code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import gpytorch
import matplotlib.pyplot as plt
import numpy as np
from gpytorch.means import Mean
import seaborn as sns
sns.set_style('white')
palette = ["#1b4079", "#C6DDF0", "#048A81", "#B9E28C", "#8C2155", "#AF7595", "#E6480F", "#FA9500"]
sns.set(palette = palette, font_scale=2.0, style="white", rc={"lines.linewidth": 4.0})
# +
class FeatureExtractor(torch.nn.Sequential):
def __init__(self, widths=[50, 50]):
super(FeatureExtractor, self).__init__()
self.add_module('linear0', torch.nn.Linear(1, widths[0]))
self.add_module('relu0', torch.nn.ReLU())
for lyr in range(1, len(widths)):
self.add_module('linear' + str(lyr),
torch.nn.Linear(widths[lyr-1], widths[lyr]))
self.add_module('relu' + str(lyr), torch.nn.ReLU())
self.add_module('linear' + str(len(widths)+1),
torch.nn.Linear(widths[-1], 1))
class NNMean(Mean):
def __init__(self, input_size, batch_shape=torch.Size(), widths=[50,50]):
super().__init__()
self.net = FeatureExtractor(widths)
def forward(self, x):
return self.net(x).squeeze()
class NNGPModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood, widths=[50,50]):
super(NNGPModel, self).__init__(train_x, train_y, likelihood)
self.mean_module = NNMean(train_x.shape[-1], widths)
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
class ExactGPModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(ExactGPModel, self).__init__(train_x, train_y, likelihood)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
# -
torch.manual_seed(85234)
gen_lh = gpytorch.likelihoods.GaussianLikelihood()
gen_lh.noise = torch.tensor([0.02])
gen_model = ExactGPModel(None, None, gen_lh)
gen_model.covar_module.base_kernel.lengthscale = torch.tensor([0.75])
gen_model.eval();
# +
ntest = 50
ntrain = 50
full_x = torch.linspace(0, 10, ntrain + ntest).unsqueeze(-1)
# -
full_y = gen_lh(gen_model(full_x)).sample()
plt.scatter(full_x, full_y)
train_x = full_x[:ntrain]
train_y = full_y[:ntrain]
test_x = full_x[ntrain:]
test_y = full_y[ntrain:]
# ## Fit NN Model
lh = gpytorch.likelihoods.GaussianLikelihood()
model = NNGPModel(train_x, train_y, lh)
# +
training_iter = 500
# Find optimal model hyperparameters
model.train()
lh.train()
# Use the adam optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=0.01) # Includes GaussianLikelihood parameters
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(lh, model)
for i in range(training_iter):
# Zero gradients from previous iteration
optimizer.zero_grad()
# Output from model
output = model(train_x)
# Calc loss and backprop gradients
loss = -mll(output, train_y)
loss.backward()
if i % 50 == 0:
print('Iter %d/%d - Loss: %.3f lengthscale: %.3f noise: %.3f' % (
i + 1, training_iter, loss.item(),
model.covar_module.base_kernel.lengthscale.item(),
model.likelihood.noise.item()
))
optimizer.step()
# -
plt.plot(train_x, model(train_x).mean.detach())
plt.scatter(train_x, train_y)
# +
lh.eval();
model.eval();
pred_ = model(full_x)
pred_lh_ = lh(pred_)
pred_mean = pred_.mean.detach()
pred_lower, pred_upper = pred_.confidence_region()
# -
plt.scatter(train_x, train_y, label="Train", color='k')
plt.scatter(test_x, test_y, label="Test", color='OrangeRed')
plt.plot(full_x, pred_mean, label="GP Posterior", color='steelblue')
plt.fill_between(full_x.squeeze(), pred_lower.detach(), pred_upper.detach(),
alpha=0.1)
plt.legend()
plt.scatter(train_x, train_y, label="Train", color=palette[4])
plt.scatter(test_x, test_y, label="Test", color=palette[7])
plt.plot(full_x, pred_mean, label="GP Posterior", color=palette[0],
lw=3.5)
plt.fill_between(full_x.squeeze(), pred_lower.detach(), pred_upper.detach(),
alpha=0.4, color=palette[1])
plt.legend(bbox_to_anchor=(1., 0.75),
frameon=False)
plt.xlabel("X")
plt.ylabel("Y")
sns.despine()
plt.savefig("./gp-overfitting.pdf", bbox_inches='tight')
np.savez("../../plots/data/gp_mean_overfitting.npz",
train_x=train_x.detach().numpy()[:, 0],
train_y=train_y.detach().numpy(),
test_x=test_x.detach().numpy()[:, 0],
test_y=test_y.detach().numpy(),
full_x=full_x.detach().numpy()[:, 0],
pred_mean=pred_mean.detach().numpy(),
pred_sigma=pred_.stddev.detach().numpy(),
pred_sigma_wnoise=pred_lh_.stddev.detach().numpy(),
)
# ## Regular GP
lh = gpytorch.likelihoods.GaussianLikelihood()
model = ExactGPModel(train_x, train_y, lh)
# +
training_iter = 500
# Find optimal model hyperparameters
model.train()
lh.train()
# Use the adam optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=0.01) # Includes GaussianLikelihood parameters
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(lh, model)
for i in range(training_iter):
# Zero gradients from previous iteration
optimizer.zero_grad()
# Output from model
output = model(train_x)
# Calc loss and backprop gradients
loss = -mll(output, train_y)
loss.backward()
if i % 50 == 0:
print('Iter %d/%d - Loss: %.3f lengthscale: %.3f noise: %.3f' % (
i + 1, training_iter, loss.item(),
model.covar_module.base_kernel.lengthscale.item(),
model.likelihood.noise.item()
))
optimizer.step()
# +
lh.eval();
model.eval();
pred_ = model(full_x)
pred_lh_ = lh(pred_)
pred_mean = pred_.mean.detach()
pred_lower, pred_upper = pred_.confidence_region()
# -
plt.scatter(train_x, train_y, label="Train", color='k')
plt.scatter(test_x, test_y, label="Test", color='OrangeRed')
plt.plot(full_x, pred_mean, label="GP Posterior", color='steelblue')
plt.fill_between(full_x.squeeze(), pred_lower.detach(), pred_upper.detach(),
alpha=0.1)
plt.legend()
plt.scatter(train_x, train_y, label="Train", color=palette[4])
plt.scatter(test_x, test_y, label="Test", color=palette[7])
plt.plot(full_x, pred_mean, label="GP Posterior", color=palette[0],
lw=3.5)
plt.fill_between(full_x.squeeze(), pred_lower.detach(), pred_upper.detach(),
alpha=0.4, color=palette[1])
plt.legend(bbox_to_anchor=(1., 0.75),
frameon=False)
plt.xlabel("X")
plt.ylabel("Y")
sns.despine()
plt.savefig("./gp-overfitting.pdf", bbox_inches='tight')
np.savez("../../plots/data/gp_mean_overfitting_baseline.npz",
train_x=train_x.detach().numpy()[:, 0],
train_y=train_y.detach().numpy(),
test_x=test_x.detach().numpy()[:, 0],
test_y=test_y.detach().numpy(),
full_x=full_x.detach().numpy()[:, 0],
pred_mean=pred_mean.detach().numpy(),
pred_sigma=pred_.stddev.detach().numpy(),
pred_sigma_wnoise=pred_lh_.stddev.detach().numpy(),
)
# ## Analysis
print(gen_model.covar_module.base_kernel.lengthscale)
print(model.covar_module.base_kernel.lengthscale)
print(gen_model.likelihood.noise)
print(model.likelihood.noise)
plt.scatter(train_x, train_y, label="Train Data", color=palette[4])
plt.plot(train_x, model.mean_module(train_x).detach(), label="GP Mean Function", color=palette[0])
plt.legend(bbox_to_anchor=(1., 0.75),
frameon=False)
plt.xlabel("X")
plt.ylabel("Y")
sns.despine()
# plt.savefig("./gp-overfitting.pdf", bbox_inches='tight')
| GP_experiments/GP_Overfitting/mlp_gp_mean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
import os
import tqdm
import time
import spectrum
from sklearn.decomposition import PCA
from sklearn import svm
import warnings
warnings.filterwarnings("ignore")
# -
# ## process training datasets => flatten
BASE_DIR= "data/数据挖掘题目/近红外试题1/"
TRAINING_DIR= os.path.join(BASE_DIR, "建模集光谱")
chemical_excel = os.path.join(BASE_DIR, "建模集化学值.xlsx")
TEST_DIR= "验证集光谱"
SHEET_NAME="初始化学值"
TEST_DIR= os.path.join(BASE_DIR, "验证集光谱")
# +
# add a helper func to read Excel file
def get_chemical_source(file_path: str, sheet_name=None)-> pd.DataFrame:
chemical_data = pd.read_excel(file_path, sheet_name)
return chemical_data
chemical_data = get_chemical_source(chemical_excel, "初始化学值")
chemical_data.tail(2)
# -
def flatten_speactrum(training_dir: str)-> pd.DataFrame:
spectrum_data= list()
file_number= list()
for file in os.listdir(training_dir):
try:
data = (pd.read_csv(os.path.join(training_dir,file), \
header= None, \
float_precision='high', \
index_col= None))
spectrum_data.append(data.iloc[:,1].values)
file_number.append(file.split(".")[0])
except Exception as e:
print(str(e))
pass
return pd.DataFrame({"file_number":file_number, "spectrum_data": spectrum_data})
flatten_speactrum(TRAINING_DIR)
def process_flatten_data(df_flatten, column_name):
df_flatten["file_number"]= df_flatten.iloc[:, 0].apply(lambda x: int(x))
df_flatten.rename(columns={"file_number":"样本序号"}, inplace= True)
merged_df= chemical_data.merge(df_flatten, on="样本序号").rename(columns={"spectrum_data": "光谱数据"})
return merged_df
merged_flatten_df= process_flatten_data(flatten_speactrum(TRAINING_DIR), "file_number")
merged_flatten_df.tail(2)
# spectrum_display
def plot_spectrum(file):
plt.figure(figsize=(8, 6), dpi=100)
for i in range(len(file.index)):
plt.xlabel("Wavelength") #波长
plt.ylabel("Absorbance (AU)") #吸光度
plt.plot(merged_flatten_df["光谱数据"][i])
plt.grid(True)
plt.tight_layout()
plot_spectrum(merged_flatten_df["光谱数据"])
# #### Pre_processing - standardization
# - normalisation, smoothing or derivatives were applied to its training data, as well as the spectrum range and sampling frequency of the training data.
# +
from sklearn.preprocessing import StandardScaler
def standard_row(row):
scaler = StandardScaler()
return scaler.fit_transform(row.reshape(-1,1))
merged_flatten_df["标准化光谱数据"]= merged_flatten_df.loc[:,"光谱数据"].apply(lambda x: standard_row(x))
merged_flatten_df.tail(2)
# -
sns.distplot(merged_flatten_df["标准化光谱数据"][0])
# ## Baseline
# #### PCA to reduce dims then use classification model
# +
def flatten2row(training_dir: str)-> pd.DataFrame:
spectrum_data= list()
file_name = list()
for file in os.listdir(training_dir):
try:
data = (pd.read_csv(os.path.join(training_dir,file), \
header= None, \
index_col= None))
spectrum_data.append(data.iloc[:,1].values)
file_name.append(file.split(".")[0])
except Exception as e:
pass
return pd.DataFrame(spectrum_data), file_name
train_data, file_name= flatten2row(TRAINING_DIR)
test_data, _= flatten2row(TEST_DIR)
# -
train_data.tail(2)
# #### Find peak time of each spectrum, during certain frequency
plt.plot(train_data[200])
plt.plot(train_data[0])
plt.plot(train_data[10])
train_data["样本序号"]= file_name
train_data["样本序号"]= train_data["样本序号"].apply(lambda x: int(x))
train_data.tail(2)
# +
from spectrum import Periodogram, data_cosine
p = Periodogram(train_data)
p.plot(marker='o')
# -
merged_df= chemical_data.merge(train_data, on="样本序号", how= "left")
merged_df.tail(2)
# #### SVM
# +
from sklearn import svm
#task 1
cls = svm.SVC(kernel="linear")
#train the model
merged_df.dropna(inplace= True)
cls.fit(merged_df.iloc[:, 4:], merged_df["等级"])
#predict the response
pred = cls.predict(test_data)
pred
# -
cls.fit(merged_df.iloc[:, 4:], merged_df["样本序号"])
#predict the response
pred_sample = cls.predict(test_data)
pred_sample
pca = PCA(n_components=100)
principalComponents = pd.DataFrame(pca.fit_transform(merged_df.iloc[:, 4:]))
# #### KNN
# +
from sklearn.neighbors import KNeighborsClassifier
neigh = KNeighborsClassifier(n_neighbors=100)
neigh.fit(merged_df.iloc[:, 4:], merged_df.loc[:, "样本序号"])
# -
neigh.predict(test_data)
# ## Task 2
BASE_DIR_2= "data/数据挖掘题目/近红外试题2/"
TRAINING_DIR_2= os.path.join(BASE_DIR_2, "建模光谱/")
chemical_excel_2 = os.path.join(BASE_DIR_2, "建模化学值.xlsx")
TEST_DIR_2= "验证光谱/"
TEST_DIR_2= os.path.join(BASE_DIR_2, "验证光谱/")
chemical_data_2= pd.read_excel(chemical_excel_2)
chemical_data_2.tail(2)
chemical_data_2["sample_type"]= chemical_data_2["名称"].apply(lambda title: title.split(".")[0])
def get_data(train_dir):
col1= list()
col2= list()
file_number= list()
start = time.time()
for file in os.listdir(train_dir):
data= pd.read_csv(train_dir+ file, header=None, names=["col", "col2"])
for i,k in data.loc[:, ["col","col2"]].values:
col1.append(i)
col2.append(k)
file_number.append(file.split(".")[0])
end = time.time() -start
print(f"process time costs {end}")
return pd.DataFrame({"sample_type": file_number, "col":col1, "col2": col2})
train_data_2 = get_data(TRAINING_DIR_2)
train_data_2.tail(2)
megered_df_2= train_data_2.merge(chemical_data_2.iloc[:, [2,3]], on="sample_type")
megered_df_2.tail(2)
megered_df_2["烟碱"] = megered_df_2["烟碱"].apply(lambda x: str(x))
megered_df_2["Y"]= megered_df_2[["sample_type","烟碱"]].apply(lambda x: '_'.join(x), axis=1)
# +
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=500)
model.fit(megered_df_2[["col", "col2"]],megered_df_2["Y"])
# -
predicted= model.predict([["4532.51627173342","0.5663"]])
print(predicted)
test_data_2= get_data(TEST_DIR_2)
test_data_2.dropna(inplace= True)
try:
test_data_2["pred_label"]= test_data_2[["col", "col2"]].apply(model.predict)
except Exception as e:
print(e)
pass
# +
# test_data_2[test_data_2["col"]=="\x06"]
# -
predicated5= model.predict([["8205.73","0.2867"]])
predicated5
# +
#encoding categorical labels
from sklearn.preprocessing import OneHotEncoder
ohe = OneHotEncoder()
ohe.fit_transform(megered_df_2[["sample_type"]])
# -
# #### use SVM for these all tasks
# - after testing with PCA, which didn't generated good model result by random checking, not apply model inference yet by checking model accuracy using ground truth label.
# +
# # for task2
# cls = svm.SVC(kernel="linear")
# cls.fit(megered_df_2.iloc[:, [1,2]], megered_df_2["烟碱"])
# #predict the tabacco
# pred_2= cls.predict(test_data_2.iloc[:, [1,2]])
# pred_2
# +
BASE_DIR_3= "data/数据挖掘题目/近红外试题3/"
TRAINING_DIR_3= os.path.join(BASE_DIR_3, "建模集光谱/")
chemical_excel_3 = os.path.join(BASE_DIR_3, "建模集化学值.xlsx")
TEST_DIR_3= "验证集光谱/"
TEST_DIR_3= os.path.join(BASE_DIR_3, "验证集光谱/")
chemical_data_3= pd.read_excel(chemical_excel_3)[["Sample", "化学值"]]
chemical_data_3.tail(2)
chemical_data_3["样本序号"]= chemical_data_3["Sample"].apply(lambda x: int(x.split("-")[-1]))
df_flatten= flatten_speactrum(TRAINING_DIR_3)
df_flatten["file_number"]= df_flatten.iloc[:, 0].apply(lambda x: int(x))
df_flatten= df_flatten.rename(columns={"file_number": "样本序号"})
merged_df= chemical_data_3.merge(df_flatten, on="样本序号").rename(columns={"spectrum_data": "光谱数据"})
merged_df
# -
train_data3, file_name3= flatten2row(TRAINING_DIR_3)
test_data3, _= flatten2row(TEST_DIR_3)
train_data3.head(2)
train_data["样本序号"]= file_name
train_data["样本序号"]= train_data["样本序号"].apply(lambda x: int(x))
train_data.tail(2)
merged_df_3= chemical_data_3.merge(train_data, on="样本序号")
#task 3
cls = svm.SVC(kernel="linear")
#train the model
merged_df.dropna(inplace= True)
cls.fit(merged_df_3.iloc[:, 3:], merged_df_3["Sample"])
#predict the response
pred = cls.predict(test_data)
pred
| spectral_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/agemagician/Prot-Transformers/blob/master/Benchmark/Bert.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="jKLvLEnt0aw7" colab_type="text"
# <h3> Benchmark ProtBert Model using GPU or CPU <h3>
# + [markdown] id="3jXYUooT0aw9" colab_type="text"
# <b>1. Load necessry libraries including huggingface transformers<b>
# + id="DpoYBdbs0jSS" colab_type="code" colab={}
# !pip install -q transformers
# + id="GnGZuNIm0axA" colab_type="code" colab={}
import torch
from transformers import BertModel
import time
from datetime import timedelta
import os
import requests
from tqdm.auto import tqdm
# + [markdown] id="XqmVMZYw0axG" colab_type="text"
# <b>2. Set the url location of ProtBert and the vocabulary file<b>
# + id="D9-pJtmU0axH" colab_type="code" colab={}
modelUrl = 'https://www.dropbox.com/s/dm3m1o0tsv9terq/pytorch_model.bin?dl=1'
configUrl = 'https://www.dropbox.com/s/d3yw7v4tvi5f4sk/bert_config.json?dl=1'
vocabUrl = 'https://www.dropbox.com/s/jvrleji50ql5m5i/vocab.txt?dl=1'
# + [markdown] id="Cv_-QMOE1EME" colab_type="text"
# <b>3. Download ProtBert models and vocabulary files<b>
# + id="GpJpbyfJ1LSU" colab_type="code" colab={}
downloadFolderPath = 'models/ProtBert/'
# + id="hhrYlqAu1LbS" colab_type="code" colab={}
modelFolderPath = downloadFolderPath
modelFilePath = os.path.join(modelFolderPath, 'pytorch_model.bin')
configFilePath = os.path.join(modelFolderPath, 'config.json')
vocabFilePath = os.path.join(modelFolderPath, 'vocab.txt')
# + id="9RgoS6bj1Lga" colab_type="code" colab={}
if not os.path.exists(modelFolderPath):
os.makedirs(modelFolderPath)
# + id="5nQI_lPu1ZBx" colab_type="code" colab={}
def download_file(url, filename):
response = requests.get(url, stream=True)
with tqdm.wrapattr(open(filename, "wb"), "write", miniters=1,
total=int(response.headers.get('content-length', 0)),
desc=filename) as fout:
for chunk in response.iter_content(chunk_size=4096):
fout.write(chunk)
# + id="c7UfPSyL1ZHM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 164, "referenced_widgets": ["2bf939cf51634a2891810201a22b3ed6", "8b575ac374044f919678aa4239d0e118", "f9ddd85e5db144f4a102245a4be9bb15", "69c86dd28d3549969f18721bb7543047", "9f77377124ea4ad29aa936d83132e881", "0601b95b5292494e86d450032e13b027", "9c01724ea4e84ff49652981c9b3e30ce", "<KEY>", "<KEY>", "<KEY>", "54bddeac0f8d4a2a87f74395e9415e12", "<KEY>", "<KEY>", "d0287581330d4a42be15d24f8c264824", "b7392b1ae6244e298440be33e03997c5", "<KEY>", "5385f58962f64a528d2f36d84249cd93", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "bb0e6825d00249129b6db2708f84efb9", "<KEY>", "250af8ec98654827a917f86f587b927b"]} outputId="de11ba9a-6e64-43ae-a97c-3030094020aa"
if not os.path.exists(modelFilePath):
download_file(modelUrl, modelFilePath)
if not os.path.exists(configFilePath):
download_file(configUrl, configFilePath)
if not os.path.exists(vocabFilePath):
download_file(vocabUrl, vocabFilePath)
# + [markdown] id="DA8VJMAk0axN" colab_type="text"
# <b>4. Load ProtBert Model<b>
# + id="hjXcKjqg0axP" colab_type="code" colab={}
model = BertModel.from_pretrained(modelFolderPath)
# + [markdown] id="hKD2Qvde0axT" colab_type="text"
# <b>5. Load the model into the GPU if avilabile and switch to inference mode<b>
# + id="ksASffyj0axU" colab_type="code" colab={}
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# + id="hxf0RX9o0axX" colab_type="code" colab={}
model = model.to(device)
model = model.eval()
# + [markdown] id="JGRdbkad0axb" colab_type="text"
# <b>6. Benchmark Configuration<b>
# + id="JFkGZ1gT0axl" colab_type="code" colab={}
min_batch_size = 8
max_batch_size = 32
inc_batch_size = 8
min_sequence_length = 64
max_sequence_length = 512
inc_sequence_length = 64
iterations = 10
# + [markdown] id="atUiO9Y10axr" colab_type="text"
# <b>7. Start Benchmarking<b>
# + id="xbAIdSYl0axs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 748} outputId="072af120-6da7-4bbc-ab2a-a45e407e8e70"
device_name = torch.cuda.get_device_name(device.index) if device.type == 'cuda' else 'CPU'
with torch.no_grad():
print((' Benchmarking using ' + device_name + ' ').center(80, '*'))
print(' Start '.center(80, '*'))
for sequence_length in range(min_sequence_length,max_sequence_length+1,inc_sequence_length):
for batch_size in range(min_batch_size,max_batch_size+1,inc_batch_size):
start = time.time()
for i in range(iterations):
input_ids = torch.randint(1, 20, (batch_size,sequence_length)).to(device)
results = model(input_ids)[0].cpu().numpy()
end = time.time()
ms_per_protein = (end-start)/(iterations*batch_size)
print('Sequence Length: %4d \t Batch Size: %4d \t Ms per protein %4.2f' %(sequence_length,batch_size,ms_per_protein))
print(' Done '.center(80, '*'))
print(' Finished '.center(80, '*'))
| Benchmark/ProtBert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
# +
from tensorflow.keras import layers
from tensorflow.keras.regularizers import l2
from tensorflow.keras.layers import Activation, Conv1D, Conv2D, Input, Lambda
from tensorflow.keras.layers import BatchNormalization, Flatten, Dense, Reshape
from tensorflow.keras.layers import (
MaxPooling2D,
AveragePooling2D,
GlobalAveragePooling2D,
)
import tensorflow.keras as keras
import tensorflow as tf
import tensorflow.keras.backend as K
weight_decay = 1e-4
def identity_block_2D(
input_tensor, kernel_size, filters, stage, block, trainable = True
):
"""The identity block is the block that has no conv layer at shortcut.
# Arguments
input_tensor: input tensor
kernel_size: default 3, the kernel size of middle conv layer at main path
filters: list of integers, the filterss of 3 conv layer at main path
stage: integer, current stage label, used for generating layer names
block: 'a','b'..., current block label, used for generating layer names
# Returns
Output tensor for the block.
"""
filters1, filters2, filters3 = filters
bn_axis = 3
conv_name_1 = 'conv' + str(stage) + '_' + str(block) + '_1x1_reduce'
bn_name_1 = 'conv' + str(stage) + '_' + str(block) + '_1x1_reduce/bn'
x = Conv2D(
filters1,
(1, 1),
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_1,
)(input_tensor)
x = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_1
)(x)
x = Activation('relu')(x)
conv_name_2 = 'conv' + str(stage) + '_' + str(block) + '_3x3'
bn_name_2 = 'conv' + str(stage) + '_' + str(block) + '_3x3/bn'
x = Conv2D(
filters2,
kernel_size,
padding = 'same',
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_2,
)(x)
x = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_2
)(x)
x = Activation('relu')(x)
conv_name_3 = 'conv' + str(stage) + '_' + str(block) + '_1x1_increase'
bn_name_3 = 'conv' + str(stage) + '_' + str(block) + '_1x1_increase/bn'
x = Conv2D(
filters3,
(1, 1),
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_3,
)(x)
x = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_3
)(x)
x = layers.add([x, input_tensor])
x = Activation('relu')(x)
return x
def conv_block_2D(
input_tensor,
kernel_size,
filters,
stage,
block,
strides = (2, 2),
trainable = True,
):
"""A block that has a conv layer at shortcut.
# Arguments
input_tensor: input tensor
kernel_size: default 3, the kernel size of middle conv layer at main path
filters: list of integers, the filterss of 3 conv layer at main path
stage: integer, current stage label, used for generating layer names
block: 'a','b'..., current block label, used for generating layer names
# Returns
Output tensor for the block.
Note that from stage 3, the first conv layer at main path is with strides=(2,2)
And the shortcut should have strides=(2,2) as well
"""
filters1, filters2, filters3 = filters
bn_axis = 3
conv_name_1 = 'conv' + str(stage) + '_' + str(block) + '_1x1_reduce'
bn_name_1 = 'conv' + str(stage) + '_' + str(block) + '_1x1_reduce/bn'
x = Conv2D(
filters1,
(1, 1),
strides = strides,
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_1,
)(input_tensor)
x = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_1
)(x)
x = Activation('relu')(x)
conv_name_2 = 'conv' + str(stage) + '_' + str(block) + '_3x3'
bn_name_2 = 'conv' + str(stage) + '_' + str(block) + '_3x3/bn'
x = Conv2D(
filters2,
kernel_size,
padding = 'same',
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_2,
)(x)
x = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_2
)(x)
x = Activation('relu')(x)
conv_name_3 = 'conv' + str(stage) + '_' + str(block) + '_1x1_increase'
bn_name_3 = 'conv' + str(stage) + '_' + str(block) + '_1x1_increase/bn'
x = Conv2D(
filters3,
(1, 1),
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_3,
)(x)
x = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_3
)(x)
conv_name_4 = 'conv' + str(stage) + '_' + str(block) + '_1x1_proj'
bn_name_4 = 'conv' + str(stage) + '_' + str(block) + '_1x1_proj/bn'
shortcut = Conv2D(
filters3,
(1, 1),
strides = strides,
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = trainable,
kernel_regularizer = l2(weight_decay),
name = conv_name_4,
)(input_tensor)
shortcut = BatchNormalization(
axis = bn_axis, trainable = trainable, name = bn_name_4
)(shortcut)
x = layers.add([x, shortcut])
x = Activation('relu')(x)
return x
def resnet_2D_v1(inputs, mode = 'train'):
bn_axis = 3
# if mode == 'train':
# inputs = Input(shape=input_dim, name='input')
# else:
# inputs = Input(shape=(input_dim[0], None, input_dim[-1]), name='input')
# ===============================================
# Convolution Block 1
# ===============================================
x1 = Conv2D(
64,
(7, 7),
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = True,
kernel_regularizer = l2(weight_decay),
padding = 'same',
name = 'conv1_1/3x3_s1',
)(inputs)
x1 = BatchNormalization(
axis = bn_axis, name = 'conv1_1/3x3_s1/bn', trainable = True
)(x1)
x1 = Activation('relu')(x1)
x1 = MaxPooling2D((2, 2), strides = (2, 2))(x1)
# ===============================================
# Convolution Section 2
# ===============================================
x2 = conv_block_2D(
x1,
3,
[48, 48, 96],
stage = 2,
block = 'a',
strides = (1, 1),
trainable = True,
)
x2 = identity_block_2D(
x2, 3, [48, 48, 96], stage = 2, block = 'b', trainable = True
)
# ===============================================
# Convolution Section 3
# ===============================================
x3 = conv_block_2D(
x2, 3, [96, 96, 128], stage = 3, block = 'a', trainable = True
)
x3 = identity_block_2D(
x3, 3, [96, 96, 128], stage = 3, block = 'b', trainable = True
)
x3 = identity_block_2D(
x3, 3, [96, 96, 128], stage = 3, block = 'c', trainable = True
)
# ===============================================
# Convolution Section 4
# ===============================================
x4 = conv_block_2D(
x3, 3, [128, 128, 256], stage = 4, block = 'a', trainable = True
)
x4 = identity_block_2D(
x4, 3, [128, 128, 256], stage = 4, block = 'b', trainable = True
)
x4 = identity_block_2D(
x4, 3, [128, 128, 256], stage = 4, block = 'c', trainable = True
)
# ===============================================
# Convolution Section 5
# ===============================================
x5 = conv_block_2D(
x4, 3, [256, 256, 512], stage = 5, block = 'a', trainable = True
)
x5 = identity_block_2D(
x5, 3, [256, 256, 512], stage = 5, block = 'b', trainable = True
)
x5 = identity_block_2D(
x5, 3, [256, 256, 512], stage = 5, block = 'c', trainable = True
)
y = MaxPooling2D((3, 1), strides = (2, 1), name = 'mpool2')(x5)
return inputs, y
def resnet_2D_v2(inputs, mode = 'train'):
bn_axis = 3
# if mode == 'train':
# inputs = Input(shape=input_dim, name='input')
# else:
# inputs = Input(shape=(input_dim[0], None, input_dim[-1]), name='input')
# ===============================================
# Convolution Block 1
# ===============================================
x1 = Conv2D(
64,
(7, 7),
strides = (2, 2),
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = True,
kernel_regularizer = l2(weight_decay),
padding = 'same',
name = 'conv1_1/3x3_s1',
)(inputs)
x1 = BatchNormalization(
axis = bn_axis, name = 'conv1_1/3x3_s1/bn', trainable = True
)(x1)
x1 = Activation('relu')(x1)
x1 = MaxPooling2D((2, 2), strides = (2, 2))(x1)
# ===============================================
# Convolution Section 2
# ===============================================
x2 = conv_block_2D(
x1,
3,
[64, 64, 256],
stage = 2,
block = 'a',
strides = (1, 1),
trainable = True,
)
x2 = identity_block_2D(
x2, 3, [64, 64, 256], stage = 2, block = 'b', trainable = True
)
x2 = identity_block_2D(
x2, 3, [64, 64, 256], stage = 2, block = 'c', trainable = True
)
# ===============================================
# Convolution Section 3
# ===============================================
x3 = conv_block_2D(
x2, 3, [128, 128, 512], stage = 3, block = 'a', trainable = True
)
x3 = identity_block_2D(
x3, 3, [128, 128, 512], stage = 3, block = 'b', trainable = True
)
x3 = identity_block_2D(
x3, 3, [128, 128, 512], stage = 3, block = 'c', trainable = True
)
# ===============================================
# Convolution Section 4
# ===============================================
x4 = conv_block_2D(
x3,
3,
[256, 256, 1024],
stage = 4,
block = 'a',
strides = (1, 1),
trainable = True,
)
x4 = identity_block_2D(
x4, 3, [256, 256, 1024], stage = 4, block = 'b', trainable = True
)
x4 = identity_block_2D(
x4, 3, [256, 256, 1024], stage = 4, block = 'c', trainable = True
)
# ===============================================
# Convolution Section 5
# ===============================================
x5 = conv_block_2D(
x4, 3, [512, 512, 2048], stage = 5, block = 'a', trainable = True
)
x5 = identity_block_2D(
x5, 3, [512, 512, 2048], stage = 5, block = 'b', trainable = True
)
x5 = identity_block_2D(
x5, 3, [512, 512, 2048], stage = 5, block = 'c', trainable = True
)
y = MaxPooling2D((3, 1), strides = (2, 1), name = 'mpool2')(x5)
return inputs, y
class VladPooling(keras.layers.Layer):
"""
This layer follows the NetVlad, GhostVlad
"""
def __init__(self, mode, k_centers, g_centers = 0, **kwargs):
self.k_centers = k_centers
self.g_centers = g_centers
self.mode = mode
super(VladPooling, self).__init__(**kwargs)
def build(self, input_shape):
self.cluster = self.add_weight(
shape = [self.k_centers + self.g_centers, input_shape[0][-1]],
name = 'centers',
initializer = 'orthogonal',
)
self.built = True
def compute_output_shape(self, input_shape):
assert input_shape
return (input_shape[0][0], self.k_centers * input_shape[0][-1])
def call(self, x):
# feat : bz x W x H x D, cluster_score: bz X W x H x clusters.
feat, cluster_score = x
num_features = feat.shape[-1]
# softmax normalization to get soft-assignment.
# A : bz x W x H x clusters
max_cluster_score = K.max(cluster_score, -1, keepdims = True)
exp_cluster_score = K.exp(cluster_score - max_cluster_score)
A = exp_cluster_score / K.sum(
exp_cluster_score, axis = -1, keepdims = True
)
# Now, need to compute the residual, self.cluster: clusters x D
A = K.expand_dims(A, -1) # A : bz x W x H x clusters x 1
feat_broadcast = K.expand_dims(
feat, -2
) # feat_broadcast : bz x W x H x 1 x D
feat_res = (
feat_broadcast - self.cluster
) # feat_res : bz x W x H x clusters x D
weighted_res = tf.multiply(
A, feat_res
) # weighted_res : bz x W x H x clusters x D
cluster_res = K.sum(weighted_res, [1, 2])
if self.mode == 'gvlad':
cluster_res = cluster_res[:, : self.k_centers, :]
cluster_l2 = K.l2_normalize(cluster_res, -1)
outputs = K.reshape(
cluster_l2, [-1, int(self.k_centers) * int(num_features)]
)
return outputs
def amsoftmax_loss(y_true, y_pred, scale = 30, margin = 0.35):
y_pred = y_true * (y_pred - margin) + (1 - y_true) * y_pred
y_pred *= scale
return K.categorical_crossentropy(y_true, y_pred, from_logits = True)
def vggvox_resnet2d_icassp(
inputs, num_class = 8631, mode = 'train', args = None
):
net = 'resnet34s'
loss = 'softmax'
vlad_clusters = 8
ghost_clusters = 2
bottleneck_dim = 512
aggregation = 'gvlad'
mgpu = 0
if net == 'resnet34s':
inputs, x = resnet_2D_v1(inputs, mode = mode)
else:
inputs, x = resnet_2D_v2(inputs, mode = mode)
x_fc = keras.layers.Conv2D(
bottleneck_dim,
(7, 1),
strides = (1, 1),
activation = 'relu',
kernel_initializer = 'orthogonal',
use_bias = True,
trainable = True,
kernel_regularizer = keras.regularizers.l2(weight_decay),
bias_regularizer = keras.regularizers.l2(weight_decay),
name = 'x_fc',
)(x)
# ===============================================
# Feature Aggregation
# ===============================================
if aggregation == 'avg':
if mode == 'train':
x = keras.layers.AveragePooling2D(
(1, 5), strides = (1, 1), name = 'avg_pool'
)(x)
x = keras.layers.Reshape((-1, bottleneck_dim))(x)
else:
x = keras.layers.GlobalAveragePooling2D(name = 'avg_pool')(x)
x = keras.layers.Reshape((1, bottleneck_dim))(x)
elif aggregation == 'vlad':
x_k_center = keras.layers.Conv2D(
vlad_clusters,
(7, 1),
strides = (1, 1),
kernel_initializer = 'orthogonal',
use_bias = True,
trainable = True,
kernel_regularizer = keras.regularizers.l2(weight_decay),
bias_regularizer = keras.regularizers.l2(weight_decay),
name = 'vlad_center_assignment',
)(x)
x = VladPooling(
k_centers = vlad_clusters, mode = 'vlad', name = 'vlad_pool'
)([x_fc, x_k_center])
elif aggregation == 'gvlad':
x_k_center = keras.layers.Conv2D(
vlad_clusters + ghost_clusters,
(7, 1),
strides = (1, 1),
kernel_initializer = 'orthogonal',
use_bias = True,
trainable = True,
kernel_regularizer = keras.regularizers.l2(weight_decay),
bias_regularizer = keras.regularizers.l2(weight_decay),
name = 'gvlad_center_assignment',
)(x)
x = VladPooling(
k_centers = vlad_clusters,
g_centers = ghost_clusters,
mode = 'gvlad',
name = 'gvlad_pool',
)([x_fc, x_k_center])
else:
raise IOError('==> unknown aggregation mode')
x = keras.layers.Dense(
bottleneck_dim,
activation = 'relu',
kernel_initializer = 'orthogonal',
use_bias = True,
trainable = True,
kernel_regularizer = keras.regularizers.l2(weight_decay),
bias_regularizer = keras.regularizers.l2(weight_decay),
name = 'fc6',
)(x)
if loss == 'softmax':
y = keras.layers.Dense(
num_class,
activation = 'softmax',
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = True,
kernel_regularizer = keras.regularizers.l2(weight_decay),
bias_regularizer = keras.regularizers.l2(weight_decay),
name = 'prediction',
)(x)
trnloss = 'categorical_crossentropy'
elif loss == 'amsoftmax':
x_l2 = keras.layers.Lambda(lambda x: K.l2_normalize(x, 1))(x)
y = keras.layers.Dense(
num_class,
kernel_initializer = 'orthogonal',
use_bias = False,
trainable = True,
kernel_constraint = keras.constraints.unit_norm(),
kernel_regularizer = keras.regularizers.l2(weight_decay),
bias_regularizer = keras.regularizers.l2(weight_decay),
name = 'prediction',
)(x_l2)
trnloss = amsoftmax_loss
else:
raise IOError('==> unknown loss.')
return y
# -
class Model:
def __init__(self):
self.X = tf.placeholder(tf.float32, [None, 257, None, 1])
params = {'dim': (257, None, 1),
'nfft': 512,
'spec_len': 250,
'win_length': 400,
'hop_length': 160,
'n_classes': 5994,
'sampling_rate': 16000,
'normalize': True,
}
self.logits = vggvox_resnet2d_icassp(self.X, num_class=2, mode='eval')
self.logits = tf.identity(self.logits, name = 'logits')
ckpt_path = 'output-vggvox-v2-vad/model.ckpt-300000'
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model()
sess.run(tf.global_variables_initializer())
var_lists = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES)
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, ckpt_path)
saver = tf.train.Saver()
saver.save(sess, 'out/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
and 'Assign' not in n.name
]
)
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('out', strings)
# +
# def load_graph(frozen_graph_filename):
# with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
# graph_def = tf.GraphDef()
# graph_def.ParseFromString(f.read())
# with tf.Graph().as_default() as graph:
# tf.import_graph_def(graph_def)
# return graph
def load_graph(frozen_graph_filename, **kwargs):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
# https://github.com/onnx/tensorflow-onnx/issues/77#issuecomment-445066091
# to fix import T5
for node in graph_def.node:
if node.op == 'RefSwitch':
node.op = 'Switch'
for index in xrange(len(node.input)):
if 'moving_' in node.input[index]:
node.input[index] = node.input[index] + '/read'
elif node.op == 'AssignSub':
node.op = 'Sub'
if 'use_locking' in node.attr:
del node.attr['use_locking']
elif node.op == 'AssignAdd':
node.op = 'Add'
if 'use_locking' in node.attr:
del node.attr['use_locking']
elif node.op == 'Assign':
node.op = 'Identity'
if 'use_locking' in node.attr:
del node.attr['use_locking']
if 'validate_shape' in node.attr:
del node.attr['validate_shape']
if len(node.input) == 2:
node.input[0] = node.input[1]
del node.input[1]
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
# -
g = load_graph('out/frozen_model.pb')
x = g.get_tensor_by_name('import/Placeholder:0')
logits = g.get_tensor_by_name('import/logits:0')
# !tar -czvf output-vggvox-v2-vad-300k.tar.gz output-vggvox-v2-vad
| session/vad/export/vggvox-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import os
from pathlib import Path
import tqdm
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### Reading
def txt_to_matrix(filename, line_skip = 5):
f = open (filename, 'r')
# Lineskip, cleaning, conversion
data = f.readlines()[line_skip:]
return np.asarray(
[l.replace("\n", "").split() for l in data]
).astype(np.float32)
def get_time_step(root, index):
wse = txt_to_matrix(root + '/decoded--' + index + '.WSE')
dep = txt_to_matrix(root + '/decoded--' + index + '.DEP')
vvx = txt_to_matrix(root + '/decoded--' + index + '.VVX')
vvy = txt_to_matrix(root + '/decoded--' + index + '.VVY')
# timestep: matrice 801 rows x 4 misurazioni x 1256 colonne (valori)
return np.array(list(zip(wse, dep, vvx, vvy)))
# +
rootdir = '../output/'
timesteps = []
paths = [p for p in sorted(os.listdir('../output'))]
x = 0
ceiling = 50
# Read all dirs and process them
for path in tqdm.tqdm(paths):
if x >= ceiling: break
# Processing
path = rootdir + path
timesteps.append(
get_time_step(
path, ("{:04d}".format(x))
)
)
x += 1
timesteps = np.asarray(timesteps).astype(np.float32)
# -
timesteps.shape
# ### Processing
# +
import matplotlib.animation as animation
# WSE animation
data = timesteps[:,:,0,:]
ims = []
for frame in data:
im = plt.imshow(frame)
ims.append([im])
ani = animation.ArtistAnimation(fig, ims, interval=50, blit=True,
repeat_delay=1000)
# ani.save('dynamic_images.mp4')
plt.show()
# -
plt.plot(dep_progression)
| notebooks/.ipynb_checkpoints/00-exploratory-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="0O88LX8mS6tq"
# Importing Libraries
import tensorflow as tf
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Dense
from tensorflow.keras.callbacks import EarlyStopping
import torch
import pickle
from sklearn.metrics import classification_report, roc_auc_score, confusion_matrix, precision_recall_curve
# -
tf.config.list_physical_devices('GPU')
# +
# embedding_dir = "/home/andrewngo/Desktop/MLTracker/model_20220217192920"
embedding_dir = "/home/andrewngo/Desktop/MLTracker/model_CUC_20220311120344" # model folder
path = torch.load(embedding_dir + "/path.pt")
path_embedding = torch.load(embedding_dir + "/out_1.pt")
out_index_path = torch.load(embedding_dir + "/out_index_path_1.pt")
out_mal_train_val_dict = torch.load(embedding_dir + "/out_mal_train_val_1.pt")
out_mal_train_val_index_path = torch.load(embedding_dir + "/out_mal_train_val_index_path_1.pt")
out_mal_test_dict = torch.load(embedding_dir + "/out_mal_test_1.pt")
out_mal_test_index_path = torch.load(embedding_dir + "/out_mal_test_index_path_1.pt")
out_normal_dict = torch.load(embedding_dir + "/out_normal_1.pt")
out_normal_index_path = torch.load(embedding_dir + "/out_normal_index_path_1.pt")
labels = torch.load(embedding_dir + "/path_labels_1.pt")
model_metapath2vec = torch.load(embedding_dir + "/model.pt")
# +
dir_graph = '/home/andrewngo/Desktop/MLTracker/graph_data_20220305160351'
computer2nodeid = torch.load(dir_graph + "/computer2nodeid.pt")
user2nodeid = torch.load(dir_graph + "/user2nodeid.pt")
# -
# # Random shuffling testing strategy with benign sample sampling from graph (Mentioned in paper)
# +
out_mal_test_keys = out_mal_test_dict
out_mal_val_keys = out_mal_train_val_dict
out_normal_keys = out_normal_dict
seed = 40
# out_mal_test_keys = list(out_mal_test_dict.keys())
# out_mal_val_keys = list(out_mal_train_val_dict.keys())
# out_normal_keys = list(out_normal_dict.keys())
# +
# len(path_embedding.values[: int(len(path_embedding.values)/2)])
benign_path = path_embedding.values[:labels.count(0)]
malicious_path = path_embedding.values[labels.count(0):]
benign_path
training_sample = 100000
val_normal_sample = 5000
test_normal_sample = 5000
#validation + testing = 1 for easier code, the correct ratio value should be validation + testing + training = 1
validation_ratio = 0.5
test_ratio = 0.5
from numpy.random import permutation
np.random.seed(20)
perm = permutation(len(benign_path))
# normal_train_data_idx =
normal_train_data_idx = perm[:training_sample]
perm = permutation(len(out_normal_keys))
perm_val = perm[:val_normal_sample]
perm_test = perm[val_normal_sample:(val_normal_sample+test_normal_sample)]
perm = permutation(len(malicious_path))
mal_val_data_idx = perm[:int(len(perm)*test_ratio)]
mal_test_data_idx = perm[int(len(perm)*test_ratio):]
# normal_train_data_idx
# normal_val_data_idx
# normal_test_data_idx
# mal_val_data_idx
# mal_test_data_idx
normal_train_data = np.asarray([benign_path[i] for i in normal_train_data_idx])
out_test_keys_sample = [out_normal_keys[i] for i in perm_test]
normal_test_data = np.asarray([out_normal_dict[i].tolist() for i in out_test_keys_sample])
out_val_keys_sample = [out_normal_keys[i] for i in perm_val]
normal_val_data = np.asarray([out_normal_dict[i].tolist() for i in out_val_keys_sample])
mal_val_data = np.asarray([malicious_path[i] for i in mal_val_data_idx])
mal_test_data = np.asarray([malicious_path[i] for i in mal_test_data_idx])
# print(normal_train_data)
# Initializing a MinMax Scaler
scaler = MinMaxScaler()
# Fitting the train data to the scaler
data_scaled = scaler.fit(normal_train_data)
normal_train_data = data_scaled.transform(normal_train_data)
normal_val_data = data_scaled.transform(normal_val_data)
normal_test_data = data_scaled.transform(normal_test_data)
mal_val_data = data_scaled.transform(mal_val_data)
mal_test_data = data_scaled.transform(mal_test_data)
# normal_train_data
test_data = np.concatenate((normal_test_data,mal_test_data), axis=0)
val_data = np.concatenate((normal_val_data, mal_val_data), axis=0)
labels_test = [0 for i in range(len(normal_test_data))] + [1 for i in range(len(mal_test_data))]
labels_val = [0 for i in range(len(normal_val_data))] + [1 for i in range(len(mal_val_data))]
# normal_train_data, normal_test_data, train_labels, test_labels = train_test_split(benign_path, , test_size = 0.2, random_state = 111)
# -
(0.82+0.93+0.86+0.92+0.88)/5
(0.94+0.93+0.97+0.91+0.94)/5
# # Random split testing with benign sampling on log file
# +
len(path_embedding.values[: int(len(path_embedding.values)/2)])
benign_path = path_embedding.values[:labels.count(0)]
malicious_path = path_embedding.values[labels.count(0):]
benign_path
training_sample = 10000
val_normal_sample = 5000
test_normal_sample = 5000
#validation + testing = 1 for easier code, the correct ratio value should be validation + testing + training = 1
validation_ratio = 0.5
test_ratio = 0.5
from numpy.random import permutation
np.random.seed(20)
perm = permutation(len(out_normal_keys))
# normal_train_data_idx =
perm_val = perm[:val_normal_sample]
perm_test = perm[val_normal_sample:(val_normal_sample+test_normal_sample)]
perm_train = perm[(val_normal_sample+test_normal_sample):(val_normal_sample+test_normal_sample+training_sample)]
perm = permutation(len(malicious_path))
mal_val_data_idx = perm[:int(len(perm)*test_ratio)]
mal_test_data_idx = perm[int(len(perm)*test_ratio):]
# normal_train_data_idx
# normal_val_data_idx
# normal_test_data_idx
# mal_val_data_idx
# mal_test_data_idx
out_train_keys_sample = [out_normal_keys[i] for i in perm_train]
normal_train_data = np.asarray([out_normal_dict[i].tolist() for i in out_train_keys_sample])
out_test_keys_sample = [out_normal_keys[i] for i in perm_test]
normal_test_data = np.asarray([out_normal_dict[i].tolist() for i in out_test_keys_sample])
out_val_keys_sample = [out_normal_keys[i] for i in perm_val]
normal_val_data = np.asarray([out_normal_dict[i].tolist() for i in out_val_keys_sample])
mal_val_data = np.asarray([malicious_path[i] for i in mal_val_data_idx])
mal_test_data = np.asarray([malicious_path[i] for i in mal_test_data_idx])
# print(normal_train_data)
# Initializing a MinMax Scaler
scaler = MinMaxScaler()
# Fitting the train data to the scaler
data_scaled = scaler.fit(normal_train_data)
normal_train_data = data_scaled.transform(normal_train_data)
normal_val_data = data_scaled.transform(normal_val_data)
normal_test_data = data_scaled.transform(normal_test_data)
mal_val_data = data_scaled.transform(mal_val_data)
mal_test_data = data_scaled.transform(mal_test_data)
# normal_train_data
test_data = np.concatenate((normal_test_data,mal_test_data), axis=0)
val_data = np.concatenate((normal_val_data, mal_val_data), axis=0)
labels_test = [0 for i in range(len(normal_test_data))] + [1 for i in range(len(mal_test_data))]
labels_val = [0 for i in range(len(normal_val_data))] + [1 for i in range(len(mal_val_data))]
# normal_train_data, normal_test_data, train_labels, test_labels = train_test_split(benign_path, , test_size = 0.2, random_state = 111)
# -
# # Split by day testing strategy
# +
training_sample = 100000
val_normal_sample = 5000
test_normal_sample = 5000
# Randomly select normal sample for training, testing, validation sets
from numpy.random import permutation
np.random.seed(seed)
perm = permutation(len(out_normal_keys))
perm_val = perm[:val_normal_sample]
perm_test = perm[(val_normal_sample):(val_normal_sample+test_normal_sample)]
perm_train = perm[(val_normal_sample+test_normal_sample):(training_sample+val_normal_sample+test_normal_sample)]
# out_train_keys_sample =
normal_train_data = np.asarray([out_normal_keys[i].tolist() for i in perm_train])
# out_test_keys_sample =
normal_test_data = np.asarray([out_normal_keys[i].tolist() for i in perm_test])
# out_val_keys_sample =
normal_val_data = np.asarray([out_normal_keys[i].tolist() for i in perm_val])
mal_val_data = np.asarray([i.tolist() for i in out_mal_train_val_dict])
mal_test_data = np.asarray([i.tolist() for i in out_mal_test_dict])
normal_train_data_path = [out_normal_index_path[i] for i in perm_train]
normal_test_data_path = [out_normal_index_path[i] for i in perm_test]
normal_val_data_path = [out_normal_index_path[i] for i in perm_val]
mal_val_data_path = [out_mal_train_val_index_path[i] for i in out_mal_train_val_index_path]
mal_test_data_path = [out_mal_test_index_path[i] for i in out_mal_test_index_path]
scaler = MinMaxScaler()
# Fitting the train data to the scaler
data_scaled = scaler.fit(normal_train_data)
normal_train_data = data_scaled.transform(normal_train_data)
normal_val_data = data_scaled.transform(normal_val_data)
normal_test_data = data_scaled.transform(normal_test_data)
mal_val_data = data_scaled.transform(mal_val_data)
mal_test_data = data_scaled.transform(mal_test_data)
test_data = np.concatenate((normal_test_data,mal_test_data), axis=0)
val_data = np.concatenate((normal_val_data, mal_val_data), axis=0)
val_data_path = normal_train_data_path + mal_val_data_path
test_data_path = normal_test_data_path + mal_test_data_path
labels_test = [0 for i in range(len(normal_test_data))] + [1 for i in range(len(mal_test_data))]
labels_val = [0 for i in range(len(normal_val_data))] + [1 for i in range(len(mal_val_data))]
# +
# Train
# -
print("Train Data: " + str(len(normal_train_data)))
print("Validation Normal: " + str(len(normal_val_data)))
print("Validation Malicious: " + str(len(mal_val_data)))
print("Test Normal: " + str(len(normal_test_data)))
print("Test Malicious: " + str(len(mal_test_data)))
# +
# # Initializing a MinMax Scaler
# scaler = MinMaxScaler()
# # Fitting the train data to the scaler
# data_scaled = scaler.fit(normal_train_data)
# normal_train_data = data_scaled.transform(normal_train_data)
# normal_test_data = data_scaled.transform(normal_test_data)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="S2ligR14O2Yk" outputId="da6da3d0-72ca-4220-de6e-5035c3d1e5e8"
# plotting the first three normal data points
plt.plot(mal_test_data[0])
plt.plot(mal_test_data[1])
plt.plot(mal_test_data[3])
# plt.plot(mal_test_data[4])
plt.plot(mal_test_data[5])
# plt.plot(mal_test_data[2])
# -
normal_test_data[4]
plt.plot(normal_test_data[0])
plt.plot(normal_test_data[1])
plt.plot(normal_test_data[2])
plt.plot(normal_test_data[3])
plt.plot(normal_test_data[4])
plt.plot(normal_test_data[5])
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="MTzbDn_XPVJP" outputId="760b543d-dcd2-4e67-b6b9-3541e29d5f69"
# plotting the first three anomaly data points
plt.plot(normal_train_data[0])
plt.plot(normal_train_data[1])
plt.plot(normal_train_data[2])
# -
class Autoencoder(Model):
def __init__(self):
super(Autoencoder, self).__init__()
self.encoder = Sequential([
Dense(128, activation='tanh'),
# Dense(32, activation='relu'),
Dense(64, activation='tanh'),
# Dense(8, activation='relu')
])
self.decoder = Sequential([
Dense(64, activation='tanh'),
# Dense(32, activation='rel'),
Dense(128, activation='tanh'),
])
def call(self,x):
encoded = self.encoder(x)
decoded = self.decoder(encoded)
return decoded
# +
# class Autoencoder(Model):
# def __init__(self):
# super(Autoencoder, self).__init__()
# self.encoder = Sequential([
# Dense(128, activation='tanh'),
# Dense(64, activation='tanh'),
# Dense(32, activation='tanh'),
# Dense(16, activation='tanh'),
# Dense(8, activation='tanh')
# ])
# self.decoder = Sequential([
# Dense(16, activation='tanh'),
# Dense(32, activation='tanh'),
# Dense(64, activation='tanh'),
# Dense(128, activation='tanh')
# ])
# def call(self,x):
# encoded = self.encoder(x)
# decoded = self.decoder(encoded)
# return decoded
# + id="EaejgzKQfrGC"
# Instantiating the Autoencoder
model = Autoencoder()
# creating an early_stopping
# early_stopping = EarlyStopping(monitor='val_loss',
# patience = 20,
# mode = 'min')
early_stopping = EarlyStopping(
monitor='val_loss',
min_delta=0.0001,
patience=20,
verbose=1,
mode='min',
restore_best_weights=True)
# Compiling the model
model.compile(optimizer = 'adam',
loss = 'mae')
# model.compile(optimizer = 'adam',
# loss = 'mse')
# -
len(normal_train_data)
# +
# Training the model
validate = np.concatenate((normal_train_data, normal_val_data), axis=0)
history = model.fit(normal_train_data,normal_train_data,
epochs = 500,
batch_size = 128,
validation_data = (validate,validate),
shuffle = True,
callbacks = [early_stopping])
# -
plt.plot(history.history['loss'], linewidth=2, label='Train')
plt.plot(history.history['val_loss'], linewidth=2, label='Test')
plt.legend(loc='upper right')
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
#plt.ylim(ymin=0.70,ymax=1)
plt.show()
# predictions for normal test data points
encoder_out = model.encoder(normal_val_data).numpy()
decoder_out = model.decoder(encoder_out).numpy()
# plotting normal test data point and its predictiction by the autoencoder
plt.plot(normal_test_data[0], 'b')
plt.plot(decoder_out[0], 'r')
# predictions for anomaly test data points
encoder_out_a = model.encoder(mal_test_data).numpy()
decoder_out_a = model.decoder(encoder_out_a).numpy()
# plotting anomaly test data point and its predictiction by the autoencoder
plt.plot(mal_test_data[0], 'b')
plt.plot(decoder_out_a[0], 'r')
# +
# reconstruction loss for anomaly test data
reconstructions_a = model.predict(val_data)
val_loss = tf.keras.losses.mae(reconstructions_a, val_data)
# Plotting histogram for recontruction loss for anomaly test data
# plt.hist(train_loss_a, bins = 20)
# +
# feature scaling
from sklearn.metrics import roc_curve, roc_auc_score
from matplotlib import pyplot
prob_val = (val_loss-min(val_loss))/(max(val_loss)-min(val_loss))
fpr, tpr, thresholds = roc_curve(labels_val,prob_val)
# plot the roc curve for the model
pyplot.plot([0,1], [0,1], linestyle='--', label='No Skill')
pyplot.plot(fpr, tpr, marker='.', label='Logistic')
# axis labels
pyplot.xlabel('False Positive Rate')
pyplot.ylabel('True Positive Rate')
pyplot.legend()
# show the plot
pyplot.show()
# -
from numpy import sqrt
from numpy import argmax
gmeans = sqrt(tpr * (1-fpr))
# locate the index of the largest g-mean
ix = argmax(gmeans)
print('Best Threshold=%f, G-Mean=%.3f' % (thresholds[ix], gmeans[ix]))
# plot the roc curve for the model
pyplot.plot([0,1], [0,1], linestyle='--', label='No Skill')
pyplot.plot(fpr, tpr, marker='.', label='Logistic')
pyplot.scatter(fpr[ix], tpr[ix], marker='o', color='black', label='Best')
# axis labels
pyplot.xlabel('False Positive Rate')
pyplot.ylabel('True Positive Rate')
pyplot.legend()
# show the plot
pyplot.show()
threshold = thresholds[ix]
auc = roc_auc_score(labels_val, prob_val, average=None)
auc
# Number of correct predictions for Normal test data
from sklearn.metrics import classification_report, roc_auc_score, confusion_matrix
pred_val = [0 if prob_val[i] < threshold else 1 for i in range(len(prob_val))]
print(roc_auc_score(labels_val, pred_val, average=None))
print(classification_report(labels_val, pred_val, labels=[0, 1], target_names=['benign', 'malicious']))
print(confusion_matrix(labels_val, pred_val))
# +
threshold_fixed = 0.001
val_x_predictions = model.predict(val_data)
mse = np.mean(np.power(val_data - val_x_predictions, 2), axis=1)
error_df = pd.DataFrame({'Reconstruction_error': mse,
'True_class': labels_val})
groups = error_df.groupby('True_class')
fig, ax = plt.subplots()
for name, group in groups:
ax.plot(group.index, group.Reconstruction_error, marker='o', ms=0.3, linestyle='',
label= "Malicious" if name == 1 else "Normal")
ax.hlines(threshold_fixed, ax.get_xlim()[0], ax.get_xlim()[1], colors="r", zorder=100, label='Threshold')
ax.legend()
plt.title("Reconstruction error for normal and malicious data")
plt.ylabel("Reconstruction error")
plt.xlabel("Data point index")
plt.show();
# -
reconstructions_a = model.predict(test_data)
test_loss = tf.keras.losses.mae(reconstructions_a, test_data)
prob_test = (test_loss-min(test_loss))/(max(test_loss)-min(test_loss))
precision, recall, thresholds = precision_recall_curve(labels_test,prob_test)
f1_scores = 2*recall*precision/(recall+precision)
f1_scores = np.nan_to_num(f1_scores)
print('Best threshold: ', thresholds[np.argmax(f1_scores)])
print('Best F1-Score: ', np.max(f1_scores))
threshold = thresholds[np.argmax(f1_scores)]
# +
# feature scaling
from sklearn.metrics import roc_curve, roc_auc_score
from matplotlib import pyplot
prob_test = (test_loss-min(test_loss))/(max(test_loss)-min(test_loss))
fpr, tpr, thresholds = roc_curve(labels_test,prob_test)
# plot the roc curve for the model
pyplot.plot([0,1], [0,1], linestyle='--', label='No Skill')
pyplot.plot(fpr, tpr, marker='.', label='Logistic')
# axis labels
pyplot.xlabel('False Positive Rate')
pyplot.ylabel('True Positive Rate')
pyplot.legend()
# show the plot
pyplot.show()
from numpy import sqrt
from numpy import argmax
gmeans = sqrt(tpr * (1-fpr))
# locate the index of the largest g-mean
ix = argmax(gmeans)
print('Best Threshold=%f, G-Mean=%.3f' % (thresholds[ix], gmeans[ix]))
# plot the roc curve for the model
pyplot.plot([0,1], [0,1], linestyle='--', label='No Skill')
pyplot.plot(fpr, tpr, marker='.', label='Logistic')
pyplot.scatter(fpr[ix], tpr[ix], marker='o', color='black', label='Best')
# axis labels
pyplot.xlabel('False Positive Rate')
pyplot.ylabel('True Positive Rate')
pyplot.legend()
# show the plot
pyplot.show()
threshold = thresholds[ix]
# -
threshold
# Number of correct predictions for Normal test data
prob_test = (test_loss-min(test_loss))/(max(test_loss)-min(test_loss))
print(roc_auc_score(labels_test, prob_test, average=None))
pred_test = [0 if prob_test[i] < threshold else 1 for i in range(len(prob_test))]
len(pred_test)
# # original result
from sklearn.metrics import classification_report, roc_auc_score, confusion_matrix
print("Test Evaluation")
print(roc_auc_score(labels_test, pred_test, average=None))
print(classification_report(labels_test ,pred_test, labels=[0, 1]))
print(confusion_matrix(labels_test ,pred_test))
# # improve with port profile
# +
# add filter step
# open clustering profile
df_profile = pd.read_csv("/home/andrewngo/Desktop/MLTracker/server_profile_prop_labels_27.csv")
host_profile = df_profile[["profile_labels", "computer"]]
host_profile = host_profile.set_index('computer').to_dict()['profile_labels']
# create a dict of path and labels
label_dict = dict()
for i in range(len(pred_test)):
label_dict[test_data_path[i]] = i
path_to_index = convert_nodeid_to_user_CUC(label_dict, computer2nodeid, user2nodeid)
index_to_path = {path_to_index[i]:i for i in path_to_index}
# filter the false positive (bruce force)
new_pred = []
for i in range(len(pred_test)):
if pred_test[i] == 1:
try:
if host_profile[index_to_path[i][0]] == host_profile[index_to_path[i][2]]:
new_pred.append(0)
else:
new_pred.append(pred_test[i])
except:
new_pred.append(pred_test[i])
else:
new_pred.append(pred_test[i])
temp = new_pred
new_pred = []
for i in range(len(pred_test)):
if pred_test[i] == 1:
try:
if host_profile[index_to_path[i][0]] == host_profile[index_to_path[i][2]]:
new_pred.append(0)
else:
new_pred.append(pred_test[i])
except:
new_pred.append(pred_test[i])
else:
new_pred.append(pred_test[i])
# label_dict
# -
from sklearn.metrics import classification_report, roc_auc_score, confusion_matrix
print("Test Evaluation")
print(roc_auc_score(labels_test, new_pred, average=None))
print(classification_report(labels_test ,new_pred, labels=[0, 1]))
print(confusion_matrix(labels_test ,new_pred))
# +
# add filter step
# open clustering profile
df_profile = pd.read_csv("/home/andrewngo/Desktop/MLTracker/server_profile_prop_labels_30.csv")
host_profile = df_profile[["profile_labels", "computer"]]
host_profile = host_profile.set_index('computer').to_dict()['profile_labels']
# create a dict of path and labels
label_dict = dict()
for i in range(len(pred_test)):
label_dict[test_data_path[i]] = i
path_to_index = convert_nodeid_to_user_CUC(label_dict, computer2nodeid, user2nodeid)
index_to_path = {path_to_index[i]:i for i in path_to_index}
# filter the false positive (bruce force)
new_pred = []
for i in range(len(pred_test)):
if pred_test[i] == 1:
try:
if host_profile[index_to_path[i][0]] == host_profile[index_to_path[i][2]]:
new_pred.append(0)
else:
new_pred.append(pred_test[i])
except:
new_pred.append(pred_test[i])
else:
new_pred.append(pred_test[i])
temp = new_pred
new_pred = []
for i in range(len(pred_test)):
if pred_test[i] == 1:
try:
if host_profile[index_to_path[i][0]] == host_profile[index_to_path[i][2]]:
new_pred.append(0)
else:
new_pred.append(pred_test[i])
except:
new_pred.append(pred_test[i])
else:
new_pred.append(pred_test[i])
# label_dict
# -
from sklearn.metrics import classification_report, roc_auc_score, confusion_matrix
print("Test Evaluation")
print(roc_auc_score(labels_test, new_pred, average=None))
print(classification_report(labels_test ,new_pred, labels=[0, 1]))
print(confusion_matrix(labels_test ,new_pred))
# # Evaluate
# +
# find the false positive
def false_positive_path(path, labels, pred, prob):
return_path = dict()
for i in range(len(pred)):
if pred[i] == 0 and labels[i] == 1:
return_path[path[i]] = float(prob[i])
return return_path
def true_positive_path(path, labels, pred, prob):
return_path = dict()
for i in range(len(pred)):
if pred[i] == 0 and labels[i] == 0:
return_path[path[i]] = float(prob[i])
return return_path
def false_negative_path(path, labels, pred, prob):
return_path = dict()
for i in range(len(pred)):
if pred[i] == 1 and labels[i] == 0:
return_path[path[i]] = float(prob[i])
return return_path
def true_negative_path(path, labels, pred, prob):
return_path = dict()
for i in range(len(pred)):
if pred[i] == 1 and labels[i] == 1:
return_path[path[i]] = float(prob[i])
return return_path
#convert nodeid to user
def convert_nodeid_to_user_CUC(path_list, computer2nodeid, user2nodeid):
return_path = dict()
nodeid2computer = {computer2nodeid[i]:i for i in computer2nodeid}
nodeid2user = {user2nodeid[i]:i for i in user2nodeid}
for i in path_list:
i_convert = (nodeid2computer[i[0]], nodeid2user[i[1] - model_metapath2vec.start["User"]], nodeid2computer[i[2]])
return_path[i_convert] = path_list[i]
return return_path
# +
false_positive_path_test = false_positive_path(test_data_path, labels_test, pred_test, prob_test)
false_negative_path_test = false_negative_path(test_data_path, labels_test, pred_test, prob_test)
true_positive_path_test = true_positive_path(test_data_path, labels_test, pred_test, prob_test)
true_negative_path_test = true_negative_path(test_data_path, labels_test, pred_test, prob_test)
false_negative_path_test = convert_nodeid_to_user_CUC(false_negative_path_test, computer2nodeid, user2nodeid)
false_positive_path_test = convert_nodeid_to_user_CUC(false_positive_path_test, computer2nodeid, user2nodeid)
true_negative_path_test = convert_nodeid_to_user_CUC(true_negative_path_test, computer2nodeid, user2nodeid)
true_positive_path_test = convert_nodeid_to_user_CUC(true_positive_path_test, computer2nodeid, user2nodeid)
test_result = dict()
test_result["false_negative"] = false_negative_path_test
test_result["false_positive"] = false_positive_path_test
test_result["true_negative"] = true_negative_path_test
test_result["true_positive"] = true_positive_path_test
# -
test_result["false_negative"]
test_result["false_positive"]
test_result["true_negative"]
host_profile["C687"]
# +
# add filter step
# open clustering profile
df_profile = pd.read_csv("/home/andrewngo/Desktop/MLTracker/server_profile_prop_labels_27.csv")
host_profile = df_profile[["profile_labels", "computer"]]
host_profile = host_profile.set_index('computer').to_dict()['profile_labels']
# create a dict of path and labels
label_dict = dict()
for i in range(len(pred_test)):
label_dict[test_data_path[i]] = i
path_to_index = convert_nodeid_to_user_CUC(label_dict, computer2nodeid, user2nodeid)
index_to_path = {path_to_index[i]:i for i in path_to_index}
# filter the false positive (bruce force)
new_pred = []
for i in range(len(pred_test)):
if pred_test[i] == 1:
try:
if host_profile[index_to_path[i][0]] == host_profile[index_to_path[i][2]]:
new_pred.append(0)
else:
new_pred.append(pred_test[i])
except:
new_pred.append(pred_test[i])
else:
new_pred.append(pred_test[i])
temp = new_pred
new_pred = []
for i in range(len(pred_test)):
if pred_test[i] == 1:
try:
if host_profile[index_to_path[i][0]] == host_profile[index_to_path[i][2]]:
new_pred.append(0)
else:
new_pred.append(pred_test[i])
except:
new_pred.append(pred_test[i])
else:
new_pred.append(pred_test[i])
# -
test_result["true_negative"]
# +
test_result_dir = "test_result__outc2" + str(seed) + ".pkl"
with open(test_result_dir, 'wb') as f:
pickle.dump(test_result, f)
# -
# false_positive_path_test = convert_nodeid_to_user_CUC(false_positive_path_test, computer2nodeid, user2nodeid)
false_positive_path_test
threshold
nodeid2computer = {computer2nodeid[i]:i for i in computer2nodeid}
len((nodeid2computer))
# +
normal_train_dict = {out_normal_dict[i]:i for i in out_train_keys_sample}
normal_test_dict = {out_normal_dict[i]:i for i in out_test_keys_sample}
normal_val_dict = {out_normal_dict[i]:i for i in out_val_keys_sample}
mal_test_dict = {out_mal_test_dict[i]:i for i in out_mal_test_dict}
mal_val_dict = {out_mal_train_val_dict[i]:i for i in out_mal_train_val_dict}
data_dict = {}
data_dict = {**normal_train_dict, **normal_test_dict}
data_dict = {**data_dict, **normal_val_dict}
data_dict = {**data_dict, **mal_test_dict}
data_dict = {**data_dict, **mal_val_dict}
train_path = [data_dict[i] for i in normal_train_data]
# out_train_dict_sample = {out_normal_dict[i]:i for i in out_train_keys_sample}
train_path
# -
temp = torch.Tensor(3,2)
temp
for i in temp:
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="5iLAZNQNg3m1" outputId="2e0ef3e6-a0db-4e7f-9ffc-ae9f82596e2d"
# Training the model
history = model.fit(normal_train_data,normal_train_data,
epochs = 50,
batch_size = 128,
validation_data = (train_data_scaled[:,1:], train_data_scaled[:,1:]),
shuffle = True,
callbacks = [early_stopping])
# + id="sJaXZqEpg8GE"
# predictions for normal test data points
encoder_out = model.encoder(normal_val_data).numpy()
decoder_out = model.decoder(encoder_out).numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="zLNcwREih5BC" outputId="83f2ed6f-92c2-4824-806e-2de7151dc934"
encoder_out.shape
# + colab={"base_uri": "https://localhost:8080/"} id="V6m0WV80h6rW" outputId="4f2cc0dd-e582-428e-e33a-3a114bf5eec2"
decoder_out.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="lyGlYWPah7_h" outputId="876c8dd2-1a50-4150-d8da-20dc9c62df3a"
# plotting normal test data point and its predictiction by the autoencoder
plt.plot(normal_test_data[0], 'b')
plt.plot(decoder_out[0], 'r')
# + id="m7ZTj6saiJPr"
# predictions for anomaly test data points
encoder_out_a = model.encoder(mal_test_data).numpy()
decoder_out_a = model.decoder(encoder_out_a).numpy()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="P2UI_yvOi33V" outputId="31551fd0-5eaa-42b1-c74e-67982c0b541b"
# plotting anomaly test data point and its predictiction by the autoencoder
plt.plot(mal_test_data[0], 'b')
plt.plot(decoder_out_a[0], 'r')
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="WKWvXpCFjAKu" outputId="a0c48128-28de-44bb-8fe9-ecc107616a09"
# reconstruction loss for normal test data
reconstructions = model.predict(normal_test_data)
train_loss = tf.keras.losses.mae(reconstructions, normal_test_data)
# Plotting histogram for recontruction loss for normal test data
plt.hist(train_loss, bins = 10)
# + colab={"base_uri": "https://localhost:8080/"} id="AdmXLSBaja5j" outputId="4550b752-0670-4a3c-a4fc-c9908f2ff246"
np.mean(train_loss)
# + colab={"base_uri": "https://localhost:8080/"} id="vtW3CBUjjiIU" outputId="e766c857-2dc6-492d-dd45-275cb5962f1b"
np.std(train_loss)
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="-r38UR6Qjw8M" outputId="ffd1278a-b299-402a-f573-65a9b7f74bc0"
# reconstruction loss for anomaly test data
reconstructions_a = model.predict(mal_test_data)
train_loss_a = tf.keras.losses.mae(reconstructions_a, mal_test_data)
# Plotting histogram for recontruction loss for anomaly test data
plt.hist(train_loss_a, bins = 10)
# + colab={"base_uri": "https://localhost:8080/"} id="K_6QYRT2kpJQ" outputId="fa0ef513-0b2e-4928-8eb0-169b68bd0d2e"
np.mean(train_loss_a)
# + colab={"base_uri": "https://localhost:8080/"} id="XQikvTQuky7-" outputId="3f139441-04ca-4166-bf53-09388142f1c2"
np.std(train_loss_a)
# -
np.mean(train_loss_a)
# + id="dy7SKgIojlJA"
# setting threshold
# threshold = np.mean(train_loss) + 2*np.std(train_loss)
threshold = (np.mean(train_loss) +np.mean(train_loss_a))/2
threshold
# + colab={"base_uri": "https://localhost:8080/"} id="yU2D1OA1ju3-" outputId="7f41e25e-0a15-435c-8141-e742c133a8b1"
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="CA9baHE3j-NT" outputId="4caee125-6bbe-449f-c519-c5ffdf8c862a"
# Plotting the normal and anomaly losses with the threshold
plt.hist(train_loss, bins = 10, label = 'Normal')
plt.hist(train_loss_a, bins = 10, label = 'Anomaly')
plt.axvline(threshold, color='r', linewidth = 3, linestyle = 'dashed', label = '{:0.3f}'.format(threshold))
plt.legend(loc = 'upper right')
plt.show()
# + id="YhkCxKbTk1eh"
# Number of correct predictions for Normal test data
preds = tf.math.less(train_loss, threshold)
preds
# + colab={"base_uri": "https://localhost:8080/"} id="6B9m-mfnk-HH" outputId="fc3cdba3-9d8a-4aae-8b78-3e73d98c74b1"
tf.math.count_nonzero(preds)
# + id="c-zCwgsYlI0v"
# Number of correct predictions for Anomaly test data
preds_a = tf.math.greater(train_loss_a, threshold)
# + colab={"base_uri": "https://localhost:8080/"} id="DH0nTPxrlSTi" outputId="4f4af546-41b4-4bc8-8647-c98ad30c9f06"
tf.math.count_nonzero(preds_a)
# + colab={"base_uri": "https://localhost:8080/"} id="w3lKPX8tlaiW" outputId="1d5da799-7a6d-4a97-f312-3a6f1d68e7ef"
preds_a.shape
# -
| autoencoder_keras/Anomaly_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### 1. Which of the following is NOT considered an expensive operation?
# - [ ] Parsing a file
# - [ ] Downloading data over the network
# - [ ] Going through a list
# - [x] **Using a dictionary**
#
# ### 2. Which of the following may be the most expensive to carry out in most automation tasks in a script?
# - [x] **Loops**
# - [ ] Lists
# - [ ] Vector
# - [ ] Hash
#
# ### 3. Which of the following statements represents the most sound advice when writing scripts?
# - [ ] Aim for every speed advantage you can get in your code
# - [ ] Use expensive operations often
# - [x] **Start by writing clear code, then speed it up only if necessary**
# - [ ] Use loops as often as possible
#
# ### 4. In Python, what is a data structure that stores multiple pieces of data, in order, which can be changed later?
# - [ ] A hash
# - [ ] Dictionaries
# - [x] **Lists**
# - [ ] Tuples
#
# ### 5. What command, keyword, module, or tool can be used to measure the amount of time it takes for an operation or program to execute? (Check all that apply)
# - [x] **time**
# - [x] **kcachegrind**
# - [x] **cProfile**
# - [ ] break
| troubleshooting-debugging-techniques/week-2/quiz-slow-code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false
# # **Tutorial 02: OOP in Python (Part 1)** 👀
# ##### Abstraction and Encapsulation
# + [markdown] deletable=false editable=false
# <a id='t2toc'></a>
# #### Contents: ####
# - **[Initialization](#t2init)**
# - [Recall](#t2recall)
# - [`__init__` method](#t2__init)
# - *[Exercise 1](#t2ex1)*
# - **[Abstraction and Encapsulation](#t2abs_enc)**
# - *[Exercise 2](#t2ex2)*
# - [Access Modifiers](#t2accessmod)
# - *[Exercise 3](#t2ex3)*
# - [Exercises Solutions](#t2sol)
#
# + [markdown] deletable=false editable=false
# 💡 <b>TIP</b><br>
# > <i>In Exercises, when time permits, try to write the codes yourself, and do not copy it from the other cells.</i>
#
# + [markdown] deletable=false editable=false
# <br><br><a id='t2init'></a>
# ## ▙▂ **🄸NITIALIZATION (CONSTRCUCTOR) ▂▂**
# + [markdown] deletable=false editable=false
# Let's understand the meaning of initializtion using a simple example, and a recall from the previous lesson.
# + [markdown] deletable=false editable=false
# <a id='t2recall'></a>
# #### **▇▂ Recall ▂▂**
# In the previous lesson, we learnt how to define a simple class, including some attributes and methods.
# + deletable=false
class eagle:
species = 'bird'
def can():
print('fly')
def describe(self):
print('Eagle is the common name for many large birds of prey of the family Accipitridae.')
print('Eagles belong to several groups of genera, not all of which are closely related.')
print('Most of the 60 species of eagle are from Eurasia and Africa. ')
# + deletable=false
Goldie = eagle()
Goldie.describe()
# + [markdown] deletable=false editable=false
# Eventhough the method `describe()` is known as an instance method, but it is not performing any specific operation on an instance.<br>
# What is the benefit of `describe()` compare to `can()`?<br>
# Is it really useful.<br>
# Why?<br>
#
# It would be a useful method, if it can perform a task for a specific instance. But how?
# + [markdown] deletable=false editable=false
# To do that, we need to make some instance attributes. Then the instance method can perform a task on the specific instance. <br>
# + deletable=false
class eagle:
species = 'bird'
def can():
print('fly')
def describe(self):
print('{} is a {} eagle with {} color, and born on {}.'.
format(self.name, self.gender, self.color, self.birth_year))
e1= eagle()
e1.name = 'Goldie'
e1.color = 'White'
e1.gender = 'Male'
e1.birth_year = 2015
e2= eagle()
e2.name = 'Remo'
e2.color = 'Black'
e2.gender = 'Female'
e2.birth_year = 2018
# + deletable=false
e1.describe()
# + deletable=false
e2.describe()
# + [markdown] deletable=false editable=false
# Wouldn't be more intersting, if we can define the attributes of instance, when we are creating the instance?
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# <a id='t2__init'></a>
# #### **▇▂ `__init__` ▂▂**
# The `__init__` method (is similar to constructors in C++ and Java) is used to initialize the object’s state. The task of constructors is to initialize (assign values) to the data members of the class when an object of class is created. Like methods, a constructor also contains collection of statements (i.e. instructions) that are executed at the time of Object creation. It is run as soon as an object of a class is instantiated. The method is useful to do any initialization you want to do with your object.
# + [markdown] deletable=false editable=false
# In the previous example, we can use `__init__` to initialize the attributes of an instances.
# + deletable=false
class eagle:
species = 'bird'
def can():
print('fly')
# init method or constructor
def __init__(self, e_name, e_color, e_gender, e_birth_year):
self.name = e_name
self.color = e_color
self.gender = e_gender
self.birth_year = e_birth_year
def describe(self):
print('{} is a {} eagle with {} color, and born on {}.'.
format(self.name, self.gender, self.color, self.birth_year))
# + [markdown] deletable=false editable=false
# So, we pass the values as parameter when an instances is created:
# + deletable=false
e1 = eagle('Goldie', 'White', 'Male', 2015)
e2 = eagle('Remo', 'Black', 'Female', 2018)
# + deletable=false
e1.describe()
# + deletable=false
e2.describe()
# + deletable=false
e1.species
# + deletable=false
e1.name
# + [markdown] deletable=false editable=false
# If you like to call `can()` method using an instance, you can add `self` as an argument of the method:
# + deletable=false
class eagle:
species = 'bird'
def can(self):
print('fly')
def __init__(self, e_name, e_color, e_gender, e_birth_year):
self.name = e_name
self.color = e_color
self.gender = e_gender
self.birth_year = e_birth_year
def describe(self):
print('{} is a {} eagle with {} color, and born on {}'.
format(self.name, self.gender, self.color, self.birth_year))
e1= eagle('Goldie', 'White', 'Male', 2015)
e2= eagle('Remo', 'Black', 'Female', 2018)
# + deletable=false
e1.can()
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# <br><br><a id='t2ex1'></a>
# ◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾
# + [markdown] deletable=false editable=false
# **✎ Exercise 𝟙**<br> <br> ▙ ⏰ ~ 3 min. ▟ <br>
# + [markdown] deletable=false editable=false
# ❶ Add a new method `age()` to the class `eagle`, which accept the current year as an argument and print the age of the eagle with an appropriate formatted message.<br>
# + deletable=false
# Exercise 1.1
# + [markdown] deletable=false editable=false
# ❷ Define two new instances and print the age of each instance. <br>
# + deletable=false
# Exercise 1.2
# + [markdown] deletable=false editable=false
# ◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# <br><br><a id='t2abs_enc'></a>
# ## **▙▂ 🄰BSTRACTION AND 🄴NCAPSULATION ▂▂**
# + [markdown] deletable=false editable=false
# We use a simple example to understand the concept of abstraction and encapsulation. We start by using a procedural programming approach, and then move to a basic level of abstraction and encapsulation using OOP approach.
# + [markdown] deletable=false editable=false
# #### Example 1: Bank Account
# + [markdown] deletable=false editable=false
# **A poor solution.** Let's make a global variable to store the balance of the bank account and define functions to deposit to or withdraw from the bank account:
# + deletable=false
balance = 0
def deposit(amount):
global balance
balance += amount
return balance
def withdraw(amount):
global balance
balance -= amount
return balance
# + deletable=false
deposit(10)
print(balance)
# + deletable=false
withdraw(5)
print(balance)
# + [markdown] deletable=false editable=false
# **Add a little bit more.** We can use a separate variable for each person to manage multiple accounts:
# + deletable=false
def make_account():
return {'balance': 0}
def deposit(account, amount):
account['balance'] += amount
return account['balance']
def withdraw(account, amount):
account['balance'] -= amount
return account['balance']
# + deletable=false
John_acc = make_account()
print(John_acc)
# + deletable=false
Mike_acc = make_account()
print(Mike_acc)
# + deletable=false
deposit(John_acc,10)
print(John_acc)
# + deletable=false
withdraw(John_acc,5)
print(John_acc)
# + deletable=false
deposit(Mike_acc,25)
print(Mike_acc)
# + deletable=false
withdraw(Mike_acc,8)
print(Mike_acc)
# + [markdown] deletable=false editable=false
# In the above examples, we did not use OOP approach. There are different **global variables** used to store the account information. There is no abstraction and encapsulation.
# + [markdown] deletable=false editable=false
# **OOP Approach.** Now, let's use OOP approach to define a class for an account as an abstract data type and encapsulate data and methods in an object.
# + deletable=false
class BankAccount:
def __init__(self, acc_owner_name, initial_inves_value):
self.name = acc_owner_name
self.balance = initial_inves_value
def withdraw(self, amount):
self.balance -= amount
return self.balance
def deposit(self, amount):
self.balance += amount
return self.balance
def show_balance(self):
print(self.name, ":", self.balance)
# + deletable=false
John_acc = BankAccount('John', 10)
Mike_acc = BankAccount('Mike', 25)
# + deletable=false
John_acc.deposit(4)
John_acc.show_balance()
# + deletable=false
John_acc.withdraw(5)
John_acc.show_balance()
# + [markdown] deletable=false editable=false
# Do some extra practice by making new objects and calling methods of the class.
# + deletable=false
# + [markdown] deletable=false editable=false
# <br>🔴 Discuss the benefits of this approach.
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# #### Example 2: Card Deck
# + [markdown] deletable=false editable=false
# First, we can define a class for a card:
# + deletable=false
class Card:
def __init__(self, suit, value):
self.suit = suit
self.value = value
# + deletable=false
card1 = Card('Hearts', '8')
# + [markdown] deletable=false editable=false
# A low level of abstraction is implemented to define a card with the required details, as a class. However, we still need to know internal structure of the `card` class in order to represent the card:
# + deletable=false
print(card1.value, card1.suit)
# + [markdown] deletable=false editable=false
# Let's implement more abstraction, by adding an interface to represent the card:
# + deletable=false
class Card:
def __init__(self, suit, value):
self.suit = suit
self.value = value
def represent(self):
return '{} {}'.format(self.value, self.suit)
# + [markdown] deletable=false editable=false
# With this definition, we just need to know the name of the method for the representation of an object:
# + deletable=false
card1 = Card('Hearts', '8')
print(card1.represent())
# + [markdown] deletable=false editable=false
# Now, let's extend it to create a deck of cards:
# + deletable=false
from random import shuffle
class Card:
def __init__(self, suit, value):
self.suit = suit
self.value = value
def represent(self):
return '{} {}'.format(self.value, self.suit)
class Deck:
def __init__(self):
suits = ['♥','♦','♣','♠']
# suits = ['Hearts','Diamonds','Clubs','Spades']
values = ['A','2','3','4','5','6','7','8','9','10','J','Q','K']
self.cards = [Card(suit, value) for suit in suits for value in values]
def deal(self):
if len(self.cards) == 0:
raise ValueError("All cards have been dealt")
return self.cards.pop().represent()
def shuffle(self):
if len(self.cards) < 52:
raise ValueError("Only full decks can be shuffled")
shuffle(self.cards)
return self.represent()
def represent(self):
return "Cards remaining in deck: {}".format(len(self.cards))
# + deletable=false
c1 = Deck()
print(c1.represent())
for _ in range(52):
print(c1.deal())
# + deletable=false
c2 = Deck()
print(c2.represent())
c2.shuffle()
for _ in range(52):
print('your card: ',c2.deal())
print(c2.represent())
# + deletable=false
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# <br><br><a id='t2ex2'></a>
# ◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾
# + [markdown] deletable=false editable=false
# **✎ Exercise 𝟚** <br> <br> ▙ ⏰ 10 min. ▟ <br>
# + [markdown] deletable=false editable=false
# ❶ Define a class for a player. The class should contain the name of player and set of their cards. Add a represent method to show the player's name with the cards in their hand.<br>
# + deletable=false
# Exercise 2.1
# + [markdown] deletable=false editable=false
# ❷ Add another method to the player's class to get a number of cards from the deck.
# + deletable=false
# Exercise 2.2
# + [markdown] deletable=false editable=false
# ❸ In the previous exercise, make sure that the number of requested cards is available on the deck, before dealing cards to the player.
