prior language representation models, BERT is designed to pre-train deep bidirectional representations from unlabeled text by jointly conditioning on both left and right context in all layers. As a result, the pre-trained BERT model can be fine-tuned with just one additional output layer to create state-of-the-art models for a wide range of natural language processing (NLP) tasks without substantial architecture modifications.
#
# ## Why use BERT model?
# [Introduction of BERT model](https://arxiv.org/pdf/1810.04805.pdf) changed the world of NLP. Many NLP problems that before relied on specialized models to achive state of the art performance are now solved with BERT better and with more generic approach.
#
# If we look at the leaderboards on such popular NLP problems as GLUE and SQUAD, most of the top models are based on BERT:
# * [GLUE Benchmark Leaderboard](https://gluebenchmark.com/leaderboard/)
# * [SQuAD Benchmark Leaderboard](https://rajpurkar.github.io/SQuAD-explorer/)
#
# Recently, Allen Institue for AI announced new language understanding system called Aristo [https://allenai.org/aristo/](https://allenai.org/aristo/). The system has been developed for 20 years, but it's performance was stuck at 60% on 8th grade science test. The result jumped to 90% once researchers adopted BERT as core language understanding component. With BERT Aristo now solves the test with A grade.
# ## Quick Overview of How BERT model works
#
# The foundation of BERT model is Transformer model, which was introduced in [Attention Is All You Need paper](https://arxiv.org/abs/1706.03762). Before that event the dominant way of processing language was Recurrent Neural Networks (RNNs). Let's start our overview with RNNs.
#
# ## RNNs
#
# RNNs were powerful way of processing language due to their ability to memorize its previous state and perform sophisticated inference based on that.
#
# <img src="https://miro.medium.com/max/400/1*L38xfe59H5tAgvuIjKoWPg.png" alt="Drawing" style="width: 100px;"/>
#
# _Taken from [1](https://towardsdatascience.com/transformers-141e32e69591)_
#
# Applied to language translation task, the processing dynamics looked like this.
#
# 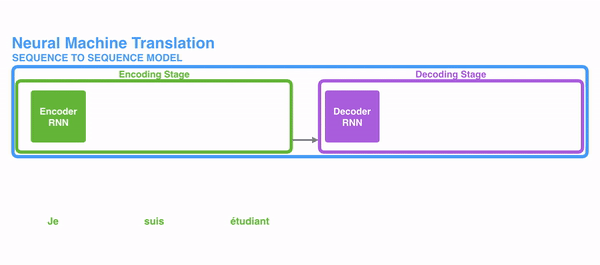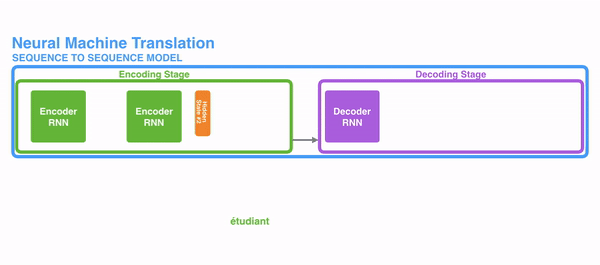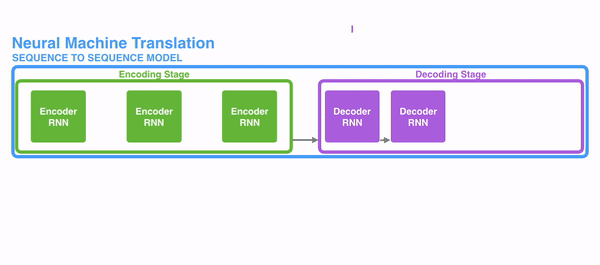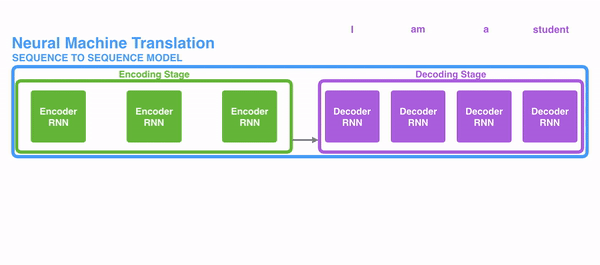
# _Taken from [2](https://jalammar.github.io/visualizing-neural-machine-translation-mechanics-of-seq2seq-models-with-attention/)_
#
# But RNNs suffered from 2 disadvantes:
# 1. Sequential computation put a limit on parallelization, which limited effectiveness of larger models.
# 2. Long term relationships between words were harder to detect.
# ## Transformers
#
# Transformers were designed to address these two limitations of RNNs.
#
# <img src="https://miro.medium.com/max/2436/1*V2435M1u0tiSOz4nRBfl4g.png" alt="Drawing" style="width: 500px;"/>
#
# _Taken from [3](http://jalammar.github.io/illustrated-transformer/)_
#
# In each Encoder layer Transformer performs Self-Attention operation which detects relationships between all word embeddings in one matrix multiplication operation.
#
# <img src="https://miro.medium.com/max/2176/1*fL8arkEFVKA3_A7VBgapKA.gif" alt="Drawing" style="width: 500px;"/>
#
# _Taken from [4](https://towardsdatascience.com/deconstructing-bert-part-2-visualizing-the-inner-workings-of-attention-60a16d86b5c1)_
#
# ## BERT Model
#
# BERT is a very large network with multiple layers of Transformers (12 for BERT-base, and 24 for BERT-large). The model is first pre-trained on large corpus of text data (WikiPedia + books) using un-superwised training (predicting masked words in a sentence). During pre-training the model absorbs significant level of language understanding.
#
# <img src="http://jalammar.github.io/images/bert-output-vector.png" alt="Drawing" style="width: 700px;"/>
#
# _Taken from [5](http://jalammar.github.io/illustrated-bert/)_
#
# Pre-trained network then can easily be fine-tuned to solve specific language task, like answering questions, or categorizing spam emails.
#
# <img src="http://jalammar.github.io/images/bert-classifier.png" alt="Drawing" style="width: 700px;"/>
#
# _Taken from [5](http://jalammar.github.io/illustrated-bert/)_
#
# ## What is Azure Machine Learning Service?
# Azure Machine Learning service is a cloud service that you can use to develop and deploy machine learning models. Using Azure Machine Learning service, you can track your models as you build, train, deploy, and manage them, all at the broad scale that the cloud provides.
# 
#
#
# #### How can we use it for training machine learning models?
# Training machine learning models, particularly deep neural networks, is often a time- and compute-intensive task. Once you've finished writing your training script and running on a small subset of data on your local machine, you will likely want to scale up your workload.
#
# To facilitate training, the Azure Machine Learning Python SDK provides a high-level abstraction, the estimator class, which allows users to easily train their models in the Azure ecosystem. You can create and use an Estimator object to submit any training code you want to run on remote compute, whether it's a single-node run or distributed training across a GPU cluster.
# ## Connect To Workspace
#
# The [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class)?view=azure-ml-py) is the top-level resource for Azure Machine Learning, providing a centralized place to work with all the artifacts you create when you use Azure Machine Learning. The workspace holds all your experiments, compute targets, models, datastores, etc.
#
# You can [open ml.azure.com](https://ml.azure.com) to access your workspace resources through a graphical user interface of **Azure Machine Learning studio**.
#
# 
#
# **You will be asked to login in the next step. Use your Microsoft AAD credentials.**
# +
from azureml.core import Workspace
workspace = Workspace.from_config()
print('Workspace name: ' + workspace.name,
'Azure region: ' + workspace.location,
'Subscription id: ' + workspace.subscription_id,
'Resource group: ' + workspace.resource_group, sep = '\n')
# -
# ## Create Compute Target
#
# A [compute target](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.computetarget?view=azure-ml-py) is a designated compute resource/environment where you run your training script or host your service deployment. This location may be your local machine or a cloud-based compute resource. Compute targets can be reused across the workspace for different runs and experiments.
#
# For this tutorial, we will create an auto-scaling [Azure Machine Learning Compute](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.compute.amlcompute?view=azure-ml-py) cluster, which is a managed-compute infrastructure that allows the user to easily create a single or multi-node compute. To create the cluster, we need to specify the following parameters:
#
# - `vm_size`: The is the type of GPUs that we want to use in our cluster. For this tutorial, we will use **Standard_NC12s_v3 (NVIDIA V100) GPU Machines** .
# - `idle_seconds_before_scaledown`: This is the number of seconds before a node will scale down in our auto-scaling cluster. We will set this to **6000** seconds.
# - `min_nodes`: This is the minimum numbers of nodes that the cluster will have. To avoid paying for compute while they are not being used, we will set this to **0** nodes.
# - `max_modes`: This is the maximum number of nodes that the cluster will scale up to. Will will set this to **2** nodes.
#
# **When jobs are submitted to the cluster it takes approximately 5 minutes to allocate new nodes**
# +
from azureml.core.compute import AmlCompute, ComputeTarget
cluster_name = 'v100cluster'
compute_config = AmlCompute.provisioning_configuration(vm_size='Standard_NC12s_v3',
idle_seconds_before_scaledown=6000,
min_nodes=0,
max_nodes=2)
compute_target = ComputeTarget.create(workspace, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# To ensure our compute target was created successfully, we can check it's status.
compute_target.get_status().serialize()
# #### If the compute target has already been created, then you (and other users in your workspace) can directly run this cell.
compute_target = workspace.compute_targets['v100cluster']
# ## Prepare Data Using Apache Spark
#
# To train our model, we used the Stackoverflow data dump from [Stack exchange archive](https://archive.org/download/stackexchange). Since the Stackoverflow _posts_ dataset is 12GB, we prepared the data using [Apache Spark](https://spark.apache.org/) framework on a scalable Spark compute cluster in [Azure Databricks](https://azure.microsoft.com/en-us/services/databricks/).
#
# For the purpose of this tutorial, we have processed the data ahead of time and uploaded it to an [Azure Blob Storage](https://azure.microsoft.com/en-us/services/storage/blobs/) container. The full data processing notebook can be found in the _spark_ folder.
#
# * **ACTION**: Open and explore [data preparation notebook](spark/stackoverflow-data-prep.ipynb).
#
# ## Register Datastore
# A [Datastore](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.datastore.datastore?view=azure-ml-py) is used to store connection information to a central data storage. This allows you to access your storage without having to hard code this (potentially confidential) information into your scripts.
#
# In this tutorial, the data was been previously prepped and uploaded into a central [Blob Storage](https://azure.microsoft.com/en-us/services/storage/blobs/) container. We will register this container into our workspace as a datastore using a [shared access signature (SAS) token](https://docs.microsoft.com/en-us/azure/storage/common/storage-sas-overview).
# +
from azureml.core import Datastore, Dataset
datastore_name = 'tfworld'
container_name = 'azureml-blobstore-7c6bdd88-21fa-453a-9c80-16998f02935f'
account_name = 'tfworld6818510241'
sas_token = <PASSWORD>&se=2019-11-08T05:12:15Z&st=2019-10-23T20:12:15Z&spr=https&sig=eDqnc51TkqiIklpQfloT5vcU70pgzDuKb5PAGTvCdx4%3D'
datastore = Datastore.register_azure_blob_container(workspace=workspace,
datastore_name=datastore_name,
container_name=container_name,
account_name=account_name,
sas_token=sas_token)
# -
# #### If the datastore has already been registered, then you (and other users in your workspace) can directly run this cell.
datastore = workspace.datastores['tfworld']
# #### What if my data wasn't already hosted remotely?
# All workspaces also come with a blob container which is registered as a default datastore. This allows you to easily upload your own data to a remote storage location. You can access this datastore and upload files as follows:
# ```
# datastore = workspace.get_default_datastore()
# ds.upload(src_dir='<LOCAL-PATH>', target_path='<REMOTE-PATH>')
# ```
#
# ## Register Dataset
#
# Azure Machine Learning service supports first class notion of a Dataset. A [Dataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.dataset.dataset?view=azure-ml-py) is a resource for exploring, transforming and managing data in Azure Machine Learning. The following Dataset types are supported:
#
# * [TabularDataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) represents data in a tabular format created by parsing the provided file or list of files.
#
# * [FileDataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.filedataset?view=azure-ml-py) references single or multiple files in datastores or from public URLs.
#
# First, we will use visual tools in Azure ML studio to register and explore our dataset as Tabular Dataset.
#
# * **ACTION**: Follow [create-dataset](images/create-dataset.ipynb) guide to create Tabular Dataset from our training data.
# #### Use created dataset in code
# +
from azureml.core import Dataset
# Get a dataset by name
tabular_ds = Dataset.get_by_name(workspace=workspace, name='Stackoverflow dataset')
# Load a TabularDataset into pandas DataFrame
df = tabular_ds.to_pandas_dataframe()
df.head(10)
# -
# ## Register Dataset using SDK
#
# In addition to UI we can register datasets using SDK. In this workshop we will register second type of Datasets using code - File Dataset. File Dataset allows specific folder in our datastore that contains our data files to be registered as a Dataset.
#
# There is a folder within our datastore called **azure-service-data** that contains all our training and testing data. We will register this as a dataset.
# +
azure_dataset = Dataset.File.from_files(path=(datastore, 'azure-service-classifier/data'))
azure_dataset = azure_dataset.register(workspace=workspace,
name='Azure Services Dataset',
description='Dataset containing azure related posts on Stackoverflow')
# -
# #### If the dataset has already been registered, then you (and other users in your workspace) can directly run this cell.
azure_dataset = workspace.datasets['Azure Services Dataset']
# ## Explore Training Code
# In this workshop the training code is provided in [train.py](./train.py) and [model.py](./model.py) files. The model is based on popular [huggingface/transformers](https://github.com/huggingface/transformers) libary. Transformers library provides performant implementation of BERT model with high level and easy to use APIs based on Tensorflow 2.0.
#
# 
#
# * **ACTION**: Explore _train.py_ and _model.py_ using [Azure ML studio > Notebooks tab](images/azuremlstudio-notebooks-explore.png)
# * NOTE: You can also explore the files using Jupyter or Jupyter Lab UI.
# ## Test Locally
#
# Let's try running the script locally to make sure it works before scaling up to use our compute cluster. To do so, you will need to install the transformers libary.
# %%pip install transformers==2.0.0
# We have taken a small partition of the dataset and included it in this repository. Let's take a quick look at the format of the data.
data_dir = './data'
import os
import pandas as pd
data = pd.read_csv(os.path.join(data_dir, 'train.csv'), header=None)
data.head(5)
# Now we know what the data looks like, let's test out our script!
import sys
# !{sys.executable} train.py --data_dir {data_dir} --max_seq_length 128 --batch_size 16 --learning_rate 3e-5 --steps_per_epoch 5 --num_epochs 1 --export_dir ../outputs/model
# ## Debugging in TensorFlow 2.0 Eager Mode
#
# Eager mode is new feature in TensorFlow 2.0 which makes understanding and debugging models easy. Let's start by configuring our remote debugging environment.
#
# #### Configure VS Code Remote connection to Notebook VM
#
# * **ACTION**: Install [Microsoft VS Code](https://code.visualstudio.com/) on your local machine.
#
# * **ACTION**: Follow this [configuration guide](https://github.com/danielsc/azureml-debug-training/blob/master/Setting%20up%20VSCode%20Remote%20on%20an%20AzureML%20Notebook%20VM.md) to setup VS Code Remote connection to Notebook VM.
#
# #### Debug training code using step-by-step debugger
#
# * **ACTION**: Open Remote VS Code session to your Notebook VM.
# * **ACTION**: Open file `/home/azureuser/cloudfiles/code/<username>/bert-stack-overflow/1-Training/train_eager.py`.
# * **ACTION**: Set break point in the file and start Python debugging session.
#
# On a CPU machine training on a full dataset will take approximatly 1.5 hours. Although it's a small dataset, it still takes a long time. Let's see how we can speed up the training by using latest NVidia V100 GPUs in the Azure cloud.
# ## Perform Experiment
#
# Now that we have our compute target, dataset, and training script working locally, it is time to scale up so that the script can run faster. We will start by creating an [experiment](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment.experiment?view=azure-ml-py). An experiment is a grouping of many runs from a specified script. All runs in this tutorial will be performed under the same experiment.
# +
from azureml.core import Experiment
experiment_name = 'azure-service-classifier'
experiment = Experiment(workspace, name=experiment_name)
# -
# #### Create TensorFlow Estimator
#
# The Azure Machine Learning Python SDK Estimator classes allow you to easily construct run configurations for your experiments. They allow you too define parameters such as the training script to run, the compute target to run it on, framework versions, additional package requirements, etc.
#
# You can also use a generic [Estimator](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.estimator.estimator?view=azure-ml-py) to submit training scripts that use any learning framework you choose.
#
# For popular libaries like PyTorch and Tensorflow you can use their framework specific estimators. We will use the [TensorFlow Estimator](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.dnn.tensorflow?view=azure-ml-py) for our experiment.
# +
from azureml.train.dnn import TensorFlow
estimator1 = TensorFlow(source_directory='.',
entry_script='train.py',
compute_target=compute_target,
script_params = {
'--data_dir': azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length': 128,
'--batch_size': 32,
'--learning_rate': 3e-5,
'--steps_per_epoch': 150,
'--num_epochs': 3,
'--export_dir':'./outputs/model'
},
framework_version='2.0',
use_gpu=True,
pip_packages=['transformers==2.0.0', 'azureml-dataprep[fuse,pandas]==1.1.22'])
# -
# A quick description for each of the parameters we have just defined:
#
# - `source_directory`: This specifies the root directory of our source code.
# - `entry_script`: This specifies the training script to run. It should be relative to the source_directory.
# - `compute_target`: This specifies to compute target to run the job on. We will use the one created earlier.
# - `script_params`: This specifies the input parameters to the training script. Please note:
#
# 1) *azure_dataset.as_named_input('azureservicedata').as_mount()* mounts the dataset to the remote compute and provides the path to the dataset on our datastore.
#
# 2) All outputs from the training script must be outputted to an './outputs' directory as this is the only directory that will be saved to the run.
#
#
# - `framework_version`: This specifies the version of TensorFlow to use. Use Tensorflow.get_supported_verions() to see all supported versions.
# - `use_gpu`: This will use the GPU on the compute target for training if set to True.
# - `pip_packages`: This allows you to define any additional libraries to install before training.
# #### 1) Submit First Run
#
# We can now train our model by submitting the estimator object as a [run](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.run.run?view=azure-ml-py).
run1 = experiment.submit(estimator1)
# We can view the current status of the run and stream the logs from within the notebook.
from azureml.widgets import RunDetails
RunDetails(run1).show()
# You cancel a run at anytime which will stop the run and scale down the nodes in the compute target.
run1.cancel()
# While we wait for the run to complete, let's go over how a Run is executed in Azure Machine Learning.
#
# 
# #### 2) Add Metrics Logging
#
# So we were able to clone a Tensorflow 2.0 project and run it without any changes. However, with larger scale projects we would want to log some metrics in order to make it easier to monitor the performance of our model.
#
# We can do this by adding a few lines of code into our training script:
#
# ```python
# # 1) Import SDK Run object
# from azureml.core.run import Run
#
# # 2) Get current service context
# run = Run.get_context()
#
# # 3) Log the metrics that we want
# run.log('val_accuracy', float(logs.get('val_accuracy')))
# run.log('accuracy', float(logs.get('accuracy')))
# ```
# We've created a *train_logging.py* script that includes logging metrics as shown above.
#
# * **ACTION**: Explore _train_logging.py_ using [Azure ML studio > Notebooks tab](images/azuremlstudio-notebooks-explore.png)
# We can submit this run in the same way that we did before.
#
# *Since our cluster can scale automatically to two nodes, we can run this job simultaneously with the previous one.*
# +
estimator2 = TensorFlow(source_directory='.',
entry_script='train_logging.py',
compute_target=compute_target,
script_params = {
'--data_dir': azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length': 128,
'--batch_size': 32,
'--learning_rate': 3e-5,
'--steps_per_epoch': 150,
'--num_epochs': 3,
'--export_dir':'./outputs/model'
},
framework_version='2.0',
use_gpu=True,
pip_packages=['transformers==2.0.0', 'azureml-dataprep[fuse,pandas]==1.1.22'])
run2 = experiment.submit(estimator2)
# -
# Now if we view the current details of the run, you will notice that the metrics will be logged into graphs.
from azureml.widgets import RunDetails
RunDetails(run2).show()
# #### 3) Monitoring metrics with Tensorboard
#
# Tensorboard is a popular Deep Learning Training visualization tool and it's built-in into TensorFlow framework. We can easily add tracking of the metrics in Tensorboard format by adding Tensorboard callback to the **fit** function call.
# ```python
# # Add callback to record Tensorboard events
# model.fit(train_dataset, epochs=FLAGS.num_epochs,
# steps_per_epoch=FLAGS.steps_per_epoch, validation_data=valid_dataset,
# callbacks=[
# AmlLogger(),
# tf.keras.callbacks.TensorBoard(update_freq='batch')]
# )
# ```
#
# #### Launch Tensorboard
# Azure ML service provides built-in integration with Tensorboard through **tensorboard** package.
#
# While the run is in progress (or after it has completed), we can start Tensorboard with the run as its target, and it will begin streaming logs.
# +
from azureml.tensorboard import Tensorboard
# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here
tb = Tensorboard([run2])
# If successful, start() returns a string with the URI of the instance.
tb.start()
# -
# #### Stop Tensorboard
# When you're done, make sure to call the stop() method of the Tensorboard object, or it will stay running even after your job completes.
tb.stop()
# ## Check the model performance
#
# Last training run produced model of decent accuracy. Let's test it out and see what it does. First, let's check what files our latest training run produced and download the model files.
#
# #### Download model files
run2.get_file_names()
# +
run2.download_files(prefix='outputs/model')
# If you haven't finished training the model then just download pre-made model from datastore
datastore.download('./',prefix="azure-service-classifier/model")
# -
# #### Instantiate the model
#
# Next step is to import our model class and instantiate fine-tuned model from the model file.
# +
from model import TFBertForMultiClassification
from transformers import BertTokenizer
import tensorflow as tf
def encode_example(text, max_seq_length):
# Encode inputs using tokenizer
inputs = tokenizer.encode_plus(
text,
add_special_tokens=True,
max_length=max_seq_length
)
input_ids, token_type_ids = inputs["input_ids"], inputs["token_type_ids"]
# The mask has 1 for real tokens and 0 for padding tokens. Only real tokens are attended to.
attention_mask = [1] * len(input_ids)
# Zero-pad up to the sequence length.
padding_length = max_seq_length - len(input_ids)
input_ids = input_ids + ([0] * padding_length)
attention_mask = attention_mask + ([0] * padding_length)
token_type_ids = token_type_ids + ([0] * padding_length)
return input_ids, attention_mask, token_type_ids
labels = ['azure-web-app-service', 'azure-storage', 'azure-devops', 'azure-virtual-machine', 'azure-functions']
# Load model and tokenizer
loaded_model = TFBertForMultiClassification.from_pretrained('azure-service-classifier/model', num_labels=len(labels))
tokenizer = BertTokenizer.from_pretrained('bert-base-cased')
print("Model loaded from disk.")
# -
# #### Define prediction function
#
# Using the model object we can interpret new questions and predict what Azure service they talk about. To do that conveniently we'll define **predict** function.
# Prediction function
def predict(question):
input_ids, attention_mask, token_type_ids = encode_example(question, 128)
predictions = loaded_model.predict({
'input_ids': tf.convert_to_tensor([input_ids], dtype=tf.int32),
'attention_mask': tf.convert_to_tensor([attention_mask], dtype=tf.int32),
'token_type_ids': tf.convert_to_tensor([token_type_ids], dtype=tf.int32)
})
prediction = labels[predictions[0].argmax().item()]
probability = predictions[0].max()
result = {
'prediction': str(labels[predictions[0].argmax().item()]),
'probability': str(predictions[0].max())
}
print('Prediction: {}'.format(prediction))
print('Probability: {}'.format(probability))
# #### Experiement with our new model
#
# Now we can easily test responses of the model to new inputs.
# * **ACTION**: Invent yout own input for one of the 5 services our model understands: 'azure-web-app-service', 'azure-storage', 'azure-devops', 'azure-virtual-machine', 'azure-functions'.
# Route question
predict("How can I specify Service Principal in devops pipeline when deploying virtual machine")
# Now more tricky cae - the opposite
predict("How can virtual machine trigger devops pipeline")
# ## Distributed Training Across Multiple GPUs
#
# Distributed training allows us to train across multiple nodes if your cluster allows it. Azure Machine Learning service helps manage the infrastructure for training distributed jobs. All we have to do is add the following parameters to our estimator object in order to enable this:
#
# - `node_count`: The number of nodes to run this job across. Our cluster has a maximum node limit of 2, so we can set this number up to 2.
# - `process_count_per_node`: The number of processes to enable per node. The nodes in our cluster have 2 GPUs each. We will set this value to 2 which will allow us to distribute the load on both GPUs. Using multi-GPUs nodes is benefitial as communication channel bandwidth on local machine is higher.
# - `distributed_training`: The backend to use for our distributed job. We will be using an MPI (Message Passing Interface) backend which is used by Horovod framework.
#
# We use [Horovod](https://github.com/horovod/horovod), which is a framework that allows us to easily modifying our existing training script to be run across multiple nodes/GPUs. The distributed training script is saved as *train_horovod.py*.
#
# * **ACTION**: Explore _train_horovod.py_ using [Azure ML studio > Notebooks tab](images/azuremlstudio-notebooks-explore.png)
# We can submit this run in the same way that we did with the others, but with the additional parameters.
# +
from azureml.train.dnn import Mpi
estimator3 = TensorFlow(source_directory='./',
entry_script='train_horovod.py',compute_target=compute_target,
script_params = {
'--data_dir': azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length': 128,
'--batch_size': 32,
'--learning_rate': 3e-5,
'--steps_per_epoch': 150,
'--num_epochs': 3,
'--export_dir':'./outputs/model'
},
framework_version='2.0',
node_count=1,
distributed_training=Mpi(process_count_per_node=2),
use_gpu=True,
pip_packages=['transformers==2.0.0', 'azureml-dataprep[fuse,pandas]==1.1.22'])
run3 = experiment.submit(estimator3)
# -
# Once again, we can view the current details of the run.
from azureml.widgets import RunDetails
RunDetails(run3).show()
# Once the run completes note the time it took. It should be around 5 minutes. As you can see, by moving to the cloud GPUs and using distibuted training we managed to reduce training time of our model from more than an hour to 5 minutes. This greatly improves speed of experimentation and innovation.
# ## Tune Hyperparameters Using Hyperdrive
#
# So far we have been putting in default hyperparameter values, but in practice we would need tune these values to optimize the performance. Azure Machine Learning service provides many methods for tuning hyperparameters using different strategies.
#
# The first step is to choose the parameter space that we want to search. We have a few choices to make here :
#
# - **Parameter Sampling Method**: This is how we select the combinations of parameters to sample. Azure Machine Learning service offers [RandomParameterSampling](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.randomparametersampling?view=azure-ml-py), [GridParameterSampling](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.gridparametersampling?view=azure-ml-py), and [BayesianParameterSampling](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.bayesianparametersampling?view=azure-ml-py). We will use the `GridParameterSampling` method.
# - **Parameters To Search**: We will be searching for optimal combinations of `learning_rate` and `num_epochs`.
# - **Parameter Expressions**: This defines the [functions that can be used to describe a hyperparameter search space](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.parameter_expressions?view=azure-ml-py), which can be discrete or continuous. We will be using a `discrete set of choices`.
#
# The following code allows us to define these options.
# +
from azureml.train.hyperdrive import GridParameterSampling
from azureml.train.hyperdrive.parameter_expressions import choice
param_sampling = GridParameterSampling( {
'--learning_rate': choice(3e-5, 3e-4),
'--num_epochs': choice(3, 4)
}
)
# -
# The next step is to a define how we want to measure our performance. We do so by specifying two classes:
#
# - **[PrimaryMetricGoal](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.primarymetricgoal?view=azure-ml-py)**: We want to `MAXIMIZE` the `val_accuracy` that is logged in our training script.
# - **[BanditPolicy](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.banditpolicy?view=azure-ml-py)**: A policy for early termination so that jobs which don't show promising results will stop automatically.
# +
from azureml.train.hyperdrive import BanditPolicy
from azureml.train.hyperdrive import PrimaryMetricGoal
primary_metric_name='val_accuracy'
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE
early_termination_policy = BanditPolicy(slack_factor = 0.1, evaluation_interval=1, delay_evaluation=2)
# -
# We define an estimator as usual, but this time without the script parameters that we are planning to search.
estimator4 = TensorFlow(source_directory='./',
entry_script='train_logging.py',
compute_target=compute_target,
script_params = {
'--data_dir': azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length': 128,
'--batch_size': 32,
'--steps_per_epoch': 150,
'--export_dir':'./outputs/model',
},
framework_version='2.0',
use_gpu=True,
pip_packages=['transformers==2.0.0', 'azureml-dataprep[fuse,pandas]==1.1.22'])
# Finally, we add all our parameters in a [HyperDriveConfig](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.hyperdriveconfig?view=azure-ml-py) class and submit it as a run.
# +
from azureml.train.hyperdrive import HyperDriveConfig
hyperdrive_run_config = HyperDriveConfig(estimator=estimator4,
hyperparameter_sampling=param_sampling,
policy=early_termination_policy,
primary_metric_name=primary_metric_name,
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,
max_total_runs=10,
max_concurrent_runs=2)
run4 = experiment.submit(hyperdrive_run_config)
# -
# When we view the details of our run this time, we will see information and metrics for every run in our hyperparameter tuning.
from azureml.widgets import RunDetails
RunDetails(run4).show()
# We can retrieve the best run based on our defined metric.
best_run = run4.get_best_run_by_primary_metric()
# ## Register Model
#
# A registered [model](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model(class)?view=azure-ml-py) is a reference to the directory or file that make up your model. After registering a model, you and other people in your workspace can easily gain access to and deploy your model without having to run the training script again.
#
# We need to define the following parameters to register a model:
#
# - `model_name`: The name for your model. If the model name already exists in the workspace, it will create a new version for the model.
# - `model_path`: The path to where the model is stored. In our case, this was the *export_dir* defined in our estimators.
# - `description`: A description for the model.
#
# Let's register the best run from our hyperparameter tuning.
model = best_run.register_model(model_name='azure-service-classifier',
model_path='./outputs/model',
datasets=[('train, test, validation data', azure_dataset)],
description='BERT model for classifying azure services on stackoverflow posts.')
# We have registered the model with Dataset reference.
# * **ACTION**: Check dataset to model link in **Azure ML studio > Datasets tab > Azure Service Dataset**.
# In the [next tutorial](), we will perform inferencing on this model and deploy it to a web service.
| 1-Training/AzureServiceClassifier_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Что нейронные сети знают о наших лицах?
#
# 
#
# ### [Школа GoTo](https://goto.msk.ru)
# [<NAME>](https://github.com/TIXFeniks) <br>
# [<NAME>](https://tvorog.me)
#
# ### Подключим необходимые библиотеки
# %load_ext autoreload
# %autoreload 2
import numpy as np
from sklearn.model_selection import train_test_split
from helpers.lfw_dataset import load_lfw_dataset
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.neighbors import LSHForest
from IPython.display import display
from ipywidgets import widgets
from helpers.autoencoder import load_autoencoder
import skimage
from skimage import io
import matplotlib.patches as patches
from skimage import transform
#
# # Загрузим датасет
# Данные были уже загружены специально для вас. Ссылки (на всякий случай):
# - http://www.cs.columbia.edu/CAVE/databases/pubfig/download/lfw_attributes.txt
# - http://vis-www.cs.umass.edu/lfw/lfw-deepfunneled.tgz
# - http://vis-www.cs.umass.edu/lfw/lfw.tgz
# +
## Загружаем датасет
X, attr = load_lfw_dataset(use_raw=True,dimx=38,dimy=38)
# Представляем ввиде матрицы
m_attr = attr.as_matrix()
# Делаем непонятно что
X = X.astype('float32') / 255.0
# Смотрим размерность картинки
img_shape = X.shape[1:]
# Делим на трейн, тест
X_train, X_test, attr_train, attr_test = train_test_split(X, m_attr, test_size=0.1, random_state=42)
# -
# ## Посмотрим на имеющиеся изображения
# +
plt.title('sample image')
for i in range(6):
plt.subplot(2,3,i+1)
plt.imshow(X[i])
print("X shape:",X.shape)
print("attr shape:",attr.shape)
# -
# ## Модель
#
# Для манипуляций с лицами мы будем использовать автокодировщик. Эту модель мы учим сжимать картинку до вектора малой размерности и разжимать её обратно, теряя как можно меньше инфромации.
#
# <img src="https://blog.keras.io/img/ae/autoencoder_schema.jpg">
autoencoder = load_autoencoder(img_shape, weights_file= 'model_weights/deep_weights_64.pkl')
# gan4.pkl - это весело! Попробуй и другие .pkl файлы из папки с этой тетрадкой
# За время мастер-класса мы не успеем обучить модель с нуля, так что мы предобучили модель заранее, код на предыдущей клетке загружает модель.
#
# 
# ## Визуализация
#
# Используем нашу модель для того, чтобы сжать картинки. Затем разожмём их обратно
def visualize(img,autoencoder):
"""Draws original, encoded and decoded images"""
code = autoencoder.encode(img[None])[0]
reco = autoencoder.decode(code[None])[0]
plt.subplot(1,3,1)
plt.title("Original")
plt.imshow(img)
plt.subplot(1,3,2)
plt.title("Code")
plt.imshow(code.reshape([code.shape[-1]//8,-1]))
plt.subplot(1,3,3)
plt.title("Reconstructed")
plt.imshow(reco.clip(0,1))
plt.show()
for i in range(10):
img = X_test[i]
visualize(img,autoencoder)
# ## Поиск картинок с помощью автокодировщиков
# Нам удалось научить модель сжимать картинки и восстанавливать их неточно. С первого взгляда, решение этой задачи не приносит большой пользы, но, решив её, мы получили несколько интересных побочных эффектов.
#
# Первым полезным применением нашей модели является поиск схожих изображений по сгенерированным кодам картинок.
#
# Сперва закодируем наши изображения(не разкодируя обратно в картинки). Затем найдём близкие векторы-коды в нашей базе и покажем соответствующие им изображения как поисковую выдачу.
#
# Импользуем локально чувствительное хэширование(LSH) для ускорения процесса поиска. Для простоты, возьмём <a href="http://scikit-learn.org/0.18/modules/generated/sklearn.neighbors.LSHForest.html#sklearn.neighbors.LSHForest"> реализацию из scikit-learn</a>
# закодируем изображения
images = X_train
codes = autoencoder.encode(images, batch_size=10)
# build hashes
lshf = LSHForest(n_estimators=50).fit(codes)
# Функция нахождения схожих изображений
def get_similar(image, n_neighbors=5):
assert image.ndim==3,"image must be [batch,height,width,3]"
code = autoencoder.encode(image[None])
(distances,),(idx,) = lshf.kneighbors(code,n_neighbors=n_neighbors)
return distances,images[idx]
# + code_folding=[]
# Визуализация похожих изображений
def show_similar(image):
distances,neighbors = get_similar(image,n_neighbors=11)
plt.figure(figsize=[8,6])
plt.subplot(3,4,1)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom='off', # ticks along the bottom edge are off
top='off', # ticks along the top edge are off
labelbottom='off')
plt.imshow(image)
plt.title("Original image")
for i in range(11):
plt.subplot(3,4,i+2)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom='off', # ticks along the bottom edge are off
top='off', # ticks along the top edge are off
labelbottom='off')
plt.imshow(neighbors[i])
plt.title("Dist=%.3f"%distances[i])
plt.show()
# -
# улыбки
show_similar(X_test[1])
# Национальность
show_similar(X_test[499])
# очки
show_similar(X_test[63])
# ## Преобразование картинок
# Вторым, менее полезным, но не менее наглядным примером использования нашей модели будет нахождение промежуточных изображений при переходе от одной картинки к другой
N_INTERMEDIATE = 8
for _ in range(5):
image1,image2 = X_test[np.random.randint(0,len(X_test),size=2)]
code1, code2 = autoencoder.encode(np.stack([image1,image2]))
plt.figure(figsize=[10,4])
plt.subplot(1,N_INTERMEDIATE+2,1)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom='off', # ticks along the bottom edge are off
top='off', # ticks along the top edge are off
labelbottom='off')
plt.imshow(image1)
plt.title("original")
for i,a in enumerate(np.linspace(0,1,endpoint=True,num=N_INTERMEDIATE)):
output_code = code1*(1-a) + code2*(a)
output_image = autoencoder.decode(output_code[None])[0]
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom='off', # ticks along the bottom edge are off
top='off', # ticks along the top edge are off
labelbottom='off')
plt.subplot(1,N_INTERMEDIATE+2,i+2)
plt.imshow(output_image)
plt.title("a=%.2f"%a)
plt.subplot(1,N_INTERMEDIATE+2,N_INTERMEDIATE+1)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom='off', # ticks along the bottom edge are off
top='off', # ticks along the top edge are off
labelbottom='off')
plt.imshow(image2)
plt.title("target")
plt.show()
# ## Преобразуем изображения на основе побочных данных картинок
# Наша модель умеет восстанавливать изображение, по его закодированному вектору. Кодированный вектор несёт в себе много осмысленной информации. Мы можем манипулировать таким вектором, чтобы манипулировать хранимой в нём информацией.
#
# Помимо лиц, наш датасет имеет набор атрибутов - значений, характеризующих дополнительную инфрмацию о картинке
#
# Используем эту информацию для осмысленной манипуляцией над изображениями
# <img src="http://www.samyzaf.com/ML/nlp/word2vec2.png">
# пример представления связей объектов в векторном пространстве
#
# + run_control={"marked": false}
# какие есть атрибуты
attr.columns
# -
# закодируем изображения
encoded = autoencoder.encode(X)
# +
attribute = 'Smiling' # аттрибут, который мы будем менять
# Попробуй 'Smiling', 'Strong Nose-Mouth Lines','Male', 'Black', 'Asian', 'Attractive Woman', 'Big Nose', Mustache'
mean_featured = (encoded * attr[attribute].as_matrix()[:encoded.shape[0],None].astype('float32')).mean(axis = 0)
mean_code = encoded.mean(axis = 0)
featured_direction = mean_featured - mean_code
# -
attr['Mustache'].astype('float32').idxmax()
# выберем фото из датасета
def plot_morphing(factor, index):
#factor = 2 # насколько сильно мы меняем картинку
img = X[index]
code = encoded[index]
plt.subplot(1,3,1)
plt.imshow(img) # выводим оригинальное изображение
code_open = code + featured_direction*factor
plt.subplot(1,3,2)
plt.imshow(autoencoder.decode([code])[0])
plt.subplot(1,3,3)
plt.imshow(autoencoder.decode([code_open])[0]);
layout = widgets.Layout(width='100%', height='80px')
widgets.interact(plot_morphing, factor = widgets.FloatSlider(min=-10.0,max=10.,step= 0.1,layout=layout),
index = widgets.IntSlider(min=0,max=X.shape[0], step=1, layout=layout));
# ## загрузи свою картинку
# загрузи её в папку с тетрадкой и укажи путь к ней в поле внизу
_left = _right = _bottom = _top = 0.5 # значения по умолчанию
img = X[0]
def load_photo(path, left, bottom, right, top):
#try:
if True:
pic = io.imread(path)
global _left;global _right; global _bottom; global _top
_left = left
_right = right
_bottom = bottom
_top = top
left = int(pic.shape[1] * left)
top = int(pic.shape[0] * top)
right = left + int((pic.shape[1] - left) * right)
bottom = top + int((pic.shape[0] - top) * bottom)
cropped = pic[top:bottom,left:right]
cropped = transform.resize(cropped, img_shape, anti_aliasing=True)
#pic = skimage.util.(pic, img_shape)
fig,ax = plt.subplots(1)
# Display the image
ax.imshow(pic)
# Create a Rectangle patch
rect = patches.Rectangle((left,top),right - left, bottom - top,linewidth=1,edgecolor='r',facecolor='none')
# Add the patch to the Axes
ax.add_patch(rect)
plt.show()
plt.imshow(cropped)
global img;
img = cropped
#except:
# pass
widgets.interact(load_photo, path = "your_image.jpg",
left = widgets.FloatSlider(value = _left,min=0.,max=1.,step= 0.01,continuous_update=False,layout=layout),
right = widgets.FloatSlider(value = _right,min=0.01,max=1.,step= 0.01,continuous_update=False,layout=layout),
bottom = widgets.FloatSlider(value = _bottom,min=0,max=1.,step= 0.01,continuous_update=False,layout=layout),
top = widgets.FloatSlider(value = _top,min=0.01,max=1.,step= 0.01,continuous_update=False,layout=layout))
visualize(img, autoencoder)
<Попробуй сделать преобразования с лицом c твоей картинки>
plt.imshow(img)
# # Ура!
#
# ## Что делать дальше?
# 1. Можно посмотреть в эти же тетрадки дома и разобраться более детально. [Вот](https://github.com/tvorogme/digitalfest) репозиторий!
# 2. Прочитай [блог школы GoTo](https://habrahabr.ru/company/goto/blog/339050/), рассказывающий, с чего начинать изучать анализ данных и машинное обучение.
# 3. Когда ты наберёшься знаний и тебе захочется проверить свои силы, попробуй поучаствовать в соревнованиях на [kaggle](https://www.kaggle.com/competitions)
#
# 4. Когда ты научишься самостоятельно обучать нейронные сети, CPU для вычислений начнёт не хватать. Подумай о покупке GPU. Подойдёт любая CUDA-совместимая видеокарта, но чем мощнее - тем лучше
# <br><br>
# 
| Master Class 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction to Chinese NLP using Jieba
# Jieba is an useful library performing Chinese word segmentation.
#
# * ```pip install jieba```
# Jieba supports three segmentation methods:
#
# - Accurate Mode (精確模式),試圖將句子最精確地切開,適合文本分析: ```jieba.cut(sentence, cut_all=False)```
#
# - Full Mode (全模式),把句子中所有的可以成詞的詞語都掃描出來, 速度非常快,但是不能解決歧義: ```jieba.cut(sentence, cut_all=True)```
#
# - Search Engine Mode (搜索引擎模式),在精確模式的基礎上,對長詞再次切分,提高召回率,適合用於搜索引擎分詞: ```jieba.cut_for_search```
#
# Use Hidden Markov Model as default. Alter by changing the keyword ```HMM=False```.
import jieba
sentence = "獨立音樂需要大家一起來推廣,歡迎加入我們的行列!"
print ("Example:", sentence)
words = jieba.cut(sentence, cut_all=False)
print("Default/Accurate Mode:" + "/ ".join(words))
sentence_2 = "独立音乐需要大家一起来推广,欢迎加入我们的行列!"
words_2 = jieba.cut(sentence_2, cut_all=False)
l_2 = []
for x in words_2:
l_2.append(x)
"/".join(l_2)
# ### Compare results: Trad. vs Simp. Chinese
#
# There are a slight difference in the segmentation results between Traditional vs Simplified Chinese.
#
# This is because the original dictionary is built upon a simplified one.
# ## Chinese Lyrics Segmentation
#
# This example we will use ***Remeber Me***, the famous theme song from the movie **Coco**.
lyrics = '''請記住我 雖然再見必須說
請記住我 眼淚不要墜落
我雖然要離你遠去 你住在我心底
在每個分離的夜裡 為你唱一首歌
請記住我 雖然我要去遠方
請記住我 當聽見吉他的悲傷
這就是我跟你在一起唯一的憑據
直到我再次擁抱你 請記住我
你閉上眼睛音樂就會響起 不停的愛就永不會流失
你閉上眼睛音樂就會響起 要不停的愛
請記住我 雖然再見必須說
請記住我 眼淚不要墜落
我雖然要離你遠去 你住在我心底
在每個分離的夜裡 為你唱一首歌
請記住我 我即將會消失
請記住我 我們的愛不會消失
我用我的辦法跟你一起不離不棄
直到我再次擁抱你 請記住我'''
words_3 = jieba.cut(lyrics, cut_all=False)
for x in words_3:
print(x, end='/')
# Satisfied with the result or not? Evalute the result with the next session!
# ## Use Custom Dictionary:
# We can change the dictionary to a Traditional Chinese one in hopes of getting better performance.
#
# Download an example trad Chinese dict: https://github.com/fxsjy/jieba/raw/master/extra_dict/dict.txt.big
jieba.set_dictionary('dict.txt.big')
words_4 = jieba.cut(lyrics, cut_all=False)
for x in words_4:
print(x, end='/')
# ### Hola! Very small but important differences:
#
# * 憑/據/ is now grouped as 憑據
# * 閉上/眼睛/ is now grouped as 閉上眼睛
# * 再/見 is grouped as 再見
# * 我/要/ is grouped as 我要
# * /當聽/見/ is grouped as /當/聽見/
# * 就/會響/起/ is grouped as 就/會/響起/
# * /我/即/將會/消失/ is grouped as /我/即將/會/消失/
# * /我們/的/愛不會/消失/ is grouped as /我們/的/愛/不會/消失/
# * 不離/不棄/ is grouped as 不/離/不棄/ *(The only worsened example)*
# ## Load Custom Dictionary to add self-defined new words
#
# This function allows user adding new words on top of the default (base) dictionary. Although Jieba has HMM to identify new words, it is more accurate to input on our own.
# First, we have to create a file in the same format as the dictionary above. Each line with a word, word frequency, and POS tag.
#
# > POS Tag refers to https://blog.csdn.net/kevin_darkelf/article/details/39520881
#
# > Here, i means idiom
# !echo '不離不棄 2 i' | tee userdict.txt
# Using the command line, we got a file named userdict.txt with the new word we want.
#
# Load the dict using `jieba.load_userdict()`
jieba.load_userdict('userdict.txt')
words_5 = jieba.cut(lyrics, cut_all=False)
for x in words_5:
print(x, end='/')
# See? 不離不棄 is identified as a single group!
# ## Returns words with the Part of Speech
from jieba import posseg as pseg
words_pseg = pseg.cut('''你閉上眼睛音樂就會響起 不停的愛就永不會流失
你閉上眼睛音樂就會響起 要不停的愛''')
for x in words_pseg:
print(x)
# ## Returns words with Position
word_token = jieba.tokenize('''你閉上眼睛音樂就會響起 不停的愛就永不會流失
你閉上眼睛音樂就會響起 要不停的愛''')
for x in word_token:
print('word: %s \t\t start: %d \t\t end: %d' % (x[0],x[1],x[2]))
# ## Extracting keywords
#
# Built-in IDF corpus comes in handy!
import jieba.analyse
tags = jieba.analyse.extract_tags(lyrics, topK=10, withWeight=True)
tags
# Remember, this result is based on the trained idf comes along with the jieba library. In practice, we might want to use different idf in different semantics environment.
#
# If we want to learn the idf vector for specific corpus, try using scikit-learn `sklearn.feature_extraction.text.TfidfVectorizer` and then load it with `jieba.analyse.set_idf_path(file_name)`.
#
# Same function available for stop words: `jieba.analyse.set_stop_words(file_name)`
# * Keywords extraction using TextRank algorithm is available as well! (TextRank is an algorithm developed by Mihalcea & Tarau (2004))
jieba.analyse.textrank(lyrics, withWeight=True)
# Additional Reading Materials:
# 1. http://blogs.lessthandot.com/index.php/artificial-intelligence/automated-keyword-extraction-tf-idf-rake-and-textrank/
# References:
# 1. https://github.com/fxsjy/jieba
# 2. http://blog.fukuball.com/ru-he-shi-yong-jieba-jie-ba-zhong-wen-fen-ci-cheng-shi/
| jieba_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Randomized Benchmarking
#
# A randomized benchmarking (RB) experiment consists of the generation of random Clifford circuits on the given qubits such that the unitary computed by the circuits is the identity. After running the circuits, the number of shots resulting in an error (i.e. an output different than the ground state) are counted, and from this data one can infer error estimates for the quantum device, by calculating the Error Per Clifford.
# See [Qiskit Textbook](https://qiskit.org/textbook/ch-quantum-hardware/randomized-benchmarking.html) for an explanation on the RB method, which is based on Ref. [1, 2].
# +
import numpy as np
from qiskit_experiments.library import StandardRB, InterleavedRB
from qiskit_experiments.framework import ParallelExperiment
from qiskit_experiments.library.randomized_benchmarking import RBUtils
import qiskit.circuit.library as circuits
# For simulation
from qiskit.providers.aer import AerSimulator
from qiskit.test.mock import FakeParis
backend = AerSimulator.from_backend(FakeParis())
# -
# ## Standard RB experiment
#
# To run the RB experiment we need to provide the following RB parameters, in order to generate the RB circuits and run them on a backend:
#
#
# - `qubits`: The number of qubits or list of physical qubits for the experiment
#
# - `lengths`: A list of RB sequences lengths
#
# - `num_samples`: Number of samples to generate for each sequence length
#
# - `seed`: Seed or generator object for random number generation. If `None` then `default_rng` will be used
#
# - `full_sampling`: If `True` all Cliffords are independently sampled for all lengths. If `False` for sample of lengths longer sequences are constructed by appending additional Clifford samples to shorter sequences. The default is `False`
#
# The analysis results of the RB Experiment includes:
#
# - `EPC`: The estimated Error Per Clifford
#
# - `alpha`: The depolarizing parameter. The fitting function is $a \cdot \alpha^m + b$, where $m$ is the Clifford length
#
# - `EPG`: The Error Per Gate calculated from the EPC, only for 1-qubit or 2-qubit quantum gates (see Ref. [3])
# ### Running a 1-qubit RB experiment
# +
lengths = np.arange(1, 1000, 100)
num_samples = 10
seed = 1010
qubits = [0]
# Run an RB experiment on qubit 0
exp1 = StandardRB(qubits, lengths, num_samples=num_samples, seed=seed)
expdata1 = exp1.run(backend).block_for_results()
results1 = expdata1.analysis_results()
# View result data
display(expdata1.figure(0))
for result in results1:
print(result)
# -
# ### Running a 2-qubit RB experiment
#
# Running a 1-qubit RB experiment and a 2-qubit RB experiment, in order to calculate the gate error (EPG) of the `cx` gate:
# +
lengths = np.arange(1, 200, 30)
num_samples = 10
seed = 1010
qubits = (1,4)
# Run a 1-qubit RB expriment on qubits 1, 4 to determine the error-per-gate of 1-qubit gates
expdata_1q = {}
epg_1q = []
lengths_1_qubit = np.arange(1, 1000, 100)
for qubit in qubits:
exp = StandardRB([qubit], lengths_1_qubit, num_samples=num_samples, seed=seed)
expdata = exp.run(backend).block_for_results()
expdata_1q[qubit] = expdata
epg_1q += expdata.analysis_results()
# +
# Run an RB experiment on qubits 1, 4
exp2 = StandardRB(qubits, lengths, num_samples=num_samples, seed=seed)
# Use the EPG data of the 1-qubit runs to ensure correct 2-qubit EPG computation
exp2.set_analysis_options(epg_1_qubit=epg_1q)
# Run the 2-qubit experiment
expdata2 = exp2.run(backend).block_for_results()
# View result data
results2 = expdata2.analysis_results()
# -
# View result data
display(expdata2.figure(0))
for result in results2:
print(result)
# +
# Compare the computed EPG of the cx gate with the backend's recorded cx gate error:
expected_epg = RBUtils.get_error_dict_from_backend(backend, qubits)[(qubits, 'cx')]
exp2_epg = expdata2.analysis_results("EPG_cx").value
print("Backend's reported EPG of the cx gate:", expected_epg)
print("Experiment computed EPG of the cx gate:", exp2_epg)
# -
# ### Displaying the RB circuits
#
# Generating an example RB circuit:
# Run an RB experiment on qubit 0
exp = StandardRB(qubits=[0], lengths=[10], num_samples=1, seed=seed)
c = exp.circuits()[0]
# We transpile the circuit into the backend's basis gate set:
from qiskit import transpile
basis_gates = backend.configuration().basis_gates
print(transpile(c, basis_gates=basis_gates))
# ## Interleaved RB experiment
#
# Interleaved RB experiment is used to estimate the gate error of the interleaved gate (see Ref. [4]).
#
# In addition to the usual RB parameters, we also need to provide:
#
# - `interleaved_element`: the element to interleave, given either as a group element or as an instruction/circuit
#
# The analysis results of the RB Experiment includes the following:
#
# - `EPC`: The estimated error of the interleaved gate
#
# - `alpha` and `alpha_c`: The depolarizing parameters of the original and interleaved RB sequences respectively
#
# Extra analysis results include
#
# - `EPC_systematic_err`: The systematic error of the interleaved gate error (see Ref. [4])
#
# - `EPC_systematic_bounds`: The systematic error bounds of the interleaved gate error (see Ref. [4])
#
# ### Running a 1-qubit interleaved RB experiment
# +
lengths = np.arange(1, 1000, 100)
num_samples = 10
seed = 1010
qubits = [0]
# Run an Interleaved RB experiment on qubit 0
# The interleaved gate is the x gate
int_exp1 = InterleavedRB(
circuits.XGate(), qubits, lengths, num_samples=num_samples, seed=seed)
# Run
int_expdata1 = int_exp1.run(backend).block_for_results()
int_results1 = int_expdata1.analysis_results()
# -
# View result data
display(int_expdata1.figure(0))
for result in int_results1:
print(result)
# ### Running a 2-qubit interleaved RB experiment
# +
lengths = np.arange(1, 200, 30)
num_samples = 10
seed = 1010
qubits = [1,4]
# Run an Interleaved RB experiment on qubits 1, 4
# The interleaved gate is the cx gate
int_exp2 = InterleavedRB(
circuits.CXGate(), qubits, lengths, num_samples=num_samples, seed=seed)
# Run
int_expdata2 = int_exp2.run(backend).block_for_results()
int_results2 = int_expdata2.analysis_results()
# -
# View result data
display(int_expdata2.figure(0))
for result in int_results2:
print(result)
# ## Running a simultaneous RB experiment
#
# We use `ParallelExperiment` to run the RB experiment simultaneously on different qubits (see Ref. [5])
# +
lengths = np.arange(1, 1000, 100)
num_samples = 10
seed = 1010
qubits = range(3)
# Run a parallel 1-qubit RB experiment on qubits 0, 1, 2
exps = [StandardRB([i], lengths, num_samples=num_samples, seed=seed + i)
for i in qubits]
par_exp = ParallelExperiment(exps)
par_expdata = par_exp.run(backend).block_for_results()
par_results = par_expdata.analysis_results()
# View result data
for result in par_results:
print(result)
print("\nextra:")
print(result.extra)
# -
# ### Viewing sub experiment data
#
# The experiment data returned from a batched experiment also contains individual experiment data for each sub experiment which can be accessed using `child_data`
# Print sub-experiment data
for i, sub_data in enumerate(par_expdata.child_data):
print(f"Component experiment {i}")
display(sub_data.figure(0))
for result in sub_data.analysis_results():
print(result)
# ## References
#
# [1] <NAME>, <NAME>, and <NAME>, *Robust randomized benchmarking of quantum processes*, https://arxiv.org/pdf/1009.3639
#
# [2] <NAME>, <NAME>, and <NAME>, *Characterizing Quantum Gates via Randomized Benchmarking*, https://arxiv.org/pdf/1109.6887
#
# [3] <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, *Three Qubit Randomized Benchmarking*, https://arxiv.org/pdf/1712.06550
#
# [4] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>,
# *Efficient measurement of quantum gate error by interleaved randomized benchmarking*,
# https://arxiv.org/pdf/1203.4550
#
# [5] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, *Characterization of addressability by simultaneous randomized benchmarking*, https://arxiv.org/pdf/1204.6308
#
#
import qiskit.tools.jupyter
# %qiskit_copyright
| docs/tutorials/randomized_benchmarking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Project: Investigating No-Show Appointments in Brazil
#
# ## Table of Contents
# <ul>
# <li><a href="#intro">Introduction</a></li>
# <li><a href="#wrangling">Data Wrangling</a></li>
# <li><a href="#eda">Exploratory Data Analysis</a></li>
# <li><a href="#conclusions">Conclusions</a></li>
# </ul>
# <a id='intro'></a>
# ## Introduction
#
# > This data set represents appointments that patients did or did not show up for in Brazil. This categorizes patients by multiple conditions, ages, gender and even if they receive aid.
# >
# > For this data set, there will be three questions addressed. First, it will address whether or not alcoholism leads to more missed appointments. Second, it will determine if any age groups miss more appointments than others. Finally, it will look at differences in gender.
# > This section includes: loading the packages needed for the analysis.
# +
#import the packages and assign an alias:
import unicodecsv
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
#Show graphs inline
# %matplotlib inline
# -
# <a id='wrangling'></a>
# ## Data Wrangling
#
# > This section of the report will load the data, check for cleanliness, trim and clean the dataset.
#
# +
#Open and Read the file.
noshow_filename = '/Users/knico/OneDrive/Documents/Python Scripts/noshowappointments-kagglev2-may-2016.csv'
def open_files(my_name):
with open(my_name , 'rb') as f:
reader = unicodecsv.DictReader(f)
return list(reader)
#use the open_files function to opent the file.
no_show_appointments = open_files(noshow_filename)
# Print a line of the no show appointment file.
print('No Show Appointments:', no_show_appointments[0:5])
# -
# Create a pandas dataframe and view the first 3 lines.
#have Pandas return the first 3 rows.
df_noshow = pd.read_csv(noshow_filename)
print(df_noshow.head(3))
# ### Data Cleaning
#
# > 1) Review then Update the Data Types <br>
# > 2) Find and clean bad data.<br>
#
# <ul>
# <li><a href="#DataTypes">Data Types</a></li>
# <li><a href="#BadData">Find and Clean Bad Data</a></li>
# </ul>
#
#
# <a id='DataTypes'></a>
# #### Data Types
# >This section the data types are reviewed to see how they converted. <br>
# >There are some fields that will be used that are objects or numbers (int, float) and need to be converted to strings. <br>
# >When looking at the first few lines of data, the appointment ID and the Patient ID are changing to scientific formats. These items will not be used for any math. They will be converted to strings so they look neater.
# Display the data types and columns
df_noshow.info()
# This function will convert the data types
def change_var_type(col_as_string, ctype):
if ctype == 'string':
new_type = df_noshow[col_as_string].astype("|S")
if ctype == 'number':
new_type = df_noshow[col_as_string].astype(int)
return new_type
# +
# This will convert the first few object, integer and float variables to string.
df_noshow["PatientId"] = change_var_type("PatientId",'string')
df_noshow["AppointmentID"] = change_var_type("AppointmentID",'string')
df_noshow["Gender"] = change_var_type("Gender",'string')
# -
# The neighborhood is more difficult to convert. First, the non-ascii characters need to be removed.
#Once removed, then the type can be changed.
df_noshow.Neighbourhood.replace({r'[^\x00-\x7F]+':''}, regex=True, inplace=True)
df_noshow["Neighbourhood"] = change_var_type("Neighbourhood",'string')
# +
#A new column will be generated on Noshow as an integer so that is can be used for correlations later.
df_noshow["Noshow2"] = df_noshow["No-show"].map({'Yes': '1', 'No': '0'})
df_noshow["Noshow2"] = pd.to_numeric(df_noshow["Noshow2"])
print(df_noshow.loc[1])
print(df_noshow.head())
# -
#The name No-show is causing issues because of the dash. This will rename the column in the existing dataframe.
#Then object type can be change to string, and clean up can be done.
#Once complete, we will run info to verify it is complete.
df_noshow.rename(columns = {'No-show' : 'Noshow' , 'Hipertension':'Hypertension', 'Handcap':'Handicap'}, inplace = True )
df_noshow["Noshow"] = change_var_type("Noshow", 'string')
df_noshow.info()
# <a id='BadData'></a>
# ### Find and Clean Bad Data
# > This section will review the data to find and clean up data that is not valid. <br>
# > First, a review of some statistics will help to find the issue.
# +
# Review some statistics on the data frame.
df_noshow.describe()
# -
# > There are no null values to be concerned with, but the minimum age is listed as a -1. <br>
# > Because the data set is large, it makes more sense to delete it than to put in an average value, or guess the typo.<br>
# > Then display the minimum age again to make sure they are all 0 or over.<br>
# +
#Loop through the values and make sure that any values less than 0 will be deleted. This will alter the original data frame.
for x in df_noshow.index:
if df_noshow.loc[x, "Age"] < 0:
df_noshow.drop(x, inplace=True)
print("Minimum Age: " , df_noshow["Age"].min())
# -
# <a id='eda'></a>
# ## Exploratory Data Analysis
#
# > This section will explore the data set and look for insights in the data. Because this has not been tested with a scientific experiment, this section cannot state causation between variables. It will explore three questions and show the correlations and more ideas to explore.
#
# #### Correlation and the Variables
# >First, let's explore correlation between different variables. The independent variable is if the patient showed up for their appointment or not. The dependent variables include alcoholism, gender, various diseases and conditions, and age. <br>
# >The correlation function can be used to get a quick look at all the numberical variables.
# >From the Statology website listed below, this explains the strength of the relationships in the tables and heat maps in the next two sections.
# > If we take the absolute value less than .25 is considered to have no relationship.<br>
# > A weak relationship would be less than .5, but greater than .25 <br>
# > A moderate relationship should be between .5 and .75 <br>
# > A strong relationship will be over .75 <br>
# https://www.statology.org/what-is-a-strong-correlation/
#
# > For the correlation function, a 1 is a perfect relationship, and it will be a one when compared with itself.
# Show the correlation between variables.
df_noshow.corr()
# ### Research Question 1: Alcoholism
# #### Are Alcoholics More Likely to Miss Their Appointments?
# >Per the Valley Hope website listed below, two symptoms of being an alcoholic are "Drinking at inappropriate times or alone."
# > and "Drinking with the intention of getting drunk."<br>
# >Because someone that suffers from alcoholism may be drinking at inappropriate times, and could be at a level of intoxication, it could be unsafe for them to drive to appointments. They may also feel ashamed, or be worried about being judged by the Dr. This will use a heat map to quickly consume the data listed in the table above.
#
# https://valleyhope.org/am-i-an-alcoholic/?gclid=Cj0KCQjwtMCKBhDAARIsAG-2Eu-fbNnCXP2UDs3qlLtZmKwl6isq0sZilas442qdDCb1pMOofD1rTQQaAklFEALw_wcB
# +
#Research question 1- Heat map to show correlation between alcoholism.
f, ax = plt.subplots(figsize= (10,8))
corr = df_noshow.corr()
sns.heatmap(corr, mask=np.zeros_like(corr, dtype= bool) , cmap= sns.diverging_palette(220, 10, as_cmap = True), square = True, ax= ax)
plt.title("Correlation on Appointment Data")
plt.ylabel("Patient Variables")
plt.xlabel("Patient Variables")
plt.show()
# -
# >Surprisingly, there is no relationship between not showing up for appointments and suffering from alcoholism.
# >There does seem to be a weak to modeate relationship between hypertension and diabetes, and a moderate relationship between age and hypertension. These could be investigated at another time.
# ### Research Question 2: Age
# #### What age ranges are most likely to miss appointments?
# >Those very young or very old would have to rely on others for rides. There are other differences between age groups as well, so let's explore the data to see which age groups tend to miss the most appointments.
# +
# Make a new dataframe consisting of only the no show appointments.
i = 0
df_noshow_appointments = pd.DataFrame()
appointment_list = [1]
df_noshow_appointments = df_noshow[df_noshow.Noshow2.isin(appointment_list)]
print("Total Missed Appointments: ", len(df_noshow_appointments))
# +
#Histogram to show only missed appointments.
df_noshow_appointments.plot(kind='hist', y = 'Age', x='Noshow', bins = 11)
plt.title('Age Ranges of Missed Appointments')
plt.xlabel("Age")
plt.show()
#Histogram to show ages of all appointments mage
df_noshow.plot(kind='hist', y = 'Age', x ='Noshow', bins = 11)
plt.title('Age Ranges of All Appointments')
plt.xlabel("Age")
plt.show()
# -
# >The first histogram shows that there are higher frequencies of missed appointments for all age buckets for patients 0-10 and 20-30. When looking at the entire dataset by age, it shows that children ages 0-10 have a higher number of appointments overall, so it makes sense that they would miss more. However, the 20-30 group, while on the larger size, seems to have a large amount of missed appointments in comparison with total appointments. This should be explored further, but the doctor's staff could set up an experiment to see if additional SMS message warnings would help to have less no shows.
# ### Research Question 3: Gender
# #### Does one gender miss more appointments than the other?
# > Finally, this will explore whether or not one gender is canceling more appointments than the other. This will use a bar chart to count the missed appointments by gender.
# +
#Female vs. Male appointments
Genderindexes_NoShow = df_noshow_appointments['Gender'].value_counts().index.tolist()
Gendercount_NoShow = df_noshow_appointments['Gender'].value_counts().values.tolist()
plt.bar(Genderindexes_NoShow, Gendercount_NoShow, color='r', label='Missed Appointments')
plt.title("Appointments Missed by Gender")
plt.xlabel("Gender")
plt.ylabel("Number of Missed Appointments")
plt.legend()
plt.show()
Genderindexes_All = df_noshow['Gender'].value_counts().index.tolist()
Gendercount_All = df_noshow['Gender'].value_counts().values.tolist()
print('Total Appointments for Females: ' , Gendercount_All[0])
print('Total Appointments for Males: ' , Gendercount_All[1])
def ratio_Missed(indexnum):
my_ratio = Gendercount_NoShow[indexnum] / Gendercount_All[indexnum]
return my_ratio
print("Ratio for Missed Appointment Females: " , ratio_Missed(0))
print("Ratio for Missed Appointment Males: " , ratio_Missed(1))
# -
# >Although females appear to miss a lot more appointments than men, when comparing the amount of missed appointments to total appointments, they are both missing about one in five appointments, with men just a small amount less.
# <a id='conclusions'></a>
# ## Conclusions
#
# > This dataset was imported into a pandas dataset and then cleaned. The cleansing involved removing bad data in the form of a negative age, a symbol in a column, and non-ascii characters. It also involved changing the data types and formatting the no show column to be able to demonstrate correlations. <br>
# > The data was explored for correlations, and no strong correlations were found between the variables and missing an appointment. People with alcoholism do not miss a significant amount of appointments less or more than people without that condition. Gender seems to have no correlation with missed appointments, though in this dataset, more females book appointments than males. In addition, while age did not have a correlation to missed appointments, the doctors' office may want to experiment on ways to target the 20-30 age range, since they missed a larger percentage of their appointments. Causation cannot be determined at this time.
# >
# ### Limitations
# >Some of the limitations included were that the younger ages and oldest ages could not bring themselves to the appointment. Another analysis could be done eliminating those ranges, or also considering the information on the person responsible for bringing them.
# ### Sites and Materials Referenced:
#
# >W3 Schools <br>
# >"Python All-In-One for Dummies" <br>
# https://www.kite.com/python/answers/how-to-convert-a-column-of-objects-to-strings-in-a-pandas-dataframe-in-python
# https://datatofish.com/string-to-integer-dataframe/
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.map.#
# https://www.geeksforgeeks.org/how-to-rename-columns-in-pandas-dataframe/
# https://www.youtube.com/watch?v=71FoexyFCXE
# https://www.askpython.com/python/examples/subset-a-dataframe
# https://newbedev.com/remove-non-ascii-characters-from-pandas-column
# https://www.statology.org/what-is-a-strong-correlation/
# https://www.python-graph-gallery.com/4-add-title-and-axis-label
# https://www.geeksforgeeks.org/create-a-stacked-bar-plot-in-matplotlib/
# https://valleyhope.org/am-i-an-alcoholic/?gclid=Cj0KCQjwtMCKBhDAARIsAG-2Eu-fbNnCXP2UDs3qlLtZmKwl6isq0sZilas442qdDCb1pMOofD1rTQQaAklFEALw_wcB
#
# ###### <NAME> 9/26/21
| KLYNCH Final No Show Appointments 2021-09-29.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Python for Research Homework: Week 3, Case Study 1
#
# A cipher is a secret code for a language. In this case study, we will explore a cipher that is reported by contemporary Greek historians to have been used by <NAME>ar to send secret messages to generals during times of war.
# ### Exercise 1
#
# A cipher is a secret code for a language. In this case study, we will explore a cipher that is reported by contemporary Greek historians to have been used by <NAME> to send secret messages to generals during times of war.
#
# The Caesar cipher shifts each letter of a message to another letter in the alphabet located a fixed distance from the original letter. If our encryption key were `1`, we would shift `h` to the next letter `i`, `i` to the next letter `j`, and so on. If we reach the end of the alphabet, which for us is the space character, we simply loop back to `a`. To decode the message, we make a similar shift, except we move the same number of steps backwards in the alphabet.
#
# Over the next five exercises, we will create our own Caesar cipher, as well as a message decoder for this cipher. In this exercise, we will define the alphabet used in the cipher.
#
# #### Instructions
# - The `string` library has been imported. Create a string called `alphabet` consisting of the space character `' '` followed by (concatenated with) the lowercase letters. Note that we're only using the lowercase letters in this exercise.
import string
# write your code here!
alphabet =' ' + string.ascii_lowercase
print(alphabet)
# ### Exercise 2
#
# In this exercise, we will define a dictionary that specifies the index of each character in `alphabet`.
#
# #### Instructions
# - `alphabet` has already defined in the last exercise. Create a dictionary with keys consisting of the characters in alphabet and values consisting of the numbers from 0 to 26.
# - Store this as `positions`.
# write your code here!
positions = {}
for i in range(len(alphabet)):
positions[alphabet[i]] = i
print(positions)
# ### Exercise 3
#
# In this exercise, we will encode a message with a Caesar cipher.
#
# #### Instructions
#
# - `alphabet` and `positions` have already been defined in previous exercises. Use `positions` to create an encoded message based on message where each character in message has been shifted forward by 1 position, as defined by positions.
# - **Note that you can ensure the result remains within 0-26 using result % 27**
# - Store this as `encoded_message`.
# +
message = "hi my name is caesar"
# write your code here!
encoded_message = ''
for char in message:
for key,values in positions.items():
if values == ((positions[char]+ 1)%27):
encoded_message += key
print(encoded_message)
# -
# ### Exercise 4
#
# In this exercise, we will define a function that encodes a message with any given encryption key.
#
# #### Instructions
# - `alphabet`, `position` and `message` remain defined from previous exercises. Define a function `encoding` that takes a message as input as well as an int encryption key `key` to encode a message with the Caesar cipher by shifting each letter in message by key positions.
# - Your function should return a string consisting of these encoded letters.
# - Use `encoding` to encode message using `key = 3` and save the result as `encoded_message`.
# Print `encoded_message`.
# +
# write your code here
message = "hi my name is caesar"
def encoding(message, ke):
encoded_message =''
for c in message:
for key,values in positions.items():
if values == (positions[c] + ke)%27:
encoded_message += key
return encoded_message
encoded_message = encoding(message,3)
print(encoded_message)
# -
# ### Exercise 5
#
# In this exercise, we will decode an encoded message.
#
# #### Instructions
# - Use `encoding` to decode `encoded_message`.
# - Store your encoded message as `decoded_message`.
# - Print `decoded_message`. Does this recover your original message?
# +
# write your code here!
def decoding(encoded_message , ke):
decoded_message = ''
for i in encoded_message:
for key,values in positions.items():
if values == (positions[i] - ke) %27:
decoded_message +=key
return decoded_message
decoded_message = decoding(encoded_message,3)
print(decoded_message)
# -
| HW3-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table width="100%"><tr style="background-color:white;">
# <td style="text-align:left;padding:0px;width:142px'">
# <a href="https://qworld.net" target="_blank">
# <img src="qworld/images/QWorld.png"></a></td>
# <td width="*"> </td>
# <!-- ############################################# -->
# <td style="padding:0px;width:90px;">
# <img align="right" src="qworld/images/follow_us.png" height="40px"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://twitter.com/QWorld19" target="_blank">
# <img align="right" src="qworld/images/Twitter.png" width="40px"></a> </td>
# <td style="padding:0px;width:5px;"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://www.facebook.com/qworld19/" target="_blank">
# <img align="right" src="qworld/images/Fb.png"></a></td>
# <td style="padding:0px;width:5px;"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://www.linkedin.com/company/qworld19" target="_blank">
# <img align="right" src="qworld/images/LinkedIn.png"></a></td>
# <td style="padding:0px;width:5px;"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://youtube.com/QWorld19?sub_confirmation=1" target="_blank">
# <img align="right" src="qworld/images/YT.png"></a></td>
# <!-- ############################################# -->
# <td style="padding:0px;width:60px;">
# <img align="right" src="qworld/images/join.png" height="40px"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://join.slack.com/t/qworldworkspace/shared_invite/zt-lrkilojl-HPsirAdvGoDHR8NhJpo75A"
# target="_blank">
# <img align="right" src="qworld/images/Slack.png"></a></td>
# <!-- ############################################# -->
# <td style="padding:0px;width:72px;">
# <img align="right" src="qworld/images/w3.png" height="40px"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://qworld.net" target="_blank">
# <img align="right" src="qworld/images/www.png"></a></td>
# </tr></table>
# <h2 align="left" style="color: #cd7f32;"> Before Workshop </h2>
#
# 1. Check the first notebook below to complete the installation of Qiskit and also test your system.
#
# 1. Complete all notebooks listed under section "Python review" if you are not familiar with Python.
#
# 1. Review all notebooks listed under section "Basic math" before the workshop starts.
# ### Installation and Test
#
# <a href="quantum-with-qiskit/Q04_Qiskit_installation_and_test.ipynb" target="_blank">Qiskit installation and test</a>
#
# ### Python review
#
# [Jupyter notebooks](python/Python02_Into_Notebooks.ipynb) |
# [Variables](python/Python08_Basics_Variables.ipynb) |
# [Loops](python/Python12_Basics_Loops.ipynb) |
# [Conditionals](python/Python16_Basics_Conditionals.ipynb) |
# [Lists](python/Python20_Basics_Lists.ipynb) |
# [Python Reference](python/Python04_Quick_Reference.ipynb) |
# [Drawing Reference](python/Python06_Drawing.ipynb)
#
# ### Basic math
#
# [Vectors](math/Math20_Vectors.ipynb) |
# [Dot Product](math/Math24_Dot_Product.ipynb) |
# [Matrices](math/Math28_Matrices.ipynb) |
# [Tensor Product](math/Math32_Tensor_Product.ipynb) |
# [Exercises](math/Exercises_Basic_Math.ipynb)
| Bronze/before-workshop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# (examples)=
# # RST Conversion Gallery
#
# ```{note}
# A minimum configured sphinx repo is available [here](https://github.com/QuantEcon/sphinx-tojupyter.minimal)
# which generates a [sample notebook](https://github.com/QuantEcon/sphinx-tojupyter.minimal#simple_notebookrst)
# ```
#
# ```{contents} Examples
# ```
#
# The test suite, located [here](https://github.com/QuantEcon/sphinx-tojupyter/tree/master/tests)
# provides examples of conversions between RST and the Jupyter notebook which form the test cases
# for this extension. It can be a useful resource to check how elements are converted if they are not
# contained in this gallery.
#
# ## code-blocks
#
# The following code in the **.rst** file
#
# ```{code-block} rst
# Code blocks
# -----------
#
# This is a collection to test various code-blocks
#
# This is a **.. code::** directive
#
# .. code:: python
#
# this = 'is a code block'
# x = 1
# no = 'really!'
# p = argwhere(x == 2)
#
# This is another **.. code::** directive
#
# .. code:: python
#
# from pylab import linspace
# t = linspace(0, 1)
# x = t**2
#
# This is a **::** directive
#
# ::
#
# from pylab import *
# x = logspace(0, 1)
# y = x**2
# figure()
# plot(x, y)
# show()
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/code-block.png
#
# ```
#
# ## images and figures
#
# The following code in the .rst file
#
# ```{code-block} rst
# Images
# ======
#
# Collection of tests for **.. image::** and **.. figure::** directives
#
# Image
# -----
#
# `Docutils Reference <http://docutils.sourceforge.net/docs/ref/rst/directives.html#images>`__
#
# Most basic image directive
#
# .. image:: _static/hood.jpg
#
# A scaled down version with 25 % width
#
# .. image:: _static/hood.jpg
# :width: 25 %
#
# A height of 50px
#
# .. image:: _static/hood.jpg
# :height: 50px
#
# Figure
# ------
#
# `Docutils Reference <http://docutils.sourceforge.net/docs/ref/rst/directives.html#figure>`__
#
# Testing the **.. figure::** directive
#
# .. figure:: _static/hood.jpg
# :scale: 50 %
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/images.png
#
# ```
#
# ```{image} img/figure.png
#
# ```
#
# ```{warning}
# if `jupyter_images_markdown = True` then the `:scale:`, `:height:`
# and `:width:` attributes will be ignored.
# ```
#
# ## jupyter-directive
#
# The following code in the .rst file
#
# ```{code-block} rst
# Jupyter Directive
# =================
#
# This is a set of tests related to the Jupyter directive
#
#
# The following jupyter directive with cell-break option should
# split this text and the text that follows into different IN
# blocks in the notebook
#
# .. jupyter::
# :cell-break:
#
# This text should follow in a separate cell.
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/jupyter-directive.png
#
# ```
#
# ## links
#
# The following code in the .rst file
#
# ```{code-block} rst
# .. _links:
#
# Links
# -----
#
# Links are generated as markdown references to jump between notebooks and
# the sphinx link machinery is employed to track links across documents.
#
# An external link to another `notebook (as full file) <links_target.ipynb>`_
#
# This is a paragraph that contains `a google hyperlink`_.
#
# .. _a google hyperlink: https://google.com.au
#
# - An inline reference to :ref:`another document <links_target>`
#
# Special Cases
# -------------
#
# The following link has ( and ) contained within them that doesn't render nicely in markdown. In this case the extension will substitute ( with `%28` and ) with `%29`
#
# Thinking back to the mathematical motivation, a `Field <https://en.wikipedia.org/wiki/Field_\(mathematics\)>`_ is an `Ring` with a few additional properties
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/links.png
#
# ```
#
# ## math
#
# The following code in the .rst file
#
# ```{code-block} rst
# Math
# ----
#
# Inline maths with inline role: :math:`x^3+\frac{1+\sqrt{2}}{\pi}`
#
# Inline maths using dollar signs (not supported yet): $x^3+\frac{1+\sqrt{2}}{\pi}$ as the
# backslashes are removed.
#
# .. math::
#
# x^3+\frac{1+\sqrt{2}}{\pi}
#
# check math with some more advanced LaTeX, previously reported as an issue.
#
# .. math::
#
# \mathbb P\{z = v \mid x \}
# = \begin{cases}
# f_0(v) & \mbox{if } x = x_0, \\
# f_1(v) & \mbox{if } x = x_1
# \end{cases}
#
# and labeled test cases
#
# .. math::
# :label: firsteq
#
# \mathbb P\{z = v \mid x \}
# = \begin{cases}
# f_0(v) & \mbox{if } x = x_0, \\
# f_1(v) & \mbox{if } x = x_1
# \end{cases}
#
# Further Inline
# --------------
#
# A continuation Ramsey planner at :math:`t \geq 1` takes
# :math:`(x_{t-1}, s_{t-1}) = (x_-, s_-)` as given and before
# :math:`s` is realized chooses
# :math:`(n_t(s_t), x_t(s_t)) = (n(s), x(s))` for :math:`s \in {\cal S}`
#
# Referenced Math
# ---------------
#
# Simple test case with reference in text
#
# .. math::
# :label: test
#
# v = p + \beta v
#
# this is a reference to :eq:`test` which is the above equation
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/math.png
#
# ```
#
# ## block-quote
#
# The following code in the .rst file
#
# ```{code-block} rst
# Quote
# -----
#
# This is some text
#
# This is a quote!
#
# and this is not
#
# Epigraph
# --------
#
# An epigraph is a special block-quote node
#
# .. epigraph::
#
# "Debugging is twice as hard as writing the code in the first place.
# Therefore, if you write the code as cleverly as possible, you are, by definition,
# not smart enough to debug it."
#
# -- <NAME>
#
# and one that is technically malformed
#
# .. epigraph::
#
# "Debugging is twice as hard as writing the code in the first place.
# Therefore, if you write the code as cleverly as possible, you are, by definition,
# not smart enough to debug it." -- <NAME>
#
# with some final text
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/quote.png
#
# ```
#
# ## slides
#
# The following code in the .rst file
#
# ```{code-block} rst
# Slide option activated
# ----------------------
#
# .. jupyter::
# :slide: enable
#
# This is a collection of different types of cells where the toolbar: Slideshow has been activated
#
# .. jupyter::
# :cell-break:
# :slide-type: subslide
#
# The idea is that eventually we will assign a type (*slide*, *subslide*, *skip*, *note*) for each one. We used our **jupyter** directive to break the markdown cell into two different cells.
#
#
# .. code:: python3
#
# import numpy as np
#
# x = np.linspace(0, 1, 5)
# y = np.sin(4 * np.pi * x) * np.exp(-5 * x)
#
# print(y)
#
# .. code:: python3
#
# import numpy as np
#
# z = np.cos(3 * np.pi * x) * np.exp(-2 * x)
# w = z*y
#
# print(w)
#
# Math
# ++++
#
# The previous function was
#
# .. math:: f(x)=\sin(4\pi x)\cos(4\pi x)e^{-7x}
#
#
# .. jupyter::
# :cell-break:
# :slide-type: fragment
#
# We can also include the figures from some folder
#
#
# .. figure:: _static/hood.jpg
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/slides.png
#
# ```
#
# ## footnotes
#
# The following code in the .rst file
#
# ```{code-block} rst
# Rubric
# ======
#
# Define the government's one-period loss function [#f1]_
#
# .. math::
# :label: target
#
# r(y, u) = y' R y + u' Q u
#
#
# History dependence has two sources: (a) the government's ability to commit [#f2]_ to a sequence of rules at time :math:`0`
#
#
# .. rubric:: Footnotes
#
# .. [#f1] The problem assumes that there are no cross products between states and controls in the return function. A simple transformation converts a problem whose return function has cross products into an equivalent problem that has no cross products.
#
# .. [#f2] The government would make different choices were it to choose sequentially, that is, were it to select its time :math:`t` action at time :math:`t`.
