code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Use Decision Optimization to plan your diet with `ibm-watson-machine-learning`
#
# This notebook facilitates Decision Optimization and Watson Machine Learning services. It contains steps and code to work with [ibm-watson-machine-learning](https://pypi.python.org/pypi/ibm-watson-machine-learning) library available in PyPI repository. It also introduces commands for getting model and training data, persisting model, deploying model and scoring it.
#
# Some familiarity with Python is helpful. This notebook uses Python 3.8.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Learning goals
#
# The learning goals of this notebook are:
#
# - Load a DO model file into an Watson Machine learning repository.
# - Prepare data for training and evaluation.
# - Create an DO machine learning job.
# - Persist a DO model Watson Machine Learning repository.
# - Deploy a model for batch scoring using Wastson Machine Learning API.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Contents
#
# This notebook contains the following parts:
#
# 1. [Setup](#setup)
# 2. [Download externally created DO model](#download)
# 3. [Persist externally created DO model](#upload)
# 4. [Deploy](#deploy)
# 5. [Create job](#score)
# 6. [Clean up](#cleanup)
# 7. [Summary and next steps](#summary)
#
#
# -
# <a id="setup"></a>
# ## 1. Set up the environment
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# - Contact with your Cloud Pack for Data administrator and ask him for your account credentials
# ### Connection to WML
#
# Authenticate the Watson Machine Learning service on IBM Cloud Pack for Data. You need to provide platform `url`, your `username` and `api_key`.
username = 'PASTE YOUR USERNAME HERE'
api_key = 'PASTE YOUR API_KEY HERE'
url = 'PASTE THE PLATFORM URL HERE'
wml_credentials = {
"username": username,
"apikey": api_key,
"url": url,
"instance_id": 'openshift',
"version": '4.0'
}
# Alternatively you can use `username` and `password` to authenticate WML services.
#
# ```
# wml_credentials = {
# "username": ***,
# "password": ***,
# "url": ***,
# "instance_id": 'openshift',
# "version": '4.0'
# }
#
# ```
# ### Install and import the `ibm-watson-machine-learning` package
# **Note:** `ibm-watson-machine-learning` documentation can be found <a href="http://ibm-wml-api-pyclient.mybluemix.net/" target="_blank" rel="noopener no referrer">here</a>.
# !pip install -U ibm-watson-machine-learning
# +
from ibm_watson_machine_learning import APIClient
client = APIClient(wml_credentials)
# -
# ### Working with spaces
#
# First of all, you need to create a space that will be used for your work. If you do not have space already created, you can use `{PLATFORM_URL}/ml-runtime/spaces?context=icp4data` to create one.
#
# - Click New Deployment Space
# - Create an empty space
# - Go to space `Settings` tab
# - Copy `space_id` and paste it below
#
# **Tip**: You can also use SDK to prepare the space for your work. More information can be found [here](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Space%20management.ipynb).
#
# **Action**: Assign space ID below
space_id = 'PASTE YOUR SPACE ID HERE'
# You can use `list` method to print all existing spaces.
client.spaces.list(limit=10)
# To be able to interact with all resources available in Watson Machine Learning, you need to set **space** which you will be using.
client.set.default_space(space_id)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### <a id="download"></a>
# ## 2. Download externally created DO model and data
#
#
# In this section, you will download externally created DO models and data used for training it.
#
#
# **Action**: Get your DO model.
# -
# !wget https://github.com/IBM/watson-machine-learning-samples/raw/master/cpd4.0/models/decision_optimization/do-model.tar.gz \
# -O do-model.tar.gz
# + pycharm={"name": "#%%\n"}
model_path = 'do-model.tar.gz'
# -
#
# <a id="upload"></a>
# ## 3. Persist externally created DO model
# In this section, you will learn how to store your model in Watson Machine Learning repository by using the Watson Machine Learning Client.
#
# ### 3.1: Publish model
#
# #### Publish model in Watson Machine Learning repository on Cloud.
# Define model name, autor name and email.
#
# + [markdown] pycharm={"name": "#%% md\n"}
# Get software specification for DO model
# + pycharm={"name": "#%%\n"}
sofware_spec_uid = client.software_specifications.get_uid_by_name("do_12.9")
# + [markdown] pycharm={"name": "#%% md\n"}
# Output data schema for storing model in WML repository
# + pycharm={"name": "#%%\n"}
output_data_schema = [{'id': 'stest',
'type': 'list',
'fields': [{'name': 'age', 'type': 'float'},
{'name': 'sex', 'type': 'float'},
{'name': 'cp', 'type': 'float'},
{'name': 'restbp', 'type': 'float'},
{'name': 'chol', 'type': 'float'},
{'name': 'fbs', 'type': 'float'},
{'name': 'restecg', 'type': 'float'},
{'name': 'thalach', 'type': 'float'},
{'name': 'exang', 'type': 'float'},
{'name': 'oldpeak', 'type': 'float'},
{'name': 'slope', 'type': 'float'},
{'name': 'ca', 'type': 'float'},
{'name': 'thal', 'type': 'float'}]
}, {'id': 'teste2',
'type': 'test',
'fields': [{'name': 'age', 'type': 'float'},
{'name': 'sex', 'type': 'float'},
{'name': 'cp', 'type': 'float'},
{'name': 'restbp', 'type': 'float'},
{'name': 'chol', 'type': 'float'},
{'name': 'fbs', 'type': 'float'},
{'name': 'restecg', 'type': 'float'},
{'name': 'thalach', 'type': 'float'},
{'name': 'exang', 'type': 'float'},
{'name': 'oldpeak', 'type': 'float'},
{'name': 'slope', 'type': 'float'},
{'name': 'ca', 'type': 'float'},
{'name': 'thal', 'type': 'float'}]}]
# + pycharm={"name": "#%%\n"}
model_meta_props = {
client.repository.ModelMetaNames.NAME: "LOCALLY created DO model",
client.repository.ModelMetaNames.TYPE: "do-docplex_12.9",
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: sofware_spec_uid,
client.repository.ModelMetaNames.OUTPUT_DATA_SCHEMA: output_data_schema
}
published_model = client.repository.store_model(model=model_path, meta_props=model_meta_props)
# -
# **Note:** You can see that model is successfully stored in Watson Machine Learning Service.
# ### 3.2: Get model details
#
# +
import json
published_model_uid = client.repository.get_model_uid(published_model)
model_details = client.repository.get_details(published_model_uid)
print(json.dumps(model_details, indent=2))
# -
# ### 3.3 Get all models
# + pycharm={"name": "#%%\n"}
client.repository.list_models()
# -
# <a id="deploy"></a>
# ## 4. Deploy
# In this section you will learn how to create batch deployment to create job using the Watson Machine Learning Client.
#
# You can use commands bellow to create batch deployment for stored model (web service).
#
# ### 4.1: Create model deployment
#
# + pycharm={"name": "#%%\n"}
meta_data = {
client.deployments.ConfigurationMetaNames.NAME: "deployment_DO",
client.deployments.ConfigurationMetaNames.BATCH: {},
client.deployments.ConfigurationMetaNames.HARDWARE_SPEC: {"name": "S", "num_nodes": 1}
}
deployment_details = client.deployments.create(published_model_uid, meta_props=meta_data)
# -
# **Note**: Here we use deployment url saved in published_model object. In next section, we show how to retrive deployment url from Watson Machine Learning instance.
#
deployment_uid = client.deployments.get_uid(deployment_details)
# Now, You can list all deployments.
# + pycharm={"name": "#%%\n"}
client.deployments.list()
# -
# ### 4.2: Get deployment details
#
# + pycharm={"name": "#%%\n"}
client.deployments.get_details(deployment_uid)
# -
# <a id="score"></a>
# ## 5. Create job
#
# You can create job to web-service deployment using `create_job` method.
# Prepare test data
# +
import pandas as pd
diet_food = pd.DataFrame([["Roasted Chicken", 0.84, 0, 10],
["Spaghetti W/ Sauce", 0.78, 0, 10],
["Tomato,Red,Ripe,Raw", 0.27, 0, 10],
["Apple,Raw,W/Skin", 0.24, 0, 10],
["Grapes", 0.32, 0, 10],
["Chocolate Chip Cookies", 0.03, 0, 10],
["Lowfat Milk", 0.23, 0, 10],
["Raisin Brn", 0.34, 0, 10],
["Hotdog", 0.31, 0, 10]], columns=["name", "unit_cost", "qmin", "qmax"])
diet_food_nutrients = pd.DataFrame([
["Spaghetti W/ Sauce", 358.2, 80.2, 2.3, 3055.2, 11.6, 58.3, 8.2],
["Roasted Chicken", 277.4, 21.9, 1.8, 77.4, 0, 0, 42.2],
["Tomato,Red,Ripe,Raw", 25.8, 6.2, 0.6, 766.3, 1.4, 5.7, 1],
["Apple,Raw,W/Skin", 81.4, 9.7, 0.2, 73.1, 3.7, 21, 0.3],
["Grapes", 15.1, 3.4, 0.1, 24, 0.2, 4.1, 0.2],
["Chocolate Chip Cookies", 78.1, 6.2, 0.4, 101.8, 0, 9.3, 0.9],
["Lowfat Milk", 121.2, 296.7, 0.1, 500.2, 0, 11.7, 8.1],
["Raisin Brn", 115.1, 12.9, 16.8, 1250.2, 4, 27.9, 4],
["Hotdog", 242.1, 23.5, 2.3, 0, 0, 18, 10.4]
], columns=["Food", "Calories", "Calcium", "Iron", "Vit_A", "Dietary_Fiber", "Carbohydrates", "Protein"])
diet_nutrients = pd.DataFrame([
["Calories", 2000, 2500],
["Calcium", 800, 1600],
["Iron", 10, 30],
["Vit_A", 5000, 50000],
["Dietary_Fiber", 25, 100],
["Carbohydrates", 0, 300],
["Protein", 50, 100]
], columns=["name", "qmin", "qmax"])
# -
job_payload_ref = {
client.deployments.DecisionOptimizationMetaNames.INPUT_DATA: [
{
"id": "diet_food.csv",
"values": diet_food
},
{
"id": "diet_food_nutrients.csv",
"values": diet_food_nutrients
},
{
"id": "diet_nutrients.csv",
"values": diet_nutrients
}
],
client.deployments.DecisionOptimizationMetaNames.OUTPUT_DATA: [
{
"id": ".*.csv"
}
]
}
# Create job using Watson Machine Learning client
job = client.deployments.create_job(deployment_uid, meta_props=job_payload_ref)
# Checking created job status and calculated KPI.
# +
import time
job_id = client.deployments.get_job_uid(job)
elapsed_time = 0
while client.deployments.get_job_status(job_id).get('state') != 'completed' and elapsed_time < 300:
elapsed_time += 10
time.sleep(10)
if client.deployments.get_job_status(job_id).get('state') == 'completed':
job_details_do = client.deployments.get_job_details(job_id)
kpi = job_details_do['entity']['decision_optimization']['solve_state']['details']['KPI.Total Calories']
print(f"KPI: {kpi}")
else:
print("Job hasn't completed successfully in 5 minutes.")
# -
# <a id="cleanup"></a>
# ## 6. Clean up
# If you want to clean up all created assets:
# - experiments
# - trainings
# - pipelines
# - model definitions
# - models
# - functions
# - deployments
#
# please follow up this sample [notebook](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Machine%20Learning%20artifacts%20management.ipynb).
# <a id="summary"></a>
# ## 7. Summary and next steps
# You successfully completed this notebook! You learned how to use DO as well as Watson Machine Learning for model creation and deployment.
#
# Check out our _[Online Documentation](https://dataplatform.cloud.ibm.com/docs/content/analyze-data/wml-setup.html)_ for more samples, tutorials, documentation, how-tos, and blog posts.
#
# ### Authors
#
# **<NAME>**, Software Engineer
# Copyright © 2020, 2021 IBM. This notebook and its source code are released under the terms of the MIT License.
#
| cpd4.0/notebooks/python_sdk/deployments/decision_optimization/Use Decision Optimization to plan your diet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Seminar and homework (10 points total)
#
# Today we shall compose encoder-decoder neural networks and apply them to the task of machine translation.
#
# 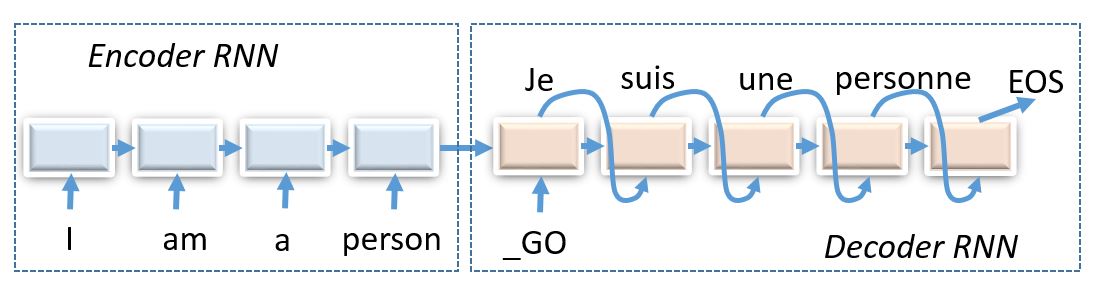
# _(img: esciencegroup.files.wordpress.com)_
#
#
# Encoder-decoder architectures are about converting anything to anything, including
# * Machine translation and spoken dialogue systems
# * [Image captioning](http://mscoco.org/dataset/#captions-challenge2015) and [image2latex](https://openai.com/requests-for-research/#im2latex) (convolutional encoder, recurrent decoder)
# * Generating [images by captions](https://arxiv.org/abs/1511.02793) (recurrent encoder, convolutional decoder)
# * Grapheme2phoneme - convert words to transcripts
# ## Our task: machine translation
#
# We gonna try our encoder-decoder models on russian to english machine translation problem. More specifically, we'll translate hotel and hostel descriptions. This task shows the scale of machine translation while not requiring you to train your model for weeks if you don't use GPU.
#
# Before we get to the architecture, there's some preprocessing to be done. ~~Go tokenize~~ Alright, this time we've done preprocessing for you. As usual, the data will be tokenized with WordPunctTokenizer.
#
# However, there's one more thing to do. Our data lines contain unique rare words. If we operate on a word level, we will have to deal with large vocabulary size. If instead we use character-level models, it would take lots of iterations to process a sequence. This time we're gonna pick something inbetween.
#
# One popular approach is called [Byte Pair Encoding](https://github.com/rsennrich/subword-nmt) aka __BPE__. The algorithm starts with a character-level tokenization and then iteratively merges most frequent pairs for N iterations. This results in frequent words being merged into a single token and rare words split into syllables or even characters.
#
#
# !pip3 install subword-nmt &> log
# !wget https://raw.githubusercontent.com/yandexdataschool/nlp_course/master/week04_seq2seq/data.txt -O data.txt 2> log
# !wget https://github.com/yandexdataschool/nlp_course/raw/master/week04_seq2seq/utils.py -O utils.py 2> log
# !wget https://github.com/yandexdataschool/nlp_course/raw/master/week04_seq2seq/dummy_checkpoint.npz -O dummy_checkpoint.npz 2> log
#thanks to tilda and deephack teams for the data
# +
from nltk.tokenize import WordPunctTokenizer
from subword_nmt.learn_bpe import learn_bpe
from subword_nmt.apply_bpe import BPE
tokenizer = WordPunctTokenizer()
def tokenize(x):
return ' '.join(tokenizer.tokenize(x.lower()))
# split and tokenize the data
with open('train.en', 'w') as f_src, open('train.ru', 'w') as f_dst:
for line in open('data.txt'):
src_line, dst_line = line.strip().split('\t')
f_src.write(tokenize(src_line) + '\n')
f_dst.write(tokenize(dst_line) + '\n')
# build and apply bpe vocs
bpe = {}
for lang in ['en', 'ru']:
learn_bpe(open('./train.' + lang), open('bpe_rules.' + lang, 'w'), num_symbols=8000)
bpe[lang] = BPE(open('./bpe_rules.' + lang))
with open('train.bpe.' + lang, 'w') as f_out:
for line in open('train.' + lang):
f_out.write(bpe[lang].process_line(line.strip()) + '\n')
# -
# ### Building vocabularies
#
# We now need to build vocabularies that map strings to token ids and vice versa. We're gonna need these fellas when we feed training data into model or convert output matrices into words.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
data_inp = np.array(open('./train.bpe.ru').read().split('\n'))
data_out = np.array(open('./train.bpe.en').read().split('\n'))
from sklearn.model_selection import train_test_split
train_inp, dev_inp, train_out, dev_out = train_test_split(data_inp, data_out, test_size=3000,
random_state=42)
for i in range(3):
print('inp:', train_inp[i])
print('out:', train_out[i], end='\n\n')
# -
from utils import Vocab
inp_voc = Vocab.from_lines(train_inp)
out_voc = Vocab.from_lines(train_out)
# +
# Here's how you cast lines into ids and backwards.
batch_lines = sorted(train_inp, key=len)[5:10]
batch_ids = inp_voc.to_matrix(batch_lines)
batch_lines_restored = inp_voc.to_lines(batch_ids)
print("lines")
print(batch_lines)
print("\nwords to ids (0 = bos, 1 = eos):")
print(batch_ids)
print("\nback to words")
print(batch_lines_restored)
# -
# Draw source and translation length distributions to estimate the scope of the task.
# +
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.title("source length")
plt.hist(list(map(len, map(str.split, train_inp))), bins=20);
plt.subplot(1, 2, 2)
plt.title("translation length")
plt.hist(list(map(len, map(str.split, train_out))), bins=20);
# -
# ### Encoder-decoder model
#
# The code below contas a template for a simple encoder-decoder model: single GRU encoder/decoder, no attention or anything. This model is implemented for you as a reference and a baseline for your homework assignment.
import tensorflow as tf
import keras.layers as L
from utils import infer_length, infer_mask
class BasicModel:
def __init__(self, name, inp_voc, out_voc, emb_size=64, hid_size=128):
"""
A simple encoder-decoder model
"""
self.name, self.inp_voc, self.out_voc = name, inp_voc, out_voc
with tf.variable_scope(name):
self.emb_inp = L.Embedding(len(inp_voc), emb_size)
self.emb_out = L.Embedding(len(out_voc), emb_size)
self.enc0 = tf.nn.rnn_cell.GRUCell(hid_size)
self.dec_start = L.Dense(hid_size)
self.dec0 = tf.nn.rnn_cell.GRUCell(hid_size)
self.logits = L.Dense(len(out_voc))
# prepare to translate_lines
self.inp = tf.placeholder('int32', [None, None])
self.initial_state = self.prev_state = self.encode(self.inp)
self.prev_tokens = tf.placeholder('int32', [None])
self.next_state, self.next_logits = self.decode(self.prev_state, self.prev_tokens)
self.weights = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope=name)
def encode(self, inp, **flags):
"""
Takes symbolic input sequence, computes initial state
:param inp: matrix of input tokens [batch, time]
:returns: initial decoder state tensors, one or many
"""
inp_lengths = infer_length(inp, self.inp_voc.eos_ix)
inp_emb = self.emb_inp(inp)
with tf.variable_scope('enc0'):
_, enc_last = tf.nn.dynamic_rnn(
self.enc0, inp_emb,
sequence_length=inp_lengths,
dtype = inp_emb.dtype)
dec_start = self.dec_start(enc_last)
return [dec_start]
def decode(self, prev_state, prev_tokens, **flags):
"""
Takes previous decoder state and tokens, returns new state and logits for next tokens
:param prev_state: a list of previous decoder state tensors
:param prev_tokens: previous output tokens, an int vector of [batch_size]
:return: a list of next decoder state tensors, a tensor of logits [batch, n_tokens]
"""
[prev_dec] = prev_state
prev_emb = self.emb_out(prev_tokens[:,None])[:,0]
with tf.variable_scope('dec0'):
new_dec_out, new_dec_state = self.dec0(prev_emb, prev_dec)
output_logits = self.logits(new_dec_out)
return [new_dec_state], output_logits
def translate_lines(self, inp_lines, max_len=100):
"""
Translates a list of lines by greedily selecting most likely next token at each step
:returns: a list of output lines, a sequence of model states at each step
"""
state = sess.run(self.initial_state, {self.inp: inp_voc.to_matrix(inp_lines)})
outputs = [[self.out_voc.bos_ix] for _ in range(len(inp_lines))]
all_states = [state]
finished = [False] * len(inp_lines)
for t in range(max_len):
state, logits = sess.run([self.next_state, self.next_logits], {**dict(zip(self.prev_state, state)),
self.prev_tokens: [out_i[-1] for out_i in outputs]})
next_tokens = np.argmax(logits, axis=-1)
all_states.append(state)
for i in range(len(next_tokens)):
outputs[i].append(next_tokens[i])
finished[i] |= next_tokens[i] == self.out_voc.eos_ix
return out_voc.to_lines(outputs), all_states
# +
tf.reset_default_graph()
sess = tf.InteractiveSession()
# ^^^ if you get "variable *** already exists": re-run this cell again - it will clear all tf operations youve 'built
model = BasicModel('model', inp_voc, out_voc)
sess.run(tf.global_variables_initializer())
# -
# ### Training loss (2 points)
#
# Our training objetive is almost the same as it was for neural language models:
# $$ L = {\frac1{|D|}} \sum_{X, Y \in D} \sum_{y_t \in Y} - \log p(y_t \mid y_1, \dots, y_{t-1}, X, \theta) $$
#
# where $|D|$ is the __total length of all sequences__, including BOS and first EOS, but excluding PAD.
# +
def compute_logits(model, inp, out, **flags):
"""
:param inp: input tokens matrix, int32[batch, time]
:param out: reference tokens matrix, int32[batch, time]
:returns: logits of shape [batch, time, voc_size]
* logits must be a linear output of your neural network.
* logits [:, 0, :] should always predic BOS
* logits [:, -1, :] should be probabilities of last token in out
This function should NOT return logits predicted when taking out[:, -1] as y_prev
"""
batch_size = tf.shape(inp)[0]
# Encode inp, get initial state
first_state = model.encode(inp)
# initial logits: always predict BOS
first_logits = tf.log(tf.one_hot(tf.fill([batch_size], model.out_voc.bos_ix),
len(model.out_voc)) + 1e-30)
# Decode step
def step(prev_state, y_prev):
# Given previous state, obtain next state and next token logits
return model.decode(prev_state[0], y_prev)
# You can now use tf.scan to run step several times.
# use tf.transpose(out) as elems (to process one time-step at a time)
# docs: https://www.tensorflow.org/api_docs/python/tf/scan
# <YOUR CODE>
# a =
logits_seq = tf.scan(step, tf.transpose(out), (first_state, first_logits))[1]
# prepend first_logits to logits_seq
logits_seq = tf.concat([first_logits[None, ...], logits_seq], axis=0)[:-1, ...]
# Make sure you convert logits_seq from [time, batch, voc_size] to [batch, time, voc_size]
# logits_seq = <...>
logits_seq = tf.transpose(logits_seq, [1, 0, 2])
return logits_seq
# -
from utils import load
load(tf.trainable_variables(), 'dummy_checkpoint.npz')
dummy_inp = tf.constant(inp_voc.to_matrix(train_inp[:3]))
dummy_out = tf.constant(out_voc.to_matrix(train_out[:3]))
dummy_logits = sess.run(compute_logits(model, dummy_inp, dummy_out))
dummy_ref = np.array([-0.13257082, -0.11084784, -0.09024167, -0.14910498], dtype='float32')
assert np.allclose(dummy_logits.sum(-1)[0, 1:5], dummy_ref)
ref_shape = (dummy_out.shape[0], dummy_out.shape[1], len(out_voc))
assert dummy_logits.shape == ref_shape, "Your logits shape should be {} but got {}".format(dummy_logits.shape, ref_shape)
assert all(dummy_logits[:, 0].argmax(-1) == out_voc.bos_ix), "first step must always be BOS"
# +
from utils import select_values_over_last_axis
def compute_loss(model, inp, out, **flags):
"""
Compute loss (float32 scalar) as in the formula above
:param inp: input tokens matrix, int32[batch, time]
:param out: reference tokens matrix, int32[batch, time]
In order to pass the tests, your function should
* include loss at first EOS but not the subsequent ones
* divide sum of losses by a sum of input lengths (use infer_length or infer_mask)
"""
mask = infer_mask(out, out_voc.eos_ix)
logits_seq = compute_logits(model, inp, out, **flags)
# Compute loss as per instructions above
losses = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits_seq, labels=out) * mask
return tf.reduce_sum(losses) / tf.reduce_sum(mask)
# -
dummy_loss = sess.run(compute_loss(model, dummy_inp, dummy_out))
print("Loss:", dummy_loss)
assert np.allclose(dummy_loss, 8.425, rtol=0.1, atol=0.1), "We're sorry for your loss"
# ### Evaluation: BLEU
#
# Machine translation is commonly evaluated with [BLEU](https://en.wikipedia.org/wiki/BLEU) score. This metric simply computes which fraction of predicted n-grams is actually present in the reference translation. It does so for n=1,2,3 and 4 and computes the geometric average with penalty if translation is shorter than reference.
#
# While BLEU [has many drawbacks](http://www.cs.jhu.edu/~ccb/publications/re-evaluating-the-role-of-bleu-in-mt-research.pdf), it still remains the most commonly used metric and one of the simplest to compute.
# __Note:__ in this assignment we measure token-level bleu with bpe tokens. Most scientific papers report word-level bleu. You can measure it by undoing BPE encoding before computing BLEU. Please stay with the token-level bleu for this assignment, however.