# + deletable=false
# Exercise 2.3
# + [markdown] deletable=false editable=false
# ◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# <a id='t2accessmod'></a>
# #### **▇▂ Access Modifiers in Python ▂▂**
# Various object-oriented languages like C++, Java, Python control access modifications which are used to restrict access to the variables and methods of the class. Most programming languages has three forms of access modifiers, which are Public, Protected and Private in a class.
#
# Python uses ‘_’ symbol to determine the access control for a specific data member or a member function of a class. Access specifiers in Python have an important role to play in securing data from unauthorized access and in preventing it from being exploited.
#
# A Class in Python has three types of access modifiers –
#
# - **Public** Access Modifier
# - **Protected** Access Modifier
# - **Private** Access Modifier
# + [markdown] deletable=false editable=false
# #### Public Access Modifier
# The members of a class that are declared public are easily accessible from any part of the program. All data members and member functions of a class are **public** *by default*.
# + deletable=false
class Student:
# constructor
def __init__(self, name, age):
# public data mambers
self.studentName = name
self.studentAge = age
# public memeber function
def displayAge(self):
# accessing public data member
print("Age: ", self.studentAge)
# creating object of the class
obj = Student("Raymond", 20)
# accessing public data member
print("Name: ", obj.studentName)
# calling public member function of the class
obj.displayAge()
# + [markdown] deletable=false editable=false
# In the above program, `studentName` and `studentAge` are public data members and `displayAge()` method is a public member function of the class `Student`. These data members of the class `Student' can be accessed from anywhere in the program.
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# #### Protected Access Modifier
# The members of a class that are declared protected are only accessible to a class derived from it. Data members of a class are declared protected by adding a single underscore `_` symbol before the data member of that class.
# + [markdown] deletable=false editable=false
# <br>⚠ <b>NOTE</b><br>
# > We did not introduce the concepts of inheritance and derived class, yet. Those will be discussed in details in the next tutorial.<br>
# At the moment, the general information about the inheritanc explained by your teacher is sufficient to understand the concept of the Protected Access Modifier.
# + deletable=false
# super class
class Student:
# protected data members
_name = None
_student_number = None
_study_program = None
# constructor
def __init__(self, name, student_number, study_program):
self._name = name
self._student_number = student_number
self._study_program = study_program
# protected member function
def _displayStudent(self):
# accessing protected data members
print("Student Number: ", self._student_number)
print("Study Program: ", self._study_program)
# derived class
class PeerCoach(Student):
# constructor
def __init__(self, name, student_number, study_program):
Student.__init__(self, name, student_number, study_program)
# public member function
def displayDetails(self):
# accessing protected data members of super class
print("Name: ", self._name)
# accessing protected member functions of super class
self._displayStudent()
# creating objects of the derived class
obj = PeerCoach("Raymond", 1706256, "Computer Science")
# calling public member functions of the class
obj.displayDetails()
# + [markdown] deletable=false editable=false
# In the above program, `_name`, `_student_number` and `_study_program` are protected data members and `_displayStudent()` method is a protected method of the super class Student. The `displayDetails()` method is a public member function of the class `PeerCoach` which is derived from the `Student` class, the `displayDetails()` method in `PeerCoach` class accesses the protected data members of the `Student` class.
# + [markdown] deletable=false editable=false
# <br>⚠ <b>NOTE</b><br>
# > In fact, this does not really prevent instance variables from accessing or modifying the instance. You can still perform the following operations:
# + deletable=false
obj = Student("Elizabeth", 1811123, "Electrical Engineering")
print(obj._student_number)
# + deletable=false
obj._student_number = 1811126
print(obj._student_number)
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# #### Private Access Modifier
# The members of a class that are declared private are accessible within the class only, private access modifier is the most secure access modifier. Data members of a class are declared private by adding a double underscore `__` symbol before the data member of that class.
# + deletable=false
class Student:
# private members
__name = None
__student_number = None
__study_program = None
# constructor
def __init__(self, name, student_number, study_program):
self.__name = name
self.__student_number = student_number
self.__study_program = study_program
# private member function
def __displayDetails(self):
# accessing private data members
print("Name: ", self.__name)
print("Student Number: ", self.__student_number)
print("Study Program: ", self.__study_program)
# public member function
def accessPrivateFunction(self):
# accesing private member function
self.__displayDetails()
# creating object
obj = Student("Raymond", 1706256, "Computer Science")
# calling public member function of the class
obj.accessPrivateFunction()
# + [markdown] deletable=false editable=false
# In the above program, `__name`,` __student_number` and `__study_program` are private members, `__displayDetails()` method is a private member function (these can only be accessed within the class) and `accessPrivateFunction()` method is a public member function of the class `Student` which can be accessed from anywhere within the program. The `accessPrivateFunction()` method accesses the private members of the class `Student`.
# + [markdown] deletable=false editable=false
# 🔴 What happens if you try to directly access the private members or methods?
# + deletable=false
obj.__displayDetails()
# + deletable=false
print(obj.__student_number)
# + [markdown] deletable=false editable=false
# <br>⚠ <b>NOTE</b><br>
# > Python does not have any mechanism that effectively restricts access to any instance variable or method. <br>
# **We can say that Python prescribes a convention of prefixing the name of the variable/method with a single or double underscore to emulate the behavior of protected and private access specifiers.**<br>
# It performs name mangling of private variables. Every member with a double underscore will be changed to `object._class__variable`. So, it can still be accessed from outside the class, but the practice should be refrained.
# + deletable=false
print(obj._Student__student_number)
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# Below is a program to illustrate the use of all the above three access modifiers (public, protected and private) of a class in Python:
#
#
# + deletable=false
# super class
class Super:
# public data member
var1 = None
# protected data member
_var2 = None
# private data member
__var3 = None
# constructor
def __init__(self, var1, var2, var3):
self.var1 = var1
self._var2 = var2
self.__var3 = var3
# public member function
def displayPublicMembers(self):
# accessing public data members
print("Public Data Member: ", self.var1)
# protected member function
def _displayProtectedMembers(self):
# accessing protected data members
print("Protected Data Member: ", self._var2)
# private member function
def __displayPrivateMembers(self):
# accessing private data members
print("Private Data Member: ", self.__var3)
# public member function
def accessPrivateMembers(self):
# accessing private memeber function
self.__displayPrivateMembers()
# derived class
class Sub(Super):
# constructor
def __init__(self, var1, var2, var3):
Super.__init__(self, var1, var2, var3)
# public member function
def accessProtectedMemebers(self):
# accessing protected member functions of super class
self._displayProtectedMembers()
# creating objects of the derived class
obj = Sub("Oliver", 4455667, "Microbiology")
# calling public member functions of the class
obj.displayPublicMembers()
obj.accessProtectedMemebers()
obj.accessPrivateMembers()
# + deletable=false
# Object can access protected member
print("Object is accessing protected member:", obj._var2)
# + deletable=false
# object can not access private member, so it will generate Attribute error
print(obj.__var3)
# + [markdown] deletable=false editable=false
# <br><br><a id='t2ex3'></a>
# ◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾
# + [markdown] deletable=false editable=false
# **✎ Exercise 𝟛** <br> <br> ▙ ⏰ 1 min. ▟ <br>
# + [markdown] deletable=false editable=false
# ❶ Write a piece of code to get direct access to `__var3` .<br>
# + deletable=false
# Exercise 3.1
# + [markdown] deletable=false editable=false
# ❷ Write a piece of code to get direct access to `__displayPrivateMembers()` method.<br>
# + deletable=false
# Exercise 3.2
# + [markdown] deletable=false editable=false
# ◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾◾
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# <br><br><a id='t2sol'></a>
# ◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼<br>
# ◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼
# + [markdown] deletable=false editable=false
# #### 🔑 **Exercises Solutions** ####
# + [markdown] deletable=false editable=false
# **Exercise 1.1:**
# + deletable=false
class eagle:
species = 'bird'
def can(self):
print('fly')
# init method or constructor
def __init__(self, e_name, e_color, e_gender, e_birth_year):
self.name = e_name
self.color = e_color
self.gender = e_gender
self.birth_year = e_birth_year
def describe(self):
print('{} is a {} eagle with {} color, and born on {}.'.
format(self.name, self.gender, self.color, self.birth_year))
def age(self, current_year):
print('{} is {} years old.'.format(self.name, current_year - self.birth_year))
# + [markdown] deletable=false editable=false
# <br>[back to the Exercise 1 ↥](#t2ex1)
# + [markdown] deletable=false editable=false
# **Exercise 1.2:**
# + deletable=false
e1 = eagle('Cleo', 'Grey', 'Male', 2014)
e2 = eagle('Ava', 'Gold', 'Female', 2019)
e1.age(2021)
e2.age(2021)
# + [markdown] deletable=false editable=false
# <br>[back to the Exercise 1 ↥](#t2ex1)
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# **Exercise 2.1:**
# + deletable=false
class Player:
def __init__(self, name):
self.name = name
self.cards = []
def represent(self):
return "Cards in {}'s hand: {}".format(self.name, self.cards)
# + deletable=false
p1 = Player('Ronald')
# + [markdown] deletable=false editable=false
# **Exercise 2.2:**
# + deletable=false
class Player:
def __init__(self, name):
self.name = name
self.cards = []
def get_cards(self, deck, number_of_cards):
self.cards = [deck.deal() for i in range(4)]
def represent(self):
return "Cards in {}'s hand: {}".format(self.name, self.cards)
# + deletable=false
from random import shuffle
class Card:
def __init__(self, suit, value):
self.suit = suit
self.value = value
def represent(self):
return '{} {}'.format(self.value, self.suit)
class Deck:
def __init__(self):
suits = ['♥','♦','♣','♠']
# suits = ['Hearts','Diamonds','Clubs','Spades']
values = ['A','2','3','4','5','6','7','8','9','10','J','Q','K']
self.cards = [Card(suit, value) for suit in suits for value in values]
def deal(self):
if len(self.cards) == 0:
raise ValueError("All cards have been dealt")
return self.cards.pop().represent()
def shuffle(self):
if len(self.cards) < 52:
raise ValueError("Only full decks can be shuffled")
shuffle(self.cards)
return self.represent()
def represent(self):
return "Cards remaining in deck: {}".format(len(self.cards))
# + deletable=false
c1 = Deck()
c1.shuffle()
p1 = Player('Ronald')
p1.get_cards(c1,4)
print(p1.represent())
# + [markdown] deletable=false editable=false
# <br>[back to the Exercise 2 ↥](#t2ex2)
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# **Exercise 3.1:**
# + deletable=false
print(obj._Super__var3)
# + [markdown] deletable=false editable=false
# **Exercise 3.2:**
# + deletable=false
obj._Super__displayPrivateMembers()
# + [markdown] deletable=false editable=false
# <br>[back to the Exercise 3 ↥](#t2ex3)
# + [markdown] deletable=false editable=false
# <br>[back to top ↥](#t2toc)
# + [markdown] deletable=false editable=false
# ◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼<br>
# ◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼◼
| ipynb/T02-OOP-in-Python-Part-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:41.313099", "end_time": "2020-10-29T03:00:41.336098", "duration": 0.022999, "status": "completed"} tags=[]
# # Text Preprocessing
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:41.360099", "end_time": "2020-10-29T03:00:45.234098", "duration": 3.873999, "status": "completed"} tags=[]
import pandas as pd
import itertools
from data_describe.text.text_preprocessing import *
from data_describe.misc.load_data import load_data
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:45.257098", "end_time": "2020-10-29T03:00:45.280098", "duration": 0.023, "status": "completed"} tags=[]
# ## Load Data
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:45.302099", "end_time": "2020-10-29T03:00:45.856143", "duration": 0.554044, "status": "completed"} tags=[]
from sklearn.datasets import fetch_20newsgroups
categories = ['alt.atheism']
newsgroups = fetch_20newsgroups(subset='train', categories=categories)['data']
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:45.877168", "end_time": "2020-10-29T03:00:45.913168", "duration": 0.036, "status": "completed"} tags=[]
newsgroups[0][:100]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:45.936173", "end_time": "2020-10-29T03:00:45.958141", "duration": 0.021968, "status": "completed"} tags=[]
# ## Tokenize
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:45.981143", "end_time": "2020-10-29T03:00:46.625174", "duration": 0.644031, "status": "completed"} tags=[]
newsgroups_tokens = tokenize(newsgroups)
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:46.648142", "end_time": "2020-10-29T03:00:48.956979", "duration": 2.308837, "status": "completed"} tags=[]
to_list(newsgroups_tokens)[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:48.980978", "end_time": "2020-10-29T03:00:49.004988", "duration": 0.02401, "status": "completed"} tags=[]
# ## Change to all lowercase
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:49.027988", "end_time": "2020-10-29T03:00:51.417571", "duration": 2.389583, "status": "completed"} tags=[]
newsgroups_tokens = tokenize(newsgroups)
newsgroups_lower = to_lower(newsgroups_tokens)
to_list(newsgroups_lower)[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:51.445570", "end_time": "2020-10-29T03:00:51.470567", "duration": 0.024997, "status": "completed"} tags=[]
# ## Run a preprocessing pipeline in one line
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:51.493569", "end_time": "2020-10-29T03:00:53.864232", "duration": 2.370663, "status": "completed"} tags=[]
to_list(preprocess_texts(newsgroups, custom_pipeline=['tokenize', 'to_lower']))[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:53.888196", "end_time": "2020-10-29T03:00:53.912227", "duration": 0.024031, "status": "completed"} tags=[]
# ## Remove punctuation
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:53.935226", "end_time": "2020-10-29T03:00:57.506240", "duration": 3.571014, "status": "completed"} tags=[]
to_list(preprocess_texts(newsgroups, custom_pipeline=['tokenize', 'remove_punct']))[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:57.531266", "end_time": "2020-10-29T03:00:57.556239", "duration": 0.024973, "status": "completed"} tags=[]
# ## Remove digits
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:57.580240", "end_time": "2020-10-29T03:00:57.615271", "duration": 0.035031, "status": "completed"} tags=[]
digits_test_list = [['this', 'is', '3', 'a', 'test', '2c', 'if', 'it', 'works']]
to_list(preprocess_texts(digits_test_list, custom_pipeline=['remove_digits']))[0]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:57.640276", "end_time": "2020-10-29T03:00:57.667245", "duration": 0.026969, "status": "completed"} tags=[]
# ## Remove single characters and spaces
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:57.697293", "end_time": "2020-10-29T03:00:57.733268", "duration": 0.035975, "status": "completed"} tags=[]
single_char_spaces_test_list = [['this', 'is', ' ', 'a', 'test', ' ', 'b']]
to_list(preprocess_texts(single_char_spaces_test_list, custom_pipeline=['remove_single_char_and_spaces']))[0]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:00:57.758273", "end_time": "2020-10-29T03:00:57.785274", "duration": 0.027001, "status": "completed"} tags=[]
# ## Remove stopwords
# + papermill={"exception": false, "start_time": "2020-10-29T03:00:57.810275", "end_time": "2020-10-29T03:01:01.483073", "duration": 3.672798, "status": "completed"} tags=[]
to_list(preprocess_texts(newsgroups, custom_pipeline=['tokenize', 'remove_punct', 'remove_stopwords']))[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:01.509072", "end_time": "2020-10-29T03:01:01.536075", "duration": 0.027003, "status": "completed"} tags=[]
# ## Stem words
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:01.565073", "end_time": "2020-10-29T03:01:08.164895", "duration": 6.599822, "status": "completed"} tags=[]
to_list(preprocess_texts(newsgroups, custom_pipeline=['tokenize', 'remove_punct', 'remove_stopwords', 'stem']))[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:08.195870", "end_time": "2020-10-29T03:01:08.221866", "duration": 0.025996, "status": "completed"} tags=[]
# ## Lemmatize words
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:08.246894", "end_time": "2020-10-29T03:01:14.840998", "duration": 6.594104, "status": "completed"} tags=[]
to_list(preprocess_texts(newsgroups, custom_pipeline=['tokenize', 'remove_punct', 'remove_stopwords', 'lemmatize']))[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:14.867980", "end_time": "2020-10-29T03:01:14.894969", "duration": 0.026989, "status": "completed"} tags=[]
# ## Custom Function
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:14.919970", "end_time": "2020-10-29T03:01:17.440015", "duration": 2.520045, "status": "completed"} tags=[]
def shout(text_docs_bow):
return ((word.upper() for word in doc) for doc in text_docs_bow)
to_list(preprocess_texts(newsgroups, custom_pipeline=['tokenize', shout]))[0][:10]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:17.476015", "end_time": "2020-10-29T03:01:17.508015", "duration": 0.032, "status": "completed"} tags=[]
# ## Convert back to a single string
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:17.540015", "end_time": "2020-10-29T03:01:22.116117", "duration": 4.576102, "status": "completed"} tags=[]
to_list(preprocess_texts(newsgroups, custom_pipeline=[
'tokenize',
'remove_punct',
'remove_stopwords',
'lemmatize',
'remove_digits',
'bag_of_words_to_docs'
]))[0][:1000]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:22.144092", "end_time": "2020-10-29T03:01:22.173127", "duration": 0.029035, "status": "completed"} tags=[]
# ## Create a document-word frequency matrix
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:22.199126", "end_time": "2020-10-29T03:01:26.927102", "duration": 4.727976, "status": "completed"} tags=[]
newsgroups_docs = preprocess_texts(newsgroups, custom_pipeline=[
'tokenize',
'remove_punct',
'remove_stopwords',
'lemmatize',
'remove_digits',
'bag_of_words_to_docs'
])
create_doc_term_matrix(newsgroups_docs).iloc[:5, 10:]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:26.955074", "end_time": "2020-10-29T03:01:26.983102", "duration": 0.028028, "status": "completed"} tags=[]
# ## Create a TF-IDF matrix
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:27.011104", "end_time": "2020-10-29T03:01:32.965454", "duration": 5.95435, "status": "completed"} tags=[]
newsgroups_docs = preprocess_texts(newsgroups, custom_pipeline=[
'tokenize',
'remove_punct',
'remove_stopwords',
'lemmatize',
'remove_digits',
'bag_of_words_to_docs'
])
create_tfidf_matrix(newsgroups_docs).iloc[:5, 10:]
# + [markdown] papermill={"exception": false, "start_time": "2020-10-29T03:01:32.994422", "end_time": "2020-10-29T03:01:33.023451", "duration": 0.029029, "status": "completed"} tags=[]
# ## Ngrams Frequency
# + papermill={"exception": false, "start_time": "2020-10-29T03:01:33.050448", "end_time": "2020-10-29T03:01:37.305524", "duration": 4.255076, "status": "completed"} tags=[]
newsgroups_ngrams = preprocess_texts(newsgroups, custom_pipeline=[
'tokenize',
'remove_punct',
'remove_digits',
'remove_stopwords',
'ngram_freq'
])
newsgroups_ngrams
| examples/Text_Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Prepare train_data and test_data
import pandas as pd
from sklearn import datasets
from sklearn.model_selection import train_test_split
from hypergbm import make_experiment
from hypernets.tabular.metrics import metric_to_scoring
from sklearn.metrics import get_scorer
train_data = pd.read_csv('datasets/Magic/train.csv.gz')
test_data = pd.read_csv('datasets/Magic/test.csv.gz')
X_train = train_data.copy()
y_train = X_train.pop('Class')
X_test = test_data.copy()
y_test = X_test.pop('Class')
# # Without pseudo_labeling
# _best_reward is 0.6666
experiment = make_experiment(train_data.copy(),test_data=test_data.copy(),target='Class',
random_state=8888,cv=True,early_stopping_rounds=0,
max_trials=10)
estimator = experiment.run()
scorer = get_scorer(metric_to_scoring('accuracy'))
score = scorer(estimator, X_test, y_test)
score
# # Use pseudo_labeling
# Pseudo labeling is a semi-supervised learning technique, instead of manually labeling the unlabelled data, we give approximate labels on the basis of the labelled data. Pseudo-labeling can sometimes improve the generalization capabilities of the model.
#
# _best_reward is 0.7812
experiment = make_experiment(train_data.copy(),test_data=test_data.copy(), target='Class',
random_state=8888,max_trials=10,cv=True,early_stopping_rounds=0,
pseudo_labeling=True,
)
estimator = experiment.run()
scorer = get_scorer(metric_to_scoring('accuracy'))
score = scorer(estimator, X_test, y_test)
score
# # Set more params of pseudo_labeling
# _best_reward is 0.7789
experiment = make_experiment(train_data.copy(),test_data=test_data.copy(), target='Class',
random_state=8888,max_trials=10,cv=True,early_stopping_rounds=0,
pseudo_labeling=True,
pseudo_labeling_proba_threshold=0.5,
pseudo_labeling_proba_quantile=0.9,
pseudo_labeling_sample_number=0.9,
)
estimator = experiment.run()
scorer = get_scorer(metric_to_scoring('accuracy'))
score = scorer(estimator, X_test, y_test)
score
| hypergbm/examples/18.pseudo_labeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# language: python
# name: python37464bitd04ad80605dc4165a042c77f86d6bacf
# ---
from numpy import vstack
from pandas import read_csv
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import accuracy_score
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torch.utils.data import random_split
from torch import Tensor
from torch.nn import Linear
from torch.nn import ReLU
from torch.nn import Sigmoid
from torch.nn import Module
from torch.optim import SGD
from torch.nn import BCELoss
from torch.nn.init import kaiming_uniform_
from torch.nn.init import xavier_uniform_
from cortx_jupyter import read_data, write_data,write_model
# # Load Data from Cortx
class CSVDataset(Dataset):
def __init__(self, path):
# Loading data from cortx
data = read_data(path)
df = read_csv(data, header=None)
self.X = df.values[:, :-1]
self.y = df.values[:, -1]
self.X = self.X.astype('float32')
self.y = LabelEncoder().fit_transform(self.y)
self.y = self.y.astype('float32')
self.y = self.y.reshape((len(self.y), 1))
def __len__(self):
return len(self.X)
def __getitem__(self, idx):
return [self.X[idx], self.y[idx]]
def get_splits(self, n_test=0.33):
test_size = round(n_test * len(self.X))
train_size = len(self.X) - test_size
return random_split(self, [train_size, test_size])
def prepare_data(path):
dataset = CSVDataset(path)
train, test = dataset.get_splits()
train_dl = DataLoader(train, batch_size=32, shuffle=True)
test_dl = DataLoader(test, batch_size=1024, shuffle=False)
return train_dl, test_dl
# # Defining the Model
class MLP(Module):
def __init__(self, n_inputs):
super(MLP, self).__init__()
self.hidden1 = Linear(n_inputs, 10)
kaiming_uniform_(self.hidden1.weight, nonlinearity='relu')
self.act1 = ReLU()
self.hidden2 = Linear(10, 8)
kaiming_uniform_(self.hidden2.weight, nonlinearity='relu')
self.act2 = ReLU()
self.hidden3 = Linear(8, 1)
xavier_uniform_(self.hidden3.weight)
self.act3 = Sigmoid()
def forward(self, X):
X = self.hidden1(X)
X = self.act1(X)
X = self.hidden2(X)
X = self.act2(X)
X = self.hidden3(X)
X = self.act3(X)
return X
# # Model Training
def train_model(train_dl, model):
criterion = BCELoss()
optimizer = SGD(model.parameters(), lr=0.01, momentum=0.9)
for epoch in range(100):
for i, (inputs, targets) in enumerate(train_dl):
optimizer.zero_grad()
yhat = model(inputs)
loss = criterion(yhat, targets)
loss.backward()
optimizer.step()
# # Model Evaluation
# +
#https://machinelearningmastery.com/pytorch-tutorial-develop-deep-learning-models/
def evaluate_model(test_dl, model):
predictions, actuals = list(), list()
for i, (inputs, targets) in enumerate(test_dl):
yhat = model(inputs)
yhat = yhat.detach().numpy()
actual = targets.numpy()
actual = actual.reshape((len(actual), 1))
yhat = yhat.round()
predictions.append(yhat)
actuals.append(actual)
predictions, actuals = vstack(predictions), vstack(actuals)
acc = accuracy_score(actuals, predictions)
return acc
# -
# # Model Prediction
# +
def predict(row, model):
row = Tensor([row])
yhat = model(row)
yhat = yhat.detach().numpy()
return yhat
path = 'ionosphere.csv'
train_dl, test_dl = prepare_data(path)
print(len(train_dl.dataset), len(test_dl.dataset))
model = MLP(34)
train_model(train_dl, model)
acc = evaluate_model(test_dl, model)
print('Accuracy: %.3f' % acc)
row = [1,0,0.99539,-0.05889,0.85243,0.02306,0.83398,-0.37708,1,0.03760,0.85243,-0.17755,0.59755,-0.44945,0.60536,-0.38223,0.84356,-0.38542,0.58212,-0.32192,0.56971,-0.29674,0.36946,-0.47357,0.56811,-0.51171,0.41078,-0.46168,0.21266,-0.34090,0.42267,-0.54487,0.18641,-0.45300]
yhat = predict(row, model)
print('Predicted: %.3f (class=%d)' % (yhat, yhat.round()))
# -
| doc/integrations/cortx_jupyter_integration/Examples/Pytorch-Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
# Mimi Meeting 4/26/2021
# +
@defcomp Foo
p1 = Parameter()
p2 = Parameter(default = 1)
p3 = Parameter()
p4 = Parameter(default = 2)
end
@defcomp Bar
p1 = Parameter()
p2 = Parameter()
p5 = Parameter(default = 5)
p6 = Parameter(default = 6)
end
# -
# Simple Cases:
# +
m = Model()
add_comp!(m, Foo)
# above the add_comp! call calls create_nonshared_param! for each parameter, and for the parameters with
# default values sets a value, otherwise it sets to a sentinal value or type for a missing parameter value
update_param!(m, :Foo, :p1, 5) # updates nonshared param Foo.p1 to 5
set_param!(m, :p2, 10) # now we create a new shared model parameter called p2
update_param!(m, :Foo, :p2, 7) # Errors with a message that Foo.p2 is connected to a shared model
# parameter, and you can't use the comp.param method of update_param!
# in that case
# -
# Old Way to Handle Parameters
# +
m = Model()
add_comp!(m, Foo)
# at this point there are no shared model parameters, everything is unconnected
set_param!(m, :Foo, :p1, 5) # now there is a shared model parameter with the name :p1 connected to Foo
set_param!(m, :Bar, :p1, 5) # errors because we already have a :p1 model parameter
update_param!(m, :p1, 5)
set_param!(m, :p2, 8) # now there is a shared model parameter with the name :p2 connected to Foo and Bar
update_param!(m, :p2, 5)
# defaults handled at runtime
# -
# Old Way to Handle Parameters
# +
m = Model()
add_comp!(m, Foo)
# at this point there are nonshared model parameters for each component/parameter pair, and the ones with
# defaults have values while the others have sentinal NaN or missing types
update_param!(m, :Foo, :p1, 5)
update_param!(m, :p1, 5) # errors because there is no shared :p1
create_shared_param!(m, :p2_shared, 5) # create's a shared parameter :p2
connect_param!(m, :Foo, :p2, :p2_shared) # connects Foo's :p2 to m's :p2_shared
connect_param!(m, :Bar, :p2, :p2_shared) # connects Bar's :p2 to m's :p2_shared
# -
| docs/src/internals/Mimi Meeting_5_26_2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# This notebook is intented for the users that experience difficulties executing the voila dashboard contained in UI.ipynb. It provides the same fonctionalities that you can run and use by clicking on :
# ```
# Kernel -> Restart & Run All
# ```
# %run 'file_ui.ipynb'
# %run 'bands_ui.ipynb'
# %run 'mspa_ui.ipynb'
# %run 'about_ui.ipynb'
fi_tile
bi_tile
mp_widget
mp_results
ma_about
ma_disclaimer
| no_ui.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
from collections import Counter
from datetime import datetime
with open('telegramHistory.json') as json_file:
data = json.load(json_file)
valid_symbols = " ?!@#$%^&*()><_-+=абвгдаеёжзийклмнопрстуфхцчшщъыьэюяabcdefghijklmnopqrstuvwxyz0123456789.,:;/n"
def checkSymbs(text):
global badSymbols
for c in text:
if c not in valid_symbols:
badSymbols += c
return False
return True
# +
myMessages = []
lastDate = None
for chat in data['chats']['list']:
if ('name' in chat and 'messages' in chat and len(chat['messages']) > 1000):
#print(chat['name'] + " " + str(len(chat['messages'])))
for msg in chat['messages']:
if ('type' in msg and msg['type'] == 'message'):
if ('text' in msg and len(msg['text']) >= 2):
date = datetime.strptime(msg['date'], "%Y-%m-%dT%H:%M:%S").timestamp()
if lastDate != None and (date - lastDate > 600):
myMessages.append("===")
lastDate = date;
prefix = '> '
if ('from_id' in msg and msg['from_id'] == 47173181):
prefix = '< ';
text = str(msg['text'])
date
if not text.startswith('[') and not """://""" in text:
t = str(msg['text']).lower()
t = t.replace('\n', ' ').replace('\r', ' ').replace('\t', ' ').replace('\"', '').replace('\'', '')
t = t.replace(' ', ' ')
if checkSymbs(t):
myMessages.append(prefix + t)
print(len(myMessages))
# -
print(len(myMessages))
print(myMessages[:20])
with open('myMessages.txt', mode='wt', encoding='utf-8') as myfile:
myfile.write('\n'.join(str(line) for line in myMessages))
| ConvertChatHistory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lady Tasting Tea - An Investigation
#
# About the Lady Tasting tea problem goes here.
# ## Number of ways of selecting cups
# In this section I will investigate the number of ways in which the Lady can arrange the cups into two sets of four- the cups with milk first and the cups with tea first.
import numpy as np
import matplotlib.pyplot as plt
cups = list(range(8))
cups
#Using Combinations the number of arrangements is 8x7x6x5x4x3x2 or 8!, as order doesn't matter
#Assuming the lady only has to pick 4 cups,
8*7*6*5
#But order does matter in this question. So we need to divide by 4x3x2
1680/(4*3*2)
# If the subject is picking cups randomly, they can't tell which cup has the milk in first. Then they are randomly picking one of 70 different outcomes. They have a probability of 1/70 of been correct.
#google python3 combinations for more tools
import itertools
pos = list(itertools.combinations(cups,4))
pos
#We can also use itertools to verify the number of possible outcomes
len(list(itertools.combinations(cups,4)))
# ## My Hypothesis
# **H0:** My hypothesis is that the Lady cannot tell if the tea was made using milk first or last.
#
# **H1:** The person can tell
1/70
# If the null hypothesis is true then the chance that they guess correctly is 1.4%. So if they guess correctly I will accept the alternative hypothesis.
# ## Distribution
(1, 2, 3, 4, 4)
#This is a list(tuples?) with repeated elements which is fine, as each element identified by a position
{(1, 2, 3, 4, 4)}
#This is a set, see the curley brackets.
{1, 2, 3, 4, 4} & {2, 5, 6, 7}
#This will give me the intersection of the two sets
# #### Back to my investigation
import random
milkfirst = set(random.choice(pos))
milkfirst
# So I have randomly chosen a correct answer that can now be used in my investigation.
counts = [len(milkfirst & set(i)) for i in itertools.combinations(cups, 4)]
counts
import seaborn as sns
sns.countplot(counts)
# ## References
#
# - https://github.com/ianmcloughlin/jupyter-teaching-notebooks/blob/master/lady-tasting-tea.ipynb
#
| lecture_notes/4lady-tasting-tea.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# http://nflsavant.com/pbp_data.php?year=2019
# http://nflsavant.com/pbp_data.php?year=2018
# http://nflsavant.com/pbp_data.php?year=2017
# http://nflsavant.com/pbp_data.php?year=2016
# http://nflsavant.com/pbp_data.php?year=2015
# http://nflsavant.com/pbp_data.php?year=2014
# http://nflsavant.com/pbp_data.php?year=2013
#
# https://sportsdata.io/developers/data-dictionary/nfl
#
# https://www.nfl.com/stats/player-stats/
| notebooks/data_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SparkContext - number of workers and lazy evaluation
# ## Checking the impact of number of workers
# While initializing the `SparkContext`, we can specify number of worker nodes. Generally, it is recommended to have one worker per core of the machine. But it can be smaller or larger. In the following code, we will examine the impact of number of worker cores on some parallelized operation.
from time import time
from pyspark import SparkContext
for j in range(1,5):
sc= SparkContext(master = "local[%d]"%(j))
t0=time()
for i in range(10):
sc.parallelize([1,2]*10000).reduce(lambda x,y:x+y)
print(f"{j} executors, time = {time()-t0}")
sc.stop()
# #### We observe that it takes almost double time for 1 worker, and after that time reduces to a flat level for 2,3,4 workers etc. This is because this code run on a Linux virtual box using only 2 cores from the host machine. If you run this code on a machine with 4 cores, you will see benefit upto 4 cores and then the flattening out of the time taken. It also become clear that using more than one worker per core is not beneficial as it just does context-switching in that case and does not speed up the parallel computation.
# ## Showing the essence of _lazy_ evaluation
# 
sc = SparkContext(master="local[2]")
# ### Make a RDD with 1 million elements
# %%time
rdd1 = sc.parallelize(range(1000000))
# ### Some computing function - `taketime`
from math import cos
def taketime(x):
[cos(j) for j in range(100)]
return cos(x)
# ### Check how much time is taken by `taketime` function
# %%time
taketime(2)
# ### Now do the `map` operation on the function
# %%time
interim = rdd1.map(lambda x: taketime(x))
# #### How come each taketime function takes 45.8 us but the map operation with a 10000 element RDD also took similar time?<br><br>Because of _lazy_ evaluation i.e. nothing was computed in the previous step, just a plan of execution was made. The variable `interim` does not point to a data structure, instead it points to a plan of execution, expressed as a dependency graph. The dependency graph defines how RDDs are computed from each other.
# ### Let's see the "Dependency Graph" using `toDebugString` method
print(interim.toDebugString().decode())
# 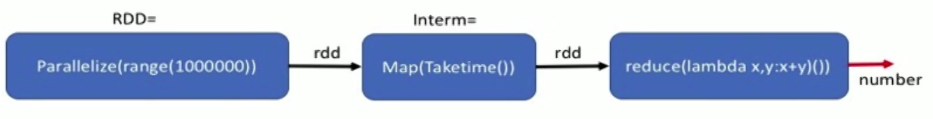
# ### The actual execution by `reduce` method
# %%time
print('output =',interim.reduce(lambda x,y:x+y))
1000000*31e-6
# #### It is less than what we would have expected considering 1 million operations with the `taketime` function. This is the result of parallel operation of 2 cores.
# ### Now, we have not saved (materialized) any intermediate results in `interim`, so another simple operation (e.g. counting elements > 0) will take almost same time
# %%time
print(interim.filter(lambda x:x>0).count())
# ## Caching to reduce computation time on similar operation (spending memory)
# ### Run the same computation as before with `cache` method to tell the dependency graph to plan for caching
# %%time
interim = rdd1.map(lambda x: taketime(x)).cache()
print(interim.toDebugString().decode())
# %%time
print('output =',interim.reduce(lambda x,y:x+y))
# ### Now run the same `filter` method with the help of cached result
# %%time
print(interim.filter(lambda x:x>0).count())
# #### This time it took much shorter time due to cached result, which it could use to compare to 0 and count easily.
| SparkContext_Workers_Lazy_Evaluations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import geopandas as gpd
import cobra.postgres.interface as pgi
import cobra.utils.filehandling as fh
import cobra.utils.jobhandling as jh
import geopandas as gpd
import cobra.utils.geodatahandling as gdh
#download_file = 'https://download.geofabrik.de/europe/switzerland-latest.osm.pbf'
#country = 'swiss'
#download_file = 'https://download.geofabrik.de/europe/great-britain-latest.osm.pbf'
#country = 'gb'
#download_file = 'https://download.geofabrik.de/europe/portugal-latest.osm.pbf'
country = 'portugal'
#download_file = 'https://download.geofabrik.de/europe/france-latest.osm.pbf'
#country = 'france'
#download_file = 'https://download.geofabrik.de/europe/germany-latest.osm.pbf'
#country = 'Germany'
fileman = fh.Filemanager()
jobman = jh.Jobmanager()
# +
#jobman.delete_jobs()
# -
fileman.download_plain_file(download_file, country)
datasets = fileman.get_data(datatype='OSM PBF')
dataset = datasets[datasets['Dataset'] == country].iloc[0]
dataset
jobman.create_import_job_from_dataset(dataset, f'{country}power', style='power.style')
jobman.get_jobs(df=True)
sql = f"SELECT osm.way AS geom, osm.* FROM {country}power.planet_osm_line osm WHERE power IS NOT NULL"
pg_interface = pgi.PgInterface()
conn = pg_interface.get_connection()
powerlines = gpd.GeoDataFrame.from_postgis(sql, conn)
powerlines.plot(figsize=(16,16))
sql = f"SELECT osm.way AS geom, osm.* FROM {country}power.planet_osm_line osm WHERE power IS NOT NULL AND (power = 'line')"
jobman.create_new_pg_to_x(sql=sql, format='GPKG', filename=f'{country}_powerline.gpkg')
jobman.get_jobs(df=True)
sql = f"SELECT osm.way AS geom, osm.* FROM {country}power.planet_osm_point osm WHERE power IS NOT NULL"
i = pgi.PgInterface()
gdf_powerpoint = i.get_gdf(sql)
gdf_powerpoint.plot(figsize=(16, 16))
sql = f"SELECT osm.way AS geom, osm.* FROM {country}power.planet_osm_point osm WHERE power IS NOT NULL"
jobman.create_new_pg_to_x(sql=sql, format='GPKG', filename=f'{country}_powerpoint.gpkg')
jobman.get_jobs(df=True)
| data/jupyter/Samples/CountryGrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import time
import json
import pandas as pd
import re
driver = webdriver.Chrome('C:/Users/user/Downloads/chromedriver.exe')
driver.get("http://www.megastudy.net/teacher_v2/teacher_main.asp")
def crawling():
# 과목 수
for sub_num in range(6):
# 과목 선택
subject = driver.find_elements_by_xpath("//*[@id=\"container\"]/div/div[2]/ul/li/a")[sub_num].text
subject_btn = driver.find_elements_by_xpath("//*[@id=\"container\"]/div/div[2]/ul/li/a")[sub_num]
subject_btn.send_keys(Keys.ENTER)
time.sleep(2)
teacher_cnt = range(0, len(driver.find_elements_by_xpath("//*[@id=\"tchContArea\"]/div/div/ul/li/a[1]")))
for tch_num in teacher_cnt:
# 선생님 선택
teacher_btn = driver.find_elements_by_xpath("//*[@id=\"tchContArea\"]/div/div/ul/li/a[1]")[tch_num]
teacher_btn.send_keys(Keys.ENTER)
time.sleep(2)
all_class = driver.find_element_by_xpath("//*[@id=\"container\"]/div[1]/div[1]/ul/li[2]/ul/li[1]/a")
all_class.send_keys(Keys.ENTER)
time.sleep(2)
all_class = driver.find_element_by_xpath("//*[@id=\"iMenuList1\"]")
all_class.send_keys(Keys.ENTER)
time.sleep(2)
for j in range(len(driver.find_elements_by_xpath("//*[@id=\"divChrTabArea\"]/ul/li/a"))):
cate = driver.find_elements_by_xpath("//*[@id=\"divChrTabArea\"]/ul/li/a")[j]
cate.send_keys(Keys.ENTER)
time.sleep(2)
for i in range(3):
megastudy['teacher'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[1]/a")[i].text)
megastudy['title'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[2]/div/p[3]/a")[i].text)
megastudy['subject'].append(subject)
megastudy['grade'].append(re.findall('\[(\S*)\].*', (driver.find_elements_by_css_selector("span.lstedu_bookinfo--class__txt")[i].text))[0])
megastudy['link'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[2]/div/p[3]/a")[i].get_attribute("href"))
driver.get("http://www.megastudy.net/teacher_v2/teacher_main.asp")
time.sleep(2)
subject_btn = driver.find_elements_by_xpath("//*[@id=\"container\"]/div/div[2]/ul/li/a")[sub_num]
subject_btn.send_keys(Keys.ENTER)
time.sleep(2)
if sub_num != 3:
# 고1, 고2 전용
teacher_cnt = range(len(driver.find_elements_by_xpath("//*[@id=\"tchContArea\"]/div/div[1]/ul/li/a[1]")))
for tch_num in teacher_cnt:
# 선생님 선택
teacher_btn = driver.find_elements_by_xpath("//*[@id=\"tchContArea\"]/div/div[1]/ul/li/a[1]")[tch_num]
teacher_btn.send_keys(Keys.ENTER)
time.sleep(2)
all_class = driver.find_element_by_xpath("//*[@id=\"container\"]/div[1]/div[1]/ul/li[2]/ul/li[1]/a")
all_class.send_keys(Keys.ENTER)
time.sleep(2)
all_class = driver.find_element_by_xpath("//*[@id=\"iMenuList1\"]")
all_class.send_keys(Keys.ENTER)
time.sleep(2)
for i in range(3):
megastudy['teacher'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[1]/a")[i].text)
megastudy['title'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[2]/div/p[3]/a")[i].text)
megastudy['subject'].append(subject)
megastudy['grade'].append(re.findall('\[(\S*)\].*', (driver.find_elements_by_css_selector("span.lstedu_bookinfo--class__txt")[i].text))[0])
megastudy['link'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[2]/div/p[3]/a")[i].get_attribute('href'))
all_class = driver.find_element_by_xpath("//*[@id=\"divChrTabArea\"]/ul/li[2]/a")
all_class.send_keys(Keys.ENTER)
time.sleep(2)
for i in range(3):
megastudy['teacher'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[1]/a")[i].text)
megastudy['title'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[2]/div/p[3]/a")[i].text)
megastudy['subject'].append(subject)
megastudy['grade'].append(re.findall('\[(\S*)\].*', (driver.find_elements_by_css_selector("span.lstedu_bookinfo--class__txt")[i].text))[0])
megastudy['link'].append(driver.find_elements_by_xpath("//*[@id=\"iChrList\"]/table/tbody/tr/td[2]/div/p[3]/a")[i].get_attribute('href'))
driver.get("http://www.megastudy.net/teacher_v2/teacher_main.asp")
time.sleep(2)
subject_btn = driver.find_elements_by_xpath("//*[@id=\"container\"]/div/div[2]/ul/li/a")[sub_num]
subject_btn.send_keys(Keys.ENTER)
time.sleep(2)
megastudy = {'title': [], 'teacher': [], 'subject': [], 'grade': [], 'link': []}
crawling()
df = pd.DataFrame.from_dict(megastudy)
df.to_json('megastudy.json', orient = 'records')
| crawling_code/megastudy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
# +
import ipywidgets as widgets
from IPython.display import display
def cb(elem):
print("hi")
button = widgets.Button(description="Connect")
button.on_click(cb)
display(button)
# -
# !pip install pandas
# %connect
# +
import ipywidgets as widgets
from IPython.display import display
def cb(elem):
print("hi")
button = widgets.Button(description="Connect")
button.on_click(cb)
display(button)
# -
# %disconnect
# +
import pandas as pd
import matplotlib.pyplot as plt
from ipywidgets import *
from IPython.display import display
from IPython.html import widgets
plt.style.use('ggplot')
NUMBER_OF_PINGS = 4
# displaying the text widget
text = widgets.Text(description="Domain to ping", width=200)
display(text)
# preparing the plot
data = pd.DataFrame()
x = range(1,NUMBER_OF_PINGS+1)
plots = dict()
fig, ax = plt.subplots()
plt.xlabel('iterations')
plt.ylabel('ms')
plt.xticks(x)
plt.show()
# preparing a container to put in created checkbox per domain
checkboxes = []
cb_container = widgets.HBox()
display(cb_container)
# add button that updates the graph based on the checkboxes
button = widgets.Button(description="Update the graph")
# function to deal with the added domain name
def handle_submit(sender):
# a part of the magic inside python : pinging
# res = !ping -c {NUMBER_OF_PINGS} {text.value}
hits = res.grep('64 bytes').fields(-2).s.replace("time=","").split()
if len(hits) == 0:
print ("Domain gave error on pinging")
else:
# rebuild plot based on ping result
data = hits
data = data.astype(float)
plots, = ax.plot(x, data, label=text.value)
plt.legend()
plt.draw()
# add a new checkbox for the new domain
checkboxes.append(widgets.Checkbox(description = text.value, value=True, width=90))
cb_container.children=[i for i in checkboxes]
if len(checkboxes) == 1:
display(button)
# function to deal with the checkbox update button
def on_button_clicked(b):
for c in cb_container.children:
if not c.value:
plots.set_visible(False)
else:
plots.set_visible(True)
plt.legend()
plt.draw()
button.on_click(on_button_clicked)
text.on_submit(handle_submit)
plt.show()
from IPython.display import HTML
# +
import ipywidgets as widgets
out = widgets.Output(layout={'border': '1px solid black'})
@out.capture(clear_output=True, wait=True)
def handle_click(button):
kernel.disconnect()
kernel.connect(copy.deepcopy(kernel.last_used_backend))
kernel.do_execute(code, True)
widgs = []
widgs.append(button)
ipydisplay(widgets.VBox(widgs))
button = widgets.Button(description="Next Step")
button.on_click(handle_click)
widgs = []
widgs.append(button)
ipydisplay(widgets.VBox(widgs))
# +
import ipywidgets as widgets
out = widgets.Output(layout={'border': '1px solid black'})
def function_with_captured_output():
print('This goes into the output widget')
raise Exception('As does this')
function_with_captured_output()
# -
from io import BytesIO
from PIL import Image
from numpy import ones, zeros
import matplotlib
from IPython.display import display, Image
matplotlib.use('Agg')
from matplotlib import pyplot as plt
try:
from base64 import encodebytes, decodebytes
except ImportError:
from base64 import encodestring as encodebytes, decodebytes
from IPython.display import display, Image
import ipywidgets as widgets
from traitlets import Unicode, validate, Int
# +
def to_img_buffer(arr, fmt="JPEG"):
im = plt.imshow(arr)
bio = BytesIO()
im.write_png(bio)
bio.seek(0)
return bio.read()
def display_raw_png(data):
display(Image(data, format="png"))
# -
import ipywidgets as widgets
from traitlets import Unicode, validate, Int
class TrackballWidget(widgets.DOMWidget):
_view_name = Unicode('TrackballView').tag(sync=True)
_view_module = Unicode('trackball').tag(sync=True)
_view_module_version = Unicode('0.0.0').tag(sync=True)
width = Int(400).tag(sync=True)
height = Int(400).tag(sync=True)
image = Unicode('').tag(sync=True)
def __init__(self, *args, **kwargs):
widgets.DOMWidget.__init__(self, *args, **kwargs)
self.on_msg(self._handle_msg)
self.state = zeros([self.width, self.height])
self.state[20:30,:] = 1
self._update_image()
def _update_image(self):
self.image = encodebytes(to_img_buffer(self.state));
def _add_cube(self, x0, y0, x1, y1, val=1):
x_min, x_max = (x0,x1) if x0 < x1 else (x1,x0)
y_min, y_max = (y0,y1) if y0 < y1 else (y1,y0)
self.state[int(y_min):int(y_max), int(x_min):int(x_max)] = val
def _handle_msg(self, msg, *args, **kwargs):
content = msg["content"]["data"]["content"]
print(content)
if "move" in content:
prev = content["move"]["prev"]
cur = content["move"]["cur"]
self._add_cube(prev["x"], prev["y"], cur["x"], cur["y"])
self._update_image()
elif 'keypress' in content:
code = content['code']
if code == 87: #W
elif code == 65: #A
elif code == 83: #S
elif code == 68: #D
def on_key_press(self, content):
print("Key {c}".format(c=content['code']))
# + language="javascript"
# require.undef('trackball')
#
# define('trackball', ['@jupyter-widgets/base', ], function(widgets) {
# var trackball_view = widgets.DOMWidgetView.extend({
# render: function() {
# this.canvas = document.createElement('canvas');
# this.context = this.canvas.getContext('2d');
# this.canvas.width = this.model.get('width');
# this.canvas.height = this.model.get('height');
# this.canvas.tabIndex = 1;
#
# let is_mouse_down = false;
# let prev_coords = {x: 0, y: 0};
# var that = this;
# // send less update events
# var update_delay = 2;
# var updates = 0;
#
#
# function get_canvas_xy(e) {
# let x = e.clientX;
# let y = e.clientY;
# let bound = that.canvas.getBoundingClientRect();
#
# return {x: x-bound.left, y: y-bound.top};
# }
#
# this.canvas.addEventListener('mousedown', function(e) {
# is_mouse_down = true;
# prev_coords = get_canvas_xy(e);
# });
#
# this.canvas.addEventListener('mouseup', function(e) {
# is_mouse_down = false;
# });
#
# this.canvas.addEventListener('mousemove', function(e) {
# if (!is_mouse_down) { return; }
# if (updates < update_delay) {
# updates++;
# return;
# }
# let coords = get_canvas_xy(e);
# that.send({'move': {'cur': coords, 'prev': prev_coords}});
# prev_coords = coords;
# updates = 0;
# });
#
# this.el.append(this.canvas);
# this.model.on('change:image', this.update, this);
# this.update();
# },
# update: function() {
# let buffer = this.model.get('image');
# var img = new Image;
# img.src = "data:image/png;base64," + buffer;
# var that = this;
# img.onload = function() {
# that.context.clearRect(0, 0, that.canvas.width, that.canvas.height);
# that.context.drawImage(img, 0, 0, that.canvas.width, that.canvas.width);
# };
# },
# events: {
# 'keydown': 'keydown',
# },
# keydown: function(e) {
# var code = e.keyCode || e.which;
# this.send({event: 'keypress', code: code});
# }
# });
#
# return {TrackballView: trackball_view};
# })
# -
s = TrackballWidget()
s
# +
import os.path
from jupyter_core.paths import jupyter_config_dir
jupyter_dir = jupyter_config_dir()
custom_js_path = os.path.join(jupyter_dir, 'custom', 'custom.js')
# my custom js
if os.path.isfile(custom_js_path):
with open(custom_js_path) as f:
print(f.read())
else:
print("You don't have a custom.js file")
# -
| src/ascent/python/ascent_jupyter_bridge/notebooks/Widget Tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/souravgopal25/Data-Structure-Algorithm-Nanodegree/blob/master/Queue.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pNgWyGGbooin" colab_type="code" colab={}
class Queue:
def __init__(self, initial_size=10):
self.arr = [0 for _ in range(initial_size)]
self.next_index = 0
self.front_index = -1
self.queue_size = 0
def enqueue(self, value):
# TODO: Check if the queue is full; if it is, call the _handle_queue_capacity_full method
# enqueue new element
self.arr[self.next_index] = value
self.queue_size += 1
self.next_index = (self.next_index + 1) % len(self.arr)
if self.front_index == -1:
self.front_index = 0
def dequeue(self):
# check if queue is empty
if self.is_empty():
self.front_index = -1 # resetting pointers
self.next_index = 0
return None
# dequeue front element
value = self.arr[self.front_index]
self.front_index = (self.front_index + 1) % len(self.arr)
self.queue_size -= 1
return value
def size(self):
return self.queue_size
def is_empty(self):
return self.size() == 0
def front(self):
# check if queue is empty
if self.is_empty():
return None
return self.arr[self.front_index]
def _handle_queue_capacity_full(self):
old_arr = self.arr
self.arr = [0 for _ in range(2 * len(old_arr))]
index = 0
# copy all elements from front of queue (front-index) until end
for i in range(self.front_index, len(old_arr)):
self.arr[index] = old_arr[i]
index += 1
# case: when front-index is ahead of next index
for i in range(0, self.front_index):
self.arr[index] = old_arr[i]
index += 1
# reset pointers
self.front_index = 0
self.next_index = index
# + id="Dj5UbfmZplU9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 115} outputId="d865016d-4acf-4255-8ab8-2b9d3f3c7ddc"
# Setup
q = Queue()
q.enqueue(1)
q.enqueue(2)
q.enqueue(3)
# Test size
print ("Pass" if (q.size() == 3) else "Fail")
# Test dequeue
print ("Pass" if (q.dequeue() == 1) else "Fail")
# Test enqueue
q.enqueue(4)
print ("Pass" if (q.dequeue() == 2) else "Fail")
print ("Pass" if (q.dequeue() == 3) else "Fail")
print ("Pass" if (q.dequeue() == 4) else "Fail")
q.enqueue(5)
print ("Pass" if (q.size() == 1) else "Fail")
# + id="xlAynWx4pq24" colab_type="code" colab={}
| Queue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/juunnn/DTSAI2019/blob/master/D26_JunaediFahmi.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="juRiKg7hhcwl" colab_type="text"
# # Natural Language Proccessing
# ------
# Why? Selama ada manusia yang berkomunikasi dengan komputer maka jenis interface akan selalu mengalami perubahan mengikuti jenis komunikasi antara manusia dengan manusia (**Natural**).
# + id="UhN-jqRghQ-L" colab_type="code" colab={}
import nltk
# + id="Fbg4Fga9wMDR" colab_type="code" outputId="ad228156-755c-46f7-cd0f-f0076abe7039" colab={"base_uri": "https://localhost:8080/", "height": 68}
nltk.download('punkt')
# + id="M6Svf-howbfI" colab_type="code" colab={}
text = """ Hello, Mr. Smith, how are you doing today? the weather is great and city is awesome. The sky is pinkish-blue. You shouldn't eat cardboard """
# + id="N3G5rPEnwbcI" colab_type="code" outputId="56e1683e-1b50-467f-d88f-455b32f27d9c" colab={"base_uri": "https://localhost:8080/", "height": 85}
from nltk.tokenize import sent_tokenize
tokenized_text = sent_tokenize(text)
tokenized_text
# + id="fmlEz3h3ysee" colab_type="code" outputId="0aaff4a0-4b6b-4e98-bc53-a344c509ca09" colab={"base_uri": "https://localhost:8080/", "height": 527}
from nltk.tokenize import word_tokenize
tokenized_words = word_tokenize(text)
tokenized_words
# + id="3kwLGtIIy7eJ" colab_type="code" outputId="86bd1e8a-f673-45a9-e430-e05f939aaa09" colab={"base_uri": "https://localhost:8080/", "height": 425}
text.split()
# + [markdown] id="ddQBMhY4OWMr" colab_type="text"
# ## Excercise 1
# + id="_w7Hm5PdzkwG" colab_type="code" outputId="6c175efc-944d-4795-a682-ba1b3563e169" colab={"base_uri": "https://localhost:8080/", "height": 255}
def tokenized_words_sent(text):
return [word_tokenize(x) for x in sent_tokenize(text)]
sword = tokenized_words_sent(text)
sword
# + id="Synen-GU0amN" colab_type="code" outputId="efcb3e38-d202-40c4-9a0a-715d855a1eb3" colab={"base_uri": "https://localhost:8080/", "height": 510}
from nltk.util import bigrams, trigrams, ngrams
list(bigrams(tokenized_words))
# + id="xOFRxuiE05J0" colab_type="code" outputId="04ba7d7c-06e2-45a9-cc47-52aa0425912f" colab={"base_uri": "https://localhost:8080/", "height": 493}
list(trigrams(tokenized_words))
# + id="u0IwjFuP1QPC" colab_type="code" outputId="dea2e0c0-16c6-4004-ca7f-32bae9932eeb" colab={"base_uri": "https://localhost:8080/", "height": 459}
list(ngrams(tokenized_words, 5))
# + id="niHQ0yTT1gUI" colab_type="code" outputId="98f0275d-2644-4dcf-a0e8-fffb68948c11" colab={"base_uri": "https://localhost:8080/", "height": 459}
from nltk import FreqDist
freq_dist = FreqDist(tokenized_words)
freq_dist
# + id="gb_3Qzh710n3" colab_type="code" outputId="d715dcc1-2637-40c3-ac78-2a50a371f60a" colab={"base_uri": "https://localhost:8080/", "height": 34}
freq_dist.most_common(3)
# + id="yNXVYP6s17Yf" colab_type="code" outputId="20923dd6-2f56-4075-b6be-59e40f2f8340" colab={"base_uri": "https://localhost:8080/", "height": 334}
import matplotlib.pyplot as plt
# %matplotlib inline
freq_dist.plot()
# + id="4dBUxVBI3LXq" colab_type="code" outputId="6404455f-0690-4209-e6e1-d180182cf891" colab={"base_uri": "https://localhost:8080/", "height": 68}
nltk.download("stopwords")
# + id="NpvTMfUi2RvW" colab_type="code" outputId="ff8332c1-2fa1-4774-c285-ba28bd16c83c" colab={"base_uri": "https://localhost:8080/", "height": 3060}
from nltk.corpus import stopwords
stop_words = set(stopwords.words("english"))
stop_words
# + id="I5iaSxNR2hwT" colab_type="code" outputId="8beda17c-769e-4cfd-9864-92b1d6b96269" colab={"base_uri": "https://localhost:8080/", "height": 357}
filtered_word = []
for w in tokenized_words:
if w not in stop_words:
filtered_word.append(w)
filtered_word
# + [markdown] id="ke-WG8RvIbGL" colab_type="text"
# ## Excercise 2
# + id="ZFeA1gR93WQ-" colab_type="code" outputId="68a91b87-f9cb-4ae2-f5b4-3cc5ab576d2a" colab={"base_uri": "https://localhost:8080/", "height": 54}
def remove_sent(text):
sword = []
for sent in sent_tokenize(text):
sent = word_tokenize(sent)
# print(sent)
nsent = []
for w in sent:
if w not in stop_words:
nsent.append(w)
# print()
sword.append(" ".join(nsent))
return sword
sword = remove_sent(text)
print(sword)
# + id="CzaSqEPC_CnD" colab_type="code" outputId="55eecac3-7cbc-4718-e3fd-db64ba1bf34d" colab={"base_uri": "https://localhost:8080/", "height": 85}
nltk.download("averaged_perceptron_tagger")
# + id="bVqcsCSNIGsj" colab_type="code" outputId="4d162dbe-3b57-403d-ac11-1a6ad1fa5980" colab={"base_uri": "https://localhost:8080/", "height": 527}
nltk.pos_tag(tokenized_words)
# + id="yj8WDfWgJogd" colab_type="code" outputId="35429833-ea72-49ed-f843-71b2ce5da2a8" colab={"base_uri": "https://localhost:8080/", "height": 85}
def filtered_sent3(tok_text):
tags = ['NN', 'NNP', 'VBP', 'VBZ', 'VBG', 'JJ']
pos_tagged = dict(nltk.pos_tag(word_tokenize(tok_text)))
filt = []
for y in pos_tagged:
if pos_tagged[y] in tags:
filt.append(y)
return filt
filted_sent3 = []
for text in tokenized_text:
filted_sent3.append(filtered_sent3(text))
filted_sent3
# + [markdown] id="fQU1XcRFKgHd" colab_type="text"
# ## Wordnet
# + id="xc908fZ4JPBQ" colab_type="code" outputId="ffe51012-dca0-404d-d298-a9c4a5633f79" colab={"base_uri": "https://localhost:8080/", "height": 68}
nltk.download('wordnet')
# + id="QEzAH2RyKkkh" colab_type="code" outputId="ad3fd9b0-76a0-49d2-c71b-913ddce35f3c" colab={"base_uri": "https://localhost:8080/", "height": 51}
from nltk.stem.wordnet import WordNetLemmatizer
lem = WordNetLemmatizer()
from nltk.stem.porter import PorterStemmer
stem = PorterStemmer()
word = 'flying'
print("Lemmatization of word",lem.lemmatize(word,'v'))
print("Stemming of word",stem.stem(word))
# + id="YxsDEyWYLYJF" colab_type="code" colab={}
text2 = "Based on these preliminary experiments, we decided to limit the final evaluation on the unseen test set to the logistic regression model, as its probability prediction allows us to rank chiasmi easily.In addition, its linear implementation allows us toobserve the learned feature weights and comparethem to those of the earlier hand-tuned systems.For the linear logistic regression implementationwe used scikit-learn (Pedregosa et al., 2011)"
# + id="aLJwva-oMLPi" colab_type="code" colab={}
pattern = 'NP: {<DT>?<JJ>*<NN>}'
# + id="p2eQmY07QZkD" colab_type="code" colab={}
cp = nltk.RegexpParser(pattern)
# + id="FoYDScVVR3ow" colab_type="code" colab={}
cs = cp.parse(nltk.pos_tag(nltk.word_tokenize(text2)))
# + id="McnX-STIR_gQ" colab_type="code" outputId="e5e5c11a-e67c-4427-c32c-f48ed7704009" colab={"base_uri": "https://localhost:8080/", "height": 969}
print(cs)
# + [markdown] id="zTbXLMkB9FWi" colab_type="text"
# ## IOB Tag
# + id="H3j8LcihSGTq" colab_type="code" outputId="10fb034e-2b2e-4eb7-aa64-35aa2cfdaa29" colab={"base_uri": "https://localhost:8080/", "height": 1190}
from nltk.chunk import conlltags2tree, tree2conlltags
from pprint import pprint
iob_tagged = tree2conlltags(cs)
pprint(iob_tagged)
# + [markdown] id="R94t4DPy-BOH" colab_type="text"
# ## Named Entity Tagger
# + id="kpBJThGPDWcQ" colab_type="code" outputId="1fa73442-de99-4c22-899e-7f1225980726" colab={"base_uri": "https://localhost:8080/", "height": 119}
nltk.download('maxent_ne_chunker')
nltk.download('words')
# + id="__jixuwL9_LT" colab_type="code" colab={}
text3 = "<NAME> went to Bandung"
sentences = nltk.sent_tokenize(text3)
sentences = [nltk.word_tokenize(sent) for sent in sentences]
sentences = [nltk.pos_tag(sent) for sent in sentences]
tree = [nltk.ne_chunk(sent) for sent in sentences]
# + id="CgvEqEBB-3oE" colab_type="code" outputId="7b224f02-1789-41e6-c830-7bfe597f33c3" colab={"base_uri": "https://localhost:8080/", "height": 102}
sentences
# + id="qJKPpmqL-5hS" colab_type="code" outputId="1acb788d-04cd-48e6-b718-db3f52575ed4" colab={"base_uri": "https://localhost:8080/", "height": 54}
pprint(tree)
# + id="vFMySYSR_dOV" colab_type="code" colab={}
entity_names = []
for sent in nltk.sent_tokenize(text):
for chunk in nltk.ne_chunk(nltk.pos_tag(nltk.word_tokenize(sent))):
if hasattr(chunk, 'label') == 'PERSON':
name_value = ' '.join(child[0] for child in chuck.level())
if name_value not in entity_names:
entitiy_names.append(name_value)
# + id="YJ8izfXaBPDY" colab_type="code" outputId="a5ab733e-aec6-4185-96e7-6ed110a1cb1d" colab={"base_uri": "https://localhost:8080/", "height": 34}
entity_names
# + [markdown] id="EgTdhBJp_Up8" colab_type="text"
# ## Parser
# + id="PXCxBF6HBSbr" colab_type="code" colab={}
grammar = nltk.CFG.fromstring("""
S -> NP VP
PP -> P NP
NP -> Det N | Det N PP | 'I'
Det -> 'an' | 'my'
N -> 'elephant' | 'pajamas'
V -> 'shot'
P -> 'in'
""")
# + id="IGN6khpyC5e0" colab_type="code" colab={}
sent = ['I', 'shot', 'an','elephant','in','my','pajamas']
parser = nltk.ChartParser(grammar)
for tree in parser.parse(sent):
print(tree)
| D26_JunaediFahmi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# # !pip install -e ../ # If not done yet...
# +
import os
import matplotlib.pyplot as plt
from pubrecon.data import DataFrame, ImagesData
from pubrecon.model import RCNN
from pubrecon.hyper import Hyper
# +
# General
data_path = "../data/in/" # Where is your input data
work_path = "../data/out/" # Where everything will be saved
seed = 1337 # Random seed
verbose = 1 # 0: no output; 1: normal informations; 2: e v e r y th i n g
# DataFrame
dataframe_pickle_path = os.path.join(work_path, "dataframe.pickle") # Where will the DataFrame be saved
force_preparation = True # Do you want to bypassed the saved DataFrame
subsamples = 128 # Number of samples to use for the DataFrame; -1: Use all of them
# ImagesData
imagesdata_pickle_path = os.path.join(work_path, 'imagesdata.pickle') # Where will the ImagesData be saved
number_of_results = 2500 # How many samples will selective search use
iou_threshold = 0.85 # What is the percent of precision required
max_samples = 15 # How many class samples do you want
show_infos = True # Show information for images output
show_labels = True # Show labels for images output
# RCNN
model_and_weights_path = "../data/out/" # Where will the model and weights be saved/loaded
loss = None # Loss function; None: Use
opt = None # Optimization function; None: Use Adame
lr = 0.001 # Learning rate
epochs = 5 # Number of epochs
batch_size = 16
split_size = 0.15 # Test/Train proportion
checkpoint_path = os.path.join(work_path, 'checkpoint.h5') # Where will the checkpoints be saved; None: No checkpoint (don't.)
early_stopping = True # Should the learning stop if no more improvment is done
threshold = 0.85 # Threshold used for the recognition
# -
dataframe = DataFrame(data_path, pickle_path=dataframe_pickle_path)
dataframe.prepare_data(force_preparation=force_preparation, subsamples=subsamples, verbose=verbose)
dataframe.summary()
imagesdata = ImagesData(dataframe, pickle_path=imagesdata_pickle_path)
# That part is quite long, beware!
imagesdata.prepare_images_and_labels(number_of_results=number_of_results, iou_threshold=iou_threshold,
max_samples=max_samples, verbose=verbose)
x = imagesdata.images
y = imagesdata.labels
# +
np.save("x", x)
np.save("y", y)
# -
import numpy as np
x = np.load("x.npy")
y = np.load("y.npy")
def rcnn(x_train, y_train, x_val, y_val, param):
vggmodel = VGG16(weights='imagenet', include_top=True) # https://keras.io/applications/
# Freeze first 15 layers
for i, layers in enumerate(vggmodel.layers[:15]):
layers.trainable = False
# Add a {number of classes} unit softmax dense layer
predictions = Dense(len(set(y_train)), activation="softmax")(
vggmodel.layers[-2].output) # Maybe not all labels
model = Model(input=vggmodel.input, output=predictions)
# Compile the model using Adam optimizer with learning rate of 0.001 by default
# We are using categorical_crossentropy as loss by default since the output of the model is categorical
model.compile(loss=param['loss'], optimizer=param['opt'](lr=lr_normalizer(param['lr'], param['opt'])), metrics=["acc"])
class MyLabelBinarizer(LabelBinarizer):
def transform(self, y):
Y = super().transform(y)
if self.y_type_ == 'binary':
return np.hstack((Y, 1 - Y))
else:
return Y
def inverse_transform(self, Y, threshold=None):
if self.y_type_ == 'binary':
return super().inverse_transform(Y[:, 0], threshold)
else:
return super().inverse_transform(Y, threshold)
chosen_binarizer = MyLabelBinarizer()
train_labels_fit = chosen_binarizer.fit_transform(y_train)
classes = chosen_binarizer.classes_
# Dataset augmentation
# This may not be needed following some magazines, as we do not often have rotated texts...
# ... Or do we? Anyway it applies for the pictures so there's that.
imgdatagen = ImageDataGenerator(horizontal_flip=True, vertical_flip=True, rotation_range=90)
train_data = imgdatagen.flow(x=x_train, y=y_train, batch_size=param['batch_size'])
imgdatagen = ImageDataGenerator(horizontal_flip=True, vertical_flip=True, rotation_range=90)
test_data = imgdatagen.flow(x=x_val, y=y_val, batch_size=param['batch_size'])
print(train_datata)
# FINALLY train the model. https://keras.io/models/sequential/#fit_generator
steps = ceil(len(train_data) / param['batch_size'])
hist = model.fit(x_train, y_train, batch_size=param['batch_size'], steps_per_epoch=steps, epochs=param['epochs'], verbose=0,
validation_data=[x_val, y_val], validation_steps=steps)
return hist, model
p = {'loss': ['categorical_crossentropy'],
'lr': (0.1, 10, 10),
'epochs': [200],
'opt': [Adam, Nadam],
'batch_size': [2, 3, 4],
'split_size': [0.1, 0.15],
}
t = talos.Scan(x=x, y=y, params=p, model=rcnn, experiment_name='diabetes')
from keras.optimizers import Adam, Nadam
import talos
from keras.applications.vgg16 import VGG16
from keras.layers import Dense
from keras import Model
from talos.utils import lr_normalizer
from sklearn.preprocessing import LabelBinarizer
from keras.preprocessing.image import ImageDataGenerator
from keras.losses import categorical_crossentropy
from math import ceil
analyze_object = talos.Analyze(t)
# get the highest result for any metric
analyze_object.high('val_acc')
# get the round with the best result
analyze_object.rounds2high('val_acc')
# get the best paramaters
analyze_object.best_params('val_acc', ['acc', 'loss', 'val_loss'])
# +
# heatmap correlation
analyze_object.plot_corr('val_loss', ['acc', 'loss', 'val_loss'])
# -
def model(x_train, y_train, x_val, y_val, params):
vggmodel = VGG16(weights='imagenet', include_top=True) # https://keras.io/applications/
# Freeze first 15 layers
for i, layers in enumerate(vggmodel.layers[:15]):
layers.trainable = False
# Add a {number of classes} unit softmax dense layer
predictions = Dense(imagesdata.get_num_classes(), activation="softmax")(
vggmodel.layers[-2].output) # Maybe not all labels
self.model = Model(input=vggmodel.input, output=predictions)
# Compile the model using Adam optimizer with learning rate of 0.001 by default
# We are using categorical_crossentropy as loss by default since the output of the model is categorical
self.model.compile(loss=param['loss'], optimizer=param['opt'], metrics=["acc"])
class MyLabelBinarizer(LabelBinarizer):
def transform(self, y):
Y = super().transform(y)
if self.y_type_ == 'binary':
return np.hstack((Y, 1 - Y))
else:
return Y
def inverse_transform(self, Y, threshold=None):
if self.y_type_ == 'binary':
return super().inverse_transform(Y[:, 0], threshold)
else:
return super().inverse_transform(Y, threshold)
chosen_binarizer = MyLabelBinarizer()
train_labels_fit = chosen_binarizer.fit_transform(labels)
self.classes = chosen_binarizer.classes_
# Dataset augmentation
# This may not be needed following some magazines, as we do not often have rotated texts...
# ... Or do we? Anyway it applies for the pictures so there's that.
imgdatagen = ImageDataGenerator(horizontal_flip=True, vertical_flip=True, rotation_range=90)
train_data = imgdatagen.flow(x=x_train, y=y_train, batch_size=batch_size)
imgdatagen = ImageDataGenerator(horizontal_flip=True, vertical_flip=True, rotation_range=90)
test_data = imgdatagen.flow(x=x_test, y=y_test, batch_size=batch_size)
# FINALLY train the model. https://keras.io/models/sequential/#fit_generator
steps = ceil(len(train_data) / param['batch_size'])
self.hist = self.model.fit_generator(generator=train_data, steps_per_epoch=steps, epochs=param['epochs'], verbose=0,
validation_data=test_data, validation_steps=steps)
import talos
scan_object = talos.Scan(images,
labels,
model=model,
params=params,
experiment_name='aaa',
fraction_limit=.001)
| notebooks/Tests and research/4. Hyperparameters tunning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import scipy
from scipy import stats
d = stats.norm.rvs(loc = 5., scale = 0.01, size = 100000)
fig, ax = plt.subplots(1, 1)
ax.hist(d,50, density=True)
plt.tick_params(labelsize = 15)
plt.xlim([4.95,5.05])
x = np.linspace(4.95,5.05,1000)
ax.plot(x,stats.norm.pdf(x,loc = 5., scale = 0.01),linewidth = 8,alpha = 0.7)
plt.show()
ax = plt.hist(d,50)
plt.yscale('log')
plt.tick_params(labelsize = 10)
plt.xlim([4.95,5.05])
plt.show()
fig, ax = plt.subplots(1, 1)
ax.hist(d,50, density=True)
plt.yscale('log')
plt.tick_params(labelsize = 15)
plt.xlim([4.95,5.05])
x = np.linspace(4.95,5.05,1000)
ax.plot(x,stats.norm.pdf(x,loc = 5., scale = 0.01),linewidth = 8,alpha = 0.7)
plt.show()
| HW#1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# Powerlaw Sphere
# ===============
# -
import flaremodel as fm
import numpy as np
import matplotlib.pyplot as plt
SAVE_PLOT = True
# +
nus = np.logspace(7, 13, 100)
SgrA = fm.SgrA()
B, ne, R, x, p, g_min, g_max, incang, rsteps = 10, 1e7, SgrA.Rs, 50., 1, 3., 1e4, -1, 50
edist = "powerlaw"
R2 = R
R1 = R2/x
params = [p, g_min, g_max]
# +
n = 2.
m = 1.
p = 3. # equivalent to a=1
params = [p, g_min, g_max]
r_p_fun = lambda r, n: (r/R1)**-n
r_pm_fun = lambda r: r_p_fun(r, m) # Radial B field
r_pn_fun = lambda r: r_p_fun(r, n) # Radial edist
# +
def L_nu_bg_fun(nus):
s = n + m*((p-1)/2.+1)
j_nu_pl_br = fm.j_nu_brute(nus, ne, B, params, "powerlaw", incang=-1)
return 4*np.pi*4*np.pi*R1**3*j_nu_pl_br*(x**(3-s)-1)/(3-s)
L_nu_bg = L_nu_bg_fun(nus)
# -
plt.figure(figsize=(6,5))
for i in [4, 2, 1]:
sr_sp = fm.RadialSphere(n_r_fun=r_pn_fun, B_r_fun=r_pm_fun,
edist=edist, rsteps=int((i)*x))
sr_sp_syn = sr_sp.compute_synchrotron(nus, ne, [R2, R1, -1], B, params)
if i == 4:
nu_t = nus[np.argmax(sr_sp_syn)]
syn_p = np.max(sr_sp_syn)
plt.loglog(nus/nu_t, sr_sp_syn/syn_p/2, label=r"$R/\Delta R$=%d" % (50*i))
plt.loglog(nus/nu_t, L_nu_bg/syn_p/2, 'k--')
plt.legend(loc=2)
plt.ylim([1e-2, 1])
plt.xlim([1e-2, 10])
plt.ylabel(r"$L_\nu$ [arb.]")
plt.xlabel(r"Frequency $[\nu/\nu_T]$")
if SAVE_PLOT: plt.savefig("BG85_pl.eps", bbox_inches="tight", dpi=300)
| notebooks/PowerlawSphere.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <small><i>This notebook was prepared by [<NAME>](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).</i></small>
# # Challenge Notebook
# ## Problem: Remove duplicates from a linked list
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Is this a singly or doubly linked list?