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/footnotes.png
#
# ```
#
# ## solutions
#
# The following code in the .rst file
#
# ```{code-block} rst
# Notebook without solutions
# ==========================
#
# The idea is with the use of classes, we can decide whether to show or not the solutions
# of a particular lecture, creating two different types of jupyter notebooks. For now it only
# works with *code blocks*, you have to include **:class: solution**, and set in the conf.py file
# *jupyter_drop_solutions=True*.
#
#
# Here is a small example
#
# Question 1
# ----------
#
# Plot the area under the curve
#
# .. math::
#
# f(x)=\sin(4\pi x) exp(-5x)
#
# when :math:`x \in [0,1]`
#
# .. code-block:: python3
# :class: solution
#
# import numpy as np
# import matplotlib.pyplot as plt
#
# x = np.linspace(0, 1, 500)
# y = np.sin(4 * np.pi * x) * np.exp(-5 * x)
#
# fig, ax = plt.subplots()
#
# ax.fill(x, y, zorder=10)
# ax.grid(True, zorder=5)
# plt.show()
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/solutions.png
#
# ```
#
# ```{todo}
# ## Todo
#
# Currently generating the two sets of notebooks requires two separate
# runs of sphinx which is incovenient. It would be better to develop a set
# of notebooks without solutions (as Default) and a set of notebooks with
# solutions in a subdir.
#
# ```
#
# ## tables
#
# Basic table support is provided by this extension.
#
# ```{note}
# Complex tables are **not** currently supported.
# See Issue [#54]([https://github.com/QuantEcon/sphinx-tojupyter/issues/54](https://github.com/QuantEcon/sphinx-tojupyter/issues/54))
# ```
#
# The following code in the .rst file
#
# ```{code-block} rst
# Table
# =====
#
# These tables are from the `RST specification <http://docutils.sourceforge.net/docs/ref/rst/restructuredtext.html#grid-tables>`__:
#
# Grid Tables
# -----------
#
# A simple rst table with header
#
# +------+------+
# | C1 | C2 |
# +======+======+
# | a | b |
# +------+------+
# | c | d |
# +------+------+
#
# **Note:** Tables without a header are currently not supported as markdown does
# not support tables without headers.
#
#
# Simple Tables
# -------------
#
# ===== ===== =======
# A B A and B
# ===== ===== =======
# False False False
# True False False
# False True False
# True True True
# ===== ===== =======
#
# Directive Table Types
# ---------------------
#
# These table types are provided by `sphinx docs <http://www.sphinx-doc.org/en/master/rest.html#directives>`__
#
#
# List Table directive
# ~~~~~~~~~~~~~~~~~~~~
#
# .. list-table:: Frozen Delights!
# :widths: 15 10 30
# :header-rows: 1
#
# * - Treat
# - Quantity
# - Description
# * - Albatross
# - 2.99
# - On a stick!
# * - Crunchy Frog
# - 1.49
# - If we took the bones out, it wouldn't be crunchy, now would it?
# * - Gannet Ripple
# - 1.99
# - On a stick!
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/tables.png
#
# ```
#
# ## tests
#
# The following code in the .rst file
#
# ```{code-block} rst
# Notebook without Tests
# ======================
#
# This is an almost exact analogue to the solutions class. The idea is that we can include test blocks using **:class: test** that we can toggle on or off with *jupyter_drop_tests = True*. A primary use case is for regression testing for the 0.6 => 1.0 port, which we will not want to show to the end user.
#
# Here is a small example:
#
# Question 1
# ------------
#
# .. code-block:: julia
#
# x = 3
# foo = n -> (x -> x + n)
#
# .. code-block:: julia
# :class: test
#
# import Test
# @test x == 3
# @test foo(3) isa Function
# @test foo(3)(4) == 7
# ```
#
# will look as follows in the jupyter notebook
#
# ```{image} img/tests.png
#
# ```
#
# ```{note}
# inclusion of tests in the generated notebook can be controlled in the `conf.py`
# file using `jupyter_drop_tests = False`. This is useful when using the
# `coverage` build pathway.
# ```
| docs/_build/jupyter_execute/examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 08 REGEX START ^ AND END $
import re
# This article is all about the start of line ^ and end of line $ regular expressions in Python's re library.
#
# These two regexes are fundamental to all regular expressions---even outside the Python world. So invest 5 minutes now and master them once and for all!
#
# **Python Re Start-of-String (^) Regex**
#
# You can use the caret operator ^ to match the beginning of the string. For example, this is useful if you want to ensure that a pattern appears at the beginning of a string.
re.findall('^PYTHON', 'PYTHON is fun.')
# The **findall(pattern, string)** method finds all occurrences of the pattern in the string.
#
# The caret at the beginning of the pattern **'^PYTHON'** ensures that you match the word Python only at the beginning of the string. In the previous example, this doesn't make any difference. But in the next example, it does:
#
re.findall('^PYTHON', 'PYTHON! PYTHON is fun')
# Although there are two occurrences of the substring **'PYTHON'**, there's only one matching substring---at the beginning of the string.
#
# But what if you want to match not only at the beginning of the string but at the beginning of each line in a multi-line string?
#
# **Python Re Start-of-Line (^) Regex**
#
# The caret operator, per default, only applies to the start of a string. So if you've got a multi-line string---for example, when reading a text file---it will still only match once: at the beginning of the string.
#
# However, you may want to match at the beginning of each line. For example, you may want to find all lines that start with **'Python'** in a given string.
#
# You can specify that the caret operator matches the beginning of each line via the **re.MULTILINE** flag. Here's an example showing both usages---without and with setting the **re.MULTILINE** flag:
#
text = '''
Python is great.
Python is the fastest growing
major programming language in
the world.
Pythonistas thrive.'''
re.findall('^Python', text)
re.findall('^Python', text, re.MULTILINE)
# The first output is the empty list because the string **'Python'** does not appear at the beginning of the string.
#
# The second output is the list of three matching substrings because the string **'Python'** appears three times at the beginning of a line.
#
# **Python Re End of String ($) Regex**
#
# Similarly, you can use the dollar-sign operator **$** to match the end of the string. Here's an example:
re.findall('fun$', 'PYTHON is fun')
# The **findall()** method finds all occurrences of the pattern in the string---although the trailing dollar-sign $ ensures that the regex matches only at the end of the string.
#
# This can significantly alter the meaning of your regex as you can see in the next example:
re.findall('fun$', 'fun fun fun')
# Although, there are three occurrences of the substring **'fun'**, there's only one matching substring---at the end of the string.
#
# But what if you want to match not only at the end of the string but at the end of each line in a multi-line string?
#
# **Python Re End of Line ($)**
#
# The dollar-sign operator, per default, only applies to the end of a string. So if you've got a multi-line string---for example, when reading a text file---it will still only match once: at the end of the string.
#
# However, you may want to match at the end of each line. For example, you may want to find all lines that end with **'.py'**.
#
# To achieve this, you can specify that the dollar-sign operator matches the end of each line via the **re.MULTILINE** flag. Here's an example showing both usages---without and with setting the **re.MULTILINE** flag:
text = '''
Coding is fun
Python is fun
Games are fun
Agreed?'''
re.findall('fun$', text)
re.findall('fun$', text, flags=re.MULTILINE)
# The first output is the empty list because the string **'fun'** does not appear at the end of the string.
#
# The second output is the list of three matching substrings because the string **'fun'** appears three times at the end of a line.
| Data Science and Machine Learning/Machine-Learning-In-Python-THOROUGH/EXAMPLES/DIFFERENT/FINXTER/REGEX/08_REGEX_START_AND_END.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import casadi as ca
import numpy as np
import control
import matplotlib.pyplot as plt
from casadi.tools.graph import dotgraph
from IPython.display import Image
def draw_graph(expr):
return Image(dotgraph(expr).create_png())
# make matrix printing prettier
np.set_printoptions(precision=3, suppress=True)
# -
def rhs(x, u): # EOMs
s = 2170
cbar = 17.5
mass = 5.0e3
iyy = 4.1e6
tstat = 6.0e4
dtdv = -38.0
ze = 2.0
cdcls = 0.042
cla = 0.085
cma = -0.022
cmde = -0.016
cmq = -16.0
cmadot = -6.0
cladot = 0.0
rtod = 57.29578
gd = 32.17
thtl = u[0]
elev_deg = u[1]
xcg = u[2]
land = u[3]
phi = u[4]
vt = x[0] # velocity, ft/s
alpha = x[1]
alpha_deg = rtod*alpha # angle of attack, deg
theta = x[2] # pitch angle, rad
q = x[3] # pitch rate, rad/s
h = x[4] # altitude, ft
pos = x[5] # horizontal position from origin, ft (not used in dynamics)
r0 = 2.377e-3
tfac = 1.0 - 0.703e-5*h
temperature = ca.if_else(h > 35000, 390.0, 519.0*tfac)
rho = r0*(tfac**4.14)
mach = vt/ca.sqrt(1.4*1716.3*temperature)
qbar = 0.5*rho*vt**2
qs = qbar*s
salp = ca.sin(alpha + phi)
calp = ca.cos(alpha + phi)
gam = theta - alpha
sgam = ca.sin(gam)
cgam = ca.cos(gam)
aero_p = ca.if_else(
land,
(1.0, 0.08, -0.20, 0.02, -0.05),
(0.2, 0.016, 0.05, 0.0, 0.0))
cl0 = aero_p[0]
cd0 = aero_p[1]
cm0 = aero_p[2]
dcdg = aero_p[3]
dcmg = aero_p[4]
thr = (tstat + vt*dtdv)*ca.fmax(thtl, 0)
cl = cl0 + cla*alpha_deg
cm = dcmg + cm0 + cma*alpha_deg + cmde*elev_deg + cl*(xcg - 0.25)
cd = dcdg + cd0 + cdcls*cl**2
x_dot = ca.SX.zeros(6)
x_dot[0] = (thr*calp - qs*cd)/mass - gd*sgam
x_dot[1] = (-thr*salp - qs*cl + mass*(vt*q + gd*cgam))/(mass*vt + qs*cladot)
x_dot[2] = q
d = 0.5*cbar*(cmq*q + cmadot*x_dot[1])/vt
x_dot[3] = (qs*cbar*(cm + d) + thr*ze)/iyy
x_dot[4] = vt*sgam
x_dot[5] = vt*cgam
return x_dot
# +
def constrain(s, vt, h, q, gamma):
# s is our design vector:
# s = [thtl, elev_deg, alpha]
thtl = s[0]
elev_deg = s[1]
alpha = s[2]
phi = s[3]
pos = 0 # we don't care what horiz. position we are at
xcg = 0.25 # we assume xcg at 1/4 chord
land = 0 # we assume we do not have flaps/gear deployed
theta = alpha + gamma
# vt, alpha, theta, q, h, pos
x = ca.vertcat(vt, alpha, theta, q, h, pos) # create column vector
# thtl, elev_deg, xcg, land
u = ca.vertcat(thtl, elev_deg, xcg, land, phi)
return x, u
def trim_cost(x, u):
x_dot = rhs(x, u)
return x_dot[0]**2 + 100*x_dot[1]**2 + 10*x_dot[3]**2
def objective(s, vt, h, q, gamma): # trimming fun
x, u = constrain(s, vt, h, q, gamma)
return trim_cost(x, u)
# -
def trim(vt, h, q, gamma):
s = ca.SX.sym('s', 4)
nlp = {'x': s, 'f': objective(s, vt=vt, h=h, q=q, gamma=gamma)}
S = ca.nlpsol('S', 'ipopt', nlp, {
'print_time': 0,
'ipopt': {
'sb': 'yes',
'print_level': 0,
}
})
# s = [thtl, elev_deg, alpha]
s0 = [0.293, 2.46, np.deg2rad(0.58), np.deg2rad(1)]
res = S(x0=s0, lbg=0, ubg=0, lbx=[0, -60, -np.deg2rad(5), np.deg2rad(0)], ubx=[10, 60, np.deg2rad(18),np.deg2rad(90)]) # boundaries
trim_cost = res['f']
trim_tol = 1e-10
if trim_cost > trim_tol:
raise ValueError('Trim failed to converge', trim_cost)
assert np.abs(float(res['f'])) < trim_tol
s_opt = res['x']
x0, u0 = constrain(s_opt, vt, h, q, gamma)
return {
'x0': np.array(x0).reshape(-1),
'u0': np.array(u0).reshape(-1),
's': np.array(s_opt).reshape(-1)
}
trim(500, 0, 0, 0)
# +
def power_required_curve():
throttle = []
vt_list = np.arange(190, 500, 5)
for vt in vt_list:
res = trim(vt=vt, h=0, q=0, gamma=0)
throttle.append(res['s'][0])
plt.plot(vt_list, 100*np.array(throttle))
plt.grid()
plt.ylabel(r'throttle, %')
plt.xlabel('VT, ft/s')
plt.title('power required curve')
power_required_curve()
# -
# Part 1)
trim(500, 10, 0, 0)
trim(100, 10, 0, 0)
def linearize(trim):
x0 = trim['x0']
u0 = trim['u0']
x = ca.SX.sym('x', 6)
u = ca.SX.sym('u', 5)
y = x
A = ca.jacobian(rhs(x, u), x)
B = ca.jacobian(rhs(x, u), u)
C = ca.jacobian(y, x)
D = ca.jacobian(y, u)
f_ss = ca.Function('ss', [x, u], [A, B, C, D])
return control.ss(*f_ss(x0, u0))
def pitch_rate_control_design(vt, H, xlim, ylim, tf=10):
trim_state = trim(vt=vt, h=0, q=0, gamma=0)
print(trim_state)
sys = linearize(trim_state)
G = control.minreal(control.tf(sys[3, 1]), 1e-2)
control.rlocus(G*H, kvect=np.linspace(0, 1, 1000), xlim=xlim, ylim=ylim);
Go = G*H
Gc = control.feedback(Go)
plt.plot([0, -3], [0, 3*np.arccos(0.707)], '--')
#plt.axis('equal')
plt.grid()
plt.figure())
control.bode(Go, margins=True, dB=True, Hz=True, omega_limits=[1e-2, 1e2], omega_num=1000);
plt.grid(
plt.figure()
t = np.linspace(0, tf, 1000)
r = np.array(t > 0.1, dtype=float)
t, y, x = control.forced_response(Gc, T=t, U=r)
_, u, _ = control.forced_response((1/G)*Gc, T=t, U=r)
u_norm = np.abs(u)
max_u_norm = np.max(u_norm)
print('u_norm max', max_u_norm)
plt.plot(t, y, label='x')
plt.plot(t, r, label='r')
plt.plot(t, u_norm/max_u_norm, label='u normalized')
plt.gca().set_ylim(0, 1.5)
plt.grid()
plt.legend()
s = control.tf([1, 0], [0, 1])
# this is a PID controller with an extra pole at the origin
pitch_rate_control_design(500, -900*((s + 2 + 1j)*(s + 2 - 1j)/s**2), [-8, 2], [-4, 4]) # v, H, xlim, ylim
pitch_rate_control_design(100, -850*((s + 0.75)*(s + 1)/s**2), [-8, 2], [-4, 4]) # v, H, xlim, ylim
# +
#(100, -900*((s + 2 + 1j)*(s + 2 - 1j)/s**2), [-8, 2], [-4, 4]) # v, H, xlim, ylim
# -
| homework/Sam Hazel HW3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="85bf6121" outputId="9d9f6ee0-32ec-4efc-eb15-77c18883b1e2" colab={"base_uri": "https://localhost:8080/"}
# !pip install pytorch-adapt
# + [markdown] id="64a83b07"
# ### Inputs to hooks
# Every hook takes in 2 arguments that represent the current context:
#
# - A dictionary of models and tensors.
# - An optional dictionary of losses.
# + [markdown] id="12f30a44"
# ### FeaturesHook
# + id="5a58f4a5" outputId="317cb30d-91b6-45a8-8e2e-295c79c90ad0" colab={"base_uri": "https://localhost:8080/"}
import torch
from pytorch_adapt.hooks import FeaturesHook
def forward_count(self, *_):
self.count += 1
def print_keys_and_count(inputs, outputs, models):
print("Inputs", list(inputs.keys()))
print("Outputs", list(outputs.keys()))
for k, v in models.items():
print(f"{k}.count = {v.count}")
print("")
G = torch.nn.Linear(1000, 100)
G.register_forward_hook(forward_count)
G.count = 0
models = {"G": G}
data = {
"src_imgs": torch.randn(32, 1000),
"target_imgs": torch.randn(32, 1000),
}
hook = FeaturesHook()
inputs = data
outputs, losses = hook({**models, **inputs})
# Outputs contains src_imgs_features and target_imgs_features.
print_keys_and_count(inputs, outputs, models)
inputs = {**data, **outputs}
outputs, losses = hook({**models, **inputs})
# Outputs is empty because the required outputs are already in the inputs.
# G.count remains the same because G wasn't used for anything.
print_keys_and_count(inputs, outputs, models)
hook = FeaturesHook(detach=True)
outputs, losses = hook({**models, **inputs})
# Detached data is kept separate.
# G.count remains the same because the existing tensors
# were simply detached, and this requires no computation.
print_keys_and_count(inputs, outputs, models)
inputs = data
hook = FeaturesHook(detach=True)
outputs, losses = hook({**models, **inputs})
# G.count increases because the undetached data wasn't passed in
# so it has to be computed
print_keys_and_count(inputs, outputs, models)
inputs = {**data, **outputs}
hook = FeaturesHook()
outputs, losses = hook({**models, **inputs})
# Even though detached data is passed in,
# G.count increases because you can't get undetached data from detached data
print_keys_and_count(inputs, outputs, models)
# + [markdown] id="05f3d1ea"
# ### LogitsHook
#
# ```LogitsHook``` works the same as ```FeaturesHook```, but expects features as input.
# + id="458726af" outputId="156a582b-4a31-450d-f537-4bc8e5eaf6c8" colab={"base_uri": "https://localhost:8080/"}
from pytorch_adapt.hooks import LogitsHook
C = torch.nn.Linear(100, 10)
C.register_forward_hook(forward_count)
C.count = 0
models = {"C": C}
data = {
"src_imgs_features": torch.randn(32, 100),
"target_imgs_features": torch.randn(32, 100),
}
hook = LogitsHook()
inputs = data
outputs, losses = hook({**models, **inputs})
print_keys_and_count(inputs, outputs, models)
# + [markdown] id="c8257928"
# ### FeaturesAndLogitsHook
#
# ```FeaturesAndLogitsHook``` combines ```FeaturesHook``` and ```LogitsHook```.
# + id="0dd4ec18" outputId="622b689c-5a33-4510-e483-6de5a52f6308" colab={"base_uri": "https://localhost:8080/"}
from pytorch_adapt.hooks import FeaturesAndLogitsHook
G.count, C.count = 0, 0
models = {"G": G, "C": C}
data = {
"src_imgs": torch.randn(32, 1000),
"target_imgs": torch.randn(32, 1000),
}
hook = FeaturesAndLogitsHook()
inputs = data
outputs, losses = hook({**models, **inputs})
print_keys_and_count(inputs, outputs, models)
# + [markdown] id="3ccd7789"
# ### ChainHook
#
# ```ChainHook``` allows you to chain together an arbitrary number of hooks. The hooks are run sequentially, with the outputs of hook ```n``` being added to the context so that they become part of the inputs to hook ```n+1```.
# + id="a856c4a3" outputId="e3845d1c-a7b4-45f0-c0fd-13cb0ce8ba2b" colab={"base_uri": "https://localhost:8080/"}
from pytorch_adapt.hooks import ChainHook
G.count, C.count = 0, 0
hook = ChainHook(FeaturesHook(), LogitsHook())
inputs = data
outputs, losses = hook({**models, **inputs})
print_keys_and_count(inputs, outputs, models)
# + id="28d4dd1d"
| examples/in_depth/Hooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practical 1: Create a block from scratch using Python
# This jupyter notebook can be downloaded and run locally with anaconda. Jupyter and anaconda should be installed in all AUT engineering and computer science labs. The benefit of using jupyter is that code snippets can be run live (python is running in the background).
#
# A static version can be found on github at https://github.com/millecodex/COMP842/. All code can be copied and pasted into your favourite text editior or IDE and *might* run with Python 3.x.
#
# You are encouraged to use any programming language you feel comfortable with, this is simply an example using python (and jupyter is designed for python demonstrations). AUT lab computers also have a java interpreter (and maybe a C++?) installed.
# the hash library has many built-in hash functions such as SHA-1 and MD5
import hashlib as hash
# Our block of data will contain many fields such as:<br>
# - identifier
# - time
# - previous hash
# - merkle root
# - list of transactions
# These can be stored in a python dictionary which is a key-value structure
#
# `dict = {key_1:value_1,
# key_2:value_2,
# .
# .
# .
# key_n:value_n
# }`
# ## Initialize a new block. This one will be the _genesis_ block
# initialize a block. Note 'transactions' is initialized as an empty list
block = {
'height':1,
'time':0,
'prevHash':'this is the genesis block',
'merkleRoot': '',
'transactions': []
}
print(block)
# ## Create a transaction and hash it
# Let's create a transaction to store in our blockchain. Remember a transaction is just data; this can be anything represented as a digital object.
# create a transaction (string)
transaction='Pay $1,000,000 to Jeff'
print(transaction)
# To store the transaction object, we will hash it to create a unique identifier of the information
# +
#hashed_tx = hash.sha1(transaction)
#print(hashed_tx)
# -
# ```p
# ---------------------------------------------------------------------------
# TypeError Traceback (most recent call last)
# <ipython-input-34-6b646aa88a56> in <module>
# ----> 1 hashed_tx = hash.sha1(transaction)
# 2 print(hashed_tx)
#
# TypeError: Unicode-objects must be encoded before hashing
# ```
# The error message above is telling us that we cannot hash a string object such as 'Pay $1,000,000 to Jeff'. (Why not?)
# First is must be encoded.
encoded_tx = transaction.encode()
print(encoded_tx)
# the 'b' is telling us that the string is now a byte object
hashed_tx = hash.sha1(encoded_tx)
print(hashed_tx)
# This shows a SHA-1 hash object at the specified memory address. Unfortuantely this isn't human-readable and we can't copy and paste it for verification elsewhere.
#
# The `digest()` and `hexdigest()` methods will output byte objects and hex strings respectively.
print(hashed_tx.digest())
print(hashed_tx.hexdigest())
# ## Add the transaction to the block
hex_tx = hashed_tx.hexdigest()
block["transactions"].append(hex_tx)
print(block)
# ## Create a new block and append it to the chain
# This block only has a single transaction (perhaps its the block reward to Jeff ;) Now we will create a new block and append it to the chain. The block is created in the same manner, except we must update the prevHash field with the hash of the genesis block. This will ensure the state of the blockchain is preserved moving forward.
# some attributes have been hard-coded for simplicity
block2 = {
'height':2,
'time':1,
'prevHash':'null',
'merkleRoot': 'null',
'transactions': []
}
# create a transaction and add it to the block
tx = hash.sha1('Alice +10'.encode()).hexdigest()
block2["transactions"].append(tx)
block2["merkleRoot"] = tx
print(block2)
# The only thing left is to link the blocks. For this we need to hash the entire genesis block object. Proceeding as before:
hash_block_1 = hash.sha1(block.encode())
# This is a uniquely python error. We need to convert the block (dictionary) into a byte object. To do this we need to use the pickle functionality that is built in. You may know this as serialization. Once pickled, we can hash and store as a hex digest.
# +
import pickle
# convert to a byte object
byte_genesis = pickle.dumps(block)
print(byte_genesis)
# compress to a human-readable SHA-1 digest
hash_genesis = hash.sha1(byte_genesis).hexdigest()
print('\n')
print(hash_genesis)
# -
# The byte_genesis output is much longer than our previous byte outputs. Hashing is advantageous because the output is always a fixed length.
#
# Set the prevHash pointer in block2 to the hash of the genesis block.
# set the prevHash and print the block
block2["prevHash"] = hash_genesis
for key, value in block2.items():
print(key+': '+str(value))
# # Modify a transaction to attack the chain
# A hash produces randomized output without any discernable patter relating to the original data. Let test this by modifying the transaction in the genesis block, rehashing, and comparing to the prevHash pointer in block2.
# +
# changing a single transaction modifies the block hash and will invalidate the entire chain
#
# change the dollar sign to a negative sign in the original transaction
new_transaction = 'Pay -1,000,000 to Jeff'
hashed_new_tx=hash.sha1(new_transaction.encode()).hexdigest()
# update the block with the new tx
block["transactions"][0]=hashed_new_tx
# hash the updated block
import pickle
byte_genesis_new = pickle.dumps(block)
hash_genesis = hash.sha1(byte_genesis_new).hexdigest()
# compare hashes
if block2["prevHash"] != hash_genesis:
print('Your chain has been attacked!!')
# -
# ## Summary
# In this tutorial we have:<br>
# - created a block structure including a list of transactions (data)
# - hashed the transaction and added it to the block
# - hashed the entire block
# - added a new block
# - linked the two blocks with a previous hash field to create a block chain
#
# What we have __not__ done is:<br>
# - use a merkle tree to store the transactions
# - store the merkle root in our block structure
#
# Python libraries that this code depends on:
# - __[hashlib](https://docs.python.org/3/library/hashlib.html)__
# - __[pickle](https://docs.python.org/3/library/pickle.html)__
# ## Exercise
# Create a merkle root of the transactions from bitcon block 566446. A \*.csv file can be downloaded from blackboard or this __[repo](https://github.com/millecodex/COMP842/blob/master/tx_list_bitcoin_566446.csv)__.
| Tutorial 1 - Create a block from scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **List:**
# - Name
# - Title
# - Service Year (calc years of service)
# - Job move next 18 months?
# - Organization
# - Interests (list)
# - Job History (list)
mike = {
"Name": "Michael",
"Title": "Site Reliability Engineer",
"Service Year": 2010,
"Job Move": False,
"Organization": "ITC-ITSD-NA",
"Interests": [ "Cycling", "Home Lab" ],
"Job History": [
"Process Analyst",
"Security & Access Management Analyst"
]
}
eric = ["Eric", "Digital Advisor", 1990, False, 'ETC', ["Hiking", "Reading"], ["Application Team Lead", "Solution Architect", "Software Engineer"]]
monty = ['Monty','Data Engineer',2012,False,'ITC',['Umpire Baseball','Referee Football','Woodworking'],['Database Consultant','Data Architect','SCADA Specialist','Systems Engineer','Anti-Money Laundering Consultant']]
nick = ['Nick','Network Security Operations Engineer',2008,False,'ITC',['Gaming','Movies','Technology'],['Mobility Analyst','Business Analyst','Windows Server Analyst']]
satish = ['Satish','IRM Coordinator',2007,True,'ITC',['DJing','Data Engineering'],['Tech Team Lead','IT Analyst']]
Jay_dict= {'name': 'Jay', 'Title': 'EA, Data Science and Analytics', 'Service': 2006, 'Job Move': True, 'Org': 'TSS', 'Interest': ['Coffee'], 'JobHistory': ['IAM', 'SAPM', 'S&T', 'TSS Architecture']}
brian = ['Brian', 'Data Engineer', 2009, False, 'ITC', ['Antiques', 'Birding'], ['SCADA Communications', 'Data Engineer']]
students = [ list(mike.values()) , eric , monty , nick , brian , satish , list(Jay_dict.values()) ]
students_list = []
for student in students:
stdict = {}
stdict = { "Name": student[0],
"Title": student[1],
"Service Year": student[2],
"Years of Service": 2019 - int(student[2]),
"Job Move": bool(student[3]),
"Org": student[4],
"Interests": student[5],
"Job History": student[6]
}
students_list.append(stdict)
for student in students_list:
print(student['Name'].title() + ' - ' + student['Title'].title() + ' in ' + student['Org'])
print('\t' + str(2019 - int(student['Service Year'])) + ' years of service')
if(bool(student['Job Move'])):
print('\tPlans to move jobs within 18 months')
print('\t' + 'Previous Jobs:')
for job in student['Job History']:
print('\t\t' + job)
print('\t' + 'Interests:')
for interest in student['Interests']:
print('\t\t' + interest)
import pandas as pd
students_df = pd.DataFrame(data=students_list)
# +
print(' Average years of serivce: ' + str(students_df['Years of Service'].mean()))
students_df['Years of Service'].hist()
# -
| RedRising/team-rr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import glob
import numpy as np
def tanimoto_dissimilarity(X, Y, X_batch_size=50, Y_batch_size=50):
n_features = X.shape[-1]
if X.ndim == 1:
X = X.reshape(-1, n_features)
if Y.ndim == 1:
Y = Y.reshape(-1, n_features)
tan_sim = []
X_total_batches = X.shape[0] // X_batch_size + 1
Y_total_batches = Y.shape[0] // Y_batch_size + 1
for X_batch_i in range(X_total_batches):
X_start_idx = X_batch_i*X_batch_size
X_end_idx = min((X_batch_i+1)*X_batch_size, X.shape[0])
X_batch = X[X_start_idx:X_end_idx,:]
for Y_batch_i in range(Y_total_batches):
Y_start_idx = Y_batch_i*Y_batch_size
Y_end_idx = min((Y_batch_i+1)*Y_batch_size, Y.shape[0])
Y_batch = Y[Y_start_idx:Y_end_idx,:]
# adapted from: https://github.com/deepchem/deepchem/blob/2531eca8564c1dc68910d791b0bcd91fd586afb9/deepchem/trans/transformers.py#L752
numerator = np.dot(X_batch, Y_batch.T).flatten() # equivalent to np.bitwise_and(X_batch, Y_batch), axis=1)
denominator = n_features - np.dot(1-X_batch, (1-Y_batch).T).flatten() # np.sum(np.bitwise_or(X_rep, Y_rep), axis=1)
tan_sim.append(numerator / denominator)
tan_sim = np.hstack(tan_sim)
return 1.0 - tan_sim
num_files = len(glob.glob('../datasets/lc_clusters_cv_96/unlabeled_*.csv'))
csv_files_list = ['../datasets/lc_clusters_cv_96/unlabeled_{}.csv'.format(i) for i in range(num_files)]
df = pd.concat([pd.read_csv(f) for f in csv_files_list])
X_train = np.vstack([np.fromstring(x, 'u1') - ord('0') for x in df['Morgan FP_2_1024']]).astype(float)
# +
X_train = np.vstack([np.fromstring(x, 'u1') - ord('0') for x in df['Morgan FP_2_1024']]).astype(float)
c2 = np.memmap('../datasets/clustering/cluster_assigment_vector_0.2.dat',
mode='r', dtype='int32', shape=(df.shape[0],))
c3 = np.memmap('../datasets/clustering/cluster_assigment_vector_0.3.dat',
mode='r', dtype='int32', shape=(df.shape[0],))
c4 = np.memmap('../datasets/clustering/cluster_assigment_vector_0.4.dat',
mode='r', dtype='int32', shape=(df.shape[0],))
dissimilarity_matrix = np.memmap('../datasets/dissimilarity_matrix_94857_94857.dat',
shape=(94857,94857), mode='r', dtype='float16')
c2_df = pd.DataFrame(data=np.vstack([c2, np.arange(c2.shape[0])]).T,
columns=['Cluster_0.2', 'Index ID'])
c3_df = pd.DataFrame(data=np.vstack([c3, np.arange(c3.shape[0])]).T,
columns=['Cluster_0.3', 'Index ID'])
c4_df = pd.DataFrame(data=np.vstack([c4, np.arange(c4.shape[0])]).T,
columns=['Cluster_0.4', 'Index ID'])
cl2 = np.memmap('../datasets/clustering/cluster_leader_idx_vector_0.2.dat',
mode='r', dtype='int32', shape=(df.shape[0],))
cl3 = np.memmap('../datasets/clustering/cluster_leader_idx_vector_0.3.dat',
mode='r', dtype='int32', shape=(df.shape[0],))
cl4 = np.memmap('../datasets/clustering/cluster_leader_idx_vector_0.4.dat',
mode='r', dtype='int32', shape=(df.shape[0],))
cl2_df = pd.DataFrame(data=np.vstack([cl2, np.arange(cl2.shape[0])]).T,
columns=['Cluster_0.2_leader_idx', 'Index ID'])
cl3_df = pd.DataFrame(data=np.vstack([cl3, np.arange(cl3.shape[0])]).T,
columns=['Cluster_0.3_leader_idx', 'Index ID'])
cl4_df = pd.DataFrame(data=np.vstack([cl4, np.arange(cl4.shape[0])]).T,
columns=['Cluster_0.4_leader_idx', 'Index ID'])
# -
u2, cc2 = np.unique(c2, return_counts=True)
u3, cc3 = np.unique(c3, return_counts=True)
u4, cc4 = np.unique(c4, return_counts=True)
u2.shape, u3.shape, u4.shape, np.where(cc2==1)[0].shape, np.where(cc3==1)[0].shape, np.where(cc4==1)[0].shape
import scipy.spatial.distance
h_list = np.where(np.in1d(c3, u3[np.where(cc3 == 1)[0]]))[0][2100:2200]
h_list = df.reset_index()[df.reset_index()['Index ID'].isin(h_list)].index.values
cnidx = -1
for h in h_list:
mint = 1000
for i in range(df.shape[0]):
if i != h:
curr_min = scipy.spatial.distance.jaccard(X_train[h], X_train[i])
if curr_min < mint:
mint = curr_min
print(h, mint)
assert(mint >= 0.3)
# +
import pandas as pd
import glob
import numpy as np
new_fmt = '../datasets/lc_clusters_cv_96_new/unlabeled_{}.csv'
num_files = len(glob.glob('../datasets/lc_clusters_cv_96/unlabeled_*.csv'))
csv_files_list = ['../datasets/lc_clusters_cv_96/unlabeled_{}.csv'.format(i) for i in range(num_files)]
for i, f in enumerate(csv_files_list):
df = pd.read_csv(f)
merge_df = pd.merge(df.drop('Cluster_0.2', axis=1), c2_df, how='inner', on='Index ID')
assert np.array_equal(df['Index ID'].values, merge_df['Index ID'].values)
merge_df = pd.merge(merge_df.drop('Cluster_0.3', axis=1), c3_df, how='inner', on='Index ID')
assert np.array_equal(df['Index ID'].values, merge_df['Index ID'].values)
merge_df = pd.merge(merge_df.drop('Cluster_0.4', axis=1), c4_df, how='inner', on='Index ID')
assert np.array_equal(df['Index ID'].values, merge_df['Index ID'].values)
merge_df.to_csv(new_fmt.format(i), index=False)
# +
import pandas as pd
import glob
import numpy as np
new_fmt = '../datasets/lc_clusters_cv_96_new/unlabeled_{}.csv'
num_files = len(glob.glob('../datasets/lc_clusters_cv_96/unlabeled_*.csv'))
csv_files_list = ['../datasets/lc_clusters_cv_96/unlabeled_{}.csv'.format(i) for i in range(num_files)]
for i, f in enumerate(csv_files_list):
df = pd.read_csv(f)
merge_df = pd.merge(df, cl2_df, how='inner', on='Index ID')
assert np.array_equal(df['Index ID'].values, merge_df['Index ID'].values)
merge_df = pd.merge(merge_df, cl3_df, how='inner', on='Index ID')
assert np.array_equal(df['Index ID'].values, merge_df['Index ID'].values)
merge_df = pd.merge(merge_df, cl4_df, how='inner', on='Index ID')
assert np.array_equal(df['Index ID'].values, merge_df['Index ID'].values)
merge_df.to_csv(new_fmt.format(i), index=False)
# +
import pandas as pd
import glob
import numpy as np
new_fmt = '../datasets/lc_clusters_cv_96_new/unlabeled_{}.csv'
num_files = len(glob.glob(new_fmt.format('*')))
csv_files_list = [new_fmt.format(i) for i in range(num_files)]
df = pd.concat([pd.read_csv(f) for f in csv_files_list])
# -
df.to_csv('../datasets/all_data.csv.gz', compression='gzip', index=False)
df[df['Cluster_0.2'] == 3333]
| analysis_notebooks/Analyze Clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Import Libraries
from sklearn.linear_model import SGDRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
from sklearn.metrics import median_absolute_error
#Applying SGDRegressor Model
'''
#sklearn.linear_model.SGDRegressor(loss='squared_loss’, penalty=’l2’, alpha=0.0001,
# l1_ratio=0.15, fit_intercept=True, max_iter=None,
# tol=None, shuffle=True, verbose=0, epsilon=0.1,
# random_state=None, learning_rate='invscaling’,
# eta0=0.01, power_t=0.25, early_stopping=False,
# validation_fraction=0.1, n_iter_no_change=5,
# warm_start=False, average=False, n_iter=None)
'''
SGDRegressionModel = SGDRegressor(alpha=0.1,random_state=33,penalty='l2',loss = 'huber')
SGDRegressionModel.fit(X_train, y_train)
#Calculating Details
print('SGD Regression Train Score is : ' , SGDRegressionModel.score(X_train, y_train))
print('SGD Regression Test Score is : ' , SGDRegressionModel.score(X_test, y_test))
print('SGD Regression Coef is : ' , SGDRegressionModel.coef_)
print('SGD Regression intercept is : ' , SGDRegressionModel.intercept_)
print('----------------------------------------------------')
#Calculating Prediction
y_pred = SGDRegressionModel.predict(X_test)
print('Predicted Value for SGD Regression is : ' , y_pred[:10])
#----------------------------------------------------
#Calculating Mean Absolute Error
MAEValue = mean_absolute_error(y_test, y_pred, multioutput='uniform_average') # it can be raw_values
print('Mean Absolute Error Value is : ', MAEValue)
#----------------------------------------------------
#Calculating Mean Squared Error
MSEValue = mean_squared_error(y_test, y_pred, multioutput='uniform_average') # it can be raw_values
print('Mean Squared Error Value is : ', MSEValue)
#----------------------------------------------------
#Calculating Median Squared Error
MdSEValue = median_absolute_error(y_test, y_pred)
print('Median Squared Error Value is : ', MdSEValue )
# +
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
#load boston data
BostonData = load_boston()
#X Data
X = BostonData.data
#y Data
y = BostonData.target
# +
#Splitting data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=44, shuffle =True)
# +
#Applying SGDRegressor Model
'''
#sklearn.linear_model.SGDRegressor(loss='squared_loss’, penalty=’l2’, alpha=0.0001,
# l1_ratio=0.15, fit_intercept=True, max_iter=None,
# tol=None, shuffle=True, verbose=0, epsilon=0.1,
# random_state=None, learning_rate='invscaling’,
# eta0=0.01, power_t=0.25, early_stopping=False,
# validation_fraction=0.1, n_iter_no_change=5,
# warm_start=False, average=False, n_iter=None)
'''
SGDRegressionModel = SGDRegressor(alpha=0.1,random_state=33,penalty='l2',loss = 'huber')
SGDRegressionModel.fit(X_train, y_train)
# -
#Calculating Details
print('SGD Regression Train Score is : ' , SGDRegressionModel.score(X_train, y_train))
print('SGD Regression Test Score is : ' , SGDRegressionModel.score(X_test, y_test))
print('SGD Regression Coef is : ' , SGDRegressionModel.coef_)
print('SGD Regression intercept is : ' , SGDRegressionModel.intercept_)
#Calculating Prediction
y_pred = SGDRegressionModel.predict(X_test)
print('Predicted Value for SGD Regression is : ' , y_pred[:10])
#Calculating Mean Absolute Error
MAEValue = mean_absolute_error(y_test, y_pred, multioutput='uniform_average') # it can be raw_values
print('Mean Absolute Error Value is : ', MAEValue)
#Calculating Mean Squared Error
MSEValue = mean_squared_error(y_test, y_pred, multioutput='uniform_average') # it can be raw_values
print('Mean Squared Error Value is : ', MSEValue)
#Calculating Median Squared Error
MdSEValue = median_absolute_error(y_test, y_pred)
print('Median Squared Error Value is : ', MdSEValue )
| Sklearn/2.1 Linear Regression/.ipynb_checkpoints/SGD Regressor-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] colab_type="text" id="UIp998OHnZSN"
# ##### Copyright 2018 The TensorFlow Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + colab={} colab_type="code" id="li5wNGR6naj0"
#@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="ykgJW69K4TNL"
# # Linear Mixed-Effect Regression in {TF Probability, R, Stan}
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/probability/blob/master/tensorflow_probability/examples/jupyter_notebooks/HLM_TFP_R_Stan.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/probability/blob/master/tensorflow_probability/examples/jupyter_notebooks/HLM_TFP_R_Stan.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
#
# + [markdown] colab_type="text" id="epdzLgKQRfAI"
# ## 1 Introduction
# + [markdown] colab_type="text" id="bMstt69hR44D"
# In this colab we will fit a linear mixed-effect regression model to a popular, toy dataset. We will make this fit thrice, using R's `lme4`, Stan's mixed-effects package, and TensorFlow Probability (TFP) primitives. We conclude by showing all three give roughly the same fitted parameters and posterior distributions.
#
# Our main conclusion is that TFP has the general pieces necessary to fit HLM-like models and that it produces results which are consistent with other software packages, i.e.., `lme4`, `rstanarm`. This colab is not an accurate reflection of the computational efficiency of any of the packages compared.
# + colab={} colab_type="code" id="0axKjgZvRtL9"
# %matplotlib inline
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
from six.moves import urllib
import numpy as np
import pandas as pd
import warnings
from matplotlib import pyplot as plt
import seaborn as sns
from IPython.core.pylabtools import figsize
figsize(11, 9)
import tensorflow as tf
import tensorflow_probability as tfp
# + [markdown] colab_type="text" id="IFC9r-h0XlQ3"
# ## 2 Hierarchical Linear Model
#
# + [markdown] colab_type="text" id="wqS-HhvhDlno"
# For our comparison between R, Stan, and TFP, we will fit a [Hierarchical Linear Model](https://en.wikipedia.org/wiki/Multilevel_model) (HLM) to the [Radon dataset](http://www.stat.columbia.edu/~gelman/arm/examples/radon/) made popular in [_Bayesian Data Analysis_ by Gelman, et. al.](http://www.stat.columbia.edu/~gelman/book/) (page 559, second ed; page 250, third ed.).
#
# + [markdown] colab_type="text" id="fAD8am2a4TaY"
# We assume the following generative model:
#
# $$\begin{align*}
# \text{for } & c=1\ldots \text{NumCounties}:\\
# & \beta_c \sim \text{Normal}\left(\text{loc}=0, \text{scale}=\sigma_C \right) \\
# \text{for } & i=1\ldots \text{NumSamples}:\\
# &\eta_i = \underbrace{\omega_0 + \omega_1 \text{Floor}_i}_\text{fixed effects} + \underbrace{\beta_{ \text{County}_i} \log( \text{UraniumPPM}_{\text{County}_i}))}_\text{random effects} \\
# &\log(\text{Radon}_i) \sim \text{Normal}(\text{loc}=\eta_i , \text{scale}=\sigma_N)
# \end{align*}$$
#
# + [markdown] colab_type="text" id="5styKLl_MyWu"
# In R's `lme4` "tilde notation", this model is equivalent to:
# > `log_radon ~ 1 + floor + (0 + log_uranium_ppm | county)`
#
# + [markdown] colab_type="text" id="SpurhP2gP_T8"
# We will find MLE for $\omega, \sigma_C, \sigma_N$ using the posterior distribution (conditioned on evidence) of $\{\beta_c\}_{c=1}^\text{NumCounties}$.
# + [markdown] colab_type="text" id="Nj6adNwgPTUP"
# For essentially the same model but _with_ a random intercept, see _[Appendix A](#scrollTo=tsXhZ4rtNUXL)_.
#
# For a more general specification of HLMs, see _[Appendix B](#scrollTo=H0w7ofFvNsxi)_.
# + [markdown] colab_type="text" id="LR0ZC0dE4MWb"
# ## 3 Data Munging
# + [markdown] colab_type="text" id="OdboSs9G3JlE"
# In this section we obtain the [`radon` dataset](http://www.stat.columbia.edu/~gelman/arm/examples/radon/) and do some minimal preprocessing to make it comply with our assumed model.
# + colab={} colab_type="code" id="4LjOBqLDV0IQ"
# We'll use the following directory to store files we download as well as our
# preprocessed dataset.
CACHE_DIR = os.path.join(os.sep, 'tmp', 'radon')
def cache_or_download_file(cache_dir, url_base, filename):
"""Read a cached file or download it."""
filepath = os.path.join(cache_dir, filename)
if tf.gfile.Exists(filepath):
return filepath
if not tf.gfile.Exists(cache_dir):
tf.gfile.MakeDirs(cache_dir)
url = os.path.join(url_base, filename)
print("Downloading {url} to {filepath}.".format(url=url, filepath=filepath))
urllib.request.urlretrieve(url, filepath)
return filepath
def download_radon_dataset(cache_dir=CACHE_DIR):
"""Download the radon dataset and read as Pandas dataframe."""
url_base = 'http://www.stat.columbia.edu/~gelman/arm/examples/radon/'
# Alternative source:
# url_base = ('https://raw.githubusercontent.com/pymc-devs/uq_chapter/'
# 'master/reference/data/')
srrs2 = pd.read_csv(cache_or_download_file(cache_dir, url_base, 'srrs2.dat'))
srrs2.rename(columns=str.strip, inplace=True)
cty = pd.read_csv(cache_or_download_file(cache_dir, url_base, 'cty.dat'))
cty.rename(columns=str.strip, inplace=True)
return srrs2, cty
def preprocess_radon_dataset(srrs2, cty, state='MN'):
"""Preprocess radon dataset as done in "Bayesian Data Analysis" book."""
srrs2 = srrs2[srrs2.state==state].copy()
cty = cty[cty.st==state].copy()
# We will now join datasets on Federal Information Processing Standards
# (FIPS) id, ie, codes that link geographic units, counties and county
# equivalents. http://jeffgill.org/Teaching/rpqm_9.pdf
srrs2['fips'] = 1000 * srrs2.stfips + srrs2.cntyfips
cty['fips'] = 1000 * cty.stfips + cty.ctfips
df = srrs2.merge(cty[['fips', 'Uppm']], on='fips')
df = df.drop_duplicates(subset='idnum')
df = df.rename(index=str, columns={'Uppm': 'uranium_ppm'})
# For any missing or invalid activity readings, we'll use a value of `0.1`.
df['radon'] = df.activity.apply(lambda x: x if x > 0. else 0.1)
# Remap categories to start from 0 and end at max(category).
county_name = sorted(df.county.unique())
df['county'] = df.county.astype(
pd.api.types.CategoricalDtype(categories=county_name)).cat.codes
county_name = map(str.strip, county_name)
df['log_radon'] = df['radon'].apply(np.log)
df['log_uranium_ppm'] = df['uranium_ppm'].apply(np.log)
df = df[['log_radon', 'floor', 'county', 'log_uranium_ppm']]
return df, county_name
# + colab={} colab_type="code" id="hJE3-eC0I-Lm"
radon, county_name = preprocess_radon_dataset(*download_radon_dataset())
# + colab={} colab_type="code" id="nV-IAEW2FIqX"
# Save processed data. (So we can later read it in R.)
with tf.gfile.Open(os.path.join(CACHE_DIR, 'radon.csv'), 'w') as f:
radon.to_csv(f, index=False)
# + [markdown] colab_type="text" id="ubvf1vHenyCx"
# ### 3.1 Know Thy Data
# + [markdown] colab_type="text" id="r39MiQV2zUz0"
# In this section we explore the `radon` dataset to get a better sense of why the proposed model might be reasonable.
# + colab={"height": 204} colab_type="code" id="GRCyjhSknu9z" outputId="879816fb-1512-42c9-80cc-e0e966e5ed50"
radon.head()
# + colab={"height": 362} colab_type="code" id="gdASxsWHjvw4" outputId="97e80a33-a8bd-4969-8239-2dfca8c18050"
fig, ax = plt.subplots(figsize=(22, 5));
county_freq = radon['county'].value_counts()
county_freq.plot(kind='bar', color='#436bad');
plt.xlabel('County index')
plt.ylabel('Number of radon readings')
plt.title('Number of radon readings per county', fontsize=16)
county_freq = np.array(zip(county_freq.index, county_freq.values)) # We'll use this later.
# + colab={"height": 291} colab_type="code" id="k9yN_rGLlqHE" outputId="90ef0661-6d70-4bc6-99c0-d261ca962c00"
fig, ax = plt.subplots(ncols=2, figsize=[10, 4]);
radon['log_radon'].plot(kind='density', ax=ax[0]);
ax[0].set_xlabel('log(radon)')
radon['floor'].value_counts().plot(kind='bar', ax=ax[1]);
ax[1].set_xlabel('Floor');
ax[1].set_ylabel('Count');
fig.subplots_adjust(wspace=0.25)
# + [markdown] colab_type="text" id="vuy-t8oNoDTH"
# Conclusions:
# - There's a long tail of 85 counties. (A common occurrence in GLMMs.)
# - Indeed $\log(\text{Radon})$ is unconstrained. (So linear regression might make sense.)
# - Readings are most made on the $0$-th floor; no reading was made above floor $1$. (So our fixed effects will only have two weights.)
#
# + [markdown] colab_type="text" id="zSCWOamv3nXU"
# ## 4 HLM In R
# + [markdown] colab_type="text" id="ZPtvJXUdn6an"
# In this section we use R's [`lme4`](https://cran.r-project.org/web/packages/lme4/index.html) package to fit probabilistic model described above.
# + [markdown] colab_type="text" id="3iX5L0srRGIQ"
# **NOTE: To execute this section, you must switch to an `R` colab runtime.**
# + colab={} colab_type="code" id="ZBqZjyHdsPIB"
suppressMessages({
library('bayesplot')
library('data.table')
library('dplyr')
library('gfile')
library('ggplot2')
library('lattice')
library('lme4')
library('plyr')
library('rstanarm')
library('tidyverse')
RequireInitGoogle()
})
# + colab={"height": 143} colab_type="code" id="Lq3_yATCshI-" outputId="989dbf1a-2ee3-44f4-9391-f40d73856bc2"
data = read_csv(gfile('/tmp/radon/radon.csv'))
# + colab={"height": 179} colab_type="code" id="_3gj9hfxshE8" outputId="61582bc1-2f65-4c2d-917a-eb49a106bf10"
head(data)
# + colab={} colab_type="code" id="uRqAdn3WsoN-"
# https://github.com/stan-dev/example-models/wiki/ARM-Models-Sorted-by-Chapter
radon.model <- lmer(log_radon ~ 1 + floor + (0 + log_uranium_ppm | county), data = data)
# + colab={"height": 449} colab_type="code" id="MuMBVnkAsoMS" outputId="ab284dea-3748-411a-c1a6-a41b69ddecfe"
summary(radon.model)
# + colab={"height": 515} colab_type="code" id="0qZXx27dp7aZ" outputId="088c0305-72ee-42be-cbe2-cebc00faca53"
qqmath(ranef(radon.model, condVar=TRUE))
# + colab={} colab_type="code" id="nCsGcLnP40Lg"
write.csv(as.data.frame(ranef(radon.model, condVar = TRUE)), '/tmp/radon/lme4_fit.csv')
# + [markdown] colab_type="text" id="2XrrSLW43pHL"
# ## 5 HLM In Stan
#
# + [markdown] colab_type="text" id="-ddXXuiWnv2-"
# In this section we use [rstanarm](http://mc-stan.org/users/interfaces/rstanarm) to fit a Stan model using the same formula/syntax as the `lme4` model above.
#
# Unlike `lme4` and the TF model below, `rstanarm` is a fully Bayesian model, i.e., all parameters are presumed drawn from a Normal distribution with parameters themselves drawn from a distribution.
# + [markdown] colab_type="text" id="c6IpkPOOnsmQ"
# **NOTE: To execute this section, you must switch an `R` colab runtime.**
# + colab={"height": 1385} colab_type="code" id="s-p-rAMZuaGh" outputId="e2d4f709-b81d-434c-c829-50829e69348f"
fit <- stan_lmer(log_radon ~ 1 + floor + (0 + log_uranium_ppm | county), data = data)
# + [markdown] colab_type="text" id="KZNNvBB8TWTW"
# **Note**: The runtimes are from a single CPU core. (This colab is not intended to be a faithful representation of Stan or TFP runtime.)
# + colab={"height": 395} colab_type="code" id="yNKzX9rnuz9H" outputId="91df9ccb-d977-4b85-e8bc-42f12f1c8ccc"
fit
# + colab={"height": 497} colab_type="code" id="iO6In1K3uz7B" outputId="7f0248bb-6e0d-4ed2-feab-a887e5b9904f"
color_scheme_set("red")
ppc_dens_overlay(y = fit$y,
yrep = posterior_predict(fit, draws = 50))
# + colab={"height": 497} colab_type="code" id="ZAMwe8rWvId4" outputId="280ddaa7-7377-4d80-fc73-508a9732baf7"
color_scheme_set("brightblue")
ppc_intervals(
y = data$log_radon,
yrep = posterior_predict(fit),
x = data$county,
prob = 0.8
) +
labs(
x = "County",
y = "log radon",
title = "80% posterior predictive intervals \nvs observed log radon",
subtitle = "by county"
) +
panel_bg(fill = "gray95", color = NA) +
grid_lines(color = "white")
# + colab={} colab_type="code" id="h9HtqG65x1a6"
# Write the posterior samples (4000 for each variable) to a CSV.
write.csv(tidy(as.matrix(fit)), "/tmp/radon/stan_fit.csv")
# + [markdown] colab_type="text" id="FedP5SMQ3u4z"
# **Note: Switch back to the Python TF kernel runtime.**
# + colab={} colab_type="code" id="wwhJD-t86Dnq"
with tf.gfile.Open('/tmp/radon/lme4_fit.csv', 'r') as f:
lme4_fit = pd.read_csv(f, index_col=0)
# + colab={"height": 204} colab_type="code" id="Qs9VpUOz6LZR" outputId="6871c6ff-d570-4e4b-cc49-b9242e0f7ce1"
lme4_fit.head()
# + [markdown] colab_type="text" id="EJktXVyR6zK6"
# Retrieve the point estimates and conditional standard deviations for the group random effects from lme4 for visualization later.
# + colab={"height": 35} colab_type="code" id="le7XkSvL6a2Z" outputId="3196b30b-6950-4812-b07f-1ee1b0401fc2"
posterior_random_weights_lme4 = np.array(lme4_fit.condval, dtype=np.float32)
lme4_prior_scale = np.array(lme4_fit.condsd, dtype=np.float32)
print(posterior_random_weights_lme4.shape, lme4_prior_scale.shape)
# + [markdown] colab_type="text" id="42fI-VbduCXy"
# Draw samples for the county weights using the lme4 estimated means and standard deviations.
# + colab={} colab_type="code" id="S8TQNRaKFecg"
with tf.Session() as sess:
lme4_dist = tfp.distributions.Independent(
tfp.distributions.Normal(
loc=posterior_random_weights_lme4,
scale=lme4_prior_scale),
reinterpreted_batch_ndims=1)
posterior_random_weights_lme4_final_ = sess.run(lme4_dist.sample(4000))
# + colab={"height": 35} colab_type="code" id="KA2OZUp3FltF" outputId="6228936d-7e06-412c-d857-2166970569b6"
posterior_random_weights_lme4_final_.shape
# + [markdown] colab_type="text" id="y8GpeGOauTar"
# We also retrieve the posterior samples of the county weights from the Stan fit.
# + colab={} colab_type="code" id="YxXhcMfG3uoX"
with tf.gfile.Open('/tmp/radon/stan_fit.csv', 'r') as f:
samples = pd.read_csv(f, index_col=0)
# + colab={"height": 248} colab_type="code" id="Arc-QdJ33ukk" outputId="e972b3ee-0bb3-4f09-d858-794064903344"
samples.head()
# + colab={"height": 35} colab_type="code" id="-TdxDTcJ40QY" outputId="18146805-d370-487d-dfed-091337a52247"
posterior_random_weights_cols = [
col for col in samples.columns if 'b.log_uranium_ppm.county' in col
]
posterior_random_weights_final_stan = samples[
posterior_random_weights_cols].values
print(posterior_random_weights_final_stan.shape)
# + [markdown] colab_type="text" id="Qv_caP1t4FVF"
# [This Stan example](https://github.com/stan-dev/example-models/blob/master/ARM/Ch.16/radon.3.stan) shows how one would implement LMER in a style closer to TFP, i.e., by directly specifying the probabilistic model.
# + [markdown] colab_type="text" id="QkchUh3V382r"
# ## 6 HLM In TF Probability
# + [markdown] colab_type="text" id="Ywj6S9iQ0aZe"
# In this section we will use low-level TensorFlow Probability primitives (`Distributions`) to specify our Hierarchical Linear Model as well as fit the unkown parameters.
# + colab={} colab_type="code" id="TOh_69los9gK"
# Handy snippet to reset the global graph and global session.
with warnings.catch_warnings():
warnings.simplefilter('ignore')
tf.reset_default_graph()
try:
sess.close()
except:
pass
sess = tf.InteractiveSession()
# + [markdown] colab_type="text" id="g6xl7I6XTTg5"
# ### 6.1 Specify Model
# + [markdown] colab_type="text" id="GjzA6-vAXXLS"
# In this section we specify the [radon linear mixed-effect model](#scrollTo=IFC9r-h0XlQ3) using TFP primitives. To do this, we specify two functions which produce two TFP distributions:
# - `make_weights_prior`: A multivariate Normal prior for the random weights (which are multiplied by $\log(\text{UraniumPPM}_{c_i})$ to compue the linear predictor).
# - `make_log_radon_likelihood`: A batch of `Normal` distributions over each observed $\log(\text{Radon}_i)$ dependent variable.
#
#
# + [markdown] colab_type="text" id="_QtzWC-ZZ9U-"
# Since we will be fitting the parameters of each of these distributions we must use TF variables (i.e., [`tf.get_variable`](https://www.tensorflow.org/api_docs/python/tf/get_variable)). However, since we wish to use unconstrained optimzation we must find a way to constrain real-values to achieve the necessary semantics, eg, postives which represent standard deviations.
# + colab={} colab_type="code" id="NzpFXkvOXMav"
inv_scale_transform = lambda y: np.log(y) # Not using TF here.
fwd_scale_transform = tf.exp
# + [markdown] colab_type="text" id="817V1h2_aCFp"
# The following function constructs our prior, $p(\beta|\sigma_C)$ where $\beta$ denotes the random-effect weights and $\sigma_C$ the standard deviation.
#
# We use `tf.make_template` to ensure that the first call to this function instantiates the TF variables it uses and all subsequent calls _reuse_ the variable's current value.
# + colab={} colab_type="code" id="JnPFL-pKXMRl"
def _make_weights_prior(num_counties, dtype):
"""Returns a `len(log_uranium_ppm)` batch of univariate Normal."""
raw_prior_scale = tf.get_variable(
name='raw_prior_scale',
initializer=np.array(inv_scale_transform(1.), dtype=dtype))
return tfp.distributions.Independent(
tfp.distributions.Normal(
loc=tf.zeros(num_counties, dtype=dtype),
scale=fwd_scale_transform(raw_prior_scale)),
reinterpreted_batch_ndims=1)
make_weights_prior = tf.make_template(
name_='make_weights_prior', func_=_make_weights_prior)
# + [markdown] colab_type="text" id="3-aeEIbmaJQ1"
# The following function constructs our likelihood, $p(y|x,\omega,\beta,\sigma_N)$ where $y,x$ denote response and evidence, $\omega,\beta$ denote fixed- and random-effect weights, and $\sigma_N$ the standard deviation.
#
# Here again we use `tf.make_template` to ensure the TF variables are reused across calls.
# + colab={} colab_type="code" id="wNQTcHcQXMIp"
def _make_log_radon_likelihood(random_effect_weights, floor, county,
log_county_uranium_ppm, init_log_radon_stddev):
raw_likelihood_scale = tf.get_variable(
name='raw_likelihood_scale',
initializer=np.array(
inv_scale_transform(init_log_radon_stddev), dtype=dtype))
fixed_effect_weights = tf.get_variable(
name='fixed_effect_weights', initializer=np.array([0., 1.], dtype=dtype))
fixed_effects = fixed_effect_weights[0] + fixed_effect_weights[1] * floor
random_effects = tf.gather(
random_effect_weights * log_county_uranium_ppm,
indices=tf.to_int32(county),
axis=-1)
linear_predictor = fixed_effects + random_effects
return tfp.distributions.Normal(
loc=linear_predictor, scale=fwd_scale_transform(raw_likelihood_scale))
make_log_radon_likelihood = tf.make_template(
name_='make_log_radon_likelihood', func_=_make_log_radon_likelihood)
# + [markdown] colab_type="text" id="_dGxuoLPbRMs"
# Finally we use the prior and likelihood generators to construct the joint log-density.
# + colab={} colab_type="code" id="_UBayNK538JD"
def joint_log_prob(random_effect_weights, log_radon, floor, county,
log_county_uranium_ppm, dtype):
num_counties = len(log_county_uranium_ppm)
rv_weights = make_weights_prior(num_counties, dtype)
rv_radon = make_log_radon_likelihood(
random_effect_weights,
floor,
county,
log_county_uranium_ppm,
init_log_radon_stddev=radon.log_radon.values.std())
return (rv_weights.log_prob(random_effect_weights)
+ tf.reduce_sum(rv_radon.log_prob(log_radon), axis=-1))
# + [markdown] colab_type="text" id="xkeKH0rTTWDo"
# ### 6.2 Training (Stochastic Approximation of Expectation Maximization)
# + [markdown] colab_type="text" id="h7Xr0X4Qbe9C"
# To fit our linear mixed-effect regression model, we will use a stochastic approximation version of the Expectation Maximization algorithm (SAEM). The basic idea is to use samples from the posterior to approximate the expected joint log-density (E-step). Then we find the parameters which maximize this calculation (M-step). Somewhat more concretely, the fixed-point iteration is given by:
#
# $$\begin{align*}
# \text{E}[ \log p(x, Z | \theta) | \theta_0]
# &\approx \frac{1}{M} \sum_{m=1}^M \log p(x, z_m | \theta), \quad Z_m\sim p(Z | x, \theta_0) && \text{E-step}\\
# &=: Q_M(\theta, \theta_0) \\
# \theta_0 &= \theta_0 - \eta \left.\nabla_\theta Q_M(\theta, \theta_0)\right|_{\theta=\theta_0} && \text{M-step}
# \end{align*}$$
#
# where $x$ denotes evidence, $Z$ some latent variable which needs to be marginalized out, and $\theta,\theta_0$ possible parameterizations.
#
# For a more thorough explanation, see [_Convergence of a stochastic approximation version of the EM algorithms_ by <NAME>, <NAME>, <NAME> (Ann. Statist., 1999)](https://projecteuclid.org/euclid.aos/1018031103).
# + [markdown] colab_type="text" id="hwEoELEueaeZ"
# To compute the E-step, we need to sample from the posterior. Since our posterior is not easy to sample from, we use Hamiltonian Monte Carlo (HMC). HMC is a Monte Carlo Markov Chain procedure which uses gradients (wrt state, not parameters) of the unnormalized posterior log-density to propose new samples.
#
#
# + [markdown] colab_type="text" id="JOTzK4qne9Qr"
# Specifying the unnormalized posterior log-density is simple--it is merely the joint log-density "pinned" at whatever we wish to condition on.
# + colab={} colab_type="code" id="FSwVJAkNEx6Y"
# Specify unnormalized posterior.
dtype = np.float32
log_county_uranium_ppm = radon[
['county', 'log_uranium_ppm']].drop_duplicates().values[:, 1]
log_county_uranium_ppm = log_county_uranium_ppm.astype(dtype)
def unnormalized_posterior_log_prob(random_effect_weights):
return joint_log_prob(
random_effect_weights=random_effect_weights,
log_radon=dtype(radon.log_radon.values),
floor=dtype(radon.floor.values),
county=np.int32(radon.county.values),
log_county_uranium_ppm=log_county_uranium_ppm,
dtype=dtype)
# + [markdown] colab_type="text" id="khZHTgVYfASP"
# We now complete the E-step setup by creating an HMC transition kernel.
#
# Notes:
#
# - We use `state_stop_gradient=True`to prevent the M-step from backpropping through draws from the MCMC. (Recall, we needn't backprop through because our E-step is intentionally parameterized at the _previous_ best known estimators.)
#
# - We use [`tf.placeholder`](https://www.tensorflow.org/api_docs/python/tf/placeholder) so that when we eventually execute our TF graph, we can feed the previous iteration's random MCMC sample as the the next iteration's chain's value.
#
# - We use TFP's adaptive `step_size` heuristic, `tfp.mcmc.hmc_step_size_update_fn`.
# + colab={} colab_type="code" id="WnZ_KMP0E0ot"
# Set-up E-step.
step_size = tf.get_variable(
'step_size',
initializer=np.array(0.2, dtype=dtype),
trainable=False)
hmc = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=unnormalized_posterior_log_prob,
num_leapfrog_steps=2,
step_size=step_size,
step_size_update_fn=tfp.mcmc.make_simple_step_size_update_policy(
num_adaptation_steps=None),
state_gradients_are_stopped=True)
init_random_weights = tf.placeholder(dtype, shape=[len(log_county_uranium_ppm)])
posterior_random_weights, kernel_results = tfp.mcmc.sample_chain(
num_results=3,
num_burnin_steps=0,
num_steps_between_results=0,
current_state=init_random_weights,
kernel=hmc)
# + [markdown] colab_type="text" id="-GmtUwoLff1y"
# We now set-up the M-step. This is essentially the same as an optimization one might do in TF.
# + colab={} colab_type="code" id="wceMwnGwvUfF"
# Set-up M-step.
loss = -tf.reduce_mean(kernel_results.accepted_results.target_log_prob)
global_step = tf.train.get_or_create_global_step()
learning_rate = tf.train.exponential_decay(
learning_rate=0.1,
global_step=global_step,
decay_steps=2,
decay_rate=0.99)
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss, global_step=global_step)
# + [markdown] colab_type="text" id="s_ykFN0Cfoel"
# We conclude with some housekeeping tasks. We must tell TF that all variables are initialized. We also create handles to our TF variables so we can `print` their values at each iteration of the procedure.
# + colab={} colab_type="code" id="PakV59O8E3m5"
# Initialize all variables.
init_op = tf.initialize_all_variables()
# + colab={} colab_type="code" id="FziBCkW_NXFF"
# Grab variable handles for diagnostic purposes.
with tf.variable_scope('make_weights_prior', reuse=True):
prior_scale = fwd_scale_transform(tf.get_variable(
name='raw_prior_scale', dtype=dtype))
with tf.variable_scope('make_log_radon_likelihood', reuse=True):
likelihood_scale = fwd_scale_transform(tf.get_variable(
name='raw_likelihood_scale', dtype=dtype))
fixed_effect_weights = tf.get_variable(
name='fixed_effect_weights', dtype=dtype)
# + [markdown] colab_type="text" id="pwjiJLywlZgF"
# ### 6.3 Execute
# + [markdown] colab_type="text" id="ouO-E4Ncf0KE"
# In this section we execute our SAEM TF graph. The main trick here is to feed our last draw from the HMC kernel into the next iteration. This is achieved through our use of `feed_dict` in the `sess.run` call.
# + colab={} colab_type="code" id="Cy36-LMMNbTc"
init_op.run()
w_ = np.zeros([len(log_county_uranium_ppm)], dtype=dtype)
# + colab={"height": 361} colab_type="code" id="OrzwMVaoE0i2" outputId="8ff7881a-e249-448c-b048-7d9e3a8b286c"
# %%time
maxiter = int(1500)
num_accepted = 0
num_drawn = 0
for i in range(maxiter):
[
_,
global_step_,
loss_,
posterior_random_weights_,
kernel_results_,
step_size_,
prior_scale_,
likelihood_scale_,
fixed_effect_weights_,
] = sess.run([
train_op,
global_step,
loss,
posterior_random_weights,
kernel_results,
step_size,
prior_scale,
likelihood_scale,
fixed_effect_weights,
], feed_dict={init_random_weights: w_})
w_ = posterior_random_weights_[-1, :]
num_accepted += kernel_results_.is_accepted.sum()
num_drawn += kernel_results_.is_accepted.size
acceptance_rate = num_accepted / num_drawn
if i % 100 == 0 or i == maxiter - 1:
print('global_step:{:>4} loss:{: 9.3f} acceptance:{:.4f} '
'step_size:{:.4f} prior_scale:{:.4f} likelihood_scale:{:.4f} '
'fixed_effect_weights:{}'.format(
global_step_, loss_.mean(), acceptance_rate, step_size_,
prior_scale_, likelihood_scale_, fixed_effect_weights_))
# + [markdown] colab_type="text" id="QR50yLYygWdg"
# Looks like after ~1500 steps, our estimates of the parameters have stabilized.
# + [markdown] colab_type="text" id="x2BtkWEIVsB9"
# ### 6.4 Results
# + [markdown] colab_type="text" id="HYs17VUto_te"
# Now that we've fit the parameters, let's generate a large number of posterior samples and study the results.
# + colab={"height": 53} colab_type="code" id="v-X0DhqHjdue" outputId="ae3dbb66-843a-4e90-b94f-e34eea0e7ffb"
# %%time
posterior_random_weights_final, kernel_results_final = tfp.mcmc.sample_chain(
num_results=int(15e3),
num_burnin_steps=int(1e3),
current_state=init_random_weights,
kernel=tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=unnormalized_posterior_log_prob,
num_leapfrog_steps=2,
step_size=step_size))
[
posterior_random_weights_final_,
kernel_results_final_,
] = sess.run([
posterior_random_weights_final,
kernel_results_final,
], feed_dict={init_random_weights: w_})
# + colab={"height": 89} colab_type="code" id="LCECfwexk1DN" outputId="557e2f55-865e-4086-f243-b44fba016984"
print('prior_scale: ', prior_scale_)
print('likelihood_scale: ', likelihood_scale_)
print('fixed_effect_weights: ', fixed_effect_weights_)
print('acceptance rate final: ', kernel_results_final_.is_accepted.mean())
# + [markdown] colab_type="text" id="qbzMVKPykLYq"
# We now construct a box and whisker diagram of the $\beta_c \log(\text{UraniumPPM}_c)$ random-effect. We'll order the random-effects by decreasing county frequency.
# + colab={"height": 393} colab_type="code" id="vjA7TVT4wuQa" outputId="fefa381f-379b-439b-d603-3fc8cc364bfc"
x = posterior_random_weights_final_ * log_county_uranium_ppm
I = county_freq[:, 0]
x = x[:, I]
cols = np.array(county_name)[I]
pw = pd.DataFrame(x)
pw.columns = cols
fig, ax = plt.subplots(figsize=(25, 4))
ax = pw.boxplot(rot=80, vert=True);
# + [markdown] colab_type="text" id="DXIqPxN1j3wI"
# From this box and whisker diagram, we observe that the variance of the county-level $\log(\text{UraniumPPM})$ random-effect increases as the county is less represented in the dataset. Intutively this makes sense--we should be less certain about the impact of a certain county if we have less evidence for it.
# + [markdown] colab_type="text" id="DtlOjtAkyE76"
# ## 7 Side-by-Side-by-Side Comparison
# + [markdown] colab_type="text" id="QF6OwHJSk7xM"
# We now compare the results of all three procedures. To do this, we will compute non-parameteric estimates of the posterior samples as generated by Stan and TFP. We will also compare against the parameteric (approximate) estimates produced by R's `lme4` package.
# + [markdown] colab_type="text" id="Qg8pYIRQy9Ea"
# The following plot depicts the posterior distribution of each weight for each county in Minnesota. We show results for Stan (red), TFP (blue), and R's `lme4` (orange). We shade results from Stan and TFP thus expect to see purple when the two agree. For simplicity we do not shade results from R. Each subplot represents a single county and are ordered in descending frequency in raster scan order (i.e., from left-to-right then top-to-bottom).
# + colab={"height": 1254} colab_type="code" id="9vhm-sNWrhmB" outputId="d6890814-1334-4bff-ae5f-8a03cfc43287"
nrows = 17
ncols = 5
fig, ax = plt.subplots(nrows, ncols, figsize=(18, 21), sharey=True, sharex=True)
with warnings.catch_warnings():
warnings.simplefilter('ignore')
ii = -1
for r in range(nrows):
for c in range(ncols):
ii += 1
idx = county_freq[ii, 0]
sns.kdeplot(
posterior_random_weights_final_[:, idx] * log_county_uranium_ppm[idx],
color='blue',
alpha=.3,
shade=True,
label='TFP',
ax=ax[r][c])
sns.kdeplot(
posterior_random_weights_final_stan[:, idx] *
log_county_uranium_ppm[idx],
color='red',
alpha=.3,
shade=True,
label='Stan/rstanarm',
ax=ax[r][c])
sns.kdeplot(
posterior_random_weights_lme4_final_[:, idx] *
log_county_uranium_ppm[idx],
color='#F4B400',
alpha=.7,
shade=False,
label='R/lme4',
ax=ax[r][c])
ax[r][c].vlines(
posterior_random_weights_lme4[idx] * log_county_uranium_ppm[idx],
0,
5,
color='#F4B400',
linestyle='--')
ax[r][c].set_title(county_name[idx] + ' ({})'.format(idx), y=.7)
ax[r][c].set_ylim(0, 5)
ax[r][c].set_xlim(-1., 1.)
ax[r][c].get_yaxis().set_visible(False)
if ii == 2:
ax[r][c].legend(bbox_to_anchor=(1.4, 1.7), fontsize=20, ncol=3)
else:
ax[r][c].legend_.remove()
fig.subplots_adjust(wspace=0.03, hspace=0.1)
# + [markdown] colab_type="text" id="bv4rVc4Mye7J"
# ## 8 Conclusion
# + [markdown] colab_type="text" id="jNtzJUQXksvZ"
# In this colab we fit a linear mixed-effect regression model to the radon dataset. We tried three different software packages: R, Stan, and TensorFlow Probability. We concluded by plotting the 85 posterior distributions as computed by the three different software packages.
# + [markdown] colab_type="text" id="tsXhZ4rtNUXL"
# ## Appendix A: Alternative Radon HLM (Add Random Intercept)
# + [markdown] colab_type="text" id="F9PqNJQK002P"
# In this section we describe an alternative HLM which also has a random intercept associated with each county.
# + [markdown] colab_type="text" id="qt8a50GYSqbe"
#
# $$\begin{align*}
# \text{for } & c=1\ldots \text{NumCounties}:\\
# & \beta_c \sim \text{MultivariateNormal}\left(\text{loc}=\left[ \begin{array}{c} 0 \\ 0 \end{array}\right] , \text{scale}=\left[\begin{array}{cc} \sigma_{11} & 0 \\ \sigma_{12} & \sigma_{22} \end{array}\right] \right) \\
# \text{for } & i=1\ldots \text{NumSamples}:\\
# & c_i := \text{County}_i \\
# &\eta_i = \underbrace{\omega_0 + \omega_1\text{Floor}_i \vphantom{\log( \text{CountyUraniumPPM}_{c_i}))}}_{\text{fixed effects}} + \underbrace{\beta_{c_i,0} + \beta_{c_i,1}\log( \text{CountyUraniumPPM}_{c_i}))}_{\text{random effects}} \\
# &\log(\text{Radon}_i) \sim \text{Normal}(\text{loc}=\eta_i , \text{scale}=\sigma)
# \end{align*}$$
#
# + [markdown] colab_type="text" id="oI-DkfJWxK5K"
# In R's `lme4` "tilde notation", this model is equivalent to:
# > `log_radon ~ 1 + floor + (1 + log_county_uranium_ppm | county)`
# + [markdown] colab_type="text" id="H0w7ofFvNsxi"
# ## Appendix B: Generalized Linear Mixed-Effect Models
# + [markdown] colab_type="text" id="5g4VAJP5xOPZ"
# In this section we give a more general characterization of Hierarchical Linear Models than what is used in the main body. This more general model is known as a [generalized linear mixed-effect model](https://en.wikipedia.org/wiki/Generalized_linear_mixed_model) (GLMM).
#
# + [markdown] colab_type="text" id="lA1xwCdENyTx"
# GLMMs are generalizations of [generalized linear models](https://en.wikipedia.org/wiki/Generalized_linear_model) (GLMs). GLMMs extend GLMs by incorporating sample specific random noise into the predicted linear response. This is useful in part because it allows rarely seen features to share information with more commonly seen features.
#
#
#
# + [markdown] colab_type="text" id="9RJ6ryvUvJ5O"
# As a generative process, a Generalized Linear Mixed-effects Model (GLMM) is characterized by:
#
# \begin{align}
# \text{for } & r = 1\ldots R: \hspace{2.45cm}\text{# for each random-effect group}\\
# &\begin{aligned}
# \text{for } &c = 1\ldots |C_r|: \hspace{1.3cm}\text{# for each category ("level") of group $r$}\\
# &\begin{aligned}
# \beta_{rc}
# &\sim \text{MultivariateNormal}(\text{loc}=0_{D_r}, \text{scale}=\Sigma_r^{1/2})
# \end{aligned}
# \end{aligned}\\\\
# \text{for } & i = 1 \ldots N: \hspace{2.45cm}\text{# for each sample}\\
# &\begin{aligned}
# &\eta_i = \underbrace{\vphantom{\sum_{r=1}^R}x_i^\top\omega}_\text{fixed effects} + \underbrace{\sum_{r=1}^R z_{r,i}^\top \beta_{r,C_r(i) }}_\text{random effects} \\
# &Y_i|x_i,\omega,\{z_{r,i} , \beta_r\}_{r=1}^R \sim \text{Distribution}(\text{mean}= g^{-1}(\eta_i))
# \end{aligned}
# \end{align}
#
# + [markdown] colab_type="text" id="ycA3XujsN5FJ"
# where:
#
# \begin{align}
# R &= \text{number of random-effect groups}\\
# |C_r| &= \text{number of categories for group $r$}\\
# N &= \text{number of training samples}\\
# x_i,\omega &\in \mathbb{R}^{D_0}\\
# D_0 &= \text{number of fixed-effects}\\
# C_r(i) &= \text{category (under group $r$) of the $i$th sample}\\
# z_{r,i} &\in \mathbb{R}^{D_r}\\
# D_r &= \text{number of random-effects associated with group $r$}\\
# \Sigma_{r} &\in \{S\in\mathbb{R}^{D_r \times D_r} : S \succ 0 \}\\
# \eta_i\mapsto g^{-1}(\eta_i) &= \mu_i, \text{inverse link function}\\
# \text{Distribution} &=\text{some distribution parameterizable solely by its mean}
# \end{align}
#
# + [markdown] colab_type="text" id="-JAvS_UfN7gl"
# In words, this says that every category of each group is associated with an iid MVN, $\beta_{rc}$. Although the $\beta_{rc}$ draws are always independent, they are only indentically distributed for a group $r$; notice there is exactly one $\Sigma_r$ for each $r\in\{1,\ldots,R\}$.
#
# When affinely combined with a sample's group's features, $z_{r,i}$, the result is sample-specific noise on the $i$-th predicted linear response (which is otherwise $x_i^\top\omega$).
#
# + [markdown] colab_type="text" id="G4W5ijp1OFPW"
# When we estimate $\{\Sigma_r:r\in\{1,\ldots,R\}\}$ we're essentially estimating the amount of noise a random-effect group carries which would otherwise drown out the signal present in $x_i^\top\omega$.