#
from nltk.translate.bleu_score import corpus_bleu
def compute_bleu(model, inp_lines, out_lines, **flags):
""" Estimates corpora-level BLEU score of model's translations given inp and reference out """
translations, _ = model.translate_lines(inp_lines, **flags)
# Note: if you experience out-of-memory error, split input lines into batches and translate separately
return corpus_bleu([[ref] for ref in out_lines], translations) * 100
compute_bleu(model, dev_inp, dev_out)
# ### Training loop
#
# Training encoder-decoder models isn't that different from any other models: sample batches, compute loss, backprop and update
# +
inp = tf.placeholder('int32', [None, None])
out = tf.placeholder('int32', [None, None])
loss = compute_loss(model, inp, out)
train_step = tf.train.AdamOptimizer().minimize(loss)
# +
from IPython.display import clear_output
from tqdm import tqdm, trange
metrics = {'train_loss': [], 'dev_bleu': [] }
sess.run(tf.global_variables_initializer())
batch_size = 32
# +
for _ in trange(25000):
step = len(metrics['train_loss']) + 1
batch_ix = np.random.randint(len(train_inp), size=batch_size)
feed_dict = {
inp: inp_voc.to_matrix(train_inp[batch_ix]),
out: out_voc.to_matrix(train_out[batch_ix]),
}
loss_t, _ = sess.run([loss, train_step], feed_dict)
metrics['train_loss'].append((step, loss_t))
if step % 100 == 0:
metrics['dev_bleu'].append((step, compute_bleu(model, dev_inp, dev_out)))
clear_output(True)
plt.figure(figsize=(12,4))
for i, (name, history) in enumerate(sorted(metrics.items())):
plt.subplot(1, len(metrics), i + 1)
plt.title(name)
plt.plot(*zip(*history))
plt.grid()
plt.show()
print("Mean loss=%.3f" % np.mean(metrics['train_loss'][-10:], axis=0)[1], flush=True)
# Note: it's okay if bleu oscillates up and down as long as it gets better on average over long term (e.g. 5k batches)
# -
assert np.mean(metrics['dev_bleu'][-10:], axis=0)[1] > 35, "We kind of need a higher bleu BLEU from you. Kind of right now."
for inp_line, trans_line in zip(dev_inp[::500], model.translate_lines(dev_inp[::500])[0]):
print(inp_line)
print(trans_line)
print()
# ### Your Attention Required (4 points)
#
# In this section we want you to improve over the basic model by implementing a simple attention mechanism.
#
# This is gonna be a two-parter: building the __attention layer__ and using it for an __attentive seq2seq model__.
# ### Attention layer
#
# Here you will have to implement a layer that computes a simple additive attention:
#
# Given encoder sequence $ h^e_0, h^e_1, h^e_2, ..., h^e_T$ and a single decoder state $h^d$,
#
# * Compute logits with a 2-layer neural network
# $$a_t = linear_{out}(tanh(linear_{e}(h^e_t) + linear_{d}(h_d)))$$
# * Get probabilities from logits,
# $$ p_t = {{e ^ {a_t}} \over { \sum_\tau e^{a_\tau} }} $$
#
# * Add up encoder states with probabilities to get __attention response__
# $$ attn = \sum_t p_t \cdot h^e_t $$
#
# You can learn more about attention layers in the leture slides or [from this post](https://distill.pub/2016/augmented-rnns/).
class AttentionLayer:
def __init__(self, name, enc_size, dec_size, hid_size, activ=tf.tanh,):
""" A layer that computes additive attention response and weights """
self.name = name
self.enc_size = enc_size # num units in encoder state
self.dec_size = dec_size # num units in decoder state
self.hid_size = hid_size # attention layer hidden units
self.activ = activ # attention layer hidden nonlinearity
with tf.variable_scope(name):
# YOUR CODE - create layer variables
self.l_encoder = L.Dense(hid_size)
self.l_decoder = L.Dense(hid_size)
self.l_output = L.Dense(1)
def __call__(self, enc, dec, inp_mask):
"""
Computes attention response and weights
:param enc: encoder activation sequence, float32[batch_size, ninp, enc_size]
:param dec: single decoder state used as "query", float32[batch_size, dec_size]
:param inp_mask: mask on enc activatons (0 after first eos), float32 [batch_size, ninp]
:returns: attn[batch_size, enc_size], probs[batch_size, ninp]
- attn - attention response vector (weighted sum of enc)
- probs - attention weights after softmax
"""
with tf.variable_scope(self.name):
# Compute logits
key = self.l_encoder(enc)
query = self.l_decoder(dec)[:, None, :]
logits = self.l_output(self.activ(key + query))[..., 0]
# Apply mask - if mask is 0, logits should be -inf or -1e9
# You may need tf.where
logits = inp_mask * logits + (1 - inp_mask) * (-1e9)
# Compute attention probabilities (softmax)
probs = tf.nn.softmax(logits)
# Compute attention response using enc and probs
attn = tf.reduce_sum(probs[..., None] * enc, axis=1)
return attn, probs
# ### Seq2seq model with attention
#
# You can now use the attention layer to build a network. The simplest way to implement attention is to use it in decoder phase:
# 
# _image from distill.pub [article](https://distill.pub/2016/augmented-rnns/)_
#
# On every step, use __previous__ decoder state to obtain attention response. Then feed concat this response to the inputs of next attetion layer.
#
# The key implementation detail here is __model state__. Put simply, you can add any tensor into the list of `encode` outputs. You will then have access to them at each `decode` step. This may include:
# * Last RNN hidden states (as in basic model)
# * The whole sequence of encoder outputs (to attend to) and mask
# * Attention probabilities (to visualize)
#
# _There are, of course, alternative ways to wire attention into your network and different kinds of attention. Take a look at [this](https://arxiv.org/abs/1609.08144), [this](https://arxiv.org/abs/1706.03762) and [this](https://arxiv.org/abs/1808.03867) for ideas. And for image captioning/im2latex there's [visual attention](https://arxiv.org/abs/1502.03044)_
class AttentiveModel(BasicModel):
def __init__(self, name, inp_voc, out_voc,
emb_size=64, hid_size=128, attn_size=128):
""" Translation model that uses attention. See instructions above. """
self.name = name
self.inp_voc = inp_voc
self.out_voc = out_voc
with tf.variable_scope(name):
# YOUR CODE - define model layers
self.l_emb_in = L.Embedding(len(inp_voc), emb_size)
self.l_emb_out = L.Embedding(len(out_voc), emb_size)
self.l_encoder_rnn = L.GRU(hid_size, return_sequences=True, return_state=True)
self.l_decoder_rnn = L.GRU(hid_size, return_sequences=True, return_state=True)
self.l_attention = AttentionLayer("attention", hid_size, hid_size, attn_size)
self.l_initial_decoder_state = L.Dense(hid_size)
self.l_output = L.Dense(len(out_voc))
# END OF YOUR CODE
# prepare to translate_lines
self.inp = tf.placeholder('int32', [None, None])
self.initial_state = self.prev_state = self.encode(self.inp)
self.prev_tokens = tf.placeholder('int32', [None])
self.next_state, self.next_logits = self.decode(self.prev_state, self.prev_tokens)
self.weights = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope=name)
def encode(self, inp, **flags):
"""
Takes symbolic input sequence, computes initial state
:param inp: matrix of input tokens [batch, time]
:return: a list of initial decoder state tensors
"""
# encode input sequence, create initial decoder states
encoder_states, last_state = self.l_encoder_rnn(self.l_emb_in(inp))
decoder_init_state = self.l_initial_decoder_state(last_state)
# apply attention layer from initial decoder hidden state
mask = tf.sequence_mask(infer_length(inp, self.inp_voc.eos_ix), dtype=tf.float32)
first_attn_response, first_attn_probas = self.l_attention(encoder_states, decoder_init_state, mask)
# Build first state: include
# * initial states for decoder recurrent layers
# * encoder sequence and encoder attn mask (for attention)
# * make sure that last state item is attention probabilities tensor
first_state = [decoder_init_state, encoder_states, mask, first_attn_probas]
return first_state
def decode(self, prev_state, prev_tokens, **flags):
"""
Takes previous decoder state and tokens, returns new state and logits
:param prev_state: a list of previous decoder state tensors
:param prev_tokens: previous output tokens, an int vector of [batch_size]
:return: a list of next decoder state tensors, a tensor of logits [batch,n_tokens]
"""
# Unpack your state: you will get tensors in the same order that you've packed in encode
[decoder_prev_state, encoder_states, mask, prev_attn_probas] = prev_state
# Perform decoder step
# * predict next attn response and attn probas given previous decoder state
# * use prev token embedding and attn response to update decoder states (concatenate and feed into decoder cell)
# * predict logits
self.l_emb_out(prev_tokens[:, None])
next_attn_response, next_attn_probas = self.l_attention(encoder_states, decoder_prev_state, mask)
prev_emb = self.l_emb_out(prev_tokens)
concat_emb = tf.concat([next_attn_response, prev_emb], axis=-1)
new_dec_out, new_dec_state = self.l_decoder_rnn(concat_emb[:, None, :], decoder_prev_state)
output_logits = self.l_output(new_dec_out)[:, 0, :]
# Pack new state:
# * replace previous decoder state with next one
# * copy encoder sequence and mask from prev_state
# * append new attention probas
next_state = [new_dec_state, encoder_states, mask, next_attn_probas]
return next_state, output_logits
# WARNING! this cell will clear your TF graph from the regular model. All trained variables will be gone!
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = AttentiveModel('model_attn', inp_voc, out_voc)
# ### Training attentive model
#
# We'll reuse the infrastructure you've built for the regular model. I hope you didn't hard-code anything :)
# +
inp = tf.placeholder('int32', [None, None])
out = tf.placeholder('int32', [None, None])
loss = compute_loss(model, inp, out)
train_step = tf.train.AdamOptimizer().minimize(loss)
# -
metrics = {'train_loss': [], 'dev_bleu': []}
sess.run(tf.global_variables_initializer())
batch_size = 32
# +
for _ in trange(25000):
step = len(metrics['train_loss']) + 1
batch_ix = np.random.randint(len(train_inp), size=batch_size)
feed_dict = {
inp: inp_voc.to_matrix(train_inp[batch_ix]),
out: out_voc.to_matrix(train_out[batch_ix]),
}
loss_t, _ = sess.run([loss, train_step], feed_dict)
metrics['train_loss'].append((step, loss_t))
if step % 100 == 0:
metrics['dev_bleu'].append((step, compute_bleu(model, dev_inp, dev_out)))
clear_output(True)
plt.figure(figsize=(12,4))
for i, (name, history) in enumerate(sorted(metrics.items())):
plt.subplot(1, len(metrics), i + 1)
plt.title(name)
plt.plot(*zip(*history))
plt.grid()
plt.show()
print("Mean loss=%.3f" % np.mean(metrics['train_loss'][-10:], axis=0)[1], flush=True)
# Your model may train slower than the basic one. check that it's at least >30 bleu by 5k steps
# Also: you don't have to train for 25k steps. It was chosen by a squirrel.
# -
assert np.mean(metrics['dev_bleu'][-10:], axis=0)[1] > 45, "Something might be wrong with the model..."
# +
import bokeh.plotting as pl
import bokeh.models as bm
from bokeh.io import output_notebook, show
output_notebook()
def draw_attention(inp_line, translation, probs):
""" An intentionally ambiguous function to visualize attention weights """
inp_tokens = inp_voc.tokenize(inp_line)
trans_tokens = out_voc.tokenize(translation)
probs = probs[:len(trans_tokens), :len(inp_tokens)]
fig = pl.figure(x_range=(0, len(inp_tokens)), y_range=(0, len(trans_tokens)),
x_axis_type=None, y_axis_type=None, tools=[])
fig.image([probs[::-1]], 0, 0, len(inp_tokens), len(trans_tokens))
fig.add_layout(bm.LinearAxis(axis_label='source tokens'), 'above')
fig.xaxis.ticker = np.arange(len(inp_tokens)) + 0.5
fig.xaxis.major_label_overrides = dict(zip(np.arange(len(inp_tokens)) + 0.5, inp_tokens))
fig.xaxis.major_label_orientation = 45
fig.add_layout(bm.LinearAxis(axis_label='translation tokens'), 'left')
fig.yaxis.ticker = np.arange(len(trans_tokens)) + 0.5
fig.yaxis.major_label_overrides = dict(zip(np.arange(len(trans_tokens)) + 0.5, trans_tokens[::-1]))
show(fig)
# +
inp = dev_inp[::500]
trans, states = model.translate_lines(inp)
# select attention probs from model state (you may need to change this for your custom model)
attention_probs = np.stack([state[-1] for state in states], axis=1)
# -
for i in range(5):
draw_attention(inp[i], trans[i], attention_probs[i])
# ## Grand Finale (4+ points)
#
# We want you to find the best model for the task. Use everything you know.
#
# * different recurrent units: rnn/gru/lstm; deeper architectures
# * bidirectional encoder, different attention methods for decoder
# * word dropout, training schedules, anything you can imagine
#
# As usual, we want you to describe what you tried and what results you obtained.
# `[your report/log here or anywhere you please]`
| week04_seq2seq/practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Detect Model Bias with Amazon SageMaker Clarify
#
# ## Amazon Science: _[How Clarify helps machine learning developers detect unintended bias](https://www.amazon.science/latest-news/how-clarify-helps-machine-learning-developers-detect-unintended-bias)_
#
# [<img src="img/amazon_science_clarify.png" width="100%" align="left">](https://www.amazon.science/latest-news/how-clarify-helps-machine-learning-developers-detect-unintended-bias)
# # Terminology
#
# * **Bias**:
# An imbalance in the training data or the prediction behavior of the model across different groups, such as age or income bracket. Biases can result from the data or algorithm used to train your model. For instance, if an ML model is trained primarily on data from middle-aged individuals, it may be less accurate when making predictions involving younger and older people.
#
# * **Bias metric**:
# A function that returns numerical values indicating the level of a potential bias.
#
# * **Bias report**:
# A collection of bias metrics for a given dataset, or a combination of a dataset and a model.
#
# * **Label**:
# Feature that is the target for training a machine learning model. Referred to as the observed label or observed outcome.
#
# * **Positive label values**:
# Label values that are favorable to a demographic group observed in a sample. In other words, designates a sample as having a positive result.
#
# * **Negative label values**:
# Label values that are unfavorable to a demographic group observed in a sample. In other words, designates a sample as having a negative result.
#
# * **Facet**:
# A column or feature that contains the attributes with respect to which bias is measured.
#
# * **Facet value**:
# The feature values of attributes that bias might favor or disfavor.
# # Posttraining Bias Metrics
# https://docs.aws.amazon.com/sagemaker/latest/dg/clarify-measure-post-training-bias.html
#
# * **Difference in Positive Proportions in Predicted Labels (DPPL)**:
# Measures the difference in the proportion of positive predictions between the favored facet a and the disfavored facet d.
#
# * **Disparate Impact (DI)**:
# Measures the ratio of proportions of the predicted labels for the favored facet a and the disfavored facet d.
#
# * **Difference in Conditional Acceptance (DCAcc)**:
# Compares the observed labels to the labels predicted by a model and assesses whether this is the same across facets for predicted positive outcomes (acceptances).
#
# * **Difference in Conditional Rejection (DCR)**:
# Compares the observed labels to the labels predicted by a model and assesses whether this is the same across facets for negative outcomes (rejections).
#
# * **Recall Difference (RD)**:
# Compares the recall of the model for the favored and disfavored facets.
#
# * **Difference in Acceptance Rates (DAR)**:
# Measures the difference in the ratios of the observed positive outcomes (TP) to the predicted positives (TP + FP) between the favored and disfavored facets.
#
# * **Difference in Rejection Rates (DRR)**:
# Measures the difference in the ratios of the observed negative outcomes (TN) to the predicted negatives (TN + FN) between the disfavored and favored facets.
#
# * **Accuracy Difference (AD)**:
# Measures the difference between the prediction accuracy for the favored and disfavored facets.
#
# * **Treatment Equality (TE)**:
# Measures the difference in the ratio of false positives to false negatives between the favored and disfavored facets.
#
# * **Conditional Demographic Disparity in Predicted Labels (CDDPL)**:
# Measures the disparity of predicted labels between the facets as a whole, but also by subgroups.
#
# * **Counterfactual Fliptest (FT)**:
# Examines each member of facet d and assesses whether similar members of facet a have different model predictions.
#
# +
import boto3
import sagemaker
import pandas as pd
import numpy as np
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sm = boto3.Session().client(service_name="sagemaker", region_name=region)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
# -
# # Test data for bias
#
# We created test data in JSONLines format to match the model inputs.
test_data_bias_path = "./data-clarify/test_data_bias.jsonl"
# !head -n 1 $test_data_bias_path
# ### Upload the data
test_data_bias_s3_uri = sess.upload_data(bucket=bucket, key_prefix="bias/test_data_bias", path=test_data_bias_path)
test_data_bias_s3_uri
# !aws s3 ls $test_data_bias_s3_uri
# %store test_data_bias_s3_uri
# # Run Posttraining Model Bias Analysis
# %store -r pipeline_name
print(pipeline_name)
# +
# %%time
import time
from pprint import pprint
executions_response = sm.list_pipeline_executions(PipelineName=pipeline_name)["PipelineExecutionSummaries"]
pipeline_execution_status = executions_response[0]["PipelineExecutionStatus"]
print(pipeline_execution_status)
while pipeline_execution_status == "Executing":
try:
executions_response = sm.list_pipeline_executions(PipelineName=pipeline_name)["PipelineExecutionSummaries"]
pipeline_execution_status = executions_response[0]["PipelineExecutionStatus"]
except Exception as e:
print("Please wait...")
time.sleep(30)
pprint(executions_response)
# -
# # List Pipeline Execution Steps
#
pipeline_execution_status = executions_response[0]["PipelineExecutionStatus"]
print(pipeline_execution_status)
pipeline_execution_arn = executions_response[0]["PipelineExecutionArn"]
print(pipeline_execution_arn)
# +
from pprint import pprint
steps = sm.list_pipeline_execution_steps(PipelineExecutionArn=pipeline_execution_arn)
pprint(steps)
# -
# # View Created Model
# _Note: If the trained model did not pass the Evaluation step (> accuracy threshold), it will not be created._
# +
for execution_step in steps["PipelineExecutionSteps"]:
if execution_step["StepName"] == "CreateModel":
model_arn = execution_step["Metadata"]["Model"]["Arn"]
break
print(model_arn)
pipeline_model_name = model_arn.split("/")[-1]
print(pipeline_model_name)
# -
# # SageMakerClarifyProcessor
# +
from sagemaker import clarify
clarify_processor = clarify.SageMakerClarifyProcessor(
role=role, instance_count=1, instance_type="ml.c5.2xlarge", sagemaker_session=sess
)
# -
# # Writing DataConfig and ModelConfig
# A `DataConfig` object communicates some basic information about data I/O to Clarify. We specify where to find the input dataset, where to store the output, the target column (`label`), the header names, and the dataset type.
#
# Similarly, the `ModelConfig` object communicates information about your trained model and `ModelPredictedLabelConfig` provides information on the format of your predictions.
#
# **Note**: To avoid additional traffic to your production models, SageMaker Clarify sets up and tears down a dedicated endpoint when processing. `ModelConfig` specifies your preferred instance type and instance count used to run your model on during Clarify's processing.
# ## DataConfig
# +
bias_report_prefix = "bias/report-{}".format(pipeline_model_name)
bias_report_output_path = "s3://{}/{}".format(bucket, bias_report_prefix)
data_config = clarify.DataConfig(
s3_data_input_path=test_data_bias_s3_uri,
s3_output_path=bias_report_output_path,
label="star_rating",
features="features",
# label must be last, features in exact order as passed into model
headers=["review_body", "product_category", "star_rating"],
dataset_type="application/jsonlines",
)
# -
# ## ModelConfig
model_config = clarify.ModelConfig(
model_name=pipeline_model_name,
instance_type="ml.m5.4xlarge",
instance_count=1,
content_type="application/jsonlines",
accept_type="application/jsonlines",
# {"features": ["the worst", "Digital_Software"]}
content_template='{"features":$features}',
)
# ## _Note: `label` is set to the JSON key for the model prediction results_
predictions_config = clarify.ModelPredictedLabelConfig(label="predicted_label")
# ## BiasConfig
bias_config = clarify.BiasConfig(
label_values_or_threshold=[
5,
4,
], # needs to be int or str for continuous dtype, needs to be >1 for categorical dtype
facet_name="product_category",
# facet_values_or_threshold=['Gift Card'],
group_name="product_category",
)
# # Run Clarify Job
clarify_processor.run_post_training_bias(
data_config=data_config,
data_bias_config=bias_config,
model_config=model_config,
model_predicted_label_config=predictions_config,
# methods='all', # FlipTest requires all columns to be numeric
methods=["DPPL", "DI", "DCA", "DCR", "RD", "DAR", "DRR", "AD", "CDDPL", "TE"],
wait=False,
logs=False,
)
run_post_training_bias_processing_job_name = clarify_processor.latest_job.job_name
run_post_training_bias_processing_job_name
# +
from IPython.core.display import display, HTML
display(
HTML(
'<b>Review <a target="blank" href="https://console.aws.amazon.com/sagemaker/home?region={}#/processing-jobs/{}">Processing Job</a></b>'.format(
region, run_post_training_bias_processing_job_name
)
)
)
# +
from IPython.core.display import display, HTML
display(
HTML(
'<b>Review <a target="blank" href="https://console.aws.amazon.com/cloudwatch/home?region={}#logStream:group=/aws/sagemaker/ProcessingJobs;prefix={};streamFilter=typeLogStreamPrefix">CloudWatch Logs</a> After About 5 Minutes</b>'.format(
region, run_post_training_bias_processing_job_name
)
)
)
# +
from IPython.core.display import display, HTML
display(
HTML(
'<b>Review <a target="blank" href="https://s3.console.aws.amazon.com/s3/buckets/{}?prefix={}/">S3 Output Data</a> After The Processing Job Has Completed</b>'.format(
bucket, bias_report_prefix
)
)
)
# +
from pprint import pprint
running_processor = sagemaker.processing.ProcessingJob.from_processing_name(
processing_job_name=run_post_training_bias_processing_job_name, sagemaker_session=sess
)
processing_job_description = running_processor.describe()
pprint(processing_job_description)
# -
running_processor.wait(logs=False)
# # Download Report From S3
# !aws s3 ls $bias_report_output_path/
# !aws s3 cp --recursive $bias_report_output_path ./generated_bias_report/
# +
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="./generated_bias_report/report.html">Bias Report</a></b>'))
# -
# # View Bias Report in Studio
# In Studio, you can view the results under the experiments tab.
#
# <img src="img/bias_report.gif">
#
# Each bias metric has detailed explanations with examples that you can explore.
#
# <img src="img/bias_detail.gif">
#
# You could also summarize the results in a handy table!
#
# <img src="img/bias_report_chart.gif">
# # Release Resources
# + language="html"
#
# <p><b>Shutting down your kernel for this notebook to release resources.</b></p>
# <button class="sm-command-button" data-commandlinker-command="kernelmenu:shutdown" style="display:none;">Shutdown Kernel</button>
#
# <script>
# try {
# els = document.getElementsByClassName("sm-command-button");
# els[0].click();
# }
# catch(err) {
# // NoOp
# }
# </script>
| 00_quickstart/wip/09_Detect_Model_Bias_Clarify.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Remove input cells at runtime (nbsphinx)
import IPython.core.display as d
d.display_html('<script>jQuery(function() {if (jQuery("body.notebook_app").length == 0) { jQuery(".input_area").toggle(); jQuery(".prompt").toggle();}});</script>', raw=True)
# # Performance poster layout
# This notebook produces a nice poster layout about the comparison between pipelines.
# It is just a refurbished version of the DL3-level "IRF and sensitivity" notebook.
#
# Latest performance results cannot be shown on this public documentation and are therefore hosted at [this RedMine page](https://forge.in2p3.fr/projects/benchmarks-reference-analysis/wiki/Protopipe_performance_data) .