# * Singly
# * Can you insert None values in the list?
# * No
# * Can you use additional data structures?
# * Implement both solutions
# * Can we assume we already have a linked list class that can be used for this problem?
# * Yes
# ## Test Cases
#
# * Empty linked list -> []
# * One element linked list -> [element]
# * General case with no duplicates
# * General case with duplicates
# ## Algorithm
#
# Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/linked_lists/remove_duplicates/remove_duplicates_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
# %run ../linked_list/linked_list.py
# %load ../linked_list/linked_list.py
class MyLinkedList(LinkedList):
def remove_dupes(self):
# TODO: Implement me
pass
# ## Unit Test
#
#
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_remove_duplicates.py
from nose.tools import assert_equal
class TestRemoveDupes(object):
def test_remove_dupes(self, linked_list):
print('Test: Empty list')
linked_list.remove_dupes()
assert_equal(linked_list.get_all_data(), [])
print('Test: One element list')
linked_list.insert_to_front(2)
linked_list.remove_dupes()
assert_equal(linked_list.get_all_data(), [2])
print('Test: General case, duplicates')
linked_list.insert_to_front(1)
linked_list.insert_to_front(3)
linked_list.insert_to_front(1)
linked_list.insert_to_front(1)
linked_list.remove_dupes()
assert_equal(linked_list.get_all_data(), [1, 3, 2])
print('Test: General case, no duplicates')
linked_list.remove_dupes()
assert_equal(linked_list.get_all_data(), [1, 3, 2])
print('Success: test_remove_dupes\n')
def main():
test = TestRemoveDupes()
linked_list = MyLinkedList(None)
test.test_remove_dupes(linked_list)
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/linked_lists/remove_duplicates/remove_duplicates_solution.ipynb) for a discussion on algorithms and code solutions.
| linked_lists/remove_duplicates/remove_duplicates_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from yelpapi import YelpAPI
# +
# Creating an empty list to store dictionary
# -
dict_list=[]
# +
# API call to search based on term=['cafes','restaurants','bars'] and
# location=['Brooklyn,NY','Bronx, NY','Manhattan, NY','Queens, NY', 'Staten Island, NY']
# As we can only fetch 1000 rows at a time, try different combination( e.g: term= cafe and location=Manhattan,NY)
# We run a for loop on the offset (pagination- Going from 1-50, 51-100...) and store our result in a dictionary
# +
import requests
import json
import pandas as pd
from pandas.io.json import json_normalize
from pprint import pprint
#file = open('YelpAPI_Data.json', 'w+')
api_key= #put your key
url="https://api.yelp.com/v3/businesses/search"
# n=['Brooklyn,NY','Bronx, NY','Manhattan, NY','Queens, NY', 'Staten Island, NY']
neighborhood= 'Manhattan, NY'
for x in range(20):
params={'term':'bars','location':neighborhood,'limit':50,'offset':x*50}
headers= {'Authorization': 'bearer %s' % api_key}
response=requests.get(url, headers=headers, params=params)
# proceed only if the status code is 200
print('The status code is {}'.format(response.status_code))
d1=response.json()
dict_list.append(d1)
# +
# We separate the key-value of the dictionary to columns names and values using indexing.. and store the cleaned data in a list
# +
from pprint import pprint
import re
s=['coordinates','location']
#customized header
header =['id', 'alias', 'name', 'image_url', 'is_closed', 'url', 'review_count', 'category alias', 'category title', 'rating',
'coordinates', 'latitude', 'longitude',
'street', 'apt', 'area','city','pincode','country','state','address1','address2',
'phone', 'display_phone', 'distance', 'price']
csv_data = []
csv_data.append(header)
for city in dict_list:
for n, item in enumerate(city):
if item == 'businesses':
for value in city[item]:
row = []
price = ' '
for each in value:
if each in s:
for key in value[each]:
if type(value[each][key]) is list:
row.append("-".join(value[each][key]))
else:
row.append(value[each][key])
elif each=='categories':
for j in value[each][0]:
row.append(value[each][0][j])
elif each=='transactions':
row.append("-".join(value[each]))
elif each=='price':
price = value[each]
else:
val = re.sub(r'[^\x00-\x7F]+','', str(value[each]))
row.append('"' + val + '"')
row.append(price)
#print(row)
csv_data.append(row)
# +
# Now we have lists instead list (csv_data), so we can convert them into a csv file..
# -
import csv
from csv import DictWriter
with open('cafes_mn.csv', 'w', encoding='utf8') as f:
for row in csv_data:
f.write(",".join([str(val) for val in row]) + "\n")
# +
## We have collected listings of restaurants, bars and cafes based on the neighborhood,
# let's group them into bars, retsaurants,cafes
# -
import pandas as pd
# +
# All restaurants in NY
# -
rest_mn=pd.read_csv('restaurants/rest_mn.csv',error_bad_lines=False)
rest_qn=pd.read_csv('restaurants/rest_qn.csv',error_bad_lines=False)
rest_bk=pd.read_csv('restaurants/rest_bk.csv',error_bad_lines=False)
rest_bx=pd.read_csv('restaurants/rest_bx.csv',error_bad_lines=False)
rest_si=pd.read_csv('restaurants/rest_si.csv',error_bad_lines=False)
df_rest=[rest_mn,rest_qn,rest_bk,rest_bx,rest_si]
rest=pd.concat(df_rest)
# +
# Total restaurnats collected
# -
print(rest.shape)
# +
# How many of them are unque? It's possible that restaurants in Bronx might be in brooklyn too
# -
print(rest.id.nunique())
# +
# All cafes in NY
# -
cafes_mn=pd.read_csv('cafes/cafes_mn.csv',error_bad_lines=False)
cafes_qn=pd.read_csv('cafes/cafes_qn.csv',error_bad_lines=False)
cafes_bk=pd.read_csv('cafes/cafes_bk.csv',error_bad_lines=False)
cafes_bx=pd.read_csv('cafes/cafes_bx.csv',error_bad_lines=False)
cafes_si=pd.read_csv('cafes/cafes_si.csv',error_bad_lines=False)
df_cafes=[cafes_mn,cafes_qn,cafes_bk,cafes_bx,cafes_si]
cafes=pd.concat(df_cafes)
print(cafes.shape)
print(cafes.id.nunique())
# +
# All bars in NY
# -
bars_mn=pd.read_csv('bars/bars_mn.csv',error_bad_lines=False)
bars_qn=pd.read_csv('bars/bars_qn.csv',error_bad_lines=False)
bars_bk=pd.read_csv('bars/bars_bk.csv',error_bad_lines=False)
bars_bx=pd.read_csv('bars/bars_bx.csv',error_bad_lines=False)
bars_si=pd.read_csv('bars/bars_si.csv',error_bad_lines=False)
df_bars=[bars_mn,bars_qn,bars_bk,bars_bx,bars_si]
bars=pd.concat(df_bars)
print(bars.shape)
print(bars.id.nunique())
# +
# Now let's merge all the dataframes
# -
df_final=[bars,rest,cafes]
final=pd.concat(df_final)
# +
# Total listings collected..
# -
print(final.shape)
# +
# But how many are unique?? Are bars listed separately? Or Bars are also shown in Restaurants?
# DO cafes come in Restaurnats as well?
# -
print(final.id.nunique())
# +
## Clearly Restaurants, bars and cafes are inter-shown when you searh for either of them..
# +
# So, let's get rid of duplicate values! Here, we know we have 8462 unique values..
# -
final.drop_duplicates(subset='id',keep="last",inplace=True)
print(final.shape)
print(final.id.nunique())
# +
# let's save this dataframe to csv for future use
# -
final.to_csv('final_yelp.csv',index=False)
| Jupyter Notebook/Yelp/Yelp-API-Data-Collection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import networkx as nx
import SimpleITK as sitk
import scipy
import skimage
from utils import (
preprocessing, rashno_ns_mapper, get_ilm_line, get_rpe_line, rpe_flattening,
get_retina_mask, ns_mapper, clustering, select_seeds, get_graph_flow_adj_matrix,
post_processing, save_segmentation, alpha_mean_beta_enh, image_from_minimum_cut
)
import time
def plot_slices(im, ref):
plt.figure(figsize=(20,5))
for k, i in enumerate(range(0, ref.shape[0],10)):
plt.subplot(1,5,k+1)
plt.title(f'Slice N°{i}')
plt.imshow(im[i,:,:], cmap='gray')
plt.imshow(ref[i,:,:], alpha=0.5)
plt.axis('off')
plt.show()
def plot_results(im, ref, retina_mask, result_clust, min_cut, segmentation):
plt.figure(figsize=(20,5))
plt.subplot(1,5,1)
plt.title('Preprocessed + Reference')
plt.imshow(im, cmap='gray')
plt.imshow(ref, alpha=0.5)
plt.axis('off')
plt.subplot(1,5,2)
plt.title('Retina Mask')
plt.imshow(im, cmap='gray')
plt.imshow(retina_mask, alpha=0.5)
plt.axis('off')
plt.subplot(1,5,3)
plt.title('Clustering for seeds')
plt.imshow(im, cmap='gray')
plt.imshow(result_clust, alpha=0.5)
plt.axis('off')
plt.subplot(1,5,4)
plt.title('Minimum Cut')
plt.imshow(im, cmap='gray')
plt.imshow(min_cut, alpha=0.5)
plt.axis('off')
plt.subplot(1,5,5)
plt.title('Postprocessed Segmentation')
plt.imshow(im, cmap='gray')
plt.imshow(segmentation, alpha=0.5)
plt.axis('off')
plt.show()
# +
# for i in range(26,49):
i=31
case = f'TRAIN0{i}'
# Image Loading:
manufacturer = 'Spectralis'
itkimage = sitk.ReadImage(f'../data/RETOUCH-TrainingSet-{manufacturer}/{case}/oct.mhd')
ref_itkimage = sitk.ReadImage(f'../data/RETOUCH-TrainingSet-{manufacturer}/{case}/reference.mhd')
oct_scan = sitk.GetArrayFromImage(itkimage)
ref_oct_scan = sitk.GetArrayFromImage(ref_itkimage)
spacing = itkimage.GetSpacing()
complete_segmentation = np.zeros(ref_oct_scan.shape)
start = time.time()
for j in range(0, ref_oct_scan.shape[0]):
print(f'Processing Slice: {j}')
slice_ = j
if j==0:
retina_mask_prev = np.ones(ref_oct_scan[slice_, :, :].shape)
retina_mask_2prev = np.ones(ref_oct_scan[slice_, :, :].shape)
# Preprocess:
im, ref, indx_start, indx_end = preprocessing(
manufacturer, oct_scan[slice_, :, :], ref_oct_scan[slice_, :, :], itkimage.GetSpacing()
)
# Get the NS tranform
T, I = rashno_ns_mapper(im)
# Get ILM line
g1, R, V, ilm_image = get_ilm_line(im, T, r_window=50)
# Get RPE line and flatten()
g2, R, U, V2, rpe_image, indices = get_rpe_line(im, T, ilm_image, r_window=150, u_window=20)
rpe_image, _, _, _, _ = \
rpe_flattening(rpe_image, peak_window_size=500, tr=80, k_size=101, iterations=3)
# Get retina mask
retina_mask = get_retina_mask(ilm_image.copy(), rpe_image.copy())
alt_retina_mask = get_retina_mask(ilm_image.copy(), rpe_image.copy(), alt=True)
# Resize to reduce computation
rel = spacing[1]/spacing[0]
im_or = im.copy()
ref_or = ref.copy()
im = scipy.ndimage.zoom(im, (1, rel), order=3, mode='reflect', prefilter=False)
ref = scipy.ndimage.zoom(ref, (1, rel), order=0, prefilter=False)
retina_mask = scipy.ndimage.zoom(retina_mask, (1, rel), order=0, prefilter=False)
alt_retina_mask = scipy.ndimage.zoom(alt_retina_mask, (1, rel), order=0, prefilter=False)
# Neutrosophic transform and alpha mean beta enhacement
im_NS = ns_mapper(im, inversion=True)
_, T_am_be, betaI = alpha_mean_beta_enh(im_NS)
# Kmeans clustering
result_clust = clustering(T_am_be, alt_retina_mask, 6)
# Seeds sampling
fluid_seeds, bkg_seeds, clusters_centers, indices = \
select_seeds(T_am_be, result_clust, FN=500, TN=700, n_clust_object=1)
indices = indices-1
# Obtain the full weigthed graph
graph_, nodes = get_graph_flow_adj_matrix(
T_am_be, alt_retina_mask, betaI, fluid_seeds,
bkg_seeds, lamda1=1e8, lamda2=1e8, directed=False
)
# Get mincut
G = nx.from_scipy_sparse_matrix(graph_, create_using=nx.Graph, edge_attribute='capacity')
cut_value, partition = nx.minimum_cut(G, 0, graph_.shape[0]-1)
min_cut = image_from_minimum_cut(partition, T_am_be.shape, nodes)
# Post process segmentation
segmentation, segm, labels, final_ = \
post_processing(min_cut, T_am_be, result_clust, retina_mask, indices, Tr=50)
# Save the slice
complete_segmentation[slice_, indx_start:indx_end, :] = segmentation
complete_segmentation[slice_, :, :] = \
complete_segmentation[slice_, :, :]*retina_mask_2prev
# Acumulate previous retina masks
retina_mask_2prev = retina_mask_prev
retina_mask_prev = np.zeros(complete_segmentation[slice_,:,:].shape)
retina_mask_prev[indx_start:indx_end, :] = retina_mask
filename = f'../data/RETOUCH-TrainingSet-{manufacturer}/{case}/segmentation_.mhd'
save_segmentation(complete_segmentation, ref_itkimage, filename)
print(f'The whole processing of the oct study took: {time.time()-start}')
| MAIA_image_processing_demo_complete_case.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
new_arr = np.array([[5,1,9,11],[2,4,8,10],[13,3,6,7],[15,14,12,16]])
inverse_arr = np.array([[15,13,2,5],[14,3,4,1],[12,6,8,9],[16,7,10,11]])
new_arr
inverse_arr
# +
# for i in range(0,len(new_arr)):
# for j in range(0,len(new_arr)):
# print((i,j),new_arr[i,j])
# -
print("INPUT: \n")
print(new_arr)
new_list = []
new_list_1 = []
for i in range(0,len(new_arr)):
for j in range(0,len(new_arr)):
# print((i,j),new_arr[i,j])
temp = new_arr[i,j]
new_arr[i,j] = new_arr[j,i]
new_arr[j,i] = temp
new_list.append(temp)
print("OUTPUT:\n")
print(new_list)
| Linked_list/rotate_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="dCqpeEVNImBz"
# This code generates **Figure 7**
# + id="OAQwi-EHroP4"
import torch
import torch
import torchvision
import torchvision.transforms as transforms
import numpy as np
import scipy as sc
from matplotlib import pyplot as plt
import torch.nn as nn
# + colab={"base_uri": "https://localhost:8080/"} id="kf3mhKg1JGYC" outputId="b0fc281a-0567-4c40-b8f3-fa10f9e42f9f"
# # !nvidia-smi --query-gpu=gpu_name,driver_version,memory.total --format=csv
# + colab={"base_uri": "https://localhost:8080/", "height": 100, "referenced_widgets": ["1d2e9de93e914aad87f8b5b3834cf7e9", "1a28c279c37541a0bd1ebee7dd093cf4", "a9c7574f7269494b828fc791ae347a1c", "b34934a9dfea430d9bc4100a376496f6", "fa6941aa382f48979c08291723d244e2", "c398ab9408c94279806243f9100e447e", "3c483155b0d546d88a248807f9fa2d9e", "21e06bfaec6642f4b56d78ef15884aed"]} id="eG_gGxuP6Tvl" outputId="7f9ea9ec-b1bf-4aaa-aac4-0426a977f7db"
min_batch_size = 30
#transforms.Normalize((0.1307,), (0.3081,))
transform = transforms.Compose(
[transforms.ToTensor()])
train_dataset = torchvision.datasets.CIFAR10(root='CIFAR', train=True,
download=True, transform=transform)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=min_batch_size,
shuffle=True, num_workers=1)
# + id="DOV3c0HZ6VnZ"
dtype = torch.cuda.FloatTensor
dtype_labels = torch.cuda.LongTensor
# + id="wl7C45aH6jeX"
import torch.nn as nn
class AlexNet(torch.nn.Module):
def __init__(self, output_dim,num_layers = 0,level = 3, withbn = False):
super().__init__()
filters = [64, 192 , 385, 256, 256 ]
layers = []
layers += [nn.Conv2d(3, 64, 3, 2, 1)] #in_channels, out_channels, kernel_size, stride, padding
layers += [nn.MaxPool2d(2)] #kernel_size
layers += [nn.ReLU(inplace = True)]
layers += [nn.Conv2d(64, 192, 3, padding = 1)]
layers += [nn.MaxPool2d(2)]
layers += [nn.ReLU(inplace = True)]
# -----------
layers += [nn.Conv2d(192, 192, 3, padding = 1)]
layers += [nn.ReLU(inplace = True)]
if level == 2:
for i in range(num_layers):
layers += [nn.Conv2d(192, 192, 3, padding = 1,bias=False),nn.ReLU(inplace=True)]
if withbn:
layers += [nn.BatchNorm2d(192)]
# ------------
layers += [nn.Conv2d(192, 256, 3, padding = 1)]
layers += [nn.Conv2d(256, 256, 3, padding = 1),nn.ReLU(inplace=True)]
if level == 3:
for i in range(num_layers):
layers += [nn.Conv2d(256, 256, 3, padding = 1),nn.ReLU(inplace=True)]
if withbn:
layers += [nn.BatchNorm2d(256)]
layers += [nn.MaxPool2d(2)]
layers += [nn.ReLU(inplace = True)]
self.features = nn.Sequential(*layers)
self.classifier = nn.Sequential(
nn.Dropout(0.5),
nn.Linear(1024, 4*1024),
nn.ReLU(inplace = True),
nn.Dropout(0.5),
nn.Linear(4*1024, 4*1024),
nn.ReLU(inplace = True),
nn.Linear(4*1024, output_dim),
)
def forward(self, x):
x = self.features(x)
h = x.view(x.shape[0], -1)
x = self.classifier(h)
return x
def forward_noclassi(self, x):
x = self.features(x)
h = x.view(x.shape[0], -1)
return h
def weights_init(m):
if isinstance(m, torch.nn.Linear):
torch.nn.init.xavier_uniform_(m.weight,gain=torch.nn.init.calculate_gain('relu'))
if m.bias is not None:
m.bias.data.fill_(0)
if isinstance(m, torch.nn.Conv2d):
torch.nn.init.xavier_uniform_(m.weight,gain=torch.nn.init.calculate_gain('relu'))
if m.bias is not None:
m.bias.data.fill_(0)
# + id="yULi_YB27iGB"
def train(net,epoch_num = 10,stepsize = 0.01):
criterion = nn.CrossEntropyLoss(size_average=True)
opt= torch.optim.SGD(net.parameters(), lr=stepsize)
output = []
num_total = len(train_dataset.targets)
for j in range(epoch_num):
loss_epoch = 0
for i, data in enumerate(train_loader):
x, y = data
x = x.type(dtype)
y = y.type(dtype_labels)
opt.zero_grad()
outx = net.forward(x)
loss = criterion(outx, y)
loss.backward()
loss_epoch += loss.item()*min_batch_size/num_total
# if i % 1000 ==0:
# print(i)
opt.step()
print(loss_epoch)
output.append(loss_epoch)
return output
# + [markdown] id="Wu6kV587I9KQ"
# The following function extend the svd based initialization to convolutional networks. When samples are pass throught the network layers. We use SVD decomposition of hidden representations to recurrently initialize the weights through the layers.
# + id="kcho4VL3-OKB"
def novel_initialization(anet_im): # the input is a network
# picks a batch of samples
bs = 256
train_loader2 = torch.utils.data.DataLoader(train_dataset, batch_size=bs,
shuffle=True, num_workers=1)
dataiter = iter(train_loader2)
images, labels = dataiter.next()
images = images.type(dtype)
out = anet_im.forward_noclassi(images)
M = out.t().mm(out)
print(torch.norm(M)/torch.trace(M))
x = images
steps = 1200
for unit in anet_im.features:
if isinstance(unit,nn.Conv2d):
if unit.in_channels == unit.out_channels:
zp = torch.nn.ZeroPad2d(unit.padding[0])
input = x.data
print(input.size())
input = zp(input)
w = unit.weight
d2 = input.size(2)
d3 = input.size(3)
### here we reshape representations to implemnet the convolution using matrix multiplication
inp_unf = torch.nn.functional.unfold(input,kernel_size=unit.kernel_size,stride=unit.stride)
inp_unf_size = inp_unf.size()
inp_unf = inp_unf.transpose(1,2)
inp_unf = inp_unf.reshape(inp_unf.size(0)*inp_unf.size(1),inp_unf.size(2))
### svd decomposition of the reshaped representations
u,s,v = torch.svd(inp_unf)
wd = w.view(w.size(0), -1).t().size(1)
w1 = u[0:wd,0:wd].mm(torch.diag(1/torch.pow(s[0:wd],0.5))).mm(v.t()[0:wd,:])
unit.weight.data = w1.reshape(w.size())
unit.weight.data = unit.weight.data/torch.norm(unit.forward(x))
print(torch.norm(unit.weight.data))
x = unit(x)
H = x.flatten(1)
M = H.mm(H.t())
# print(torch.norm(M)/torch.trace(M))
# print('=======')
out = anet_im.forward_noclassi(images)
M = out.t().mm(out)
print(torch.norm(M)/torch.trace(M))
return anet_im
# + id="PDdjwkU59hx0"
layers = [15,75]
epochs = 20
repeat = 4
lr = 0.001
results_xavier = []
results_novel = []
for i in range(repeat):
result_xavier = []
result_novel = []
for layer in layers:
print(layer,'============')
anet = AlexNet(10,num_layers=layer,withbn = False)
anet = anet.cuda()
anet = anet.apply(weights_init)
conv = train(anet,epoch_num=epochs,stepsize=lr)
result_xavier.append(conv)
anet_novel = AlexNet(10,num_layers=layer,withbn=False)
anet_novel = anet_novel.cuda()
anet_novel.apply(weights_init)
anet_novel = novel_initialization(anet_novel)
conv_novel = train(anet_novel,epoch_num=epochs,stepsize=lr)
result_novel.append(conv_novel)
results_novel.append(result_novel)
results_xavier.append(result_xavier)
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="bLgMcEsyKN7h" outputId="90bb61bd-ab5f-41e2-9878-19bfa5fed2cf"
# np.save('novelinit_conv',[results_novel,results_xavier])
# files.download('novelinit_conv.npy')
# + id="byUQHpYQi2YA"
import pandas as pd
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="TOTOcflQfW_7" outputId="d5adb612-fc07-4891-c2f9-b5cdfcc15247"
result_array_novel = np.asarray(results_novel)
result_array_xav = np.asarray(results_xavier)
results_plot=[]
D = []
for i in range(repeat):
run_dict={'run_id': np.ones(epochs,dtype=np.int8)*i,'training loss novel':result_array_novel[i][0],'training loss':result_array_xav[i][0]}
for j in range(epochs):
D.append(j+1)
results_plot.append(pd.DataFrame(run_dict))
results_plot=pd.concat(results_plot)
p2 = sns.lineplot(data=results_plot,x=D,y="training loss", marker='s',ci=95,color='blue',label='xavier')
p1 = sns.lineplot(data=results_plot,x=D,y="training loss novel", marker = 'o', ci=95,color='red',label='orthogonal')
# ax2 = plt.twinx()
# p2 = sns.lineplot(data=results_plot,x=D,y="training loss", marker='s',ci=95,color='blue',ax = ax2)
p2.set(xlabel='epochs')
fig = p2.get_figure()
fig.savefig('convolutional_15.pdf',format='pdf')
files.download('convolutional_15.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="uHVdEycw4qFA" outputId="4459d052-ba6a-4019-f742-1a53cd609fa2"
results_plot=[]
D = []
for i in range(repeat):
run_dict={'run_id': np.ones(epochs,dtype=np.int8)*i,'training loss novel':result_array_novel[i][1],'training loss':result_array_xav[i][1]}
for j in range(epochs):
D.append(j+1)
results_plot.append(pd.DataFrame(run_dict))
results_plot=pd.concat(results_plot)
p2 = sns.lineplot(data=results_plot,x=D,y="training loss", marker='s',ci=95,color='blue',label='xavier')
p1 = sns.lineplot(data=results_plot,x=D,y="training loss novel", marker = 'o', ci=95,color='red',label='orthogonal')
# ax2 = plt.twinx()
# p2 = sns.lineplot(data=results_plot,x=D,y="training loss", marker='s',ci=95,color='blue',ax = ax2)
p2.set(xlabel='epochs')
fig = p2.get_figure()
fig.savefig('convolutional_75.pdf',format='pdf')
files.download('convolutional_75.pdf')
# + id="3yoYaIx52VZh"
| Figure_7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="5y9PlRXkiZ6o"
# # Installation
# + colab={"base_uri": "https://localhost:8080/"} id="lJk6YjCagMU7" outputId="df1b1c2b-bf29-4da4-efac-012562a7ae75"
# Please visit https://github.com/rusty1s/pytorch_geometric#pip-wheels for lastest installation instruction
# !pip install torch-scatter -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html -U
# !pip install torch-sparse -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html -U
# !pip install torch-cluster -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html -U
# !pip install torch-spline-conv -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html -U
# !pip install torch-geometric -U
# + [markdown] id="MmtN4k9gpT61"
# # Introduction
# In this notebook we are going to predict movie ratings in the [MovieLens 100K Dataset](https://grouplens.org/datasets/movielens/100k/). The dataset contains around 100,000 ratings from around 1000 users on 1700 movies. The users and movies have features associated with them.
#
# For this task, we are going to build a graph containing two types of nodes; one representing users, and another representing movies. The graph will contain edges connecting users to the movies they rated. Each edge will contain one attribute, i.e. the rating.
#
# We will be going through the following steps:
#
#
# 1. Downloading and processing the dataset
# 2. Converting the dataset to a PyG Data object
# 3. Training a simple Graph Neural Network to predict movie ratings
# + [markdown] id="5jb6a6wDiepZ"
# # Downloading and Processing Dataset
# + [markdown] id="f-wkDfcedfdX"
# ### Download dataset
# We will first download and unzip the movielens 100k dataset. A full description of the files contained in this dataset can be found [here](http://files.grouplens.org/datasets/movielens/ml-100k-README.txt).
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="vYhmIXp1xJqX" outputId="4f3ad323-0ba0-428f-d88c-4713377bc5e4"
from six.moves import urllib
url ="http://files.grouplens.org/datasets/movielens/ml-100k.zip"
filename = url.rpartition("/")[2] # = ml-100k.zip
data = urllib.request.urlopen(url)
with open(filename, 'wb') as f:
f.write(data.read())
folder=filename[:-4] # ml-100k
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="0cfGawoygEAN" outputId="d2028d57-b560-41ed-940d-b3ae89898eb2"
import zipfile
zip = zipfile.ZipFile(filename)
zip.extractall()
# !ls ml-100k
# + [markdown] id="_0JOYu6Sjnaq"
# ### Loading Edges
# The movie ratings are stored in the file *u.data*. The file contains user ids, movie ids, ratings and timestamps associated with each rating
#
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="wXCuH7vCjrT7" outputId="88b7318a-24ee-4611-ff17-6172618fbd37"
import pandas as pd
filename_edges= folder+"/u.data"
# Load file using pandas , and specify seperator as "|", and provide names of the columns.
# Discard the timestamp
df_edges = pd.read_csv(filename_edges, sep="\t",
header=None, names=["user_id", "movie_id", "rating", "timestamp"],
usecols = ["user_id", "movie_id", "rating"])
df_edges.head()
# + [markdown] id="tdPuooCtk7QF"
# ### Loading User features
#
# Next we will load the user features which are stored in *u.user*. This file contains the age, gender, occupation and zipcode of each user.
# + id="PgvaUYRBncbu"
filename_users = folder +"/u.user"
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="hc1fdF9nk54d" outputId="ccd5ee35-bfac-4a46-a064-6e12f0488d41"
# Load file using pandas , and specify seperator as "|", and provide names of the columns.
# Keep the user_id , age and gender columns and discard the zipcode column
df_users = pd.read_csv(
filename_users,
sep="|",
header=None,
names=["user_id", "age", "gender", "occupation", "zipcode"],
usecols=["user_id", "age", "gender", "occupation"],
)
df_users.head()
# + [markdown] id="VfPsSkcYw2P_"
# We will be ignoring the zipcode, and one-hot encoding the gender and occupation features. We can compute the one-hot encoded features using the pandas *get_dummies* function. We will also standardize the age of the users
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="stET6vsynsvN" outputId="35470da3-a7a6-4bc1-d637-15dac4629b75"
df_gender_onehot = pd.get_dummies(df_users["gender"])
df_occupation_onehot = pd.get_dummies(df_users["occupation"])
age = df_users["age"]
age_standard = (age -age.mean())/age.std()
# post-processed user features
df_users_pp = pd.concat( [df_users["user_id"], age_standard, df_occupation_onehot, df_gender_onehot], axis=1)
df_users_pp.head()
# + [markdown] id="UEzJh3Ljnt56"
# ### Loading Movie features
# Finally we will load the movie features contained in the file *u.item* and *u.genre*
#
# Quoting from the [readme](http://files.grouplens.org/datasets/movielens/ml-100k-README.txt) of the dataset:
#
# ---
#
#
# u.item -- Information about the items (movies); this is a tab separated
# list of
# movie id | movie title | release date | video release date |
# IMDb URL | unknown | Action | Adventure | Animation |
# Children's | Comedy | Crime | Documentary | Drama | Fantasy |
# Film-Noir | Horror | Musical | Mystery | Romance | Sci-Fi |
# Thriller | War | Western |
# The last 19 fields are the genres, a 1 indicates the movie
# is of that genre, a 0 indicates it is not; movies can be in
# several genres at once.
# The movie ids are the ones used in the u.data data set.
#
# u.genre -- A list of the genres.
#
# + id="oqZ9y-fTn_mm"
filename_movies = folder + "/u.item"
# + colab={"base_uri": "https://localhost:8080/"} id="GdGZVb2xq1lF" outputId="7606275f-b3bf-4fd3-9e58-76d7d93ac384"
filename_genre = folder + "/u.genre"
df_genre = pd.read_csv(filename_genre, header=None, sep="|", names=["genre", "id"])
list_genre = df_genre.genre.to_list()
list_genre
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="NJjXhH_7sAOJ" outputId="3a94c12d-8a33-408e-c33c-1cb9164decf2"
movie_columns = ["movie_id", "title", "_", "year", "url"] + list_genre
#Load file using pandas , and specify seperator as "|", and provide names of the columns
df_movies = pd.read_csv(filename_movies, sep="|", header=None, names=movie_columns,
usecols= ["movie_id"] + list_genre )
df_movies
# + [markdown] id="36FjWyGj2h-d"
# ### Summary summarizing the dataset we loaded so far:
# + colab={"base_uri": "https://localhost:8080/", "height": 443} id="p7ChmQn-2jiC" outputId="119c4b1a-369a-4607-9f3d-c0a894cb996c"
df_users_pp # User features
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="gruw--NQ2m2L" outputId="8aefd61a-66b4-4043-b6b2-30040c43a97d"
df_movies # Movie features
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="EYw3L6zb57kb" outputId="09541fb1-ff84-4710-86bc-ae2d25b63289"
df_edges.head()
# + [markdown] id="mpFK06a5CUCx"
# # Converting to a Pytorch Geometric Dataset
# + [markdown] id="j9eSVpdVbldG"
# ## Heterogeneous Graphs in Pytorch Geometric
#
# The dataframe *df_edges* contains an edge list, with the first column referring to an id of the user, and the second column referring to an id of a movie. Pytorch Geometric does not (currently) natively handle this format. Currently PyG expects that the source and target indices in the edge list belong to the same set.
#
# However there is a simple workaround
#
# Consider this edge list, where the first row contains the user ids and the second row contains the movie ids:
#
# ```
# 0 5 4 2 1 3
# 0 1 2 1 2 1
# ```
#
# We can shift the movie ids by the number of users (i.e. 6) to form this edge list
#
# ```
# 0 5 4 2 1 3
# 6 7 8 7 8 7
# ```
#
# We can pass this edge list to existing convolutional operators in PyG without modifying them or writing our own convolutional operator.
#
# However we have to keep in mind that the two node types (users and movies) still represent fundamentally different entities and they both have features of different sizes. We will come back to this issue when we create the graph neural network.
#
#
#
#
# + [markdown] id="-2Rf1ePNCrDd"
# ## Converting to Pytorch Geometric Format
# We will now convert the dataset format to that of Pytorch Geometric. We will maintain the edge index as two separate arrays in order to facilate future calculations related to the heterogeneous nature of the problem which will become clear in the next section
# + id="j6XlMEJGUyQ_"
import torch
from torch_geometric.data import Data
assert (df_movies["movie_id"].isin(df_edges["movie_id"])).all()
assert (df_users_pp["user_id"].isin(df_edges["user_id"])).all()
assert (df_edges["movie_id"].isin(df_movies["movie_id"])).all()
assert (df_edges["user_id"].isin(df_users_pp["user_id"])).all()
# User and Movie IDs
x_user = torch.tensor(df_users_pp.drop(columns=["user_id"]).values)
x_movie = torch.tensor(df_movies.drop(columns=["movie_id"]).values)
# Ids start at 1 in the original dataset.
# Shift the ids back so that they start a 0.
edge_index_user = torch.tensor(df_edges["user_id"].values) - 1
edge_index_movie = torch.tensor(df_edges["movie_id"].values) - 1
# Attributes of edge . ie. ratings
edge_ratings = torch.tensor(df_edges["rating"].values)
# Number of edges
n_edges = edge_ratings.shape[0]
# Checks
assert len(edge_index_user.unique()) == len(x_user)
assert len(edge_index_movie.unique()) == len(x_movie)
# + id="_6wBshuDcz_f"
# Define train and test split for later model validation
test_size = 0.2
train_size= 0.8
ind_cut = int(train_size*n_edges)
edge_ratings_train = edge_ratings[:ind_cut]
edge_ratings_test = edge_ratings[ind_cut:]
edge_index_user_train = edge_index_user[:ind_cut]
edge_index_user_test = edge_index_user[ind_cut:]
edge_index_movie_train = edge_index_movie[:ind_cut]
edge_index_movie_test = edge_index_movie[ind_cut:]
# Package attributes into Data object
device = "cuda"
data = Data(edge_index_user_train = edge_index_user_train.to(device),
edge_index_user_test = edge_index_user_test.to(device),
edge_index_movie_train = edge_index_movie_train.to(device),
edge_index_movie_test = edge_index_movie_test.to(device),
edge_ratings_train = edge_ratings_train.to(device),
edge_ratings_test = edge_ratings_test.to(device),
n_edges = n_edges,
x_user=x_user.float().to(device), x_movie=x_movie.float().to(device),
n_user = x_user.shape[0],
n_movie=x_movie.shape[0]).to(device)
# + [markdown] id="Ebz1nNOqIhhk"
# # Graph Neural Network for Heterogeneous Data
#
# Now that we have defined our data format, we now design a graph neural network to input this data and give out a prediction of movie ratings. The general idea of this graph neural network is
#
#
#
# 1. Map the user and movie feature vectors into feature vectors of the same dimension. This makes it straight forward to use convolutional operators that expect homogeneous nodal features. For this purpose we will use two different linear layers. Note the user nodes have features of 24 dimensions and movie nodes have features of 19 dimensions. But note even if they both had the same dimensions, this step is still necessary since the features have a different meaning!
# 2. Construct an edge list with movie ids shifted as described above
# 3. Pass the features from step 1 and edge list from step 2 through multiple graph convolutional operators
# 4. For each edge, concatenate the features of the adjacent nodes (one of which will be a user node and the other will be a movie node) and pass this concatenated vector through a linear layer to predict the user's rating of the movie
#
#
# + id="PxeOUoafuoGn"
from torch.nn import Linear
from torch_geometric.nn import SGConv
import torch.nn.functional as F
# + id="3NIs8XDFqA0g"
class MovieNet(torch.nn.Module):
def __init__(self,data):
super().__init__()
# Encoder of the user features
self.user_encoder = Linear(data.x_user.shape[1], 5)
# Encoder of the movie features
self.movie_encoder = Linear(data.x_movie.shape[1], 5)
#First convolutional layer
self.conv1 = SGConv(5, 5)
#Second convolutional layer
self.conv2 = SGConv(5, 5)
#Linear layer to predict movie rating
self.regr = Linear(10,1)
def convolutional_operators(self, x, edge_index):
# Pass the encoded features through multiple convolutional layers
x = self.conv1(x, edge_index)
x = F.relu(x)
x = self.conv2(x,edge_index)
x = F.relu(x)
return x
def encode_features(self, x_user, x_movie):
"""
Function that encodes the user and movie features
"""
x_user_enc = self.user_encoder(x_user)
x_user_enc = F.relu(x_user_enc)
x_movie_enc = self.movie_encoder(x_movie)
x_movie_enc = F.relu(x_movie_enc)
return x_user_enc, x_movie_enc
def construct_edge_index(self, data):
# Training edge indices. These are the edges that will be passed to the
# convolutional operators
edge_index_user_train = data.edge_index_user_train
edge_index_movie_train = data.edge_index_movie_train
# Combine the edge indices into the usual format expected by the convolutional operators
edge_index = torch.stack([edge_index_user_train,
edge_index_movie_train])
# Shift the node indices by the number of users, such as the indices
# in edge_index correspond to the indices in *x_combined* feature matrix
# defined above
edge_index[1] += data.n_user
# Convert directed graph to undirected graph to allow information
# to propagate in both direction (movie to user and user to movie)
edge_index_undirected = torch.cat([edge_index, edge_index.flip(dims=(0,1)) ],dim=1)
return edge_index, edge_index_undirected
def forward(self, data):
# Encoded user and movie features
x_user_enc, x_movie_enc = self.encode_features(data.x_user, data.x_movie)
# Concatenate the user and movie encodings
x_combined = torch.cat([x_user_enc,x_movie_enc])
edge_index, edge_index_undirected = self.construct_edge_index(data)
x = self.convolutional_operators(x_combined, edge_index_undirected)
# Extract the features corresponding to user nodes
# and features corresponding to movie nodes using the indices in the
# edge list
user_feature_i = x[edge_index[0]]
movie_feature_j = x[edge_index[1]]
x = torch.cat([user_feature_i,movie_feature_j],dim=1)
y = self.regr(x)
return y.reshape(-1)
def predict(self, edge_index_user, edge_index_movie, data):
"""
Similar function to forward, except that the predictions are made on
the specified user/movie pairs
"""
x_user_enc, x_movie_enc = self.encode_features(data.x_user, data.x_movie)
x_combined = torch.cat([x_user_enc,x_movie_enc])
edge_index, edge_index_undirected = self.construct_edge_index(data)
x = self.convolutional_operators(x_combined, edge_index_undirected)
user_feature_i = x[edge_index_user]
movie_feature_j = x[edge_index_movie+ data.n_user]
x = torch.cat([user_feature_i,movie_feature_j],dim=1)
y = self.regr(x)
return y.reshape(-1)
# + [markdown] id="Ty7ic360gbth"
# # Training and evaluation
#
# Finally we train and evaluate the prediction capability of the Graph Neural Network on the 80/20 split defined earlier.
#
# + colab={"base_uri": "https://localhost:8080/"} id="wbJZb3h7FXfc" outputId="6190020f-06a9-4109-abd8-4b1c4c7bcf5f"
model = MovieNet(data).to(device)
optimizer = torch.optim.Adam(params=model.parameters(), lr=0.01,
weight_decay=0.001)
for epoch in range(5000):
model.train()
optimizer.zero_grad()
y_pred_train = model(data)
# Compute mean-square error
mse_train = torch.mean((y_pred_train-data.edge_ratings_train)**2)
mse_train.backward()
optimizer.step()
if epoch %200 ==0:
model.eval()
y_pred_test = model.predict(data.edge_index_user_test,
data.edge_index_movie_test,
data)
mse_test = torch.mean((y_pred_test-data.edge_ratings_test)**2)
print(" Epoch: {}, RMSE Train: {}, RMSE Test: {}".format(
epoch, torch.sqrt(mse_train),torch.sqrt(mse_test) )
)
# + [markdown] id="klJ3spoHraAO"
#
# ## Optional Exercise
# Try obtaining a Root mean square error (RMSE) test below 1.0 by modifying the GNN defined above by e.g. using dropout layer, using another convolutional operator.
# For reference, a list of RMSE obtained by classical ML methods on the Movie-lens 100k can be found at http://surpriselib.com/. Results by GNN based approaches can be found here https://paperswithcode.com/sota/collaborative-filtering-on-movielens-100k.
# + [markdown] id="a2DPiDiqT0OE"
# # References
#
# [Heterogeneous Graph Neural Network](https://dl.acm.org/doi/pdf/10.1145/3292500.3330961)
#
#
| graph neural networks/6_heterogeneous_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling Off-Axis Seamounts; Expediting insight into near-ridge mantle heterogeneity
# - Part Three - IN PROGRESS -
# - Workflow
# #### By: <NAME>
# ### I. Problem Statement
# #### In an age where seeing the unseen is commonplace, scientists still have surprisingly few constraints on mantle dynamics beneath our feet. Studies performed at mid-ocean ridges attempt to address this gap in knowledge by examining chemistry of basalts produced at ridge axes where distance to the mantle is minimal (often less than 1 km). Ideally, those basalts are as close to representing the mantle as a geochemist can access. However, at fast-spreading ridges a continuous thermal barrier allows for efficient mixing of mantle material prior to eruption, so materials collected on-axis (the most common site for sampling) only represent an average composition. To probe any heterogeneities in the mantle, ~300 basalt samples were collected from a long seamount chain perpendicular to the fast-spreading East Pacific Rise in an attempt to by-pass homogenization mixing and see into the mantle.
#
# #### To assess heterogeneity, fractional crystallization, source mixing, and mantle melting models are compared to the 300 basalt samples. However, the stages from inputing parameters into the melting program (alphaMELTs) to producing visualizations of the results are arduous and time-consuming. This results in week-long modeling sessions that are frequently inadequate and need adjusted and re-run. To expedite the melt model results, this study will incorporate the computing power of Python to read the alphaMELTs output results, process the data, feed it into a script which plots the results in real-time without all of the intermediate, time-consuming steps. This will effectively reduce the time required to visualize the results so I can immediately decide which parameters to adjust, quickly adjust those, and automate getting updated results.
# ### II. Import libraries and Upload seamount data
import pandas as pd # standard library for data analysis
import numpy as np # library for scientific computing
import matplotlib.pyplot as plt # library for plotting values and generating visualizations
import subprocess # library for executing an external program through Jupyter notebooks
from subprocess import call # use subprocess call to execute external program MELTs...
#it seems that Jupyter already has this capability by just using an exclamation point
Elements = pd.read_csv('Documents/EPR/Code/MajorTrace.csv') #read in seamount data file and set variable Elements for table
# ### III. Choose melting parameters
# * Edit batch file to reflect melting parameters for the program MELTs
# * Set variables for specifying titles, etc in the plots later so that labels accurately reflect the parameters modelled
# %load C:\Users\Molly\Documents\bin\batch.txt #adjust batch file to appropriate composition txt file, temp, pressure conditions
# %%write editedbatch.txt #save changes as the batch we apply later
# ### IV. Execute MELTs Program
# * Call batch file which executes MELTs and provides the right flags (f for environment, b for batch, p for output)
# (tells MELTs which environment file, batch file (the one adjusted above), and output location)
# * Produce an output in the form of a txt file (ends in traceint)
# +
# !C:/Users/Molly/Documents/bin/test.bat #exclamation point to call the batch which executes MELTs program
print('done') #since the program is finnicky and long, have Jupyter output 'done' when it is complete and successful
# -
# ### V. Upload Output Results as CSV
# * Remove the upper header rows
# * Convert to csv
# * Upload csv into Jupyter notebooks
Output_TE = pd.read('Documents/bin/Output.traceint') #read trace element file of MELTs output (an example for this part of the project)
# ### VI. Calculate Trace Element Ratios Normalized to Mantle --> create new columns
# * Use La/SmN formula and put calculated results for each row in an additional column in the table
# * Use Nb/La formula and put calculated results of each row in an additional column in the table
# did not get to this section because Anaconda crashed on my computer. Due to installation issues the project changed here
# ### VII. Plot Results of Model with Seamount Data for Visual Comparison
# * Plot La/SmN versus Nb/La showing 1. the seamount data as points, and 2. the model results in the form of a line
# * Include axes titles, labels, a figure title, and a legend
LaSmN_Data = Elements['La.SmN'].values #set variables for columns of seamount data
NbLa_Data = Elements['Nb.La'].values
LaSmN_Model = Output_TE['La/Sm_1D'].values #set variables for columns of modelled data
NbLa_Model = Output_TE['Nb/La_1D'].values
plt.plot(LaSmN_Data, NbLa_Data, 'ro', label = "Seamount Data") #plot the seamount data as red points, title for legend
plt.plot(LaSmN_Model, NbLa_Model, 'bl', label = "Model Results") #plot the model results as a blue line, title for legend
plt.title('Melting models') #title of the plot
plt.xlabel('La/SmN') #x-axis label
plt.ylabel('Nb/La') #y-axis label
plt.legend() #automatically produces a legend based on the two plotted series
plt.text(2,0.6, 'Depleted Depleted Mantle') #add text to the plot to describe the line and what model it represents
| SeamountGeochemistry_Workflow_Update.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + uuid="fb1a49fb-2775-43ed-ba80-cd439465770d"
import pandas as pd
from nltk.translate.bleu_score import sentence_bleu
references = ["吴承恩是著名文学家"] * 6
candidates = """
吴承恩是著名文学家
吴承恩是大作家
吴承恩是作曲家
吴承恩是运动员
吴承恩不是著名文学家
风马牛不相及
""".strip().split("\n")
bleu_score = []
for ref, cand in zip(references, candidates):
bleu_score.append(sentence_bleu([ref], cand))
bleu_score = pd.Series(bleu_score)
print(bleu_score)
bleurt_score = pd.Series([0.8896132111549377, 0.22806376218795776, 0.22379213571548462, 0.21101689338684082, 0.5763261318206787, -0.007796883583068848])
bleu_score - bleurt_score
| pretrain/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 3: Activation Functions
#
# 
#
# **Filled notebook:**
# [](https://github.com/phlippe/uvadlc_notebooks/blob/master/docs/tutorial_notebooks/tutorial3/Activation_Functions.ipynb)
# [](https://colab.research.google.com/github/phlippe/uvadlc_notebooks/blob/master/docs/tutorial_notebooks/tutorial3/Activation_Functions.ipynb)
# **Empty notebook:**
# [](https://github.com/phlippe/uvadlc_notebooks/blob/master/docs/tutorial_notebooks/tutorial3/Activation_Functions_empty.ipynb)
# [](https://colab.research.google.com/github/phlippe/uvadlc_notebooks/blob/master/docs/tutorial_notebooks/tutorial3/Activation_Functions_empty.ipynb)
# **Pre-trained models:**
# [](https://github.com/phlippe/saved_models/tree/main/tutorial3)
# [](https://drive.google.com/drive/folders/1sFpZUpDJVjiYEvIqISqfkFizfsTnPf4s?usp=sharing)
# In this tutorial, we will take a closer look at (popular) activation functions and investigate their effect on optimization properties in neural networks.
# Activation functions are a crucial part of deep learning models as they add the non-linearity to neural networks.
# There is a great variety of activation functions in the literature, and some are more beneficial than others.
# The goal of this tutorial is to show the importance of choosing a good activation function (and how to do so), and what problems might occur if we don't.
#
# Before we start, we import our standard libraries and set up basic functions:
# +
## Standard libraries
import os
import json
import math
import numpy as np
## Imports for plotting
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('svg', 'pdf') # For export
import seaborn as sns
sns.set()
## Progress bar
from tqdm.notebook import tqdm
## PyTorch
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data as data
import torch.optim as optim
# -
# We will define a function to set a seed on all libraries we might interact with in this tutorial (here numpy and torch). This allows us to make our training reproducible. However, note that in contrast to the CPU, the same seed on different GPU architectures can give different results. All models here have been trained on an NVIDIA GTX1080Ti, which is also the GPU type provided by the Lisa gpu_shared_course partition.
#
# Additionally, the following cell defines two paths: `DATASET_PATH` and `CHECKPOINT_PATH`. The dataset path is the directory where we will download datasets used in the notebooks. It is recommended to store all datasets from PyTorch in one joined directory to prevent duplicate downloads. The checkpoint path is the directory where we will store trained model weights and additional files. The needed files will be automatically downloaded. In case you are on Google Colab, it is recommended to change the directories to start from the current directory (i.e. remove `../` for both dataset and checkpoint path).
# +
# Path to the folder where the datasets are/should be downloaded (e.g. MNIST)
DATASET_PATH = "../data"
# Path to the folder where the pretrained models are saved
CHECKPOINT_PATH = "../saved_models/tutorial3"
# Function for setting the seed
def set_seed(seed):
np.random.seed(seed)
torch.manual_seed(seed)
if torch.cuda.is_available(): # GPU operation have separate seed
torch.cuda.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
set_seed(42)
# Additionally, some operations on a GPU are implemented stochastic for efficiency
# We want to ensure that all operations are deterministic on GPU (if used) for reproducibility
torch.backends.cudnn.determinstic = True
torch.backends.cudnn.benchmark = False
# Fetching the device that will be used throughout this notebook
device = torch.device("cpu") if not torch.cuda.is_available() else torch.device("cuda:0")
print("Using device", device)
# -
# The following cell downloads all pretrained models we will use in this notebook. The files are stored on a separate [repository](https://github.com/phlippe/saved_models) to reduce the size of the notebook repository, especially for building the documentation on ReadTheDocs. In case the download below fails, you can download the models from a [Google Drive folder](https://drive.google.com/drive/folders/1sFpZUpDJVjiYEvIqISqfkFizfsTnPf4s?usp=sharing). Please let me (Phillip) know if an error occurs so it can be fixed for all students.
# +
import urllib.request
from urllib.error import HTTPError
# Github URL where saved models are stored for this tutorial
base_url = "https://raw.githubusercontent.com/phlippe/saved_models/main/tutorial3/"
# Files to download
pretrained_files = ["FashionMNIST_elu.config", "FashionMNIST_elu.tar",
"FashionMNIST_leakyrelu.config", "FashionMNIST_leakyrelu.tar",
"FashionMNIST_relu.config", "FashionMNIST_relu.tar",
"FashionMNIST_sigmoid.config", "FashionMNIST_sigmoid.tar",
"FashionMNIST_swish.config", "FashionMNIST_swish.tar",
"FashionMNIST_tanh.config", "FashionMNIST_tanh.tar"]
# Create checkpoint path if it doesn't exist yet
os.makedirs(CHECKPOINT_PATH, exist_ok=True)
# For each file, check whether it already exists. If not, try downloading it.
for file_name in pretrained_files:
file_path = os.path.join(CHECKPOINT_PATH, file_name)
if not os.path.isfile(file_path):
file_url = base_url + file_name
print("Downloading %s..." % file_url)
try:
urllib.request.urlretrieve(file_url, file_path)
except HTTPError as e:
print("Something went wrong. Please try to download the file from the GDrive folder, or contact the author with the full output including the following error:\n", e)
# -
# ## Common activation functions
# As a first step, we will implement some common activation functions by ourselves. Of course, most of them can also be found in the `torch.nn` package (see the [documentation](https://pytorch.org/docs/stable/nn.html#non-linear-activations-weighted-sum-nonlinearity) for an overview).
# However, we'll write our own functions here for a better understanding and insights.
#
# For an easier time of comparing various activation functions, we start with defining a base class from which all our future modules will inherit:
class ActivationFunction(nn.Module):
def __init__(self):
super().__init__()
self.name = self.__class__.__name__
self.config = {"name": self.name}
# Every activation function will be an `nn.Module` so that we can integrate them nicely in a network. We will use the `config` dictionary to store adjustable parameters for some activation functions.
#
# Next, we implement two of the "oldest" activation functions that are still commonly used for various tasks: sigmoid and tanh.
# Both the sigmoid and tanh activation can be also found as PyTorch functions (`torch.sigmoid`, `torch.tanh`) or as modules (`nn.Sigmoid`, `nn.Tanh`).
# Here, we implement them by hand:
# +
##############################
class Sigmoid(ActivationFunction):
def forward(self, x):
## TODO: Implement sigmoid
return ...
##############################
class Tanh(ActivationFunction):
def forward(self, x):
## TODO: Implement tanh
return ...
##############################
# -
# Another popular activation function that has allowed the training of deeper networks, is the Rectified Linear Unit (ReLU).
# Despite its simplicity of being a piecewise linear function, ReLU has one major benefit compared to sigmoid and tanh: a strong, stable gradient for a large range of values.
# Based on this idea, a lot of variations of ReLU have been proposed, of which we will implement the following three: LeakyReLU, ELU, and Swish.
# LeakyReLU replaces the zero settings in the negative part with a smaller slope to allow gradients to flow also in this part of the input.
# Similarly, ELU replaces the negative part with an exponential decay.
# The third, most recently proposed activation function is Swish, which is actually the result of a large experiment with the purpose of finding the "optimal" activation function.
# Compared to the other activation functions, Swish is both smooth and non-monotonic (i.e. contains a change of sign in the gradient).
# This has been shown to prevent dead neurons as in standard ReLU activation, especially for deep networks.
# If interested, a more detailed discussion of the benefits of Swish can be found in [this paper](https://arxiv.org/abs/1710.05941) [1].
#
# Let's implement the four activation functions below:
# +
##############################
class ReLU(ActivationFunction):
def forward(self, x):
## TODO: Implement ReLU
return ...
##############################
class LeakyReLU(ActivationFunction):
def __init__(self, alpha=0.1):
super().__init__()
self.config["alpha"] = alpha
def forward(self, x):
## TODO: Implement LeakyReLU with alpha as negative slope
return ...
##############################
class ELU(ActivationFunction):
def forward(self, x):
## TODO: Implement ELU
return ...
##############################
class Swish(ActivationFunction):
def forward(self, x):
## TODO: Implement Swish
return ...
##############################
# -
# For later usage, we summarize all our activation functions in a dictionary mapping the name to the class object. In case you implement a new activation function by yourself, add it here to include it in future comparisons as well:
act_fn_by_name = {
"sigmoid": Sigmoid,
"tanh": Tanh,
"relu": ReLU,
"leakyrelu": LeakyReLU,
"elu": ELU,
"swish": Swish
}
# ### Visualizing activation functions
#
# To get an idea of what each activation function actually does, we will visualize them in the following.
# Next to the actual activation value, the gradient of the function is an important aspect as it is crucial for optimizing the neural network.
# PyTorch allows us to compute the gradients simply by calling the `backward` function:
def get_grads(act_fn, x):
"""
Computes the gradients of an activation function at specified positions.
Inputs:
act_fn - An object of the class "ActivationFunction" with an implemented forward pass.
x - 1D input tensor.
Output:
A tensor with the same size of x containing the gradients of act_fn at x.
"""
x = x.clone()
## TODO: Calculate and return the gradients of the activation function for each point of x
return ...
# Now we can visualize all our activation functions including their gradients:
# +
def vis_act_fn(act_fn, ax, x):
# Run activation function
y = act_fn(x)
y_grads = get_grads(act_fn, x)
# Push x, y and gradients back to cpu for plotting
x, y, y_grads = x.cpu().numpy(), y.cpu().numpy(), y_grads.cpu().numpy()
## Plotting
ax.plot(x, y, linewidth=2, label="ActFn")
ax.plot(x, y_grads, linewidth=2, label="Gradient")
ax.set_title(act_fn.name)
ax.legend()
ax.set_ylim(-1.5, x.max())
# Add activation functions if wanted
act_fns = [act_fn() for act_fn in act_fn_by_name.values()]
x = torch.linspace(-5, 5, 1000) # Range on which we want to visualize the activation functions
## Plotting
rows = math.ceil(len(act_fns)/2.0)
fig, ax = plt.subplots(rows, 2, figsize=(8, rows*4))
for i, act_fn in enumerate(act_fns):
vis_act_fn(act_fn, ax[divmod(i,2)], x)
fig.subplots_adjust(hspace=0.3)
plt.show()
# -
# ## Analysing the effect of activation functions
# After implementing and visualizing the activation functions, we are aiming to gain insights into their effect.
# We do this by using a simple neural network trained on [FashionMNIST](https://github.com/zalandoresearch/fashion-mnist) and examine various aspects of the model, including the performance and gradient flow.
# ### Setup
# Firstly, let's set up a neural network. The chosen network views the images as 1D tensors and pushes them through a sequence of linear layers and a specified activation function. Feel free to experiment with other network architectures.
class BaseNetwork(nn.Module):
def __init__(self, act_fn, input_size=784, num_classes=10, hidden_sizes=[512, 256, 256, 128]):
"""
Inputs:
act_fn - Object of the activation function that should be used as non-linearity in the network.
input_size - Size of the input images in pixels
num_classes - Number of classes we want to predict
hidden_sizes - A list of integers specifying the hidden layer sizes in the NN
"""
super().__init__()
# Create the network based on the specified hidden sizes
layers = []
layer_sizes = [input_size] + hidden_sizes
for layer_index in range(1, len(layer_sizes)):
layers += [nn.Linear(layer_sizes[layer_index-1], layer_sizes[layer_index]),
act_fn]
layers += [nn.Linear(layer_sizes[-1], num_classes)]
self.layers = nn.Sequential(*layers) # nn.Sequential summarizes a list of modules into a single module, applying them in sequence
# We store all hyperparameters in a dictionary for saving and loading of the model
self.config = {"act_fn": act_fn.config, "input_size": input_size, "num_classes": num_classes, "hidden_sizes": hidden_sizes}
def forward(self, x):
## TODO: Transform images into vector + batch dimension, and apply layers on it
return ...
# We also add functions for loading and saving the model. The hyperparameters are stored in a configuration file (simple json file):
# +
def _get_config_file(model_path, model_name):
# Name of the file for storing hyperparameter details
return os.path.join(model_path, model_name + ".config")
def _get_model_file(model_path, model_name):
# Name of the file for storing network parameters
return os.path.join(model_path, model_name + ".tar")
def load_model(model_path, model_name, net=None):
"""
Loads a saved model from disk.
Inputs:
model_path - Path of the checkpoint directory
model_name - Name of the model (str)
net - (Optional) If given, the state dict is loaded into this model. Otherwise, a new model is created.
"""
config_file, model_file = _get_config_file(model_path, model_name), _get_model_file(model_path, model_name)
assert os.path.isfile(config_file), "Could not find the config file \"%s\". Are you sure this is the correct path and you have your model config stored here?" % (config_file)
assert os.path.isfile(model_file), "Could not find the model file \"%s\". Are you sure this is the correct path and you have your model stored here?" % (model_file)
with open(config_file, "r") as f:
config_dict = json.load(f)
if net is None:
act_fn_name = config_dict["act_fn"].pop("name").lower()
act_fn = act_fn_by_name[act_fn_name](**config_dict.pop("act_fn"))
net = BaseNetwork(act_fn=act_fn, **config_dict)
net.load_state_dict(torch.load(model_file, map_location=device))
return net
def save_model(model, model_path, model_name):
"""
Given a model, we save the state_dict and hyperparameters.
Inputs:
model - Network object to save parameters from
model_path - Path of the checkpoint directory
model_name - Name of the model (str)
"""
config_dict = model.config
os.makedirs(model_path, exist_ok=True)
config_file, model_file = _get_config_file(model_path, model_name), _get_model_file(model_path, model_name)
with open(config_file, "w") as f:
json.dump(config_dict, f)
torch.save(model.state_dict(), model_file)
# -
# We also set up the dataset we want to train it on, namely [FashionMNIST](https://github.com/zalandoresearch/fashion-mnist). FashionMNIST is a more complex version of MNIST and contains black-and-white images of clothes instead of digits. The 10 classes include trousers, coats, shoes, bags and more. To load this dataset, we will make use of yet another PyTorch package, namely `torchvision` ([documentation](https://pytorch.org/docs/stable/torchvision/index.html)). The `torchvision` package consists of popular datasets, model architectures, and common image transformations for computer vision. We will use the package for many of the notebooks in this course to simplify our dataset handling.
#
# Let's load the dataset below, and visualize a few images to get an impression of the data.
# +
import torchvision
from torchvision.datasets import FashionMNIST
from torchvision import transforms
# Transformations applied on each image => first make them a tensor, then normalize them in the range -1 to 1
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
# Loading the training dataset. We need to split it into a training and validation part
train_dataset = FashionMNIST(root=DATASET_PATH, train=True, transform=transform, download=True)
train_set, val_set = torch.utils.data.random_split(train_dataset, [50000, 10000])
# Loading the test set
test_set = FashionMNIST(root=DATASET_PATH, train=False, transform=transform, download=True)
# We define a set of data loaders that we can use for various purposes later.
# Note that for actually training a model, we will use different data loaders
# with a lower batch size.
train_loader = data.DataLoader(train_set, batch_size=1024, shuffle=True, drop_last=False)
val_loader = data.DataLoader(val_set, batch_size=1024, shuffle=False, drop_last=False)
test_loader = data.DataLoader(test_set, batch_size=1024, shuffle=False, drop_last=False)
# +
exmp_imgs = [train_set[i][0] for i in range(16)]
# Organize the images into a grid for nicer visualization
img_grid = torchvision.utils.make_grid(torch.stack(exmp_imgs, dim=0), nrow=4, normalize=True, pad_value=0.5)
img_grid = img_grid.permute(1, 2, 0)
plt.figure(figsize=(8,8))
plt.title("FashionMNIST examples")
plt.imshow(img_grid)
plt.axis('off')
plt.show()
plt.close()
# -
# ### Visualizing the gradient flow after initialization
#
# As mentioned previously, one important aspect of activation functions is how they propagate gradients through the network. Imagine we have a very deep neural network with more than 50 layers. The gradients for the input layer, i.e. the very first layer, have passed >50 times the activation function, but we still want them to be of a reasonable size. If the gradient through the activation function is (in expectation) considerably smaller than 1, our gradients will vanish until they reach the input layer. If the gradient through the activation function is larger than 1, the gradients exponentially increase and might explode.
#
# To get a feeling of how every activation function influences the gradients, we can look at a freshly initialized network and measure the gradients for each parameter for a batch of 256 images:
def visualize_gradients(net, color="C0"):
"""
Inputs:
net - Object of class BaseNetwork
color - Color in which we want to visualize the histogram (for easier separation of activation functions)
"""
net.eval()
small_loader = data.DataLoader(train_set, batch_size=256, shuffle=False)
imgs, labels = next(iter(small_loader))
imgs, labels = imgs.to(device), labels.to(device)
net.zero_grad()
# TODO: Pass one batch through the network, and calculate the gradients for the weights
...
# We limit our visualization to the weight parameters and exclude the bias to reduce the number of plots
grads = {name: params.grad.data.view(-1).cpu().clone().numpy() for name, params in net.named_parameters() if "weight" in name}
net.zero_grad()
## Plotting
columns = len(grads)
fig, ax = plt.subplots(1, columns, figsize=(columns*3.5, 2.5))
fig_index = 0
for key in grads:
key_ax = ax[fig_index%columns]
sns.histplot(data=grads[key], bins=30, ax=key_ax, color=color, kde=True)
key_ax.set_title("%s" % key)
key_ax.set_xlabel("Grad magnitude")
fig_index += 1
fig.suptitle("Gradient magnitude distribution for activation function %s" % (net.config["act_fn"]["name"]), fontsize=14, y=1.05)
fig.subplots_adjust(wspace=0.45)
plt.show()
plt.close()
# Seaborn prints warnings if histogram has small values. We can ignore them for now
import warnings
warnings.filterwarnings('ignore')
## Create a plot for every activation function
for i, act_fn_name in enumerate(act_fn_by_name):
set_seed(42) # Setting the seed ensures that we have the same weight initialization for each activation function
act_fn = act_fn_by_name[act_fn_name]()
net_actfn = BaseNetwork(act_fn=act_fn).to(device)
visualize_gradients(net_actfn, color="C%i"%i)
# The sigmoid activation function shows a clearly undesirable behavior. While the gradients for the output layer are very large with up to 0.1, the input layer has the lowest gradient norm across all activation functions with only 1e-5. This is due to its small maximum gradient of 1/4, and finding a suitable learning rate across all layers is not possible in this setup.
# All the other activation functions show to have similar gradient norms across all layers. Interestingly, the ReLU activation has a spike around 0 which is caused by its zero-part on the left, and dead neurons (we will take a closer look at this later on).
#
# Note that additionally to the activation, the initialization of the weight parameters can be crucial. By default, PyTorch uses the [Kaiming](https://pytorch.org/docs/stable/nn.init.html#torch.nn.init.kaiming_uniform_) initialization for linear layers optimized for Tanh activations. In Tutorial 4, we will take a closer look at initialization, but assume for now that the Kaiming initialization works for all activation functions reasonably well.
# ### Training a model
#
# Next, we want to train our model with different activation functions on FashionMNIST and compare the gained performance. All in all, our final goal is to achieve the best possible performance on a dataset of our choice.
# Therefore, we write a training loop in the next cell including a validation after every epoch and a final test on the best model:
# +
def train_model(net, model_name, max_epochs=50, patience=7, batch_size=256, overwrite=False):
"""
Train a model on the training set of FashionMNIST
Inputs:
net - Object of BaseNetwork
model_name - (str) Name of the model, used for creating the checkpoint names
max_epochs - Number of epochs we want to (maximally) train for
patience - If the performance on the validation set has not improved for #patience epochs, we stop training early
batch_size - Size of batches used in training
overwrite - Determines how to handle the case when there already exists a checkpoint. If True, it will be overwritten. Otherwise, we skip training.
"""
file_exists = os.path.isfile(_get_model_file(CHECKPOINT_PATH, model_name))
if file_exists and not overwrite:
print("Model file already exists. Skipping training...")
else:
if file_exists:
print("Model file exists, but will be overwritten...")
# Defining optimizer, loss and data loader
optimizer = optim.SGD(net.parameters(), lr=1e-2, momentum=0.9) # Default parameters, feel free to change
loss_module = nn.CrossEntropyLoss()
train_loader_local = data.DataLoader(train_set, batch_size=batch_size, shuffle=True, drop_last=True, pin_memory=True)
val_scores = []
best_val_epoch = -1
for epoch in range(max_epochs):
##############
## Training ##
##############
net.train()
true_preds, count = 0., 0
for imgs, labels in tqdm(train_loader, desc="Epoch %i" % (epoch+1), leave=False):
## TODO: Take an optimization step on the batch. Use "preds" for naming the outputs of the networks
...
# Record statistics during training
true_preds += (preds.argmax(dim=-1) == labels).sum()
count += labels.shape[0]
train_acc = true_preds / count
################
## Validation ##
################
val_acc = test_model(net, val_loader)
val_scores.append(val_acc)
print("[Epoch %2i] Training accuracy: %05.2f%%, Validation accuracy: %05.2f%%" % (epoch+1, train_acc*100.0, val_acc*100.0))
if len(val_scores) == 1 or val_acc > val_scores[best_val_epoch]:
print("\t (New best performance, saving model...)")
save_model(net, CHECKPOINT_PATH, model_name)
best_val_epoch = epoch
elif best_val_epoch <= epoch - patience:
print("Early stopping due to no improvement over the last %i epochs" % (patience))
break
# Plot a curve of the validation accuracy
plt.plot([i for i in range(1,len(val_scores)+1)], val_scores)
plt.xlabel("Epochs")
plt.ylabel("Validation accuracy")
plt.title("Validation performance of %s" % model_name)
plt.show()
plt.close()
load_model(CHECKPOINT_PATH, model_name, net=net)
test_acc = test_model(net, test_loader)
print((" Test accuracy: %4.2f%% " % (test_acc*100.0)).center(50, "=")+"\n")
return test_acc
def test_model(net, data_loader):
"""
Test a model on a specified dataset.
Inputs:
net - Trained model of type BaseNetwork
data_loader - DataLoader object of the dataset to test on (validation or test)
"""
net.eval()
true_preds, count = 0., 0
for imgs, labels in data_loader:
## TODO: Predict the labels for the test data, and record number of correct predictions in "true_preds",
## and number of predictions overall in "count"
...
test_acc = true_preds / count
return test_acc
# -
# We train one model for each activation function. We recommend using the pretrained models to save time if you are running this notebook on CPU.
for act_fn_name in act_fn_by_name:
print("Training BaseNetwork with %s activation..." % act_fn_name)
set_seed(42)
act_fn = act_fn_by_name[act_fn_name]()
net_actfn = BaseNetwork(act_fn=act_fn).to(device)
train_model(net_actfn, "FashionMNIST_%s" % act_fn_name, overwrite=False)
# Not surprisingly, the model using the sigmoid activation function shows to fail and does not improve upon random performance (10 classes => 1/10 for random chance).
#
# All the other activation functions gain similar performance.
# To have a more accurate conclusion, we would have to train the models for multiple seeds and look at the averages.
# However, the "optimal" activation function also depends on many other factors (hidden sizes, number of layers, type of layers, task, dataset, optimizer, learning rate, etc.) so that a thorough grid search would not be useful in our case.
# In the literature, activation functions that have shown to work well with deep networks are all types of ReLU functions we experiment with here, with small gains for specific activation functions in specific networks.
# ### Visualizing the activation distribution
# After we have trained the models, we can look at the actual activation values that find inside the model. For instance, how many neurons are set to zero in ReLU? Where do we find most values in Tanh?
# To answer these questions, we can write a simple function which takes a trained model, applies it to a batch of images, and plots the histogram of the activations inside the network:
def visualize_activations(net, color="C0"):
activations = {}
net.eval()
small_loader = data.DataLoader(train_set, batch_size=1024)
imgs, labels = next(iter(small_loader))
with torch.no_grad():
layer_index = 0
imgs = imgs.to(device)
imgs = imgs.view(imgs.size(0), -1)
# We need to manually loop through the layers to save all activations
for layer_index, layer in enumerate(net.layers[:-1]):
imgs = layer(imgs)
activations[layer_index] = imgs.view(-1).cpu().numpy()
## Plotting
columns = 4
rows = math.ceil(len(activations)/columns)
fig, ax = plt.subplots(rows, columns, figsize=(columns*2.7, rows*2.5))
fig_index = 0
for key in activations:
key_ax = ax[fig_index//columns][fig_index%columns]
sns.histplot(data=activations[key], bins=50, ax=key_ax, color=color, kde=True, stat="density")
key_ax.set_title("Layer %i - %s" % (key, net.layers[key].__class__.__name__))
fig_index += 1
fig.suptitle("Activation distribution for activation function %s" % (net.config["act_fn"]["name"]), fontsize=14)
fig.subplots_adjust(hspace=0.4, wspace=0.4)
plt.show()
plt.close()
for i, act_fn_name in enumerate(act_fn_by_name):
net_actfn = load_model(model_path=CHECKPOINT_PATH, model_name="FashionMNIST_%s" % act_fn_name).to(device)
visualize_activations(net_actfn, color="C%i" % i)
# As the model with sigmoid activation was not able to train properly, the activations are also less informative and all gathered around 0.5 (the activation at input 0).
#
# The tanh shows a more diverse behavior. While for the input layer we experience a larger amount of neurons to be close to -1 and 1, where the gradients are close to zero, the activations in the two consecutive layers are closer to zero. This is probably because the input layers look for specific features in the input image, and the consecutive layers combine those together. The activations for the last layer are again more biased to the extreme points because the classification layer can be seen as a weighted average of those values (the gradients push the activations to those extremes).
#
# The ReLU has a strong peak at 0, as we initially expected. The effect of having no gradients for negative values is that the network does not have a Gaussian-like distribution after the linear layers, but a longer tail towards the positive values.
# The LeakyReLU shows a very similar behavior while ELU follows again a more Gaussian-like distribution.
# The Swish activation seems to lie in between, although it is worth noting that Swish uses significantly higher values than other activation functions (up to 20).
#
# As all activation functions show slightly different behavior although obtaining similar performance for our simple network, it becomes apparent that the selection of the "optimal" activation function really depends on many factors, and is not the same for all possible networks.
# ### Finding dead neurons in ReLU networks
# One known drawback of the ReLU activation is the occurrence of "dead neurons", i.e. neurons with no gradient for any training input.
# The issue of dead neurons is that as no gradient is provided for the layer, we cannot train the parameters of this neuron in the previous layer to obtain output values besides zero.
# For dead neurons to happen, the output value of a specific neuron of the linear layer before the ReLU has to be negative for all input images.
# Considering the large number of neurons we have in a neural network, it is not unlikely for this to happen.
#
# To get a better understanding of how much of a problem this is, and when we need to be careful, we will measure how many dead neurons different networks have. For this, we implement a function which runs the network on the whole training set and records whether a neuron is exactly 0 for all data points or not:
def measure_number_dead_neurons(net):
# For each neuron, we create a boolean variable initially set to 1. If it has an activation unequals 0 at any time,
# we set this variable to 0. After running through the whole training set, only dead neurons will have a 1.
neurons_dead = [
torch.ones(layer.weight.shape[0], device=device, dtype=torch.bool) for layer in net.layers[:-1] if isinstance(layer, nn.Linear)
] # Same shapes as hidden size in BaseNetwork
net.eval()
with torch.no_grad():
for imgs, labels in tqdm(train_loader, leave=False): # Run through whole training set
layer_index = 0
imgs = imgs.to(device)
imgs = imgs.view(imgs.size(0), -1)
for layer in net.layers[:-1]:
imgs = layer(imgs)
if isinstance(layer, ActivationFunction):
## TODO: Update the buffer neurons_dead[layer_index] with the activations on the new batch
neurons_dead[layer_index] = ...
layer_index += 1
number_neurons_dead = [t.sum().item() for t in neurons_dead]
print("Number of dead neurons:", number_neurons_dead)
print("In percentage:", ", ".join(["%4.2f%%" % (100.0 * num_dead / tens.shape[0]) for tens, num_dead in zip(neurons_dead, number_neurons_dead)]))
# First, we can measure the number of dead neurons for an untrained network:
set_seed(42)
net_relu = BaseNetwork(act_fn=ReLU()).to(device)
measure_number_dead_neurons(net_relu)
# We see that only a minor amount of neurons are dead, but that they increase with the depth of the layer.
# However, this is not a problem for the small number of dead neurons we have as the input to later layers is changed due to updates to the weights of previous layers. Therefore, dead neurons in later layers can potentially become "alive"/active again.
#
# How does this look like for a trained network (with the same initialization)?
net_relu = load_model(model_path=CHECKPOINT_PATH, model_name="FashionMNIST_relu").to(device)
measure_number_dead_neurons(net_relu)
# The number of dead neurons indeed decreased in the later layers. However, it should be noted that dead neurons are especially problematic in the input layer. As the input does not change over epochs (the training set is kept as it is), training the network cannot turn those neurons back active. Still, the input data has usually a sufficiently high standard deviation to reduce the risk of dead neurons.
#
# Finally, we check how the number of dead neurons behaves with increasing layer depth. For instance, let's take the following 10-layer neural network:
set_seed(42)
net_relu = BaseNetwork(act_fn=ReLU(), hidden_sizes=[256, 256, 256, 256, 256, 128, 128, 128, 128, 128]).to(device)
measure_number_dead_neurons(net_relu)
# The number of dead neurons is significantly higher than before which harms the gradient flow especially in the first iterations. For instance, more than 56% of the neurons in the pre-last layer are dead which creates a considerable bottleneck.
# Hence, it is advisible to use other nonlinearities like Swish for very deep networks.
# ## Conclusion
#
# In this notebook, we have reviewed a set of six activation functions (sigmoid, tanh, ReLU, LeakyReLU, ELU, and Swish) in neural networks, and discussed how they influence the gradient distribution across layers. Sigmoid tends to fail deep neural networks as the highest gradient it provides is 0.25 leading to vanishing gradients in early layers. All ReLU-based activation functions have shown to perform well, and besides the original ReLU, do not have the issue of dead neurons. When implementing your own neural network, it is recommended to start with a ReLU-based network and select the specific activation function based on the properties of the network.
# ## References
#
# [1] Ramachandran, Prajit, <NAME>, and <NAME>. "Searching for activation functions." arXiv preprint arXiv:1710.05941 (2017). [Paper link](https://arxiv.org/abs/1710.05941)
| docs/tutorial_notebooks/tutorial3/Activation_Functions_empty.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Version 1.2
import logging, os, re, datetime
from IPython.display import display
from arcgis.gis import GIS
''' ********************** SCRIPT CONFIGURATION START ********************** '''
#What is the ID of the Feature Layer you want to download attachments from?
FeatureLayerId = '092d075f4b3a40f78cf1329b20b0d5e7'
#What are your ArcGIS Enterprise/ArcGIS Online credentials? This is case sensitive.
PortalUserName = ''
PortalPassword = ''
PortalUrl = 'https://www.arcgis.com'
#Where do you want your attachments stored?
SaveAttachmentsTo = 'C:\ScriptDownloads'
SaveLogsTo = 'Logging'
#How do you want your attachments stored? Options are GroupedFolder and IndividualFolder
#GroupedFolder - Attachments from every feature in each layer is stored in the same folder - attachments are renamed in the format OBJECTID-ATTACHMENTID-OriginalFileName
#IndividualFolder - A new folder is created for each OBJECTID, and associated attachments are stored in that folder - attachments are renamed in the format ATTACHMENTID-OriginalFileName
AttachmentStorage = 'GroupedFolder'
#Set to False if ArcGIS Enterprise cert is not valid
PortalCertVerification = True
#Setup logging - levels are DEBUG,INFO,WARNING,ERROR,CRITICAL
logging.basicConfig(level=logging.INFO)
''' ********************** SCRIPT CONFIGURATION END ********************** '''
#https://stackoverflow.com/questions/273192/how-can-i-create-a-directory-if-it-does-not-exist
def createFolder(folderPath):
if not os.path.exists(folderPath):
try:
os.makedirs(folderPath)
except OSError as e:
if e.errno != errno.EEXIST:
raise
def renameFile(currentAttachmentPath, newAttachmentPath):
#Rename file - ensure new attachment path does not exist already
if not os.path.exists(newAttachmentPath):
os.rename(currentAttachmentPath, newAttachmentPath)
logger.info('{} being renamed as {}'.format(currentAttachmentPath, newAttachmentPath))
else:
logger.warning('Not able to rename {} as {} because file already exists. Removing {}'.format(currentAttachmentPath, newAttachmentPath, currentAttachmentPath))
os.remove(currentAttachmentPath)
#Create specified folder if it does not exist already
createFolder(SaveAttachmentsTo)
createFolder(SaveLogsTo)
#Logging level specified in script configuration
logger = logging.getLogger(__name__)
logFileName = datetime.datetime.now().strftime('%Y-%m-%d %H-%M-%S')
fileHandler = logging.handlers.RotatingFileHandler('{}/{}.log'.format(SaveLogsTo, logFileName), maxBytes=100000, backupCount=5)
formatter = logging.Formatter('%(asctime)s %(levelname)s %(relativeCreated)d \n%(filename)s %(module)s %(funcName)s %(lineno)d \n%(message)s\n')
fileHandler.setFormatter(formatter)
logger.addHandler(fileHandler)
logger.info('Script Starting at {}'.format(str(datetime.datetime.now())))
#Connect to GIS, and get Feature Layer information
if PortalUserName == '' and PortalPassword == '':
gis = GIS()
else:
gis = GIS(PortalUrl, PortalUserName, PortalPassword, verify_cert=PortalCertVerification)
downloadCounter = 0
nonDownloadCounter = 0
downloadSizeCounter = 0
itemObject = gis.content.get(FeatureLayerId)
logger.info('Iterating through layers in Feature Layer "{}"'.format(itemObject.name))
display(itemObject)
#Loop through layers in Feature Layer
for i in range(len(itemObject.layers)):
featureLayer = itemObject.layers[i]
#Skip layer if attachments are not enabled
if featureLayer.properties.hasAttachments == True:
#Remove any characters from feature layer name that may cause problems and ensure it's unique...
featureLayerName = '{}-{}'.format(str(i), re.sub(r'[^A-Za-z0-9]+', '', featureLayer.properties.name))
featureLayerFolder = SaveAttachmentsTo + r'\\' + featureLayerName
createFolder(featureLayerFolder)
#Query to get list of object ids in layer
featureObjectIds = featureLayer.query(where='1=1', return_ids_only=True)
#Provide some updates to user...
logger.info('Time: {}'.format(str(datetime.datetime.now())))
logger.info('Currently looping through feature attachments in layer {} of {}: storing in folder named "{}"'.format(str(i + 1), str(len(itemObject.layers)), featureLayerName))
logger.info('There are {} features to iterate in this layer'.format(str(len(featureObjectIds['objectIds']))))
#Loop through features in layer
emptyAttachments = 0
for j in range(len(featureObjectIds['objectIds'])):
currentObjectId = featureObjectIds['objectIds'][j]
currentObjectIdAttachments = featureLayer.attachments.get_list(oid=currentObjectId)
if len(currentObjectIdAttachments) > 0:
#Loop through feature attachments and download to appropriate folder
for k in range(len(currentObjectIdAttachments)):
attachmentId = currentObjectIdAttachments[k]['id']
attachmentName = currentObjectIdAttachments[k]['name']
attachmentSize = currentObjectIdAttachments[k]['size']
if AttachmentStorage == 'IndividualFolder':
currentFolder = featureLayerFolder + r'\\' + str(currentObjectId)
#Create a folder for attachments
createFolder(currentFolder)
fileName = '{}-{}'.format(attachmentId, attachmentName)
newAttachmentPath = '{}\\{}'.format(currentFolder, fileName)
if not os.path.isfile(newAttachmentPath):
logger.info('The size of the current attachment being downloaded is {}MB'.format((attachmentSize/1000000)))
currentAttachmentPath = featureLayer.attachments.download(oid=currentObjectId, attachment_id=attachmentId, save_path=currentFolder)
#Rename to ensure file name is unique
renameFile(currentAttachmentPath, newAttachmentPath)
downloadCounter += 1
downloadSizeCounter += attachmentSize
else:
logger.info('File {} already exists. Not downloading again!'.format(newAttachmentPath))
nonDownloadCounter += 1
elif AttachmentStorage == 'GroupedFolder':
fileName = '{}-{}-{}'.format(currentObjectId, attachmentId, attachmentName)
newAttachmentPath = '{}\\{}'.format(featureLayerFolder, fileName)
if not os.path.isfile(newAttachmentPath):
logger.info('The size of the current attachment being downloaded is {}MB'.format((attachmentSize/1000000)))
currentAttachmentPath = featureLayer.attachments.download(oid=currentObjectId, attachment_id=attachmentId, save_path=featureLayerFolder)
#Rename to ensure file name is unique
renameFile(currentAttachmentPath, newAttachmentPath)
downloadCounter += 1
downloadSizeCounter += attachmentSize
else:
logger.info('File {} already exists. Not downloading again!'.format(newAttachmentPath))
nonDownloadCounter += 1
else:
logger.error('AttachmentStorage option not valid: {}. Valid options are IndividualFolder and GroupedFolder'.format(AttachmentStorage))
else:
emptyAttachments += 1
logger.info('{} of these features do not contain attachments'.format(str(emptyAttachments)))
else:
logger.info('Layer {} does not have attachments enabled'.format(featureLayer.properties.name))
logger.info('Summary: {} new files have been downloaded totalling {}MB in size'.format(downloadCounter, (downloadSizeCounter/1000000)))
logger.info('Summary: {} attachments already existed so were not downloaded again'.format(nonDownloadCounter))
| python/general-python/download-attachments/DownloadAttachments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
import os
os.listdir("../Spam Classifier /smsspamcollection")
# +
# Dataset is taken from UCI Machine Learning Repository.
data = pd.read_table("../Spam Classifier /smsspamcollection/SMSSpamCollection",header=None)
data.head()
# -
data.columns = ["label", "message"]
data.head()
data.describe()
data.label.value_counts()
# +
label_dict = {
"ham" : 0,
"spam" : 1
}
data.label = data.label.map(label_dict)
data.head()
# -
# <hr>
#
# ## Get bag of words
#
# <hr>
# #### Order does not matter in bag of words.
# #### By default, it removes punctuation marks, treat them as delimters, convert all words to lower case.
# +
from sklearn.feature_extraction.text import CountVectorizer
count_vector = CountVectorizer()
count_vector
# -
# <hr>
#
# ## Sample data
#
# <hr>
# +
documents = [
"Are you bored?",
"Do ypu want to earn BILLIONS?",
"How are you you?"
]
count_vector.fit(documents)
count_vector.get_feature_names()
# -
doc_array = count_vector.transform(documents).toarray()
doc_array
# <hr>
#
# ## Make a dataframe out of document array.
#
# <hr>
# +
doc_df = pd.DataFrame(doc_array, columns = count_vector.get_feature_names())
doc_df.head()
# -
# <hr>
#
# ## Splitting data
#
# <hr>
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data["message"], data["label"], random_state=0, test_size=0.2)
# +
count_vector = CountVectorizer(stop_words="english")
train_data = count_vector.fit_transform(X_train)
test_data = count_vector.transform(X_test)
# -
# <hr>
#
# ## Building model
#
# <hr>
# ##### We choose to use multinomail Naive Bayesian classifier as we have selected features based on bag of words.
# ##### Multinomial Bayesian classifier is suitable for discrete features.
# ##### Gaussian(normal distribution) Naive Bayesian is suitable for continuous data.
# +
from sklearn.naive_bayes import MultinomialNB
naive_bayes = MultinomialNB()
naive_bayes.fit(train_data, y_train)
# -
y_pred = naive_bayes.predict(test_data)
y_pred
# +
from sklearn.metrics import accuracy_score, precision_score, confusion_matrix, precision_recall_fscore_support, classification_report
print("Accuracy Score : ", accuracy_score(y_test, y_pred))
print("Precision Score : ", precision_score(y_test, y_pred))
print("Precision, Recall, Fscore, Support : \n", precision_recall_fscore_support(y_test, y_pred))
print("Classification Report : \n", classification_report(y_test, y_pred))
print("Confusion Matrix : \n", confusion_matrix(y_test, y_pred))
# -
| Spam Classifier using Naive Bayesian Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# +
import json
from collections import Counter
from keras.models import Model
from keras.layers import Embedding, Input, Reshape
from keras.layers.merge import Dot
from sklearn.linear_model import LinearRegression
import numpy as np
import random
from sklearn import svm
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# -
with open('data/wp_movies_10k.ndjson') as fin:
movies = [json.loads(l) for l in fin]
link_counts = Counter()
for movie in movies:
link_counts.update(movie[2])
link_counts.most_common(10)
movies[1][0]
movies[1][1]
movies[1][2]
# +
top_links = [link for link, c in link_counts.items() if c >= 3]
link_to_idx = {link: idx for idx, link in enumerate(top_links)}
idx_to_link = {idx: link for link, idx in link_to_idx.items()}
movie_to_idx = {movie[0]: idx for idx, movie in enumerate(movies)}
idx_to_movie = {idx: movie for movie, idx in movie_to_idx.items()}
pairs = []
for movie in movies:
pairs.extend((link_to_idx[link], movie_to_idx[movie[0]]) for link in movie[2] if link in link_to_idx)
pairs_set = set(pairs)
len(pairs), len(top_links), len(movie_to_idx)
# -
len(link_to_idx)
# +
pairs[1]
idx_to_link[1]
idx_to_movie[0]
# -
pairs[1000]
idx_to_link[564]
idx_to_movie[3]
pairs[:10]
len(pairs_set)
# +
def movie_embedding_model(embedding_size=50):
link = Input(name='link', shape=(1,))
movie = Input(name='movie', shape=(1,))
link_embedding = Embedding(name='link_embedding',
input_dim=len(top_links),
output_dim=embedding_size)(link)
movie_embedding = Embedding(name='movie_embedding',
input_dim=len(movie_to_idx),
output_dim=embedding_size)(movie)
dot = Dot(name='dot_product', normalize=True, axes=2)([link_embedding, movie_embedding])
merged = Reshape((1,))(dot)
model = Model(inputs=[link, movie], outputs=[merged])
model.compile(optimizer='nadam', loss='mse')
return model
model = movie_embedding_model()
model.summary()
# +
random.seed(5)
def batchifier(pairs, positive_samples=50, negative_ratio=10):
"""Generate batches of positive and negative samples"""
batch_size = positive_samples * (1 + negative_ratio)
batch = np.zeros((batch_size, 3))
# Generator yields batches
while True:
# Sample random positive samples
for idx, (link_id, movie_id) in enumerate(random.sample(pairs, positive_samples)):
# The labels for these samples are positive (1)
batch[idx, :] = (link_id, movie_id, 1)
idx = positive_samples
# Continue adding negative samples until batch size is reached
while idx < batch_size:
# Sample a random movie
movie_id = random.randrange(len(movie_to_idx))
# Sample a random link
link_id = random.randrange(len(top_links))
# If the link is not on the movie page, this is a negative sample
if not (link_id, movie_id) in pairs_set:
# Set the label as negative (-1)
batch[idx, :] = (link_id, movie_id, -1)
idx += 1
# Make sure to shuffle the order of the positive and negative samples
np.random.shuffle(batch)
# Yield the links, movies, and labels
yield {'link': batch[:, 0], 'movie': batch[:, 1]}, batch[:, 2]
next(batchifier(pairs, positive_samples=3, negative_ratio=2))
# +
positive_samples_per_batch = 512
# Fit the model to samples from the generator
model.fit_generator(
batchifier(pairs, positive_samples=positive_samples_per_batch, negative_ratio=10),
epochs=15,
# Number of batches to grab every epoch
steps_per_epoch=len(pairs) // positive_samples_per_batch,
verbose=2
)
# +
movie = model.get_layer('movie_embedding')
movie_weights = movie.get_weights()[0]
movie_lengths = np.linalg.norm(movie_weights, axis=1)
normalized_movies = (movie_weights.T / movie_lengths).T
def similar_movies(movie):
dists = np.dot(normalized_movies, normalized_movies[movie_to_idx[movie]])
closest = np.argsort(dists)[-10:]
for c in reversed(closest):
print(c, 'Movie:', movies[c][0], 'Rating:', movies[c][-1], 'Distance:', dists[c])
return dists
d = similar_movies('Rogue One')
# +
from sklearn.manifold import TSNE
tsne = TSNE(n_components = 2, perplexity = 10, verbose = 1).fit_transform(normalized_movies)
# -
ratings = [int(r[3][:-1]) if r[3] is not None else np.nan for r in movies]
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('fivethirtyeight')
import seaborn as sns
xs = tsne[:, 0]
ys = tsne[:, 1]
plt.figure(figsize = (10, 8))
plt.scatter(xs, ys, c = ratings)
plot_movies = random.sample(movie_to_idx.keys(), 20)
for m in plot_movies:
index = movie_to_idx[m]
plt.text(x = tsne[index, 0], y = tsne[index, 1], s = m, fontsize = 10);
plt.colorbar();
# -
link_counts
movies[1]
list(reversed(np.argsort(d)[-10:]))
# +
link = model.get_layer('link_embedding')
link_weights = link.get_weights()[0]
# Find magnitudes
link_lengths = np.linalg.norm(link_weights, axis=1)
# Divide by magnitudes (normalization)
normalized_links = (link_weights.T / link_lengths).T
def similar_links(link: str) -> None:
"""Find the most similar links"""
dists = np.dot(normalized_links, normalized_links[link_to_idx[link]])
closest = np.argsort(dists)[-10:]
for c in reversed(closest):
print(c, top_links[c], dists[c])
similar_links('<NAME>')
# -
similar_links('Category:Films based on American novels')
best = ['Star Wars: The Force Awakens', 'The Martian (film)', 'Tangerine (film)', 'Straight Outta Compton (film)',
'Brooklyn (film)', 'Carol (film)', 'Spotlight (film)']
worst = ['American Ultra', 'The Cobbler (2014 film)', 'Entourage (film)', 'Fantastic Four (2015 film)',
'Get Hard', 'Hot Pursuit (2015 film)', 'Mortdecai (film)', 'Serena (2014 film)', 'Vacation (2015 film)']
y = np.asarray([1 for _ in best] + [0 for _ in worst])
X = np.asarray([normalized_movies[movie_to_idx[movie]] for movie in best + worst])
X.shape
clf = svm.SVC(kernel='linear')
clf.fit(X, y)
# +
estimated_movie_ratings = clf.decision_function(normalized_movies)
best = np.argsort(estimated_movie_ratings)
print('best:')
for c in reversed(best[-5:]):
print(c, movies[c][0], estimated_movie_ratings[c])
print('worst:')
for c in best[:5]:
print(c, movies[c][0], estimated_movie_ratings[c])
# -
rotten_y = np.asarray([float(movie[-2][:-1]) / 100 for movie in movies if movie[-2]])
rotten_X = np.asarray([normalized_movies[movie_to_idx[movie[0]]] for movie in movies if movie[-2]])
TRAINING_CUT_OFF = int(len(rotten_X) * 0.8)
regr = LinearRegression()
regr.fit(rotten_X[:TRAINING_CUT_OFF], rotten_y[:TRAINING_CUT_OFF])
error = (regr.predict(rotten_X[TRAINING_CUT_OFF:]) - rotten_y[TRAINING_CUT_OFF:])
'mean square error %2.2f' % np.mean(error ** 2)
error = (np.mean(rotten_y[:TRAINING_CUT_OFF]) - rotten_y[TRAINING_CUT_OFF:])
'mean square error %2.2f' % np.mean(error ** 2)
# +
def gross(movie):
v = movie[1].get('gross')
if not v or not ' ' in v:
return None
v, unit = v.split(' ', 1)
unit = unit.lower()
if not unit in ('million', 'billion'):
return None
if not v.startswith('$'):
return None
try:
v = float(v[1:])
except ValueError:
return None
if unit == 'billion':
v *= 1000
return v
movie_gross = [gross(m) for m in movies]
movie_gross = np.asarray([gr for gr in movie_gross if gr is not None])
highest = np.argsort(movie_gross)[-10:]
for c in reversed(highest):
print(c, movies[c][0], movie_gross[c])
# -
gross_y = np.asarray([gr for gr in movie_gross if gr])
gross_X = np.asarray([normalized_movies[movie_to_idx[movie[0]]] for movie, gr in zip(movies, movie_gross) if gr])
TRAINING_CUT_OFF = int(len(gross_X) * 0.8)
regr = LinearRegression()
regr.fit(gross_X[:TRAINING_CUT_OFF], gross_y[:TRAINING_CUT_OFF])
error = (regr.predict(gross_X[TRAINING_CUT_OFF:]) - gross_y[TRAINING_CUT_OFF:])
'mean square error %2.2f' % np.mean(error ** 2)
error = (np.mean(gross_y[:TRAINING_CUT_OFF]) - gross_y[TRAINING_CUT_OFF:])
'mean square error %2.2f' % np.mean(error ** 2)
| 04.2 Build a recommender system based on outgoing Wikipedia links.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ###### Content provided under a Creative Commons Attribution license, CC-BY 4.0; code under BSD 3-Clause License. (c)2014 <NAME>, <NAME>. Thanks: NSF for support via CAREER award #1149784.