# + [markdown] colab_type="text" id="7Dl2iZkAODfG"
# There are a variety of options for the $\text{Distribution}$ and [inverse link function](https://en.wikipedia.org/wiki/Generalized_linear_model#Link_function), $g^{-1}$. Common choices are:
# - $Y_i\sim\text{Normal}(\text{mean}=\eta_i, \text{scale}=\sigma)$,
# - $Y_i\sim\text{Binomial}(\text{mean}=n_i \cdot \text{sigmoid}(\eta_i), \text{total_count}=n_i)$, and,
# - $Y_i\sim\text{Poisson}(\text{mean}=\exp(\eta_i))$.
#
# For more possibilities, see the [`tfp.glm`](https://github.com/tensorflow/probability/tree/master/tensorflow_probability/python/glm) module.
| tensorflow_probability/examples/jupyter_notebooks/HLM_TFP_R_Stan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import libraries
import numpy as np
import pandas as pd
from sklearn.datasets import load_iris
import matplotlib.pyplot as plt
from sklearn.manifold import MDS
from sklearn.preprocessing import MinMaxScaler
import seaborn as sns
# ## First a tutorial making MDS plots with the iris dataset
# https://towardsdatascience.com/visualize-multidimensional-datasets-with-mds-64d7b4c16eaa
# load in the data
data = load_iris()
X = data.data
# 0-1 scaling
scaler = MinMaxScaler()
X_scaled = scaler.fit_transform(X)
# apply MDS to get a two-dimensional dataset
mds = MDS(2,random_state=0)
X_2d = mds.fit_transform(X_scaled)
# plot the data
colors = ['red','green','blue']
plt.rcParams['figure.figsize'] = [7, 7]
plt.rc('font', size=14)
for i in np.unique(data.target):
subset = X_2d[data.target == i]
x = [row[0] for row in subset]
y = [row[1] for row in subset]
plt.scatter(x,y,c=colors[i],label=data.target_names[i])
plt.legend()
plt.show()
# ## Running MDS with our test data
test = pd.read_csv('result_table.csv')
test.head()
# keep rows with only certain tissue types for now - this will change with the clean dataset
test = test[(test == 'leaves').any(axis=1)|(test == 'root').any(axis=1)|(test == 'whole plant').any(axis=1)|(test == 'seedlings').any(axis=1)|(test == 'rosette leaf').any(axis=1)]
test['Tissue'].unique()
# taking a random 20% sample of the data to see if it speeds up processing time
# also making a variable with the tissue column names to merge back in with the MDS dataframe later on
test_tissue_20 = test.sample(frac=0.2, replace=False, random_state=1)
sample_names = test_tissue_20["Tissue"]
test_tissue_20.shape
#test_tissue_20.to_csv('data_subset.csv')
# transform data to contain only genes as columns and tissue types as rows
# first, a dataframe with just tissue types for testing - this will definitely change w/ the entire data set
test_tissue = test_tissue_20[['AT1G22630','AT1G22620','AT1G22610','Tissue']]
test_tissue = test_tissue.set_index('Tissue')
test_tissue = test_tissue.rename_axis('').rename_axis('Gene', axis='columns')
test_tissue.head()
test_tissue.shape
# Log transform data - do we need to do this? will the data cleaning group do any transformations?
for c in [c for c in test_tissue.columns if np.issubdtype(test_tissue[c].dtype , np.number)]:
test_tissue[c] += 1
for c in [c for c in test_tissue.columns if np.issubdtype(test_tissue[c].dtype , np.number)]:
test_tissue[c] = np.log(test_tissue[c])
test_tissue.head()
# apply MDS to get a two-dimensional dataset
mds = MDS(n_components=2, random_state=3)
test_mds = mds.fit_transform(test_tissue)
# make dataframe with MDS values
real_mds_df = pd.DataFrame(data = test_mds, columns = ["MDS1", "MDS2"])
real_mds_df.shape
# check that tissue column is good
sample_names.head()
sample_names.shape
# add tissue names back in
real_mds_df.reset_index(drop=True, inplace=True)
sample_names.reset_index(drop=True, inplace=True)
final_real_df = pd.concat([sample_names, real_mds_df], axis = 1)
final_real_df.head()
# plot
sns.lmplot('MDS1', 'MDS2', height=10, data=final_real_df, hue="Tissue", fit_reg=False)
#plt.savefig('plot.png')
| Dimension_Reduction/MDS_test_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fully Convolutional Network - Semantic Segmentation
#
# 
# 
# +
import os
import os.path as osp
import pytz
import torch
import warnings
warnings.filterwarnings('ignore')
configurations = {
# same configuration as original work
# https://github.com/shelhamer/fcn.berkeleyvision.org
1: dict(
max_iteration=100000,
lr=1.0e-10,
momentum=0.99,
weight_decay=0.0005,
interval_validate=4000,
)
}
# -
from types import SimpleNamespace
opts = SimpleNamespace()
opts.cfg = configurations[1]
opts.resume = ''
print(opts.cfg)
from utils import get_log_dir
opts.out = get_log_dir('vgg8s', 1, opts.cfg)
print(opts.out)
gpu = 1
os.environ['CUDA_VISIBLE_DEVICES'] = str(gpu)
cuda = torch.cuda.is_available()
print('Cuda: {}'.format(cuda))
opts.cuda = 'cuda' if cuda else 'cpu'
opts.mode = 'train'
opts.backbone = 'vgg'
opts.fcn = '8s'
# ## PascalVOC Dataset - Downloaded on _`root`_ variable
root = './data/Pascal_VOC'
print(root)
from data_loader import Pascal_Data
kwargs = {'num_workers': 4} if cuda else {}
train_loader = torch.utils.data.DataLoader(
Pascal_Data(root, image_set='train', backbone='vgg'),
batch_size=1, shuffle=True, **kwargs)
val_loader = torch.utils.data.DataLoader(
Pascal_Data(root, image_set='val', backbone='vgg'),
batch_size=1, shuffle=False, **kwargs)
data_loader = [train_loader, val_loader]
# +
# %matplotlib inline
import matplotlib.pyplot as plt
for data, target in train_loader: break
print(data.shape)
print(target.shape)
data.min()
data_show, label_show = train_loader.dataset.untransform(data[0].cpu().clone(), target[0].cpu().clone())
plt.imshow(data_show)
plt.show()
def imshow_label(label_show):
import matplotlib
import numpy as np
cmap = plt.cm.jet
# extract all colors from the .jet map
cmaplist = [cmap(i) for i in range(cmap.N)]
cmaplist[0] = (0.0,0.0,0.0,1.0)
cmap = cmap.from_list('Custom cmap', cmaplist, cmap.N)
# define the bins and normalize
bounds = np.arange(0,len(train_loader.dataset.class_names))
norm = matplotlib.colors.BoundaryNorm(bounds, cmap.N)
plt.imshow(label_show, cmap=cmap, norm=norm)
cbar = plt.colorbar(ticks=bounds)
cbar.ax.set_yticklabels(train_loader.dataset.class_names)
plt.show()
imshow_label(label_show)
# -
# ## FCN - Model
# +
import numpy as np
import torch.nn as nn
class FCN8s(nn.Module):
def __init__(self, n_class=21):
super(FCN8s, self).__init__()
# conv1
self.conv1_1 = nn.Conv2d(3, 64, 3, padding=100)
self.relu1_1 = nn.ReLU(inplace=True)
self.conv1_2 = nn.Conv2d(64, 64, 3, padding=1)
self.relu1_2 = nn.ReLU(inplace=True)
self.pool1 = nn.MaxPool2d(2, stride=2, ceil_mode=True) # 1/2
# conv2
self.conv2_1 = nn.Conv2d(64, 128, 3, padding=1)
self.relu2_1 = nn.ReLU(inplace=True)
self.conv2_2 = nn.Conv2d(128, 128, 3, padding=1)
self.relu2_2 = nn.ReLU(inplace=True)
self.pool2 = nn.MaxPool2d(2, stride=2, ceil_mode=True) # 1/4
# conv3
self.conv3_1 = nn.Conv2d(128, 256, 3, padding=1)
self.relu3_1 = nn.ReLU(inplace=True)
self.conv3_2 = nn.Conv2d(256, 256, 3, padding=1)
self.relu3_2 = nn.ReLU(inplace=True)
self.conv3_3 = nn.Conv2d(256, 256, 3, padding=1)
self.relu3_3 = nn.ReLU(inplace=True)
self.pool3 = nn.MaxPool2d(2, stride=2, ceil_mode=True) # 1/8
# conv4
self.conv4_1 = nn.Conv2d(256, 512, 3, padding=1)
self.relu4_1 = nn.ReLU(inplace=True)
self.conv4_2 = nn.Conv2d(512, 512, 3, padding=1)
self.relu4_2 = nn.ReLU(inplace=True)
self.conv4_3 = nn.Conv2d(512, 512, 3, padding=1)
self.relu4_3 = nn.ReLU(inplace=True)
self.pool4 = nn.MaxPool2d(2, stride=2, ceil_mode=True) # 1/16
# conv5
self.conv5_1 = nn.Conv2d(512, 512, 3, padding=1)
self.relu5_1 = nn.ReLU(inplace=True)
self.conv5_2 = nn.Conv2d(512, 512, 3, padding=1)
self.relu5_2 = nn.ReLU(inplace=True)
self.conv5_3 = nn.Conv2d(512, 512, 3, padding=1)
self.relu5_3 = nn.ReLU(inplace=True)
self.pool5 = nn.MaxPool2d(2, stride=2, ceil_mode=True) # 1/32
# fc6
self.fc6 = nn.Conv2d(512, 4096, 7)
self.relu6 = nn.ReLU(inplace=True)
self.drop6 = nn.Dropout2d()
# fc7
self.fc7 = nn.Conv2d(4096, 4096, 1)
self.relu7 = nn.ReLU(inplace=True)
self.drop7 = nn.Dropout2d()
self.score_fr = nn.Conv2d(4096, n_class, 1)
self.score_pool3 = nn.Conv2d(256, n_class, 1)
self.score_pool4 = nn.Conv2d(512, n_class, 1)
self.upscore2 = nn.ConvTranspose2d(n_class,
n_class,
4,
stride=2,
bias=False)
self.upscore_pool4 = nn.ConvTranspose2d(n_class,
n_class,
4,
stride=2,
bias=False)
self.upscore8 = nn.ConvTranspose2d(n_class,
n_class,
16,
stride=8,
bias=False)
self._initialize_weights()
def _initialize_weights(self):
from models.vgg.helpers import get_upsampling_weight
for m in self.modules():
if isinstance(m, nn.Conv2d):
m.weight.data.zero_()
if m.bias is not None:
m.bias.data.zero_()
if isinstance(m, nn.ConvTranspose2d):
assert m.kernel_size[0] == m.kernel_size[1]
initial_weight = get_upsampling_weight(m.in_channels,
m.out_channels,
m.kernel_size[0])
m.weight.data.copy_(initial_weight)
def forward(self, x, debug=False):
h = x
if debug:
print(h.data.shape)
h = self.relu1_1(self.conv1_1(h))
if debug:
print(h.data.shape)
h = self.relu1_2(self.conv1_2(h))
if debug:
print(h.data.shape)
h = self.pool1(h)
if debug:
print(h.data.shape)
h = self.relu2_1(self.conv2_1(h))
if debug:
print(h.data.shape)
h = self.relu2_2(self.conv2_2(h))
if debug:
print(h.data.shape)
h = self.pool2(h)
if debug:
print(h.data.shape)
h = self.relu3_1(self.conv3_1(h))
if debug:
print(h.data.shape)
h = self.relu3_2(self.conv3_2(h))
if debug:
print(h.data.shape)
h = self.relu3_3(self.conv3_3(h))
if debug:
print(h.data.shape)
h = self.pool3(h)
if debug:
print('pool3: {}'.format(h.data.shape))
pool3 = h # 1/8
h = self.relu4_1(self.conv4_1(h))
if debug:
print(h.data.shape)
h = self.relu4_2(self.conv4_2(h))
if debug:
print(h.data.shape)
h = self.relu4_3(self.conv4_3(h))
if debug:
print(h.data.shape)
h = self.pool4(h)
if debug:
print('pool4: {}'.format(h.data.shape))
pool4 = h # 1/16 #<------------------------------------
h = self.relu5_1(self.conv5_1(h))
if debug:
print(h.data.shape)
h = self.relu5_2(self.conv5_2(h))
if debug:
print(h.data.shape)
h = self.relu5_3(self.conv5_3(h))
if debug:
print(h.data.shape)
h = self.pool5(h)
if debug:
print(h.data.shape)
h = self.relu6(self.fc6(h))
if debug:
print(h.data.shape)
h = self.drop6(h)
if debug:
print(h.data.shape)
h = self.relu7(self.fc7(h))
if debug:
print(h.data.shape)
h = self.drop7(h)
if debug:
print(h.data.shape)
h = self.score_fr(h)
if debug:
print(h.data.shape)
h = self.upscore2(h)
if debug:
print('upscore2: {}'.format(h.data.shape))
upscore2 = h # 1/16
h = self.score_pool4(pool4)
if debug:
print('score_pool4: {}'.format(h.data.shape))
h = h[:, :, 5:5 + upscore2.size()[2], 5:5 + upscore2.size()[3]]
if debug:
print('score_pool4c: {}'.format(h.data.shape))
score_pool4c = h # 1/16
h = upscore2 + score_pool4c
if debug:
print('upscore2+score_pool4c: {}'.format(h.data.shape))
h = self.upscore_pool4(h)
if debug:
print('upscore_pool4: {}'.format(h.data.shape))
upscore_pool4 = h # 1/8
h = self.score_pool3(pool3)
if debug:
print('score_pool3: {}'.format(h.data.shape))
h = h[:, :, 9:9 + upscore_pool4.size()[2], 9:9 +
upscore_pool4.size()[3]]
if debug:
print('score_pool3c: {}'.format(h.data.shape))
score_pool3c = h # 1/8
h = upscore_pool4 + score_pool3c # 1/8
if debug:
print('upscore_pool4+score_pool3c: {}'.format(h.data.shape))
h = self.upscore8(h)
if debug:
print('upscore8: {}'.format(h.data.shape))
h = h[:, :, 31:31 + x.size()[2], 31:31 + x.size()[3]].contiguous()
if debug:
print('upscore8 rearranged: {}'.format(h.data.shape))
return h
def copy_params_from_fcn16s(self, fcn16s):
for name, l1 in fcn16s.named_children():
try:
l2 = getattr(self, name)
l2.weight # skip ReLU / Dropout
except Exception:
continue
assert l1.weight.size() == l2.weight.size()
l2.weight.data.copy_(l1.weight.data)
if l1.bias is not None:
assert l1.bias.size() == l2.bias.size()
l2.bias.data.copy_(l1.bias.data)
# -
# https://github.com/shelhamer/fcn.berkeleyvision.org/blob/master/surgery.py
def get_upsampling_weight(in_channels, out_channels, kernel_size):
"""Make a 2D bilinear kernel suitable for upsampling"""
factor = (kernel_size + 1) // 2
if kernel_size % 2 == 1:
center = factor - 1
else:
center = factor - 0.5
og = np.ogrid[:kernel_size, :kernel_size]
filt = (1 - abs(og[0] - center) / factor) * \
(1 - abs(og[1] - center) / factor)
weight = np.zeros((in_channels, out_channels, kernel_size, kernel_size),
dtype=np.float64)
weight[range(in_channels), range(out_channels), :, :] = filt
return torch.from_numpy(weight).float()
# ## From fcn16s weights
model = FCN8s(n_class=21)
model.to(opts.cuda)
model
iter_loader=iter(train_loader)
data, target = next(iter_loader)
data = data.to(opts.cuda)
with torch.no_grad():
output = model(data)
print('input: ', data.shape)
print('output: ', output.data.shape)
data, target = next(iter_loader)
data = data.to(opts.cuda)
with torch.no_grad():
output = model(data, debug=True)
data, target = next(iter_loader)
data = data.to(opts.cuda)
with torch.no_grad():
output = model(data, debug=True)
if opts.resume:
print('Loading checkpoint from: '+resume)
checkpoint = torch.load(resume)
model.load_state_dict(checkpoint['model_state_dict'])
else:
from models.vgg.fcn16s import FCN as FCN16
fcn16s = FCN16()
fcn16s_weights = FCN16.download() # Original FCN16 pretrained model
fcn16s.load_state_dict(torch.load(fcn16s_weights))
model.copy_params_from_fcn16s(fcn16s)
# %matplotlib inline
from trainer import Trainer
trainer = Trainer(data_loader, opts)
print(opts.cfg.get('interval_validate', len(train_loader))) #Validate every 4000 iterations
print(opts.out)
start_epoch = 0
start_iteration = 0
if opts.resume:
start_epoch = checkpoint['epoch']
start_iteration = checkpoint['iteration']
trainer.epoch = start_epoch
trainer.iteration = start_iteration
trainer.Train()
| vgg8s_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../')
import soynlp
print(soynlp.__version__)
# -
# soynlp 0.0.46+ 에서는 soynlp.noun.LRNounExtractor 를 보완한 LRNounExtractor_v2 를 제공합니다.
#
# version 2 에서는 (1) 명사 추출의 정확성을 높였으며, (2) 합성명사의 인식이 가능합니다. 또한 (3) 명사의 빈도를 정확히 계산합니다.
#
# 사용법은 version 1 과 비슷합니다. train_extract 함수를 통하여 명사 점수를 계산할 수 있습니다. verbose mode 일 경우에는 학습 과정의 진행 상황이 출력됩니다.
#
# 더 자세한 version 1 과 2 의 차이점은 [lovit.github.io/nlp/2018/05/08/noun_extraction_ver2](https://lovit.github.io/nlp/2018/05/08/noun_extraction_ver2/) 에 적어뒀습니다.
# +
from soynlp.utils import DoublespaceLineCorpus
from soynlp.noun import LRNounExtractor_v2
corpus_path = '2016-10-20-news'
sents = DoublespaceLineCorpus(corpus_path, iter_sent=True)
# -
# train(), extract() 를 따로 진행할 수 있습니다.
# %%time
noun_extractor = LRNounExtractor_v2(verbose=True)
noun_extractor.train(sents)
nouns = noun_extractor.extract()
# 이 과정은 한 번에 일어나는 경우가 많기 때문에 train_extract() 함수를 이용해도 됩니다.
#
# version 2 에서는 soynlp.utils 의 EojeolCounter 와 LRGraph 를 이용합니다. 명사가 포함되었다고 인식된 어절의 빈도수 비율이 출력됩니다. 71.69 % 의 어절에 추출된 명사가 포함되어 있습니다. 총 86,133 개의 명사가 추출되었으며, 이 중 35,044 개는 복합명사입니다.
#
# 복합 명사의 경우에는 min_count 가 1 일 경우에도 탐지가 될 수 있도록 하였습니다. min_count 와 minimum_noun_score 는 extract, train_extract 함수의 argument 로 조절할 수 있습니다.
# %%time
nouns = noun_extractor.train_extract(sents)
# nouns = noun_extractor.train_extract(sents, min_count=1, minimum_noun_score=0.3)
# nouns 는 {str: NounScore} 형식의 dict 입니다. 추출된 명사 단어에 대한 빈도수와 명사 점수가 namedtuple 인 NounScore 로 저장되어 있습니다.
#
# version 1 의 명사 추출기에서는 '뉴스'라는 left-side substring 의 빈도수를 명사의 빈도수로 이용하였습니다만, version 2 에서는 어절에서 '뉴스'가 실제로 명사로 이용된 경우만 카운팅 됩니다. '뉴스방송'과 같은 복합명사의 빈도수는 '뉴스'에 포함되지 않습니다.
nouns['뉴스']
# LRNounExtractor_v2._compounds_components 에는 복합 명사의 components 가 저장되어 있습니다. _compounds_components 는 {str:tuple of str} 형식입니다.
for word in ['두바이월', '두바이월드', '두바이월드센터', '연합뉴스', '연합뉴스자료사', '연합뉴스자료사진', '군사기술']:
print(word, noun_extractor._compounds_components.get(word, None))
list(noun_extractor._compounds_components.items())[:10]
# 복합 명사도 nouns 에 포함되어 출력됩니다.
nouns['두바이월드센터']
# LRNounExtractor_v2.decompose_compound 는 입력된 str 가 복합 명사일 경우, 이를 단일 명사의 tuple 로 분해합니다.
noun_extractor.decompose_compound('두바이월드센터시카고옵션거래소')
# 복합명사가 아닌 경우에는 길이가 1 인 tuple 로 출력됩니다.
noun_extractor.decompose_compound('두바이월드센터시카고옵션거래소라라라라라')
# LRNounExtractor_v2 는 soynlp.utils 의 LRGraph 를 이용합니다. 데이터의 L-R 구조를 살펴볼 수 있습니다.
noun_extractor.lrgraph.get_r('아이오아이')
# topk=10 으로 설정되어 있습니다. topk < 0 으로 설정하면 모든 R set 이 출력됩니다.
noun_extractor.lrgraph.get_r('아이오아이', topk=-1)
# L-R 구조의 L parts 도 확인할 수 있습니다. 이 역시 topk=10 으로 기본값이 설정되어 있습니다.
noun_extractor.lrgraph.get_l('었다고')
| SoyNLP/tutorials/nounextractor-v2_usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multilayer perceptron
# +
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Prettier plots
sns.set()
# +
class NeuronLayer():
def __init__(self, n_inputs, n_neurons):
self.weights = 2 * np.random.random((n_inputs, n_neurons)) - 1
self.output = np.zeros(n_neurons)
self.bias = np.zeros(n_neurons)
self.deltas = np.zeros(n_neurons)
def print(self):
print("Weights:", self.weights)
class NeuralNetwork:
def __init__(self, layers):
self.layers = layers
self.errors = []
def activation_function(self, x):
return 1 / (1 + np.exp(-x)) # Sigmoid
def activation_function_derivative(self, x):
return x * (1 - x) # Sigmoid derivative
def train(self, inputs, expected, learning_rate, n_epochs):
for epoch in range(n_epochs):
self.forward_propagation(inputs)
self.backward_propagation(expected)
self.update_weights(inputs, learning_rate)
def forward_propagation(self, inputs):
for layer in self.layers:
layer.outputs = self.activation_function(np.dot(inputs, layer.weights))
inputs = layer.outputs
return inputs
def backward_propagation(self, expected):
for i in reversed(range(len(self.layers))):
layer = self.layers[i]
errors = 0
if i == len(self.layers) - 1:
errors = expected - layer.outputs
self.errors.append(errors)
else:
errors = np.dot(self.layers[i+1].deltas, self.layers[i+1].weights.T)
layer.deltas = errors * self.activation_function_derivative(layer.outputs)
def update_weights(self, inputs, learning_rate):
adjustments = [];
for i in range(len(self.layers)):
if i != 0:
inputs = self.layers[i-1].outputs
adjustments.append(inputs.T.dot(self.layers[i].deltas) * learning_rate)
for i in range(len(self.layers)):
self.layers[i].weights += adjustments[i]
# -
data_set = np.array([[0, 0, 1], [0, 1, 1], [1, 0, 1], [1, 1, 1]])
xor_set = np.array([[0], [1], [1], [0]])
# +
for n_hidden in range(1, 5):
hl = NeuronLayer(3, n_hidden)
ol = NeuronLayer(n_hidden, 1)
nn = NeuralNetwork((hl, ol))
nn.train(data_set, xor_set, 0.1, 20000)
plt.plot(np.sum(np.squeeze(nn.errors)**2,1), label=n_hidden)
plt.title("XOR with Multilayer Perceptron")
plt.ylabel("SSE")
plt.xlabel("Epochs")
plt.legend()
plt.show()
nn.forward_propagation(data_set)
# -
| Multilayer_perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Proteomic and 3D Structural Evidence of Cysteine Oxidative PTMs.
#
# Under oxidative stress Cysteines can undergo oxidative post-translational modifications (PTMs).
#
# In this study we compare the results of a proteomics study with observed oxidized forms of Cysteines in 3D structures from the Protein Data Bank (PDB).
#
# * Proteomics Dataset
#
# The study by Akter et al. compares the differences between S-Sulfinylations (R-SO2H) and S-Sulfenylations (R-SOH) in A549 and HeLa cell lines.
#
# Chemical proteomics reveals new targets of cysteine sulfinic acid reductase.
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>.
# Nat Chem Biol. 2018 Sep 3. doi: [10.1038/s41589-018-0116-2](https://doi.org/10.1038/s41589-018-0116-2)
#
# * PDB Dataset
#
# BioJava-ModFinder: identification of protein modifications in 3D structures from the Protein Data Bank. <NAME>, <NAME>, <NAME> <NAME>, <NAME>, <NAME> Bourne, PE, Rose PW, Bioinformatics 2017, 33: 2047–2049. [doi: doi.org/10.1093/bioinformatics/btx101](https://doi.org/10.1093/bioinformatics/btx101)
# + code_folding=[0]
import pandas as pd
import numpy as np
from io import BytesIO
import xlrd
from ipywidgets import interact, IntSlider, widgets
import py3Dmol
from pyspark.sql import SparkSession
from pyspark.sql.functions import asc, collect_set, collect_list, col, concat_ws, sort_array
from mmtfPyspark.datasets import pdbToUniProt, pdbPtmDataset
# + code_folding=[]
# setup checkboxes for datasets
w1 = widgets.Checkbox(value=True, description='A549-RSO2H',disabled=False)
w2 = widgets.Checkbox(value=False, description='HeLa-RSO2H',disabled=False)
w3 = widgets.Checkbox(value=True, description='A549-RSOH',disabled=False)
w4 = widgets.Checkbox(value=False, description='HeLa-RSOH',disabled=False)
# -
# ## Select one or more datasets (cell line-PTM)
display(w1, w2, w3, w4)
# ## Read and process datasets from supplementary materials
# + code_folding=[]
def read_datasets():
dfs = []
if w1.value:
df1 = pd.read_excel('https://static-content.springer.com/esm/art%3A10.1038%2Fs41589-018-0116-2/MediaObjects/41589_2018_116_MOESM32_ESM.xlsx', sheet_name='A549', dtype=str)
df1 = df1.assign(ptms=np.full((df1.shape[0], 1), "A549-RSO2H"))
df1 = df1.rename(index=str, columns={"Modified site": "modifiedSite", "Uniprot Accession #": "uniprotAccession"})
dfs.append(df1)
if w2.value:
df2 = pd.read_excel('https://static-content.springer.com/esm/art%3A10.1038%2Fs41589-018-0116-2/MediaObjects/41589_2018_116_MOESM32_ESM.xlsx', sheet_name='HeLa', dtype=str)
df2 = df2.assign(ptms=np.full((df2.shape[0], 1), "HeLa-RSO2H"))
df2 = df2.rename(index=str, columns={"Modified site": "modifiedSite", "Uniprot Accession #": "uniprotAccession"})
dfs.append(df2)
if w3.value:
df3 = pd.read_excel('https://static-content.springer.com/esm/art%3A10.1038%2Fs41589-018-0116-2/MediaObjects/41589_2018_116_MOESM33_ESM.xlsx', sheet_name='A549', dtype=str)
df3 = df3.assign(ptms=np.full((df3.shape[0], 1), "A549-RSOH"))
df3 = df3.rename(index=str, columns={"Site #": "modifiedSite", "Uniprot Accession #": "uniprotAccession"})
dfs.append(df3)
if w4.value:
df4 = pd.read_excel('https://static-content.springer.com/esm/art%3A10.1038%2Fs41589-018-0116-2/MediaObjects/41589_2018_116_MOESM33_ESM.xlsx', sheet_name='HeLa', dtype=str)
df4 = df4.assign(ptms=np.full((df4.shape[0], 1), "HeLa-RSOH"))
df4 = df4.rename(index=str, columns={"Site #": "modifiedSite", "Uniprot Accession #": "uniprotAccession"})
dfs.append(df4)
return dfs
# + code_folding=[0]
# concatenate and process dataset
dfs = read_datasets()
df = pd.concat(dfs, ignore_index=True, sort=False)
#display(df)
df = df[['ptms', 'modifiedSite', 'uniprotAccession', 'Description']]
df['modifiedSite'] = df['modifiedSite'].astype(np.int64)
df.head()
# -
# ## Map PTM locations to residues in PDB structures
# + code_folding=[]
# convert Pandas dataframe to a Spark dataframe
spark = SparkSession.builder.appName("CysOxydationProteomicAndStructuralEvidence").getOrCreate()
ds = spark.createDataFrame(df)
ds = ds.sort(ds.uniprotAccession, ds.modifiedSite)
# -
# Download PDB to UniProt mappings and filter out residues that were not observed in the 3D structure.
up = pdbToUniProt.get_cached_residue_mappings().filter("pdbResNum IS NOT NULL")
# Joint PTM with PDB data if the UniProt Id and UniProt residue numbers match
st = up.join(ds, (up.uniprotId == ds.uniprotAccession) & (up.uniprotNum == ds.modifiedSite))
# ## Get PTMs present in 3D Structure of the PDB
# We retrieve oxidated forms of L-cysteine in the PDB using the [PSI-MOD Ontology](https://www.ebi.ac.uk/ols/ontologies/mod)
# * [MOD:00210](https://www.ebi.ac.uk/ols/ontologies/mod/terms?iri=http%3A%2F%2Fpurl.obolibrary.org%2Fobo%2FMOD_00210) - oxydation to L-cysteine sulfenic acid (RSOH)
# * [MOD:00267](https://www.ebi.ac.uk/ols/ontologies/mod/terms?iri=http%3A%2F%2Fpurl.obolibrary.org%2Fobo%2FMOD_00267) - oxydation to L-cysteine sulfinic acid (RSO2H)
# * [MOD:00460](https://www.ebi.ac.uk/ols/ontologies/mod/terms?iri=http%3A%2F%2Fpurl.obolibrary.org%2Fobo%2FMOD_00460) - oxydation to L-cysteine sulfonic acid (RSO3H)
# + code_folding=[0]
# get PTM dataset
pt = pdbPtmDataset.get_ptm_dataset()
pt = pt.filter("psimodId = 'MOD:00210' OR psimodId = 'MOD:00267' OR psimodId = 'MOD:00460'")
print("Total number of oxidized cysteines in PDB: ", pt.count())
pt.limit(5).toPandas()
# -
pt = pt.withColumnRenamed("pdbResNum", "resNum") # avoid two columns with identical names
st = st.join(pt, (st.pdbResNum == pt.resNum) & (st.structureChainId == pt.pdbChainId))
# ## Aggregate PTM data on a per residue and per chain basis
# + code_folding=[0]
# Aggregate data
st = st.groupBy("structureChainId","pdbResNum","uniprotAccession","uniprotNum","Description").agg(collect_list("ptms").alias("ptms"))
st = st.withColumn("ptms", concat_ws((","), col("ptms")))
st = st.groupBy("structureChainId","uniprotAccession","Description").agg(collect_list("ptms").alias("ptms"), collect_list("pdbResNum").alias("pdbResNum"), collect_list("uniprotNum").alias("uniprotNum"))
# + [markdown] code_folding=[]
# Keep only a single structural representative
# -
st = st.drop_duplicates(["uniprotAccession","uniprotNum"])
# ## Show Table with PDB mappings
# PDB residue numbers do not always match UniProt residue numbers. The table below shows the mapping for each protein chain.
# + code_folding=[]
# convert Spark dataframe back to a Pandas dataframe
sp = st.toPandas()
sp.head()
# + code_folding=[0] jupyter={"source_hidden": true}
def view_modifications(df, cutoff_distance, *args):
def view3d(show_labels=True,show_bio_assembly=False, show_surface=False, i=0):
pdb_id, chain_id = df.iloc[i]['structureChainId'].split('.')
res_num = df.iloc[i]['pdbResNum']
labels = df.iloc[i]['ptms']
# print header
print ("PDB Id: " + pdb_id + " chain Id: " + chain_id)
# print any specified additional columns from the dataframe
for a in args:
print(a + ": " + df.iloc[i][a])
mod_res = {'chain': chain_id, 'resi': res_num}
# select neigboring residues by distance
surroundings = {'chain': chain_id, 'resi': res_num, 'byres': True, 'expand': cutoff_distance}
viewer = py3Dmol.view(query='pdb:' + pdb_id, options={'doAssembly': show_bio_assembly})
# polymer style
viewer.setStyle({'cartoon': {'color': 'spectrum', 'width': 0.6, 'opacity':0.8}})
# non-polymer style
viewer.setStyle({'hetflag': True}, {'stick':{'radius': 0.3, 'singleBond': False}})
# style for modifications
viewer.addStyle(surroundings,{'stick':{'colorscheme':'orangeCarbon', 'radius': 0.15}})
viewer.addStyle(mod_res, {'stick':{'colorscheme':'redCarbon', 'radius': 0.4}})
viewer.addStyle(mod_res, {'sphere':{'colorscheme':'gray', 'opacity': 0.7}})
# set residue labels
if show_labels:
for residue, label in zip(res_num, labels):
viewer.addLabel(residue + ": " + label, \
{'fontColor':'black', 'fontSize': 9, 'backgroundColor': 'lightgray'}, \
{'chain': chain_id, 'resi': residue})
viewer.zoomTo(surroundings)
if show_surface:
viewer.addSurface(py3Dmol.SES,{'opacity':0.8,'color':'lightblue'})
return viewer.show()
s_widget = IntSlider(min=0, max=len(df)-1, description='Structure', continuous_update=False)
return interact(view3d, show_labels=True, show_bio_assembly=False, show_surface=False, i=s_widget)
# -
# ## Visualize Results
# Residues with reported modifications are shown in an all atom prepresentation as red sticks with transparent spheres. Each modified residue position is labeled by the PDB residue number and the type of the modification. Residues surrounding modified residue (within 6 A) are highlighted as yellow sticks. Small molecules within the structure are rendered as gray sticks.
#
# * Move slider to browse through the results
# * To rotate the structure, hold down the left mouse button and move the mouse.
view_modifications(sp, 6, 'uniprotAccession', 'Description');
# ## Matching evidence was found for the following proteins
rs = sp[['uniprotAccession', 'Description']].drop_duplicates()
display(rs)
spark.stop()
| notebooks/CysOxidationProteomicAndStructuralEvidence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classifying Yelp Reviews
# ## Imports
# +
from argparse import Namespace
from collections import Counter
import json
import os
import re
import string
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
from tqdm import tqdm_notebook
# -
# ## Data Vectorization classes
# + [markdown] heading_collapsed=true
# ### The Vocabulary
# + code_folding=[] hidden=true
class Vocabulary(object):
"""Class to process text and extract vocabulary for mapping"""
def __init__(self, token_to_idx=None, add_unk=True, unk_token="<UNK>"):
"""
Args:
token_to_idx (dict): a pre-existing map of tokens to indices
add_unk (bool): a flag that indicates whether to add the UNK token
unk_token (str): the UNK token to add into the Vocabulary
"""
if token_to_idx is None:
token_to_idx = {}
self._token_to_idx = token_to_idx
self._idx_to_token = {idx: token
for token, idx in self._token_to_idx.items()}
self._add_unk = add_unk
self._unk_token = unk_token
self.unk_index = -1
if add_unk:
self.unk_index = self.add_token(unk_token)
def to_serializable(self):
""" returns a dictionary that can be serialized """
return {'token_to_idx': self._token_to_idx,
'add_unk': self._add_unk,
'unk_token': self._unk_token}
@classmethod
def from_serializable(cls, contents):
""" instantiates the Vocabulary from a serialized dictionary """
return cls(**contents)
def add_token(self, token):
"""Update mapping dicts based on the token.
Args:
token (str): the item to add into the Vocabulary
Returns:
index (int): the integer corresponding to the token
"""
if token in self._token_to_idx:
index = self._token_to_idx[token]
else:
index = len(self._token_to_idx)
self._token_to_idx[token] = index
self._idx_to_token[index] = token
return index
def add_many(self, tokens):
"""Add a list of tokens into the Vocabulary
Args:
tokens (list): a list of string tokens
Returns:
indices (list): a list of indices corresponding to the tokens
"""
return [self.add_token(token) for token in tokens]
def lookup_token(self, token):
"""Retrieve the index associated with the token
or the UNK index if token isn't present.
Args:
token (str): the token to look up
Returns:
index (int): the index corresponding to the token
Notes:
`unk_index` needs to be >=0 (having been added into the Vocabulary)
for the UNK functionality
"""
if self.unk_index >= 0:
return self._token_to_idx.get(token, self.unk_index)
else:
return self._token_to_idx[token]
def lookup_index(self, index):
"""Return the token associated with the index
Args:
index (int): the index to look up
Returns:
token (str): the token corresponding to the index
Raises:
KeyError: if the index is not in the Vocabulary
"""
if index not in self._idx_to_token:
raise KeyError("the index (%d) is not in the Vocabulary" % index)
return self._idx_to_token[index]
def __str__(self):
return "<Vocabulary(size=%d)>" % len(self)
def __len__(self):
return len(self._token_to_idx)
# + [markdown] heading_collapsed=true
# ### The Vectorizer
# + code_folding=[] hidden=true
class ReviewVectorizer(object):
""" The Vectorizer which coordinates the Vocabularies and puts them to use"""
def __init__(self, review_vocab, rating_vocab):
"""
Args:
review_vocab (Vocabulary): maps words to integers
rating_vocab (Vocabulary): maps class labels to integers
"""
self.review_vocab = review_vocab
self.rating_vocab = rating_vocab
def vectorize(self, review):
"""Create a collapsed one-hit vector for the review
Args:
review (str): the review
Returns:
one_hot (np.ndarray): the collapsed one-hot encoding
"""
one_hot = np.zeros(len(self.review_vocab), dtype=np.float32)
for token in review.split(" "):
if token not in string.punctuation:
one_hot[self.review_vocab.lookup_token(token)] = 1
return one_hot
@classmethod
def from_dataframe(cls, review_df, cutoff=25):
"""Instantiate the vectorizer from the dataset dataframe
Args:
review_df (pandas.DataFrame): the review dataset
cutoff (int): the parameter for frequency-based filtering
Returns:
an instance of the ReviewVectorizer
"""
review_vocab = Vocabulary(add_unk=True)
rating_vocab = Vocabulary(add_unk=False)
# Add ratings
for rating in sorted(set(review_df.rating)):
rating_vocab.add_token(rating)
# Add top words if count > provided count
word_counts = Counter()
for review in review_df.review:
for word in review.split(" "):
if word not in string.punctuation:
word_counts[word] += 1
for word, count in word_counts.items():
if count > cutoff:
review_vocab.add_token(word)
return cls(review_vocab, rating_vocab)
@classmethod
def from_serializable(cls, contents):
"""Instantiate a ReviewVectorizer from a serializable dictionary
Args:
contents (dict): the serializable dictionary
Returns:
an instance of the ReviewVectorizer class
"""
review_vocab = Vocabulary.from_serializable(contents['review_vocab'])
rating_vocab = Vocabulary.from_serializable(contents['rating_vocab'])
return cls(review_vocab=review_vocab, rating_vocab=rating_vocab)
def to_serializable(self):
"""Create the serializable dictionary for caching
Returns:
contents (dict): the serializable dictionary
"""
return {'review_vocab': self.review_vocab.to_serializable(),
'rating_vocab': self.rating_vocab.to_serializable()}
# + [markdown] heading_collapsed=true
# ### The Dataset
# + code_folding=[65] hidden=true
class ReviewDataset(Dataset):
def __init__(self, review_df, vectorizer):
"""
Args:
review_df (pandas.DataFrame): the dataset
vectorizer (ReviewVectorizer): vectorizer instantiated from dataset
"""
self.review_df = review_df
self._vectorizer = vectorizer
self.train_df = self.review_df[self.review_df.split=='train']
self.train_size = len(self.train_df)
self.val_df = self.review_df[self.review_df.split=='val']
self.validation_size = len(self.val_df)
self.test_df = self.review_df[self.review_df.split=='test']
self.test_size = len(self.test_df)
self._lookup_dict = {'train': (self.train_df, self.train_size),
'val': (self.val_df, self.validation_size),
'test': (self.test_df, self.test_size)}
self.set_split('train')
@classmethod
def load_dataset_and_make_vectorizer(cls, review_csv):
"""Load dataset and make a new vectorizer from scratch
Args:
review_csv (str): location of the dataset
Returns:
an instance of ReviewDataset
"""
review_df = pd.read_csv(review_csv)
train_review_df = review_df[review_df.split=='train']
return cls(review_df, ReviewVectorizer.from_dataframe(train_review_df))
@classmethod
def load_dataset_and_load_vectorizer(cls, review_csv, vectorizer_filepath):
"""Load dataset and the corresponding vectorizer.
Used in the case in the vectorizer has been cached for re-use
Args:
review_csv (str): location of the dataset
vectorizer_filepath (str): location of the saved vectorizer
Returns:
an instance of ReviewDataset
"""
review_df = pd.read_csv(review_csv)
vectorizer = cls.load_vectorizer_only(vectorizer_filepath)
return cls(review_df, vectorizer)
@staticmethod
def load_vectorizer_only(vectorizer_filepath):
"""a static method for loading the vectorizer from file
Args:
vectorizer_filepath (str): the location of the serialized vectorizer
Returns:
an instance of ReviewVectorizer
"""
with open(vectorizer_filepath) as fp:
return ReviewVectorizer.from_serializable(json.load(fp))
def save_vectorizer(self, vectorizer_filepath):
"""saves the vectorizer to disk using json
Args:
vectorizer_filepath (str): the location to save the vectorizer
"""
with open(vectorizer_filepath, "w") as fp:
json.dump(self._vectorizer.to_serializable(), fp)
def get_vectorizer(self):
""" returns the vectorizer """
return self._vectorizer
def set_split(self, split="train"):
""" selects the splits in the dataset using a column in the dataframe
Args:
split (str): one of "train", "val", or "test"
"""
self._target_split = split
self._target_df, self._target_size = self._lookup_dict[split]
def __len__(self):
return self._target_size
def __getitem__(self, index):
"""the primary entry point method for PyTorch datasets
Args:
index (int): the index to the data point
Returns:
a dictionary holding the data point's features (x_data) and label (y_target)
"""
row = self._target_df.iloc[index]
review_vector = \
self._vectorizer.vectorize(row.review)
rating_index = \
self._vectorizer.rating_vocab.lookup_token(row.rating)
return {'x_data': review_vector,
'y_target': rating_index}
def get_num_batches(self, batch_size):
"""Given a batch size, return the number of batches in the dataset
Args:
batch_size (int)
Returns:
number of batches in the dataset
"""
return len(self) // batch_size
def generate_batches(dataset, batch_size, shuffle=True,
drop_last=True, device="cpu"):
"""
A generator function which wraps the PyTorch DataLoader. It will
ensure each tensor is on the write device location.
"""
dataloader = DataLoader(dataset=dataset, batch_size=batch_size,
shuffle=shuffle, drop_last=drop_last)
for data_dict in dataloader:
out_data_dict = {}
for name, tensor in data_dict.items():
out_data_dict[name] = data_dict[name].to(device)
yield out_data_dict
# + [markdown] heading_collapsed=true
# ## The Model: ReviewClassifier
# + code_folding=[] hidden=true
class ReviewClassifier(nn.Module):
""" a simple perceptron based classifier """
def __init__(self, num_features):
"""
Args:
num_features (int): the size of the input feature vector
"""
super(ReviewClassifier, self).__init__()
self.fc1 = nn.Linear(in_features=num_features,
out_features=1)
def forward(self, x_in, apply_sigmoid=False):
"""The forward pass of the classifier
Args:
x_in (torch.Tensor): an input data tensor.
x_in.shape should be (batch, num_features)
apply_sigmoid (bool): a flag for the sigmoid activation
should be false if used with the Cross Entropy losses
Returns:
the resulting tensor. tensor.shape should be (batch,)
"""
y_out = self.fc1(x_in).squeeze()
if apply_sigmoid:
y_out = torch.sigmoid(y_out)
return y_out
# -
# ## Training Routine
# ### Helper functions
# + code_folding=[]
def make_train_state(args):
return {'stop_early': False,
'early_stopping_step': 0,
'early_stopping_best_val': 1e8,
'learning_rate': args.learning_rate,
'epoch_index': 0,
'train_loss': [],
'train_acc': [],
'val_loss': [],
'val_acc': [],
'test_loss': -1,
'test_acc': -1,
'model_filename': args.model_state_file}
def update_train_state(args, model, train_state):
"""Handle the training state updates.
Components:
- Early Stopping: Prevent overfitting.
- Model Checkpoint: Model is saved if the model is better
:param args: main arguments
:param model: model to train
:param train_state: a dictionary representing the training state values
:returns:
a new train_state
"""
# Save one model at least
if train_state['epoch_index'] == 0:
torch.save(model.state_dict(), train_state['model_filename'])
train_state['stop_early'] = False
# Save model if performance improved
elif train_state['epoch_index'] >= 1:
loss_tm1, loss_t = train_state['val_loss'][-2:]
# If loss worsened
if loss_t >= train_state['early_stopping_best_val']:
# Update step
train_state['early_stopping_step'] += 1
# Loss decreased
else:
# Save the best model
if loss_t < train_state['early_stopping_best_val']:
torch.save(model.state_dict(), train_state['model_filename'])
# Reset early stopping step
train_state['early_stopping_step'] = 0
# Stop early ?
train_state['stop_early'] = \
train_state['early_stopping_step'] >= args.early_stopping_criteria
return train_state
def compute_accuracy(y_pred, y_target):
y_target = y_target.cpu()
y_pred_indices = (torch.sigmoid(y_pred)>0.5).cpu().long()#.max(dim=1)[1]
n_correct = torch.eq(y_pred_indices, y_target).sum().item()
return n_correct / len(y_pred_indices) * 100
# -
# #### General utilities
# +
def set_seed_everywhere(seed, cuda):
np.random.seed(seed)
torch.manual_seed(seed)
if cuda:
torch.cuda.manual_seed_all(seed)
def handle_dirs(dirpath):
if not os.path.exists(dirpath):
os.makedirs(dirpath)
# -
# ### Settings and some prep work
# +
args = Namespace(
# Data and Path information
frequency_cutoff=25,
model_state_file='model.pth',
review_csv='data/yelp/reviews_with_splits_lite.csv',
# review_csv='data/yelp/reviews_with_splits_full.csv',
save_dir='model_storage/ch3/yelp/',
vectorizer_file='vectorizer.json',
# No Model hyper parameters
# Training hyper parameters
batch_size=128,
early_stopping_criteria=5,
learning_rate=0.001,
num_epochs=100,
seed=1337,
# Runtime options
catch_keyboard_interrupt=True,
cuda=True,
expand_filepaths_to_save_dir=True,
reload_from_files=False,
)
if args.expand_filepaths_to_save_dir:
args.vectorizer_file = os.path.join(args.save_dir,
args.vectorizer_file)
args.model_state_file = os.path.join(args.save_dir,
args.model_state_file)
print("Expanded filepaths: ")
print("\t{}".format(args.vectorizer_file))
print("\t{}".format(args.model_state_file))
# Check CUDA
if not torch.cuda.is_available():
args.cuda = False
print("Using CUDA: {}".format(args.cuda))
args.device = torch.device("cuda" if args.cuda else "cpu")
# Set seed for reproducibility
set_seed_everywhere(args.seed, args.cuda)
# handle dirs
handle_dirs(args.save_dir)
# -
# ### Initializations
# +
if args.reload_from_files:
# training from a checkpoint
print("Loading dataset and vectorizer")
dataset = ReviewDataset.load_dataset_and_load_vectorizer(args.review_csv,
args.vectorizer_file)
else:
print("Loading dataset and creating vectorizer")
# create dataset and vectorizer
dataset = ReviewDataset.load_dataset_and_make_vectorizer(args.review_csv)
dataset.save_vectorizer(args.vectorizer_file)
vectorizer = dataset.get_vectorizer()
classifier = ReviewClassifier(num_features=len(vectorizer.review_vocab))
# -
# ### Training loop
# + code_folding=[30]
classifier = classifier.to(args.device)
loss_func = nn.BCEWithLogitsLoss()
optimizer = optim.Adam(classifier.parameters(), lr=args.learning_rate)
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer=optimizer,
mode='min', factor=0.5,
patience=1)
train_state = make_train_state(args)
epoch_bar = tqdm_notebook(desc='training routine',
total=args.num_epochs,
position=0)
dataset.set_split('train')
train_bar = tqdm_notebook(desc='split=train',
total=dataset.get_num_batches(args.batch_size),
position=1,
leave=True)
dataset.set_split('val')
val_bar = tqdm_notebook(desc='split=val',
total=dataset.get_num_batches(args.batch_size),
position=1,
leave=True)
try:
for epoch_index in range(args.num_epochs):
train_state['epoch_index'] = epoch_index
# Iterate over training dataset
# setup: batch generator, set loss and acc to 0, set train mode on
dataset.set_split('train')
batch_generator = generate_batches(dataset,
batch_size=args.batch_size,
device=args.device)
running_loss = 0.0
running_acc = 0.0
classifier.train()
for batch_index, batch_dict in enumerate(batch_generator):
# the training routine is these 5 steps:
# --------------------------------------
# step 1. zero the gradients
optimizer.zero_grad()
# step 2. compute the output
y_pred = classifier(x_in=batch_dict['x_data'].float())
# step 3. compute the loss
loss = loss_func(y_pred, batch_dict['y_target'].float())
loss_t = loss.item()
running_loss += (loss_t - running_loss) / (batch_index + 1)
# step 4. use loss to produce gradients
loss.backward()
# step 5. use optimizer to take gradient step
optimizer.step()
# -----------------------------------------
# compute the accuracy
acc_t = compute_accuracy(y_pred, batch_dict['y_target'])
running_acc += (acc_t - running_acc) / (batch_index + 1)
# update bar
train_bar.set_postfix(loss=running_loss,
acc=running_acc,
epoch=epoch_index)
train_bar.update()
train_state['train_loss'].append(running_loss)
train_state['train_acc'].append(running_acc)
# Iterate over val dataset
# setup: batch generator, set loss and acc to 0; set eval mode on
dataset.set_split('val')
batch_generator = generate_batches(dataset,
batch_size=args.batch_size,
device=args.device)
running_loss = 0.
running_acc = 0.
classifier.eval()
for batch_index, batch_dict in enumerate(batch_generator):
# compute the output
y_pred = classifier(x_in=batch_dict['x_data'].float())
# step 3. compute the loss
loss = loss_func(y_pred, batch_dict['y_target'].float())
loss_t = loss.item()
running_loss += (loss_t - running_loss) / (batch_index + 1)
# compute the accuracy
acc_t = compute_accuracy(y_pred, batch_dict['y_target'])
running_acc += (acc_t - running_acc) / (batch_index + 1)
val_bar.set_postfix(loss=running_loss,
acc=running_acc,
epoch=epoch_index)
val_bar.update()
train_state['val_loss'].append(running_loss)
train_state['val_acc'].append(running_acc)
train_state = update_train_state(args=args, model=classifier,
train_state=train_state)
scheduler.step(train_state['val_loss'][-1])
train_bar.n = 0
val_bar.n = 0
epoch_bar.update()
if train_state['stop_early']:
break
train_bar.n = 0
val_bar.n = 0
epoch_bar.update()
except KeyboardInterrupt:
print("Exiting loop")
# +
# compute the loss & accuracy on the test set using the best available model
classifier.load_state_dict(torch.load(train_state['model_filename']))
classifier = classifier.to(args.device)
dataset.set_split('test')
batch_generator = generate_batches(dataset,
batch_size=args.batch_size,
device=args.device)
running_loss = 0.
running_acc = 0.
classifier.eval()
for batch_index, batch_dict in enumerate(batch_generator):
# compute the output
y_pred = classifier(x_in=batch_dict['x_data'].float())
# compute the loss
loss = loss_func(y_pred, batch_dict['y_target'].float())
loss_t = loss.item()
running_loss += (loss_t - running_loss) / (batch_index + 1)
# compute the accuracy
acc_t = compute_accuracy(y_pred, batch_dict['y_target'])
running_acc += (acc_t - running_acc) / (batch_index + 1)
train_state['test_loss'] = running_loss
train_state['test_acc'] = running_acc
# -
print("Test loss: {:.3f}".format(train_state['test_loss']))
print("Test Accuracy: {:.2f}".format(train_state['test_acc']))
# ### Inference
def preprocess_text(text):
text = text.lower()
text = re.sub(r"([.,!?])", r" \1 ", text)
text = re.sub(r"[^a-zA-Z.,!?]+", r" ", text)
return text
def predict_rating(review, classifier, vectorizer, decision_threshold=0.5):
"""Predict the rating of a review
Args:
review (str): the text of the review
classifier (ReviewClassifier): the trained model
vectorizer (ReviewVectorizer): the corresponding vectorizer
decision_threshold (float): The numerical boundary which separates the rating classes
"""
review = preprocess_text(review)
vectorized_review = torch.tensor(vectorizer.vectorize(review))
result = classifier(vectorized_review.view(1, -1))
probability_value = F.sigmoid(result).item()
index = 1
if probability_value < decision_threshold:
index = 0
return vectorizer.rating_vocab.lookup_index(index)
# +
test_review = "this is a pretty awesome book"
classifier = classifier.cpu()
prediction = predict_rating(test_review, classifier, vectorizer, decision_threshold=0.5)
print("{} -> {}".format(test_review, prediction))
# -
# ### Interpretability
classifier.fc1.weight.shape
# +
# Sort weights
fc1_weights = classifier.fc1.weight.detach()[0]
_, indices = torch.sort(fc1_weights, dim=0, descending=True)
indices = indices.numpy().tolist()
# Top 20 words
print("Influential words in Positive Reviews:")
print("--------------------------------------")
for i in range(20):
print(vectorizer.review_vocab.lookup_index(indices[i]))
print("====\n\n\n")
# Top 20 negative words
print("Influential words in Negative Reviews:")
print("--------------------------------------")
indices.reverse()
for i in range(20):
print(vectorizer.review_vocab.lookup_index(indices[i]))
# -
# ### End
| chapters/chapter_3/3_5_Classifying_Yelp_Review_Sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# ## Vetores e Matrizes
import numpy as np
A=np.array([[5.90,7.10],[2.38,2.52],[2.90,2.00]])
B=4.10*A
print(B)
# ### Dados os vetores u=(3,-5,2) e v=(4,6,3), calcule u.v e uXv
import numpy as np
u=np.array([[3,-5,2]])
v=np.array([[4,6,3]])
uv=np.inner(u,v)
uXv=np.cross(u,v)
print(uv)
print(uXv)
import numpy as np
u=np.array([[7,1,-9]])
v=np.array([[3,-5,-4]])
uv=np.inner(u,v)
uXv=np.cross(u,v)
print(uv)
print(uXv)
# ## Sistema Lineares
import numpy as np
A=np.array([[4,2,-1],[3,3,2],[0,5,2]])
b=np.array([[7],[20],[-1]])
x=np.linalg.solve(A,b)
print(x)
import numpy as np
A=np.array([[4,-3,1],[1,1,3],[2,3,-4]])
b=np.array([[15],[27],[31]])
x=np.linalg.solve(A,b)
print(x)
import numpy as np
A=np.array([[5,1,3],[-1,2,5],[4,-5,1]])
b=np.array([[76],[35],[22]])
x=np.linalg.solve(A,b)
print(x)
A=np.array([[1,3,4],[2,-1,1],[-4,2,-2]])
b=np.array([[18],[10],[-7]])
x=np.linalg.solve(A,b)
print(x)
# #### É UMA MATRIZ SINGULAR (NÃO TEM SOLUÇÃO)
import numpy as np
A=np.array([[10,120],[8,80]])
b=np.array([[844],[576]])
x=np.linalg.solve(A,b)
print(x)
# # Funções Trigonomêtricas
# ### Qual é o seno de 57° ?
import numpy as np
arco=np.deg2rad(57)
np.sin(arco)
# ### Qual é o ângulo cujo cosseno é igual a 0,7?
import numpy as np
angulo=np.arccos(0.7)
np.rad2deg(angulo)
# # Números complexos
# #### Obs: separando raiz em duas
# Número Complexo na forma algébrica: z=a+bi
# a=parte real:
# a=Re(z)
#
#
#
#
# b=parte imaginária:
# b=Im(z)
# ### Escreva o número complexo z=3+5j utilizando o Python
z=complex(3,5)
print(z)
# ### Obtenha o módulo do número complexo z=3+5j
z=complex(3,5)
abs(z)
z1=complex(3,5)
z2=complex(7,-3)
soma=z1+z2
produto=z1*z2
divisao=z1/z2
print(soma)
print(produto)
print(divisao)
# # Sistemas Lineares com números complexos
import numpy as np
A=np.array([[complex(2,1),complex(5,-1)],[complex(5,1),complex(7,4)]])
b=np.array([[complex(3,8)],[complex(2,5)]])
np.linalg.solve(A,b)
import numpy as np
A=np.array([[complex(3,2),complex(-2,-6)],[complex(-2,-6),complex(10,1)]])
x1=np.cos(np.deg2rad(0))
y1=np.sin(np.deg2rad(0))
x2=np.cos(np.deg2rad(30))
y2=np.sin(np.deg2rad(30))
b=np.array([[70*complex(x1,y1)],[110*complex(x2,y2)]])
np.linalg.solve(A,b)
import numpy as np
A=np.array([[complex(1,5),complex(3,-2)],[complex(3,-2),complex(4,1)]])
x1=np.cos(np.deg2rad(23))
y1=np.sin(np.deg2rad(37))
b=np.array([[30*complex(x1,y1)],[52*complex(x2,y2)]])
np.linalg.solve(A,b)
| Operations/Vetores,Matrizes,Sistemas Lineares.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="MhoQ0WE77laV"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="_ckMIh7O7s6D"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" colab={} colab_type="code" id="vasWnqRgy1H4"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] colab_type="text" id="jYysdyb-CaWM"
# # Regressão: preveja consumo de combustível
# + [markdown] colab_type="text" id="S5Uhzt6vVIB2"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/pt-br/r1/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Execute em Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/pt-br/r1/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Veja a fonte em GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="UA1pKuH8wqxz"
# Note: A nossa comunidade TensorFlow traduziu estes documentos. Como as traduções da comunidade são *o melhor esforço*, não há garantias de que sejam uma reflexão exata e atualizada da [documentação oficial em Inglês](https://www.tensorflow.org/?hl=en). Se tem alguma sugestão para melhorar esta tradução, por favor envie um pull request para o repositório do GitHub [tensorflow/docs](https://github.com/tensorflow/docs). Para se voluntariar para escrever ou rever as traduções da comunidade, contacte a [lista <EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs).
# + [markdown] colab_type="text" id="OezgXCpHi4v_"
# Em um problema de regressão, o objetivo é prever as saídas (*outputs*) de um valor contínuo, como um preço ou probabilidade. Em contraste de problemas de classificação, onde temos o propósito de escolher uma classe em uma lista de classificações (por exemplo, se uma imagem contém uma maçã ou laranja, assim reconhecendo qual fruta é representada na imagem).
#
# Este *notebook* usa a clássica base de dados [Auto MPG](https://archive.ics.uci.edu/ml/datasets/auto+mpg) e constrói um modelo para prever a economia de combustíveis de automóveis do final dos anos 1970, início dos anos 1980. Para isso, forneceremos um modelo com descrição de vários automóveis desse período. Essa descrição inclui atributos como: cilindros, deslocamento, potência do motor, e peso.
#
# Este exemplo usa a API `tf.keras`. Veja [este guia](https://www.tensorflow.org/r1/guide/keras) para mais detalhes.
# + colab={} colab_type="code" id="dzLKpmZICaWN"
# Use seaborn para pairplot
# !pip install seaborn
# + colab={} colab_type="code" id="gOL_C-OkBKva"
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import tensorflow.compat.v1 as tf
from tensorflow import keras
from tensorflow.keras import layers
print(tf.__version__)
# + [markdown] colab_type="text" id="yR0EdgrLCaWR"
# ## Base de dados Auto MPG
#
# A base de dados está disponível em [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/).
#
# + [markdown] colab_type="text" id="DLdCchMdCaWQ"
# ### Pegando os dados
# Primeiro baixe a base de dados dos automóveis.
# + colab={} colab_type="code" id="7MqDQO0KCaWS"
dataset_path = keras.utils.get_file("auto-mpg.data", "http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
dataset_path
# + [markdown] colab_type="text" id="t9FDsUlxCaWW"
# Utilizando o pandas, impoorte os dados:
# + colab={} colab_type="code" id="IjnLH5S2CaWx"
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Model Year', 'Origin']
raw_dataset = pd.read_csv(dataset_path, names=column_names,
na_values = "?", comment='\t',
sep=" ", skipinitialspace=True)
dataset = raw_dataset.copy()
dataset.tail()
# + [markdown] colab_type="text" id="Brm0b_KACaWX"
# ### Limpe os dados
#
# Esta base contém alguns valores não conhecidos (*unknown*).
# + colab={} colab_type="code" id="zW5k_xz1CaWX"
dataset.isna().sum()
# + [markdown] colab_type="text" id="cIAcvQqMCaWf"
# Para manter esse tutorial básico, remova as linhas com esses valores não conhecidos.
# + colab={} colab_type="code" id="TRFYHB2mCaWb"
dataset = dataset.dropna()
# + [markdown] colab_type="text" id="YSlYxFuRCaWk"
# A coluna "Origin" é uma coluna categórica e não numérica. Logo converta para *one-hot* :
# + colab={} colab_type="code" id="XKnCTHz4CaWg"
origin = dataset.pop('Origin')
# + colab={} colab_type="code" id="X_PB-wCUHgxU"
dataset['USA'] = (origin == 1)*1.0
dataset['Europe'] = (origin == 2)*1.0
dataset['Japan'] = (origin == 3)*1.0
dataset.tail()
# + [markdown] colab_type="text" id="TMPI88iZpO2T"
# ### Separando dados de treinamento e teste
#
# Agora separe os dados em um conjunto de treinamento e outro teste.
#
# Iremos utilizar o de conjunto de teste no final da análise do model.
# + colab={} colab_type="code" id="2KFnYlcwCaWl"
train_dataset = dataset.sample(frac=0.8,random_state=0)
test_dataset = dataset.drop(train_dataset.index)
# + [markdown] colab_type="text" id="rd0A0Iu0CaWq"
# ### Inspecione o dado
#
# Dê uma rápida olhada em como está a distribuição de algumas colunas do conjunto de treinamento.
# + colab={} colab_type="code" id="iJmPr5-ACaWn"
sns.pairplot(train_dataset[["MPG", "Cylinders", "Displacement", "Weight"]], diag_kind="kde")
# + [markdown] colab_type="text" id="ES6uQoLKCaWr"
# Repare na visão geral dos estatísticas:
# + colab={} colab_type="code" id="m4VEw8Ud9Quh"
train_stats = train_dataset.describe()
train_stats.pop("MPG")
train_stats = train_stats.transpose()
train_stats
# + [markdown] colab_type="text" id="Wz7l27Lz9S1P"
# ### Separe features de labels
#
# Separe o valor alvo (*labels*), das *features*. Essa label é o valor no qual o modelo é treinado para prever.
# + colab={} colab_type="code" id="bW5WzIPlCaWv"
train_labels = train_dataset.pop('MPG')
test_labels = test_dataset.pop('MPG')
# + [markdown] colab_type="text" id="Ee638AlnCaWz"
# ### Normalize os dados
#
# Observe novamente o `train_stats` acima e note quão diferente são os intervalos de uma feature e outra.
# + [markdown] colab_type="text" id="cVdIpzAdnxew"
# Uma boa prática é normalizar as *features* que usam diferentes escalas e intervalos. Apesar do modelo poder convergir sem a normalização, isso torna o treinamento mais difícil, e torna o resultado do modelo dependente da escolha das unidades da entrada.
#
# Observação: embora geramos intencionalmente essas estatísticas para os dados de treinamento, essas estatísticas serão usadas também para normalizar o conjunto de teste. Precisamos delinear o conjunto de teste na mesma distribuição que o modelo foi treinado.
# + colab={} colab_type="code" id="oZTImqg_CaW1"
def norm(x):
return (x - train_stats['mean']) / train_stats['std']
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
# + [markdown] colab_type="text" id="JoBpCiVPwwck"
# Esse dado normalizado é o que usaremos para treinar o modelo.
#
# Atenção: As estatísticas usadas para normalizar as entradas aqui (média e desvio padrão) precisa ser aplicada em qualquer outro dado que alimenta o modelo, junto com o código *one-hot* que fizemos anteriormente. Isso inclui o conjunto de teste e os dados que o modelo usará em produção.
# + [markdown] colab_type="text" id="59veuiEZCaW4"
# ## O Modelo
#
# + [markdown] colab_type="text" id="Gxg1XGm0eOBy"
# ### Construindo o modelo
#
# Vamos construir o modelo. Aqui usaremos o modelo `Sequential` com duas camadas *densely connected*, e a camada de saída que retorna um único valor contínuo. Os passos de construção do modelo são agrupados em uma função, build_model, já que criaremos um segundo modelo mais tarde.
# + colab={} colab_type="code" id="9ODch-OFCaW4"
def build_model():
model = keras.Sequential([
layers.Dense(64, activation=tf.nn.relu, input_shape=[len(train_dataset.keys())]),
layers.Dense(64, activation=tf.nn.relu),
layers.Dense(1)
])
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=['mean_absolute_error', 'mean_squared_error'])
return model
# + colab={} colab_type="code" id="Lhan11blCaW7"
model = build_model()
# + [markdown] colab_type="text" id="qKF6uW-BCaW-"
# ## Examine o modelo
#
# Use o método `.summary` para exibir uma descrição simples do modelo.
# + colab={} colab_type="code" id="xvwvpA64CaW_"
model.summary()
# + [markdown] colab_type="text" id="W3ZVOhugCaXA"
# Agora teste o modelo. Pegue um batch de de 10 exemplos do conjunto de treinamento e chame `model.predict`nestes.
# + colab={} colab_type="code" id="PcJrGsO5hZzK"
example_batch = normed_train_data[:10]
example_result = model.predict(example_batch)
example_result
# + [markdown] colab_type="text" id="oEw4bZgGCaXB"
# Parece que está funcionando e ele produz o resultado de forma e tipo esperados.
# + [markdown] colab_type="text" id="yWfgsmVXCaXG"
# ### Treinando o modelo
#
# Treine o modelo com 1000 *epochs*, e grave a acurácia do treinamento e da validação em um objeto `history`.
# + colab={} colab_type="code" id="a_MWJBuaC8EM"
# Mostra o progresso do treinamento imprimindo um único ponto para cada epoch completada
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 100 == 0: print('')
print('.', end='')
EPOCHS = 1000
history = model.fit(
normed_train_data, train_labels,
epochs=EPOCHS, validation_split = 0.2, verbose=0,
callbacks=[PrintDot()])
# + [markdown] colab_type="text" id="OgQZyNaWEALn"
# Visualize o progresso do modelo de treinamento usando o estados armazenados no objeto `history`
# + colab={} colab_type="code" id="Gl91RPhdCaXI"
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
# + colab={} colab_type="code" id="dsKd_b-ZEfKe"
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Abs Error [MPG]')
plt.plot(hist['epoch'], hist['mean_absolute_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mean_absolute_error'],
label = 'Val Error')
plt.ylim([0,5])
plt.legend()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Square Error [$MPG^2$]')
plt.plot(hist['epoch'], hist['mean_squared_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mean_squared_error'],
label = 'Val Error')
plt.ylim([0,20])
plt.legend()
plt.show()
plot_history(history)
# + [markdown] colab_type="text" id="x9Kk1voUCaXJ"
# Este grafo mostra as pequenas melhoras, ou mesmo a diminuição do `validation error` após 100 *epochs*. Vamos atualizar o `model.fit` para que pare automatixamente o treinamento quando o `validation score` não aumentar mais. Usaremos o `EarlyStopping callback` que testa a condição do treinamento a cada `epoch`. Se um grupo de `epochs` decorre sem mostrar melhoras, o treinamento irá parar automaticamente.
#
# Você pode aprender mais sobre este callback [aqui](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping)
# + colab={} colab_type="code" id="fvV-BxG6FiaZ"
model = build_model()
# The patience parameter is the amount of epochs to check for improvement
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
history = model.fit(normed_train_data, train_labels, epochs=EPOCHS,
validation_split = 0.2, verbose=0, callbacks=[early_stop, PrintDot()])
plot_history(history)
# + [markdown] colab_type="text" id="-hw1hgeSCaXN"
# O gráfico mostra que no conjunto de validação, a média de erro é próximo de +/- 2MPG. Isso é bom? Deixaremos essa decisão a você.
#
# Vamos ver quão bem o modelo generaliza usando o conjunto de **teste**, que não usamos para treinar o modelo. Isso diz quão bem podemos esperar que o modelo se saia quando usarmos na vida real.
# + colab={} colab_type="code" id="prh4jLRTJ_Rc"
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=2)
print("Testing set Mean Abs Error: {:5.2f} MPG".format(mae))
# + [markdown] colab_type="text" id="3IfAjv3BJleB"
# ### Make predictions
# Finalmente, prevejamos os valores MPG usando o conjunto de teste.
# + colab={} colab_type="code" id="qsqenuPnCaXO"
test_predictions = model.predict(normed_test_data).flatten()
plt.scatter(test_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
plt.axis('equal')
plt.axis('square')
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100, 100], [-100, 100])
# + [markdown] colab_type="text" id="E51yS7iCCaXO"
# Parece que o nosso modelo prediz razoavelmente bem. Vamos dar uma olhada na distribuição dos erros.
# + colab={} colab_type="code" id="Sd7Pgsu6CaXP"
error = test_predictions - test_labels
plt.hist(error, bins = 25)
plt.xlabel("Prediction Error [MPG]")
_ = plt.ylabel("Count")
# + [markdown] colab_type="text" id="ygh2yYC972ne"
# Não é tão gaussiana, porém podemos esperar que por conta do número de exemplo é bem pequeno.
# + [markdown] colab_type="text" id="YFc2HbEVCaXd"
# ## Conclusão
#
# Este notebook introduz algumas técnicas para trabalhar com problema de regressão.
#
# * Mean Sqaured Error(MSE), é uma função comum de *loss* usada para problemas de regressão (diferentes funçẽso de *loss* são usadas para problemas de classificação).
# * Similarmente, as métricas de evolução usadas na regressão são diferentes da classificação. Uma métrica comum de regressão é Mean Absolute Error (MAE).
# * Quando o dado de entrada de *features* tem diferentes intervalos, cada *feature* deve ser escalada para o mesmo intervalo.
# * Se não possui muitos dados de treinamento, uma técnica é preferir uma pequena rede com poucas camadas para evitar *overfitting*.
# * *Early stopping* é uma boa técnica para evitar *overfitting*.
| site/pt-br/r1/tutorials/keras/basic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# To install the [bonobo](https://www.bonobo-project.org/) jupyter extension: https://www.bonobo-project.org/with/jupyter
# +
import bonobo
import pandas as pd
def get_csv():
""" Method to retrieve a csv file """
yield pd.read_csv('/home/pybokeh/Dropbox/data_sets/sales_data.csv')
def add_column(df):
""" Method to add a new column to the dataframe """
print("Original dataframe:")
print(df.head())
df = df.assign(New_column=df['Sales'] + 200)
yield df
def print_dataframe(df):
""" Method to print the first 5 rows of the dataframe """
print("Modified dataframe:")
print(df.head())
# Define the order the methods will be executed
graph = bonobo.Graph(
get_csv,
add_column,
print_dataframe
)
if __name__ == '__main__':
""" Execute the methods in the specified order """
bonobo.run(graph)
| jupyter_notebooks/ETL/bonobo/Simple_Example.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: ScalaTion
// language: scala
// name: scalation
// ---
// # Deriving Multiple Linear Regression
// In this notebook, we demonstrate how to derive the least squares solution for multiple linear regression.
// ## Derivation
// Suppose you want to predict the vector $y$ using variables $x_1, x_2, \dots, x_n$ stored as the columns of the design matrix $X$. This suggests the following model $ y = X\beta + e $ where $\beta$ denotes a vector of coefficients and $e$ denotes a vector of error terms not explained by the rest of the model (i.e., the residuals). Our goal is to determine the $\beta$ values that minimize the sum of squared error. This approach is often called [*least squares*](https://en.wikipedia.org/wiki/Least_squares).
// * The error is simply $e = Xb$.
// * We want to find the $\beta$ that minimizes $e \cdot e = e'e = (y-X\beta)'(y-X\beta) = y'y - 2b'X'y + b'X'Xb$.
// * To mimimize, we take the derivative and set it equal to zero: $\frac{\partial(e \cdot e)}{\partial b} = -2X'y + 2X'Xb = 0$
// * Solving for $\beta$ gives $X'Xb=X'y$ and finally $b=(X'X)^{-1}X'y$.
//
// Therefore, when trying to predict $y$ by $\hat{y} = X\hat{\beta}$, our estimate for $\beta$ should be $\hat{\beta} = (X'X)^{-1}X'y$.
// ## Application
import scalation.linalgebra._
val x = new MatrixD((8, 2), 1, 1.1, 2, 2.2, 3, 3.3, 4, 4.4, 5, 5.5, 6, 6.5, 7, 7.5, 8, 8.8)
val actual = VectorD(2, 3)
val rng = scalation.random.Normal(0, 0.01) // random number generator
val noise = VectorD(for (i <- x.range1) yield rng.gen) // make some noise
val y = (x * actual) + noise // make noisy response vector
val b = (x.t * x).inverse * x.t * y
val e = y - x * b
val sse = e dot e
| notebooks/regression2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial of hdfs python library
# References:
# - [PyHive Documentation](https://github.com/dropbox/PyHive)
# - [Using Hive External Tables](https://docs.cloudera.com/HDPDocuments/HDP3/HDP-3.1.5/using-hiveql/content/hive_create_an_external_table.html)
# Install hdfs library from pyPi repo:
# !pip install --user pyhive
# References:
# - [PyHive Documentation](https://github.com/dropbox/PyHive)
# - [Using Hive External Tables](https://docs.cloudera.com/HDPDocuments/HDP3/HDP-3.1.5/using-hiveql/content/hive_create_an_external_table.html)
# !pip show parquet
# ## HDFS DATASET
# Subimos el dataset a hdfs
# +
from hdfs import InsecureClient
hdfs_client = InsecureClient('http://hive-hdfs:50070', user='hdfs')
if not(hdfs_client.status('/datasets/world_cities', strict=False)):
# Download a file or folder locally.
hdfs_client.makedirs('/datasets/world_cities')
hdfs_client.upload('/datasets/world_cities','world-cities_csv.csv', n_threads=5)
hdfs_client.list('/datasets/world_cities',status=True)
# -
from pyhive import hive
conn = hive.Connection(host="hive-server", port=10000, username="hive")
cursor = conn.cursor()
# ## Consultas SQL con Hive
#
# Directamente sobre los archivos csv almacenados en hdfs
# Crear base de datos de prueba en el metastore
# +
cursor.execute("CREATE DATABASE IF NOT EXISTS hive_tutorial")
cursor.execute("USE hive_tutorial")
conn.commit()
cursor.execute("SHOW DATABASES")
conn.commit()
print(cursor.fetchall())
# +
cursor.execute("DROP table world_cities")
query = """
CREATE EXTERNAL TABLE IF NOT EXISTS world_cities (
name STRING,
country STRING,
subcountry STRING,
geonameid INT)
COMMENT 'database with information about world cities'
ROW FORMAT DELIMITED
FIELDS TERMINATED BY ','
STORED AS TEXTFILE
location 'hdfs://hive-hdfs/datasets/world_cities/'
tblproperties ("skip.header.line.count"="1")
"""
cursor.execute(query)
conn.commit()
# -
cursor.execute("SHOW TABLES")
conn.commit()
print(cursor.fetchall())
cursor.execute("SELECT * FROM world_cities limit 100")
conn.commit()
size = 10
records = cursor.fetchmany(size)
print("Fetching Total ", size," rows")
print("Printing each row")
for row in records:
print(row)
cursor.close()
cursor.execute("SELECT name FROM world_cities")
conn.commit()
size = 10
records = cursor.fetchmany(size)
print("Fetching Total ", size," rows")
print("Printing each row")
for row in records:
print(row)
cursor.close()
| doc/notebooks/hive/hive-101.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py36
# language: python
# name: py36
# ---
# %load_ext autoreload
# +
#coding=utf-8
# %autoreload
import numpy as np
from tqdm import tqdm
import pickle
from gensim.models import Word2Vec
import node2vec_layout
import networkx as nx
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# -
adj_list = pickle.load(open('adj_list.pkl', 'rb'))
directed = True
G=nx.DiGraph()
list_nodes = []
node_index_in_G = {}
for v in adj_list:
G.add_node(v)
node_index_in_G[v] = len(node_index_in_G)
list_nodes.append(v)
for u in adj_list[v]:
G.add_edge(v, u, weight = 1)
poses = node2vec_layout.node2vec_layout(adj_list)
plt.figure(figsize = (7, 7))
nx.draw(G, poses, node_size=1, width=0.1, with_labels=False, edge_color='gray');
| fragments/test_node2vec_layout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Solving the SLIM MIP
# -
# This script shows how to solve the SLIM MIP using CPLEX. Here, we solve the MIP, run unit tests on the solution, and output the resulting scoring system model as well as some statistics about the MIP solver. Edit the variables in the following cell in order to pick a different instance.
# + deletable=true editable=true
data_name = 'breastcancer'
instance_name = data_name + '_max_5_features'
repository_dir = '/Users/berk/Desktop/Dropbox (MIT)/Research/SLIM/Toolboxes/miplib2017-slim'
data_file = repository_dir + '/models/data/' + data_name + '_processed.csv'
instance_file = repository_dir + '/instances/' + instance_name + '.mps'
instance_info_file = repository_dir + '/misc/' + instance_name + '.p'
# + [markdown] deletable=true editable=true
# The next cell loads packages and files needed to run the script. Use ``slim_mip.parameters`` to pass parameters for CPLEX.
# + deletable=true editable=true
import os
import sys
import numpy as np
import cplex as cpx
import pickle
import slim as slim
#load IP
slim_mip = cpx.Cplex(instance_file)
slim_info = pickle.load(open(instance_info_file))
data = slim.load_data_from_csv(data_file)
#set CPLEX IP parameters
slim_mip.parameters.timelimit.set(60)
slim_mip.parameters.randomseed.set(0)
slim_mip.parameters.output.clonelog.set(0)
slim_mip.parameters.threads.set(1)
slim_mip.parameters.parallel.set(1)
slim_mip.parameters.mip.tolerances.mipgap.set(np.finfo(np.float).eps)
slim_mip.parameters.mip.tolerances.absmipgap.set(np.finfo(np.float).eps)
slim_mip.parameters.mip.tolerances.integrality.set(np.finfo(np.float).eps)
# + [markdown] deletable=true editable=true
# We now solve the slim_mip, and use ``slim.check_ip_solution`` to make sure that the solution passes unit tests.
# + deletable=true editable=true
slim_IP.solve()
slim.check_slim_ip_solution(slim_mip, slim_info, data)
# -
Get statistics for the MIP as well as the scoring system
# + deletable=true editable=true
slim_results = slim.get_slim_summary(slim_IP, slim_info, data)
print(slim_results)
# -
| models/solve_slim_instance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparing color-color tracks of the stellar templates
#
# The goal of this notebook is to compare the stellar loci (in various color-color spaces) of the (theoretical) templates to the observed loci.
# +
import os
import numpy as np
import fitsio
import matplotlib.pyplot as plt
from speclite import filters
from astropy import constants
import astropy.units as u
from desisim.io import read_basis_templates
import seaborn as sns
# %pylab inline
# -
sns.set(style='white', font_scale=1.8, font='sans-serif', palette='Set2')
setcolors = sns.color_palette()
# ### Read a random sweep, select stars, and correct the observed fluxes for reddening
def read_and_dered():
bright, faint = 18, 19.5
sweepfile = 'sweep-240p000-250p005.fits'
print('Reading {}...'.format(sweepfile))
cat = fitsio.read(sweepfile, ext=1, upper=True)
these = np.where( (np.char.strip(cat['TYPE'].astype(str)) == 'PSF') *
(cat['DECAM_FLUX'][..., 2] > 1e9 * 10**(-0.4*faint)) *
(cat['DECAM_FLUX'][..., 2] < 1e9 * 10**(-0.4*bright))
)[0]
cat = cat[these]
print('...and selected {} stars with {} < r < {}.'.format(len(cat), bright, faint))
for prefix in ('DECAM', 'WISE'):
cat['{}_FLUX'.format(prefix)] = ( cat['{}_FLUX'.format(prefix)] /
cat['{}_MW_TRANSMISSION'.format(prefix)] )
cat['{}_FLUX_IVAR'.format(prefix)] = ( cat['{}_FLUX_IVAR'.format(prefix)] *
cat['{}_MW_TRANSMISSION'.format(prefix)]**2 )
return cat
cat = read_and_dered()
# ## Load the filter curves, the stellar templates, and get synthetic colors.
def obsflux2colors(cat):
"""Convert observed DECam/WISE fluxes to magnitudes and colors."""
cc = dict()
with warnings.catch_warnings(): # ignore missing fluxes (e.g., for QSOs)
warnings.simplefilter('ignore')
for ii, band in zip((1, 2, 4), ('g', 'r', 'z')):
cc[band] = 22.5 - 2.5 * np.log10(cat['DECAM_FLUX'][..., ii].data)
for ii, band in zip((0, 1), ('W1', 'W2')):
cc[band] = 22.5 - 2.5 * np.log10(cat['WISE_FLUX'][..., ii].data)
cc['gr'] = cc['g'] - cc['r']
cc['gz'] = cc['g'] - cc['z']
cc['rz'] = cc['r'] - cc['z']
cc['rW1'] = cc['r'] - cc['W1']
cc['zW1'] = cc['z'] - cc['W1']
cc['W1W2'] = cc['W1'] - cc['W2']
return cc
def synthflux2colors(synthflux):
"""Convert the synthesized DECam/WISE fluxes to colors."""
cc = dict(
r = 22.5 - 2.5 * np.log10(synthflux[1, :]),
gr = -2.5 * np.log10(synthflux[0, :] / synthflux[1, :]),
rz = -2.5 * np.log10(synthflux[1, :] / synthflux[2, :]),
gz = -2.5 * np.log10(synthflux[0, :] / synthflux[2, :]),
rW1 = -2.5 * np.log10(synthflux[1, :] / synthflux[3, :]),
zW1 = -2.5 * np.log10(synthflux[2, :] / synthflux[3, :]),
)
return cc
def star_synthflux():
"""Read the DESI stellar templates and synthesize photometry."""
flux, wave, meta = read_basis_templates(objtype='STAR')
nt = len(meta)
print('Read {} DESI templates.'.format(nt))
phot = filt.get_ab_maggies(flux, wave, mask_invalid=False)
synthflux = np.vstack( [phot[ff].data for ff in filts] )
return synthflux
def pickles_synthflux():
"""Read the Pickles+98 stellar templates and synthesize photometry."""
picklefile = os.path.join(os.getenv('CATALOGS_DIR'), '98pickles', '98pickles.fits')
data = fitsio.read(picklefile, ext=1)
print('Read {} Pickles templates.'.format(len(data)))
wave = data['WAVE'][0, :]
flux = data['FLUX']
padflux, padwave = filt.pad_spectrum(flux, wave, method='edge')
phot = filt.get_ab_maggies(padflux, padwave, mask_invalid=False)
synthflux = np.vstack( [phot[ff].data for ff in filts] )
return synthflux
filts = ('decam2014-g', 'decam2014-r', 'decam2014-z', 'wise2010-W1', 'wise2010-W2')
filt = filters.load_filters(*filts)
starcol = synthflux2colors(star_synthflux())
picklecol = synthflux2colors(pickles_synthflux())
obscol = obsflux2colors(cat)
# ## Generate color-color plots.
grrange = (-0.6, 2.2)
gzrange = (0.0, 4.0)
rzrange = (-0.6, 2.8)
zW1range = (-2.5, 0.0)
def grz(pngfile=None):
fig, ax = plt.subplots(figsize=(10, 6))
if False:
hb = ax.scatter(obscol['rz'], obscol['gr'], c=obscol['r'], s=1,
edgecolor='none')
else:
hb = ax.hexbin(obscol['rz'], obscol['gr'], mincnt=5,
bins='log', gridsize=150)
ax.scatter(picklecol['rz'], picklecol['gr'], marker='s',
s=40, linewidth=1, alpha=0.5, label='Pickles+98', c='r')
ax.scatter(starcol['rz'], starcol['gr'], marker='o',
s=10, linewidth=1, alpha=0.8, label='STAR Templates', c='b')
ax.set_xlabel('r - z')
ax.set_ylabel('g - r')
ax.set_xlim(rzrange)
ax.set_ylim(grrange)
lgnd = ax.legend(loc='upper left', frameon=False, fontsize=18)
lgnd.legendHandles[0]._sizes = [100]
lgnd.legendHandles[1]._sizes = [100]
cb = fig.colorbar(hb, ax=ax)
cb.set_label(r'log$_{10}$ (Number of 18<r<19.5 Stars per Bin)')
if pngfile:
fig.savefig(pngfile)
def gzW1(pngfile=None):
fig, ax = plt.subplots(figsize=(10, 6))
hb = ax.hexbin(obscol['zW1'], obscol['gz'], mincnt=10,
bins='log', gridsize=150)
ax.scatter(starcol['zW1'], starcol['gz'], marker='o',
s=10, alpha=0.5, label='STAR Templates', c='b')
ax.set_xlabel('z - W1')
ax.set_ylabel('g - z')
ax.set_ylim(gzrange)
ax.set_xlim(zW1range)
lgnd = ax.legend(loc='upper left', frameon=False, fontsize=18)
lgnd.legendHandles[0]._sizes = [100]
cb = fig.colorbar(hb, ax=ax)
cb.set_label(r'log$_{10}$ (Number of 18<r<19.5 Stars per Bin)')
if pngfile:
fig.savefig(pngfile)
gzW1()
grz()
| doc/nb/color-color-templates-star.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linked List Reference Implementations
# ### Operations:
#
# **insert_front(data)**
# * Insert data to the front of list
# * List may be empty
#
#
# **append(data)**
# * Append data to the end of list
# * List may be empty
# * precondition: the data to be appended must be valid
#
#
# **find(data)**
# * Returns True if data exists in the list
# * List may be empty
# * precondition: the value to be searched must be valid
#
#
# **delete(data)**
# * Delete a node with node.data == data in the list
# * List may be empty
# * precondition: the value to be deleted must exists in the list
#
#
# **print_list()**
# * Print all the elements in the list
# * List may be empty
#
#
# **get_length()**
# * Returns the length of the list
#
#
# **insert_after(val, data)**:
# * Insert val after data
# * precondition: List must be not empty, the ***data*** must exists in the list
#
#
# **insert_before(val, data)**:
# * Insert val before data
# * precondition: List must be not empty, the ***data** must exists in the list
# # 1. Singly Linked List
class Node(object):
def __init__(self, data, next=None):
self.data = data
self.next = next
def __str__(self):
return str(self.data)
class SinglyLinkedList(object):
def __init__(self, head=None):
self.head = head
self.tail = head
if self.head:
self.length = 1
else:
self.length = 0
def insert_front(self, data):
if self.length == 0:
self.head = Node(data)
self.tail = self.head
else:
temp = Node(data, self.head)
self.head = temp
self.length += 1
def append(self, data):
if self.length == 0:
self.head = Node(data)
self.tail = self.head
else:
temp = Node(data)
self.tail.next = temp
self.tail = temp
self.length += 1
def find(self, data):
if self.length == 0:
return False
if self.length == 1 and self.head.data != data:
return False
p = self.head
while p is not None:
if p.data == data:
return True
p = p.next
return False
def delete(self, data):
if self.head.data == data:
self.length -= 1
self.head = self.head.next
if self.head is None:
self.tail = None
elif self.length > 0:
p = self.head
pprev = self.head
while p is not None:
if p.data == data:
pprev.next = p.next
self.length -= 1
if p.next is None:
self.tail = pprev
return
pprev = p
p = p.next
def print_list(self):
p = self.head
while p is not None:
print(p.data, " ")
p = p.next
def get_length(self):
return self.length
def insert_after(self, val, data):
p = self.head
while p is not None:
if p.data == data:
self.length += 1
temp = Node(val, p.next)
p.next = temp
if p == self.tail:
self.tail = temp
return
p = p.next
def insert_before(self, val, data):
if self.head.data == data:
temp = Node(val, self.head)
self.head = temp
self.length += 1
else:
p = self.head
pprev = None
while p is not None:
if p.data == data:
temp = Node(val, p)
pprev.next = temp
self.length += 1
return
pprev = p
p = p.next
a = Node(12)
b = Node(10)
c = Node(5)
l = SinglyLinkedList(a)
print("Length: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
l.delete(12)
print("Length after delete 12: ", l.get_length())
l.append(50)
l.append(100)
print("Length after append 50 and 100: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
print("Does 100 appear in the list? ", l.find(100))
print("Does 12 appear in the list? ", l.find(12))
l.delete(100)
print("Length after delete 100: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
print("Lets print the list:")
l.print_list()
l.insert_after(150, 50)
l.insert_before(10, 50)
print("The list after inserting 150 after 50 and 10 before 50:")
l.print_list()
print("Length: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
# # 2. Doubly Linked List
class Node2(object):
def __init__(self, data, previous=None, next=None):
self.data = data
self.previous = previous
self.next = next
def __str__(self):
return str(self.data)
class DoublyLinkedList(object):
def __init__(self, head=None):
self.head = head
self.tail = self.head
if self.head:
self.length = 1
else:
self.length = 0
def insert_front(self, data):
if self.length == 0:
self.head = Node2(data)
self.tail = self.head
else:
temp = Node2(data, next=self.head)
self.head.previous = temp
self.head = temp
self.length += 1
def append(self, data):
if self.length == 0:
self.head = Node2(data)
self.tail = self.head
else:
temp = Node2(data, previous=self.tail)
self.tail.next = temp
self.tail = temp
self.length += 1
def find(self, data):
if self.length == 0:
return False
if self.length == 1 and self.head.data != data:
return False
p = self.head
while p is not None:
if p.data == data:
return True
p = p.next
return False
def delete(self, data):
if self.head.data == data:
self.length -= 1
self.head = self.head.next
if self.head is None:
self.tail = None
elif self.length > 0:
p = self.head
pprev = self.head
while p is not None:
if p.data == data:
pprev.next = p.next
self.length -= 1
if p.next is None:
self.tail = pprev
else:
p.next.previous = pprev
return
pprev = p
p = p.next
def print_list(self):
p = self.head
while p is not None:
print(p.data, " ")
p = p.next
def get_length(self):
return self.length
def insert_after(self, val, data):
p = self.head
while p is not None:
if p.data == data:
self.length += 1
temp = Node2(val, p, p.next)
if p.next is None:
self.tail = temp
else:
p.next.previous = temp
p.next = temp
return
p = p.next
def insert_before(self, val, data):
if self.head.data == data:
temp = Node2(val, next=self.head)
self.head.previous = temp
self.head = temp
self.length += 1
else:
p = self.head
pprev = None
while p is not None:
if p.data == data:
temp = Node2(val, pprev, p)
p.previous = temp
pprev.next = temp
self.length += 1
return
pprev = p
p = p.next
l = DoublyLinkedList(Node2(12,None,None))
print("Length: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
l.delete(12)
print("Length after delete 12: ", l.get_length())
l.append(50)
l.append(100)
print("Length after append 50 and 100: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
print("Does 100 appear in the list? ", l.find(100))
print("Does 12 appear in the list? ", l.find(12))
l.delete(100)
print("Length after delete 100: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
print("Lets print the list:")
l.print_list()
l.insert_after(150, 50)
l.insert_before(10, 50)
print("The list after inserting 150 after 50 and 10 before 50:")
l.print_list()
print("Length: ", l.get_length())
print("Head Now: {}, tail now: {}".format(l.head.data, l.tail.data))
print("Tail.previous: {} Tail.previous.previous: {}".format(l.tail.previous, l.tail.previous.previous))
| DataStructures/Linked List.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from y0.dsl import One, P, A, B, C, D, Q, R, S, T, W, X, Y, Z, Sum, Variable, Product
One()
P(A)
P(A @ W)
P(A @ ~W)
P(A @ ~W @ X)
P(A, B)
P(A | B)
One() / P(A | B)
(One() / P(A | B)) * P(A)
One() * P(A | B)
P(A | B) * P(B)
Sum[B](P(A | B))
Sum(P(A | B) * Sum(P(C | D), [R]), [S, T])
Sum[W](P((Y @ ~Z @ W) & X) * P(D) * P(Z @ D) * P(W @ ~X))
(P(A) / P(B)) / (P(C) / P(D))
Q[A](B)
P(Variable(v) for v in ['A', 'B', 'C'])
Product([P(A), P(B), P(C)])
Sum(P(Variable(v) for v in ['A','B','C']), [Variable(v) for v in ['A', 'B']])
| notebooks/DSL Demo.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] colab_type="text" id="JndnmDMp66FL"
# #### Copyright 2017 Google LLC.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="hMqWDc_m6rUC"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="zbIgBK-oXHO7"
# # Validation
# + [markdown] colab_type="text" id="WNX0VyBpHpCX"
# **Learning Objectives:**
# * Use multiple features, instead of a single feature, to further improve the effectiveness of a model
# * Debug issues in model input data
# * Use a test data set to check if a model is overfitting the validation data
# + [markdown] colab_type="text" id="za0m1T8CHpCY"
# As in the prior exercises, we're working with the [California housing data set](https://developers.google.com/machine-learning/crash-course/california-housing-data-description), to try and predict `median_house_value` at the city block level from 1990 census data.