# + [markdown] nbsphinx="hidden"
# ## Imports
# +
## From the standard library
import os
from pathlib import Path
# From pyirf
import pyirf
from pyirf.binning import bin_center
from pyirf.utils import cone_solid_angle
# From other 3rd-party libraries
from yaml import load, FullLoader
import numpy as np
import astropy.units as u
from astropy.io import fits
from astropy.table import QTable, Table, Column
import uproot
import matplotlib.pyplot as plt
from matplotlib.ticker import ScalarFormatter
# %matplotlib inline
plt.rcParams['axes.labelsize'] = 15
plt.rcParams['xtick.labelsize'] = 15
plt.rcParams['ytick.labelsize'] = 15
# + [markdown] nbsphinx="hidden"
# ## Functions
# -
def load_config(name):
"""Load YAML configuration file."""
try:
with open(name, "r") as stream:
cfg = load(stream, Loader=FullLoader)
except FileNotFoundError as e:
print(e)
raise
return cfg
# + [markdown] nbsphinx="hidden"
# ## Input data
# + [markdown] nbsphinx="hidden"
# ### Protopipe
# -
# EDIT THIS CELL WITH YOUR LOCAL SETUP INFORMATION
parent_dir = "" # path to 'analyses' folder
analysisName = ""
infile = "performance_protopipe_Prod3b_CTANorth_baseline_full_array_Zd20deg_180deg_Time50.00h.fits.gz"
config_performance = load_config(f"{parent_dir}/{analysisName}/configs/performance.yaml")
obs_time = f'{config_performance["analysis"]["obs_time"]["value"]}{config_performance["analysis"]["obs_time"]["unit"]}'
production = infile.split("protopipe_")[1].split("_Time")[0]
protopipe_file = Path(parent_dir, analysisName, "data/DL3", infile)
# + [markdown] nbsphinx="hidden"
# ### ASWG
# +
# EDIT THIS CELL WITH YOUR LOCAL SETUP INFORMATION
parent_dir_aswg = ""
# MARS performance (available here: https://forge.in2p3.fr/projects/step-by-step-reference-mars-analysis/wiki)
indir_CTAMARS = ""
infile_CTAMARS = "SubarrayLaPalma_4L15M_south_IFAE_50hours_20190630.root"
MARS_label = "CTAMARS (2019)"
# ED performance (available here: https://forge.in2p3.fr/projects/cta_analysis-and-simulations/wiki/Prod3b_based_instrument_response_functions)
indir_ED = ""
infile_ED = "CTA-Performance-North-20deg-S-50h_20181203.root"
ED_label = "EventDisplay (2018)"
# -
MARS_performance = uproot.open(Path(parent_dir_aswg, indir_CTAMARS, infile_CTAMARS))
ED_performance = uproot.open(Path(parent_dir_aswg, indir_ED, infile_ED))
# + [markdown] nbsphinx="hidden"
# ### Requirements
# -
# EDIT THIS CELL WITH YOUR LOCAL SETUP INFORMATION
indir_requirements = ''
site = 'North'
obs_time = '50h'
# +
infile_requirements = {"sens" : f'/{site}-{obs_time}.dat',
"AngRes" : f'/{site}-{obs_time}-AngRes.dat',
"ERes" : f'/{site}-{obs_time}-ERes.dat'}
requirements = {}
for key in infile_requirements.keys():
requirements[key] = Table.read(indir_requirements + infile_requirements[key], format='ascii')
requirements['sens'].add_column(Column(data=(10**requirements['sens']['col1']), name='ENERGY'))
requirements['sens'].add_column(Column(data=requirements['sens']['col2'], name='SENSITIVITY'))
# + [markdown] nbsphinx="hidden"
# ## Poster plot
# -
# First we check if a _plots_ folder exists already.
# If not, we create it.
Path("./plots").mkdir(parents=True, exist_ok=True)
# +
fig = plt.figure(figsize = (20, 10), constrained_layout=True)
gs = fig.add_gridspec(3, 3, figure=fig)
# ==========================================================================================================
#
# SENSITIVITY
#
# ==========================================================================================================
ax1 = fig.add_subplot(gs[0:-1, 0:-1])
# [1:-1] removes under/overflow bins
sensitivity_protopipe = QTable.read(protopipe_file, hdu='SENSITIVITY')[1:-1]
unit = u.Unit('erg cm-2 s-1')
# Add requirements
ax1.plot(requirements['sens']['ENERGY'],
requirements['sens']['SENSITIVITY'],
color='black',
ls='--',
lw=2,
label='Requirements'
)
# protopipe
e = sensitivity_protopipe['reco_energy_center']
w = (sensitivity_protopipe['reco_energy_high'] - sensitivity_protopipe['reco_energy_low'])
s_p = (e**2 * sensitivity_protopipe['flux_sensitivity'])
ax1.errorbar(
e.to_value(u.TeV),
s_p.to_value(unit),
xerr=w.to_value(u.TeV) / 2,
ls='',
label='protopipe',
color='DarkOrange'
)
# ED
s_ED, edges = ED_performance["DiffSens"].to_numpy()
yerr = ED_performance["DiffSens"].errors()
bins = 10**edges
x = bin_center(bins)
width = np.diff(bins)
ax1.errorbar(
x,
s_ED,
xerr=width/2,
yerr=yerr,
label=ED_label,
ls='',
color='DarkGreen'
)
# MARS
s_MARS, edges = MARS_performance["DiffSens"].to_numpy()
yerr = MARS_performance["DiffSens"].errors()
bins = 10**edges
x = bin_center(bins)
width = np.diff(bins)
ax1.errorbar(
x,
s_MARS,
xerr=width/2,
yerr=yerr,
label=MARS_label,
ls='',
color='DarkBlue'
)
# Style settings
ax1.set_xscale("log")
ax1.set_yscale("log")
ax1.set_ylabel(rf"$(E^2 \cdot \mathrm{{Flux Sensitivity}}) /$ ({unit.to_string('latex')})")
ax1.grid(which="both")
ax1.legend()
# ==========================================================================================================
#
# SENSITIVITY RATIO
#
# ==========================================================================================================
ax2 = fig.add_subplot(gs[2, 0])
ax2.errorbar(
e.to_value(u.TeV),
s_p.to_value(unit) / s_ED,
xerr=w.to_value(u.TeV)/2,
ls='',
label = "",
color='DarkGreen'
)
ax2.errorbar(
e.to_value(u.TeV),
s_p.to_value(unit) / s_MARS,
xerr=w.to_value(u.TeV)/2,
ls='',
label = "",
color='DarkBlue'
)
ax2.axhline(1, color = 'DarkOrange')
ax2.set_xscale('log')
ax2.set_yscale('log')
ax2.set_xlabel("Reconstructed energy [TeV]")
ax2.set_ylabel('Sensitivity ratio')
ax2.grid()
ax2.yaxis.set_major_formatter(ScalarFormatter())
ax2.set_ylim(0.5, 2.0)
ax2.set_yticks([0.5, 2/3, 1, 3/2, 2])
ax2.set_yticks([], minor=True)
# ==========================================================================================================
#
# EFFECTIVE COLLECTION AREA
#
# ==========================================================================================================
ax3 = fig.add_subplot(gs[0, 2])
# protopipe
# uncomment the other strings to see effective areas
# for the different cut levels. Left out here for better
# visibility of the final effective areas.
suffix =''
#'_NO_CUTS'
#'_ONLY_GH'
#'_ONLY_THETA'
area = QTable.read(protopipe_file, hdu='EFFECTIVE_AREA' + suffix)[0]
ax3.errorbar(
0.5 * (area['ENERG_LO'] + area['ENERG_HI']).to_value(u.TeV)[1:-1],
area['EFFAREA'].to_value(u.m**2).T[1:-1, 0],
xerr=0.5 * (area['ENERG_LO'] - area['ENERG_HI']).to_value(u.TeV)[1:-1],
ls='',
label='protopipe ' + suffix,
color='DarkOrange'
)
# ED
y, edges = ED_performance["EffectiveAreaEtrue"].to_numpy()
yerr = ED_performance["EffectiveAreaEtrue"].errors()
x = bin_center(10**edges)
xerr = 0.5 * np.diff(10**edges)
ax3.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=ED_label,
color='DarkGreen'
)
# MARS
y, edges = MARS_performance["EffectiveAreaEtrue"].to_numpy()
yerr = MARS_performance["EffectiveAreaEtrue"].errors()
x = bin_center(10**edges)
xerr = 0.5 * np.diff(10**edges)
ax3.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=MARS_label,
color='DarkBlue'
)
# Style settings
ax3.set_xscale("log")
ax3.set_yscale("log")
ax3.set_xlabel("True energy [TeV]")
ax3.set_ylabel("Effective area [m²]")
ax3.grid(which="both")
# ==========================================================================================================
#
# ANGULAR RESOLUTION
#
# ==========================================================================================================
ax4 = fig.add_subplot(gs[2, 1])
# protopipe
ang_res = QTable.read(protopipe_file, hdu='ANGULAR_RESOLUTION')[1:-1]
ax4.errorbar(
0.5 * (ang_res['reco_energy_low'] + ang_res['reco_energy_high']).to_value(u.TeV),
ang_res['angular_resolution'].to_value(u.deg),
xerr=0.5 * (ang_res['reco_energy_high'] - ang_res['reco_energy_low']).to_value(u.TeV),
ls='',
label='protopipe',
color='DarkOrange'
)
# ED
y, edges = ED_performance["AngRes"].to_numpy()
yerr = ED_performance["AngRes"].errors()
x = bin_center(10**edges)
xerr = 0.5 * np.diff(10**edges)
ax4.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=ED_label,
color='DarkGreen')
# MARS
y, edges = MARS_performance["AngRes"].to_numpy()
yerr = MARS_performance["AngRes"].errors()
x = bin_center(10**edges)
xerr = 0.5 * np.diff(10**edges)
ax4.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=MARS_label,
color='DarkBlue')
# Requirements
ax4.plot(10**requirements['AngRes']['col1'],
requirements['AngRes']['col2'],
color='black',
ls='--',
lw=2,
label='Requirements'
)
# Style settings
ax4.set_xscale("log")
ax4.set_yscale("log")
ax4.set_xlabel("Reconstructed energy [TeV]")
ax4.set_ylabel("Angular resolution [deg]")
ax4.grid(which="both")
None # to remove clutter by mpl objects
# ==========================================================================================================
#
# ENERGY RESOLUTION
#
# ==========================================================================================================
ax5 = fig.add_subplot(gs[2, 2])
# protopipe
bias_resolution = QTable.read(protopipe_file, hdu='ENERGY_BIAS_RESOLUTION')[1:-1]
ax5.errorbar(
0.5 * (bias_resolution['reco_energy_low'] + bias_resolution['reco_energy_high']).to_value(u.TeV),
bias_resolution['resolution'],
xerr=0.5 * (bias_resolution['reco_energy_high'] - bias_resolution['reco_energy_low']).to_value(u.TeV),
ls='',
label='protopipe',
color='DarkOrange'
)
# ED
y, edges = ED_performance["ERes"].to_numpy()
yerr = ED_performance["ERes"].errors()
x = bin_center(10**edges)
xerr = np.diff(10**edges) / 2
ax5.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=ED_label,
color='DarkGreen'
)
# MARS
y, edges = MARS_performance["ERes"].to_numpy()
yerr = MARS_performance["ERes"].errors()
x = bin_center(10**edges)
xerr = np.diff(10**edges) / 2
ax5.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=MARS_label,
color='DarkBlue'
)
# Requirements
ax5.plot(10**requirements['ERes']['col1'],
requirements['ERes']['col2'],
color='black',
ls='--',
lw=2,
label='Requirements'
)
# Style settings
ax5.set_xlabel("Reconstructed energy [TeV]")
ax5.set_ylabel("Energy resolution")
ax5.grid(which="both")
ax5.set_xscale('log')
None # to remove clutter by mpl objects
# ==========================================================================================================
#
# BACKGROUND RATE
#
# ==========================================================================================================
ax6 = fig.add_subplot(gs[1, 2])
from pyirf.utils import cone_solid_angle
# protopipe
rad_max = QTable.read(protopipe_file, hdu='RAD_MAX')[0]
bg_rate = QTable.read(protopipe_file, hdu='BACKGROUND')[0]
reco_bins = np.append(bg_rate['ENERG_LO'], bg_rate['ENERG_HI'][-1])
# first fov bin, [0, 1] deg
fov_bin = 0
rate_bin = bg_rate['BKG'].T[:, fov_bin]
# interpolate theta cut for given e reco bin
e_center_bg = 0.5 * (bg_rate['ENERG_LO'] + bg_rate['ENERG_HI'])
e_center_theta = 0.5 * (rad_max['ENERG_LO'] + rad_max['ENERG_HI'])
theta_cut = np.interp(e_center_bg, e_center_theta, rad_max['RAD_MAX'].T[:, 0])
# undo normalization
rate_bin *= cone_solid_angle(theta_cut)
rate_bin *= np.diff(reco_bins)
ax6.errorbar(
0.5 * (bg_rate['ENERG_LO'] + bg_rate['ENERG_HI']).to_value(u.TeV)[1:-1],
rate_bin.to_value(1 / u.s)[1:-1],
xerr=np.diff(reco_bins).to_value(u.TeV)[1:-1] / 2,
ls='',
label='protopipe',
color='DarkOrange'
)
# ED
y, edges = ED_performance["BGRate"].to_numpy()
yerr = ED_performance["BGRate"].errors()
x = bin_center(10**edges)
xerr = np.diff(10**edges) / 2
ax6.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=ED_label,
color="DarkGreen")
# MARS
y, edges = MARS_performance["BGRate"].to_numpy()
yerr = MARS_performance["BGRate"].errors()
x = bin_center(10**edges)
xerr = np.diff(10**edges) / 2
ax6.errorbar(x,
y,
xerr=xerr,
yerr=yerr,
ls='',
label=MARS_label,
color="DarkBlue")
# Style settings
ax6.set_xscale("log")
ax6.set_xlabel("Reconstructed energy [TeV]")
ax6.set_ylabel("Background rate [s⁻¹ TeV⁻¹] ")
ax6.grid(which="both")
ax6.set_yscale('log')
fig.suptitle(f'{production} - {obs_time}', fontsize=25)
fig.savefig(f"./plots/protopipe_{production}_{obs_time}.png")
None # to remove clutter by mpl objects
# -
| docs/contribute/benchmarks_latest_results/Prod3b/CTAN_Zd20_AzSouth_NSB1x_baseline_pointsource/DL3/overall_performance_plot_CTA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Transfer Learning and Fine Tuning
# * Train a simple convnet on the MNIST dataset the first 5 digits [0..4].
# * Freeze convolutional layers and fine-tune dense layers for the classification of digits [5..9].
# #### Using GPU (highly recommended)
#
# -> If using `theano` backend:
#
# `THEANO_FLAGS=mode=FAST_RUN,device=gpu,floatX=float32`
# +
import numpy as np
import datetime
np.random.seed(1337) # for reproducibility
# +
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.utils import np_utils
from keras import backend as K
from numpy import nan
import keras
print keras.__version__
now = datetime.datetime.now
# -
# ### Settings
# +
now = datetime.datetime.now
batch_size = 128
nb_classes = 5
nb_epoch = 5
# input image dimensions
img_rows, img_cols = 28, 28
# number of convolutional filters to use
nb_filters = 32
# size of pooling area for max pooling
pool_size = 2
# convolution kernel size
kernel_size = 3
# -
if K.image_data_format() == 'channels_first':
input_shape = (1, img_rows, img_cols)
else:
input_shape = (img_rows, img_cols, 1)
def train_model(model, train, test, nb_classes):
X_train = train[0].reshape((train[0].shape[0],) + input_shape)
X_test = test[0].reshape((test[0].shape[0],) + input_shape)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
print('X_train shape:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
Y_train = np_utils.to_categorical(train[1], nb_classes)
Y_test = np_utils.to_categorical(test[1], nb_classes)
model.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
t = now()
model.fit(X_train, Y_train,
batch_size=batch_size, nb_epoch=nb_epoch,
verbose=1,
validation_data=(X_test, Y_test))
print('Training time: %s' % (now() - t))
score = model.evaluate(X_test, Y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
# ## Dataset Preparation
# +
# the data, shuffled and split between train and test sets
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# create two datasets one with digits below 5 and one with 5 and above
X_train_lt5 = X_train[y_train < 5]
y_train_lt5 = y_train[y_train < 5]
X_test_lt5 = X_test[y_test < 5]
y_test_lt5 = y_test[y_test < 5]
X_train_gte5 = X_train[y_train >= 5]
y_train_gte5 = y_train[y_train >= 5] - 5 # make classes start at 0 for
X_test_gte5 = X_test[y_test >= 5] # np_utils.to_categorical
y_test_gte5 = y_test[y_test >= 5] - 5
# -
# define two groups of layers: feature (convolutions) and classification (dense)
feature_layers = [
Convolution2D(nb_filters, kernel_size, kernel_size,
border_mode='valid',
input_shape=input_shape),
Activation('relu'),
Convolution2D(nb_filters, kernel_size, kernel_size),
Activation('relu'),
MaxPooling2D(pool_size=(pool_size, pool_size)),
Dropout(0.25),
Flatten(),
]
classification_layers = [
Dense(128),
Activation('relu'),
Dropout(0.5),
Dense(nb_classes),
Activation('softmax')
]
# +
# create complete model
model = Sequential(feature_layers + classification_layers)
# train model for 5-digit classification [0..4]
train_model(model,
(X_train_lt5, y_train_lt5),
(X_test_lt5, y_test_lt5), nb_classes)
# +
# freeze feature layers and rebuild model
for l in feature_layers:
l.trainable = False
# transfer: train dense layers for new classification task [5..9]
train_model(model,
(X_train_gte5, y_train_gte5),
(X_test_gte5, y_test_gte5), nb_classes)
# -
# ## Your Turn
# Try to Fine Tune a VGG16 Network
# +
from keras.applications import VGG16
from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
from keras.applications.vgg16 import preprocess_input
from keras.layers import Input, Flatten, Dense
from keras.models import Model
import numpy as np
#Get back the convolutional part of a VGG network trained on ImageNet
model_vgg16_conv = VGG16(weights='imagenet', include_top=False)
model_vgg16_conv.summary()
#Create your own input format (here 3x200x200)
inp = Input(shape=(48,48,3),name = 'image_input')
#Use the generated model
output_vgg16_conv = model_vgg16_conv(inp)
#Add the fully-connected layers
x = Flatten(name='flatten')(output_vgg16_conv)
x = Dense(4096, activation='relu', name='fc1')(x)
x = Dense(4096, activation='relu', name='fc2')(x)
x = Dense(5, activation='softmax', name='predictions')(x)
#Create your own model
my_model = Model(input=inp, output=x)
#In the summary, weights and layers from VGG part will be hidden, but they will be fit during the training
my_model.summary()
# -
# ```python
# ...
# ...
# # Plugging new Layers
# model.add(Dense(768, activation='sigmoid'))
# model.add(Dropout(0.0))
# model.add(Dense(768, activation='sigmoid'))
# model.add(Dropout(0.0))
# model.add(Dense(n_labels, activation='softmax'))
# ```
# +
import scipy
new_shape = (48,48)
X_train_new = np.empty(shape=(X_train_gte5.shape[0],)+(48,48,3))
for idx in xrange(X_train_gte5.shape[0]):
X_train_new[idx] = np.resize(scipy.misc.imresize(X_train_gte5[idx], (new_shape)), (48, 48, 3))
X_train_new[idx] = np.resize(X_train_new[idx], (48, 48, 3))
#X_train_new = np.expand_dims(X_train_new, axis=-1)
print X_train_new.shape
X_train_new = X_train_new.astype('float32')
X_train_new /= 255
print('X_train shape:', X_train_new.shape)
print(X_train_new.shape[0], 'train samples')
print(X_train_new.shape[0], 'test samples')
# convert class vectors to binary class matrices
Y_train = np_utils.to_categorical(y_train_gte5, nb_classes)
Y_test = np_utils.to_categorical(y_test_gte5, nb_classes)
print y_train.shape
my_model.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
my_model.fit(X_train_new, Y_train,
batch_size=batch_size, nb_epoch=nb_epoch,
verbose=1)
#print('Training time: %s' % (now() - t))
#score = my_model.evaluate(X_test, Y_test, verbose=0)
#print('Test score:', score[0])
#print('Test accuracy:', score[1])
#train_model(my_model,
# (X_train_new, y_train_gte5),
# (X_test_gte5, y_test_gte5), nb_classes)
# -
| 2.4 Transfer Learning & Fine-Tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot the Env and some trees
# %load_ext autoreload
# %autoreload 2
import jpy_canvas
import random
import time
import sys
# in case you need to tweak your PYTHONPATH...
sys.path.append("../flatland")
import flatland.core.env
import flatland.utils.rendertools as rt
from flatland.envs.rail_env import RailEnv, random_rail_generator
from flatland.envs.observations import TreeObsForRailEnv
from flatland.envs.predictions import ShortestPathPredictorForRailEnv
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:90% !important; }</style>"))
# # Generate
nAgents = 3
fnMethod = random_rail_generator(cell_type_relative_proportion=[1] * 11)
env = RailEnv(width=20,
height=10,
rail_generator=fnMethod,
number_of_agents=nAgents,
obs_builder_object=TreeObsForRailEnv(max_depth=3, predictor=ShortestPathPredictorForRailEnv()))
env.reset()
# # Render
oRT = rt.RenderTool(env,gl="PILSVG")
oRT.render_env(show_observations=False,show_predictions=True)
img = oRT.get_image()
jpy_canvas.Canvas(img)
oRT = rt.RenderTool(env,gl="PIL")
oRT.render_env(show_observations=False,show_predictions=True)
img = oRT.get_image()
jpy_canvas.Canvas(img)
| notebooks/Simple_Rendering_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cycleways and their impact in New Zealand
#
# ### Aim:
# * Illustrate the current status and historical development of cycleways in New Zealand
# * Investigate the impact of cycleway development on the popularity of cycling (recreation or utility)
# * Develop my skills using Python in the Jupyter Notebooks framework to process, display and analyse data
# * Begin to develop my own processes for projects of interest such as these
#
# ### Background
#
# ###### My own learning
# This will be my first personal research project using Python on Jupyter Notebooks. I'm also generally inexperienced with programming and as a result I fully expect this to be a slow and frustrating process as I get to grips with the intricacies. However I hope that this will ultimately be a rewarding and interesting piece of work. By documenting my steps I will intend to consolidate the learning that I do, aiding me in the future. If this can be of interest to anyone else, well all the better.
#
# Although I will tidy up this notebook in order to share it, I intend to not make wholesale changes to how I undertook my analysis - that is, as I begin to understand my findings, I will develop on them as I did in the order that I did in reality, for the most part.
#
# ###### Why cycleways?
# * I am an advocate for cycling. I enjoy it for recreation and utility; I commute to work by bicycle most days and use my bike to get around the city by default. For me there are many more reasons _to_ cycle than _not to_ cycle.
#
# * However I understand that I am lucky enough to have grown up cycling; it was the default mode of transport when I was young too. Among other things, I recognise that this has given me confidence to cycle in traffic without major concern.
#
# * For me, and I think it safe to assume most people, cycling away from (particularly busy) traffic is a more positive and enjoyable experience. It is also much more accessible for those newer to cycling or less accustomed to cycling in traffic.
#
# * There is a commonly held belief that New Zealand is a forward thinking, relatively liberal country. It places high value on sustainaibility and the natural environment; as well as its citizen's health, clean air and encouraging physical exercise.
#
# * I thought it would be interesting to investigate whether or not this would translate into significant investment into cycling infrastructure. Additionally whether or not increased levels of infrastructure would deliver a discernible uptake in cycling.
# ## Datasets used
#
# ### From the New Zealand government transport data portal:
# https://www.nzta.govt.nz/planning-and-investment/learning-and-resources/transport-data/data-and-tools/
#
# ###### Physical statistics - cycleways
# "The length of cycleways within Road Controlling Authority areas, each region and NZ...over the last 10 years"
# Uploaded to my github here:
# https://github.com/phtevegibson/data-projects/blob/main/datasets/PSCycleways.xlsx?raw=true
#
# ###### Physical statistics - roads
# "The length of urban and rural, sealed and unsealed roads within Road Controlling Authority areas, each region and NZ...over the last 10 years"
# Uploaded to my github here:
# https://github.com/phtevegibson/data-projects/blob/main/datasets/PSRoads.xlsx?raw=true
#
# ### From stats and insights section of the New Zealand government transport website:
# https://www.transport.govt.nz/statistics-and-insights/household-travel
#
# ###### Regional travel: 2015-1018
# "The New Zealand Household Travel Survey is an ongoing survey of household travel conducted for the Ministry of Transport. Between 2015 and 2018, each year people in over 2,000 households throughout New Zealand were invited to participate in the survey, by recording all their travel over a 7-day period. Each person in the household was asked about their travel and other related information.
#
# Note that this travel survey captures travel in the road/footpath environment (and domestic flights and ferries) — off-road activities such as mountain biking and hiking are not included in these estimates. The 2015–2018 data is from 8,267 people in 4,144 households, collected between October 2015 and July 2018. Professional driver trips (including cycling trips such as mail and pamphlet delivery) have been excluded from the analysis."
# Uploaded to my github here:
# https://github.com/phtevegibson/data-projects/blob/main/datasets/Regions_travel-by-residents-2015_2018_revised.xlsx?raw=true
# ## Import modules to setup notebook
# +
#import useful modules
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#set the seaborn style to my preference
sns.set_style('darkgrid')
# -
# ## Physical statistics - cycleways
# First steps:
# * Read in dataset
# * Have a look at data
# * Prepare it for use
# * Quick summary
# +
#read cycleways excel data file from my github
url_git_pscycleways = 'https://github.com/phtevegibson/data-projects/blob/main/datasets/PSCycleways.xlsx?raw=true'
df_cycleways = pd.read_excel(url_git_pscycleways,sheet_name=1,header=0,engine='openpyxl')
#show size of dataframe and display first 5 rows to get an idea of what we are dealing with
print (df_cycleways.shape)
df_cycleways.head()
# +
#measure column contains only 1 unique value (kms), therefore drop this column to simplify
df_cycleways=df_cycleways.drop(columns=['Measure'])
#melt column year columns to make df narrow and deep - this makes it easier to plot
#the id_vars from column title that we don't specify (all the years) become values in a new column (name: variable)
df_cycleways=df_cycleways.melt(id_vars=['Region/Authority','Region/Authority Type'])
#rename columns to make them more user-friendly
df_cycleways=df_cycleways.rename(columns={"Region/Authority": "Area", "Region/Authority Type": "Type",
"variable": "Year", "value": "Cycleways(km)"})
#check we did what we intended
print (df_cycleways.shape)
df_cycleways.head()
# -
# We can partly summarise this data by plotting the total length of cycleways in New Zealand over the last 10 years
# +
#set a decent plot size and font scale so its readable
size = 8
fig, ax = plt.subplots(figsize=(size, size))
sns.set_theme(font_scale=0.8)
#plot just the national (New Zealand) data
sns.lineplot(data=df_cycleways.query("Area=='New Zealand'"), x="Year", y="Cycleways(km)")
#set the y axis to 0 for clarity
plt.ylim(0)
# -
# Well that's a positive for a start. We can see very clearly that the length of total cycleway in New Zealand is increasing pretty fast. That's good, but I don't really know how useful 1000, 2000 or 3000kms of cycleway really is for getting around.
# ## Accessibility
# Instead of the total length of cycleways, a more useful measure could be how much of New Zealand's road network is also serviced by a cycle specific route. That is, how much of what is accessibile by car can also be accessed by cycleway.
#
# In order to understand this, I will need to obtain the equivalent data for roads in New Zealand as we have for cycleways above.