# # Flow over a cylinder with source-panels
# In previous lessons, we used potential-flow singularities of defined strength to represent the shape of simple geometries, such as a [Rankine oval](02_Lesson02_sourceSinkFreestream.ipynb) or a [circular cylinder](03_Lesson03_doublet.ipynb), immersed in a free stream. We were rather lucky that when superposing a few fundamental potential-flow solutions, the stream-line pattern that resulted had a closed dividing stream line that we could interpret as a solid body.
#
# But what if we want to represent the stream lines around an *arbitrary* geometry? Would you be able to define the combination of fundamental solutions to get the expected result? *How could you do that?* Trial and error? It would take enormous luck and a lot of work to get a geometry we want.
#
# In this lesson, the objective is to calculate the source-strength distribution that can produce potential flow around a given geometry: a circular cylinder. We know that we can get the flow around a cylinder by superposing a doublet in a free stream, but here we want to develop a more general approach that can later be extended to *different* shapes.
#
# The method we will use consists of representing the geometry of the body by a series of short linear segments, called *panels*, that correspond to [source sheets](08_Lesson08_sourceSheet.ipynb) like we studied in the previous lesson.
#
# What we are aiming for is a method that starts with a defined body geometry, then works out the strength of sources needed in each panel representing that geometry to get a dividing streamline right on the body boundary. We will have to *solve* for the source strengths by specifying that the body be a streamline, i.e., the velocity be tangent there.
#
# Let's start by loading the Python libraries that we will need in this notebook.
import math
import numpy
from scipy import integrate
from matplotlib import pyplot
# embed the figures into the notebook
# %matplotlib inline
# We will add a uniform horizontal flow of magnitude `u_inf`, so let's make that equal to 1:
u_inf = 1.0 # free-stream speed
# ### Definition of the geometry
# The geometry considered here will be a circular cylinder of unit radius. We can define this geometry very easily by a set of points going around the angular range between $0$ and $2\pi$.
# +
# define the cylinder of untit radius centered at (0, 0)
R = 1.0
x_center, y_center = 0.0, 0.0
theta = numpy.linspace(0.0, 2 * math.pi, 100)
x_cylinder, y_cylinder = (x_center + R * numpy.cos(theta),
y_center + R * numpy.sin(theta))
# plot the cylinder
size = 4
pyplot.figure(figsize=(size, size))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('y', fontsize=16)
pyplot.plot(x_cylinder, y_cylinder, color='b', linestyle='-', linewidth=2)
pyplot.xlim(-1.1, 1.1)
pyplot.ylim(-1.1, 1.1);
# -
# ### Discretization into panels
# A panel, which represents a source sheet, is defined by its two end-points (`xa`, `ya`) and (`xb`, `yb`) and its strength `sigma`. We'll also need its center point (`xc`, `yc`) and its length for further calculations. The orientation of the panel is defined by the angle between the $x$-axis and its normal in the counter-clockwise sense.
#
# What information do we need to compute on each panel? First of all, we will need the strength of the source sheet that will lead to the correct streamlines. In addition, we'll also want the tangential velocity (the normal velocity on the body is zero for an inviscid flow) and the pressure coefficient.
#
# In this lesson, you'll really appreciate having learned about classes. It will make the code so much easier to manage. We create a class named `Panel` containing all the geometry data related to one panel. With a start- and end-point, the class internally calculates the center-point, length and normal vector. It also initializes to zero the source strength, tangential velocity and pressure coefficient. (These will be updated later.)
class Panel:
"""
Contains information related to a panel.
"""
def __init__(self, xa, ya, xb, yb):
"""
Initializes the panel.
Sets the end-points and calculates the center, length, and angle
(with the x-axis) of the panel.
Initializes the strength of the source-sheet, the tangential velocity,
and the pressure coefficient to zero.
Parameters
----------
xa: float
x-coordinate of the first end-point.
ya: float
y-coordinate of the first end-point.
xb: float
x-coordinate of the second end-point.
yb: float
y-coordinate of the second end-point.
"""
self.xa, self.ya = xa, ya
self.xb, self.yb = xb, yb
self.xc, self.yc = (xa + xb) / 2, (ya + yb) / 2 # control-point (center-point)
self.length = math.sqrt((xb - xa)**2 + (yb - ya)**2) # length of the panel
# orientation of the panel (angle between x-axis and panel's normal)
if xb - xa <= 0.:
self.beta = math.acos((yb - ya) / self.length)
elif xb - xa > 0.:
self.beta = math.pi + math.acos(-(yb - ya) / self.length)
self.sigma = 0.0 # source strength
self.vt = 0.0 # tangential velocity
self.cp = 0.0 # pressure coefficient
# To store all the discretization, we create a NumPy array of size `N_panels` where each item in the array is an object of type `Panel`.
# +
N_panels = 10 # number of panels desired
# define the end-points of the panels
x_ends = R * numpy.cos(numpy.linspace(0.0, 2 * math.pi, N_panels + 1))
y_ends = R * numpy.sin(numpy.linspace(0.0, 2 * math.pi, N_panels + 1))
# define the panels
panels = numpy.empty(N_panels, dtype=object)
for i in range(N_panels):
panels[i] = Panel(x_ends[i], y_ends[i], x_ends[i + 1], y_ends[i + 1])
# plot the panels
size = 6
pyplot.figure(figsize=(size, size))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('y', fontsize=16)
pyplot.plot(x_cylinder, y_cylinder,
label='cylinder',
color='b', linestyle='-', linewidth=1)
pyplot.plot(x_ends, y_ends,
label='panels',
color='#CD2305', linestyle='-', linewidth=2)
pyplot.scatter([p.xa for p in panels], [p.ya for p in panels],
label='end-points',
color='#CD2305', s=40)
pyplot.scatter([p.xc for p in panels], [p.yc for p in panels],
label='center-points',
color='k', s=40, zorder=3)
pyplot.legend(loc='best', prop={'size':16})
pyplot.xlim(-1.1, 1.1)
pyplot.ylim(-1.1, 1.1);
# -
# ### Flow-tangency boundary condition
# In [Lesson 1](01_Lesson01_sourceSink.ipynb), you worked out the velocity potential of a single source as part of the final *Challenge Task.* It followed simply from integrating the radial velocity, $u_r=\frac{\sigma}{2\pi r}$, that
#
# $$\phi=\frac{\sigma}{2\pi}\ln r$$
#
# (The integration also gives a function of $\theta$ that is seen to be a constant because $u_{\theta}=0$; we take this constant to be zero.)
#
# We will use the velocity potential in this lesson to easily express that the velocity be tangent at the panel, i.e., that $u_n=0$ with:
#
# $$u_n(x,y)=\frac{\partial \phi}{\partial n}(x,y)$$
#
# at a given point on the panel. We choose the point to enforce that velocity-tangency condition as the center of the panel (and we call it the *control point*).
#
# The velocity potential in Cartesian coordinates of a [source sheet](08_Lesson08_sourceSheet.ipynb) on a panel is
#
# $$\phi\left(x,y\right) = \frac{\sigma}{2\pi} \int_\text{panel} \ln \sqrt{ \left(x-x(s)\right)^2 + \left(y-y(s)\right)^2 } {\rm d}s$$
#
# where $s$ is the running coordinate along the panel and $\left(x(s),y(s)\right)$ are the Cartesian coordinates of $s$.
#
# Superposition of the potential of each panel gives the total potential at any point $\left(x,y\right)$, so we make a sum of all the panel contributions as follows (moving the $\frac{1}{2}$ exponent in the logarithmic term as a factor outside the integral):
#
# $$\phi\left(x,y\right) = \sum_{j=1}^{N_p} \frac{\sigma_j}{4\pi} \int \ln \left( \left(x-x_j(s_j)\right)^2 + \left(y-y_j(s_j)\right)^2 \right) {\rm d}s_j$$
#
# By finally superposing the free stream, the flow around an immersed circular cylinder will be represented by the following velocity potential:
#
# $$\phi\left(x,y\right) = U_\infty x + \sum_{j=1}^{N_p} \frac{\sigma_j}{4\pi} \int \ln \left( \left(x-x_j(s_j)\right)^2 + \left(y-y_j(s_j)\right)^2 \right) {\rm d}s_j$$
# Enforcing the flow-tangency condition on each *control point* approximately makes the body geometry correspond to a dividing streamline (and the approximation improves if we represent the body with more and more panels). So, for each panel $i$, we make $u_n=0$ at $(x_{c_i},y_{c_i})$:
#
# $$u_{n_i} = \frac{\partial}{\partial n_i}\left\lbrace \phi\left(x_{c_i},y_{c_i}\right) \right\rbrace = 0$$
#
# which leads to
#
# $$
# 0 = U_\infty \cos\beta_i + \sum_{j=1}^{N_p} \frac{\sigma_j}{2\pi} \int \frac{\left(x_{c_i}-x_j(s_j)\right) \frac{\partial x_{c_i}}{\partial n_i} + \left(y_{c_i}-y_j(s_j)\right) \frac{\partial y_{c_i}}{\partial n_i}} {\left(x_{c_i}-x_j(s)\right)^2 + \left(y_{c_i}-y_j(s)\right)^2} {\rm d}s_j
# $$
#
# where $\beta_i$ is the angle that the panel's normal makes with the $x$-axis, so
#
# $$\frac{\partial x_{c_i}}{\partial n_i} = \cos\beta_i \quad \text{and}\quad\frac{\partial y_{c_i}}{\partial n_i} = \sin\beta_i$$
#
# and
#
# $$x_j(s_j) = x_{a_j} - \sin\left(\beta_j\right) s_j$$
# $$y_j(s_j) = y_{a_j} + \cos\left(\beta_j\right) s_j$$
#
# But, there is still a problem to handle when $i=j$. From the previous notebook, we have seen that the strength of the [source sheet](08_Lesson08_sourceSheet.ipynb) should be a specific value so that the streamlines do not penetrate the panel. This helps us determine that the contribution of the $i$-th panel to itself is $\frac{\sigma_i}{2}$.
#
# Finally, the boundary condition at the center point of the $i$-th panel gives
#
# $$
# 0 = U_\infty \cos\beta_i + \frac{\sigma_i}{2} + \sum_{j=1,j\neq i}^{N_p} \frac{\sigma_j}{2\pi} \int \frac{\left(x_{c_i}-x_j(s_j)\right) \cos\beta_i + \left(y_{c_i}-y_j(s_j)\right) \sin\beta_i} {\left(x_{c_i}-x_j(s)\right)^2 + \left(y_{c_i}-y_j(s)\right)^2} {\rm d}s_j
# $$
#
# From the equation above, we understand that we will have to compute integrals using the SciPy function `integrate.quad()`. We define a function `integral_normal()` that will do the job.
def integral_normal(p_i, p_j):
"""
Evaluates the contribution of a panel at the center-point of another,
in the normal direction.
Parameters
----------
p_i: Panel object
Panel on which the contribution is calculated.
p_j: Panel object
Panel from which the contribution is calculated.
Returns
-------
Integral over the panel at the center point of the other.
"""
def integrand(s):
return (((p_i.xc - (p_j.xa - math.sin(p_j.beta) * s)) * math.cos(p_i.beta) +
(p_i.yc - (p_j.ya + math.cos(p_j.beta) * s)) * math.sin(p_i.beta)) /
((p_i.xc - (p_j.xa - math.sin(p_j.beta) * s))**2 +
(p_i.yc - (p_j.ya + math.cos(p_j.beta) * s))**2))
return integrate.quad(integrand, 0.0, p_j.length)[0]
# ### Solving the system of equations
# We just developed an equation to enforce a flow-tangency condition on the $i$-th panel. There are `N_panels` panels $i$ and `N_panels` unknown strengths $\sigma_i$. Therefore, the problem represents solving a linear system of equations of the form
#
# $$[A][\sigma] = [b]$$
#
# where
#
# $$
# A_{ij} = \begin{cases}
# \begin{matrix}
# \frac{1}{2} & \mbox{, if } i=j \cr
# \frac{1}{2\pi} \int \frac{\left(x_{c_i}-x_j(s_j)\right) \cos\beta_i + \left(y_{c_i}-y_j(s_j)\right) \sin\beta_i} {\left(x_{c_i}-x_j(s)\right)^2 + \left(y_{c_i}-y_j(s)\right)^2} ds_j & \mbox{, if } i\neq j
# \end{matrix}
# \end{cases}
# $$
#
# and
#
# $$b_i = - U_\infty \cos\beta_i$$
#
# for $1\leq i,j \leq N_p$. Let's fill a matrix $A$ and a right-hand side vector $b$ with the necessary values:
# +
# compute the source influence matrix
A = numpy.empty((N_panels, N_panels), dtype=float)
numpy.fill_diagonal(A, 0.5)
for i, p_i in enumerate(panels):
for j, p_j in enumerate(panels):
if i != j:
A[i, j] = 0.5 / math.pi * integral_normal(p_i, p_j)
# compute the RHS of the linear system
b = - u_inf * numpy.cos([p.beta for p in panels])
# -
# Hey! We just used a new Python built-in function: [enumerate()](https://docs.python.org/2/library/functions.html#enumerate). It allows us to have access to each element `panel` in the array `panels` while keeping a count `i` (that starts from `0`) to locate the element of `A` to fill.
# Now, we can easily solve the linear system of equations using the function [`linalg.solve()`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.solve.html) from NumPy, and assign each source-panel its appropriate strength:
# +
# solve the linear system
sigma = numpy.linalg.solve(A, b)
for i, panel in enumerate(panels):
panel.sigma = sigma[i]
# -
# ### Pressure coefficient on the surface
# At this point, we have the source strength distribution required to compute the streamlines around our geometry. A very useful measurement of the results is the pressure coefficient along the surface of the geometry.
#
# From Bernoulli's equation, the pressure coefficient on the $i$-th panel is
#
# $$C_{p_i} = 1-\left(\frac{u_{t_i}}{U_\infty}\right)^2$$
#
# where $u_{t_i}$ is the tangential component of the velocity at the center point of the $i$-th panel,
#
# $$u_{t_i} = \frac{\partial}{\partial t_i}\left\lbrace \phi\left(x_{c_i},y_{c_i}\right) \right\rbrace$$
#
# which we can obtain as:
#
# $$
# u_{t_i} = -U_\infty \sin\beta_i + \sum_{j=1}^{N_p} \frac{\sigma_j}{2\pi} \int \frac{\left(x_{c_i}-x_j(s_j)\right) \frac{\partial x_{c_i}}{\partial t_i} + \left(y_{c_i}-y_j(s_j)\right) \frac{\partial y_{c_i}}{\partial t_i}} {\left(x_{c_i}-x_j(s)\right)^2 + \left(y_{c_i}-y_j(s)\right)^2} {\rm d}s_j
# $$
#
# with
#
# $$\frac{\partial x_{c_i}}{\partial t_i} = -\sin\beta_i \quad\text{and} \quad \frac{\partial y_{c_i}}{\partial t_i} = \cos\beta_i$$
#
# Note that the contribution to the tangential velocity at a source panel from its own velocity potential is zero, which makes sense because streamlines go *outwards* from a source.
#
# We define a function `integral_tangential()` that will compute the integrals above using the SciPy function `integrate.quad()` once again:
def integral_tangential(p_i, p_j):
"""
Evaluates the contribution of a panel at the center-point of another,
in the tangential direction.
Parameters
----------
p_i: Panel object
Panel on which the contribution is calculated.
p_j: Panel object
Panel from which the contribution is calculated.
Returns
-------
Integral over the panel at the center point of the other.
"""
def integrand(s):
return ((-(p_i.xc - (p_j.xa - math.sin(p_j.beta) * s)) * math.sin(p_i.beta) +
(p_i.yc - (p_j.ya + math.cos(p_j.beta) * s)) * math.cos(p_i.beta)) /
((p_i.xc - (p_j.xa - math.sin(p_j.beta) * s))**2 +
(p_i.yc - (p_j.ya + math.cos(p_j.beta) * s))**2))
return integrate.quad(integrand, 0.0, p_j.length)[0]
# +
# compute the matrix of the linear system
A = numpy.empty((N_panels, N_panels), dtype=float)
numpy.fill_diagonal(A, 0.0)
for i, p_i in enumerate(panels):
for j, p_j in enumerate(panels):
if i != j:
A[i, j] = 0.5 / math.pi * integral_tangential(p_i, p_j)
# compute the RHS of the linear system
b = - u_inf * numpy.sin([panel.beta for panel in panels])
# compute the tangential velocity at each panel center-point
vt = numpy.dot(A, sigma) + b
for i, panel in enumerate(panels):
panel.vt = vt[i]
# -
# Once we have computed the tangential velocity on each panel, we can calculate the pressure coefficient.
# calculate the surface pressure coefficient
for panel in panels:
panel.cp = 1.0 - (panel.vt / u_inf)**2
# Alright! It is time to plot the surface pressure coefficient.
#
# Just before this, we should remember that in the lesson on the [doublet](03_Lesson03_doublet.ipynb), we found that the exact pressure coefficient on the surface on a cylinder was
#
# $$Cp = 1 - 4\sin^2 \theta$$
#
# i.e.
#
# $$Cp = 1 - 4\left(\frac{y}{R}\right)^2$$
#
# We can use this to compare with the results obtained with our source-panel code.
# +
# calculate the analytical surface pressure coefficient
cp_analytical = 1.0 - 4 * (y_cylinder / R)**2
# plot the surface pressure coefficient
pyplot.figure(figsize=(10, 6))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('$C_p$', fontsize=16)
pyplot.plot(x_cylinder, cp_analytical,
label='analytical',
color='b', linestyle='-', linewidth=1, zorder=1)
pyplot.scatter([p.xc for p in panels], [p.cp for p in panels],
label='source-panel method',
color='#CD2305', s=40, zorder=2)
pyplot.title('Number of panels : %d' % N_panels, fontsize=16)
pyplot.legend(loc='best', prop={'size':16})
pyplot.xlim(-1.0, 1.0)
pyplot.ylim(-4.0, 2.0);
# -
# ##### Challenge task
# Now that we have computed the pressure coefficient on the surface of the cylinder, it will be interesting to visualize what the streamlines look like.
#
# To do that, we use the function `streamplot()` from Matplotlib, requiring the Cartesian velocity components (`u`,`v`) on a mesh grid (`X`,`Y`). Therefore, the first step is to derive the equations for the velocity components.
#
# The potential at point $\left(x,y\right)$ of the $N_p$ source sheets in a uniform horizontal flow $U_\infty$ is
#
# $$\phi\left(x,y\right) = U_\infty x + \sum_{j=1}^{N_p} \frac{\sigma_j}{4\pi} \int \ln \left( \left(x-x_j(s_j)\right)^2 + \left(y-y_j(s_j)\right)^2 \right) {\rm d}s_j$$
#
# And the velocity field at point $\left(x,y\right)$ is
#
# $$u\left(x,y\right) = \frac{\partial}{\partial x}\left\lbrace \phi\left(x,y\right) \right\rbrace$$
#
# $$v\left(x,y\right) = \frac{\partial}{\partial y}\left\lbrace \phi\left(x,y\right) \right\rbrace$$
#
# Your task will be to:
#
# * derive the Cartesian velocity components
# * create a mesh grid
# * compute the velocity field on the mesh grid
# * plot the results
# * change the number of panels to improve the visualization
# ---
# + active=""
# Please ignore the cell below. It just loads our style for the notebook.
# -
from IPython.core.display import HTML
def css_styling(filepath):
styles = open(filepath, 'r').read()
return HTML(styles)
css_styling('../styles/custom.css')
| lessons/09_Lesson09_flowOverCylinder.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// # Set
// https://en.cppreference.com/w/cpp/container/set
//
// ## Table of Contents
// - [set definition](#set)
// - [Declare](#declare)
// - [Access elements](#access)
// - [Modifiers](#modifiers)
// - [Comparisons](#comparisons)
// - [Iterators](#iterators)
// - [Lookup operations](#operations)
// <a id="set"></a>
// ## set
// - associative container that contains a sorted set of unique objects of some type called key
// - implemented as [red-black trees](https://en.wikipedia.org/wiki/Red%E2%80%93black_tree)
// - the complexity (efficiency) of common operations on set such as search, removal, and insertion operations `O(lg n)`
// <a id="declare"></a>
// ## declare set
// - must include set header file and use namespace std;
// - a template class designed to store any data type that can be comapred
// + code_folding=[]
// include header files
#include <iostream>
#include <string>
#include <set>
using namespace std;
// + code_folding=[]
// operator<< overloaded to print a list container
template<class T>
ostream& operator<<(ostream& out, const set<T>& v) {
char comma[3] = {'\0', ' ', '\0'};
out << '[';
for (auto& e: v) {
out << comma << e;
comma[0] = ',';
}
out << ']';
return out;
}
// + code_folding=[]
// declare
set<string> names;
set<float> tests;
set<int> numbers;
// + code_folding=[]
// declare and initialize
set<string> words = {"i", "love", "C++", "!"};
set<float> prices = {1.99, 199, 2.99, 200.85, 45.71};
set<float> dupPrices = prices;
// -
// print contents using cout operator<< overloaded function
cout << "names contents: " << names << endl;
cout << "words contents: "<< words << endl;
cout << prices << endl;
// <a id="access"></a>
// ## access elements
// - no direct method or operator to access elements
// <a id="capacity"></a>
// ## capacity
// - empty : checks whethere the container is empty
// - size : returns the number of elements
// - max_size : returns the maximum possible number of elements
cout << boolalpha; // convert boolean to text true/false
cout << "is names set empty? " << names.empty() << endl;
cout << "is prices set empty? " << prices.empty() << endl;
cout << "size of prices: " << prices.size() << endl;
cout << "max_size of prices: " << prices.max_size() << endl;
// <a id="modifiers"></a>
// ## modifiers
// - **clear** : clears the contents
// - **insert** : inserts elements into the container, if the container doesn't already contain an element with an quivalent key
// - **erase** : erases elements at the specified location or key
// - **swap** : swaps the contents
set<int> age = {21, 74, 46, 48, 46};
cout << age << endl;
age.clear();
cout << age << endl;
age.insert(10);
cout << age << endl;
age.insert(12);
cout << age;
auto it = age.begin();
cout << *it << endl;
age.erase(it);
cout << "age: " << age;
age.insert(15);
cout << age << endl;
age.insert(100);
cout << age << endl;
age.insert(50);
age.insert(55);
cout << age;
// erase key 12 if exists; nothing happes if doesn't exist
age.erase(12);
cout << age << endl;
age.erase(999);
set<int> age2 = {1, 2, 3, 4, 5};
cout << age << endl;
cout << age2 << endl;
// inorder to swap, number of elements between the containers have to be equal
age2.swap(age);
cout << age << endl;
cout << age2 << endl;
// <a id="comparisons"></a>
// ## aggregate comparisons
// - comparison operators ==, !=, <, <=, and >= are overloaded
// - elements are compared lexicographically
if (age == age2)
cout << "age and age2 contains same elements compared lexicographically!" << endl;
else
cout << "age and age2 do not contain same element(s)" << endl;
// <a id="iterators"></a>
// ## iterators
// - **begin** - returns an iterator to the beginning
// - **rbegin** - returns a reverse iterator to the beginning
// - **end** - returns an iterator to the end (past the last element)
// - **rend** - returns a reverse iterator to the end
// <img src="./resources/range-rbegin-rend.svg" />
set<int> nums = {10, 15, 20, 30, 35};
for(auto iterator = nums.begin(); iterator != nums.end(); iterator++)
cout << *iterator << " ";
cout << endl;
using si = set<int>;
si aset = {1, 2, 3, 4, 5};
cout << aset << endl;
// move iterators using next function
auto iter = aset.begin();
cout << *iter << endl;
// move iterator 1 element ahead
iter = next(iter);
cout << *iter << endl;
// move iterator 3 elements ahead
iter = next(iter, 3);
cout << *iter << endl;
// <a id="operations"></a>
// ## Lookup operations
// - **count** : returns the number of elements matching specific key (always 1 if exists, 0 otherwise)
// - **find** : finds elements with specific key, returns iterator
set<int> set1 = {5, 9, 100, 15, 75};
set<int> set2 = {30, 11, 11, 7, 75};
cout << set1 << endl;
cout << set2 << endl;
cout << set1.count(100) << endl;
cout << set1.count(99) << endl;
if (set1.count(15) == 1)
cout << "Found!";
else
cout << "Not found!";
set<int> set3 = {90, 11, 0, 7, 75};
// find method
auto search = set3.find(7);
if (search != set3.end())
cout << "found " << *search << endl;
else
cout << "NOT found!";
| Set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Load the "autoreload" extension so that code can change
# %load_ext autoreload
# %reload_ext autoreload
from pathlib import Path
# always reload modules so that as you change code in src, it gets loaded
# %autoreload 2
# %matplotlib inline
import sys
sys.path.append('../')
from src.imports import *
from src.data.download_data import *
from src.data.read_data import *
from src.data.fire_data import *
from src.data.vn_data import *
from src.data.weather_data import *
from src.gen_functions import *
from src.features.dataset import Dataset
import scrapy
from scrapy.crawler import CrawlerProcess
from twisted.internet import reactor
import scrapy
from scrapy.crawler import CrawlerRunner
from crochet import setup
import logging
# -
# Scrape location of VN power plants using scrapy
# + [markdown] heading_collapsed=true
# # Example
# + hidden=true
class JsonWriterPipeline(object):
def open_spider(self, spider):
self.file = open('quoteresult.jl', 'w')
def close_spider(self, spider):
self.file.close()
def process_item(self, item, spider):
line = json.dumps(dict(item)) + "\n"
self.file.write(line)
return item
# + hidden=true
import logging
class QuotesSpider(scrapy.Spider):
name = "quotes"
start_urls = [
'http://quotes.toscrape.com/page/1/',
'http://quotes.toscrape.com/page/2/',
]
custom_settings = {
'LOG_LEVEL': logging.WARNING,
'ITEM_PIPELINES': {'__main__.ret': 1}, # Used for pipeline 1
'FEED_FORMAT':'json', # Used for pipeline 2
'FEED_URI': '../data/hanoi/plants_info.json' # Used for pipeline 2
}
def parse(self, response):
for quote in response.css('div.quote'):
yield {
'text': quote.css('span.text::text').extract_first(),
'author': quote.css('span small::text').extract_first(),
'tags': quote.css('div.tags a.tag::text').extract(),
}
# + hidden=true
runner = CrawlerRunner() #from Scrapy docs
runner.crawl(QuotesSpider)
# -
# # Wikipedia Page on Coal Power Plants in Vietnam
class WikiSpider(scrapy.Spider):
name = "wiki"
custom_settings = {
'LOG_LEVEL': logging.WARNING,
'FEED_FORMAT':'json', # Used for pipeline 2
'FEED_URI': '../data/hanoi/plants_info.json' }
start_urls = ['https://en.wikipedia.org/wiki/List_of_power_stations_in_Vietnam#For_gas,_coal-fired_power_plants']
def parse(self, response):
links = response.xpath('//*[@id="mw-content-text"]/div[1]/div[3]/div/ol/li/span[2]/cite[1]/a[1]/@href')
idxs = np.arange(len(links)) + 1
for i, link in zip(idxs, links):
if 'sourcewatch.org' in link.get():
yield scrapy.Request(link.get(), callback=self.re_direct, cb_kwargs=dict(ref=i))
def re_direct(self, response, ref):
link = response.xpath('//*[@id="softredirect"]/a/@href').get()
print(ref, link)
yield scrapy.Request(link, callback=self.parse_loc, cb_kwargs=dict(ref=ref))
def parse_loc(self, response, ref):
info = response.xpath('//*[@id="mw-content-text"]/div/ul[1]/li')
plant_info = {'ref':str(ref)}
for sub_info in info:
k = sub_info.xpath('b/text()').get()
k = k.replace(':','')
try:
content = sub_info.xpath('text()').get().lstrip()
except:
content = ''
plant_info[k] = content
print(plant_info)
yield plant_info
runner = CrawlerRunner() #from Scrapy docs
runner.crawl(WikiSpider)
# +
with open('../data/hanoi/plants_info.json') as f:
coals = json.load(f)
coals = pd.DataFrame(coals)
coals.head(2)
# -
coals['latitude'] = coals['Coordinates'].str.extract('(\d+.\d+)').astype(float)
coals['longitude'] = coals['Coordinates'].str.extract(', (\d+.\d+)').astype(float)
coals.to_csv('../data/hanoi/coat_plants.csv', index= False)
# +
coals = pd.read_csv('../data/hanoi/coat_plants.csv')
coals.head()
# -
# ## Other Power Plants
dfs = pd.read_html('https://en.wikipedia.org/wiki/List_of_power_stations_in_Vietnam#For_gas,_coal-fired_power_plants')
label_list = ['coal','gas','solar', 'wind', 'biomass', 'hydroelectricity']
# +
power_plants = []
for df, label in zip(dfs, label_list):
df['type'] = label
power_plants.append(df)
power_plants= pd.concat(power_plants)
# -
power_plants['Status'] = power_plants['Status'].fillna(power_plants['Unnamed: 7'])
#power_plants['Capacity (MW)'] = power_plants['Capacity (MW)'].fillna(power_plants['Capacity (MWp)'])
power_plants.to_csv('C:/Users/Benny/Documents/Fern/aqi_thailand2/data/hanoi/power_plants.csv',index=False)
power_plants[~power_plants['Unnamed: 7'].isna()]
power_plants.dropna(axis=1,how='all')
power_plants[(power_plants['type']=='gas') & (power_plants['Status']=='Operating')]
df2 = pd.read_html('https://en.wikipedia.org/wiki/List_of_gas_power_plants_in_Vietnam')[0]
df2.head()
df2[df2['Status']=='operating']
| notebooks/1.1_vn_power_plants.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
from numpy import sin, cos
import bokeh
from bokeh.io import output_file, show
from bokeh.plotting import figure
from bokeh.transform import linear_cmap
from bokeh.util.hex import hexbin
def clifford(xy,a,b,c,d):
x = xy[0]
y = xy[1]
return sin(a*y) + c*(cos(b*x)), sin(b*x) + d*(cos(a*y))
CI = [1,1]
l = []
for i in range(10000000):
CI = clifford(CI,-1.4,1.6,1,.7)
l.append(CI)
l = np.array(l)
plt.scatter(l.T[0],l.T[1])
bins = hexbin(l.T[0],l.T[1], 0.025)
# +
p = figure(tools="wheel_zoom,reset", match_aspect=True, background_fill_color='#440154')
p.grid.visible = False
p.hex_tile(q="q", r="r", size=0.1, line_color=None, source=bins,
fill_color=linear_cmap('counts', 'Viridis256', 0, max(bins.counts)))
show(p)
# -
CI = [1,1]
l = []
for i in range(5000000):
CI = clifford(CI,1.7,1.7,0.6,1.2)
l.append(CI)
l = np.array(l)
plt.scatter(l.T[0],l.T[1])
bins = hexbin(l.T[0],l.T[1], 0.025)
# +
p = figure(tools="wheel_zoom,reset", match_aspect=True, background_fill_color='#440154')
p.grid.visible = False
p.hex_tile(q="q", r="r", size=0.1, line_color=None, source=bins,
fill_color=linear_cmap('counts', 'Viridis256', 0, max(bins.counts)))
show(p)
# -
CI = [1,1]
l = []
for i in range(5000000):
CI = clifford(CI,-1.7,1.3,-0.1,-1.2)
l.append(CI)
l = np.array(l)
plt.scatter(l.T[0],l.T[1])
bins = hexbin(l.T[0],l.T[1], 0.025)
# +
p = figure(tools="wheel_zoom,reset", match_aspect=True, background_fill_color='#440154')
p.grid.visible = False
p.hex_tile(q="q", r="r", size=0.1, line_color=None, source=bins,
fill_color=linear_cmap('counts', 'Viridis256', 0, max(bins.counts)))
show(p)
# -
CI = [1,1]
l = []
for i in range(500000):
CI = clifford(CI,-1.7,2.9,-2.1,-3.2)
l.append(CI)
l = np.array(l)
plt.scatter(l.T[0],l.T[1])
bins = hexbin(l.T[0],l.T[1], 0.025)
# +
p = figure(tools="wheel_zoom,reset", match_aspect=True, background_fill_color='#440154')
p.grid.visible = False
p.hex_tile(q="q", r="r", size=0.1, line_color=None, source=bins,
fill_color=linear_cmap('counts', 'Viridis256', 0, max(bins.counts)))
show(p)
# -
| Clifford.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MJaiman/ML/blob/main/Headbrain_Simple_linear_regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="R_RZRC5Upo_6"
#importing necessary libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="bjeA3BLTqKsk" outputId="2645bc55-e74d-4484-b2d9-c243414815ce"
df=pd.read_csv('headbrain.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="VBqF7IrdqXCb" outputId="94ba06ab-8c5f-4052-d78a-731ce37a8158"
df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="6aKCTnO0qbGj" outputId="f43ffa7e-b0b7-4cd2-8077-eb175f59dfdd"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="9tOQsYU7qr_O" outputId="c14ad8d2-6575-436b-f12e-9510c099af68"
df.shape
# + id="m6egdt7Gq0ZX"
X = df['Head Size(cm^3)'].values
Y = df['Brain Weight(grams)'].values
# + colab={"base_uri": "https://localhost:8080/"} id="F-CLbaA6q9Lj" outputId="c43c5d3b-2d4a-4649-8192-377ce6337e21"
mean_x=np.mean(X)
mean_y=np.mean(Y)
n=len(X)
num =0
denom = 0
for i in range(n):
num+=(X[i]-mean_x)*(Y[i]-mean_y)
denom+=(X[i]-mean_x)**2
m=num/denom
c=mean_y-(m*mean_x)
print(m,',',c)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="fAI67BN8tCBV" outputId="bb892069-bd8d-4a49-ded4-a857562dfcf7"
plt.scatter(X,Y)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="-7askxRJxQsV" outputId="f5e764a3-4a16-4235-b2d4-4bd29e64b058"
x=X
y=yp
plt.scatter(X,Y,color='g')
plt.plot(x,y,color='r')
plt.title('Simple Linear Regression')
plt.xlabel('Head size cm^3')
plt.ylabel('Brain weight in grams')
# + colab={"base_uri": "https://localhost:8080/"} id="mXl5Elw5t19g" outputId="599cc4b4-1b48-4dc7-f878-47d2ba48de26"
#Calculating the error
nu=0
den=0
for i in range(n):
nu+=(yp[i]-mean_y)**2
den+=(Y[i]-mean_y)**2
R=nu/den
print(R)
# + id="sHAs9VeEwWIO"
def predict(x):
y = m*x + c
print(y)
# + colab={"base_uri": "https://localhost:8080/"} id="JawrHb8kwYiw" outputId="72ed515c-8b97-4344-ba54-2a93df722d79"
predict(4177)
| Headbrain_Simple_linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('/Users/lindsaybassman/Documents/LBL/ArQTiC')
from arqtic.simulation_generator import Simulation_Generator
#create simulation object
sim_obj = Simulation_Generator("TFIM_input.txt")
#connect to IBM
#if this is your first run enter your API key
#sim_obj.connect_IBM(api_key="your_key_here")
sim_obj.connect_IBM()
#generate circuits
sim_obj.generate_circuits()
#run circuits on backend
sim_obj.run_circuits()
# +
results=sim_obj.result_matrix
import matplotlib.pyplot as plt
plt.plot(results)
plt.xlabel("Simulation Timestep")
plt.ylabel("Average Magnetization")
plt.show()
# -
| examples/TFIM_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false}
# %matplotlib inline
# -
#
# Cold Magnetized Plasma Waves Tensor Elements (S, D, P in Stix's notation)
# =========================================================================
#
# This example shows how to calculate the values of the cold plasma tensor
# elements for various electromagnetic wave frequencies.
#
# + jupyter={"outputs_hidden": false}
# First, import some basics (and `PlasmaPy`!)
import numpy as np
import matplotlib.pyplot as plt
from astropy import units as u
from plasmapy.formulary import (cold_plasma_permittivity_SDP,
cold_plasma_permittivity_LRP)
# -
# Let's define some parameters, such as the magnetic field magnitude,
# the plasma species and densities and the frequency band of interest
#
#
# + jupyter={"outputs_hidden": false}
B = 2 * u.T
species = ['e', 'D+']
n = [1e18 * u.m ** -3, 1e18 * u.m ** -3]
f = np.logspace(start=6, stop=11.3, num=3001) # 1 MHz to 200 GHz
omega_RF = f * (2 * np.pi) * (u.rad / u.s)
# + jupyter={"outputs_hidden": false}
help(cold_plasma_permittivity_SDP)
# + jupyter={"outputs_hidden": false}
S, D, P = cold_plasma_permittivity_SDP(B, species, n, omega_RF)
# -
# Filter positive and negative values, for display purposes only.
# Still for display purposes, replace 0 by NaN to NOT plot 0 values
#
#
# + jupyter={"outputs_hidden": false}
S_pos = S * (S > 0)
D_pos = D * (D > 0)
P_pos = P * (P > 0)
S_neg = S * (S < 0)
D_neg = D * (D < 0)
P_neg = P * (P < 0)
S_pos[S_pos == 0] = np.NaN
D_pos[D_pos == 0] = np.NaN
P_pos[P_pos == 0] = np.NaN
S_neg[S_neg == 0] = np.NaN
D_neg[D_neg == 0] = np.NaN
P_neg[P_neg == 0] = np.NaN
# + jupyter={"outputs_hidden": false} tags=["nbsphinx-thumbnail"]
plt.figure(figsize=(12, 6))
plt.semilogx(f, abs(S_pos),
f, abs(D_pos),
f, abs(P_pos), lw=2)
plt.semilogx(f, abs(S_neg), '#1f77b4',
f, abs(D_neg), '#ff7f0e',
f, abs(P_neg), '#2ca02c', lw=2, ls='--')
plt.yscale('log')
plt.grid(True, which='major')
plt.grid(True, which='minor')
plt.ylim(1e-4, 1e8)
plt.xlim(1e6, 200e9)
plt.legend(('S > 0', 'D > 0', 'P > 0', 'S < 0', 'D < 0', 'P < 0'),
fontsize=16, ncol=2)
plt.xlabel('RF Frequency [Hz]', size=16)
plt.ylabel('Absolute value', size=16)
plt.tick_params(labelsize=14)
# -
# Cold Plasma tensor elements in the rotating basis
#
#
# + jupyter={"outputs_hidden": false}
L, R, P = cold_plasma_permittivity_LRP(B, species, n, omega_RF)
# + jupyter={"outputs_hidden": false}
L_pos = L * (L > 0)
R_pos = R * (R > 0)
L_neg = L * (L < 0)
R_neg = R * (R < 0)
L_pos[L_pos == 0] = np.NaN
R_pos[R_pos == 0] = np.NaN
L_neg[L_neg == 0] = np.NaN
R_neg[R_neg == 0] = np.NaN
plt.figure(figsize=(12, 6))
plt.semilogx(f, abs(L_pos),
f, abs(R_pos),
f, abs(P_pos), lw=2)
plt.semilogx(f, abs(L_neg), '#1f77b4',
f, abs(R_neg), '#ff7f0e',
f, abs(P_neg), '#2ca02c', lw=2, ls='--')
plt.yscale('log')
plt.grid(True, which='major')
plt.grid(True, which='minor')
plt.xlim(1e6, 200e9)
plt.legend(('L > 0', 'R > 0', 'P > 0', 'L < 0', 'R < 0', 'P < 0'),
fontsize=16, ncol=2)
plt.xlabel('RF Frequency [Hz]', size=16)
plt.ylabel('Absolute value', size=16)
plt.tick_params(labelsize=14)
# -
# Checks if the values obtained are coherent. They should satisfy
# S = (R+L)/2 and D = (R-L)/2
#
#
# + jupyter={"outputs_hidden": false}
try:
np.testing.assert_allclose(S, (R + L) / 2)
np.testing.assert_allclose(D, (R - L) / 2)
except AssertionError as e:
print(e)
# Checks for R=S+D and L=S-D
try:
np.testing.assert_allclose(R, S + D)
np.testing.assert_allclose(L, S - D)
except AssertionError as e:
print(e)
| docs/notebooks/cold_plasma_tensor_elements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv("train_.csv")
# -
df.head()
df = df[["Gender", "Married", "Education", "ApplicantIncome", "LoanAmount", "Loan_Amount_Term", "Credit_History", "Loan_Status"]]
df.head()
df = df[df["Loan_Status"].notnull()]
df.head()
df.info()
df = df.dropna()
df.isnull().sum()
df['Loan_Status'].value_counts()
fig, ax = plt.subplots(1,1, figsize=(12, 7))
df.boxplot('LoanAmount', 'Loan_Status', ax=ax)
plt.suptitle('LoanAmount v Loan_Status')
plt.title('')
plt.ylabel('LoanAmount')
plt.xticks(rotation=90)
plt.show()
from sklearn.preprocessing import LabelEncoder
le_education = LabelEncoder()
df['Education'] = le_education.fit_transform(df['Education'])
df['Education'].unique()
le_married = LabelEncoder()
df['Married'] = le_married.fit_transform(df['Married'])
df['Married'].unique()
le_gender = LabelEncoder()
df['Gender'] = le_gender.fit_transform(df['Gender'])
df['Gender'].unique()
le_lstatus = LabelEncoder()
df['Loan_Status'] = le_lstatus.fit_transform(df['Loan_Status'])
df['Loan_Status'].unique()
x = df.drop("Loan_Status", axis=1)
y = df["Loan_Status"]
from sklearn.linear_model import LinearRegression
linear_reg = LinearRegression()
linear_reg.fit(x, y.values)
y_pred = linear_reg.predict(x)
from sklearn.metrics import mean_squared_error, mean_absolute_error
import numpy as np
error = np.sqrt(mean_absolute_error(y, y_pred))
error = np.sqrt(mean_absolute_error(y, y_pred))
error
x
x = np.array([["Male", "Yes", "Not Graduate", 4333, 222, 180, 1]])
x
x[:, 0] = le_gender.transform(x[:, 0])
x[:, 1] = le_married.transform(x[:, 1])
x[:, 2] = le_education.transform(x[:, 2])
x = x.astype(float)
x
import pickle
data = {"model" : linear_reg, "le_gender": le_gender, "le_married": le_married, "le_education": le_education}
with open('saved_steps_polynomial.pkl', 'wb') as file:
pickle.dump(data, file)
# +
with open('saved_steps.pkl', 'rb') as file:
data = pickle.load(file)
linear_reg_load = data["model"]
le_gender = data["le_gender"]
le_married = data["le_married"]
le_education = data["le_education"]
# -
y_pred = linear_reg_load.predict(x)
y_pred
| Polynomial_Regression_Loan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6.0 64-bit
# name: python360jvsc74a57bd0aab07c05e18e3fae3dc841dc401f8bbd0373bf54f494cf150722bab0ea91cfef
# ---
# # 翻譯
#
# 與其他人交流的能力是使人類文明得以發展的驅動力之一。在多數人的努力嘗試中,交流是關鍵。
#
# 
#
# 人工智慧 (AI) 能透過翻譯不同語言間的文字或語音協助簡化交流,還能協助打破跨越國家/地區和文化的交流屏障。
#
# ## 建立認知服務資源
#
# 在 Azure 中,您可以使用認知服務在多個語言間的進行翻譯。
#
# 如果您還沒有該資源,請使用以下步驟在您的 Azure 訂用帳戶中建立一個**認知服務**資源:
#
# > **備註**:若您已經有認知服務資源,只需在 Azure 入口網站中開啟其 **[快速入門]**頁面並將其金鑰和端點複製到下面的儲存格。否則,可追隨下面的步驟來建立一個認知服務資源。
#
# 1.在其它瀏覽器索引標籤中,透過 https://portal.azure.com 開啟 Azure 入口網站,並用您的 Microsoft 帳戶登入。
# 2.按一下 **[+ 建立資源]** 按鈕,搜尋*認知服務*,並建立包含以下設定的**認知服務**資源:
# - **訂用帳戶**: *您的 Azure 訂用帳戶*。
# - **資源群組**: *選取或建立具有唯一名稱的資源群組*。
# - **區域**: *選擇任一可用區域*:
# - **名稱**: *輸入唯一名稱*。
# - **定價層** :S0
# - **我確認已閱讀通知並理解通知內容**: 已選取。
# 3.等待部署完成。然後前往您的認知服務資源,在 **[概觀]** 頁面上,按一下連結以管理服務金鑰。您將需要端點和金鑰,以便從用戶端應用程式連線到您的認知服務資源。
#
# ### 獲取適用於認知服務資源的金鑰和位置
#
# 若要使用您的認知服務資源,用戶端應用程式需要其驗證金鑰和位置:
#
# 1.在 Azure 入口網站中,您的認知服務資源之 **[金鑰和端點]** 頁面上,複製您的資源之**金鑰 1**並將其貼上到下面的程式碼,取代 **YOUR_COG_KEY**。
# 2.為您的資源複製**位置**並將其貼上到下方程式碼中,取代 **YOUR_COG_LOCATION**。
# >**備註**:留在 **[金鑰和端點]** 頁面並從此頁面複製**位置**(範例:_westus_)。請 _勿_ 在 [位置] 欄位的文字之間增加空格。
# 3.透過按一下儲存格左側的 **[執行儲存格]** (▷) 按鈕執行下方程式碼。
# + gather={"logged": 1599695377020}
cog_key = 'YOUR_COG_KEY'
cog_location = 'YOUR_COG_LOCATION'
print('Ready to use cognitive services in {} using key {}'.format(cog_location, cog_key))
# -
# ## 翻譯文字
#
# 顧名思義,**翻譯文字**服務可以讓您將文字從一種語言翻譯為另一種語言。
#
# 沒有適用於此服務的 Python SDK,但您可以使用它的 REST 介面透過 HTTP 向端點提交要求,這在 Python 中透過使用**要求**程式庫相對容易操作。關於待翻譯文字和產生的已翻譯文字之資訊以 JSON 格式交換。
#
# 執行下列儲存格以建立執行此操作的函式,然後用一個從英文到法文的簡單翻譯來測試此函式。
# + gather={"logged": 1599695393341}
# Create a function that makes a REST request to the Text Translation service
def translate_text(cog_location, cog_key, text, to_lang='fr', from_lang='en'):
import requests, uuid, json
# Create the URL for the Text Translator service REST request
path = 'https://api.cognitive.microsofttranslator.com/translate?api-version=3.0'
params = '&from={}&to={}'.format(from_lang, to_lang)
constructed_url = path + params
# Prepare the request headers with Cognitive Services resource key and region
headers = {
'Ocp-Apim-Subscription-Key': cog_key,
'Ocp-Apim-Subscription-Region':cog_location,
'Content-type': 'application/json',
'X-ClientTraceId': str(uuid.uuid4())
}
# Add the text to be translated to the body
body = [{
'text': text
}]
# Get the translation
request = requests.post(constructed_url, headers=headers, json=body)
response = request.json()
return response[0]["translations"][0]["text"]
# Test the function
text_to_translate = "Hello"
translation = translate_text(cog_location, cog_key, text_to_translate, to_lang='fr', from_lang='en')
print('{} -> {}'.format(text_to_translate,translation))
# -
# 此服務應將英文文字 "Hello" 翻譯為法文 "Bonjour"。
#
# 注意,這些語言透過使用語言縮寫標準體系來指定,*en* 表示英文,*fr* 表示法文。您也可以使用包含特定文化的縮寫,當同一語言用於不同地理區域時 (通常拼字不同),這類縮寫較為實用。例如 *en-US* 代表美式英文,而 *en-GB* 表示英式英文。
#
# 執行下列儲存格,以便在英式英文和義大利文間進行翻譯。
# + gather={"logged": 1599695400335}
text_to_translate = "Hello"
translation = translate_text(cog_location, cog_key, text_to_translate, to_lang='it-IT', from_lang='en-GB')
print('{} -> {}'.format(text_to_translate,translation))
# -
# 讓我們嘗試另一種翻譯 - 這次將美式英文翻譯為中文。
# + gather={"logged": 1599695403076}
text_to_translate = "Hello"
translation = translate_text(cog_location, cog_key, text_to_translate, to_lang='zh-CN', from_lang='en-US')
print('{} -> {}'.format(text_to_translate,translation))
# -
# ## 語音翻譯
#
# 您可以使用**語音**服務以翻譯語音語言。
#
# 現在您可以執行下列儲存格以建立和測試使用語音 SDK 翻譯聲音語音的函式。
# + gather={"logged": 1599695532629}
# 建立函數,以便將一種語言的音訊翻譯為另一種語言的文字
def translate_speech(cog_location, cog_key, audio_file=None, to_lang='fr-FR', from_lang='en-US'):
from azure.cognitiveservices.speech import SpeechConfig, AudioConfig, ResultReason
from azure.cognitiveservices.speech.translation import SpeechTranslationConfig, TranslationRecognizer
# Configure the speech translation service
translation_config = SpeechTranslationConfig(subscription=cog_key, region=cog_location)
translation_config.speech_recognition_language = from_lang
translation_config.add_target_language(to_lang)
# Configure audio input
if audio_file is None:
audio_config = AudioConfig() # Use default input (microphone)
else:
audio_config = AudioConfig(filename=audio_file) # Use file input
# Create a translation recognizer and use it to translate speech input
recognizer = TranslationRecognizer(translation_config, audio_config)
result = recognizer.recognize_once()
# Did we get it?
translation = ''
speech_text = ''
if result.reason == ResultReason.TranslatedSpeech:
speech_text = result.text
translation = result.translations[to_lang]
elif result.reason == ResultReason.RecognizedSpeech:
speech_text = result.text
translation = 'Unable to translate speech'
else:
translation = 'Unknown'
speech_text = 'Unknown'
# rturn the translation
return speech_text, translation
# Test the function
import os
file_name = 'english.wav'
file_path = os.path.join('data', 'translation', file_name)
speech, translated_speech = translate_speech(cog_location, cog_key, file_path, to_lang='es', from_lang='en-US')
result = '{} -> {}'.format(speech, translated_speech)
# Show translated text
print(result)
# -
# 注意,必須使用 2 個字元的語言程式碼來識別目標語言 (例如 *en*),而源語言必須包括文化指示 (例如 *en-US*)。
#
# 讓我們嘗試將法文翻譯為英文。
# + gather={"logged": 1599695542192}
import os
file_name = 'french.wav'
file_path = os.path.join('data', 'translation', file_name)
speech, translated_speech = translate_speech(cog_location, cog_key, file_path, to_lang='en', from_lang='fr-FR')
result = '{} -> {}'.format(speech, translated_speech)
# Show translated text
print(result)
# -
# ## 了解更多資訊
#
# 您可以在服務文件中了解關於[翻譯工具文字](https://docs.microsoft.com/azure/cognitive-services/translator/)和[使用語音服務翻譯](https://docs.microsoft.com/azure/cognitive-services/speech-service/index-speech-translation)的更多資訊。
| 09 - Translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="UxrVblUVob8t"
# # Clase 4: rudimentos de python (III)
# + [markdown] id="7XM91O8Lob8u"
# ## Errores
# + colab={"base_uri": "https://localhost:8080/"} id="Wqew-dMuob8u" outputId="0d14631a-71dc-4b4e-abb7-5669ceb48af7"
#try, except
while True:
try:
x = int(input('Por favor, ingrese un número: '))
#print(a)
break
except (ValueError, TypeError):
print('Hay un error. El dato no es un número. Escriba otro.')
except NameError:
print('La variable a no está definida.')
# + colab={"base_uri": "https://localhost:8080/"} id="av0NYALt2lSn" outputId="4abcb2f6-2232-4b51-d3f2-0536ce028134"
try:
print(a)
except NameError as e:
print(f'La variable a no está definida. Ocurrió un {e}')
# + colab={"base_uri": "https://localhost:8080/"} id="kvwLjn132vOz" outputId="eb595bcb-4270-43f3-9cb8-8b6cf73a6e3a"
try:
print(x)
except:
print('Se presentó un error.')
else:
print(f'El valor de x es {x}.')
# + colab={"base_uri": "https://localhost:8080/"} id="FMw-iP-Z7hxX" outputId="2c2857aa-81e8-471d-f20d-2bb6220d94fb"
try:
print(a)
except:
print('Se presentó un error.')
finally:
print('Esto se ejecutará sin importar qué pase.')
# + [markdown] id="_3HvcAcoob8v"
#
# ## Programación orientada a objetos
# + id="eqAAPxR-ob8w"
class Mesa:
def __init__(self, n_patas, color: str, capacidad: int, rota=False):
self.n_patas = n_patas
self.rota = rota
self.color = color
self.capacidad = capacidad
def romper(self):
if self.rota == False:
self.rota = True
else:
print('La mesa ya está rota.')
def pintar(self, color: str):
self.color = color
def apoyar(self, numero_cosas: int):
if self.capacidad >= numero_cosas:
self.capacidad = self.capacidad - numero_cosas
elif self.capacidad == 0:
print('La mesa está llena.')
else:
print(f'Solo se pueden apoyar {self.capacidad} cosas.')
def retirar(self, numero_cosas: int):
espacio_libre = self.capacidad + numero_cosas
if espacio_libre > 20:
print(f'No es posible. Solo hay {20-self.capacidad} cosas.')
else:
self.capacidad = espacio_libre
# + id="jiJ9UpfbSFMh"
| clase_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vishnu89chand/python_solve_maze/blob/master/python_solve_maze.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lh2xlq5yJW7d" colab_type="text"
# # Python solve Maze!!!
#
# * Solving maze without machine learning, in-fact no learning :P
#
# * A simple image processing approach to solve maze using morphological operations
#
# reference: [Guide to Signals and Patterns in Image Processing](https://link.springer.com/book/10.1007/978-3-319-14172-5)
#
# + id="rpI2P8BaJW7f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="072a9876-85d1-44d8-e3f7-d56748ca3d8a"
# download and install all the requirements
# set autoreload 2 to reload .py script changes
# %load_ext autoreload
# %autoreload 2
# !git clone https://github.com/vishnu89chand/python_solve_maze.git
import cv2
import numpy as np
from glob import glob
import matplotlib.pyplot as plt
from ipywidgets import interact
# + id="oMVKhD6RJW7m" colab_type="code" colab={}
# conver the image to binary
# crop maze from image
# get the thickness of maze path in pixel
def preProcessMaze(oimg):
img = cv2.cvtColor(oimg, cv2.COLOR_BGR2GRAY)
# create binary image using otsu adaptive thresholding
img = cv2.threshold(img, 0, 255, cv2.THRESH_BINARY_INV + cv2.THRESH_OTSU)[1]
# trim the image border
r,c = np.where(img==255)
x0,x1, y0, y1 = r[0], r[-1], c[0], c[-1]
img = img[y0:y1, x0:x1] # crop roi
# on four sides search no of zero pixel on border
mazePathSize = max([(img[0] == 0).sum(),(img[:,0] == 0).sum(),
(img[1] == 0).sum(), (img[:, 1] == 0).sum()])
return img, mazePathSize
# + [markdown] id="ikmTCMQcJW7q" colab_type="text"
# Hints:
# * A maze consists of walls (255) and paths (0)
# * Where the correct path split the maze into two parts.
# * Dead ends are basically paths (0) surrounded by wall (255) can be detected using closing operations
#
# Algorithm:
# 1. Select any one contour and choose a ksize (for good result choose kernel size == pathSize).
# 2. Dilate the contour. This will generate a mask with all possible paths
# 3. Perform closing operation on the image. This mask shows all the dead ends
# 4. Now find the final path by removing dead ends from the all possible paths.
#
# + id="mnej4eR5JW7r" colab_type="code" colab={}
def solveMaze(img, kSize, contourIx):
contours, hierarchy = cv2.findContours(img, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
contourImg = cv2.drawContours(np.zeros_like(img), contours, contourIx, 255, 5)
kernel = np.ones((kSize, kSize), 'u1')
# to solve maze subtract dilation with close
dilation = cv2.morphologyEx(contourImg, cv2.MORPH_DILATE, kernel)
close = cv2.morphologyEx(dilation, cv2.MORPH_ERODE, kernel) # perform closing
diff = cv2.bitwise_xor(dilation, close) # bitwise subraction
return diff, contourImg, dilation, close
# + id="7-Dphd2fJW7w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["61fdc6886382480196c213659d95ec81", "72a13b8005c24c8098d4c33b47ce16c6", "115d9cf6ad024beaa262cf32b1e51f50", "54c389de3c6a4d7d8f6f3c80ff19f888", "913db2f2b91c484b9122b4a5ff873e3e", "b474a7ad2ac4486b873bc07239d33cee", "cc4d531ebe2243c88abee49d572f78ac", "b005f2d835e1412a9998f5e28e722548", "859dcd7927c74e2ea19174c3ec3a887e", "1247edb3708e476b993f6b51563ab40f", "dec03d14f36f42359f01d01741609bd6", "<KEY>", "<KEY>", "<KEY>", "774b0ea3c3f944dbafe6a43d5762aeba", "<KEY>", "dd7651ed393545cca0a0ecbcadc00cf9", "df18fddcca3b4707bb6d3500f6a31992", "5f80f3105ec74f93b35a946d976797a2"]} outputId="1a0e0506-164c-4694-8950-7a8481515426"
def main(imPath, contourIx, enablePathSize, pathSize, figsize):
fig, axs = plt.subplots(nrows=2, ncols=3, figsize=(figsize, figsize))
oimg = cv2.imread(imPath)
dispImgs = []
img, mazePathSize = preProcessMaze(oimg)
if enablePathSize:
mazePathSize = pathSize
path, contourImg, dilation , close = solveMaze(img, mazePathSize, contourIx)
dispImgs.append(["Contour Image", contourImg.copy()])
dispImgs.append(["Dilated Image", dilation.copy()])
dispImgs.append(["Close Image", close.copy()])
dispImgs.append(["Input", oimg])
dispImgs.append(["Path", path.copy()])
img[path == 255] = 196
dispImgs.append(["Solution [size: %s]" % mazePathSize, img.copy()])
for ax, (title, img) in zip(axs.flat, dispImgs):
ax.imshow(img, interpolation='bilinear')
ax.set_title(title)
ax.axis('off')
plt.show()
print("Before using track bar please enablePathSize")
a = interact(main, imPath=glob('python_solve_maze/maze*.*'),contourIx=[0,1], enablePathSize=False, pathSize=(1,100), figsize=(10, 26))
# + id="i6QF3UfxJW70" colab_type="code" colab={}
| python_solve_maze.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Intro to Jupyter Notebooks
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### `Jupyter` is a project for developing open-source software
# ### `Jupyter Notebooks` is a `web` application to create scripts
# ### `Jupyter Lab` is the new generation of web user interface for Jypyter
# + [markdown] slideshow={"slide_type": "subslide"}
# ### But it is more than that
# #### It lets you insert and save text, equations & visualizations ... in the same page!
# 
# -
# ***
# + [markdown] slideshow={"slide_type": "slide"}
# # Notebook dashboard
# When you launch the Jupyter notebook server in your computer, you would see a dashboard like this:
#
# 
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Saving your own script
# All scripts we are showing here today are running online & we will make changes through the workshop. To keep your modified script for further reference, you will need to save a copy on your own computer at the end.
#
# <div class="alert alert-block alert-info">
# <b>Try it out! </b>
# <br><br>
# Go to <b>File</b> in the top menu -> Download As -> Notebook </div>
# <br>
# Any changes made online, even if saved (not downloaded) will be lost once the binder connection is closed.
# -
# ***
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Two type of cells
# ### `Code` Cells: execute code
# ### `Markdown` Cells: show formated text
#
# There are two ways to change the type on a cell:
# - Cliking on the scroll-down menu on the top
#
# - using the shortcut `Esc-y` for code and `Esc-m` for markdown types
# <br>
# <div class="alert alert-block alert-info"><b>Try it out! </b>
# <bR>
# <br>- Click on the next cell
# <br>- Change the type using the scroll-down menu & select <b>Code</b>
# <br>- Change it back to <b>Markdown</b>
# </div>
# -
# ## This is a simple operation
# y = 4 + 6
# print(y)
# ## <i>Note the change in format of the first line & the text color in the second line</i>
# <div class="alert alert-block alert-info"><b>Try it out!</br>
# <br><br>In the next cell:
# <br>- Double-Click on the next cell
# <br>- Press <b> Esc</b> (note to blue color of the left border)
# <br>- Type <b>y</b> to change it to <b>Code</b> type
# <br>- Use <b>m</b> to change it back to <b>Markdown</b> type
# </div>
# This is a simple operation
y = 4 + 6
print(y)
# ***
# + [markdown] slideshow={"slide_type": "slide"}
# # To execute commands
#
# ## - `Shift-Enter` : executes cell & advance to next
# ## - `Control-enter` : executes cell & stay in the same cell
#
# <div class="alert alert-block alert-info"><b>Try it out!</b>
# <br>
# <br>In the previous cell:
# <br>- Double-Click on the previous cell
# <br>- Use <b>Shift-Enter</b> to execute
# <br>- Double-Click on the in the previous cell again
# <br>- This time use <b>Control-Enter</b> to execute
# <br>
# <br>- Now change the type to <b>Code</b> & execute the cell
# </div>
# -
# ## You could also execute the entire script use the `Run` tab in the top menu
#
#
# ## Or even the entire script from the `Cell` menu at the top
# ***
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Other commands
# ### From the icon menu:
# ### Save, Add Cell, Cut Cell, Copy Cell, Paste Cell, Move Cell Up, Move Cell Down
#
# 
#
# ### or the drop down menu 'command palette'
# -
# <div class="alert alert-block alert-info"><b>Try them out!</b>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Now, the keyboard shortcuts
# #### First press `Esc`, then:
# - `s` : save changes
# <br>
# - `a`, `b` : create cell above and below
# <br>
# - `dd` : delete cell
# <br>
# - `x`, `c`, `v` : cut, copy and paste cell
# <br>
# - `z` undo last change
# + [markdown] slideshow={"slide_type": "fragment"}
# <div class="alert alert-block alert-info">
# <b> Let's practice!</b>
# <br>
# <br>- Create a cell bellow with <b>Esc-b</b>, and click on it
# <br>- Type print('Hello world!') and execute it using <b>Control-Enter</b>
# <br>- Copy-paste the cell to make a duplicate by typing <b>Esc-c</b> & <b>Esc-v</b>
# <br>- Cut the first cell using <b>Esc-x</b>
# </div>
# -
# ## And the last one: adding line numbers
# - `Esc-l` : in Jupyter Notebooks
# - `Esc-Shift-l`: in Jupyter Lab
#
# <div class="alert alert-block alert-info">
# <b>Try it out!</b>
# <br><br>
# - Try it in a code cell
# <br>- And now try it in the markdown cell
# </div>
y = 5
print(y + 4)
x = 8
print(y*x)
# ***
# ## Last note about the `Kernel`
# #### That little program that is running in the background & let you run your notebook
# <div class="alert alert-block alert-danger">
# Once in a while the <b>kernel</b> will die or your program will get stucked, & like everything else in the computer world.... you'll have to restart it.
# </div>
#
# ### You can do this by going to the `Kernel` menu -> Restart, & then you'll have to run all your cells (or at least the ones above the one you're working on (use `Cell` menu -> Run all Above.
| notebooks/.ipynb_checkpoints/Intro_02_JupyterNotebooks-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classifier comparison
# +
print(__doc__)
# Code source: <NAME>
# <NAME>
# Modified for documentation by <NAME>
# License: BSD 3 clause
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.datasets import make_moons, make_circles, make_classification
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
h = .02 # step size in the mesh
names = ["Nearest Neighbors", "Linear SVM", "RBF SVM", "Gaussian Process",
"Decision Tree", "Random Forest", "Neural Net", "AdaBoost",
"Naive Bayes", "QDA"]
classifiers = [
KNeighborsClassifier(3),
SVC(kernel="linear", C=0.025),
SVC(gamma=2, C=1),
GaussianProcessClassifier(1.0 * RBF(1.0)),
DecisionTreeClassifier(max_depth=5),
RandomForestClassifier(max_depth=5, n_estimators=10, max_features=1),
MLPClassifier(alpha=1, max_iter=1000),
AdaBoostClassifier(),
GaussianNB(),
QuadraticDiscriminantAnalysis()]
X, y = make_classification(n_features=2, n_redundant=0, n_informative=2,
random_state=1, n_clusters_per_class=1)
rng = np.random.RandomState(2)
X += 2 * rng.uniform(size=X.shape)
linearly_separable = (X, y)
datasets = [make_moons(noise=0.3, random_state=0),
make_circles(noise=0.2, factor=0.5, random_state=1),
linearly_separable
]
figure = plt.figure(figsize=(27, 9))
i = 1
# iterate over datasets
for ds_cnt, ds in enumerate(datasets):
# preprocess dataset, split into training and test part
X, y = ds
X = StandardScaler().fit_transform(X)
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=.4, random_state=42)
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
# just plot the dataset first
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
if ds_cnt == 0:
ax.set_title("Input data")
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.6,
edgecolors='k')
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
i += 1
# iterate over classifiers
for name, clf in zip(names, classifiers):
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
ax.contourf(xx, yy, Z, cmap=cm, alpha=.8)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright,
edgecolors='k', alpha=0.6)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
if ds_cnt == 0:
ax.set_title(name)
ax.text(xx.max() - .3, yy.min() + .3, ('%.2f' % score).lstrip('0'),
size=15, horizontalalignment='right')
i += 1
plt.tight_layout()
plt.show()
# -
| pittsburgh-bridges-data-set-analysis/resources/scikit-learn-examples/Classifier comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Perform $k$-means on the data
# +
# Load packages
import numpy as np
import pickle
from FDApy.representation.functional_data import DenseFunctionalData
from FDApy.representation.functional_data import MultivariateFunctionalData
from skfda import FDataGrid
from skfda.ml.clustering import KMeans
# -
# Load data
with open('./data/canadian_smooth.pkl', 'rb') as f:
data_fd = pickle.load(f)
# +
# Format data for skfda
temperature = data_fd[0].values
precipitation = data_fd[1].values
# skfda only accept data with same shape
new_prec = np.hstack([precipitation,
precipitation[:, -1][:, np.newaxis]])
# -
# Create FDataGrid object
data_matrix = np.stack([temperature, new_prec], axis=-1)
sample_points = data_fd[0].argvals['input_dim_0']
fdata = FDataGrid(data_matrix, sample_points)
# Compute derivatives
fdata_derivatives = fdata.derivative(order=1)
# Perform k-means
res = {}
for i in np.arange(2, 9, 1):
kmeans = KMeans(n_clusters=i)
kmeans.fit(fdata)
res[i] = kmeans.predict(fdata)
res[4]
with open('./results/results_weather_kmeans_d1.pkl', 'wb') as f:
pickle.dump(res, f)
# Perform k-means on derivatives
res_derivative = {}
for i in np.arange(2, 9, 1):
kmeans = KMeans(n_clusters=i)
kmeans.fit(fdata_derivatives)
res_derivative[i] = kmeans.predict(fdata_derivatives)
res_derivative[4]
with open('./results/results_weather_kmeans_d2.pkl', 'wb') as f:
pickle.dump(res_derivative, f)
| canadian_weather/04-kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.12 ('flyai_pytorch1_5')
# language: python
# name: python3
# ---
# # 上下文管理器和else
#
# with语句可以创建一个临时的上下文管理器对象管理上下文。with语句使得一些代码的运行更安全,例如最常见的,利用with语句读写文件,with语句能够正确的启动以及关闭数据流。
#
# else子句除了在if语句中使用外,还能够和for、while以及try语句搭配使用。for/else、while/else和try/else中的else子句和if/else中的else子句相差甚远,因此有必要详细讨论这一区别。
# ## else子句
#
# else子句和不同的语句搭配有不同的效果:
#
# 1. if
# 仅当if块以及elif块的条件均不满足时才运行else子句
# 2. for
# 仅当for循环运行完毕时(for循环没有被break终止)才运行else子句
# 3. while
# 仅当while循环正常退出时(while循环没有被break终止)才运行else子句
# 4. try
# 仅当try块中没有异常抛出时才运行else块,并且else块中抛出的异常不会由前面的except子句处理
#
# 从上述描述可以发现,仅有if语句中else子块表示互斥的概念,在if/else中else子块具有排他性。其他三个语句中均表示了先后的关系,即先完成前置的循环/测试然后再运行else子句中的代码
#
# else子句在除if/else之外的语句中使用略显多余,并且其语义和惯常理解的语义有非常大的区别。当然,else子句还是有一些用途的
#
# 例如在循环中可以使用else语句代替if,以避免设置多余的控制标识,例如下述的char_check例子。
# +
def char_check1(input_text):
char_flag = False
for _char in input_text:
if _char == "a":
char_flag = True
break
if not char_flag:
print("没有发现字母a")
def char_check2(input_text):
for _char in input_text:
if _char == "a":
break
else:
print("没有发现字母a")
test_text = "dsfcvsd"
char_check1(test_text)
char_check2(test_text)
# -
# 在try语句中else子句承担逻辑上的要求:try语句用于捕获特定代码可能抛出的错误,因此try块不应该捕获后续代码抛出的错误,此时应当在else语句中运行后续代码。本书有一个非常好的例子用于说明try/else语句。下述伪代码中dangerous_call()表示应当捕获错误的代码,after_call()表示后续代码:
#
# 不正确的使用方式如下:
# ```Python
# try:
# dangerous_call()
# after_call()
# except ErrorType:
# ...
# ```
# 正确的使用方式:
# ```Python
# try:
# dangerous_call()
# except ErrorType:
# ...
# else:
# after_call()
# ```
# ## EAFP & LBYL
#
# EAFP和LBYL是两种风格,表示了处理错误以及控制流程的方式。
#
# * EAFP(easier to ask for forgiveness than permission)
# 不管如何,先运行,然后处理错误。
# Python中常用这种方式。很多Python程序会首先假定代码能够正常运行,若假定不成立,抛出错误,那就捕获这个错误,然后处理。
# * LBYL(look before you leap)
# 不同于EAFP,LBYL在运行特定代码前会执行各类检查以及分派以尽可能确保代码能够正常运行。
#
# LBYL在多线程中可能引起意外错误。例如通过if检查某个键是否存在,然后从字典中读取该键对应的值;对于多线程,很可能在线程A中通过了if检查,但是线程B删除了该键值对导致运行失败
# ## with与上下文管理器
#
# with语句旨在简化try/finally模式。with语句能够保证一段代码在运行前以及运行后执行某些操作,无论这段代码是否正常运行。显然,这种逻辑能够释放一些重要的资源,或者还原临时变更的状态。
#
# with语句会创建一个上下文管理器用于管理和实施上述操作。具体来说,上下文管理器会在with语句开始时调用__enter__方法,并在with语句结束时调用__exit__方法。值得注意的是,不同于函数,with语句并没有定义新的作用域,with语句中创建的变量在with语句结束后依然可以调用。
#
# 本章给出了一个非常好的例子,该例子用于说明with语句的应用以及一些注意事项:
# +
class LookingGlass:
def __enter__(self):
import sys
self.original_write = sys.stdout.write
sys.stdout.write = self.reverse_write
return "JABBERWOCKY"
def reverse_write(self, text):
self.original_write(text[::-1])
def __exit__(self, exc_type, exc_value, traceback):
import sys
sys.stdout.write = self.original_write
if exc_type is ZeroDivisionError:
print("Please DO NOT divide by zero!")
return True
with LookingGlass() as what:
print("Alice, Kitty and snowdrop")
print(what)
print("Alice, Kitty and snowdrop")
print(what)
# -
# 上述例子揭示了with语句的很多注意事项:
# 1. with语句开始时会创建上下文管理器并且调用上下文管理器的__enter__方法
# 2. __enter__方法完全可以修改环境,而不仅仅是进行一些数据/数据流上的工作,例如上述例子中利用自定义的函数接管了标准输出方式
# 3. __enter__方法的返回值可以任选,并不一定要返回上下文管理器自身,甚至可以不返回任何东西,虽然在很多应用中都是返回self
# 4. __exit__方法可以接收with语句运行过程中抛出的错误并对这些错误进行处理
# ## contextlib中的实用工具
#
# contextlib模块提供了一些适用于特定场合的类和函数,本章列举了一些。
#
# 1. closing
# 若对象提供了close()方法,而没有实现__enter__/__exit__,则可以使用closing函数构建一个上下文管理器。显然closing()函数适用于仅在退出with语句时进行特殊处理的应用场合
# 2. suppress
# suppress可用于构建临时忽略指定异常的上下文管理器
# 3. @contextmanager
# 该装饰器可以将生成器函数变为上下文管理器,从而避免创建类
# 4. ContextDecorator
# ContextDecorator是个基类,用于定义基于类的上下文管理器
# 5. ExitStack
# 能够管理多个上下文管理器的__exit__运行顺序 —— 按照先进后出的顺序调用各上下文管理器的__exit__方法。具体来说ExitStack可以创建管理上下文管理器的对象,该对象具有enter_context方法用于启动一个上下文管理器,在退出with语句时会按照先进后出的顺序调用这个上下文管理器的__exit__方法
#
# ### @contextmanager
#
# @contextmanager能够将一个生成器“变为”一个上下文管理器。具体来说,被@contextmanager装饰的生成器会以yield为界,yield语句前的部分相当于__enter__方法,会在with语句开始时被调用;yield语句后的部分相当于__exit__方法,会在with语句结束时调用(吐槽:更像断点调试了)
#
# 利用@contextmanager可以将上述的LookingGlass改写为如下函数,功能完全一致。
# +
import contextlib
@contextlib.contextmanager
def looking_glass():
import sys
original_write = sys.stdout.write
def reverse_write(text):
original_write(text[::-1])
sys.stdout.write = reverse_write
msg = ""
try:
yield "JABBERWOCKY"
except ZeroDivisionError:
msg = "Please DO NOT divide by zero!"
finally:
sys.stdout.write = original_write
if msg:
print(msg)
with looking_glass() as what:
print("Alice, Kitty and snowdrop")
print(what)
print("Alice, Kitty and snowdrop")
print(what)
# -
# 上述with语句的调用过程如下:
#
# 1. @contextmanager会将生成器包装成实现__enter__和__exit__方法
# 2. 对于__enter__方法
# 首先会调用生成器函数并保存生成器对象gen
# 然后调用next(gen)并且获取返回值以绑定到with/as中
# 3. 对于__exit__方法
# 首先会检查是否有抛出异常,若抛出异常则调用gen.throw(exception)并在yield位置抛出
# 若没有抛出异常则继续运行next(gen)以运行yield语句后的部分
#
# 值得注意的是,若在生成器中没有使用try/finally语句处理在yield位置抛出的错误,那么yield后续的代码不会运行。对于looking_glass,yield后续代码无法运行意味着无法恢复标准输出方式,这可能导致致命问题。
# ## 总结
#
# 1. else子句和不同的语句搭配会有不同的效果,在if语句中else子句表示了一种互斥的概念,而在for/while/try语句中else子句则表示了一种先后关系
# 2. try语句中的else子句有特殊的作用。为了尽可能仅对特定代码抛出的错误进行捕获和处理,try语句中应当仅放入特定代码,其余代码则放置到else语句
# 3. EAFP和LBYL是两种代码风格,Python作为一种弱类型语言,其趋向于使用EAFP,即首先假定代码能正常运行,若运行过程中报错则处理错误
# 4. with语句的正确使用能够创建上下文管理器接管一些重要设置的开启和关闭。with语句能够保证在一段代码执行前后执行一些必要操作,这些操作能够获取/释放一些重要资源,或者改变/还原临时变更的状态
# 5. with语句创建的上下文管理器依靠__enter__和__exit__执行相应的功能。在进入with语句时,解释器会自动调用并运行上下文管理器的__enter__方法,在退出with语句时,解释器则会自动调用并运行上下文管理器的__exit__方法
# 6. __enter__方法可以有返回值,也可以没有返回值;__exit__方法则可以接收并处理with语句运行过程中抛出的错误
# 7. contextlib模块中提供了一些适用于特定场合的类以及函数。其中@contextmanager管理器能够将一个生成器变为一个上下文管理器。具体来说,以yield为界,yield语句前的部分相当于__enter__方法,会在with语句开始时被调用;yield语句后的部分相当于__exit__方法,会在with语句结束时调用。值得注意的是,若yield之前的语句抛出错误,yield之后的语句不会运行,这在一些应用中可能导致致命错误
| 15_ContextManagersAndSomethingElse/15_ContextManagersAndSomethingElse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing Step
#
# ! pip install -U pip
# ! pip install -U clearml==0.16.2rc0
# ! pip install -U pandas==1.0.4
# ! pip install -U numpy==1.18.4
# +
import pandas as pd
import numpy as np
from collections import Counter
from clearml import Task
# -
# ## Configure Task
# Instantiate a ClearML Task using `Task.init`.
#
# A Configuration dictionary is connected to the task using `Task.connect`. This will enable the [pipeline controller](https://github.com/allegroai/clearml/blob/master/examples/frameworks/pytorch/notebooks/table/tabular_ml_pipeline.ipynb) to access this task's configurations and override the value when the pipeline is executed.
task = Task.init(project_name="Tabular Example", task_name="tabular preprocessing")
logger = task.get_logger()
configuration_dict = {
"data_task_id": "39fbf86fc4a341359ac6df4aa70ff91b",
"fill_categorical_NA": True,
"fill_numerical_NA": True,
}
configuration_dict = task.connect(
configuration_dict
) # enabling configuration override by clearml
print(
configuration_dict
) # printing actual configuration (after override in remote mode)
# ## Get Data
#
# ClearML retrieves that data which will be processed. First, the data task is fetched using `Task.get_task` and inputting the task's ID from the configuration dictionary. Then the data task's artifacts are accessed in order to retrieve the training and validations sets.