# + [markdown] colab_type="text" id="r2zgMfWDWF12"
# ## Setup
# + [markdown] colab_type="text" id="8jErhkLzWI1B"
# First off, let's load up and prepare our data. This time, we're going to work with multiple features, so we'll modularize the logic for preprocessing the features a bit:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="PwS5Bhm6HpCZ"
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_train.csv", sep=",")
# california_housing_dataframe = california_housing_dataframe.reindex(
# np.random.permutation(california_housing_dataframe.index))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="J2ZyTzX0HpCc"
def preprocess_features(california_housing_dataframe):
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
["latitude",
"longitude",
"housing_median_age",
"total_rooms",
"total_bedrooms",
"population",
"households",
"median_income"]]
processed_features = selected_features.copy()
# Create a synthetic feature.
processed_features["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] /
california_housing_dataframe["population"])
return processed_features
def preprocess_targets(california_housing_dataframe):
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = pd.DataFrame()
# Scale the target to be in units of thousands of dollars.
output_targets["median_house_value"] = (
california_housing_dataframe["median_house_value"] / 1000.0)
return output_targets
# + [markdown] colab_type="text" id="sZSIaDiaHpCf"
# For the **training set**, we'll choose the first 12000 examples, out of the total of 17000.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="P9wejvw7HpCf"
training_examples = preprocess_features(california_housing_dataframe.head(12000))
training_examples.describe()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="JlkgPR-SHpCh"
training_targets = preprocess_targets(california_housing_dataframe.head(12000))
training_targets.describe()
# + [markdown] colab_type="text" id="5l1aA2xOHpCj"
# For the **validation set**, we'll choose the last 5000 examples, out of the total of 17000.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="fLYXLWAiHpCk"
validation_examples = preprocess_features(california_housing_dataframe.tail(5000))
validation_examples.describe()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="oVPcIT3BHpCm"
validation_targets = preprocess_targets(california_housing_dataframe.tail(5000))
validation_targets.describe()
# + [markdown] colab_type="text" id="z3TZV1pgfZ1n"
# ## Task 1: Examine the Data
# Okay, let's look at the data above. We have `9` input features that we can use.
#
# Take a quick skim over the table of values. Everything look okay? See how many issues you can spot. Don't worry if you don't have a background in statistics; common sense will get you far.
#
# After you've had a chance to look over the data yourself, check the solution for some additional thoughts on how to verify data.
# + [markdown] colab_type="text" id="4Xp9NhOCYSuz"
# ### Solution
#
# Click below for the solution.
# + [markdown] colab_type="text" id="gqeRmK57YWpy"
# Let's check our data against some baseline expectations:
#
# * For some values, like `median_house_value`, we can check to see if these values fall within reasonable ranges (keeping in mind this was 1990 data — not today!).
#
# * For other values, like `latitude` and `longitude`, we can do a quick check to see if these line up with expected values from a quick Google search.
#
# If you look closely, you may see some oddities:
#
# * `median_income` is on a scale from about 3 to 15. It's not at all clear what this scale refers to—looks like maybe some log scale? It's not documented anywhere; all we can assume is that higher values correspond to higher income.
#
# * The maximum `median_house_value` is 500,001. This looks like an artificial cap of some kind.
#
# * Our `rooms_per_person` feature is generally on a sane scale, with a 75th percentile value of about 2. But there are some very large values, like 18 or 55, which may show some amount of corruption in the data.
#
# We'll use these features as given for now. But hopefully these kinds of examples can help to build a little intuition about how to check data that comes to you from an unknown source.
# + [markdown] colab_type="text" id="fXliy7FYZZRm"
# ## Task 2: Plot Latitude/Longitude vs. Median House Value
# + [markdown] colab_type="text" id="aJIWKBdfsDjg"
# Let's take a close look at two features in particular: **`latitude`** and **`longitude`**. These are geographical coordinates of the city block in question.
#
# This might make a nice visualization — let's plot `latitude` and `longitude`, and use color to show the `median_house_value`.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "test": {"output": "ignore", "timeout": 600}} colab_type="code" id="5_LD23bJ06TW"
plt.figure(figsize=(13, 8))
ax = plt.subplot(1, 2, 1)
ax.set_title("Validation Data")
ax.set_autoscaley_on(False)
ax.set_ylim([32, 43])
ax.set_autoscalex_on(False)
ax.set_xlim([-126, -112])
plt.scatter(validation_examples["longitude"],
validation_examples["latitude"],
cmap="coolwarm",
c=validation_targets["median_house_value"] / validation_targets["median_house_value"].max())
ax = plt.subplot(1,2,2)
ax.set_title("Training Data")
ax.set_autoscaley_on(False)
ax.set_ylim([32, 43])
ax.set_autoscalex_on(False)
ax.set_xlim([-126, -112])
plt.scatter(training_examples["longitude"],
training_examples["latitude"],
cmap="coolwarm",
c=training_targets["median_house_value"] / training_targets["median_house_value"].max())
_ = plt.plot()
# + [markdown] colab_type="text" id="32_DbjnfXJlC"
# Wait a second...this should have given us a nice map of the state of California, with red showing up in expensive areas like the San Francisco and Los Angeles.
#
# The training set sort of does, compared to a [real map](https://www.google.com/maps/place/California/@37.1870174,-123.7642688,6z/data=!3m1!4b1!4m2!3m1!1s0x808fb9fe5f285e3d:0x8b5109a227086f55), but the validation set clearly doesn't.
#
# **Go back up and look at the data from Task 1 again.**
#
# Do you see any other differences in the distributions of features or targets between the training and validation data?
# + [markdown] colab_type="text" id="pECTKgw5ZvFK"
# ### Solution
#
# Click below for the solution.
# + [markdown] colab_type="text" id="49NC4_KIZxk_"
# Looking at the tables of summary stats above, it's easy to wonder how anyone would do a useful data check. What's the right 75<sup>th</sup> percentile value for total_rooms per city block?
#
# The key thing to notice is that for any given feature or column, the distribution of values between the train and validation splits should be roughly equal.
#
# The fact that this is not the case is a real worry, and shows that we likely have a fault in the way that our train and validation split was created.
# + [markdown] colab_type="text" id="025Ky0Dq9ig0"
# ## Task 3: Return to the Data Importing and Pre-Processing Code, and See if You Spot Any Bugs
# If you do, go ahead and fix the bug. Don't spend more than a minute or two looking. If you can't find the bug, check the solution.
# + [markdown] colab_type="text" id="JFsd2eWHAMdy"
# When you've found and fixed the issue, re-run `latitude` / `longitude` plotting cell above and confirm that our sanity checks look better.
#
# By the way, there's an important lesson here.
#
# **Debugging in ML is often *data debugging* rather than code debugging.**
#
# If the data is wrong, even the most advanced ML code can't save things.
# + [markdown] colab_type="text" id="dER2_43pWj1T"
# ### Solution
#
# Click below for the solution.
# + [markdown] colab_type="text" id="BnEVbYJvW2wu"
# Take a look at how the data is randomized when it's read in.
#
# If we don't randomize the data properly before creating training and validation splits, then we may be in trouble if the data is given to us in some sorted order, which appears to be the case here.
# + [markdown] colab_type="text" id="xCdqLpQyAos2"
# ## Task 4: Train and Evaluate a Model
#
# **Spend 5 minutes or so trying different hyperparameter settings. Try to get the best validation performance you can.**
#
# Next, we'll train a linear regressor using all the features in the data set, and see how well we do.
#
# Let's define the same input function we've used previously for loading the data into a TensorFlow model.
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="rzcIPGxxgG0t"
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model of one feature.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified
if shuffle:
ds = ds.shuffle(10000)
# Return the next batch of data
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# + [markdown] colab_type="text" id="CvrKoBmNgRCO"
# Because we're now working with multiple input features, let's modularize our code for configuring feature columns into a separate function. (For now, this code is fairly simple, as all our features are numeric, but we'll build on this code as we use other types of features in future exercises.)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="wEW5_XYtgZ-H"
def construct_feature_columns(input_features):
"""Construct the TensorFlow Feature Columns.
Args:
input_features: The names of the numerical input features to use.
Returns:
A set of feature columns
"""
return set([tf.feature_column.numeric_column(my_feature)
for my_feature in input_features])
# + [markdown] colab_type="text" id="D0o2wnnzf8BD"
# Next, go ahead and complete the `train_model()` code below to set up the input functions and calculate predictions.
#
# **NOTE:** It's okay to reference the code from the previous exercises, but make sure to call `predict()` on the appropriate data sets.
#
# Compare the losses on training data and validation data. With a single raw feature, our best root mean squared error (RMSE) was of about 180.
#
# See how much better you can do now that we can use multiple features.
#
# Check the data using some of the methods we've looked at before. These might include:
#
# * Comparing distributions of predictions and actual target values
#
# * Creating a scatter plot of predictions vs. target values
#
# * Creating two scatter plots of validation data using `latitude` and `longitude`:
# * One plot mapping color to actual target `median_house_value`
# * A second plot mapping color to predicted `median_house_value` for side-by-side comparison.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "test": {"output": "ignore", "timeout": 600}} colab_type="code" id="UXt0_4ZTEf4V"
def train_model(
learning_rate,
steps,
batch_size,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a linear regression model of one feature.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `LinearRegressor` object trained on the training data.
"""
periods = 10
steps_per_period = steps / periods
# Create a linear regressor object.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
linear_regressor = tf.estimator.LinearRegressor(
feature_columns=construct_feature_columns(training_examples),
optimizer=my_optimizer
)
# 1. Create input functions.
training_input_fn = # YOUR CODE HERE
predict_training_input_fn = # YOUR CODE HERE
predict_validation_input_fn = # YOUR CODE HERE
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print "Training model..."
print "RMSE (on training data):"
training_rmse = []
validation_rmse = []
for period in range (0, periods):
# Train the model, starting from the prior state.
linear_regressor.train(
input_fn=training_input_fn,
steps=steps_per_period,
)
# 2. Take a break and compute predictions.
training_predictions = # YOUR CODE HERE
validation_predictions = # YOUR CODE HERE
# Compute training and validation loss.
training_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(training_predictions, training_targets))
validation_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(validation_predictions, validation_targets))
# Occasionally print the current loss.
print " period %02d : %0.2f" % (period, training_root_mean_squared_error)
# Add the loss metrics from this period to our list.
training_rmse.append(training_root_mean_squared_error)
validation_rmse.append(validation_root_mean_squared_error)
print "Model training finished."
# Output a graph of loss metrics over periods.
plt.ylabel("RMSE")
plt.xlabel("Periods")
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(training_rmse, label="training")
plt.plot(validation_rmse, label="validation")
plt.legend()
return linear_regressor
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="zFFRmvUGh8wd"
linear_regressor = train_model(
# TWEAK THESE VALUES TO SEE HOW MUCH YOU CAN IMPROVE THE RMSE
learning_rate=0.00001,
steps=100,
batch_size=1,
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
# + [markdown] colab_type="text" id="I-La4N9ObC1x"
# ### Solution
#
# Click below for a solution.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Xyz6n1YHbGef"
def train_model(
learning_rate,
steps,
batch_size,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a linear regression model of one feature.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `LinearRegressor` object trained on the training data.
"""
periods = 10
steps_per_period = steps / periods
# Create a linear regressor object.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
linear_regressor = tf.estimator.LinearRegressor(
feature_columns=construct_feature_columns(training_examples),
optimizer=my_optimizer
)
# Create input functions.
training_input_fn = lambda: my_input_fn(
training_examples,
training_targets["median_house_value"],
batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(
training_examples,
training_targets["median_house_value"],
num_epochs=1,
shuffle=False)
predict_validation_input_fn = lambda: my_input_fn(
validation_examples, validation_targets["median_house_value"],
num_epochs=1,
shuffle=False)
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print "Training model..."
print "RMSE (on training data):"
training_rmse = []
validation_rmse = []
for period in range (0, periods):
# Train the model, starting from the prior state.
linear_regressor.train(
input_fn=training_input_fn,
steps=steps_per_period,
)
# Take a break and compute predictions.
training_predictions = linear_regressor.predict(input_fn=predict_training_input_fn)
training_predictions = np.array([item['predictions'][0] for item in training_predictions])
validation_predictions = linear_regressor.predict(input_fn=predict_validation_input_fn)
validation_predictions = np.array([item['predictions'][0] for item in validation_predictions])
# Compute training and validation loss.
training_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(training_predictions, training_targets))
validation_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(validation_predictions, validation_targets))
# Occasionally print the current loss.
print " period %02d : %0.2f" % (period, training_root_mean_squared_error)
# Add the loss metrics from this period to our list.
training_rmse.append(training_root_mean_squared_error)
validation_rmse.append(validation_root_mean_squared_error)
print "Model training finished."
# Output a graph of loss metrics over periods.
plt.ylabel("RMSE")
plt.xlabel("Periods")
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(training_rmse, label="training")
plt.plot(validation_rmse, label="validation")
plt.legend()
return linear_regressor
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="i1imhjFzbWwt"
linear_regressor = train_model(
learning_rate=0.00003,
steps=500,
batch_size=5,
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
# + [markdown] colab_type="text" id="65sin-E5NmHN"
# ## Task 5: Evaluate on Test Data
#
# **In the cell below, load in the test data set and evaluate your model on it.**
#
# We've done a lot of iteration on our validation data. Let's make sure we haven't overfit to the pecularities of that particular sample.
#
# Test data set is located [here](https://storage.googleapis.com/mledu-datasets/california_housing_test.csv).
#
# How does your test performance compare to the validation performance? What does this say about the generalization performance of your model?
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "test": {"output": "ignore", "timeout": 600}} colab_type="code" id="icEJIl5Vp51r"
california_housing_test_data = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_test.csv", sep=",")
#
# YOUR CODE HERE
#
# + [markdown] colab_type="text" id="yTghc_5HkJDW"
# ### Solution
#
# Click below for the solution.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="_xSYTarykO8U"
california_housing_test_data = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_test.csv", sep=",")
test_examples = preprocess_features(california_housing_test_data)
test_targets = preprocess_targets(california_housing_test_data)
predict_test_input_fn = lambda: my_input_fn(
test_examples,
test_targets["median_house_value"],
num_epochs=1,
shuffle=False)
test_predictions = linear_regressor.predict(input_fn=predict_test_input_fn)
test_predictions = np.array([item['predictions'][0] for item in test_predictions])
root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(test_predictions, test_targets))
print "Final RMSE (on test data): %0.2f" % root_mean_squared_error
| ml/cc/exercises/validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# # Amazon SageMaker Model Monitor
# This notebook shows how to:
# * Host a machine learning model in Amazon SageMaker and capture inference requests, results, and metadata
# * Analyze a training dataset to generate baseline constraints
# * Monitor a live endpoint for violations against constraints
#
# ---
# ## Background
#
# Amazon SageMaker provides every developer and data scientist with the ability to build, train, and deploy machine learning models quickly. Amazon SageMaker is a fully-managed service that encompasses the entire machine learning workflow. You can label and prepare your data, choose an algorithm, train a model, and then tune and optimize it for deployment. You can deploy your models to production with Amazon SageMaker to make predictions and lower costs than was previously possible.
#
# In addition, Amazon SageMaker enables you to capture the input, output and metadata for invocations of the models that you deploy. It also enables you to analyze the data and monitor its quality. In this notebook, you learn how Amazon SageMaker enables these capabilities.
#
# ---
# ## Setup
#
# To get started, make sure you have these prerequisites completed.
#
# * Specify an AWS Region to host your model.
# * An IAM role ARN exists that is used to give Amazon SageMaker access to your data in Amazon Simple Storage Service (Amazon S3). See the documentation for how to fine tune the permissions needed.
# * Create an S3 bucket used to store the data used to train your model, any additional model data, and the data captured from model invocations. For demonstration purposes, you are using the same bucket for these. In reality, you might want to separate them with different security policies.
# + isConfigCell=true
# cell 01
# %%time
# Handful of configuration
import os
import boto3
import re
import json
from sagemaker import get_execution_role, session
region= boto3.Session().region_name
role = get_execution_role()
print("RoleArn: {}".format(role))
# You can use a different bucket, but make sure the role you chose for this notebook
# has the s3:PutObject permissions. This is the bucket into which the data is captured
bucket = session.Session(boto3.Session()).default_bucket()
print("Demo Bucket: {}".format(bucket))
prefix = 'sagemaker/DEMO-ModelMonitor'
data_capture_prefix = '{}/datacapture'.format(prefix)
s3_capture_upload_path = 's3://{}/{}'.format(bucket, data_capture_prefix)
reports_prefix = '{}/reports'.format(prefix)
s3_report_path = 's3://{}/{}'.format(bucket,reports_prefix)
code_prefix = '{}/code'.format(prefix)
s3_code_preprocessor_uri = 's3://{}/{}/{}'.format(bucket,code_prefix, 'preprocessor.py')
s3_code_postprocessor_uri = 's3://{}/{}/{}'.format(bucket,code_prefix, 'postprocessor.py')
print("Capture path: {}".format(s3_capture_upload_path))
print("Report path: {}".format(s3_report_path))
print("Preproc Code path: {}".format(s3_code_preprocessor_uri))
print("Postproc Code path: {}".format(s3_code_postprocessor_uri))
# -
# You can quickly verify that the execution role for this notebook has the necessary permissions to proceed. Put a simple test object into the S3 bucket you specified above. If this command fails, update the role to have `s3:PutObject` permission on the bucket and try again.
# cell 02
# Upload some test files
boto3.Session().resource('s3').Bucket(bucket).Object("test_upload/test.txt").upload_file('test_data/upload-test-file.txt')
print("Success! You are all set to proceed.")
# # PART A: Capturing real-time inference data from Amazon SageMaker endpoints
# Create an endpoint to showcase the data capture capability in action.
#
# ### Upload the pre-trained model to Amazon S3
# This code uploads a pre-trained XGBoost model that is ready for you to deploy. This model was trained using the XGB Churn Prediction Notebook in SageMaker. You can also use your own pre-trained model in this step. If you already have a pretrained model in Amazon S3, you can add it instead by specifying the s3_key.
# cell 03
model_file = open("model/xgb-churn-prediction-model.tar.gz", 'rb')
s3_key = os.path.join(prefix, 'xgb-churn-prediction-model.tar.gz')
boto3.Session().resource('s3').Bucket(bucket).Object(s3_key).upload_fileobj(model_file)
# ### Deploy the model to Amazon SageMaker
# Start with deploying a pre-trained churn prediction model. Here, you create the model object with the image and model data.
# +
# cell 04
from time import gmtime, strftime
from sagemaker.model import Model
from sagemaker.image_uris import retrieve
model_name = "DEMO-xgb-churn-pred-model-monitor-" + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
model_url = 'https://{}.s3-{}.amazonaws.com/{}/xgb-churn-prediction-model.tar.gz'.format(bucket, region, prefix)
image_uri = retrieve(region=boto3.Session().region_name, framework='xgboost', version='0.90-2')
model = Model(image_uri=image_uri, model_data=model_url, role=role)
# -
# To enable data capture for monitoring the model data quality, you specify the new capture option called `DataCaptureConfig`. You can capture the request payload, the response payload or both with this configuration. The capture config applies to all variants. Go ahead with the deployment.
# +
# cell 05
from sagemaker.model_monitor import DataCaptureConfig
endpoint_name = 'DEMO-xgb-churn-pred-model-monitor-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("EndpointName={}".format(endpoint_name))
data_capture_config = DataCaptureConfig(
enable_capture=True,
sampling_percentage=100,
destination_s3_uri=s3_capture_upload_path)
predictor = model.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge',
endpoint_name=endpoint_name,
data_capture_config=data_capture_config)
# -
# ## Invoke the deployed model
#
# You can now send data to this endpoint to get inferences in real time. Because you enabled the data capture in the previous steps, the request and response payload, along with some additional metadata, is saved in the Amazon Simple Storage Service (Amazon S3) location you have specified in the DataCaptureConfig.
# This step invokes the endpoint with included sample data for about 2 minutes. Data is captured based on the sampling percentage specified and the capture continues until the data capture option is turned off.
# +
# cell 06
from sagemaker.predictor import Predictor
import sagemaker
import time
predictor = Predictor(endpoint_name=endpoint_name, serializer=sagemaker.serializers.CSVSerializer())
# get a subset of test data for a quick test
# !head -120 test_data/test-dataset-input-cols.csv > test_data/test_sample.csv
print("Sending test traffic to the endpoint {}. \nPlease wait...".format(endpoint_name))
with open('test_data/test_sample.csv', 'r') as f:
for row in f:
payload = row.rstrip('\n')
response = predictor.predict(data=payload)
time.sleep(0.5)
print("Done!")
# -
# ## View captured data
#
# Now list the data capture files stored in Amazon S3. You should expect to see different files from different time periods organized based on the hour in which the invocation occurred. The format of the Amazon S3 path is:
#
# `s3://{destination-bucket-prefix}/{endpoint-name}/{variant-name}/yyyy/mm/dd/hh/filename.jsonl`
# cell 07
s3_client = boto3.Session().client('s3')
current_endpoint_capture_prefix = '{}/{}'.format(data_capture_prefix, endpoint_name)
result = s3_client.list_objects(Bucket=bucket, Prefix=current_endpoint_capture_prefix)
capture_files = [capture_file.get("Key") for capture_file in result.get('Contents')]
print("Found Capture Files:")
print("\n ".join(capture_files))
# Next, view the contents of a single capture file. Here you should see all the data captured in an Amazon SageMaker specific JSON-line formatted file. Take a quick peek at the first few lines in the captured file.
# +
# cell 08
def get_obj_body(obj_key):
return s3_client.get_object(Bucket=bucket, Key=obj_key).get('Body').read().decode("utf-8")
capture_file = get_obj_body(capture_files[-1])
print(capture_file[:2000])
# -
# Finally, the contents of a single line is present below in a formatted JSON file so that you can observe a little better.
# cell 09
import json
print(json.dumps(json.loads(capture_file.split('\n')[0]), indent=2))
# As you can see, each inference request is captured in one line in the jsonl file. The line contains both the input and output merged together. In the example, you provided the ContentType as `text/csv` which is reflected in the `observedContentType` value. Also, you expose the encoding that you used to encode the input and output payloads in the capture format with the `encoding` value.
#
# To recap, you observed how you can enable capturing the input or output payloads to an endpoint with a new parameter. You have also observed what the captured format looks like in Amazon S3. Next, continue to explore how Amazon SageMaker helps with monitoring the data collected in Amazon S3.
# # PART B: Model Monitor - Baseling and continuous monitoring
# In addition to collecting the data, Amazon SageMaker provides the capability for you to monitor and evaluate the data observed by the endpoints. For this:
# 1. Create a baseline with which you compare the realtime traffic.
# 1. Once a baseline is ready, setup a schedule to continously evaluate and compare against the baseline.
# ## 1. Constraint suggestion with baseline/training dataset
# The training dataset with which you trained the model is usually a good baseline dataset. Note that the training dataset data schema and the inference dataset schema should exactly match (i.e. the number and order of the features).
#
# From the training dataset you can ask Amazon SageMaker to suggest a set of baseline `constraints` and generate descriptive `statistics` to explore the data. For this example, upload the training dataset that was used to train the pre-trained model included in this example. If you already have it in Amazon S3, you can directly point to it.
# +
# cell 10
# # copy over the training dataset to Amazon S3 (if you already have it in Amazon S3, you could reuse it)
baseline_prefix = prefix + '/baselining'
baseline_data_prefix = baseline_prefix + '/data'
baseline_results_prefix = baseline_prefix + '/results'
baseline_data_uri = 's3://{}/{}'.format(bucket,baseline_data_prefix)
baseline_results_uri = 's3://{}/{}'.format(bucket, baseline_results_prefix)
print('Baseline data uri: {}'.format(baseline_data_uri))
print('Baseline results uri: {}'.format(baseline_results_uri))
# -
# cell 11
training_data_file = open("test_data/training-dataset-with-header.csv", 'rb')
s3_key = os.path.join(baseline_prefix, 'data', 'training-dataset-with-header.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(s3_key).upload_fileobj(training_data_file)
# ### Create a baselining job with training dataset
# Now that you have the training data ready in Amazon S3, start a job to `suggest` constraints. `DefaultModelMonitor.suggest_baseline(..)` starts a `ProcessingJob` using an Amazon SageMaker provided Model Monitor container to generate the constraints.
# +
# cell 12
from sagemaker.model_monitor import DefaultModelMonitor
from sagemaker.model_monitor.dataset_format import DatasetFormat
my_default_monitor = DefaultModelMonitor(
role=role,
instance_count=1,
instance_type='ml.m5.xlarge',
volume_size_in_gb=20,
max_runtime_in_seconds=3600,
)
my_default_monitor_baseline = my_default_monitor.suggest_baseline(
baseline_dataset=baseline_data_uri+'/training-dataset-with-header.csv',
dataset_format=DatasetFormat.csv(header=True),
output_s3_uri=baseline_results_uri,
wait=True
)
# -
# ### Explore the generated constraints and statistics
# cell 13
s3_client = boto3.Session().client('s3')
result = s3_client.list_objects(Bucket=bucket, Prefix=baseline_results_prefix)
report_files = [report_file.get("Key") for report_file in result.get('Contents')]
print("Found Files:")
print("\n ".join(report_files))
# +
# cell 14
import pandas as pd
baseline_job = my_default_monitor.latest_baselining_job
schema_df = pd.json_normalize(baseline_job.baseline_statistics().body_dict["features"])
schema_df.head(10)
# -
# cell 15
constraints_df = pd.json_normalize(baseline_job.suggested_constraints().body_dict["features"])
constraints_df.head(10)
# ## 2. Analyzing collected data for data quality issues
#
# When you have collected the data above, analyze and monitor the data with Monitoring Schedules
# ### Create a schedule
# cell 16
# First, copy over some test scripts to the S3 bucket so that they can be used for pre and post processing
boto3.Session().resource('s3').Bucket(bucket).Object(code_prefix+"/preprocessor.py").upload_file('preprocessor.py')
boto3.Session().resource('s3').Bucket(bucket).Object(code_prefix+"/postprocessor.py").upload_file('postprocessor.py')
# You can create a model monitoring schedule for the endpoint created earlier. Use the baseline resources (constraints and statistics) to compare against the realtime traffic.
# +
# cell 17
from sagemaker.model_monitor import CronExpressionGenerator
from time import gmtime, strftime
mon_schedule_name = 'DEMO-xgb-churn-pred-model-monitor-schedule-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
my_default_monitor.create_monitoring_schedule(
monitor_schedule_name=mon_schedule_name,
endpoint_input=predictor.endpoint_name,
#record_preprocessor_script=pre_processor_script,
post_analytics_processor_script=s3_code_postprocessor_uri,
output_s3_uri=s3_report_path,
statistics=my_default_monitor.baseline_statistics(),
constraints=my_default_monitor.suggested_constraints(),
schedule_cron_expression=CronExpressionGenerator.hourly(),
enable_cloudwatch_metrics=True,
)
# -
# ### Start generating some artificial traffic
# The cell below starts a thread to send some traffic to the endpoint. Note that you need to stop the kernel to terminate this thread. If there is no traffic, the monitoring jobs are marked as `Failed` since there is no data to process.
# +
# cell 18
from threading import Thread
from time import sleep
import time
endpoint_name=predictor.endpoint_name
runtime_client = boto3.client('runtime.sagemaker')
# (just repeating code from above for convenience/ able to run this section independently)
def invoke_endpoint(ep_name, file_name, runtime_client):
with open(file_name, 'r') as f:
for row in f:
payload = row.rstrip('\n')
response = runtime_client.invoke_endpoint(EndpointName=ep_name,
ContentType='text/csv',
Body=payload)
response['Body'].read()
time.sleep(1)
def invoke_endpoint_forever():
while True:
invoke_endpoint(endpoint_name, 'test_data/test-dataset-input-cols.csv', runtime_client)
thread = Thread(target = invoke_endpoint_forever)
thread.start()
# Note that you need to stop the kernel to stop the invocations
# -
# ### Describe and inspect the schedule
# Once you describe, observe that the MonitoringScheduleStatus changes to Scheduled.
# cell 19
desc_schedule_result = my_default_monitor.describe_schedule()
print('Schedule status: {}'.format(desc_schedule_result['MonitoringScheduleStatus']))
# ### List executions
# The schedule starts jobs at the previously specified intervals. Here, you list the latest five executions. Note that if you are kicking this off after creating the hourly schedule, you might find the executions empty. You might have to wait until you cross the hour boundary (in UTC) to see executions kick off. The code below has the logic for waiting.
#
# Note: Even for an hourly schedule, Amazon SageMaker has a buffer period of 20 minutes to schedule your execution. You might see your execution start in anywhere from zero to ~20 minutes from the hour boundary. This is expected and done for load balancing in the backend.
# +
# cell 20
mon_executions = my_default_monitor.list_executions()
print("We created a hourly schedule above and it will kick off executions ON the hour (plus 0 - 20 min buffer.\nWe will have to wait till we hit the hour...")
while len(mon_executions) == 0:
print("Waiting for the 1st execution to happen...")
time.sleep(60)
mon_executions = my_default_monitor.list_executions()
# -
# ### Inspect a specific execution (latest execution)
# In the previous cell, you picked up the latest completed or failed scheduled execution. Here are the possible terminal states and what each of them mean:
# * Completed - This means the monitoring execution completed and no issues were found in the violations report.
# * CompletedWithViolations - This means the execution completed, but constraint violations were detected.
# * Failed - The monitoring execution failed, maybe due to client error (perhaps incorrect role premissions) or infrastructure issues. Further examination of FailureReason and ExitMessage is necessary to identify what exactly happened.
# * Stopped - job exceeded max runtime or was manually stopped.
# +
# cell 21
latest_execution = mon_executions[-1] # latest execution's index is -1, second to last is -2 and so on..
time.sleep(60)
latest_execution.wait(logs=False)
print("Latest execution status: {}".format(latest_execution.describe()['ProcessingJobStatus']))
print("Latest execution result: {}".format(latest_execution.describe()['ExitMessage']))
latest_job = latest_execution.describe()
if (latest_job['ProcessingJobStatus'] != 'Completed'):
print("====STOP==== \n No completed executions to inspect further. Please wait till an execution completes or investigate previously reported failures.")
# -
# cell 22
report_uri=latest_execution.output.destination
print('Report Uri: {}'.format(report_uri))
# ### List the generated reports
# +
# cell 23
from urllib.parse import urlparse
s3uri = urlparse(report_uri)
report_bucket = s3uri.netloc
report_key = s3uri.path.lstrip('/')
print('Report bucket: {}'.format(report_bucket))
print('Report key: {}'.format(report_key))
s3_client = boto3.Session().client('s3')
result = s3_client.list_objects(Bucket=report_bucket, Prefix=report_key)
report_files = [report_file.get("Key") for report_file in result.get('Contents')]
print("Found Report Files:")
print("\n ".join(report_files))
# -
# ### Violations report
# If there are any violations compared to the baseline, they will be listed here.
# cell 24
violations = my_default_monitor.latest_monitoring_constraint_violations()
pd.set_option('display.max_colwidth', None)
constraints_df = pd.json_normalize(violations.body_dict["violations"])
constraints_df.head(10)
# ### Other commands
# We can also start and stop the monitoring schedules.
# +
# cell 25
#my_default_monitor.stop_monitoring_schedule()
#my_default_monitor.start_monitoring_schedule()
# -
# ## Delete the resources
#
# You can keep your endpoint running to continue capturing data. If you do not plan to collect more data or use this endpoint further, you should delete the endpoint to avoid incurring additional charges. Note that deleting your endpoint does not delete the data that was captured during the model invocations. That data persists in Amazon S3 until you delete it yourself.
#
# But before that, you need to delete the schedule first.
# cell 26
my_default_monitor.delete_monitoring_schedule()
time.sleep(60) # actually wait for the deletion
# cell 27
predictor.delete_endpoint()
# cell 28
predictor.delete_model()
| SageMaker-ModelMonitoring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
import torchvision.models as models
import torch.nn as nn
from subspace_wrapper import to_subspace_class
import numpy as np
# +
SubLinear = to_subspace_class(nn.Linear, num_vertices=3, verbose=True)
l = SubLinear(5,5)
l(torch.rand(5))
# SubResNet = to_subspace_class(models.ResNet, verbose=True)
# sub_resnet18 = SubResNet(models.resnet.BasicBlock, [2, 2, 2, 2])
# sub_resnet18.to('cuda:3')
# out = sub_resnet18(torch.rand(100, 3, 224, 224).to('cuda:3'))
# -
ln = nn.Linear(5,5)
l.state_dict().keys()
ln.state_dict().keys()
incompatible_keys = l.load_state_dict(ln.state_dict())
incompatible_keys
# + pycharm={"name": "#%%\n"}
l.orig_parameter_names
# + pycharm={"name": "#%%\n"}
l.state_dict()['parametrization_points.4']
# + pycharm={"name": "#%%\n"}
ln.state_dict()[l.param_point_keys_to_orig_state_keys['parametrization_points.4']]
# -
device = 'cuda:3'
SubspaceTransformer = to_subspace_class(nn.Transformer, num_vertices=1, verbose=True)
transformer_model = SubspaceTransformer(nhead=16, num_encoder_layers=12).to(device)
src = torch.rand((10, 32, 512)).to(device)
tgt = torch.rand((20, 32, 512)).to(device)
out = transformer_model(src, tgt)
out.shape
[1,2,3][np.floor(5/3).astype(int)]
| subspace_wrapper_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train, hyperparameter tune and test the TV Script Generation model
# ## Prerequisites
# If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, you have to install the Azure Machine Learning Python SDK and create an Azure ML Workspace first.
# +
# Check core SDK version number
import azureml.core
print("Azure SDK version:", azureml.core.VERSION)
# -
# ## Initialize workspace
#
# Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. Workspace.from_config() creates a workspace object from the details stored in config.json.
# +
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# -
# ## Create or Attach existing AmlCompute
#
# **Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace, this code will skip the creation process.
#
# As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
cluster_name = "gpu-cluster"
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing compute target')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = AmlCompute.provisioning_configuration(
vm_size='Standard_NC6',
vm_priority="dedicated",
min_nodes = 0,
max_nodes = 12,
idle_seconds_before_scaledown=300
)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it uses the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# use get_status() to get a detailed status for the current cluster.
print(compute_target.get_status().serialize())
# -
# ## Train model on the remote compute
#
# I need to setup my training environment with the following steps
# - Create a project directory & add training assets like scripts and data
# - Create an Azure ML experiment
# - Create an environment
# - Configure & submit the training job
#
# ### Create a project directory & add training assets
#
# Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on.
# +
import os
import shutil
import glob
project_folder = './train'
os.makedirs(project_folder, exist_ok=True)
for script in glob.glob('../*.py'):
shutil.copy(script, project_folder)
for script in glob.glob('*.py'):
shutil.copy(script, project_folder)
data_folder = project_folder + '/data'
os.makedirs(data_folder, exist_ok=True)
for txt in glob.glob('../data/*.txt'):
shutil.copy(txt, data_folder)
# -
# ### Create an experiment
# Create an Experiment to track all the runs in your workspace.
# +
from azureml.core import Experiment
experiment_name = 'tv-script-generation'
experiment = Experiment(ws, name=experiment_name)
# -
# ### Create an environment
#
# Define a conda environment YAML file with your training script dependencies and create an Azure ML environment.
# +
# %%writefile conda_dependencies.yml
channels:
- conda-forge
dependencies:
- python=3.6.2
- pip:
- azureml-defaults
- torch==1.6.0
- torchvision==0.7.0
- future==0.17.1
- torchsummary
- torchsummaryX
- pillow
# +
from azureml.core import Environment
pytorch_env = Environment.from_conda_specification(name = 'pytorch-1.6-gpu', file_path = './conda_dependencies.yml')
# Specify a GPU base image
#pytorch_env.docker.enabled = True
pytorch_env.docker.base_image = 'mcr.microsoft.com/azureml/openmpi3.1.2-cuda10.1-cudnn7-ubuntu18.04'
# -
# ### Configure the training job
# Create a ScriptRunConfig object to specify the configuration details of your training job, including your training script, environment to use, and the compute target to run on.
# +
from azureml.core import ScriptRunConfig
from azureml.core.runconfig import DockerConfiguration
args = [
'--num_epochs', 15,
'--batch_size', 256,
'--learning_rate', 0.0005,
'--sequence_length', 10,
'--embedding_dim', 300,
'--hidden_dim', 400,
'--num_layers', 2,
'--output_dir', './outputs'
]
docker_config = DockerConfiguration(use_docker=True)
script_run_config = ScriptRunConfig(source_directory=project_folder,
script='train.py',
arguments=args,
compute_target=compute_target,
environment=pytorch_env,
docker_runtime_config=docker_config)
# -
# ### Submit job
# Run your experiment by submitting your ScriptRunConfig object. Note that this call is asynchronous.
test_run = experiment.submit(script_run_config)
# ### Monitor your run
# You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
# +
from azureml.widgets import RunDetails
RunDetails(test_run).show()
# -
# Alternatively, you can block until the script has completed training before running more code.
test_run.wait_for_completion(show_output=True)
# ## Hyperparameter Tuning
#
# Run 3 Hyperparameter Tuning experiments:
# - Combintation of Embedding & Hidden dimensions
# - Sequence Length
# - Number of Hidden Layers
# +
from azureml.train.hyperdrive import GridParameterSampling
from azureml.train.hyperdrive import HyperDriveConfig, PrimaryMetricGoal
from azureml.train.hyperdrive import choice
param_sampling = GridParameterSampling( {
"embedding_dim": choice(range(200, 601, 50)),
"hidden_dim": choice(range(200, 601, 50))
}
)
hyperdrive_config = HyperDriveConfig(run_config=script_run_config,
hyperparameter_sampling=param_sampling,
primary_metric_name='train_loss',
primary_metric_goal=PrimaryMetricGoal.MINIMIZE,
max_total_runs=100,
max_concurrent_runs=4)
hyperdrive_run_embedding_hidden_dim = experiment.submit(hyperdrive_config)
# -
# 
#
# This showed that fewer layers produced the better results in this case.
# +
from azureml.train.hyperdrive import GridParameterSampling
from azureml.train.hyperdrive import HyperDriveConfig, PrimaryMetricGoal
from azureml.train.hyperdrive import choice
param_sampling = GridParameterSampling( {
"sequence_length": choice(range(5, 51, 5))
}
)
hyperdrive_config = HyperDriveConfig(run_config=script_run_config,
hyperparameter_sampling=param_sampling,
primary_metric_name='train_loss',
primary_metric_goal=PrimaryMetricGoal.MINIMIZE,
max_total_runs=100,
max_concurrent_runs=4)
hyperdrive_run_sequence_length = experiment.submit(hyperdrive_config)
# -
# 
#
# The sweet spot here seemed to be 25 or 50 words.
#
# However, since the training times increased with the corresponding sequence length, I opted for a length of 25 words here.
# +
from azureml.train.hyperdrive import GridParameterSampling
from azureml.train.hyperdrive import HyperDriveConfig, PrimaryMetricGoal
from azureml.train.hyperdrive import choice
param_sampling = GridParameterSampling( {
"num_layers": choice(range(1, 4))
}
)
hyperdrive_config = HyperDriveConfig(run_config=script_run_config,
hyperparameter_sampling=param_sampling,
primary_metric_name='train_loss',
primary_metric_goal=PrimaryMetricGoal.MINIMIZE,
max_total_runs=100,
max_concurrent_runs=4)
hyperdrive_run_num_layers = experiment.submit(hyperdrive_config)
# -
# 
#
# As it turned out, the model converged best with the highest values, so I did another HyperDrive experiment with correspondingly higher ranges.
hyperdrive_run_num_layers.wait_for_completion()
hyperdrive_run_sequence_length.wait_for_completion()
hyperdrive_run_embedding_hidden_dim.wait_for_completion()
# ## Re-Train
#
# Re-Train the RNN model based on the outcome of the first round of tuning.
# +
from azureml.core import ScriptRunConfig
from azureml.core.runconfig import DockerConfiguration
args = [
'--num_epochs', 15,
'--batch_size', 256,
'--learning_rate', 0.001,
'--sequence_length', 25,
'--embedding_dim', 600,
'--hidden_dim', 600,
'--num_layers', 1,
'--output_dir', './outputs'
]
docker_config = DockerConfiguration(use_docker=True)
script_run_config = ScriptRunConfig(source_directory=project_folder,
script='train.py',
arguments=args,
compute_target=compute_target,
environment=pytorch_env,
docker_runtime_config=docker_config)
# -
test_run = experiment.submit(script_run_config)
# +
from azureml.widgets import RunDetails
RunDetails(test_run).show()
# -
test_run.wait_for_completion()
# +
from azureml.train.hyperdrive import BayesianParameterSampling
from azureml.train.hyperdrive import HyperDriveConfig, PrimaryMetricGoal
from azureml.train.hyperdrive import choice, quniform, qnormal
param_sampling = BayesianParameterSampling( {
"embedding_dim": quniform(400, 1101, 10),
"hidden_dim": quniform(400, 1101, 10),
"num_layers": choice(range(1,3))
}
)
hyperdrive_config = HyperDriveConfig(run_config=script_run_config,
hyperparameter_sampling=param_sampling,
policy=None,
primary_metric_name='train_loss',
primary_metric_goal=PrimaryMetricGoal.MINIMIZE,
max_total_runs=96,
max_duration_minutes=1440,
max_concurrent_runs=12)
# -
# start the HyperDrive run
hyperdrive_run = experiment.submit(hyperdrive_config)
RunDetails(hyperdrive_run).show()
# 
#
| hyperparameter_tuning/hyperparameter_tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 根据fake_tweets产生的数据文件进行分析
from my_weapon import *
from collections import defaultdict, Counter
from IPython.display import display
from tqdm import tqdm_notebook as tqdm
# %time groups_url_tweets = [json.load(open("disk/url_ts_media_{}.json".format(i))) for i in range(8)]
# +
map_labels = {
"0": "fake",
"1": "extreme bias (right)",
"2": "right",
"3": "right leaning",
"4": "center",
"5": "left leaning",
"6": "left",
"7": "extreme bias (left)"
}
def get_cdf_pdf(data, mini, maxi, step, norm=True):
x = np.arange(mini, maxi, step)
# print(x, len(x))
cdf = np.zeros(len(x))
pdf = np.zeros(len(x))
cnt = 0
for d in data:
if d < mini or d >= maxi:
continue
cnt += 1
for i, _x in enumerate(x):
if d >= _x:
cdf[i] += 1
for i, _x in enumerate(x):
if i == (len(x) - 1):
pdf[i] += 1
elif _x <= d < x[i+1]:
pdf[i] += 1
break
# print(cdf[0], cnt, sum(pdf))
if norm:
cdf = cdf / cnt
pdf = pdf / cnt
# print(cdf)
return x, cdf, pdf
num_of_tweets = {}
plt.figure(figsize=(13, 8))
for i in range(8):
print(i, "...")
url_tweets = groups_url_tweets[i]
data = pd.Series([len(url_tweet["tweets"]) for url_tweet in url_tweets])
num_of_tweets[map_labels[str(i)]] = data
x, cdf, pdf = get_cdf_pdf(data, 0, 20000, 500)
_zero_i = -1
for j in range(len(cdf)):
if cdf[j] == 0:
_zero_i = j
break
x = x[:_zero_i]
cdf = cdf[:_zero_i]
plt.plot(x, cdf, label=map_labels[str(i)], lw=2)
# data.hist(bins=50)
plt.title('Distribution of number of tweets in each news', fontsize=15)
plt.ylabel('CCDF', fontsize=14)
plt.xlabel('N of tweets in this news', fontsize=14)
plt.xticks(fontsize=13); plt.yticks(fontsize=13)
plt.legend()
plt.yscale("log")
# plt.xscale("log")
# plt.savefig('fig/distribution_size_fake_news.pdf', dpi=300)
plt.show()
plt.close()
# -
# ## 针对上图的描述性统计
pd.DataFrame(num_of_tweets).describe()
# +
num_of_sources = {}
plt.figure(figsize=(13, 8))
for i in range(8):
print(i, "...")
url_tweets = groups_url_tweets[i]
data = []
for url_tweet in url_tweets:
_len = len([t for t in url_tweet["tweets"] if t["is_source"]==1])
data.append(_len)
if _len == 0:
print(url_tweet["URL"], len(url_tweet["tweets"]))
data = pd.Series(data)
num_of_sources[map_labels[str(i)]] = data
x, cdf, pdf = get_cdf_pdf(data, 0, 10000, 500)
_zero_i = -1
for j in range(len(cdf)):
if cdf[j] == 0:
_zero_i = j
break
x = x[:_zero_i]
cdf = cdf[:_zero_i]
plt.plot(x, cdf, label=map_labels[str(i)], lw=2)
# data.hist(bins=50)
plt.title('Distribution of number of source tweets in each news', fontsize=15)
plt.ylabel('N of news', fontsize=14)
plt.xlabel('N of tweets referring to this news', fontsize=14)
plt.xticks(fontsize=13); plt.yticks(fontsize=13)
plt.legend()
plt.yscale("log")
# plt.xscale("log")
# plt.savefig('fig/distribution_size_fake_news.pdf', dpi=300)
plt.show()
plt.close()
# -
pd.DataFrame(num_of_sources).describe()
# ### 专门针对用户的分析,放到analyze_users.ipy 和 analyze_IRAs.ipynb 中~
# +
import pendulum
def plot_day(i, url, sorted_dts, sorted_dts2=None, save=False):
"""
包含了两条线!
"""
plt.figure(figsize=(10, 6))
ts = cal_ts_day(sorted_dts)
ts.plot()
if sorted_dts2:
ts2 = cal_ts_day(sorted_dts2)
ts2.plot()
# configure
plt.ylabel('N of tweets with this fake news', fontsize=15)
plt.xticks(fontsize=11); plt.yticks(fontsize=11)
# plt.xlabel('$Date$', fontsize=15)
# plt.title(url)
if save:
plt.savefig('fig/{}-{}-overall-spread.pdf'.format(i, url), dpi=300)
else:
plt.show()
plt.close()
def plot_48hours(i, url, sorted_dts, sorted_dts2=None, save=False):
"""
包含了两条线!
"""
# print(url)
# print("实际传播开始和结束时间:", sorted_dts[0], sorted_dts[-1])
plt.figure(figsize=(10, 6))
ts = cal_ts_48hours(sorted_dts)
ts.plot()
if sorted_dts2:
ts2 = cal_ts_48hours(sorted_dts2)
ts2.plot()
# configure
plt.ylabel('N of tweets with this fake news', fontsize=15)
plt.xticks(fontsize=11); plt.yticks(fontsize=11)
# plt.xlabel('$Date$', fontsize=15)
# plt.title(url)
if save:
plt.savefig('fig/{}-{}-first-48-hours.pdf'.format(i, url), dpi=300)
else:
plt.show()
plt.close()
# +
for i, url_tweet in enumerate(url_tweets):
if i == 0:
continue
if i >= 3:
break
print("plot {} ~".format(i))
url = url_tweet["url"]
tweets_list = url_tweet["tweets"]
dts = [t["dt"] for t in tweets_list]
dts2 = [t["dt"] for t in tweets_list if t["is_source"]==1]
plot_day(i, url, dts, dts2)
plot_48hours(i, url, dts, dts2)
# plot_day(i, url, dts)
# plot_48hours(i, url, dts)
# -
# ---
#
# ## 意见领袖定义
# +
user_data = {}
x = []
y = []
for line in tqdm(open("disk/user_info.json")):
w = line.strip().split(",")
if int(w[2]) >= 1 and int(w[3]) >= 1:
user_data[w[0]] = {
"name": w[1],
"fol": int(w[2]),
"fri": int(w[3]),
}
x.append(int(w[2]))
y.append(int(w[3]))
# -
x = pd.Series(x).apply(np.log10)
y = pd.Series(y).apply(np.log10)
# +
sns.set_style("darkgrid")
# sns.jointplot(x, y, kind="hex")
# plt.xlim(0.5, 4.5)
# plt.ylim(0.5, 4.5)
# plt.legend()
plt.figure(figsize=(10, 10))
ax = plt.subplot(211)
x.hist(bins=50)
ax = plt.subplot(212)
y.hist(bins=50)
plt.show()
# -
| analyze_spreading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Armenian Data Cube - Shoreline Delineation Notebook
#
# **Description:**
# This Python notebook allows users to monitor small water area and particularly study of changes of shorelines combining Sentinel 2 and UAV photogrammetry. The script is structured as follows:
#
# **Step 1: Preprocessing**
# - Read RGB image from UAV image using bands and take from metadata the boundaries.
# - Using that boundaries, fetch data from Datacube for Sentinel 2 for the closest date to the UAV image data. The used bands are ['green','red', ‘blue’, 'nir', 'swir1', 'swir2' ].
#
#
# **Step 2: To identify and extract water bodies**
# - BandRatio = B3/B8
# - McFeeters = (B3 - B8) / (B3 + B8)
# - MSAVI12 = (2*B8 + 1 - xu.sqrt((2*B8)*(2*B8) - 8*(B8 - B4))) / 2
# - MNDWI1 = (B3 - B11) / (B3 + B11)
# - MNDWI2 = (B3 - B12) / (B3 + B12)
#
# **Step 3: To extract coastline**
# - K-means clustering method
# - Gaussian Blur
# - Canny edge detection
#
# After these steps we will have the shoreline matrix where in the positions of the line will have values 1 and the other positions 0.
#
# **Step 4: To compere UAV and Satelite images**
# - Mask the shoreline matrix with initial images
# - Get the coordinates in (lat,long) pairs
# - For each Satellite image shoreline point to find the nearest points from the UAV image with Euclidean distance and calculate the RMSE for the whole line
# **Import necessary Data Cube libraries and dependencies.**
from logic import *
show()
| ShorelineDetectionUAV/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def eq(x,y,z,r):
return x**1 + y**1< z**2 #replace with your equation
def cordfind(x0, y0, z0, r):
import math;
points =[];
#build blocks in z-axis
for x in range(x0-r-1, x0+r+1):
for y in range(y0-r-1, y0+r+1):
prev_inside = False
for z in range(z0-r-1, z0+r+1):
inside = eq(x,y,z,r)
if inside != prev_inside:
points.append((x,y,z))
prev_inside = inside
#build blocks in x-axis
for y in range(x0-r-1, x0+r+1):
for z in range(y0-r-1, y0+r+1):
prev_inside = False
for x in range(z0-r-1, z0+r+1):
inside = eq(x,y,z,r)
if inside != prev_inside:
points.append((x,y,z))
prev_inside = inside
#build blocks in y-axis
for z in range(x0-r-1, x0+r+1):
for x in range(y0-r-1, y0+r+1):
prev_inside = False
for y in range(z0-r-1, z0+r+1):
inside = eq(x,y,z,r)
if inside != prev_inside:
points.append((x,y,z))
prev_inside = inside
res = []
for i in points:
if i not in res:
res.append(i)
return(res);
# -
#generate minecraft function
points = cordfind(0,0,0,20)
print(len(points))
with open('C:\Users\user-name\AppData\Roaming\.minecraft\saves\world-name\datapacks\datapack-name\data\namespace\functions\function-name.mcfunction', 'w') as towrite:
for point in points:
command = "setblock ~%d ~%d ~%d oak_wood\n" %(point[0],point[1],point[2]); #replace oak wood with a needed block
towrite.write(command);
| implicitbuilder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Object Detection and Bounding Boxes
# :label:`sec_bbox`
#
#
# In the previous section, we introduced many models for image classification. In image classification tasks, we assume that there is only one main target in the image and we only focus on how to identify the target category. However, in many situations, there are multiple targets in the image that we are interested in. We not only want to classify them, but also want to obtain their specific positions in the image. In computer vision, we refer to such tasks as object detection (or object recognition).
#
# Object detection is widely used in many fields. For example, in self-driving technology, we need to plan routes by identifying the locations of vehicles, pedestrians, roads, and obstacles in the captured video image. Robots often perform this type of task to detect targets of interest. Systems in the security field need to detect abnormal targets, such as intruders or bombs.
#
# In the next few sections, we will introduce multiple deep learning models used for object detection. Before that, we should discuss the concept of target location. First, import the packages and modules required for the experiment.
#
# + origin_pos=2 tab=["pytorch"]
# %matplotlib inline
import torch
from d2l import torch as d2l
# + [markdown] origin_pos=4
# Next, we will load the sample images that will be used in this section. We can see there is a dog on the left side of the image and a cat on the right. They are the two main targets in this image.
#
# + origin_pos=6 tab=["pytorch"]
d2l.set_figsize()
img = d2l.plt.imread('../img/catdog.jpg')
d2l.plt.imshow(img);
# + [markdown] origin_pos=7
# ## Bounding Box
#
# In object detection, we usually use a bounding box to describe the target location.
# The bounding box is a rectangular box that can be determined by the $x$ and
# $y$ axis coordinates in the upper-left corner and the $x$ and $y$ axis
# coordinates in the lower-right corner of the rectangle.
# Another commonly used bounding box representation is the $x$ and $y$ axis
# coordinates of the bounding box center, and its width and height.
# Here we define functions to convert between these two
# representations, `box_corner_to_center` converts from the two-corner
# representation to the center-width-height presentation,
# and `box_center_to_corner` vice verse.
# The input argument `boxes` can be either a length $4$ tensor,
# or a $(N, 4)$ 2-dimensional tensor.
#
# + origin_pos=8 tab=["pytorch"]
#@save
def box_corner_to_center(boxes):
"""Convert from (upper_left, bottom_right) to (center, width, height)"""
x1, y1, x2, y2 = boxes[:, 0], boxes[:, 1], boxes[:, 2], boxes[:, 3]
cx = (x1 + x2) / 2
cy = (y1 + y2) / 2
w = x2 - x1
h = y2 - y1
boxes = torch.stack((cx, cy, w, h), axis=-1)
return boxes
#@save
def box_center_to_corner(boxes):
"""Convert from (center, width, height) to (upper_left, bottom_right)"""
cx, cy, w, h = boxes[:, 0], boxes[:, 1], boxes[:, 2], boxes[:, 3]
x1 = cx - 0.5 * w
y1 = cy - 0.5 * h
x2 = cx + 0.5 * w
y2 = cy + 0.5 * h
boxes = torch.stack((x1, y1, x2, y2), axis=-1)
return boxes
# + [markdown] origin_pos=9
# We will define the bounding boxes of the dog and the cat in the image based
# on the coordinate information. The origin of the coordinates in the image
# is the upper left corner of the image, and to the right and down are the
# positive directions of the $x$ axis and the $y$ axis, respectively.
#
# + origin_pos=10 tab=["pytorch"]
# bbox is the abbreviation for bounding box
dog_bbox, cat_bbox = [60.0, 45.0, 378.0, 516.0], [400.0, 112.0, 655.0, 493.0]
# + [markdown] origin_pos=11
# We can verify the correctness of box conversion functions by converting twice.
#
# + origin_pos=12 tab=["pytorch"]
boxes = torch.tensor((dog_bbox, cat_bbox))
box_center_to_corner(box_corner_to_center(boxes)) - boxes
# + [markdown] origin_pos=13
# We can draw the bounding box in the image to check if it is accurate. Before drawing the box, we will define a helper function `bbox_to_rect`. It represents the bounding box in the bounding box format of `matplotlib`.
#
# + origin_pos=14 tab=["pytorch"]
#@save
def bbox_to_rect(bbox, color):
"""Convert bounding box to matplotlib format."""
# Convert the bounding box (top-left x, top-left y, bottom-right x,
# bottom-right y) format to matplotlib format: ((upper-left x,
# upper-left y), width, height)
return d2l.plt.Rectangle(xy=(bbox[0], bbox[1]), width=bbox[2] - bbox[0],
height=bbox[3] - bbox[1], fill=False,
edgecolor=color, linewidth=2)
# + [markdown] origin_pos=15
# After loading the bounding box on the image, we can see that the main outline of the target is basically inside the box.
#
# + origin_pos=16 tab=["pytorch"]
fig = d2l.plt.imshow(img)
fig.axes.add_patch(bbox_to_rect(dog_bbox, 'blue'))
fig.axes.add_patch(bbox_to_rect(cat_bbox, 'red'));
# + [markdown] origin_pos=17
# ## Summary
#
# * In object detection, we not only need to identify all the objects of interest in the image, but also their positions. The positions are generally represented by a rectangular bounding box.
#
# ## Exercises
#
# 1. Find some images and try to label a bounding box that contains the target. Compare the difference between the time it takes to label the bounding box and label the category.
#
# + [markdown] origin_pos=19 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1527)
#
| scripts/d21-en/pytorch/chapter_computer-vision/bounding-box.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ** Today Concepts **
#
# 1.Multi Linear Regression (applying linear Regression with multiple features)
#
# 2.Polynomial Regression(polynomial features)
# ## Multi Linear Regression
# -- Applying linear regression algorithm with multiple featutres
# ### prediction of the house price of boston dataset
# #### 1.get the data
#
from sklearn.datasets import load_boston
boston = load_boston()
boston.keys()
print(boston['DESCR'])
print(boston['target'])
boston['feature_names']
import pandas as pd
df = pd.DataFrame(boston['data'])
df
df.head()
df.tail()
df.sample(5)
df.columns = boston['feature_names']
df.head(3)
df['target'] = boston['target']
df.head(3)
df.shape
# ### 2.Pre processing
## is there any missing values
df.isna().sum()
df.info()
# i am taking RM column randomly as my feature
# and we check the performence of the model .if performence not good we can improve the model
X = df[['RM']]
y = df['target']
# it is better to separate the data for training data and testing data
#
# we can say 70% data for training and 30% data for testing
#
# we have 506 rows available
#
# in that how many rows for training and how many rows for testing
70*506/100 # training rows
506-354 # testing rows
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,train_size = 0.7)
X_train.shape
X_test.shape
# ### train the model
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_train,y_train)
print('training score',model.score(X_train,y_train)*100)
print('testing score',model.score(X_test,y_test)*100)
# **improve the model**
# above score very low so want improve the model
#
# 1.by giving more examples
#
# 2.by taking different features
#
# 3.by parameter tuning
#
# **improving score by different feature selection**
#
# df.corr() gives co relation between the features
#
# if corelation values near to 1 then we can say positive linearity
#
# if corealtion values near to -1 then we can -ve linearity
#
# if corelation 0 then we can say no relation
df.corr()
# RM -->0.69
# LSTAT -->-0.73
#
# We can take RM&LSTAT as features and again check performance.
# if not uptomark we can apply improve the model
#
X = df[['RM','LSTAT']]
y = df['target']
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,train_size=0.7)
X_train.shape
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_train,y_train)
print('train score',model.score(X_train,y_train)*100)
print('test score',model.score(X_test,y_test)*100)
# ## Applying polynomial features to linear regression
# **underfitting** model not able to recognize the pattern(student not able to learn from the training data)
#
# **overfitting** model capturing all the points but failed in testing.
#
# **best fit** model recognizing the pattern and giving good performence in training and testing
#
#
# **prediction of salay of employee with his experience**
# without polynomial features and with polynomial feature and observe the scores.
#
# #### 1.get the data
# +
experience = [0,1,2,3,4,5,6,7,8]
salary = [5000,6000,7000,8000,15000,25000,40000,55000,70000]
import pandas as pd
df = pd.DataFrame({'experience':experience,'salary':salary})
# -
df
df.shape
## 2.pre-processing
df.isna().sum()
df.info()
X =df[['experience']]
y=df['salary']
# 7 rows data for training and 2 rows data for testing
X_train =X.head(7)
X_test = X.tail(2)
y_train=y.head(7)
y_test = y.tail(2)
X_train.shape
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_train,y_train)
print('train score',model.score(X_train,y_train)*100)
print('test score',model.score(X_test,y_test)*100)
import matplotlib.pyplot as plt
plt.figure()
plt.scatter(df['experience'],df['salary'],c='blue',label='true values')
plt.plot(df['experience'],model.predict(X),c='red',label='predicted line')
plt.legend()
plt.show()
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures()
X_poly_train = poly.fit_transform(X_train)
X_poly_test = poly.transform(X_test)
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_poly_train,y_train)
print('train score',model.score(X_poly_train,y_train)*100)
print('test score',model.score(X_poly_test,y_test)*100)
import matplotlib.pyplot as plt
plt.figure()
plt.scatter(df['experience'],df['salary'],c='green',label='true values')
plt.plot(df['experience'],model.predict(poly.transform(X)),c='red',label='predictedline')
plt.xlabel('experience')
plt.ylabel('salary')
plt.legend()
plt.show()
| Day-3/Day3_18Nov2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="xNBS3nVFkmsO" colab_type="text"
# カブを予測する
# + [markdown] id="HxtrAXVWoYPF" colab_type="text"
# 1.データ読み込み
# + id="SecHomufkiQG" colab_type="code" outputId="2518d8f5-bfc1-47a4-93b8-f8e7472d4c29" colab={"base_uri": "https://localhost:8080/", "height": 255}
import numpy as np
import pandas as pd
dir = ''
#テーブル読み込み
df = pd.read_csv(directory+'data/oimori.csv')
print(df)
# + [markdown] id="7Cm53Oviug22" colab_type="text"
# 2.ラベルエンコーディング
# + id="8l6sH6XiufKX" colab_type="code" outputId="c9949028-a38b-432f-dff6-71a3632b2cfd" colab={"base_uri": "https://localhost:8080/", "height": 204}
#カテゴリ変数の作成
#cat_df = pd.get_dummies(df,columns=['結果'])
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
encoded = le.fit_transform(df['結果'].values)
df['結果'] = encoded
cat_df = df
cat_df.head()
# + [markdown] id="xt0hr7_tpyBH" colab_type="text"
# 3.歯抜けデータの抽出
# + id="TRz-bmuqpBgF" colab_type="code" outputId="549db2eb-fd8d-42f2-d0b8-f425f1fbfff2" colab={"base_uri": "https://localhost:8080/", "height": 204}
columns = ['月曜午前','月曜午後','火曜午前','火曜午後','水曜午前','水曜午後',
'木曜午前','木曜午後','金曜午前','金曜午後','土曜午前','土曜午後'
]
#欠損値を含むデータ
nan_df = cat_df[cat_df.isnull().any(axis=1)]
#欠損がないデータ
cat_df = cat_df.dropna().astype(int)
nan_df.head()
#cat_df.head()
# + [markdown] id="QQIeT8XFw_XC" colab_type="text"
# 3.5.可視化
# + id="UirVadPZxBaC" colab_type="code" outputId="04980ee9-7524-4caa-bf7e-89e98ba4ee6e" colab={"base_uri": "https://localhost:8080/", "height": 717}
import matplotlib.pyplot as plt
#ignoring warnings
import warnings
warnings.filterwarnings('ignore')
#Decision Boundary viewer
# %matplotlib inline
plt.style.use('ggplot')
import itertools
plotarr = cat_df[columns].values
labelarr = cat_df['結果'].values
#create graph
#2images each row
dev = int((np.max(labelarr)+np.max(labelarr)%2)/2)
plots = []
fig, axes = plt.subplots(nrows=dev, ncols= 2, figsize=(6*2,6*dev))
t = np.arange(0,12)
for i,grid in zip(range(0,np.max(labelarr)),itertools.product(list(range(dev)),[0,1])):
plot = plotarr[labelarr == i]
for j in plot:
axes[grid].plot(t,j[:])
#axes[grid].xlabel('youbi')
#axes[grid].ylabel('bell')
axes[grid].set_title(i)
# + [markdown] id="Hln9c1Pzws5E" colab_type="text"
# 4.データの正規化
# + id="KVqYt_4TwsV7" colab_type="code" outputId="6dd05622-1193-4157-fdb8-fc491219b9d7" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.preprocessing import MinMaxScaler
#データ
data = cat_df.values[:,1:-1]
print(data[0].shape)
#ラベル
data_labels = cat_df.values[:,-1:]
#欠損データ
nan = nan_df.values[:,1:-1]
#欠損データラベル
nan_labels = nan_df.values[:,-1:]
#正規化
scaler = MinMaxScaler().fit(data)
data = scaler.transform(data)
nan = scaler.transform(nan)
# + [markdown] id="-BIw8vrCMnyS" colab_type="text"
# 5.npyに出力
# + id="9rXnkSlkKFNZ" colab_type="code" outputId="d466c160-2629-44b6-9037-f54072fdfd80" colab={"base_uri": "https://localhost:8080/", "height": 88}
#保存ディレクトリ
#データ
np.save(directory+'data/data.npy',data)
np.save(directory+'data/data_labels.npy',data_labels)
#欠損
np.save(directory+'data/nan.npy',nan)
np.save(directory+'data/nan_labels.npy',nan_labels)
#minmaxscaler
from sklearn.externals import joblib
joblib.dump(scaler,directory+'data/scaler.save')
#labelencoder
joblib.dump(le,directory+'data/labelencoder.save')
| 1_KabuPreprocessor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%time
import pandas as pd
import numpy as np
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from imblearn.over_sampling import SMOTE,RandomOverSampler
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import RobustScaler
from sklearn.preprocessing import Normalizer
from itertools import combinations
from mlxtend.classifier import StackingClassifier
from sklearn import model_selection
df=pd.read_csv('60s_window_wrist_chest.csv',index_col=0)
df
# +
features=df.columns.tolist()
features
removed = ['label']
for rem in removed:
features.remove(rem)
features_with_sub=[]
features_with_sub[:]=features
removed = ['subject']
for rem in removed:
features.remove(rem)
feature=features
print(len(feature))
len(features_with_sub)
# -
sm = SMOTE(random_state=2)
X, y= sm.fit_sample(df[features_with_sub], df['label'])
df_new=pd.concat([pd.DataFrame(X,columns=features_with_sub),pd.DataFrame(y,columns=['label'])],axis=1)
df_new
for i in range (len(list(df_new['subject']))):
df_new['subject'][i] = min([2,3,4,5,6,7,8,9,10,11,13,14,15,16,17], key=lambda x:abs(x-df_new['subject'][i]))
df_new['subject']=df_new['subject'].astype(int)
p_d=pd.read_csv('personal_detail.csv',index_col=0)
df_new_1=df_new.merge(p_d,on='subject')
df_new_1
df_new_1['label'].value_counts()
# +
features=df_new_1.columns.tolist()
features
removed = ['label']
for rem in removed:
features.remove(rem)
features_with_sub=[]
features_with_sub[:]=features
removed = ['subject']
for rem in removed:
features.remove(rem)
feature=features
print(len(feature))
len(features_with_sub)
# -
train=df_new_1[df_new_1['subject']<=9]
test=df_new_1[df_new_1['subject']>9]
scaler = Normalizer()
scaled_data_train = scaler.fit_transform(train[feature])
scaled_data_test = scaler.transform(test[feature])
rf = RandomForestClassifier(n_estimators=100,n_jobs=10,random_state=56)
rf.fit(scaled_data_train,train['label'])
y_pred=et.predict(scaled_data_test)
print(classification_report(test['label'],y_pred))
# # STACKED
#
classification_report(test['label'],y_pred,output_dict = True)['accuracy']
rnd_st = []
for i in range (501):
rf = RandomForestClassifier(n_estimators=100,n_jobs=10,random_state=i ,)
rf.fit(scaled_data_train,train['label'])
y_pred=et.predict(scaled_data_test)
if ((classification_report(test['label'],y_pred,output_dict=True)['0']['recall'])>.60 and (classification_report(test['label'],y_pred,output_dict=True)['2']['recall'])>.60 and (classification_report(test['label'],y_pred,output_dict=True)['3']['recall'])>.60 and (classification_report(test['label'],y_pred,output_dict=True)['accuracy'])>.74):
print(i)
rnd_st.append(i)
clf = []
for st in rnd_st:
clf.append(RandomForestClassifier(n_estimators=100,n_jobs=10,random_state=st))
meta = ExtraTreesClassifier(n_estimators=100,n_jobs=10,random_state=56)
sclf = StackingClassifier(classifiers=clf, meta_classifier=meta)
sclf.fit(scaled_data_train,train['label'])
y_pred_sta=sclf.predict(scaled_data_test)
print(classification_report(test['label'],y_pred_sta))
# +
# Stacked random state:
# 72
# 86
# 235
# 388
# 396
for i in range (500):
meta = RandomForestClassifier(n_estimators=100,n_jobs=10,random_state=i)
sclf = StackingClassifier(classifiers=clf, meta_classifier=meta)
sclf.fit(scaled_data_train,train['label'])
y_pred_sta=sclf.predict(scaled_data_test)
if (classification_report(test['label'],y_pred_sta,output_dict=True)['accuracy']>.75):
print (i)
print(classification_report(test['label'],y_pred_sta))
# -
| User Independence Analysis/ipynb/.ipynb_checkpoints/Stacked model (Random Forest)-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# %matplotlib inline
# +
from sklearn import datasets
iris = datasets.load_iris()
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
y_pred = gnb.fit(iris.data, iris.target).predict(iris.data)
y_pred
print("Number of mislabeled points out of a total %d points : %d"
% (iris.data.shape[0],(iris.target != y_pred).sum()))
for i in range(len(y_pred)):
if(iris.target[i] != y_pred[i]):
print("Actual target %d and Predicted target %d "%(iris.target.data[i],y_pred[i]))
correct = (iris.target == y_pred).sum()
accuracy = (correct/150)*100
accuracy
# -
| Naive Bayes using sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
df = pd.read_csv("dataset.csv")
df.head()
df.describe()
df.info()
df[df.isnull().any(axis=1)]
tenure_meean = df["tenure"].mean()
df.fillna({"SeniorCitizen":0.0,"tenure":tenure_meean},inplace=True)
df.info()
X = df.iloc[:,:-1].values
Y = df.iloc[:,-1]
print(X)
print(Y)
## another way to fill NaN using Imputer
from sklearn.preprocessing import Imputer
imputer = Imputer(missing_values="NaN",strategy="mean",axis=0)
#imputer = Imputer(missing_values="NaN",startegy="medain",axis=0)
#imputer = Imputer(missing_values="NaN",startegy="most_frequent",axis=0
imputer = imputer.fit(X[:,5:6])
tenure_new = imputer.transform(X[:,5:6])
print(tenure_new)
imputer2 = Imputer(missing_values="NaN",strategy="most_frequent",axis=0)
imputer2 = imputer2.fit(X[:,2:3])
seniorCitizen_new = imputer.transform(X[:,2:3])
print(seniorCitizen_new)
## label encoding
from sklearn.preprocessing import LabelEncoder
label = LabelEncoder()
gender = label.fit_transform(df["gender"].values)
gender # based on alphapetical characters f before m
from sklearn.preprocessing import OneHotEncoder
ohe = OneHotEncoder()
new_df = df[['MultipleLines','PaymentMethod']]
new_df.head()
new_df_enc = ohe.fit_transform(new_df).toarray()
# standardization
# X standard = x - mean / standard devation
x = pd.DataFrame({"age":[20,25,30],"salary":[2000,3000,5000]})
# + active=""
# Normalization
# x-min(x) \ max(x) - min(x)
# -
#using library
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
# ## Test
# ### Original Dataset
df = pd.read_csv("dataset.csv")
df.head()
# ### Feature Enginering
my_df = df[["customerID","gender","SeniorCitizen","tenure","Dependents","MonthlyCharges","TotalCharges","Churn"]]
my_df.head()
# ### fill Missing Values
tenure_meean = my_df["tenure"].mean()
my_df.fillna({"SeniorCitizen":0.0,"tenure":tenure_meean},inplace=True)
print(my_df.info())
my_df.head()
# ### Encoding
from sklearn.preprocessing import LabelEncoder
my_df[['gender']] = LabelEncoder().fit_transform(df[['gender']])
#______________________________
new_SeniorCitizen = pd.DataFrame(my_df[ 'SeniorCitizen' ], dtype=int)
my_df["SeniorCitizen"] = new_SeniorCitizen
#_______________________________
my_df[['Dependents']] = LabelEncoder().fit_transform(df[['Dependents']])
#_______________________________
my_df[['Churn']] = LabelEncoder().fit_transform(df[['Churn']])
my_df.head()
# ### Standardization
from sklearn.preprocessing import StandardScaler
my_df[['tenure', 'TotalCharges','MonthlyCharges']] = StandardScaler().fit_transform(my_df[['tenure', 'TotalCharges','MonthlyCharges']])
my_df.head()
| Pre-Processing Case/Pre-Processing .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Estilos y color
#
# En este ejercicio se demostrara como controlar la estetica de las graficas en seaborn:
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
tips = sns.load_dataset('tips')
# ## Estilos
#
# Se puede implementar un estilo en particular:
sns.countplot(x='sex',data=tips)
sns.set_style('white')
sns.countplot(x='sex',data=tips)
sns.set_style('ticks')
sns.countplot(x='sex',data=tips,palette='deep')
# ## Remover contorno
sns.countplot(x='sex',data=tips)
sns.despine()
sns.countplot(x='sex',data=tips)
sns.despine(left=True)
# ## Tamanio y aspecto
# Se puede utilizar la funcion de matplotlib **plt.figure(figsize=(width,height) ** para cambiar el tamano de la mayoria de las grafiacas de seaborn.
#
# Para controlar el tamanio y el aspecto solo es necesario pasar los parametros: size y aspect, por ejemplo:
# Grafica sin grid
plt.figure(figsize=(12,3))
sns.countplot(x='sex',data=tips)
# Grafica con Grid
sns.lmplot(x='total_bill',y='tip',size=2,aspect=4,data=tips)
# ## Escala y contexto
#
# La funcion **set_context()** permite reescribir los parametros base:
sns.set_context('poster',font_scale=4)
sns.countplot(x='sex',data=tips,palette='coolwarm')
# Para mayor informacion puedes consultar la informacion que se encuentra disponible en linea en:
# https://stanford.edu/~mwaskom/software/seaborn/tutorial/aesthetics.html
| ejercicios/5_Seaborn/6_Color_Estilo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
# %matplotlib inline
# %config IPython.matplotlib.backend = "retina"
import matplotlib.pyplot as plt
import numpy as np
import sys
import astropy.units as u
from astropy.io import ascii, fits
from astropy.modeling.blackbody import blackbody_lambda
sys.path.insert(0, '../')
from libra import Spot, Star
# + deletable=true editable=true
times = np.arange(0, 3.3, 0.1)
contrasts = [0.1, 0.5]
spots = [Spot.from_latlon(0, 330, 0.02, contrast=10),
Spot.from_latlon(0, 50, 0.008, contrast=10),
Spot.from_latlon(0, 210, 0.008, contrast=10)]
star = Star(rotation_period=3.3*u.day, spots=spots)
fluxes1 = star.flux(times)
fluxes1 /= np.median(fluxes1)
spots = [Spot.from_latlon(0, 330, 0.02, contrast=2),
Spot.from_latlon(0, 50, 0.008, contrast=2),
Spot.from_latlon(0, 210, 0.008, contrast=2)]
star = Star(rotation_period=3.3*u.day, spots=spots)
fluxes2 = star.flux(times)
fluxes2 /= np.median(fluxes2)
plt.plot(times, fluxes1)
plt.plot(times, fluxes2)
#plt.plot(times, (fluxes1 - np.median(fluxes1)) * 0.5/0.1)
# + deletable=true editable=true
| notebooks/sanity_check_amps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/antnewman/nlp-textgenerator-gpt2-notebook/blob/main/Text_Generator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="IHBWIhFWqHBg"
# # Text Generator
# + id="uCJkCmadqEed"
# %tensorflow_version 1.x
# !pip install -q gpt-2-simple
import gpt_2_simple as gpt2
# + [markdown] id="ic5dQ-IEr0CD"
# There are three released sizes of GPT-2:
# - 124M (default): the "small" model, 500MB on disk.
# - 355M: the "medium" model, 1.5GB on disk.
# - 774M: the "large" model
# - 1558M: the "extra large"
#
# The large model cannot currently be finetuned with Colaboratory but can be used to generate text from a pretrained model.
#
# The extra large is the true model. It will not work if a K80/P4 GPU is attached to the notebook and as with the large model, it cannot be finetuned.
#
# The larger the model, the greater the knowledge but the longer the processing time.
#
# Use *model_name* to cahnge the base model.
# + id="Qk2nMu8_qwFD"
# Check GPU
# !nvidia-smi
# + id="zkyLjeNFswC7"
gpt2.download_gpt2(model_name="124M")
# + id="DePAZN6uXuvt"
from google.colab import drive
drive.mount('/content/drive')
# + id="mHzP1O7PXDhV"
txt = '/content/drive/MyDrive/data/Text/Down To The Sunless Sea.txt'
# + id="mmO6f-8JX5NP"
sess = gpt2.start_tf_sess()
gpt2.finetune(sess,
dataset=txt,
model_name='124M',
steps=1000,
restore_from='fresh',
run_name='run1',
print_every=10,
sample_every=200,
save_every=500
)
# + id="fnkJ7UVWYKsx"
gpt2.copy_checkpoint_to_gdrive(run_name='run1')
gpt2.copy_checkpoint_from_gdrive(run_name='run1')
# + id="uOFtLLfXeDI6"
# sess = gpt2.start_tf_sess()
# gpt2.load_gpt2(sess, run_name='run1')
# + [markdown] id="CLbNFUjBeOkw"
# After training the model or loading a retrained model from checkpoint, we can generate text. *generate* generates a single text from the loaded model.
# + id="eO0Xn7cLeEOa"
gpt2.generate(sess, run_name='run1')
| Text_Generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimization in Python
#
# You might have noticed that we didn't do anything related to sparsity with scikit-learn models. A lot of the work we covered in the machine learning class is very recent research, and as such is typically not implemented by the popular libraries.
#
# If we want to do things like sparse regression, we're going to have to roll up our sleeves and do it ourselves. For that, we need to be able to solve optimization problems. In Julia, we did this with JuMP. In Python, we'll use a similar library called *pyomo*.
# # Installing pyomo
# You can run the following command to install pyomo if you haven't already.
# !pip install pyomo --user
# # Intro to pyomo
#
# Let's see how we translate a simple, 2 variable LP to pyomo code.
#
# $$
# \begin{align*}
# \max_{x,y} \quad& x + 2y \\
# \text{s.t.}\quad& x + y \leq 1 \\
# & x, y \geq 0.
# \end{align*}
# $$
# First thing is to import the pyomo functions:
from pyomo.environ import *
from pyomo.opt import SolverFactory
# Next, we construct a model object. This is a container for everything in our optimization problem: variables, constraints, solver options, etc.
m = ConcreteModel()
# Next, we define the two decision variables in our optimization problem. We use the ``Var`` function to create the variables. The `within` keyword is used to specify the bounds on the variables, or equivalently the `bounds` keyword. The variables are added to the model object with names `x` and `y`.
m.x = Var(within=NonNegativeReals)
m.y = Var(bounds=(0, float('inf')))
# We now add the single constraint of our problem using the ``Constraint`` function. We write it algebraically, and save the result to the model.
m.con = Constraint(expr=m.x + m.y <= 1)
# We specify the objective function with the `Objective` function:
m.obj = Objective(sense=maximize, expr=m.x + 2 * m.y)
# We solve the optimization problem by first specifying a solver using `SolverFactory` and then using this solver to solve the model:
solver = SolverFactory('gurobi')
solver.solve(m)
# We can now inspect the solution values and optimal cost.
m.obj()
m.x.value
m.y.value
# Let's put it all together to compare with Julia/JuMP
# Create model
m = ConcreteModel()
# Add variables
m.x = Var(within=NonNegativeReals)
m.y = Var(bounds=(0, float('inf')))
# Add constraint
m.con = Constraint(expr=m.x + m.y <= 1)
# Add objective
m.obj = Objective(sense=maximize, expr=m.x + 2 * m.y)
# Solve model
solver = SolverFactory('gurobi')
solver.solve(m)
# Inspect solution
print(m.obj())
print(m.x.value)
print(m.y.value)
# ```julia
# # Create model
# m = Model(solver=GurobiSolver())
# # Add variables
# @variable(m, x >= 0)
# @variable(m, y >= 0)
# # Add constraint
# @constraint(m, x + y <= 1)
# # Add objective
# @objective(m, Max, x + 2y)
# # Solve model
# solve(m)
# # Inspect solution
# @show getobjectivevalue(m)
# @show getvalue(x)
# @show getvalue(y)
# ```
# ### Exercise
#
# Code and solve the following optimization problem:
#
# $$
# \begin{align*}
# \min_{x,y} \quad& 3x - y \\
# \text{s.t.}\quad& x + 2y \geq 1 \\
# & x \geq 0 \\
# & 0 \leq y \leq 1.
# \end{align*}
# $$
# +
# Create the model
m = ConcreteModel()
# Add the variables
m.x = Var(within=NonNegativeReals)
m.y = Var(bounds=(0, 1))
# Add the constraint
m.con = Constraint(expr=m.x + 2 * m.y >= 1)
# Add the objective
m.obj = Objective(sense=minimize, expr=3 * m.x - m.y)
solver = SolverFactory('gurobi')
solver.solve(m)
print(m.x.value, m.y.value)
# -
for v in m.component_data_objects(Var, active=True):
print(v, value(v)) # doctest: +SKIP
m.pprint()
# # Index sets
#
# Let's now move to a more complicated problem. We'll look at a transportation problem:
#
# $$
# \begin{align}
# \min & \sum\limits_{i = 1}^{m} \sum\limits_{j = 1}^{n} c_{ij} x_{ij}\\
# & \sum\limits_{j = 1}^{n} x_{ij} \leq b_i && i = 1, \ldots, m\\
# & \sum\limits_{i = 1}^{m} x_{ij} = d_j && j = 1, \ldots, n\\
# & x_{ij} \ge 0 && i = 1, \ldots, m, j = 1, \ldots, n
# \end{align}
# $$
#
# And with some data:
# +
import numpy as np
m = 2 # Number of supply nodes
n = 5 # Number of demand nodes
# Supplies
b = np.array([1000, 4000])
# Demands
d = np.array([500, 900, 1800, 200, 700])
# Costs
c = np.array([[2, 4, 5, 2, 1],
[3, 1, 3, 2, 3]])
# -
# Now we can formulate the problem with pyomo
model = ConcreteModel()
# First step is adding variables. We can add variables with indices by passing the relevant index sets to the `Var` constructor. In this case, we need a $m$-by$n$ matrix of variables:
model.x = Var(range(m), range(n), within=NonNegativeReals)
# Now to add the constraints. We have to add one supply constraint for each factory, so we might try something like:
for i in range(m):
model.supply = Constraint(expr=sum(model.x[i, j] for j in range(n)) <= b[i])
# Can you see the problem? We are overwriting `model.supply` in each iteration of the loop, and so only the last constraint is applied.