#
# I'm making a few assumptions here which could lead to some inaccuracies. I'll briefly discuss these towards the end of the notebook.
df_cycleways=df_cycleways.groupby(by=['Type','Area','Year']).sum()
df_cycleways=df_cycleways.reset_index()
print(df_cycleways.shape)
df_cycleways.head()
# ## Physical statistics - roads
# First steps:
# * Read in dataset
# * Have a look at data
# * Prepare it for use
# * Quick summary
# +
#read roads dataset from my github
url_git = 'https://github.com/phtevegibson/data-projects/blob/main/datasets/PSRoads.xlsx?raw=true'
df_roads = pd.read_excel(url_git, engine='openpyxl', sheet_name=1,header=0)
#show size and first 5 rows
print (df_roads.shape)
df_roads.head()
# +
#melt year columns to make df narrow and deep as before
df_roads=df_roads.melt(id_vars=['Region/Authority','Region/Authority Type','UR','Measure'])
#rename columns to make them more user-friendly and consistent with cycleways df
df_roads=df_roads.rename(columns={"Region/Authority": "Area", "Region/Authority Type": "Type",
"variable": "Year", "value": "Roads(km)"})
#check
print (df_roads.shape)
df_roads.head()
# -
# ###### Lane kms
# * From a quick web search I found that _lane_ kms refers to the total length of single lane width road surface
# * That is, for a 1km long stretch of 3 lane wide motorway, the number of lane kms will be 3km, whereas the number of plain vanilla kms will be 1km
# * For this project I am only interested the length of road in terms of accessibility and how far that road can take you, therefore I will discount lane kms
# +
#drop 'lane km' as this type of measure isn't relevant to our analysis
df_roads = df_roads.drop(df_roads[df_roads['Measure']=='Lane km'].index)
#can now drop Measure column as it only contains 1 unique value (km)
df_roads=df_roads.drop(columns=['Measure'])
#our cycleways dataframe does not have values for 09/10 or the Chatham Islands so drop these too
df_roads = df_roads.drop(df_roads[df_roads['Year']=='2009/10'].index)
df_roads = df_roads.drop(df_roads[df_roads['Area']=='Chatham Islands'].index)
#check
print (df_roads.shape)
df_roads.head()
# -
# ###### UR
# For this project I have no requirement to discern between different types of road (UR column), and have no equivalent to compare this measure to in the cycleways df. I could revisit this at later stage. For now I will sum the count of kms to give us a dataframe of the same form as the cycleways data
# +
#sum types of road for each area and year. use reset index to turn the df back into a standard df from a 'groupby' type
df_roads=df_roads.groupby(by=['Type','Area','Year']).sum()
df_roads=df_roads.reset_index()
#check
print (df_roads.shape)
df_roads.head()
# -
# We can get a reasonable picture of this data by plotting all the regional data on a line plot
# +
#set a decent plot size and font scale so its readable
size = 8
fig, ax = plt.subplots(figsize=(size, size))
sns.set_theme(font_scale=0.8)
#plot just the regional data
sns.lineplot(data=df_roads.query("Type=='Regional'"), hue="Area", x="Year", y="Roads(km)")
#set the y axis to 0 for clarity
plt.ylim(0)
# -
# This gives us at least an idea of the scale of roads in New Zealand and that some regions have more roads than others (unsurprising). Perhaps more interest would be to understand the percentage increase in road length for each region, year on year.
#
# Turns out implementing this idea was a little more difficult than I anticipated but turned out to be a useful learning exercise.
#use the shift function to add a new column of the previous years' road length so that we can perform operations using it next
df_roads["Roads(km)_previous_year"] = df_roads["Roads(km)"].shift(periods=1)
df_roads.head()
# +
#function to calculate the percentage increase in road length from last year to this year
def percentage_change(year, length_current, length_previous):
#can't calculate this for our first year of data
if year == "2010/11":
return 0
#catch any dividing by 0 errors
elif length_previous == 0:
return 0
else:
return ((length_current - length_previous)/length_previous)*100
#use of apply function to apply our function above
df_roads["Road length % change"] = df_roads.apply(lambda x: percentage_change(x["Year"], x["Roads(km)"], x["Roads(km)_previous_year"]),axis=1)
#remove extra column which is no longer needed
df_roads=df_roads.drop(columns=["Roads(km)_previous_year"])
df_roads.head()
# +
#set a decent plot size and font scale so its readable
size = 4
sns.set_theme(font_scale=1)
g = sns.FacetGrid(df_roads[df_roads['Type']!='Territorial'], col="Type", hue="Area", height=size, aspect=1)
g.map(sns.lineplot, "Year", "Road length % change")
g.set_xticklabels(rotation=45)
g.add_legend()
# -
#mean change in road length nationwide:
df_roads[df_roads['Type']=='National']["Road length % change"].mean()
#largest % change, excluding territorial:
df_roads.iloc[df_roads[df_roads['Type']!='Territorial']["Road length % change"].idxmax()]
# The regional plot is quite busy so I included this as a facetgrid plot; showing the overall New Zealand trend more clearly: fairly steady year on year increase of 0.26%, except in 2018/19.
#
# A large increase is seen across the country in this year, the largest percentage increase is 4.2% on the West Coast. Whether this really was a big year for road building, or there was some change to how the road length is calculated I'm not sure. Although the fact that the % change drops back down to around its average value the following year suggests the value is accurate.
# ### Cycleways length % change
df_cycleways.head()
# +
#use the shift function to add a new column of the previous years' cycleway length
df_cycleways["Cycleways(km)_previous_year"] = df_cycleways["Cycleways(km)"].shift(periods=1)
#use of apply function to apply our function above
df_cycleways["Cycleways length % change"] = df_cycleways.apply(lambda x: percentage_change(x["Year"],
x["Cycleways(km)"],
x["Cycleways(km)_previous_year"]),
axis=1)
#remove extra column which is no longer needed
df_cycleways=df_cycleways.drop(columns=["Cycleways(km)_previous_year"])
df_cycleways.head()
# -
df_cycleways[df_cycleways['Type']=='Territorial']['Area'].nunique()
# There are 63 different territorial areas, which can make plots too busy and difficult to understand
# +
#set a decent plot size and font scale so its readable
size = 4
sns.set_theme(font_scale=1)
#don't plot territorial areas
g = sns.FacetGrid(df_cycleways.query("Type!='Territorial'"), #df_cycleways[df_cycleways['Type']!='Territorial'], ***same thing
col="Type", hue="Area", height=size, aspect=1)
g.map(sns.lineplot, "Year", "Cycleways length % change")
g.set_xticklabels(rotation=45)
g.add_legend()
# -
# Have a quick look at that big spike in 2011/12:
#largest year on year percentage change in cycleway length:
df_cycleways.iloc[df_cycleways["Cycleways length % change"].idxmax()]
# And a more general look at the Taranaki data behind that:
df_cycleways.query("Area=='Taranaki'")
# It seems the problem with this measure is how easily most areas can be dwarfed by large increases in cycleway, relative to what already existed. Makes more sense to show just the national cycleway % change, year on year, as below:
# +
#set a decent plot size and font scale so its readable
size = 5
fig, ax = plt.subplots(figsize=(size, size))
sns.set_theme(font_scale=0.8)
#plot just the regional data
sns.lineplot(data=df_cycleways.query("Type=='National'"), hue="Area", x="Year", y="Cycleways length % change")
plt.xticks(rotation=45)
plt.ylim(0,100)
# -
df_cycleways[df_cycleways['Type']=='National']['Cycleways length % change'].mean()
# ## Combine cycleways and roads data
#
# To combine the cycleways and roads datasets we'll need to be sure we are combine the right rows with each other - therefore we need to check they have the same number of rows
print(df_cycleways.shape)
print(df_roads.shape)
df_cycleways['Type'].value_counts()
df_roads['Type'].value_counts()
# It seems we have some additional territorial type areas in the roads dataset. For simplicity we will just drop all territorial type areas from both datasets
df_cycleways = df_cycleways.drop(df_cycleways[df_cycleways['Type']=='Territorial'].index)
df_roads = df_roads.drop(df_roads[df_roads['Type']=='Territorial'].index)
#Check:
print(df_cycleways.shape)
print(df_roads.shape)
# +
#create new infrastructure dataframe to combine datasets
df_infrastructure = pd.merge(df_cycleways,df_roads)
#create new column as a ratio of cycleway to road length for each area and year
df_infrastructure['Cycleway to road length(%)'] = (df_infrastructure['Cycleways(km)']/df_infrastructure['Roads(km)'])*100
df_infrastructure.head()
# -
#plot the national trend
size=5
fig, ax = plt.subplots(figsize=(size, size))
sns.set_theme(font_scale=1.05)
sns.lineplot(data=df_infrastructure.query("Area=='New Zealand'"), x="Year", y="Cycleway to road length(%)",
hue='Area',style='Area',legend=False)
plt.xticks(rotation=45)
plt.ylim(0)
# A nice positive trend here again, cycleways steadily increasing against road length year on year
# <h3>Travel section</h3>
# In order to see if we could link increasing cycleways with an uptake in cycling, I found a dataset containing the results from a survey performed by the New Zealand Government on the modes of tranport used by a selection of a households over a period
url_git = 'https://github.com/phtevegibson/data-projects/blob/main/datasets/Regions_travel-by-residents-2015_2018_revised.xlsx?raw=true'
#read excel file, each sheet becomes a value in dictionary
df_dict = pd.read_excel(url_git,engine='openpyxl',sheet_name=None)
#create list of sheet names
sheet_names = list(df_dict.keys())
sheet_names
#compare to place names in infrastructure df
list(df_infrastructure['Area'].unique())
# +
#remove first 2 sheets as not part of dataset
del(sheet_names[0:2])
#check
sheet_names
# -
# Next we better see what sort of state the data is in, on each sheet:
df_dict['Auckland'].head(20)
# I'd like to compile all sheets into 1 dataframe, therefore there are a number of steps we need to perform for each sheet. As we can see from the Auckland sheet above:
# * Make index 8 the column names
# * Remove rows preceeding index 8
# * Remove rows below index 16
# * Remove columns at positions 3, 7, 11
# * Add column specifying location (Area, for consistency)
# +
#create a new, empty df
df_travel=pd.DataFrame()
#iterate over dictionary values (dfs), performing each step above and adding it to the new travel df
for sheet in sheet_names:
df_sheet = df_dict[sheet]
df_sheet.columns = df_sheet.iloc[8]
df_sheet=df_sheet.drop(df_sheet.index[0:9])
df_sheet=df_sheet.drop(df_sheet.index[8:])
df_sheet.insert(0,column='Area',value=sheet)
df_travel=pd.concat([df_travel,df_sheet])
df_travel.columns = df_travel.columns.fillna('to_drop')
df_travel.drop('to_drop', axis = 1, inplace = True)
df_travel=df_travel.reset_index(drop=True)
df_travel.head(10)
# -
# So that we can combine this travel df with our previous infrastructure df, we need to:
# * Swap underscores for hyphens in place names
# * Re-order to 'Marlborough-Nelson-Tasman'
# * Alphabetise list
# * Put New Zealand back to the top of the list after alphabetising
# +
for i in list(df_travel[df_travel['Area']=='Manawatu_Wanganui'].index):
df_travel['Area'][i] = 'Manawatu-Whanganui'
for i in list(df_travel[df_travel['Area']=='Nelson_Marlborough_Tasman'].index):
df_travel['Area'][i] = 'Marlborough-Nelson-Tasman'
#store just NZ values as a new df
df_travel_NZ = df_travel[df_travel['Area']=='New Zealand']
#drop NZ rows
df_travel = df_travel.drop(df_travel[df_travel['Area']=='New Zealand'].index)
#sort remaining rows alphabetically
df_travel = df_travel.sort_values(by=['Area','Mode of travel'])
#return NZ rows to top of df
df_travel=pd.concat([df_travel_NZ,df_travel], ignore_index=True)
# -
# It seems important to highlight here, I would like to see if the length of road also accessed by cycleway increases the amount of people cycling over people driving. Other measures could be investigated, but for now that is the question I will try and answer
#
# I'm sure I should have highlighted this as my focus or question from the offset
# +
#to answer the above, we will only compare cyclist numbers against driver numbers - all over modes of travel will be discounted
rows_mask=df_travel['Mode of travel'].isin(['Total','2.Car/van passgr','3.Pedestrian',
'5.PT (bus/train/ferry)','6.Motorcyclist','7.Other household travel'])
df_travel=df_travel[~rows_mask]
#for simplicity, we will also only look at the counts of people with any trips, and trip legs in sample - all other counts will be discounted
columns_drop = ['Million km per year','Million hours per year','Million trip legs per year',
'Mode share of distance','Mode share of duration','Mode share of trip legs',
'Km per person per year','Trip legs per person per year','Hours per person per year']#,'Trip legs in sample'
df_travel = df_travel.drop(columns=columns_drop)
df_travel.head()
# +
#create ratio of cyclist/cycle trips to motorists/car trips
#create just car and just cycling dfs
df_travel_driver = df_travel.drop(df_travel[df_travel['Mode of travel']=='4.Cyclist'].index)
df_travel_cycle = df_travel.drop(df_travel[df_travel['Mode of travel']!='4.Cyclist'].index)
#create df combining 2 dfs above, reset their indices first so we can join them correctly
df_travel_driver=df_travel_driver.reset_index()
df_travel_cycle=df_travel_cycle.reset_index()
df_cycle_ratio = df_travel_driver.join(df_travel_cycle,lsuffix='_driver',rsuffix='_cycle')
df_cycle_ratio.head()
# +
#we have quite a lot of extra columns that we no longer need, remove them here
columns_drop = ['index_driver','Mode of travel_driver','index_cycle','Mode of travel_cycle']
df_cycle_ratio = df_cycle_ratio.drop(columns=columns_drop)
df_cycle_ratio.head()
# -
# Now we will calculate our ratios; of people who cycle to people who drive; of trips taken cycling to trips taken driving
df_cycle_ratio['people_%']=(df_cycle_ratio['Sample: People with any trips_cycle']/df_cycle_ratio['Sample: People with any trips_driver'])*100
df_cycle_ratio['trips_%']=(df_cycle_ratio['Trip legs in sample_cycle']/df_cycle_ratio['Trip legs in sample_driver'])*100
df_cycle_ratio.head()
# +
#again, remove unnecessary columns, and rename Area column
columns_drop = ['Sample: People with any trips_driver','Trip legs in sample_driver','Area_cycle',
'Sample: People with any trips_cycle', 'Trip legs in sample_cycle']
df_cycle_ratio = df_cycle_ratio.drop(columns=columns_drop)
df_cycle_ratio=df_cycle_ratio.rename(columns={"Area_driver": "Area"})
df_cycle_ratio.head()
# -
# <h3>Combine cycleways and cycle trips data</h3>
#
# Finally we can combine this snapshot of travel data with our infrastructure data to see if areas with more cycleways result in a higher proportion of cycling journeys
#
# Again, we should check our Area labels are consistent
print(df_infrastructure['Area'])
print(df_cycle_ratio['Area'])
#drop the New Zealand entry for now
df_cycle_ratio = df_cycle_ratio.drop(df_cycle_ratio[df_cycle_ratio['Area']=='New Zealand'].index)
# +
#use just regional areas (same as cycle ration df)
df_infrastructure = df_infrastructure.query("Type=='Regional'")
#use just 2016/17 values as that was the date of the most recent travel survey conducted
df_infrastructure = df_infrastructure.drop(df_infrastructure[df_infrastructure['Year']!='2016/17'].index)
df_infrastructure = df_infrastructure.reset_index(drop=True)
# +
#add the cycleway to road ratio to cycle travel ratio df
df_cycle_ratio['Cycleway to road length(%)']=df_infrastructure['Cycleway to road length(%)'].values
df_cycle_ratio.head()
# -
# ### Plot some data
#
# Now lets see if anything interesting appears:
sns.scatterplot(data=df_cycle_ratio, x="Cycleway to road length(%)", y="people_%")
plt.xlim(0)
plt.ylim(0)
# Well it was never going to be so simple and clear as we would hope. It seems there is a fairly stable proportion of people across the country, of between 5% and 10% who regularly cycle
sns.scatterplot(data=df_cycle_ratio, x="Cycleway to road length(%)", y="trips_%")
plt.xlim(0)
plt.ylim(0)
# We can begin to see somewhat more of a trend here, that could indicate that an increased ratio of cycleways to road length may result in an increase proportion of journeys being taken by bike than by driving a car
# ### Assumptions and other limitations
#
# The measure of cycleway to road length ratio as road accessibility by bike is not strictly accurate. That is, cycleways need not follow roads directly and can take their own route. However it still seems appropriate for a measure of cycleway density for an area, scaled by the amount of roads in that area
#
# There was no information available for me in terms of what constitutes a cycleway. That is, whether this measure includes any or all of:
# * cyclelanes segregated from traffic
# * on road painted cyclelanes
# * great ride style gravel cycle routes
#
# There are numerous other factors that could influence the uptake of cycling in different areas. Examples include how hilly an area is, or the typical weather there
# ## Personal evaluation
#
# As a learning exercise, the main takeaway for me here is the amount of data wrangling and manipulation that can be required. This was by far the most time consuming and frustrating part for me. I'm sure some of what I have done here could be much more elegant and efficiently performed. It was at least satisfying to see myself become more and more proficient over time
#
# It would have been satisfying to try more data analysis now that I have a better grasp of my datasets, and I may come back to this in the future. However for now I am interested in looking at other areas
#
# Ideally, it would be best to have a well defined question to answer when beginning a project such as this. In this case, I didn't realise what I would be able to do, how far I would get, or understand what sort of data I would be able to find before I began
| notebooks/new_zealand_cycleways.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Wrangle Report
# The objective of this report is to describe the wrangling process for the we rate dogs data. The process is divided into three steps:
# - Gather
# - Assess
# - Clean
#
# # Gather
# Three files are required for this step.
# - The twitter archive is downloaded manually from a link and saved to the workbook space.
# - The predictions file is hosted on Udacity’s servers and contains the output of a neural network that predicts dog breed. This one is downloaded programmatically by using the requests library.
# - The JSON file is created using Twitter’s API. It contains the tweet ID, text, retweets, and other relevant data.
#
# # Assess
# After all files are gathered, the contents are evaluated. The first method is to display a random sample of the file. Other methods include looking at data types and maximum/minimum values.
# ## Quality
# For each folder some quality issues stand out. These are issues related to completness, accuracy, validity and consistency.
#
# ### Quality issues from Twitter Archive:
# 1. in_reply_to_status_id, in_reply_to_user_id, retweeted_status_id, andretweeted_status_user_id should be integers.
# 2. timestamp should have date time attribute
# 3. Numerator/Denominator values look incorrect.
# 4. The dog types should have NaN instead of None
# 5. Some tweets have no dog names (a, the) and might be a retweet
# ### Quality issues from Image Predictions
#
# 6. 2075 rows in images versus 2356 in the twitter archive
# 7. Some jpgs are repeated
# ### Quality issues in JSON tweets
# 8. Some tweets are repeated.
# ## Tidiness
# Tidiness is related to the presentation of the data
# 1. All tables should be made into one dataset
# 2. Dog stages should be in one column
# # Clean
# The quality and tidiness issues listed in the previous section are addressed by programatic and sometimes manual methods. The first step was to merge all folders into one data with the tweet ID as the primary key.
# <br> For this step is is always useful to work outisde the main dataframe by creating copies or arrays before modifying the contents.
# # Conclusion
# The data needed for analyzing is often located somewhere else and in an ufamiliar format. Our objective is to extract, assess, and clean it for our use. In some instances it is made easy by the host's API. After gathering the data, exploration is useful to identify quality and tidiness issues. Finally, the data is cleaned based on the latter step.
# <br> In order for the results to be valid, the steps must be reproducible for users who come after us. It is also useful to work on copies of the main dataframe before modifying.