#
data_task = Task.get_task(configuration_dict.get("data_task_id"))
train_set = data_task.artifacts["train_data"].get().drop(columns=["Unnamed: 0"])
val_set = data_task.artifacts["val_data"].get().drop(columns=["Unnamed: 0"])
logger.report_table(
title="Trainset - raw",
series="pandas DataFrame",
iteration=0,
table_plot=train_set.head(),
)
# ## Preprocess Data
# +
# Remove hour and year from DateTime data
def change_time_format(data_frame):
timestamp = pd.to_datetime(data_frame["DateTime"])
months = [d.month for d in timestamp]
data_frame["Month"] = pd.DataFrame(months).astype("object")
data_frame.drop(columns=["DateTime"], inplace=True)
return data_frame
train_set = change_time_format(train_set)
val_set = change_time_format(val_set)
# +
def change_age_format(data_frame):
age = data_frame["AgeuponOutcome"]
months_age = []
for val in age:
if pd.isnull(val):
months_age.append(val)
else:
amount, time_type = val.split(" ")
if "day" in time_type:
mult = 1.0 / 30
if "week" in time_type:
mult = 1.0 / 4
if "month" in time_type:
mult = 1.0
if "year" in time_type:
mult = 12.0
months_age.append(int(amount) * mult)
data_frame["Age"] = pd.DataFrame(months_age).astype(np.float32)
data_frame.drop(columns=["AgeuponOutcome"], inplace=True)
return data_frame
train_set = change_age_format(train_set)
val_set = change_age_format(val_set)
# +
def change_sex_format(data_frame):
sex_neutered = data_frame["SexuponOutcome"]
sex = []
neutered = []
for val in sex_neutered:
if pd.isnull(val):
sex.append(val)
neutered.append(val)
elif "Unknown" in val:
sex.append(np.nan)
neutered.append(np.nan)
else:
n, s = val.split(" ")
if n in ["Neutered", "Spayed"]:
neutered.append("Yes")
else:
neutered.append("No")
sex.append(s)
data_frame["Sex"] = pd.DataFrame(sex)
data_frame["Neutered"] = pd.DataFrame(neutered)
data_frame.drop(columns=["SexuponOutcome"], inplace=True)
return data_frame
train_set = change_sex_format(train_set)
val_set = change_sex_format(val_set)
# +
# Remove irrelevant columns
def remove_columns(data_frame, list_columns_names=None):
if list_columns_names is not None:
data_frame.drop(columns=list_columns_names, inplace=True)
return data_frame
train_set = remove_columns(train_set, ["Name", "OutcomeSubtype", "AnimalID"])
val_set = remove_columns(val_set, ["Name", "OutcomeSubtype", "AnimalID"])
logger.report_table(
title="Trainset - after preprocessing",
series="pandas DataFrame",
iteration=0,
table_plot=train_set.head(),
)
# -
# ## *Fill NA Values*
object_columns = train_set.select_dtypes(include=["object"]).copy()
numerical_columns = train_set.select_dtypes(include=["number"]).copy()
# Notice that the configuration dictionary is accessed below to access `fill_categorical_NA`'s value. This value can be overridden by the pipeline controller.
if configuration_dict.get("fill_categorical_NA", True):
for col in object_columns.columns:
if object_columns[col].isnull().sum() > 0:
most_common = Counter(object_columns[col]).most_common(1)[0][0]
print(
'Column "{}": replacing null values with "{}"'.format(col, most_common)
)
train_set[col].fillna(most_common, inplace=True)
val_set[col].fillna(most_common, inplace=True)
# Notice that the configuration dictionary is accessed below to access `fill_numerical_NA`'s value. This value can be overridden by the pipeline controller.
if configuration_dict.get("fill_numerical_NA", True):
for col in numerical_columns.columns:
if numerical_columns[col].isnull().sum() > 0:
median_val = numerical_columns[col].median()
print(
'Column "{}": replacing null values with "{}"'.format(col, median_val)
)
train_set[col].fillna(median_val, inplace=True)
val_set[col].fillna(median_val, inplace=True)
# Drop rows with NA values if were chosen not to be filled
train_set.dropna(inplace=True)
val_set.dropna(inplace=True)
if configuration_dict.get("fill_categorical_NA", True) or configuration_dict.get(
"fill_numerical_NA", True
):
logger.report_table(
title="Trainset - after filling missing values",
series="pandas DataFrame",
iteration=0,
table_plot=train_set.head(),
)
# ## *Labels Encoding*
all_data = pd.concat([train_set, val_set])
outcome_categories = all_data["OutcomeType"].astype("category").cat.categories
outcome_dict = {key: val for val, key in enumerate(outcome_categories)}
task.upload_artifact("Outcome dictionary", outcome_dict)
for col in object_columns.columns:
all_data[col] = all_data[col].astype("category").cat.codes
train_set = all_data.iloc[: len(train_set.index), :]
val_set = all_data.iloc[len(train_set.index) :, :]
logger.report_table(
title="Trainset - after labels encoding",
series="pandas DataFrame",
iteration=0,
table_plot=train_set.head(),
)
# making all variables categorical
object_columns_names = object_columns.drop(columns=["OutcomeType"]).columns
for col in object_columns_names:
all_data[col] = all_data[col].astype("category")
columns_categories = {
col: len(all_data[col].cat.categories) for col in object_columns_names
}
task.upload_artifact("Categories per column", columns_categories)
task.upload_artifact("train_data", artifact_object=train_set)
task.upload_artifact("val_data", artifact_object=val_set)
| examples/frameworks/pytorch/notebooks/table/preprocessing_and_encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# orphan: true
# ---
# (tune-rllib-example)=
#
# # Using RLlib with Tune
#
# ```{image} /rllib/images/rllib-logo.png
# :align: center
# :alt: RLlib Logo
# :height: 120px
# :target: https://docs.ray.io
# ```
#
# ```{contents}
# :backlinks: none
# :local: true
# ```
#
# ## Example
#
# Example of using PBT with RLlib.
#
# Note that this requires a cluster with at least 8 GPUs in order for all trials
# to run concurrently, otherwise PBT will round-robin train the trials which
# is less efficient (or you can set {"gpu": 0} to use CPUs for SGD instead).
#
# Note that Tune in general does not need 8 GPUs, and this is just a more
# computationally demanding example.
# +
import random
from ray import tune
from ray.tune.schedulers import PopulationBasedTraining
if __name__ == "__main__":
# Postprocess the perturbed config to ensure it's still valid
def explore(config):
# ensure we collect enough timesteps to do sgd
if config["train_batch_size"] < config["sgd_minibatch_size"] * 2:
config["train_batch_size"] = config["sgd_minibatch_size"] * 2
# ensure we run at least one sgd iter
if config["num_sgd_iter"] < 1:
config["num_sgd_iter"] = 1
return config
pbt = PopulationBasedTraining(
time_attr="time_total_s",
perturbation_interval=120,
resample_probability=0.25,
# Specifies the mutations of these hyperparams
hyperparam_mutations={
"lambda": lambda: random.uniform(0.9, 1.0),
"clip_param": lambda: random.uniform(0.01, 0.5),
"lr": [1e-3, 5e-4, 1e-4, 5e-5, 1e-5],
"num_sgd_iter": lambda: random.randint(1, 30),
"sgd_minibatch_size": lambda: random.randint(128, 16384),
"train_batch_size": lambda: random.randint(2000, 160000),
},
custom_explore_fn=explore,
)
analysis = tune.run(
"PPO",
name="pbt_humanoid_test",
scheduler=pbt,
num_samples=1,
metric="episode_reward_mean",
mode="max",
config={
"env": "Humanoid-v1",
"kl_coeff": 1.0,
"num_workers": 8,
"num_gpus": 0, # number of GPUs to use
"model": {"free_log_std": True},
# These params are tuned from a fixed starting value.
"lambda": 0.95,
"clip_param": 0.2,
"lr": 1e-4,
# These params start off randomly drawn from a set.
"num_sgd_iter": tune.choice([10, 20, 30]),
"sgd_minibatch_size": tune.choice([128, 512, 2048]),
"train_batch_size": tune.choice([10000, 20000, 40000]),
},
)
print("best hyperparameters: ", analysis.best_config)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## More RLlib Examples
#
# - {doc}`/tune/examples/includes/pb2_ppo_example`:
# Example of optimizing a distributed RLlib algorithm (PPO) with the PB2 scheduler.
# Uses a small population size of 4, so can train on a laptop.
| doc/source/tune/examples/pbt_ppo_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hypothesis and Inference
# ## Example: FLipping a Coin
from typing import Tuple
import numpy as np
import math
# +
# Distribucion noramal de una variable aleatoria
def normal_cdf(x:float, mu: float = 0, sigma: float = 1) -> float:
return (1 + math.erf((x-mu)/math.sqrt(2)/sigma)) /2
def inverse_normal_cdf(p:float,
mu: float = 0,
sigma: float = 1,
tolerance: float = 0.00001) -> float:
"""find approximate inverse using binary search"""
# if not standard, compute standard and rescale
if mu != 0 or sigma != 1:
return mu + sigma * inverse_normal_cdf(p, tolerance=tolerance)
low_z, low_p = -10.0, 0 # normal_cdf(-10) is (very close to) 0
hi_z, hi_p = 10.0, 1 # normal_cdf(10) is (very close to) 1
while hi_z - low_z > tolerance:
mid_z = (low_z + hi_z) / 2 # consider the midpoint
mid_p = normal_cdf(mid_z) # and the cdf's value there
if mid_p < p:
# midpoint is still too low, search above it
low_z, low_p = mid_z, mid_p
elif mid_p > p:
# midpoint is still too high, search below it
hi_z, hi_p = mid_z, mid_p
else:
break
return mid_z
# +
def normal_approximation_to_binomial(n, p):
"""finds mu and sigma corresponding to a Binomial(n, p)"""
mu = p * n
sigma = math.sqrt(p * (1 - p) * n)
return mu, sigma
#####
#
# probabilities a normal lies in an interval
#
######
# the normal cdf _is_ the probability the variable is below a threshold
normal_probability_below = normal_cdf
# it's above the threshold if it's not below the threshold
def normal_probability_above(lo, mu=0, sigma=1):
return 1 - normal_cdf(lo, mu, sigma)
# it's between if it's less than hi, but not less than lo
def normal_probability_between(lo, hi, mu=0, sigma=1):
return normal_cdf(hi, mu, sigma) - normal_cdf(lo, mu, sigma)
# it's outside if it's not between
def normal_probability_outside(lo, hi, mu=0, sigma=1):
return 1 - normal_probability_between(lo, hi, mu, sigma)
######
#
# normal bounds
#
######
def normal_upper_bound(probability, mu=0, sigma=1):
"""returns the z for which P(Z <= z) = probability"""
return inverse_normal_cdf(probability, mu, sigma)
def normal_lower_bound(probability, mu=0, sigma=1):
"""returns the z for which P(Z >= z) = probability"""
return inverse_normal_cdf(1 - probability, mu, sigma)
def normal_two_sided_bounds(probability, mu=0, sigma=1):
"""returns the symmetric (about the mean) bounds
that contain the specified probability"""
tail_probability = (1 - probability) / 2
# upper bound should have tail_probability above it
upper_bound = normal_lower_bound(tail_probability, mu, sigma)
# lower bound should have tail_probability below it
lower_bound = normal_upper_bound(tail_probability, mu, sigma)
return lower_bound, upper_bound
def two_sided_p_value(x, mu=0, sigma=1):
if x >= mu:
# if x is greater than the mean, the tail is above x
return 2 * normal_probability_above(x, mu, sigma)
else:
# if x is less than the mean, the tail is below x
return 2 * normal_probability_below(x, mu, sigma)
def count_extreme_values():
extreme_value_count = 0
for _ in range(100000):
num_heads = sum(1 if random.random() < 0.5 else 0 # count # of heads
for _ in range(1000)) # in 1000 flips
if num_heads >= 530 or num_heads <= 470: # and count how often
extreme_value_count += 1 # the # is 'extreme'
return extreme_value_count / 100000
upper_p_value = normal_probability_above
lower_p_value = normal_probability_below
# -
mu_0, sigma_0 = normal_approximation_to_binominal(1000, 0.5)
lower_bound, upper_bound = normal_two_sided_bounds(0.95, mu_0, sigma_0)
print(lower_bound, upper_bound)
# ## p-Values
# +
def two_sided_p_values(x: float, mu: float = 0, sigma: float = 1) -> float:
if x >= mu:
# Si x es mayor que el promedio, entonces la cola es todo mayor que x
return 2 * normal_probability_above(x, mu, sigma)
else:
# Si x es menor que el promedio, entonces la cosa es toda menor que x
return 2 * normal_probability_below(x, mu, sigma)
result = two_sided_p_value(529.5, mu_0, sigma_0)
print(result)
# -
import random
extreme_value_count = 0
for _ in range(1000):
num_heads = sum(1 if random.random() < 0.5 else 0
for _ in range(1000))
if num_heads
| python/data_scientist_from_scratch/7_hypothesis_and_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Constantes en Python
import math
math.pi
math.tau
math.e
2*math.pi
tau = 2*math.pi
math.inf
-math.inf
float('inf') # el float tiene 8 dígitos exactos mientras que los dbls tiene doble precisión
nan
math.nan # nan: not a number
float('nan')
sqrt(-1,0) # ojo con los errores
math.log(10)
math.exp(10) # exponente
math.pow(4,2) # potencia
2 + 4
math.hypot(math.nan, math.inf)
# # Representación numérica
2+2
3*5
3/5
# +
# $\int_0^1 x^2 dx$
# -
math.ceil(3.4523)
math.floor(3.4523)
math.trunc(9.145)
math.copysign(3, -2)
math.copysign(3, -0.0) # colocar los números con punto
math.fabs(-5)
2**3 # '**' es la potencia
math.factorial(4)
# +
x = 5
y = 2
math.factorial(x)/ (math.factorial(y) * math.factorial(x - y)) # ojo que hay librerías como el choose de r
# -
math.fmod(7,3) # me da el resto de la división
7%3 # trabaja con números enteros
7//3 # cociente
math.remainder(7,3) # el resto de la división
math.modf(-4.25) # separa la parte entera de la decimal
math.gcd(24,36) # Máximo común divisor
math.isfinite(4.6)
math.isinf(4.5)
math.isnan(4.5)
math.sqrt(2)**2 == 2
math.isclose(math.sqrt(2)**2,2, rel_tol = 1e-09) # rel_tol define la precisión o cercanía
math.sqrt(2)**2
# # Funciones Matemáticas
math.exp(3) # fn exponencial, esta función es más precisa que hacer la potencia
math.e**3 # diferencias en los resultados
math.pow(math.e, 3) # fíjate que los resultados son similares
math.expm1(1) # corresponde al número exponencial menos 1
math.exp(1) - 1 # es una manera de hacerlo, pero se pierde precisión (según el instructor, se ve que es igual; USA LA FN ANTERIOR)
math.expm1(1e-05) # esta es más precisa
math.exp(1e-05) - 1 # acá se puede notar digferencias con respecto a la expresión anterior
math.log(12) # si no definimos el argumento en esta fn el programa lo tomará como el log neperiano
math.log(1000,10) # esta es la manera correcta de precisarlo log(arg, base)
math.log1p(1e-5) # esta es más precisa en caso de ln(1 + x)
math.log2(32) # log(32,2)
math.log(32,2) #
math.log10(1000000)
math.sin(180) # ojo que está en radianes
math.cos(180)
math.cos(math.pi)
math.tan(math.pi/2) # es un número muy grande, fíjate que se acerca a infinito
math.asin(1) # Resultado en radianes
math.acos(1)
math.atan(1) # resultado en radianes
math.degrees(0.7853981633974483) # convierte los RADIANES a GRADOS
math.radians(60) # Una transformación de GRADOS a Radianes
math.cos(math.radians(60)) # Una transformación conjunta
math.hypot(3,4) # qué tan largoi es un vector. es la norma raiz de la suma de los cuadrados
math.sqrt(3**2 + 4**2)
math.atan2(4,3) # hay que dar el eje de la y y luego el de la x, el resultado está entre [-pi, pi]
math.degrees(math.atan2(4,3)) # construir el angulo de coordenadas (y,x)
math.sinh(0) # funciones hiperbólicas
math.cosh(0)
math.tanh(0)
math.erf(0) # función de error
math.erf(math.pi)
math.erfc(math.pi) # función complementaria
math.gamma(6) # factorial generalizado gamma
math.lgamma(5)
| scripts/tema1/05-math-pythonADIPIZ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyspark.sql
import pyspark.sql.functions as sf
spark = pyspark.sql.SparkSession.Builder().getOrCreate()
# # 1 Fictional Sales Data
#
# In this example we use a fictional data set of company revenues. The special property of this data set is that a company can have a different company as its parent company. Eventually a business expert wants to see the whole revenue of a company including all child companies. This requires that we build up an additional table containing all children (direct and indirect) for every company, such that we can join the revenues against this table and then aggregate over all direct and indirect children for each parent.
# Let's start by loading and inspecting the data.
basedir = "s3://dimajix-training/data"
# +
data = spark.read \
.option("header", True) \
.option("inferSchema", True) \
.csv(basedir + "/global-sales.csv")
data.printSchema()
# -
data.toPandas()
# # 2 Single Step of transitive parent-child relations
#
# In the next step we want to build the helper table containing all children for every company. We will calculate this table using an iterative algorithm which adds the next level of children in every iteration. We first implement a single iteration, which will add the next level of children to each parent company.
# Remove all records without a parent company for the algorithm
cleaned_df = data \
.filter(data["parent_company"].isNotNull()) \
.select(data["company"], data["parent_company"])
def iterate_parent_child(df):
# Denote the incoming table "parent" and "child", since we will do a self-join and the join condition would be ambigious without aliases otherwise
parent_df = df.alias("parent")
child_df = df.alias("child")
# Calculate next levels of indirect children by joining the table to itself and by retrieving the child of each child of each parent
next_level = parent_df.join(child_df, sf.col("parent.company") == sf.col("child.parent_company"), "inner") \
.select(sf.col("parent.parent_company"), sf.col("child.company"))
# Add current relations, otherwise they will be lost
cur_level = parent_df.select(parent_df["parent_company"], parent_df["company"])
# Return union of next indirection and current relations
return next_level.union(cur_level).distinct()
# ### Perform one iteration
#
# Now let us perform a single iteration and inspect the result.
next = iterate_parent_child(cleaned_df)
next.orderBy("parent_company","company").toPandas()
# # 3 Iterative Algorithm
#
# Now that we can add one level of indirection to our table of parent-child relations, we simply need to apply this algorithm as often as new records are created. We also add a reflective relation of each company to itself at the end, such that when using the table for aggregating all children, the revenue of each company itself will also be added up in addition to its children.
def calc_transitive_children(df):
# Remove records without a parent
cleaned_df = data \
.filter(data["parent_company"].isNotNull()) \
.select(data["company"], data["parent_company"])
# Iterate as long as new records are created
cur_df = cleaned_df
cur_count = cur_df.count()
while (True):
next_df = iterate_parent_child(cur_df)
next_count = next_df.count()
# If no new records are created, we are finished
if next_count == cur_count:
break
# This would be a good place to perform a checkpoint
cur_df = next_df
cur_count = next_count
# Create additional reflective relation of each company to irself
self_df = data.select(sf.col("company").alias("parent_company"), sf.col("company"))
return self_df.union(cur_df).distinct()
# ### Run Algorithm
#
# Now let us run the whole algorithm on the original data set and inspect the result.
# +
relations = calc_transitive_children(data)
relations.orderBy("parent_company","company").toPandas()
# -
# ### Inspect execution plan
relations.explain()
# # 4 Perform Aggregation
#
# Now let us perform the final aggregation, such that we can calculate the revenue of each company including each direct and indirect child. This can be performed by joining the `relations` data frame to the original `data` data frame and then grouping on the `parent_company` column of the `relations` data frame and adding up the revenue.
hierarchical_revenue = relations \
.join(data, ["company"]) \
.groupby(relations["parent_company"]) \
.agg(sf.sum(sf.col("revenue")).alias("total_revenue"))
hierarchical_revenue.toPandas()
# ### Check Totals
#
# Just to verify the result, let us compare the result of company 1 ("Earth") with a simple sum over all revenues.
totals = data.select(sf.sum(data["revenue"]))
totals.toPandas()
| spark-training/spark-python/jupyter-advanced-caching/Iterative Algorithms - Full.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/appliedaitest/UCI_templates/blob/main/Bank.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="MAwwF-AjuANR"
# # Welcome - Bank assessment
#
# The data you will load is related with direct marketing phonecalls campaigns of a Portuguese banking institution.
#
# The idea is to build a classification model to predict if the client will subscribe a term deposit ( variable: y ).
#
# Please be aware that final dataset is a Pandas Dataframe named final_df and it has 41188 rows and 15 columns. GOOD LUCK !
#
# ---
#
# Attribute Information:
#
# 1- age (numeric)
#
# 2- job : type of job (categorical: 'admin.','blue-collar','entrepreneur','housemaid','management','retired','self-employed','services','student','technician','unemployed','unknown')
#
# 3- marital : marital status (categorical: 'divorced','married','single','unknown'; note: 'divorced' means divorced or widowed)
#
# 4- education (categorical: 'basic.4y','basic.6y','basic.9y','high.school','illiterate','professional.course','university.degree','unknown')
#
# 5- default: has credit in default? (categorical: 'no','yes','unknown')
#
# 6- housing: has housing loan? (categorical: 'no','yes','unknown')
#
# 7- loan: has personal loan? (categorical: 'no','yes','unknown')
#
#
# **related with the last contact of the current campaign:**
#
# 8- contact: contact communication type (categorical: 'cellular','telephone')
#
# 9- month: last contact month of year (categorical: 'jan', 'feb', 'mar', ..., 'nov', 'dec')
#
# 10- day_of_week: last contact day of the week (categorical: 'mon','tue','wed','thu','fri')
#
# 11- duration: last contact duration, in seconds (numeric).
#
# **other attributes:**
#
# 12- campaign: number of contacts performed during this campaign and for this client (numeric, includes last contact)
#
# 13- pdays: number of days that passed by after the client was last contacted from a previous campaign (numeric; 999 means client was not previously contacted)
#
# 14- previous: number of contacts performed before this campaign and for this client (numeric)
#
# 15- poutcome: outcome of the previous marketing campaign (categorical: 'failure','nonexistent','success')
#
# **social and economic context attributes**
#
# 16- emp.var.rate: employment variation rate - quarterly indicator (numeric)
#
# 17- cons.price.idx: consumer price index - monthly indicator (numeric)
#
# 18- cons.conf.idx: consumer confidence index - monthly indicator (numeric)
#
# 19- euribor3m: euribor 3 month rate - daily indicator (numeric)
#
# 20- nr.employed: number of employees - quarterly indicator (numeric)
#
# **Output variable (desired target):**
#
# 21- y: has the client subscribed a term deposit? (binary: 'yes','no')
#
# + [markdown] id="BkPs95X-uFrA"
# ## Setup
#
# Run all the below
# + id="o97XPieIst6K"
## You can install libraries here
# # !pip install requests
# # !pip install pandas
# # !pip install scikit-learn
# + colab={"base_uri": "https://localhost:8080/"} id="gGffYbafamOw" outputId="6006f412-9d29-4192-fa95-0851df6c1147"
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/00222/bank-additional.zip
# !unzip bank-additional.zip
# !ls bank-additional/*.csv
# + id="0TbkCmAhs2D-" colab={"base_uri": "https://localhost:8080/"} outputId="92189527-7965-4aef-cb58-ed946747452c"
import pandas as pd
original_df = pd.read_csv( "./bank-additional/bank-additional-full.csv", sep= ";")
print ("original_df.shape: ", original_df.shape)
selected_cols = ['age', 'marital', 'default', 'housing', 'loan',
'duration', 'campaign', 'pdays', 'previous',
'emp.var.rate', 'cons.price.idx', 'cons.conf.idx', 'euribor3m',
'nr.employed', 'y']
final_df = original_df.loc[:, selected_cols]
print("final_df.shape: ",final_df.shape) #print(final_df.dtypes)
# + [markdown] id="IjAp1SnOuKn5"
# # Go
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="ZHI4u-7qtIat" outputId="7040f217-b62d-4385-8de5-e459426c3b76"
final_df.head()
# + id="3XuJPABetS1k"
| Bank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# name: python382jvsc74a57bd031f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# ---
# #### This notebook contains examples for the exceptions described in the manuscript
# #### Notice that we use scipy.sparse.linalg.eigsh for eigen calculation since it adopts the same underling routine as SpecHap
import numpy as np
from scipy.sparse.linalg import eigsh
# #### Here we provide an simple illustration when Fiedler vector contains entries approximating zero
# #### The matrix demonstrate the linkage graph of three variants loci with equal likelihood for conflicting haplotypes 000 and 010
mat = np.array([[0,0, 0.1,0.1, 0.1,0],
[0,0, 0.1,0.1, 0,0.1],
[0.1,0.1, 0,0, 0.1,0.1],
[0.1,0.1, 0,0, 0.1,0.1],
[0.1,0, 0.1,0.1, 0,0],
[0,0.1, 0.1,0.1, 0,0],], dtype='f')
# #### Notice the entries corresponding to the second variant locus contains value approximating zero
D = np.diag(np.sum(mat, axis= 0))
L = np.matrix(D - mat)
vals, vecs = eigsh(L, k=2, which='SM')
fiedler_vec = vecs[:,[1]]
fiedler_vec
# #### Now we provide an example for Fiedler vector that guides the partitioning of variants.
# #### The matrix demonstrate the linkage graph for 6 variants loci
# #### The first and last three variants loci are fully connected correspondingly
# #### The two fully connected blocks are connected through a relatively low-weight edge between the third and fourth locus
# +
mat = np.array([[0,0, 0.1,0.05, 0.1,0.05, 0,0, 0,0 , 0,0],
[0,0, 0.05,0.1, 0.05,0.1, 0,0, 0,0 , 0,0],
[0.1,0.05, 0,0, 0.1,0.05, 0,0, 0,0, 0,0],
[0.05,0.1, 0,0, 0.05,0.1, 0,0, 0,0, 0,0],
[0.1,0.05, 0.1,0.05, 0,0, 0.01,0, 0,0, 0,0],
[0.05,0.1, 0.05,0.1, 0,0, 0,0.01, 0,0, 0,0],
[0,0, 0,0, 0.01,0, 0,0, 0.1,0.05, 0.1,0.05],
[0,0, 0,0, 0,0.01, 0,0, 0.05,0.1, 0.05,0.1],
[0,0, 0,0, 0,0, 0.1,0.05, 0,0, 0.1,0.05,],
[0,0, 0,0, 0,0, 0.05,0.1, 0,0, 0.05,0.1,],
[0,0, 0,0, 0,0, 0.1,0.05, 0.1,0.05, 0,0,],
[0,0, 0,0, 0,0, 0.05,0.1, 0.05,0.1, 0,0,],]
, dtype='f')
# -
# #### Notice that the Fielder vector partition the variants loci into two groups
D = np.diag(np.sum(mat, axis= 0))
L = np.matrix(D - mat)
vals, vecs = eigsh(L, k=2, which='SM')
fiedler_vec = vecs[:,[1]]
fiedler_vec
| scripts/Fiedler_vec_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/davidalejandromiranda/fisicoquimica/blob/master/impedancia/Espectroscopia_de_Capacitancia.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# <h1 align="center">Propiedades eléctricas</h1>
# <div align="right">Por <NAME>, PhD<br>2020</div>
# <h2>1. Importa las librerias</h2>
# + id="B829joPepENE" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
f = np.logspace(-3, 6, 1000)
w = 2 * np.pi * f
# + [markdown] id="mCWind29qlHC" colab_type="text"
# # 1. Modelo circuital impedimétrico
# + id="x_BeotqvpMhD" colab_type="code" colab={}
Cd = 100e-6 # F
Rd = 100 # Ohm
Rr = 1 / 7.748091729e-5 # Ohm -> 1/R = 2 q^2 / h
Rm = 100 # Ohm
w = 2 * np.pi * f
Z_Cd = -1j/(w * Cd)
Zd = Rd + Z_Cd
Z1 = 1 / (1/Zd + 1/Rr)
Z = Rm + Z1
C = Z / (1j*w)
# + id="bh6vzrKpqYtZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 806} outputId="77ca0ad3-6776-4b42-ec33-e3789a67056a"
plt.figure(dpi=200)
plt.plot(Z.real, -Z.imag)
plt.xlabel('real{Z}')
plt.ylabel('-imag{Z}')
plt.title('Diagrama de Nyquist')
# + id="7wO8YP92p3Y-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 806} outputId="626e93d2-f73d-4223-beaa-8943f60f2d13"
plt.figure(dpi=200)
plt.plot(C.real, -C.imag)
plt.xlabel('real{C}')
plt.ylabel('-imag{C}')
plt.title('Diagrama de Nyquist')
# + [markdown] id="-XxP6VaLquLf" colab_type="text"
# # 2. Modelo de Cole-Cole para elemento capacitivo
# + id="Sr5pHwXPqHPv" colab_type="code" colab={}
Co = 120e-6 # F
Ci = 15e-6 # F
tau = 95e-3 # s/rad
c = 0.5
C = Ci + (Co - Ci) / ( 1+ (1j*w*tau)**c )
Z = 1 / (1j*w*C)
# + id="1IwmLpMwrc9_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 806} outputId="c215a85b-2546-4313-fb77-7ba39a04e385"
plt.figure(dpi=200)
plt.plot(Z.real, -Z.imag)
plt.xlabel('real{Z}')
plt.ylabel('-imag{Z}')
plt.title('Diagrama de Nyquist')
# + id="oSY7n4ZVrd39" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 806} outputId="ddc20070-052b-42b0-aa8e-e01a837fbcfd"
plt.figure(dpi=200)
plt.plot(C.real, -C.imag)
plt.xlabel('real{C}')
plt.ylabel('-imag{C}')
plt.title('Diagrama de Nyquist')
# + [markdown] id="ObWVzQr3rh8O" colab_type="text"
# End!
| impedancia/Espectroscopia_de_Capacitancia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SolbiChoi/test_deeplearning/blob/master/single_perceptron.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Yl7jTCY0Pzjs"
import tensorflow as tf
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="muPK3CHDpgws" outputId="b36f6e75-15f9-40dc-f1e4-c29292461850"
tf.__version__
# + colab={"base_uri": "https://localhost:8080/"} id="E3oSlyX5qGTC" outputId="46b7f534-1b70-4445-c152-0de2f2f09491"
x_data = [[0,0],
[1,0],
[0,1],
[1,1]]
y_data = [[0],
[1],
[1],
[1]]
type(x_data), type(y_data)
# + [markdown] id="z9L68iFurl3X"
# list는 tensorflow로 사용 못하므로 numpy로 변환해주어야한다.
# + colab={"base_uri": "https://localhost:8080/"} id="Nbd4QmH4rWIf" outputId="5e9f85d0-1013-4f80-82e9-3eb151c7ca2d"
import numpy as np
x_train = np.array(x_data)
y_train = np.array(y_data)
x_train.shape, y_train.shape
# + id="QNU7kZzxr9ic"
model = tf.keras.models.Sequential()
# + colab={"base_uri": "https://localhost:8080/"} id="2TM444LluB3w" outputId="48972762-d460-48ed-a8ba-a055b1e3517c"
model.add(tf.keras.Input(shape=(2,))) # shape=(첫번째 넣어주는 값의 모양/x열의 갯수)
model.add(tf.keras.layers.Dense(1)) # shape=(y열의 갯수)
model.compile(optimizer='sgd', loss='mse')
# + colab={"base_uri": "https://localhost:8080/"} id="QXSeXNetyeeC" outputId="03f23759-46e5-4c4d-cda9-5cea47f3706c"
model.fit(x_train,y_train, epochs=500)
# + [markdown] id="MvFlwrS5Ea-s"
# y = ax + bx +c
#
# y = -1.0857741 + 1.0996372x + 0.0047582 (밑에 나온 계수들 대입한 방정식..?)
# + colab={"base_uri": "https://localhost:8080/"} id="aachd_nSMSOQ" outputId="4169e3bb-6f2d-46fc-a599-f467670fceeb"
model.predict([[0,1]]) # epoch = 50
# + colab={"base_uri": "https://localhost:8080/"} id="69n9wfEYVyYJ" outputId="8ca94754-3b07-402f-eb72-57ec0cab493f"
model.predict([[0,1]]) # epoch = 500
# + colab={"base_uri": "https://localhost:8080/"} id="BMVTVSqeDmrK" outputId="68466cda-abc2-4548-f5f0-296ea0179599"
model.get_weights()
# + colab={"base_uri": "https://localhost:8080/", "height": 201} id="-PvDMDv4EMby" outputId="e31949ed-fe90-4d9d-89e0-11b4e62f8900"
tf.keras.utils.plot_model(model,show_shapes=True) # model:변수
# + id="wpidfP0dGQ0q"
| single_perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Clipping with a Surface {#clip_with_surface_example}
# =======================
#
# Clip any PyVista dataset by a `pyvista.PolyData`{.interpreted-text
# role="class"} surface mesh using the
# `pyvista.DataSetFilters.clip_surface`{.interpreted-text role="func"}
# filter.
#
# Note that we first demonstrate how the clipping is performed by
# computing an implicit distance and thresholding the mesh. This
# thresholding is one approach to clip by a surface, and preserve the
# original geometry of the given mesh, but many folks leverage the
# `clip_surface` filter to triangulate/tessellate the mesh geometries
# along the clip.
#
# sphinx_gallery_thumbnail_number = 4
import pyvista as pv
from pyvista import examples
import numpy as np
# +
surface = pv.Cone(direction=(0,0,-1), height=3.0, radius=1,
resolution=50, capping=False)
# Make a gridded dataset
n = 51
xx = yy = zz = 1 - np.linspace(0, n, n) * 2 / (n-1)
dataset = pv.RectilinearGrid(xx, yy, zz)
# Preview the problem
p = pv.Plotter()
p.add_mesh(surface, color='w', label='Surface')
p.add_mesh(dataset, color='gold', show_edges=True,
opacity=0.75, label='To Clip')
p.add_legend()
p.show()
# -
# Take a look at the implicit function used to perform the surface
# clipping by using the
# `pyvista.DataSetFilters.compute_implicit_distance`{.interpreted-text
# role="func"} filter. The clipping operation field is performed where the
# `implicit_distance` field is zero and the `invert` flag controls which
# sides of zero to preserve.
#
# +
dataset.compute_implicit_distance(surface, inplace=True)
inner = dataset.threshold(0.0, scalars="implicit_distance", invert=True)
outer = dataset.threshold(0.0, scalars="implicit_distance", invert=False)
p = pv.Plotter()
p.add_mesh(surface, color='w', label='Surface', opacity=0.75)
p.add_mesh(inner, scalars="implicit_distance", show_edges=True,
opacity=0.75, label='Inner region', clim=[-1,1], cmap="bwr")
p.add_legend()
p.enable_depth_peeling()
p.show()
# -
p = pv.Plotter()
p.add_mesh(surface, color='w', label='Surface', opacity=0.75)
p.add_mesh(outer, scalars="implicit_distance", show_edges=True,
opacity=0.75, label='Outer region', clim=[-1,1], cmap="bwr")
p.add_legend()
p.enable_depth_peeling()
p.show()
# Clip the rectilinear grid dataset using the
# `pyvista.PolyData`{.interpreted-text role="class"} surface mesh via the
# `pyvista.DataSetFilters.clip_surface`{.interpreted-text role="func"}
# filter. This will triangulate/tessellate the mesh geometries along the
# clip.
#
# +
clipped = dataset.clip_surface(surface, invert=False)
# Visualize the results
p = pv.Plotter()
p.add_mesh(surface, color='w', opacity=0.75, label='Surface')
p.add_mesh(clipped, color='gold', show_edges=True, label="clipped", opacity=0.75)
p.add_legend()
p.enable_depth_peeling()
p.show()
# -
# Here is another example of clipping a mesh by a surface. This time,
# we\'ll generate a `pyvista.UniformGrid`{.interpreted-text role="class"}
# around a topography surface and then clip that grid using the surface to
# create a closed 3D model of the surface
#
# +
surface = examples.load_random_hills()
# Create a grid around that surface
grid = pv.create_grid(surface)
# Clip the grid using the surface
model = grid.clip_surface(surface)
# Compute height and display it
model.elevation().plot()
| locale/examples/01-filter/clipping-with-surface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Create 5000 user profiles based on CS and LSA requirements
# +
import pandas as pd
import pickle
import numpy as np
import numpy as np
import json
import glob
import nltk
from nltk.corpus import stopwords
from top2vec import Top2Vec
# -
cs_students = pd.read_pickle('CS requirement/cs_students.pickle')
qr1_students = pd.read_pickle('LSA requirements/qr1_students.pickle')
qr2_students = pd.read_pickle('LSA requirements/qr2_students.pickle')
# + jupyter={"outputs_hidden": true}
# student_profiles = {}
# for i in range(2500):
# student_profiles[i] = pd.concat([cs_students[i], qr1_students[i]], ignore_index=True)
# student_profiles[i+2500] = pd.concat([cs_students[i+2500], qr2_students[i]], ignore_index=True)
# +
# pickle.dump(student_profiles, open("student_profiles.pickle", "wb"))
# -
student_profiles = pd.read_pickle('student_profiles.pickle')
pd.DataFrame.from_dict(student_profiles.items())
student_profiles[0]
f_21 = pd.read_csv('LSA requirements/f_21_merge.csv')
w_22 = pd.read_csv('LSA requirements/w_22_merge.csv')
df = pd.concat([f_21, w_22])
df.head()
df.columns
df['course_full_text'] = df['Course Title'] + df['sub_title'] + df['description'] + df['Acad Group'] + df['Subject']
len(df)
df.dropna(subset=['course_full_text'], inplace=True)
df.reset_index(inplace=True)
texts = df.course_full_text.tolist()
len(texts)
model = Top2Vec(texts, embedding_model='universal-sentence-encoder', min_count=5, speed='learn', workers=8)
model.save('course_topics_model')
model = Top2Vec.load('course_topics_model')
model.get_num_topics()
topic_sizes, topic_nums = model.get_topic_sizes()
df = pd.DataFrame(model.document_vectors)
from sklearn.cluster import DBSCAN
from matplotlib import pyplot
from numpy import unique
from collections import Counter
import hdbscan
hdbscan_args = {'min_cluster_size': 100,
'metric': 'euclidean',
'cluster_selection_method': 'eom',
'min_samples': 10}
clustering = hdbscan.HDBSCAN(**hdbscan_args).fit(df)
clustering.labels_
cluster_list = clustering.labels_.tolist()
Counter(cluster_list)
clusters = unique(clustering)
clusters
| User Profiles/.~user_profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
import squad_utils as utils
# +
# # !wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v2.0.json
# # !wget https://rajpurkar.github.io/SQuAD-explorer/dataset/dev-v2.0.json
# # !wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v1.1.json
# # !wget https://rajpurkar.github.io/SQuAD-explorer/dataset/dev-v1.1.json
# -
import json
import time
import subprocess
import csv
from tqdm import tqdm
from collections import defaultdict
import pickle
import argparse
from tqdm import tqdm
import logging
import malaya
# +
with open('dev-v1.1.json') as hn:
content = json.load(hn)
squad_version = content['version']
titles = [data['title'] for data in content['data']]
context_sentences = [
context_sentence
for data in content['data']
for paragraph in data['paragraphs']
for context_sentence in malaya.text.function.split_into_sentences(
utils.remove_line_breaks(paragraph['context']), minimum_length = 2
)
if context_sentence
]
questions = [
qa['question']
for data in content['data']
for paragraph in data['paragraphs']
for qa in paragraph['qas']
if qa['question']
]
answers = [
answer['text']
for data in content['data']
for paragraph in data['paragraphs']
for qa in paragraph['qas']
for answer in qa['answers']
if answer['text']
]
if squad_version == 'v2.0':
plausible_answers = []
for data in content['data']:
for paragraph in data['paragraphs']:
for qa in paragraph['qas']:
if qa['is_impossible']:
for answer in qa['plausible_answers']:
plausible_answers.append(answer['text'])
else:
plausible_answers = []
content = titles + context_sentences + questions + answers + plausible_answers
content = set(content)
# +
file = 'translated-dev-1.1.json'
batch_size = 10
if os.path.exists(file):
with open(file) as fopen:
data = json.load(fopen)
list_content, translated = data
else:
list_content = list(content)
transformer = malaya.translation.en_ms.transformer()
translated = []
for i in tqdm(range(0, len(list_content), batch_size)):
translated.extend(transformer.greedy_decoder(list_content[i: i + batch_size]))
with open(file, 'w') as fopen:
json.dump([list_content, translated], fopen)
# +
file = 'content_translations_alignments-dev-1.1.json'
if os.path.exists(file):
with open(file) as fopen:
content_translations_alignments = json.load(fopen)
else:
context_sentence_questions_answers_alignments = utils.compute_alignment(
list_content, 'en', translated, 'ms', 'forward', 'dev-v1.1.json', 'out'
)
content_translations_alignments = {}
for sentence, sentence_translated, alignment in zip(
list_content, translated, context_sentence_questions_answers_alignments
):
content_translations_alignments[sentence] = {
'translation': sentence_translated,
'alignment': alignment,
}
with open(file, 'w') as fopen:
json.dump(content_translations_alignments, fopen)
# +
answers_from_alignment = True
with open('dev-v1.1.json') as hn:
content = json.load(hn)
squad_version = content['version']
for data in tqdm(content['data']):
title = data['title']
data['title'] = content_translations_alignments[title]['translation']
for paragraphs in data['paragraphs']:
context = paragraphs['context']
context_sentences = [
s
for s in malaya.text.function.split_into_sentences(
utils.remove_line_breaks(context), minimum_length = 2
)
]
context_translated = ' '.join(
content_translations_alignments[s]['translation']
for s in context_sentences
)
context_alignment_tok = utils.compute_context_alignment(
[
content_translations_alignments[s]['alignment']
for s in context_sentences
]
)
paragraphs['context'] = context_translated
for qa in paragraphs['qas']:
question = qa['question']
question_translated = content_translations_alignments[question][
'translation'
]
qa['question'] = question_translated
# Translate answers and plausible answers for SQUAD v2.0
if squad_version == 'v2.0':
if not qa['is_impossible']:
for answer in qa['answers']:
answer_translated = content_translations_alignments[
answer['text']
]['translation']
answer_translated, answer_translated_start = utils.extract_answer_translated(
answer,
answer_translated,
context,
context_translated,
context_alignment_tok,
answers_from_alignment,
)
answer['text'] = answer_translated
answer['answer_start'] = answer_translated_start
else:
for plausible_answer in qa['plausible_answers']:
plausible_answer_translated = content_translations_alignments[
plausible_answer['text']
][
'translation'
]
answer_translated, answer_translated_start = utils.extract_answer_translated(
plausible_answer,
plausible_answer_translated,
context,
context_translated,
context_alignment_tok,
answers_from_alignment,
)
plausible_answer['text'] = answer_translated
plausible_answer[
'answer_start'
] = answer_translated_start
# Translate answers for SQUAD v1.1
else:
for answer in qa['answers']:
answer_translated = content_translations_alignments[
answer['text']
]['translation']
answer_translated, answer_translated_start = utils.extract_answer_translated(
answer,
answer_translated,
context,
context_translated,
context_alignment_tok,
answers_from_alignment,
)
answer['text'] = answer_translated
answer['answer_start'] = answer_translated_start
# -
content_translated = content
content_cleaned = {'version': content['version'], 'data': []}
total_answers = 0
total_correct_plausible_answers = 0
total_correct_answers = 0
for idx_data, data in tqdm(enumerate(content_translated['data'])):
content_title = content_translated['data'][idx_data]['title']
content_cleaned['data'].append({'title': content_title, 'paragraphs': []})
for par in data['paragraphs']:
qas_cleaned = []
for idx_qa, qa in enumerate(par['qas']):
question = qa['question']
# Extract answers and plausible answers for SQUAD v2.0
if squad_version == 'v2.0':
if not qa['is_impossible']:
correct_answers = []
for a in qa['answers']:
total_answers += 1
if a['text']:
total_correct_answers += 1
correct_answers.append(a)
correct_plausible_answers = []
else:
correct_plausible_answers = []
for pa in qa['plausible_answers']:
total_answers += 1
if pa['text']:
total_correct_plausible_answers += 1
correct_plausible_answers.append(pa)
correct_answers = []
# add answers and plausible answers to the content cleaned
if correct_answers:
content_qas_id = qa['id']
content_qas_is_impossible = qa['is_impossible']
correct_answers_from_context = []
for a in qa['answers']:
start = a['answer_start']
correct_answers_from_context.append(
{
'text': par['context'][
start : start + len(a['text'])
],
'answer_start': start,
}
)
qa_cleaned = {
'question': question,
'answers': correct_answers_from_context,
'id': content_qas_id,
'is_impossible': content_qas_is_impossible,
}
qas_cleaned.append(qa_cleaned)
if correct_plausible_answers and not correct_answers:
content_qas_id = qa['id']
content_qas_is_impossible = qa['is_impossible']
correct_answers_from_context = []
for a in qa['answers']:
start = a['answer_start']
correct_answers_from_context.append(
{
'text': par['context'][
start : start + len(a['text'])
],
'answer_start': start,
}
)
qa_cleaned = {
'question': question,
'answers': correct_answers,
'plausible_answers': correct_plausible_answers,
'id': content_qas_id,
'is_impossible': content_qas_is_impossible,
}
qas_cleaned.append(qa_cleaned)
# Extract answers for SQUAD v1.0
else:
correct_answers = []
for a in qa['answers']:
total_answers += 1
if a['text']:
total_correct_answers += 1
correct_answers.append(a)
# add answers and plausible answers to the content cleaned
if correct_answers:
content_qas_id = qa['id']
correct_answers_from_context = []
for a in qa['answers']:
start = a['answer_start']
correct_answers_from_context.append(
{
'text': par['context'][
start : start + len(a['text'])
],
'answer_start': start,
}
)
qa_cleaned = {
'question': question,
'answers': correct_answers_from_context,
'id': content_qas_id,
}
qas_cleaned.append(qa_cleaned)
# Add the paragraph only if there are non-empty question-answer examples inside
if qas_cleaned:
content_context = par['context']
content_cleaned['data'][idx_data]['paragraphs'].append(
{'context': content_context, 'qas': qas_cleaned}
)
with open('ms-dev-1.1.json', 'w') as fn:
json.dump(content_cleaned, fn)
# +
if squad_version == 'v2.0':
total_correct = total_correct_answers + total_correct_plausible_answers
accuracy = round((total_correct / total_answers) * 100, 2)
print(
'Percentage of translated examples (correct answers/total answers): {}/{} = {}%\n'
'No. of answers: {}\n'
'No. of plausible answers: {}'.format(
total_correct,
total_answers,
accuracy,
total_correct_answers,
total_correct_plausible_answers,
)
)
# Count correct answers
else:
total_correct = total_correct_answers
accuracy = round((total_correct / total_answers) * 100, 2)
print(
'Percentage of translated examples (correct answers/total answers): {}/{} = {}%\n'
'No. of answers: {}'.format(
total_correct, total_answers, accuracy, total_correct_answers
)
)
# -
| question-answer/squad/notebook/squad-dev-1.1-ms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Description
# This example performs a small Monte Carlo simulation, using UQpy's RunModel module and a Matlab model that takes the sum of Normal random variables.
#
# <NAME>
# December 9, 2018
# ## Required files
# To run this example, the following files must be present in the same directory as this Jupyter script:
#
# 1) matlab_model.py
# 2) dummy_model.m
# 3) process_matlab_output.py
# ## Structure of files
# 1) "matlab_model.py" is the model script. It contains the Python commands to run the Matlab model.
# 2) "dummy_model.m" is a template input file is used to create matlab input files (.m) files for each simulation. This text file has placeholders for placement of the sampled variables.
# 3) "process_matlab_output.py" is the output script. It is a Python script which processes the output of the Matlab simulations.
# +
from UQpy.SampleMethods import MCS
from UQpy.RunModel import RunModel
import os
import time
# Add MATLAB to $PATH - modify and use the command in the next line if necessary
# os.system("export PATH=$PATH:'/Applications/MATLAB_R2018a.app/bin'")
# -
# ## Sample generation
# +
# Call MCS to generate samples
x_mcs = MCS(dist_name=['Normal','Normal','Normal'], dist_params=[[0,1],[0,1],[0,1]], nsamples=3,
var_names = ['var1', 'var2', 'var3'])
print("Three Monte Carlo samples from a trivariate standard normal distribution.")
print(x_mcs.samples)
# -
# ## Calling RunModel in Serial
# +
# Call to RunModel
t = time.time()
m = RunModel(samples=x_mcs.samples, ntasks=1, model_script='matlab_model.py', input_template='dummy_model.m',
var_names=x_mcs.var_names, model_object_name="matlab", output_script='process_matlab_output.py',
output_object_name='read_output', resume=False)
t_ser_matlab = time.time() - t
print("\nTime for serial execution:")
print(t_ser_matlab)
print("The values retured from the Matlab simulation:")
print(m.qoi_list)
# -
# ## Calling RunModel in Parallel
# Note that GNU parallel must be installed for this to run.
# +
# Call to RunModel
t = time.time()
m = RunModel(samples=x_mcs.samples, ntasks=4, model_script='matlab_model.py', input_template='dummy_model.m',
var_names=x_mcs.var_names, model_object_name="matlab", output_script='process_matlab_output.py',
output_object_name='read_output', resume=False)
t_ser_matlab = time.time() - t
print("\nTime for parallel execution:")
print(t_ser_matlab)
print("The values retured from the Matlab simulation:")
print(m.qoi_list)
# -
| example/RunModel/Matlab_Model_Example/Matlab_Model_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
with open('day3.input') as fp:
data = fp.readlines()
wire1 = data[0].strip().split(',')
wire2 = data[1].strip().split(',')
# ## Part 1 ##
testwire0a = 'R8,U5,L5,D3'.split(',')
testwire0b = 'U7,R6,D4,L4'.split(',')
testwire1a = 'R75,D30,R83,U83,L12,D49,R71,U7,L72'.split(',')
testwire1b = 'U62,R66,U55,R34,D71,R55,D58,R83'.split(',')
testwire2a = 'R98,U47,R26,D63,R33,U87,L62,D20,R33,U53,R51'.split(',')
testwire2b = 'U98,R91,D20,R16,D67,R40,U7,R15,U6,R7'.split(',')
def walkpath(wire):
pos = (0, 0)
points = {pos}
for segment in wire:
d = segment[0]
steps = int(segment[1:])
if d == 'R':
for i in range(steps):
pos = (pos[0]+1, pos[1])
points.add(pos)
elif d == 'L':
for i in range(steps):
pos = (pos[0]-1, pos[1])
points.add(pos)
elif d == 'U':
for i in range(steps):
pos = (pos[0], pos[1]+1)
points.add(pos)
elif d == 'D':
for i in range(steps):
pos = (pos[0], pos[1]-1)
points.add(pos)
else:
raise ValueError(f'Bad direction: {d}')
return points
def taxicab_dist(pt):
return abs(pt[0]) + abs(pt[1])
tst0apts = walkpath(testwire0a)
tst0bpts = walkpath(testwire0b)
intersections = tst0apts & tst0bpts
intersections.remove((0,0))
intersections
6 == min(taxicab_dist(x) for x in intersections)
tst1apts = walkpath(testwire1a)
tst1bpts = walkpath(testwire1b)
intersections = tst1apts & tst1bpts
intersections.remove((0,0))
159 == min(taxicab_dist(x) for x in intersections)
tst2apts = walkpath(testwire2a)
tst2bpts = walkpath(testwire2b)
intersections = tst2apts & tst2bpts
intersections.remove((0,0))
135 == min(taxicab_dist(x) for x in intersections)
wire1pts = walkpath(wire1)
wire2pts = walkpath(wire2)
intersections = wire1pts & wire2pts
intersections.remove((0,0))
min(taxicab_dist(x) for x in intersections)
# ## Part 2 ##
#
# For each wire, store the position of each point along the wire in a list, in order. It's not efficient, but it's simple.
# The length along the wire corresponding to each intersection is then just the index of the intersection point in
# the list.
def walkpath2(wire):
pos = (0, 0)
points = [pos]
for segment in wire:
d = segment[0]
steps = int(segment[1:])
if d == 'R':
for i in range(steps):
pos = (pos[0]+1, pos[1])
points.append(pos)
elif d == 'L':
for i in range(steps):
pos = (pos[0]-1, pos[1])
points.append(pos)
elif d == 'U':
for i in range(steps):
pos = (pos[0], pos[1]+1)
points.append(pos)
elif d == 'D':
for i in range(steps):
pos = (pos[0], pos[1]-1)
points.append(pos)
else:
raise ValueError(f'Bad direction: {d}')
return points
tst0apts = walkpath2(testwire0a)
tst0bpts = walkpath2(testwire0b)
intersections = set(tst0apts) & set(tst0bpts)
intersections.remove((0,0))
intersections
for x in intersections:
stepsa = tst0apts.index(x)
stepsb = tst0bpts.index(x)
print(x, stepsa+stepsb)
wire1pts = walkpath2(wire1)
wire2pts = walkpath2(wire2)
intersections = set(wire1pts) & set(wire2pts)
intersections.remove((0,0))
minsteps = None
for x in intersections:
steps1 = wire1pts.index(x)
steps2 = wire2pts.index(x)
totsteps = steps1 + steps2
if minsteps is None:
minsteps = totsteps
continue
if totsteps < minsteps:
minsteps = totsteps
minsteps
| day3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# <center>
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
# </center>
#
# # Conditions in Python
#
# Estimated time needed: **20** minutes
#
# ## Objectives
#
# After completing this lab you will be able to:
#
# * work with condition statements in Python, including operators, and branching.
#
# <h2>Table of Contents</h2>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <ul>
# <li>
# <a href="https://#cond">Condition Statements</a>
# <ul>
# <li><a href="https://comp/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Comparison Operators</a></li>
# <li><a href="https://branch/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Branching</a></li>
# <li><a href="https://logic/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Logical operators</a></li>
# </ul>
# </li>
# <li>
# <a href="https://#quiz">Quiz on Condition Statement</a>
# </li>
# </ul>
#
# </div>
#
# <hr>
#
# <h2 id="cond">Condition Statements</h2>
#
# <h3 id="comp">Comparison Operators</h3>
#
# Comparison operations compare some value or operand and based on a condition, produce a Boolean. When comparing two values you can use these operators:
#
# <ul>
# <li>equal: <b>==</b></li>
# <li>not equal: <b>!=</b></li>
# <li>greater than: <b>></b></li>
# <li>less than: <b><</b></li>
# <li>greater than or equal to: <b>>=</b></li>
# <li>less than or equal to: <b><=</b></li>
# </ul>
#
# Let's assign <code>a</code> a value of 5. Use the equality operator denoted with two equal <b>==</b> signs to determine if two values are equal. The case below compares the variable <code>a</code> with 6.
#
# +
# Condition Equal
a = 5
a == 6
# -
# The result is <b>False</b>, as 5 does not equal to 6.
#
# Consider the following equality comparison operator: <code>i > 5</code>. If the value of the left operand, in this case the variable <b>i</b>, is greater than the value of the right operand, in this case 5, then the statement is <b>True</b>. Otherwise, the statement is <b>False</b>. If <b>i</b> is equal to 6, because 6 is larger than 5, the output is <b>True</b>.
#
# +
# Greater than Sign
i = 6
i > 5
# -
# Set <code>i = 2</code>. The statement is False as 2 is not greater than 5:
#
# +
# Greater than Sign
i = 2
i > 5
# -
# Let's display some values for <code>i</code> in the figure. Set the values greater than 5 in green and the rest in red. The green region represents where the condition is **True**, the red where the statement is **False**. If the value of <code>i</code> is 2, we get **False** as the 2 falls in the red region. Similarly, if the value for <code>i</code> is 6 we get a **True** as the condition falls in the green region.
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsGreater.gif" width="650" />
#
# The inequality test uses an exclamation mark preceding the equal sign, if two operands are not equal then the condition becomes **True**. For example, the following condition will produce **True** as long as the value of <code>i</code> is not equal to 6:
#
# +
# Inequality Sign
i = 2
i != 6
# -
# When <code>i</code> equals 6 the inequality expression produces <b>False</b>.
#
# +
# Inequality Sign
i = 6
i != 6
# -
# See the number line below. When the condition is **True**, the corresponding numbers are marked in green and for where the condition is **False** the corresponding number is marked in red. If we set <code>i</code> equal to 2 the operator is true, since 2 is in the green region. If we set <code>i</code> equal to 6, we get a **False**, since the condition falls in the red region.
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsIneq.gif" width="650" />
#
# We can apply the same methods on strings. For example, we can use an equality operator on two different strings. As the strings are not equal, we get a **False**.
#
# +
# Use Equality sign to compare the strings
"ACDC" == "<NAME>"
# -
# If we use the inequality operator, the output is going to be **True** as the strings are not equal.
#
# +
# Use Inequality sign to compare the strings
"ACDC" != "<NAME>"
# -
# The inequality operation is also used to compare the letters/words/symbols according to the ASCII value of letters. The decimal value shown in the following table represents the order of the character:
#
# <style type="text/css">
# .tg {border-collapse:collapse;border-spacing:0;}
# .tg td{border-color:black;border-style:solid;border-width:1px;font-family:Arial, sans-serif;font-size:14px;
# overflow:hidden;padding:10px 5px;word-break:normal;}
# .tg th{border-color:black;border-style:solid;border-width:1px;font-family:Arial, sans-serif;font-size:14px;
# font-weight:normal;overflow:hidden;padding:10px 5px;word-break:normal;}
# .tg .tg-baqh{text-align:center;vertical-align:top}
# .tg .tg-7geq{background-color:#ffffc7;text-align:center;vertical-align:top}
# .tg .tg-1cln{background-color:#ffcc67;font-size:100%;font-weight:bold;text-align:center;vertical-align:top}
# .tg .tg-xozw{background-color:#ffcc67;font-weight:bold;text-align:center;vertical-align:top}
# </style>
#
# <table class="tg">
# <thead>
# <tr>
# <th class="tg-1cln">Char.</th>
# <th class="tg-xozw">ASCII</th>
# <th class="tg-xozw">Char.</th>
# <th class="tg-xozw">ASCII</th>
# <th class="tg-xozw">Char.</th>
# <th class="tg-xozw">ASCII</th>
# <th class="tg-xozw">Char.</th>
# <th class="tg-xozw">ASCII</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td class="tg-7geq">A</td>
# <td class="tg-baqh">65</td>
# <td class="tg-7geq">N</td>
# <td class="tg-baqh">78</td>
# <td class="tg-7geq">a</td>
# <td class="tg-baqh">97</td>
# <td class="tg-7geq">n</td>
# <td class="tg-baqh">110</td>
# </tr>
# <tr>
# <td class="tg-7geq">B</td>
# <td class="tg-baqh">66</td>
# <td class="tg-7geq">O</td>
# <td class="tg-baqh">79</td>
# <td class="tg-7geq">b</td>
# <td class="tg-baqh">98</td>
# <td class="tg-7geq">o</td>
# <td class="tg-baqh">111</td>
# </tr>
# <tr>
# <td class="tg-7geq">C</td>
# <td class="tg-baqh">67</td>
# <td class="tg-7geq">P</td>
# <td class="tg-baqh">80</td>
# <td class="tg-7geq">c</td>
# <td class="tg-baqh">99</td>
# <td class="tg-7geq">p</td>
# <td class="tg-baqh">112</td>
# </tr>
# <tr>
# <td class="tg-7geq">D</td>
# <td class="tg-baqh">68</td>
# <td class="tg-7geq">Q</td>
# <td class="tg-baqh">81</td>
# <td class="tg-7geq">d</td>
# <td class="tg-baqh">100</td>
# <td class="tg-7geq">q</td>
# <td class="tg-baqh">113</td>
# </tr>
# <tr>
# <td class="tg-7geq">E</td>
# <td class="tg-baqh">69</td>
# <td class="tg-7geq">R</td>
# <td class="tg-baqh">82</td>
# <td class="tg-7geq">e</td>
# <td class="tg-baqh">101</td>
# <td class="tg-7geq">r</td>
# <td class="tg-baqh">114</td>
# </tr>
# <tr>
# <td class="tg-7geq">F</td>
# <td class="tg-baqh">70</td>
# <td class="tg-7geq">S</td>
# <td class="tg-baqh">83</td>
# <td class="tg-7geq">f</td>
# <td class="tg-baqh">102</td>
# <td class="tg-7geq">s</td>
# <td class="tg-baqh">115</td>
# </tr>
# <tr>
# <td class="tg-7geq">G</td>
# <td class="tg-baqh">71</td>
# <td class="tg-7geq">T</td>
# <td class="tg-baqh">84</td>
# <td class="tg-7geq">g</td>
# <td class="tg-baqh">103</td>
# <td class="tg-7geq">t</td>
# <td class="tg-baqh">116</td>
# </tr>
# <tr>
# <td class="tg-7geq">H</td>
# <td class="tg-baqh">72</td>
# <td class="tg-7geq">U</td>
# <td class="tg-baqh">85</td>
# <td class="tg-7geq">h</td>
# <td class="tg-baqh">104</td>
# <td class="tg-7geq">u</td>
# <td class="tg-baqh">117</td>
# </tr>
# <tr>
# <td class="tg-7geq">I</td>
# <td class="tg-baqh">73</td>
# <td class="tg-7geq">V</td>
# <td class="tg-baqh">86</td>
# <td class="tg-7geq">i</td>
# <td class="tg-baqh">105</td>
# <td class="tg-7geq">v</td>
# <td class="tg-baqh">118</td>
# </tr>
# <tr>
# <td class="tg-7geq">J</td>
# <td class="tg-baqh">74</td>
# <td class="tg-7geq">W</td>
# <td class="tg-baqh">87</td>
# <td class="tg-7geq">j</td>
# <td class="tg-baqh">106</td>
# <td class="tg-7geq">w</td>
# <td class="tg-baqh">119</td>
# </tr>
# <tr>
# <td class="tg-7geq">K</td>
# <td class="tg-baqh">75</td>
# <td class="tg-7geq">X</td>
# <td class="tg-baqh">88</td>
# <td class="tg-7geq">k</td>
# <td class="tg-baqh">107</td>
# <td class="tg-7geq">x</td>
# <td class="tg-baqh">120</td>
# </tr>
# <tr>
# <td class="tg-7geq">L</td>
# <td class="tg-baqh">76</td>
# <td class="tg-7geq">Y</td>
# <td class="tg-baqh">89</td>
# <td class="tg-7geq">l</td>
# <td class="tg-baqh">108</td>
# <td class="tg-7geq">y</td>
# <td class="tg-baqh">121</td>
# </tr>
# <tr>
# <td class="tg-7geq">M</td>
# <td class="tg-baqh">77</td>
# <td class="tg-7geq">Z</td>
# <td class="tg-baqh">90</td>
# <td class="tg-7geq">m</td>
# <td class="tg-baqh">109</td>
# <td class="tg-7geq">z</td>
# <td class="tg-baqh">122</td>
# </tr>
# </tbody>
# </table>
#
# For example, the ASCII code for <b>!</b> is 33, while the ASCII code for <b>+</b> is 43. Therefore <b>+</b> is larger than <b>!</b> as 43 is greater than 33.
#
# Similarly, from the table above we see that the value for <b>A</b> is 65, and the value for <b>B</b> is 66, therefore:
#
# +
# Compare characters
'B' > 'A'
# -
# When there are multiple letters, the first letter takes precedence in ordering:
#
# +
# Compare characters
'BA' > 'AB'
# -
# <b>Note</b>: Upper Case Letters have different ASCII code than Lower Case Letters, which means the comparison between the letters in Python is case-sensitive.
#
# <h3 id="branch">Branching</h3>
#
# Branching allows us to run different statements for different inputs. It is helpful to think of an **if statement** as a locked room, if the statement is **True** we can enter the room and your program will run some predefined tasks, but if the statement is **False** the program will ignore the task.
#
# For example, consider the blue rectangle representing an ACDC concert. If the individual is older than 18, they can enter the ACDC concert. If they are 18 or younger, they cannot enter the concert.
#
# We can use the condition statements learned before as the conditions that need to be checked in the **if statement**. The syntax is as simple as <code> if <i>condition statement</i> :</code>, which contains a word <code>if</code>, any condition statement, and a colon at the end. Start your tasks which need to be executed under this condition in a new line with an indent. The lines of code after the colon and with an indent will only be executed when the **if statement** is **True**. The tasks will end when the line of code does not contain the indent.
#
# In the case below, the code <code>print(“you can enter”)</code> is executed only if the variable <code>age</code> is greater than 18 is a True case because this line of code has the indent. However, the execution of <code>print(“move on”)</code> will not be influenced by the if statement.
#
# +
# If statement example
age = 19
#age = 18
#expression that can be true or false
if age > 18:
#within an indent, we have the expression that is run if the condition is true
print("you can enter" )
#The statements after the if statement will run regardless if the condition is true or false
print("move on")
# -
# <i>Try uncommenting the age variable</i>
#
# It is helpful to use the following diagram to illustrate the process. On the left side, we see what happens when the condition is <b>True</b>. The person enters the ACDC concert representing the code in the indent being executed; they then move on. On the right side, we see what happens when the condition is <b>False</b>; the person is not granted access, and the person moves on. In this case, the segment of code in the indent does not run, but the rest of the statements are run.
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsIf.gif" width="650" />
#
# The <code>else</code> statement runs a block of code if none of the conditions are **True** before this <code>else</code> statement. Let's use the ACDC concert analogy again. If the user is 17 they cannot go to the ACDC concert, but they can go to the Meatloaf concert.
# The syntax of the <code>else</code> statement is similar as the syntax of the <code>if</code> statement, as <code>else :</code>. Notice that, there is no condition statement for <code>else</code>.
# Try changing the values of <code>age</code> to see what happens:
#
# +
# Else statement example
age = 18
# age = 19
if age > 18:
print("you can enter" )
else:
print("go see Meat Loaf" )
print("move on")
# -
# The process is demonstrated below, where each of the possibilities is illustrated on each side of the image. On the left is the case where the age is 17, we set the variable age to 17, and this corresponds to the individual attending the Meatloaf concert. The right portion shows what happens when the individual is over 18, in this case 19, and the individual is granted access to the concert.
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsElse.gif" width="650" />
#
# The <code>elif</code> statement, short for else if, allows us to check additional conditions if the condition statements before it are <b>False</b>. If the condition for the <code>elif</code> statement is <b>True</b>, the alternate expressions will be run. Consider the concert example, where if the individual is 18 they will go to the Pink Floyd concert instead of attending the ACDC or Meat-loaf concert. A person that is 18 years of age enters the area, and as they are not older than 18 they can not see ACDC, but since they are 18 years of age, they attend Pink Floyd. After seeing Pink Floyd, they move on. The syntax of the <code>elif</code> statement is similar in that we merely change the <code>if</code> in the <code>if</code> statement to <code>elif</code>.
#
# +
# Elif statment example
age = 18
if age > 18:
print("you can enter" )
elif age == 18:
print("go see Pink Floyd")
else:
print("go see Meat Loaf" )
print("move on")
# -
# The three combinations are shown in the figure below. The left-most region shows what happens when the individual is less than 18 years of age. The central component shows when the individual is exactly 18. The rightmost shows when the individual is over 18.
#
# <img src ="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsElif.gif" width="650" />
#
# Look at the following code:
#
# +
# Condition statement example
album_year = 1983
album_year = 1970
if album_year > 1980:
print("Album year is greater than 1980")
print('do something..')
# -
# Feel free to change <code>album_year</code> value to other values -- you'll see that the result changes!
#
# Notice that the code in the above <b>indented</b> block will only be executed if the results are <b>True</b>.
#
# As before, we can add an <code>else</code> block to the <code>if</code> block. The code in the <code>else</code> block will only be executed if the result is <b>False</b>.
#
# <b>Syntax:</b>
#
# if (condition):
# \# do something
# else:
# \# do something else
#
# If the condition in the <code>if</code> statement is <b>False</b>, the statement after the <code>else</code> block will execute. This is demonstrated in the figure:
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsLogicMap.png" width="650" />
#
# +
# Condition statement example
album_year = 1983
#album_year = 1970
if album_year > 1980:
print("Album year is greater than 1980")
else:
print("less than 1980")
print('do something..')
# -
# Feel free to change the <code>album_year</code> value to other values -- you'll see that the result changes based on it!
#
# <h3 id="logic">Logical operators</h3>
#
# Sometimes you want to check more than one condition at once. For example, you might want to check if one condition and another condition are both **True**. Logical operators allow you to combine or modify conditions.
#
# <ul>
# <li><code>and</code></li>
# <li><code>or</code></li>
# <li><code>not</code></li>
# </ul>
#
# These operators are summarized for two variables using the following truth tables:
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsTable.png" width="650" />
#
# The <code>and</code> statement is only **True** when both conditions are true. The <code>or</code> statement is True if one condition, or both are **True**. The <code>not</code> statement outputs the opposite truth value.
#
# Let's see how to determine if an album was released after 1979 (1979 is not included) and before 1990 (1990 is not included). The time periods between 1980 and 1989 satisfy this condition. This is demonstrated in the figure below. The green on lines <strong>a</strong> and <strong>b</strong> represents periods where the statement is **True**. The green on line <strong>c</strong> represents where both conditions are **True**, this corresponds to where the green regions overlap.
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsEgOne.png" width="650" />
#
# The block of code to perform this check is given by:
#
# +
# Condition statement example
album_year = 1980
if(album_year > 1979) and (album_year < 1990):
print ("Album year was in between 1980 and 1989")
print("")
print("Do Stuff..")
# -
# To determine if an album was released before 1980 (1979 and earlier) or after 1989 (1990 and onward ), an or statement can be used. Periods before 1980 (1979 and earlier) or after 1989 (1990 and onward) satisfy this condition. This is demonstrated in the following figure, the color green in <strong>a</strong> and <strong>b</strong> represents periods where the statement is true. The color green in **c** represents where at least one of the conditions
# are true.
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%203/images/CondsEgTwo.png" width="650" />
#
# The block of code to perform this check is given by:
#
# +
# Condition statement example
album_year = 1990
if(album_year < 1980) or (album_year > 1989):
print ("Album was not made in the 1980's")
else:
print("The Album was made in the 1980's ")
# -
# The <code>not</code> statement checks if the statement is false:
#
# +
# Condition statement example
album_year = 1983
if not (album_year == '1984'):
print ("Album year is not 1984")
# -
# <hr>
#
# <h2 id="quiz">Quiz on Conditions</h2>
#
# Write an if statement to determine if an album had a rating greater than 8. Test it using the rating for the album <b>“Back in Black”</b> that had a rating of 8.5. If the statement is true print "This album is Amazing!"
#
# Write your code below and press Shift+Enter to execute
rating = 8.5
if rating > 8:
print ("This album is Amazing!")
# <details><summary>Click here for the solution</summary>
#
# ```python
# rating = 8.5
# if rating > 8:
# print ("This album is Amazing!")
# ```
#
# </details>
#
# <hr>
#
# Write an if-else statement that performs the following. If the rating is larger then eight print “this album is amazing”. If the rating is less than or equal to 8 print “this album is ok”.
#
# Write your code below and press Shift+Enter to execute
rating = 8.5
if rating > 8:
print ("this album is amazing")
else:
print ("this album is ok")
# <details><summary>Click here for the solution</summary>
#
# ```python
# rating = 8.5
# if rating > 8:
# print ("this album is amazing")
# else:
# print ("this album is ok")
#
# ```
#
# </details>
#
# <hr>
#
# Write an if statement to determine if an album came out before 1980 or in the years: 1991 or 1993. If the condition is true print out the year the album came out.
#
# +
# Write your code below and press Shift+Enter to execute
album_year = 1979
if album_year < 1980 or album_year == 1991 or album_year == 1993:
print("This album came out in year", album_year)
# -
# <details><summary>Click here for the solution</summary>
#
# ```python
# album_year = 1979
#
# if album_year < 1980 or album_year == 1991 or album_year == 1993:
# print("This album came out in year", album_year)
#
# ```
#
# </details>
#
# <hr>
# <h2>The last exercise!</h2>
# <p>Congratulations, you have completed your first lesson and hands-on lab in Python. However, there is one more thing you need to do. The Data Science community encourages sharing work. The best way to share and showcase your work is to share it on GitHub. By sharing your notebook on GitHub you are not only building your reputation with fellow data scientists, but you can also show it off when applying for a job. Even though this was your first piece of work, it is never too early to start building good habits. So, please read and follow <a href="https://cognitiveclass.ai/blog/data-scientists-stand-out-by-sharing-your-notebooks/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01" target="_blank">this article</a> to learn how to share your work.
# <hr>
#
# ## Author
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01" target="_blank"><NAME></a>
#
# ## Other contributors
#
# <a href="https://www.linkedin.com/in/jiahui-mavis-zhou-a4537814a?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01"><NAME></a>
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ---------- | ---------------------------------- |
# | 2020-08-26 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
# | | | | |
# | | | | |
#
# <hr/>
#
# ## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
#
| 1 - Python for Data Science/Module 3 - Python Programming Fundamentals/code/1-PY0101EN-3-1-Conditions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Packages
import torch
from functionalities import dataloader as dl
from functionalities import tracker as tk
from architecture import INN as inn
from functionalities import MMD_autoencoder_loss as mmd_loss
from functionalities import trainer as tr
from functionalities import filemanager as fm
from functionalities import plot as pl
from functionalities import gpu
# # Pretraining Setup
# +
num_epoch = 10
batch_size = 128
latent_dim_lst = [1, 2, 3, 4, 6, 8, 12, 16, 24, 32, 48, 64]
number_dev = 0
lr_init = 1e-3
l2_reg = 1e-6
milestones = [8, 10]
modelname = 'mnist_INN_glow_com_bottleneck'
get_model = inn.mnist_inn_com
device = gpu.get_device(number_dev)
print(device)
# -
trainset, testset, classes = dl.load_mnist()
trainloader, validloader, testloader = dl.make_dataloaders(trainset, testset, batch_size)
# # Training
model = tr.train_bottleneck(num_epoch, get_model, 'l1', modelname, milestones, latent_dim_lst, trainloader, None,
testloader, a_distr=0, a_disen=0, lr_init=lr_init, l2_reg=l2_reg, device=device, save_model=True)
# # Plot Reconstruction and Difference Images Examples
pl.plot_diff_all(get_model, modelname, num_epoch, testloader, latent_dim_lst, device='cpu', num_img=1, grid_row_size=10, figsize=(30, 30),
filename=None, conditional=False)
# # Plot Recontruction Loss against Bottleneck Size
# +
_, l1_rec_test, _, _, _ = fm.load_variable('bottleneck_test_loss_{}'.format(modelname), modelname)
_, l1_rec_train, _, _, _ = fm.load_variable('bottleneck_train_loss_{}'.format(modelname), modelname)
pl.plot(latent_dim_lst, [l1_rec_train, l1_rec_test], 'bottleneck size', 'loss', ['train', 'test'], 'Test Reconstruction Loss History', '{}_bottleneck_History'.format(modelname))
# -
| INN_VAE/INN_autoencoder_example/INN Autoencoder_mnist-glow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Reproducibility Verification, and Validation
# + [markdown] slideshow={"slide_type": "subslide"}
# Brief definitions...
#
# **Reproducibility**: Can someone reproduce a given result.
#
# **Verification**: Can we verify that a code solves what it is supposed to, say a model set of equations.
#
# **Validation**: Can the numerical model approximate reality.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Reproducibility
#
# Can someone reproduce your results?
# + [markdown] slideshow={"slide_type": "subslide"}
# Levels of reproducibility:
# - Within numerical approximation, e.g. $\mathcal{O}(\Delta x)$
# - Within numerical precision (machine epsilon)
# - It is bit-wise reproducible
# + [markdown] slideshow={"slide_type": "subslide"}
# How do we achieve reproducibility:
# - Open source software
# - Open data
# - Libraries
# - Hardware
# - Containers
# - Testing
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Testing
#
# **Types**
# - Unit testing
# - Regression testing
#
# **Tools**
# - Continuous integration services
# + [markdown] slideshow={"slide_type": "slide"}
# ## Verification and Validation
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Official Definitions
#
# **Software Validation**: The process of evaluating software during or at the end of the development process to determine whether it satisfies specified requirements. [IEEE-STD-610]
#
# **Software Verification**: The process of evaluating software to determine whether the products of a given development phase satisfy the conditions imposed at the start of that phase. [IEEE-STD-610]
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Perhaps more relevant
#
# **Verification** is the process of determining that a computer model, simulation, or federation of models and simulations implementations and their associated data accurately represent the developer's conceptual description and specifications.
#
# **Validation** is the process of determining the degree to which a model, simulation, or federation of models and simulations, and their associated data are accurate representations of the real world from the perspective of the intended use(s).
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Verification - Example
#
# **Given:**
# You have the linear advection PDE
# $$
# u_t + u_x = 0
# $$
# and have chosen a numerical method that will solve it theoretically with $\mathcal{O}(\Delta x, \Delta t)$ convergence rate.
#
# Verification of a software to solve this would mean that it converges to the true solution of the PDE at the expected rates.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Validation - Example
#
# **Given:**
# You need to compute a prediction of a fluid going down a pipe and have chosen the linear advection PDE to represent the real fluid flow.
#
# Validation would entail using your verified software to compare how well the real-world fluid flow behaved.
| 02_reproducibility_V_and_V.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##Import Libraries
from fuzzywuzzy import fuzz
##Build out Standard List
SL=['PRG','AFT','MEL','ANE','LVL','CustOps','EDS','Paulo']
def fm(x,SL):
scores={}
for i in SL:
scores.update({i:fuzz.token_set_ratio(x,i)+fuzz.partial_ratio(x,i)+fuzz.ratio(x,i)})
score_max = max(scores.keys(), key=(lambda k: scores[k]))
return score_max
##Test
fm('cust',SL)
| Gadgets/Fuzzy_String_Matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2: First look at data
# In this lesson we will look at a toy dataset simulating $J/\psi \rightarrow \mu^+ \mu^-$ events. We will discuss ways of loading the data in python, data formats and plotting with ```matplotlib```.
# ### Recap: Importing modules
#
# It's generally seen as good practice to put imports at the top of your file:
from matplotlib import pyplot as plt
import uproot
import numpy as np
import pandas as pd
from xgboost import XGBClassifier
from sklearn.ensemble import AdaBoostClassifier, GradientBoostingClassifier
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import KFold
# ## 5. The toy dataset
# We're going to look at some fake $J/\psi \rightarrow \mu^+ \mu^-$ data, the variables available are:
#
# - `Jpsi_M` `Jpsi_P` `Jpsi_PT` `Jpsi_PE` `Jpsi_PX` `Jpsi_PY` `Jpsi_PZ`
# - `mum_M` `mum_PT` `mum_eta` `mum_PE` `mum_PX` `mum_PY` `mum_PZ` `mum_IP` `mum_ProbNNmu` `mum_ProbNNpi`
# - `mup_M` `mup_PT` `mup_eta` `mup_PE` `mup_PX` `mup_PY` `mup_PZ` `mup_IP` `mup_ProbNNmu` `mup_ProbNNpi`
# - `nTracks`
#
# The meanings of the suffixes are as follows:
#
# - `_M`: Invarient mass of the particle (fixed to the PDG value for muons)
# - `_P`: Absolute value of the particle's three momentum
# - `_PT`: Absolute value of the particle's momentum in the `x`-`y` plane
# - `_PE`, `_PX`, `_PY`, `_PZ`: Four momentum of the particle
# - `_IP`: Impact parameter, i.e. the distance of closest approach between the reconstructed particle and the primary vertex
# - `ProbNNmu`, `ProbNNpi`: Particle identificaton variables which corrospond to how likely is it that the particle is really a muon or a pion
# - `nTracks`: The total number of tracks in the event
# ### Loading data
#
# - `root_numpy` and `root_pandas` are a way of reading+writing ROOT files
# - `uproot` is a way of reading ROOT files without having ROOT installed, see the github reposity [here](https://github.com/scikit-hep/uproot)
# - We can look at the objects that are available in the file and access objects using dictionary style syntax
# - The tree class contains converters to a varity of common Python libraries, such as numpy
# - We will also use `pandas DataFrames` to load data in a table like format
#
# First let's load the data using `uproot`.
#
# Often it is convenient to access data stored on the *grid* at CERN so you don't have to keep it locally. This can be done using the *XRootD* protocol:
#
# ```python
# my_file = uproot.open('root://eosuser.cern.ch//eos/user/l/lhcbsk/advanced-python/data/real_data.root')
# ```
#
# Accessing data this way requires you to have valid CERN credentials to access it. If authenication fails you will see an error message like:
#
# ```
# OSError: [ERROR] Server responded with an error: [3010] Unable to give access - user access restricted - unauthorized identity used ; Permission denied
# ```
#
# Credentials can be obtained by typing `kinit <EMAIL>` in your terminal and entering your CERN password.
#
# For this tutorial we will use a publically accessible file instead, using HTTPS to access it remotely. This is significantly slower then using XRootD.
my_file = uproot.open('https://cern.ch/starterkit/data/advanced-python-2018/real_data.root',
httpsource={'chunkbytes': 1024*1024, 'limitbytes': 33554432, 'parallel': 64})
my_file.keys()
tree = my_file['DecayTree']
# Get a numpy array containing the J/Ψ mass
tree.array('Jpsi_M')
# Load data as a pandas DataFrame
data_df = tree.pandas.df()
# Show the first 5 lines of the DataFrame
data_df.head()
# ## 6. Plotting a histogram with `matplotlib`
# Start with a basic histogram
plt.hist(data_df['Jpsi_M'])
plt.xlabel('Jpsi mass')
# That's okay but we could use some more bins, lets make it tidier and turn it into a function we can use later.
#
# Take a look at the `matplotlib` documentation:
# - https://matplotlib.org/api/_as_gen/matplotlib.pyplot.hist.html
# - It returns an array of counts, an array of bins and an array of patches. We don't care about the patches so we put them into a junk variable `_`.
# - Lets also set `histtype="step"` so we can plot multiple datasets on the same axis easily
# +
def plot_mass(df):
counts, bins, _ = plt.hist(df['Jpsi_M'], bins=100, range=[2.75, 3.5], histtype='step')
# You can also use LaTeX in the axis label
plt.xlabel('$J/\\psi$ mass [GeV]')
plt.xlim(bins[0], bins[-1])
plot_mass(data_df)
# -
# ### Adding variables
# When making the ROOT file we forgot to add some variables, no bother lets add them now!
data_df.eval('Jpsi_eta = arctanh(Jpsi_PZ/Jpsi_P)', inplace=True)
data_df.head()['Jpsi_eta']
# **Exercise:** Add `mu_P` and `mum_P` columns to the DataFrame.
data_df.eval('mup_P = sqrt(mup_PX**2 + mup_PY**2 + mup_PZ**2)', inplace=True)
data_df.eval('mum_P = sqrt(mum_PX**2 + mum_PY**2 + mum_PZ**2)', inplace=True)
# We can also get multiple columns at the same time
data_df.head()[['mum_P', 'mup_P']]
# ## Using rectangular cuts
# * We want to increase the 'signal significance' of our sample - this means more signal events with respect to background
# * To do this we can cut on certain discriminating variables
# * Here we will make cuts on the `Jpsi_PT` and **PID** (Particle Identification) variables
plot_mass(data_df)
data_with_cuts_df = data_df.query('Jpsi_PT > 4')
plot_mass(data_with_cuts_df)
plot_mass(data_df)
data_with_cuts_df = data_df.query('Jpsi_PT > 4')
plot_mass(data_with_cuts_df)
# Lets add some PID cuts as well
data_with_cuts_df = data_df.query('(Jpsi_PT > 4) & ((mum_ProbNNmu > 0.9) & (mup_ProbNNmu > 0.9))')
plot_mass(data_with_cuts_df)
# Let's go back and add a label argument to our plot function. This makes it easier to identify each line.
# We can also use the `density` argument in `matplotlib.hist` to plot all the histograms as the same scale.
# +
def plot_mass(df, **kwargs):
counts, bins, _ = plt.hist(df['Jpsi_M'], bins=100, range=[2.75, 3.5], histtype='step', **kwargs)
plt.xlabel('$J/\\psi$ mass [GeV]')
plt.xlim(bins[0], bins[-1])
plot_mass(data_df, label='No cuts', density=1)
data_with_cuts_df = data_df.query('Jpsi_PT > 4')
plot_mass(data_with_cuts_df, label='$J/\\psi$ p$_T$ only', density=1)
data_with_cuts_df = data_df.query('(Jpsi_PT > 4) & ((mum_ProbNNmu > 0.9) & (mup_ProbNNmu > 0.9))')
plot_mass(data_with_cuts_df, label='$J/\\psi$ p$_T$ and muon PID', density=1)
plt.legend(loc='best')
# -
# For this tutorial we have a special function for testing the significance of the signal in our dataset. There are many different ways to do this with real data, though we will not cover them here.
from python_lesson import check_truth
data_df.columns
# +
print('Originally the significance is')
check_truth(data_df)
print('\nCutting on pT gives us')
check_truth(data_df.query('Jpsi_PT > 4'))
print('\nCutting on pT and ProbNNmu gives us')
check_truth(data_df.query('(Jpsi_PT > 4) & ((mum_ProbNNmu > 0.9) & (mup_ProbNNmu > 0.9))'))
# -
# ### Comparing distributions
#
# Before we just used the cuts that were told to you but how do we pick them?
#
# One way is to get a sample of simulated data, we have a file in `data/simulated_data.root`.
#
# **Exercise:** Load it into a pandas `DataFrame` called `mc_df`. Don't forget to add the `Jpsi_eta`, `mup_P` and `mum_P` columns!
mc_df = uproot.open('https://cern.ch/starterkit/data/advanced-python-2018/simulated_data.root',
httpsource={'chunkbytes': 1024*1024, 'limitbytes': 33554432, 'parallel': 64}
)['DecayTree'].pandas.df()
mc_df.eval('Jpsi_eta = arctanh(Jpsi_PZ/Jpsi_P)', inplace=True)
mc_df.eval('mup_P = sqrt(mum_PX**2 + mum_PY**2 + mum_PZ**2)', inplace=True)
mc_df.eval('mum_P = sqrt(mum_PX**2 + mum_PY**2 + mum_PZ**2)', inplace=True)
# **QUESTION:** What can we to get a background sample?
#
# Sidebands, we know the peak is only present in $3.0~\text{GeV} < M(J/\psi) < 3.2~\text{GeV}$. If we select events outside the region we know it's a pure background sample.
#
# **Exercise:** Make a new `DataFrame` called `bkg_df` containing only events outside $3.0~\text{GeV} < M(J/\psi) < 3.2~\text{GeV}$.
bkg_df = data_df.query('~(3.0 < Jpsi_M < 3.2)')
plot_mass(bkg_df)
# **QUESTION:** Why is there a step at 3 GeV on this plot?
#
# It's a binning effect, we've appled a cut at 3.0 GeV but the nearest bin is $[2.9975, 3.005]$ so it is only partially filled.
# Now let's plot the variables in MC and background to see what they look like?
var = 'Jpsi_PT'
_, bins, _ = plt.hist(mc_df[var], bins=100, histtype='step', label='MC')
_, bins, _ = plt.hist(bkg_df[var], bins=bins, histtype='step', label='Background')
plt.xlabel(var)
plt.xlim(bins[0], bins[-1])
plt.legend(loc='best')
# Those are hard to compare!!!
# We should add the density keyword argument to normalise the distributions
var = 'Jpsi_PT'
_, bins, _ = plt.hist(mc_df[var], bins=100, histtype='step', label='MC', density=1)
_, bins, _ = plt.hist(bkg_df[var], bins=bins, histtype='step', label='Background', density=1)
plt.xlabel(var)
plt.xlim(bins[0], bins[-1])
plt.legend(loc='best')
# **Exercise:** Make a function which plots both variables with the signature `plot_comparision(var, mc_df, bkg_df)`.
def plot_comparision(var, mc_df, bkg_df):
_, bins, _ = plt.hist(mc_df[var], bins=100, histtype='step', label='MC', density=1)
_, bins, _ = plt.hist(bkg_df[var], bins=bins, histtype='step', label='Background', density=1)
plt.xlabel(var)
plt.xlim(bins[0], bins[-1])
plt.legend(loc='best')
# We can now use this function to plot all of the variables available in the data using `data_df.columns`:
for var in data_df.columns:
plt.figure()
plot_comparision(var, mc_df, bkg_df)
# Things to note:
#
# - This doesn't work for the $J/\psi$ mass variable: We can only rely on this method if the variable is independent of mass. Fortunately we often want to do this as if a variable is heavily dependent on mass it can shape our distributions and can make it hard to know what is signal and what is background.
# - Muon mass is a fixed value for all muons: In this sample we have assumed the PDG value of the muon mass to allow us to calculate the energy component using only the information from the tracking detectors. This is often more precise than using calorimeters to measure $P_E$.
# - We got a warning about `More than 20 figures have been opened.`: Opening plots uses memory so if you open too many at the same time your scripts can be become slow or even crash. In this case we can ignore it as we only produce 30 plots but be careful if you ever make thousands of plots.
# - Pseudorapidity (eta) only goes between about 1 and 6: This dataset is supposed to look like vaugely like LHCb data where the detector only covers that region.
# **Exercise:** Look at the variables above and try to get a clean $J/\psi$ mass peak and use the significance function to see how well you do.
# **Aside:** We will want to use some of the variables and dataframes in the next lesson. In order to do this we will *store* them in this session and reload them in the next lesson.
# %store bkg_df
# %store mc_df
# %store data_df
| advanced-python/2DataAndPlotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Image smoothing via 2D convolution
#
# In section 10.3.1, we discussed how convolving an image with a smoothing kernel can help eliminate noisy artifacts and produce a smooth output. This notebook contains the fully functional code for the same.
import torch
import torch.nn.functional as F
import cv2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# +
def load_grayscale_image(img_path):
"""
Takes an image path, reads the image as a gray scale image and converts
the resulting image to a tensor
"""
return cv2.imread(img_path, cv2.IMREAD_GRAYSCALE)
def img_to_tensor(img):
"""
Takes a numpy array corresponding to an image and converts it to a Pytorch tensor
"""
return torch.from_numpy(img).float()
def tensor_to_img(x):
"""
Takes a tensor, does min-max normalization to scale the tensor to a value between 0 and 255
and returns the resulting numpy array
"""
x_norm = ((x - x.min()) / (x.max() - x.min())) * 255
return x_norm.to(torch.uint8).numpy()
# -
# First, let us load an image that contains noise. If you look carefully at the image shown below, you can see small white dots present across the entire image. Our goal is to remove these white dots to obtain a cleaner image.
noisy_img = load_grayscale_image("Figures/ml_noisy_full_res.png")
# Let us take a look at the image and plot it
fig = plt.figure(figsize = (20, 10))
plt.xticks([])
plt.yticks([])
imgplot = plt.imshow(noisy_img.astype(np.float32), cmap="gray")
noisy_img_tensor = img_to_tensor(noisy_img)
# Let us now define a 3x3 smoothing kernel that we will convolve over the entire image. The smoothing kernel is a uniform matrix with equal weights, i.e.
# $$
# \begin{bmatrix}
# \frac{1}{9} & \frac{1}{9} & \frac{1}{9}\\
# \frac{1}{9} & \frac{1}{9} & \frac{1}{9}\\
# \frac{1}{9} & \frac{1}{9} & \frac{1}{9}\\
# \end{bmatrix}
# $$
#
# This means that the output value at any given location is the average of neighbouring 3x3 input pixels. The noisy white pixels are surrounded by black pixels in the input image. Hence, the output pixel would be much closer to a black pixel than a white pixel, resulting in a de-noised output.
# Let us define the smoothing kernel
kernel = torch.tensor([[1, 1, 1], [1, 1, 1], [1, 1, 1]], dtype=torch.float32)
# Conv 2D kernels need to be of shape (out_channel, in_channel, k_x, k_y). Since we are defining a single kernel
# out_channel = in_channel = 1. So we unsqueeze to add those extra dimensions
kernel = kernel.unsqueeze(0).unsqueeze(0)
kernel = kernel/ 9
print(f"Kernel shape {kernel.shape}\nKernel: {kernel}") # 1x1x3x3
weight = torch.nn.Parameter(data=kernel, requires_grad=False)
# Let us now convolve the smoothing kernel over the noisy input image. 2D Convolution operations work on 4D tensors of the form (N, C, H, W), where N is the batch size, C is the number of channels, H is height and W is width. Since we are dealing with a single grayscale image, N and C are both 1.