#
# Luckily, pyomo has a (not-so-easy) way to add multiple constraints at a time. We first define a *rule* that takes in the model and any required indices, and then returns the expression for the constraint:
def supply_rule(model, i):
return sum(model.x[i, j] for j in range(n)) <= b[i]
# We then add the constraint by referencing this rule along with the index set we want the constraint to be defined over:
model.supply2 = Constraint(range(m), rule=supply_rule)
# We then apply the same approach for the demand constraints
# +
def demand_rule(model, j):
return sum(model.x[i, j] for i in range(m)) == d[j]
model.demand = Constraint(range(n), rule=demand_rule)
# -
# Finally, we add the objective:
model.obj = Objective(sense=minimize,
expr=sum(c[i, j] * model.x[i, j]
for i in range(m) for j in range(n)))
# Now we can solve the problem
solver = SolverFactory('gurobi')
solver.solve(model)
# It solved, so we can extract the results
flows = np.array([[model.x[i, j].value for j in range(n)] for i in range(m)])
flows
# We can also check the objective value for the cost of this flow
model.obj()
# For simplicity, here is the entire formulation and solving code together:
model = ConcreteModel()
# Variables
model.x = Var(range(m), range(n), within=NonNegativeReals)
# Supply constraint
def supply_rule(model, i):
return sum(model.x[i, j] for j in range(n)) <= b[i]
model.supply2 = Constraint(range(m), rule=supply_rule)
# Demand constraint
def demand_rule(model, j):
return sum(model.x[i, j] for i in range(m)) == d[j]
model.demand = Constraint(range(n), rule=demand_rule)
# Objective
model.obj = Objective(sense=minimize,
expr=sum(c[i, j] * model.x[i, j]
for i in range(m) for j in range(n)))
# Solve
solver = SolverFactory('gurobi')
solver.solve(model)
# Get results
flows = np.array([[model.x[i, j].value for j in range(n)] for i in range(m)])
model.obj()
# # Machine Learning
#
# Now let's put our pyomo knowledge to use and implement some of the same methods we saw in the machine learning class
# First, specify your solver executable location:
executable='C:/Users/omars/.julia/v0.6/Ipopt/deps/usr/bin/ipopt.exe'
# To use the version left over from Julia
# ### On MacOS and Linux
#
# `executable="~/.julia/v0.6/Homebrew/deps/usr/Cellar/ipopt/3.12.4_1/bin/ipopt")`
#
# ### On Windows
#
# The path is probably under WinRPM:
#
# `executable='%HOME%/.julia/v0.6/WinRPM/...')")`
#
# # Linear Regression
#
# Let's just try a simple linear regression
def linear_regression(X, y):
n, p = X.shape
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
# Add constraints
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)), 2)
for i in range(n)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return [m.beta[j].value for j in range(p)]
# Let's load up some data to test it out on:
from sklearn.datasets import load_boston
data = load_boston()
X = data.data
y = data.target
# Try our linear regression function:
print(linear_regression(X, y))
# We can compare with sklearn to make sure it's right:
from sklearn.linear_model import LinearRegression
m = LinearRegression(fit_intercept=False)
m.fit(X, y)
m.coef_
# Just for reference, let's look back at how we do the same thing in JuMP!
#
# ```julia
# using JuMP, Gurobi
# function linear_regression(X, y)
# n, p = size(X)
# m = Model(solver=GurobiSolver())
# @variable(m, beta[1:p])
# @objective(m, Min, sum((y[i] - sum(X[i, j] * beta[j] for j = 1:p)) ^ 2 for i = 1:n))
# solve(m)
# getvalue(beta)
# end
# ```
#
# or even
#
# ```julia
# using JuMP, Gurobi
# function linear_regression(X, y)
# n, p = size(X)
# m = Model(solver=GurobiSolver())
# @variable(m, beta[1:p])
# @objective(m, Min, sum((y - X * beta) .^ 2))
# solve(m)
# getvalue(beta)
# end
# ```
#
# Much simpler!
# ### Exercise
#
# Modify the linear regression formulation to include an intercept term, and compare to scikit-learn's LinearRegression with `fit_intercept=False` to make sure it's the same
# +
def linear_regression_intercept(X, y):
n, p = X.shape
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
m.b0 = Var()
# Add constraints
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)) - m.b0, 2)
for i in range(n)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return [m.beta[j].value for j in range(p)]
linear_regression_intercept(X, y)
# -
m = LinearRegression(fit_intercept=True)
m.fit(X, y)
m.coef_
# # Robust Regression
#
# We saw in the class that both ridge and lasso regression were robust versions of linear regression. Both of these are provided by `sklearn`, but we need to know how to implement them if we want to extend regression ourselves
def ridge_regression(X, y, rho):
n, p = X.shape
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)),2)
for i in range(n)) + rho * sum(pow(m.beta[j], 2) for j in range(p)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return [m.beta[j].value for j in range(p)]
ridge_regression(X, y, 100000)
def lasso(X, y, rho):
n, p = X.shape
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)),2)
for i in range(n)) + rho * sum(pow(m.beta[j], 2) for j in range(p)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return [m.beta[j].value for j in range(p)]
# ### Exercise
#
# Implement Lasso regression
def lasso_regression(X, y, rho):
n, p = X.shape
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
m.absb = Var(range(p))
# Add constraints
def absbeta1(m, j):
return m.beta[j] <= m.absb[j]
m.absb1 = Constraint(range(p), rule=absbeta1)
def absbeta2(m, j):
return -m.beta[j] <= m.absb[j]
m.absb2 = Constraint(range(p), rule=absbeta2)
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)), 2)
for i in range(n)) + rho * sum(m.absb[j] for j in range(p)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return [m.beta[j].value for j in range(p)]
lasso_regression(X, y, 1000)
# # Sparse Regression
def sparse_regression(X, y, k):
n, p = X.shape
M = 1000
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
m.z = Var(range(p), within=Binary)
# Add constraints
def bigm1(m, j):
return m.beta[j] <= M * m.z[j]
m.bigm1 = Constraint(range(p), rule=bigm1)
def bigm2(m, j):
return m.beta[j] >= -M * m.z[j]
m.bigm2 = Constraint(range(p), rule=bigm2)
m.sparsity = Constraint(expr=sum(m.z[j] for j in range(p)) <= k)
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)), 2)
for i in range(n)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return [m.beta[j].value for j in range(p)]
sparse_regression(X, y, 10)
# +
import numpy as np
l = np.array([1,2,3,4])
print(l**2)
print([sqrt(i) for i in l])
# -
# ### Exercise
#
# Try implementing the algorithmic framework for linear regression:
# - sparsity constraints
# - lasso regularization
# - restrict highly correlated pairs of features
# - nonlinear transformations (just $\sqrt(x)$ and $x^2$)
# +
import numpy as np
from sklearn.preprocessing import normalize
def all_regression(X_orig, y, k, rho):
n, p_orig = X_orig.shape
M = 10
X = np.concatenate(
[X_orig, np.sqrt(X_orig), np.square(X_orig)], axis=1
)
p = X.shape[1]
# Normalize data
X = normalize(X, axis=0)
y = (y - np.mean(y)) / np.linalg.norm(y)
# Create model
m = ConcreteModel()
# Add variables
m.beta = Var(range(p))
m.z = Var(range(p), within=Binary)
m.absb = Var(range(p))
# Sparsity constraints
def bigm1(m, j):
return m.beta[j] <= M * m.z[j]
m.bigm1 = Constraint(range(p), rule=bigm1)
def bigm2(m, j):
return m.beta[j] >= -M * m.z[j]
m.bigm2 = Constraint(range(p), rule=bigm2)
m.sparsity = Constraint(expr=sum(m.z[j] for j in range(p)) <= k)
# Lasso constraints
def absbeta1(m, j):
return m.beta[j] <= m.absb[j]
m.absb1 = Constraint(range(p), rule=absbeta1)
def absbeta2(m, j):
return -m.beta[j] <= m.absb[j]
m.absb2 = Constraint(range(p), rule=absbeta2)
# Correlation constraints
corX = np.corrcoef(np.transpose(X))
def cor_rule(m, i, j):
if i > j and abs(corX[i, j]) > 0.8:
return (sum(m.z[k] for k in range(i, p, p_orig)) +
sum(m.z[k] for k in range(j, p, p_orig)) <= 1)
else:
return Constraint.Skip
m.cor = Constraint(range(p_orig), range(p_orig), rule=cor_rule)
# Nonlinear constraints
def nl_rule(m, i):
return sum(m.z[k] for k in range(i, p, p_orig)) <= 1
m.nl = Constraint(range(p_orig), rule=nl_rule)
# Add objective
m.obj = Objective(sense=minimize, expr=sum(
pow(y[i] - sum(X[i, j] * m.beta[j] for j in range(p)), 2)
for i in range(n)) + rho * sum(m.absb[j] for j in range(p)))
solver = SolverFactory('ipopt', executable=executable)
## tee=True enables solver output
# results = solver.solve(m, tee=True)
results = solver.solve(m, tee=False)
return np.array([m.beta[j].value for j in range(p)]).reshape(-1, p_orig)
# -
all_regression(X, y, 6, 0)
# # Logistic Regression
# Like JuMP, we need to use a new solver for the nonlinear problem. We can use Ipopt as before, except we have to set it up manually. You'll need to download Ipopt and add it to the PATH.
#
# On Mac, you can do this with Homebrew if you have it:
# The other way is to download a copy of ipopt and specify the path to it exactly when creating the solver. For example, I have a copy of Ipopt left over from JuMP, which I can use by modifying the SolverFactory line as indicated below:
def logistic_regression(X, y):
n, p = X.shape
# Convert y to (-1, +1)
assert np.min(y) == 0
assert np.max(y) == 1
Y = y * 2 - 1
assert np.min(Y) == -1
assert np.max(Y) == 1
# Create the model
m = ConcreteModel()
# Add variables
m.b = Var(range(p))
m.b0 = Var()
# Set nonlinear objective function
m.obj = Objective(sense=maximize, expr=-sum(
log(1 + exp(-Y[i] * (sum(X[i, j] * m.b[j] for j in range(p)) + m.b0)))
for i in range(n)))
# Solve the model and get the optimal solutions
solver = SolverFactory('ipopt', executable=executable)
solver.solve(m)
return [m.b[j].value for j in range(p)], m.b0.value
# Load up some data
from sklearn.datasets import load_breast_cancer
data = load_breast_cancer()
X = data.data
y = data.target
logistic_regression(X, y)
# ### Exercise
#
# Implement the regularized versions of logistic regression that scikit-learn provides:
#
# 
#
# 
def logistic_regression_l1(X, y, C):
n, p = X.shape
# Convert y to (-1, +1)
assert np.min(y) == 0
assert np.max(y) == 1
Y = y * 2 - 1
assert np.min(Y) == -1
assert np.max(Y) == 1
# Create the model
m = ConcreteModel()
# Add variables
m.b = Var(range(p))
m.b0 = Var()
# Lasso constraints
m.absb = Var(range(p))
def absbeta1(m, j):
return m.b[j] <= m.absb[j]
m.absb1 = Constraint(range(p), rule=absbeta1)
def absbeta2(m, j):
return -m.b[j] <= m.absb[j]
m.absb2 = Constraint(range(p), rule=absbeta2)
# Set nonlinear objective function
m.obj = Objective(sense=minimize, expr=sum(m.absb[j] for j in range(p)) + C * sum(
log(1 + exp(-Y[i] * (sum(X[i, j] * m.b[j] for j in range(p)) + m.b0)))
for i in range(n)))
# Solve the model and get the optimal solutions
solver = SolverFactory('ipopt', executable=executable)
solver.solve(m)
return [m.b[j].value for j in range(p)], m.b0.value
logistic_regression_l1(X, y, 100)
def logistic_regression_l2(X, y, C):
n, p = X.shape
# Convert y to (-1, +1)
assert np.min(y) == 0
assert np.max(y) == 1
Y = y * 2 - 1
assert np.min(Y) == -1
assert np.max(Y) == 1
# Create the model
m = ConcreteModel()
# Add variables
m.b = Var(range(p))
m.b0 = Var()
# Set nonlinear objective function
m.obj = Objective(sense=minimize, expr=0.5 * sum(pow(m.b[j], 2) for j in range(p)) + C * sum(
log(1 + exp(-Y[i] * (sum(X[i, j] * m.b[j] for j in range(p)) + m.b0)))
for i in range(n)))
# Solve the model and get the optimal solutions
solver = SolverFactory('ipopt', executable=executable)
solver.solve(m)
return [m.b[j].value for j in range(p)], m.b0.value
logistic_regression_l2(X, y, 1000)
| ML3 - Optimization Modeling (Complete).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"is_executing": false, "name": "#%% md\n"}
# ## Introduction
# The transformers library is an open-source, community-based repository to train, use and share models based on
# the Transformer architecture [(Vaswani & al., 2017)](https://arxiv.org/abs/1706.03762) such as Bert [(Devlin & al., 2018)](https://arxiv.org/abs/1810.04805),
# Roberta [(Liu & al., 2019)](https://arxiv.org/abs/1907.11692), GPT2 [(Radford & al., 2019)](https://cdn.openai.com/better-language-models/language_models_are_unsupervised_multitask_learners.pdf),
# XLNet [(Yang & al., 2019)](https://arxiv.org/abs/1906.08237), etc.
#
# Along with the models, the library contains multiple variations of each of them for a large variety of
# downstream-tasks like **Named Entity Recognition (NER)**, **Sentiment Analysis**,
# **Language Modeling**, **Question Answering** and so on.
#
# ## Before Transformer
#
# Back to 2017, most of the people using Neural Networks when working on Natural Language Processing were relying on
# sequential processing of the input through [Recurrent Neural Network (RNN)](https://en.wikipedia.org/wiki/Recurrent_neural_network).
#
# 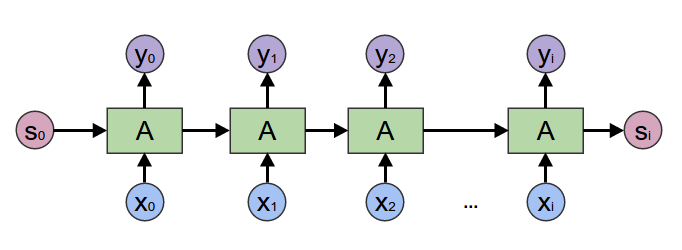
#
# RNNs were performing well on large variety of tasks involving sequential dependency over the input sequence.
# However, this sequentially-dependent process had issues modeling very long range dependencies and
# was not well suited for the kind of hardware we're currently leveraging due to bad parallelization capabilities.
#
# Some extensions were provided by the academic community, such as Bidirectional RNN ([Schuster & Paliwal., 1997](https://www.researchgate.net/publication/3316656_Bidirectional_recurrent_neural_networks), [Graves & al., 2005](https://mediatum.ub.tum.de/doc/1290195/file.pdf)),
# which can be seen as a concatenation of two sequential process, one going forward, the other one going backward over the sequence input.
#
# 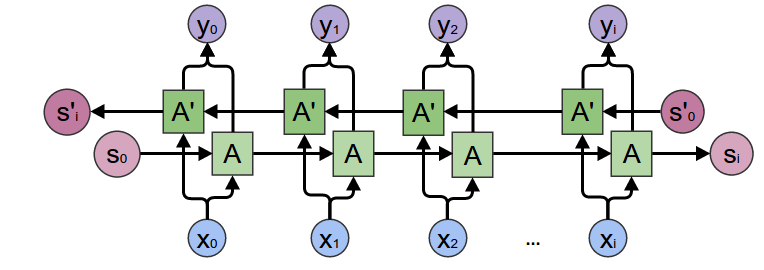
#
#
# And also, the Attention mechanism, which introduced a good improvement over "raw" RNNs by giving
# a learned, weighted-importance to each element in the sequence, allowing the model to focus on important elements.
#
# 
#
# ## Then comes the Transformer
#
# The Transformers era originally started from the work of [(Vaswani & al., 2017)](https://arxiv.org/abs/1706.03762) who
# demonstrated its superiority over [Recurrent Neural Network (RNN)](https://en.wikipedia.org/wiki/Recurrent_neural_network)
# on translation tasks but it quickly extended to almost all the tasks RNNs were State-of-the-Art at that time.
#
# One advantage of Transformer over its RNN counterpart was its non sequential attention model. Remember, the RNNs had to
# iterate over each element of the input sequence one-by-one and carry an "updatable-state" between each hop. With Transformer, the model is able to look at every position in the sequence, at the same time, in one operation.
#
# For a deep-dive into the Transformer architecture, [The Annotated Transformer](https://nlp.seas.harvard.edu/2018/04/03/attention.html#encoder-and-decoder-stacks)
# will drive you along all the details of the paper.
#
# 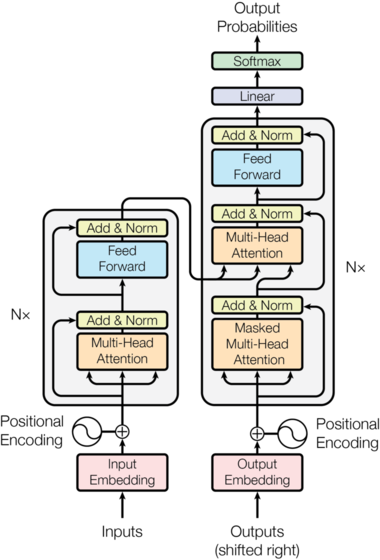
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Getting started with transformers
#
# For the rest of this notebook, we will use the [BERT (Devlin & al., 2018)](https://arxiv.org/abs/1810.04805) architecture, as it's the most simple and there are plenty of content about it
# over the internet, it will be easy to dig more over this architecture if you want to.
#
# The transformers library allows you to benefits from large, pretrained language models without requiring a huge and costly computational
# infrastructure. Most of the State-of-the-Art models are provided directly by their author and made available in the library
# in PyTorch and TensorFlow in a transparent and interchangeable way.
# + pycharm={"is_executing": false, "name": "#%% code\n"}
# !pip install transformers
# !pip install tensorflow==2.1.0
# + pycharm={"is_executing": false, "name": "#%% code\n"}
import torch
from transformers import AutoModel, AutoTokenizer, BertTokenizer
torch.set_grad_enabled(False)
# + pycharm={"is_executing": false, "name": "#%% code\n"}
# Store the model we want to use
MODEL_NAME = "bert-base-cased"
# We need to create the model and tokenizer
model = AutoModel.from_pretrained(MODEL_NAME)
tokenizer = AutoTokenizer.from_pretrained(MODEL_NAME)
# + [markdown] pycharm={"name": "#%% md\n"}
# With only the above two lines of code, you're ready to use a BERT pre-trained model.
# The tokenizers will allow us to map a raw textual input to a sequence of integers representing our textual input
# in a way the model can manipulate.
# + pycharm={"is_executing": false, "name": "#%% code\n"}
# Tokens comes from a process that splits the input into sub-entities with interesting linguistic properties.
tokens = tokenizer.tokenize("This is an input example")
print("Tokens: {}".format(tokens))
# This is not sufficient for the model, as it requires integers as input,
# not a problem, let's convert tokens to ids.
tokens_ids = tokenizer.convert_tokens_to_ids(tokens)
print("Tokens id: {}".format(tokens_ids))
# Add the required special tokens
tokens_ids = tokenizer.build_inputs_with_special_tokens(tokens_ids)
# We need to convert to a Deep Learning framework specific format, let's use PyTorch for now.
tokens_pt = torch.tensor([tokens_ids])
print("Tokens PyTorch: {}".format(tokens_pt))
# Now we're ready to go through BERT with out input
outputs, pooled = model(tokens_pt)
print("Token wise output: {}, Pooled output: {}".format(outputs.shape, pooled.shape))
# + [markdown] pycharm={"name": "#%% md\n"}
# As you can see, BERT outputs two tensors:
# - One with the generated representation for every token in the input `(1, NB_TOKENS, REPRESENTATION_SIZE)`
# - One with an aggregated representation for the whole input `(1, REPRESENTATION_SIZE)`
#
# The first, token-based, representation can be leveraged if your task requires to keep the sequence representation and you
# want to operate at a token-level. This is particularly useful for Named Entity Recognition and Question-Answering.
#
# The second, aggregated, representation is especially useful if you need to extract the overall context of the sequence and don't
# require a fine-grained token-level. This is the case for Sentiment-Analysis of the sequence or Information Retrieval.
# + [markdown] pycharm={"name": "#%% md\n"}
# The code you saw in the previous section introduced all the steps required to do simple model invocation.
# For more day-to-day usage, transformers provides you higher-level methods which will makes your NLP journey easier.
# Let's improve our previous example
# + pycharm={"is_executing": false, "name": "#%% code\n"}
# tokens = tokenizer.tokenize("This is an input example")
# tokens_ids = tokenizer.convert_tokens_to_ids(tokens)
# tokens_pt = torch.tensor([tokens_ids])
# This code can be factored into one-line as follow
tokens_pt2 = tokenizer.encode_plus("This is an input example", return_tensors="pt")
for key, value in tokens_pt2.items():
print("{}:\n\t{}".format(key, value))
outputs2, pooled2 = model(**tokens_pt2)
print("Difference with previous code: ({}, {})".format((outputs2 - outputs).sum(), (pooled2 - pooled).sum()))
# -
# As you can see above, the method `encode_plus` provides a convenient way to generate all the required parameters
# that will go through the model.
#
# Moreover, you might have noticed it generated some additional tensors:
#
# - token_type_ids: This tensor will map every tokens to their corresponding segment (see below).
# - attention_mask: This tensor is used to "mask" padded values in a batch of sequence with different lengths (see below).
# + pycharm={"is_executing": false}
# Single segment input
single_seg_input = tokenizer.encode_plus("This is a sample input")
# Multiple segment input
multi_seg_input = tokenizer.encode_plus("This is segment A", "This is segment B")
print("Single segment token (str): {}".format(tokenizer.convert_ids_to_tokens(single_seg_input['input_ids'])))
print("Single segment token (int): {}".format(single_seg_input['input_ids']))
print("Single segment type : {}".format(single_seg_input['token_type_ids']))
# Segments are concatened in the input to the model, with
print()
print("Multi segment token (str): {}".format(tokenizer.convert_ids_to_tokens(multi_seg_input['input_ids'])))
print("Multi segment token (int): {}".format(multi_seg_input['input_ids']))
print("Multi segment type : {}".format(multi_seg_input['token_type_ids']))
# + pycharm={"is_executing": false}
# Padding highlight
tokens = tokenizer.batch_encode_plus(
["This is a sample", "This is another longer sample text"],
pad_to_max_length=True # First sentence will have some PADDED tokens to match second sequence length
)
for i in range(2):
print("Tokens (int) : {}".format(tokens['input_ids'][i]))
print("Tokens (str) : {}".format([tokenizer.convert_ids_to_tokens(s) for s in tokens['input_ids'][i]]))
print("Tokens (attn_mask): {}".format(tokens['attention_mask'][i]))
print()
# -
# ## Frameworks interoperability
#
# One of the most powerfull feature of transformers is its ability to seamlessly move from PyTorch to Tensorflow
# without pain for the user.
#
# We provide some convenient methods to load TensorFlow pretrained weight insinde a PyTorch model and opposite.
# + pycharm={"is_executing": false}
from transformers import TFBertModel, BertModel
# Let's load a BERT model for TensorFlow and PyTorch
model_tf = TFBertModel.from_pretrained('bert-base-cased')
model_pt = BertModel.from_pretrained('bert-base-cased')
# + pycharm={"is_executing": false}
# transformers generates a ready to use dictionary with all the required parameters for the specific framework.
input_tf = tokenizer.encode_plus("This is a sample input", return_tensors="tf")
input_pt = tokenizer.encode_plus("This is a sample input", return_tensors="pt")
# Let's compare the outputs
output_tf, output_pt = model_tf(input_tf), model_pt(**input_pt)
# Models outputs 2 values (The value for each tokens, the pooled representation of the input sentence)
# Here we compare the output differences between PyTorch and TensorFlow.
for name, o_tf, o_pt in zip(["output", "pooled"], output_tf, output_pt):
print("{} differences: {:.5}".format(name, (o_tf.numpy() - o_pt.numpy()).sum()))
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Want it lighter? Faster? Let's talk distillation!
#
# One of the main concerns when using these Transformer based models is the computational power they require. All over this notebook we are using BERT model as it can be run on common machines but that's not the case for all of the models.
#
# For example, Google released a few months ago **T5** an Encoder/Decoder architecture based on Transformer and available in `transformers` with no more than 11 billions parameters. Microsoft also recently entered the game with **Turing-NLG** using 17 billions parameters. This kind of model requires tens of gigabytes to store the weights and a tremendous compute infrastructure to run such models which makes it impracticable for the common man !
#
# 
#
# With the goal of making Transformer-based NLP accessible to everyone we @huggingface developed models that take advantage of a training process called **Distillation** which allows us to drastically reduce the resources needed to run such models with almost zero drop in performances.
#
# Going over the whole Distillation process is out of the scope of this notebook, but if you want more information on the subject you may refer to [this Medium article written by my colleague <NAME>, author of DistilBERT paper](https://medium.com/huggingface/distilbert-8cf3380435b5), you might also want to directly have a look at the paper [(Sanh & al., 2019)](https://arxiv.org/abs/1910.01108)
#
# Of course, in `transformers` we have distilled some models and made them available directly in the library !
# + pycharm={"is_executing": false}
from transformers import DistilBertModel
bert_distil = DistilBertModel.from_pretrained('distilbert-base-cased')
input_pt = tokenizer.encode_plus(
'This is a sample input to demonstrate performance of distiled models especially inference time',
return_tensors="pt"
)
# %time _ = bert_distil(input_pt['input_ids'])
# %time _ = model_pt(input_pt['input_ids'])
# -
# ## Community provided models
#
# Last but not least, earlier in this notebook we introduced Hugging Face `transformers` as a repository for the NLP community to exchange pretrained models. We wanted to highlight this features and all the possibilities it offers for the end-user.
#
# To leverage community pretrained models, just provide the organisation name and name of the model to `from_pretrained` and it will do all the magic for you !
#
#
# We currently have more 50 models provided by the community and more are added every day, don't hesitate to give it a try !
# + pycharm={"is_executing": false}
# Let's load German BERT from the Bavarian State Library
de_bert = BertModel.from_pretrained("dbmdz/bert-base-german-cased")
de_tokenizer = BertTokenizer.from_pretrained("dbmdz/bert-base-german-cased")
de_input = de_tokenizer.encode_plus(
"Hugging Face ist eine französische Firma mit Sitz in New-York.",
return_tensors="pt"
)
print("Tokens (int) : {}".format(de_input['input_ids'].tolist()[0]))
print("Tokens (str) : {}".format([de_tokenizer.convert_ids_to_tokens(s) for s in de_input['input_ids'].tolist()[0]]))
print("Tokens (attn_mask): {}".format(de_input['attention_mask'].tolist()[0]))
print()
output_de, pooled_de = de_bert(**de_input)
print("Token wise output: {}, Pooled output: {}".format(outputs.shape, pooled.shape))
| notebooks/02-transformers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Injured in Madrid due to traffic accidents during 2019</h1>
# <h4>by <NAME></h4>
#
#
# ### Table of Contents
#
# 1. [Business Understanding](#Business)
# 2. [Data Understanding](#Data)
# 3. [Data Preparation](#Preparation)
# 4. [Modeling](#Modeling)
# 5. [Evaluation](#Evaluation)
#
# ## Business Understanding <a name="Business"></a>
#
# **Study on traffic accidents in Madrid**
#
# For this project, I was interestested in using El Ayuntamiento de Madrid data from 2019 about traffic accidents to better understand:
#
# 1. How many traffic accident occurred in Madrid during 2019?,
# How many people are involved? and
# What is the average number of people involved in a traffic accident?
#
# 2. What day of the week are there the most accidents?
#
# 3. What time do most accidents occur?
#
# 4. At what age are more traffic accidents suffered?
# ## Data Understanding <a name="Data"></a>
#
# The dataset can be find in:
# https://datos.madrid.es/portal/site/egob/menuitem.c05c1f754a33a9fbe4b2e4b284f1a5a0/?vgnextoid=7c2843010d9c3610VgnVCM2000001f4a900aRCRD&vgnextchannel=374512b9ace9f310VgnVCM100000171f5a0aRCRD&vgnextfmt=default
# First the code to import the necesary **libraries** and the **dataset**
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
df = pd.read_csv('2019_Accidentalidad.csv', sep=';' , encoding='latin-1')
df.head()
# -
# ### Rename the columns seems a good idea in orther to create a dataset understable in English languaje
df.rename(columns={'N_EXPEDIENTE': 'exp','FECHA': 'date'
,'CALLE': 'street' ,'NUMERO': 'number'
,'DISTRITO': 'district' ,'TIPO VEHICULO': 'vehType'
,'ESTADO METEREOLOGICO': 'weather' ,'TIPO ACCIDENTE': 'accType'
,'TIPO PERSONA': 'person' ,'RANGO EDAD': 'age'
,'SEXO': 'sex' ,'LESIVIDAD': 'injuries' ,'HORA': 'time'}, inplace=True)
df.columns
# ## Data Preparation <a name="Preparation"></a>
# ### Let's create some aditional columns based on the date of the accident and the hour
df['date'] = pd.to_datetime(df['date'], format="%d/%m/%Y") #This gives datetime format to the 'date' variable
df['hour'] = pd.to_datetime(df['time']).dt.hour #This create a variable named 'hour' with the accident hour
df['year'] = pd.DatetimeIndex(df['date']).year #This create a variable named 'year' with the year of the accident
df['month'] = pd.DatetimeIndex(df['date']).month #This create a variable named 'month' with the month of the accident
df['day'] = pd.DatetimeIndex(df['date']).day #This create a variable named 'day' with the day of the accident
df['day_of_week'] = df['date'].dt.weekday_name #This create a variable named 'day_of_week' with the day of the week
df.head()
# ### Using dictionaries, rename the columns values in orther to create a dataset understable in English languaje
# +
#create de dictionaries
di = {'Hombre': "Man",
'Mujer': "Woman"}
di2 = {'Despejado': "Clear",
'Lluvia débil': "Rain",
'Nublado': "Cloudy",
'Se desconoce': "Other" ,
'LLuvia intensa': "HeavyRain",
'Granizando': "Hail",
'Nevando': "Snow",}
di3 = {'DE 0 A 5 AÑOS': "a.[0,5]"
,'DE 6 A 9 AÑOS': "b.[6,9]"
,'DE 10 A 14 AÑOS': "c.[10,14]"
,'DE 15 A 17 AÑOS': "d.[15,17]"
,'DE 18 A 20 AÑOS': "e.[18,20]"
,'DE 21 A 24 AÑOS': "f.[21,24]"
,'DE 25 A 29 AÑOS': "g.[25,29]"
,'DE 30 A 34 AÑOS': "h.[30,34]"
,'DE 35 A 39 AÑOS': "i.[35,39]"
,'DE 40 A 44 AÑOS': "j.[40,44]"
,'DE 45 A 49 AÑOS': "k.[45,49]"
,'DE 50 A 54 AÑOS': "l.[50,54]"
,'DE 55 A 59 AÑOS': "m.[55,59]"
,'DE 60 A 64 AÑOS': "n.[60,64]"
,'DE 65 A 69 AÑOS': "o.[65,69]"
,'DE 70 A 74 AÑOS': "p.[70,74]"
,'MAYOR DE 74 AÑOS': "q.[75,.)"
,'DESCONOCIDA': "w.[Unknown]"}
di4 = {'Conductor': "Driver",
'Pasajero': "Passenger",
'Peatón': "Pedestrian"}
# -
# There are many ways to deal with missing values.
#
# When I do an Exploratory Data Analisys.[here](https://en.wikipedia.org/wiki/Exploratory_data_analysis)
#
# I usually follow the next rules (which I find really useful in my daily profesional life):
# 1. Try to understand the variable before dealing with missing values.(how is it collected?, Expected values...)
# 2. Only delete a variable if it's clear that it gives no relevant information.
# 3. Only delete a row with a missing value if it gives no relevant infomation.
# 4. Fill NaN with another value of the variable only if its clear that the value chosen is the correct one.
# 5. If point 4 can not be applied, then fill NaN with a new value that allows us to now that it is a missing value.
#
# Why do I use this rules?
# 1. If the missing values are generated by a IT mistake, they will be easier to report and solve.
# 2. This way it is easier to find out if the missing values have an effect on other variables.
# +
def use_dict(df,vari, dicti, comple):
"""
Summary line.
This function has been created to deal with the variables that I want to modify using a dictionary
also to deal with the missing values filling them with a value choosen by me.
In this case I think that the best option is the fifth rule.
Parameters:
- df: the dataframe in which we want to apply the dictionary and to fill NaN
- vari: variable
- dicti: the dictionary we want to apply
- comple: the value chosen to fill NaN
Returns:
df where the chosen variable has replaced its values with the dictionary ones
also the missing values have been replaced by the chosen value 'comple'.
"""
df=df.replace({vari: dicti})
df[vari].fillna(comple, inplace=True)
return df
# variables data = [variable, dictionary, value for NaN]
subjects =[["sex", di,"Other"], # variable1
["weather",di2,"Other"], # variable2
["age", di3,"w.[Unknown]"], # variable3
["person", di4,"Other"]] # variable4
for sub in subjects:
df = use_dict(df,sub[0], sub[1], sub[2])
# -
df.head()
# ## Modeling <a name="Modeling"></a>
# Before dealing with the questions weshould keep in mind this dataset includes:
#
# - Traffic accidents in the City of Madrid registered by the Municipal Police in 2019.
# - One record is included per person involved in the accident.
# - Witness records not included.
# ### Q1.1. - How many people are involved in a traffic accident in Madrid during 2019?
# One record is included per person involved in the accident. To answer this quetion we have to know the number of rows in our dataset.
#number of people involved in a car accident ocurred during 2019
df.shape[0]
# ### Q1.2. - How many car accident occurred during 2019?
# One record is included per person involved in the accident.
# That means that one single accident could have more than one record.
# The variable exp is unique for every accident, so to answer this quetion we have to know the number of different exp in our dataset.
#number of accident ocurred during 2019
df1 = df.drop_duplicates(subset=['exp'])
df.drop_duplicates(subset=['exp']).shape[0]
# ### Q1.3. - What is the average number of people involved in a traffic accident?
# On average, at least two people are affected in each accident
#Average number of people affected
df.shape[0]/df1.shape[0]
# ### Q2 - What day of the week are there the most accidents?
# Taking into account the number of files:
#
# **Friday** is the day with the highest number of accidents
# +
day_order = ["Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"]
count_vals = df.day_of_week.value_counts()
(count_vals/df.shape[0]).loc[day_order].plot(kind="bar");
plt.title("day_of_week");
# -
# Taking into account the number of files created by the police:
#
# **Friday** is the day with the highest number of accidents
# +
day_order = ["Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"]
count_vals = df1.day_of_week.value_counts()
(count_vals/df1.shape[0]).loc[day_order].plot(kind="bar");
plt.title("day_of_week");
# -
# ### Q3 - What time do most accidents occur?
# Most claims occur **at 6 p.m.**, coinciding with time to **return** home from work or school.
#
# It is clear that the frequency increases at **8 and 9 a.m.** coinciding with the time to **go to work or school.**
# +
count_vals = df1.hour.value_counts().sort_index()
(count_vals/df1.shape[0]).plot(kind="bar");
plt.title("Hour");
# -
# #### Does the pattern change over the weekend?
# +
#code to create a dataframe with the week involved in accidents
#I will consider Friday as part of the weekend, because is when the party stars
df_week = df[df['day_of_week'].isin(["Monday", "Tuesday", "Wednesday", "Thursday"])]
# % of the total people involved in an accident by hour
dd=(df_week.hour.value_counts().sort_index()/df_week.shape[0]*100).rename_axis('hour').reset_index(name='perc_week')
dd.set_index('hour', inplace=True)
# -
#code to create a dataframe with the week involved in accidents
df_weekend = df[df['day_of_week'].isin(["Friday", "Saturday", "Sunday"])]
dd2=(df_weekend.hour.value_counts().sort_index()/df_weekend.shape[0]*100).rename_axis('hour').reset_index(name='perc_weekend')
dd2.set_index('hour', inplace=True)
# Clearly, accident hours change over the weekend.
#
# Accidents increase during party hours which is from 10 p.m. until 6 a.m.
comp_df = pd.merge(dd, dd2, left_index=True, right_index=True)
comp_df.plot.bar(rot=0)
# #### How much the pattern change over the weekend?
# The number of people affected by accidents drops 50% at 9 a.m.
#
# On the other hand, at 4 a.m. the number of people affected by accidents increases by 232%
comp_df = pd.merge(dd, dd2, left_index=True, right_index=True) # merge of the two database that I want to compare
comp_df.columns = ['perc_week', 'perc_weekend']
comp_df['Diff_perc_point'] = comp_df['perc_weekend'] / comp_df['perc_week']-1 #% of increase or decrease from week %
comp_df.style.bar(subset=['Diff_perc_point'], align='mid', color=['#d65f5f', '#5fba7d'])
# ### Q4 - At what age are more traffic accidents suffered?
# Continuing with the study we are going to separate the working hours from the party hours
df_day = df[df['hour'].isin([7,8,9,10,11,12,13,14,15,16,17,18,19,20])]
df_day = df_day[~df_day['age'].isin(['w.[Unknown]'])]
dd=(df_day.age.value_counts().sort_index()/df_day.shape[0]*100).rename_axis('age').reset_index(name='perc_day')
dd.set_index('age', inplace=True)
df_party = df[df['hour'].isin([21,22,23,0,1,2,3,4,5,6])]
df_party = df_party[~df_party['age'].isin(['w.[Unknown]'])]
dd2=(df_party.age.value_counts().sort_index()/df_party.shape[0]*100).rename_axis('age').reset_index(name='perc_party')
dd2.set_index('age', inplace=True)
# During the working hours, people between **40 and 44** years old are the ones who suffer the most traffic accidents
#
# During party hours, people between the ages of **25 and 29** suffer the most traffic accidents
comp_df = pd.merge(dd, dd2, left_index=True, right_index=True)
comp_df.plot.bar()
# #### How much the pattern change?
# The greatest increase is seen in those between **18 and 20** years old with an increase of 134%
#
# It is observed that the percentage of children between **6 and 9** years old decreases by 75%
comp_df = pd.merge(dd, dd2, left_index=True, right_index=True) # merge of the two database that I want to compare
comp_df['Diff_perc_point'] = comp_df['perc_party'] / comp_df['perc_day']-1
comp_df.style.bar(subset=['Diff_perc_point'], align='mid', color=['#d65f5f', '#5fba7d'])
# ## Evaluation <a name="Evaluation"></a>
# ### Results
#
# The main findings of the code can be found at the post available [here](https://i-lucas.medium.com/traffic-accidents-are-as-different-as-day-and-night-51b52458646d).
| Traffic_accident_Madrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd07945e9a82d7512fbf96246d9bbc29cd2f106c1a4a9cf54c9563dadf10f2237d4
# ---
# # 05 - Apply
# ### Step 1. Import the necessary libraries
import pandas as pd
import json
import requests
import sys
# ### Step 2. This time you need to download a dataset from [Kaggle](https://www.kaggle.com/uciml/student-alcohol-consumption)
json_fullpath = "kaggle.json"
with open(json_fullpath, "r+") as outfile:
json_readed=json.load(outfile)
json_readed
print()
# ### Step 3. Assign it to a variable called df.
df=pd.read_csv("student-mat.csv")
df
# ### Step 4. For the purpose of this exercise slice the dataframe from 'school' until the 'guardian' column
df2=df.loc[:,"school":"guardian"]
df2
# ### Step 5. Create a lambda function that will capitalize strings.
cap=lambda x:x.capitalize()
# ### Step 6. Capitalize both Mjob and Fjob
df2["Mjob"]=df2["Mjob"].apply(cap)
df2["Fjob"]=df2["Fjob"].apply(cap)
df2
# ### Step 7. Print the last elements of the data set.
df2.tail()
# ### Step 8. Did you notice the original dataframe is still lowercase? Why is that? Fix it and capitalize Mjob and Fjob.
# +
#esta en el 6
# -
# ### Step 9. Create a function called majority that returns a boolean value to a new column called legal_drinker (Consider majority as older than 17 years old)
def majority (df2):
if df2 >17:
return True
else:
return False
df2["legal_drinker"]=df2["age"].apply(majority)
df2
# ### Step 10. Multiply every number of the dataset by 10.
# ##### I know this makes no sense, don't forget it is just an exercise
# %%timeit
def por10 (x):
if type(x)==int:
return x*10
return x
df2.applymap(por10)
| week4_EDA_np_pd_json_apis_regex/day5_matplotlib_I_api/exercises/pandas/05_Apply/Students_Alcohol_Consumption/Students Alcohol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%writefile prime.py
'''
its is a prime number function
'''
def prime():
'''
taking input here
'''
num = int(input("Enter a number:"))
if num > 1:
for i in range(2, num):
if (num % i) == 0:
print(num, "is not a prime number")
break
else:
print(num, "is a prime number")
else:
print(num, "is not a prime number")
# ! pylint "prime.py"
def prime():
'''
taking input here
'''
num = int(input("Enter a number:"))
if num > 1:
for i in range(2, num):
if (num % i) == 0:
print(num, "is not a prime number")
break
else:
print(num, "is a prime number")
else:
print(num, "is not a prime number")
prime()
# +
# %%writefile testprime.py
import prime
import unittest
class testprime(unittest.TestCase):
def testnumber(self):
num = int(input("Enter a number:"))
result = num, "is a prime number"
self.assertEquals(result,"num")
def testingnumbers(self):
num = int(input("Enter a number:"))
result = num, "is a prime number"
self.assertEquals(result,"num")
if __name__ == "__main__":
unittest.main()
# -
def armstrong():
for num in range(1, 1000):
# order of number
order = len(str(num))
# initialize sum
sum = 0
temp = num
while temp > 0:
digit = temp % 10
sum += digit ** order
temp //= 10
if num == sum:
yield num
print(list(armstrong()))
| assignment day 9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NIAID DATA HUB: Mycobacteria drug resistance prediction
# ---
# ## Setup
# ---
# We are using Gen3 SDK to query structure data and retrieve object data. After installing the gen3 package using pip and using the import statements to import the classes and functions from the package, we need to set and endpoint variable and an auth variable to initialize instances of the classes we just imported. The endpoint should be the url of the commons you would like to interact with, and the refresh_file should contain your API key, which you can obtain by logging into the commons and going to the **Profile** page to create an API key.
# installing packages
# !pip install gen3
# !pip install --force --upgrade gen3
# !pip install flatten_json
# !pip install pandas
# !pip install requests
# !pip install sh
from gen3.auth import Gen3Auth
from gen3.submission import Gen3Submission
from gen3.file import Gen3File
import subprocess
import pandas as pd
import nde_tb_function as nde
endpoint = "https://tb.niaiddata.org/"
auth = Gen3Auth(endpoint, refresh_file = "/home/jovyan/pd/credentials.json")
sub = Gen3Submission(endpoint, auth)
file = Gen3File(endpoint, auth)
# ## Query
# We will use Gen3 Python SDK to run GraphQL queries on NIAID Data Hub using the Gen3Submission class. You can pass your query as a string and use the Gen3Submission.query() function to receive the results of your query.
object_dict = nde.query_file("TB-PATRIC",10,2,{"isoniazid_res_phenotype":"Resistant","amikacin_res_phenotype":"Resistant"})
df = nde.parse_json(object_dict,10)
# ### Run Ariba for drug resistance prediction
# We are getting reference data from CARD as an example. Ariba getref generates reference fasta file and reference metadata file for drug resistance prediction. User can use customized reference fasta file and reference metadata file to improve prediction accuracy.
subprocess.run(["ariba","getref","card","/home/jovyan/pd/nb_output/tb/ariba/reference"])
# After getting reference fasta and reference metadata files, Ariba prepareref generates gene clusters or variants clusters
subprocess.run(["ariba","prepareref","-f","/home/jovyan/pd/nb_output/tb/ariba/reference.fa","-m","/home/jovyan/pd/nb_output/tb/ariba/reference.tsv","/home/jovyan/pd/nb_output/tb/ariba/prepareref.out"])
# Ariba run runs local assembly to map raw sequences to gene clusters/variant clusters conveying drug resistance
nde.runAriba(df)
# Ariba summary creates a summary matrix from individual report files to give an overview of gene cluster/variant clusters occurrance among all the samples tested.
nde.extract_ariba_predict("/home/jovyan/pd/nb_output/tb/ariba/output")
# ### Run Mykrobe for drug resistance prediction
nde.runMykrobe(df)
# #### Extract Mykrobe resistant prediction
nde.extract_mykrobe_predict(df)
# ### Submission of Ariba and Mykrobe to Sheepdog
data = nde.extract_ariba_predict("/home/jovyan/pd/nb_output/tb/ariba/output")
nde.submit_results(data,"Ariba")
| demo/TB_notebook/nde_tb_pynb.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from fastai2.text.all import *
from nbdev.showdoc import *
# +
# all_slow
# -
# # Transfer learning in text
#
# > How to fine-tune a language model and train a classifier
# ## Finetune a pretrained Language Model
# First we get our data and tokenize it.
path = untar_data(URLs.IMDB_SAMPLE)
df = pd.read_csv(path/'texts.csv')
# Then we put it in a `Datasets`. For a language model, we don't have targets, so there is only one transform to numericalize the texts.
splits = ColSplitter()(df)
tfms = [attrgetter("text"), Tokenizer.from_df("text"), Numericalize()]
dsets = Datasets(df, [tfms], splits=splits, dl_type=LMDataLoader)
# Then we use that `Datasets` to create a `DataLoaders`. Here the class of `TfmdDL` we need to use is `LMDataLoader` which will concatenate all the texts in a source (with a shuffle at each epoch for the training set), split it in `bs` chunks then read continuously through it.
dls = dsets.dataloaders(bs=64, seq_len=72)
# Or more simply with a factory method:
dls = TextDataLoaders.from_df(df, text_col='text', is_lm=True, valid_col='is_valid')
dls.show_batch(max_n=2)
# Then we have a convenience method to directly grab a `Learner` from it, using the `AWD_LSTM` architecture.
learn = language_model_learner(dls, AWD_LSTM, metrics=[accuracy, Perplexity()], path=path, wd=0.1).to_fp16()
learn.freeze()
learn.fit_one_cycle(1, 1e-2)
learn.unfreeze()
learn.fit_one_cycle(4, 1e-2)
# Once we have fine-tuned the pretrained language model to this corpus, we save the encoder since we will use it for the classifier.
learn.show_results()
learn.save_encoder('enc1')
# ## Use it to train a classifier
# For classification, we need to use two set of transforms: one to numericalize the texts and the other to encode the labels as categories. Note that we have to use the same vocabulary as the one used in fine-tuning the language model.
lm_vocab = dls.vocab
splits = ColSplitter()(df)
x_tfms = [attrgetter("text"), Tokenizer.from_df("text"), Numericalize(vocab=lm_vocab)]
dsets = Datasets(df, splits=splits, tfms=[x_tfms, [attrgetter("label"), Categorize()]], dl_type=SortedDL)
# We once again use a subclass of `TfmdDL` for the dataloaders, since we want to sort the texts (sortish for the training set) by order of lengths. We also use `pad_collate` to create batches form texts of different lengths.
dls = dsets.dataloaders(before_batch=pad_input_chunk)
# And there is a factory method, once again:
dls = TextDataLoaders.from_df(df, text_col="text", text_vocab=lm_vocab, label_col='label', valid_col='is_valid', bs=32)
dls.show_batch(max_n=2, trunc_at=60)
# Then we once again have a convenience function to create a classifier from this `DataLoaders` with the `AWD_LSTM` architecture.
learn = text_classifier_learner(dls, AWD_LSTM, metrics=[accuracy], path=path,drop_mult=0.5)
learn = learn.load_encoder('enc1')
# Then we can train with gradual unfreezing and differential learning rates.
learn.fit_one_cycle(4)
learn.unfreeze()
learn.opt = learn.create_opt()
learn.fit_one_cycle(8, slice(1e-5,1e-3))
learn.show_results(max_n=4, trunc_at=60)
learn.predict("This was a good movie")
from fastai2.interpret import *
interp = Interpretation.from_learner(learn)
interp.plot_top_losses(6)
| nbs/38_tutorial.ulmfit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gather background data on the Holocaust
# ## Dates, people, places and events
#
# Material is extracted from the glossary page of the United States Holocaust Memorial Museum, https://www.ushmm.org/.
#
# The page was downloaded as a flat HTML file, and processed using the code below.
#
# Due to the difficulties in correctly sub-classing noun types, an interactive spreadsheet is used in this notebook ... for the ontologist to insert the appropriate superclasses and to indicate if a term is a type or instance.
#
# Note that this notebook must be executed in Jupyter Notebook (vs JupyterLab due to problems with qgrid in JupyterLab).
# +
## Imports
from bs4 import BeautifulSoup
import string
import pandas as pd
import qgrid
# -
## Constants
ttl_prefix = '@prefix : <urn:ontoinsights:ontology:dna:> . \n'\
'@prefix dna: <urn:ontoinsights:ontology:dna:> . \n'\
'@prefix owl: <http://www.w3.org/2002/07/owl#> . \n'\
'@prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> . \n'\
'@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> . \n'\
'@prefix xsd: <http://www.w3.org/2001/XMLSchema#> . \n\n'
def fix_up_name(text: str) -> (str, str):
"""
Fix the word(s) that will become the class name to remove quotation marks, be upper camel-cased,
handle parentheses and commas, and remove spaces.
The input parameter is the text that will become the class name.
The output parameter is the 'label' for the text.
"""
label = text.replace('"', '') # Remove quotation marks
label = label.replace('“', '')
label = label.replace('”', '')
label = label.title() # Upper camel-cased
if '(' in label:
paren_index = label.find('(')
class_name = label[0:paren_index].strip() # Get rid of anything after parentheses
label_text = f'{label[0:paren_index]}; {label[paren_index + 1:label.find(')')]}'
else:
class_name = label
label_text = label
class_name = class_name.replace(',', '') # Remove commas
class_name = class_name.replace(' ', '') # Remove spaces
return class_name, label_text
# +
# Process the glossary
with open('Glossary _ Holocaust Encyclopedia.html', 'r') as gloss_in:
gloss_page = gloss_in.read()
soup = BeautifulSoup(gloss_page, 'html.parser')
terms = soup.find('div', {'class': 'article-main-text'})
# print(terms) results are:
# <div class="article-main-text" id="story-text">
# <p><strong><a href="/narrative/3225/en">Antisemitism</a>:</strong> hostility toward or hatred of Jews as a religious or ethnic group, often accompanied by social, economic, or political discrimination.</p>
# <p><strong><em>Appellplatz</em>:</strong> German word for roll call square where prisoners were forced to assemble.</p>
# ...
# </div>
# Create lists of glossary terms, their labels and their definitions
gloss_terms = list()
gloss_labels = list()
gloss_defns = list()
gloss = ''
for term in terms.find_all('p'):
found_colon = term.get_text().find(':', 0, term.find('</strong>'))
if found_colon > 0:
# Colon indicates that there is a new term being defined
# Save the info
if gloss:
gloss_terms.append(gloss)
gloss_labels.append(label)
gloss_defns.append(new_defn)
text = term.get_text().split(':')
gloss, label = fix_up_name(text[0])
defn = text[1].strip()
# Make sure that the first character of the defn is upper case and that double quotes are escaped
new_defn = (defn[0].upper() + defn[1:]).replace('"', '\\"')
else:
# Another paragraph but not a new term
defn = term.get_text().replace('"', '\\"')
new_defn += defn
# When finished, write out the last term
gloss_terms.append(gloss)
gloss_labels.append(label)
gloss_defns.append(new_defn)
# Turn the lists into a dataframe
dict = {'Term': gloss_terms, 'Label': gloss_labels, 'Defn': gloss_defns}
gloss_df = pd.DataFrame(dict)
# Add columns to be hand-edited (using qgrid, next)
gloss_df['Superclass'] = ''
gloss_df['IsInstance'] = 'False'
# +
# To run qgrid, need to 1) import it and 2) have executed:
# jupyter nbextension enable --py --sys-prefix qgrid
# jupyter nbextension enable --py --sys-prefix widgetsnbextension # only required if you have not enabled the ipywidgets nbextension yet
# Currently qgrid does not work in JupyterLab 3
# QGrid is used to add superclass and instance info for the new concepts
# Also, used to fix up any acronyms (e.g., 'SS') since they are not capitalized correctly due to .title()
# And, added synonyms from the text to the labels (will be done automatically later)
grid_widget = qgrid.show_grid(gloss_df, show_toolbar=True)
grid_widget
# -
updated_gloss = grid_widget.get_changed_df()
print(updated_gloss.iloc[[25]])
# Write out the dataframe as turtle
with open('holocaust-gloss.ttl', 'w') as gloss_out:
# Write the prefix details
gloss_out.write(ttl_prefix)
# Write out each gloss term
for index, row in updated_gloss.iterrows():
superclass_text = row['Superclass']
if superclass_text == 'XXX': # Term will be addressed manually
continue
if ',' in superclass_text:
superclass_text = superclass_text.replace(', ', ', :')
term_text = row['Term']
label_text = row['Label']
if ';' in label_text:
label_text = label_text.replace('; ', '", "')
defn_text = row['Defn']
instance_text = row['IsInstance']
if instance_text == 'True':
gloss_out.write(f':{term_text} a :{superclass_text} ;\n')
else:
gloss_out.write(f':{term_text} a owl:Class ;\n rdfs:subClassOf :{superclass_text} ;\n')
gloss_out.write(f' rdfs:label "{label_text}" ;\n :defn "{defn_text}" .\n\n')
# +
# Note that the resulting Turtle is then further hand-edited and stored in the /ontologies directory
# -
| notebooks/Backgrd_Holocaust_Glossary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming Microblaze Subsystems from Jupyter
#
# In the Base I/O overlays that accompany the PYNQ release Microblazes are used to control peripherals attached to the various connectors. These can either be programmed with existing programs compiled externally or from within Jupyter. This notebook explains how the Microblazes can be integrated into Jupyter and Python.
#
# The Microblaze is programmed in C as the limited RAM available (64 KB) limits what runtimes can be loaded - as an example, the MicroPython runtime requires 256 KB of code and data space. The PYNQ framework provides a mechanism to write the C code inside Jupyter, compile it, load it on to the Microblaze and then execute and interact with it.
#
# The first stage is to load an overlay.
# +
from pynq.overlays.base import BaseOverlay
base = BaseOverlay('base.bit')
# -
# Now we can write some C code. The `%%microblaze` magic provides an environment where we can write the code and it takes a single argument - the Microblaze we wish to target this code at. This first example simply adds two numbers together and returns the result.
# +
# %%microblaze base.PMODA
int add(int a, int b) {
return a + b;
}
# -
# The functions we defined in the magic are now available for us to interact with in Python as any other function.
add(4,6)
# ## Data Motion
#
# The main purpose of the Python bindings it to transfer data between the host and slave processors. For simple cases, any primitive C type can be used as function parameters and return values and Python values will be automatically converted as necessary.
# +
# %%microblaze base.PMODA
float arg_passing(float a, char b, unsigned int c) {
return a + b + c;
}
# -
arg_passing(1, 2, 3)
# Arrays can be passed in two different way. If a type other than `void` is provided then the data will be copied to the microblaze and if non-`const` the data will be copied back as well. And iterable and modifiable object can be used as the argument in this case.
# +
# %%microblaze base.PMODA
int culm_sum(int* val, int len) {
int sum = 0;
for (int i = 0; i < len; ++i) {
sum += val[i];
val[i] = sum;
}
return sum;
}
# -
numbers = [i for i in range(10)]
culm_sum(numbers, len(numbers))
print(numbers)
# Finally we can pass a `void` pointer which will allow the Microblaze to directly access the memory of the host processing system for transferring large quantities of data. In Python these blocks of memory should be allocated using the `pynq.allocate` function and it is the responsibility of the programmer to make sure that the Python and C code agree on the types used.
# +
# %%microblaze base.PMODA
long long big_sum(void* data, int len) {
int* int_data = (int*)data;
long long sum = 0;
for (int i = 0; i < len; ++i) {
sum += int_data[i];
}
return sum;
}
# +
from pynq import allocate
buffer = allocate(shape=(1024 * 1024), dtype='i4')
buffer[:] = range(1024*1024)
big_sum(buffer, len(buffer))
# -
# ## Debug printing
#
# One unique feature of the PYNQ Microblaze environment is the ability to print debug information directly on to the Jupyter or Python console using the new `pyprintf` function. This functions acts like `printf` and `format` in Python and allows for a format string and variables to be passed back to Python for printing. In this release on the `%d` format specifier is supported but this will increase over time.
#
# To use `pyprintf` first the appropriate header needs to be included
# +
# %%microblaze base.PMODA
#include <pyprintf.h>
int debug_sum(int a, int b) {
int sum = a + b;
pyprintf("Adding %d and %d to get %d\n", a, b, sum);
return sum;
}
# -
debug_sum(1,2)
# ## Long running processes
#
# So far all of the examples presented have been synchronous with the Python code with the Python code blocking until a result is available. Some applications call instead for a long-running process which is periodically queried by other functions. If a C function return `void` then the Python process will resume immediately leaving the function running on its own.
#
# Other functions can be run while the long-running process is active but as there is no pre-emptive multithreading the persistent process will have to `yield` at non-timing critical points to allow other queued functions to run.
#
# In this example we launch a simple counter process and then pull the value using a second function.
# +
# %%microblaze base.PMODA
#include <yield.h>
static int counter = 0;
void start_counter() {
while (1) {
++counter;
yield();
}
}
int counter_value() {
return counter;
}
# -
# We can now start the counter going.
start_counter()
# And interrogate its current value
counter_value()
# There are some limitations with using `pyprintf` inside a persistent function in that the output will not be displayed until a subsequent function is called. If the buffer fills in the meantime this can cause the process to deadlock.
#
# Only one persistent process can be called at once - if another is started it will block the first until it returns. If two many processes are stacked in this way a stack overflow may occur leading to undefined results.
# ## Creating class-like objects
#
# In the C code `typedef`s can be used to create psuedo classes in Python. If you have a `typedef` called `my_class` then any functions that being `my_class_` are assumed to be associated with it. If one of those functions takes `my_class` as the first argument it is taken to be equivalent to `self`. Note that the `typedef` can only ultimately refer a primitive type. The following example does some basic modular arithmetic base 53 using this idiom.
# +
# %%microblaze base.PMODA
typedef unsigned int mod_int;
mod_int mod_int_create(int val) { return val % 53; }
mod_int mod_int_add(mod_int lhs, int rhs) { return (lhs + rhs) % 53; }
# -
# We can now create instances using our `create` function and call the `add` method on the returned object. The underlying value of the typedef instance can be retrieved from the `.val` attribute.
a = mod_int_create(63)
b = a.add(4)
print(b)
print(b.val)
# ## Coding Guidelines for Microblaze Interfacing Code
#
# There are some limitations to be aware of in the Jupyter integration with the Microblaze subsystem in particular the following things are unsupported and will result in the function not being available.
#
# * `struct`s or `union`s of any kind
# * Pointers to pointers
# * returning pointers
#
# All non `void*` paramters are passed on the stack so beware of passing large arrays in this fashion or a stack overflow will result.
| boards/Pynq-Z1/base/notebooks/microblaze/microblaze_programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib as mpl
mpl.rcParams['text.usetex'] = True
mpl.rcParams['text.latex.unicode'] = True
blues = mpl.cm.get_cmap(mpl.pyplot.get_cmap('Blues'))
greens = mpl.cm.get_cmap(mpl.pyplot.get_cmap('Greens'))
reds = mpl.cm.get_cmap(mpl.pyplot.get_cmap('Reds'))
oranges = mpl.cm.get_cmap(mpl.pyplot.get_cmap('Oranges'))
purples = mpl.cm.get_cmap(mpl.pyplot.get_cmap('Purples'))
greys = mpl.cm.get_cmap(mpl.pyplot.get_cmap('Greys'))
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import warnings
warnings.filterwarnings('ignore')
# !radical-stack
# +
Des1TilingParameters = [1.92036472e-02, 6.04868300e+01]
Des2TilingParameters = [3.17470564e-02, 6.46812975e+01]
Des2ATilingParameters = [0.08647082, -9.00736028]
Des2ATilingParameters2 = [0.03841578, 37.914536091]
Des1PredParameters = [5.21328399e-02, 1.28531895e+02]
Des2PredParameters = [4.71315045e-02, 9.58382425e+01]
Des2aPredParameters = [4.79876605e-02, 8.73630794e+01]
x = np.array(range(50,2950,50))
def func_lin(x,a,b):
return a*x+b
# -
fig,axis = mpl.pyplot.subplots(nrows=1,ncols=1,figsize=(11,5),sharex='row')
_ = axis.plot(x, func_lin(x,*Des1TilingParameters), 'r-', label="Design 1 Fitted Linear Curve")
_ = axis.fill_between(x, func_lin(x,*Des1TilingParameters) - 1.9257976881740269, func_lin(x,*Des1TilingParameters) + 1.9257976881740269, color=reds(250), alpha=0.2)
_ = axis.plot(x, func_lin(x,*Des2TilingParameters), 'g-', label="Design 2 Fitted Linear Curve")
_ = axis.fill_between(x, func_lin(x,*Des2TilingParameters) - 5.496694974632333, func_lin(x,*Des2TilingParameters) + 5.496694974632333, color=greens(250), alpha=0.2)
_ = axis.plot(x, func_lin(x,*Des2ATilingParameters), 'b-', label="Design 2A Fitted Linear Curve")
_ = axis.fill_between(x, func_lin(x,*Des2ATilingParameters) - 30.83509658381304, func_lin(x,*Des2ATilingParameters) + 30.83509658381304, color=blues(250), alpha=0.2)
_ = axis.set_ylabel('Execution Time in seconds',fontsize=20)
_ = axis.set_xlabel('Image Size in MBs',fontsize=20)
_ = axis.set_xticks(np.array(range(50,3050,125)))
_ = axis.set_xticklabels(axis.get_xticks().astype('int').tolist(),fontsize=18,rotation=45)
_ = axis.set_yticklabels(axis.get_yticks().astype('int').tolist(),fontsize=18)
_ = axis.grid('on')
_ = axis.legend(fontsize=18)
# _ = axis[1].plot(x, funcP(x,*Des1PredParameters), 'r-', label="Design 1 Fitted Linear Curve")
# _ = axis[1].plot(x, funcP(x,*Des2PredParameters), 'g-', label="Design 2 Fitted Linear Curve")
# _ = axis[1].set_ylabel('Execution Time in seconds',fontsize=14)
# _ = axis[1].set_xlabel('Image Size in MBs',fontsize=14)
# _ = axis[1].set_xticks(np.array(range(50,3050,125)))
# _ = axis[1].set_xticklabels(axis[0].get_xticks().astype('int').tolist(),fontsize=14,rotation=45)
# _ = axis[1].set_yticklabels(axis[0].get_yticks().astype('int').tolist(),fontsize=14)
# _ = axis[1].grid('on')
# _ = axis[1].legend(fontsize=14)
# _ = axis[0].set_title('Design 1 and 2 Model comparison for Seals use case',fontsize=20)
# fig.savefig('ModelComparison.pdf',dpi=800,bbox_inches='tight')
fig,axis = mpl.pyplot.subplots(nrows=1,ncols=1,figsize=(11,5),sharex='row')
_ = axis.plot(x, func_lin(x,*Des1TilingParameters), 'r-', label="Design 1 Fitted Linear Curve")
_ = axis.fill_between(x, func_lin(x,*Des1TilingParameters) - 1.9257976881740269, func_lin(x,*Des1TilingParameters) + 1.9257976881740269, color=reds(250), alpha=0.2)
_ = axis.plot(x, func_lin(x,*Des2TilingParameters), 'g-', label="Design 2 Fitted Linear Curve")
_ = axis.fill_between(x, func_lin(x,*Des2TilingParameters) - 5.496694974632333, func_lin(x,*Des2TilingParameters) + 5.496694974632333, color=greens(250), alpha=0.2)
_ = axis.plot(x, func_lin(x,*Des2ATilingParameters2), 'b-', label="Design 2A Fitted Linear Curve")
_ = axis.fill_between(x, func_lin(x,*Des2ATilingParameters2) - 3.889984729022672, func_lin(x,*Des2ATilingParameters2) + 3.889984729022672, color=blues(250), alpha=0.2)
_ = axis.set_ylabel('Execution Time in seconds',fontsize=20)
_ = axis.set_xlabel('Image Size in MBs',fontsize=20)
_ = axis.set_xticks(np.array(range(50,3050,125)))
_ = axis.set_xticklabels(axis.get_xticks().astype('int').tolist(),fontsize=18,rotation=45)
_ = axis.set_yticklabels(axis.get_yticks().astype('int').tolist(),fontsize=18)
_ = axis.grid('on')
_ = axis.legend(fontsize=18)
# _ = axis[1].plot(x, funcP(x,*Des1PredParameters), 'r-', label="Design 1 Fitted Linear Curve")
# _ = axis[1].plot(x, funcP(x,*Des2PredParameters), 'g-', label="Design 2 Fitted Linear Curve")
# _ = axis[1].set_ylabel('Execution Time in seconds',fontsize=14)
# _ = axis[1].set_xlabel('Image Size in MBs',fontsize=14)
# _ = axis[1].set_xticks(np.array(range(50,3050,125)))
# _ = axis[1].set_xticklabels(axis[0].get_xticks().astype('int').tolist(),fontsize=14,rotation=45)
# _ = axis[1].set_yticklabels(axis[0].get_yticks().astype('int').tolist(),fontsize=14)
# _ = axis[1].grid('on')
# _ = axis[1].legend(fontsize=14)
# _ = axis[0].set_title('Design 1 and 2 Model comparison for Seals use case',fontsize=20)
# fig.savefig('ModelComparison.pdf',dpi=800,bbox_inches='tight')
# +
# fig,axis = mpl.pyplot.subplots(nrows=1,ncols=1,figsize=(13,6),sharex='row')
# _ = axis.plot(x, func_lin(x,*Des2TilingParameters) - func_lin(x,*Des1TilingParameters), 'r-', label="Models Diff")
# _ = axis.set_xlabel('Image Size in MBs',fontsize=14)
# _ = axis.set_xticks(np.array(range(50,3050,125)))
# _ = axis.set_xticklabels(axis.get_xticks().astype('int').tolist(),fontsize=14,rotation=45)
# _ = axis.set_yticklabels(axis.get_yticks().astype('int').tolist(),fontsize=14)
# _ = axis.grid('on')
# _ = axis.legend(fontsize=14)
# +
# fig,axis = mpl.pyplot.subplots(nrows=1,ncols=1,figsize=(11,5),sharex='row')
# _ = axis.plot(x, func_lin(x,*Des1PredParameters), 'r-', label="Design 1 Fitted Linear Curve")
# _ = axis.fill_between(x, func_lin(x,*Des1PredParameters) - 5.731348094430763, func_lin(x,*Des1PredParameters) + 5.731348094430763, color=reds(250), alpha=0.2)
# _ = axis.plot(x, func_lin(x,*Des2PredParameters), 'g-', label="Design 2 Fitted Linear Curve")
# _ = axis.fill_between(x, func_lin(x,*Des2PredParameters) - 5.960059379846854, func_lin(x,*Des2PredParameters) + 5.960059379846854, color=greens(250), alpha=0.2)
# _ = axis.plot(x, func_lin(x,*Des2aPredParameters), 'b-', label="Design 2 Fitted Linear Curve")
# _ = axis.fill_between(x, func_lin(x,*Des2aPredParameters) - 6.1860055267227185, func_lin(x,*Des2aPredParameters) + 6.1860055267227185, color=blues(250), alpha=0.2)
# _ = axis.set_ylabel('Execution Time in seconds',fontsize=20)
# _ = axis.set_xlabel('Image Size in MBs',fontsize=20)
# _ = axis.set_xticks(np.array(range(50,3050,125)))
# _ = axis.set_xticklabels(axis.get_xticks().astype('int').tolist(),fontsize=18,rotation=45)
# _ = axis.set_yticklabels(axis.get_yticks().astype('int').tolist(),fontsize=18)
# _ = axis.grid('on')
# _ = axis.legend(fontsize=18)
# # _ = axis[1].plot(x, funcP(x,*Des1PredParameters), 'r-', label="Design 1 Fitted Linear Curve")
# # _ = axis[1].plot(x, funcP(x,*Des2PredParameters), 'g-', label="Design 2 Fitted Linear Curve")
# # _ = axis[1].set_ylabel('Execution Time in seconds',fontsize=14)
# # _ = axis[1].set_xlabel('Image Size in MBs',fontsize=14)
# # _ = axis[1].set_xticks(np.array(range(50,3050,125)))
# # _ = axis[1].set_xticklabels(axis[0].get_xticks().astype('int').tolist(),fontsize=14,rotation=45)
# # _ = axis[1].set_yticklabels(axis[0].get_yticks().astype('int').tolist(),fontsize=14)
# # _ = axis[1].grid('on')
# # _ = axis[1].legend(fontsize=14)
# # _ = axis[0].set_title('Design 1 and 2 Model comparison for Seals use case',fontsize=20)
# fig.savefig('ModelComparisonCounting.pdf',dpi=800,bbox_inches='tight')
| Seals/Notebooks/Des1Des2TTXModelingComp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from keras.utils import np_utils
import os
import matplotlib.pyplot as plt
import seaborn as sns
import cv2
import tensorflow
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense,Dropout,Flatten,Conv2D,MaxPooling2D,BatchNormalization
import time
import keras
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
dirdata1='forehand_train_img/'
dirdata2='backhand_train_img/'
def data_img(dirdata1,dirdata2):
data = [] # 1 step
label = []
height=120
BLACK = [0,0,0]
for i in os.listdir(dirdata1): # 2 step
img = cv2.imread(dirdata1+i)
x,y,z = img.shape
if x<=height:
constant = cv2.copyMakeBorder(img,0,height-x,0,0,cv2.BORDER_CONSTANT,value=BLACK)
data.append(constant) # 2-2
label.append(0) # 2-3
for i in os.listdir(dirdata2):
img = cv2.imread(dirdata2+i)
x,y,z = img.shape
if x<=height:
constant = cv2.copyMakeBorder(img,0,height-x,0,0,cv2.BORDER_CONSTANT,value=BLACK)
data.append(constant) # 2-2
label.append(1) # 2-3
data = np.array(data) # 3
label = np.array(label) #3
label_one_hot = np.eye(2)[label] # 4 one-hot encoding
data_norm = data / 255 #5
return data_norm,label_one_hot
data, label=data_img(dirdata1,dirdata2)
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(data, label, test_size=0.5, random_state=42)
from keras.applications import vgg16
#讀取vgg16模型並重設輸入層
vgg16=vgg16.VGG16(weights='imagenet', include_top=False, input_shape=(120,50,3))
vgg16.summary()
#建立新的模型
model = Sequential()
#將vgg16的訓練層讀進新模型
for layer in vgg16.layers:
model.add(layer)
model.summary()
#將vgg16的訓練層參數凍結
for layer in model.layers:
layer.trainable = False
#在新模型後加入新的輸出
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.25))
model.add(Dense(2, activation='softmax'))
model.summary()
# +
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
start_time = time.time()
# 開始訓練
train_history = model.fit(X_train,Y_train,validation_split=0.2,epochs=100, batch_size=30, verbose=1)
end_time = time.time()
# -
execution_time = (end_time - start_time)
print("執行時間: ",execution_time,"s")
# +
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
t = f.suptitle('class model evaluate', fontsize=12)
f.subplots_adjust(top=0.85, wspace=0.3)
class_epoch_list = list(range(1,101))
ax1.plot(class_epoch_list, train_history.history['accuracy'], label='class_Train Accuracy')
ax1.plot(class_epoch_list, train_history.history['val_accuracy'], label='class_Validation Accuracy')
ax1.set_xticks(np.arange(0, 101, 5))
ax1.set_ylabel('class_Accuracy Value')
ax1.set_xlabel('class_Epoch')
ax1.set_title('class_Accuracy')
l1 = ax1.legend(loc="best")
ax2.plot(class_epoch_list, train_history.history['loss'], label='class_Train Loss')
ax2.plot(class_epoch_list, train_history.history['val_loss'], label='class_Validation Loss')
ax2.set_xticks(np.arange(0, 101, 5))
ax2.set_ylabel('class_Loss Value')
ax2.set_xlabel('class_Epoch')
ax2.set_title('class_Loss')
l2 = ax2.legend(loc="best")
# +
#model.save('2d_cnn_transfer_vgg16.h5')
# -
modello = keras.models.load_model('2d_cnn_transfer_vgg16.h5')
test_dirdata1='forehand_test_img/'
test_dirdata2='backhand_test_img/'
test_data, test_label=data_img(test_dirdata1,test_dirdata2)
time_pre=modello.predict(test_data,verbose=2)
Y_test = np.argmax(test_label,axis=1)
Y_pred = np.argmax(time_pre,axis=1)
# +
mat = confusion_matrix(Y_test,Y_pred)
sns.set(font_scale=1.5)
plt.subplot(1,2,1)
sns.heatmap(mat, square=True, annot=True, fmt='d', cbar=False)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
plt.ylabel('true label')
plt.xlabel('predicted label');
target_names = ["forehand","backhand"]
print(classification_report(Y_test,Y_pred,target_names=target_names))
# -
| transfer_2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="2KQ6GDYcPpEK" colab_type="code" outputId="000d56ae-606c-44dc-82c4-19eb0a078c43" executionInfo={"status": "ok", "timestamp": 1587696521258, "user_tz": -480, "elapsed": 24040, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgtI1uuN3eLJSIU9wQoDnkXliOrNcbvHO6SpNS6Kg=s64", "userId": "03474220991191819164"}} colab={"base_uri": "https://localhost:8080/", "height": 190}
#Mount gdrive and change directory to project location.
from google.colab import drive
drive.mount('/content/drive')
# %cd 'drive/My Drive/cs6240 - media/vqa/TVQAplus/tvqa_plus_stage_features'
#Check CUDA availability
# !nvidia-smi
import torch
torch.cuda.is_available()
# + id="B63l5ahlq--v" colab_type="code" colab={}
# # %pip install torch==1.2.0+cu92 torchvision==0.4.0+cu92 -f https://download.pytorch.org/whl/torch_stable.html
# %pip install --upgrade tqdm
# %pip install h5py numpy
# + [markdown] id="wlLkVvEc-gEP" colab_type="text"
#
# + id="_xHo_5NRBuJA" colab_type="code" colab={}
# !python tvqa_small_dataset.py
# # %cd tvqa_plus_stage_features/
# # !ls
# # !python tvqa_small_dataset.py
# # %cd sample3
# # !ls
# # !cp eval_object_vocab.json /sample3
# # !cp frm_cnt_cache.json /sample
# # !cp frm_cnt_cache.json /sample3
# # !cp tvqa_bbt_bottom_up_pool5_hq_20_100_pca.h5 /sample3
# + id="tzxUqh7F_jD_" colab_type="code" outputId="c81239b9-f545-4d72-9331-b6f38c9e2856" executionInfo={"status": "ok", "timestamp": 1587696593187, "user_tz": -480, "elapsed": 3034, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgtI1uuN3eLJSIU9wQoDnkXliOrNcbvHO6SpNS6Kg=s64", "userId": "03474220991191819164"}} colab={"base_uri": "https://localhost:8080/", "height": 340}
# # %cd ..
# %cd '../tvqa_original_videos/tvqa_frames_hq (1)'
# !ls
# + id="DwQCxVp1A0JM" colab_type="code" outputId="9b565449-5616-4709-b0cd-5be00ba64f49" executionInfo={"status": "ok", "timestamp": 1587695549775, "user_tz": -480, "elapsed": 5917, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgtI1uuN3eLJSIU9wQoDnkXliOrNcbvHO6SpNS6Kg=s64", "userId": "03474220991191819164"}} colab={"base_uri": "https://localhost:8080/", "height": 61}
# %mkdir frames
# !cat tvqa_video_frames_fps3_hq.tar.gz.* | tar xz -C frames
# + id="R4Z_x2dwBN65" colab_type="code" colab={}
# !df -h .
# + id="S54pmxI5FGky" colab_type="code" colab={}
| TVQAplus/Preprocess.ipynb |
# ---
# layout: post
# title: "손실함수 이야기"
# author: <NAME>
# date: 2017-01-27 23:08:00
# categories: Keras
# comments: true
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# * mean_squared_error / mse
# * mean_absolute_error / mae
# * mean_absolute_percentage_error / mape
# * mean_squared_logarithmic_error / msle
# * squared_hinge
# * hinge
# * binary_crossentropy: Also known as logloss.
# * categorical_crossentropy: Also known as multiclass logloss. Note: using this objective requires that your labels are binary arrays of shape (nb_samples, nb_classes).
# * sparse_categorical_crossentropy: As above but accepts sparse labels. Note: this objective still requires that your labels have the same number of dimensions as your outputs; you may need to add a length-1 dimension to the shape of your labels, e.g with np.expand_dims(y, -1).
# * kullback_leibler_divergence / kld: Information gain from a predicted probability distribution Q to a true probability distribution P. Gives a measure of difference between both distributions.
# * poisson: Mean of (predictions - targets * log(predictions))
# * cosine_proximity: The opposite (negative) of the mean cosine proximity between predictions and targets.
| _writing/Temp 2017-1-27-LossFuncion_Talk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
import plotly
plotly.__version__
# ### Named Colors
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
mpl_fig = plt.figure()
ax = mpl_fig.add_subplot(111)
color_names = ["r", "g", "b", "peachpuff", "fuchsia"] # Some of the colors
ax.set_title('Named Colors in Matplotlib')
for i in range(1,6):
x = np.linspace(0,10,1000)
y = np.sin(x*(np.pi/i))
line, = ax.plot(x, y, lw=2, c=color_names[i-1],label='color:'+ color_names[i-1])
plotly_fig = tls.mpl_to_plotly( mpl_fig )
plotly_fig.layout.showlegend = True
plotly_fig.layout.width = 550
plotly_fig.layout.height = 400
py.iplot(plotly_fig)
# -
# ### Matplotlib Colormap
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
mpl_fig = plt.figure()
num = 1000
s = 121
x1 = np.linspace(-0.5,1,num) + (0.5 - np.random.rand(num))
y1 = np.linspace(-5,5,num) + (0.5 - np.random.rand(num))
x2 = np.linspace(-0.5,1,num) + (0.5 - np.random.rand(num))
y2 = np.linspace(5,-5,num) + (0.5 - np.random.rand(num))
x3 = np.linspace(-0.5,1,num) + (0.5 - np.random.rand(num))
y3 = (0.5 - np.random.rand(num))
ax1 = mpl_fig.add_subplot(221)
cb1 = ax1.scatter(x1, y1, c=x1, cmap=plt.cm.get_cmap('Blues'))
plt.colorbar(cb1, ax=ax1)
ax1.set_title('Blues')
ax2 = mpl_fig.add_subplot(222)
cb2 = ax2.scatter(x2, y2, c=x2, cmap=plt.cm.get_cmap('RdBu'))
plt.colorbar(cb2, ax=ax2)
ax2.set_title('RdBu')
ax3 = mpl_fig.add_subplot(223)
cb3 = ax3.scatter(x3, y3, c=x3, cmap=plt.cm.get_cmap('Dark2'))
plt.colorbar(cb3, ax=ax3)
ax3.set_xlabel('Dark2')
mpl_fig = plt.gcf()
plotly_fig = tls.mpl_to_plotly(mpl_fig)
py.iplot(plotly_fig)
# -
# ### Matplotlib Colormap Reversed
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
mpl_fig = plt.figure()
num = 1000
s = 121
x1 = np.linspace(-0.5,1,num) + (0.5 - np.random.rand(num))
y1 = np.linspace(-5,5,num) + (0.5 - np.random.rand(num))
x2 = np.linspace(-0.5,1,num) + (0.5 - np.random.rand(num))
y2 = np.linspace(5,-5,num) + (0.5 - np.random.rand(num))
x3 = np.linspace(-0.5,1,num) + (0.5 - np.random.rand(num))
y3 = (0.5 - np.random.rand(num))
ax1 = mpl_fig.add_subplot(221)
cb1 = ax1.scatter(x1, y1, c=x1, cmap=plt.cm.get_cmap('Blues_r'))
#plt.colorbar(cb1, ax=ax1)
ax1.set_title('Reversed Blues')
ax2 = mpl_fig.add_subplot(222)
cb2 = ax2.scatter(x2, y2, c=x2, cmap=plt.cm.get_cmap('RdBu_r'))
#plt.colorbar(cb2, ax=ax2)
ax2.set_title('Reversed RdBu')
ax3 = mpl_fig.add_subplot(223)
cb3 = ax3.scatter(x3, y3, c=x3, cmap=plt.cm.get_cmap('Dark2_r'))
#plt.colorbar(cb3, ax=ax3)
ax3.set_xlabel('Reversed Dark2')
mpl_fig = plt.gcf()
plotly_fig = tls.mpl_to_plotly(mpl_fig)
py.iplot(plotly_fig)
# -
# ### Setting Colormap Range
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
mpl_fig = plt.figure()
ax1 = mpl_fig.add_subplot(121)
x = np.linspace(1,10,100)
y = np.random.randint(1,10,100)
ax1.scatter(x,y, c=x, s=100, cmap=plt.cm.get_cmap('RdBu'))
ax1.set_title('Colormap range varying in X Direction')
ax2 = mpl_fig.add_subplot(122)
ax2.scatter(x,y, c=y, s=100, cmap=plt.cm.get_cmap('RdBu'))
ax2.set_title('Colormap range varying in Y Direction')
plotly_fig = tls.mpl_to_plotly(mpl_fig)
plotly_fig.layout.width = 500
plotly_fig.layout.height = 300
py.iplot(plotly_fig)
# -
# ### Colorbar Custom Range
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
delta = 0.025
x = np.arange(-3.0, 3.0, delta)
y = np.arange(-2.0, 2.0, delta)
X, Y = np.meshgrid(x, y)
Z = np.sin(8*X) + np.cos(8*Y)
mpl_fig = plt.figure()
plt.title('Setting Colorbar range Manually')
plotly_fig = tls.mpl_to_plotly(mpl_fig)
plotly_fig.add_trace(dict(type='contour',
x=x,
y=y,
z=Z,
colorbar=dict(nticks=10,
tickmode='array',
tickvals=[-2,-1,0,1,2]),
colorscale='Viridis'
)
)
plotly_fig.layout.width = 500
plotly_fig.layout.height = 400
py.iplot(plotly_fig)
# -
# ### Colorbar Custom Size And Ticks
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
delta = 0.025
x = np.arange(-3.0, 3.0, delta)
y = np.arange(-2.0, 2.0, delta)
X, Y = np.meshgrid(x, y)
Z = np.sin(8*X) + np.cos(8*Y)
mpl_fig = plt.figure()
plt.title('Simple Example with Custom Colorbar')
plotly_fig = tls.mpl_to_plotly(mpl_fig)
custom_colorbar = dict(nticks=10,
tickangle=20,
#titlefont=dict(family=Arial, type=sans-serif),
title="Custom Colorbar Title",
thickness=50,
len=1,
outlinewidth=2.2)
plotly_fig.add_traces([dict(type='contour', x=x, y=y, z=Z, colorbar=custom_colorbar)])
plotly_fig.layout.width = 500
plotly_fig.layout.height = 300
py.iplot(plotly_fig)
# -
# ### Matplotlib Colormap With Legend
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import matplotlib.pyplot as plt
import numpy as np
mpl_fig = plt.figure()
ax = mpl_fig.add_subplot(111)
for i in range(10):
x = np.random.normal(loc=i, size=100)
y = np.random.normal(loc=i, size=100)
ax.scatter(x,y,c=y, cmap=plt.cm.get_cmap('RdBu'), label='Trace {}'.format(i))
plotly_fig = tls.mpl_to_plotly(mpl_fig)
plotly_fig.layout.showlegend = True
plotly_fig.layout.width = 500
plotly_fig.layout.height = 400
py.iplot(plotly_fig)
# -
# #### Reference
# See [https://plot.ly/python/reference/](https://plot.ly/python/reference/) for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'colors-and-colormaps.ipynb', 'matplotlib/colors-and-colormaps/', 'Colors and Colormaps',
'Colors and Colorscale options in matplotlib. Examples of different colors and colormaps available in matplotlib.',
title = 'Matplotlib Colors and Colormaps | Examples | Plotly',
has_thumbnail='true', thumbnail='thumbnail/colors-and-colormaps.jpg',
language='matplotlib',
page_type='example_index',
display_as='basic', order=8,
ipynb= '~notebook_demo/228')
# -
| _posts/matplotlib/colors-and-colormaps/colors-and-colormaps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Atividades
# 1) Leia uma imagem colorida, transforme a mesma em níveis de cinza e salve o resultado em disco.
# +
from skimage.io import imread,imsave, imshow
from skimage.color import rgb2gray
image = imread('dog.jpeg')
img = rgb2gray(image)
imsave('cinza.png', img)
# -
# 2) Leia uma imagem colorida, transforme a mesma em níveis de cinza. Após isso, atribua o valor zero a todos os pixels cujo vizinho à direita tenha o mesmo valor de intensidade. Salve o resultado em disco.