| wrangle_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="6VXPfRp-tARR" outputId="6c27acaf-2c6a-467f-fd7a-68efce98e30b"
import pandas as pd
import matplotlib.pyplot as plt
import re
import time
import warnings
import sqlite3
from sqlalchemy import create_engine # database connection
import csv
import os
warnings.filterwarnings("ignore")
import datetime as dt
import numpy as np
from nltk.corpus import stopwords
from sklearn.decomposition import TruncatedSVD
from sklearn.preprocessing import normalize
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.manifold import TSNE
import seaborn as sns
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score, log_loss
from sklearn.feature_extraction.text import TfidfVectorizer
from collections import Counter
from scipy.sparse import hstack
from sklearn.multiclass import OneVsRestClassifier
from sklearn.svm import SVC
from sklearn.model_selection import StratifiedKFold
from collections import Counter, defaultdict
from sklearn.calibration import CalibratedClassifierCV
from sklearn.naive_bayes import MultinomialNB
from sklearn.naive_bayes import GaussianNB
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
import math
from sklearn.metrics import normalized_mutual_info_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import SGDClassifier
from mlxtend.classifier import StackingClassifier
from sklearn import model_selection
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import precision_recall_curve, auc, roc_curve
# + [markdown] colab_type="text" id="ZihvUPvHtARd"
# <h1>4. Machine Learning Models </h1>
# + [markdown] colab_type="text" id="CtN9VBPutARf"
# <h2> 4.1 Reading data from file and storing into sql table </h2>
# + colab={} colab_type="code" id="owBQdjY1tARh"
#Creating db file from csv
if not os.path.isfile('train.db'):
disk_engine = create_engine('sqlite:///train.db')
start = dt.datetime.now()
chunksize = 180000
j = 0
index_start = 1
for df in pd.read_csv('final_features.csv', names=['Unnamed: 0','id','is_duplicate','cwc_min','cwc_max','csc_min','csc_max','ctc_min','ctc_max','last_word_eq','first_word_eq','abs_len_diff','mean_len','token_set_ratio','token_sort_ratio','fuzz_ratio','fuzz_partial_ratio','longest_substr_ratio','freq_qid1','freq_qid2','q1len','q2len','q1_n_words','q2_n_words','word_Common','word_Total','word_share','freq_q1+q2','freq_q1-q2','0_x','1_x','2_x','3_x','4_x','5_x','6_x','7_x','8_x','9_x','10_x','11_x','12_x','13_x','14_x','15_x','16_x','17_x','18_x','19_x','20_x','21_x','22_x','23_x','24_x','25_x','26_x','27_x','28_x','29_x','30_x','31_x','32_x','33_x','34_x','35_x','36_x','37_x','38_x','39_x','40_x','41_x','42_x','43_x','44_x','45_x','46_x','47_x','48_x','49_x','50_x','51_x','52_x','53_x','54_x','55_x','56_x','57_x','58_x','59_x','60_x','61_x','62_x','63_x','64_x','65_x','66_x','67_x','68_x','69_x','70_x','71_x','72_x','73_x','74_x','75_x','76_x','77_x','78_x','79_x','80_x','81_x','82_x','83_x','84_x','85_x','86_x','87_x','88_x','89_x','90_x','91_x','92_x','93_x','94_x','95_x','96_x','97_x','98_x','99_x','100_x','101_x','102_x','103_x','104_x','105_x','106_x','107_x','108_x','109_x','110_x','111_x','112_x','113_x','114_x','115_x','116_x','117_x','118_x','119_x','120_x','121_x','122_x','123_x','124_x','125_x','126_x','127_x','128_x','129_x','130_x','131_x','132_x','133_x','134_x','135_x','136_x','137_x','138_x','139_x','140_x','141_x','142_x','143_x','144_x','145_x','146_x','147_x','148_x','149_x','150_x','151_x','152_x','153_x','154_x','155_x','156_x','157_x','158_x','159_x','160_x','161_x','162_x','163_x','164_x','165_x','166_x','167_x','168_x','169_x','170_x','171_x','172_x','173_x','174_x','175_x','176_x','177_x','178_x','179_x','180_x','181_x','182_x','183_x','184_x','185_x','186_x','187_x','188_x','189_x','190_x','191_x','192_x','193_x','194_x','195_x','196_x','197_x','198_x','199_x','200_x','201_x','202_x','203_x','204_x','205_x','206_x','207_x','208_x','209_x','210_x','211_x','212_x','213_x','214_x','215_x','216_x','217_x','218_x','219_x','220_x','221_x','222_x','223_x','224_x','225_x','226_x','227_x','228_x','229_x','230_x','231_x','232_x','233_x','234_x','235_x','236_x','237_x','238_x','239_x','240_x','241_x','242_x','243_x','244_x','245_x','246_x','247_x','248_x','249_x','250_x','251_x','252_x','253_x','254_x','255_x','256_x','257_x','258_x','259_x','260_x','261_x','262_x','263_x','264_x','265_x','266_x','267_x','268_x','269_x','270_x','271_x','272_x','273_x','274_x','275_x','276_x','277_x','278_x','279_x','280_x','281_x','282_x','283_x','284_x','285_x','286_x','287_x','288_x','289_x','290_x','291_x','292_x','293_x','294_x','295_x','296_x','297_x','298_x','299_x','300_x','301_x','302_x','303_x','304_x','305_x','306_x','307_x','308_x','309_x','310_x','311_x','312_x','313_x','314_x','315_x','316_x','317_x','318_x','319_x','320_x','321_x','322_x','323_x','324_x','325_x','326_x','327_x','328_x','329_x','330_x','331_x','332_x','333_x','334_x','335_x','336_x','337_x','338_x','339_x','340_x','341_x','342_x','343_x','344_x','345_x','346_x','347_x','348_x','349_x','350_x','351_x','352_x','353_x','354_x','355_x','356_x','357_x','358_x','359_x','360_x','361_x','362_x','363_x','364_x','365_x','366_x','367_x','368_x','369_x','370_x','371_x','372_x','373_x','374_x','375_x','376_x','377_x','378_x','379_x','380_x','381_x','382_x','383_x','0_y','1_y','2_y','3_y','4_y','5_y','6_y','7_y','8_y','9_y','10_y','11_y','12_y','13_y','14_y','15_y','16_y','17_y','18_y','19_y','20_y','21_y','22_y','23_y','24_y','25_y','26_y','27_y','28_y','29_y','30_y','31_y','32_y','33_y','34_y','35_y','36_y','37_y','38_y','39_y','40_y','41_y','42_y','43_y','44_y','45_y','46_y','47_y','48_y','49_y','50_y','51_y','52_y','53_y','54_y','55_y','56_y','57_y','58_y','59_y','60_y','61_y','62_y','63_y','64_y','65_y','66_y','67_y','68_y','69_y','70_y','71_y','72_y','73_y','74_y','75_y','76_y','77_y','78_y','79_y','80_y','81_y','82_y','83_y','84_y','85_y','86_y','87_y','88_y','89_y','90_y','91_y','92_y','93_y','94_y','95_y','96_y','97_y','98_y','99_y','100_y','101_y','102_y','103_y','104_y','105_y','106_y','107_y','108_y','109_y','110_y','111_y','112_y','113_y','114_y','115_y','116_y','117_y','118_y','119_y','120_y','121_y','122_y','123_y','124_y','125_y','126_y','127_y','128_y','129_y','130_y','131_y','132_y','133_y','134_y','135_y','136_y','137_y','138_y','139_y','140_y','141_y','142_y','143_y','144_y','145_y','146_y','147_y','148_y','149_y','150_y','151_y','152_y','153_y','154_y','155_y','156_y','157_y','158_y','159_y','160_y','161_y','162_y','163_y','164_y','165_y','166_y','167_y','168_y','169_y','170_y','171_y','172_y','173_y','174_y','175_y','176_y','177_y','178_y','179_y','180_y','181_y','182_y','183_y','184_y','185_y','186_y','187_y','188_y','189_y','190_y','191_y','192_y','193_y','194_y','195_y','196_y','197_y','198_y','199_y','200_y','201_y','202_y','203_y','204_y','205_y','206_y','207_y','208_y','209_y','210_y','211_y','212_y','213_y','214_y','215_y','216_y','217_y','218_y','219_y','220_y','221_y','222_y','223_y','224_y','225_y','226_y','227_y','228_y','229_y','230_y','231_y','232_y','233_y','234_y','235_y','236_y','237_y','238_y','239_y','240_y','241_y','242_y','243_y','244_y','245_y','246_y','247_y','248_y','249_y','250_y','251_y','252_y','253_y','254_y','255_y','256_y','257_y','258_y','259_y','260_y','261_y','262_y','263_y','264_y','265_y','266_y','267_y','268_y','269_y','270_y','271_y','272_y','273_y','274_y','275_y','276_y','277_y','278_y','279_y','280_y','281_y','282_y','283_y','284_y','285_y','286_y','287_y','288_y','289_y','290_y','291_y','292_y','293_y','294_y','295_y','296_y','297_y','298_y','299_y','300_y','301_y','302_y','303_y','304_y','305_y','306_y','307_y','308_y','309_y','310_y','311_y','312_y','313_y','314_y','315_y','316_y','317_y','318_y','319_y','320_y','321_y','322_y','323_y','324_y','325_y','326_y','327_y','328_y','329_y','330_y','331_y','332_y','333_y','334_y','335_y','336_y','337_y','338_y','339_y','340_y','341_y','342_y','343_y','344_y','345_y','346_y','347_y','348_y','349_y','350_y','351_y','352_y','353_y','354_y','355_y','356_y','357_y','358_y','359_y','360_y','361_y','362_y','363_y','364_y','365_y','366_y','367_y','368_y','369_y','370_y','371_y','372_y','373_y','374_y','375_y','376_y','377_y','378_y','379_y','380_y','381_y','382_y','383_y'], chunksize=chunksize, iterator=True, encoding='utf-8', ):
df.index += index_start
j+=1
print('{} rows'.format(j*chunksize))
df.to_sql('data', disk_engine, if_exists='append')
index_start = df.index[-1] + 1
# + colab={} colab_type="code" id="4hpD3aBktARn"
#http://www.sqlitetutorial.net/sqlite-python/create-tables/
def create_connection(db_file):
""" create a database connection to the SQLite database
specified by db_file
:param db_file: database file
:return: Connection object or None
"""
try:
conn = sqlite3.connect(db_file)
return conn
except Error as e:
print(e)
return None
def checkTableExists(dbcon):
cursr = dbcon.cursor()
str = "select name from sqlite_master where type='table'"
table_names = cursr.execute(str)
print("Tables in the databse:")
tables =table_names.fetchall()
print(tables[0][0])
return(len(tables))
# + colab={} colab_type="code" id="nR8ZIUnttARs" outputId="810fb3fb-7da2-4b78-9e29-9edabbf68cf6"
read_db = 'train.db'
conn_r = create_connection(read_db)
checkTableExists(conn_r)
conn_r.close()
# + colab={} colab_type="code" id="SZq5gaaztARy"
# try to sample data according to the computing power you have
if os.path.isfile(read_db):
conn_r = create_connection(read_db)
if conn_r is not None:
# for selecting first 1M rows
# data = pd.read_sql_query("""SELECT * FROM data LIMIT 100001;""", conn_r)
# for selecting random points
data = pd.read_sql_query("SELECT * From data ORDER BY RANDOM() LIMIT 100001;", conn_r)
conn_r.commit()
conn_r.close()
# + colab={} colab_type="code" id="ZkeBKktKtAR3"
# remove the first row
data.drop(data.index[0], inplace=True)
y_true = data['is_duplicate']
data.drop(['Unnamed: 0', 'id','index','is_duplicate'], axis=1, inplace=True)
# + colab={} colab_type="code" id="QKSenpsmtAR9" outputId="81d890ce-df79-4402-9324-84817dbd5a7d"
data.head()
# + [markdown] colab_type="text" id="KaWHDzqUtASD"
# <h2> 4.2 Converting strings to numerics </h2>
# + colab={} colab_type="code" id="iLV60gkptASD" outputId="f297e0f4-52d5-4ab4-8a43-f0ff82f63698"
# after we read from sql table each entry was read it as a string
# we convert all the features into numaric before we apply any model
cols = list(data.columns)
for i in cols:
data[i] = data[i].apply(pd.to_numeric)
print(i)
# + colab={} colab_type="code" id="_LpfQwc9tASJ"
# https://stackoverflow.com/questions/7368789/convert-all-strings-in-a-list-to-int
y_true = list(map(int, y_true.values))
# -
data=pd.read_csv("final_features.csv")
data.head()
print("Number of data points and columns in the final dataframe:",data.shape)
#droppin the using columns and separating the features and the target variable
y_true=data.is_duplicate
data.drop(['Unnamed: 0','id','is_duplicate'],axis=1,inplace=True)
# ## 4.3 Train-test split construction
# + colab={} colab_type="code" id="3Rat2obGtASP"
#splitting the data by 70:30 ratio
X_train,X_test, y_train, y_test = train_test_split(data, y_true, stratify=y_true, test_size=0.3)
# + colab={} colab_type="code" id="1Iw9zCHqtASS" outputId="910b684b-0876-4dd8-e0d9-457846236833"
print("Number of data points in train data :",X_train.shape)
print("Number of data points in test data :",X_test.shape)
# + colab={} colab_type="code" id="0oDV15LJtASY" outputId="70a1e4eb-3f31-4f1e-a53b-ad972978505d"
print("-"*10, "Distribution of output variable in train data", "-"*10)
train_distr = Counter(y_train)
train_len = len(y_train)
print("Class 0: ",int(train_distr[0])/train_len,"Class 1: ", int(train_distr[1])/train_len)
print("\n")
print("-"*10, "Distribution of output variable in test data", "-"*10)
test_distr = Counter(y_test)
test_len = len(y_test)
print("Class 0: ",int(test_distr[0])/test_len, "Class 1: ",int(test_distr[1])/test_len)
# + colab={} colab_type="code" id="XfxcPT6jtASg"
# This function plots the confusion matrices given y_i, y_i_hat.
def plot_confusion_matrix(test_y, predict_y):
C = confusion_matrix(test_y, predict_y)
# C = 9,9 matrix, each cell (i,j) represents number of points of class i are predicted class j
A =(((C.T)/(C.sum(axis=1))).T)
#divid each element of the confusion matrix with the sum of elements in that column
# C = [[1, 2],
# [3, 4]]
# C.T = [[1, 3],
# [2, 4]]
# C.sum(axis = 1) axis=0 corresonds to columns and axis=1 corresponds to rows in two diamensional array
# C.sum(axix =1) = [[3, 7]]
# ((C.T)/(C.sum(axis=1))) = [[1/3, 3/7]
# [2/3, 4/7]]
# ((C.T)/(C.sum(axis=1))).T = [[1/3, 2/3]
# [3/7, 4/7]]
# sum of row elements = 1
B =(C/C.sum(axis=0))
#divid each element of the confusion matrix with the sum of elements in that row
# C = [[1, 2],
# [3, 4]]
# C.sum(axis = 0) axis=0 corresonds to columns and axis=1 corresponds to rows in two diamensional array
# C.sum(axix =0) = [[4, 6]]
# (C/C.sum(axis=0)) = [[1/4, 2/6],
# [3/4, 4/6]]
plt.figure(figsize=(20,4))
labels = [1,2]
# representing A in heatmap format
cmap=sns.light_palette("blue")
plt.subplot(1, 3, 1)
sns.heatmap(C, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.title("Confusion matrix")
plt.subplot(1, 3, 2)
sns.heatmap(B, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.title("Precision matrix")
plt.subplot(1, 3, 3)
# representing B in heatmap format
sns.heatmap(A, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.title("Recall matrix")
plt.show()
# + [markdown] colab_type="text" id="UStQJ5F_tASk"
# <h2> 4.4 Building a random model (Finding worst-case log-loss) </h2>
# + colab={} colab_type="code" id="qwMDqcU7tASl" outputId="c1e90d53-25ec-445b-e33a-299538520e32"
# we need to generate 9 numbers and the sum of numbers should be 1
# one solution is to genarate 9 numbers and divide each of the numbers by their sum
# ref: https://stackoverflow.com/a/18662466/4084039
# we create a output array that has exactly same size as the CV data
predicted_y = np.zeros((test_len,2))
for i in range(test_len):
rand_probs = np.random.rand(1,2)
predicted_y[i] = ((rand_probs/sum(sum(rand_probs)))[0])
print("Log loss on Test Data using Random Model",log_loss(y_test, predicted_y, eps=1e-15))
predicted_y =np.argmax(predicted_y, axis=1)
plot_confusion_matrix(y_test, predicted_y)
# + [markdown] colab_type="text" id="YgY29g_qtASq"
# <h2> 4.4 Logistic Regression with hyperparameter tuning </h2>
# -
from sklearn.linear_model import LogisticRegression
# + colab={} colab_type="code" id="Wb2tOE3GtASr" outputId="d7e4fc88-7d4e-4313-cda7-462a2409292e"
alpha = [10 ** x for x in range(-5, 2)] # hyperparam for SGD classifier.
# read more about SGDClassifier() at http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html
# ------------------------------
# default parameters
# SGDClassifier(loss=’hinge’, penalty=’l2’, alpha=0.0001, l1_ratio=0.15, fit_intercept=True, max_iter=None, tol=None,
# shuffle=True, verbose=0, epsilon=0.1, n_jobs=1, random_state=None, learning_rate=’optimal’, eta0=0.0, power_t=0.5,
# class_weight=None, warm_start=False, average=False, n_iter=None)
# some of methods
# fit(X, y[, coef_init, intercept_init, …]) Fit linear model with Stochastic Gradient Descent.
# predict(X) Predict class labels for samples in X.
#-------------------------------
# video link:
#------------------------------
log_error_array=[]
for i in alpha:
clf = LogisticRegression(C=i, penalty='l2', class_weight='balanced',n_jobs=-1,random_state=42)
clf.fit(X_train, y_train)
sig_clf = CalibratedClassifierCV(clf, method="sigmoid")
sig_clf.fit(X_train, y_train)
predict_y = sig_clf.predict_proba(X_test)
log_error_array.append(log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
print('For values of alpha = ', i, "The log loss is:",log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
fig, ax = plt.subplots()
ax.plot(alpha, log_error_array,c='g')
for i, txt in enumerate(np.round(log_error_array,3)):
ax.annotate((alpha[i],np.round(txt,3)), (alpha[i],log_error_array[i]))
plt.grid()
plt.title("Cross Validation Error for each alpha")
plt.xlabel("Alpha i's")
plt.ylabel("Error measure")
plt.show()
best_alpha = np.argmin(log_error_array)
clf = SGDClassifier(alpha=alpha[best_alpha], penalty='l2', loss='log', random_state=42)
clf.fit(X_train, y_train)
sig_clf = CalibratedClassifierCV(clf, method="sigmoid")
sig_clf.fit(X_train, y_train)
predict_y = sig_clf.predict_proba(X_train)
print('For values of best alpha = ', alpha[best_alpha], "The train log loss is:",log_loss(y_train, predict_y, labels=clf.classes_, eps=1e-15))
predict_y = sig_clf.predict_proba(X_test)
print('For values of best alpha = ', alpha[best_alpha], "The test log loss is:",log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
predicted_y =np.argmax(predict_y,axis=1)
print("Total number of data points :", len(predicted_y))
plot_confusion_matrix(y_test, predicted_y)
# + [markdown] colab_type="text" id="ouQSEnr3tASy"
# <h2> 4.5 Linear SVM with hyperparameter tuning </h2>
# -
from tqdm import tqdm
# + colab={} colab_type="code" id="AOFfZ5PLtAS0" outputId="d31eb598-e275-48cb-c49b-98e9eb76d8ba"
alpha = [10 ** x for x in range(-5, 2)] # hyperparam for SGD classifier.
# read more about SGDClassifier() at http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html
# ------------------------------
# default parameters
# SGDClassifier(loss=’hinge’, penalty=’l2’, alpha=0.0001, l1_ratio=0.15, fit_intercept=True, max_iter=None, tol=None,
# shuffle=True, verbose=0, epsilon=0.1, n_jobs=1, random_state=None, learning_rate=’optimal’, eta0=0.0, power_t=0.5,
# class_weight=None, warm_start=False, average=False, n_iter=None)
# some of methods
# fit(X, y[, coef_init, intercept_init, …]) Fit linear model with Stochastic Gradient Descent.
# predict(X) Predict class labels for samples in X.
#-------------------------------
# video link:
#------------------------------
log_error_array=[]
for i in tqdm(alpha):
clf = SGDClassifier(alpha=i, penalty='l1', loss='hinge', random_state=42)
clf.fit(X_train, y_train)
sig_clf = CalibratedClassifierCV(clf, method="sigmoid")
sig_clf.fit(X_train, y_train)
predict_y = sig_clf.predict_proba(X_test)
log_error_array.append(log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
print('For values of alpha = ', i, "The log loss is:",log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
fig, ax = plt.subplots()
ax.plot(alpha, log_error_array,c='g')
for i, txt in enumerate(np.round(log_error_array,3)):
ax.annotate((alpha[i],np.round(txt,3)), (alpha[i],log_error_array[i]))
plt.grid()
plt.title("Cross Validation Error for each alpha")
plt.xlabel("Alpha i's")
plt.ylabel("Error measure")
plt.show()
best_alpha = np.argmin(log_error_array)
clf = SGDClassifier(alpha=alpha[best_alpha], penalty='l1', loss='hinge', random_state=42)
clf.fit(X_train, y_train)
sig_clf = CalibratedClassifierCV(clf, method="sigmoid")
sig_clf.fit(X_train, y_train)
predict_y = sig_clf.predict_proba(X_train)
print('For values of best alpha = ', alpha[best_alpha], "The train log loss is:",log_loss(y_train, predict_y, labels=clf.classes_, eps=1e-15))
predict_y = sig_clf.predict_proba(X_test)
print('For values of best alpha = ', alpha[best_alpha], "The test log loss is:",log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
predicted_y =np.argmax(predict_y,axis=1)
print("Total number of data points :", len(predicted_y))
plot_confusion_matrix(y_test, predicted_y)
# + [markdown] colab_type="text" id="ZhTJgclztAS6"
# <h2> 4.6 XGBoost </h2>
# + colab={} colab_type="code" id="9U367-xetAS7" outputId="167e8588-2ac4-4c6d-ac22-f56a2fce5657"
import xgboost as xgb
params = {}
params['objective'] = 'binary:logistic'
params['eval_metric'] = 'logloss'
params['eta'] = 0.02
params['max_depth'] = 4
d_train = xgb.DMatrix(X_train, label=y_train)
d_test = xgb.DMatrix(X_test, label=y_test)
watchlist = [(d_train, 'train'), (d_test, 'valid')]
bst = xgb.train(params, d_train, 400, watchlist, early_stopping_rounds=20, verbose_eval=10)
xgdmat = xgb.DMatrix(X_train,y_train)
predict_y = bst.predict(d_test)
print("The test log loss is:",log_loss(y_test, predict_y, labels=clf.classes_, eps=1e-15))
# +
#https://blog.cambridgespark.com/hyperparameter-tuning-in-xgboost-4ff9100a3b2f
# + colab={} colab_type="code" id="6U5b17AatAS_" outputId="ca83b680-023b-4bc5-f499-8d8d85c2ff5e"
predicted_y =np.array(predict_y>0.5,dtype=int)
print("Total number of data points :", len(predicted_y))
plot_confusion_matrix(y_test, predicted_y)
# -
# https://blog.cambridgespark.com/hyperparameter-tuning-in-xgboost-4ff9100a3b2f
| 4.ML_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Extinction risks for cartilaginous fish
#
# An exploration of some of the results in [Extinction risk is most acute for the world’s largest and smallest vertebrates](https://www.pnas.org/content/114/40/10678), Ripple et al., PNAS October 3, 2017 114 (40) 10678-10683
#
# Specifically, we'll investigate how extinction risks vary by weight for cartilaginous fish. This provides some nice practice with simple linear and logistic regression, with the overall goal of explaining basic diagnostics for both methods.
#
# All of this (and more!) is in Chapters 2-5 of my Manning book, [Regression: A friendly guide](https://www.manning.com/books/regression-a-friendly-guide).
#
# This notebook and the relevant CSVs are available in my [regression repo on github](https://github.com/mbrudd/regression), along with other code and data for the book. Clone and fork at will!
# ### Imports and settings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
import statsmodels.formula.api as smf
from scipy.stats import chi2
from sklearn import metrics
import warnings
warnings.filterwarnings('ignore')
sns.set_theme()
plt.rcParams['figure.figsize'] = [8,8]
# ### Data
#
# First things first: we need data! Thanks to the good people at [ROpenSci](https://ropensci.org/), the data from [Fishbase.org](https://www.fishbase.se/search.php) is easily available in the [rfishbase](https://docs.ropensci.org/rfishbase/index.html) package.
#
fish = pd.read_csv("fish.csv")
fish.shape
fish.columns
fish = fish.filter(["Species","Length","Weight"])
fish
train = fish.dropna(axis='index')
train
sns.scatterplot(data=train, x="Length", y="Weight")
plt.title("Fish weights versus fish lengths")
train = train.assign(LogLength = np.log(train.Length), LogWeight = np.log(train.Weight))
sns.scatterplot(data=train, x="LogLength", y="LogWeight")
plt.title("Log(Weight) versus Log(Length)")
plt.axvline( np.mean( train["LogLength"] ), linestyle='--')
plt.axhline( np.mean( train["LogWeight"] ), linestyle='--')
# ### Linear regression
#
# The basic metric for the strength of a _linear_ relationship is the _correlation coefficient_:
#
train.LogLength.corr( train.LogWeight )
# This is a very strong correlation! In real life, especially in the social sciences, correlations between .3 and .7 in magnitude are much more common. Having checked the linear relationship, let's fit the regression line:
#
train_model = smf.ols( "LogWeight ~ LogLength", data=train)
train_fit = train_model.fit()
train_fit.params
#
# This model says that
#
# $$ \log{ \left( \text{Weight} \right) } ~ = ~ -3.322617 + 2.681095 \times \log{ \left( \text{Length} \right) } \ ,$$
#
# which is easier to digest after exponentiating:
#
# $$ \text{Weight} ~ = ~ e^{-3.322617} \times \text{Length}^{2.681095} ~ = ~ 0.036 \times \text{Length}^{2.681095} \ .$$
#
# This _power law relationship_ says that weight is roughly proportional to the cube of the length!
#
# ***
#
# The _null model_ predicts that _every_ needed/unseen weight equals the average of the known weights:
#
np.mean( fish["Weight"] )
np.log( np.mean( fish["Weight"] ) )
# Is the regression model better than this at predicting weights? Answering this specific question is the job of the _coefficient of determination_, denoted $R^2$.
#
# $$R^2 ~ = ~ \frac{ \text{TSS} - \text{SSR} }{ \text{TSS} } ~ = ~ 1 - \frac{ \text{SSR} }{ \text{TSS} }$$
#
# You could compute it this way...
( train_fit.centered_tss - train_fit.ssr) / train_fit.centered_tss
#
# but don't! It's already provided:
#
train_fit.rsquared
# ### Sharks!
#
# The information we need about [cartilaginous fish](https://en.wikipedia.org/wiki/Chondrichthyes) (sharks, rays, skates, sawfish, ghost sharks) comes from the [IUCN Red List](https://www.iucnredlist.org/):
sharks = pd.read_csv("chondrichthyes.csv")
sharks
sharks = sharks.join( fish.set_index("Species"), on="Species")
sharks
sharks = sharks[ sharks.Length.notna() ]
sharks = sharks[ sharks.Category != "Data Deficient" ]
sharks
# ### Data imputation
#
# Use the power law relationship to _impute_ the missing weights:
#
imp = np.exp( train_fit.params.Intercept )*np.power( sharks.Length, train_fit.params.LogLength )
sharks.Weight = sharks.Weight.where( sharks.Weight.notna(), imp )
sharks
sharks = sharks.assign(LogLength = np.log(sharks.Length),
LogWeight = np.log(sharks.Weight))
sns.scatterplot( data=sharks, x="LogLength", y="LogWeight")
plt.title("Log(Weight) versus Log(Length) for sharks")
sharks
threatened = ["Critically Endangered","Endangered","Vulnerable"]
sharks["Threatened"] = sharks["Category"].isin( threatened ).astype('int')
sharks = sharks.drop(columns = "Category")
sharks
null_prob = np.mean(sharks["Threatened"])
null_prob
sharks_model = smf.glm("Threatened ~ LogWeight", data=sharks, family=sm.families.Binomial())
sharks_fit = sharks_model.fit()
sharks_fit.params
# This model says that
#
# $$\log{ \left( \text{Odds of being threatened} \right) } ~ = ~ -3.173571 + 0.293120 \times \log{\left( \text{Weight} \right) } \ ,$$
#
# which is equivalent to a power law:
#
# $$\text{Odds of being threatened} ~ = ~ .042 \times \text{Weight}^{.293120} \ .$$
#
# In other words, bigger fish are more likely to be threatened.
#
# ***
#
# - For logistic models, the _deviance_ is analogous to the sum of squared residuals in linear regression analysis; logistic model coefficients minimize the deviance.
#
# - The _likelihood ratio statistic_ compares the deviances of the simple logistic model and the null model; it's analogous to the coefficient of determination.
#
# - Unlike the coefficient of determination, the likelihood ratio statistic defies easy interpretation. It's easy to gauge its size, though: it's a $\chi^2$ statistic with $df=1$ (_why_ this is true is another story...).
#
sharks_fit.null_deviance - sharks_fit.deviance
1 - chi2.cdf(sharks_fit.null_deviance - sharks_fit.deviance, df=1)
#
# This is astronomically small -- the logistic model that includes `LogLength` is better than the null model that ignores it!
#
# And if we plot the results, things look pretty good:
#
sns.regplot(data=sharks, x="LogWeight", y="Threatened", logistic=True, ci=None)
plt.savefig("sharks_fit.png")
# ### Model assessment
#
# Ripple et al. stop here with this particular model, but they should have assessed it carefully! We'll look at two options for what to do next.
#
# #### Binary classification and the ROC curve
#
# The naive thing to do is to compare the model's fitted probabilities to a threshold of 50% : classify the fish as `Threatened` if the fitted probability is higher than 50%, as `Not threatened` otherwise.
#
sharks["Class"] = (sharks_fit.fittedvalues > 0.50).astype(int)
sharks
pd.crosstab(sharks["Threatened"], sharks["Class"])
np.mean( sharks["Threatened"] == sharks["Class"] )
fpr, tpr, thresholds = metrics.roc_curve(sharks["Threatened"], sharks_fit.fittedvalues)
chronic_auc = metrics.auc(fpr, tpr)
chronic_auc
plt.figure()
plt.plot(fpr, tpr, label='ROC curve AUC: %0.2f' % chronic_auc)
plt.plot([0,1], [0,1], 'r--', label='Random classification')
# plt.xlim([0, 1])
# plt.ylim([0, 1.05])
plt.xlabel('False Positive Rate (1-Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.title('ROC curve for shark extinction risk classifier')
plt.legend(loc="lower right")
# #### Logistic analogues of $R^2$
# _McFadden's pseudo_-$R^2$ : replace sums of squares with deviances to measure the proportional reduction in the deviance
R2_M = 1 - (sharks_fit.deviance / sharks_fit.null_deviance)
R2_M
#
# Or use the native sums of squares in this context:
#
sharks["Null_residual"] = sharks["Threatened"] - null_prob
sharks["Residual"] = sharks["Threatened"] - sharks_fit.fittedvalues
sharks["Difference"] = sharks_fit.fittedvalues - null_prob
R2_S = np.sum(sharks["Difference"]**2) / np.sum(sharks["Null_residual"]**2)
R2_S
#
# Or compute _Tjur's coefficient of discrimination_: a good model should, on average, assign high probabilities to observed successes (1's) and low probabilities to observed failures (0's)
#
sharks["Fit_prob"] = sharks_fit.fittedvalues
sns.displot( data=sharks, x="Fit_prob", col="Threatened", binwidth=0.2)
fit_avgs = sharks.groupby("Threatened").agg(Fit_average=('Fit_prob','mean'))
fit_avgs
R2_D = fit_avgs["Fit_average"][1] - fit_avgs["Fit_average"][0]
R2_D
# Yikes! Not a very good model after all. :(
| twitch/.ipynb_checkpoints/Simple regression diagnostics-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://raw.githubusercontent.com/Qiskit/qiskit-tutorials/master/images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# # Ask the Quantum 8-ball
# Sometimes we let chance decide. Quantum uncertainty is the cornerstone design feature of Quantum computers. Once the program is executed on a Quantum computer, you will receive the most unbiased random answer, a *True* chance that classical computers incapable to provide. Think about a question that can be answered "yes or no" and run the code. Quantum 8-ball returns one of the 8 possible answers to your question. You might be already familiar with some parts of the code used in other Qiskit tutorials.