# +
# Convert tensors to N x C x H x W
noisy_img_batch = noisy_img_tensor.unsqueeze(0).unsqueeze(0)
# Convolution can be done in 2 ways.
# 1) Using conv2d function
smooth_img_tensor_1 = F.conv2d(noisy_img_batch, weight, padding=1)
# 2) Using Conv2d layer
conv2d = torch.nn.Conv2d(1, 1, kernel_size=[3, 3], stride=1, padding=0, bias=False)
conv2d.weight = weight
with torch.no_grad():
smooth_img_tensor_2 = conv2d(noisy_img_batch)
# -
# Let us assert both the ways yielded the same result
assert torch.allclose(smooth_img_tensor_1, smooth_img_tensor_1)
smooth_img_tensor = smooth_img_tensor_2[0][0]
smooth_img = tensor_to_img(smooth_img_tensor)
# Let us take a look at the image and plot it
fig = plt.figure(figsize = (20, 10))
# For better visualization, let us threshold the smoothened image before plotting
_, smooth_img = cv2.threshold(smooth_img, 120, 255, cv2.THRESH_TOZERO)
plt.xticks([])
plt.yticks([])
imgplot = plt.imshow(smooth_img.astype(np.float32), cmap="gray")
| python/ch10/10.3.1-2dconv-image-smoothing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from sklearn.neural_network import MLPClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import classification_report,confusion_matrix,accuracy_score
import pandas as pd
from pandas import DataFrame,Series
from matplotlib.colors import ListedColormap
import numpy as np
from sklearn.metrics import accuracy_score,confusion_matrix,classification_report
import matplotlib.pyplot as plt
from pandas.plotting import scatter_matrix
from random import sample
# +
multi_layer_dup_train = pd.read_csv('../FeaturesCsvFile/featuresfile.csv')
multi_layer_dup_test = pd.read_csv('../FeaturesCsvFile/featuresfile_10.csv')
multi_layer_train = multi_layer_dup_train.drop_duplicates(subset=['User', 'Timestamp'])
multi_layer_unique_test = multi_layer_dup_test.drop_duplicates(subset=['User', 'Timestamp'])
multi_layer_test = multi_layer_unique_test.iloc[sample(range(len(multi_layer_unique_test)), 40), :]
print ('(#row,#column) of train dataset' , multi_layer_train.shape)
print ('(#row,#column) of test dataset' , multi_layer_test.shape)
# -
X_train = multi_layer_train.values[:, 2:45]
y_train = multi_layer_train.values[:, 45]
X_test = multi_layer_test.values[:, 2:45]
y_test = multi_layer_test.values[:, 45]
scaler = StandardScaler()
scaler.fit(X_train)
StandardScaler(copy=True, with_mean=True, with_std=True)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mlp = MLPClassifier(hidden_layer_sizes=(20,),max_iter=60)
mlp_pred=mlp.fit(X_train,y_train)
y_pred = mlp.predict(X_test)
print(confusion_matrix(y_test,y_pred))
print(classification_report(y_test,y_pred))
print('\nAccuracy of Multi-layer Perceptron Score: %.2f' % mlp.score(X_test,y_test))
print('\nAccuracy of Accuracy Score : %.2f' % accuracy_score(y_test,y_pred))
for i in range(0,len(mlp.coefs_[0])):
print mlp.coefs_[0][i]
avg_weight = []
for i in range(0,len(mlp.coefs_[0])):
avg_weight.append(np.mean(mlp.coefs_[0][i]))
print ('Important features (featureName, weigh of important, #column)')
header = list(multi_layer_train.head(1))
important_feature = []
for i in range(0,len(avg_weight)):
important_feature.append((header[i+2],avg_weight[i],i+2))
sorted_list = sorted(important_feature,key=lambda important_feature: important_feature[1],reverse=True)
for j in range(0,len(sorted_list)):
first_imp_fea = sorted_list[0]
second_imp_fea = sorted_list[1]
print sorted_list[j]
# +
from sklearn import metrics
def plot_roc_curve(Y_predict,Y_test,name_graph):
num_predns = []
for i in range(0,len(Y_predict)):
if Y_predict[i] == "walking":
num_predns.append(0)
else:
num_predns.append(1)
num_labels = []
for i in range(0,len(Y_test)):
if Y_test[i] == "walking":
num_labels.append(0)
else:
num_labels.append(1)
predns = np.array(num_predns)
labels = np.array(num_labels)
fpr, tpr, thresholds = metrics.roc_curve(labels, predns)
roc_auc = metrics.auc(fpr, tpr)
plt.title('Area under ROC Curve')
plt.plot(fpr, tpr, 'grey', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
# plt.show()
plt.savefig('./image/roc_multipc.png', dpi=100)
plot_roc_curve(y_pred,y_test,"Area_under_roc_pc")
# +
import itertools
import numpy as np
import matplotlib.pyplot as plt
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Greens):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes)
plt.yticks(tick_marks, classes, rotation=90)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
# plt.figure()
class_names = ["walking", "running"]
plot_confusion_matrix(cnf_matrix, classes=["walking", "running"],
title='Confusion matrix, without normalization')
plt.savefig('./image/confusion_matrix_multipc.png', dpi=100)
# plt.savefig('H:/mastersProject/activity_analyzer/LogisticRegression/cm_lr', dpi=1000)
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=class_names, normalize=True,
title='Normalized confusion matrix')
plt.show()
| NeuronBinaryClassification/Jupyter_Notebook/MultiLayerPerceptrop_3_train_10_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import querries
import os
import sys
sys.path.insert(1, '../')
from data_quality.scripts.utils_cleaning import clean_data
from data_quality.scripts.utils import enhance_data, clean_dates
df = pd.read_csv("./Data/msda_data_clinicians.csv")
#Needed to interface with cleaning scripts
df["id"] = np.arange(df.shape[0])
date_cols = [c for c in df.columns if "date" in c]
for date_col in date_cols:
df[date_col] = pd.to_datetime(df[date_col],errors="coerce")
#Clean
print("Cleaning data ....")
df = clean_data(df,None,None)
#Augment
print("Creating new variables ....")
df = clean_dates(df)
df["report_source"]="clinicians"
df = enhance_data(df)
df["secret_name"] = np.arange(df.shape[0])
#Compute the tables
print("Computing tables ....")
querries.compute_tables(df, report_source = "clinicians")
print("Done !")
os.makedirs("./Outputs", exist_ok=True)
file_name = "clinicians_query3_bmi_in_cat2"
outcome_types = ["covid19_admission_hospital","covid19_icu_stay","covid19_ventilation","covid19_outcome_death","covid19_outcome_ventilation_or_ICU", "covid19_outcome_levels_1", "covid19_outcome_levels_2"]
for outcome_type in outcome_types:
df = pd.read_csv(f"./results/{file_name}_{outcome_type}.csv")
if outcome_type=="covid19_admission_hospital":
variables_list = ["dmt_type_overall","age_in_cat","ms_type2","sex_binary","edss_in_cat2"]
variables_list += ["covid19_diagnosis"]
variables_list += [outcome_type]
else:
variables_list = [outcome_type]
for variable in variables_list:
# print(df.groupby(variable)["secret_name"].sum()) #tables to return.
result = (df.groupby(variable)["secret_name"].sum())
result.to_json(f"./Outputs/clinicians-{variable}.json")
with open("DoneC.txt", "w") as file:
file.write("query3_clinicians Process is Done")
| MSDA_Query/query3_clinicians.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NobuoTsukamoto/tensorrt-examples/blob/main/python/detection/Add_TFLiteNMS_Plugin.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="f598iir_p1U4"
# # Convert a SSDLite MobileNet V2 TFLite model to ONNX and Add TFLite NMS Plugin.
#
# This notebook converts from TensorFlow Lite model to ONNX model and replaces NonMax Suppression with TensorRT's TF-Lite NMS Plugin.
# The model uses [SSD Lite MobileNet V2](https://github.com/tensorflow/models/blob/a4fd64722dcdd42361beb1be478ad8fdb10bde31/research/object_detection/g3doc/tf1_detection_zoo.md) and modifies the ONNX model with [ONNX GraphSurgeon](https://github.com/NVIDIA/TensorRT/tree/master/tools/onnx-graphsurgeon).
# + [markdown] id="heEyNGCd-dgM"
# Copyright (c) 2021 <NAME>
#
# This software is released under the MIT License.
# See the LICENSE file in the project root for more information.
#
# + [markdown] id="6VghN88vg2NM"
# ## Reference
# - [onnx/tensorflow-onnx - Convert a mobiledet tflite model to ONNX](https://github.com/onnx/tensorflow-onnx/blob/de67f2051d1f036b29901e2910c8eb41a1c71b6e/tutorials/mobiledet-tflite.ipynb)
# - [TensorFlow Model Garden - TensorFlow 1 Detection Model Zoo](https://github.com/tensorflow/models/blob/a4fd64722dcdd42361beb1be478ad8fdb10bde31/research/object_detection/g3doc/tf1_detection_zoo.md)
# - [TensorFlow Model Garden - Running on mobile with TensorFlow Lite](https://github.com/tensorflow/models/blob/a4fd64722dcdd42361beb1be478ad8fdb10bde31/research/object_detection/g3doc/running_on_mobile_tensorflowlite.md)
# - [TensorRT Backend For ONNX](https://github.com/onnx/onnx-tensorrt/tree/868e636f51f0d7e61df340371303275265146fe0)
# - [ONNX GraphSurgeon](https://github.com/NVIDIA/TensorRT/tree/0953f2ff8762b28e0f1bef0582b6ca3d7a12fcaa/tools/onnx-graphsurgeon)
# + [markdown] id="92EB-Un1htTK"
# ## Convert the TensorFlow Lite Model.
# + id="XDwBImKugaIU" colab={"base_uri": "https://localhost:8080/"} outputId="6e96a5b3-32dc-43e8-e388-c262f10105a8"
# %tensorflow_version 1.x
# + [markdown] id="EEd6ynRMiQLJ"
# Clone [tensorflow/models](https://github.com/tensorflow/models) repository and install dependency.
# + colab={"base_uri": "https://localhost:8080/"} id="8qHlK2mdgp4X" outputId="fcec66f7-fe20-4e84-97c5-aa0c8c7720be" language="bash"
#
# pip install tensorflow-addons
# git clone https://github.com/tensorflow/models.git
# cd models
# git checkout a4fd64722dcdd42361beb1be478ad8fdb10bde31
#
# cd research
# protoc object_detection/protos/*.proto --python_out=.
#
# cp object_detection/packages/tf1/setup.py .
# python -m pip install .
# + colab={"base_uri": "https://localhost:8080/"} id="qJiVt0nbgxYj" outputId="5a61c165-504f-4a6e-ed89-ec3a1b09fa94"
import os
os.environ['PYTHONPATH'] = '/content/models:' + os.environ['PYTHONPATH']
print(os.environ['PYTHONPATH'])
# + colab={"base_uri": "https://localhost:8080/"} id="I5PPsFUUhNAQ" outputId="a01e0bfe-d3c8-42cc-8d7d-bde9cbc7a2ef"
# %cd models/research/
# !python object_detection/builders/model_builder_tf1_test.py
# + [markdown] id="kvLFSF2hivaA"
# Download SSDLite MobileNet V2 checkpoint and export TensorFlow Lite model.
# + colab={"base_uri": "https://localhost:8080/"} id="AZUJyMjEm5d6" outputId="028b8817-a091-4bf2-dd64-024441e0a229"
# !wget http://download.tensorflow.org/models/object_detection/ssdlite_mobilenet_v2_coco_2018_05_09.tar.gz -P /content
# !tar xf /content/ssdlite_mobilenet_v2_coco_2018_05_09.tar.gz -C /content
# + colab={"base_uri": "https://localhost:8080/"} id="fJEzEHIMgWhV" outputId="950dd63f-9e5b-4e5f-846d-79434f8edcc3"
# !python object_detection/export_tflite_ssd_graph.py \
# --pipeline_config_path="/content/ssdlite_mobilenet_v2_coco_2018_05_09/pipeline.config" \
# --trained_checkpoint_prefix="/content/ssdlite_mobilenet_v2_coco_2018_05_09/model.ckpt" \
# --output_directory="/content/ssdlite_mobilenet_v2_coco_2018_05_09/tflite" \
# --add_postprocessing_op=true
# + colab={"base_uri": "https://localhost:8080/"} id="5z0bcYh4iE8a" outputId="19c31a5d-64d6-4176-dbf7-be0f7d7968b2"
# !tflite_convert \
# --enable_v1_converter \
# --graph_def_file="/content/ssdlite_mobilenet_v2_coco_2018_05_09/tflite/tflite_graph.pb" \
# --output_file="/content/ssdlite_mobilenet_v2_coco_2018_05_09/tflite/ssdlite_mobilenet_v2_320x320.tflite" \
# --inference_input_type=FLOAT \
# --inference_type=FLOAT \
# --input_arrays="normalized_input_image_tensor" \
# --output_arrays="TFLite_Detection_PostProcess,TFLite_Detection_PostProcess:1,TFLite_Detection_PostProcess:2,TFLite_Detection_PostProcess:3" \
# --input_shapes=1,300,300,3 \
# --allow_nudging_weights_to_use_fast_gemm_kernel=true \
# --allow_custom_op
# + [markdown] id="DfAP2oH6qTaU"
# ## Export ONNX Model
# + [markdown] id="faSTJYvxnKch"
# Install dependency.
# + colab={"base_uri": "https://localhost:8080/"} id="knvpxQlzjfOP" outputId="b72de806-2193-479d-8150-4eb9395688c6" language="bash"
# pip3 install onnxruntime
# pip3 install tf2onnx
# + [markdown] id="I10zwTrynWYJ"
# Note: TensorRT 7.2 supports operators up to Opset 13.
# - [onnx/onnx-tensorrt - Supported ONNX Operators](https://github.com/onnx/onnx-tensorrt/blob/868e636f51f0d7e61df340371303275265146fe0/docs/operators.md)
# + colab={"base_uri": "https://localhost:8080/"} id="fbTs3y5nj_dX" outputId="e2bc164e-4802-4918-b2a0-7ff83f15c1fd"
# !python3 -m tf2onnx.convert --opset 11 \
# --tflite /content/ssdlite_mobilenet_v2_coco_2018_05_09/tflite/ssdlite_mobilenet_v2_320x320.tflite \
# --output /content/ssdlite_mobilenet_v2_coco_2018_05_09/onnx/ssdlite_mobilenet_v2_320x320.onnx
# + [markdown] id="KN5E-Q5uqo3y"
# ## Add TF-Lite NMS Plugin
# + id="kCQ1qTMvk1nc" colab={"base_uri": "https://localhost:8080/"} outputId="2d386cdb-d2ff-4769-cd67-cdaf8a014901"
# !python3 -m pip install onnx_graphsurgeon --index-url https://pypi.ngc.nvidia.com
# + colab={"base_uri": "https://localhost:8080/"} id="USJqsm-7lRK1" outputId="6d8b59e3-adfd-4149-c739-0a87a7f75e45"
# %cd /content/
# !git clone https://github.com/NobuoTsukamoto/tensorrt-examples
# + id="EZSIts0rlciG" colab={"base_uri": "https://localhost:8080/"} outputId="58d0957f-907d-439c-b356-b8371c4a76d5"
# %cd /content/tensorrt-examples/python/detection
# + id="2nOXo6yJnfD-" colab={"base_uri": "https://localhost:8080/"} outputId="7cd6c371-d8b2-40ae-c56d-234ffcf0921b"
# !python3 add_tensorrt_tflitenms_plugin.py \
# --input /content/ssdlite_mobilenet_v2_coco_2018_05_09/onnx/ssdlite_mobilenet_v2_320x320.onnx \
# --output /content/ssdlite_mobilenet_v2_coco_2018_05_09/onnx/ssdlite_mobilenet_v2_320x320_gs.onnx
| python/detection/Add_TFLiteNMS_Plugin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # TEXT MINING for PRACTICE
# - 본 자료는 텍스트 마이닝을 활용한 연구 및 강의를 위한 목적으로 제작되었습니다.
# - 본 자료를 강의 목적으로 활용하고자 하시는 경우 꼭 아래 메일주소로 연락주세요.
# - 본 자료에 대한 허가되지 않은 배포를 금지합니다.
# - 강의, 저작권, 출판, 특허, 공동저자에 관련해서는 문의 바랍니다.
# - **Contact : ADMIN(<EMAIL>)**
#
# ---
# ## WEEK 04-3. 동적 페이지 수집하기: 네이버 뉴스기사
# - Python을 활용해 가상의 브라우저를 띄워 웹페이지에서 데이터를 크롤링하는 방법에 대해 다룹니다.
#
# ---
# > **\*\*\* 주의사항 \*\*\***
# 본 자료에서 설명하는 웹크롤링하는 방법은 해당 기법에 대한 이해를 돕고자하는 교육과 이를 활용한 연구 목적으로 사용되었으며, 대량의 무단 크롤링 및 상업적 활용을 금합니다.
# ## 1. 데이터 수집 준비하기
# ### 1-1. Selenium 라이브러리 설치
# +
# 가상의 브라우저를 컨트롤 할 수 있도록 도와주는 selenium 패키지를 설치합니다.
# 아래 주석을 해지하고 셀을 실행합니다.
# 설치는 한번만 수행하면 되며, 재설치시 Requirement already satisfied: ~ 라는 메시지가 출력됩니다.
# #!pip install selenium
# -
# ### 1-2. 라이브러리 Import 및 Chrome Driver 실행
# +
# Python 코드를 통해 가상의 브라우저를 띄우기 위해 selenium 패키지를 import 합니다.
import selenium
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.chrome.service import Service
# selenium을 활용해 브라우저를 직접 띄우는 경우, 실제 웹서핑을 할때처럼 로딩시간이 필요합니다.
# 로딩시간 동안 대기하도록 코드를 구성하기위해 time 패키지를 import 합니다.
import time
# Python 코드를 통해 웹페이지에 정보를 요청하기 위해 BeautifulSoup, urllib 패키지를 import 합니다.
from bs4 import BeautifulSoup
import requests
# +
# Chrome Driver를 호출합니다.
chrome_options = webdriver.ChromeOptions()
# 브라우저에 임의로 User-agent 옵션을 넣어 Python 코드로 접속함을 숨깁니다.
chrome_options.add_argument('--user-agent="Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_4) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/66.0.3359.139 Safari/537.36"')
# Chrome Driver 파일의 경로를 지정하고 실행합니다.
# Chrome Driver는 아래 링크에서 다운로드 가능합니다.
# 본 Jupyter Notebook 파일과 동일한 경로에 Chrome Driver가 존재하는 경우 아래 경로를 그대로 사용합니다.
#service = Service("./chromedriver") # Windows 운영체제
service = Service("./chromedriver") # MAC, Linux 운영체제
# 경고메시지 출력시 조치 : [시스템 환경설정] > [보안 및 개인정보 보호] > "Chrome Drive ~ 확인없이 허용"
driver = webdriver.Chrome(service=service, options=chrome_options)
# -
# ## 2. 뉴스기사 및 댓글 수집하기
#
# ---
# ### 2-1. 뉴스기사 정보 입력
# 수집할 뉴스기사 정보를 입력합니다.
QUERY = "코로나19" # 필터링 키워드
START_DATE = "2022.01.01" # 필터링 일자 (작성일 기준)
END_DATE = "2022.01.30"
START_PAGE = 1 # 검색결과 저장 페이지 범위 (네이버 뉴스기사는 검색결과 중 최대 4,000페이지만 제공)
END_PAGE = 2
# 기사와 댓글을 저장할 파일명
article_filename = "article_" + QUERY + "_" + START_DATE + "_" + END_DATE + "_" + str(START_PAGE) + ".txt"
comment_filename = "comment_" + QUERY + "_" + START_DATE + "_" + END_DATE + "_" + str(START_PAGE) + ".txt"
# ### 2-2. 뉴스기사 수집
# +
fa = open(article_filename, "w", encoding="utf-8")
fc = open(comment_filename, "w", encoding="utf-8")
news_count = 0
for page in range(START_PAGE, END_PAGE+1, 10):
#print(page, "번째 기시부터 최대 10개 수집중...", end="\r")
URL = "https://search.naver.com/search.naver?&where=news&query=" + QUERY
URL += "&sm=tab_pge&sort=2&photo=0&field=0&reporter_article=&pd=3&ds="
URL += START_DATE + "&de=" + END_DATE + "&docid=&&start=" + str(page) + "&refresh_start=0"
driver.get(URL)
time.sleep(2)
try:
news_list = driver.find_element(By.CLASS_NAME, "list_news").find_elements(By.CLASS_NAME, "bx")
except:
break
news_count += len(news_list)
for news in news_list[:]:
link_list = news.find_element(By.CLASS_NAME, "info_group").find_elements(By.TAG_NAME, "a")
if len(link_list) == 1:
continue
article_url = link_list[1].get_attribute("href").strip()
link_list[1].click()
time.sleep(3)
current_window = driver.current_window_handle
try:
new_window = [window for window in driver.window_handles if window != current_window][0]
driver.switch_to.window(new_window)
except:
driver.switch_to.window(current_window)
continue
time.sleep(4)
try:
try:
source_label = driver.find_element(By.CLASS_NAME, "press_logo")
except:
source_label = driver.find_element(By.ID, "pressLogo")
source_img = source_label.find_element(By.TAG_NAME, "img")
source = source_img.get_attribute("alt").strip()
datetime = ""
content = ""
title = ""
etc_good_count = ""
etc_warm_count = ""
etc_sad_count = ""
etc_angry_count = ""
etc_want_count = ""
ent_good_count = ""
ent_cheer_count = ""
ent_congrats_count = ""
ent_expect_count = ""
ent_suprise_count = ""
ent_sad_count = ""
basic_good_count = ""
basic_sad_count = ""
basic_angry_count = ""
basic_fan_count = ""
basic_want_count = ""
try:
# 기타 섹션 기사
datetime = driver.find_element(By.CLASS_NAME, "t11").text.strip()
content = driver.find_element(By.CLASS_NAME, "_article_body_contents").text.strip().replace("\n", " ")
title = driver.find_element(By.CLASS_NAME, "tts_head").text.strip()
reaction_list = driver.find_element(By.CLASS_NAME, "end_btn").find_element(By.CLASS_NAME, "_reactionModule").find_elements(By.TAG_NAME, "a")
etc_good_count = reaction_list[0].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
etc_warm_count = reaction_list[1].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
etc_sad_count = reaction_list[2].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
etc_angry_count = reaction_list[3].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
etc_want_count = reaction_list[4].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
except:
try:
# 연예 섹션 기사
datetime = driver.find_element(By.CLASS_NAME, "author")
datetime = datetime.find_element(By.TAG_NAME, "em").text.strip()
content = driver.find_element(By.ID, "articeBody").text.strip().replace("\n", " ")
title = driver.find_element(By.CLASS_NAME, "end_tit").text.strip()
reaction_list = driver.find_element(By.CLASS_NAME, "end_btn").find_element(By.CLASS_NAME, "_reactionModule").find_elements(By.TAG_NAME, "a")
ent_good_count = reaction_list[0].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
ent_cheer_count = reaction_list[1].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
ent_congrats_count = reaction_list[2].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
ent_expect_count = reaction_list[3].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
ent_suprise_count = reaction_list[4].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
ent_sad_count = reaction_list[5].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
except:
# 일반 섹션 기사
datetime = driver.find_element(By.CLASS_NAME, "info").find_element(By.TAG_NAME, "span").text.replace("기사입력", "").strip()
content = driver.find_element(By.CLASS_NAME, "news_end").text.strip().replace("\n", " ")
title = driver.find_element(By.CLASS_NAME, "title").text.strip()
reaction_list = driver.find_element(By.CLASS_NAME, "news_end_btn").find_element(By.CLASS_NAME, "_reactionModule").find_elements(By.TAG_NAME, "a")
basic_good_count = reaction_list[0].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
basic_sad_count = reaction_list[1].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
basic_angry_count = reaction_list[2].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
basic_fan_count = reaction_list[3].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
basic_want_count = reaction_list[4].find_element(By.CLASS_NAME, "u_likeit_list_count").text.strip()
review_count_list = driver.find_elements(By.CLASS_NAME, "u_cbox_count")
if len(review_count_list) > 0:
review_count = review_count_list[0].text.replace(",", "")
else:
review_count = "0"
#print(source + "\t" + datetime + "\t" + review_count + "\t" +
# good_count + "\t" + warm_count + "\t" + sad_count + "\t" +
# angry_count + "\t" + want_count + "\t" + article_url + "\t" +
# title + "\t" + content)
print(" "*100, end="\r")
print(str(page)+"/"+str(END_PAGE+1)+" Page,", title + " (댓글: "+str(review_count)+"건)", end="\r")
fa.write(source + "\t" + datetime + "\t" + review_count + "\t" +
ent_good_count + "\t" + ent_cheer_count + "\t" + ent_congrats_count + "\t" +
ent_expect_count + "\t" + ent_suprise_count + "\t" + ent_sad_count + "\t" +
basic_good_count + "\t" + basic_sad_count + "\t" + basic_angry_count + "\t" +
basic_fan_count + "\t" + basic_want_count + "\t" +
etc_good_count + "\t" + etc_warm_count + "\t" + etc_sad_count + "\t" +
etc_angry_count + "\t" + etc_want_count + "\t" +
article_url + "\t" +
title + "\t" + content + "\n")
# 댓글 더보기 클릭
if int(review_count) > 0:
driver.find_element(By.CLASS_NAME, "u_cbox_in_view_comment").click()
time.sleep(3)
# 더보기 버튼 클릭
for i in range(10):
try:
more_button_status = driver.find_element(By.CLASS_NAME, "u_cbox_paginate").get_attribute("style").strip()
if more_button_status == '':
driver.find_element(By.CLASS_NAME, "u_cbox_more_wrap").click()
time.sleep(1)
except:
continue
# 답글 클릭
comment_list = driver.find_elements(By.CLASS_NAME, "u_cbox_area")
driver.execute_script('window.scrollTo(0, 0);')
for comment in comment_list:
reply_count_list = comment.find_elements(By.CLASS_NAME, "u_cbox_btn_reply")
if len(reply_count_list) > 0:
reply_count = reply_count_list[0].text.strip()
if reply_count != "답글0":
#reply_button = comment.find_element(By.CLASS_NAME, "u_cbox_btn_reply")
#reply_button.click()
reply_count_list[0].click()
time.sleep(3)
# 댓글 수집
driver.execute_script('window.scrollTo(0, 0);')
comment_box_list = driver.find_elements(By.CLASS_NAME, "u_cbox_area")
for i in range(len(comment_box_list)):
comment_box = comment_box_list[i]
comment_nick = comment_box.find_element(By.CLASS_NAME, "u_cbox_nick").text
comment_content = comment_box.find_element(By.CLASS_NAME, "u_cbox_text_wrap").text
comment_datetime = comment_box.find_element(By.CLASS_NAME, "u_cbox_date").text
try:
comment_good = comment_box.find_element(By.CLASS_NAME, "u_cbox_cnt_recomm").text
comment_bed = comment_box.find_element(By.CLASS_NAME, "u_cbox_cnt_unrecomm").text
except:
comment_good = "0"
comment_bed = "0"
reply_count_list = comment_box.find_elements(By.CLASS_NAME, "u_cbox_reply_cnt")
if len(reply_count_list) > 0:
is_reply = "1"
parent_id = "-1"
temp_parent_id = i
reply_count = reply_count_list[0].text
else:
is_reply = "0"
parent_id = str(temp_parent_id)
reply_count = "0"
#print(str(i) + "\t" + is_reply + "\t" + reply_count + "\t" +
# parent_id + "\t" + comment_nick + "\t" + comment_datetime + "\t" +
# comment_good + "\t" + comment_bed + "\t" + comment_content + "\t" +
# article_url)
fc.write(str(i) + "\t" + is_reply + "\t" + reply_count + "\t" +
parent_id + "\t" + comment_nick + "\t" + comment_datetime + "\t" +
comment_good + "\t" + comment_bed + "\t" + comment_content + "\t" +
article_url + "\n")
except:
driver.close()
time.sleep(3)
driver.switch_to.window(current_window)
fa.flush()
fc.flush()
continue
driver.close()
time.sleep(3)
driver.switch_to.window(current_window)
fa.flush()
fc.flush()
fa.close()
fc.close()
# 수집종료
print()
print("* 최대", news_count, "개 기사 수집이 완료되었습니다.")
print("* 수집된 기사는 아래 파일에 저장되었습니다.")
print(" - 기사본문 :", article_filename)
print(" - 댓글 :", comment_filename)
# -
driver.close()
# > **\*\*\* TIP \*\*\***
# 새탭에서 기사가 열린 후 수집이 종료된 경우 아래 셀을 실행합니다.
driver.switch_to.window(current_window)
| practice-note/week_04/W04-3_text-mining-for-practice_python-crawling-practice-4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="CXwaJiZXC-QZ"
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy import stats
import seaborn as sns
import scipy.stats as ss
from pandas_datareader import DataReader
from datetime import datetime
# Make plots larger
plt.rcParams['figure.figsize'] = (15, 9)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="mfn2jAjlC-Qe" outputId="a1b4c203-7361-4532-f580-8f77f88535a6"
facebook = DataReader('FB', 'yahoo', datetime(2016,4,1), datetime(2019,9,1));
facebook.reset_index(inplace=True,drop=False)
facebook.set_index('Date')
facebook.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="zX_IclzpC-Qi" outputId="dd84379e-4455-4852-cecc-29f9bc8afcec"
# Plot of Open vs date
plt.plot(facebook['Date'], facebook['Open'])
plt.show()
# + colab={} colab_type="code" id="M8lVoz0sC-Ql"
openmean= (facebook['Open']+ facebook['Close'])/2
#print(openmean)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="L7iYo4qaC-Qo" outputId="c254830a-1f67-4cd0-b419-d4d8af62f3c6"
Y=openmean
print(openmean.var())
# + [markdown] colab_type="text" id="3tVoXP7QC-Qt"
# ### KALMAN FILTER
# + colab={} colab_type="code" id="OZlDEjcnC-Qt"
import numpy as np
from scipy.optimize import minimize
import matplotlib.pyplot as plt
def Kalman_Filter(Y):
S = Y.shape[0]
S = S + 1
Z = param0[0]
T = param0[1]
H = param0[2]
Q = param0[3]
u_predict = np.zeros(S)
u_update = np.zeros(S)
P_predict = np.zeros(S)
P_update = np.zeros(S)
v = np.zeros(S)
F = np.zeros(S)
KF_Dens = np.zeros(S)
for s in range(1,S):
if s == 1:
P_update[s] = 1000
P_predict[s] = T*P_update[1]*np.transpose(T)+Q
else:
F[s] = Z*P_predict[s-1]*np.transpose(Z)+H
v[s]=Y[s-1]-Z*u_predict[s-1]
u_update[s] = u_predict[s-1]+P_predict[s-1]*np.transpose(Z)*(1/F[s])*v[s]
u_predict[s] = T*u_update[s];
P_update[s] = P_predict[s-1]-P_predict[s-1]*np.transpose(Z)*(1/F[s])*Z*P_predict[s-1];
P_predict[s] = T*P_update[s]*np.transpose(T)+Q
KF_Dens[s] = (1/2)*np.log(2*np.pi)+(1/2)*np.log(abs(F[s]))+(1/2)*np.transpose(v[s])*(1/F[s])*v[s]
Likelihood = np.sum(KF_Dens[1:-1])
return Likelihood
def Kalman_Smoother(params, Y):
S = Y.shape[0]
S = S + 1
"Initialize Params:"
Z = params[0]
T = params[1]
H = params[2]
Q = params[3]
"Kalman Filter Starts:"
u_predict = np.zeros(S)
u_update = np.zeros(S)
P_predict = np.zeros(S)
P_update = np.zeros(S)
v = np.zeros(S)
F = np.zeros(S)
for s in range(1,S):
if s == 1:
P_update[s] = 1000
P_predict[s] = T*P_update[1]*np.transpose(T)+Q
else:
F[s] = Z*P_predict[s-1]*np.transpose(Z)+H
v[s]=Y[s-1]-Z*u_predict[s-1]
u_update[s] = u_predict[s-1]+P_predict[s-1]*np.transpose(Z)*(1/F[s])*v[s]
u_predict[s] = T*u_update[s];
P_update[s] = P_predict[s-1]-P_predict[s-1]*np.transpose(Z)*(1/F[s])*Z*P_predict[s-1];
P_predict[s] = T*P_update[s]*np.transpose(T)+Q
u_smooth = np.zeros(S)
P_smooth = np.zeros(S)
u_smooth[S-1] = u_update[S-1]
P_smooth[S-1] = P_update[S-1]
for t in range(S-1,0,-1):
u_smooth[t-1] = u_update[t] + P_update[t]*np.transpose(T)/P_predict[t]*(u_smooth[t]-T*u_update[t])
P_smooth[t-1] = P_update[t] + P_update[t]*np.transpose(T)/P_predict[t]*(P_smooth[t]-P_predict[t])/P_predict[t]*T*P_update[t]
u_smooth = u_smooth[0:-1]
return u_smooth
# + colab={"base_uri": "https://localhost:8080/", "height": 690} colab_type="code" id="M2Su4fyyC-Qw" outputId="797eadff-81e1-4625-b0b8-f80e0a5c2473"
T = 861
Y = openmean
param0 = np.array([0.5, 0.5, openmean.var()*100, 100*openmean.var()])
param_star = minimize(Kalman_Filter, param0, method='BFGS', options={'xtol': 1e-8, 'disp': True})
Y_update = Kalman_Smoother(param_star.x, Y)
timevec = np.linspace(1,T,T)
plt.plot(timevec[3:-1], Y_update[3:-1],'r',timevec, Y,'b:')
# + colab={} colab_type="code" id="PvZG5VvpC-Qz"
def rmseCalc(Y,Y_hat):
rmse = np.sqrt(np.mean((Y_update-Y)**2))
print(rmse)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="2jXKnl40C-Q2" outputId="ecd0d031-c1b7-4c2b-8ed3-440f33fbd3fc"
rmseCalc(Y,Y_update)
# + [markdown] colab_type="text" id="-1GDpu2iC-Q5"
# ### GARCH
# + colab={} colab_type="code" id="4JnOIOhXC-Q6"
T = 861;
mu = openmean.mean();
sig = openmean.var();
Y =openmean
# + colab={} colab_type="code" id="nwz9O_itC-Q_"
def GARCH(Y):
mu = param0[0]
omega = np.exp(param0[1])
alpha = (1-np.exp(-param0[2])) / (1+np.exp(-param0[2]))
beta = param0[3]
T = Y.shape[0]
GARCH_Dens = np.zeros(T)
sigma2 = np.zeros(T)
F = np.zeros(T)
v = np.zeros(T)
for t in range(1,T):
sigma2[t] = omega+alpha*((Y[t-1]-mu)**2)+beta*(sigma2[t-1]);
F[t] = Y[t] - mu-np.sqrt(sigma2[t])*np.random.normal(0,1,1);
v[t] = sigma2[t];
GARCH_Dens[t] = (1/2)*np.log(2*np.pi)+(1/2)*np.log(v[t])+(1/2)*(F[t]/v[t])
Likelihood = np.sum(GARCH_Dens[1:-1])
return Likelihood
# + colab={} colab_type="code" id="aiOIO9HkC-RC"
def GARCH_PROD(params, Y0, T):
mu = params[0]
omega = np.exp(params[1])
alpha = (1-np.exp(-params[2])) / (1+np.exp(-params[2]))
beta = params[3]
Y = np.zeros(T)
sigma2 = np.zeros(T)
Y[0] = Y0
sigma2[0] = omega/(1-alpha)
for t in range (1,T):
sigma2[t] = omega+alpha*((Y[t-1]-mu)**2)+beta*(sigma2[t-1]);
Y[t] = mu-np.sqrt(sigma2[t])*np.random.normal(0,1,1);
return Y
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="Lk-MlfpvC-RE" outputId="4e907bac-46bb-4ba5-fb94-d1a33005c378"
param_star = minimize(GARCH, param0, method = 'BFGS',options = {'xtol':1e-8,'disp':True})
#param_star = minimize(Kalman_Filter, param0, method='BFGS', options={'xtol':1e-8, 'disp': True})
# + colab={"base_uri": "https://localhost:8080/", "height": 690} colab_type="code" id="8J83UOg6C-RI" outputId="204ac374-c5e7-40fa-b619-f4fcd462ea38"
param0 = np.array([mu,2.5,0.3,0.5])
param_star = minimize(GARCH, param0, method = 'BFGS',options = {'xtol':1e-8,'disp':True})
Y_GARCH = GARCH_PROD(param_star.x,Y[0],T)
#Y_GARCH = GARCH_PROD(param_star.x,Y[0],T)
timevec = np.linspace(1,T,T)
plt.plot(timevec, Y ,'b', timevec[5:-1], Y_GARCH[5:-1],'r:')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="r_dPGVHwC-RK" outputId="c2ad17e6-edb8-4bde-a3e1-333cde531afd"
rmse = np.sqrt(np.mean((Y_GARCH-Y)**2))
print(rmse)
# + [markdown] colab_type="text" id="NpfxSmbOC-RN"
# ### 2 weeks data fitting for Garch and Garch-T
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="zZ3uCmmnC-RN" outputId="7ef7a224-806b-4b29-8d15-72cd73088446"
fb=facebook[350:379]
fb.head()
fb.shape
openmean1= (fb['Open']+ fb['Close'])/2
#openmean
def GARCH(Y):
mu = param0[0]
omega = np.exp(param0[1])
alpha = (1-np.exp(-param0[2])) / (1+np.exp(-param0[2]))
beta = param0[3]
T = Y.shape[0]
GARCH_Dens = np.zeros(T)
sigma2 = np.zeros(T)
F = np.zeros(T)
v = np.zeros(T)
for t in range(1,T):
sigma2[t] = omega+alpha*((Y[t-1]-mu)**2)+beta*(sigma2[t-1]);
F[t] = Y[t] - mu-np.sqrt(sigma2[t])*np.random.normal(0,1,1);
v[t] = sigma2[t];
GARCH_Dens[t] = (1/2)*np.log(2*np.pi)+(1/2)*np.log(v[t])+(1/2)*(F[t]/v[t])
Likelihood = np.sum(GARCH_Dens[1:-1])
return Likelihood
def GARCH_PROD(params, Y0, T):
mu = params[0]
omega = np.exp(params[1])
alpha = (1-np.exp(-params[2])) / (1+np.exp(-params[2]))
beta = params[3]
Y = np.zeros(T)
sigma2 = np.zeros(T)
Y[0] = Y0
sigma2[0] = omega/(1-alpha)
for t in range (1,T):
sigma2[t] = omega+alpha*((Y[t-1]-mu)**2)+beta*(sigma2[t-1]);
Y[t] = mu-np.sqrt(sigma2[t])*np.random.normal(0,1,1);
return Y
param_star = minimize(GARCH, param0, method = 'BFGS',options = {'xtol':1e-8,'disp':True})
T = 29;
mu = openmean1.mean();
sig = openmean1.var();
Y=openmean1
param0 = np.array([mu,2.5,0.3,0.5])
param_star = minimize(GARCH, param0, method = 'BFGS',options = {'xtol':1e-8,'disp':True})
Y_GARCH = GARCH_PROD(param_star.x,Y[350],T)
#Y_GARCH = GARCH_PROD(param_star.x,Y[0],T)
timevec = np.linspace(1,T,T)
plt.plot(timevec, Y ,'b', timevec[5:-1], Y_GARCH[5:-1],'r:')
rmse = np.sqrt(np.mean((Y_GARCH-Y)**2))
print(rmse)## 2 week data fitting for Garch and Garch-T
fb=facebook[350:379]
fb.head()
fb.shape
openmean1= (fb['Open']+ fb['Close'])/2
#openmean
def GARCH(Y):
mu = param0[0]
omega = np.exp(param0[1])
alpha = (1-np.exp(-param0[2])) / (1+np.exp(-param0[2]))
beta = param0[3]
T = Y.shape[0]
GARCH_Dens = np.zeros(T)
sigma2 = np.zeros(T)
F = np.zeros(T)
v = np.zeros(T)
for t in range(1,T):
sigma2[t] = omega+alpha*((Y[t-1]-mu)**2)+beta*(sigma2[t-1]);
F[t] = Y[t] - mu-np.sqrt(sigma2[t])*np.random.normal(0,1,1);
v[t] = sigma2[t];
GARCH_Dens[t] = (1/2)*np.log(2*np.pi)+(1/2)*np.log(v[t])+(1/2)*(F[t]/v[t])
Likelihood = np.sum(GARCH_Dens[1:-1])
return Likelihood
def GARCH_PROD(params, Y0, T):
mu = params[0]
omega = np.exp(params[1])
alpha = (1-np.exp(-params[2])) / (1+np.exp(-params[2]))
beta = params[3]
Y = np.zeros(T)
sigma2 = np.zeros(T)
Y[0] = Y0
sigma2[0] = omega/(1-alpha)
for t in range (1,T):
sigma2[t] = omega+alpha*((Y[t-1]-mu)**2)+beta*(sigma2[t-1]);
Y[t] = mu-np.sqrt(sigma2[t])*np.random.normal(0,1,1);
return Y
param_star = minimize(GARCH, param0, method = 'BFGS',options = {'xtol':1e-8,'disp':True})
T = 29;
mu = openmean1.mean();
sig = openmean1.var();
Y=openmean1
param0 = np.array([mu,2.5,0.3,0.5])
param_star = minimize(GARCH, param0, method = 'BFGS',options = {'xtol':1e-8,'disp':True})
Y_GARCH = GARCH_PROD(param_star.x,Y[350],T)
#Y_GARCH = GARCH_PROD(param_star.x,Y[0],T)
timevec = np.linspace(1,T,T)
plt.plot(timevec, Y ,'b', timevec[5:-1], Y_GARCH[5:-1],'r:')
rmse = np.sqrt(np.mean((Y_GARCH-Y)**2))
print(rmse)
# + colab={} colab_type="code" id="cjzuHrdBC-RQ"
df = facebook[['Open']]
df.reset_index(level=0, inplace=True)
df.columns=['ds','y']
# + [markdown] colab_type="text" id="30pAhUWnC-RS"
# ### Simple Moving Average
# + colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="Am2mOBfUC-RS" outputId="9cf13d38-57dc-49e5-c6d3-caab0d323fef"
rolAvg = df.y.rolling(window=5).mean()
rolAvg2 = df.y.rolling(window=25).mean()
plt.plot(df.ds, df.y, label='FB')
plt.plot(df.ds, rolAvg, label='FB 5 Day', color='orange')
plt.plot(df.ds, rolAvg2, label='FB 25 Day', color='magenta')
plt.legend(loc='upper left')
plt.show()
# + [markdown] colab_type="text" id="TkmqKT9QC-RU"
# ### Exponential Moving Average
# + colab={"base_uri": "https://localhost:8080/", "height": 267} colab_type="code" id="wWr23-TRC-RV" outputId="43d60fed-3e4b-429f-c00e-f0abef8ab2cc"
movAvg = df.y.ewm(span=5, adjust=False).mean()
movAvg2 = df.y.ewm(span=25, adjust=False).mean()
plt.plot(df.ds, df.y, label='FB')
plt.plot(df.ds, movAvg, label='FB 5 Day')
plt.plot(df.ds, movAvg2, label='FB 25 Day')
plt.legend(loc='upper left')
plt.show()
# + [markdown] colab_type="text" id="92qsdChQC-RX"
# We can see that using this signal we could have predicted the price trend of FB. When short-term crosses above long-term we get a buy signal. When short-term passes below the longer-term we get a sell signal.
# + [markdown] colab_type="text" id="f9c76H3IC-RY"
# ### Linear Regression of Simple Moving Average
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="1p2rKK-eC-RY" outputId="d9828c09-900d-4d44-af77-8fb15ff97854"
movAverage= rolAvg
movAverage.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="xAUJTV6QC-Ra" outputId="610ead1a-09c0-4255-9387-287ba505098d"
# Sample for X,Y
T = 861
#mu = (facebook['Open '].mean(),facebook['Open '].mean(),facebook['Open '].mean())
#cov = [[1,0.75,-0.35],[0.75,1,0.9],[-0.35,0.9,1]]
#F = np.random.multivariate_normal(mu,cov,T)
#Sample for Y,X
X = movAvg
X = np.column_stack([np.ones((T,1)),X])
#X=pd.DataFrame(movAverage[4:])
#print(X)
N = X.shape
#beta = np.array([0.56,2.53,2.05,1.78])
#beta.shape=(N[1],1)
#Y =X@beta+np.random.normal(0,1,(T,1))
Y=facebook['Open'].values
print(Y.shape)
invXX = np.linalg.inv(X.transpose()@X)
beta_hat = invXX@X.transpose()@Y
y_hat = X@beta_hat
residuals = Y-y_hat
sigma2 = (1/T)*residuals.transpose()@residuals
sigma = np.sqrt(sigma2)
#variance - covariance of beta_hat
varcov_beta_hat = (sigma2)*invXX
std_beta_hat = np.sqrt(T*np.diag(varcov_beta_hat))
R_square = 1-(residuals.transpose()@residuals)/(T*np.var(Y))
adj_R_square = 1-(1-R_square)*(T-1)/(T - N[1])
#Testing Coefficents:beta_i
#Null Hypotesis
t_stat = (beta_hat.transpose()-0)/std_beta_hat
p_val = 1-ss.norm.cdf(t_stat)
#Test of joint significance
F_stat= (beta_hat.transpose()@np.linalg.inv(varcov_beta_hat)@beta_hat/N[1])/(residuals.transpose()@residuals/(T-N[1]))
p_val_F= 1 - ss.f.cdf(F_stat,N[1]-1, T-N[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="IFRcBQ-0C-Rc" outputId="bd60de4e-f13c-4eec-ec1b-ee6684851d4a"
plt.plot(facebook['Date'],y_hat,color='magenta')
plt.plot(facebook['Date'],Y, color='black')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="mFd2lHyZC-Re" outputId="d750b9fd-c5be-4f27-aa2c-d39b9238f103"
rmse = np.sqrt(np.mean((y_hat-Y)**2))
print(rmse)
# + [markdown] colab_type="text" id="rAaR9qW_C-Rh"
# ### Linear Regression of Exponential Moving Average
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="fR2fWANvC-Rh" outputId="09c0159d-01d1-4581-cc2e-21ef5d1b490b"
movAverage1= (movAvg)
movAverage1.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="1kiy9bMhC-Rk" outputId="13aaec46-da93-421f-d0ef-0cef79f23c61"
# Sample for X,Y
T = 861
#mu = (facebook['Open '].mean(),facebook['Open '].mean(),facebook['Open '].mean())
#cov = [[1,0.75,-0.35],[0.75,1,0.9],[-0.35,0.9,1]]
#F = np.random.multivariate_normal(mu,cov,T)
#Sample for Y,X
X=movAverage1
X = np.column_stack([np.ones((T,1)),X])
N = X.shape
#beta = np.array([0.56,2.53,2.05,1.78])
#beta.shape=(N[1],1)
#Y =X@beta+np.random.normal(0,1,(T,1))
Y=facebook['Open'].values
print(X)
invXX = np.linalg.inv(X.transpose()@X)
beta_hat = invXX@X.transpose()@Y
y_hat = X@beta_hat
residuals = Y-y_hat
sigma2 = (1/T)*residuals.transpose()@residuals
sigma = np.sqrt(sigma2)
#variance - covariance of beta_hat
varcov_beta_hat = (sigma2)*invXX
std_beta_hat = np.sqrt(T*np.diag(varcov_beta_hat))
R_square = 1-(residuals.transpose()@residuals)/(T*np.var(Y))
adj_R_square = 1-(1-R_square)*(T-1)/(T - N[1])
#Testing Coefficents:beta_i
#Null Hypotesis
t_stat = (beta_hat.transpose()-0)/std_beta_hat
p_val = 1-ss.norm.cdf(t_stat)
#Test of joint significance
F_stat= (beta_hat.transpose()@np.linalg.inv(varcov_beta_hat)@beta_hat/N[1])/(residuals.transpose()@residuals/(T-N[1]))
p_val_F= 1 - ss.f.cdf(F_stat,N[1]-1, T-N[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="CqxlnChdC-Rn" outputId="09aa2a63-d311-4f0b-ef3e-63a45b2c9bfc"
plt.plot(facebook['Date'],y_hat,color='orange')
plt.plot(facebook['Date'],Y, color='black')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="oG-p53w3C-Rp" outputId="85636eff-1498-4e3d-b4c2-389e69494a99"
rmse = np.sqrt(np.mean((y_hat-Y)**2))
print(rmse)
# + [markdown] colab_type="text" id="WCMMKbhYC-Rr"
# ## Autoregression
# + colab={} colab_type="code" id="jUXw_MJAC-Rr"
facebook['y_hat'] = y_hat
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="W9--eJSRC-Rt" outputId="208de171-c455-4986-e3ec-3d1dac3de6b8"
facebook.head()
# + colab={} colab_type="code" id="gtshkNJ_C-Rz"
#facebook = facebook.set_index(pd.DatetimeIndex(facebook['Date']))
# + colab={} colab_type="code" id="VvodHZHJC-R1"
#facebook.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="ZYqR36HgC-R4" outputId="0de57a2d-c18a-451d-c7df-ef568e7a689c"
from pandas import Series
from pandas import DataFrame
from pandas import concat
from matplotlib import pyplot
values = DataFrame(facebook['y_hat'].values)
dataframe = concat([values.shift(1), values], axis=1)
dataframe.columns = ['t-1', 't']
result = dataframe.corr()
print(result)
# + colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="PSav_634C-R6" outputId="c1dfb3c9-6583-4336-b6e5-4bc4fcf921bf"
from pandas import Series
from matplotlib import pyplot
from statsmodels.tsa.ar_model import AR
# split dataset
X = facebook['y_hat']
#def AR(X):
train, test = X[1:len(X)-10], X[len(X)-10:]
# train autoregression
model = AR(train)
model_fit = model.fit()
#print('Lag: %s' % model_fit.k_ar)
#print('Coefficients: %s' % model_fit.params)
predictions = model_fit.predict(start=800, end=861, dynamic=False)
#print(len(predictions))
#rmseCalc()
#return predictions
# error = mean_squared_error(test, predictions)
# print('Test MSE: %.3f' % error)
# plot results
#X = facebook['y_hat']
#predictions = AR(X)
pyplot.plot(test)
pyplot.plot(predictions, color='red')
pyplot.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="KFd7wQQCC-R8" outputId="24d04632-3dcf-4549-f6f2-597a187c4661"
model_fit.predict(600,610)
# + [markdown] colab_type="text" id="cqaEvQhdC-R_"
# ## CAPM
# + colab={"base_uri": "https://localhost:8080/", "height": 576} colab_type="code" id="sq74WDunC-SA" outputId="d92635b2-a783-4cdf-86a2-229c753bc87e"
import pandas_datareader as pdr
from pandas_datareader import data, wb
from datetime import date
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
def capm(start_date, end_date, ticker1, ticker2):
df = pdr.get_data_yahoo(ticker1, start_date, end_date)
dfb = pdr.get_data_yahoo(ticker2, start_date, end_date)
rts = df.resample('M').last()
rbts = dfb.resample('M').last()
dfsm = pd.DataFrame({'s_open' : rts['Open'], 'b_open' : rbts['Open']}, index=rts.index)
dfsm[['s_returns', 'b_returns']] = dfsm[['s_open','b_open']]/dfsm[['s_open','b_open']].shift(1) -1
dfsm = dfsm.dropna()
covmat = np.cov(dfsm["s_returns"], dfsm["b_returns"])
beta = covmat[0,1]/covmat[1,1]
alpha = np.mean(dfsm["s_returns"])-beta*np.mean(dfsm["b_returns"])
y = beta * dfsm["b_returns"] + alpha
SS_res = np.sum(np.power(y - dfsm["s_returns"],2))
SS_tot = covmat[0,0]*(len(dfsm) - 1) # SS_tot is sample_variance*(n-1)
r_squared = 1.0 - SS_res/SS_tot
volatility = np.sqrt(covmat[0,0])
momentum = np.prod(1+dfsm["s_returns"].tail(12).values) - 1.0
prd = 12.0
alpha = alpha*prd
volatility = volatility*np.sqrt(prd)
print ("Beta, alpha, r_squared, volatility, momentum:")
print (beta, alpha, r_squared, volatility, momentum)
# %matplotlib inline
fig,ax = plt.subplots(1,figsize=(20,10))
ax.scatter(dfsm["b_returns"], dfsm['s_returns'], label="Data points")
beta,alpha = np.polyfit(dfsm["b_returns"], dfsm['s_returns'], deg=1)
ax.plot(dfsm["b_returns"], beta*dfsm["b_returns"] + alpha, color='red', label="CAPM line")
plt.title('Capital Asset Pricing Model, finding alphas and betas')
plt.xlabel('Market return $R_m$', fontsize=14)
plt.ylabel('Stock return $R_i$')
plt.legend()
plt.grid(True)
plt.show()
return beta, y
beta, y = capm('2016-01-04', '2019-08-30','FB', '^GSPC')
# + [markdown] colab_type="text" id="tq5oAKBZC-Sr"
# ### SVM
# + colab={} colab_type="code" id="xmbv0aVOC-Sr"
X = facebook[['Open']]
Y = facebook[['Open']].values
# + colab={} colab_type="code" id="LaZL2mxyC-St"
from sklearn.svm import SVR
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="Q2EMFP_PC-Sv" outputId="06359df0-a20d-4bab-c9dc-d14a522b9747"
svr_model = SVR(kernel='rbf', gamma=0.0005)
result = svr_model.fit(X, Y)
y_hat = result.predict(X)
# + colab={"base_uri": "https://localhost:8080/", "height": 284} colab_type="code" id="LTUr3zAJC-Sx" outputId="a95815a5-0c3c-4278-eee9-e91bc08dc475"
plt.plot(y_hat)
plt.plot(Y)
# + [markdown] colab={} colab_type="code" id="iQA9iKHOC-S3"
# ## Professionalism
# 50% Minghao Ru %50 Tong Yang
#
#
# ## Licensing
# Copyright <2020> <NAME>, <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# + colab={} colab_type="code" id="tnPscfjQC-S5"
# + colab={} colab_type="code" id="CkW60DoNC-S_"
| Project/Facebook_Portfolio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''.venv'': poetry)'
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append("../../")
import os
import yaml
import numpy as np
import textwrap
import json
import seaborn as sns
import matplotlib.pyplot as plt
plt.rcParams['pdf.fonttype'] = 42
import matplotlib.gridspec as gridspec
plt.rcParams["font.family"] = "Optima"
plt.rcParams["font.weight"] = "light"
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import scipy.stats
import arviz as az
from epimodel import preprocess_data, run_model, EpidemiologicalParameters
from epimodel.plotting.intervention_plots import combine_npi_samples
# -
B_REGIONS_ISO = ['AL', 'AD', 'AT', 'BE', 'BA', 'BG', 'HR',
'CZ', 'DK', 'EE', 'FR', 'DE', 'GR', 'HU', 'IE', 'IT',
'LV', 'LT', 'MT', 'NL', 'PL', 'PT', 'RO', 'RS', 'SK',
'SI', 'ES', 'CH', 'GB']
B_LAT = [41.32, 42.5, 48.2, 50.85, 43.87, 42.7, 45.82,
50.08, 55.67, 59.43, 48.85, 52.52, 37.97, 47.47, 53.33, 41.9,
56.93, 54.68, 35.88, 52.37, 52.23, 38.7, 44.42, 44.82, 48.13,
46.05, 40.38, 46.95, 51.5]
B_REGIONS = ['AL', 'AD', 'Austria', 'BE', 'BA', 'BG', 'HR', 'Czech Rep.', 'DK', 'EE', 'FR', 'Germany', 'GR', 'HU', 'IE', 'Italy', 'LV', 'LT', 'MT', 'Netherlands', 'PL', 'PT', 'RO', 'RS', 'SK', 'SI', 'ES', 'Switzerland', 'England']
S_REGIONS = ['Austria', 'Czech Rep.', 'England', 'Germany', 'Italy', 'Netherlands', 'Switzerland']
def load_json(path, vars=("seasonality_beta1", "seasonality_max_R_day")):
with open('../../'+fn) as f:
d = json.load(f)
d["MODEL"] = re.search('model(.*)_', d['model_config_name']).groups()[0]
d["DATA"] = re.search('data(.*)', d['model_config_name']).groups()[0]
d["LABEL"] = f"Seasonal {d['MODEL']} et al." #\n{d['DATA']} data" # NB: Change for 2x2 plots
if d['DATA'] == "BraunerTE":
d["LABEL"] += "\n(temperate Europe)"
print(f"Loaded {d['MODEL']} model, {d['DATA']} data. Rhat: {d['rhat']}")
cols = {v: np.array(d[v]) for v in vars}
cols["label"] = d["LABEL"]
return d, pd.DataFrame(cols)
ds = []
for fn in [
"sensitivity_final/modelBrauner_dataBraunerTE/seasonality_local/complex_seasonal_2021-06-27-214513_pid47284_summary.json",
"sensitivity_final/modelBrauner_dataBraunerTE/seasonality_local/complex_seasonal_2021-06-27-214413_pid46689_summary.json",
"sensitivity_final/modelBrauner_dataBraunerTE/seasonality_local/complex_seasonal_2021-06-27-214443_pid47122_summary.json",
"sensitivity_final/modelBrauner_dataBraunerTE/seasonality_local/complex_seasonal_2021-06-27-214543_pid47441_summary.json",
"sensitivity_final/modelBrauner_dataBraunerTE/seasonality_local/complex_seasonal_2021-06-27-214614_pid47588_summary.json",
"sensitivity_final/default_cmodelSharma_dataSharma/seasonality_local/20210628-002851-52446_summary.json",
"sensitivity_final/default_cmodelSharma_dataSharma/seasonality_local/20210628-002856-52455_summary.json",
"sensitivity_final/default_cmodelSharma_dataSharma/seasonality_local/20210628-002901-52575_summary.json",
"sensitivity_final/default_cmodelSharma_dataSharma/seasonality_local/20210628-002906-52694_summary.json",
"sensitivity_final/default_cmodelSharma_dataSharma/seasonality_local/20210628-002911-52834_summary.json",
]:
d, df0 = load_json(fn)
d["df"] = df0
d["Rs"] = {"BraunerTE": B_REGIONS, "Sharma": S_REGIONS}[d['DATA']]
d["fn"] = fn
ds.append(d)
# +
for d in ds:
local_beta1 = np.array(d["seasonality_local_beta1"])
dfs = []
for i, r in enumerate(d['Rs']):
dfs.append(pd.DataFrame({"Country": r, "Local gamma": local_beta1[:,i]}))
dfs.sort(key=lambda df: df["Local gamma"].mean())
dfs.append(pd.DataFrame({"Country": "Base\ngamma", "Local gamma": np.array(d["seasonality_beta1"])}))
df = pd.concat(dfs, axis=0, ignore_index=True)
#sns.kdeplot(data=df, x="local_beta1", hue="Country", multiple="stack")
if d['DATA'] == "Sharma":
plt.figure(figsize=(6,4))
else:
plt.figure(figsize=(6,8))
sns.boxplot(data=df, x="Local gamma", y="Country", fliersize=0)
local_sd = d['exp_config']['local_seasonality_sd']
plt.title(f"Local seasonal amplitudes, sd={local_sd:.2f}")
plt.xlim(-0.2, 0.8)
sns.despine()
plt.savefig(f'figures/Fig_seasonality_local_{d["DATA"]}_{local_sd:.2f}.pdf', bbox_inches='tight')
plt.close()
# +
bd = [d for d in ds if d['exp_config']['local_seasonality_sd'] == local_sd and d["DATA"] == "BraunerTE"][0]
dfs = []
for i, r in enumerate(bd['Rs']):
r2 = B_REGIONS_ISO[B_REGIONS.index(r)]
dfs.append(pd.DataFrame({"Country": r2, "Model": "Brauner",
"Local gamma": np.array(bd["seasonality_local_beta1"])[:, i]}))
df = pd.concat(dfs, axis=0, ignore_index=True)
df.groupby(["Country", "Model"]).median().to_csv("tmp.csv")
# +
SDs = sorted(set(d['exp_config']['local_seasonality_sd'] for d in ds))
pal = sns.color_palette()
for local_sd in SDs:
print(local_sd)
bd = [d for d in ds if d['exp_config']['local_seasonality_sd'] == local_sd and d["DATA"] == "BraunerTE"][0]
sd = [d for d in ds if d['exp_config']['local_seasonality_sd'] == local_sd and d["DATA"] == "Sharma"][0]
b_local_beta1 = np.array(bd["seasonality_local_beta1"])
s_local_beta1 = np.array(sd["seasonality_local_beta1"])
dfs = []
for i, r in enumerate(bd['Rs']):
dfs.append(pd.DataFrame({"Country": r, "Model": "Brauner", "Local gamma": b_local_beta1[:,i]}))
for i, r in enumerate(sd['Rs']):
dfs.append(pd.DataFrame({"Country": r, "Model": "Sharma", "Local gamma": s_local_beta1[:,i]}))
dfs.sort(key=lambda df: df["Local gamma"].mean())
dfs.append(pd.DataFrame(
{"Country": "Base\ngamma", "Model": "Brauner", "Local gamma": np.array(bd["seasonality_beta1"])}))
dfs.append(pd.DataFrame(
{"Country": "Base\ngamma", "Model": "Sharma", "Local gamma": np.array(sd["seasonality_beta1"])}))
df = pd.concat(dfs, axis=0, ignore_index=True)
plt.figure(figsize=(5,10))
Rs = list(reversed(df['Country'].unique()))
Rs[1:] = sorted(Rs[1:],
key=lambda r: df[df['Country'] == r]["Local gamma"].median(), reverse=True)
plt.yticks(range(len(Rs)), Rs)
plt.ylim(-0.5, len(Rs) -0.5)
for i, r in enumerate(Rs):
df2 = df[df['Country'] == r]
#print(df2)
#sns.kdeplot(data=df2, x="Local gamma", y=np.full(len(df2), i),hue="Model", multiple="stack")
bx = df2[df2['Model']=='Brauner']["Local gamma"].values
sx = df2[df2['Model']=='Sharma']["Local gamma"].values
x = np.concatenate([bx, sx])
x0, xA0, xB0, xM, xB1, xA1, x1 = np.quantile(x,
[0.025, 0.025, 0.25, 0.5, 0.75, 0.975, 0.975])
yoff = i - 0
xs = np.linspace(x0, x1, 500)
M = 0.15
bkde = scipy.stats.gaussian_kde(bx, 0.2)
bxs = bkde(xs) * M
if len(sx) > 0:
skde = scipy.stats.gaussian_kde(sx, 0.2)
sxs = skde(xs) * M / 2
#bxs = bxs / 2
else:
skde = lambda x: 0.0
sxs = skde(xs) * M
#plt.plot([x0, x1], [y, y], color=pal[y])
for axx in [0.0, 0.1, 0.2, 0.3, 0.4, 0.5]:
plt.axvline(x=axx, lw=0.2, zorder=-2, c="#777")
plt.fill_between(xs, yoff, yoff + bxs, color=pal[0], alpha=1)
plt.fill_between(xs, yoff + bxs, yoff + bxs + sxs, color=pal[1], alpha=1)
plt.plot(xs, yoff + bxs + sxs, color='k')
plt.scatter(xM, yoff, marker="+", color='k', s=60)
plt.plot([xA0, xA1], [yoff, yoff], color='k', lw=2, alpha=0.5)
plt.plot([xB0, xB1], [yoff, yoff], color='k', lw=3, alpha=1.0)
#g = sns.FacetGrid(df, row="Country", hue="Model", aspect=15, height=.5)#, palette=pal)
#g.map_dataframe(sns.kdeplot, "Local gamma",# multiple="stack",
#bw_adjust=.5, clip_on=False,
#fill=True, alpha=1, linewidth=1.5)
#sns.violinplot(data=df, y="Country", x="Local gamma", hue="Model", split=True)
#sns.kdeplot(data=df, x="local_beta1", hue="Country", multiple="stack")
#sns.boxplot(data=df, x="Local gamma", y="Country", fliersize=0)
#local_sd = d['exp_config']['local_seasonality_sd']
plt.title(f"Local seasonal amplitudes, sd={local_sd:.2f}")
plt.xlim(-0.2, 0.8)
sns.despine()
plt.savefig(f'figures/Fig_seasonality_local_kdes_{local_sd:.2f}.pdf', bbox_inches='tight')
plt.close()
plt.figure(figsize=(5,10))
Rs = df['Country'].unique()
Rs[:-1] = sorted(Rs[:-1],
key=lambda r: df[df['Country'] == r]["Local gamma"].median(), reverse=True)
#plt.yticks(range(len(Rs)), Rs)
#plt.ylim(-0.5, len(Rs) -0.5)
for i, r in enumerate(B_REGIONS):
df2 = df[df['Country'] == r]
bx = df2[df2['Model']=='Brauner']["Local gamma"].values
sx = df2[df2['Model']=='Sharma']["Local gamma"].values
x = np.concatenate([bx, sx])
x0, xA0, xB0, xM, xB1, xA1, x1 = np.quantile(x,
[0.025, 0.025, 0.25, 0.5, 0.75, 0.975, 0.975])
yoff = B_LAT[i]
xs = np.linspace(x0, x1, 500)
M = 0.15
bkde = scipy.stats.gaussian_kde(bx, 0.2)
bxs = bkde(xs) * M
if len(sx) > 0:
skde = scipy.stats.gaussian_kde(sx, 0.2)
sxs = skde(xs) * M / 2
bxs = bxs / 2
else:
skde = lambda x: 0.0
sxs = skde(xs) * M
#plt.plot([x0, x1], [y, y], color=pal[y])
for axx in [0.0, 0.1, 0.2, 0.3, 0.4, 0.5]:
plt.axvline(x=axx, lw=0.2, zorder=-2, c="#777")
#plt.fill_between(xs, yoff, yoff + bxs, color=pal[0], alpha=1)
#plt.fill_between(xs, yoff + bxs, yoff + bxs + sxs, color=pal[1], alpha=1)
#plt.plot(xs, yoff + bxs + sxs, color='k')
plt.scatter(xM, yoff, marker="+", color='k', s=60)
plt.plot([xA0, xA1], [yoff, yoff], color='k', lw=2, alpha=0.5)
plt.plot([xB0, xB1], [yoff, yoff], color='k', lw=3, alpha=1.0)
plt.title(f"Local seasonal amplitudes, sd={local_sd:.2f}")
plt.xlim(-0.2, 0.8)
sns.despine()
plt.savefig(f'figures/Fig_seasonality_local_latplot_{local_sd:.2f}.pdf', bbox_inches='tight')
plt.close()
# -
| notebooks/final_results/seasonality_result_local_plotter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Fno6-pp9h7J6" colab_type="code" outputId="d204f8d5-88b2-4b11-d7fb-56c4727016d0" colab={"base_uri": "https://localhost:8080/", "height": 204}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score, roc_auc_score, roc_curve
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import OrdinalEncoder
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import SimpleImputer
from sklearn.inspection import partial_dependence, PartialDependenceDisplay
from sklearn.model_selection import train_test_split, GridSearchCV, RandomizedSearchCV
# !pip install eli5
import eli5
from eli5.sklearn import PermutationImportance
from xgboost import XGBClassifier
import seaborn as sns
pd.set_option('float_format', '{:f}'.format)
# + id="7ADhAmUZh7KC" colab_type="code" outputId="e3e799cb-cd91-4ffb-85fe-4b5c7a32afd5" colab={"base_uri": "https://localhost:8080/", "height": 292}
names = ['age', 'workclass', 'fnlwgt', 'education', 'education_num',
'marital_status', 'occupation', 'relationship', 'race', 'sex',
'capital_gain', 'capital_loss', 'hours_per_week',
'native_country', 'salary'
]
df = pd.read_csv('/content/adult.data', names=names)
df.head()
# + id="tqz9ay_qh7KH" colab_type="code" outputId="d3a1d39c-6e68-4718-8026-91536b936353" colab={"base_uri": "https://localhost:8080/", "height": 54}
'''
Explanation of the column 'fnlwgt':
| Description of fnlwgt (final weight)
|
| The weights on the CPS files are controlled to independent estimates of the
| civilian noninstitutional population of the US. These are prepared monthly
| for us by Population Division here at the Census Bureau. We use 3 sets of
| controls.
| These are:
| 1. A single cell estimate of the population 16+ for each state.
| 2. Controls for Hispanic Origin by age and sex.
| 3. Controls by Race, age and sex.
|
| We use all three sets of controls in our weighting program and "rake" through
| them 6 times so that by the end we come back to all the controls we used.
|
| The term estimate refers to population totals derived from CPS by creating
| "weighted tallies" of any specified socio-economic characteristics of the
| population.
|
| People with similar demographic characteristics should have
| similar weights. There is one important caveat to remember
| about this statement. That is that since the CPS sample is
| actually a collection of 51 state samples, each with its own
| probability of selection, the statement only applies within
| state.
'''
# + id="JyjQHi6Fh7KL" colab_type="code" outputId="9d3608bc-d0b9-41ff-ad6d-0fb1dbe94773" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head() # Missing values are '?'
# + id="Qpu2aeQ_h7KP" colab_type="code" outputId="4f21d25e-70f9-42ab-80b9-99ac233e5fd1" colab={"base_uri": "https://localhost:8080/", "height": 34}
df['workclass'][27] # Question mark has a leading space
# + id="MZ-otQX4h7KT" colab_type="code" outputId="7264dcba-9b98-47a5-b16f-e05314e9c1cf" colab={"base_uri": "https://localhost:8080/", "height": 289}
df = df.replace({' ?': np.nan})
df.loc[27,:]
# + id="B4J_IaCch7KZ" colab_type="code" outputId="6c8a796f-50be-4a34-df61-476f7945573e" colab={"base_uri": "https://localhost:8080/", "height": 173}
df.describe(exclude='number')
# df.describe(exclude='number').columns
# + id="VlNF5wKkh7Kd" colab_type="code" colab={}
for col in df.describe(exclude='number').columns:
df[col].replace({np.nan: 'Unknown'}, inplace=True)
# + id="9cmOvm0qh7Kg" colab_type="code" outputId="f8176604-37a1-4ded-9ce5-9246cd6bea01" colab={"base_uri": "https://localhost:8080/", "height": 173}
df.describe(exclude='number')
# + id="9h1B86KCh7Km" colab_type="code" outputId="1996dd42-a23f-4d18-faf1-c7adf05f6f8d" colab={"base_uri": "https://localhost:8080/", "height": 297}
# From this data, we can see that there are columns containing values which are poorly
# distributed such as 'capital_gain', 'capital_loss' which, for the most part, are
# mostly 0s. I think we might be able to curtail 'age' and 'hours_per_week' as well.
df.describe(exclude=['category', 'object'])
# + id="r569Avxeh7Kr" colab_type="code" outputId="0c6b62b4-157d-4629-d138-960d5e632c13" colab={"base_uri": "https://localhost:8080/", "height": 297}
# For curtailing purposes, I would like to only see results from the United States
# since 29170 of the 32561 observations are for this location.
# The value United-States has a leading space
us_df = df[df['native_country']==' United-States']
us_df.describe()
# + id="mADt2YRvh7Ku" colab_type="code" outputId="e53f56e8-2082-4982-f95e-e1d0e410a997" colab={"base_uri": "https://localhost:8080/", "height": 34}
# For the purposes of being consistant, I would keep the hours_per_week limited to
# 40 hours in a work week (avg = 39.2h in 1995) source: https://www.bls.gov/opub/mlr/1997/04/art1full.pdf
# Since 75% of the data is from 1-45, I'd like to make the max just above that at 50h
# At the other side, the government, in 1996, defined part time as working less than 30-35h
# per work week so, I will have the lower end be 30h
hour_mask = (us_df['hours_per_week']>=30) & (us_df['hours_per_week']<=50)
len(us_df['hours_per_week'][(us_df['hours_per_week']<30) & (us_df['hours_per_week']>50)])
# + id="WmynSkhQh7Ky" colab_type="code" colab={}
hour_df = us_df[hour_mask]
# + id="9Y1hdNnhh7K1" colab_type="code" outputId="2dd855a0-27c8-4047-8c4f-acaedefedb3f" colab={"base_uri": "https://localhost:8080/", "height": 34}
# I chose 62 since, back in the early 90s, the early retirement age was 61 and, today
# it is now 62 (full 65-67).
age_mask = hour_df['age']<=62
print(len(hour_df['age'][hour_df['age']>62]))
# + id="lBTF1Zu4h7K4" colab_type="code" colab={}
age_df = hour_df[age_mask]
# + id="q_TkB7sgh7K-" colab_type="code" outputId="aa494ac5-1467-4b58-e25e-542ddeb9ca14" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Next I will drop trivial columns and columns with many 0s
df = age_df.drop(columns=['fnlwgt', 'capital_gain', 'capital_loss', 'native_country'])
df.shape
# + id="UIknzstNh7LD" colab_type="code" outputId="b9be37ab-7db2-456b-9083-109616bde015" colab={"base_uri": "https://localhost:8080/", "height": 297}
df.describe()
# + id="plk8XLI-h7LG" colab_type="code" outputId="f262a72e-f7c9-4297-b084-4607c69af219" colab={"base_uri": "https://localhost:8080/", "height": 173}
df.describe(exclude='number')
# + id="_bAzbWYAh7LJ" colab_type="code" colab={}
# This distribution tapers off at 500000 so I will have that be the limit.
# sns.distplot(df['fnlwgt']);
# + id="oAVWZcg-h7LN" colab_type="code" colab={}
df['salary_over_50k'] = df['salary'].replace({' <=50K': 0, ' >50K': 1})
# + id="yiJa9dSxh7LQ" colab_type="code" outputId="e3fb1159-be79-48ff-8bc7-4c06d1bcc269" colab={"base_uri": "https://localhost:8080/", "height": 68}
df['salary_over_50k'].value_counts(normalize=True)
# + id="Zyn6L_Suh7LV" colab_type="code" outputId="28310938-bd75-40f7-cae5-8dae07ddbfb2" colab={"base_uri": "https://localhost:8080/", "height": 204}
df = df.drop(columns='salary')
df.head()
# + id="2I1r_iREh7LZ" colab_type="code" outputId="fac81809-90ca-4e73-ae98-283a688ea614" colab={"base_uri": "https://localhost:8080/", "height": 34}
train, test = train_test_split(df, test_size=0.15, random_state=42)
train, val = train_test_split(train, test_size=0.20, random_state=42)
print(train.shape, val.shape, test.shape)
# + id="puHdjupgh7Ld" colab_type="code" colab={}
target = 'salary_over_50k'
features = train.drop(columns=target).columns
X_train = train[features]
X_val = val[features]
X_test = test[features]
y_train = train[target]
y_val = val[target]
y_test = test[target]
# + id="MG8lzXMTh7Lg" colab_type="code" outputId="ec36a631-5f76-46cc-91c0-eb41e48ac4cc" colab={"base_uri": "https://localhost:8080/", "height": 306}
pipeline = make_pipeline(
OrdinalEncoder(),
RandomForestClassifier(n_estimators=100, random_state=42)
)
pipeline.fit(X_train, y_train)
# + id="IDHbgF10h7Lk" colab_type="code" outputId="921b1a63-d1ee-427c-d8f4-7f30c4879e7d" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_pred_proba = pipeline.predict_proba(X_val)[:, -1]
roc_auc_score(y_val, y_pred_proba)
# y_pred_proba
# + id="erujt6TMh7Lp" colab_type="code" outputId="839445e5-9e79-4d3b-af2a-21969344b52f" colab={"base_uri": "https://localhost:8080/", "height": 1000}
fpr, tpr, thresholds = roc_curve(y_val, y_pred_proba)
(fpr, tpr, thresholds)
# + id="IEo7IGFnh7Ls" colab_type="code" outputId="f144bfae-a5f9-45a6-f8f5-6062e0db68a4" colab={"base_uri": "https://localhost:8080/", "height": 312}
plt.scatter(fpr, tpr)
plt.plot(fpr, tpr)
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
# + id="5FuYgwD8h7Lv" colab_type="code" outputId="c967ea67-4f58-46a8-d1df-c1a71537917a" colab={"base_uri": "https://localhost:8080/", "height": 276}
from sklearn.metrics import plot_confusion_matrix
y_pred = pipeline.predict(X_test)
plot_confusion_matrix(pipeline, X_test, y_pred, values_format='.0f', xticks_rotation='vertical');
# + id="7_K938i9h7Ly" colab_type="code" outputId="71174a5c-3f77-4373-e2d1-f3e34542060c" colab={"base_uri": "https://localhost:8080/", "height": 153}
transformers = make_pipeline(
OrdinalEncoder()
)
X_train_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.transform(X_val)
model = RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
model.fit(X_train_transformed, y_train)
# + id="sKQXZzPFh7L2" colab_type="code" outputId="593f0428-5ff3-4d11-be42-4fedf801c833" colab={"base_uri": "https://localhost:8080/", "height": 374}
permuter = PermutationImportance(
model,
scoring='roc_auc',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed, y_val)
# + id="af_zVXlih7L5" colab_type="code" outputId="31e0223b-ec34-4ceb-c16a-3b32b2c22b11" colab={"base_uri": "https://localhost:8080/", "height": 204}
feature_names = X_val.columns.tolist()
pd.Series(permuter.feature_importances_, feature_names).sort_values()
# + id="MCnkHAGkh7L9" colab_type="code" outputId="92f32385-d351-46e0-bb07-704ba758b8b8" colab={"base_uri": "https://localhost:8080/", "height": 204}
eli5.show_weights(
permuter,
top=None,
feature_names=feature_names
)
# + id="BhK50UPRh7MB" colab_type="code" colab={}
minimum_importance = 0
mask = permuter.feature_importances_ > minimum_importance
features = X_train.columns[mask]
X_train = X_train[features]
# + id="jUvV10SSh7ME" colab_type="code" outputId="19bbb2a6-0cdd-4d3f-9043-c631dda07a3d" colab={"base_uri": "https://localhost:8080/", "height": 309}
# !pip install category_encoders
import category_encoders as ce
X_val = X_val[features]
pipeline = make_pipeline(
ce.OrdinalEncoder(),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
# Fit on train, score on val
pipeline.fit(X_train, y_train)
print('Validation Accuracy', pipeline.score(X_val, y_val))
# + id="8osmFD4Zh7MI" colab_type="code" outputId="2ce6d455-c5fb-472a-b75f-12abb3bcf1f3" colab={"base_uri": "https://localhost:8080/", "height": 68}
features
# + id="1PGg3cA9h7MK" colab_type="code" outputId="1c2f6973-c524-4efe-a4dc-a11fad292104" colab={"base_uri": "https://localhost:8080/", "height": 445}
plt.rcParams['figure.dpi'] = 72
# !pip install pdpbox
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'hours_per_week'
isolated = pdp_isolate(
model=pipeline,
dataset=X_val,
model_features=X_val.columns,
num_grid_points=50,
feature=feature
)
# + id="xTx9YVaOh7MP" colab_type="code" outputId="8c6efa0f-8122-42d0-d8f6-c86e129ef895" colab={"base_uri": "https://localhost:8080/", "height": 595}
pdp_plot(isolated, feature_name=feature);
# + id="YBoq6bWGh7MS" colab_type="code" colab={}
feature = 'age'
isolated = pdp_isolate(
model=pipeline,
dataset=X_val,
model_features=X_val.columns,
num_grid_points=50,
feature=feature
)
# + id="XP9H7EySh7MV" colab_type="code" outputId="145572f8-150a-4b28-8a29-64061f628022" colab={"base_uri": "https://localhost:8080/", "height": 578}
pdp_plot(isolated, feature_name=feature);
# + id="cmurS3Ych7MZ" colab_type="code" outputId="57f08ca5-0a88-4a8a-a14b-613f2ec87819" colab={"base_uri": "https://localhost:8080/", "height": 598}
from pdpbox.pdp import pdp_interact, pdp_interact_plot
features = ['age', 'hours_per_week']
interaction = pdp_interact(
model=pipeline,
dataset=X_val,
model_features=X_val.columns,
features=features
)
# pdp_interact_plot(interaction, plot_type='grid', feature_names=features);
pdp_interact_plot(interaction, plot_type='grid', feature_names=features);
# + id="_O-7OjKo7eJE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="2c5f7028-908f-4a44-a48d-e7c67b0a2b66"
# !pip install shap
import shap
# + id="e0frcL248Isx" colab_type="code" colab={}
| practice_unit-2-sprint_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
import os
import trako as TKO
# %load_ext autoreload
# %autoreload 2
# -
DATADIR = '/home/d/DATA/Dropbox/ADHD_STFC_diff_for_Daniel/'
SUBJECTS = ['101','201']
GROUP = ['tracts_left_hemisphere']
files = os.listdir(os.path.join(DATADIR,SUBJECTS[0],GROUP[0]))
files[0]
vtp_file = os.path.join(DATADIR,SUBJECTS[0],GROUP[0],files[0])
gltf_w_tko = TKO.Encoder.fromVtp(vtp_file)
tko_file = '/tmp/test.tko'
gltf_w_tko.save(tko_file)
gltf = TKO.Encoder.fromVtp(vtp_file, draco=False)
gltf_file = '/tmp/test.gltf'
gltf.save(gltf_file)
# +
import plotly
import plotly.graph_objs as go
x = ['VTP','GLTF','TKO']
y = [os.path.getsize(v) for v in [vtp_file, gltf_file, tko_file]]
fig = go.Figure(data=[go.Bar(
x=x, y=y,
text=y,
textposition='auto',
)])
fig.show()
# -
| IPY/TKO_ADHD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ### Data Mining and Machine Learning
# ### <NAME>
# #### September 2021
# ## Data preparation I: Reading the data file and counting the missing values
# #### Dataset: Adult also known as Census, available at the UCI
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
# ### Reading the data from the UCI
# +
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data',
header=None, sep=',',na_values=[' ?'])
df.columns=['v1', 'v2', 'v3', 'v4', 'v5','v6','v7','v8','v9','v10','v11','v12','v13','v14','class']
# -
# ### Finding information about the features of the dataset.
df.info()
#Reading the first five rows of the dataset
df.head()
# + active=""
# #Another way to read the data
# # Load CSV using Pandas from URL
# url = "https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data"
# names = ['v1', 'v2', 'v3', 'v4', 'v5', 'v6', 'v7', 'v8','v9','v10','v11','v12','v13','v14','class']
# data = pd.read_csv(url, names=names,na_values=[' ?'])
# print(data.shape) #imprimiendo la dimension de los datos
# -
# ## Reading the data in csv format
#df = pd.read_csv('c://PW-PR/census.csv',sep=',',na_values=[' ?'])
df=pd.read_csv('http://academic.uprm.edu/eacuna/census.csv',sep=",",na_values=[' ?'])
#information about the dataset allowing us to notice that there are missing values
df.info()
# Notice that columns: v2(employment), v7(job) y v14 (native.country) have a different number of entries than the other columns because they have missing values.