# +
image = imread('dog.jpeg')
img = rgb2gray(image)
for i in range(img.shape[0]):
for j in range(img.shape[1]-1):
if(img[i][j] == img[i][j+1]):
img[i][j] = 0
imsave('img2questao.png', img)
# -
# 3) Leia uma imagem colorida, calcule as componentes no modelo de cores RGB. Calcule a maior e menor intensidade dos pixels em cada componente.
# +
def maiorMenor(img):
maior = 0
menor = 9999
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if(img[i][j] < menor):
menor = img[i][j]
if(img[i][j] > maior):
maior = img[i][j]
return maior,menor
image = imread('dog.jpeg')
red = image[:,:,0]
green = image[:,:,1]
blue = image[:,:,2]
maior, menor = maiorMenor(red)
print("Red: " + str(maior) + "; " + str(menor))
maior, menor = maiorMenor(green)
print("Green: " + str(maior) + "; " + str(menor))
maior, menor = maiorMenor(blue)
print("Blue: " + str(maior) + "; " + str(menor))
# -
# 4) Leia uma imagem colorida, calcule as componentes no modelo de cores RGB. Atribua o valor 255 a todos os pixels com intensidade maior que 150, faça isso em cada componente. Após isso, junte as componentes para formar uma imagem colorida novamente e salve o resultado.
# +
import numpy as np
image = imread('dog.jpeg')
red = image[:,:,0]
green = image[:,:,1]
blue = image[:,:,2]
for i in range(image.shape[0]):
for j in range(image.shape[1]):
if(red[i][j] > 150):
red[i][j] = 255
if(green[i][j] > 150):
green[i][j] = 255
if(blue[i][j] > 150):
blue[i][j] = 255
img = np.zeros((image.shape[0],image.shape[1],3),'uint8')
img[:,:,0] = red
img[:,:,1] = green
img[:,:,2] = blue
imsave('juntaCanais.png', img)
# -
# 5) Aplique ruído em uma imagem utilizando a função "skimage.util.random_noise" com os seguites parâmentros: ‘s&p’,‘gaussian’ e ‘speckle’. Cada um desses parâmetros irá gerar uma imagem com ruído. Após isso, aplique funções disponíveis em 'skimage.restoration' para recuperar as imagens.
# +
from skimage.util import random_noise
from skimage import restoration
import matplotlib.pyplot as plt
image = imread('dog.jpeg')
img = rgb2gray(image)
sp = random_noise(img, 's&p')
gaussian = random_noise(img, 'gaussian')
speckle = random_noise(img, 'speckle')
denoiseSp = restoration.denoise_tv_chambolle(sp)
denoiseGaussian = restoration.denoise_tv_chambolle(gaussian)
denoiseSpeckle = restoration.denoise_tv_chambolle(speckle)
plt.figure()
plt.imshow(denoiseSp,cmap = 'gray')
plt.figure()
plt.imshow(denoiseGaussian,cmap = 'gray')
plt.figure()
plt.imshow(denoiseSpeckle,cmap = 'gray')
# -
# 6) Aplique pelo menos 4 métodos diferentes para binarizar uma imagem ('skimage.filters').
# +
from skimage.io import imread,imsave, imshow
from skimage.color import rgb2gray
import matplotlib.pyplot as plt
from skimage.filters import threshold_otsu, threshold_yen, threshold_isodata, threshold_li
def binThresholdOtsu(img):
thresh = threshold_otsu(img)
binary = img < thresh
return binary
def binThresholdYen(img):
thresh = threshold_yen(img)
binary = img < thresh
return binary
def binThresholdIsodata(img):
thresh = threshold_isodata(img)
binary = img < thresh
return binary
def binThresholdLi(img):
thresh = threshold_li(img)
binary = img < thresh
return binary
image = imread('dog.jpeg')
img = rgb2gray(image)
otsu = binThresholdOtsu(img)
yen = binThresholdYen(img)
isodata = binThresholdIsodata(img)
li = binThresholdLi(img)
plt.figure()
plt.imshow(otsu,cmap = 'gray')
plt.figure()
plt.imshow(yen,cmap = 'gray')
plt.figure()
plt.imshow(isodata,cmap = 'gray')
plt.figure()
plt.imshow(li,cmap = 'gray')
# -
# 7) Aplique pelo menos 4 métodos diferentes ('skimage.filters') para calcular as bordas de uma imagem utilizando como entrada o resultado da questão anterior.
# +
from skimage.filters import sobel,roberts,prewitt,scharr
bordaOtsu = sobel(otsu)
bordaYen = roberts(yen)
bordaIsoData = prewitt(isodata)
bordaLi = scharr(li)
plt.figure()
plt.imshow(bordaOtsu,cmap = 'gray')
plt.figure()
plt.imshow(bordaYen,cmap = 'gray')
plt.figure()
plt.imshow(bordaIsoData,cmap = 'gray')
plt.figure()
plt.imshow(bordaLi,cmap = 'gray')
# -
# 8) Recorte uma imagem da seguinte forma:
# - O corte será feito no canal de cor com a maior média de pixels.
# - O centro do corte deve ser o pixel com o maior desvio padrão em uma janela 3x3.
# - O tamanho do recorte será 1/4 do tamanho da imagem. Ex: Imagem 100 x 80 -> Recorte 25 x 20
# - Cuidado com as bordas! Caso uma janela fique fora da imagem, o centro do recorte deverá ser alterado.
# +
from skimage.io import imread,imsave, imshow
from skimage.color import rgb2gray
import matplotlib.pyplot as plt
import numpy as np
img = imread('dog.jpeg')
r = img[:,:,0]
g = img[:,:,1]
b = img[:,:,2]
print(r.shape)
mediaR = r.max()
mediaG = g.max()
mediaB = b.max()
maior = -1
if(mediaR > mediaG) and (mediaR > mediaB):
maior = r
elif(mediaG > mediaR) and (mediaG > mediaB):
maior = g
elif(mediaB > mediaR) and (mediaB > mediaG):
maior = b
else:
maior = r #considerei que no caso dos canais terem a mesma média, o corte será feito no red
maiorDp = -1
posicoes = [-1, -1]
for i in range(1, (maior.shape[0])-1):
for j in range(1, (maior.shape[1])-1):
janela = []
janela.append(maior[i][j])
janela.append(maior[i-1][j-1])
janela.append(maior[i-1][j])
janela.append(maior[i-1][j+1])
janela.append(maior[i][j-1])
janela.append(maior[i][j+1])
janela.append(maior[i+1][j-1])
janela.append(maior[i+1][j])
janela.append(maior[i+1][j+1])
desvio = np.std(janela)
if(desvio > maiorDp):
maiorDp = desvio
posicoes[0] = i
posicoes[1] = j
tamLin = int(r.shape[0]/4)
tamCol = int(r.shape[1]/4)
corteX = int(tamLin/2)
corteY = int(tamCol/2)
l = posicoes[0] - corteX
c = posicoes[1] - corteY
print(tamLin, tamCol)
print(posicoes)
print(l, c)
if(posicoes[0]+corteX) > (maior.shape[0]):
sobraX = posicoes[0]+corteX-maior.shape[0]
posicoes[0] = posicoes[0] - sobraX
l -= sobraX
#corteX -= sobraX
if(posicoes[1]+corteY) > (maior.shape[1]):
sobraY = posicoes[1]+corteY-maior.shape[1]
posicoes[1] = posicoes[1] - sobraY
c -= sobraY
#corteY -= sobraY
cont1 = 0
cont2 = 0
for i in range(l, posicoes[0]+corteX):
cont1 += 1
for j in range(c, posicoes[1]+corteY):
maior[i][j] = 0
cont2 = j
plt.figure()
plt.imshow(img)
plt.figure()
plt.imshow(img[l:posicoes[0]+corteX,c:posicoes[1]+corteY,:])
# -
| Trabalho 1 - Manipulando Imagens/manipulandoImgs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="LnhjV7zhmafJ" colab_type="code" outputId="26e7848e-bba9-4901-b190-1b41da7dfcff" executionInfo={"status": "ok", "timestamp": 1574220901670, "user_tz": 300, "elapsed": 2737, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import json, re
from tqdm import tqdm_notebook
from uuid import uuid4
from google.colab import drive
from sklearn.model_selection import train_test_split
## Torch Modules
import torch
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
from torch.utils.data import Dataset, DataLoader
print(torch.__version__)
# https://colab.research.google.com/drive/1xg4UMQmXjDik3v9w-dAsk4kq7dXX_0Fm#scrollTo=Pexochza5lZq
# + id="MBNLJDLlmlBU" colab_type="code" outputId="c65a5c3e-4b78-49c7-c985-17d943e86ee4" executionInfo={"status": "ok", "timestamp": 1574220908634, "user_tz": 300, "elapsed": 9689, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 649}
# !pip install pytorch-transformers
# + id="dwRPMe1xnklx" colab_type="code" colab={}
## PyTorch Transformer
from pytorch_transformers import RobertaModel, RobertaTokenizer
from pytorch_transformers import RobertaForSequenceClassification, RobertaConfig
# + id="PMNwWjXlrP58" colab_type="code" outputId="90e6b68f-efc6-4be6-c363-e4ddf9a2c163" executionInfo={"status": "ok", "timestamp": 1574220908637, "user_tz": 300, "elapsed": 9678, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
## Check if Cuda is Available
print(torch.cuda.is_available())
# + [markdown] id="TAQ-od7CxvXH" colab_type="text"
# ## Loading data from Github
# + id="5eqIHmo1x1cJ" colab_type="code" colab={}
# main training dataset
cbert_train_url = 'https://raw.githubusercontent.com/1024er/cbert_aug/develop/datasets/subj/train.tsv'
cbert_test_url = 'https://raw.githubusercontent.com/1024er/cbert_aug/develop/datasets/subj/test.tsv'
cbert_train = pd.read_csv(cbert_train_url, sep='\t')
cbert_test = pd.read_csv(cbert_test_url, sep='\t')
# + id="F33ly4cnyOFk" colab_type="code" outputId="25cfbc9f-b2af-41a8-b1a9-fb65c39e6351" executionInfo={"status": "ok", "timestamp": 1574220909765, "user_tz": 300, "elapsed": 10791, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
print(cbert_train.shape, cbert_test.shape)
cbert_test.head()
# + id="v3iV7ZQCNvxT" colab_type="code" colab={}
# subj_train_samples = subj_train.sample(frac=0.5, random_state=0).reset_index(drop=True)
# subj_test_samples = subj_test.sample(frac=0.5, random_state=0).reset_index(drop=True)
# + id="fR0-HDH34Xlt" colab_type="code" colab={}
# dataset 2
mqpq_train_url = "https://raw.githubusercontent.com/1024er/cbert_aug/develop/datasets/mpqa/train.tsv"
mqpq_test_url = "https://raw.githubusercontent.com/1024er/cbert_aug/develop/datasets/mpqa/test.tsv"
mpqa_train = pd.read_csv(mqpq_train_url, sep='\t')
mpqa_test = pd.read_csv(mqpq_test_url, sep='\t')
# + id="6I4dAGli5jke" colab_type="code" outputId="c16ba406-c372-4504-80c8-b570666e6ea4" executionInfo={"status": "ok", "timestamp": 1574220910015, "user_tz": 300, "elapsed": 10992, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
print(mpqa_train.shape, mpqa_test.shape)
mpqa_train.head()
# + id="5D0ANBsD4saW" colab_type="code" outputId="bc9db4dd-6490-4add-b566-c99da4e2a670" executionInfo={"status": "ok", "timestamp": 1574220963616, "user_tz": 300, "elapsed": 64584, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 275}
# dataset 3
drive.mount('/gdrive')
# %cd '/gdrive/My Drive/281 Project'
# !ls
# + id="aBBox63H5-le" colab_type="code" outputId="08cf9107-7257-4f2b-ecfa-5af42b5e4f0e" executionInfo={"status": "ok", "timestamp": 1574220963619, "user_tz": 300, "elapsed": 64574, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
cornell_df= pd.read_csv('dataset3-movies-cornell.csv')
cornell_df.head()
# + id="IvEQvs6x8HcA" colab_type="code" outputId="2aee0db6-9948-46f2-addb-4bd0b9b9ddf9" executionInfo={"status": "ok", "timestamp": 1574220963622, "user_tz": 300, "elapsed": 64574, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 122}
cornell_train, cornell_test = train_test_split(cornell_df, test_size=0.2, shuffle=True, random_state=0)
cornell_train.reset_index(inplace=True)
cornell_train.drop(columns=['index'], inplace=True)
cornell_test.reset_index(inplace=True)
cornell_test.drop(columns=['index'], inplace=True)
# + id="1aLnF3Hr9LCi" colab_type="code" outputId="fe0fb4cf-f345-4ee8-b4be-c135be136fd1" executionInfo={"status": "ok", "timestamp": 1574220963623, "user_tz": 300, "elapsed": 64566, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
print(cornell_train.shape, cornell_test.shape)
cornell_test.head()
# + [markdown] id="nS9B3eqVrQ8H" colab_type="text"
# ## Loading RoBERTa classes
# + id="Tv0y2PGMrZ2e" colab_type="code" outputId="7f535442-e27d-4705-aaf2-998ef6e75324" executionInfo={"status": "ok", "timestamp": 1574220964776, "user_tz": 300, "elapsed": 65706, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 391}
# subjective: 0
# objective: 1
label_to_ix = {0: 0, 1: 1}
print(label_to_ix)
config = RobertaConfig.from_pretrained('roberta-base')
config.num_labels = len(list(label_to_ix.values()))
config
# + id="DwAL-AcDra5C" colab_type="code" outputId="2913354d-26fb-46af-daac-24d829f6df12" executionInfo={"status": "ok", "timestamp": 1574220970691, "user_tz": 300, "elapsed": 71618, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
tokenizer = RobertaTokenizer.from_pretrained('roberta-base')
model = RobertaForSequenceClassification(config)
# + [markdown] id="g_dClEc2rk_x" colab_type="text"
# ## Feature Preparation
# + id="YOxH_cb1r13a" colab_type="code" colab={}
def prepare_features(seq_1, max_seq_length = 300,
zero_pad = False, include_CLS_token = True, include_SEP_token = True):
## Tokenzine Input
tokens_a = tokenizer.tokenize(seq_1)
## Truncate
if len(tokens_a) > max_seq_length - 2:
tokens_a = tokens_a[0:(max_seq_length - 2)]
## Initialize Tokens
tokens = []
if include_CLS_token:
tokens.append(tokenizer.cls_token)
## Add Tokens and separators
for token in tokens_a:
tokens.append(token)
if include_SEP_token:
tokens.append(tokenizer.sep_token)
# adding sep token to end and padding up to max_seq_length - 1
input_ids = tokenizer.convert_tokens_to_ids(tokens)
## Input Mask
input_mask = [1] * len(input_ids)
## Zero-pad sequence length
# zero_pad must be set to true for batch size > 1 per https://discuss.pytorch.org/t/dataloaders-problem-with-batch-size-for-customized-data-in-nlp/33572
if zero_pad:
while len(input_ids) < max_seq_length:
input_ids.append(pad_char)
input_mask.append(pad_char)
return torch.tensor(input_ids).unsqueeze(0), input_mask
# + [markdown] id="ve5joHp4r5i4" colab_type="text"
# ## Dataset Loader Classes
# + id="zhwgCIHPr-ND" colab_type="code" colab={}
class Intents(Dataset):
def __init__(self, dataframe):
self.len = len(dataframe)
self.data = dataframe
def __getitem__(self, index):
sentence = self.data.sentence[index]
label = self.data.label[index]
X, _ = prepare_features(sentence)
y = label_to_ix[self.data.label[index]]
return X, y
def __len__(self):
return self.len
# + id="24D1UoxhsMHZ" colab_type="code" colab={}
training_set = Intents(mpqa_train)
testing_set = Intents(mpqa_test)
# + [markdown] id="opbAhvgcsMwO" colab_type="text"
# ## Training Parameters
# + id="-2s8v845sPHA" colab_type="code" colab={}
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = model.cuda()
# + id="Qg78mQ_7spHc" colab_type="code" colab={}
# Parameters
params = {'batch_size': 1,
'shuffle': True,
'drop_last': False,
'num_workers': 1}
# + id="wEV44ZmSssTL" colab_type="code" colab={}
training_loader = DataLoader(training_set, **params)
testing_loader = DataLoader(testing_set, **params)
# + id="ftlW8mRLsuWF" colab_type="code" colab={}
loss_function = nn.CrossEntropyLoss()
learning_rate = 1e-05
optimizer = optim.Adam(params = model.parameters(), lr=learning_rate)
# + id="c1wyr6YYswRc" colab_type="code" outputId="b5e55b71-44de-4d25-e347-e181c2afa656" executionInfo={"status": "ok", "timestamp": 1574220978903, "user_tz": 300, "elapsed": 79785, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
## Test Forward Pass
inp = training_set.__getitem__(0)[0].cuda()
output = model(inp)[0]
print(output.shape)
# + id="7gZX7XnuoGBo" colab_type="code" colab={}
loss_arr =[]
acc_arr = []
max_epochs = 4
iteration_evaluation_intervals = 1000
# + id="XFql5IcHsy-W" colab_type="code" outputId="4ee8509f-1ed7-439d-a497-550affb492a4" executionInfo={"status": "ok", "timestamp": 1574224384130, "user_tz": 300, "elapsed": 3397330, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 748, "referenced_widgets": ["128b64c3543043e5896e8bf1a234f237"]}
# %%time
model = model.train()
for epoch in tqdm_notebook(range(max_epochs)):
print("EPOCH -- {}".format(epoch))
loss_arr.append([])
acc_arr.append([])
for i, (sent, label) in enumerate(training_loader):
optimizer.zero_grad()
sent = sent.squeeze(0)
if torch.cuda.is_available():
sent = sent.cuda()
label = label.cuda()
output = model.forward(sent)[0]
_, predicted = torch.max(output, 1)
loss = loss_function(output, label)
loss.backward()
optimizer.step()
if i%iteration_evaluation_intervals == 0:
correct = 0
total = 0
for sent, label in testing_loader:
sent = sent.squeeze(0)
if torch.cuda.is_available():
sent = sent.cuda()
label = label.cuda()
output = model.forward(sent)[0]
_, predicted = torch.max(output.data, 1)
total += label.size(0)
correct += (predicted.cpu() == label.cpu()).sum()
accuracy = 100.00 * correct.numpy() / total
print('Iteration: {}. Loss: {}. Accuracy: {}%'.format(i, loss.item(), accuracy))
loss_arr[-1].append(loss.item())
acc_arr[-1].append(accuracy)
if accuracy > 85:
break
# + [markdown] id="Sv-diHqH0ezE" colab_type="text"
# ### Baselines
# First check what the accuracy would be if we labeled everything the majority label.
# + id="x7atE9cc0vv0" colab_type="code" colab={}
cbert_naive = np.sum(cbert_test[['label']]) / len(cbert_test[['label']])
mpqa_train_naive = np.sum(mpqa_train[['label']]) / len(mpqa_train[['label']])
mpqa_test_naive = np.sum(mpqa_test[['label']]) / len(mpqa_test[['label']])
cornell_naive = np.sum(cornell_test[['label']]) / len(cornell_test[['label']])
# + id="pAVKjFr91q6Y" colab_type="code" outputId="51b44462-b843-4168-a551-273654f65f3c" executionInfo={"status": "ok", "timestamp": 1574224630375, "user_tz": 300, "elapsed": 1065, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
mpqa_train_naive, mpqa_test_naive, cbert_naive, cornell_naive
# + [markdown] id="B9AyUwdfAZc7" colab_type="text"
# ##Graphs
# + id="voD7yga3Acrb" colab_type="code" colab={}
epoch_colors = ["#deebf7", "#9ecae1", "#4292c6", "#08519c"]
def saveGraph(vals, is_loss=False, split_epoch=False, dataset="mpqa"):
if split_epoch:
x_vals = [iteration_evaluation_intervals * x for x in range(len(vals[0]))]
for i, y_vals in enumerate(vals):
plt.plot(x_vals, y_vals, color=epoch_colors[i], label='epoch {}'.format(i))
else:
x_vals = [iteration_evaluation_intervals * x for x in range(len(vals) * len(vals[0]))]
y_vals = [elt for arr in vals for elt in arr]
plt.plot(x_vals, y_vals)
title = "RoBERTa Test Acccuracy (" + dataset + ")"
save_label = "roberta_acc"
plt.ylabel("test accuracy")
if is_loss:
plt.ylabel("training loss")
title = "RoBERTa Training Loss (" + dataset + ")"
save_label = "roberta_loss"
title += "\n" + str(max_epochs) + " Epochs, batch size = 1, lr = " + str(learning_rate)
plt.title(title)
plt.xlabel("iteration")
save_label += "_" + dataset + "_" + str(max_epochs) + "epochs"
if split_epoch:
save_label += "_split"
plt.legend()
save_label += ".png"
print(save_label)
plt.savefig(save_label)
# + id="B1m-W5bJTTE4" colab_type="code" outputId="c54ec4b4-06b7-4ec0-dee1-616552523f3b" executionInfo={"status": "ok", "timestamp": 1574225392819, "user_tz": 300, "elapsed": 1659, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 327}
saveGraph(loss_arr, is_loss=True, split_epoch=True)
# + id="-DTiKeYpTTWg" colab_type="code" outputId="ea1f60e3-f3c6-4cd2-c893-87b5149479e2" executionInfo={"status": "ok", "timestamp": 1574225404167, "user_tz": 300, "elapsed": 1526, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 327}
saveGraph(loss_arr, is_loss=True, split_epoch=False)
# + id="A3v_0I-GTTwG" colab_type="code" outputId="9159c29f-7d45-4727-d044-cc21f79479b7" executionInfo={"status": "ok", "timestamp": 1574225418560, "user_tz": 300, "elapsed": 1917, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 327}
saveGraph(acc_arr, is_loss=False, split_epoch=True)
# + id="ktHDyfrRWHjq" colab_type="code" outputId="1578a7ee-ed0e-4825-d8da-7880b714bbb5" executionInfo={"status": "ok", "timestamp": 1574225427463, "user_tz": 300, "elapsed": 1340, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 327}
saveGraph(acc_arr, is_loss=False, split_epoch=False)
# + [markdown] id="IeGp-u5cW_zl" colab_type="text"
# ### Cross Data Accuracy
# + id="hJvHMlyTXewz" colab_type="code" colab={}
def evaluateModel(data_loader):
correct = 0
total = 0
for sent, label in data_loader:
sent = sent.squeeze(0)
if torch.cuda.is_available():
sent = sent.cuda()
label = label.cuda()
output = model.forward(sent)[0]
_, predicted = torch.max(output.data, 1)
total += label.size(0)
correct += (predicted.cpu() == label.cpu()).sum()
accuracy = 100.00 * correct.numpy() / total
print('Accuracy: {}%'.format(accuracy))
return accuracy
# + id="OclgGZpFXDMn" colab_type="code" colab={}
cbert_testDataLoader = DataLoader(Intents(cbert_test), **params)
cornell_testDataLoader = DataLoader(Intents(cornell_test), **params)
# + id="OCwTjq8FXDj5" colab_type="code" outputId="bf2467f1-464b-4bb1-d9a5-f8727d0822db" executionInfo={"status": "ok", "timestamp": 1574225517812, "user_tz": 300, "elapsed": 18195, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
cbert_acc = evaluateModel(cbert_testDataLoader)
# + id="wtB3oUF8X7uQ" colab_type="code" outputId="7ffcad78-d807-414a-9358-5af5988f9e46" executionInfo={"status": "ok", "timestamp": 1574225555472, "user_tz": 300, "elapsed": 34951, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
cornell_acc = evaluateModel(cornell_testDataLoader)
# + id="fRzK6638zuvU" colab_type="code" outputId="805c486c-e24b-4e71-a0e9-4fbfeb3ffa42" executionInfo={"status": "ok", "timestamp": 1574225580753, "user_tz": 300, "elapsed": 18533, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
final_test_acc = evaluateModel(testing_loader)
# + id="sHVA3mitz2Bh" colab_type="code" outputId="d123aabb-3982-4b1e-af9f-c1e06abec789" executionInfo={"status": "ok", "timestamp": 1574225723087, "user_tz": 300, "elapsed": 139051, "user": {"displayName": "e l", "photoUrl": "", "userId": "17360354320395717431"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
final_train_acc = evaluateModel(training_loader)
# + [markdown] id="mW8J4rB3s-6K" colab_type="text"
# ## Load model
# + id="Vad17Wx10IrI" colab_type="code" colab={}
torch.save(model.cpu().state_dict(), "roBERTa_mpqa_data_4epochs.pth")
# + id="bdTEpN1CKl8t" colab_type="code" colab={}
| Roberta_mqpa_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.11 ('base')
# language: python
# name: python3
# ---
# Sometimes, scipy.optimize.root does not perform well enough, or you simply want a jacobian function
# +
import numpy as np
import itertools
def myfunc(x):
rows,cols = x.shape
eqs = np.zeros((rows,cols))
eqs[:,0] = x[:,2]-x[:,0]
eqs[:,1:-1] = x[:,2::]+x[:,0:-2]-2*x[:,1:-1]
eqs[:,-1] = x[:,-1]-x[:,-3]
return eqs
def jacobian(func, x, args=None, delta=1e-6):
"""
Returns 2d jacobian of 1 or 2 dimensional function
Can we do 3d jacobian of 2d problem?
"""
dims = x.shape
ndims = len(dims)
npoints = np.prod(dims)
jac = np.zeros((npoints, npoints))
if args is None:
args = ()
if ndims==1:
dx = np.eye(npoints) * delta
for j in range(npoints):
jac[:,j] = ( func(x+dx[j,:], *args) - func(x-dx[j,:], *args) ) / (2*delta)
elif ndims>1:
Z = np.zeros_like(x)
diffpoints = [i for i in itertools.product(*[range(dim) for dim in dims])]
cunts = [i for i,_ in enumerate(diffpoints)]
for cunt,point in zip(cunts,diffpoints):
dx = Z.copy()
dx[point] += delta
dF = (func(x+dx, *args) - func(x-dx, *args) ) / (2*delta)
jac[:,cunt] = dF.ravel()
return jac
x = np.random.random((2,11))
jacobian(myfunc, x)
# -
| numerikk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem setup:
#
# The last step in most machine learning problems is to tune a model with a grid search. However, you have to be careful how you evaluate the results of the search.
# +
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import KFold
from sklearn.ensemble import GradientBoostingRegressor
from scipy.stats import randint
import numpy as np
# Load the data
X, y = load_boston(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Define (roughly) our hyper parameters
hyper = {
'max_depth': randint(3, 10),
'n_estimators': randint(25, 250),
'learning_rate': np.linspace(0.001, 0.01, 20),
'min_samples_leaf': [1, 5, 10]
}
# Define our CV class (remember to always shuffle!)
cv = KFold(shuffle=True, n_splits=3, random_state=1)
# Define our estimator
search = RandomizedSearchCV(GradientBoostingRegressor(random_state=42),
scoring='neg_mean_squared_error', n_iter=25,
param_distributions=hyper, cv=cv,
random_state=12, n_jobs=4)
# Fit the grid search
search.fit(X_train, y_train)
# -
# Now we want to know if the model is good enough. __Does this model meet business requirements?__
#
# ## Wrong approach:
#
# If you repeatedly expose your model to your test set, you risk "p-hacking":
# +
from sklearn.metrics import mean_squared_error
# Evaluate:
print("Test MSE: %.3f" % mean_squared_error(y_test, search.predict(X_test)))
# -
# This is the wrong approach since you've now gained information that could cause model leakage. If you decide to make adjustments to your model to improve the test score, you're effectively fitting the test set indirectly.
#
# The more appropriate approach is to examine the CV scores of the model.
#
# ## Better approach:
# +
import pandas as pd
pd.DataFrame(search.cv_results_)\
.sort_values('mean_test_score',
# descend since neg MSE
ascending=False)\
.head()
# -
# ## CV outside scope of grid search:
#
# You typically don't go straight into a grid search. First, you try several models. Scikit allows us to fit a model in the context of cross validation and examine the fold scores. This
# is useful for determining whether a model will perform in the ballpark of business requirements before a lengthy tuning process:
# +
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
# Set our CV seed
cv = KFold(n_splits=3, random_state=0, shuffle=True)
# Fit and score a model in CV:
cross_val_score(GradientBoostingRegressor(random_state=42),
X_train, y_train, cv=cv, scoring='neg_mean_squared_error')
# -
| code/2018-08-23-data-dredging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dapopov-st/python-youtube-code/blob/master/C3_W4_Lab_3_Fairness_Indicators.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YwpE8n4v_JeT"
# # Ungraded Lab: Fairness Indicators
#
# In this colab notebook, you will use [Fairness Indicators](https://www.tensorflow.org/tfx/guide/fairness_indicators) to explore the `Smiling` attribute in a large-scale face image dataset. Fairness Indicators is a suite of tools built on top of [TensorFlow Model Analysis](https://www.tensorflow.org/tfx/model_analysis/get_started) that enable regular evaluation of fairness metrics in product pipelines. This [Introductory Video](https://www.youtube.com/watch?v=pHT-ImFXPQo) provides more details and context on the real-world scenario presented here, one of primary motivations for creating Fairness Indicators. This notebook will teach you to:
#
#
# * Train a simple neural network model to detect a person's smile in images using [TF Keras](https://www.tensorflow.org/guide/keras) and the [CelebFaces Attributes (CelebA)](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) dataset.
# * Evaluate model performance against a commonly used fairness metric across age groups using Fairness Indicators.
#
# *Credits: Some of the code and discussions are taken from this [Tensorflow tutorial](https://colab.research.google.com/github/tensorflow/fairness-indicators/blob/master/g3doc/tutorials/Fairness_Indicators_TFCO_CelebA_Case_Study.ipynb).*
# + [markdown] id="QTvlORA3TuTz"
# ## Install Fairness Indicators
#
# This will install all related libraries such as TFMA and TFDV.
# + id="t1ZWcTAvoi_o"
# !pip install fairness-indicators
# + [markdown] id="joQr8PWXUWon"
# *Note: In Google Colab, you need to restart the runtime at this point to finalize updating the packages you just installed. You can do so by clicking the `Restart Runtime` at the end of the output cell above (after installation), or by selecting `Runtime > Restart Runtime` in the Menu bar. **Please do not proceed to the next section without restarting.** You can also ignore the errors about version incompatibility of some of the bundled packages because we won't be using those in this notebook.*
# + [markdown] id="fiqohiU73cRf"
# ## Import packages
#
# Next, you will import the main packages and some utilities you will need in this notebook. Notice that you are not importing `fairness-indicators` directly. As mentioned in the intro, this suite of tools is built on top of TFMA so you can just import TFMA to access it.
# + id="A66hFOyMorfQ"
import tensorflow as tf
import tensorflow_model_analysis as tfma
import tensorflow_datasets as tfds
from tensorflow import keras
# + [markdown] id="LwJ63Ay0WUjP"
# The code below should not produce any error. Otherwise, please restart the installation.
# + id="WzZNQrUCpEs6"
print("TensorFlow " + tf.__version__)
print("TFMA " + tfma.VERSION_STRING)
# + [markdown] id="aLzC_ZvSEM_C"
# ## Download and prepare the dataset
#
# [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) is a large-scale face attributes dataset with more than 200,000 celebrity images, each with 40 attribute annotations (such as hair type, fashion accessories, facial features, etc.) and 5 landmark locations (eyes, mouth and nose positions). For more details, you can read more in [this paper](https://liuziwei7.github.io/projects/FaceAttributes.html).
#
# With the permission of the owners, this dataset is stored on Google Cloud Storage (GCS) and mostly accessed via [TensorFlow Datasets(`tfds`)](https://www.tensorflow.org/datasets). To save on download time and disk space, you will use the GCS bucket specified below as your data directory. This already contains the TFRecords. If you want to download it to your workspace, you can pass a local directory to the `data_dir` argument. Just take note that it will take time to complete the download.
#
#
#
# + id="_tRPCjI9YxKN"
# URI of the dataset in Goocle Cloud Storage
GCS_BASE_DIR = "gs://celeb_a_dataset/"
# Load the data using TFDS
data, data_info = tfds.load("celeb_a", data_dir=GCS_BASE_DIR, with_info=True, builder_kwargs={'version':'2.0.0'})
# + [markdown] id="5c8Rsf-WamtK"
# You can preview some of the images in the dataset.
# + id="8r0ZMW3_ZnZU"
# Take 6 examples and preview images
fig = tfds.show_examples(data['train'].take(6), data_info)
# + [markdown] id="Nt8Ahn9fbBTh"
# You can also view the dataset as a dataframe to preview the other attributes in tabular format.
# + id="S7Ndy-sKbpbk"
# Take 4 examples as a dataframe
df = tfds.as_dataframe(data['train'].take(4), data_info)
# View the dataframe
df.head()
# + [markdown] id="NqoM3nazby8C"
# Let's list the column header so you can see the attribute names in the dataset. For this notebook, you will just examine the `attributes/Young` and `attributes/Smiling` features but feel free to pick other features once you've gone over the whole exercise.
# + id="icDbLpkaGN0S"
# List dataframe header
df.columns
# + [markdown] id="BO5Ld9oOYAvZ"
# In this notebook:
# * Your model will attempt to classify whether the subject of the image is smiling, as represented by the `Smiling` attribute<sup>*</sup>.
# * Images will be resized from 218x178 to 28x28 to reduce the execution time and memory when training.
# * Your model's performance will be evaluated across age groups, using the binary `Young` attribute. You will call this "age group" in this notebook.
#
# ___
#
# <sup>*</sup> *While there is little information available about the labeling methodology for this dataset, you will assume that the "Smiling" attribute was determined by a pleased, kind, or amused expression on the subject's face. For the purpose of this example, you will take these labels as ground truth.*
# + [markdown] id="1uzyvjt8EyLT"
# ### Caveats
# Before moving forward, there are several considerations to keep in mind when using CelebA:
# * Although, in principle, this notebook could use any dataset of face images, CelebA was chosen because it contains public domain images of public figures.
# * All of the attribute annotations in CelebA are operationalized as binary categories. For example, the `Young` attribute (as determined by the dataset labelers) is denoted as either present or absent in the image.
# * CelebA's categorizations do not reflect real human diversity of attributes.
# * For the purposes of this notebook, the feature containing the `Young` attribute is referred to as "age group". A `True` will put the image as a member of the `Young` age group and a `False` will put the image as a member of the `Not Young` age group. These are assumptions made as this information is not mentioned in the [original paper](http://openaccess.thecvf.com/content_iccv_2015/html/Liu_Deep_Learning_Face_ICCV_2015_paper.html).
# * As such, performance in the models trained in this notebook is tied to the ways the attributes have been operationalized and annotated by the authors of CelebA.
# * This model should not be used for commercial purposes as that would violate [CelebA's non-commercial research agreement](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html).
# + [markdown] id="6yDEIn3cE6Uo"
# ### Setting Up Input Functions
#
# Now, you will define the preprocessing functions to prepare your data as model inputs. These include resizing images, normalizing pixels, casting to the right data type, and grouping the features and labels.
# + id="3PSk2eUUx9L8"
# Define Constants
ATTR_KEY = "attributes"
IMAGE_KEY = "image"
LABEL_KEY = "Smiling"
GROUP_KEY = "Young"
IMAGE_SIZE = 28
# Define Preprocessing Function
def preprocess_input_dict(feat_dict):
''' Picks the attributes to study and resizes the images
Args:
feat_dict (dictionary): features from the dataset
Returns:
dictionary containing the resized image, label, and age group
'''
# Separate out the image and target variable from the feature dictionary.
image = feat_dict[IMAGE_KEY]
label = feat_dict[ATTR_KEY][LABEL_KEY]
group = feat_dict[ATTR_KEY][GROUP_KEY]
# Resize and normalize image.
image = tf.cast(image, tf.float32)
image = tf.image.resize(image, [IMAGE_SIZE, IMAGE_SIZE])
image /= 255.0
# Cast label and group to float32.
label = tf.cast(label, tf.float32)
group = tf.cast(group, tf.float32)
# Put the computed values in a dictionary
feat_dict[IMAGE_KEY] = image
feat_dict[ATTR_KEY][LABEL_KEY] = label
feat_dict[ATTR_KEY][GROUP_KEY] = group
return feat_dict
# Define lambda functions to group features and labels for training and evaluation
get_image_and_label = lambda feat_dict: (feat_dict[IMAGE_KEY], feat_dict[ATTR_KEY][LABEL_KEY])
get_image_label_and_group = lambda feat_dict: (feat_dict[IMAGE_KEY], feat_dict[ATTR_KEY][LABEL_KEY], feat_dict[ATTR_KEY][GROUP_KEY])
# + [markdown] id="IQOsJKPshhmJ"
# ## Prepare train and test splits
#
# This next helper function will help split, shuffle, batch and preprocess your training data. For this notebook, you will just develop a model that accepts the image as input and outputs the `Smiling` attribute (i.e. label).
# + id="-CfC6ZF4pc7I"
def celeb_a_train_data_wo_group(data, batch_size):
'''
Args:
data (TF dataset) - dataset to preprocess
batch_size (int) - batch size
Returns:
Batches of preprocessed datasets containing tuples with (image, label)
'''
celeb_a_train_data = data.shuffle(1024).repeat().batch(batch_size).map(preprocess_input_dict)
return celeb_a_train_data.map(get_image_and_label)
# + [markdown] id="GZYWhFB-lal-"
# The `test` split does not need to be shuffled so you can just preprocess it like below.
# + id="hLkOPoVwdhKh"
# Prepare test data
celeb_a_test_data = data['test'].batch(1).map(preprocess_input_dict).map(get_image_label_and_group)
# + [markdown] id="xcQwGY4JnsT6"
# As a sanity check, you can examine the contents of a one example in the test data. You should see that it is successfully reshaped and the pixels should be normalized.
# + id="svHIiXWPmfy4"
# Print info about the test data records
for sample in celeb_a_test_data.take(1):
print(f'Data type: {type(sample)}')
print(f'Number of elements: {len(sample)}')
print(f'Shape of 1st element: {sample[0].shape}')
print(f'Shape of 2nd element: {sample[1].shape}')
print(f'Shape of 3rd element: {sample[2].shape}')
print(f'Contents: \n{sample}')
# + [markdown] id="bZ7SCKIEF1IC"
# ## Build a simple DNN Model
#
# With the dataset prepared, you will now assemble a simple `tf.keras.Sequential` model to classify your images. The model consists of:
#
# 1. An input layer that represents the flattened 28x28x3 image.
# 2. A fully connected layer with 64 units activated by a ReLU function.
# 3. A single-unit readout layer to output real-scores instead of probabilities.
#
# You may be able to greatly improve model performance by adding some complexity (e.g., more densely-connected layers, exploring different activation functions, increasing image size), but that may distract from the goal of demonstrating how easy it is to apply the indicators when working with Keras. For that reason, you will first keep the model simple — but feel free to explore this space later.
# + id="tpNlbyyapfbL"
def create_model():
'''Builds the simple DNN binary classifier'''
# Build the model using the Sequential API
model = keras.Sequential([
keras.layers.Flatten(input_shape=(IMAGE_SIZE, IMAGE_SIZE, 3), name='image'),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(1, activation=None)
])
# Compile the model with hinge loss and binary accuracy metric
model.compile(
optimizer=tf.keras.optimizers.Adam(0.001),
loss='hinge',
metrics='binary_accuracy')
return model
# + [markdown] id="zLVPrZGYGwiI"
# ## Train & Evaluate Model
#
# You’re now ready to train your model. To cut back on the amount of execution time and memory, you will train the model by slicing the data into small batches with only a few repeated iterations.
# + id="spMUm9wNp0nW"
BATCH_SIZE = 32
# Build the model
model = create_model()
# Train the model
model.fit(celeb_a_train_data_wo_group(data['train'], BATCH_SIZE), epochs=5, steps_per_epoch=1000)
# + [markdown] id="CgYTmRdcHYOK"
# Evaluating the model on the test data should result in a final accuracy score of just over 85%. Not bad for a simple model with no fine tuning.
# + id="tlPUI-ovqL-F"
# Evaluate trained model on the test data
results = model.evaluate(celeb_a_test_data)
# + [markdown] id="HKdvynqKrWfn"
# You will then save the model so you can analyze it in the next section.
# + id="Jk06C_O3SIkx"
# Define model directory
MODEL_LOCATION = 'saved_model'
# Save the model
model.save(MODEL_LOCATION, save_format='tf')
# + [markdown] id="WGJl6pJeGYzD"
# ## Model Analysis
#
# As you already know, it is usually not enough to just measure your model's performance on global metrics. For instance, performance evaluated across age groups may reveal some shortcomings.
#
# To explore this further, you will evaluate the model with Fairness Indicators via TFMA. In particular, you will see whether there is a significant gap in performance between "Young" and "Not Young" categories when evaluated on false positive rate (FPR).
#
# A false positive error occurs when the model incorrectly predicts the positive class. In this context, a false positive outcome occurs when the ground truth is an image of a celebrity 'Not Smiling' and the model predicts 'Smiling'. While this seems like a relatively mundane error, false positive errors can sometimes cause more problematic behaviors when deployed in a real world application. For instance, a false positive error in a spam classifier could cause a user to miss an important email.
#
# You will mostly follow the same steps as you did in the first ungraded lab of this week. Namely, you will:
#
# * Create a TFRecord of the test dataset.
# * Write an `EvalConfig` file
# * Create an `EvalSharedModel`
# * Define a `Schema` message
# * Run model analysis with TFMA
# + [markdown] id="kuPDsIdmClFa"
# ### Create TFRecord
#
# You will need to serialize the preprocessed test dataset so it can be read by TFMA. We've provided a helper function to do just that. Notice that the age group feature is transformed into a string ('Young' or 'Not Young'). This will come in handy in the visualization so the tags are easier to interpret (compared to just 1 or 0).
# + id="-zrBYHAFteT6"
# Define filename
TFRECORD_FILE = 'celeb_a_test.tfrecord'
# + id="LY1QtzKBtG7p"
def celeb_ds_to_tfrecord(dataset, tfrecord_file):
''' Helper function to convert a TF Dataset to TFRecord
Args:
dataset (TF Dataset) - dataset to save as TFRecord
tfrecord_file (string) - filename to use when writing the TFRecord
'''
# Initialize examples list
examples = []
for row in dataset:
# Get image, label, and group tensors
image = row[0]
label = row[1]
group = row[2]
# Flatten image
image = tf.reshape(image, [-1])
# Instantiate Example
output = tf.train.Example()
# Assign features' numpy arrays to the Example feature values
output.features.feature[IMAGE_KEY].float_list.value.extend(image.numpy().tolist())
output.features.feature[LABEL_KEY].float_list.value.append(label.numpy())
output.features.feature[GROUP_KEY].bytes_list.value.append(b"Young" if group.numpy() else b'Not Young')
# Append to examples list
examples.append(output)
# Serialize examples and save as tfrecord
with tf.io.TFRecordWriter(tfrecord_file) as writer:
for example in examples:
writer.write(example.SerializeToString())
# + id="hPsF2Uu3uh7U"
# Use the helper function to serialize the test dataset
celeb_ds_to_tfrecord(celeb_a_test_data, TFRECORD_FILE)
# + [markdown] id="KYEFWyrZHqyV"
# ### Write EvalConfig file
#
# Next, you will define the model, metrics, and slicing specs in an eval config file. As mentioned, you will slice the data across age groups to see if there is an underlying problem. For metrics, you will include the `FairnessIndicators` class. These are commonly-identified fairness metrics for binary and multiclass classifiers. Moreover, you will configure a list of thresholds. These will allow you to observe if the model predicts better when the threshold to determine between the two classes is changed (e.g. will the FPR be lower if the model predicts "Smiling" for outputs greater than 0.22?).
#
# + id="KNM8_wn1S5-P"
# Import helper module
from google.protobuf import text_format
# Write EvalConfig string
eval_config_pbtxt = """
model_specs {
label_key: "%s"
}
metrics_specs {
metrics {
class_name: "FairnessIndicators"
config: '{ "thresholds": [0.22, 0.5, 0.75] }'
}
metrics {
class_name: "ExampleCount"
}
}
slicing_specs {}
slicing_specs { feature_keys: "%s" }
""" % (LABEL_KEY, GROUP_KEY)
# Parse as a Message
eval_config = text_format.Parse(eval_config_pbtxt, tfma.EvalConfig())
# + [markdown] id="iN-67hysIycZ"
# ### Create EvalSharedModel
#
# This will be identical to the command you ran in an earlier lab. This is needed so TFMA will know how to load and configure your model from disk.
# + id="_3cBZNlOvGC1"
# Create EvalSharedModel
eval_shared_model = tfma.default_eval_shared_model(
eval_saved_model_path=MODEL_LOCATION, eval_config=eval_config)
# + [markdown] id="5DOHO4VoJ99b"
# ### Create a Schema
#
# This is an additional step from your previous TFMA workflow. It is needed particularly because, unlike the TFMA ungraded lab, you didn't include a serving signature with the model. If you remember, the function called by that signature took care of parsing the tfrecords, converting them to the correct data type, and preprocessing. Since that part is not included in this lab, you will need to provide a schema so TFMA will know what data types are in the serialized examples when it parses the tfrecord into a dictionary of features. You will also need to define the dimensions of the image since that is expected by your model input. That is handled by the `tensor_representation_group` below.
# + id="BtiJuufKvDV8"
from tensorflow_metadata.proto.v0 import schema_pb2
from google.protobuf import text_format
# Define Schema message as string
schema_pbtxt = """
tensor_representation_group {
key: ""
value {
tensor_representation {
key: "%s"
value {
dense_tensor {
column_name: "%s"
shape {
dim { size: 28 }
dim { size: 28 }
dim { size: 3 }
}
}
}
}
}
}
feature {
name: "%s"
type: FLOAT
}
feature {
name: "%s"
type: FLOAT
}
feature {
name: "%s"
type: BYTES
}
""" % (IMAGE_KEY, IMAGE_KEY, IMAGE_KEY, LABEL_KEY, GROUP_KEY)
# Parse the schema string to a message
schema = text_format.Parse(schema_pbtxt, schema_pb2.Schema())
# + [markdown] id="N8GUBgybKq39"
# ### Run TFMA
#
# You will pass the objects you created in the previous sections to `tfma.run_model_analysis()`. As you've done previously, this will take care of loading the model and data, and computing the metrics on the data slices you specified.
# + id="dbpqTuHbTKFH"
# Define output directory
OUTPUT_PATH = 'tfma_output'
# Run model analysis
eval_results = tfma.run_model_analysis(
eval_shared_model=eval_shared_model,
eval_config=eval_config,
data_location=TFRECORD_FILE,
schema=schema,
output_path=OUTPUT_PATH
)
# + [markdown] id="i5BvsF8OH3CE"
# Now you can view the fairness metrics you specified. The FPR should already be selected and you can see that it is considerably higher for the `Not Young` age group. Try to explore the widget and see if you can make other findings. Here are some suggestions:
#
# * Toggle the threshold drop down and see how the FPR changes with different thresholds
#
# * Change the baseline to `Young: Young` so the percentage difference (in the table below the chart) will be measured against it.
#
# * Deselect the `Overall` slice so you can compare the two age groups side by side.
#
# * Select other metrics to display and observe their charts.
# + id="Q0R5OlNDqWwS"
# Visualize the fairness metrics
tfma.addons.fairness.view.widget_view.render_fairness_indicator(eval_results)
# + [markdown] id="Pr5DrvGZO2B6"
# After studying the discrepancies in your predictions, you can then investigate why that happens and have a plan on remidiating it. Aside from changing your model architecture, you can also look first at your training data. `fairness-indicators` is also packaged with TFDV so you can use it to generate statistics from your data. Here is a short review on how to do that.
# + id="Rd-oUa1z-yjQ"
import tensorflow_data_validation as tfdv
# Define training directory
TRAIN_DIR = f'{GCS_BASE_DIR}celeb_a/2.0.0/celeb_a-train.tfrecord*'
# View tfrecord filenames in GCS
# !gsutil ls {TRAIN_DIR}
# + id="xGteF9i1FyhA"
# Filter features to observe
stats_options = tfdv.StatsOptions(feature_allowlist=['attributes/Young'])
# Compute the statistics
statistics = tfdv.generate_statistics_from_tfrecord(TRAIN_DIR, stats_options=stats_options)
# Visualize the statistics
tfdv.visualize_statistics(statistics)
# + [markdown] id="q6huPG-SQNyV"
# The statistics show that the `Not Young` age group (i.e. `0` in the `attributes/Young` column) has very few images compared to the `Young` age group. Maybe that's why the model learns on the `Young` images better. You could try adding more `Not Young` images and see if your model performs better on this slice.
# + [markdown] id="6hhF68-0Q0nZ"
# ## Wrap Up
#
# In this lab, you prepared an image dataset and trained a model to predict one of its attributes (i.e. `Smiling`). You then sliced the data based on age groups and computed fairness metrics from the `Fairness Indicators` package via TFMA. Though the outcome looks simple, it is an important step in production ML projects because not detecting these problems can greatly affect the experience of your users. Improving these metrics will help you commit to fairness in your applications. We encourage you to try exploring more slices of the dataset and see what findings you can come up with.
#
# For more practice, [here](https://colab.research.google.com/github/tensorflow/fairness-indicators/blob/master/g3doc/tutorials/Fairness_Indicators_Example_Colab.ipynb) is an official tutorial that uses fairness indicators on text data. It uses the [What-If-Tool](https://pair-code.github.io/what-if-tool/) which is another package that comes with `Fairness Indicators`. You will also get to explore that in this week's programming assignment.
| C3_W4_Lab_3_Fairness_Indicators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3
# language: python
# name: python3
# ---
# setup
from mlwpy import *
diabetes = datasets.load_diabetes()
# %matplotlib inline
# +
N = 20
ftr = np.linspace(-10, 10, num=N) # ftr values
tgt = 2*ftr**2 - 3 + np.random.uniform(-2, 2, N) # tgt = func(ftr)
(train_ftr, test_ftr,
train_tgt, test_tgt) = skms.train_test_split(ftr, tgt, test_size=N//2)
display(pd.DataFrame({"ftr":train_ftr,
"tgt":train_tgt}).T)
# -
plt.plot(train_ftr, train_tgt, 'bo')
plt.plot(test_ftr, np.zeros_like(test_ftr), 'r+');
# note: sklearn *really* wants 2D inputs (a table)
# so we use rehape here.
sk_model = linear_model.LinearRegression()
sk_model.fit(train_ftr.reshape(-1,1), train_tgt)
sk_preds = sk_model.predict(test_ftr.reshape(-1,1))
sk_preds[:3]
# fit predict evaluate a 1-D polynomial (a line)
model_one = np.poly1d(np.polyfit(train_ftr, train_tgt, 1))
preds_one = model_one(test_ftr)
print(preds_one[:3])
# +
# the predictions come back the same
print("all close?", np.allclose(sk_preds, preds_one))
# and we can still use sklearn to evaluate it
mse = metrics.mean_squared_error
print("RMSE:", np.sqrt(mse(test_tgt, preds_one)))
# -
# fit predict evaluate a 2-D polynomial (a parabola)
model_two = np.poly1d(np.polyfit(train_ftr, train_tgt, 2))
preds_two = model_two(test_ftr)
print("RMSE:", np.sqrt(mse(test_tgt, preds_two)))
model_three = np.poly1d(np.polyfit(train_ftr, train_tgt, 9))
preds_three = model_three(test_ftr)
print("RMSE:", np.sqrt(mse(test_tgt, preds_three)))
# +
fig, axes = plt.subplots(1,2, figsize=(6,3), sharey=True)
labels = ['line', 'parabola', 'nonic']
models = [model_one, model_two, model_three]
train = (train_ftr, train_tgt)
test = (test_ftr, test_tgt)
for ax, (ftr, tgt) in zip(axes, [train, test]):
ax.plot(ftr, tgt, 'k+')
for m, lbl in zip(models, labels):
ftr = sorted(ftr)
ax.plot(ftr, m(ftr), '-', label=lbl)
axes[1].set_ylim(-20, 200)
axes[0].set_title("Train")
axes[1].set_title("Test");
axes[0].legend(loc='upper center');
# +
results = []
for complexity in [1,2,6,9]:
model = np.poly1d(np.polyfit(train_ftr, train_tgt, complexity))
train_error = np.sqrt(mse(train_tgt, model(train_ftr)))
test_error = np.sqrt(mse(test_tgt, model(test_ftr)))
results.append((complexity, train_error, test_error))
columns = ["Complexity", "Train Error", "Test Error"]
results_df = pd.DataFrame.from_records(results,
columns=columns,
index="Complexity")
results_df
# -
results_df.plot();
# +
def training_loss(loss, model, training_data):
' total training_loss on train_data with model under loss'
return sum(loss(model.predict(x.reshape(1,-1)), y)
for x,y in training_data)
def squared_error(prediction, actual):
' squared error on a single example '
return (prediction-actual)**2
# could be used like:
# my_training_loss = training_loss(squared_error, model, training_data)
# +
knn = neighbors.KNeighborsRegressor(n_neighbors=3)
fit = knn.fit(diabetes.data, diabetes.target)
training_data = zip(diabetes.data, diabetes.target)
my_training_loss = training_loss(squared_error,
knn,
training_data)
print(my_training_loss)
# -
mse = metrics.mean_squared_error(diabetes.target,
knn.predict(diabetes.data))
print(mse*len(diabetes.data))
# +
def complexity(model):
return model.complexity
def cost(model, training_data, loss, _lambda):
return training_loss(m,D) + _lambda * complexity(m)
# -
# data, model, fit & cv-score
model = neighbors.KNeighborsRegressor(10)
skms.cross_val_score(model,
diabetes.data,
diabetes.target,
cv=5,
scoring='neg_mean_squared_error')
# notes:
# defaults for cross_val_score are
# cv=3 fold, no shuffle, stratified if classifier
# model.score by default (regressors: r2, classifiers: accuracy)
iris = datasets.load_iris()
model = neighbors.KNeighborsClassifier(10)
skms.cross_val_score(model, iris.data, iris.target, cv=5)
# +
# not stratified
pet = np.array(['cat', 'dog', 'cat',
'dog', 'dog', 'dog'])
list_folds = list(skms.KFold(2).split(pet))
training_idxs = np.array(list_folds)[:,0,:]
print(pet[training_idxs])
# -
# stratified
# note: typically this is behind the scenes
# making StratifiedKFold produce readable output
# requries some trickery. feel free to ignore.
pet = np.array(['cat', 'dog', 'cat', 'dog', 'dog', 'dog'])
idxs = np.array(list(skms.StratifiedKFold(2)
.split(np.ones_like(pet), pet)))
training_idxs = idxs[:,0,:]
print(pet[training_idxs])
# running non-stratified CV
iris = datasets.load_iris()
model = neighbors.KNeighborsClassifier(10)
non_strat_kf = skms.KFold(5)
skms.cross_val_score(model,
iris.data,
iris.target,
cv=non_strat_kf)
# +
# as a reminder, these are some of the imports
# that are hidden behind: from mlwpy import *
# from sklearn import (datasets, neighbors,
# model_selection as skms,
# linear_model, metrics)
# see Appendix A for details
linreg = linear_model.LinearRegression()
diabetes = datasets.load_diabetes()
scores = []
for r in range(10):
tts = skms.train_test_split(diabetes.data,
diabetes.target,
test_size=.25)
(diabetes_train_ftrs, diabetes_test_ftrs,
diabetes_train_tgt, diabetes_test_tgt) = tts
fit = linreg.fit(diabetes_train_ftrs, diabetes_train_tgt)
preds = fit.predict(diabetes_test_ftrs)
score = metrics.mean_squared_error(diabetes_test_tgt, preds)
scores.append(score)
scores = pd.Series(np.sqrt(sorted(scores)))
df = pd.DataFrame({'RMSE':scores})
df.index.name = 'Repeat'
display(df.T)
# -
ax = plt.figure(figsize=(4,3)).gca()
sns.swarmplot(y='RMSE', data=df, ax=ax)
ax.set_xlabel('Over Repeated\nTrain-Test Splits');
display(df.describe().T)
# +
def tts_fit_score(model, data, msr, test_size=.25):
' apply a train-test split to fit model on data and eval with msr '
tts = skms.train_test_split(data.data,
data.target,
test_size=test_size)
(train_ftrs, test_ftrs, train_tgt, test_tgt) = tts
fit = linreg.fit(train_ftrs, train_tgt)
preds = fit.predict(test_ftrs)
score = msr(test_tgt, preds)
return score
linreg = linear_model.LinearRegression()
diabetes = datasets.load_diabetes()
scores = [tts_fit_score(linreg, diabetes,
metrics.mean_squared_error) for i in range(10)]
print(np.mean(scores))
# +
linreg = linear_model.LinearRegression()
diabetes = datasets.load_diabetes()
# non-default cv= argument
ss = skms.ShuffleSplit(test_size=.25) # default, 10 splits
scores = skms.cross_val_score(linreg,
diabetes.data, diabetes.target,
cv=ss,
scoring='neg_mean_squared_error')
scores = pd.Series(np.sqrt(-scores))
df = pd.DataFrame({'RMSE':scores})
df.index.name = 'Repeat'
display(df.describe().T)
ax = sns.swarmplot(y='RMSE', data=df)
ax.set_xlabel('Over Repeated\nTrain-Test Splits');
# +
ss = skms.ShuffleSplit(test_size=.25, random_state=42)
# note:
# look at the first split (next)
# look at training set
# look at first 10 examples
train, test = 0,1
next(ss.split(diabetes.data))[train][:10]
# -
ss = skms.ShuffleSplit(test_size=.25, random_state=42)
next(ss.split(diabetes.data))[train][:10]
train, test = 0, 1
kf = skms.KFold(5)
next(kf.split(diabetes.data))[train][:10]
kf = skms.KFold(5)
next(kf.split(diabetes.data))[train][:10]
# +
pet = np.array(['cat', 'dog', 'cat',
'dog', 'dog', 'dog'])
kf = skms.KFold(3, shuffle=True)
train, test = 0, 1
split_1_group_1 = next(kf.split(pet))[train]
split_2_group_1 = next(kf.split(pet))[train]
print(split_1_group_1,
split_2_group_1)
# +
kf = skms.KFold(3, shuffle=True, random_state=42)
split_1_group_1 = next(kf.split(pet))[train]
split_2_group_1 = next(kf.split(pet))[train]
print(split_1_group_1,
split_2_group_1)
# +
linreg = linear_model.LinearRegression()
diabetes = datasets.load_diabetes()
loo = skms.LeaveOneOut()
scores = skms.cross_val_score(linreg,
diabetes.data, diabetes.target,
cv=loo,
scoring='neg_mean_squared_error')
scores = pd.Series(np.sqrt(-scores))
df = pd.DataFrame({'RMSE':scores})
df.index.name = 'Repeat'
display(df.describe().T)
ax = sns.swarmplot(y='RMSE', data=df)
ax.set_xlabel('Over LOO\nTrain-Test Splits');
# +
iris = datasets.load_iris()
# 10 data set sizes: 10% - 100%
# (that much data is piped to a 5-fold CV)
train_sizes = np.linspace(.1,1.0,10)
nn = neighbors.KNeighborsClassifier()
(train_N,
train_scores,
test_scores) = skms.learning_curve(nn,
iris.data, iris.target,
cv=5,
train_sizes=train_sizes)
# collapse across the 5 CV scores; one result for each data set size
df = pd.DataFrame(test_scores, index=(train_sizes*100).astype(np.int))
df['Mean 5-CV'] = df.mean(axis='columns')
df.index.name = "% Data Used"
display(df)
# +
neat_sizes = (train_sizes*100).astype(np.int)
labels = np_cartesian_product(neat_sizes, [0,1], np.arange(5))
score = np.concatenate([train_scores.flatten(),
test_scores.flatten()], axis=0)
assert len(score) == len(labels)
df = pd.DataFrame.from_records(labels)
df.columns = ['pct', 'set', 'fold']
df.set = df.set.replace({0:'Train', 1:'Test'})
df['score'] = score
# +
ax = sns.lineplot(x='pct', y='score', hue='set', data=df)
ax.set_title("Learning Curve for 5-NN Classifier")
ax.set_xlabel("Percent of Data used for Training")
ax.set_ylabel("Accuracy");
# -
# tidying the numpy array is a bit of a pain
# xarray is designed to do this "natively" but
# i don't want to introduce that dependency
# [seems like there could be a better broadcasting
# solution to this]
def sk_graph_to_tidy(train_test_scores, # y values
eval_points, # x values
eval_label, # x column name
num_folds): # could be inferred
train_scores, test_scores = train_test_scores
# humph, didn't know np_cartesian was order sensitive
labels = np_cartesian_product(eval_points,
[0,1], # surrogates for train/test
np.arange(num_folds))
score = np.concatenate([train_scores.flatten(),
test_scores.flatten()], axis=0)
df = pd.DataFrame.from_records(labels)
df.columns = [eval_label, 'set', 'fold']
df.set = df.set.replace({0:'Train', 1:'Test'})
df['score'] = score
return df
# +
neat_sizes = (train_sizes*100).astype(np.int)
tidy_df = sk_graph_to_tidy([train_scores, test_scores],
neat_sizes, 'pct', 5)
ax = sns.lineplot(x='pct', y='score', hue='set', data=tidy_df)
ax.set_title("Learning Curve for 5-NN Classifier")
ax.set_xlabel("Percent of Data used for Training")
ax.set_ylabel("Accuracy");
# -
# deprecated in recent seaborn; kept for comparison with above
if False:
joined = np.array([train_scores, test_scores]).transpose()
ax = sns.tsplot(joined,
time=train_sizes,
condition = ['Train', 'Test'],
interpolate=False)
ax.set_title("Learning Curve for 5-NN Classifier")
ax.set_xlabel("Percent of Data used for Training")
ax.set_ylabel("Accuracy");
num_neigh = [1,3,5,10,15,20]
KNC = neighbors.KNeighborsClassifier
tt = skms.validation_curve(KNC(),
iris.data, iris.target,
param_name='n_neighbors',
param_range=num_neigh,
cv=5)
# +
tidy_df = sk_graph_to_tidy(tt, num_neigh, 'k', 5)
ax = sns.lineplot(x='k', y='score', hue='set', data=tidy_df)
ax.set_title('5-fold CV Performance for k-NN')
ax.set_xlabel("\n".join(['k for k-NN',
'lower k, more complex',
'higher k, less complex']))
ax.set_ylim(.9, 1.01)
ax.set_ylabel('Accuracy');
# -
# tsplot deprecated; kept for comparison with above
if False:
# stack and transpose trick (as above)
ax = sns.tsplot(np.array(tt).transpose(),
time=num_neigh,
condition=['Train', 'Test'],
interpolate=False)
ax.set_title('5-fold CV Performance for k-NN')
ax.set_xlabel("\n".join(['k for k-NN',
'lower k, more complex',
'higher k, less complex']))
ax.set_ylim(.9, 1.01)
ax.set_ylabel('Accuracy');
# +
classifiers = {'gnb' : naive_bayes.GaussianNB(),
'5-NN' : neighbors.KNeighborsClassifier(n_neighbors=5)}
iris = datasets.load_iris()
fig, ax = plt.subplots(figsize=(6,4))
for name, model in classifiers.items():
cv_scores = skms.cross_val_score(model,
iris.data, iris.target,
cv=10,
scoring='accuracy',
n_jobs=-1) # use all cores
my_lbl = "{} {:.3f}".format(name, cv_scores.mean())
ax.plot(cv_scores, '-o', label=my_lbl) # marker=next(markers)
ax.set_ylim(0.0, 1.1)
ax.set_xlabel('Fold')
ax.set_ylabel('Accuracy')
ax.legend(ncol=2);
| 05_EvaluatingAndComparingLearners_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from src.utils import *
import re
puzzle_input = parse_puzzle_input(5)
puzzle_input[:3]
sample_input = [
'0,9 -> 5,9',
'8,0 -> 0,8',
'9,4 -> 3,4',
'2,2 -> 2,1',
'7,0 -> 7,4',
'6,4 -> 2,0',
'0,9 -> 2,9',
'3,4 -> 1,4',
'0,0 -> 8,8',
'5,5 -> 8,2'
]
def line_to_coordinates(line_str):
pattern = '([0-9]+),([0-9]+) -> ([0-9]+),([0-9]+)'
re_search = re.search(pattern, line_str)
x1 = re_search.group(1)
y1 = re_search.group(2)
x2 = re_search.group(3)
y2 = re_search.group(4)
return int(x1), int(y1), int(x2), int(y2)
def check_horizontal_or_vertical(x1, y1, x2, y2):
return x1 == x2 or y1 == y2
def get_line_set(puzzle_input, ignore_non_vertical=True):
line_set = set()
for line_str in puzzle_input:
x1, y1, x2, y2 = line_to_coordinates(line_str)
if not ignore_non_vertical or check_horizontal_or_vertical(x1, y1, x2, y2):
line_set.add((x1, y1, x2, y2))
return line_set
def find_line_ranges(line_set):
x_values = [line[0] for line in line_set]
x_values.extend([line[2] for line in line_set])
y_values = [line[1] for line in line_set]
y_values.extend([line[3] for line in line_set])
return min(x_values), max(x_values), min(y_values), max(y_values)
def create_coordinate_dict(min_x, max_x, min_y, max_y):
coordinate_list = []
for x in range(min_x, max_x + 1):
for y in range(min_y, max_y + 1):
coordinate_list.append((x, y))
return {coordinate: 0 for coordinate in coordinate_list}
def find_coordinates_to_update(x1, y1, x2, y2):
coordinates_to_update = []
for x in range(min(x1, x2), max(x1, x2) + 1):
for y in range(min(y1, y2), max(y1, y2) + 1):
coordinates_to_update.append((x, y))
return coordinates_to_update
def update_coordinate_dict(x1, y1, x2, y2, coordinate_dict):
for coordinate in find_coordinates_to_update(x1, y1, x2, y2):
coordinate_dict[coordinate] += 1
return coordinate_dict
def get_final_coordinate_dict(line_set):
min_x, max_x, min_y, max_y = find_line_ranges(line_set)
coordinate_dict = create_coordinate_dict(min_x, max_x, min_y, max_y)
for line in line_set:
x1, y1, x2, y2 = line
coordinate_dict = update_coordinate_dict(x1, y1, x2, y2, coordinate_dict)
return coordinate_dict
def count_overlap(coordinate_dict, min_overlap = 2):
return len([x for x in coordinate_dict.values() if x >= min_overlap])
def count_hydrothermal_vents(puzzle_input, ignore_non_vertical=True, min_overlap = 2):
line_set = get_line_set(puzzle_input, ignore_non_vertical)
coordinate_dict = get_final_coordinate_dict(line_set)
return count_overlap(coordinate_dict, min_overlap)
count_hydrothermal_vents(sample_input)
count_hydrothermal_vents(puzzle_input)
# ## Part 2
def find_direction(a, b):
if a > b:
return -1
elif a < b:
return 1
return 0
def find_coordinates_to_update(x1, y1, x2, y2):
coordinates_to_update = []
if check_horizontal_or_vertical(x1, y1, x2, y2):
for x in range(min(x1, x2), max(x1, x2) + 1):
for y in range(min(y1, y2), max(y1, y2) + 1):
coordinates_to_update.append((x, y))
else:
x_direction = find_direction(x1, x2)
y_direction = find_direction(y1, y2)
coordinates_to_update = list(
zip(
range(x1, x2 + x_direction, x_direction),
range(y1, y2 + y_direction, y_direction)
)
)
return coordinates_to_update
count_hydrothermal_vents(sample_input, ignore_non_vertical=False)
count_hydrothermal_vents(puzzle_input, ignore_non_vertical=False)
| notebooks/day_05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="g1DHUvdAwC-P" colab_type="text"
# # Imports
# + id="DY7MtVPwx912" colab_type="code" outputId="0b74291a-3991-4c6e-ba21-eea420024023" colab={"base_uri": "https://localhost:8080/", "height": 34}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import patches,patheffects
import torch
from torchvision import datasets,transforms,models
from torch import nn
import torch.nn.functional as F
from torch.utils.data import DataLoader , Dataset
import torchvision.transforms.functional as FT
from PIL import Image
import xml.etree.ElementTree as ET
from pathlib import Path
import json
import os
import random
from tqdm import tqdm,trange
import time
from math import sqrt
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(f"{torch.__version__} and {device}")
# + id="Z1S3fcynC9Js" colab_type="code" colab={}
def xy_to_cxcy(xy):
"""
Convert bounding boxes from boundary coordinates (x_min, y_min, x_max, y_max) to center-size coordinates (c_x, c_y, w, h).
:param xy: bounding boxes in boundary coordinates, a tensor of size (n_boxes, 4)
:return: bounding boxes in center-size coordinates, a tensor of size (n_boxes, 4)
"""
return torch.cat([(xy[:, 2:] + xy[:, :2]) / 2, # c_x=(x_max+x_min)/2, similarly for c_y = (y_max+y_min)/2
xy[:, 2:] - xy[:, :2]], 1) # w = x_max-x_min, similarly for h
def cxcy_to_xy(cxcy):
"""
Convert bounding boxes from center-size coordinates (c_x, c_y, w, h) to boundary coordinates (x_min, y_min, x_max, y_max).
:param cxcy: bounding boxes in center-size coordinates, a tensor of size (n_boxes, 4)
:return: bounding boxes in boundary coordinates, a tensor of size (n_boxes, 4)
"""
return torch.cat([cxcy[:, :2] - (cxcy[:, 2:] / 2), # c_x-w=x_min, c_y-h=y_min
cxcy[:, :2] + (cxcy[:, 2:] / 2)], 1) # x_max, y_max
def cxcy_to_gcxgcy(cxcy, priors_cxcy):
"""
Encode bounding boxes (that are in center-size form) w.r.t. the corresponding prior boxes (that are in center-size form).
For the center coordinates, find the offset with respect to the prior box, and scale by the size of the prior box.
For the size coordinates, scale by the size of the prior box, and convert to the log-space.
In the model, we are predicting bounding box coordinates in this encoded form.
:param cxcy: bounding boxes in center-size coordinates, a tensor of size (n_priors, 4)
:param priors_cxcy: prior boxes with respect to which the encoding must be performed, a tensor of size (n_priors, 4)
:return: encoded bounding boxes, a tensor of size (n_priors, 4)
"""
# The 10 and 5 below are referred to as 'variances' in the original Caffe repo, completely empirical
# They are for some sort of numerical conditioning, for 'scaling the localization gradient'
# See https://github.com/weiliu89/caffe/issues/155
return torch.cat([(cxcy[:, :2] - priors_cxcy[:, :2]) / (priors_cxcy[:, 2:] / 10), # g_c_x, g_c_y
torch.log(cxcy[:, 2:] / priors_cxcy[:, 2:]) * 5], 1) # g_w, g_h
def gcxgcy_to_cxcy(gcxgcy, priors_cxcy):
"""
Decode bounding box coordinates predicted by the model, since they are encoded in the form mentioned above.
They are decoded into center-size coordinates.
This is the inverse of the function above.
:param gcxgcy: encoded bounding boxes, i.e. output of the model, a tensor of size (n_priors, 4)
:param priors_cxcy: prior boxes with respect to which the encoding is defined, a tensor of size (n_priors, 4)
:return: decoded bounding boxes in center-size form, a tensor of size (n_priors, 4)
"""
return torch.cat([gcxgcy[:, :2] * priors_cxcy[:, 2:] / 10 + priors_cxcy[:, :2], # c_x, c_y
torch.exp(gcxgcy[:, 2:] / 5) * priors_cxcy[:, 2:]], 1) # w, h
# + id="FwOrD4TIPAwk" colab_type="code" outputId="fac330b0-0444-45d3-dd48-816e20bcce8b" colab={"base_uri": "https://localhost:8080/", "height": 87}
s1=[[0,0,3,4],[0,0,3,4]]
set1=torch.tensor(s1,dtype=torch.float)
cxcy=xy_to_cxcy(set1)
print("Cx_C_y",cxcy)
gcxgcy=cxcy_to_gcxgcy(cxcy,set1)
print("G_cx_G_cy",gcxgcy)
# + id="5QSn3FX7LcIb" colab_type="code" outputId="46a18f69-449e-4c37-ec58-e426703bd31e" colab={"base_uri": "https://localhost:8080/", "height": 122}
s1=[[0,0,3,4],[2,1,3,4]]
set1=torch.tensor(s1,dtype=torch.float)
cxcy=xy_to_cxcy(set1)
print("Cx_C_y",cxcy)
print("xy",cxcy_to_xy(cxcy))
gcxgcy=cxcy_to_gcxgcy(cxcy,set1)
print("G_cx_G_cy",gcxgcy)
# + [markdown] id="EUxtXRyBwK6A" colab_type="text"
# # Jaccard Overlap
# + id="ywkvHv0oxzMh" colab_type="code" colab={}
def find_intersection(set_1, set_2):
"""
Find the intersection of every box combination between two sets of boxes that are in boundary coordinates.
:param set_1: set 1, a tensor of dimensions (n1, 4)
:param set_2: set 2, a tensor of dimensions (n2, 4)
:return: intersection of each of the boxes in set 1 with respect to each of the boxes in set 2, a tensor of dimensions (n1, n2)
"""
# PyTorch auto-broadcasts singleton dimensions
lower_bounds = torch.max(set_1[:, :2].unsqueeze(1), set_2[:, :2].unsqueeze(0)) # (n1, n2, 2)
upper_bounds = torch.min(set_1[:, 2:].unsqueeze(1), set_2[:, 2:].unsqueeze(0)) # (n1, n2, 2)
intersection_dims = torch.clamp(upper_bounds - lower_bounds, min=0) # (n1, n2, 2)
return intersection_dims[:, :, 0] * intersection_dims[:, :, 1] # (n1, n2)
def find_jaccard_overlap(set_1, set_2):
"""
Find the Jaccard Overlap (IoU) of every box combination between two sets of boxes that are in boundary coordinates.
:param set_1: set 1, a tensor of dimensions (n1, 4)
:param set_2: set 2, a tensor of dimensions (n2, 4)
:return: Jaccard Overlap of each of the boxes in set 1 with respect to each of the boxes in set 2, a tensor of dimensions (n1, n2)
"""
# Find intersections
intersection = find_intersection(set_1, set_2) # (n1, n2)
# Find areas of each box in both sets
areas_set_1 = (set_1[:, 2] - set_1[:, 0]) * (set_1[:, 3] - set_1[:, 1]) # (n1)
areas_set_2 = (set_2[:, 2] - set_2[:, 0]) * (set_2[:, 3] - set_2[:, 1]) # (n2)
# Find the union
# PyTorch auto-broadcasts singleton dimensions
union = areas_set_1.unsqueeze(1) + areas_set_2.unsqueeze(0) - intersection # (n1, n2)
return intersection / union # (n1, n2)
# + id="8buqtswvydy9" colab_type="code" outputId="7cccebd6-28ca-4372-e225-fdba43689879" colab={"base_uri": "https://localhost:8080/", "height": 105}
s1=[[0,0,3,4],[0,0,3,4]]
set1=torch.tensor(s1,dtype=torch.float)
s2=[[2,0,5,4],[1,3,3,6],[1,0,2,2]]
set2=torch.tensor(s2,dtype=torch.float)
#find_jaccard_overlap(set1,set2)
print("Bounding boxes",set1)
print("Anchor Boxes",set2)
# + id="L3aYjnr1rXVl" colab_type="code" outputId="a598315b-6513-47b5-e887-f2c59d01261b" colab={"base_uri": "https://localhost:8080/", "height": 34}
set1.shape
# + id="OMSiczgfzUus" colab_type="code" outputId="65b7d2be-239a-4b96-870f-5a5002700f4d" colab={"base_uri": "https://localhost:8080/", "height": 70}
set1[:, :2].unsqueeze(dim=1) #2x1x2
# + id="hoP-YSWS3qz3" colab_type="code" outputId="b7a6e835-9c79-4e83-e383-586c3a790b28" colab={"base_uri": "https://localhost:8080/", "height": 70}
set2[:, :2].unsqueeze(dim=0) #1x3x2
# + id="cBF8uPAwz_P-" colab_type="code" outputId="59280a7f-8618-4a0e-c5f3-7826d30ea916" colab={"base_uri": "https://localhost:8080/", "height": 140}
#lower_bound
torch.max(set1[:, :2].unsqueeze(1), set2[:, :2].unsqueeze(0))
# + id="dNlMoABJ47FR" colab_type="code" outputId="adce1ea0-d1d9-4e95-e15f-66442a613b34" colab={"base_uri": "https://localhost:8080/", "height": 140}
#upper_bound
torch.min(set1[:, 2:].unsqueeze(1), set2[:, 2:].unsqueeze(0))
# + id="GqJbAThj4wNS" colab_type="code" outputId="00a0f9cd-dd8a-4dd8-9ea0-c43451f66647" colab={"base_uri": "https://localhost:8080/", "height": 140}
lower_bounds = torch.max(set1[:, :2].unsqueeze(1), set2[:, :2].unsqueeze(0)) # (n1, n2, 2)
upper_bounds = torch.min(set1[:, 2:].unsqueeze(1), set2[:, 2:].unsqueeze(0)) # (n1, n2, 2)
intersection_dims = torch.clamp(upper_bounds - lower_bounds, min=0)
print("Width and Height of Intersection",intersection_dims)
# + id="xNgKO2Yr1hAE" colab_type="code" outputId="2ceed21a-52ce-4748-85dd-49a709e9c70e" colab={"base_uri": "https://localhost:8080/", "height": 52}
x=find_intersection(set1,set2)
print("Area of intersection",x)
# + id="gVxbV275Uwhg" colab_type="code" outputId="cf5dd89f-e96d-42bd-c41f-57f016049187" colab={"base_uri": "https://localhost:8080/", "height": 70}
areas_set_1 = (set1[:, 2] - set1[:, 0]) * (set1[:, 3] - set1[:, 1]) # (n1)
print("Area Set1 :",areas_set_1[0])
areas_set_2 = (set2[:, 2] - set2[:, 0]) * (set2[:, 3] - set2[:, 1]) # (n2)
print("Area Set2 :",areas_set_2)
union = areas_set_1.unsqueeze(1) + areas_set_2.unsqueeze(0) - x # (n1, n2)
print("Union: ",union[0])
# + id="MNEGMHmk1n-K" colab_type="code" outputId="f1b1ddb8-a76e-4f09-d417-3e3d6122feb6" colab={"base_uri": "https://localhost:8080/", "height": 52}
find_jaccard_overlap(set1,set2)
# + [markdown] id="5EkNDC2FwUpo" colab_type="text"
# # Decimate
# + id="J6oK3ou96va9" colab_type="code" colab={}
def decimate(tensor, m):
"""
Decimate a tensor by a factor 'm', i.e. downsample by keeping every 'm'th value.
This is used when we convert FC layers to equivalent Convolutional layers, BUT of a smaller size.
:param tensor: tensor to be decimated
:param m: list of decimation factors for each dimension of the tensor; None if not to be decimated along a dimension
:return: decimated tensor
"""
assert tensor.dim() == len(m)
for d in range(tensor.dim()):
if m[d] is not None:
print("Index:",d)
print("Number of elements in d = {} (dimension) : {}".format(d,tensor.size(d)))
print("Step Size:",m[d])
'''
Returns a new tensor which indexes the input tensor along dimension dim using the entries in index which is a LongTensor.