#
# The latest version of this notebook is available on https://github.com/qiskit/qiskit-tutorial.
#
# ***
# ### Contributors
# <NAME>, MBA
# begin with importing essential libraries for IBM Q
from qiskit import IBMQ, BasicAer
from qiskit import ClassicalRegister, QuantumRegister, QuantumCircuit
# To obtain $2^3 = 8$ outcomes with the equal likelyhood we need to generate 3 qubits and put them into superposition using The Hadamard gate $H$.
# set up Quantum Register and Classical Register for 3 qubits
q = QuantumRegister(3)
c = ClassicalRegister(3)
# Create a Quantum Circuit
qc = QuantumCircuit(q, c)
qc.h(q)
qc.measure(q, c)
# Visualize the circuit. Note: you need LaTeX to make visualizations. Details: https://www.latex-project.org/latex3/
qc.draw()
# Before we move on, let's assign predictions to interpret the outcomes. The particle can be observed in one of 8 states: [000, 001, 010, 011, 100, 101, 110, 111]. We link each state to the particular "answer".
def answer(result):
for key in result.keys():
state = key
print('The Quantum 8-ball says:')
if state == '000':
print('It is certain.')
elif state == '001':
print('Without a doubt.')
elif state == '010':
print('Yes - definitely.')
elif state == '011':
print('Most likely.')
elif state == '100':
print("Don't count on it.")
elif state == '101':
print('My reply is no.')
elif state == '110':
print('Very doubtful.')
else:
print('Concentrate and ask again.')
# ## Simulation
# First, we launch the program on a classical computer you are currently using (or cloud service), launch one particle, interpret and display the result.
from qiskit import execute
job = execute(qc, backend=BasicAer.get_backend('qasm_simulator'), shots=1)
result = job.result().get_counts(qc)
answer(result)
# Try to run the code multiple times to obtain different answer. The simulation is mimiking the behavior of quantum computer. States of the particle are generated using pseudo-random numbers, which give you the impression of being random, yet they are following certain prescribed rules and algorithms. Classical machines, unlike quantum computers, are inherently incapable of producing true random numbers.
# ## Running the program on real quantum computer
# We are almost set to run the program on a real quantum computer.
# load IBM Q account
IBMQ.load_accounts()
# define the least busy device
from qiskit.providers.ibmq import least_busy
backend = least_busy(IBMQ.backends(simulator=False))
print("The least busy device:",backend.name())
# Once we successfully loaded IBM Q account and connected to the least busy quantum computer, it's time to run the program on real device.
job = execute(qc, backend=backend, shots=1)
result = job.result().get_counts(qc)
answer(result)
| hello_world/quantum_8ball.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data preparation for tutorial
#
# This notebook contains the code to convert raw downloaded external data into a cleaned or simplified version for tutorial purposes.
#
#
# The raw data is expected to be in the `./raw` sub-directory (not included in the git repo).
# +
# %matplotlib inline
import geopandas
# -
# ## Countries dataset
#
# http://www.naturalearthdata.com/downloads/110m-cultural-vectors/110m-admin-0-countries/
countries = geopandas.read_file("zip://./raw/original_data_ne/ne_110m_admin_0_countries.zip")
countries.head()
len(countries)
countries_subset = countries[['ADM0_A3', 'NAME', 'CONTINENT', 'POP_EST', 'GDP_MD_EST', 'geometry']]
countries_subset.columns = countries_subset.columns.str.lower()
countries_subset = countries_subset.rename(columns={'adm0_a3': 'iso_a3'})
countries_subset.head()
countries_subset.to_file("ne_110m_admin_0_countries.shp")
# ## Natural Earth - Cities dataset
#
# http://www.naturalearthdata.com/downloads/110m-cultural-vectors/110m-populated-places/ (simple, version 4.0.0, downloaded May 2018)
cities = geopandas.read_file("zip://./raw/original_data_ne/ne_110m_populated_places_simple.zip")
cities.head()
len(cities)
cities_subset = cities[['name', 'geometry']]
cities_subset.head()
cities_subset.to_file("ne_110m_populated_places.shp")
# ## Natural Earth - Rivers dataset
#
# http://www.naturalearthdata.com/downloads/50m-physical-vectors/50m-rivers-lake-centerlines/ (version 4.0.0, downloaded May 2018)
rivers = geopandas.read_file("zip://./raw/ne_50m_rivers_lake_centerlines.zip")
rivers.head()
# Remove rows with missing geometry:
len(rivers)
rivers = rivers[~rivers.geometry.isna()].reset_index(drop=True)
len(rivers)
# Subset of the columns:
rivers_subset = rivers[['featurecla', 'name_en', 'geometry']].rename(columns={'name_en': 'name'})
rivers_subset.head()
rivers_subset.to_file("ne_50m_rivers_lake_centerlines.shp")
| data/data-preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import joblib
objects = []
with (open("../npy/E2.npy_rmses_dict.pkl", "rb")) as openfile:
while True:
try:
objects.append(pickle.load(openfile))
except EOFError:
break
# +
import re
objects = []
with (open("../npy/E2.npy_rmses_dict.pkl", "rb")) as openfile:
while True:
try:
objects.append(pickle.load(openfile))
except EOFError:
break
my_dict=objects[0]
reObj = re.compile('npy/E\w+')
ls=[]
for key in my_dict.keys():
if(reObj.match(key)):
ls.append(key)
# -
temp=[w[4:]for w in ls]
ls = [w[:-4]for w in temp]
# ls
def generate (row):
if(df['log key'].isin(ls)).any():
df['anomalies'] = 1
else:
df['anomalies'] = 0
# +
import pandas as pd
import numpy as np
df = pd.read_csv('log_value_vector.csv')
df['anomalies'] = 0
fin = pd.DataFrame
for i in ls:
df.loc[df['log key'] == i, "anomalies"] = 0
# -
df.to_csv('anomaly.csv')
objects
objects1 = []
with (open("suspicious_anomaly.pkl", "rb")) as openfile:
while True:
try:
objects1.append(pickle.load(openfile))
except EOFError:
break
objects1
import numpy as np
data = np.load('E109.npy')
# +
#data
# -
data=np.load('RMSE.csv')
num_records = len(data['npy/E0.npy'])
log_value = pd.read_csv("log_value_vector.csv")
log_value = log_value.groupby('log key')
for k, gr in log_value:
# do your stuff instead of print
print(k)
print(type(gr)) # This will output <class 'pandas.core.frame.DataFrame'>
print(gr)
# You can save each 'gr' in a csv as follows
gr.to_csv('temp/{}.csv'.format(k),index=False)
# +
import glob
log_anomaly = pd.DataFrame()
path = "temp/*.csv"
cmp = pd.read_csv('RMSE.csv')
for fname in glob.glob(path):
df = pd.read_csv(fname)
count = df[df.columns[0]].count()
count = count -1
#print(count)
for column in cmp:
columnSeriesObj = cmp[column]
if count == columnSeriesObj.count():
df['RMSE'] = cmp[column]
break
log_anomaly= log_anomaly.append(df)
#print (log_anomaly)
# -
log_anomaly.to_csv('log_anomaly.csv', index = False)
| deepgravewell/dns-query-log/result/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monoecious Hydrilla
# #### *Hydrilla verticillata [monoecious]*
# ---
# %load_ext autoreload
# %autoreload 2
# +
import sys
import numpy as np
import pandas as pd
import flbs_ais.nas as nas
# -
# #### Database Species Search Results
search_result = nas.species_search('Hydrilla', '')
search_string = nas.species_string(search_result)
print(search_string)
# ---
# ## API Query Dataframe for Monoecious Hydrilla
api_key = ""
with open("../tmp/api_key.txt", 'r') as file:
api_key = file.readline()
api_df = nas.getdf(2943, keep_columns=None, limit=-1, api_key=api_key)
print(f"Results: {len(api_df)}")
api_df.head()
# ---
# ## Most Common References for API Query Dataframe
api_ref_count = nas.ref_counts(api_df)
print(nas.ref_string(api_ref_count, limit=5))
# ---
# ## CSV File Dataframe for Monoecious Hydrilla
csv_df = nas.process_csv_df("../tmp/NAS-Data-MonoeciousHydrilla.csv")
print(f"Results: {len(csv_df)}")
csv_df.head()
# ---
# ## Most Common References for CSV File Dataframe
csv_ref_count = nas.ref_counts(csv_df)
print(nas.ref_string(csv_ref_count, limit=20))
| notebooks/MonoeciousHydrilla.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Manipulação de dados - III
# + [markdown] slideshow={"slide_type": "slide"}
# ## Agregação e agrupamento
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Agregando informações de linhas ou colunas
#
# Para agregar informações (p.ex. somar, tomar médias etc.) de linhas ou colunas podemos utilizar alguns métodos específicos já existentes em *DataFrames* e *Series*, tais como `sum`, `mean`, `cumsum` e `aggregate` (ou equivalentemente `agg`):
# + slideshow={"slide_type": "subslide"}
import pandas as pd
import numpy as np
dados_covid_PB = pd.read_csv('https://superset.plataformatarget.com.br/superset/explore_json/?form_data=%7B%22slice_id%22%3A1550%7D&csv=true',
sep=',', index_col=0)
# + slideshow={"slide_type": "fragment"}
dados_covid_PB.agg(lambda vetor: np.sum(vetor))[['casosNovos','obitosNovos']].astype('int')
# + [markdown] slideshow={"slide_type": "subslide"}
# Podemos conferir esta agregação resultante com o número de casos acumulados e óbitos acumulados
# + slideshow={"slide_type": "fragment"}
dados_covid_PB.head(9)
# + [markdown] slideshow={"slide_type": "subslide"}
# Isto também pode ser obtido utilizando o método `sum` de *DataFrames* e *Series*:
# + slideshow={"slide_type": "fragment"}
dados_covid_PB[['casosNovos','obitosNovos']].sum()
# + [markdown] slideshow={"slide_type": "subslide"}
# Podemos recriar a coluna `'obitosAcumulados'` com o método `cumsum` (soma cumulativa):
# + slideshow={"slide_type": "fragment"}
dados_covid_PB.obitosNovos.sort_index().cumsum()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Selecionando entradas distintas
#
# Para selecionar entradas distintas utilizamos o método `drop_duplicate`. Aqui, para exemplificar, vamos utilizar o banco de dados oficial sobre COVID no Brasil:
# + slideshow={"slide_type": "subslide"}
# pode levar um tempo para ler...
covid_BR = pd.read_excel('../database/HIST_PAINEL_COVIDBR_18jul2020.xlsx')
# + slideshow={"slide_type": "subslide"}
covid_BR.tail(3)
# + slideshow={"slide_type": "subslide"}
# resumo da tabela
covid_BR.info()
# + slideshow={"slide_type": "subslide"}
# todos os estados únicos
covid_BR.estado.drop_duplicates().array
# + slideshow={"slide_type": "fragment"}
# ordena alfabeticamente
covid_BR.estado.drop_duplicates().dropna().sort_values().array
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Agrupando dados por valores em colunas e agregando os resultados
#
# Vamos determinar uma coluna para agrupar. Consideraremos o *DataFrame* `covid_BR`e selecionaremos os estados *PB*, *PE*, *RJ* e *SP* para realizar análises agrupando os resultados por estados.
# + slideshow={"slide_type": "subslide"}
covid_BR.query('estado in ["PB", "PE", "RJ", "SP"]')
# + [markdown] slideshow={"slide_type": "subslide"}
# Inspecionando o conjunto de dados, observamos que os dados para estado são apresentados com o valor `NaN` para `codmun` e quando `codmun` possui um valor diferente de `NaN`, o resultado é apenas para o município do código em questão.
#
# Como estamos interessados nos valores por estado, vamos selecionar apenas os dados com `codmun` contendo `NaN`.
# + slideshow={"slide_type": "fragment"}
covid_estados = covid_BR.query('estado in ["PB", "PE", "RJ", "SP"]')
covid_apenas_estados = covid_estados.loc[covid_estados['codmun'].isna()]
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos agora selecionar apenas as colunas de interesse. Para tanto, vejamos os nomes das colunas:
# + slideshow={"slide_type": "fragment"}
covid_apenas_estados.columns
# + slideshow={"slide_type": "fragment"}
covid_apenas_estados = covid_apenas_estados[['estado', 'data', 'casosNovos', 'obitosNovos']]
# + [markdown] slideshow={"slide_type": "subslide"}
# A data parece ser o *index* natural, já que o *index* atual não representa nada. Observe que teremos *index* repetidos, pois teremos as mesmas datas em estados diferentes.
# + slideshow={"slide_type": "fragment"}
covid_apenas_estados
# + slideshow={"slide_type": "subslide"}
covid_apenas_estados = covid_apenas_estados.set_index('data')
# + slideshow={"slide_type": "fragment"}
covid_apenas_estados
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Agrupando com o método *groupby*
#
# Podemos escolher uma (ou mais colunas, incluindo o índice) para agrupar os dados. Ao agruparmos os dados, receberemos um objeto do tipo `DataFrameGroupBy`. Para vermos os resultados, devemos agregar os valores:
# + slideshow={"slide_type": "fragment"}
covid_estados_agrupado = covid_apenas_estados.groupby('estado')
# + slideshow={"slide_type": "fragment"}
covid_estados_agrupado.sum().rename({'casosNovos':'Casos Totais', 'obitosNovos':'Obitos Totais'},axis=1)
# + [markdown] slideshow={"slide_type": "subslide"}
# Podemos agrupar por mais de uma coluna. Vamos fazer dois grupos. *grupo_1* formado por PB e PE e *grupo_2* formado por RJ e SP. Em seguida, vamos agrupar por grupo e por data:
# + slideshow={"slide_type": "fragment"}
covid_estados_grupos = covid_apenas_estados.copy()
col_grupos = covid_estados_grupos.estado.map(lambda estado: 'grupo_1' if estado in ['PB','PE']
else 'grupo_2')
covid_estados_grupos['grupo'] = col_grupos
# + slideshow={"slide_type": "fragment"}
covid_estados_grupos
# + [markdown] slideshow={"slide_type": "subslide"}
# Agora vamos agrupar e agregar:
# + slideshow={"slide_type": "fragment"}
covid_grupo_agrupado = covid_estados_grupos.groupby(['grupo','data'])
# + slideshow={"slide_type": "fragment"}
covid_grupo_agrupado.sum()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Mesclando *DataFrames*
#
# Vamos agora ver algumas formas de juntar dois ou mais *DataFrames* com *index* ou colunas em comum para formar um novo *DataFrame*.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Mesclando *DataFrames* através de concatenações
#
# Concatenar nada mais é do que "colar" dois ou mais *DataFrames*. Podemos concatenar por linhas ou por colunas.
#
# A função que realiza a concatenação é `concat`. Os dois argumentos mais utilizados são a lista de *DataFrames* a serem concatenados e `axis`, onde `axis = 0` indica concatenação por linha (um *DataFrame* "embaixo" do outro) e `axis=1` indica concatenação por coluna (um *DataFrame* ao lado do outro).
# + [markdown] slideshow={"slide_type": "subslide"}
# Relembre do *DataFrame* `df_dict_series`:
# + slideshow={"slide_type": "fragment"}
df_dict_series = pd.read_csv('../database/df_dict_series.csv')
df_dict_series
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos criar um novo, com novas pessoas:
# + slideshow={"slide_type": "fragment"}
serie_Idade_nova = pd.Series({'Augusto':13, 'André': 17, 'Alexandre': 45}, name="Idade")
serie_Peso_novo = pd.Series({'Augusto':95, 'André': 65, 'Alexandre': 83}, name="Peso")
serie_Altura_nova = pd.Series({'Augusto':192, 'André': 175, 'Alexandre': 177}, name="Altura")
serie_sobrenome = pd.Series({'Augusto':'Castro', 'André':'Castro', 'Alexandre':'Castro'}, name='Sobrenome')
dicionario_novo = {'Sobrenome':serie_sobrenome, 'Peso': serie_Peso_novo,
'Idade': serie_Idade_nova, 'Altura': serie_Altura_nova}
df_novo = pd.DataFrame(dicionario_novo)
df_novo = df_novo.assign(IMC=round(df_novo.eval('Peso/(Altura/100)**2'),2))
# + slideshow={"slide_type": "subslide"}
df_novo
# + [markdown] slideshow={"slide_type": "subslide"}
# Agora vamos concatená-los:
# + slideshow={"slide_type": "fragment"}
pd.concat([df_dict_series,df_novo])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Concatenando por coluna
#
# Para exemplificar vamos considerar os dados de COVID da Paraíba, selecionando casos novos e óbitos novos, e vamos obter dos dados do Brasil apenas os casos e óbitos diários do país, e vamos concatená-los por coluna.
# + slideshow={"slide_type": "fragment"}
covid_PB_casos_obitos = dados_covid_PB[['casosNovos','obitosNovos']]
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos tratar os dados do Brasil:
# + slideshow={"slide_type": "fragment"}
covid_BR_casos_obitos = covid_BR.query('regiao=="Brasil"')
covid_BR_casos_obitos = covid_BR_casos_obitos.set_index('data')
covid_BR_casos_obitos = covid_BR_casos_obitos[['casosNovos','obitosNovos']].rename({
'casosNovos':'casosBR', 'obitosNovos':'obitosBR'}, axis=1)
# + slideshow={"slide_type": "subslide"}
covid_PB_casos_obitos
# + slideshow={"slide_type": "fragment"}
covid_BR_casos_obitos
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos agora concatená-los por coluna:
# + slideshow={"slide_type": "subslide"}
pd.concat([covid_PB_casos_obitos, covid_BR_casos_obitos], axis=1)
# + [markdown] slideshow={"slide_type": "subslide"}
# Para um polimento final, vamos substituir os valores `NaN` que ocorreram antes do dia 13 de julho por 0. Para tanto, a forma ideal é utilizando o método `map`:
# + slideshow={"slide_type": "subslide"}
dados_PB_BR = pd.concat([covid_PB_casos_obitos, covid_BR_casos_obitos], axis=1)
dados_PB_BR['casosNovos'] = dados_PB_BR.casosNovos.map(lambda caso: 0 if np.isnan(caso) else caso).astype('int')
dados_PB_BR['obitosNovos'] = dados_PB_BR.obitosNovos.map(lambda obito: 0 if np.isnan(obito) else obito).astype('int')
dados_PB_BR
# + [markdown] slideshow={"slide_type": "slide"}
# ### Mesclando *DataFrames* através de *joins*
#
# Para realizar *joins* iremos utilizar a função `merge` do *pandas*. *joins* tomam duas tabelas, uma tabela à esquerda e uma à direita e retornam uma terceira tabela contendo a união das colunas das duas tabelas.
# + [markdown] slideshow={"slide_type": "subslide"}
# Existem 4 tipos de *joins*:
#
# * *left join*: Apenas irão aparecer os *index* (da linha) que existem na tabela à esquerda;
# * *right join*: Apenas irão aparecer os *index* (da linha) que existem na tabela à direita;
# * *inner join*: Apenas irão aparecer os *index* que existem nas duas tabelas;
# * *full join* ou *outer join*: irão aparecer todos os *index* das duas tabelas.
# + [markdown] slideshow={"slide_type": "subslide"}
# Para exemplificar vamos considerar dois *DataFrames* (aqui teremos menos linhas, com nomes e dados fictícios). O primeiro *DataFrame* consistirá de nomes de alunos, CPF e matrícula da UFPB (*nome_cpf_mat*). O segundo *DataFrame* consistirá de nome, CPF e e-mail (*nome_cpf_email*). Nosso objetivo é criar um novo *DataFrame* contendo Nome, CPF, matrícula e e-mail.
# + [markdown] slideshow={"slide_type": "subslide"}
# Temos ainda as seguintes situações:
#
# - No *DataFrame* *nome_cpf_mat* existem alunos que não estão presentes no *nome_cpf_email*, pois não enviaram esta informação.
#
# - No *DataFrame* *nome_cpf_email* existem alunos que não estão presentes no *nome_cpf_mat* pois estes não são alunos da UFPB.
# + slideshow={"slide_type": "subslide"}
nome_cpf_mat = pd.read_csv('../database/nome_cpf_mat.csv')
nome_cpf_email = pd.read_csv('../database/nome_cpf_email.csv')
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos agora examinar os *DataFrames*. Como são bem simples, basta realizar *prints* deles.
# + slideshow={"slide_type": "subslide"}
nome_cpf_mat
# + slideshow={"slide_type": "subslide"}
nome_cpf_email
# + [markdown] slideshow={"slide_type": "subslide"}
# Tipicamente é bom possuir *index* únicos. Neste sentido, vamos definir o CPF como *index*:
# + slideshow={"slide_type": "fragment"}
nome_cpf_mat = nome_cpf_mat.set_index('CPF')
nome_cpf_email = nome_cpf_email.set_index('CPF')
nome_cpf_mat, nome_cpf_email
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos agora realizar um **left** join com o *DataFrame* **nome_cpf_mat** ficando à esquerda (neste caso, apenas alunos com matrícula irão aparecer):
# + slideshow={"slide_type": "fragment"}
pd.merge(nome_cpf_mat, nome_cpf_email, how = 'left', on = ['Nome','CPF'])
# + [markdown] slideshow={"slide_type": "subslide"}
# - Na opção *how* dizemos qual o tipo de *join* que queremos realizar.
#
# - Na opção *on* dizemos quais as colunas que existem em comum nos *DataFrames*.
#
# Veja o que aconteceria se informássemos apenas que o *CPF* está presente nos dois *DataFrames*:
# + slideshow={"slide_type": "fragment"}
pd.merge(nome_cpf_mat, nome_cpf_email, how = 'left', on = 'CPF')
# + [markdown] slideshow={"slide_type": "subslide"}
# Observe que os nomes dos alunos que estão na segunda tabela ficam indeterminados na coluna *Nome_y*.
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos agora realizar um **right** join com o *DataFrame* **nome_cpf_mat** ficando à esquerda (neste caso, apenas alunos **com e-mail** irão aparecer):
# + slideshow={"slide_type": "subslide"}
pd.merge(nome_cpf_mat, nome_cpf_email, how = 'right', on = ['Nome','CPF'])
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos agora realizar um **inner** join com o *DataFrame* **nome_cpf_mat** ficando à esquerda (neste caso, apenas alunos **com matrícula e com e-mail** irão aparecer):
# + slideshow={"slide_type": "fragment"}
pd.merge(nome_cpf_mat, nome_cpf_email, how = ' ', on = ['Nome','CPF'])
# + [markdown] slideshow={"slide_type": "subslide"}
# Por fim, vamos agora realizar um **outer** ou **full** join com o *DataFrame* **nome_cpf_mat** ficando à esquerda (neste caso, **todos** os alunos irão aparecer):
# + slideshow={"slide_type": "fragment"}
pd.merge(nome_cpf_mat, nome_cpf_email, how = 'outer', on = ['Nome','CPF'])
# + [markdown] slideshow={"slide_type": "slide"}
# ### Os métodos *apply*, *map* e *applymap*
#
# A ideia é relativamente simples. Os três métodos são vetorizados e aplicam uma função ou uma substituição via dicionário de tal forma que:
#
# * *apply* é realizado via linha ou coluna em um *DataFrame*;
# * *map* é aplicado a cada elemento de uma *Series*;
# * *applymap* é aplicado a cada elemento de um *DataFrame*.
#
# Já vimos diversos exemplos de uso de `map`. Vejamos exemplos de `applymap` e `apply`.
# + [markdown] slideshow={"slide_type": "subslide"}
# * Neste exemplo vamos retomar a concatenação entre os dados da Paraíba e do Brasil, porém iremos substituir *todos* os valores de `NaN` por zero, usando o métodp `applymap`.