#Printing the first 20 rows
df.head(20)
#printing the last five rows
df.tail()
#Printing the dimension of the dataset
print(df.shape)
# The dataset has 32561 instances amd 15 attributes
# #### Finding some statistics of the continuous features
df.describe()
# #### Finding some statistics of the nominal features
df.describe(include=['O'])
# ## Filtering:
# #### Finding the percentage of subjects for education level
tabla=pd.crosstab(index=df['education'],columns='counts')
tabla['frec.relativa.porc']=tabla*100/tabla.sum()
tabla
# #### Finding the frequency of subjetcs by gender and salary level
df.groupby(by =['salary', 'gender']).size()
# #### Extracting from the dataset a subset including subjects with age less than 30 years
df1=df[df['age']<30]
df2=df[df.age<30]
df2.shape
df3=df.query('age<30')
df3.shape
# #### Extracting from the dataset a subset including subjects with age greater than 80 years
df4=df.query('age>80')
df4.shape
# #### Drawing from the dataset a subset including subjects working more than 60 hours a week
df5=df[df['hours.per.week']>60]
df5.shape
# #### Drawing from the dataset a subset including subjects older than 80 years and working more than 60 hours a week
df6=df[(df['age']>80) & (df['hours.per.week']>60)]
df6.info()
# #### The last case is very unsual. The subject is a 90 years man working 99 hours per week as a trucker
# #### Finding a crosstabulation table relating gender and salary
p_table = pd.pivot_table(df,index='salary',columns='gender',aggfunc='size')
p_table
# ## Visualization:
#Clustered bars using matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
p_table.plot.bar(stacked=False)
#Stacked bars using matplotlib
p_table.plot.bar(stacked=True)
#Building a stacked bars from the raw data
p_table1 = pd.pivot_table(df,index='education',columns='salary',aggfunc='size')
plt.figure()
p_table1.plot.bar(stacked=True, width=0.9)
# ## Handling Missing Values
#Checking the existence of missing values
df.isnull()
# Finding the number of columns with missings
df.isnull().any().sum()
# Finding the columns containing missing values
colmiss=df.columns[df.isnull().any()].tolist()
print(colmiss)
# Finding the number of missings per column
df[colmiss].isnull().sum()
# Finding the percentage of missings per column
df[colmiss].isnull().sum()*100/len(df)
#Another way to find the number of missing values in each column of the dataframe
df.apply(lambda x: sum(x.isnull().values), axis = 0)
# Finding rows containing missing values
rowmiss=df.index[df.isnull().T.any()].tolist()
# Finding the number of rows containing missing values
df.isnull().T.any().sum()
# Finding the percentage of rows with missing values
df.isnull().T.any().sum()*100/len(df)
# Second method:
sum(df.apply(lambda x: sum(x.isnull().values), axis = 1)>0)
# Third method:
sum([True for idx,row in df.iterrows() if any(row.isnull())])
# Finding the total number of cells containing missing values
df.isnull().sum().sum()
# ## Handling of missing values
# Eliminating all the rows containing at least one missing value
dfclean=df.dropna()
dfclean.info()
# ### Removing columns having at least 5 percent of missing values
from math import floor
bound=floor(len(df)*.95)
dfclean=df.dropna(thresh=bound,axis='columns')
print(dfclean.shape)
dfclean.head()
# Two columns were eliminated
#
# Retaining only the rows with at least two missing values
dfclean=df.dropna(thresh=13,axis='rows')
print(dfclean.shape)
# Size of table after eliminating records withn misssing values
dfclean.info()
# Eliminating sequentially columns with at least 5% de missing values and then rows with at least two missings
dfclean1=df.dropna(thresh=bound,axis='columns').dropna(thresh=13,axis='rows')
dfclean1.info()
print(dfclean1.shape)
| notebooks/dprep12021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# default_exp dwpc
# -
# # DWPC
#
# > DWPC calculation
#hide
from nbdev.showdoc import *
from neo4jtools import SimpleNeo4jHTTPAPIClient
# +
#export
def make_dwpc_query(genes, reltype, hops, dwpc_score_prop_name='dwpc_score', only_relations_with_pmid=False):
'''
Make a cypher query to calculate DWPC
input parameters
genes : gene list
reltype : edge type
hops : number of hops to calculate
dwpc_score_prop_name : name of dwpc score property, default dwpc_score
only_relations_with_pmid : include only relations with pubmed_id
output
query:string
'''
qry_template="""MATCH p=(n0:Gene)-[:{reltype}*..{hops}]->(n1:Gene)
WHERE n0.name in {genestr}
AND n1.name in {genestr}
AND n0 <> n1
{conditionstr}
WITH
n0.name as source_name
, n1.name as target_name
, p as path
, [r in relationships(p)|r.{dwpc_score_prop_name}] as dwpc_scores
, [r in relationships(p) | r.pubmed_id] as pubmed_ids
WITH
source_name
, target_name
, length(path) as len
, reduce(acc=1, d in dwpc_scores|acc*d) as dwpc_score
, [n IN nodes(path) | labels(n)[0]] as node_labels
, [n IN nodes(path) | n.identifier] as node_identifiers
, [r IN relationships(path) | type(r)] as rel_types
, reduce(acc='', s in pubmed_ids | acc+';'+s) as pubmed_ids
WITH
source_name
, target_name
, dwpc_score as dwpc
, [r IN rel_types | split(r, '_')[1]] as rtypes
, "(:"+node_labels[0]+"{{identifier:'"+ node_identifiers[0] +"'}})" as head
, [i IN range(0,len-1)| "-[:"+ rel_types[i] +"]->(:"+node_labels[i+1]+"{{identifier:'"+ node_identifiers[i+1] +"'}})" ] as tail
, substring(pubmed_ids,1) as pubmed_ids
RETURN
source_name
, target_name
, dwpc
, head + reduce(acc='', s in tail |acc+s) as pattern
, reduce(acc='', s IN rtypes | acc + s) as type_pattern
, pubmed_ids
"""
only_if_pubmed_id_exists="AND all(r IN relationships(p) WHERE exists(r.pubmed_id)) "
genelist_str="{}{}{}".format("['", "','".join(genes), "']")
conditionstr=''
if only_relations_with_pmid:
conditionstr=conditionstr + only_if_pubmed_id_exists
qry=qry_template.format(genestr=genelist_str,
reltype=reltype,
hops=hops,
dwpc_score_prop_name=dwpc_score_prop_name,
conditionstr=conditionstr)
return qry
def patterns_to_query(patterns):
'''
Convert pattern list to cypher query
Example
input patterns:list:
["(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})",
"(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2533'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})",
"(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'7827'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})",
"(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'55243'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})"]
output query:string
"MATCH p0=(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})
MATCH p1=(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2533'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})
MATCH p2=(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'7827'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})
MATCH p3=(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'55243'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})
RETURN p0,p1,p2,p3"
'''
match_phrases=[]
for i, pattern in enumerate(patterns):
match_phrases.append('MATCH p{i}={pattern} '.format(i=i, pattern=pattern))
return_phrase='RETURN ' + ','.join(["p{}".format(i) for i in range(len(patterns))])
query= ' '.join(match_phrases) + return_phrase
return query
# -
# # Tests
# ## Test `make_dwpc_query`
genes=['LEF1','TCF7','SERINC5']
reltype='INTERACTS_GiG3'
hops=2
query=make_dwpc_query(genes, reltype, hops)
print(query)
genes=['LEF1','TCF7','SERINC5']
reltype='INTERACTS_GiG3|INTERACTS_GiG3'
hops=2
query=make_dwpc_query(genes, reltype, hops)
print(query)
client=SimpleNeo4jHTTPAPIClient('http://localhost:7474')
client
result=client.execute_read_query(query)
assert result['results'][0]['data'], 'The DWPC query is not working.'
# ## Test `patterns_to_query`
patterns=["(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})",
"(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2533'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})",
"(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'7827'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})",
"(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'55243'})-[:INTERACTS_GiG3]->(:Gene{identifier:'2534'})-[:INTERACTS_GiG3]->(:Gene{identifier:'201633'})"]
patterns
query=patterns_to_query(patterns)
query
assert client.execute_read_query(query)['results'][0]['data'], 'Converted query from patterns is not working.'
| 01_dwpc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Beispiel für eine explorative Datenanalyse: Erdbeben der letzten 7 Tage (US Geological Survey)
# ==============================================================================================
#
# Import zweier Standardpakete für die Datenanalyse: Numpy für mehrdimensionale Arrays, Pandas für Datenanalyse in Tabellen.
import pandas as pd
import numpy as np
# %matplotlib inline
# Direkter Download vom USGS, Abruf des Downloaddatums, automatischer Import in Pandas-Dataframe
fileUrl = 'http://earthquake.usgs.gov/earthquakes/feed/v1.0/summary/2.5_day.csv'
eData = pd.read_csv(fileUrl)
# dateDownloaded = !date
dateDownloaded
# 1. Darstellung als Pandas-DataFrame
# -----------------------------------
#
# Darstellung des Datensatzes als Datensatzes als Pandas-Dataframe (Tabelle der ersten und letzten 30 Einträge, Anzahl Reihen und Spalten). Konvention: Variablen sind die Spalten, einzelne Messungen die Zeilen.
eData
# Kürzere Darstellung mit head(): nur die ersten 5 Einträge des Tabellenkopfes
eData.head()
# Anzahl der Reihen und Spalten mit Numpy shape().
np.shape(eData)
# Anzeige der einzelnen Spaltennamen mit Attribut DataFrame.columns
eData.columns
# Datentyp der einzelnen Variablen mit Attribut DataFrame.dtypes
eData.dtypes
# 2. Aufbereitung des Datensatzes
# -------------------------------
#
# Überprüfen, ob Tabelle NaN enthält, mit DataFrame.isnull().any()
eData.isnull().any()
# Entfernung aller Zeilen bzw. Messungen mit NaNs durch DataFrame.dropna()
eData = eData.dropna()
eData
eData.isnull().any()
# Überprüfen, ob Zeilen bzw. Messungen doppelt vorkommen, mit DataFrame.duplicated()
eData.duplicated().any()
# Es kommen also keine Duplikate vor. Bei Bedarf mit *DataFrame.drop_duplicates()* entfernen.
#
# 3. Explorative Statistiken
# --------------------------
#
# Statistische Beschreibung der numerischen Variablen mit Dataframe.describe() (count: Anzahl Messungen, mean: Mittelwert, std: Standardabweichung, min: Minimum, 25%: 25-Perzentil, ...)
eData.describe()
# Streumatrix für alle numerischen Variablen mit Pandas *scattermatrix()*:
pd.scatter_matrix(eData, figsize=(14,14), marker='o');
# 4. Analyse von Untermengen
# --------------------------
#
# Zugriff auf die Variable 'Lat' (latitude):
eData['latitude']
# Welche Erdbeben fanden oberhalb einer geographischen Breite von 40 Grad statt?
eData['latitude'] > 40.0
# Gab es überhaupt Erdbeben oberhalb 50 Grad Breite?
(eData['latitude'] > 40.0).any()
# Gab es also. Haben alle verzeichneten Erdbeben eine Breite größer als 18 Grad?
(eData['latitude'] > 18.0).all()
# Es sind also auch Erdbeben unterhalb von 18 Grad verzeichnet.
#
# Alle unterschiedlichen Werte der kategorischen Variable 'Version' mit Dataframe['Variablenname'].*unique()*
eData['Version'].unique()
# Häufigkeit der verschiedenen Kategorien in 'Version' mit Dataframe['Variablenname'].*value_counts()*:
eData['Version'].value_counts()
# Häufigkeit von Wertepaaren der beiden kategorischen Variablen 'Version' und 'Src' mit Pandas *crosstab()*:
pd.crosstab(eData['Src'], eData['Version'])
# Darstellung der Häufigkeitsverteilung der Erdbebenstärken für die verschiedenen Quellen mit einer Kastengraphik durch Pandas *boxplot()*:
from pandas.tools.plotting import boxplot
boxplot(eData, column='Magnitude', by='Src');
| explorative_analyse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
import matplotlib.pyplot as plt
# -
# ### Data Importing and Preprocessing
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
# The entire data is stored in a numpy array. The size of the images and corresponding masks have been reduced to 380 X 194.
# Replace the path by the path to the numpy array.
# In order to work with bigger images, make a DataSet object to directly read and process from the data folders.
data = np.load("/kaggle/input/spine-dataset/dataset.npz")
# +
# These are the segmentation masks
data.files
# +
# The shape of the images
ap_shapes = data["ap"].shape
print(ap_shapes)
lat_shapes = data["lat"].shape
print(lat_shapes)
# +
# Here, we are combining all the different inputs and outputs into single arrays.
ap_inp = data["ap"]
ap_inp = np.reshape(ap_inp, (ap_shapes[0], ap_shapes[1], ap_shapes[2], 1))
lat_inp = data["lat"]
lat_inp = np.reshape(lat_inp, (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1))
## NOTE:
# In this case the code for combining all 5 output segmentations for the LAT images is commented out.
# The LAT OUT Segmentations masks do not contain the anterior and posterior lines.
# In order to include these 2 masks as well, comment the last line and uncomment the second last line
ap_out = np.concatenate((np.reshape(data["ap_pedicle"], (ap_shapes[0], ap_shapes[1], ap_shapes[2], 1)), np.reshape(data["ap_spinous"], (ap_shapes[0], ap_shapes[1], ap_shapes[2], 1)), np.reshape(data["ap_spinous"], (ap_shapes[0], ap_shapes[1], ap_shapes[2], 1))), axis = -1)
# lat_out = np.concatenate((np.reshape(data["lat_ant_line"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1)), np.reshape(data["lat_post_line"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1)), np.reshape(data["lat_disk_height"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1)), np.reshape(data["lat_spinous"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1)), np.reshape(data["lat_vertebra"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1))), axis = -1)
lat_out = np.concatenate((np.reshape(data["lat_disk_height"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1)), np.reshape(data["lat_spinous"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1)), np.reshape(data["lat_vertebra"], (lat_shapes[0], lat_shapes[1], lat_shapes[2], 1))), axis = -1)
# +
# Checking the size of the input and output arrays.
# If the ant and post lines are included in the LAT masks output, there should be 5 channels instead of 3 in the -1 dimension of the LAT output
print(ap_inp.shape)
print(lat_inp.shape)
print(ap_out.shape)
print(lat_out.shape)
# +
#Data Labels for classification
labels = data["labels"]
print(labels.shape)
# -
import torch
import torchvision.transforms as transforms
import torch.nn as nn
import torch.nn.functional as F
from torch import optim, save
from torch.utils.data import Dataset, DataLoader
from torch.utils.data.sampler import Sampler
from torch.utils.data import random_split
from torch.autograd import Variable
# ### Dataset Preparation
# +
class Full_Dataset(Dataset):
def __init__(self, ap_images, lat_images, ap_masks, lat_masks, labels, transform = None):
# Since the images are 380 X 194, we pad them accordingly so that they can be downsampled / max pooled 4 times without loss of information.
# The pad size makes the image and output dimensions divisible by 16.
self.ap_images = np.pad(ap_images, ((0, 0), (2, 2), (7, 7), (0, 0)))
self.lat_images = np.pad(lat_images, ((0, 0), (2, 2), (7, 7), (0, 0)))
self.ap_masks = np.pad(ap_masks, ((0, 0), (2, 2), (7, 7), (0, 0)))
self.lat_masks = np.pad(lat_masks, ((0, 0), (2, 2), (7, 7), (0, 0)))
self.labels = labels
self.transform = transform
def __len__(self):
return self.ap_images.shape[0]
def __getitem__(self, i):
ap_image = self.ap_images[i]
lat_image = self.ap_images[i]
ap_mask = self.ap_masks[i]
lat_mask = self.lat_masks[i]
label = self.labels[i]
# We can add a few other random transformations to make the model more robust.
sample = {"ap_image": ap_image, "lat_image": lat_image, "ap_mask": ap_mask, "lat_mask": lat_mask, "label": label}
if self.transform:
sample = self.transform(sample)
return sample
class ToTensor(object):
def __call__(self, sample):
ap_image, ap_mask, lat_image, lat_mask, label = sample['ap_image'], sample['ap_mask'], sample['lat_image'], sample['lat_mask'], sample['label']
# Making the tensors from the numpy arrays
ap_image = ap_image.transpose(2, 0, 1)
ap_mask = ap_mask.transpose(2, 0, 1)
lat_image = lat_image.transpose(2, 0, 1)
lat_mask = lat_mask.transpose(2, 0, 1)
label = label
ap_image = torch.from_numpy(ap_image)
ap_image = ap_image.type(torch.FloatTensor)
lat_image = torch.from_numpy(lat_image)
lat_image = lat_image.type(torch.FloatTensor)
ap_mask = torch.from_numpy(ap_mask)
ap_mask = ap_mask.type(torch.FloatTensor)
lat_mask = torch.from_numpy(lat_mask)
lat_mask = lat_mask.type(torch.FloatTensor)
label = torch.from_numpy(label)
label = label.type(torch.FloatTensor)
return {"ap_image": ap_image, "lat_image": lat_image, "ap_mask": ap_mask, "lat_mask": lat_mask, "label": label}
# -
dataset = Full_Dataset(ap_inp, lat_inp, ap_out, lat_out, labels, transform = transforms.Compose([ToTensor()]))
len(dataset)
# +
# Train Val Test split
train_size = 600
test_size = 0
val_size = len(dataset) - train_size - test_size
train_data, val_data, test_data = random_split(dataset, [train_size, val_size, test_size])
# -
# #### Checking Data Dimensions
# +
print("training")
for i in range(len(train_data)):
sample = train_data[i]
print(i, sample['ap_image'].size(), sample['ap_mask'].size(), sample['lat_image'].size(), sample['lat_mask'].size(), sample['label'].size())
if i == 5:
break
print("validation")
for i in range(len(val_data)):
sample = val_data[i]
print(i, sample['ap_image'].size(), sample['ap_mask'].size(), sample['lat_image'].size(), sample['lat_mask'].size(), sample['label'].size())
if i == 2:
break
# -
# ### DataLoader Construction
train_loader = DataLoader(dataset = train_data, batch_size = 16, shuffle=True, num_workers=0)
val_loader = DataLoader(dataset= val_data, batch_size = 16, shuffle=True, num_workers=0)
# test_loader = DataLoader(dataset= test_data, batch_size = 2, shuffle=True, num_workers=0)
print(len(train_loader))
print(len(val_loader))
# ## Model Construction
#
# The model would be a combination of 2 Unets and one simple 5 layer convolutional network classifier.
#
# The unet models would take in 2 input images and 2 output masks, and there would be one Unet for each of LAT and AP images.
#
# The output of the segmentations along with the original images are fed into the classifier.
# +
class conv_block(nn.Module):
def __init__(self, ch_in, ch_out, dropout_rate):
super(conv_block, self).__init__()
self.conv = nn.Sequential(nn.Conv2d(ch_in, ch_out, kernel_size = 3, padding = 1),
nn.BatchNorm2d(ch_out),
nn.Dropout2d(dropout_rate),
nn.ReLU(),
nn.Conv2d(ch_out, ch_out, kernel_size = 3, padding = 1),
nn.BatchNorm2d(ch_out),
nn.Dropout2d(dropout_rate),
nn.ReLU())
def forward(self, x):
x = self.conv(x)
return x
class deconv_block(nn.Module):
def __init__(self, ch_in, ch_out, dropout_rate):
super(deconv_block, self).__init__()
self.deconv = nn.Sequential((nn.ConvTranspose2d(ch_in, ch_out, kernel_size=2, stride = 2)), nn.Dropout2d(dropout_rate))
def forward(self, x):
x = self.deconv(x)
return x
class attention_block(nn.Module):
def __init__(self, ch_in, ch_skip, ch_out, dropout_rate):
super(attention_block, self).__init__()
self.W_skip = nn.Sequential(nn.Conv2d(ch_skip, ch_out, kernel_size = 1, padding = 0),
nn.BatchNorm2d(ch_out),
nn.Dropout2d(dropout_rate))
self.W_in = nn.Sequential(nn.Conv2d(ch_skip, ch_out, kernel_size = 1, padding = 0),
nn.BatchNorm2d(ch_out),
nn.Dropout2d(dropout_rate))
self.relu = nn.ReLU()
self.psi = nn.Sequential(nn.Conv2d(ch_out, 1, kernel_size = 1, padding = 0),
nn.BatchNorm2d(1),
nn.Dropout2d(dropout_rate),
nn.Sigmoid())
def forward(self, x, skip):
# print(x.size())
# print(skip.size())
g = self.W_skip(skip)
x = self.W_in(x)
psi = self.relu(g+x)
psi = self.psi(psi)
return skip*psi
class attention_unet_block(nn.Module):
def __init__(self, inp_channel, out_channel, multiplier, dropout_rate):
super(attention_unet_block, self).__init__()
self.Maxpool1 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Maxpool2 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Maxpool3 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Maxpool4 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Conv1 = conv_block(ch_in=inp_channel,ch_out=8 * multiplier, dropout_rate = dropout_rate)
self.Conv2 = conv_block(ch_in=8 * multiplier,ch_out=16 * multiplier, dropout_rate = dropout_rate)
self.Conv3 = conv_block(ch_in=16 * multiplier,ch_out=32 * multiplier, dropout_rate = dropout_rate)
self.Conv4 = conv_block(ch_in=32 * multiplier,ch_out=64 * multiplier, dropout_rate = dropout_rate)
self.Conv5 = conv_block(ch_in=64 * multiplier,ch_out=128 * multiplier, dropout_rate = dropout_rate)
self.Up5 = deconv_block(ch_in=128 * multiplier,ch_out=64 * multiplier, dropout_rate = dropout_rate)
self.Att5 = attention_block(ch_in=64 * multiplier,ch_skip=64 * multiplier,ch_out=32 * multiplier, dropout_rate = dropout_rate)
self.Upconv5 = conv_block(ch_in=128 * multiplier, ch_out=64 * multiplier, dropout_rate = dropout_rate)
self.Up4 = deconv_block(ch_in=64 * multiplier,ch_out=32 * multiplier, dropout_rate = dropout_rate)
self.Att4 = attention_block(ch_in=32 * multiplier,ch_skip=32 * multiplier,ch_out=16 * multiplier, dropout_rate = dropout_rate)
self.Upconv4 = conv_block(ch_in=64 * multiplier, ch_out=32 * multiplier, dropout_rate = dropout_rate)
self.Up3 = deconv_block(ch_in=32 * multiplier,ch_out=16 * multiplier, dropout_rate = dropout_rate)
self.Att3 = attention_block(ch_in=16 * multiplier,ch_skip=16 * multiplier,ch_out=8 * multiplier, dropout_rate = dropout_rate)
self.Upconv3 = conv_block(ch_in=32 * multiplier, ch_out=16 * multiplier, dropout_rate = dropout_rate)
self.Up2 = deconv_block(ch_in=16 * multiplier,ch_out=8 * multiplier, dropout_rate = dropout_rate)
self.Att2 = attention_block(ch_in=8 * multiplier,ch_skip=8 * multiplier,ch_out=4 * multiplier, dropout_rate = dropout_rate)
self.Upconv2 = conv_block(ch_in=16 * multiplier, ch_out=8 * multiplier, dropout_rate = dropout_rate)
self.Conv_1x1 = nn.Sequential(nn.Conv2d(8 * multiplier,out_channel,kernel_size=1,stride=1,padding=0), nn.Sigmoid())
def forward(self, x):
#conv path
x1 = self.Conv1(x)
x2 = self.Maxpool1(x1)
x2 = self.Conv2(x2)
x3 = self.Maxpool2(x2)
x3 = self.Conv3(x3)
x4 = self.Maxpool3(x3)
x4 = self.Conv4(x4)
x5 = self.Maxpool4(x4)
x5 = self.Conv5(x5)
# decoding + concat path
d5 = self.Up5(x5)
s4 = self.Att5(x = d5, skip = x4)
d5 = torch.cat((s4,d5),dim=1)
d5 = self.Upconv5(d5)
d4 = self.Up4(d5)
s3 = self.Att4(x=d4,skip=x3)
d4 = torch.cat((s3,d4),dim=1)
d4 = self.Upconv4(d4)
d3 = self.Up3(d4)
s2 = self.Att3(x=d3,skip=x2)
d3 = torch.cat((s2,d3),dim=1)
d3 = self.Upconv3(d3)
d2 = self.Up2(d3)
s1 = self.Att2(x=d2,skip=x1)
d2 = torch.cat((s1,d2),dim=1)
d2 = self.Upconv2(d2)
d1 = self.Conv_1x1(d2)
return d1
class classifier_block(nn.Module):
def __init__(self, dimensions, inp_channel, output_class, multiplier_classifier, dropout_rate, dropout_rate_classifier):
super(classifier_block, self).__init__()
self.multiplier_classifier = multiplier_classifier
self.dimensions = dimensions
self.Maxpool1 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Maxpool2 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Maxpool3 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Maxpool4 = nn.MaxPool2d(kernel_size=2,stride=2)
self.Conv1 = conv_block(ch_in=inp_channel,ch_out= 1 * multiplier_classifier, dropout_rate = dropout_rate)
self.Conv2 = conv_block(ch_in=1 * multiplier_classifier,ch_out=2 * multiplier_classifier, dropout_rate = dropout_rate)
self.Conv3 = conv_block(ch_in=2 * multiplier_classifier,ch_out=4 * multiplier_classifier, dropout_rate = dropout_rate)
self.Conv4 = conv_block(ch_in=4 * multiplier_classifier,ch_out=8 * multiplier_classifier, dropout_rate = dropout_rate)
self.fc1 = nn.Sequential(nn.Linear(8 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16), 4 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16)),
nn.BatchNorm1d( 4 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16)),
nn.Dropout(dropout_rate_classifier),
nn.ReLU())
self.fc2 = nn.Sequential(nn.Linear(4 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16), 2 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16)),
nn.BatchNorm1d( 2 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16)),
nn.Dropout(dropout_rate_classifier),
nn.ReLU())
self.out = nn.Sequential(nn.Linear(2 * multiplier_classifier * int(dimensions[0] * 1 / 16) * int(dimensions[1] * 1 / 16), output_class),
nn.Sigmoid())
def forward(self, x):
x1 = self.Conv1(x)
x2 = self.Maxpool1(x1)
x2 = self.Conv2(x2)
x3 = self.Maxpool2(x2)
x3 = self.Conv3(x3)
x4 = self.Maxpool3(x3)
x4 = self.Conv4(x4)
x5 = self.Maxpool4(x4)
x5 = torch.flatten(x5, start_dim = 1)
y1 = self.fc1(x5)
y2 = self.fc2(y1)
y = self.out(y2)
return y
class dual_attention_unet_classifier(nn.Module):
def __init__(self, dims, inp1_channel, out1_channel, inp2_channel, out2_channel, classifier_output, multiplier, multiplier_classifier, dropout_rate, dropout_rate_classifier):
super(dual_attention_unet_classifier, self).__init__()
self.attention_unet_1 = attention_unet_block(inp1_channel, out1_channel, multiplier, dropout_rate)
self.attention_unet_2 = attention_unet_block(inp2_channel, out2_channel, multiplier, dropout_rate)
self.classifier = classifier_block(dims, inp1_channel + out1_channel + inp2_channel + out2_channel, classifier_output, multiplier_classifier, dropout_rate, dropout_rate_classifier)
def forward(self, x1, x2):
out1 = self.attention_unet_1(x1)
out2 = self.attention_unet_2(x2)
class_out = self.classifier(torch.cat((x1, out1, x2, out2),dim=1))
return out1, out2, class_out
# -
# ### Soft Dice Loss Class
class SoftDiceLoss(nn.Module):
def __init__(self):
super(SoftDiceLoss, self).__init__()
def forward(self, probs, targets):
smooth = 0.001
num = targets.size(0)
score = 0.
for i in range(probs.shape[1]):
m1 = probs[:, i, :, :].view(num, -1)
m2 = targets[:, i, :, :].view(num, -1)
intersection = (m1 * m2)
sc = 2. * (intersection.sum(1) + smooth) / (m1.sum(1) + m2.sum(1) + smooth)
score += (1 - sc.sum() / num ) * 100. / probs.shape[1]
score = torch.mean(score)
return score
# #### Model Initialization
# +
#change the number of out2_channels to 5 if trying to segment the ant and post lines for LAT images.
#Play around with the other hyper parameters.
# The input sizes of the images after padding are the first argument to the function.
model = dual_attention_unet_classifier([384, 208], inp1_channel=1, out1_channel= 3, inp2_channel=1, out2_channel= 3, classifier_output=1, multiplier=4, multiplier_classifier=1, dropout_rate=0.2, dropout_rate_classifier=0.4)
model = model.cuda()
# -
# #### Loss and Optimizer Class Instantiation
# +
criterion_classifier = nn.BCELoss()
criterion_segmentor = SoftDiceLoss()
optimizerclass = optim.Adam(model.parameters(), lr = 0.01)
optimizerseg1 = optim.Adam(model.parameters(), lr = 0.01)
optimizerseg2 = optim.Adam(model.parameters(), lr = 0.01)
epochs = 300
use_cuda = torch.cuda.is_available()
device = torch.device("cuda:0" if use_cuda else "cpu")
# -
# ## Model Training
# +
import time
# All the below lists would store the average training and validation losses and accuracy for different epochs
seg1_train_loss_array = []
seg2_train_loss_array = []
ce_train_loss_array = []
seg1_val_loss_array = []
seg2_val_loss_array = []
ce_val_loss_array = []
correct_array = []
val_correct_array = []
for epoch in range(epochs):
start_time = time.time()
# The below arrays would store the losses and accuracies for different mini batches
seg1_temp_loss = []
seg2_temp_loss = []
ce_temp_loss = []
seg1_val_temp_loss = []
seg2_val_temp_loss = []
ce_val_temp_loss = []
correct = 0
val_correct = 0
for mini_batch_num, data in enumerate(train_loader):
torch.cuda.empty_cache()
ap_image, ap_mask, lat_image, lat_mask, label = data['ap_image'], data['ap_mask'], data['lat_image'], data['lat_mask'], data['label']
ap_image, ap_mask, lat_image, lat_mask, label = ap_image.to(device), ap_mask.to(device), lat_image.to(device), lat_mask.to(device), label.to(device)
# image, mask = image.to(device), mask.to(device)
model.train()
ap_seg, lat_seg, op = model(ap_image, lat_image)
#
loss_seg1 = criterion_segmentor(ap_seg, ap_mask)
seg1_temp_loss.append(loss_seg1.item())
optimizerseg1.zero_grad()
loss_seg1.backward(retain_graph=True)
optimizerseg1.step()
#
loss_seg2 = criterion_segmentor(lat_seg, lat_mask)
seg2_temp_loss.append(loss_seg2.item())
optimizerseg2.zero_grad()
loss_seg2.backward(retain_graph=True)
optimizerseg2.step()
#
loss_class = 100. * criterion_classifier(op, label)
ce_temp_loss.append(loss_class.item() / 100.)
optimizerclass.zero_grad()
loss_class.backward()
optimizerclass.step()
op = (op>0.5).float()
correct += (op == label).float().sum()
print("Epoch {}/{}, MiniBatch {}/{},\tSegmnetation AP Loss {},\tSegmentation LAT Loss {},\tClassifier Loss (BCE) {}".format(epoch + 1, epochs, mini_batch_num + 1, len(train_loader), round(loss_seg1.item(), 3), round(loss_seg2.item(), 3), round(loss_class.item() / 100., 3)), end = "\r", flush = True)
correct = correct.item() / train_size
correct_array.append(correct)
# Validation
with torch.no_grad():
for i, data in enumerate(val_loader):
ap_image, ap_mask, lat_image, lat_mask, label = data['ap_image'], data['ap_mask'], data['lat_image'], data['lat_mask'], data['label']
ap_image, ap_mask, lat_image, lat_mask, label = ap_image.to(device), ap_mask.to(device), lat_image.to(device), lat_mask.to(device), label.to(device)
model.eval()
ap_seg, lat_seg, op = model(ap_image, lat_image)
loss_seg1 = criterion_segmentor(ap_seg, ap_mask)
seg1_val_temp_loss.append(loss_seg1.item())
loss_seg2 = criterion_segmentor(lat_seg, lat_mask)
seg2_val_temp_loss.append(loss_seg2.item())
loss_class = criterion_classifier(op, label)
ce_val_temp_loss.append(loss_class.item())
op = (op>0.5).float()
val_correct += (op == label).float().sum()
# torch.cuda.empty_cache()
val_correct = val_correct.item() / val_size
val_correct_array.append(val_correct)
end_time = time.time()
seg1_temp_loss = np.mean(np.array(seg1_temp_loss))
seg2_temp_loss = np.mean(np.array(seg2_temp_loss))
ce_temp_loss = np.mean(np.array(ce_temp_loss))
seg1_val_temp_loss = np.mean(np.array(seg1_val_temp_loss))
seg2_val_temp_loss = np.mean(np.array(seg2_val_temp_loss))
ce_val_temp_loss = np.mean(np.array(ce_val_temp_loss))
seg1_train_loss_array.append(seg1_temp_loss)
seg2_train_loss_array.append(seg2_temp_loss)
ce_train_loss_array.append(ce_temp_loss)
seg1_val_loss_array.append(seg1_val_temp_loss)
seg2_val_loss_array.append(seg2_val_temp_loss)
ce_val_loss_array.append(ce_val_temp_loss)
epoch_time = end_time - start_time
# print(correct.item(), val_correct.item())
print("Epoch {}/{},\t Time {} seconds;\t Train Seg AP Loss {},\tTrain Seg LAT Loss {},\t Train Class Loss {},\tTrain Accuracy {},\t Val Seg AP Loss {},\t Val Seg LAT Loss {},\t Val Class Loss {},\tVal Accuracy {}".format(epoch + 1, epochs, round(epoch_time), round(seg1_temp_loss, 3), round(seg2_temp_loss, 3), round(ce_temp_loss, 3), round(correct, 3), round(seg1_val_temp_loss, 3), round(seg2_val_temp_loss, 3), round(ce_val_temp_loss, 3), round(val_correct, 3)))
torch.cuda.empty_cache()
# -
# ### Loss Tracking
# +
fig, ax = plt.subplots(2, 2, figsize = (20, 20))
ax[0, 1].plot(seg1_train_loss_array, label = "AP Segmentation Loss Training")
ax[0, 1].plot(seg1_val_loss_array, label = "AP Segmentation Loss Validation")
ax[0, 1].legend()
ax[1, 1].plot(seg2_train_loss_array, label = "LAT Segmentation Loss Training")
ax[1, 1].plot(seg2_val_loss_array, label = "LAT Segmentation Loss Validation")
ax[1, 1].legend()
ax[0, 0].plot(ce_train_loss_array, label = "Classification CE Loss Training")
ax[0, 0].plot(ce_val_loss_array, label = "Classification CE Loss Validation")
ax[0, 0].legend()
ax[1, 0].plot(correct_array, label = "Classification Accuracy Training")
ax[1, 0].plot(val_correct_array, label = "Classification Accuracy Validation")
ax[1, 0].legend()
plt.show()
# -
# ### Model Save Dict
torch.save(model.state_dict(), "total_seg_class_model_v1_3")
| assignment_1/total_segmentation_and_classification_model_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 请你给一个停车场设计一个停车系统。
# 停车场总共有三种不同大小的车位:大,中和小,每种尺寸分别有固定数目的车位。
#
# 请你实现 ParkingSystem 类:
#
# ParkingSystem(int big, int medium, int small) 初始化 ParkingSystem 类,三个参数分别对应每种停车位的数目。
# bool addCar(int carType) 检查是否有 carType 对应的停车位。 carType 有三种类型:大,中,小,分别用数字 1, 2 和 3 表示。
# 一辆车只能停在 carType 对应尺寸的停车位中。如果没有空车位,请返回 false ,否则将该车停入车位并返回 true 。
#
#
# 示例 1:
# 输入:
# ["ParkingSystem", "addCar", "addCar", "addCar", "addCar"]
# [[1, 1, 0], [1], [2], [3], [1]]
# 输出:
# [null, true, true, false, false]
# 解释:
# ParkingSystem parkingSystem = new ParkingSystem(1, 1, 0);
# parkingSystem.addCar(1); // 返回 true ,因为有 1 个空的大车位
# parkingSystem.addCar(2); // 返回 true ,因为有 1 个空的中车位
# parkingSystem.addCar(3); // 返回 false ,因为没有空的小车位
# parkingSystem.addCar(1); // 返回 false ,因为没有空的大车位,唯一一个大车位已经被占据了
#
# 提示:
# 1、0 <= big, medium, small <= 1000
# 2、carType 取值为 1, 2 或 3
# 3、最多会调用 addCar 函数 1000 次
# -
class ParkingSystem:
def __init__(self, big: int, medium: int, small: int):
self.big = big
self.med = medium
self.small = small
def addCar(self, carType: int) -> bool:
if carType == 1 and self.big > 0:
self.big -= 1
return True
if carType == 2 and self.med > 0:
self.med -= 1
return True
if carType == 3 and self.small > 0:
self.small -= 1
return True
return False
| Design/1027/1603. Design Parking System.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ''
# language: python
# name: python3
# ---
from collections import defaultdict
# +
def is_small(cave):
return cave.islower()
def search_cave_1(cave="start"):
val = 0
if cave == "end":
return 1
if is_small(cave) and cave in visited:
return 0
if is_small(cave):
visited.add(cave)
for adj_cave in caves[cave]:
val += search_cave_1(adj_cave)
if cave in visited:
visited.remove(cave)
return val
def search_cave_2(cave="start"):
val = 0
if cave == "end":
return 1
if is_small(cave):
if cave in visited_2 and visited_2[cave] > 1:
return 0
elif cave in visited_2 and visited_2[cave] > 0 and max(list(visited_2.values())) > 1:
return 0
if is_small(cave):
if cave in visited_2:
visited_2[cave] += 1
else:
visited_2[cave] = 1
for adj_cave in caves[cave]:
if adj_cave == "start":
continue
val += search_cave_2(adj_cave)
if cave in visited_2 and visited_2[cave] > 0:
visited_2[cave] -= 1
return val
def test():
input_file_name = './test_input.txt'
return main(input_file_name)
def main(input_file_name="./input.txt"):
pairs = [line.split('-') for line in open(input_file_name, 'r').read().strip().split('\n')]
global caves
caves = defaultdict(list)
for a,b in pairs:
caves[a].append(b)
caves[b].append(a)
global visited
global visited_2
visited = set()
visited_2 = {}
finalScore1,finalScore2 = search_cave_1("start"),search_cave_2("start")
return [finalScore1,finalScore2]
if __name__ == "__main__":
test = test()
answers = main()
print(f"Answer to test-question 1: {test[0]}")
assert test[0] == 226, "failed test"
print(f"Answer to question 1: {answers[0]}")
print(f"Answer to test-question 2: {test[1]}")
assert test[1] == 3509, "failed test"
print(f"Answer to question 2: {answers[1]}")
#lines = value[0]
#numbers = value[1]
# +
| 2021/12/12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Standard python helper libraries.
import os, sys, re, json, time, wget, csv, string, time, random
import itertools, collections
# NumPy
import numpy as np
# NLTK for NLP utils
import nltk
nltk.download('punkt')
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem.porter import PorterStemmer
from keras import backend as K
from keras.models import Sequential
from keras.layers import GaussianNoise, Dropout, Dense, Embedding, MaxPool1D, GlobalMaxPool1D, Conv1D, LSTM, Bidirectional
from keras.optimizers import Adam
from keras.callbacks import Callback
from sklearn.metrics import confusion_matrix, f1_score, precision_score, recall_score
from pymagnitude import *
# -
# # Categorical Classifier for Phenotypes using a Convolutional Neural Network
#
# In this notebook, we will be building a model for classifying a sentence from an EHR note for the presence of phenotypes. The model will be trained using a convoultional neural network (CNN).
#
# ## Word Embedding
#
# The first step is to load in our word embedding, which is trained on text from Wikipedia, Pubmed, and Pubmed Central. We load our word embedding through a tool called Magnitude. A full exploration of the word embedding and Magnitude can be found in the Introduction notebook.
med_vectors = Magnitude("data/wikipedia-pubmed-and-PMC-w2v.magnitude", pad_to_length=30)
# ## Text Preprocessing
# +
ehr_notes = []
with open('data/ehr_samples.csv', newline='') as csvfile:
reader = csv.DictReader(csvfile)
for row in reader:
if int(row['SpecialtyID']) in [39, 6, 16, 37, 11, 12, 29, 26, 7, 21, 19, 10, 2, 18]:
continue
else:
ehr_notes.append([row['Specialty'], row['Note']])
print(ehr_notes[0])
print(ehr_notes[2])
# -
# ## Natural Langauge Processing
# +
ehr_sentences = []
for record in ehr_notes:
sent_text = nltk.sent_tokenize(record[1])
for sent in sent_text:
tokens = word_tokenize(sent)
# convert to lower case
tokens = [w.lower() for w in tokens]
# remove punctuation from each word
table = str.maketrans('', '', string.punctuation)
tokens = [w.translate(table) for w in tokens]
# filter out stop words
stop_words = set(stopwords.words('english'))
tokens = [w for w in tokens if not w in stop_words]
# # stem words
# porter = PorterStemmer()
# tokens = [porter.stem(word) for word in tokens]
# remove blanks
tokens = [w for w in tokens if w != '']
ehr_sentences.append([record[0], ' '.join(tokens)])
random.Random(4).shuffle(ehr_sentences)
print(ehr_sentences[0])
print(ehr_sentences[1])
# -
# ## Defining our Training and Test Data
#
# Before we can start building our neural networks, we first have to define our datasets. Specifically, we have to break up our EHR data so that we have records that we can train on and records that are exclusively used to test on. Maintaining a separate set for testing ensures we avoid overfitting our data.
#
# We will use some built-in functions provided by Magnitude that helps encode our classes/categories. We then partition our data into our train and test sets. For each set we have both data and labels. Initially, we will be making these partitions small to make iterating through model development much quicker. However, once the models are developed, we will expand our datasets to include all of our data. To ensure we defined our data correctly, we can print a few lines from the two sets.
len(ehr_sentences)
# +
add_intent, intent_to_int, int_to_intent = MagnitudeUtils.class_encoding()
x_train = [ehr_sent[1].split(' ') for ehr_sent in ehr_sentences[:60000]]
x_test = [ehr_sent[1].split(' ') for ehr_sent in ehr_sentences[60001:]]
y_train = [add_intent(ehr_sent[0]) for ehr_sent in ehr_sentences[:60000]]
y_test = [add_intent(ehr_sent[0]) for ehr_sent in ehr_sentences[60001:]]
y_train = list(np.array(y_train).reshape(len(y_train)))
y_test = list(np.array(y_test).reshape(len(y_test)))
num_training = len(x_train)
num_test = len(x_test)
num_outputs = int(max(max(y_train), max(y_test))) + 1
print(int_to_intent(0))
print("First line of train/test data:")
print("\t", x_train[0])
print("\t", y_train[0], int_to_intent(y_train[0]))
print("\t", x_test[0])
print("\t", y_test[0], int_to_intent(y_test[0]))
print("Second line of train/test data:")
print("\t", x_train[1])
print("\t", y_train[1], int_to_intent(y_train[1]))
print("\t", x_test[1])
print("\t", y_test[1], int_to_intent(y_test[1]))
# -
# ## Defining Custom Callback Function
def f1(y_true, y_pred):
def recall(y_true, y_pred):
"""Recall metric.
Only computes a batch-wise average of recall.
Computes the recall, a metric for multi-label classification of
how many relevant items are selected.
"""
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
recall = true_positives / (possible_positives + K.epsilon())
return recall
def precision(y_true, y_pred):
"""Precision metric.
Only computes a batch-wise average of precision.
Computes the precision, a metric for multi-label classification of
how many selected items are relevant.
"""
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
return precision
precision = precision(y_true, y_pred)
recall = recall(y_true, y_pred)
return 2*((precision*recall)/(precision+recall+K.epsilon()))
# ## Defining Model
# +
MAX_WORDS = 30 # The maximum number of words the sequence model will consider
STD_DEV = 0.01 # Deviation of noise for Gaussian Noise applied to the embeddings
DROPOUT_RATIO = .5 # The ratio to dropout
BATCH_SIZE = 100 # The number of examples per train/validation step
EPOCHS = 100 # The number of times to repeat through all of the training data
LEARNING_RATE = .01 # The learning rate for the optimizer
HIDDEN_UNITS = 100
model = Sequential()
model.add(GaussianNoise(STD_DEV, input_shape=(MAX_WORDS, med_vectors.dim)))
model.add(Bidirectional(LSTM(HIDDEN_UNITS, activation='tanh'), merge_mode='concat'))
model.add(Dropout(DROPOUT_RATIO))
model.add(Dense(num_outputs, activation='softmax'))
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', f1])
model.summary()
# -
# ## Training Batches and Epochs
# +
training_batches = MagnitudeUtils.batchify(x_train, y_train, BATCH_SIZE) # Split the training data into batches
num_batches_per_epoch_train = int(np.ceil(num_training/float(BATCH_SIZE)))
test_batches = MagnitudeUtils.batchify(x_test, y_test, BATCH_SIZE) # Split the test data into batches
num_batches_per_epoch_test = int(np.ceil(num_test/float(BATCH_SIZE)))
# Generates batches of the transformed training data
train_batch_generator = (
(
med_vectors.query(x_train_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_train_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_train_batch, y_train_batch in training_batches
)
# Generates batches of the transformed test data
test_batch_generator = (
(
med_vectors.query(x_test_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_test_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_test_batch, y_test_batch in test_batches
)
# Start training
model.fit_generator(
generator = train_batch_generator,
steps_per_epoch = num_batches_per_epoch_train,
validation_data = test_batch_generator,
validation_steps = num_batches_per_epoch_test,
epochs = EPOCHS,
)
# -
# ## Results
# +
print("Results after training for %d epochs:" % (EPOCHS,))
train_metrics = model.evaluate_generator(
generator = train_batch_generator,
steps = num_batches_per_epoch_train,
)
print("loss: %.4f - categorical_accuracy: %.4f - f1: %.4f" % tuple(train_metrics))
val_metrics = model.evaluate_generator(
generator = test_batch_generator,
steps = num_batches_per_epoch_test,
)
print("val_loss: %.4f - val_categorical_accuracy: %.4f - f1: %.4f" % tuple(val_metrics))
# -
# ## Conclusion
# +
import matplotlib.pyplot as plt
# Plot training & validation accuracy values
plt.plot(model.history.history['acc'])
plt.plot(model.history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(model.history.history['loss'])
plt.plot(model.history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# +
from sklearn.metrics import confusion_matrix, classification_report
y_pred = np.argmax(model.predict(med_vectors.query(x_test)), axis=1)
class_labels = [int_to_intent(y) for y in set(y_test)]
report = classification_report(y_test, y_pred, target_names=class_labels)
print(class_labels)
print(confusion_matrix(y_test, y_pred))
print(report)
# +
from sklearn.utils.multiclass import unique_labels
def plot_confusion_matrix(y_true, y_pred, classes,
title=None,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
# Only use the labels that appear in the data
classes = [int_to_intent(y_class) for y_class in unique_labels(y_test, y_pred)]
fig, ax = plt.subplots()
fig.set_figheight(15)
fig.set_figwidth(15)
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
fig.tight_layout()
return ax
np.set_printoptions(precision=2)
plot_confusion_matrix(y_test, y_pred, classes=class_labels
, title='Confusion Matrix for Phenotype LSTM Classifier')
# +
correct = 0
for i, y in enumerate(y_pred):
if y == y_test[i]:
correct += 1
correct / len(y_pred)
# -
| 5.2_Phenotype-LSTM-Sentences-30WordLimit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Deep AI
# language: python
# name: dl
# ---
# +
import time
import torch
from torch.optim.lr_scheduler import ReduceLROnPlateau
from torch_geometric.nn import VGAE
from torch_geometric.loader import DataLoader
from torch_geometric.utils import degree, negative_sampling
from torch.utils.tensorboard import SummaryWriter
from gene_graph_dataset import G3MedianDataset
from phylognn_model import G3Median_GCNConv
# +
gpuid = 1
train_p, test_p = 0.8, 0.2
train_batch, test_batch = 128, 64
# -
device = torch.device('cuda:' + str(gpuid) if torch.cuda.is_available() else 'cpu')
dataset = G3MedianDataset('dataset_g3m', 100, 100)
data_size = 1000 # len(dataset)
train_size, test_size = (int)(data_size * train_p), (int)(data_size * test_p)
dataset = dataset.shuffle()
train_dataset = dataset[:train_size]
test_dataset = dataset[train_size:(train_size + test_size)]
# val_dataset = dataset[(train_size + test_size):]
train_loader = DataLoader(train_dataset, batch_size = train_batch, shuffle=True)
# test_loader = DataLoader(test_dataset, batch_size = test_batch)
# val_loader = DataLoader(val_dataset, batch_size=8)
# +
# deg = torch.zeros(5, dtype=torch.long)
# for data in train_dataset:
# d = degree(data.edge_index[1].type(torch.int64),
# num_nodes=data.num_nodes, dtype=torch.long)
# deg += torch.bincount(d, minlength=deg.numel())
# -
in_channels, out_channels = dataset.num_features, 16
model = VGAE(G3Median_GCNConv(in_channels, out_channels)).to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.5, patience=10,
min_lr=0.00001)
writer = SummaryWriter(log_dir='runs_g3m/g3median_l1000_gcn_tb1')
def train(train_loader):
model.train()
total_loss = 0
for data in train_loader:
data = data.to(device)
optimizer.zero_grad()
z = model.encode(data.x, data.edge_index)
loss = model.recon_loss(z, data.pos_edge_label_index)
loss = loss + (1 / data.num_nodes) * model.kl_loss()
loss.backward()
optimizer.step()
total_loss += loss
return total_loss/len(train_loader)
@torch.no_grad()
def test(test_loader):
model.eval()
loss, auc, ap = 0, 0, 0
for data in test_loader:
data = data.to(device)
neg_samples = negative_sampling(data.pos_edge_label_index,
data.num_nodes,
data.num_nodes*10)
z = model.encode(data.x, data.edge_index)
loss += model.recon_loss(z, data.pos_edge_label_index, neg_samples)
# negative_sampling(data.pos_edge_label_index,
# data.num_nodes,
# data.num_nodes*10))
# data.neg_edge_label_index)
tauc, tap = model.test(z, data.pos_edge_label_index, neg_samples)
# negative_sampling(data.pos_edge_label_index,
# data.num_nodes,
# data.num_nodes*10))
# data.neg_edge_label_index)
auc += tauc
ap += tap
return loss/len(test_loader), auc/len(test_loader), ap/len(test_loader)
for epoch in range(201, 600 + 1):
print(f'{time.ctime()} - Epoch: {epoch:04d}')
loss = train(train_loader)
print(f'{time.ctime()} - \t train loss: {loss:.6f}')
tloss, auc, ap = test(test_dataset)
print(f'{time.ctime()} - \t test loss: {tloss:.6f}, auc: {auc:.6f}, ap: {ap:.6f}')
scheduler.step(1 - auc)
writer.add_scalar('Loss/train', loss, epoch)
writer.add_scalar('Loss/test', tloss, epoch)
writer.add_scalar('AUC/test', auc, epoch)
writer.add_scalar('AP/test', ap, epoch)
# if epoch % 50 == 0:
# print(f'{time.ctime()} - '
# f'Epoch: {epoch:04d}, loss: {loss:.6f}, AUC: {auc:.6f}, '
# f', TL: {tloss:.6f}')
writer.close()
# +
# torch.save(model.state_dict(), 'g2g_test_model_batch')
# -
| g3median.ipynb/g3median_test_gcn-tb1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Literate programming contoh implementasinya dalam pembelajaran sains
#
# - Author: <NAME>[1, [GS](https://scholar.google.com/citations?hl=en&user=Myvc78MAAAAJ&view_op=list_works&sortby=pubdate), [ORCID](https://orcid.org/0000-0002-1526-0863)], <NAME>[2, [GS](https://scholar.google.co.id/citations?hl=en&user=NK7TDDQAAAAJ&view_op=list_works&sortby=pubdate), [ORCID](https://orcid.org/0000-0002-6694-2071), dan <NAME>[[GS](https://scholar.google.com/citations?hl=id&user=uYQgjxMAAAAJ&view_op=list_works&sortby=pubdate), [ORCID](http://orcid.org/0000-0001-8330-2095)]
# - Affiliasi:
#
# 1. Institut Teknologi Bandung
# 2. Universitas Padjadjaran
#
# - Email: <EMAIL>
# - Description: This document is a manuscript for the SNIPS 2018 ITB
# - Keywords: literate programming, reproducible research
# - Repository: [GitHub/dasaptaerwin](https://github.com/dasaptaerwin/literateprogrammingSNIPS2018)
# # Abstrak
#
# Kode sering diajarkan dalam pembelajaran sains. Selain dapat menuntun alur pikir, kode juga melatih ingatan serta kreativitas. Saat menulis laporan, sering kali kita menggabungkan narasi dengan kode dan luarannya menggunakan teknik salin tempel. Cara ini tidak praktis. *Literate programming* dapat membantu Anda menulis laporan (atau artikel) dengan menggabungkan narasi-kode-luaran secara otomatis. Dalam artikel ini, kami menayangkan aplikasi literate programming menggunakan Bahasa Python dengan Jupyter Notebook untuk melakukan analisis statistik sederhana terhadap data kualitas air tanah di Bandung. Dari hasil yang didapatkan, metode ini dapat digunakan untuk menjelaskan tahapan analisis sejak membuka data, memanipulasi data untuk menyiapkan data, visualisasi, hingga analisisnya secara naratif yang menyatu dengan kode perintah dan luaran prosesnya.
# # Abstract
#
# [translate] Kode sering diajarkan dalam pembelajaran sains. Selain dapat menuntun alur pikir, kode juga melatih ingatan serta kreativitas. Saat menulis laporan, sering kali kita menggabungkan narasi dengan kode dan luarannya menggunakan teknik salin tempel. Cara ini tidak praktis. *Literate programming* dapat membantu Anda menulis laporan (atau artikel) dengan menggabungkan narasi-kode-luaran secara otomatis. Dalam artikel ini, saya menayangkan aplikasi literate programming menggunakan Bahasa Python dengan Jupyter Notebook untuk melakukan analisis statistik sederhana terhadap data kualitas air tanah di Bandung. Dari hasil yang didapatkan, metode ini dapat digunakan untuk menjelaskan tahapan analisis sejak membuka data, memanipulasi data untuk menyiapkan data, visualisasi, hingga analisisnya secara naratif yang menyatu dengan kode perintah dan luaran prosesnya.
# # Pendahuluan
#
# Kode sering diajarkan dalam pembelajaran sains. Selain dapat menuntun alur pikir, kode juga melatih ingatan serta kreativitas ([Filiz, 2015](https://www.sciencedirect.com/science/article/pii/S0747563215004288)). Saat menulis laporan, sering kali kita menggabungkan narasi dengan kode dan luarannya menggunakan teknik salin tempel. Cara ini tidak praktis. *Literate programming* (LP) dapat membantu Anda menulis laporan (atau artikel) dengan menggabungkan narasi-kode-luaran secara otomatis. Tujuan dari artikel ini adalah untuk menjelaskan konsep LP, bagaimana metodenya, serta aplikasinya dengan studi kasus analisis statistik sederhana untuk data kualitas air tanah di Kota Bandung.
# # Sekilas tentang literate programming
#
# _Literate programming_ (LP) dikenalkan oleh <NAME>. Bila dilacak dokumentasinya, maka ide ini pertama kali terbit sebagai makalah dalam jurnal ([Knuth, 1984](https://academic.oup.com/comjnl/article/27/2/97/343244)). Dalam dokumen itu, Knuth menyampaikan bahwa dunia pemrograman secara umum memerlukan suatu cara agar kode program yang sama dapat diulang oleh orang lain, sekarang konsep ini diberi nama *reproducible research*/RR (riset yang dapat diulang). Definisi dari RR dijelaskan dengan sangat baik oleh [ROpensci (2018)](http://ropensci.github.io/reproducibility-guide/sections/introduction/), yakni suatu upaya yang bertujuan agar pihak lain dapat mengulang setiap prosedur riset yang telah kita lakukan. Tidak hanya mengulang, tapi lebih jauh lagi, yakni dapat menggunakan ulang (_reuse_) dan memodifikasinya untuk keperluan lain (_remix_), atau bahkan mengoreksi alur yang kita buat (_contribute_) ([Peng, 2011](http://science.sciencemag.org/content/334/6060/1226)),([Sandve et al., 2013](https://www.ncbi.nlm.nih.gov/pubmed/24204232)).
# # Metode
#
# Dalam artikel ini kami akan menggunakan Jupyter Notebook (JN) sebuah aplikasi LP yang awalnya dikembangkan untuk Bahasa Pemrograman Python. Bagi pengguna Linux dan MacOSX, [Python](https://www.python.org/) adalah bawaan sistem operasi (SO), walaupun demikian, direkomendasikan untuk memeriksa versi Pythonnya dan memperbaruinya. JN berjalan dengan baik pada Python versi 2.7 atau 3.x. Bagi pengguna SO Windows, Anda perlu menginstalasi Python secara terpisah. Distribusi [Continuum Anaconda](https://anaconda.org/) adalah yang kami rekomendasikan untuk SO Linux, MacOSX, dan Windows karena kemudahannya dan kelengkapan panduan instalasinya. Seluruh hasil komputasi di sini tidak dilakukan secara salin-tempel (_copy-paste_) tetapi adalah hasil dari proses pengkodean (_coding_).
#
# Kemudian kami akan membuat analisis statistik sederhana berdasarkan data terbuka kualitas air sumur di Semarang dari penelitian sebelumnya ([Triadi et al. 2016](https://doi.pangaea.de/10.1594/PANGAEA.862987)). Data, kode, dan narasi akan dikombinasikan ke dalam artikel ini secara langsung untuk mendemonstrasikan LP. Analisis statistik yang dilakukan: mendeskripsikan data, membuat histogram untuk melihat distribusi data, dan membuat beberapa grafik x-y untuk melihat beberapa korelasi yang mungkin muncul diantara parameter yang diukur.
# # Contoh aplikasi dalam analisis statistik sederhana
# ## Deskripsi data
#
# Kami akan mendeskripsikan data menggunakan fungsi dalam _library_ `Pandas`. Langkah-langkahnya adalah: (1) memuat `Pandas` ke memori, (2) membuka data menggunakan fungsi `pd.read_csv` dan menyimpannya sebagai `dataframe` bernama `data`, (3) kemudian menampilkannya sebagai tabel (10 baris pertama). Lihat Tabel 1 di bawah ini.
import pandas as pd # langkah 1
data = pd.read_csv('data.csv', sep='\t') # langkah 2
print('Tabel 1 Data')
data.head(n=10) # langkah 3
# Kemudian kita akan membuat grafik histogram parameter zat padat terlarut (_total dissolved solids_, kolom `tds`), kandungan klor (kolom `cl`), kalsium (kolom `ca`) dan elevasi sumur (kolom `elevation`). Untuk membuat grafik, kami menggunakan _library_ `Seaborn` dan memuatnya ke memori dengan nama `sns`.
# %matplotlib inline # perintah untuk memuat grafik langsung ke dalam notebook ini (inline plotting)
import numpy as np # memuat library numpy (numeric python) sebagai np
import seaborn as sns # memuat library seaborn sebagai sns
sns.distplot(data['tds'], kde=False, color='b')
print('Gambar 1 Plot histogram TDS (ppm)')
sns.distplot(data['cl'], kde=False, color='b')
print('Gambar 2 Plot histogram klor (ppm)')
sns.distplot(data['ca'], kde=False, color='b')
print('Gambar 2 Plot histogram kalsium (ppm)')
sns.distplot(data['elevation'], kde=False, color='b')
print('Gambar 3 Plot histogram elevasi posisi sumur (ppm)')
# ## Korelasi
#
# Selanjutnya kami akan mencoba membuat tabel matriks korelasi. Sebelumnya kolom dalam `dataframe` yang berisi teks (string) perlu dikeluarkan dan disimpan sebagai `dataframe` baru bernama `d` dengan fungsi `dataframe.loc()` (lihat Tabel 2). Kemudian kami buat tabel berikutnya berisi beberapa ukuran statistik dasarnya menggunakan fungsi `dataframe.describe()`(lihat Tabel 3). Pada baris berikutnya, kami menayangkan matriks korelasi dengan fungsi `dataframe.corr()` (lihat Tabel 4).
d = data.loc[:,'tds':'hco3']
d.head(n=10)
print('Tabel 2 Dataframe yang baru')
print('Tabel 3 Statistik dasar dataframe d' )
d.describe()
print('Tabel 4 Matriks korelasi dataframe d' )
d.corr()
# Dari tabel 4 di atas, dapat Anda lihat bahwa nilai TDS memiliki korelasi yang kuat dengan kandungan Ca, Mg, Na, Cl, dan HCO3, tapi memiliki korelasi lemah dengan K. Untuk memperlihatkan korelasi tersebut, kami buatkan grafik x-y antara TDS dengan Mg dan TDS dengan K (lihat Gambar 4 dan 5 berikut ini). Kedua plot menggunakan fungsi `sns.regplot()` dengan beberapa kode tambahan untuk pengaturan penamaan sumbu.
ax = sns.regplot(x='tds', y='mg', data=d)
ax.set_xlabel("TDS (ppm)")
ax.set_ylabel("konsentrasi Mg (ppm)")
plt.show()
print('Gambar 4 Plot TDS terhadap Mg')
ax = sns.regplot(x='tds', y='k', data=d)
ax.set_xlabel("TDS (ppm)")
ax.set_ylabel("konsentrasi K (ppm)")
plt.show()
print('Gambar 5 Plot TDS terhadap K')
# # Beberapa catatan
#
# Jika diperhatikan demo di atas, dapat kita lihat bahwa tahapan-tahapan dalam analisis statistik dapat dijelaskan secara naratif, bukan dengan komentar pada baris kode yang pendek-pendek. Biasanya kita memberikan komentar atau penjelasan dengan diawali karakter `#`. Dokumen JN ini, pada waktunya, dapat diekspor sebagai dokumen berformat PDF atau HTML hanya dengan memilih opsi pada menu `File` di atas. Hasilnya adalah satu file PDF atau HTML yang siap tayang berisi perintah kode, luarannya berupa tabel atau grafik, serta penjelasannya. Bahkan makalah ini pun dikonsep secara langsung dalam JN.
#
# Dengan menggunakan JN ini, proses belajar mengajar dapat menjadi lebih mudah. Pengajar hanya perlu memberikan file JN ini, disertai data mentahnya. Para siswa akan menyalin file-file yang diperlukan ke dalam folder kerjanya, maka mereka akan dapat menjalankan perintah dan menghasilkan luaran yang persis sama dengan tayangan pengajar. Piranti lunak yang diperlukanpun sangat fungsional, serta seluruhnya gratis dan _open source_, sehingga dapat menghemat biaya penyelenggaraan pendidikan.
# # Daftar pustaka
#
# 1. <NAME> (1984) Literate Programming, The Computer Journal, Volume 27, Issue 2, Pages 97–111, url: https://doi.org/10.1093/comjnl/27.2.97.
#
# 2. ROpensci (2018) Reproducibility guide, ROpensci blog, url: http://ropensci.github.io/reproducibility-guide/sections/introduction/.
#
# 3. <NAME>. (2011) Reproducible Research in Computational Science, Sciencemag blog, url: http://science.sciencemag.org/content/334/6060/1226.
#
# 4. <NAME>, <NAME>, <NAME>, <NAME>. (2013) Ten simple rules for reproducible computational research, PLOS Computational Biology, url:https://www.ncbi.nlm.nih.gov/pubmed/24204232.
#
# 5. Kalelioglu, F (2015) A new way of teaching programming skills to K-12 students, Computers in Human Behavior, Volume 52, November 2015, Pages 200-210, url: https://www.sciencedirect.com/science/article/pii/S0747563215004288.
#
# 6. Putranto, <NAME>; <NAME>; <NAME> (2016): Hydrochemical properties of groundwater samples in Semarang area, Java Island, Indonesia (1992, 1993, 2003, 2006, and 2007). PANGAEA, https://doi.org/10.1594/PANGAEA.862987.
#
| ms_snips2018_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Low-Level API
# ## Prerequisites
# If you've already completed the instructions on the **Installation** page, then let's get started.
import aiqc
from aiqc import datum
# ## 1. Ingest a `Dataset`
# ### Object Relational Model (ORM)
# 
# The `Dataset` class provides the following subclasses for working with different dimensionalities of data:
#
# - `Dataset.Tabular` - 2D constructed from either an array/dataframe or a tabular file.
# - `Dataset.Sequence` - 3D constructed from an array.
# - `Dataset.Image` - 4D constructed from either an array, a folder of files, or a list of URLs.
# During each stage of both the preprocessing and postprocessing pipelines, these shapes of data need to be handled differently. Additionally, each split and/or fold needs to be processed separately in order to avoid data leakage. You may also find yourself combining heterogenous featuresets (e.g. both tabular data and image data). If you were to derive sliding timeseries windows from a given dataset, then that would add another dimension to the data. All of these transformations needs to be recorded. The reason for this is that when making predictions during inference, the new samples must be processed identically to the original samples that the model was trained upon. Then these predictions must be decoded into their natural forms. AIQC records and coordinates all of this.
# ### a) Registering Datasets
# The `ingest:bool=True` parameter dictates whether or not the data is ingested into SQLite or if remains on disk and is accessed via `source_path`.
#
# By default the actual bytes of the file are persisted to the SQLite `BlobField`. It gets gzip compressed, reducing the size by up to 90%. Maximum BlobField size is 2.147 GB, but once you factor in compression, your bottleneck is more likely to be memory beyond that size. The bytes themselves are Parquet (single-partitioned) because, using the PyArrow engine, it preserves every dtype except certain datetimes (which are honestly better off parsed into floats/ ordinal ints). Parquet is also integrated nicely into both Spark and Dask; frameworks for distributed, in-memory computation.
#
# Persisting the file ensures reproducibility by: (a) keeping the data packaged alongside the experiment, and (b) helping entry-level users move away from relying on mutable dataframes they have had in-memory for extended periods of time or floating around on shared file systems.
#
# > *However, we realize that a different approach will be required at scale, so the `source_path` of the file is recorded whenever possible. In the future we could just read the data from that path (e.g. NFS, RDBMS, HDFS, S3) if the BlobField is none. Or just switch our data fetching/ filtering to Dask because it uses the Pandas API and Parquet.*
# #### `Dataset.Tabular`
# ##### `from_pandas()`
# +
df = datum.to_pandas('iris.tsv')
dataset = aiqc.Dataset.Tabular.from_pandas(
dataframe = df
, name = 'tab separated plants'
, dtype = None #passed to pd.Dataframe(dtype)/ inferred
, column_names = None #passed to pd.Dataframe(columns)
)
# -
# > Optionally, `dtype`, as seen in the `pandas.DataFrame.astype(dtype)` [docs](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.astype.html), can be specified as either a single type for all columns, or as a dictionary that maps a specific type to each column name. This encodes features for analysis. We read NumPy into Pandas before persisting it, so `columns` and `dtype` are read directly by `pd.DataFrame()`.
# ##### `from_numpy()`
# Must be a 2D NumPy N-Dimensional Array.
#
# > *In the future, we may add support for ingesting 3D arrays as multi-file sequences.*
#
# Regular *ndarrays* don't have column names, and I didn't like the API for *structured arrays* so you have to pass in columns names as a list. If you don't then column names will be numerically assigned in ascending order (zero-based index), but I didn't like the range object, so I stringified numerically assigned columns to string-based numbers.
# +
arr = df.to_numpy()
cols = list(df.columns)
other_dataset = aiqc.Dataset.Tabular.from_numpy(
ndarray = arr
, column_names = cols #passed to pd.Dataframe(columns)
)
# -
# ##### `from_path()`
# Intended for flat files, delimited text, and structured tabular data. It's read in via Pandas, so it supports URLs to raw data and bytes as well.
#
# The `file_path` itself can be either absolute or relative.
# +
file_path = datum.get_path('iris_10x.tsv')
# We'll keep this larger dataset handy for `Foldset` creation later.
big_dataset = aiqc.Dataset.Tabular.from_path(
file_path = file_path
, source_file_format = 'tsv'
, name = None
, dtype = None
, column_names = None
, skip_header_rows = 'infer' #passed to `pd.read_csv(header)`. Incompatible w Parquet.
)
# -
# > If you leave `name` blank, it will default to a human-readble timestamp with the appropriate file extension (e.g. '2020_10_13-01_28_13_PM.tsv').
# #### `Dataset.Sequence`
# The sequence dataset is a 3 dimensional structure intended for multi-observations per sample to enable time series analysis.
#
# Observations must be ordered in an *ascending* manner, which means:
#
# - Time-based data must be ordered from earlier to later (2010 - 2020).
# - Position-based data must be ordered from beginning to end (start - stop).
# Sequence datasets are somewhat multi-modal in that, in order to perform supervised learning on them, they require a loosely coupled `Dataset.Tabular` that contains their labels.
df = datum.to_pandas('epilepsy.parquet')
# The `['seizure']` column of this dataframe serves as the Label of that sample. We'll construct a `Dataset.Tabular` from this.
# + tags=[]
label_df = df[['seizure']]
# + tags=[]
dataset_tabular = aiqc.Dataset.Tabular.from_pandas(label_df)
# + tags=[]
tabular_label = dataset_tabular.make_label(columns='seizure')
# -
# ##### `from_numpy()`
# `ndarray3D_or_npyPath:object` requires either an in-memory 3D NumPy array or a *.npy* file. Each 2D array represents a sample.
#
# So we'll drop our seizure column from our DataFrame and reshape it into a 3D array.
seq_ndarray3D = df.drop(columns=['seizure']).to_numpy().reshape(1000,178,1)
dataset_sequence = aiqc.Dataset.Sequence.from_numpy(
ndarray3D_or_npyPath = seq_ndarray3D
, column_names = ['EEG']
)
sequence_feature = dataset_sequence.make_feature()
# Skipping forward a bit, we bring the heterogenous `Feature` and `Label` together in the `Splitset`.
sequence_splitset = aiqc.Splitset.make(
feature_ids = [sequence_feature.id]
, label_id = tabular_label.id
, size_test = 0.24
, size_validation = 0.12
)
# > It is also possible to create a `Foldset` from this splitset.
# #### `Dataset.Image`
# Image datasets are somewhat multi-modal in that, in order to perform supervised learning on them, they require a loosely coupled `Dataset.Tabular` that contains their labels.
df = datum.to_pandas(name='brain_tumor.csv')
df.head()
# The `['status']` column of this dataframe serves as the Label of that sample. We'll construct a `Dataset.Tabular` from this.
tabular_dataset = aiqc.Dataset.Tabular.from_pandas(dataframe=df)
tabular_label = tabular_dataset.make_label(columns=['status'])
# ##### `from_urls_pillow()`
# During ingestion, all image files must have the same `Image.mode` and `Image.size` according to the Pillow library.
#
# > https://pillow.readthedocs.io/en/stable/handbook/concepts.html
# `from_urls(urls:list)` needs a list of urls. In order to perform supervised learning, the order of this list must line up with the samples in your Tabular dataset.
# > We happen to have this list prepared in the `['url']` column of the dataframe above. acts as a manifest in that it contains the URL of the image file for that sample, solely for the purposes of initial ingestion. We'll construct a `Dataset.Image` from this.
image_urls = datum.get_remote_urls(manifest_name='brain_tumor.csv')
image_dataset = aiqc.Dataset.Image.from_urls_pillow(urls=image_urls)
image_feature = image_dataset.make_feature()
# Skipping forward a bit, we bring the heterogenous `Feature` and `Label` together in the `Splitset`.
image_splitset = aiqc.Splitset.make(
feature_ids = [image_feature.id]
, label_id = tabular_label.id
, size_test = 0.24
, size_validation = 0.12
)
# > It is also possible to create a `Foldset` from this splitset.
# ##### `from_folder_pillow()`
# When reading images from a locally accessible folder, the fantastic `natsort.natsorted` library is used as the source of truth for the order of the files.
# > Python reads files by insertion order rather than alpha-numerically, which isn't intuitive for humans. So make sure your tabular manifest has the same order as `natsorted`. https://natsort.readthedocs.io/en/master/api.html#natsort.natsorted
image_dataset = aiqc.Dataset.Image.from_folder_pillow("/Users/layne/desktop/MRI_scans")
# Here you can see the first 3 files that comprise that dataset.
image_dataset.files[:3]
image_feature = image_dataset.make_feature()
image_splitset = aiqc.Splitset.make(
feature_ids = [image_feature.id]
, label_id = tabular_label.id
, size_test = 0.24
, size_validation = 0.12
)
# ##### `from_numpy()`
# The `ndarray4D_or_npyPath:object` requires either an in-memory 4D NumPy array or a *.npy* file. Each 3D sequence represents an image sample.
# ```python
# Dataset.Image.from_numpy(
# ndarray4D_or_npyPath:object
# , name:str = None
# , dtype:object = None
# , column_names:list = None
# , ingest:bool = True
# )
# ```
# ### b) Reading Datasets into Memory
# All of the sample-related objects in the API have `to_numpy()` and `to_pandas()` methods that accept the following arguments:
#
# * `samples=[]` list of indices to fetch.
# * `columns=[]` list of columns to fetch.
# * In some cases you can specify a `split`/ `fold` name.
#
# For structured data, since the `Dataset` itself is fairly removed from the `File.Tabular` it creates, you can get that tabular file with `Dataset.Tabular.get_main_tabular(dataset_id)` to inspect attributes like `dtypes` and `columns`.
# Later, we'll see how these arguments allow downstream objects like `Splitset` and `Foldset` to slice up the data.
# #### `Dataset.Tabular`
# ##### `to_pandas()`
df = dataset.to_pandas()
df.head()
df = aiqc.Dataset.to_pandas(
id = dataset.id
, samples = [0,13,29,79]
, columns = ['sepal_length', 'sepal_width']
)
df.tail()
# ##### `to_numpy()`
arr = dataset.to_numpy(
samples = [0,13,29,79]
, columns = ['petal_length', 'petal_width']
)
arr[:4]
arr = aiqc.Dataset.to_numpy(id=dataset.id)
arr[:4]
# #### `Dataset.Sequence`
arr = dataset.to_numpy(
samples = [0]
, columns = None
)
arr[0][:5]
# ##### `to_numpy()`
# This essentially internal method only exists to enable Pandas-related preprocessing such as interpolation. Produces a list of dataframes.
# ##### `to_pandas()`
# This essentially internal method only exists to enable Pandas-related preprocessing such as interpolation. Produces a list of dataframes.
# ##### `to_pillow()`
# This essentially internal method only exists to fetch images in their natural form (e.g. PNG/JPG).
# #### `Dataset.Image`
# ##### `to_pillow()`
# Returns a list of `PIL.Image`'s. You can actually see the image when you call them.
images_pillow = aiqc.Dataset.Image.to_pillow(id=image_dataset.id, samples=[60,61,62])
images_pillow[1]
# ##### `to_numpy()`
# This simply performs `np.array(Pillow.Image)`. Returns an N-dimensional array where the dimensions vary based on the `mode` aka colorscale of the image. For example, it returns '3D of 2Ds for black and white' or '4D of 3Ds for colored' - which would change the class of convultional layer you would use (`Conv1D`:`Conv3D`).
#
#
# > By default, the returned arrays will be scaled by 255 because [Pillow pixel values range from 0-255](https://pillow.readthedocs.io/en/stable/handbook/concepts.html#modes).
# Let's check out one of the 2D color channels of a grayscale image.
images_pillow = aiqc.Dataset.Image.to_numpy(id=image_dataset.id, samples=[60,61,62])
images_pillow[1]
# ## 2. Selecting `Features` and `Labels`.
# ### a) Select the `Label` column(s).
# #### ORM
# From a Dataset, pick the column(s) that you want to predict/ train against. Creating a `Label` won't duplicate your data! It simply marks the Dataset `columns` to be used for supervised learning.
# Later, we'll see that a `Label` triggers:
#
# * The `supervision` attribute of a `Splitset` to be either `'unsupervised'`/`'supervised'`.
#
# * Approval/ rejection of the `Algorithm.analysis_type`. For example, you wouldn't perform regression on a string label.
# Part of the magic of this library is that it prevents you from making silly mistakes like these so that you aren't faced with some obscure NumPy/ Tensor, dtype/ dimensionality error on the nth layer of your neural network.
# For categorical labels, but not for continuous/float labels, the `Label.unique_classes` are recorded.
# #### Deriving Labels
# Keep the name of the label column handy as you may want to re-use it later when excluding features.
label_column = 'species'
# Implicit IDs
label = dataset.make_label(columns=[label_column])
# > `columns=[label_column]` is a list in case users have already OneHotEncoded (OHEd) their label. If multiple columns are provided, then they must already be in OHE format. I'm not keen on supporting multi-label/ simultaneous analysis, but that could changed based on feasibility and user demand.
# Explicit IDs
other_label = aiqc.Label.from_dataset(
dataset_id=other_dataset.id
, columns=[label_column]
)
# #### Reading Labels
# The `Label` comes in handy when we need to fetch what is traditionally referred to as '*Y*' in tutorials. It also accepts a `samples` argument, so that `Splitset` can subset it.
label.to_pandas().tail()
label.to_numpy(samples=[0,33,66,99,132])[:5]
# ### b) Select the `Feature` column(s).
# #### ORM
# Creating a Feature won't duplicate your data! It simply records the Dataset `columns` to be used as features during training.
#
# There are three ways to define which columns you want to use as features:
#
# - `exclude_columns=[]` for ruling out columns like the `Label` column.
# - `include_columns=[]` for being selective.
# - Leave both of the above blank and all columns will be used (e.g. images or unsupervised leanring).
# For structured data, since the Feature is far removed from the `File.Tabular` that it is derived from, there is a `Feature.get_dtypes()` method. This will come in handy when we are selecting dtypes/columns to include/ exclude in our `Featurecoder`(s).
# #### Specifying Features
# Via `include_columns=[]`
feature = dataset.make_feature(
include_columns = [
'sepal_length',
'petal_length',
'petal_width'
]
)
# Or via `exclude_columns=[]`
feature = dataset.make_feature(exclude_columns=[label_column])
feature.columns
# Either way, any excluded columns will be recorded since they are used for dropping.
feature.columns_excluded
# Again, for images, just perform `Dataset.Image.make_feature()` since you'll likely want to include all pixels and your label column is in a separate, coupled Dataset.
# #### Reading Features
feature.to_numpy()[:4]
feature.to_pandas(samples=[0,16,32,64]).tail()
# ## 3. Slicing samples.
# ### a) Define sliding time series `Windows`.
# #### ORM
# The `Window` object is provided in order to facilitate sliding windows for unsupervised/ self-supervised time series forecasting and backcasting. It assumes that the last time point is the most recent (aka ascending time).
# 
# **Concept of a sliding sample:**
# The 'sample' refers to the highest dimension of the data that is batched just before training.
# - `Dataset.Tabular` - The 'sample' is no longer a single image, but rather a group of images known as a window. Looking at the diagram above, there are 5 samples (*0:4* windows), not 15 samples (*0:14* time points).
# - `Dataset.Sequence` - The 'sample' is an array consisting of 1 window from each sequence. Imagine monitoring the weather of several cities at once, and taking the same window from each city as a sample.
# - `Dataset.Image` - The 'sample' is no longer a single image, but rather a group of images known as a window.
# **Parameters:**
# - `size_window:int` the number of points in time to include in a window.
# - `size_shift:int` the number of points in time to slide forward.
# - `record_shift:bool=True` used during inference. Only persists unshifted windows and while leaving shifted windows as `None`.
# #### Specifying Windows
# `window = feature.make_window(size_window=4, size_shift=2)`
# - `samples_unshifted:list of lists`
# - `samples_shifted:list of lists`
# #### Reading Windows
# In the example above, `window.samples_unshifted[0]==[1,2,3,4]`. Where *1:4* represents the raw, underlying sample indices of that window. This is used when fetching the data. First, the entire `Feature` is fetched for preprocessing, then windows are copied out of it like so:
#
# - `np.array([feature_array[w] for w in window.samples_unshifted])`
# - `np.array([feature_array[w] for w in window.samples_shifted])`
#
# Where each of the above arrays is a 3D sequence that is fed into a single recurrent model. The shifted sequence is slotted into `samples[<split>]['labels']` to facilitate self-supervision.
# ### b) Slice samples with a `Splitset`.
# A `Splitset` divides the samples of the Dataset into the following *splits* in the table below. It is the central object of the data preparation side of the ORM in that it touches `Label`, `Feature`, `Foldset`, and `Encoderset`. It is the only mandatory data preparation object required by the training `Queue`.
# Both continuous and categorical `Labels` are automatically stratified.
# | Split | Description |
# |-----------------------|---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
# | train | The samples that the model will be trained upon. <br/>Later, we’ll see how we can make *cross-folds from our training split*. <br/>Unsupervised learning will only have a training split. |
# | validation (optional) | The samples used for training evaluation. <br/>Ensures that the test set is not revealed to the model during training. |
# | test (optional) | The samples the model has never seen during training. <br/>Used to assess how well the model will perform on unobserved, natural data when it is applied in the real world aka how generalizable it is. |
# > Label-based stratification is used to ensure equally distributed label classes for both categorical and continuous data.
# >
# > If you want more control over stratification of continuous splits, specify the number of `bin_count:int` for grouping.
# > Again, creating a Splitset won't duplicate your data. It simply points to the sample indices to be used in the splits that you specify!
# #### Stratification
# All tabular splits are stratified by default in that they contain similar distributions of unique Label classes so that each split is a statistically accurate representation of the population as a whole.
#
# In order to support this process for continuous labels, binning/ discretization is utilized. For example, if 4 bins are used, values from *0.0 to 1.0* would be binned as *[0.0-0.25, 0.25-0.50, 0.50-0.75, 0.75-1.0]*. This is controlled by the `bin_count:int` argument.
#
# > Reference the handy `Pandas.qcut()` and the source code `pd.qcut(x=array_to_bin, q=bin_count, labels=False, duplicates='drop')` for more detail.
# Additionally `unsupervised_stratify_column:str` is provided for scenarios where there is no Label. For example, you may want to stratify by the month during unsupervised time series analysis.
# #### Splitting Strategies
# Notice that the `feature_ids:list` is plural. That's because we can pass a Splitset multiple Features for mixed-data analysis (e.g. using both Tabular and Image data).
#
# > If you have a feature that is capable of stratification (e.g. `dataset_type='tabular'` or `dataset_type='tabular'`), then you should make that `feature_id` the first element in the list. Only the first list element will be checked for the purposes of stratification.
# ##### i) Default supervised 70-30 split.
# If a Label is provided, then a 70:30 train:test splits will automatically be generated.
splitset = aiqc.Splitset.make(
feature_ids = [feature.id]
, label_id=label.id
)
# ##### ii) Specifying test size.
splitset = aiqc.Splitset.make(
feature_ids = [feature.id]
,label_id = label.id
, size_test = 0.22
)
# ##### iii) Specifying validation size.
# `size_validation` cannot be specified without a `size_test`.
splitset = aiqc.Splitset.make(
feature_ids = [feature.id]
, label_id = label.id
, size_test = 0.20
, size_validation = 0.12
)
# ##### iv) Unsupervised
# ```
# splitset = aiqc.Splitset.make(
# label_id = None
# , feature_ids = [feature.id]
# , size_test=0.12
# , size_validation=0.16
# )
# ```
# ##### v) Take all samples for inference
# ```
# splitset = aiqc.Splitset.make(
# feature_ids = [feature.id]
# , label_id = None # Optional for unsupervised and pure inference.
# )
# ```
# + [markdown] toc-hr-collapsed=true
# #### Reading Splitsets
# -
splitset.samples.keys()
# `.keys()` of 1st layer are referred to as "split_name" in the source code: e.g. 'train' as well as, optionally, 'validation' and 'test'.
#
# `Splitset.samples` on disk:
# ```
# {
# 'train': [<sample_indices>],
# 'validation': [<sample_indices>],
# 'test': [<sample_indices>]
# }
# ```
# You can also verify the actual size of your splits.
splitset.sizes
# The main attribute of the splitset is the `samples` dictionary. Again, on-disk this only contains sample indices. The dictionary is structured like so:
# ### c) Create a `Foldset` for cross-validation.
# #### ORM
# *Reference the [scikit-learn documentation](https://scikit-learn.org/stable/modules/cross_validation.html) to learn more about folding.*
#
# 
# We refer to the left out fold (blue) as the `fold_validation` and the remaining training data as the `folds_train_combined` (green).
# > *In the future, we may introduce more folding `strategies` aside from leave-one-out.*
# #### `Fold` objects
# For the sake of determining which samples get trained upon, the only thing that matters is the slice of data that gets left out.
# > Tip - DO NOT use a `Foldset` unless your *(total sample count / fold_count)* still gives you an accurate representation of your sample population. If you are ignoring that advice and stretching to perform cross-validation, then at least ensure that *(total sample count / fold_count)* is evenly divisible. Both of these tips help avoid poorly stratified/ undersized folds that perform either too well (only most common label class present) or poorly (handful of samples and a few inaccurate prediction on a normally good model).
# >
# > Tip - The sample indices of the validation fold are not discarded. In fact, `fold_validation` can actually be used alongside a split `validation` for double validation 🤘. However, it's more sensible to skip the validation split when cross-validating because you'll want each `fold_validation` to be as large (representative of the population) as possible. Folds naturally have fewer samples, so a handful of incorrect predictions have the potential to offset your aggregate metrics.
# >
# > Candidly, if you've ever performed cross-validation manually, let alone systematically, you'll know that, barring stratification of continuous labels, it's easy enough to construct the folds, but then it's a pain to generate performance metrics (e.g. `zero_division`, absent OHE classes) due to the absence of outlying classes and bins. Time has been invested to handle these scenarios elegantly so that folds can be treated as first-class-citizens alongside splits. That being said, if you try to do something undersized like "150 samples in their dataset and a `fold_count` > 3 with `unique_classes` > 4," then you may run into edge cases.
# Similar to `Splitset.samples`, there is a `Fold.samples` dictionary of sample indices with the following `.keys()`:
# * `samples['folds_train_combined']` - all the included folds.
# * `samples['fold_validation']` - the fold that got left out.
# 
# #### Specifying Foldsets
# We'll catch out big dataset up to the point where we can make a Foldset with it.
big_label = big_dataset.make_label(columns=[label_column])
big_fset = big_dataset.make_feature(exclude_columns=[label_column])
big_splits = aiqc.Splitset.make(
feature_ids = [big_fset.id]
, label_id = big_label.id
, size_test = 0.30
, bin_count=3
)
# Now we are ready to generate 5 `Fold` objects that belong to the `Foldset`.
foldset = big_splits.make_foldset(fold_count=5, bin_count=3)
list(foldset.folds)
# #### Reading Foldsets
# Sample indices of each Fold
foldset.folds[0].samples['folds_train_combined'][:10]
foldset.folds[0].samples['fold_validation'][:10]
# ## 4. Preprocessing - Interpolation.
# ### ORM
# If you have columns with missing data in a time series, then interpolation allows you to fill in those blanks mathematically. It does so by fitting a curve to each column. If you don't have time series data then you don't need interpolation.
# > `pandas.DataFrame.interpolate`
# >
# > https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.interpolate.html
# >
# > Is utilized due to its ease of use, variety of methods, and **support of sparse indices**. However, it does not follow the `fit/transform` pattern like many of the class-based sklearn preprocessors, so the interpolated training data is concatenated with the evalaution split during the interpolation of evaluation splits.
# The `interpolate_kwargs:dict=None` object is what gets passed to Pandas interpolation. In my experience, `method=spline` produces the best results. However, sometimes either spline cannot fit your data or you know that your pattern is linear. For those scenarios there's `method=linear`.
#
# Here the default argument that will ultimately be used in `df.interpolate(**interpolate_kwargs)` if `interpolate_kwargs=None`.