1st iteration start-0 - end-2 step_size-4 -selects just 1st element
2nd iteration Ignore
3rd iteration start-0 - end-7 step_size-3 -selects 1st , 4th and 7th element of current axis
4th iteration start-0 - end-2 step_size-3 -selects 1st , 4th and 7th element of current axis
'''
#Keep subsetting across dimensions
tensor = tensor.index_select(dim=d,
index=torch.arange(start=0, end=tensor.size(d), step=m[d]).long())
print(tensor)
return tensor
# + id="mOQV5CyA6vUH" colab_type="code" outputId="f6f880c3-f364-41a1-c44d-c9aaad705194" colab={"base_uri": "https://localhost:8080/", "height": 597}
s1=torch.randn([2,2,7,7])
print("Dimension of s1:", s1.dim())
m=[4,None,3,3] #Number of filters (Divide by 4(4096/4)),None-Input Channels,
print(s1)
# + id="17rOEglH6-gf" colab_type="code" outputId="d520ed8c-f7b2-490e-b8c4-4a3e3d86dcfc" colab={"base_uri": "https://localhost:8080/", "height": 685}
x=decimate(s1,m)
# + id="SpenaSkkvRKP" colab_type="code" outputId="e9aa9118-e0bd-4ff1-feb4-75089f2c18b4" colab={"base_uri": "https://localhost:8080/", "height": 34}
x.shape
# + [markdown] id="SqwvSuAhlChM" colab_type="text"
# # Create Prior Boxes
#
# + id="0ebq32relBsB" colab_type="code" colab={}
def create_prior_boxes(self):
"""
Create the 8732 prior (default) boxes for the SSD300, as defined in the paper.
:return: prior boxes in center-size coordinates, a tensor of dimensions (8732, 4)
"""
fmap_dims = {'conv4_3': 38,
'conv7': 19,
'conv8_2': 10,
'conv9_2': 5,
'conv10_2': 3,
'conv11_2': 1}
obj_scales = {'conv4_3': 0.1,
'conv7': 0.2,
'conv8_2': 0.375,
'conv9_2': 0.55,
'conv10_2': 0.725,
'conv11_2': 0.9}
aspect_ratios = {'conv4_3': [1., 2., 0.5],
'conv7': [1., 2., 3., 0.5, .333],
'conv8_2': [1., 2., 3., 0.5, .333],
'conv9_2': [1., 2., 3., 0.5, .333],
'conv10_2': [1., 2., 0.5],
'conv11_2': [1., 2., 0.5]}
fmaps = list(fmap_dims.keys())
prior_boxes = []
for k, fmap in enumerate(fmaps):
for i in range(fmap_dims[fmap]):
for j in range(fmap_dims[fmap]):
cx = (j + 0.5) / fmap_dims[fmap]
cy = (i + 0.5) / fmap_dims[fmap]
for ratio in aspect_ratios[fmap]:
prior_boxes.append([cx, cy, obj_scales[fmap] * sqrt(ratio), obj_scales[fmap] / sqrt(ratio)])
# For an aspect ratio of 1, use an additional prior whose scale is the geometric mean of the
# scale of the current feature map and the scale of the next feature map
if ratio == 1.:
try:
additional_scale = sqrt(obj_scales[fmap] * obj_scales[fmaps[k + 1]])
# For the last feature map, there is no "next" feature map
except IndexError:
additional_scale = 1.
prior_boxes.append([cx, cy, additional_scale, additional_scale])
prior_boxes = torch.FloatTensor(prior_boxes).to(device) # (8732, 4)
prior_boxes.clamp_(0, 1) # (8732, 4)
return prior_boxes
# + id="8E0U8uSLlJdv" colab_type="code" colab={}
def create_prior_boxes():
"""
Create the 8732 prior (default) boxes for the SSD300, as defined in the paper.
:return: prior boxes in center-size coordinates, a tensor of dimensions (8732, 4)
"""
fmap_dims = {'conv10_2': 3,
'conv11_2': 1}
obj_scales = {'conv10_2': 0.725,
'conv11_2': 0.9}
aspect_ratios = {'conv10_2': [1., 2., 0.5],
'conv11_2': [1., 2., 0.5]}
fmaps = list(fmap_dims.keys())
prior_boxes = []
for k, fmap in enumerate(fmaps):
print("For feature map = {}".format(fmap))
for i in range(fmap_dims[fmap]):
#print(i)
#print("y location on Feature Map(i) = {} and fmap = {}".format(i,fmap))
for j in range(fmap_dims[fmap]):
#print(j)
print("x,y location on Feature Map(j,i) = ({},{})".format(j,i))
cx = (j + 0.5) / fmap_dims[fmap]
cy = (i + 0.5) / fmap_dims[fmap]
print("cx = {} and cy = {}".format(cx,cy))
for m,ratio in enumerate(aspect_ratios[fmap]):
prior_boxes.append([cx, cy, obj_scales[fmap] * sqrt(ratio), obj_scales[fmap] / sqrt(ratio)])
print("For aspect ration {}".format(ratio))
print(prior_boxes)
#print("Pre\n Centre of x ={}\n Centre of y = {}\n Widht ={}\n Height {}".format(prior_boxes[m][0],prior_boxes[m][1],prior_boxes[m][2],prior_boxes[m][3]))
# For an aspect ratio of 1, use an additional prior whose scale is the geometric mean of the
# scale of the current feature map and the scale of the next feature map
if ratio == 1.:
print("Add 1 more prior box")
try:
additional_scale = sqrt(obj_scales[fmap] * obj_scales[fmaps[k + 1]])
print("Additional Scale",additional_scale)
# For the last feature map, there is no "next" feature map
except IndexError:
additional_scale = 1.
prior_boxes.append([cx, cy, additional_scale, additional_scale])
print(prior_boxes)
#print(" Post \n Modified Centre of x ={}\n Centre of y = {}\n Widht ={}\n Height {}".format(prior_boxes[m+1][0],prior_boxes[i][1],prior_boxes[i][2],prior_boxes[i][3]))
prior_boxes = torch.FloatTensor(prior_boxes).to(device) # (8732, 4)
prior_boxes.clamp_(0, 1) # (8732, 4)
return prior_boxes
# + id="GhwQc4uxlPto" colab_type="code" outputId="c4dca510-0e5c-4c7f-c7c5-6504b205b34f" colab={"base_uri": "https://localhost:8080/", "height": 2708}
print("After clamping values between 0 and 1\n",create_prior_boxes())
# + [markdown] id="8Zp8P5Gv7riV" colab_type="text"
# # MultiLoss
#
# What we have:
# 1. Two Bounding Boxes corresponding to two objects
# 2. 3 anchor boxes
# 3. True Labels for each bounding boxes
#
# What we require:
# 1. Labels for each prior boxes
#
# Steps:
# 1. Assign true labels to anchor boxes
# <br>
# For each bounding box :
# a. Calculate the overlap with the anchor boxes -Jaccard Index
# b. Identify the anchor box with the highest overlap.
# c. Assign the anchor box with the true lable of the bounding box.
# 2. Calculate the Localization Loss(Done only for positive priors)
# <br>
# Use the mask tensor to calculate loss over the relevant anchor boxes offsets.
# 3. Hard Negative Mining
# 4. Calculate the Confidence Loss
#
# + id="vQ22PSQ37qqo" colab_type="code" outputId="1b3270e8-ba35-4c01-ea16-597f863a10bd" colab={"base_uri": "https://localhost:8080/", "height": 157}
boxes=[[0,0,3,4],[0,0,3,6]]
boxes=torch.tensor(boxes,dtype=torch.float)
priors=[[0,0,5,4],[0,2.8,3,6],[0,0,4,7],[2.8,2.8,3,6]]
priors=torch.tensor(priors,dtype=torch.float)
#find_jaccard_overlap(set1,set2)
print("2 Boundin boxes\n",boxes)
print(" 3 Anchor Boxes\n",priors)
# + id="PsExCWRIDf_5" colab_type="code" outputId="c03d7908-10ee-4f8d-a2c9-87f8401afe5a" colab={"base_uri": "https://localhost:8080/", "height": 87}
priors_cxcy=xy_to_cxcy(priors)
print(priors_cxcy)
# + [markdown] id="ilq_4RBX8zMT" colab_type="text"
# ## Step1:
# 1. Assign true labels to anchor boxes
# <br>
# For each bounding box :
# a. Calculate the overlap with the anchor boxes -Jaccard Index
# b. Identify the anchor box with the highest overlap by making it 1 and the remaining to 0(masking) if overlap is less tha 0.5 .
# c. Assign the anchor box with the true lable of the bounding box.
#
# + id="ZK_saRD27L5r" colab_type="code" outputId="edd40c32-6a2a-4953-abe7-e28d6eaf7706" colab={"base_uri": "https://localhost:8080/", "height": 70}
#Bounding box = 2
labels=torch.tensor([[2],[3]])
n_objects=boxes.size(0)
overlap=find_jaccard_overlap(boxes,priors)
overlap #2 x 4
#Row is the object
#COlumn is the priors
print("Overlap between Bounding Box and Anchor Boxes\n",overlap)
#Desired Outcome -1. Assign lable of 1st Bounding box to 1st anchor box
#2. Assign label of 2nd Bounding box to 3rd anchor box
# + [markdown] id="Zr9nLATI8-28" colab_type="text"
# For each prior, find the object that has the maximum overlap
# + id="Jk_MFlHg7P2a" colab_type="code" outputId="56eb1b62-75e1-48c0-ecdc-32e101e129ef" colab={"base_uri": "https://localhost:8080/", "height": 34}
overlap_for_each_prior, object_for_each_prior=overlap.max(dim=0)
print(overlap.max(dim=0)) #4
# + [markdown] id="He8mCGXO9fFe" colab_type="text"
# For each object.find the prior that has the maximum overlap
# + id="ccZBR-oS9Lq_" colab_type="code" outputId="caeb813f-2b3b-4b45-aba3-1b5437185137" colab={"base_uri": "https://localhost:8080/", "height": 34}
_, prior_for_each_object = overlap.max(dim=1)
print(overlap.max(dim=1)) #2
# + [markdown] id="_GlJn43T91ps" colab_type="text"
# For each priors, assign each object to the corresponding maximum-overlap-prior. (This fixes 1.)
#
# + id="rLYrcttz9047" colab_type="code" outputId="27d0e6a9-4298-4808-878d-2c3d94e6b476" colab={"base_uri": "https://localhost:8080/", "height": 34}
object_for_each_prior[prior_for_each_object] = torch.LongTensor(range(n_objects)).to(device)
print("Identified object for each prior",object_for_each_prior)
# + id="UA6KyrpO-cRq" colab_type="code" outputId="9cfa196b-13b6-4462-a9f5-82819a9bd3fd" colab={"base_uri": "https://localhost:8080/", "height": 34}
overlap_for_each_prior[prior_for_each_object] = 1.
print("Overlap of Identified object for each prior", overlap_for_each_prior)
# + id="EDCFrDbKC5RT" colab_type="code" outputId="97f5ed89-e8bb-4c6c-a00a-7bcceec34890" colab={"base_uri": "https://localhost:8080/", "height": 87}
# Encode center-size object coordinates into the form we regressed predicted boxes to
true_locs = cxcy_to_gcxgcy(xy_to_cxcy(set1[object_for_each_prior]), priors_cxcy) # (8732, 4)
print(true_locs)
# + id="-xeZR5nXC3Bw" colab_type="code" colab={}
threshold=0.5
# Labels for each prior
label_for_each_prior = labels[object_for_each_prior] # (3)
# Set priors whose overlaps with objects are less than the threshold to be background (no object)
label_for_each_prior[overlap_for_each_prior < threshold] = 0 # (8732)
# Store
true_classes = label_for_each_prior
# + id="RXkptD_HGR33" colab_type="code" outputId="b6a48f25-5c55-4217-a638-a2cd0dd4ed91" colab={"base_uri": "https://localhost:8080/", "height": 87}
label_for_each_prior
# + id="Wl0RiuB6LYeD" colab_type="code" outputId="aff29083-dfcc-43f5-d1e0-da61eddc876a" colab={"base_uri": "https://localhost:8080/", "height": 87}
positive_priors = true_classes != 0 # (N, 8732)
print("Mask tensor",positive_priors)
# + [markdown] id="6nhRQCujTsNP" colab_type="text"
# Step 3: Hard Negative Mining
#
#
# + id="xRRpZ4CcRPYo" colab_type="code" outputId="588433e2-1c8c-4a76-e501-9c6e97f9f8bb" colab={"base_uri": "https://localhost:8080/", "height": 34}
positive_priors.sum(dim=1)
# + id="3yZvqRX0Q1Ap" colab_type="code" outputId="bee1f422-a066-41dc-d5c5-149492489a50" colab={"base_uri": "https://localhost:8080/", "height": 34}
neg_pos_ratio=3
n_positives = positive_priors.sum(dim=0) # (N)
n_hard_negatives = neg_pos_ratio * n_positives # (N)
print(n_positives,n_hard_negatives)
| Phase - 2/SSD/SSD_Helper_File.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Latest point anomaly detection with the Anomaly Detector API
# ### Use this Jupyter notebook to start visualizing anomalies as a batch with the Anomaly Detector API in Python.
#
# While you can detect anomalies as a batch, you can also detect the anomaly status of the last data point in the time series. This notebook iteratively sends latest-point anomaly detection requests to the Anomaly Detector API and visualizes the response. The graph created at the end of this notebook will display the following:
# * Anomalies found while in the data set, highlighted.
# * Anomaly detection boundaries
# * Anomalies seen in the data, highlighted.
#
# By calling the API on your data's latest points, you can monitor your data as it's created.
#
# The following example simulates using the Anomaly Detector API on streaming data. Sections of the example time series are sent to the API over multiple iterations, and the anomaly status of each section's last data point is saved. The data set used in this example has a pattern that repeats roughly every 7 data points (the `period` in the request's JSON file), so for best results, the data set is sent in groups of 29 points (`4 * <period> + an extra data point`. See [Best practices for using the Anomaly Detector API](https://docs.microsoft.com/azure/cognitive-services/anomaly-detector/concepts/anomaly-detection-best-practices) for more information).
# To start sending requests to the Anomaly Detector API, paste your Anomaly Detector resource access key below,
# and replace the endpoint variable with the endpoint for your region or your on-premise container endpoint.
# Endpoint examples:
# https://westus2.api.cognitive.microsoft.com/anomalydetector/v1.0/timeseries/last/detect
# http://127.0.0.1:5000/anomalydetector/v1.0/timeseries/last/detect
apikey = '[Placeholder: Your Anomaly Detector resource access key]'
endpoint_latest = '[Placeholder: Your Anomaly Detector resource endpoint]/anomalydetector/v1.0/timeseries/last/detect'
# +
import requests
import json
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings('ignore')
# Import library to display results
import matplotlib.pyplot as plt
# %matplotlib inline
# -
from bokeh.plotting import figure,output_notebook, show
from bokeh.palettes import Blues4
from bokeh.models import ColumnDataSource,Slider
import datetime
from bokeh.io import push_notebook
from dateutil import parser
from ipywidgets import interact, widgets, fixed
output_notebook()
def detect(endpoint, apikey, request_data):
headers = {'Content-Type': 'application/json', 'Ocp-Apim-Subscription-Key': apikey}
response = requests.post(endpoint, data=json.dumps(request_data), headers=headers)
if response.status_code == 200:
return json.loads(response.content.decode("utf-8"))
else:
print(response.status_code)
raise Exception(response.text)
def build_figure(result, sample_data, sensitivity):
columns = {'expectedValues': result['expectedValues'], 'isAnomaly': result['isAnomaly'], 'isNegativeAnomaly': result['isNegativeAnomaly'],
'isPositiveAnomaly': result['isPositiveAnomaly'], 'upperMargins': result['upperMargins'], 'lowerMargins': result['lowerMargins']
, 'value': [x['value'] for x in sample_data['series']], 'timestamp': [parser.parse(x['timestamp']) for x in sample_data['series']]}
response = pd.DataFrame(data=columns)
values = response['value']
label = response['timestamp']
anomalies = []
anomaly_labels = []
index = 0
anomaly_indexes = []
p = figure(x_axis_type='datetime', title="Anomaly Detection Result ({0} Sensitivity)".format(sensitivity), width=800, height=600)
for anom in response['isAnomaly']:
if anom == True and (values[index] > response.iloc[index]['expectedValues'] + response.iloc[index]['upperMargins'] or
values[index] < response.iloc[index]['expectedValues'] - response.iloc[index]['lowerMargins']):
anomalies.append(values[index])
anomaly_labels.append(label[index])
anomaly_indexes.append(index)
index = index+1
upperband = response['expectedValues'] + response['upperMargins']
lowerband = response['expectedValues'] -response['lowerMargins']
band_x = np.append(label, label[::-1])
band_y = np.append(lowerband, upperband[::-1])
boundary = p.patch(band_x, band_y, color=Blues4[2], fill_alpha=0.5, line_width=1, legend='Boundary')
p.line(label, values, legend='value', color="#2222aa", line_width=1)
p.line(label, response['expectedValues'], legend='expectedValue', line_width=1, line_dash="dotdash", line_color='olivedrab')
anom_source = ColumnDataSource(dict(x=anomaly_labels, y=anomalies))
anoms = p.circle('x', 'y', size=5, color='tomato', source=anom_source)
p.legend.border_line_width = 1
p.legend.background_fill_alpha = 0.1
show(p, notebook_handle=True)
# ### Detect latest anomaly of sample timeseries
# The following cells call the Anomaly Detector API with an example time series data set and different sensitivities for anomaly detection. Varying the sensitivity of the Anomaly Detector API can improve how well the response fits your data.
def detect_anomaly(sensitivity):
sample_data = json.load(open('univariate_sample_daily.json'))
points = sample_data['series']
skip_point = 29
result = {'expectedValues': [None]*len(points), 'upperMargins': [None]*len(points),
'lowerMargins': [None]*len(points), 'isNegativeAnomaly': [False]*len(points),
'isPositiveAnomaly':[False]*len(points), 'isAnomaly': [False]*len(points)}
anom_count = 0
for i in range(skip_point, len(points)+1):
single_sample_data = {}
single_sample_data['series'] = points[i-29:i]
single_sample_data['granularity'] = 'daily'
single_sample_data['maxAnomalyRatio'] = 0.25
single_sample_data['sensitivity'] = sensitivity
single_point = detect(endpoint_latest, apikey, single_sample_data)
if single_point['isAnomaly'] == True:
anom_count = anom_count + 1
result['expectedValues'][i-1] = single_point['expectedValue']
result['upperMargins'][i-1] = single_point['upperMargin']
result['lowerMargins'][i-1] = single_point['lowerMargin']
result['isNegativeAnomaly'][i-1] = single_point['isNegativeAnomaly']
result['isPositiveAnomaly'][i-1] = single_point['isPositiveAnomaly']
result['isAnomaly'][i-1] = single_point['isAnomaly']
build_figure(result, sample_data, sensitivity)
# 95 sensitvity
detect_anomaly(95)
# 85 sensitvity
detect_anomaly(85)
| ipython-notebook/API Sample/Latest point detection with the Anomaly Detector API.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cfcastillo/DS-6-Notebooks/blob/main/Education_Capstone_MS2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Tk2njJU2cft7"
# # TASK LIST
#
# * Find dataset(s) that offer future job popularity/need - **Amy**
# * [top 10 skills by year](https://data.world/datasets/jobs)
#
# * **IN PROCESS** SOC to OCCUP 3 pass match - take max(census) code for multiples - **Cecilia**
#
# **MS-2 requirements**
#
# Your Jupyter Notebook needs to contain the following:
# * **ONGOING** All code needed to clean and format your data
# * **DONE** A written description of the data cleaning process
# * **DONE** completed rows of the dataset, predictors and response.
#
#
# + [markdown] id="BO6cDiaVzYbK"
# # Project Definition
#
# The purpose of this project is to identify what factors influence people to choose certain professions or trades. In understanding these factors, we can help colleges like Central New Mexico College (CNM) offer courses that support those professions and better target their marketing to people who are likely to choose those professions.
#
# This project will be a supervised categorization problem using tree-based models to identify the factors that will contribute to career choice.
#
#
# + [markdown] id="Kmxlgo4Wnjgd"
# # Data Identification Process
#
# Steps:
#
# 1. We stated several questions we wanted answered (target).
# 1. After defining our problem, we listed sets of variables that we believed could answer our questions. We then put the variables and targets into a [spreadsheet](https://docs.google.com/spreadsheets/d/1bOhOBHKOae9TDN9n9-xF7ag4QW_Z0c7HXTYLXeMMLHs/edit#gid=0) to define the dataset we would need to run our analysis.
# 1. We then researched data sources such as Bureau of Labor Statistics and the US Census to locate data that supported our research.
# 1. We then mapped the columns in the data sources to the columns in our desired dataset and linked multiple datasets by target code value.
#
# *Note: The data identification process is still a work in progress. As we proceed with EDA, we will discover some columns are not needed and others are needed. As we analyzed the data during the data cleaning process, we discovered that earnings are complex, often made up of multiple jobs. Additional analysis will be needed to solidify our predictor when applying the model.*
# + [markdown] id="0I3qXXVbFUJt"
# # Data Collection
#
# The following data sources were used for this project. Data was imported into Google Drive from the below links and modified as needed to support this project.
#
# The primary datasets for this project were initially taken from the Census' [Annual Social and Economic Supplement (ASEC)](https://www.census.gov/programs-surveys/saipe/guidance/model-input-data/cpsasec.html) of the Current Population Survey (CPS) for 2020. However, because 2020 was anomalous due to Covid, we had to go back and take data from 2018 and 2019 - pre-covid to get occupation and salary information that was more stable. Per the above link, the "*ASEC data is the source of timely official national estimates of poverty levels and rates and of widely used measures of income. It provides annual estimates based on a survey of more than 75,000 households. The survey contains detailed questions covering social and economic characteristics of each person who is a household member as of the interview date. Income questions refer to income received during the previous calendar year.*"
#
# [Annual Social and Economic Survey (ASEC) All Years Data](https://www.census.gov/data/datasets/time-series/demo/cps/cps-asec.html)
#
# * Contains links to all years from 2010 to 2021. CSV format available from 2019 to 2021. Prior to 2019, fixed format file is provided so columns would need to be parsed using the available data dictionary.
# * [2021 Survey - csv](https://www.census.gov/data/datasets/time-series/demo/cps/cps-asec.2021.html)
# * [2020 Survey - csv](https://www.census.gov/data/datasets/time-series/demo/cps/cps-asec.2020.html)
# * [2019 Survey - csv](https://www.census.gov/data/datasets/time-series/demo/cps/cps-asec.2019.html)
# * [2018 Survey - dat](https://www.census.gov/data/datasets/time-series/demo/cps/cps-asec.2018.html) - Need to convert to csv
#
# [Quarterly Census of Employment and Wages](https://www.bls.gov/cew/about-data/)
#
# * Source data for OES Statistics. Can be used if detailed data is needed.
#
# [Occupational Employment Wage Statistics (OES) Data](https://data.bls.gov/oes/#/geoOcc/Multiple%20occupations%20for%20one%20geographical%20area)
#
# * Format - Excel converted to CSV
# * Contains Occupational codes and aggregated statistics on wages for those occupations.
#
# [FIPS State Codes](https://www.census.gov/library/reference/code-lists/ansi/ansi-codes-for-states.html)
#
# * Format - Copied from PDF and converted to CSV
# * Contains FIPS State codes mapped to US Postal Service (USPS) State codes.
#
# [Census Occupation Codes](https://www2.census.gov/programs-surveys/cps/techdocs/cpsmar20.pdf)
#
# * Format - Copied from PDF and converted to CSV
# * Contains Census Occupation codes mapped to Federal Standard Occupational Classification (SOC) Codes.
#
# + [markdown] id="1faCHIjobl8k"
# # Imports
# + id="QfQPgzzSer2x"
# grab the imports needed for the project
import pandas as pd
import glob
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import statsmodels.api as sm
# all
from sklearn import datasets
from sklearn import metrics
from sklearn import preprocessing
from sklearn.metrics import classification_report
import sklearn.model_selection as model_selection
# Gaussian Naive Bayes
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import MultinomialNB
# Regression
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.metrics import mean_squared_error
# + [markdown] id="2vb8BJGOadkb"
# # Data Cleaning - MS-2 - Oct 1
#
# Once we identified the data elements needed for our project and the data sources that provided those data elements, the following steps were taken to get the data into a format needed for our analysis.
#
# 1. Downloaded data from data sources and placed copies in Google Drive.
# 1. Made changes to raw data where needed to support the project.
# * Added State code to OES data and remove headers and footers from the data.
# * Created lookup data for State codes and SOC codes so secondary data sources could be merged with primary Census data. This involved cleaning the census code list so it could be properly parsed.
# 1. Converted codes in secondary datasets into Census codes.
# 1. Merged all datasets together into a single dataset.
# 1. Removed data that did not meet criteria for our analysis
# * Removed anyone under age 16.
# * Imputed null values.
# 1. Studied earnings/salary columns to determine which columns provided values that could be used for modeling. Added in columns that were missing from the initial analysis.
# 1. We were not able to reliably match the OES data to the census data using the full SOC Code because of disparities in SOC Codes. Therefore, we executed 3 matching passes reducing the SOC code by one character each time and pulling the largest Census code for the SOC code prefix. This allowed us to match XX % of the data.
#
# TODO: REFINE AS PROJECT PROGRESSES.
# + [markdown] id="rPuX27WyMnJ_"
# ## Import Data
# + colab={"base_uri": "https://localhost:8080/"} id="zN5qu2I-HlKA" outputId="e5c594c9-cddd-4c0d-fa75-956de6d4b524"
from google.colab import drive
drive.mount('/content/drive')
# + id="7cPrmvySZmj5"
# Import Census data
asec_year = '19'
asec_path = '/content/drive/MyDrive/Student Folder - Cecilia/Projects/Capstone/Data/ASEC/asecpub' + asec_year + 'csv/'
asec_data_person = pd.read_csv(asec_path + 'pppub' + asec_year + '.csv')
asec_data_household = pd.read_csv(asec_path + 'hhpub' + asec_year + '.csv')
# TODO: once all data is available, join 1x and then save combined file so don't have to join every time code is run.
# Join and import all 50 states' occupation data
oes_path = '/content/drive/MyDrive/Student Folder - Cecilia/Projects/Capstone/Data/Occupations/'
oes_file_names = glob.glob(oes_path + "*.csv")
li = []
for filename in oes_file_names:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
oes_data = pd.concat(li, axis=0, ignore_index=True)
# File path for all code conversion files.
codes_path = '/content/drive/MyDrive/Student Folder - Cecilia/Projects/Capstone/Data/Codes/'
# Import FIPS state codes so we can convert USPS state to FIPS state to match back to Census data.
fips_state_codes = pd.read_csv(codes_path + 'FIPS_STATE_CODES.csv')
# Import Census occupational codes so we can convert SOC codes into Census Occ codes.
# Is in fixed width format. Will parse out data below.
census_occ_codes = pd.read_fwf(codes_path + 'CENSUS_SOC_OCC_CODES.txt')
# + colab={"base_uri": "https://localhost:8080/"} id="8E6_TE9j9UUT" outputId="ea662ed9-0a6b-4549-b877-2d3f1d79498f"
# How many columns and rows do we have in each dataset?
print(f'Person data: {asec_data_person.shape}')
print(f'Household data: {asec_data_household.shape}')
print(f'Occupation data: {oes_data.shape}')
print(f'FIPS State Codes: {fips_state_codes.shape}')
print(f'Census Occ Codes: {census_occ_codes.shape}')
# + [markdown] id="qsyP7bCKMtEQ"
# ## ASEC Data
# + [markdown] id="nOawUJ1nQ8J9"
# ### Define ASEC Columns
#
# The following data dictionary provides details for the selected columns.
#
# [Annual Social and Economic Supplement (ASEC) 2020 Public Use Dictionary](https://www2.census.gov/programs-surveys/cps/datasets/2020/march/ASEC2020ddl_pub_full.pdf)
# + id="WDc8BIA8XttY"
# Get lists of columns for various datasets that will be used for the project
# Note: Columns can be added as needed here and will propagate through the project.
id_col = ['H_IDNUM']
person_cols = ['OCCUP','A_DTOCC','A_MJOCC','AGE1','A_AGE','A_SEX','PRDTRACE','PRCITSHP','A_HGA','A_HRLYWK', 'A_HRSPAY','A_GRSWK',
'CLWK','EARNER','HRCHECK','HRSWK','PEARNVAL','A_CLSWKR','A_DTIND']
household_cols = ['GTMETSTA','GEDIV','GESTFIPS','HEFAMINC','HHINC']
# + [markdown] id="FTp3lJSiUXLy"
# ### Get Household Id
# + id="1mKhyrDr6413"
# Extract the Household id number from the person record so we can join the household and person dataframes by this id.
asec_data_person['H_IDNUM'] = asec_data_person['PERIDNUM'].str[:20]
# + colab={"base_uri": "https://localhost:8080/"} id="VJpYTwqHsZiV" outputId="4a4ac670-a8ea-40cf-c4b0-d0a3d62f3d19"
# View Person Data
asec_data_person[person_cols].info()
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="lCGqLO4xR8Me" outputId="8a3cec73-6756-4dc8-a6f6-d9cb7d9b83f8"
# Look at first 5 records of selected columns of person data.
asec_data_person[person_cols].head()
# + colab={"base_uri": "https://localhost:8080/"} id="xc8qbSUIse8z" outputId="87c5f40e-d1d9-482a-ae34-ee89a74b3669"
# View Household Data
asec_data_household[household_cols].info()
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="f8AO0ZvtR-6w" outputId="bc7c30ae-89b8-4b32-93fa-c31351c1105f"
# Look at first 5 records of household data
asec_data_household[household_cols].head()
# + [markdown] id="lD4Z0dJhUcSF"
# ### Merge Person and Household Records
# + id="_vwS8IBWIWQ6"
# Join Household and Personal records into single dataframe
# Inner join - should not have person without household.
asec_combined = pd.merge(asec_data_household[id_col + household_cols], asec_data_person[id_col + person_cols], on=id_col)
# + colab={"base_uri": "https://localhost:8080/"} id="7vcMHyLB6cW2" outputId="20a02ad3-4fcd-4628-f611-b2d8120f741f"
# View combined result
asec_combined.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 225} id="skMqUb61iCJZ" outputId="353dc3f0-f7ef-45b0-839f-22b44d63412a"
asec_combined.head()
# + [markdown] id="2dhwZPNWM2ZN"
# ## OES Data
# + id="OIHW6Dnp7XUv"
# Shorten column names
oes_data.rename(columns={'State':'USPS_STATE',
'Occupation (SOC code)':'SOC_DESC',
'Employment(1)':'EMP',
'Employment percent relative standard error(3)':'EMP_RSDE',
'Hourly mean wage':'HOURLY_MEAN',
'Annual mean wage(2)':'ANN_MEAN',
'Wage percent relative standard error(3)':'WAGE_RSDE',
'Hourly 10th percentile wage':'HOURLY_10TH',
'Hourly 25th percentile wage':'HOURLY_25TH',
'Hourly median wage':'HOURLY_MEDIAN',
'Hourly 75th percentile wage':'HOURLY_75TH',
'Hourly 90th percentile wage':'HOURLY_90TH',
'Annual 10th percentile wage(2)':'ANN_10TH',
'Annual 25th percentile wage(2)':'ANN_25TH',
'Annual median wage(2)':'ANN_MEDIAN',
'Annual 75th percentile wage(2)':'ANN_75TH',
'Annual 90th percentile wage(2)':'ANN_90TH',
'Employment per 1,000 jobs':'EMP_PER_1000',
'Location Quotient':'LOC_QUOTIENT'}, inplace=True)
# + [markdown] id="Aw2LNu7bMKqZ"
# ### OES Column Footnotes
#
# * (1) Estimates for detailed occupations do not sum to the totals because the totals include occupations not shown separately. Estimates do not include self-employed workers.
# * (2) Annual wages have been calculated by multiplying the corresponding hourly wage by 2,080 hours.
# * (3) The relative standard error (RSE) is a measure of the reliability of a survey statistic. The smaller the relative standard error, the more precise the estimate.
# * (4) Wages for some occupations that do not generally work year-round, full time, are reported either as hourly wages or annual salaries depending on how they are typically paid.
# * (5) This wage is equal to or greater than \$100.00 per hour or \$208,000 per year.
# * (8) Estimate not released.
# + id="jnJ4w_WwajKM"
# OES columns we want to keep
# occupation_cols = ['HOURLY_MEAN','HOURLY_MEDIAN','EMP_PER_1000','LOC_QUOTIENT']
occupation_cols = ['HOURLY_MEAN','HOURLY_MEDIAN','EMP_PER_1000','LOC_QUOTIENT', 'ANN_MEAN','ANN_MEDIAN']
# 2 EMP 35874 non-null object
# 3 EMP_RSDE 35874 non-null object
# 4 HOURLY_MEAN 35874 non-null object
# 5 ANN_MEAN 35874 non-null object
# 6 WAGE_RSDE 35874 non-null object
# 7 HOURLY_10TH 35874 non-null object
# 8 HOURLY_25TH 35874 non-null object
# 9 HOURLY_MEDIAN 35874 non-null object
# 10 HOURLY_75TH 35874 non-null object
# 11 HOURLY_90TH 35874 non-null object
# 12 ANN_10TH 35874 non-null object
# 13 ANN_25TH 35874 non-null object
# 14 ANN_MEDIAN 35874 non-null object
# 15 ANN_75TH 35874 non-null object
# 16 ANN_90TH 35874 non-null object
# 17 EMP_PER_1000 35874 non-null object
# 18 LOC_QUOTIENT 35874 non-null object
# + [markdown] id="Y-KQemIYUPzd"
# ### Get Census State Codes
# + id="1BwU19nKdFeW"
# Import FIPS state codes matching on USPS state codes
# Left Join - keep OES data even if no match on state code.
oes_data = pd.merge(oes_data, fips_state_codes[['USPS_STATE','FIPS_STATE']], on='USPS_STATE', how='left')
# + colab={"base_uri": "https://localhost:8080/"} id="MxpMW58wPaTv" outputId="193115f6-bd48-4a37-d44e-feeaa5bc9328"
# Verify merge was successful - that we have expected columns and record count is unchanged.
oes_data.info()
# + colab={"base_uri": "https://localhost:8080/"} id="X8OCk1Amy9yO" outputId="c7148cf5-ecc3-417c-e188-406594211f15"
# Verify we have all states
oes_data[['USPS_STATE', 'FIPS_STATE']].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="sGW0uwIm6_RY" outputId="384df112-ad28-4ea0-ca22-20764fb9383f"
# Verify no nulls after merge.
oes_data.isnull().sum()
# + [markdown] id="_FVODfj7UDR3"
# ### Parse SOC Codes
# + id="3ovrRNV8zMGf"
# Parse out SOC code from the description. The code is inside parentheses.
def getSocCode(value):
# If not able to parse the code, then return the value from the file.
try:
return value[value.index('(')+1:value.index(')')]
except:
return value
oes_data['SOC_CODE'] = oes_data['SOC_DESC'].apply(lambda val: getSocCode(val))
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="sDFvk_x10IPL" outputId="2fb9f562-612d-4599-a9ac-4b919a841241"
# Verify codes were properly parsed
oes_data[['SOC_DESC','SOC_CODE']]
# + [markdown] id="cQVtlSg_UGs6"
# ### Get Census Occupation Codes
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="z6RmtCjqL9Wh" outputId="ab77339e-9497-4982-ea5b-bdddf4b58c99"
# Prepare Census/SOC map file. Codes are embedded in a single column so need to be parsed out.
# Parse out Census and SOC occupational codes from the description. The code has a dash in it. So locate by dash
# assuming there are no dashes in the description.
def getOccCodeSoc(value):
# If not able to parse the code, then return the value from the file.
try:
return value[value.index('-')-2:value.index('-')+5].replace('-','')
except:
return value
# Retrieve first 4 characters in the file. This is the Census code.
def getOccCodeCensus(value):
return value[:4]
census_occ_codes['OCCUP'] = census_occ_codes['CENSUS_MAP'].apply(lambda val: getOccCodeCensus(val))
census_occ_codes['SOC_CODE'] = census_occ_codes['CENSUS_MAP'].apply(lambda val: getOccCodeSoc(val))
census_occ_codes
# + id="JJ0s-S-Q24mg"
# TODO: determine if needed. This is a work in progress.
# There are some cases where we do not have an exact match on the full SOC Code.
# In such cases, try to match on the first 5 characters, taking the largest Occ code in case
# there are multiple Occ codes associated with the first 5 characters of SOC Code.
census_occ_codes['SOC_CODE5'] = census_occ_codes['SOC_CODE'].str[:5]
census_occ_codes_soc5 = census_occ_codes.groupby("SOC_CODE5").agg({
'OCCUP':['max','count']
}).reset_index()
# Repeat the process using the first 4 characters of SOC Code.
census_occ_codes['SOC_CODE4'] = census_occ_codes['SOC_CODE'].str[:4]
census_occ_codes_soc4 = census_occ_codes.groupby("SOC_CODE4").agg({
'OCCUP':['max','count']
}).reset_index()
# + id="fpe26JACQ1aP"
# Convert SOC Code into Census occupation code
# Left Join - keep OES data even if no match on state code.
oes_data = pd.merge(oes_data, census_occ_codes[['OCCUP','SOC_CODE']], on='SOC_CODE', how='left')
# + id="hhe7d0EldzDd"
# TODO: 2 more passes needed to match. If still largely unmatched with Census code, then will use census
# major and detailed categories instead and will forego OES statistics.
# Get rows that are missing OCCUP code.
df = oes_data[oes_data.isnull().any(axis=1)]
# df['SOC_CODE'].value_counts().to_csv('test.csv')
# oes_data
# + colab={"base_uri": "https://localhost:8080/", "height": 813} id="ftsNu9lggYzW" outputId="868f93c7-addb-4437-dcca-6b520e8f601e"
oes_data.info()
oes_data.head()
# + [markdown] id="pFCk35poNC2N"
# ## Combine All Data
# + id="KG00feoEfKei"
# Bring in Occupational data joining on FIPS state and full SOC code.
# Convert to numeric datatypes so data can be merged.
oes_data['FIPS_STATE'] = pd.to_numeric(oes_data['FIPS_STATE'], errors='coerce')
oes_data['OCCUP'] = pd.to_numeric(oes_data['OCCUP'], errors='coerce')
# Left Join - keep OES data even if no match on state code.
asec_oes = pd.merge(asec_combined, oes_data, left_on=['GESTFIPS','OCCUP'], right_on=['FIPS_STATE','OCCUP'], how='left')
# Only get desired columns
asec_oes = asec_oes[household_cols + person_cols + occupation_cols]
# + colab={"base_uri": "https://localhost:8080/", "height": 865} id="iTOrZhFvjM_S" outputId="be302cb2-e4a8-48d1-d31f-38683ea78d15"
# Review result of merged data
asec_oes.info()
asec_oes.head()
# + [markdown] id="6ZZwtWTaNSEN"
# ## Clean Data
# + colab={"base_uri": "https://localhost:8080/", "height": 865} id="GRPyEs4I8U7v" outputId="ea9ed1a0-6929-49f2-a5d5-803b01e6298d"
# Remove people under 15 years old because they are not relevant for this project.
# 0 = Not in universe
# 1 = 15 years
# 2 = 16 and 17 years
# 3 = 18 and 19 years
# 4 = 20 and 21 years
# 5 = 22 to 24 years
# 6 = 25 to 29 years
# 7 = 30 to 34 years
# 8 = 35 to 39 years
# 9 = 40 to 44 years
# 10 = 45 to 49 years
# 11 = 50 to 54 years
# 12 = 55 to 59 years
# 13 = 60 to 61 years
# 14 = 62 to 64 years
# 15 = 65 to 69 years
# 16 = 70 to 74 years
# 17 = 75 years and over
asec_oes = asec_oes[asec_oes['AGE1'] > 0]
asec_oes.info()
asec_oes.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 225} id="v9UCyGkdTR5W" outputId="f83fc3f4-c6d2-456a-e0af-cec25f259a04"
asec_oes.tail()
# + [markdown] id="bUT_zr2uU2Nq"
# ### Column Descriptions
#
# TODO: Create summary document that has chosen columns.
#
# **Demographic**
# * AHGA - Educational Attainment
#
# **Geo**
#
# **Earnings**
# * A_HRLYWK - Is paid by the hour
# * A_HRSPAY - If is paid by the hour, this is hourly wage
# * A_GRSWK - Gross weekly salary
# * CLWK - Longest job classification
# * EARNER - earner/non-earner
# * HRCHECK - part time/full time
# * HRSWK - how many hours does respondent work per week
# * PEARNVAL - total person earnings - can be positive or negative
# * A_CLSWKR - private/public/self employed
# * A_DTIND - Industry code - Appendix A.
# + colab={"base_uri": "https://localhost:8080/"} id="kYACiVFVfA6R" outputId="3a2b294f-93b9-49e2-8a0b-5fa7401bf2c1"
# TODO: need to switch to annual pay since A_HRSPAY is only for people who hold hourly positions.
# Convert hours pay into a float with 2 decimal places
asec_oes['A_HRSPAY'] = asec_oes['A_HRSPAY'].astype('float') / 100
# first pass - 93897/141251 nulls
asec_oes['HOURLY_MEAN'].isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="HQ8Fkx0rZzqT" outputId="f438136f-b1b1-494f-c36f-15f15f4d0938"
asec_oes.shape
# + id="vxWuX2N4-Nxr"
# TODO: Handle null or blank data - We have some "-" data in the oes file that indicates value is unavailable.
# + [markdown] id="PRkEJT36ag8H"
# # Exploratory Data Analysis (EDA) - MS-3 - Oct 15
# + [markdown] id="MK6RwwzNampl"
# # Data Processing / Models - MS-4 - Oct 29
# + [markdown] id="UDLQw48gaxPv"
# # Data Visualization and Results - MS-5 - Nov 19
# + [markdown] id="gUD7CFtATZQd"
# # Presentation and Conclusions - Final - Dec 3
#
#
# + [markdown] id="9NWc0bP5Yj4r"
#
| Education_Capstone_MS2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PE files sorter v1.2
# +
# %%time
import pefile
import os
import array
import sys
import argparse
import os
import shutil
import struct
from os import path
import glob
import time
import sys
import hashlib
from collections import defaultdict
IMAGE_FILE_MACHINE_I386=332
IMAGE_FILE_MACHINE_IA64=512
IMAGE_FILE_MACHINE_AMD64=34404
class bcolors:
BLUE = '\033[94m'
GREEN = '\033[92m'
RED = '\033[91m'
WARNING = '\033[93m'
FAIL = '\033[91m'
ENDC = '\033[0m'
def chunk_reader(fobj, chunk_size=1024):
"""Generator that reads a file in chunks of bytes"""
while True:
chunk = fobj.read(chunk_size)
if not chunk:
return
yield chunk
def md5(fname):
hash_md5 = hashlib.md5()
with open(fname, 'rb') as f:
for chunk in iter(lambda: f.read(4096), b""):
hash_md5.update(chunk)
return hash_md5.hexdigest()
def remove_duplicates(path):
"""relies on the md5 function above to remove duplicate files"""
if not os.path.isdir(path): # make sure the given directory exists
print('specified directory does not exist!')
return
md5_dict = defaultdict(list)
for root, dirs, files in os.walk(path): # the os.walk function allows checking subdirectories too...
for filename in files:
try:
filepath = os.path.join(root, filename)
print(bcolors.GREEN + "Working: %s." % filepath + bcolors.ENDC)
file_md5 = md5(filepath)
md5_dict[file_md5].append(filepath)
except:
print (bcolors.RED + "Working Error" + bcolors.ENDC)
for key in md5_dict:
try:
file_list = md5_dict[key]
while len(file_list) > 1:
item = file_list.pop()
print(bcolors.BLUE + "Duplicate: %s. (removing it...)" % item + bcolors.ENDC)
os.remove(item)
except:
print (bcolors.RED + "Removing Error" + bcolors.ENDC)
print('Done!')
def ChehckIsPEObject(filename):
"""check if object is pe or not"""
# check if the input file exists
if not os.path.exists(filename):
parser.error("cannot access input file '%s'" % args[0])
return False
# check if the input file is executable
if not os.path.isfile(filename):
print ('The given arg is not a file.')
return False
else:
try:
pe = pefile.PE(filename)
if not (pe.is_exe() or pe.is_dll()):
print ('Input file should be executable (PE format: exe or dll)')
return False
except pefile.PEFormatError as e:
print("[-] PEFormatError: %s" % e.value)
return False
return True
def GetBinaryType(filepath):
"""retrieve architecture of pe file 86\64"""
f=open(filepath, "rb")
machine = 0
s=f.read(2)
f.seek(60)
s=f.read(4)
header_offset=struct.unpack("<L", s)[0]
f.seek(header_offset+4)
s=f.read(2)
machine=struct.unpack("<H", s)[0]
f.close()
return machine
if __name__ == '__main__':
# Classify set legitimate \ malware
legitimate = True
# Copy not pe objects to not_pe \ not_pe_clean dirs
copy_not_pe = False
# Clean dublicate files matched by md5 hash
clean_dublicates = False
if legitimate == True:
root_dir = 'Z:/training_set/not_classified_clean/'
not_pe_dir = 'Z:/training_set/not_pe_clean/'
net_pe_dir = 'Z:/training_set/net_pe_clean/'
x32_dll_dir = 'Z:/training_set/x32_dll_clean/'
x64_dll_dir = 'Z:/training_set/x64_dll_clean/'
x32_exe_dir = 'Z:/training_set/x32_exe_clean/'
x64_exe_dir = 'Z:/training_set/x64_exe_clean/'
else:
root_dir = 'Z:/training_set/VirusSignList_Free_140817/'
not_pe_dir = 'Z:/training_set/not_pe/'
net_pe_dir = 'Z:/training_set/net_pe/'
x32_dll_dir = 'Z:/training_set/x32_dll/'
x64_dll_dir = 'Z:/training_set/x64_dll/'
x32_exe_dir = 'Z:/training_set/x32_exe/'
x64_exe_dir = 'Z:/training_set/x64_exe/'
# Remove dublicate files
if clean_dublicates == True:
remove_duplicates(root_dir)
exit()
# scan all files from selected dir
x = [entry for entry in os.scandir(root_dir+'.') if entry.is_file()]
i = 0
for f in x:
print("%d. %s" % (i, f))
is_dotnet = False
# check if valid PE file
if ChehckIsPEObject(f):
# load pe file to pelib
try:
pe = pefile.PE(f)
except:
print("[-] pefile open file error: %s")
# check if PE is dotnet and move to net_pe dir
try:
isDotNet = pe.OPTIONAL_HEADER.DATA_DIRECTORY[14]
if isDotNet.VirtualAddress != 0 and isDotNet.Size != 0:
print (bcolors.BLUE + "%s PE is .NET executable" % (os.path.abspath(f)) + bcolors.ENDC)
shutil.copy(f, net_pe_dir+os.path.basename(f))
is_dotnet = True
except:
print (bcolors.RED + "%s error")
# check if PE is x32 EXE OR DLL and move to dir
if GetBinaryType(f) == IMAGE_FILE_MACHINE_I386:
if(pe.is_exe() and is_dotnet == False):
try:
print (bcolors.GREEN + "%s PE is x32 EXE" % (os.path.abspath(f)) + bcolors.ENDC)
shutil.copy(f, x32_exe_dir+os.path.basename(f))
except:
print (bcolors.RED + "Error" + bcolors.ENDC)
# if PE is x32 DLL
elif(pe.is_dll() and is_dotnet == False):
try:
print (bcolors.GREEN + "%s PE is x32 DLL" % (os.path.abspath(f)) + bcolors.ENDC)
shutil.copy(f, x32_dll_dir+os.path.basename(f))
except:
print (bcolors.RED + "Error" + bcolors.ENDC)
# check if PE is x64 EXE OR DLL and move to dir
elif GetBinaryType(f) == IMAGE_FILE_MACHINE_AMD64:
if(pe.is_exe() and is_dotnet == False):
try:
print (bcolors.GREEN + "%s PE is x64 EXE" % (os.path.abspath(f))+ bcolors.ENDC)
shutil.copy(f, x64_exe_dir+os.path.basename(f))
except:
print (bcolors.RED + "Error" + bcolors.ENDC)
# if PE is x64 DLL
elif(pe.is_dll() and is_dotnet == False):
try:
print (bcolors.GREEN + "%s PE is x64 DLL" % (os.path.abspath(f))+ bcolors.ENDC)
shutil.copy(f, x64_dll_dir+os.path.basename(f))
except:
print (bcolors.RED + "Error" + bcolors.ENDC)
pe.close()
# check if PE is NOT EXE OR DLL and move to dir
else:
print (bcolors.RED + "%s is NOT a PE file" % (os.path.abspath(f))+ bcolors.ENDC)
if copy_not_pe == True:
try:
shutil.copy2(f, not_pe_dir+os.path.basename(f))
except:
print (bcolors.RED + "Error" + bcolors.ENDC)
i = i + 1
print (colors.BLUE + "DONE.")
# -
| PE_Dataset_Sorter_v_1_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Notebook with Environment Variables
#
# This python Notebook contains various environment variables to test the parser functionality.
# +
import os
os.getenv("VAR1")
os.environ["VAR2"]
os.environ.get("VAR3")
print(os.environ['VAR4'])
print(os.getenv("VAR5", 'localhost'))
# +
os.environ['VAR6'] = "value6"
print(os.environ.get('VAR7', 'value7'))
os.getenv('VAR8')
os.environ["VAR1"] = "newvalue"
| elyra/contents/tests/resources/parse_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Pub/Sub Publisher
# https://www.rabbitmq.com/tutorials/tutorial-three-python.html
#
# 
# ## Message Routing
#
# RabbitMQ uses exchanges as the message routing mechanism. In a pub/sub model a publisher will publish a message to an exchange. Subscribers will each have their own queue that they are watching for messages to appear into. Exchanges will then route messages sent by publishers to the appropriate subscriber queues.
import pika
import time
from datetime import datetime
import json
RABBIT_CONNECTION = 'amqp://guest:guest@rabbit:5672/'
EXCHANGE_NAME = 'logs'
def create_channel():
connection = pika.BlockingConnection(
pika.connection.URLParameters(RABBIT_CONNECTION))
channel = connection.channel()
return channel, connection
# ### Fanout exchange
#
# A fanout exchange will send a copy of a published method to each queue with a binding to the exchange. This is what we use for pub/sub so each subscriber will get a copy of the message.
def declare_exchange(channel):
channel.exchange_declare(exchange='logs', exchange_type='fanout')
def publish_log_messages(messages=10):
channel, connection = create_channel()
declare_exchange(channel)
for message in range(1, messages+1):
channel.basic_publish(exchange=EXCHANGE_NAME,
routing_key='',
body=f'Log Message # {message}')
print(f'Log Message # {message} sent')
connection.close()
def periodic_publish(total_messages=200):
channel, connection = create_channel()
declare_exchange(channel)
for iteration in range(1, total_messages+1):
message = {
"timestamp": datetime.now().isoformat(),
"messageNumber": iteration,
"source": "Python Log Publisher"
}
channel.basic_publish(exchange=EXCHANGE_NAME,
routing_key='',
body=json.dumps(message))
print(f'Published JSON log message # {iteration}')
time.sleep(2)
connection.close()
periodic_publish(100)
| notebooks/Pub Sub Publisher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <img src="http://sct.inf.utfsm.cl/wp-content/uploads/2020/04/logo_di.png" style="width:60%">
# <h1> INF-285 - Computación Científica </h1>
# <h2> Newton's Method in $\mathbb{R}^n$ </h2>
# <h2> <a href="#acknowledgements"> [S]cientific [C]omputing [T]eam </a> </h2>
# <h2> Version: 1.04</h2>
# </center>
# <div id='toc' />
#
# ## Table of Contents
# * [Newton's method](#newton)
# * [Python Modules and Functions](#py)
# * [Acknowledgements](#acknowledgements)
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from ipywidgets import interact
# <div id='newton' />
#
# # Newton's method
# [Back to TOC](#toc)
# Using a vectorized implementation
f1 = lambda x: x[0]**2+x[1]**2-1
f2 = lambda x: x[1]-x[0]**2
F = lambda x: np.array([f1(x),f2(x)],dtype=np.float)
J = lambda x: np.array([[2*x[0], 2*x[1]],[-2*x[0], 1.0]],dtype=np.float)
# The Newton Step takes advantage of the vectorized implementation!
# Here we use 'np.linalg.solve', but we could use LU or PALU! Or iterative methods!
NewtonStep = lambda xi: xi-np.linalg.solve(J(xi),F(xi))
# The next cell of code is just needed for plotting purposes.
# +
n_delta = 50
x = np.linspace(-1.5, 1.5, n_delta)
# We could have used 'x' since it is the same, but for completeness we will define 'y'
y = np.linspace(-1.5, 1.5, n_delta)
X, Y = np.meshgrid(x, y)
Z1 = np.zeros_like(X)
Z2 = np.zeros_like(X)
for i,xi in enumerate(x):
for j,yj in enumerate(y):
Z1[j,i] = f1([xi,yj])
Z2[j,i] = f2([xi,yj])
# -
# Here we plot the curves we will be intersecting.
plt.figure()
CS1 = plt.contour(X, Y, Z1,levels=[0])
CS2 = plt.contour(X, Y, Z2,levels=[0])
plt.grid()
plt.axis('equal')
plt.title(r'Newton $\mathbb{R}^n$')
plt.show()
# Here we implement the Newton's method in higher dimension in a widget.
# Most of the code is for plotting and showing the outcome, but the core, i.e. the Newton steps, is just 1 line of code!
def Show_Newton(x0=1.2,y0=0.3,n=0):
plt.figure()
CS1 = plt.contour(X, Y, Z1,levels=[0])
CS2 = plt.contour(X, Y, Z2,levels=[0])
plt.grid()
plt.axis('equal')
plt.title(r'Newton $\mathbb{R}^n$')
plt.plot(x0,y0,'rx')
x_previous = np.array([x0,y0])
print('Initial guess: [%.10f, %.10f]' % (x0,y0))
for i in np.arange(n):
x_next=NewtonStep(x_previous)
x1,y1 = x_next
plt.plot(x1,y1,'rx')
plt.plot([x0, x1],[y0, y1],'r')
x0=x1
y0=y1
x_previous = x_next
print('Iteration %d : [%.10f, %.10f]' % (i+1,x1,y1))
plt.show()
interact(Show_Newton,x0=(-1.4,1.4,0.1),y0=(-1.4,1.4,0.1), n=(0,100,1))
# <div id='py' />
#
# ## Python Modules and Functions
# [Back to TOC](#toc)
# **Newton's method**:
# Here is the reference to the Newton's Method in Scipy, I suggest to take a look!
#
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.newton.html
#
# **Broyden**:
# An alternative method is the Bryden's method. The Broyden's method is analogous to the Secant method in 1D, this means that we replace the approximation of the 'derivate' (which is actually the Jacobian matrix) for an approximation.
#
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.broyden1.html#scipy.optimize.broyden1
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.broyden2.html#scipy.optimize.broyden2
#
# **Minimization**:
# This is an alternative path for solving a square nonlinear system of equations, recall that if there exists a solution to a nonlinear system of equation, it means that there exists a vector $\mathbf{r}$ such that $\mathbf{F}(\mathbf{r})=\mathbf{0}$.
# So, this will be equivalent to finding the minimum of the following function $g(\mathbf{x})=\|\mathbf{F}(\mathbf{x})\|$.
# Thus, we could tackle this problen then as a minimization of $g(\mathbf{x})$.
# It depends on the context if this will be useful or not!
# But it is important to know this is an alternative.
#
# https://docs.scipy.org/doc/scipy/reference/optimize.html
# <div id='acknowledgements' />
#
# # Acknowledgements
# [Back to TOC](#toc)
# * _Material created by professor <NAME>_ (`<EMAIL>`). DI UTFSM. April 2018._
# * _Update June 2020 - v1.15 - C.Torres_ : Fixing formatting issues.
# * _Update May 2021 - v1.02 - C.Torres_ : Fixing formatting issues. Adding link to toc.
# * _Update May 2021 - v1.03 - C.Torres_ : Improving implementation usign vectorization. Adding comments in the 'Python Modules and Functions' section.
# * _Update May 2021 - v1.04 - C.Torres_ : Bug found by <NAME> in function 'Show_Newton', it was missing the line 'x_previous = x_next'. Thanks Jorge! We also updated the way the iteration umber is showed, basically we added +1 to the iterator 'i' in the 'print' function.
| SC1v2/Bonus - 04 - Newton Rn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
img = cv2.imread("dustin.jpg")
print("Resim boyutu:",img.shape)
cv2.imshow("Orijinal",img)
#resized
imgResized = cv2.resize(img,(800,800))
print("Resized Img Shape:", imgResized.shape)
cv2.imshow("Img Resized:",imgResized)
#kırp
imgCropped = img[:200,:300] # height width -->
cv2.imshow("Kirpik Resim",imgCropped)
cv2.waitKey(0)
cv2.destroyAllWindows()
| 1-) Image Processing with Cv2/3-) Resize-Crop Image with CV2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import glob
import re
ann_files = glob.glob("/data/tmp/toothpastes/US_toothpastes/*.txt")
for ann in ann_files:
try:
ann_new = re.sub("\s+", "_", ann)
jpeg_old = re.sub(".txt", ".jpeg", ann)
jpeg_new = re.sub("\s+", "_", jpeg_old)
print(ann, ann_new, jpeg_old, jpeg_new)
os.rename(ann, ann_new)
os.rename(jpeg_old, jpeg_new)
except Exception as e:
print(e)
for jpeg in glob.glob("/data/tmp/toothpastes/US_toothpastes/*.jpeg"):
try:
jpeg_new = re.sub("\s+", "_", jpeg)
os.rename(jpeg, jpeg_new)
except Exception as e:
print(e)
| rename_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import fidutils as vis
fiy=0.83
# transform INC to IND
class fd:
def __init__(self):
self.ul=0
def __call__(self,x):
return ul-x
def set_upper_limit(ul):
self.ul=ul
# -
Er = vis.Experiment_reader()
pth = "out/gn"
Er.set_path(pth)
Er.read_all_expriemts()
ank=1.0/8
ul = Er.get_data("Inc")['gn_001'][0].max()
def fd(x):
return ul-x
Er.print_param_description(0)
ank=1.0/8
# plot IND
Er.annotated_plot2(0,'Inc', zoom=0.8, pad=0, max_hight=3, xybox=None, fd=fd,
figposx=[ank,3*ank,5*ank,7*ank],
figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,16],
ylabel="IND", add_points=True)
# plot FID
Er.annotated_plot2(0,'Fid', zoom=0.8, pad=0, max_hight=300, xybox=None,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,500], ylabel="FID", add_points=True)
Er = vis.Experiment_reader()
pth = "out/sp"
Er.set_path(pth)
Er.read_all_expriemts()
ank=1.0/8
ul = Er.get_data("Inc")['sp_001'][0].max()
def fd(x):
return ul-x
Er.print_param_description(0)
ank=1.0/8
Er.annotated_plot2(0,'Inc', zoom=0.8, pad=0, max_hight=3, xybox=None, fd=fd,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,16],
ylabel="IND", add_points=True)
Er.annotated_plot2(0,'Fid', zoom=0.8, pad=0, max_hight=300, xybox=None,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,650], ylabel="FID", add_points=True)
# +
Er = vis.Experiment_reader()
pth = "out/blur"
Er.set_path(pth)
Er.read_all_expriemts()
ank=1.0/8
ul = Er.get_data("Inc")['blur_001'][0].max()
def fd(x):
return ul-x
Er.print_param_description(0)
ank=1.0/8
Er.annotated_plot2(0,'Inc', zoom=0.8, pad=0, max_hight=3, xybox=None, fd=fd,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,16],
ylabel="IND", add_points=True) #n_datapoints=4)
Er.annotated_plot2(0,'Fid', zoom=0.8, pad=0, max_hight=300, xybox=None,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,500], ylabel="FID", add_points=True)
# +
Er = vis.Experiment_reader()
pth = "out/rect"
Er.set_path(pth)
Er.read_all_expriemts()
ank=1.0/8
ul = Er.get_data("Inc")['rect_001'][0].max()
def fd(x):
return ul-x
Er.print_param_description(0)
ank=1.0/8
Er.annotated_plot2(0,'Inc', zoom=0.8, pad=0, max_hight=3, xybox=None, fd=fd,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,16],
ylabel="IND", add_points=True)
Er.annotated_plot2(0,'Fid', zoom=0.8, pad=0, max_hight=300, xybox=None,
figposx=[ank,3*ank,5*ank,7*ank], figposy=[fiy,fiy,fiy,fiy], xlim=[-0.5,3.5], ylim=[0,400], ylabel="FID", add_points=True)
# -
| FIDvsINC/create_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scratchwork 2
from finite_algebras import *
from cayley_table import *
from permutations import *
import os
aa_path = os.path.join(os.getenv("PYPROJ"), "abstract_algebra")
alg_dir = os.path.join(aa_path, "Algebras")
ex = Examples(alg_dir)
mag_id = ex[10]
mag_id.about(use_table_names=True)
mag_id.center()
n12 = generate_algebra_mod_n(12)
n12.about()
parts12 = about_subalgebras(n12)
n11 = generate_algebra_mod_n(11)
n11.about()
parts11 = about_subalgebras(n11)
# ## Generators
n = 12
alg = generate_cyclic_group(n)
alg_gens = alg.generators()
print(alg.name + " generators:")
print(alg_gens)
print(f"There are {len(alg_gens)} generators.")
n = 11
alg = generate_cyclic_group(n)
alg_gens = alg.generators()
print(alg.name + " generators:")
print(alg_gens)
print(f"There are {len(alg_gens)} generators.")
alg.is_cyclic()
for alg in ex:
print(alg.name)
print(f" {alg.is_cyclic()}")
z5 = generate_cyclic_group(5)
print(z5)
print(f"Generators: {z5.is_cyclic()}")
z5_sqr = z5 * z5
print(z5_sqr)
print(f"Generators: {z5_sqr.is_cyclic()}")
mag_id = ex[10]
mag_id.about(use_table_names=True)
mag_id.is_cyclic()
# ## Center of a Group
q8 = ex[13]
q8.about()
q8_ctr = q8.center_algebra()
q8_ctr.about()
sd16 = ex[14]
sd16.about(max_size=16)
sd16_ctr = sd16.center_algebra()
sd16_ctr.about()
ex141 = ex[11]
ex141.about()
ex141.center_algebra(verbose=True)
for x in ex:
xctr = x.center()
print(x, xctr)
# ## Cancellation
# A Magma, $M = \langle S, \circ \rangle$ is a **division Magma** if $\forall a,b \in S, \exists x,y \in S$ such that $a \circ x = b$ and $y \circ a = b$.
ex.about()
alg = ex[10]
alg.about()
# +
import itertools as it
print(f"\n{alg}\n")
for ab in it.product(alg.elements, alg.elements):
ab_ok = False
for xy in it.product(alg.elements, alg.elements):
a = ab[0]; b = ab[1]; x = xy[0]; y = xy[1]
if alg.op(a, x) == b and alg.op(y, a) == b:
print(f"{ab} & {xy}")
ab_ok = True
break
if not ab_ok:
print(f"{ab} fail")
# +
import itertools as it
def is_division_algebra(alg, verbose=False):
if verbose:
print(f"\n{alg}\n")
result = True
for ab in it.product(alg.elements, alg.elements):
ab_ok = False
for xy in it.product(alg.elements, alg.elements):
a = ab[0]; b = ab[1]; x = xy[0]; y = xy[1]
if alg.op(a, x) == b and alg.op(y, a) == b:
if verbose:
print(f"{ab} & {xy}")
ab_ok = True
break
if not ab_ok:
result = False
if verbose:
print(f"{ab} fail")
return result
# -
is_division_algebra(ex[4], verbose=True)
for alg in ex:
print(f"{alg.name}? {yes_or_no(is_division_algebra(alg))}")
ex.about()
# ## Regularity in Semigroups
# See the paper by <NAME> (ref below)
#
# A Semigroup, $\langle S, \circ \rangle$ is **regular** if $\forall a \in S, \exists x \in S$ such that $a \circ x \circ a = a$.
alg = ex[1]
alg.about()
# +
a = alg.elements[4]
print(f"a = {a}")
print(f"a_inv = {alg.inv(a)}")
[x for x in alg if alg.op(alg.op(a, x), a) == a]
print([alg.op(alg.op(a, x), a) == a for x in alg])
any([alg.op(alg.op(a, x), a) == a for x in alg])
# + active=""
# def is_regular(alg):
# return all([any([alg.op(alg.op(a, x), a) == a for x in alg]) for a in alg])
# -
for alg in ex:
#print(alg.name, all([any([alg.op(alg.op(a, x), a) == a for x in alg]) for a in alg]))
if isinstance(alg, Semigroup):
print(alg.name, alg.is_regular())
alg = generate_commutative_monoid(8)
alg.about()
# + active=""
# def weak_inverses(alg):
# return {a:[x for x in alg if alg.op(alg.op(a, x), a) == a] for a in alg}
# -
alg.is_regular()
alg.weak_inverses()
[i for i in range(1, 50) if not generate_commutative_monoid(i).is_regular()]
# + active=""
# What is the pattern here w.r.t. non-regular Monoid orders?
#
# [i for i in range(1, 50) if not generate_commutative_monoid(i).is_regular()]
#
# [4, 8, 9, 12, 16, 18, 20, 24, 25, 27, 28, 32, 36, 40, 44, 45, 48, 49]
# -
# See https://en.wikipedia.org/wiki/Cancellation_property
#
# and https://math.stackexchange.com/questions/4008196/defining-loops-why-is-divisibility-and-identitiy-implying-invertibility
#
# and ["Why Study Semigroups?" by <NAME>](http://www.thebookshelf.auckland.ac.nz/docs/Maths/PDF2/mathschron016-001.pdf)
| notebooks/scratchwork2.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GPflow 2 Upgrade Guide
#
# This is a basic guide for people who have GPflow 1 code that needs to be upgraded to GPflow 2.
# Also see the [Intro to GPflow with TensorFlow 2 notebook](intro_to_gpflow2.ipynb).
# ## Kernel Input Dims
#
# The `input_dim` parameter has been removed from the `Kernel` class’s initialiser. Therefore all calls to create a kernel must be changed to remove the `input_dim` parameter.
#
# For example:
#
# ```diff
# -gpflow.kernels.SquaredExponential(1, variance=1.0, lengthscales=0.5)
# +gpflow.kernels.SquaredExponential(variance=1.0, lengthscales=0.5)
# ```
#
# **Note**: old code may still run without obvious errors against GPflow, since many kernels take an optional numerical value as their first parameter. You may not get the result you expect though!
# ## Parameter and tf.Variable
#
# The `Parameter` class in GPflow 1 was a separate class from `tf.Variable`. The `params_as_tensors` decorator or the `params_as_tensors_for` context manager were required to turn them into a tensor that could be consumed by TensorFlow operations.
#
# In GPflow 2, `Parameter` inherits from `gpflow.Module` (a `tf.Module` subclass) that wraps a `tf.Variable`, and can directly be used in place of a tensor, so no such conversion is necessary.
#
# References to `params_as_tensors` and `params_as_tensors_for` can simply be removed.
#
# ## Parameter Assignment
#
# In GPflow 2 the semantics of assigning values to parameters has changed. It is now necessary to use the Parameter.assign method rather than assigning values directly to parameters. For example:
#
# ```diff
# # Initializations:
# -likelihood.scale = 0.1
# +likelihood.scale.assign(0.1)
# ```
#
# In the above example, the old (GPflow 1) code would have assigned the value of `likelihood.scale` to 0.1 (assuming that likelihood is a `Parameterized` object and scale is a `Parameter`), rather than replacing the `scale` attribute with a Python float (which would be the “normal” Python behaviour). This maintains the properties of the parameter. For example, it remains trainable etc.
#
# In GPflow 2, it is necessary to use the `Parameter.assign` method explicitly to maintain the same behaviour, otherwise the parameter attribute will be replaced by an (untrainable) constant value.
#
# To change other properties of the parameter (for example, to change transforms etc) you may need to replace the entire parameter object. See [this notebook](understanding/models.ipynb#Constraints-and-trainable-variables) for further details.
# ## Parameter trainable status
#
# A parameter's `trainable` attribute cannot be set. Instead, use `gpflow.set_trainable()`. E.g.:
# ```diff
# -likelihood.trainable = False
# +gpflow.set_trainable(likelihood, False)
# ```
# ## SciPy Optimizer
#
# Usage of GPflow’s Scipy optimizer has changed. It has been renamed from `gpflow.train.ScipyOptimizer` to `gpflow.optimizers.Scipy` and its `minimize` method has changed in the following ways:
#
# * Instead of a GPflow model, the method now takes a zero-argument function that returns the loss to be minimised (most GPflow models provide a `model.training_loss` method for this use-case; gpflow.models.SVGP does not encapsulate data and provides a `model.training_loss_closure(data)` closure generating method instead), as well as the variables to be optimised (typically `model.trainable_variables`).
# * The options (`disp`, `maxiter`) must now be passed in a dictionary.
#
# For example:
# ```diff
# -optimizer = gpflow.train.ScipyOptimizer()
# -optimizer.minimize(model, disp=True, maxiter=100)
# +optimizer = gpflow.optimizers.Scipy()
# +optimizer.minimize(
# + model.training_loss,
# + variables=model.trainable_variables,
# + options=dict(disp=True, maxiter=100),
# +)
# ```
#
# Any additional keyword arguments that are passed to the `minimize` method are passed directly through to the [SciPy optimizer's minimize method](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html).
# ## Model Initialisers
#
# In many cases the initialiser for the model will have changed. Typical changes include:
#
# * Instead of separate parameters for `X` and `Y`, some models now require a single `data` parameter containing a tuple of the X and Y data.
# * The `kern` parameter has been renamed to `kernel`.
#
# For example, for the `GPR` model:
# ```diff
# -model = GPR(X, Y, kern=kernel)
# +model = GPR(data=(X, Y), kernel=kernel)
# ```
#
# Models that do not take a `likelihood` argument because they hard-code a Gaussian likelihood (GPR, SGPR) now take a `noise_variance` argument that sets the initial value of the likelihood variance.
# ## SVGP Initialiser
#
# The SVGP model’s initialiser no longer accepts X and Y data. Instead this data must be passed to the various computation methods of the model (`elbo`, `training_loss` etc).
#
# In the [Introduction to GPflow 2 notebook](intro_to_gpflow2.ipynb) there is an example of how to use SVGP with optimisation using mini-batches of data.
#
# In addition, SVGP’s `Z` parameter has been removed. To pass-in inducing points use the `inducing_variable` parameter. Also `SVGP`'s `feature` attribute has been renamed to `inducing_variable`.
# ## Autoflow
#
# The `@autoflow` decorator has been removed. Since eager execution is the default in TensorFlow 2 this is no longer necessary.
#
# You may wish to consider wrapping functions that were previously wrapped in the `@autoflow` decorator in the `tf.function` decorator instead, to improve performance (but this is not necessary from a functionality point of view).
# ## Use of tf.function
#
# Wrapping compute-heavy operations such as calculating a model objective or even the optimizer steps (such as `tf.optimizers.Adam().minimize()`) with `tf.function` is crucial for efficient computation.
#
# **Note**: you should ensure that functions wrapped in `tf.function` are only passed **tensors** (not numpy arrays or other data structures, with the exception of a small number of bool or enum-style flags), or the decorator will re-compile the graph each time the function is passed new objects as its arguments. See the [TensorFlow documentation on re-tracing](https://www.tensorflow.org/guide/function#re-tracing) for further details.
#
# You can convert a numpy array to a tensor by using `tf.constant`. For example: `compiled_function(tf.constant(numpy_array))`.
# ## Model Compilation
#
# Models no longer need to be compiled before use. Remove all calls to the `compile` method.
# ## Sessions and Graphs
#
# GPflow only supports eager execution, which is the default in TensorFlow 2. It does not support graph mode, which was the default execution mode in TensorFlow 1. Therefore all references to Sessions and Graphs should be removed. You should also remove references to the `gpflow.reset_default_graph_and_session` function.
#
# **Warning**: code that creates graphs (for example `tf.Graph().as_default()`) will disable eager execution, which will not work well with GPflow 2. If you get errors like “'Tensor' object has no attribute 'numpy'” then you may not have removed all references to graphs and sessions in your code.
# ## Defer Build
#
# The `defer_build` context manager has been removed. References to it can simply be removed.
# ## Return Types from Auto-flowed Methods
#
# GPflow methods that used the `@autoflow` decorator, like for example `predict_f` and `predict_y`, will previously have returned NumPy Arrays. These now return TensorFlow tensors. In many cases these can be used like NumPy arrays (they can be passed directly to many of NumPy’s functions and even be plotted by matplotlib), but to actually turn them into numpy arrays you will need to call `.numpy()` on them.
#
# For example:
# ```diff
# def _predict(self, features: np.ndarray) -> Tuple[np.ndarray, np.ndarray]:
# mean, variance = self._model.predict_f(features)
# - return mean, variance
# + return mean.numpy(), variance.numpy()
# ```
# ## Parameter Values
#
# In GPflow 1, `Parameter.value` was a property that returned the numpy (`np.ndarray`) representation of the value of the Parameter.
#
# In GPflow 2, `Parameter` behaves similar to TensorFlow's `tf.Variable`: `Parameter.value()` is a method that returns a constant tf.Tensor with the current (constrained) value of the Parameter. To obtain the *numpy* representation, use the `Parameter.numpy()` method:
#
# For example:
# ```diff
# -std_dev = np.sqrt(model.likelihood.variance.value)
# +std_dev = np.sqrt(model.likelihood.variance.numpy())
# ```
# ## Model Class
#
# The `Model` class has been removed. A suitable replacement, for those models that do not wish to inherit from `GPModel`, may be `BayesianModel`.
# ## Periodic Base Kernel
#
# The base kernel for the `Periodic` kernel must now be specified explicitly. Previously the default was `SquaredExponential`, so to maintain the same behaviour as before this must be passed-in to the `Periodic` kernel’s initialiser (note that `active_dims` is specified in the base kernel).
#
# For example:
# ```diff
# -Periodic(1, active_dims=[2])
# +Periodic(SquaredExponential(active_dims=[2]))
# ```
# ## Predict Full Covariance
#
# The `predict_f_full_cov` method has been removed from `GPModel`. Instead, pass `full_cov=True` to the `predict_f` method.
#
# For example:
# ```diff
# -f_mean, f_cov = model.predict_f_full_cov(X)
# +f_mean, f_cov = model.predict_f(X, full_cov=True)
# ```
# ## Predictive (log)density
#
# The `predict_density` method of GPModels and Likelihoods has been renamed to `predict_log_density`. (It always returned the predictive *log*-density, so no change in behaviour.)
# ## Settings / Configuration
#
# In GPflow 2, the `gpflow.settings` module and the `gpflowrc` file have been removed. Instead, there is `gpflow.config`.
#
# `gpflow.settings.float_type` has changed to `gpflow.default_float()` and `gpflow.settings.int_type` has changed to `gpflow.default_int()`.
# `gpflow.settings.jitter`/`gpflow.settings.numerics.jitter_level` has changed to `gpflow.default_jitter()`.
#
# These default settings can be changed using environment variables (`GPFLOW_FLOAT`, `GPFLOW_INT`, `GPFLOW_JITTER`, etc.) or function calls (`gpflow.config.set_default_float()` etc.). There is also a `gpflow.config.as_context()` context manager for temporarily changing settings for only part of the code.
#
# See the `gpflow.config` API documentation for more details.
#
# ## Data Types
#
# In some cases TensorFlow will try to figure out an appropriate data type for certain variables. If Python floats have been used, TensorFlow may default these variables to `float32`, which can cause incompatibilities with GPflow, which defaults to using `float64`.
#
# To resolve this you can use `tf.constant` instead of a Python float, and explicitly specify the data type, e.g.
# ```python
# tf.constant(0.1, dtype=gpflow.default_float())
# ```
# ## Transforms
#
# These have been removed in favour of the tools in `tensorflow_probability.bijectors`. See for example [this Stackoverflow post](https://stackoverflow.com/q/58903446/5986907).
#
# GPflow 2 still provides the `gpflow.utilities.triangular` alias for `tfp.bijectors.FillTriangular`.
#
# To constrain parameters to be positive, there is `gpflow.utilities.positive` which is configurable to be either softplus or exp, with an optional shift to ensure a lower bound that is larger than zero.
# Note that the default lower bound used to be `1e-6`; by default, the lower bound if not specified explicitly is now `0.0`. Revert the previous behaviour using `gpflow.config.set_default_positive_minimum(1e-6)`.
# ## Stationary kernel subclasses
#
# Most stationary kernels are actually *isotropic*-stationary kernels, and should now subclass from `gpflow.kernels.IsotropicStationary` instead of `gpflow.kernels.Stationary`. (The `Cosine` kernel is an example of a non-isotropic stationary kernel that depends on the direction, not just the norm, of $\mathbf{x} - \mathbf{x}'$.)
# ## Likelihoods
#
# We cleaned up the likelihood API. Likelihoods now explicitly define the expected number of outputs (`observation_dim`) and latent functions (`latent_dim`), and shape-checking is in place by default.
#
# Most of the likelihoods simply broadcasted over outputs; these have now been grouped to subclass from `gpflow.likelihoods.ScalarLikelihood`, and implementations have been moved to leading-underscore functions. `ScalarLikelihood` subclasses need to implement at least `_scalar_log_prob` (previously `logp`), `_conditional_mean`, and `_conditional_variance`.
#
# The likelihood `log_prob`, `predict_log_density`, and `variational_expectations` methods now return a single value per data row; for `ScalarLikelihood` subclasses this means these methods effectively sum over the observation dimension (multiple outputs for the same input).
# ## Priors
#
# Priors used to be defined on the *unconstrained* variable. The default has changed to the prior to be defined on the *constrained* parameter value; this can be changed by passing the `prior_on` argument to `gpflow.Parameter()`. See the [MCMC notebook](advanced/mcmc.ipynb) for more details.
# ## Name Scoping
#
# The `name_scope` decorator does not exist in GPflow 2 anymore. Use TensorFlow’s [`name_scope`](https://www.tensorflow.org/api_docs/python/tf/name_scope?version=stable) context manager instead.
# ## Model Persistence
#
# Model persistence with `gpflow.saver` has been removed in GPflow 2, in favour of TensorFlow 2’s [checkpointing](https://www.tensorflow.org/guide/checkpoint) and [model persistence using the SavedModel format](https://www.tensorflow.org/guide/saved_model).
#
# There is currently a bug in saving GPflow models with TensorFlow's model persistence (SavedModels). See https://github.com/GPflow/GPflow/issues/1127 for more details; a workaround is to replace all trainable parameters with constants using `gpflow.utilities.freeze(model)`.
#
# Checkpointing works fine.
| doc/source/notebooks/gpflow2_upgrade_guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# Import necessary packages
import pandas as pd
import matplotlib.pyplot as plt
import mplleaflet, pydotplus
from math import radians, cos, sin, asin, sqrt
from ibmgeohashpy import geohash
import numpy as np
from mpl_toolkits.basemap import Basemap
from matplotlib.colors import ListedColormap
import os, sys, json, re, itertools, warnings, folium
from matplotlib import animation
from IPython.display import Image
# %matplotlib inline
from sklearn import tree
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
warnings.filterwarnings('ignore')
print('Libraries imported..')
# +
# Setup a Spark instance from Anaconda Python framework through the interface PySpark
# Use Machine Learning Library MLLib from Pyspark
spark_home = os.environ.get('SPARK_HOME', None)
sys.path.append(spark_home + "/python")
sys.path.append(os.path.join(spark_home, 'python/lib/py4j-0.9-src.zip'))
filename = os.path.join(spark_home, 'python/pyspark/shell.py')
exec(compile(open(filename, "rb").read(), filename, 'exec'))
from pyspark.mllib.fpm import FPGrowth
from pyspark.mllib.clustering import KMeans
print('Spark instance running..')
# -
train = pd.read_csv('data/gender_age_train.csv')
events_within_china = pd.read_csv('./data/events_within_china.csv')
data = events_within_china.merge(train, on = 'device_id', how = 'inner')
# Remove NA from data
data = data.dropna(how='any').drop_duplicates()
print (data.shape)
print (data.columns)
data.head()
geohashes=[]
for x,y in zip(data.latitude,data.longitude):
geohashes.append(geohash.encode(x,y,6))
data['geohash_6'] = geohashes
data = data[['device_id','gender','age','group','geohash_6','event_id']]
data.to_csv('gender_age_train_data_geohash_6_05022017.txt', sep=' ', index=False, header=False)
data.head()
data.shape
# Create training dataset with a format that FPGrowthModel understands
rdd = sc.textFile("gender_age_train_data_geohash_6_05022017.txt")
transactions = rdd.map(lambda line: line.strip().split(' '))
unique = transactions.map(lambda x: list(set(x))).cache()
print('Generating RDD in Spark for obtaining Frequent itemsets....')
numpart,minsup,validfreq = [],[],[]
for i in np.arange(2,6,2):
for j in np.arange(0.00002,0.00015,0.00001):
model = FPGrowth.train(unique, minSupport=j, numPartitions=i)
result = model.freqItemsets().collect()
filename = 'freqItemSets_train_data_05022017_ms_0_0000'+str(int(j*100000))+'_np_'+str(i)+'.txt'
outfile = open(filename, 'w')
for item in result:
outfile.write("%s\n" % str(item))
#print('Generating freqitemsets for min_support='+str(j)+'no_of_partitions='+str(i)+'..')
text_file = open(filename, "r")
freqitems = text_file.read().split('\n')
validitems = []
for item in freqitems:
if len(item)>73:
validitems.append(item)
print('Total no of valid frequent itemsets for min_support:'+str(j)+' and '+str(i)+' partitions= '+str(len(validitems)))
numpart.append(i)
minsup.append(j)
validfreq.append(len(validitems))
freqsets = []
for item in validitems:
# if the string matches the word 'freq' pick the value starting from '=' until ')'
x = re.search("freq", item).end()
# convert the extracted value to a number and check if it is > 50
item1 = int(item[x+1:len(item)-1])
#print(item1)
if item1>50:
freqsets.append(item)
newitems = []
for item in freqsets:
x = int(re.search("]",item).start())
group = item[19:x]
freq = item[x+8:len(item)-1]
newitem = group + ', ' + freq
newitems.append(newitem)
filename1 = 'formatted_'+filename
outfile1 = open(filename1, 'w')
for item in newitems:
if len(item)>53: #63 for 12-char geohash
outfile1.write("%s\n" % str(item))
outfile1.close()
filepath1 = 'formatted_freqItemSets_train_data_05022017_ms_0_0000'+str(int(j*100000))+'_np_'+str(i)+'.txt'
df = pd.read_csv(filepath1, header = None)
df1 = df
df1[0] = df1[0].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[1] = df1[1].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[2] = df1[2].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[3]= df1[3].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[4]= df1[4].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1.columns = ['geohashes','device_id','age','group','gender','freq']
for item in range(0,len(df1)):
if len(df1.iloc[item,0])>15:
#print('\n',df1.iloc[i,0])
a = df1.iloc[item,0]
b = df1.iloc[item,1]
df1.iloc[item,0] = b
df1.iloc[item,1] = a
if len(df1.iloc[item,2])>2:
#print('\n',df1.iloc[i,0])
a1 = df1.iloc[item,2]
b1 = df1.iloc[item,3]
df1.iloc[item,2] = b1
df1.iloc[item,3] = a1
#Convert the age and device_id columns to integer type and sort by frequency
df1['age'] = df1['age'].map(lambda x: int(x))
df1['device_id'] = df1['device_id'].map(lambda x: int(x))
df1 = df1.sort_values(by = 'freq', inplace = False, axis = 0, ascending= False)
# Assign numeric values to gender M := 0 and F := 1
for k in range(0,len(df1)):
if df1.iloc[k,4] == 'M':
df1.iloc[k,4] = 0
else:
df1.iloc[k,4] = 1
#Assign numeric values to groups
#print('\nUnique Groups : \n\n M22-: 1, M23-26 : 2, M27-28: 3, M29-31: 4, M32-38: 5, M39+: 6')
#print('\n F23-: 7, F24-26 : 8, F27-28: 9, F29-32: 10, F33-42: 11, F43+: 12 ')
for i1 in range(0,len(df1)):
if df1.iloc[i1,3] == 'M22-':
df1.iloc[i1,3] = 1
elif df1.iloc[i1,3] == 'M23-26':
df1.iloc[i1,3] = 2
elif df1.iloc[i1,3] == 'M27-28':
df1.iloc[i1,3] = 3
elif df1.iloc[i1,3] == 'M29-31':
df1.iloc[i1,3] = 4
elif df1.iloc[i1,3] == 'M32-38':
df1.iloc[i1,3] = 5
elif df1.iloc[i1,3] == 'M39+':
df1.iloc[i1,3] = 6
elif df1.iloc[i1,3] == 'F23-':
df1.iloc[i1,3] = 7
elif df1.iloc[i1,3] == 'F24-26':
df1.iloc[i1,3] = 8
elif df1.iloc[i1,3] == 'F27-28':
df1.iloc[i1,3] = 9
elif df1.iloc[i1,3] == 'F29-32':
df1.iloc[i1,3] = 10
elif df1.iloc[i1,3] == 'F33-42':
df1.iloc[i1,3] = 11
elif df1.iloc[i1,3] == 'F43+':
df1.iloc[i1,3] = 12
filepath2 = 'formatted_'+filepath1[28:-4]+'.csv'
df1.to_csv(filepath2, index = False)
print('Created formatted freq-itemset..')
for i in range(0,len(df)):
if len(df.iloc[i,2])!=2:
print(i)
df.to_csv('formatted_freqItemSets_train_data_05022017_ms_0_00002_np_2.txt')
# For those items in the freqitemsets where device_id comes before geohash, swap the values
for i in np.arange(2,10,2):
for j in np.arange(0.00002,0.00013,0.00001):
filepath1 = 'formatted_freqItemSets_train_data_05022017_ms_0_0000'+str(j*100000)+'_np_'+str(i)+'.txt'
df = pd.read_csv(filepath1, header = None)
df1 = df
# print('Creating a pandas dataframe using the frequently occurring geohashes..')
# print('\n Number of frequent itemsets :',df1.shape[0],'\n')
#Format the string by removing extra spaces and quotes
df1[0] = df1[0].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[1] = df1[1].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[2] = df1[2].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[3]= df1[3].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1[4]= df1[4].map(lambda x: x.lstrip(' \'').rstrip('\''))
df1.columns = ['geohashes','device_id','age','group','gender','freq']
#print(df1.head())
#print('\nRearranging data correctly from frequent itemsets..')