# + slideshow={"slide_type": "fragment"}
dados_PB_BR = pd.concat([covid_PB_casos_obitos, covid_BR_casos_obitos], axis=1)
dados_PB_BR.applymap(lambda valor: 0 if np.isnan(valor) else valor)
# + [markdown] slideshow={"slide_type": "subslide"}
# Vamos utilizar `apply` para realizar a soma de casos e óbitos através de mais de uma forma
# + slideshow={"slide_type": "fragment"}
dados_PB_BR.apply(lambda x: np.sum(x)).astype('int')
# + [markdown] slideshow={"slide_type": "subslide"}
# Se quisermos realizar a operação por linhas, basta utilizar o argumento `axis=1`:
# + slideshow={"slide_type": "fragment"}
dados_PB_BR.apply(lambda x: (x>0).all(), axis=1)
| _build/html/_sources/rise/10c-pandas-agrupamento-rise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
import numpy as np
import pandas as pd
# 統計用ツール
import statsmodels.api as sm
import statsmodels.tsa.api as tsa
from patsy import dmatrices
#描画
import matplotlib.pyplot as plt
from pandas.tools.plotting import autocorrelation_plot
#株価
import pandas as pd
import pandas_datareader.data as web
import datetime
#深層学習
import chainer
from chainer import cuda, Function, gradient_check, Variable, optimizers, serializers, utils
from chainer import Link, Chain, ChainList
import chainer.functions as F
import chainer.links as L
#k平均法
from sklearn.cluster import KMeans
# -
s = (1990, 1, 1)
e = (2015, 9, 30)
t = 'Adj Close'
start = datetime.datetime(s[0], s[1], s[2])
end = datetime.datetime(e[0], e[1], e[2])
SaP = web.DataReader('GSPC', 'yahoo', start, end)[t]
N225 = web.DataReader('^N225', 'yahoo', start, end)[t]
DJI = web.DataReader('^DJI', 'yahoo', start, end)[t]
IXIC = web.DataReader('^IXIC', 'yahoo', start, end)[t]
SaP = np.array([SaP[1:].values - SaP[:-1].values]).T*100
def fuzzy(SaP):
kms = KMeans(n_clusters=3).fit_predict(SaP)
mean1 = SaP[kms == 0].mean()
var1 = SaP[kms == 0].var()
mean2 = SaP[kms == 1].mean()
var2 = SaP[kms == 1].var()
mean3 = SaP[kms == 2].mean()
var3 = SaP[kms == 2].var()
for t in range(100):
f = SaP[t:t+20]
v = np.array([])
for i in range(20):
v1 = np.round(np.exp(-1*(f[i] - mean1)**2/var1), 5)
v2 = np.round(np.exp(-1*(f[i] - mean2)**2/var2), 5)
v3 = np.round(np.exp(-1*(f[i] - mean3)**2/var3), 5)
v = np.append(v, v1)
v = np.append(v, v2)
v = np.append(v, v3)
if t != 0:
fs = np.c_[fs, v]
else:
fs = np.array([v]).T
return fs
class DRNN(object):
def __init__(self, data):
self.data = data
self.model = DeepLearning()
self.optimizer = optimizers.Adam()
self.optimizer.setup(self.model)
def fuzzy(self):
self.fs = fuzzy(self.data).T
def autoencorder(self, num):
ft = np.array(self.fs, dtype='float32')
n, m = ft.shape
bs = 5
for j in range(num):
sffindx = np.random.permutation(n)
for i in range(0, n, bs):
self.ib = i+bs
x = Variable(ft[sffindx[i:(i+bs) if (i+bs) < n else n]])
self.model.zerograds()
loss = self.model.ae(x)
loss.backward()
self.optimizer.update()
if j % 1000 == 0:
print('epoch:', j)
print('train mean loss={}'.format(loss.data))
print(' - - - - - - - - - ')
def reinforcement(self, n):
for i in range(n):
self.optimizer = optimizers.Adam()
self.optimizer.setup(self.model)
for t in range(100):
loss = self.model.rnn(self.fs, t)
loss.backward()
self.optimizer.update()
print('epoch:', i)
print('profit={}'.format(-loss.data[0][0]))
print(' - - - - - - - - - ')
def initialization(self, n=3000, m=100):
self.fuzzy()
self.autoencorder(n)
self.reinforcement(m)
def learning(self, maxc=10):
c = 0
while c < maxc:
for t in range(100):
loss = self.model(self.fs, t)
loss.backward()
self.optimizer.update()
print('epoch:', c)
print('time:', t)
print('profit={}'.format(-loss.data[0][0]))
print(' - - - - - - - - - ')
c += 1
def strategy():
self.model.delta(self.fs)
class DeepLearning(chainer.Chain):
def __init__(self):
super(DeepLearning, self).__init__(
w = L.Linear(20, 1),
b = L.Linear(1, 1),
u = L.Linear(1, 1),
l1 = L.Linear(60, 60),
l2 = L.Linear(60, 60),
l3 = L.Linear(60, 60),
l4 = L.Linear(60, 20),
l5 = L.Linear(20, 60),
m1 = L.Linear(1, 60),
m2 = L.Linear(1, 60),
m3 = L.Linear(1, 60),
m4 = L.Linear(1, 20),
)
def ae(self, x):
bv, fv = self.ae_fwd(x)
loss = F.mean_squared_error(bv, x)
return loss
def ae_fwd(self, x):
h1 = F.sigmoid(self.l1(x))
h2 = F.sigmoid(self.l2(h1))
h3 = F.sigmoid(self.l3(h2))
fv = F.sigmoid(self.l4(h3))
bv = self.l5(fv)
return bv, fv
def rnn(self, fs, t, c=1):
fs = np.array(fs, dtype='float32')
bv, Fs = self.ae_fwd(Variable(fs))
Fs = Fs.data
d_1 = Variable(np.zeros((1, 1), dtype=np.float32))
old_grad = np.array([np.zeros(21)], dtype='float32')
one = Variable(np.array([[1]], dtype=np.float32))
zero = Variable(np.array([[0]], dtype=np.float32))
loss = 0
for i in range(len(Fs[:, 0])):
x_k = Variable(np.array([Fs[i]], dtype=np.float32))
d = F.tanh(self.w(x_k) + self.b(one) + self.u(d_1))
z = Variable(np.array([[Fs[i][-1]]], dtype=np.float32))
R = d_1.__rmatmul__(z) - c*(d - d_1).__abs__()
loss += R
d_1 = d
return -loss
def delta(self, Fs):
one = Variable(np.array([[1]], dtype=np.float32))
d_1 = Variable(np.zeros((1, 1), dtype=np.float32))
d = np.zeros(100)
d[0] = 1
for i in range(1, len(Fs[:, 0])):
Ft = Variable(np.array([Fs[i]], dtype=np.float32))
ds = F.tanh(self.w(Ft) + self.b(one) + self.u(d_1))
d[i] = ds.data[0]
return d
def __call__(self, fs, t, c=1):
fs = np.array(fs, dtype='float32')
bv, Fs = self.ae_fwd(Variable(fs))
Fs_Var = Fs
d_1 = Variable(np.zeros((1, 1), dtype=np.float32))
old_grad = np.array([np.zeros(21)], dtype='float32')
one = Variable(np.array([[1]], dtype=np.float32))
zero = Variable(np.array([[0]], dtype=np.float32))
loss = 0
Fs = Fs.data
for i in range(len(Fs[:, 0])):
d = self.delta(Fs_Var.data).data
if i == t:
ft = Variable(np.array([fs[t]], dtype=np.float32))
d = Variable(np.array([[d[t]]], dtype=np.float32))
h1 = F.sigmoid(self.l1(ft) + self.m1(d))
h2 = F.sigmoid(self.l2(h1) + self.m2(d))
h3 = F.sigmoid(self.l3(h2) + self.m3(d))
Ft = F.sigmoid(self.l4(h3) + self.m4(d))
d = F.tanh(self.w(Ft) + self.b(one) + self.u(d_1))
z = Variable(np.array([[Fs[i][-1]]], dtype=np.float32))
R = d_1.__rmatmul__(z) - c*(d - d_1).__abs__()
loss += R
d_1 = d
else:
Ft = Variable(np.array([Fs[i]], dtype=np.float32))
d = F.tanh(self.w(Ft) + self.b(one) + self.u(d_1))
z = Variable(np.array([[Fs[i][-1]]], dtype=np.float32))
R = d_1.__rmatmul__(z) - c*(d - d_1).__abs__()
loss += R
d_1 = d
return -loss
drnn = DRNN(SaP)
drnn.initialization()
drnn.learning()
bv, Ft = drnn.model.ae_fwd(np.array(drnn.fs, dtype=np.float32))
drnn.model.delta(Ft.data)
| DRNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Diversificación y fuentes de riesgo en un portafolio II - Una ilustración con mercados internacionales.
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/5/5f/Map_International_Markets.jpg" width="500px" height="300px" />
#
# > Entonces, la clase pasada vimos cómo afecta la correlación entre pares de activos en un portafolio. Dijimos que como un par de activos nunca tienen correlación perfecta, al combinarlos en un portafolio siempre conseguimos diversificación del riesgo.
#
# > Vimos también que no todo el riesgo se puede diversificar. Dos fuentes de riesgo:
# > - Sistemático: afecta de igual manera a todos los activos. No se puede diversificar.
# > - Idiosincrático: afecta a cada activo en particular por razones específicas. Se puede diversificar.
#
# En esta clase veremos un ejemplo de diversificación en un portafolio, usando datos de mercados de activos internacionales.
#
# En el camino, definiremos términos como *frontera media/varianza*, *portafolio de varianza mínima* y *portafolios eficientes*, los cuales son básicos para la construcción de la **teoría moderna de portafolios**.
#
# Estos portafolios los aprenderemos a obtener formalmente en el siguiente módulo. Por ahora nos bastará con agarrar intuición.
# **Objetivo:**
# - Ver los beneficios de la diversificación ilustrativamente.
# - ¿Qué es la frontera de mínima varianza?
# - ¿Qué son el portafolio de varianza mínima y portafolios eficientes?
#
# *Referencia:*
# - Notas del curso "Portfolio Selection and Risk Management", Rice University, disponible en Coursera.
# ___
# ## 1. Ejemplo
#
# **Los datos:** tenemos el siguiente reporte de rendimientos esperados y volatilidad (anuales) para los mercados de acciones en los países integrantes del $G5$: EU, RU, Francia, Alemania y Japón.
# Importamos pandas y numpy
import pandas as pd
import numpy as np
# +
# Resumen en base anual de rendimientos esperados y volatilidades
annual_ret_summ = pd.DataFrame(columns=['EU', 'RU', 'Francia', 'Alemania', 'Japon'], index=['Media', 'Volatilidad'])
annual_ret_summ.loc['Media'] = np.array([0.1355, 0.1589, 0.1519, 0.1435, 0.1497])
annual_ret_summ.loc['Volatilidad'] = np.array([0.1535, 0.2430, 0.2324, 0.2038, 0.2298])
annual_ret_summ.round(4)
# -
# ¿Qué podemos notar?
# - Los rendimientos esperados rondan todos por los mismos valores 14%-15%.
# - En cuanto al riesgo, la medida de riesgo del mercado de Estados Unidos es mucho menor respecto a las demás.
# Además, tenemos el siguiente reporte de la matriz de correlación:
# Matriz de correlación
corr = pd.DataFrame(data= np.array([[1.0000, 0.5003, 0.4398, 0.3681, 0.2663],
[0.5003, 1.0000, 0.5420, 0.4265, 0.3581],
[0.4398, 0.5420, 1.0000, 0.6032, 0.3923],
[0.3681, 0.4265, 0.6032, 1.0000, 0.3663],
[0.2663, 0.3581, 0.3923, 0.3663, 1.0000]]),
columns=annual_ret_summ.columns, index=annual_ret_summ.columns)
corr.round(4)
# ¿Qué se puede observar acerca de la matriz de correlación?
#
# - Los índices con mayor correlación son los de Alemania y Francia.
# - Los índices con menor correlación son los de Japón y Estados Unidos.
# Recordar: correlaciones bajas significan una gran oportunidad para diversificación.
# ### Nos enfocaremos entonces únicamente en dos mercados: EU y Japón
#
# - ¿Cómo construiríamos un portafolio que consiste de los mercados de acciones de EU y Japón?
# - ¿Cuáles serían las posibles combinaciones?
# #### 1. Supongamos que $w$ es la participación del mercado de EU en nuestro portafolio.
# - ¿Cuál es la participación del mercado de Japón entonces?: $1-w$
#
# - Luego, nuestras fórmulas de rendimiento esperado y varianza de portafolios son:
#
# $$E[r_p]=wE[r_{EU}]+(1-w)E[r_J]$$
#
# $$\sigma_p^2=w^2\sigma_{EU}^2+(1-w)^2\sigma_J^2+2w(1-w)\sigma_{EU,J}$$
# #### 2. Con lo anterior...
# - podemos variar $w$ con pasos pequeños entre $0$ y $1$, y
# - calcular el rendimiento esperado y volatilidad para cada valor de $w$.
# Vector de w variando entre 0 y 1 con n pasos
n = 100
w = np.linspace(0, 1, n)
# Rendimientos esperados individuales
# Activo1: EU, Activo2:Japon
E1 = annual_ret_summ.loc['Media', 'EU']
E2 = annual_ret_summ.loc['Media', 'Japon']
# Volatilidades individuales
s1 = annual_ret_summ.loc['Volatilidad', 'EU']
s2 = annual_ret_summ.loc['Volatilidad', 'Japon']
# Correlacion
r12 = corr.loc['Japon', 'EU']
# Crear un DataFrame cuyas columnas sean rendimiento
# y volatilidad del portafolio para cada una de las w
# generadas
portafolios = pd.DataFrame(columns=['EP', 'sp'], index=w)
portafolios['EP'] = w*E1+(1-w)*E2
portafolios['sp'] = (w**2*s1**2+(1-w)**2*s2**2+2*w*(1-w)*s1*s2*r12)**0.5
portafolios
# #### 3. Finalmente,
# - cada una de las combinaciones las podemos graficar en el espacio de rendimiento esperado (eje $y$) contra volatilidad (eje $x$).
# Importar matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# Graficar el lugar geométrico de los portafolios en el
# espacio rendimiento esperado vs. volatilidad.
# Especificar también los puntos relativos a los casos
# extremos.
plt.figure(figsize=(12,8))
plt.plot(portafolios['sp'], portafolios['EP'], label='Portafolios')
plt.plot(s1, E1, 'b*', ms=10, label='EU')
plt.plot(s2, E2, 'r*', ms=10, label='Japon')
plt.grid()
plt.legend(loc='best')
plt.xlabel('Volatilidad ($\sigma$)')
plt.ylabel('Rendimiento esperado ($E[r]$)')
# #### De la gráfica,
# 1. Ver casos extremos.
# 2. ¿Conviene invertir 100% en el mercado de EU? ¿Porqué?
# 3. ¿Porqué ocurre esto?
# 4. Definición: frontera de mínima varianza. Caso particular: dos activos.
# 5. Definición: portafolio de varianza mínima.
# 6. Definición: portafolios eficientes.
#
# #### 1. Definición (frontera de mínima varianza): es el lugar geométrico en el espacio de rendimiento esperado vs. volatilidad correspondiente a los portafolios con menor varianza.
#
# #### 2. Definición (portafolio de mínima varianza): es el portafolio con menor varianza. Es el punto que está más a la izquierda sobre la frontera de mínima varianza.
#
# #### 3. Definición (portafolios eficientes): son los puntos sobre la frontera de mínima varianza en la parte superior y a partir del portafolio de mínima varianza.
# ___
# ## 2. ¿Cómo hallar el portafolio de varianza mínima?
#
# Bien, esta será nuestra primera selección de portafolio. Si bien se hace de manera básica e intuitiva, nos servirá como introducción al siguiente módulo.
#
# **Comentario:** estrictamente, el portafolio que está más a la izquierda en la curva de arriba es el de *volatilidad mínima*. Sin embargo, como tanto la volatilidad es una medida siempre positiva, minimizar la volatilidad equivale a minimizar la varianza. Por lo anterior, llamamos a dicho portafolio, el portafolio de *varianza mínima*.
# De modo que la búsqueda del portafolio de varianza mínima corresponde a la solución del siguiente problema de optimización:
#
# - Para un portafolio con $n$ activos ($\boldsymbol{w}=[w_1,\dots,w_n]^T\in\mathbb{R}^n$):
# \begin{align*}
# &\min_{\boldsymbol{w}} & \sigma_p^2=\boldsymbol{w}^T\Sigma\boldsymbol{w}\\
# &\text{s.t.} \qquad & \boldsymbol{w}\geq0,\\
# & & w_1+\dots+w_n=1
# \end{align*}
# donde $\Sigma$ es la matriz de varianza-covarianza de los rendimientos de los $n$ activos.
# - En particular, para un portafolio con dos activos el problema anterior se reduce a:
# \begin{align*}
# &\min_{w_1,w_2}\sigma_p^2=w_1^2\sigma_1^2+w_2^2\sigma_2^2+2w_1w_2\rho_{12}\sigma_1\sigma_2\\
# &\text{s.t.} \qquad w_1,w_2\geq0,
# \end{align*}
# donde $\sigma_1,\sigma_2$ son las volatilidades de los activos individuales y $\rho_{12}$ es la correlación entre los activos. Equivalentemente, haciendo $w_1=w$ y $w_2=1-w$, el problema anterior se puede reescribir de la siguiente manera:
# \begin{align*}
# &\min_{w}\sigma_p^2=w^2\sigma_1^2+(1-w)^2\sigma_2^2+2w(1-w)\rho_{12}\sigma_1\sigma_2\\
# &\text{s.t.} \qquad 0\leq w\leq1,
# \end{align*}
# 1. Los anteriores son problemas de **programación cuadrática** (función convexa sobre dominio convexo: mínimo absoluto asegurado).
# 2. Existen diversos algoritmos para problemas de programación cuadrática. Por ejemplo, en la librería cvxopt. Más adelante la instalaremos y la usaremos.
# 3. En scipy.optimize no hay un algoritmo dedicado a la solución de este tipo de problemas de optimización. Sin embargo, la función mínimize nos permite resolver problemas de optimización en general (es un poco limitada, pero nos sirve por ahora).
# ### 2.1. Antes de resolver el problema con la función minimize: resolverlo a mano en el tablero.
# Calcular w_minvar y mostrar...
w_minvar = (s2**2-s1*s2*r12)/(s1**2+s2**2-2*s1*s2*r12)
w_minvar
# **Conclusiones:**
# - El portafolio de mínima varianza se obtiene al invertir el $75.39\%$ de la riqueza en el mercado de EU.
# ### 2.2. Ahora sí, con la función scipy.optimize.minimize
# Importar el módulo optimize
import scipy.optimize as opt
# Función minimize
help(opt.minimize)
# Función objetivo
def var(w, s1, s2, r12):
return (w*s1)**2+((1-w)*s2)**2+2*w*(1-w)*s1*s2*r12
# Dato inicial
w0 = 0
# Volatilidades individuales
s1 = annual_ret_summ.loc['Volatilidad', 'EU']
s2 = annual_ret_summ.loc['Volatilidad', 'Japon']
# Correlacion
r12 = corr.loc['Japon', 'EU']
# Cota de w
bnd = ((0, 1),)
# Solución
minvar2 = opt.minimize(var, w0, args=(s1, s2, r12), bounds=bnd)
minvar2
# Peso del portafolio de minima varianza
w_minvar2 = minvar2.x
E_minvar2 = w_minvar2*E1+(1-w_minvar2)*E2
s_minvar2 = minvar2.fun**0.5
# Graficar el portafolio de varianza mínima
# sobre el mismo gráfico realizado anteriormente
plt.figure(figsize=(12,8))
plt.plot(portafolios['sp'], portafolios['EP'], label='Portafolios')
plt.plot(s1, E1, 'b*', ms=10, label='EU')
plt.plot(s2, E2, 'r*', ms=10, label='Japon')
plt.plot(s_minvar2, E_minvar2, 'oy', ms=10, label='Port. Min. Var.')
plt.grid()
plt.legend(loc='best')
plt.xlabel('Volatilidad ($\sigma$)')
plt.ylabel('Rendimiento esperado ($E[r]$)')
# ___
# ## 3. Ahora, para tres activos, obtengamos la frontera de mínima varianza
import scipy.optimize as opt
## Construcción de parámetros
## Activo 1: EU, Activo 2: Japon, Activo 3: RU
# 1. Sigma: matriz de varianza-covarianza
s1 = annual_ret_summ['EU']['Volatilidad']
s2 = annual_ret_summ['Japon']['Volatilidad']
s3 = annual_ret_summ['RU']['Volatilidad']
s12 = corr['EU']['Japon']*s1*s2
s13 = corr['EU']['RU']*s1*s3
s23 = corr['Japon']['RU']*s2*s3
Sigma = np.array([[s1**2, s12, s13],
[s12, s2**2, s23],
[s13, s23, s3**2]])
# 2. Eind: rendimientos esperados activos individuales
E1 = annual_ret_summ['EU']['Media']
E2 = annual_ret_summ['Japon']['Media']
E3 = annual_ret_summ['RU']['Media']
Eind = np.array([E1, E2, E3])
# 3. Ereq: rendimientos requeridos para el portafolio
# Número de portafolios
N = 100
Ereq = np.linspace(Eind.min(), Eind.max(), N)
def varianza(w, Sigma):
return w.dot(Sigma).dot(w)
def rendimiento_req(w, Eind, Ereq):
return Eind.dot(w)-Ereq
# Dato inicial
w0 = np.zeros(3,)
# Cotas de las variables
bnds = ((0,None), (0,None), (0,None))
# +
def f(x):
return x**2
g = lambda x: x**2
# -
f(210), g(210)
# +
# DataFrame de portafolios de la frontera
portfolios3 = pd.DataFrame(index=range(N), columns=['w1', 'w2', 'w3', 'Ret', 'Vol'])
# Construcción de los N portafolios de la frontera
for i in range(N):
# Restricciones
cons = ({'type': 'eq', 'fun': rendimiento_req, 'args': (Eind,Ereq[i])},
{'type': 'eq', 'fun': lambda w: np.sum(w)-1})
# Portafolio de mínima varianza para nivel de rendimiento esperado Ereq[i]
min_var = opt.minimize(varianza, w0, args=(Sigma,), bounds=bnds, constraints=cons)
# Pesos, rendimientos y volatilidades de los portafolio
portfolios3.loc[i,['w1','w2','w3']] = min_var.x
portfolios3['Ret'][i] = Eind.dot(min_var.x)
portfolios3['Vol'][i] = np.sqrt(varianza(min_var.x, Sigma))
# -
# Portafolios de la frontera
portfolios3
# Portafolio de mínima varianza
cons = ({'type': 'eq', 'fun': lambda w: np.sum(w)-1},)
min_var3 = opt.minimize(varianza, w0, args=(Sigma,), bounds=bnds, constraints=cons)
min_var3
w_minvar3 = min_var3.x
E_minvar3 = Eind.dot(w_minvar3)
s_minvar3 = np.sqrt(varianza(w_minvar3, Sigma))
# Graficamos junto a los portafolios de solo EU y Japón
plt.figure(figsize=(12,8))
plt.plot(portafolios['sp'], portafolios['EP'], label='Portafolios 2 act')
plt.plot(portfolios3.Vol, portfolios3.Ret, 'k-', lw=2, label='Portafolios 3 act')
plt.plot(s1, E1, 'b*', ms=10, label='EU')
plt.plot(s2, E2, 'r*', ms=10, label='Japon')
plt.plot(s3, E3, 'c*', ms=10, label='RU')
plt.plot(s_minvar2, E_minvar2, 'oy', ms=10, label='Port. Min. Var. 2')
plt.plot(s_minvar3, E_minvar3, 'om', ms=10, label='Port. Min. Var. 3')
plt.grid()
plt.legend(loc='best')
plt.xlabel('Volatilidad ($\sigma$)')
plt.ylabel('Rendimiento esperado ($E[r]$)')
plt.axis([0.14, 0.16, 0.135, 0.14])
# **Conclusión.** Mayor diversificación.
# ___
# ## 4. Comentarios acerca de la Teoría Moderna de Portafolios.
#
# - Todo lo anterior es un abrebocas de lo que llamamos análisis de media-varianza, y que es la base de la teoría moderna de portafolios.
# - El análisis de media-varianza transformó el mundo de las inversiones cuando fué presentada por primera vez.
# - Claro, tiene ciertas limitaciones, pero se mantiene como una de las ideas principales en la selección óptima de portafolios.
# ### Historia.
#
# 1. Fue presentada por primera vez por <NAME> en 1950. Acá su [artículo](https://www.google.com/url?sa=t&rct=j&q=&esrc=s&source=web&cd=1&ved=2ahUKEwjd0cOTx8XdAhUVo4MKHcLoBhcQFjAAegQICBAC&url=https%3A%2F%2Fwww.math.ust.hk%2F~maykwok%2Fcourses%2Fma362%2F07F%2Fmarkowitz_JF.pdf&usg=AOvVaw3d29hQoNJVqXvC8zPuixYG).
# 2. Era un joven estudiante de Doctorado en la Universidad de Chicago.
# 3. Publicó su tesis doctoral en selección de portafolios en "Journal of Finance" en 1952.
# 4. Su contribución transformó por completo la forma en la que entendemos el riesgo.
# 5. Básicamente obtuvo una teoría que analiza como los inversionistas deberían escoger de manera óptima sus portafolios, en otras palabras, cómo distribuir la riqueza de manera óptima en diferentes activos.
# 6. Casi 40 años después, Markowitz ganó el Premio Nobel en economía por esta idea.
# - La suposición detrás del análisis media-varianza es que los rendimientos de los activos pueden ser caracterizados por completo por sus rendimientos esperados y volatilidad.
# - Por eso es que graficamos activos y sus combinaciones (portafolios) en el espacio de rendimiento esperado contra volatilidad.
# - El análisis media-varianza es básicamente acerca de la diversificación: la interacción de activos permite que las ganancias de unos compensen las pérdidas de otros.
# - La diversificación reduce el riesgo total mientras combinemos activos imperfectamente correlacionados.
# - En el siguiente módulo revisaremos cómo elegir portafolios óptimos como si los inversionistas sólo se preocuparan por medias y varianzas.
# - ¿Qué pasa si un inversionista también se preocupa por otros momentos (asimetría, curtosis...)?
# - La belleza del análisis media-varianza es que cuando combinamos activos correlacionados imperfectamente, las varianzas siempre decrecen (no sabemos que pasa con otras medidas de riesgo).
# - Si a un inversionista le preocupan otras medidas de riesgo, el análisis media-varianza no es el camino.
# - Además, si eres una persona que le gusta el riesgo: quieres encontrar la próxima compañía top que apenas va arrancando (como Google en los 2000) e invertir todo en ella para generar ganancias extraordinarias; entonces la diversificación no es tampoco el camino.
# - La diversificación, por definición, elimina el riesgo idiosincrático (de cada compañía), y por tanto elimina estos rendimientos altísimos que brindaría un portafolio altamente concentrado.
# # Anuncios parroquiales
#
# ## 1. Tarea 4 - segunda entrega para el miércoles 6 de Marzo.
# ## 2. Examen módulos 1 y 2 para el martes 19 de Marzo.
# ## 3. Recordar quiz la próxima clase.
# ## 4. Revisar archivo de la Tarea 5. Para el viernes 8 de Marzo.
# ## 5. La próxima clase es de repaso, sin embargo, el repaso no lo hago yo, lo hacen ustedes. Estaremos resolviendo todo tipo de dudas que ustedes planteen acerca de lo visto hasta ahora. Si no hay dudas, dedicarán el tiempo de la clase a tareas del curso.