# ```python
# interpolate_kwargs = dict(
# method = 'spline'
# , limit_direction = 'both'
# , limit_area = None
# , axis = 0
# , order = 1
# )
# ```
# ### a) Interpolate Labels with `Labelpolater`.
# Due to the fact that Labels cannot be windowed, Labelpolater simply fills in the gaps in a sequential progression.
# In scenarios where evaluation splits may not have enough data to be interpretted separately, there is the `process_separately:bool=False` argument.
# ```python
# label.make_labelpolater(
# process_separately=True
# , interpolate_kwargs=dict(
# method='spline'
# , limit_direction='both'
# , limit_area=None
# , axis=0
# , order=1
# )
# )
# ```
# ### b) Interpolate Features with `Interpolaterset`.
# 
# The fact that time series data is windowed presents challenges to preprocessing the training and evaluation splits/ folds separately in order to avoid leakage:
#
# - For 3D `Dataset.Sequence` interpolation is simply ran separately on each 2D array.
# - However, in 2D `Dataset.Tabular` different windows belong to different splits/ folds so their underlying groups of rows must be interpolated separately.
# Additionally, there can be several columns/ dtypes in the data that have completely different patterns/ curves to fit. Thus we need a chain of `Featurepolaters` that is represented by `Interpolaterset`.
# ```python
# interpolaterset = feature.make_interpolaterset()
#
# interpolaterset.make_featurepolater(
# columns = ['nox']
# , interpolate_kwargs = dict(
# method='linear'
# , limit_direction='both'
# , limit_area=None
# , axis=0
# , order=1
# )
# )
#
# interpolaterset.make_featurepolater(
# dtypes = ['float64']
# , interpolate_kwargs = dict(
# method='spline'
# , limit_direction='both'
# , limit_area=None
# , axis=0
# , order=1
# )
# )
# ```
# ## 5. Preprocessing - Encoding.
# ### Encoding
# Certain algorithms either (a) require features and/ or labels formatted a certain way, or (b) perform significantly better when their values are normalized. For example:
#
# * Scaling continuous features from (-1 to 1) or (0.0 to 1.0). Or transforming them to resemble a more Gaussian distribution.
# * Converting ordinal or categorical string data `[dog, cat, fish]` into one-hot encoded format `[[1,0,0][0,1,0][0,0,1]]`.
#
# There are two phases of encoding:
# 1. `fit` - where the encoder learns about the values of the samples made available to it. Ideally, you only want to `fit` aka learn from your training split so that you are not *"leaking"* information from your validation and test spits into your encoder!
# 2. `transform` - where the encoder transforms all of the samples in the population.
#
# AIQC has solved the following challenges related to encoding:
#
# * How does one dynamically `fit` on only the training samples in advanced scenarios like cross-validation where a different fold is used for validation each time?
#
# * For certain encoders, especially categorical ones, there is arguably no leakage. If an encoder is arbitrarilly assigning values/ tags to a sample through a process that is not aggregate-informed, then the information that is reveal to the `fit` is largely irrelevant. As an analogy, if we are examining swan color and all of a sudden there is a black swan... it's clearly not white, so slap a non-white label on it and move on. In fact, the prediction process and performance metric calucatlion may fail if it doesn't know how to handle the previously unseen category.
#
# * Certain encoders only accept certain dtypes. Certain encoders only accept certain dimensionality (e.g. 1D, 2D, 3D) or shape patterns (odd-by-odd square). Unfortunately, there is not much uniformity here.
#
# * Certain encoders output extraneous objects that don't work with deep learning libraries.
#
# > *For now, only `sklearn.preprocessing` methods are supported. That may change as we add support for more low-level tensor-based frameworks like PyTorch.*
# ### a) Encode labels with `Labelcoder`.
# The simplistic `Labelcoder` is a good warmup for the more advanced `Featurecoder`.
# Of course, you cannot encode Labels if your `Splitset` does not have labels in the first place.
# The process is straightforward. You provide an instantiated encoder [e.g. `StandardScaler()` not `StandardScaler`], and then AIQC will:
#
# * Verify that the encoder works with your `Label`'s dtype, sample values, and figure out what dimensionality it needs in order to succeed.
#
# * Automatically correct the attributes of your encoder to smooth out any common errors they would cause. For example, preventing the output of a sparse scipy matrix.
#
# * Determine whether the encoder should be `fit` either (a) exclusively on the train split, or (b) if it is not prone to leakage, inclusively on the entire dataset thereby reducing the chance of errors arising.
# #### Creating a `Labelcoder`
# AIQC only supports the uppercase `sklearn.preprocessing` methods (e.g. `RobustScaler`, but not `robust_scale`) because the lowercase methods do not separate the `fit` and `transform` steps. FYI, most of the uppercase methods have a combined `fit_transform` method if you need them.
#
# > https://scikit-learn.org/stable/modules/classes.html#module-sklearn.preprocessing
from sklearn.preprocessing import *
labelcoder = label.make_labelcoder(
sklearn_preprocess = OneHotEncoder(sparse=False)
)
# The following method is used behind the scenes to fetch the most recently create Labelcoder for your Label when it comes time to encode data during training.
label.get_latest_labelcoder()
# ### b) Encode Features sequentially with `Ensoderset`.
# The `Featurecoder` has the same validation process as the `Labelcoder`. However, it is not without its own challenges:
#
# * We want to be able to apply different encoders to columns of different dtypes.
#
# * Additionally, even within the same dtype (e.g. float/ continuous), different distributions call for different encoders.
#
# * Commonly used encoders such a `OneHotEncoder` can ouput multiple columns from a single column input. Therefore, the *shape* of the features can change during encoding.
#
# * And finally, throughout this entire process, we need to avoid data leakage.
#
# For these reasons, `Featurecoder`'s are applied sequentially; in an ordered chain, one after the other. After an encoder is applied, its columns are removed from the raw feature and placed into an intermediary cache specific to each split/ fold.
# > Right now, `Featurecoder` cannot be created for `Dataset.Image.Feature`. I'm not opposed to changing this, but I would just have to account for 3D arrays.
encoderset = feature.make_encoderset()
# #### Filtering feature columns
# The filtering mode is either:
#
# * Inclusive (`include=True`) encode columns that match the filter.
#
# * Exclusive (`include=False`) encode columns outside of the filter.
#
# Then you can select:
#
# 1. An optional list of `dtypes`.
#
# 2. An optional list of `columns` name.
#
# * The column filter is applied after the dtype filter.
#
# > You can create a filter for all columns by setting `include=False` and then seting both `dtypes` and `columns` to `None`.
# After submitting your encoder, if `verbose=True` is enabled:
# * The validation rules help determine why it may have failed.
# * The print statements help determine which columns your current filter matched, and which raw columns remain.
featurecoder = encoderset.make_featurecoder(
sklearn_preprocess = PowerTransformer(method='yeo-johnson', copy=False)
, include = True
, dtypes = ['float64']
, columns = None
, verbose = True
)
# You can also view this information via the following attributes: `matching_columns`, `leftover_dtypes`, and `leftover_columns`.
# ## 6. Preprocessing - Reshaping.
# ### ORM
# When working with architectures that are highly dimensional such convolutional and recurrent networks (Conv1D, Conv2D, Conv3D / ConvLSTM1D, ConvLSTM2D, ConvLSTM3D), you'll often find yourself needing to reshape data to fit a layer's required input shape.
#
# AIQC ingestion & preprocessing favors a *"channels_first"* (samples, channels, rows, columns) approach as opposed to *"channels_last"* (samples, rows, columns, channels).
#
# - *Reducing unused dimensions* - When working with grayscale/ single channel images (1 channel, 25 rows, 25 columns) there is no sense using Conv2D just to handle that 1 channel.
# - *Adding wrapper dimensions* - Perhaps your data is a fit for ConvLSTM1D, but that layer is only supported in the nightly TensorFlow build so you want to add a wrapper dimension in order to use the production-ready ConvLSTM2D.
# It is difficult do this on the fly during training (aka after the fact) because you need to: add reshaping layers/ views to your model, intercept and reshape the data in your post-processing functions, and, by this point, the data is in a variety of tensor formats. It's also more efficient to do this wrangling once up front rather than repeatedly on every training run.
# The `reshape_indices` argument accepts a tuple for rearranging indices in your order of choosing. Behind the scenes, it will use `np.reshape()` to rearrange the data at the end of your preprocessing pipeline. How the element is handled in that tuple is determined by its type.
# `feature.make_featureshaper(reshape_indices:tuple)`
# ```python
# # source code from the end of `feature.preprocess()`
# current_shape = feature_array.shape
#
# new shape = []
# for i in featureshaper.reshape_indices:
# if (type(i) == int):
# new_shape.append(current_shape[i])
# elif (type(i) == str):
# new_shape.append(int(i))
# elif (type(i)== tuple):
# indices = [current_shape[idx] for idx in i]
# new_shape.append(math.prod(indices))
# new_shape = tuple(new_shape)
#
# feature_array = feature_array.reshape(new_shape)
# ```
# *Warning:* if your model is unsupervised (aka generative or self-supervised), then it must output data in *"column (aka width) last"* shape. Otherwise, automated column decoding will be applied along the wrong dimension.
# ### Reshaping by Index
# Let's say we have a 4D feature consisting of 3D images `(samples * color channels * rows * columns)`. Our image is B&W, so we want to get rid of the single color channel. So we want to drop the dimension at the shape index `1`.
# ```python
# reshape_indices = (0,2,3)
# ```
# Thus we have wrangled ourselves a 3D feature consisting of 2D images `(samples * rows * columns)`.
# ### Reshaping Explicitly
# But what if the dimensions we want cannot be expressed by rearranging the existing indices? You might have been wondering why `str` appeared in the loop above. If you define a string-based number, then that number will be used as directly as the value at that position.
# So if I wanted to add an extra wrapper dimension to my data to serve as a single color channel, I would simply do:
# ```python
# reshape_indices = (0,'1',1,2)
# ```
# ### Multiplicative Reshaping
# Sometimes you need to stack/nest dimensions. This requires multiplying one shape index by another. For example, if I have a 3 separate hours worth of data and I want to treat it as 180 minutes, then I need to go from a shape of (3 hours * 60 minutes) to (180 minutes). Just provide the shape indices that you want to multiply in a `tuple` like so:
# ```python
# reshape_indices = ((0,1), 2)
# ```
# ## 7. Defining Architectures & Hyperparameters.
# ### a) Define an `Algorithm`
# Now that our data has been prepared, we transition to the other half of the ORM where the focus is the logic that will be applied to that data.
# > An `Algorithm` is our ORM's codename for a machine learning model since *Model* is the most important *reserved word* when it comes to ORMs.
# The following attributes tell AIQC how to handle the Algorithm behind the scenes:
#
# * `library` - right now, only 'keras' is supported.
#
# * Each library's model object and callbacks (history, early stopping) need to be handled differently.
#
#
# * `analysis_type` - right now, these types are supported:
#
# * `'classification_multi'`, `'classification_binary'`, `'regression'`.
#
# * Used to determine which performance metrics to run.
#
# * Must be compatible with the type of label fed to it.
# #### Model Definition
# The `Algorithm` is composed of the functions:
#
# * `fn_build`.
#
# * `fn_lose` (optional, inferred).
#
# * `fn_optimize` (optional, inferred).
#
# * `fn_train`.
#
# * `fn_predict` (optional, inferred).
#
# > May provide overridable defaults for build and train in the future.
# You can name the functions whatever you want, but do not change the predetermined arguments (e.g. `input_shape`,`**hp`, `model`, etc.) or their position.
#
# As we define these functions, we'll see that we can pass a dictionary of *hyperparameters* into these function using the `**hp` kwarg, and access them like so: `hp['<some_variable_name>']`. Later, we'll provide a list of values for each entry in the hyperparameters dictionary.
# Let's import the modules that we need.
import keras
from keras import metrics
from keras.models import Sequential
from keras.callbacks import History
from keras.layers import Dense, Dropout
# > Later, when running your `Job`'s, if you receive a "module not found" error, then you can try troubleshooting by importing that module directly within the function where it is used.
# ##### Function to build model
# You can build your topology however you like, just be sure to `return model`. Also, you don't have to use any of the hyperparameters (`**hp`) if you don't want to.
# The automatically provided `features_shape` and `label_shape` are handy because:
#
# * The number of feature/ label columns is mutable due to encoders (e.g. OHE).
#
# * Shapes can be less obvious in multi-dimensional scenarios like colored images.
# > You can customize the metrics if you so desire (e.g. change the loss or accuracy), but they will only be applied to the training process/ `History` callback. We'll see later that AIQC will calculate metrics for you automatically.
def fn_build(features_shape, label_shape, **hp):
model = Sequential()
model.add(Dense(units=hp['neuron_count'], input_shape=features_shape, activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.2))
model.add(Dense(units=hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(units=label_shape[0], activation='softmax'))
#optimizer and loss defined separately.
return model
# ##### Optional, function to calculate loss.
# We can't just specify the loss function in our training loop because we will need it later on when it comes time to produce metrics about other splits/ folds.
#
# If you do not provide an `fn_lose` then one will be automatically selected for you based on the `Algorithm.analysis_type` you are conducting and the `Algorithm.library` you are using.
def fn_lose(**hp):
loser = keras.losses.CategoricalCrossentropy()
return loser
# ##### Optional, function to optimize model.
# Some deep learning libraries persist their model and optimizer separately during checkpoint/exporting. So `fn_optimize` provides an isolated way to access the optimizer. It also allows us to automatically set the optimizer.
#
# If you do not provide an `fn_optimize` then one will be automatically selected for you based on the `Algorithm.analysis_type` you are conducting and the `Algorithm.library` you are using.
def fn_optimize(**hp):
optimizer = keras.optimizers.Adamax(learning_rate=0.01)
return optimizer
# > If you want to define your own optimizer, then you should do so within this function, rather than relying on `model.compile(optimizer='<some_optimizer_name>'`. If you do not define an optimizer, then `Adamax` will be used by default.
# ##### Function to train model
# * `samples_train` - the appropriate data will be fed into the training cycle. For example, `Foldset.samples[fold_index]['folds_train_combined']` or `Splitset.samples['train']`.
#
# * `samples_evaluate` - the appropriate data is made available for evaluation. For example, `Foldset.samples[fold_index]['fold_validation']`, `Splitset.samples['validation']`, or `Splitset.samples['test']`.
def fn_train(model, loser, optimizer, samples_train, samples_evaluate, **hp):
model.compile(
loss = loser
, optimizer = optimizer
, metrics = ['accuracy']
)
model.fit(
samples_train["features"]
, samples_train["labels"]
, validation_data = (
samples_evaluate["features"]
, samples_evaluate["labels"]
)
, verbose = 0
, batch_size = 3
, epochs = hp['epoch_count']
, callbacks=[History()]
)
return model
# ##### Optional, callback to stop training early.
# *Early stopping* isn't just about efficiency in reducing the number of `epochs`. If you've specified 300 epochs, there's a chance your model catches on to the underlying patterns early, say around 75-125 epochs. At this point, there's also good chance what it learns in the remaining epochs will cause it to overfit on patterns that are specific to the training data, and thereby and lose it's simplicity/ generalizability.
#
# > The `val_` prefix refers to the evaluation samples.
# >
# > Remember, regression does not have accuracy metrics.
# >
# > `TrainingCallback.Keras.MetricCutoff` is a custom class we wrote to make multi-metric cutoffs easier, so you won't find information about it in the official Keras documentation.
def fn_train(model, loser, optimizer, samples_train, samples_evaluate, **hp):
model.compile(
loss = loser
, optimizer = optimizer
, metrics = ['accuracy']
)
#Define one or more metrics to monitor.
metrics_cuttoffs = [
{"metric":"val_accuracy", "cutoff":0.96, "above_or_below":"above"},
{"metric":"val_loss", "cutoff":0.1, "above_or_below":"below"}
]
cutoffs = aiqc.TrainingCallback.Keras.MetricCutoff(metrics_cuttoffs)
# Remember to append `cutoffs` to the list of callbacks.
callbacks=[History(), cutoffs]
# No changes here.
model.fit(
samples_train["features"]
, samples_train["labels"]
, validation_data = (
samples_evaluate["features"]
, samples_evaluate["labels"]
)
, verbose = 0
, batch_size = 3
, epochs = hp['epoch_count']
, callbacks = callbacks
)
return model
# ##### Optional, function to predict samples
# `fn_predict` will be generated for you automatically if set to `None`. The `analysis_type` and `library` of the Algorithm help determine how to handle the predictions.
# i) Regression default.
def fn_predict(model, samples_predict):
predictions = model.predict(samples_predict['features'])
return predictions
# ii) Classification binary default.
# All classification `predictions`, both mutliclass and binary, must be returned in ordinal format.
#
# > For most libraries, classification algorithms output *probabilities* as opposed to actual predictions when running `model.predict()`. We want to return both of these object `predictions, probabilities` (the order matters) to generate performance metrics behind the scenes.
def fn_predict(model, samples_predict):
probabilities = model.predict(samples_predict['features'])
# This is the official keras replacement for binary classes `.predict_classes()`
# It returns one array per sample: `[[0][1][0][1]]`
predictions = (probabilities > 0.5).astype("int32")
return predictions, probabilities
# iii) Classification multiclass default.
def fn_predict(model, samples_predict):
import numpy as np
probabilities = model.predict(samples_predict['features'])
# This is the official keras replacement for multiclass `.predict_classes()`
# It returns one ordinal array per sample: `[[0][2][1][2]]`
predictions = np.argmax(probabilities, axis=-1)
return predictions, probabilities
# #### Group the functions together in an `Algorithm`!
algorithm = aiqc.Algorithm.make(
library = "keras"
, analysis_type = "classification_multi"
, fn_build = fn_build
, fn_train = fn_train
, fn_optimize = fn_optimize # Optional
, fn_predict = fn_predict # Optional
, fn_lose = fn_lose # Optional
)
# > <!> Remember to use `make` and not `create`. Deceptively, `create` runs because it is a standard, built-in ORM method. However, it does so without any validation logic.
# ### b) Combinations of hyperparameters with `Hyperparamset`.
# Parameters are fed into Algorithm functions.
#
# The `hyperparameters` below will be automatically fed into the functions above as `**kwargs` via the `**hp` argument we saw earlier.
#
# For example, wherever you see `hp['epoch_count']`, it will pull from the *key:value* pair `"epoch_count": [30, 60]` seen below. Where "model A" would have 30 epochs and "model B" would have 60 epochs.
hyperparameters = {
"neuron_count": [12]
, "epoch_count": [30, 60]
, "learning_rate": [0.01, 0.03]
}
# #### Hyperparameter Selection Strategies.
# ##### Grid search strategy.
#
# By default AIQC will generate all possible combinations.
#
# > With enough practice, practitioners will get a feel for what parameters and topologies make sense so you'll rely on shotgun-style approaches less and less. If you limit your experiments to 1-2 parameters at a time then it's easy to see their effect as an *independent variable*. You should really start with high-level things such as topologies (# of layers, # neurons per layer) and batch size before moving on to tuning the intra-layer nuances (activation methods, weight initialization). You're essentially testing high/ medium/ low or default/ edge case scenarios for each parameter.
# ##### Random selection strategy.
#
# Testing many different combinations in your initial runs can be a good way to get a feel for the parameter space. Although if you are doing this you'll find that many of your combinations are a bit too similar. So randomly sampling (with replacement) a few of them is a less computationally expensive way to go about this.
#
# * `pick_count:int` the fixed # of combinations to sample.
#
# * `pick_percent:float` a % of combinations to sample.
# ##### Bayesian selection strategy.
#
# "TPE (Tree-structured Parzen Estimator)" via `hyperopt` has been suggested as a future area to explore.
hyperparamset = aiqc.Hyperparamset.from_algorithm(
algorithm_id = algorithm.id
, hyperparameters = hyperparameters
, pick_count = None
, pick_percent = None
)
# #### `Hyperparamcombo` objects.
# Each unique combination of hyperparameters is recorded as a `Hyperparamcombo`.
#
# Ultimately, a training `Job` is constructed for each unique combinanation of hyperparameters aka `Hyperparamcombo`.
hyperparamset.hyperparamcombo_count
# +
hyperparamcombos = hyperparamset.hyperparamcombos
for h in hyperparamcombos:
print(h.hyperparameters)
# -
hyperparamcombos[0].get_hyperparameters(as_pandas=True)
# ## 8. `Queue` of training `Jobs`.
# The `Queue` is the central object of the "logic side" of the ORM. It ties together everything we need for training and hyperparameter tuning.
queue = aiqc.Queue.from_algorithm(
algorithm_id = algorithm.id
, splitset_id = splitset.id
, hyperparamset_id = hyperparamset.id # Optional.
, foldset_id = None # Optional.
, repeat_count = 3
, permutation_count = 3
)
# * `repeat_count:int=1` allows us to run the same `Job` multiple times. Normally, each `Job` has 1 `Predictor` associated with it upon completion. However, when `repeat_count` (> 1 of course) is used, a single `Job` will have multiple `Predictors`.
#
# > Due to the fact that training is a *nondeterministic* process, we get different weights each time we train a model, even if we use the same set of parameters. Perhaps you've have the right topology and parameters, but, this time around, the model just didn't recgonize the patterns. Similar to flipping a coin, there is a degree of chance in it, but the real trend averages out upon repetition.
# * `hide_test:bool=False` excludes the test split from the performance metrics and visualizations. This avoids data leakage by forcing the user to make decisions based on the performance on their model on the training and evaluation samples.
# * `permutation_count:int` controls the number of times each feature column is shuffled before it's impact on loss is compared to the baseline training loss before the median taken. Feature importance is calculated for each column of each `Feature.id`, except for `Feature.dataset.dataset_type=='image'`. If you are rapidly training exploratory models on many columns, then you can set `permutation_count=0`.
#
# > `[training loss - (median loss of <5> permutations)]`
# ### a) `Job` objects.
# Each `Job` in the Queue represents a `Hyperparamcombo` that needs to be trained.
#
# If a `Foldset` is used during `Queue` creation, then:
#
# - The number of jobs = `hyperparamcombo_count` * `fold_count`.
# - Each Job will have a `Fold`. Additionally, a superficial `Jobset` will be used to keep track of all Jobs related to that Foldset.
# ### b) Executing `Jobs`.
#
# There are two ways to execute a Queue of Jobs:
#
# #### i) `queue.run_jobs()`
#
# * Jobs are simply ran on a loop on the main *Process*.
#
# * Stop the Jobs with a keyboard interrupt e.g. `ctrl+Z/D/C` in Python shell or `i,i` in Jupyter.
#
# * It is the more reliable approach on Win/Mac/Lin.
#
# * Although this locks your main process (can't write more code) while models train, you can still fire up a second shell session or notebook.
#
# * Prototype your training jobs in this method so that you can see any errors that arise in the console.
#
#
# #### ii) DEPRECATED - `queue.run_jobs(in_background=True)`.
#
# *Support for background processing has not been restored after decoupling the preprocessing pipelines from the Queue/Job logic.*
#
# * The Jobs loop is executed on a separate, parallel `multiprocessing.Process`
#
# * Stop the Jobs with `queue.stop_jobs()`, which kills the parallel *Process* unless it already failed.
#
# * The benefit is that you can continue to code while your models are trained. There is no performance boost.
#
# * On Mac and Linux (Unix), `'fork'` multiprocessing is used (`force=True`), which allows us to display the progress bar. FYI, even in 'fork' mode, Python multiprocessing is much more fragile in Python 3.8, which seems to be caused by how pickling is handled in passing variables to the child process.
#
# * On Windows, `'spawn'` multiprocessing is used, which requires polling:
#
# * `queue.poll_statuses()`
#
# * `queue.poll_progress(raw:bool=False, loop:bool=False, loop_delay:int=3)` where `raw=True` is just a float, `loop=True` won't stop checking jobs until they are all complete, and `loop_delay=3` checks the progress every 3 seconds.
#
# * It is a known bug that the `aiqc.TrainingCallbacks.Keras.MetricCutoff` class does not work with `` as of Python 3.8.
#
# * Also, during stress tests, I observed that when running multiple queues at the same time, the SQLite database would lock when simultaneous writes were attempted.
queue.run_jobs()
# The queue is interuptable. You can stop the execution of a queue and resume it later.
#
# > This also comes in handy if either your machine or Python kernel either crashes or are interupted by accident. Whatever the reason, rest easy, just `run_jobs()` again to pick up where you left off. Be aware that the `tqdm` iteration time in the progress bar will be wrong because it will be divided by the jobs already ran.
# ##### Preprocessing during Job is recorded
# During the execution of a Job, the latest Labelcoder and Encoderset(s) tied to the Label and Feature(s) of the Splitset used during training will record their `fit()` to the data.
#
# - `FittedLabelcoder` for the Labelcoder used.
# - `fitted_encoders:object` to store the `fit`.
# - `FittedEncoderset` for each Feature used.
# - `fitted_encoders:list` to store the `fit`(s).
#
# This process is critical for:
#
# - `inverse_transform()`'ing aka decoding predictions.
# - Encoding new data during inference exactly the same was as the samples that the model was trained on.
# It takes a lot of joins to fetch the fitted encoders after the fact. So these methods are used behind the scenes to make it a bit easier:
#
# - `Predictor.get_fitted_encoderset(job, label)`
# - `Predictor.get_fitted_labelcoder(job, feature)`
# ### c) `Predictors` are the trained models.
# Each `Job` trains a `Predictor`. The following attributes are automatically written to the `Predictor` after training.
#
# * `model_file`: serialization varies for Keras and Pytorch deep learning framework.
#
# * `input_shapes`: used by `get_model()` during inference.
#
# * `history`: per epoch metrics recorded during training.
# #### Fetching the trained model.
compiled_model = queue.jobs[0].predictors[0].get_model()
compiled_model
# ### d) `Predictions` are the output of a Predictor.
# When you feed samples through a Predictor, you get Predictions. During training, Predictions are automatically generated for every split/fold that was tied to the Queue.
queue.jobs[0].predictors[0].predictions[0].metrics
# #### Fetching metrics.
# * `predictions`: dictionary of predictions per split/ fold. Values are `inverse_transform`'ed if Labels were encoded during training.
#
# * `probabilities`: dictionary of prediction probabilities per split/ fold. `None` for regression.
#
# * `metrics`: dictionary of metrics for each split/fold that vary based on the analysis_type.
#
# * `metrics_aggregate`: dictionary of average for each statistic across all splits/folds.
#
# * `plot_data`: metrics readily formatted for plotting.
# ## 9. Metrics & Visualization
# For more information on visualization of performance metrics, reference the [Visualization & Metrics](visualization.html) documentation.
| docs/notebooks/api_low_level.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''dev'': conda)'
# language: python
# name: python38264bitdevconda06c7c692422b4cb59f191a8cff7c413e
# ---
# # Working With Surfaces
# ## 1. Building Elemental Surfaces
#
# <b>In this lesson and in the corresponding exercises we will learn how to generate specific surfaces of elemental materials, as well as all possible surfaces up to a certain Miller index.</b>
# <br><br>
# **Copper: (1, 1, 1) surface**
#
# For our first example we will generate a copper (1, 1, 1) surface. Let's start by importing a Cu structure from a .cif file.
from pymatgen.core.structure import Structure
Cu = Structure.from_file("Cu.cif")
# <br>We can now start thinking about how to represent surfaces: in <i>pymatgen</i>, this is done through slabs. A slab is a cell containing a few layers of atoms and vacuum, where the termination corresponds to the surface we are interested in. Here is an example of a Cu slab corresponding to a (1, 1, 1) surface:
#
# 
#
# <br><br>If we visualize multiple slabs instead of just one, we can see how the surface-like structure arises. Here is the same slab from above, copied 15 times in the <i>x</i> and <i>y</i> directions, and twice in the <i>z</i> direction:
#
# 
# <br>Let's now try to generate surfaces using the <i>SlabGenerator</i> class.
from pymatgen.core.surface import SlabGenerator
# <br>Let's initialize it for the Cu structure we just created, with a Miller index of (1, 1, 1), a minimum slab height of 10 Å, and minimum 10 Å of vacuum.
slabgen = SlabGenerator(Cu, (1,1,1), 10, 10)
# <br><br>We can now use the <i>get_slabs()</i> method of the <i>SlabGenerator</i> class to create a list of <i>Slab</i> objects. The slabs returned by <i>get_slabs()</i> in this case correspond to all the unique terminations along the normal to the Miller plane we are interested in. For example, for the Cu (1, 1, 1) case, the algorithm will find all the unique terminations that are parallel to the (1, 1, 1) plane i.e. perpendicular to the normal. The different resulting slabs are characterized by different c shifts:
# 
slabs = slabgen.get_slabs()
# <br>For a fcc structure such as the Cu structure we are working with, there should only be only one unique (1, 1, 1) surface. We can check if that is the case:
print(len(slabs))
# <br>We can print the Miller index of this surface, as well as the c shift value:
# +
Cu_111 = slabs[0]
print(Cu_111.miller_index, Cu_111.shift)
# -
# <br>Let's now visualize the surface we just generated. For this, we can use the <i>plot_slab</i> function and we will also need to import <i>pyplot</i>, a python plotting framework.
# +
from pymatgen.analysis.adsorption import plot_slab
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# <br>We can now plot our surface:
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
plot_slab(Cu_111, ax, adsorption_sites=False)
ax.set_title("Cu (1, 1, 1) surface")
ax.set_xticks([])
ax.set_yticks([])
plt.show()
# <br><br>
# ## 2. Building Compound Surfaces
#
# <b>In this part of the tutorial we will extend what we learned about generating elemental surfaces to compound materials by accounting for the additional complexity of such systems.</b>
# <br><br>
# Let's try something a little more complicated, say LiFePO<sub>4</sub>:
LiFePO4 = Structure.from_file("LiFePO4.cif")
# <br>Let's add some oxidation states to LiFePO<sub>4</sub>, this will be important when we want to take surface polarity into consideration:
LiFePO4.add_oxidation_state_by_element({"Fe": 2, "Li": 1, "P": 5, "O": -2})
# <br>We can now initialize an instance of the <i>SlabGenerator</i> class, let's choose a Miller index of (0, 0, 1) and the same minimum slab size and minimum vacuum of 10 Å:
slabgen = SlabGenerator(LiFePO4, (0,0,1), 10, 10)
# <br>We can now use <i>get_slabs()</i> to generate our (0, 0, 1) slabs. Unlike before, when generating a slab of LiFePO<sub>4</sub>, we also want to be careful not break any of the P-O bonds - these bonds are strong enough that they will result in a significantly high surface energy when broken. To implement this, we add the <i>bonds</i> parameter to <i>get_slabs()</i>, a dictionary where the key will be a tuple of the two atoms whose bonds we do not want to break and the element of that value would be their maximum bond length in Å. This means that any bond between P and O less than 2 Å cannot be broken when generating slabs.
slabs = slabgen.get_slabs(bonds={("P", "O"): 2})
# <br>How many (0, 0, 1) slabs that do not break any P-O bonds are there?
print(len(slabs))
# <br>There are a couple of other things to keep in mind when working with slabs:
# <ul>
# <li>First off, if we want to calculate the surface energy, we need to ensure that the slabs have the same surface on both sides i.e. they contain inversion symmetry. We can check whether that is the case by using the <i>is_symmetric()</i> property of our slab object. If the top and bottom surface of the slab were different, we would only be able to calculate the average surface energy of the two different terminations in our slab, rather than the surface energy of the one termination we are interested in. </li>
# <li>Secondly, for structures containing oxidation states, a good thing to check for is whether our surfaces are polar. A polar termination will lead to a very high surface energy, so we might want to skip those particular structures in any calculations we want to carry out. We can check polarity using the <i>is_polar()</i> property of our slab object.</li>
# </ul>
for n, slab in enumerate(slabs):
print(n, slab.is_polar(), slab.is_symmetric())
# <br>Notice that none of the terminations in the (0, 0, 1) direction simultaneously satisfy our two criteria so a (0, 0, 1) surface with a reasonable surface energy cannot be calculated.
# <br>Now let's generate all possible slabs for a maximum Miller index of 2 for LiFePO<sub>4</sub> and see if any of these surfaces can be calculated to yield reasonable and accurate surface energy values. This may take a while.
from pymatgen.core.surface import generate_all_slabs
all_slabs = generate_all_slabs(LiFePO4, 2, 10, 10, bonds={("P", "O"): 2})
# <br>Let's now select the surfaces that satisfy our aforementioned criteria and see how many of them there are compared to the total number of generated slabs:
# +
valid_slabs = []
for slab in all_slabs:
if not slab.is_polar() and slab.is_symmetric():
valid_slabs.append(slab)
print(len(all_slabs), len(valid_slabs))
# -
# <br><br>
# ## 3. Adding adsorbates ##
#
# <b>Now that we have learned how to generate various types of surfaces, in this part of the tutorial we will learn how to find adsorption sites and generate adsorption structures.</b>
# <br><br>
# In order to find adsorption sites and generate adsorption structures, we will use the <i>AdsorbateSiteFinder</i>:
from pymatgen.analysis.adsorption import AdsorbateSiteFinder
# <br>Let's now use our Cu structure to add adsorbates to different surfaces. We can start with our previously generated Cu (1, 1, 1) surface and make an instance of the <i>AdsorbateSiteFinder</i>:
asf = AdsorbateSiteFinder(Cu_111)
# <br>By default, the three types of adsorption sites identified using <i>pymatgen</i> are:
# <ul><li><i>on top</i> - positioned on top of a surface atom</li>
# <li><i>bridge</i> - positioned between two surface atoms</li>
# <li><i>hollow</i> - positioned between three surface atoms</li>
# </ul>
# Let's use the <i>find_adsorption_sites()</i> method to find the relevant adsorption sites on our copper surface:
ads_sites = asf.find_adsorption_sites()
print(ads_sites)
# <br>You can see that there is a total of 4 adsorption sites: one "on top", one "bridge" and two "hollow".
# Let's now visualize these adsorption sites, using <i>plot_slab()</i>:
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
plot_slab(Cu_111, ax, adsorption_sites=True)
plt.show()
# <br>We can now move on to adding adsorbates. Let's start by making a H molecule, that we can use as an adsorbate:
# +
from pymatgen.core.structure import Molecule
adsorbate = Molecule("H", [[0, 0, 0]])
# -
# <br>We can now use the <i>generate_adsorption_structures()</i> method to create adsorption structures corresponding to the 4 adsorption sites we identified. Let's keep the same slab size by setting the <i>repeat</i> argument to [1, 1, 1]. We can control the distance between the adsorbate and the slab using the "distance" key in the dictionary passed to the <i>find_args</i> argument - let's make this 1.6 Å:
ads_structs = asf.generate_adsorption_structures(adsorbate, repeat=[1, 1, 1], find_args={"distance": 1.6})
# <br>Let's visualize our new adsorption structures:
fig = plt.figure(figsize=[15, 60])
for n, ads_struct in enumerate(ads_structs):
ax = fig.add_subplot(1, 4, n+1)
plot_slab(ads_struct, ax, adsorption_sites=False)
ax.set_title("Cu (1, 1, 1) + H structure %i" %(n+1))
ax.set_xticks([])
ax.set_yticks([])
ax.set_xlim(0,5)
ax.set_ylim(-1,4)
plt.show()
| workshop/lessons/03_surfaces/Working With Surfaces_lesson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 64-bit
# name: python3
# ---
import codecs
import glob
import logging
import multiprocessing
import os
import pprint
import re
import nltk
import gensim.models.word2vec as w2v
import sklearn.manifold
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import gensim
# %pylab inline
import os
import textract
import document
import glob
from os import listdir
from os.path import isfile, join
from os import walk
#read filenames for 100 sample files in BostonGlobe 2014
Path = '../../Doc2vec/model/100_articles/2014/docx/'
filenames = []
number = 0
print("Articles: ")
for file in os.listdir(Path):
if file.endswith(".doc") or file.endswith(".docx"):
filenames.append(file)
number += 1
print("(",number,")" , file)
# +
#read for 100 sample files in BostonGlobe 2014
directory = r'../../Doc2vec/model/100_articles/2014/docx/'
file_dir = []
text = ""
#for file in os.listdir(directory):
if file.endswith(".doc") or file.endswith(".docx"):
text.append(textract.process(os.path.join(directory, file)))
else:
continue
#print(text)
# -
# Tokenize each sentence
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
raw_sentences = tokenizer.tokenize(text)
| NAACP/Doc2vec/model/Doc2Vec_Hong.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
def func():
return 1
return_one = func
return_one
return_one()
# +
def new_decorator(func):
print("This is a new func")
func()
@new_decorator
def going_to_return_one():
print "Is this a decorator"
# -
new_decorator = going_to_return_one
new_decorator(lambda x, y: x**y)
def hello(name="Jose"):
print("Hello %s" % name)
hello()
greet = hello
greet()
globals()
globals()['Out']
locals()
def hello_2(name="louis"):
# local functions
print 'The hello() function has been executed'
def greet():
return '\t this is inside of the greet() function'
def welcome():
return '\t this is inside of the welcome() function'
print greet()
print welcome()
print 'Now we are back inside the hellow_2 function'
hello_2()
# returning functions
def hello_3(name="louis"):
def greet():
return '\t this is inside of the greet() function'
def welcome():
return '\t this is inside of the welcome() function'
if name == "louis":
return greet
else:
return welcome
x = hello_3()
print x()
# +
# functions as arguments
def hello_4():
return 'Hi Louis'
def other(func):
print "Other code goes here"
print func()
other(hello_4)
# +
def new_decorator(func):
def wrap_func():
print 'Code here, before executing func'
func()
print 'Code here will execute after the func'
# note: returning the function itself, not the results of the func
return wrap_func
def func_needs_decorator():
print 'This function needs a decorator'
# -
func_needs_decorator()
func_needs_decorator = new_decorator(func_needs_decorator)
func_needs_decorator()
# +
def cat_decoy(func):
def wrap_func():
print('In decoy')
func()
return wrap_func
def catnip1():
print('In catnip')
# -
catnip1 = cat_decoy(catnip1)
# +
@cat_decoy
def catnip():
print 'In cat nip 2'
@cat_decoy
def catnip_original():
print 'Original'
# -
catnip()
catnip_original()
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Making Use of Selenium
# *<NAME>*
#
# I show in this notebook how to get started using Selenium. We set up a driver and connect to a webpage.
#
# We run a Chrome browser with Selenium, so we can watch Selenium in action, but in practice we might opt for [PhantomJS](http://phantomjs.org/) (though this might not be wise if we're concerned about looking "human").
#
# The following code starts a Selenium web driver, looks at a page, then closes the driver.
from selenium import webdriver
from time import sleep
path = "chromedriver" # Depends on system/OS/etc.
driver = webdriver.Chrome(executable_path=path)
sleep(15) # For presentation purposes only
driver.get("http://pycoders.com/archive/") # Go to a website
sleep(15) # For presentation purposes only
driver.close()
| Section 5/UseSelenium.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Software Development 1
// ---
// Topics for today will include:
// - Syllabus
// - Welcome to CMPT 220
// - What's So Different?
// - Language Types
// - Different Languages for Different Problems
// - High Level vs Low Level
// - Let's Be Honest...
// - Favorite Languages
// - Programming Styles
// - Why Java?
// - JVM
// - Object Oriented Language
// - IDEs
// - Hello World!
//
//
// ## Welcome to CMPT 220!
// ---
//
// In this course I'll be teaching you introductory Java and eventually some upper level and more advanced topics related to the language.
//
// For this course lesson notes will primarily be hosted using [Jupyter Notebooks](https://jupyter.org/index.html). I'm going to expect you to be able to grasp the concepts that we'll discuss centered around them and how they work. There will be some work that needs to be done on your part since we're using a different kernel than the native one. We'll also talk about what a kernel is later! Here is the [IJava Kernel](https://github.com/SpencerPark/IJava) we're going to need this semester!
//
// With Jupyter Notebooks we can:
// - Have notes for each lesson.
// - Have a place where we can see how code executes.
// - Have an interactive space for students to be able to rerun things seen in lessons.
//
// I encourage you to get familiar with using the Jupyter Notebooks because they make an amazing playground for trying things out! Our labs will also be primarily in the Jupyter Notebooks.
System.out.println("Welcome to Soft Dev 1! I hope you're not bored!");
System.out.println("Hope you have a great time!");
// ## What's So Different
// ---
// So at this point you've already taken your first programming class and learned your first language. First off congrats! Second you're probably wondering why instead of building on top of what you already know and delving deeper, we're switching to a different language. Something that's important to understand is that in the world of computing there are several languages available to you as a programmer. These different languages are good at different things. Generally we have High Level Languages and Low Level Languages. With those come different properties typically.
//
// Coming from a language like Python you're probably wondering what other languages are like. Python was a jack of all trades. Capable of 4 different types of programming.
// - Functional
// - Procedural
// - Object Oriented
// - Imperative
//
// We may think why would we use something else with such flexibility. Well we can start with the fact that Python is a high level language. It's far away from the actual operating system commands and what not. This means that we sacrifice efficency and speed for ease of use. Think of the phrase "Jack of all trades, master of none." That applies here. Python doesn't delve all the way into all of those types.
//
// Now if we think about the real world there are a plethora of problems. Where the distinction in choosing a High Level Language and a Low Level Language is usually efficency. Does our application need to be really fast? That means we're usually going to pick a Low Level Language because comparitively it's faster than a High Level language when written well. Is it something that can be quick and dirty. Then we can use something like a Python. Not to say using Java is overkill there but that's the reasoning.
//
// For this class we're focused on Object Oriented Programming
//
// This paradigm of being an Object Oriented Language is usually associated with Low Level Languages like C++ or Java. There are several other languages that take advantage of it, High and Low. For this class we're going to learn what it's like to code in a Lower Level Language.
// ## Let's Be Honest
// ---
// I personally am not the biggest fan of Java BUT I know why it's useful and great. Personally I tend to like languages like Python more and do things over there more but the important thing that I want to get across with this is that you'll have your favorites. You'll have your dislikes. The most important thing that I want you to remember and pay attention to is that feeling. As you go through your career as a Computer Scientist you'll be faced with job roles and opportunities that are in disiplines that you like and dislike and it's important to know what you like and dislike.
//
// On the flipside!!! Be open to doing it! Using it. Knowing why it's good at what it does. Knowing Java makes me a better overall programmer and lets me do things like this and teach you guys!
//
// Stylistically what we're aiming for as a sub goal with this class is forming your identity as a programmer. Do you like High Level work more than this? Do you love Low Level programming. One thing that I see a lot are students that have no idea what they like when it comes time for **Cough Cough*\* getting internships(Start early! They pay well!!!) and interviewing for full time positions.
//
// Learning a bunch of different programming styles and methodologies is super helpful because as you get better and better the concepts start to bleed into one another and you'll be able to do things you've never imagined with ease.
// ## Why Java?
// So let's talk a little about why Java is a thing in the first place. Java is a language used everywhere and by everyone. This is mostly because of something called the JVM. So if you've never heard of the JVM or even a VM, don't worry! First JVM stands for Java Virtual Machine, and VM stands for as you can probably guess Virtual Machine. Now a Virtual Machine does it's best to take from a pool of hardware resources and make a virtualized version of a computer. Now the JVM does something similar but instead of being an instance of a computer, it's a runtime enviornment capable of running Java code and applications. Why this is important is that Java or Oracle and OpenJDK make the JVM runnable on all types of machines for the most part. So the hardware doesn't specifically need to know how to run the code. You'd just run it inside of the JVM.
//
// Nowadays that's not that impressive. When Java first came about however, this was miracaulous. This made bringing Java to different platforms a lot easier than it was for other languages. This is long before the days of package installers and easily accessible executables.
//
// In addition to being extremely portable it was an improvement on C++ or what was planned to be at least. Back then you really had to understand everything about a computer to be able to program. How to access memory. Making sure you didn't accidentally write over memory in use. Keeping programs slim and efficent. These things were difficult to grasp. Java aimed to make it a little simpler to understand than C++. Object Oriented Programming is the style that they both implement and is important to understand entering the realm of lower level programming.
//
// ## IDEs
// This will be quick. IDE stands for Integrated Development Environment. In laymen's terms it's an editor that houses a bunch of tools that you may need to develop. Unlike a VS Code where out of the box it's very lightweight, IDEs come with tools.
//
// I'm not fond of many IDEs. The one that I DO like is [IntelliJ](https://www.jetbrains.com/idea/). They just have a clean IDE and it doesn't feel clunky to me. I mostly use [VS Code](https://code.visualstudio.com) to edit things. This isn't an IDE but can be used in a similar way with certain plugins added to it.
//
// *IMPORTANT! FOR THIS CLASS YOU'RE GOING TO NEED THE INSIDER EDITION OF VS CODE FOR THE JUPYTER NOTEBOOKS! THAT CAN BE FOUND [HERE](https://code.visualstudio.com/insiders/)*
//
// The reason for this section is that these are the 2 that I use and will know the best when it comes to questions. I believe that you should do your own due dilligence though and form your own opinions.
//
// Two other IDEs that i've had to use in the past are (Not the biggest fan of either)
// - [Eclipse](https://www.eclipse.org/downloads/)
// - [NetBeans](https://netbeans.org)
//
// Some other text editors I've used (I LOVE ATOM <3 ^_^)
// - [Atom](https://atom.io)
// - [Notepad++](https://notepad-plus-plus.org/downloads/)
// ## Hello World
System.out.println("Hello there!");
| JupyterNotebooks/Lessons/Lesson 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <strong>Road networks and robustness to flooding on US Atlantic and Gulf barrier islands</strong>
# ## <strong>- Road network robustness to flooding -</strong>
# ### The purpose of this notebook is to identify, for each barrier island, the elevation and exceedance probability of the critical node that causes the network's failure and the overall robustness of each road network to flood-induced failures.
# +
# Packages
import os
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
import geopandas as gpd
import osmnx as ox
import networkx as nx
import pandas as pd
import numpy as np
import contextily as ctx
import statistics
from scipy import stats
# +
### Set working directory
path='' # introduce path to your working directory
# os.chdir(path) # In this notebook, this command cannot be used because it triggers a JSONDecodeError when GA9 is downloaded
# To avoid the error and be able to download all road networks, the path to the working directory needs to be set as an absolute path.
# +
### Explore the size decay of the GCC to identify the critical node that leads to the fragmentation of the network (road networks with more than 100 nodes) and plot maps with road networks
# Create folders if they don't exist
outdir= '{0}/Results'.format(path)
if not os.path.exists(outdir):
os.makedirs(outdir)
outdir= '{0}/Results/GCC_Plots'.format(path)
if not os.path.exists(outdir):
os.makedirs(outdir)
outdir= '{0}/Results/Networks_Maps'.format(path)
if not os.path.exists(outdir):
os.makedirs(outdir)
# Loop through files to open each barrier graphml
rootdir = '{0}/Data/Roads'.format(path)
extensions = ('.graphml')
for subdir, dirs, files in os.walk(rootdir):
for file in files:
ext = os.path.splitext(file)[-1].lower()
if ext in extensions:
file_path = os.path.join(subdir, file)
barrier = file.replace(".graphml","")
print(barrier)
G = nx.read_graphml(file_path)
N = len(G.nodes(data=True))
GCCs=[] # list with the number of connected components and its size
if N>100:
# pull out elevation attribute
Z = nx.get_node_attributes(G,'Elevations')
# convert str values in float to be able to sort them
Z = dict(zip(Z.keys(), [float(value) for value in Z.values()]))
# sort elevation values in ascending order
Sorted_Z = sorted(Z.items(), key=lambda item: item[1])
# select first element of each tuple in the list (nodes ID):
FT = [i[0] for i in Sorted_Z]
# Select second element of each tuple in the list (elevation) and convert to float
ST = [i[1] for i in Sorted_Z]
for i in range(len(ST)):
ST[i] = float(ST[i])
# create array
CCs = np.zeros([len(Sorted_Z),2])
# loop through all nodes
for i in range(0, len(FT)):
# find the node with lowest elevation from the list using i and remove it
G.remove_nodes_from(FT[0:i])
# find the number of connected components and its respective size
GCC = [len(c)
for c in sorted(nx.weakly_connected_components(G), key=len, reverse=True)]
GCCs.append(GCC)
# fill array, first column corresponds to FGC (first giant component), second column to SGC (second giant component)
if len(GCC)==1:
CCs[int(i),0]=GCC[0]/len(FT)
CCs[int(i),1]=0
else:
CCs[int(i),0]=GCC[0]/len(FT)
CCs[int(i),1]=GCC[1]/len(FT)
# find the node that, once removed, the size of the FGC drops abruptly while the size of the SGC reaches its maximum
m = max(CCs[:,1])
pos=[i for i, j in enumerate(CCs[:,1]) if j == m]
pos= pos[0] # position of max value in SGC
critical= pos-1 # position of the critical node whose removal causes the percolation transition.
elev=ST[critical] # find elevation of the critical node
removed=pos # number of nodes removed when percolation threshold occurs
# plot
col1=[]
for i in range(0,len(FT)):
if i==critical:
col1.append('#D53032')
else:
col1.append('#000000')
col2=[]
for i in range(0,len(FT)):
if i==critical:
col2.append('#D53032')
else:
col2.append('#808080')
col3=[]
for i in range(0,len(FT)):
if i==critical:
col3.append('#D53032')
else:
col3.append('#9ACD32')
f, (ax1,ax2) = plt.subplots(2, 1, sharex=True)
x_coord = 1 * np.arange(len(FT))/len(FT) # fraction of nodes removed
ax1.plot(x_coord, CCs[:,0],':ok') # FGC
for i in range (len(FT)):
ax1.plot(x_coord[i],CCs[i,0],'o', markersize=5, color=col1[i]) # plot with two colors to highlight critical node
ax1.set_ylabel("First Giant Component Size")
ax3 = ax1.twinx()
ax3.plot(x_coord, CCs[:,1],':ok') # SGC
for i in range (len(FT)):
ax3.plot(x_coord[i],CCs[i,1],'o', markersize=5, color=col2[i])
ax3.set_ylabel("Second Giant Component Size")
ax2.plot(x_coord,CCs[:,0],':ok') # FGC
for i in range (len(FT)):
ax2.plot(x_coord[i],CCs[i,0],'o', markersize=5, color=col1[i])
ax2.set_ylabel("First Giant Component Size")
ax4 = ax2.twinx()
ax4.plot(x_coord,ST,':o', color='#9ACD32') # elevation
for i in range (len(FT)):
ax4.plot(x_coord[i],ST[i],'o', markersize=5, color=col3[i])
ax2.set_ylabel("First Giant Component Size",)
ax4.set_ylabel("Elevation")
ax2.set_xlabel("Fraction of removed nodes")
legend_elements1 = [Line2D([0], [0], marker='o', color='#000000', label='FGC', markersize=10),
Line2D([0], [0], marker='o', color='#808080', label='SGC', markersize=10),
Line2D([0], [0], marker='o', color='#D53032', label='Critical node', markersize=10)]
ax1.legend(handles=legend_elements1, loc="best", frameon=False, fontsize=18)
legend_elements2 = [Line2D([0], [0], marker ='o', color='#000000', label='FGC', markersize=10),
Line2D([0], [0], marker='o', color='#9ACD32', label='Elevation', markersize=10)]
ax1.legend(handles=legend_elements1, loc="best", frameon=False, fontsize=18)
plt.rcParams["font.size"]= 20
plt.rcParams["figure.figsize"] = (15,15)
f.savefig("{0}/Results/GCC_Plots/{1}.png".format(path,barrier), dpi=500, facecolor='w')
plt.close("all")
### create maps for each network using OSM as basemap
# read polygons
poly = gpd.read_file("{0}/Data/Barriers/Barriers_AtlGulf/{1}_geo.shp".format(path,barrier))
# extract just the geometry (shapely object) part and clean it with a buffer
poly_geo = poly['geometry'].iloc[0]
poly_geo = poly_geo.buffer(0)
poly_geo.is_valid
# extract drivable network and project it
graph = ox.graph_from_polygon(poly_geo, network_type='drive', simplify=True, clean_periphery=True)
# retrieve nodes and edges as geodataframes
nodes, edges = ox.graph_to_gdfs(graph)
# create an index for the geodataframe nodes
nodes['index'] = range(0, len(nodes))
# convert Z dict in pandas dataframe and name columns
Z = pd.DataFrame(list(Z.items()),columns = ['index','elevation'])
# convert all columns in numerics so there are no errors when merging
Z = Z.apply(pd.to_numeric)
# join pandas dataframe to nodes geodataframe using 'index' so that the gdf has elevation
nodes = nodes.merge(Z, on='index')
# create new columns for color and size
def color(row):
if row['elevation'] < elev:
val = "black"
elif row['elevation']== elev:
val = "red"
else:
val = "green"
return val
def size(row):
if row['elevation'] == elev:
val = 50
else:
val = 30
return val
nodes['Color'] = nodes.apply(color, axis=1) # new column with color categories
nodes['Size'] = nodes.apply(size, axis=1) # new column with size categories
# plot map
fig, ax = plt.subplots()
nodes = nodes.to_crs(epsg=3857) # convert gdf to EPSG used by basemaps
edges = edges.to_crs(epsg=3857)
nodes.plot(ax=ax, color=nodes.Color, markersize=nodes.Size, zorder=2, legend=True) # plot nodes
edges.plot(ax=ax, alpha=0.2, color='black', zorder=1) # plot edges
ctx.add_basemap(ax, zoom=13, source=ctx.providers.OpenStreetMap.Mapnik) # add basemap (OSM)
plt.xticks(fontsize=12) # reduce fontsize of x axis
plt.yticks(fontsize=12) # reduce fontsize of y axis
legend_elements = [Line2D([0], [0], marker='o', color='black', label='Connected nodes',
markerfacecolor='g', markersize=10),
Line2D([0], [0], marker='o', color='black', label='Disconnected nodes',
markerfacecolor='b', markersize=10),
Line2D([0], [0], marker='o', color='black', label='Target node',
markerfacecolor='r', markersize=10),
] # create legend
ax.legend(handles=legend_elements, loc='best', frameon=False)
ax.set_title(barrier, fontsize=22)
ax.ticklabel_format(style='plain')
plt.rcParams["figure.figsize"] = (25,25)
plt.savefig('{0}/Results/Networks_Maps/{1}.png'.format(path,barrier), dpi=300, facecolor='w')
plt.close("all")
# +
### Create table with results for all barriers with drivable networks
barriers=[] # barrier name
n_nodes=[] # number of nodes
r=[] # robustness
min_z=[] # min node elevation in the network
max_z=[] # max node elevation in the network
mean_z=[] # mean node elevation
median_z=[] # median node elevation
critical_z=[] # elevation critical node
critical_e=[] # exceedance probability critical node (given in return period)
removed_nodes=[] # number of nodes removed when critical node is removed
removed_perc=[] # percentage of nodes removed when critical node is removed
threshold=[] # value critical threshold
rootdir = '{0}/Data/Roads'.format(path)
extensions = ('.graphml')
# Loop through files and open barrier graphml
for subdir, dirs, files in os.walk(rootdir):
for file in files:
ext = os.path.splitext(file)[-1].lower()
if ext in extensions:
file_path = os.path.join(subdir, file)
barrier = file.replace(".graphml","")
G = nx.read_graphml(file_path)
N= len(G.nodes(data=True))
if N>100:
GCCs=[]
barriers.append(barrier)
n_nodes.append(N)
# pull out elevation attribute
Z = nx.get_node_attributes(G,'Elevations')
# convert str values in float to be able to sort them
Z = dict(zip(Z.keys(), [float(value) for value in Z.values()]))
# sort it based on elevation, min first
Sorted_Z = sorted(Z.items(), key=lambda item: item[1])
CCs = np.zeros([len(Sorted_Z),2])
# select first element of each tuple in the list (nodes ID):
FT = [i[0] for i in Sorted_Z]
# select second element of each tuple in the list (elevation) and convert to float
ST = [i[1] for i in Sorted_Z]
for i in range(len(ST)):
ST[i] = float(ST[i])
# calculate elevation stats
min_elev=min(ST)
min_z.append(min_elev)
max_elev=max(ST)
max_z.append(max_elev)
mean_elev = statistics.mean(ST)
mean_z.append(mean_elev)
median_elev = statistics.median(ST)
median_z.append(median_elev)
# remove nodes by elevation and calculate size of first and second components
for i in range(0, len(FT)):
# find the node with lowest elevation from the list using i and remove it
G.remove_nodes_from(FT[0:i])
# find the number of connected components and its respective size
GCC = [len(c)
for c in sorted(nx.weakly_connected_components(G), key=len, reverse=True)]
GCCs.append(GCC) # list with the number of connected components and its size
# fill array, first column corresponds to FGC (first giant component), second column to SGC (second giant component)
if len(GCC)==1:
CCs[int(i),0]=GCC[0]/len(FT)
CCs[int(i),1]=0
else:
CCs[int(i),0]=GCC[0]/len(FT)
CCs[int(i),1]=GCC[1]/len(FT)
# find the node that, once removed, the FGC decreases and the SGC reaches its maximum (critical threshold)
m = max(CCs[:,1])
pos=[i for i, j in enumerate(CCs[:,1]) if j == m]
pos= pos[0] # position of max value in SGC
critical= pos-1 # position of the critical node whose removal causes the percolation transition.
elev=ST[critical] # find elevation of the critical node
critical_z.append(elev)
removed=pos # number of nodes removed when percolation threshold occurs
removed_nodes.append(removed)
perc_removed=int(removed)/N*100
removed_perc.append(perc_removed)
x_coord = 1 * np.arange(len(FT))/len(FT) # Fraction of nodes removed
thresh= x_coord[critical]
threshold.append(thresh)
# exceedance probability for the critical node
exceed = pd.read_csv("{0}/Data/Exceedance/Probability/{1}_Exceedance.csv".format(path,barrier), sep=",", header=0)
exceed_x= exceed.MaxWL
exceed_y= exceed.Probability
node_elev= elev
exceedance= np.interp(node_elev, exceed_x, exceed_y)
critical_e.append(exceedance)
# calculate robustness following Schneider's equation (2011)
s= sum(CCs[:,0])
rob= s/len(FT)
r.append(rob)
else:
continue
table = list(zip(barriers,n_nodes,r,min_z,max_z,mean_z,median_z,critical_z,critical_e,removed_nodes,removed_perc,threshold))
table = pd.DataFrame(table, columns=['Barrier','Nodes','Robustness','Min_elevation','Max_elevation','Mean_elevation','Median_elevation','Critical_elevation','Critical_exceedance','Removed_nodes','Removed_%','Critical_threshold'])
table.to_csv('{0}/Results/Results_AllBarriers.csv'.format(path))
# +
### For each network, calculate basic statistics using OSMnx package
# Create folders if it doesn't exist
outdir= '{0}/Results/Statistics'.format(path)
if not os.path.exists(outdir):
os.makedirs(outdir)
rootdir = "{0}/Data/Barriers/Barriers_AtlGulf".format(path)
extensions = ('.shp')
table = pd.read_csv("{0}/Results/Results_AllBarriers.csv".format(path), sep=",", header=0)
developed_barriers= list(table.Barrier) # to calculate statistics only for the selected barrier islands (those with more than 72 nodes)
for subdir, dirs, files in os.walk(rootdir):
for file in files:
ext = os.path.splitext(file)[-1].lower()
if ext in extensions:
file_path = os.path.join(subdir, file)
barrier = file.replace(".shp","")
barrier = barrier.replace("_geo","")
print(barrier)
if barrier in developed_barriers:
# read polygons
poly = gpd.read_file(file_path)
# extract just the geometry (shapely object) part and clean it with a buffer
poly_geo = poly['geometry'].iloc[0]
poly_geo = poly_geo.buffer(0)
poly_geo.is_valid
# project polygon to calculate area
poly_prj=ox.project_gdf(poly)
area=float(poly_prj.area)
# pull network
G = ox.graph_from_polygon(poly_geo, network_type='drive', simplify=True, clean_periphery=True)
if len(G.nodes(data=True))>100:
# project it and calculate statistics
G_proj = ox.project_graph(G)
stats = ox.basic_stats(G_proj, area=area, circuity_dist='euclidean')
# delete the no longer needed dict elements
del stats['streets_per_node_counts']
del stats['streets_per_node_proportion']
# load as a pandas dataframe
df = pd.DataFrame.from_dict(stats, orient='index')
df.columns= [barrier]
df.to_csv('{0}/Results/Statistics/{1}.csv'.format(path,barrier))
| Notebooks/Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pyreadstat
#conda install -c conda-forge pyreadstat
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
import datetime
#graphing
import matplotlib.pyplot as plt
#stats
import statsmodels.api as sm
from statsmodels.base.model import GenericLikelihoodModel
#import testing
import sys
sys.path.append("../")
import selection_tests
# -
dataframe, meta = pyreadstat.read_dta('Selec42.dta')
dataframe.to_csv('Selec42.csv')
print(dataframe.dropna().shape)
print(dataframe.columns)
dataframe, meta = pyreadstat.read_dta('ssdata99.dta')
dataframe.to_csv('ssdata99.csv')
print(dataframe.dropna().shape)
print(dataframe.columns)
# +
# ln st = stock
# ct = ln wholsale price
# bt = pt-1 - ct
#ln yt-1 demand? 'x',
y_names = ['tp']
x_names = ['c','s','markup']
data = dataframe[x_names+y_names]
y = data[y_names]
x = data[x_names]
probit = sm.Probit(y,x)
probit_fit = probit.fit()
probit_fit.summary()
# -
| am/summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:replay_trajectory_classification] *
# language: python
# name: conda-env-replay_trajectory_classification-py
# ---
# ### Introduction ###
#
# There are four decoders included in the **replay_trajectory_classification** package:
# + `SortedSpikesDecoder`
# + `ClusterlessDecoder`
# + `SortedSpikesClassifier`
# + `ClusterlessClassifier`.
#
# You can import them for use by running the following lines of code:
from replay_trajectory_classification import (SortedSpikesDecoder, ClusterlessDecoder,
SortedSpikesClassifier, ClusterlessClassifier)
# If `Decoder` is in the name it indicates that a single movement model is used to decode position.
#
# If `Classifier` is in the name it means that multiple movement models are used to decode position and classify the type of movement.
#
# `SortedSpikes` and `Clusterless` indicate the type of data the decoder uses:
# + `SortedSpikes` means the decoder takes in clustered single unit data.
# + `Clusterless` means the decoder takes in unsorted multiunit spikes and associated spike waveform features---like those that would usually be used to spike sort such as spike width, spike amplitude, etc.
#
#
# Next, we will explore the expected data format for `SortedSpikes` and `Clusterless` decoders in turn. The both the `Decoder` and `Classifier` take the same data format.
#
# ### Sorted Spikes Data Format ###
#
# We can see from the docstring of the fit function of `SortedSpikesDecoder` that there are two main variables that are required:
# + **position** - the 1D or 2D position of the animal at each time bin
# + **spikes** - 0 if there is no spike and 1 if there is a spike for each cell for each time bin
#
# **NOTE**: Time bins should be small (typically 1 or 2 ms)
# +
# SortedSpikesDecoder.fit?
# -
# #### Example Data ####
# Let's look at some simulated data to get an intuition for these data types. We simulate spikes of 19 cells with 360,000 time bins (360 seconds) of data:
# +
from replay_trajectory_classification.sorted_spikes_simulation import make_simulated_run_data
time, position, sampling_frequency, spikes, place_fields = make_simulated_run_data()
# -
# We can see that this numpy array is a matrix with time bins in the first dimension and cells on the second dimension
spikes.shape
# and that there is a 1 each time a cell spikes and a 0 each time there is not a spike. For example, for the first cell, there is only one spike at 17 ms in the first 100 ms of time:
spikes[:100, 0]
# We can visualize the spikes as a spike raster:
# +
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
fig, ax = plt.subplots(figsize=(12, 5))
spike_ind, neuron_ind = np.nonzero(spikes)
cmap = plt.get_cmap('tab20')
c = [cmap.colors[ind] for ind in neuron_ind]
ax.scatter(time[spike_ind], neuron_ind + 1, c=c, s=5)
ax.set_yticks((1, spikes.shape[1]))
ax.set_ylim((1, spikes.shape[1]))
ax.set_ylabel('Cells')
ax.set_xlabel('Time [s]')
ax.set_xlim((0.0, 90.0))
sns.despine(offset=5)
# -
# We also have a position variable which has the same number of time points and has one position dimension:
position.shape
# We can plot position over time:
# +
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(time, position, linewidth=2)
ax.set_ylabel("Position [cm]")
ax.set_xlabel("Time [s]")
sns.despine(offset=5)
# -
# #### Summary ####
# So we have two numpy arrays that contain the data we need to fit and evaluate the model:
spikes.shape
position.shape
# ### Clusterless Data Format ###
#
# The `Clusterless` decoders take the same information as the `SortedSpikes` decoders with one exception: the actual multiunit spikes and their associated waveform features. This variable is called **multiunits** and it has time bins on the first dimension, the marks (or associated wave form features) in the second dimension, and the electrodes in the last dimension.
# +
# ClusterlessDecoder.fit?
# -
# #### Example Data ####
#
# For the multiunits array, if there is no spike in a particular time bin, there are only NaNs. If there is a spike, then the values of the associated waveform feature are filled in. Let's loook at some example simulated data:
# +
from replay_trajectory_classification.clusterless_simulation import make_simulated_run_data
(time, position, sampling_frequency,
multiunits, _) = make_simulated_run_data()
# -
# We can see that there are 350,000 time bins, 4 mark dimensions (wave form features), and 5 electrodes:
multiunits.shape
# If we look at the first electrode, we see that there is a spike in the first time bin and then no spike for the next 9 time bins (hence they are filled with NaNs):
multiunits[:10, :, 0]
# These four values represent waveform features of this electrode at the time of the spike. Likewise, if we look at the second electrode, we can see that there was no spike in the first 10 time bins.
multiunits[:10, :, 1]
# ### Track Graph ###
#
# If you are using 1D position but want to represent an environment more complicated than a linear track, you need to constract a graph representation of the track that indicates how the 1D position can connect. We refer to this as the **track_graph**. This is done using the NetworkX package.
# Now we define the x- and y-position of the nodes. The nodes are automatically named by their order:
# + node "0" has position (0, 0)
# + node "1" has position (1, 0)
# + node "2" has position (1, 1)
# + node "3" has position (0, 1)
node_positions = [(0, 0), # xy position of node 0
(1, 0), # xy position of node 1
(1, 1), # xy position of node 2
(0, 1), # xy position of node 3
]
# Then we can define how the nodes are connected by their node names. So if we want node "0" and node "1" to be connected, then we specify (0, 1)
edges = [(0, 1), # connects node 0 and node 1
(1, 2), # connects node 1 and node 2
(2, 3), # connects node 2 and node 3
]
# Then we can construct a graph of the track by using the function `make_track_graph`, which will return a NetworkX Graph object:
# +
from replay_trajectory_classification import make_track_graph
track_graph = make_track_graph(node_positions, edges)
track_graph
# -
track_graph.nodes
track_graph.edges
# We can visualize the **track_graph** by using our `plot_track_graph` function:
# +
from replay_trajectory_classification import plot_track_graph
fig, ax = plt.subplots()
plot_track_graph(track_graph, ax=ax)
ax.tick_params(left=True, bottom=True, labelleft=True, labelbottom=True)
ax.set_xlabel("x-position")
ax.set_ylabel("y-position")
sns.despine(offset=5)
# -
# We can change how these points are connected by changing the edges:
# +
edges = [(0, 1), # connects node 0 and node 1
(0, 3), # connects node 0 and node 3
(1, 2), # connects node 1 and node 2
]
track_graph = make_track_graph(node_positions, edges)
fig, ax = plt.subplots()
plot_track_graph(track_graph, ax=ax)
ax.tick_params(left=True, bottom=True, labelleft=True, labelbottom=True)
ax.set_xlabel("x-position")
ax.set_ylabel("y-position")
sns.despine(offset=5)
# -
# We could also change the node positions:
# +
node_positions = [(5.5, 7), # xy position of node 0
(1, -5), # xy position of node 1
(1, 1), # xy position of node 2
(0, 1), # xy position of node 3
]
track_graph = make_track_graph(node_positions, edges)
fig, ax = plt.subplots()
plot_track_graph(track_graph, ax=ax)
ax.tick_params(left=True, bottom=True, labelleft=True, labelbottom=True)
ax.set_xlabel("x-position")
ax.set_ylabel("y-position")
sns.despine(offset=5)
# -
# There are two other parameters when using `track_graph` with the decoders. These specify how the edges correspond to 1D position:
# + **edge_order** -- The order of the edges in 1D position in terms of their node pairs e.g. (0, 1) represents the edge connecting node 0 to node 1.
# + **edge_spacing** -- The spacing between the edges. This is necessary when edges are not connected to each other in 2D space, because the smoothing used by the model will inappropriately smooth into a non-adjacent edge. This will always be one less than the number of edges.
#
#
# For example, we might linearize the above track by doing the following by specifying the following edge order and spacing:
edge_order = [(2, 1), # node 2 to node 1
(1, 0), # node 1 to node 0
(0, 3), # node 0 to node 3
]
edge_spacing = 0 # no spacing between edges
# We can visualize the linearization layout by using `plot_graph_as_1D`:
# +
from replay_trajectory_classification import plot_graph_as_1D
fig, ax = plt.subplots(figsize=(10, 1))
plot_graph_as_1D(track_graph, edge_order, edge_spacing,
ax=ax)
# -
# If we wanted, we could put spaces between each edge. For example if we want 5 cm spaces between edges:
# +
edge_order = [(2, 1), # node 2 to node 1
(1, 0), # node 1 to node 0
(0, 3), # node 0 to node 3
]
edge_spacing = 5
fig, ax = plt.subplots(figsize=(10, 1))
plot_graph_as_1D(track_graph, edge_order, edge_spacing,
ax=ax)
# -
# We can also change the spacing by depending on the edges:
# +
edge_order = [(2, 1), # node 2 to node 1
(1, 0), # node 1 to node 0
(0, 3), # node 0 to node 3
]
edge_spacing = [0, 5]
fig, ax = plt.subplots(figsize=(10, 1))
plot_graph_as_1D(track_graph, edge_order, edge_spacing,
ax=ax)
# -
# Now let's try a more complicated track, where you should put spacing in between edges:
# +
node_positions = [(-1, 1), # xy position of node 0
(-1, 0), # xy position of node 1
(0, 0), # xy position of node 2 *
(0, 1), # xy position of node 3
(1, 0), # xy position of node 4
(1, 1), # xy position of node 5
]
edges = [(0, 1), # connects node 0 and node 1
(1, 2), # connects node 1 and node 2
(2, 3), # connects node 2 and node 3
(2, 4), # connects node 2 and node 4
(4, 5), # connects node 4 and node 5
]
track_graph = make_track_graph(node_positions, edges)
fig, ax = plt.subplots()
plot_track_graph(track_graph, ax=ax, draw_edge_labels=True)
ax.tick_params(left=True, bottom=True, labelleft=True, labelbottom=True)
ax.set_xlabel("x-position")
ax.set_ylabel("y-position")
sns.despine(offset=5)
# -
# In this case, the edge 2 (between nodes 2 and 3) has to be placed separately from the other edges in 1D position, so a gap is needed. One possible scheme would be:
# +
center_well_id = 3
edge_order = [(3, 2),
(2, 1),
(1, 0),
(2, 4),
(4, 5)
]
edge_spacing = [0, 0, 1, 0]
fig, ax = plt.subplots(figsize=(10, 1))
plot_graph_as_1D(track_graph, edge_order, edge_spacing,
ax=ax)
# -
# The edge spacing is 0 between edges 2 and edges 1 because they are connected. Likewise the edge spacing is 0 between edges 1 and 0 because they are connected.
#
# **Importantly**, the edge spacing between edges 0 and 3 is 1 because edge 0 and 3 are not directly connected and we don't want to smooth positions on edge 0 onto edge 3. Note that this depends on the size of your bandwidth (for the `Clusterless` decoder) or the knot spacing (for the `SortedSpikesDecoder`).
#
# Finally let's try a track shaped like a circle.
# +
angle = np.linspace(-np.pi, np.pi, num=12, endpoint=False)
radius = 2
node_positions = np.stack((radius * np.cos(angle), radius * np.sin(angle)), axis=1)
node_ids = np.arange(node_positions.shape[0])
edges = np.stack((node_ids, np.roll(node_ids, shift=1)), axis=1)
track_graph = make_track_graph(node_positions, edges)
fig, ax = plt.subplots(figsize=(10, 10))
plot_track_graph(track_graph, ax=ax, draw_edge_labels=True)
ax.tick_params(left=True, bottom=True, labelleft=True, labelbottom=True)
ax.set_xlabel("x-position")
ax.set_ylabel("y-position")
sns.despine(offset=5)
# +
edge_spacing = 0
n_nodes = len(track_graph.nodes)
edge_order = np.stack((np.roll(np.arange(n_nodes-1, -1, -1), 1),
np.arange(n_nodes-1, -1, -1)), axis=1)
fig, ax = plt.subplots(figsize=(n_nodes // 2, 1))
plot_graph_as_1D(track_graph,
edge_spacing=edge_spacing,
edge_order=edge_order,
ax=ax)
# -
edge_order
| notebooks/tutorial/01-Introduction_and_Data_Format.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
from scipy import interpolate
import scipy.integrate as integrate
import pandas as pd
import cosmo
# -
import imp
imp.reload(cosmo)
cosmo.PS
my_zred_out = 6.0
imp.reload(cosmo)
aa,bb, cc = cosmo.LyALAEPkl(5.0)
plt.plot(cosmo.fk,aa)
plt.plot(cosmo.fk,bb)
plt.loglog()
# +
imp.reload(cosmo)
r2, r2xil, r2xil_nsn = cosmo.LyALAEXil_fft(6.0)
# -
plt.figure(figsize=(10,8))
plt.rcParams.update({'font.size': 22})
plt.plot(r2,r2xil_nsn)
plt.loglog()
cosmo.LyALAEPkl(zred_out=6,b_LAE=3.0,b_delta=0.707,b_Gamma=-0.173,tau_eff=6, lorder=0)
| Ly_alpha_emitters2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Notebook para o PAN - Atribuição Autoral - 2018
# +
# %matplotlib inline
#python basic libs
from __future__ import print_function
from tempfile import mkdtemp
from shutil import rmtree
import os;
from os.path import join as pathjoin;
import re;
import glob;
import json;
import codecs;
from collections import defaultdict;
import pprint;
from pprint import pprint
from time import time
import logging
#data analysis libs
import numpy as np;
import pandas as pd;
import matplotlib.pyplot as plt;
import random;
#machine learning libs
#feature extraction
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
#preprocessing and transformation
from sklearn.preprocessing import normalize, MaxAbsScaler, MinMaxScaler;
from sklearn.preprocessing import LabelBinarizer;
from sklearn.decomposition import PCA;
from sklearn.metrics.pairwise import cosine_similarity;
from sklearn.base import BaseEstimator, ClassifierMixin
#classifiers
from sklearn.svm import LinearSVC, SVC
from sklearn.multiclass import OneVsOneClassifier, OneVsRestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.neural_network import MLPClassifier
from sklearn.feature_selection import RFE,SelectFpr,SelectPercentile, chi2;
#
from sklearn.ensemble import AdaBoostClassifier, BaggingClassifier
from sklearn.ensemble import VotingClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
#model valuation
from sklearn.model_selection import train_test_split;
from sklearn.metrics import roc_auc_score, f1_score, precision_score, recall_score, accuracy_score;
# -
import platform; print(platform.platform())
print("NumPy", np.__version__)
import scipy; print("SciPy", scipy.__version__)
import sklearn; print("Scikit-Learn", sklearn.__version__)
# ### paths configuration
# +
baseDir = '/Users/joseeleandrocustodio/Dropbox/mestrado/02 - Pesquisa/code';
inputDir= pathjoin(baseDir,'pan18aa');
outputDir= pathjoin(baseDir,'out',"oficial");
if not os.path.exists(outputDir):
os.mkdir(outputDir);
# -
# ## loading the dataset
def readCollectionsOfProblems(path):
# Reading information about the collection
infocollection = path+os.sep+'collection-info.json'
with open(infocollection, 'r') as f:
problems = [
{
'problem': attrib['problem-name'],
'language': attrib['language'],
'encoding': attrib['encoding'],
}
for attrib in json.load(f)
]
return problems;
problems = readCollectionsOfProblems(inputDir);
problems[0]
def readProblem(path, problem):
# Reading information about the problem
infoproblem = path+os.sep+problem+os.sep+'problem-info.json'
candidates = []
with open(infoproblem, 'r') as f:
fj = json.load(f)
unk_folder = fj['unknown-folder']
for attrib in fj['candidate-authors']:
candidates.append(attrib['author-name'])
return unk_folder, candidates;
def read_files(path,label):
# Reads all text files located in the 'path' and assigns them to 'label' class
files = glob.glob(pathjoin(path,label,'*.txt'))
texts=[]
for i,v in enumerate(files):
f=codecs.open(v,'r',encoding='utf-8')
texts.append((f.read(),label, os.path.basename(v)))
f.close()
return texts
for index,problem in enumerate(problems):
unk_folder, candidates_folder = readProblem(inputDir, problem['problem']);
problem['candidates_folder_count'] = len(candidates_folder);
problem['candidates'] = [];
for candidate in candidates_folder:
problem['candidates'].extend(read_files(pathjoin(inputDir, problem['problem']),candidate));
problem['unknown'] = read_files(pathjoin(inputDir, problem['problem']),unk_folder);
pd.DataFrame(problems)
# +
#*******************************************************************************************************
import warnings
from sklearn.metrics import f1_score, precision_score, recall_score, accuracy_score
from sklearn.preprocessing import LabelEncoder
def eval_measures(gt, pred):
"""Compute macro-averaged F1-scores, macro-averaged precision,
macro-averaged recall, and micro-averaged accuracy according the ad hoc
rules discussed at the top of this file.
Parameters
----------
gt : dict
Ground truth, where keys indicate text file names
(e.g. `unknown00002.txt`), and values represent
author labels (e.g. `candidate00003`)
pred : dict
Predicted attribution, where keys indicate text file names
(e.g. `unknown00002.txt`), and values represent
author labels (e.g. `candidate00003`)
Returns
-------
f1 : float
Macro-averaged F1-score
precision : float
Macro-averaged precision
recall : float
Macro-averaged recall
accuracy : float
Micro-averaged F1-score
"""
actual_authors = list(gt.values())
encoder = LabelEncoder().fit(['<UNK>'] + actual_authors)
text_ids, gold_authors, silver_authors = [], [], []
for text_id in sorted(gt):
text_ids.append(text_id)
gold_authors.append(gt[text_id])
try:
silver_authors.append(pred[text_id])
except KeyError:
# missing attributions get <UNK>:
silver_authors.append('<UNK>')
assert len(text_ids) == len(gold_authors)
assert len(text_ids) == len(silver_authors)
# replace non-existent silver authors with '<UNK>':
silver_authors = [a if a in encoder.classes_ else '<UNK>'
for a in silver_authors]
gold_author_ints = encoder.transform(gold_authors)
silver_author_ints = encoder.transform(silver_authors)
# get F1 for individual classes (and suppress warnings):
with warnings.catch_warnings():
warnings.simplefilter('ignore')
f1 = f1_score(gold_author_ints,
silver_author_ints,
labels=list(set(gold_author_ints)),
average='macro')
precision = precision_score(gold_author_ints,
silver_author_ints,
labels=list(set(gold_author_ints)),
average='macro')
recall = recall_score(gold_author_ints,
silver_author_ints,
labels=list(set(gold_author_ints)),
average='macro')
accuracy = accuracy_score(gold_author_ints,
silver_author_ints)
return f1,precision,recall,accuracy
# -
def evaluate(ground_truth_file,predictions_file):
# Calculates evaluation measures for a single attribution problem
gt = {}
with open(ground_truth_file, 'r') as f:
for attrib in json.load(f)['ground_truth']:
gt[attrib['unknown-text']] = attrib['true-author']
pred = {}
with open(predictions_file, 'r') as f:
for attrib in json.load(f):
if attrib['unknown-text'] not in pred:
pred[attrib['unknown-text']] = attrib['predicted-author']
f1,precision,recall,accuracy = eval_measures(gt,pred)
return f1, precision, recall, accuracy
# +
from sklearn.base import BaseEstimator
from scipy.sparse import issparse
class DenseTransformer(BaseEstimator):
"""Convert a sparse array into a dense array."""
def __init__(self, return_copy=True):
self.return_copy = return_copy
self.is_fitted = False
def transform(self, X, y=None):
""" Return a dense version of the input array.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
Training vectors, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape = [n_samples] (default: None)
Returns
---------
X_dense : dense version of the input X array.
"""
if issparse(X):
return X.toarray()
elif self.return_copy:
return X.copy()
else:
return X
def fit(self, X, y=None):
""" Mock method. Does nothing.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
Training vectors, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape = [n_samples] (default: None)
Returns
---------
self
"""
self.is_fitted = True
return self
def fit_transform(self, X, y=None):
""" Return a dense version of the input array.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
Training vectors, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape = [n_samples] (default: None)
Returns
---------
X_dense : dense version of the input X array.
"""
return self.transform(X=X, y=y)
# +
from sklearn.base import BaseEstimator
from scipy.sparse import issparse
class ObfuscationTransformer(BaseEstimator):
def __init__(self,re_from=r'(\b)(\w{0,2})\w+(\w{1,3})(\b)', re_to=r'\1\2XX\3\4', return_copy=True):
self.re_from = re_from
self.re_to = re_to
def transform(self, X, y=None):
X = np.array(X).copy();
for i in range(len(X)):
X[i] = re.sub(self.re_from,self.re_to, X[i])
return X;
def fit(self, X, y=None):
return self
def fit_transform(self, X, y=None):
return self.transform(X=X, y=y)
# -
def runML(problem):
print ("\nProblem: %s, language: %s, " %(problem['problem'],problem['language']))
train_docs, train_labels, _ = zip(*problem['candidates'])
problem['training_docs_size'] = len(train_docs);
test_docs, _, test_filename = zip(*problem['unknown'])
cachedir = mkdtemp()
pipeline = Pipeline([
('obs',ObfuscationTransformer(re_from=r'\w',re_to='x')),
('vect', TfidfVectorizer(analyzer='char',
min_df=0.05,
max_df=1.0,
norm='l2',
lowercase =False,
sublinear_tf=True)),
('dense', DenseTransformer()),
('scaler', MaxAbsScaler()),
('transf', PCA(0.99)),
('clf', LogisticRegression(random_state=0,multi_class='multinomial', solver='newton-cg')),
], memory=cachedir)
# uncommenting more parameters will give better exploring power but will
# increase processing time in a combinatorial way
parameters = {
'vect__ngram_range':((1,2),(2,3),(2,4),(2,5),(3,5)),
'transf__n_components': (0.1,0.5,0.9,0.99),
}
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=False)
print("Performing grid search...")
t0 = time()
grid_search.fit(train_docs, train_labels)
print("done in %0.3fs" % (time() - t0))
print("Best score: %0.3f" % grid_search.best_score_)
print("Best parameters set:")
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("\t%s: %r" % (param_name, best_parameters[param_name]))
train_pred=grid_search.predict(train_docs);
test_pred=grid_search.predict(test_docs);
# Writing output file
out_data=[]
for i,v in enumerate(test_pred):
out_data.append({'unknown-text': test_filename[i],'predicted-author': v})
answerFile = pathjoin(outputDir,'answers-'+problem['problem']+'.json');
with open(answerFile, 'w') as f:
json.dump(out_data, f, indent=4)
#allProblems.extend(out_data)
#evaluation train
f1,precision,recall,accuracy=evaluate(
pathjoin(inputDir, problem['problem'], 'ground-truth.json'),
answerFile)
rmtree(cachedir)
return {
'problem-name' : problem['problem'],
"language" : problem['language'],
'AuthorCount' : len(set(train_labels)),
"train_doc_size": len(train_docs),
"train_caract_per_doc": sum([len(l) for l in train_docs])/len(train_docs),
"test_doc_size" : len(test_docs),
"test_caract_per_doc": sum([len(l) for l in test_docs])/len(test_docs),
'macro-f1' : round(f1,3),
'macro-precision': round(precision,3),
'macro-recall' : round(recall,3),
'micro-accuracy' : round(accuracy,3),
}, grid_search.cv_results_;
result = [];
cv_result = [];
for problem in problems:
r, c = runML(problem);
result.append(r);
cv_result.append(c);
df=pd.DataFrame(result)[['problem-name',
"language",
'AuthorCount',
"train_doc_size","train_caract_per_doc",
"test_doc_size", "test_caract_per_doc",
'macro-f1','macro-precision','macro-recall' ,'micro-accuracy']]
df
print(df[["macro-f1"]].reset_index().to_latex(index=False).replace(" "," "))
pd.DataFrame(result)[['macro-f1']].describe()
pd.DataFrame(result)\
.sort_values(by=['language','problem-name'])[['language','problem-name','macro-f1']]\
.plot(kind='bar', x=['language','problem-name'], legend=True, figsize=(20,5))
print(ObfuscationTransformer(re_from=r'\w',re_to='x').fit_transform(["Verdes mares bravios ç ç\n\n."])[0])
print(problems[8]['candidates'][0][0][0:500])
print(ObfuscationTransformer(re_from=r'\w',re_to='x').fit_transform([problems[8]['candidates'][0][0][0:500]])[0])
| 2018/PAN_AA_2018-Obfuscation.ipynb |