# Reshuffle geohash, device_id and age,group columns to synchronise the data
for item in range(0,len(df1)):
if len(df1.iloc[item,0])>15:
#print('\n',df1.iloc[i,0])
a = df1.iloc[item,0]
b = df1.iloc[item,1]
df1.iloc[item,0] = b
df1.iloc[item,1] = a
if len(df1.iloc[item,2])>2:
#print('\n',df1.iloc[i,0])
a1 = df1.iloc[item,2]
b1 = df1.iloc[item,3]
df1.iloc[item,2] = b1
df1.iloc[item,3] = a1
#print('\nReshuffling complete..')
#Convert the age and device_id columns to integer type and sort by frequency
df1['age'] = df1['age'].map(lambda x: int(x))
df1['device_id'] = df1['device_id'].map(lambda x: int(x))
df1 = df1.sort_values(by = 'freq', inplace = False, axis = 0, ascending= False)
#print('\nConverting categorical values to numeric values for further analyses.\n \nMale=0 & Female =1')
# Assign numeric values to gender M := 0 and F := 1
for k in range(0,len(df1)):
if df1.iloc[k,4] == 'M':
df1.iloc[k,4] = 0
else:
df1.iloc[k,4] = 1
#Assign numeric values to groups
#print('\nUnique Groups : \n\n M22-: 1, M23-26 : 2, M27-28: 3, M29-31: 4, M32-38: 5, M39+: 6')
#print('\n F23-: 7, F24-26 : 8, F27-28: 9, F29-32: 10, F33-42: 11, F43+: 12 ')
for i1 in range(0,len(df1)):
if df1.iloc[i1,3] == 'M22-':
df1.iloc[i1,3] = 1
elif df1.iloc[i1,3] == 'M23-26':
df1.iloc[i1,3] = 2
elif df1.iloc[i1,3] == 'M27-28':
df1.iloc[i1,3] = 3
elif df1.iloc[i1,3] == 'M29-31':
df1.iloc[i1,3] = 4
elif df1.iloc[i1,3] == 'M32-38':
df1.iloc[i1,3] = 5
elif df1.iloc[i1,3] == 'M39+':
df1.iloc[i1,3] = 6
elif df1.iloc[i1,3] == 'F23-':
df1.iloc[i1,3] = 7
elif df1.iloc[i1,3] == 'F24-26':
df1.iloc[i1,3] = 8
elif df1.iloc[i1,3] == 'F27-28':
df1.iloc[i1,3] = 9
elif df1.iloc[i1,3] == 'F29-32':
df1.iloc[i1,3] = 10
elif df1.iloc[i1,3] == 'F33-42':
df1.iloc[i1,3] = 11
elif df1.iloc[i1,3] == 'F43+':
df1.iloc[i1,3] = 12
filepath2 = 'formatted_'+filepath1[28:-4]+'.csv'
df1.to_csv(filepath2, index = False)
print(filepath2)
numpart
df_fpg = pd.read_csv('formatted__data_02022017_ms_0_000012.0_np_4.csv')
df_fpg.head()
import numpy as np
import seaborn as sb
numpartition = 4
freqsets1 = [25310,19258,15130,12650,10771,9135,8045,7045,6398,5594,5059]
freqsets2 = [25310,19246,15130,12653,10771,9140,8047,7045,6403,5592,5063]
minsup = np.arange(0.00002,0.00013,0.00001)
#plt.plot(minsup,freqsets1,color='k',lw=2)
plt.plot(minsup,freqsets2,color='k',lw=4)
plt.xlabel('Minimum support')
plt.ylabel('No. of Frequent Itemsets')
#plt.title('# of Frequent geohashes v/s minsupport for FPGrowth Model')
plt.show()
| Demographic_Prediction_FPGrowth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''ds'': conda)'
# metadata:
# interpreter:
# hash: 1f33512659712c6a6ac90d8f6217146f941852120cd4340b6d127417f394cbe5
# name: python3
# ---
# ## Filter Methods - Univariate feature selection - Classification
#
# Univariate feature selection works by selecting the best features based on univariate statistical tests (ANOVA). The methods based on F-test estimate the degree of linear dependency between two random variables. They assume a linear relationship between the feature and the target. These methods also assume that the variables follow a Gaussian distribution.
#
# These may not always be the case for the variables in your dataset, so if looking to implement these procedure, you will need to corroborate these assumptions.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import f_classif
from sklearn.feature_selection import SelectKBest
# -
# ### load dataset
# +
# load dataset and features from previus method
features = np.load('../features/featuresFromMIClassif.npy').tolist()
data = pd.read_pickle('../../data/features/features.pkl').loc[:,features].sample(frac=0.35).fillna(-9999)
# +
# In practice, feature selection should be done after data pre-processing,
# so ideally, all the categorical variables are encoded into numbers,
# and then you can assess how deterministic they are of the target
# here for simplicity I will use only numerical variables
# select numerical columns:
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
numerical_vars = list(data.select_dtypes(include=numerics).columns)
data = data[numerical_vars]
data.shape
# -
# ### split train - test
# +
# In all feature selection procedures, it is good practice to select the features by examining only the training set. And this is to avoid overfit.
# separate train and test sets
X_train, X_test, y_train, y_test = train_test_split(
data.drop(labels=['target'], axis=1),
data['target'],
test_size=0.3,
random_state=0)
X_train.shape, X_test.shape
# -
# ### calculate univariate statistical
# +
# calculate the univariate statistical measure between
# each of the variables and the target
# similarly to chi2, the output is the array of f-scores
# and an array of pvalues, which are the ones we will compare
univariate = f_classif(X_train.fillna(0), y_train)
univariate
# -
# let's add the variable names and order it for clearer visualisation
univariate = pd.Series(univariate[1])
univariate.index = X_train.columns
univariate.sort_values(ascending=False, inplace=True)
# and now let's plot the p values
univariate.sort_values(ascending=False).plot.bar(figsize=(20, 8))
pass
# Remember that the lower the p_value, the most predictive the feature is in principle. Features towards the left with pvalues above 0.05, which are candidates to be removed, as this means that the features do not statistically significantly discriminate the target.
# Further investigation is needed if we want to know the true nature of the relationship between feature and target.
#
# In big datasets it is not unusual that the pvalues of the different features are really small. This does not say as much about the relevance of the feature. Mostly it indicates that it is a big the dataset.
#
# ### save features
# +
# how many var would you like to keep from the previous ANOVA analysis
NNUMVAR = 10
sel_ = SelectKBest(f_classif, k=NNUMVAR).fit(X_train.fillna(0), y_train)
features_to_keep = X_train.columns[sel_.get_support()].tolist()
# -
np.save('../features/featuresFromUnivariateClassif.npy',features_to_keep)
| notebooks/feature_selection/filter_methods/univariate_anova_classif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# # SA13 Selection Functions
# ## Depth maps and selection functions
#
# The simplest selection function available is the field MOC which specifies the area for which there is Herschel data. Each pristine catalogue also has a MOC defining the area for which that data is available.
#
# The next stage is to provide mean flux standard deviations which act as a proxy for the catalogue's 5$\sigma$ depth
from herschelhelp_internal import git_version
print("This notebook was run with herschelhelp_internal version: \n{}".format(git_version()))
import datetime
print("This notebook was executed on: \n{}".format(datetime.datetime.now()))
# +
# %matplotlib inline
# #%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
import os
import time
from astropy import units as u
from astropy.coordinates import SkyCoord
from astropy.table import Column, Table, join
import numpy as np
from pymoc import MOC
import healpy as hp
#import pandas as pd #Astropy has group_by function so apandas isn't required.
import seaborn as sns
import warnings
#We ignore warnings - this is a little dangerous but a huge number of warnings are generated by empty cells later
warnings.filterwarnings('ignore')
from herschelhelp_internal.utils import inMoc, coords_to_hpidx, flux_to_mag
from herschelhelp_internal.masterlist import find_last_ml_suffix, nb_ccplots
from astropy.io.votable import parse_single_table
# -
FIELD = 'SA13'
#FILTERS_DIR = "/Users/rs548/GitHub/herschelhelp_python/database_builder/filters/"
FILTERS_DIR = "/opt/herschelhelp_python/database_builder/filters/"
# +
TMP_DIR = os.environ.get('TMP_DIR', "./data_tmp")
OUT_DIR = os.environ.get('OUT_DIR', "./data")
SUFFIX = find_last_ml_suffix()
#SUFFIX = "20171016"
master_catalogue_filename = "master_catalogue_{}_{}.fits".format(FIELD.lower(), SUFFIX)
master_catalogue = Table.read("{}/{}".format(OUT_DIR, master_catalogue_filename))
print("Depth maps produced using: {}".format(master_catalogue_filename))
ORDER = 10
#TODO write code to decide on appropriate order
field_moc = MOC(filename="../../dmu2/dmu2_field_coverages/{}_MOC.fits".format(FIELD))
# -
# ## I - Group masterlist objects by healpix cell and calculate depths
# We add a column to the masterlist catalogue for the target order healpix cell <i>per object</i>.
#Add a column to the catalogue with the order=ORDER hp_idx
master_catalogue.add_column(Column(data=coords_to_hpidx(master_catalogue['ra'],
master_catalogue['dec'],
ORDER),
name="hp_idx_O_{}".format(str(ORDER))
)
)
# +
# Convert catalogue to pandas and group by the order=ORDER pixel
group = master_catalogue.group_by(["hp_idx_O_{}".format(str(ORDER))])
# +
#Downgrade the groups from order=ORDER to order=13 and then fill out the appropriate cells
#hp.pixelfunc.ud_grade([2599293, 2599294], nside_out=hp.order2nside(13))
# -
# ## II Create a table of all Order=13 healpix cells in the field and populate it
# We create a table with every order=13 healpix cell in the field MOC. We then calculate the healpix cell at lower order that the order=13 cell is in. We then fill in the depth at every order=13 cell as calculated for the lower order cell that that the order=13 cell is inside.
depths = Table()
depths['hp_idx_O_13'] = list(field_moc.flattened(13))
depths[:10].show_in_notebook()
depths.add_column(Column(data=hp.pixelfunc.ang2pix(2**ORDER,
hp.pixelfunc.pix2ang(2**13, depths['hp_idx_O_13'], nest=True)[0],
hp.pixelfunc.pix2ang(2**13, depths['hp_idx_O_13'], nest=True)[1],
nest = True),
name="hp_idx_O_{}".format(str(ORDER))
)
)
depths[:10].show_in_notebook()
# +
for col in master_catalogue.colnames:
if col.startswith("f_"):
errcol = "ferr{}".format(col[1:])
depths = join(depths,
group["hp_idx_O_{}".format(str(ORDER)), errcol].groups.aggregate(np.nanmean),
join_type='left')
depths[errcol].name = errcol + "_mean"
depths = join(depths,
group["hp_idx_O_{}".format(str(ORDER)), col].groups.aggregate(lambda x: np.nanpercentile(x, 90.)),
join_type='left')
depths[col].name = col + "_p90"
depths[:10].show_in_notebook()
# -
# ## III - Save the depth map table
depths.write("{}/depths_{}_{}.fits".format(OUT_DIR, FIELD.lower(), SUFFIX))
# ## IV - Overview plots
#
# ### IV.a - Filters
# First we simply plot all the filters available on this field to give an overview of coverage.
tot_bands = [column[2:] for column in master_catalogue.colnames
if (column.startswith('f_') & ~column.startswith('f_ap_'))]
ap_bands = [column[5:] for column in master_catalogue.colnames
if column.startswith('f_ap_') ]
bands = set(tot_bands) | set(ap_bands)
bands
for b in bands:
plt.plot(Table(data = parse_single_table(FILTERS_DIR + b + '.xml').array.data)['Wavelength']
,Table(data = parse_single_table(FILTERS_DIR + b + '.xml').array.data)['Transmission']
, label=b)
plt.xlabel('Wavelength ($\AA$)')
plt.ylabel('Transmission')
plt.xscale('log')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('Passbands on {}'.format(FIELD))
# ### IV.a - Depth overview
# Then we plot the mean depths available across the area a given band is available
# +
average_depths = []
for b in ap_bands:
mean_err = np.nanmean(depths['ferr_ap_{}_mean'.format(b)])
print("{}: mean flux error: {}, 3sigma in AB mag (Aperture): {}".format(b, mean_err, flux_to_mag(3.0*mean_err*1.e-6)[0]))
average_depths += [('ap_' + b, flux_to_mag(1.0*mean_err*1.e-6)[0],
flux_to_mag(3.0*mean_err*1.e-6)[0],
flux_to_mag(5.0*mean_err*1.e-6)[0])]
for b in tot_bands:
mean_err = np.nanmean(depths['ferr_{}_mean'.format(b)])
print("{}: mean flux error: {}, 3sigma in AB mag (Total): {}".format(b, mean_err, flux_to_mag(3.0*mean_err*1.e-6)[0]))
average_depths += [(b, flux_to_mag(1.0*mean_err*1.e-6)[0],
flux_to_mag(3.0*mean_err*1.e-6)[0],
flux_to_mag(5.0*mean_err*1.e-6)[0])]
average_depths = np.array(average_depths, dtype=[('band', "<U16"), ('1s', float), ('3s', float), ('5s', float)])
# -
def FWHM(X,Y):
half_max = max(Y) / 2.
#find when function crosses line half_max (when sign of diff flips)
#take the 'derivative' of signum(half_max - Y[])
d = half_max - Y
#plot(X,d) #if you are interested
#find the left and right most indexes
low_end = X[np.where(d < 0)[0][0]]
high_end = X[np.where(d < 0)[0][-1]]
return low_end, high_end, (high_end - low_end)
# +
data = []
for b in ap_bands:
data += [['ap_' + b, Table(data = parse_single_table(FILTERS_DIR + b +'.xml').array.data)]]
for b in tot_bands:
data += [[b, Table(data = parse_single_table(FILTERS_DIR + b +'.xml').array.data)]]
# -
wav_range = []
for dat in data:
print(dat[0], FWHM(np.array(dat[1]['Wavelength']), np.array(dat[1]['Transmission'])))
wav_range += [[dat[0], FWHM(np.array(dat[1]['Wavelength']), np.array(dat[1]['Transmission']))]]
# +
for dat in data:
wav_deets = FWHM(np.array(dat[1]['Wavelength']), np.array(dat[1]['Transmission']))
depth = average_depths['5s'][average_depths['band'] == dat[0]]
#print(depth)
plt.plot([wav_deets[0],wav_deets[1]], [depth,depth], label=dat[0])
plt.xlabel('Wavelength ($\AA$)')
plt.ylabel('Depth')
plt.xscale('log')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('Depths on {}'.format(FIELD))
# -
# ### IV.c - Depth vs coverage comparison
#
# How best to do this? Colour/intensity plot over area? Percentage coverage vs mean depth?
# +
for dat in data:
wav_deets = FWHM(np.array(dat[1]['Wavelength']), np.array(dat[1]['Transmission']))
depth = average_depths['5s'][average_depths['band'] == dat[0]]
#print(depth)
coverage = np.sum(~np.isnan(depths['ferr_{}_mean'.format(dat[0])]))/len(depths)
plt.plot(coverage, depth, 'x', label=dat[0])
plt.xlabel('Coverage')
plt.ylabel('Depth')
#plt.xscale('log')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('Depths (5 $\sigma$) vs coverage on {}'.format(FIELD))
| dmu1/dmu1_ml_SA13/4_Selection_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import time
from torch.nn.utils.rnn import pad_packed_sequence, pack_padded_sequence
from collections import Counter
from tqdm import tqdm as tqdm
from itertools import zip_longest
from copy import deepcopy
from collections import Counter
from os.path import join
from codecs import open
from tqdm import tqdm_notebook as tqdm
def FormatTestFile(file_path,outdata_path):
trainingset = list() # store trainingset [content,content,...]
position = list() # store position [article_id, start_pos, end_pos, entity_text, entity_type, ...]
mentions = dict() # store mentions[mention] = Type
with open(file_path, 'r', encoding='utf8') as f:
file_text=f.read().encode('utf-8').decode('utf-8-sig')
datas=file_text.split('\n\n--------------------\n\n')[:-1]
output = ""
for i in tqdm(range(len(datas))):
data=datas[i].split('\n')
trainingset=data[1]
for j in range(len(trainingset)):
if(trainingset[j] == ' 'or trainingset[j] == ' '):
output += "_"+" O\n"
else:
output += trainingset[j]+" O\n"
output += "\n"
with open(outdata_path, 'w', encoding='utf-8') as f:
f.write(output)
def build_corpus(split, make_vocab=True, data_dir="./Data"):
assert split in ['train', 'dev', 'test']
word_lists = []
tag_lists = []
with open(join(data_dir, split+".char.bmes"), 'r', encoding='utf-8') as f:
word_list = []
tag_list = []
i = 0
for line in tqdm(f):
i += 1
if line != '\r\n' and line != '\n':
word, tag = line.strip('\r\n').split()
word_list.append(word)
tag_list.append(tag)
else:
word_lists.append(word_list)
tag_lists.append(tag_list)
word_list = []
tag_list = []
if make_vocab:
word2id = build_map(word_lists)
tag2id = build_map(tag_lists)
return word_lists, tag_lists, word2id, tag2id
else:
return word_lists, tag_lists
def extend_maps(word2id, tag2id, for_crf=True):
word2id['<unk>'] = len(word2id)
word2id['<pad>'] = len(word2id)
tag2id['<unk>'] = len(tag2id)
tag2id['<pad>'] = len(tag2id)
if for_crf:
word2id['<start>'] = len(word2id)
word2id['<end>'] = len(word2id)
tag2id['<start>'] = len(tag2id)
tag2id['<end>'] = len(tag2id)
return word2id, tag2id
def prepocess_data_for_lstmcrf(word_lists, tag_lists, test=False):
assert len(word_lists) == len(tag_lists)
for i in range(len(word_lists)):
word_lists[i].append("<end>")
if not test: # 如果是测试数据,就不需要加end token了
tag_lists[i].append("<end>")
return word_lists, tag_lists
def cal_lstm_crf_loss(crf_scores, targets, tag2id):
pad_id = tag2id.get('<pad>')
start_id = tag2id.get('<start>')
end_id = tag2id.get('<end>')
device = crf_scores.device
batch_size, max_len = targets.size()
target_size = len(tag2id)
mask = (targets != pad_id)
lengths = mask.sum(dim=1)
targets = indexed(targets, target_size, start_id)
targets = targets.masked_select(mask)
flatten_scores = crf_scores.masked_select(
mask.view(batch_size, max_len, 1, 1).expand_as(crf_scores)
).view(-1, target_size*target_size).contiguous()
golden_scores = flatten_scores.gather(
dim=1, index=targets.unsqueeze(1)).sum()
scores_upto_t = torch.zeros(batch_size, target_size).to(device)
for t in range(max_len):
batch_size_t = (lengths > t).sum().item()
if t == 0:
scores_upto_t[:batch_size_t] = crf_scores[:batch_size_t,
t, start_id, :]
else:
scores_upto_t[:batch_size_t] = torch.logsumexp(
crf_scores[:batch_size_t, t, :, :] +
scores_upto_t[:batch_size_t].unsqueeze(2),
dim=1
)
all_path_scores = scores_upto_t[:, end_id].sum()
loss = (all_path_scores - golden_scores) / batch_size
return loss
def tensorized(batch, maps):
PAD = maps.get('<pad>')
UNK = maps.get('<unk>')
max_len = len(batch[0])
batch_size = len(batch)
batch_tensor = torch.ones(batch_size, max_len).long() * PAD
for i, l in enumerate(batch):
for j, e in enumerate(l):
batch_tensor[i][j] = maps.get(e, UNK)
lengths = [len(l) for l in batch]
return batch_tensor, lengths
def sort_by_lengths(word_lists, tag_lists):
pairs = list(zip(word_lists, tag_lists))
indices = sorted(range(len(pairs)),
key=lambda k: len(pairs[k][0]),
reverse=True)
pairs = [pairs[i] for i in indices]
word_lists, tag_lists = list(zip(*pairs))
return word_lists, tag_lists, indices
def build_map(lists):
maps = {}
for list_ in lists:
for e in list_:
if e not in maps:
maps[e] = len(maps)
return maps
def load_model(file_name):
with open(file_name, "rb") as f:
input_model = pickle.load(f)
return input_model
# +
class BILSTM_Model(object):
def __init__(self, vocab_size, out_size, crf=True):
self.device = torch.device(
"cuda" if torch.cuda.is_available() else "cpu")
self.emb_size = LSTMConfig.emb_size
self.hidden_size = LSTMConfig.hidden_size
self.crf = crf
if not crf:
self.model = BiLSTM(vocab_size, self.emb_size, self.hidden_size, out_size).to(self.device)
self.cal_loss_func = cal_loss
else:
self.model = BiLSTM_CRF(vocab_size, self.emb_size,self.hidden_size, out_size).to(self.device)
self.cal_loss_func = cal_lstm_crf_loss
self.epoches = TrainingConfig.epoches
self.print_step = TrainingConfig.print_step
self.lr = TrainingConfig.lr
self.batch_size = TrainingConfig.batch_size
self.optimizer = optim.Adam(self.model.parameters(), lr=self.lr)
self.step = 0
self._best_val_loss = 1e18
self.best_model = None
self.model_temp[100] = None
def train(self, word_lists, tag_lists,
dev_word_lists, dev_tag_lists,
word2id, tag2id):
word_lists, tag_lists, _ = sort_by_lengths(word_lists, tag_lists)
dev_word_lists, dev_tag_lists, _ = sort_by_lengths(
dev_word_lists, dev_tag_lists)
B = self.batch_size
for e in (range(1, self.epoches+1)):
self.step = 0
losses = 0.
for ind in tqdm(range(0, len(word_lists), B)):
batch_sents = word_lists[ind:ind+B]
batch_tags = tag_lists[ind:ind+B]
losses += self.train_step(batch_sents,
batch_tags, word2id, tag2id)
if self.step % TrainingConfig.print_step == 0:
total_step = (len(word_lists) // B + 1)
losses = 0.
val_loss = self.validate(dev_word_lists, dev_tag_lists, word2id, tag2id)
print("Epoch {}/{}, Val Loss:{:.4f}".format(e, self.epoches, val_loss))
def train_step(self, batch_sents, batch_tags, word2id, tag2id):
self.model.train()
self.step += 1
tensorized_sents, lengths = tensorized(batch_sents, word2id)
tensorized_sents = tensorized_sents.to(self.device)
targets, lengths = tensorized(batch_tags, tag2id)
targets = targets.to(self.device)
scores = self.model(tensorized_sents, lengths)
self.optimizer.zero_grad()
loss = self.cal_loss_func(scores, targets, tag2id).to(self.device)
loss.backward()
self.optimizer.step()
return loss.item()
def validate(self, dev_word_lists, dev_tag_lists, word2id, tag2id):
self.model.eval()
with torch.no_grad():
val_losses = 0.
val_step = 0
for ind in range(0, len(dev_word_lists), self.batch_size):
val_step += 1
batch_sents = dev_word_lists[ind:ind+self.batch_size]
batch_tags = dev_tag_lists[ind:ind+self.batch_size]
tensorized_sents, lengths = tensorized(
batch_sents, word2id)
tensorized_sents = tensorized_sents.to(self.device)
targets, lengths = tensorized(batch_tags, tag2id)
targets = targets.to(self.device)
scores = self.model(tensorized_sents, lengths)
loss = self.cal_loss_func(
scores, targets, tag2id).to(self.device)
val_losses += loss.item()
val_loss = val_losses / val_step
if val_loss < self._best_val_loss:
print("Keep the Best...")
self.best_model = deepcopy(self.model)
self._best_val_loss = val_loss
return val_loss
def test(self, word_lists, tag_lists, word2id, tag2id):
word_lists, tag_lists, indices = sort_by_lengths(word_lists, tag_lists)
tensorized_sents, lengths = tensorized(word_lists, word2id)
tensorized_sents = tensorized_sents.to(self.device)
self.best_model.eval()
# self.model.eval()
with torch.no_grad():
batch_tagids = self.best_model.test(
tensorized_sents, lengths, tag2id)
pred_tag_lists = []
id2tag = dict((id_, tag) for tag, id_ in tag2id.items())
for i, ids in enumerate(batch_tagids):
tag_list = []
if self.crf:
for j in range(lengths[i] - 1):
tag_list.append(id2tag[ids[j].item()])
else:
for j in range(lengths[i]):
tag_list.append(id2tag[ids[j].item()])
pred_tag_lists.append(tag_list)
ind_maps = sorted(list(enumerate(indices)), key=lambda e: e[1])
indices, _ = list(zip(*ind_maps))
pred_tag_lists = [pred_tag_lists[i] for i in indices]
tag_lists = [tag_lists[i] for i in indices]
return pred_tag_lists, tag_lists
class BiLSTM_CRF(nn.Module):
def __init__(self, vocab_size, emb_size, hidden_size, out_size):
super(BiLSTM_CRF, self).__init__()
self.bilstm = BiLSTM(vocab_size, emb_size, hidden_size, out_size)
self.transition = nn.Parameter(
torch.ones(out_size, out_size) * 1/out_size)
def forward(self, sents_tensor, lengths):
emission = self.bilstm(sents_tensor, lengths)
batch_size, max_len, out_size = emission.size()
crf_scores = emission.unsqueeze(
2).expand(-1, -1, out_size, -1) + self.transition.unsqueeze(0)
return crf_scores
def test(self, test_sents_tensor, lengths, tag2id):
start_id = tag2id['<start>']
end_id = tag2id['<end>']
pad = tag2id['<pad>']
tagset_size = len(tag2id)
crf_scores = self.forward(test_sents_tensor, lengths)
device = crf_scores.device
B, L, T, _ = crf_scores.size()
viterbi = torch.zeros(B, L, T).to(device)
backpointer = (torch.zeros(B, L, T).long() * end_id).to(device)
lengths = torch.LongTensor(lengths).to(device)
for step in range(L):
batch_size_t = (lengths > step).sum().item()
if step == 0:
viterbi[:batch_size_t, step,
:] = crf_scores[: batch_size_t, step, start_id, :]
backpointer[: batch_size_t, step, :] = start_id
else:
max_scores, prev_tags = torch.max(
viterbi[:batch_size_t, step-1, :].unsqueeze(2) +
crf_scores[:batch_size_t, step, :, :],
dim=1
)
viterbi[:batch_size_t, step, :] = max_scores
backpointer[:batch_size_t, step, :] = prev_tags
backpointer = backpointer.view(B, -1)
tagids = []
tags_t = None
for step in range(L-1, 0, -1):
batch_size_t = (lengths > step).sum().item()
if step == L-1:
index = torch.ones(batch_size_t).long() * (step * tagset_size)
index = index.to(device)
index += end_id
else:
prev_batch_size_t = len(tags_t)
new_in_batch = torch.LongTensor(
[end_id] * (batch_size_t - prev_batch_size_t)).to(device)
offset = torch.cat(
[tags_t, new_in_batch],
dim=0
)
index = torch.ones(batch_size_t).long() * (step * tagset_size)
index = index.to(device)
index += offset.long()
try:
tags_t = backpointer[:batch_size_t].gather(
dim=1, index=index.unsqueeze(1).long())
except RuntimeError:
import pdb
pdb.set_trace()
tags_t = tags_t.squeeze(1)
tagids.append(tags_t.tolist())
tagids = list(zip_longest(*reversed(tagids), fillvalue=pad))
tagids = torch.Tensor(tagids).long()
return tagids
# -
class BiLSTM(nn.Module):
def __init__(self, vocab_size, emb_size, hidden_size, out_size):
super(BiLSTM, self).__init__()
self.embedding = nn.Embedding(vocab_size, emb_size)
self.bilstm = nn.LSTM(emb_size, hidden_size,
batch_first=True,
bidirectional=True)
self.lin = nn.Linear(2*hidden_size, out_size)
def forward(self, sents_tensor, lengths):
emb = self.embedding(sents_tensor) # [B, L, emb_size]
packed = pack_padded_sequence(emb, lengths, batch_first=True)
rnn_out, _ = self.bilstm(packed)
rnn_out, _ = pad_packed_sequence(rnn_out, batch_first=True)
scores = self.lin(rnn_out) # [B, L, out_size]
return scores
def test(self, sents_tensor, lengths, _):
logits = self.forward(sents_tensor, lengths) # [B, L, out_size]
_, batch_tagids = torch.max(logits, dim=2)
return batch_tagids
# +
# inputdata_path = f"test.txt" #要轉換的檔案
# outdata_path = f'./Data/test.char.bmes' #轉換輸出的檔案
# FormatTestFile(inputdata_path,outdata_path)
# -
word2id = {'醫': 0, '師': 1, ':': 2, '啊': 3, '回': 4, '去': 5, '還': 6, '好': 7, '嗎': 8, '?': 9, '民': 10, '眾': 11, '欸': 12, ',': 13, '是': 14, '虛': 15, '的': 16, '但': 17, '。': 18, '真': 19, '險': 20, '坦': 21, '白': 22, '講': 23, '我': 24, '剛': 25, '時': 26, '候': 27, '晚': 28, '上': 29, '有': 30, '盜': 31, '汗': 32, '阿': 33, '只': 34, '前': 35, '天': 36, '很': 37, '多': 38, '就': 39, '算': 40, '沒': 41, '可': 42, '一': 43, '覺': 44, '到': 45, '明': 46, '這': 47, '樣': 48, '齁': 49, '給': 50, '你': 51, '看': 52, '電': 53, '腦': 54, '斷': 55, '層': 56, '嘿': 57, '那': 58, '個': 59, '病': 60, '毒': 61, '報': 62, '告': 63, '不': 64, '知': 65, '道': 66, '出': 67, '來': 68, '對': 69, '它': 70, '幫': 71, '驗': 72, '了': 73, '少': 74, '自': 75, '體': 76, '免': 77, '疫': 78, '呢': 79, '喔': 80, '相': 81, '信': 82, '之': 83, '都': 84, '過': 85, '哇': 86, '工': 87, '作': 88, '壓': 89, '力': 90, '大': 91, '得': 92, '潰': 93, '瘍': 94, 'n': 95, 'e': 96, 'g': 97, 'a': 98, 't': 99, 'i': 100, 'v': 101, '皰': 102, '疹': 103, '抗': 104, '也': 105, '嗯': 106, 'Q': 107, '熱': 108, '陰': 109, '性': 110, '然': 111, '後': 112, '第': 113, '次': 114, '檢': 115, '查': 116, '綜': 117, '合': 118, '結': 119, '果': 120, '所': 121, '謂': 122, '種': 123, '蚊': 124, '蟲': 125, '叮': 126, '咬': 127, '跟': 128, '動': 129, '物': 130, '、': 131, '跳': 132, '蚤': 133, '些': 134, '比': 135, '較': 136, '關': 137, '解': 138, '恙': 139, '需': 140, '再': 141, '採': 142, '係': 143, '們': 144, '要': 145, '追': 146, '蹤': 147, '起': 148, '送': 149, '疾': 150, '管': 151, '區': 152, '以': 153, '機': 154, '會': 155, '因': 156, '為': 157, '在': 158, '離': 159, '島': 160, '山': 161, '斑': 162, '傷': 163, '寒': 164, '外': 165, '啦': 166, '見': 167, 'E': 168, 'B': 169, 'V': 170, 'C': 171, 'M': 172, 'O': 173, 'K': 174, '什': 175, '麼': 176, '東': 177, '西': 178, '引': 179, '肝': 180, '功': 181, '能': 182, '異': 183, '常': 184, '巨': 185, '細': 186, '胞': 187, '良': 188, '己': 189, '讓': 190, '發': 191, '燒': 192, '陣': 193, '子': 194, '直': 195, '接': 196, '調': 197, '炎': 198, '指': 199, '數': 200, '高': 201, 'R': 202, 'P': 203, '8': 204, '5': 205, '1': 206, '0': 207, '2': 208, '睡': 209, '喝': 210, '酒': 211, '應': 212, '酬': 213, '其': 214, '實': 215, '說': 216, '假': 217, '如': 218, '年': 219, '老': 220, '岸': 221, 'G': 222, 'o': 223, 'l': 224, '設': 225, '廠': 226, '客': 227, '戶': 228, '邊': 229, '做': 230, '跑': 231, '業': 232, '務': 233, 'A': 234, 'S': 235, '克': 236, '人': 237, '確': 238, '照': 239, '話': 240, '三': 241, '月': 242, '十': 243, '八': 244, '住': 245, '院': 246, '腸': 247, '點': 248, '壁': 249, '厚': 250, '正': 251, '該': 252, '像': 253, '蛋': 254, '殼': 255, '地': 256, '方': 257, '內': 258, '視': 259, '鏡': 260, '切': 261, '片': 262, '許': 263, '感': 264, '染': 265, '開': 266, '始': 267, '另': 268, '主': 269, '脈': 270, '嘛': 271, '條': 272, '旁': 273, '顆': 274, '淋': 275, '巴': 276, '怎': 277, '腫': 278, '理': 279, '論': 280, '等': 281, '於': 282, '哨': 283, '意': 284, '思': 285, '站': 286, '局': 287, '部': 288, '菌': 289, '或': 290, '周': 291, '遭': 292, '浪': 293, '誰': 294, '裡': 295, '面': 296, '兵': 297, '哥': 298, '球': 299, '奇': 300, '血': 301, '增': 302, '加': 303, '特': 304, '別': 305, '蠻': 306, '怪': 307, '通': 308, '衝': 309, '必': 310, '須': 311, '放': 312, '射': 313, '科': 314, '他': 315, '叫': 316, '反': 317, 'r': 318, 'c': 319, '消': 320, '下': 321, '定': 322, '退': 323, '惡': 324, '持': 325, '續': 326, '甚': 327, '至': 328, '變': 329, '而': 330, '且': 331, '規': 332, '則': 333, '圓': 334, '滾': 335, '長': 336, '當': 337, '最': 338, '準': 339, '深': 340, '膛': 341, '剖': 342, '肚': 343, '想': 344, '計': 345, '劃': 346, '降': 347, '段': 348, '間': 349, '兩': 350, '問': 351, '題': 352, '取': 353, '式': 354, '賈': 355, '伯': 356, '斯': 357, '門': 358, '診': 359, '禮': 360, '拜': 361, '抽': 362, '二': 363, '現': 364, '生': 365, '素': 366, '吃': 367, '預': 368, '防': 369, '綠': 370, '色': 371, '黃': 372, '先': 373, '停': 374, '橘': 375, '完': 376, '串': 377, '把': 378, '般': 379, '聽': 380, '付': 381, '難': 382, '搞': 383, '罕': 384, '掉': 385, '液': 386, '培': 387, '養': 388, '保': 389, '留': 390, '今': 391, '請': 392, '繼': 393, '班': 394, '太': 395, '累': 396, '畢': 397, '竟': 398, '居': 399, '美': 400, '國': 401, '買': 402, '房': 403, '紐': 404, '約': 405, '新': 406, '社': 407, '錯': 408, '圖': 409, '書': 410, '館': 411, '總': 412, '婚': 413, '情': 414, '路': 415, '順': 416, '遂': 417, '身': 418, '顧': 419, '據': 420, '普': 421, '拿': 422, '疼': 423, '稍': 424, '微': 425, '備': 426, '著': 427, '藥': 428, '本': 429, '用': 430, '挑': 431, '簡': 432, '單': 433, '狀': 434, '況': 435, '七': 436, '號': 437, '四': 438, '九': 439, '呀': 440, '胃': 441, '快': 442, '吧': 443, '便': 444, '幽': 445, '螺': 446, '旋': 447, '桿': 448, '純': 449, '粹': 450, '諾': 451, '羅': 452, '又': 453, '久': 454, '概': 455, '五': 456, '分': 457, '吐': 458, '瀉': 459, '嚴': 460, '重': 461, '剩': 462, '猜': 463, '哪': 464, '拖': 465, '誤': 466, '打': 467, '撞': 468, 'T': 469, '差': 470, '幹': 471, '才': 472, '中': 473, '膈': 474, '腔': 475, '刀': 476, '進': 477, '心': 478, '包': 479, '膜': 480, '肌': 481, '質': 482, '敏': 483, '哈': 484, '謝': 485, '從': 486, '走': 487, '學': 488, '弟': 489, '百': 490, '塊': 491, '車': 492, '馬': 493, '費': 494, '嬤': 495, '她': 496, '家': 497, '屬': 498, '誒': 499, '行': 500, '午': 501, '昨': 502, '半': 503, '夜': 504, '3': 505, '.': 506, '尿': 507, '恩': 508, '哼': 509, '雷': 510, '氏': 511, '肺': 512, '名': 513, '字': 514, '已': 515, '針': 516, '效': 517, '注': 518, '日': 519, '辦': 520, '法': 521, '平': 522, '營': 523, '…': 524, '突': 525, '寫': 526, '轉': 527, '全': 528, '~': 529, '慶': 530, '鴻': 531, '祝': 532, '江': 533, '鳥': 534, '健': 535, '註': 536, '冊': 537, '肉': 538, '埋': 539, '附': 540, '頭': 541, '呵': 542, '同': 543, '張': 544, '處': 545, '食': 546, '鹽': 547, '水': 548, '六': 549, '核': 550, '亂': 551, '昏': 552, '恍': 553, '神': 554, '例': 555, '低': 556, '幾': 557, '輪': 558, '整': 559, '護': 560, '7': 561, '4': 562, '眼': 563, '印': 564, '象': 565, '清': 566, '楚': 567, '入': 568, '找': 569, '緊': 570, '歷': 571, '福': 572, '療': 573, '小': 574, '穩': 575, '近': 576, '口': 577, '辛': 578, '苦': 579, '絕': 580, '症': 581, '餒': 582, '被': 583, '配': 584, '申': 585, '份': 586, '支': 587, '皮': 588, '安': 589, '怕': 590, '念': 591, '教': 592, '泰': 593, '9': 594, '事': 595, '舒': 596, '服': 597, '底': 598, '副': 599, 'k': 600, '腰': 601, '痛': 602, '脫': 603, '帶': 604, '坐': 605, '膨': 606, '瘦': 607, '骨': 608, '翻': 609, '麻': 610, '抖': 611, '萎': 612, '縮': 613, '衣': 614, '右': 615, '腿': 616, '側': 617, '隻': 618, '手': 619, '影': 620, '響': 621, '髖': 622, '黑': 623, '左': 624, '肩': 625, '稱': 626, '化': 627, '膿': 628, '止': 629, '眠': 630, '強': 631, '換': 632, 'O': 633, 'K': 634, '菸': 635, '檳': 636, '榔': 637, '原': 638, '脹': 639, '氣': 640, '鋅': 641, '補': 642, '充': 643, '劑': 644, '經': 645, '紅': 646, '類': 647, '似': 648, '愛': 649, '6': 650, '腳': 651, '何': 652, '決': 653, '復': 654, '慢': 655, '改': 656, '善': 657, '練': 658, '訓': 659, '量': 660, '撇': 661, '筷': 662, '排': 663, '士': 664, '林': 665, '裏': 666, '洗': 667, '腎': 668, '資': 669, '料': 670, '治': 671, '參': 672, '考': 673, '痠': 674, '背': 675, '躺': 676, '彎': 677, '敢': 678, '期': 679, '忙': 680, '記': 681, '敲': 682, '臟': 683, '扭': 684, '蛤': 685, '拉': 686, '網': 687, '激': 688, '烈': 689, '顯': 690, '疲': 691, '勞': 692, '畏': 693, '早': 694, '吹': 695, '風': 696, '·': 697, '哦': 698, '試': 699, '即': 700, '夠': 701, '者': 702, '冷': 703, '搬': 704, '超': 705, '流': 706, '失': 707, '活': 708, '運': 709, '更': 710, '音': 711, '波': 712, '石': 713, '悶': 714, '輸': 715, '塞': 716, '泡': 717, '造': 718, '成': 719, '鈣': 720, '組': 721, '織': 722, '吼': 723, 'D': 724, '控': 725, '制': 726, '懷': 727, '疑': 728, '胖': 729, '公': 730, '斤': 731, '目': 732, '曉': 733, '耶': 734, '壞': 735, '趕': 736, '提': 737, '醒': 738, '領': 739, '速': 740, '拍': 741, '卡': 742, '減': 743, '負': 744, '擔': 745, '無': 746, '步': 747, '喘': 748, '每': 749, '呼': 750, '吸': 751, '均': 752, 'X': 753, '光': 754, '維': 755, '謹': 756, '慎': 757, '冒': 758, '掃': 759, '描': 760, '判': 761, '亮': 762, '瘤': 763, '建': 764, '議': 765, '詳': 766, '表': 767, '示': 768, '臨': 769, '空': 770, '輕': 771, '捐': 772, '節': 773, 'H': 774, 'T': 775, 'L': 776, 'V': 777, '測': 778, '歲': 779, '倒': 780, '慮': 781, 'H': 782, '產': 783, '摸': 784, '胸': 785, '精': 786, '範': 787, '圍': 788, '讀': 789, '錢': 790, '您': 791, '煩': 792, '初': 793, '程': 794, '躲': 795, '溫': 796, '和': 797, '認': 798, '陽': 799, '曾': 800, '害': 801, '攔': 802, '束': 803, '千': 804, '癢': 805, '3': 806, '4': 807, '腹': 808, '板': 809, '升': 810, '萬': 811, '唯': 812, '往': 813, '弱': 814, '抵': 815, '移': 816, '植': 817, '恢': 818, '衛': 819, '專': 820, '審': 821, '優': 822, '場': 823, 'X': 824, '痰': 825, '項': 826, '零': 827, '任': 828, '掛': 829, '急': 830, '毛': 831, '囊': 832, '斥': 833, '擦': 834, '終': 835, '派': 836, '乾': 837, '燥': 838, '沖': 839, '爽': 840, '油': 841, '越': 842, '陳': 843, '囉': 844, '待': 845, 'B': 846, 'C': 847, '基': 848, '並': 849, '非': 850, '侯': 851, '型': 852, '託': 853, '膽': 854, '根': 855, '除': 856, '件': 857, '趟': 858, '!': 859, '硬': 860, '癌': 861, '危': 862, '丘': 863, '形': 864, '痂': 865, '箭': 866, '龜': 867, '修': 868, '抱': 869, '歉': 870, '削': 871, '膝': 872, '蓋': 873, '掀': 874, '凸': 875, '堆': 876, '抹': 877, '破': 878, '紀': 879, '夢': 880, '膚': 881, '田': 882, '操': 883, 'N': 884, '陸': 885, '幸': 886, '政': 887, '府': 888, '團': 889, '隊': 890, '希': 891, '望': 892, '星': 893, '收': 894, '洛': 895, '亞': 896, '樓': 897, '簽': 898, '籤': 899, '受': 900, '率': 901, '呃': 902, '殺': 903, '死': 904, '頑': 905, '固': 906, '⋯': 907, '忘': 908, '鑑': 909, '頻': 910, '婦': 911, '介': 912, '紹': 913, '泌': 914, '兒': 915, '賀': 916, '爾': 917, '蒙': 918, '隱': 919, '私': 920, '糖': 921, '飲': 922, '甜': 923, '標': 924, '態': 925, '落': 926, '緣': 927, '族': 928, '偏': 929, '脂': 930, '肪': 931, '度': 932, '密': 933, '醇': 934, '煮': 935, '儲': 936, '存': 937, '觀': 938, '習': 939, '慣': 940, '盡': 941, '靠': 942, '潛': 943, '遇': 944, '睛': 945, '週': 946, '避': 947, '碌': 948, '飯': 949, '透': 950, '代': 951, '衡': 952, '碗': 953, '青': 954, '菜': 955, '魚': 956, '炒': 957, '蝦': 958, '芒': 959, '蘋': 960, '餐': 961, '胰': 962, '迫': 963, '飆': 964, '瞬': 965, '伏': 966, '符': 967, '礎': 968, '隨': 969, '錄': 970, '討': 971, '研': 972, '究': 973, '患': 974, '咳': 975, '嗽': 976, 'L': 977, '嬸': 978, '酸': 979, '甘': 980, '炸': 981, '沙': 982, '筋': 983, '閃': 984, '息': 985, '術': 986, '扁': 987, '醉': 988, '鬆': 989, '姿': 990, '勢': 991, '勇': 992, '元': 993, '屎': 994, '火': 995, '龍': 996, '軟': 997, '髮': 998, '葡': 999, '萄': 1000, '櫻': 1001, '桃': 1002, '鐵': 1003, '顏': 1004, '靈': 1005, '刺': 1006, '容': 1007, '器': 1008, '裝': 1009, '櫃': 1010, '台': 1011, '爸': 1012, '膏': 1013, '灌': 1014, '雙': 1015, '樹': 1016, '仙': 1017, '草': 1018, '歐': 1019, '夏': 1020, '貝': 1021, '茶': 1022, '攝': 1023, '腺': 1024, '聖': 1025, '誕': 1026, '延': 1027, '喉': 1028, '嚨': 1029, '鼻': 1030, '擤': 1031, '涕': 1032, '濃': 1033, '淡': 1034, '跡': 1035, '罩': 1036, '孫': 1037, '傳': 1038, '足': 1039, '弓': 1040, '繁': 1041, '殖': 1042, '勒': 1043, '脖': 1044, '腋': 1045, '窩': 1046, '鼠': 1047, '蹊': 1048, '胯': 1049, '嘖': 1050, '濕': 1051, '選': 1052, 's': 1053, 'd': 1054, 'm': 1055, '乎': 1056, '滿': 1057, '勉': 1058, '授': 1059, '訊': 1060, '演': 1061, '秀': 1062, '司': 1063, '栽': 1064, '英': 1065, '文': 1066, '懂': 1067, '模': 1068, '擬': 1069, '向': 1070, '佛': 1071, '碩': 1072, '博': 1073, '材': 1074, '厲': 1075, '闆': 1076, '未': 1077, '系': 1078, '統': 1079, 'b': 1080, '譬': 1081, '推': 1082, 'ㄟ': 1083, 'W': 1084, '布': 1085, 'u': 1086, 'w': 1087, 'h': 1088, '畫': 1089, '鐘': 1090, '寬': 1091, '值': 1092, '海': 1093, '貧': 1094, '史': 1095, '痾': 1096, 'p': 1097, '積': 1098, '癒': 1099, 'y': 1100, '逐': 1101, '漸': 1102, '噬': 1103, '誇': 1104, '棉': 1105, '連': 1106, '廣': 1107, '各': 1108, '壯': 1109, '課': 1110, '擾': 1111, '劍': 1112, '橋': 1113, '某': 1114, '投': 1115, '複': 1116, '暑': 1117, '姑': 1118, '洲': 1119, '冰': 1120, '丈': 1121, '遮': 1122, '霧': 1123, '遠': 1124, '淚': 1125, '休': 1126, '粒': 1127, '拆': 1128, '床': 1129, '凌': 1130, '晨': 1131, '易': 1132, '致': 1133, '助': 1134, '褪': 1135, '利': 1136, '梅': 1137, '騎': 1138, '踏': 1139, '納': 1140, '筆': 1141, '貴': 1142, '環': 1143, '觸': 1144, '盤': 1145, '尼': 1146, '施': 1147, '黴': 1148, 'f': 1149, '交': 1150, '察': 1151, '端': 1152, '诶': 1153, '耐': 1154, '階': 1155, '瓜': 1156, '麵': 1157, '澱': 1158, '粉': 1159, '醣': 1160, '趨': 1161, '位': 1162, '置': 1163, '角': 1164, '探': 1165, '搭': 1166, '飛': 1167, '末': 1168, '噢': 1169, '銷': 1170, '品': 1171, '苗': 1172, '證': 1173, '暸': 1174, '叔': 1175, '將': 1176, '雲': 1177, '緩': 1178, '秋': 1179, '玩': 1180, '康': 1181, '技': 1182, '塗': 1183, '香': 1184, '搗': 1185, '摘': 1186, '女': 1187, '牛': 1188, '奶': 1189, '弄': 1190, '禿': 1191, '嘔': 1192, '雖': 1193, '拐': 1194, '杖': 1195, '撐': 1196, '扶': 1197, '妳': 1198, '姨': 1199, '吩': 1200, '咐': 1201, '藍': 1202, '膠': 1203, '載': 1204, '北': 1205, '市': 1206, '塭': 1207, '孩': 1208, '鄰': 1209, '騙': 1210, '王': 1211, '抓': 1212, '葉': 1213, '估': 1214, '徑': 1215, '尖': 1216, '此': 1217, '侵': 1218, '循': 1219, '擴': 1220, '散': 1221, '淨': 1222, '殘': 1223, '滲': 1224, '殊': 1225, '髒': 1226, '乘': 1227, '孔': 1228, '阻': 1229, '域': 1230, '辨': 1231, '里': 1232, '椅': 1233, '爲': 1234, '淺': 1235, '紙': 1236, '聲': 1237, '厭': 1238, '説': 1239, '麽': 1240, '呐': 1241, '着': 1242, '暈': 1243, '徹': 1244, '辣': 1245, '逆': 1246, '帕': 1247, '金': 1248, '森': 1249, '朋': 1250, '友': 1251, '男': 1252, '緒': 1253, '由': 1254, '暴': 1255, '傾': 1256, '警': 1257, '罵': 1258, '令': 1259, '憂': 1260, '鬱': 1261, '閉': 1262, '狹': 1263, '績': 1264, '桌': 1265, '寄': 1266, '朵': 1267, '花': 1268, '庭': 1269, '羞': 1270, '恥': 1271, '刻': 1272, '脆': 1273, '既': 1274, '牙': 1275, '插': 1276, '吵': 1277, '媳': 1278, '按': 1279, '摩': 1280, '李': 1281, '母': 1282, '爛': 1283, '沉': 1284, '捏': 1285, '吞': 1286, '隔': 1287, '肯': 1288, '責': 1289, '滴': 1290, '願': 1291, '線': 1292, '簿': 1293, '刷': 1294, '帳': 1295, '唉': 1296, '湄': 1297, '爆': 1298, '袋': 1299, '邏': 1300, '輯': 1301, '薄': 1302, '否': 1303, '訂': 1304, '泄': 1305, '暫': 1306, '抬': 1307, '鹹': 1308, '瓶': 1309, '礦': 1310, '泉': 1311, '6': 1312, '0': 1313, '毫': 1314, '扣': 1315, '湯': 1316, '潮': 1317, '灰': 1318, '甲': 1319, '蜂': 1320, '抑': 1321, '益': 1322, '親': 1323, '春': 1324, '痘': 1325, '洞': 1326, '癬': 1327, '故': 1328, '華': 1329, '凝': 1330, '困': 1331, '缺': 1332, '忠': 1333, '惱': 1334, '熟': 1335, '室': 1336, '混': 1337, '識': 1338, '析': 1339, '登': 1340, '吳': 1341, '樟': 1342, '芝': 1343, '彈': 1344, '集': 1345, '肛': 1346, '適': 1347, '糊': 1348, '攪': 1349, '命': 1350, '紛': 1351, '瀑': 1352, '逛': 1353, '圈': 1354, '爬': 1355, '溪': 1356, '木': 1357, '池': 1358, '夫': 1359, '邵': 1360, '承': 1361, '翰': 1362, '執': 1363, '級': 1364, '噴': 1365, '武': 1366, '漢': 1367, '媽': 1368, '限': 1369, '齊': 1370, '痔': 1371, '瘡': 1372, '曬': 1373, '嘴': 1374, '垃': 1375, '圾': 1376, '碰': 1377, '戒': 1378, '聊': 1379, '姐': 1380, '商': 1381, '冬': 1382, '雨': 1383, '源': 1384, '疣': 1385, '戴': 1386, '套': 1387, '談': 1388, '溝': 1389, '併': 1390, '牽': 1391, '涉': 1392, '溶': 1393, '委': 1394, '婉': 1395, '勸': 1396, '賠': 1397, '償': 1398, '救': 1399, '濟': 1400, '署': 1401, '境': 1402, '穿': 1403, '掌': 1404, '典': 1405, '鳳': 1406, '梨': 1407, '飽': 1408, '祕': 1409, '絞': 1410, '土': 1411, '燙': 1412, '使': 1413, '障': 1414, '褲': 1415, '富': 1416, '楷': 1417, '柔': 1418, '倍': 1419, '額': 1420, '依': 1421, '罰': 1422, '脊': 1423, '屏': 1424, '鎮': 1425, '髓': 1426, '律': 1427, '途': 1428, '義': 1429, '敗': 1430, '扎': 1431, '互': 1432, '舉': 1433, '官': 1434, '潦': 1435, '疏': 1436, '導': 1437, '供': 1438, '撥': 1439, '述': 1440, '拷': 1441, '權': 1442, '章': 1443, '蝕': 1444, '肘': 1445, '訴': 1446, '縫': 1447, '膀': 1448, '棒': 1449, '員': 1450, '督': 1451, '促': 1452, '沾': 1453, '黏': 1454, '農': 1455, '曆': 1456, '弛': 1457, '箱': 1458, '笑': 1459, '窗': 1460, '肥': 1461, '亨': 1462, '佔': 1463, '案': 1464, '短': 1465, '陪': 1466, '鍋': 1467, '醬': 1468, '歹': 1469, '漱': 1470, '滅': 1471, '罐': 1472, '妹': 1473, '雜': 1474, '慾': 1475, '梯': 1476, '幅': 1477, '省': 1478, '佐': 1479, 'I': 1480, '搓': 1481, '磨': 1482, '摳': 1483, '痱': 1484, '共': 1485, '季': 1486, '宣': 1487, '檔': 1488, '偶': 1489, '纖': 1490, '碎': 1491, '堅': 1492, '忍': 1493, '蔡': 1494, '漿': 1495, '達': 1496, '酯': 1497, '滷': 1498, '輔': 1499, '浮': 1500, '鞏': 1501, '牌': 1502, '舊': 1503, '版': 1504, '擠': 1505, '榮': 1506, '糟': 1507, '鞋': 1508, '柺': 1509, '漏': 1510, '遺': 1511, '吉': 1512, '韋': 1513, '含': 1514, '鋁': 1515, '鎂': 1516, '牠': 1517, '嗜': 1518, '貪': 1519, '尤': 1520, '忽': 1521, '粥': 1522, '竹': 1523, '炭': 1524, '倦': 1525, '濁': 1526, '氧': 1527, '嚇': 1528, '衆': 1529, '莫': 1530, '燕': 1531, '麥': 1532, '暗': 1533, '靜': 1534, '曲': 1535, '姊': 1536, '勁': 1537, '猛': 1538, '沿': 1539, '擺': 1540, '齦': 1541, '輻': 1542, '育': 1543, '玻': 1544, '璃': 1545, '娃': 1546, '侏': 1547, '儒': 1548, '竄': 1549, '勤': 1550, '奮': 1551, '軌': 1552, '遍': 1553, '痊': 1554, '宗': 1555, '碼': 1556, '惠': 1557, '縱': 1558, '校': 1559, '廖': 1560, '戚': 1561, '哩': 1562, '懶': 1563, '幻': 1564, '嫁': 1565, '氫': 1566, '屁': 1567, '茫': 1568, '丸': 1569, '序': 1570, '架': 1571, '伴': 1572, '侶': 1573, '宿': 1574, '志': 1575, '暖': 1576, '逃': 1577, '炮': 1578, '焦': 1579, '擇': 1580, '求': 1581, '貌': 1582, '釐': 1583, '寧': 1584, '扮': 1585, '喜': 1586, '歡': 1587, '填': 1588, '卷': 1589, '頂': 1590, '礙': 1591, '訪': 1592, '惶': 1593, '恐': 1594, '杯': 1595, '彼': 1596, '宴': 1597, '詢': 1598, '咧': 1599, '釋': 1600, '桶': 1601, 'Y': 1602, '匿': 1603, '篩': 1604, '格': 1605, '鬧': 1606, '滋': 1607, '懼': 1608, '浩': 1609, '址': 1610, '帥': 1611, '首': 1612, '錶': 1613, '展': 1614, 'U': 1615, '遊': 1616, '戲': 1617, '瞌': 1618, '偷': 1619, '砲': 1620, '噁': 1621, '咖': 1622, '啡': 1623, '驚': 1624, '聯': 1625, '絡': 1626, '膩': 1627, '悉': 1628, '與': 1629, '拔': 1630, '迷': 1631, '措': 1632, '矛': 1633, '盾': 1634, '歸': 1635, '潢': 1636, '熬': 1637, '羨': 1638, '慕': 1639, '興': 1640, '評': 1641, 'I': 1642, '9': 1643, '耗': 1644, '臉': 1645, '具': 1646, '咦': 1647, '碑': 1648, '努': 1649, '塑': 1650, '槍': 1651, '疙': 1652, '瘩': 1653, '哎': 1654, '革': 1655, '賣': 1656, '盒': 1657, '珍': 1658, '呦': 1659, '際': 1660, '干': 1661, '兇': 1662, '稽': 1663, '縣': 1664, '藏': 1665, '覆': 1666, '愉': 1667, '悅': 1668, '樂': 1669, '拒': 1670, '極': 1671, '尋': 1672, '諮': 1673, '癮': 1674, '鬼': 1675, '趣': 1676, '刪': 1677, '協': 1678, '壤': 1679, '韓': 1680, '儘': 1681, '2': 1682, '1': 1683, 'N': 1684, 'E': 1685, '輩': 1686, '答': 1687, '捲': 1688, '價': 1689, '購': 1690, '矩': 1691, '露': 1692, '鍵': 1693, '溯': 1694, '秒': 1695, '蹭': 1696, '丟': 1697, '割': 1698, '聼': 1699, '葩': 1700, '閡': 1701, '瘙': 1702, '熊': 1703, '饋': 1704, '内': 1705, '款': 1706, '披': 1707, '瞭': 1708, '怖': 1709, '嘉': 1710, '距': 1711, '封': 1712, '誠': 1713, '呂': 1714, '嗨': 1715, '括': 1716, '尷': 1717, '尬': 1718, '杜': 1719, '蕾': 1720, '岡': 1721, '傻': 1722, '永': 1723, '攸': 1724, '=': 1725, '擋': 1726, '7': 1727, '5': 1728, '店': 1729, '職': 1730, '傍': 1731, '糾': 1732, '截': 1733, '盯': 1734, '曼': 1735, '頓': 1736, '凍': 1737, '鏢': 1738, '唔': 1739, '妙': 1740, '阪': 1741, '并': 1742, '及': 1743, '涼': 1744, '慌': 1745, '盛': 1746, '准': 1747, '頗': 1748, '乖': 1749, '箝': 1750, '監': 1751, '豆': 1752, '貸': 1753, '債': 1754, '愧': 1755, '疚': 1756, '戀': 1757, '逼': 1758, '借': 1759, '棄': 1760, '挺': 1761, '貨': 1762, '勾': 1763, '憐': 1764, '賺': 1765, '攤': 1766, '檻': 1767, '煙': 1768, '拋': 1769, '握': 1770, '灣': 1771, '划': 1772, '嘗': 1773, '偽': 1774, '世': 1775, '界': 1776, '景': 1777, '旅': 1778, '默': 1779, '予': 1780, '郵': 1781, '朝': 1782, '替': 1783, '渣': 1784, '妖': 1785, '魔': 1786, '邀': 1787, '飄': 1788, '陌': 1789, '立': 1790, '拼': 1791, '創': 1792, '箋': 1793, '嘍': 1794, '慘': 1795, '澳': 1796, '座': 1797, '央': 1798, '淆': 1799, '頁': 1800, '威': 1801, '鋼': 1802, '鈕': 1803, '晦': 1804, '語': 1805, '拚': 1806, '喬': 1807, 'F': 1808, '糕': 1809, '.': 1810, '允': 1811, '摟': 1812, '禍': 1813, '跌': 1814, '疊': 1815, '譚': 1816, '京': 1817, '宇': 1818, '智': 1819, '列': 1820, '群': 1821, '廢': 1822, '禦': 1823, '唾': 1824, '雄': 1825, '蘭': 1826, '野': 1827, '柳': 1828, '弧': 1829, '映': 1830, '沫': 1831, '仍': 1832, '煉': 1833, '聚': 1834, '酐': 1835, '憶': 1836, '滑': 1837, '味': 1838, '獎': 1839, '夾': 1840, '鏈': 1841, '犯': 1842, '「': 1843, '」': 1844, '咯': 1845, '獲': 1846, '后': 1847, '臺': 1848, '厰': 1849, '村': 1850, '返': 1851, '潔': 1852, '癖': 1853, '欣': 1854, '賞': 1855, '潤': 1856, '佈': 1857, '億': 1858, '鼓': 1859, '傑': 1860, '徵': 1861, '挫': 1862, '枝': 1863, '紫': 1864, '堵': 1865, '薦': 1866, '寮': 1867, '彰': 1868, '戰': 1869, '藉': 1870, '吻': 1871, '玄': 1872, '尾': 1873, '賭': 1874, '挪': 1875, '曝': 1876, '曡': 1877, '嵗': 1878, '氛': 1879, '踢': 1880, '%': 1881, '﹑': 1882, '若': 1883, '橫': 1884, '鎖': 1885, '繫': 1886, '溼': 1887, '庫': 1888, '啓': 1889, '孝': 1890, '弊': 1891, '嘞': 1892, '涵': 1893, '享': 1894, '_': 1895, '耳': 1896, 'A': 1897, '井': 1898, '壽': 1899, 'R': 1900, 'P': 1901, '豐': 1902, 'Y': 1903, '嚏': 1904, '爺': 1905, '烘': 1906, '焙': 1907, '餅': 1908, '園': 1909, '樞': 1910, '磁': 1911, '振': 1912, '蹦': 1913, '眩': 1914, '疤': 1915, 'D': 1916, '凹': 1917, '槽': 1918, '綿': 1919, '屋': 1920, '盲': 1921, '譜': 1922, '儀': 1923, '糞': 1924, '挖': 1925, '繳': 1926, '尚': 1927, '禁': 1928, '餃': 1929, '宜': 1930, '錠': 1931, '荷': 1932, '肢': 1933, '搥': 1934, '欠': 1935, '漲': 1936, 'G': 1937, 'M': 1938, '劇': 1939, '躍': 1940, '童': 1941, '幼': 1942, '稚': 1943, '渾': 1944, '噩': 1945, 'S': 1946, '鹼': 1947, '彩': 1948, '齒': 1949, '曹': 1950, '蠢': 1951, '欲': 1952, '川': 1953, '龔': 1954, '鮮': 1955, '托': 1956, '抿': 1957, '蜜': 1958, '螃': 1959, '蟹': 1960, '斟': 1961, '酌': 1962, '米': 1963, '倫': 1964, '狗': 1965, '舌': 1966, '狼': 1967, '勝': 1968, '娜': 1969, '索': 1970, '隆': 1971, '釘': 1972, '燃': 1973, '稅': 1974, '郎': 1975, '雞': 1976, '穀': 1977, '糧': 1978, '饅': 1979, '爭': 1980, '搶': 1981, '瓣': 1982, '腴': 1983, '貼': 1984, '踩': 1985, '摺': 1986, '蓮': 1987, '寶': 1988, '梢': 1989, '橢': 1990, '蒜': 1991, '椎': 1992, '揍': 1993, '笨': 1994, '娘': 1995, '杉': 1996, '磯': 1997, '莊': 1998, '豪': 1999, '鄉': 2000, '瞞': 2001, '浸': 2002, '廁': 2003, '宮': 2004, '頸': 2005, '構': 2006, '摔': 2007, '洪': 2008, '澤': 2009, '恭': 2010, '酵': 2011, '憊': 2012, '唄': 2013, '矽': 2014, '晶': 2015, '脾': 2016, '奈': 2017, '娛': 2018, '寡': 2019, '契': 2020, '夕': 2021, '裸': 2022, '嘶': 2023, '幣': 2024, '策': 2025, '衍': 2026, '雅': 2027, '瞧': 2028, '南': 2029, '姓': 2030, '乏': 2031, '游': 2032, '泳': 2033, '堤': 2034, '腕': 2035, '匆': 2036, '辰': 2037, '猴': 2038, '斜': 2039, '祖': 2040, '釣': 2041, '撿': 2042, '污': 2043, '言': 2044, '挨': 2045, '閒': 2046, '蕭': 2047, '-': 2048, '煞': 2049, '惰': 2050, '役': 2051, 'U': 2052, '麗': 2053, '邂': 2054, '逅': 2055, '悟': 2056, '稠': 2057, 'F': 2058, '狂': 2059, '廝': 2060, '守': 2061, '株': 2062, '檯': 2063, '勵': 2064, '恰': 2065, '獨': 2066, '峰': 2067, '廟': 2068, '融': 2069, '諱': 2070, '銀': 2071, '悔': 2072, '折': 2073, '喲': 2074, '蛛': 2075, '絲': 2076, '蒐': 2077, '瘋': 2078, '澄': 2079, '昇': 2080, '墊': 2081, '尺': 2082, '兼': 2083, '篇': 2084, '慧': 2085, '酷': 2086, 'j': 2087, '琉': 2088, '掰': 2089, '魂': 2090, '詞': 2091, '憑': 2092, '德': 2093, '瑞': 2094, '販': 2095, '售': 2096, '街': 2097, '呆': 2098, '繞': 2099, '濾': 2100, '庚': 2101, '瘀': 2102, '矮': 2103, '跨': 2104, '撲': 2105, '仿': 2106, '酪': 2107, '乳': 2108, '窄': 2109, '虎': 2110, '餘': 2111, '疝': 2112, '磷': 2113, '堪': 2114, '卵': 2115, '孕': 2116, '盂': 2117, 'z': 2118, '唷': 2119, '腱': 2120, '蠕': 2121, '釜': 2122, '齡': 2123, '拾': 2124, '靶': 2125, '援': 2126, '搖': 2127, '祂': 2128, '惚': 2129, '婆': 2130, '咪': 2131, '駕': 2132, '災': 2133, '股': 2134, '杏': 2135, '輝': 2136, '悠': 2137, '揮': 2138, '盆': 2139, '皇': 2140, '魯': 2141, '餓': 2142, '耍': 2143, '趾': 2144, '剪': 2145, '巫': 2146, '癱': 2147, '裂': 2148, '趴': 2149, '憩': 2150, 's': 2151, 'u': 2152, 'b': 2153, 't': 2154, 'a': 2155, 'i': 2156, 'o': 2157, 'n': 2158, '悸': 2159, '昂': 2160, '栓': 2161, '邁': 2162, '丁': 2163, '彙': 2164, '鋒': 2165, '伸': 2166, '甩': 2167, '螢': 2168, '幕': 2169, '澡': 2170, '竇': 2171, '巡': 2172, '搔': 2173, '仔': 2174, '鋪': 2175, '押': 2176, '船': 2177, '聞': 2178, '颱': 2179, '鳴': 2180, '咿': 2181, '繃': 2182, '墅': 2183, '噪': 2184, '錘': 2185, '播': 2186, '誘': 2187, '昧': 2188, '蝨': 2189, '覽': 2190, '孵': 2191, '芽': 2192, '秤': 2193, '斬': 2194, '攻': 2195, '擊': 2196, '撫': 2197, '製': 2198, '濫': 2199, '贊': 2200, '銜': 2201, '閨': 2202, '刮': 2203, '鬍': 2204, '叉': 2205, '恕': 2206, '亡': 2207, '矜': 2208, '諸': 2209, '貞': 2210, '坊': 2211, '燈': 2212, '句': 2213, '競': 2214, '旺': 2215, '批': 2216, '趁': 2217, '巧': 2218, '卻': 2219, '搜': 2220, '鋌': 2221, '稀': 2222, '唱': 2223, '歌': 2224, '宵': 2225, '搏': 2226, '噎': 2227, '欄': 2228, '塵': 2229, '嗅': 2230, '喪': 2231, '怠': 2232, '畜': 2233, '牧': 2234, '瘟': 2235, '肋': 2236, '粗': 2237, '橡': 2238, '俊': 2239, '盟': 2240, '濺': 2241, '鈉': 2242, '鉀': 2243, '獻': 2244, '柏': 2245, '霉': 2246, '敷': 2247, '蕁': 2248, '鉤': 2249, '萱': 2250, '淤': 2251, '嫌': 2252, '燜': 2253, '蔬': 2254, '姪': 2255, '添': 2256, '渴': 2257, '烏': 2258, '洱': 2259, '啞': 2260, '嚥': 2261, '嗝': 2262, '鑿': 2263, '迴': 2264, '蹋': 2265, '遲': 2266, '鈍': 2267, '朗': 2268, '槌': 2269, '廳': 2270, '廚': 2271, '汽': 2272, '喊': 2273, '芳': 2274, '珠': 2275, '僵': 2276, '楊': 2277, '喂': 2278, '編': 2279, '略': 2280, '猶': 2281, '豫': 2282, '逝': 2283, '籃': 2284, '撕': 2285, '渡': 2286, '棟': 2287, '衰': 2288, '踴': 2289, '讚': 2290, '/': 2291, '劉': 2292, '捷': 2293, '‧': 2294, '徘': 2295, '徊': 2296, '鍛': 2297, '鍊': 2298, '絨': 2299, '碘': 2300, '痕': 2301, '淒': 2302, '頰': 2303, '偉': 2304, '哲': 2305, '愈': 2306, '凡': 2307, '咽': 2308, '苔': 2309, '梗': 2310, '扯': 2311, '顴': 2312, '秘': 2313, '奎': 2314, '玉': 2315, 'k': 2316, '噻': 2317, '徐': 2318, '揭': 2319, '萍': 2320, '灶': 2321, '蛀': 2322, '鄭': 2323, '捨': 2324, '佳': 2325, '拳': 2326, '胡': 2327, '賴': 2328, '剝': 2329, '怡': 2330, '痙': 2331, '攣': 2332, '臭': 2333, '蘆': 2334, '忌': 2335, '苓': 2336, '嗆': 2337, '碳': 2338, '垂': 2339, '鍾': 2340, '宏': 2341, '屜': 2342, '亭': 2343, '攙': 2344, '兄': 2345, '籬': 2346, '婷': 2347, '凱': 2348, '催': 2349, '冠': 2350, '招': 2351, '姆': 2352, '挂': 2353, '財': 2354, '牡': 2355, '蠣': 2356, '董': 2357, '脱': 2358, '碟': 2359, '伍': 2360, '鱔': 2361, '羹': 2362, '咻': 2363, '港': 2364, '胱': 2365, '斌': 2366, '垢': 2367, '熙': 2368, '芭': 2369, '艾': 2370, '萊': 2371, '躁': 2372, '筍': 2373, '豬': 2374, '糙': 2375, '咕': 2376, '嚕': 2377, '抄': 2378, '塔': 2379, '醃': 2380, '漬': 2381, '腐': 2382, '亢': 2383, '涂': 2384, '霖': 2385, '唇': 2386, '癲': 2387, '綁': 2388, 'f': 2389, 'e': 2390, 'r': 2391, 'c': 2392, 'm': 2393, 'p': 2394, 'l': 2395, 'w': 2396, 'h': 2397, 'y': 2398, '胺': 2399, '驅': 2400, '惜': 2401, '軍': 2402, '皂': 2403, '沐': 2404, '浴': 2405, '檸': 2406, '檬': 2407, '薑': 2408, '汁': 2409, '醋': 2410, '瓢': 2411, '疔': 2412, '唸': 2413, '嬰': 2414, '吊': 2415, '寵': 2416, '慈': 2417, '翹': 2418, '枕': 2419, '席': 2420, '吱': 2421, '胎': 2422, '疸': 2423, '癇': 2424, '歪': 2425, '糰': 2426, '揪': 2427, '毯': 2428, '戳': 2429, '汎': 2430, '傅': 2431, '唆': 2432, '恆': 2433, '妨': 2434, '棋': 2435, '贅': 2436, '薪': 2437, '沈': 2438, '漂': 2439, '紗': 2440, '寢': 2441, '賓': 2442, '嗦': 2443, '撮': 2444, '欽': 2445, '茄': 2446, '耀': 2447, '仁': 2448, '枚': 2449, '墾': 2450, '貢': 2451, '湖': 2452, '暢': 2453, '竭': 2454, '巾': 2455, '厠': 2456, '邦': 2457, '奧': 2458, '愫': 2459, '崩': 2460, '臀': 2461, '騷': 2462, '貓': 2463, '妝': 2464, '蔻': 2465, '蕩': 2466, '哭': 2467, '凶': 2468, '撩': 2469, '晉': 2470, '裕': 2471, '蒸': 2472, '烤': 2473, '惑': 2474, '鹿': 2475, '謀': 2476, '呈': 2477, '搂': 2478, '屯': 2479, '泛': 2480, '孳': 2481, '騰': 2482, '谷': 2483, '潑': 2484, '洋': 2485, '胚': 2486, '擁': 2487}
tag2id = {'O': 0, 'B-time': 1, 'I-time': 2, 'B-location': 3, 'I-location': 4, 'B-med_exam': 5, 'I-med_exam': 6, 'B-profession': 7, 'I-profession': 8, 'B-name': 9, 'I-name': 10, 'B-family': 11, 'I-family': 12, 'B-ID': 13, 'I-ID': 14, 'B-clinical_event': 15, 'I-clinical_event': 16, 'B-education': 17, 'I-education': 18, 'B-money': 19, 'I-money': 20, 'B-contact': 21, 'I-contact': 22, 'B-organization': 23, 'I-organization': 24, 'B-others': 25, 'I-others': 26}
# +
BiLSTMCRF_MODEL_PATH = './SavePkl/Temp/1227-2100 (0.68)/bilstm_crf.pkl'
REMOVE_O = False
print("Load File...")
# train_word_lists, train_tag_lists, word2id, tag2id = build_corpus("train", make_vocab=False)
# dev_word_lists, dev_tag_lists = build_corpus("dev", make_vocab=False)
test_word_lists, test_tag_lists = build_corpus("test", make_vocab=False)
print("Testing...")
crf_word2id, crf_tag2id = extend_maps(word2id, tag2id, for_crf=True)
bilstm_model = load_model(BiLSTMCRF_MODEL_PATH)
bilstm_model.model.bilstm.bilstm.flatten_parameters() # remove warning
test_word_lists, test_tag_lists = prepocess_data_for_lstmcrf(test_word_lists, test_tag_lists, test=True)
lstmcrf_pred, target_tag_list = bilstm_model.test(test_word_lists, test_tag_lists, crf_word2id, crf_tag2id)
print("article_id\tstart_position\tend_position\tentity_text\tentity_type")
output = ""
start = ""
end = ""
tag = ""
line = ""
output += "article_id\tstart_position\tend_position\tentity_text\tentity_type\n"
flag = 0
for chap in range(len(test_word_lists)):
for text in range(len(test_word_lists[chap][:-1])):
if lstmcrf_pred[chap][text][0] == 'B':
if line != "":
end = text
if end-start == len(line):
output += "{:}\t{:}\t{:}\t{:}\t{:}\n".format(chap, start, end, line, tag)
print("{:}\t{:}\t{:}\t{:}\t{:}".format(chap, start, end, line, tag))
flag = 0
start = text
tag = lstmcrf_pred[chap][text][2:]
line = ""
line += str(test_word_lists[chap][text])
flag = 1
elif lstmcrf_pred[chap][text][0] == 'I':
if flag == 0:
start = text
tag = lstmcrf_pred[chap][text][2:]
flag = 1
line += str(test_word_lists[chap][text])
elif lstmcrf_pred[chap][text][0] == 'O':
if line != "":
end = text
if end-start == len(line):
output += "{:}\t{:}\t{:}\t{:}\t{:}\n".format(chap, start, end, line, tag)
print("{:}\t{:}\t{:}\t{:}\t{:}".format(chap, start, end, line, tag))
line = ""
flag = 0
output_path = 'output.tsv'
with open(output_path, 'w', encoding='utf-8') as f:
f.write(output)
# -
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 测试我们的算法
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data
y = iris.target
X.shape
y.shape
# ### 1. 把数据集分为训练集和测试集
# +
# 方法一:把矩阵 X 和向量 y 合并后打乱,然后再分开
data = np.hstack([X, y.reshape(-1, 1)])
np.random.shuffle(data)
X = data[:, :4]
y = data[:, 4]
test_ratio = 0.2
test_size = int(len(X) * test_ratio)
# 获取训练集
X_train = X[test_size:]
y_train = y[test_size:]
# 获取测试集
X_test = X[:test_size]
y_test = y[:test_size]
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# +
# 方法二:利用随机索引
# shuffle_indexes = np.random.permutation(len(X)) # 生成随机索引
# test_ratio = 0.2
# test_size = int(len(X) * test_ratio)
# test_indexes = shuffle_indexes[:test_size]
# train_indexes = shuffle_indexes[test_size:]
# X_train = X[train_indexes]
# y_train = y[train_indexes]
# X_test = X[test_indexes]
# y_test = y[test_indexes]
# print(X_train.shape)
# print(y_train.shape)
# print(X_test.shape)
# print(y_test.shape)
# -
# ### 2. 使用我们的算法测试
from kNN.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
from kNN.kNN import KNNClassifier
my_knn_clf = KNNClassifier(k=3)
my_knn_clf.fit(X_train, y_train)
y_predict = my_knn_clf.predict(X_test)
y_predict
y_test
sum(y_predict == y_test)
sum(y_predict == y_test)/len(y_test)
# ### 3. sklearn 中的 train_test_split
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state= 333)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
| ML-Base-MOOC/chapt-2 KNN/02-kNN_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pathlib import Path
from sklearn.datasets import make_regression
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler, MinMaxScaler, LabelEncoder
train_df = pd.read_csv(Path('Resources/2019loans.csv'))
test_df = pd.read_csv(Path('Resources/2020Q1loans.csv'))
train_df.head()
test_df.head()
# +
# Convert categorical data to numeric and separate target feature for training data
train_df = train_df.drop(['index', 'Unnamed: 0'], axis = 1)
test_df = test_df.drop(['index', 'Unnamed: 0'], axis = 1)
X_train = train_df.drop('loan_status', axis = 1)
Y_train = train_df['loan_status']
X_test = test_df.drop('loan_status', axis = 1)
Y_test= test_df['loan_status']
X_train.shape, Y_train.shape, X_test.shape, Y_test.shape
# -
# add missing dummy variables to testing set
X_trainer = pd.get_dummies(X_train, drop_first=True)
Y_trainer = Y_train.replace({'low_risk':0, 'high_risk':1})
X_tester = pd.get_dummies(X_test, drop_first=True)
Y_tester = Y_test.replace({'low_risk':0, 'high_risk':1})
# +
X_trainer.shape, X_tester.shape
missing = []
for col in X_trainer.columns:
if col not in X_tester.columns:
X_tester[col] = 0
missing.append(col)
print(len(X_trainer.columns), len(X_tester.columns))
# -
# Train the Logistic Regression model on the unscaled data and print the model score
from sklearn.linear_model import LogisticRegression
LR = LogisticRegression(solver='lbfgs', random_state=1)
LR.fit(X_trainer, Y_trainer)
print(f"Training Score: {LR.score(X_trainer, Y_trainer)}")
print(f"Testing Score: {LR.score(X_tester, Y_tester)}")
# Train a Random Forest Classifier model and print the model score
from sklearn.ensemble import RandomForestClassifier
RF = RandomForestClassifier(random_state=1)
RF.fit(X_trainer,Y_trainer)
print(f'Training Score: {RF.score(X_trainer,Y_trainer)}')
print(f"Testing Score: {RF.score(X_tester, Y_tester)}")
# Scale the data
scaler = StandardScaler().fit(X_trainer)
X_trainer_scaled = scaler.transform(X_trainer)
X_tester_scaled = scaler.transform(X_tester)
# Train the Logistic Regression model on the scaled data and print the model score
LR = LogisticRegression(solver='lbfgs', random_state=1)
LR.fit(X_trainer_scaled, Y_trainer)
print(f"Training Score: {LR.score(X_trainer_scaled, Y_trainer)}")
print(f"Testing Score: {LR.score(X_tester_scaled, Y_tester)}")
# Train a Random Forest Classifier model on the scaled data and print the model score
rfc = RandomForestClassifier(random_state=1, n_estimators=500).fit(X_trainer_scaled, Y_trainer)
print(f'Testing Score: {rfc.score(X_trainer_scaled, Y_trainer)}')
print(f'Testing Score: {rfc.score(X_tester_scaled, Y_tester)}')
| Credit Risk Evaluator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import pyproj
import numpy as np
import geoviews as gv
import geoviews.feature as gf
from bokeh.sampledata.airport_routes import airports, routes
gv.extension('matplotlib')
gv.output(fig='svg', size=250)
# -
# ## Define data
# +
def get_circle_path(start, end, samples=200):
sx, sy = start
ex, ey = end
g = pyproj.Geod(ellps='WGS84')
(az12, az21, dist) = g.inv(sx, sy, ex, ey)
lonlats = g.npts(sx, sy, ex, ey, samples)
return np.array([(sx, sy)]+lonlats+[(ex, ey)])
# Compute great-circle paths for all US flight routes from Honolulu
paths = []
honolulu = (-157.9219970703125, 21.318700790405273)
routes = routes[routes.SourceID==3728]
airports = airports[airports.AirportID.isin(list(routes.DestinationID)+[3728])]
for i, route in routes.iterrows():
airport = airports[airports.AirportID==route.DestinationID].iloc[0]
paths.append(get_circle_path(honolulu, (airport.Longitude, airport.Latitude)))
# Define Graph from Nodes and EdgePaths
path = gv.EdgePaths(paths)
points = gv.Nodes(airports, ['Longitude', 'Latitude', 'AirportID'], ['Name', 'City'])
graph = gv.Graph((routes, points, path), ['SourceID', 'DestinationID'])
# -
# ## Plot
gf.ocean * gf.land * gf.lakes * gf.coastline * graph.opts(
node_color='black', node_size=8)
| examples/gallery/matplotlib/great_circle.ipynb |