# ## 7. Fin Módulo 2: revisar Clase0 para ver objetivos.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Modulo2/Clase10_DiversificacionFuentesRiesgoII.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import pickle
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import matplotlib.pyplot as plt
from tensorflow.keras.models import Sequential
from pandas.api.types import CategoricalDtype
import seaborn as sns
# %load_ext tensorboard
import datetime
from tensorflow.keras.utils import to_categorical
filename = "../../Datasets/WBC/data.csv"
df = pd.read_csv(filename,encoding = 'utf-8')
df.head()
# -
#df.columns.tolist()
df.rename(columns = {'diagnosis':'class_val'},inplace = True)
df.head(1)
df['class_val'].value_counts()
# +
# Create category types.
#class_type = CategoricalDtype(['Type1','Type2','Type3'], ordered=True)
# Convert all categorical values to category type.
#df.class_val = df.class_val.astype(class_type)
# -
# Convert categories into integers for each column.
df.class_val=df.class_val.replace({'M':0, 'B':1})
plt.figure(figsize=(100,60))
sns.set(font_scale=1.2)
sns.heatmap(df.corr(),annot=True, cmap='rainbow',linewidth=0.5)
plt.title('Correlation matrix');
# +
# Choose attribute columns and class column.
X=df[df.columns[1:-1]]
y=df['class_val']
y = np.array(y)
y = to_categorical(y)
df.head()
# +
initializer0 = keras.initializers.RandomUniform(minval = 0, maxval =0.005)
initializer1 = keras.initializers.RandomUniform(minval = 0, maxval =0.005)
initializer2 = keras.initializers.RandomUniform(minval = -2, maxval =-1)
class Diffact(keras.layers.Layer):
def __init__(self):
super(Diffact, self).__init__()
self.k0 = self.add_weight(name='k0', shape = (), initializer=initializer0, trainable=True)
self.k1 = self.add_weight(name='k1', shape = (), initializer=initializer1, trainable=True)
self.k2 = self.add_weight(name='k2', shape = (), initializer=initializer2, trainable=True)
def call(self, inputs):
return self.k0 + tf.multiply(inputs, self.k1) + tf.multiply(tf.multiply(inputs,inputs), self.k2)
from keras import backend as K
class Mish(keras.layers.Layer):
'''
Mish Activation Function.
.. math::
mish(x) = x * tanh(softplus(x)) = x * tanh(ln(1 + e^{x}))
Shape:
- Input: Arbitrary. Use the keyword argument `input_shape`
(tuple of integers, does not include the samples axis)
when using this layer as the first layer in a model.
- Output: Same shape as the input.
Examples:
>>> X_input = Input(input_shape)
>>> X = Mish()(X_input)
'''
def __init__(self, **kwargs):
super(Mish, self).__init__(**kwargs)
self.supports_masking = True
def call(self, inputs):
return inputs * K.tanh(K.softplus(inputs))
def get_config(self):
base_config = super(Mish, self).get_config()
return dict(list(base_config.items()) + list(config.items()))
def compute_output_shape(self, input_shape):
return input_shape
# -
from sklearn.model_selection import StratifiedKFold, cross_val_score, KFold, train_test_split
from sklearn.preprocessing import LabelEncoder, StandardScaler, MinMaxScaler
# split data into train, test
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.1,random_state=69, shuffle=True)
#kipping y since value already 1 or 0
# encoder = LabelEncoder()
# encoder.fit(Y)
# encoded_Y = encoder.transform(Y)
'''
# normalize data
scaler = MinMaxScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_train = pd.DataFrame(X_train_scaled)
X_test_scaled = scaler.fit_transform(X_test)
X_test = pd.DataFrame(X_test_scaled)
'''
print (X_train.shape, y_train.shape)
print (X_test.shape, y_test.shape)
print (df.columns)
# create model with fully connected layers with dropout regulation
model = Sequential()
model.add(layers.Dense(25, input_dim=30))
diffact=Diffact()
model.add(diffact)
model.add(layers.Dropout(0.1))
model.add(layers.Dense(2, activation = 'softmax'))
model.summary()
# +
batch_size = 5
epochs = 100
model.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
log_dir = "cancerlogs/smallk2/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
model.fit(X_train, y_train, batch_size=batch_size, epochs=epochs, validation_split=0.1 ,callbacks=[tensorboard_callback], verbose=1)
# -
score = model.evaluate(X_test, y_test, verbose=0)
print("Test loss:", score[0])
print("Test accuracy:", score[1])
print("AF coefficients (weights) {}".format(diffact.get_weights()))
# %tensorboard --logdir cancerlogs/smallk2 --port=6034
df.describe()
| Models/WBC/Diffact-keras-noncv-breastcancer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Projeto Final
#
# ## Reconhecimento de convicção através da fala para suporte ao ensino
#
# Disciplina: Computação Afetiva - IA369Y
#
# Prof. <NAME>
#
#
# Alunos:
#
# <NAME>
# <NAME>
# <NAME>
# %matplotlib inline
import pandas as pd
import csv
import matplotlib.pyplot as plt
import numpy as np
import sklearn
from sklearn import svm
from sklearn.cross_validation import train_test_split
import glob
from hmmlearn.hmm import GaussianHMM, MultinomialHMM
datas_cer = glob.glob('features/treinamento/certeza/*.csv')
datas_inc = glob.glob('features/treinamento/incerteza/*.csv')
#datas_control = glob.glob('features_projeto_IA369Y/controles/*.csv')
#df = pd.read_csv('features/treinamento/certeza/I1questao1.csv', sep=';')
#df1 = df.loc[:, 'alphaRatio_sma3': 'F3amplitudeLogRelF0_sma3nz'].values
#d = np.array(df1)
#feat_list = []
#print (d.shape)
#feat_list.append(d)
#lenghts = []
#for i in range(len(feat_list)):
# lenghts.append(len(feat_list[i]))
#print (df1)
#f = np.vstack(feat_list)
#print (len(feat_list))
def select_features(datas):
feat_list = []
for file in datas:
df = pd.read_csv(file, sep=';')
#df1 = df[df.columns[2:4]]
df1 = df.loc[:, 'alphaRatio_sma3': 'F3amplitudeLogRelF0_sma3nz'].values
d = np.array(df1)
#print (d.shape)
feat_list.append(d)
lenghts = []
for i in range(len(feat_list)):
lenghts.append(len(feat_list[i]))
#print (df1)
f = np.vstack(feat_list)
return f, lenghts, feat_list
f_cer, len_cer, list_fcer = select_features(datas_cer)
f_inc, len_inc, list_finc = select_features(datas_inc)
#f_control, len_control = select_features(datas_control)
# +
#features = np.hstack(f_cer, f_inc)
#print (type(list_fcer[1]))
#print (len(list_fcer))
#print (len_cer)
#print (f_cer.shape)
#print (type(list_finc[1]))
#print (len(list_finc))
#print (len_inc)
#print (f_inc.shape)
# -
# ## Treinamento do Classificador HMM
# +
import warnings
warnings.filterwarnings('ignore')
model1 = GaussianHMM(n_components=15, covariance_type='diag', n_iter=50)
model1.fit(f_cer, len_cer)
model1.monitor_
# -
model1.monitor_.converged
model2 = GaussianHMM(n_components=15, covariance_type='diag', n_iter=50)
model2.fit(f_inc, len_inc)
model2.monitor_
model2.monitor_.converged
# ## Salvar o modelo treinado
#Salvar o modelo treinado.
from sklearn.externals import joblib
joblib.dump(model1, "Model1_certeza.pkl")
joblib.dump(model2, "Model2_incerteza.pkl")
#Exemplo do Carregamento do modelo.
model1 = joblib.load("Model1_certeza.pkl")
model2 = joblib.load("Model2_incerteza.pkl")
# Para testar o modelo, utiliza:
#
# prob1 = model1.score(list_vcer[val])
# prob2 = model2.score(list_vcer[val])
# if prob1 > prob2:
# print (['certeza'])
# else:
# print (['incerteza'])
# ## Teste do Classificador para cada um dos áudios
datas_v_cer = glob.glob('features/validacao/certeza/*.csv')
datas_v_inc = glob.glob('features/validacao/incerteza/*.csv')
# +
v_cer, ven_cer, list_vcer = select_features(datas_v_cer)
v_inc, ven_inc, list_vinc = select_features(datas_v_inc)
#print (list_vcer[0].shape)
#print (len(list_vcer))
#print (ven_cer)
#print (v_cer.shape)
# +
# Verificar a quantidade de acerto
# Para os áudios classificados como CERTEZA
def taxa_cer(list_vcer): #taxa de erro e acerto em porcentagem
erros1 = []
acertos1 = []
for val in range(len(list_vcer)):
prob1 = model1.score(list_vcer[val])
prob2 = model2.score(list_vcer[val])
if prob1 > prob2:
acertos1.append(['certeza'])
else:
erros1.append(['incerteza'])
taxa_ac1 = (len(acertos1))/len(list_vcer)*100
taxa_er1 = (len(erros1))/len(list_vcer)*100
print ('taxa de acerto:', taxa_ac1, 'taxa de erro:', taxa_er1)
return taxa_ac1, taxa_er1
# Para os áudios classificados como INCERTEZA
def taxa_inc(list_vinc): #taxa de erro e acerto em porcentagem
erros2 = []
acertos2 = []
for val2 in range(len(list_vinc)):
prob3 = model1.score(list_vinc[val2])
prob4 = model2.score(list_vinc[val2])
if prob3 > prob4:
erros2.append(['certeza'])
else:
acertos2.append(['incerteza'])
taxa_ac2 = (len(acertos2))/len(list_vinc)*100
taxa_er2 = (len(erros2))/len(list_vinc)*100
print ('taxa de acerto:', taxa_ac2, 'taxa de erro:', taxa_er2)
return taxa_ac2, taxa_er2
print ('Validação para certeza')
ac1, er1 = taxa_cer(list_vcer)
print ('Validação para incerteza')
ac2, er2 = taxa_inc(list_vinc)
# -
# ### Audio de teste
def select_feature_test(path=''):
df = pd.read_csv(path, sep=';')
df1 = df.loc[:, 'alphaRatio_sma3': 'F3amplitudeLogRelF0_sma3nz'].values
d = np.array(df1)
#print (d.shape)
lenghts = []
for i in range(len(d)):
lenghts.append(len(d[i]))
#print (df1)
f = np.vstack(d)
return f, lenghts
# +
path_t = 'features_projeto_IA369Y/certeza/I3questao13.csv'
#path_t2 = 'features_projeto_IA369Y/incerteza/I21questao11.csv'
f_test, len_test = select_feature_test(path_t)
# Teste das perguntas de controle
prob1 = model1.score(f_test)
prob2 = model2.score(f_test)
if prob1 > prob2:
print ('certeza')
else:
print ('incerteza')
# -
# ## Referências
#
# - https://hmmlearn.readthedocs.io/en/latest/index.html
#
# - https://github.com/tiagoft/course_audio
#
# - https://github.com/naxingyu/opensmile
| scripts/Projeto_IA369_1Class_All.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# executing this cell will install all required julia packages
import Pkg
Pkg.activate(".")
Pkg.instantiate()
Pkg.precompile()
do_savefig = false
figures_dir = "insert/path/where/figures/should/be/saved/"
;
using LaTeXStrings
using GaussianProcesses
using Distributions
using LinearAlgebra
using Distances
using LibGEOS
using GeoInterface
import CSV
import GeoRDD
import PyPlot; plt=PyPlot
plt.rc("figure", dpi=200.0)
# plt.rc("figure", figsize=(6,4))
plt.rc("figure", autolayout=true)
plt.rc("savefig", dpi=200.0)
plt.rc("text", usetex=true)
plt.rc("font", family="serif")
plt.rc("font", serif="Palatino")
plt.rc("pdf", fonttype=42)
cbbPalette = ["#E69F00", "#56B4E9", "#009E73", "#F0E442", "#0072B2", "#D55E00", "#CC79A7"]
;
function read_matrix(filename)
df = CSV.read(filename; header=false)
mat = convert(Matrix{Float64}, df)
return mat
end
X_LA = read_matrix("Mississippi_data/X_LA.csv")
X_MS = read_matrix("Mississippi_data/X_MS.csv")
border_XY = read_matrix("Mississippi_data/border.csv")
border_geom = LibGEOS.LineString([border_XY[i,:] for i in 1:size(border_XY,1)])
sentinels = GeoRDD.sentinels(border_geom, 200)
;
n_MS = size(X_MS, 2)
n_LA = size(X_LA, 2)
# +
Δ = 50e3
AΔ = LibGEOS.buffer(border_geom, Δ)
AΔ_coords = GeoInterface.coordinates(AΔ)[1]
AΔ_hcat = hcat(AΔ_coords...)
;
# -
_xlim = (-1.03e7, -1.005e7)
_ylim = (3.55e6, 3.72e6)
# +
function arrow_distance(xy_start, xy_end, text;
color="black",
offset=(5, 5),
arrow_size=5.0,
shrink=0.0,
linewidth=2.0)
plt.annotate(
"",
xy=tuple(xy_start...),
xycoords="data",
xytext=tuple(xy_end...),
textcoords="data",
# shrink=1.0,
arrowprops=Dict(:arrowstyle => "<->",
:linewidth => linewidth,
:color => color,
:shrinkA => shrink,
:shrinkB => shrink,
:mutation_scale => arrow_size,
# "frac" => 0.05,
# "shrink" => 0.5,
),
zorder=10)
text_xy = tuple(((xy_start.+xy_end)/2)...)
plt.annotate(
text,
xy=text_xy,
xycoords="data",
xytext=offset,
textcoords="offset points",
arrowprops=Dict(
"color" => color,
"arrowstyle" => "-",
"linewidth" => 0.0,
),
color=color,
zorder=10,
)
end
# plot border
function plot_common(;arrow_delta=true, label_border=true, label_vicinity=true, label_obs=true)
iB_inxlim = _xlim[1] .< border_XY[:,1] .< _xlim[2]
iB_inylim = _ylim[1] .< border_XY[:,2] .< _ylim[2]
iB_inxy = iB_inxlim .& iB_inylim
border_inxy = border_XY[iB_inxy, :]
plt.plot(border_XY[:,1], border_XY[:,2], "-", color="#555555")
if label_border
iannotate = 50
plt.annotate(L"$\mathcal{B}$",
(border_inxy[iannotate, 1],
border_inxy[iannotate, 2]),
xycoords="data",
xytext=(5, 5),
textcoords="offset points",
color="#555555",
)
end
if label_obs
label="Observation (MS)"
else
label=""
end
# plot county centroids
plt.scatter(
X_MS[1,:],
X_MS[2,:],
marker="o",
color="none",
edgecolor=cbbPalette[2],
label=label,
s=20)
if label_obs
label="Observation (LA)"
else
label=""
end
plt.scatter(
X_LA[1,:],
X_LA[2,:],
marker="o",
color="none",
edgecolor=cbbPalette[3],
label=label,
s=20)
ax = plt.gca()
# BUFFER ZONE
if label_vicinity
label = L"Border vicinity $\mathcal{A}_{\Delta}$"
else
label = ""
end
AΔ_patch = plt.matplotlib.patches.Polygon(AΔ_coords,
true, # closed
color=cbbPalette[6],
alpha=0.2,
label=label,
zorder=-10)
ax.add_patch(AΔ_patch)
if arrow_delta
# Draw an arrow indicating Δ
iAΔ_inxlim = _xlim[1] .< AΔ_hcat[1,:] .< _xlim[2]
iAΔ_inylim = _ylim[1] .< AΔ_hcat[2,:] .< _ylim[2]
iAΔ_inxy = iAΔ_inxlim .& iAΔ_inylim
AΔ_inxy = AΔ_hcat[:, iAΔ_inxy]
arrow_start_xy = AΔ_inxy[:, 50]
arrow_start_point = LibGEOS.Point(arrow_start_xy...)
arrow_end_point = LibGEOS.nearestPoints(border_geom, arrow_start_point)[1]
arrow_end_xy = GeoInterface.coordinates(arrow_end_point)[1:2]
arrow_distance(
arrow_start_xy,
arrow_end_xy,
L"$\Delta$",
color=cbbPalette[6],
offset=(5, 0),
arrow_size=10,
linewidth=1.0,
)
end
plt.xlim(_xlim)
plt.ylim(_ylim)
# ax.axes.set_aspect("equal", "datalim")
end
function plot_proj_line(x, y, proj_x, proj_y, label)
plt.plot([x, proj_x], [y, proj_y],
color="black", alpha=0.05, linewidth=1,
label=label,
zorder=-10)
end
;
# +
function plot_finite_proj(X, border_geom; labeled=false, labeled_sent=false, ilabel=1)
n_X = size(X, 2)
for i in 1:n_X
# obtain coordinates for treatment point
x, y = X[:,i]
point = LibGEOS.Point(x,y)
# projection onto border (as distance along border)
proj_point = nearestPoints(border_geom,
point)[1]
# skip if far away from border
if LibGEOS.distance(point, proj_point) > Δ
continue
end
if !labeled
label="Projection line"
labeled=true
else
label=""
end
proj_x, proj_y = GeoInterface.coordinates(proj_point)
plot_proj_line(x, y, proj_x, proj_y, label)
if !labeled_sent
label="Projected sentinel"
labeled_sent=true
else
label=""
end
plt.scatter(proj_x, proj_y,
marker="D",
color="#222222",
s=5,
label=label,
zorder=5)
if i==ilabel
darkgreen = "#053021"
plt.annotate(
L"$\mathbf{s}_i$",
xy=(x, y),
xycoords="data",
xytext=(5,15),
horizontalalignment="left",
verticalalignment="bottom",
textcoords="offset points",
arrowprops=Dict(
:arrowstyle => "-",
:linewidth => 1.0,
:connectionstyle => "arc3,rad=0.3",
:color => darkgreen,
:linestyle => "dotted",
:shrinkA => 0.0,
:shrinkB => 5.0,
),
color=darkgreen,
zorder=10,
)
text = arrow_distance(
[x, y],
[proj_x, proj_y],
L"$\mathrm{dist}_\mathcal{B}\left(\mathbf{s}_i\right)$";
color=darkgreen,
linewidth=1.0,
offset=(-13,5),
arrow_size=8,
)
angle = atand((y-proj_y)/(x-proj_x))
text.set_rotation(angle)
plt.annotate(
L"$\mathrm{proj}_\mathcal{B}\left(\mathbf{s}_i\right)$",
xy=(proj_x, proj_y),
xycoords="data",
xytext=(-15,-15),
textcoords="offset points",
horizontalalignment="right",
verticalalignment="top",
arrowprops=Dict(
:arrowstyle => "-",
:linewidth => 1.0,
:connectionstyle => "arc3,rad=0.3",
:color => darkgreen,
:linestyle => "dotted",
:shrinkA => 0.0,
:shrinkB => 5.0,
),
color=darkgreen,
zorder=10,
)
end
end
end
function plot_finite_pop()
# plot projections
labeled=false
labeled_sent=false
plot_finite_proj(X_MS, border_geom)
plot_finite_proj(X_LA, border_geom; labeled=true, labeled_sent=true, ilabel=58)
end
;
# -
function plot_grid()
gridspace = 1e4
X_grid = _xlim[1]:gridspace:_xlim[2]
Y_grid = _ylim[1]:gridspace:_ylim[2]
arrow_y = 0.7*Y_grid[13] + 0.3*Y_grid[14]
arrow_distance(
[X_grid[20], arrow_y],
[X_grid[21], arrow_y],
L"$\nu$",
color=cbbPalette[5],
offset=(-1, 4),
shrink=0.0,
linewidth=1.0,
)
projected_weights = Dict{LibGEOS.Point, Float64}()
labeled = false
for x in X_grid, y in Y_grid
# convert to a point obejct
point = LibGEOS.Point(x, y)
# project the point onto the border
proj_point = nearestPoints(border_geom,
point)[1]
# Only keep points that are both within `maxdist` of
# the border, and are in the convex hull of the data.
if LibGEOS.distance(point, proj_point) > Δ
continue
end
proj_x, proj_y = GeoInterface.coordinates(proj_point)
if !labeled
label=""
grid_label=L"grid $G^\nu$"
labeled=true
else
label=""
grid_label=""
end
plt.scatter(
x,
y,
s=6,
marker="x",
color=cbbPalette[5],
label=grid_label)
plot_proj_line(x, y, proj_x, proj_y, label)
# obtain the coordinates of the projected point
proj_coords = [proj_x, proj_y]
found_nearby = false
for p in keys(projected_weights)
if LibGEOS.distance(p, proj_point) < 2000.0
projected_weights[p] += 1.0
found_nearby = true
break
end
end
if !found_nearby
# initialize
projected_weights[proj_point] = 1.0
end
end
unique_projected_points = [GeoInterface.coordinates(p) for p in keys(projected_weights)]
Xb_projected = [[p[1] for p in unique_projected_points]';
[p[2] for p in unique_projected_points]']
# And the counts as the weights.
weights = collect(values(projected_weights))
plt.scatter(
Xb_projected[1,:], Xb_projected[2, :],
s=5 .* weights,
marker="D",
color="black",
linewidths=0.0,
alpha=1.0,
zorder=5,
label="",
)
end
;
function title_in_axis(s, leftright)
if leftright == :left
plt.text(0.05, 0.95, s,
horizontalalignment="left",
verticalalignment="top",
transform = plt.gca().transAxes)
elseif leftright == :right
plt.text(0.95, 0.95, s,
horizontalalignment="right",
verticalalignment="top",
transform = plt.gca().transAxes)
else
throw("left or right?")
end
end
;
# +
plt.gcf().set_size_inches(10, 5)
plt.subplot(1,2,1)
plot_common()
plot_finite_pop()
leg = plt.legend(loc="lower left", fontsize="small")
leg.get_frame().set_alpha(1.0)
plt.xticks([]); plt.yticks([])
plt.gca().set(aspect="equal", adjustable="box")
title_in_axis("(a)", :left)
plt.subplot(1,2,2)
plot_common(; arrow_delta=false, label_border=false, label_obs=false, label_vicinity=false)
plot_grid()
leg = plt.legend(loc="lower left", fontsize="small")
leg.get_frame().set_alpha(1.0)
plt.xticks([]); plt.yticks([])
plt.gca().set(aspect="equal", adjustable="box")
title_in_axis("(b)", :left)
if do_savefig
plt.savefig(joinpath(figures_dir, "mississippi_projection_methods.png"), bbox_inches="tight")
plt.savefig(joinpath(figures_dir, "mississippi_projection_methods.pdf"), bbox_inches="tight")
end
;
| Mississippi_projection_illustration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise 19: Lambda Expression for Sorting
# 1. define a list of tuples
capitals = [("USA", "Washington"), ("India", "Delhi"), ("France", "Paris"), ("UK", "London")]
capitals
# 2. Sort this list by the name of the capitals of each country, using a simple lambda expression.
capitals.sort(key=lambda item: item[1])
capitals
| Chapter02/.ipynb_checkpoints/Exercise 19-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from pandas import Series,DataFrame
this= Series([6,5,4,9,2])
print(this)
# + active=""
# this.values
# -
this.values
this.index
# +
## create a series of us cities with their population and index is set as the city for series , the object takles the
# indexes as defined with their corresponding column values i.e population
citypopulation_2016=Series([8537673,3976322,2704958,2303454,1615017],
index=['NYC','Los Angeles','Chicago','Houston','Phoenix'])
citypopulation_2016
# +
#find the population less than 2.5 million
citypopulation_2016[citypopulation_2016<2500000]
# -
'Chicago' in citypopulation_2016
##Create a listof index
cities=['NYC','Los Angeles','Chicago','Houston','Phoenix','philadelhia']
ser= Series(citypopulation_2016,index=cities)
ser
#check null values
ser.isnull()
# +
# adding two series
ser1= ser+citypopulation_2016
ser1
# + active=""
#
#
#
#
# -
ser1.name= "us populate cities"
ser1
ser1.index.name="cities"
ser1
### dataframe
df=pd.read_csv("C:\\Users\\p#an!\\Desktop\\datascience\\DATA\\CSV FILE.csv")
df.head() ### This will give the first five rows of data
df.tail () # this will give the last 5 rows of data
df.info # this will give the information
df['Order ID'].head()
df['Order ID'].tail()
DataFrame(df,columns=['last_name','First_name','AGE'])
df.ix[4]
df= pd.read_csv("C:\\Users\\p#an!\\Desktop\\datascience\\DATA\\employees salary.csv")
df
df.head()
df.tail()
df.info()
df['Job Family'].head()
##adding columns
DataFrame(df,columns=['Job Family','Age'])
df['double_salary'] = df['Salaries']*2
df.head(5)
df['taxes']= 'base tax'
df
tax = Series(['Federal Tax','State Tax'],index=[0,6])
df['taxes'] = tax
df
# +
del df['taxes']
# +
###DaTA FRAME FROM DICTIONARY
MyFruits = {'fruits':['orange','Apple','Grapes','PineApple','Papaya'],
'Price':[100,200,80,50,150]}
dffruits= DataFrame(MyFruits)
dffruits
# -
df= Series([6,4,5,9,7],['red','violet','Green','Blue','Orange'])
df
df.reindex(['red','violet','Green','Blue','Orange','Pink'])
# +
df.reindex(['red','violet','Green','Blue','Orange','Pink'],fill_value=10)
# -
#Create a Dataframe with random integers and index value as defined below and columns also as shown
df=DataFrame(np.random.random_integers(25,size=25).reshape(5,5),index=['A','B','C','D','E'],
columns=['col1','col2','col3','col4','col5'])
df
# +
##Suming up columns and rows
df.sum(axis=1) #Sum along the rows by using axis= 1
# -
df.sum(axis=0) ## sum along the columns by using axis=0
df.describe()
# +
#Find the correlation between each of the variables of the dataframe using ‘corr’ function
corr= df.corr()
corr
# -
#Plot the Correlation heatmap matrix using Seaborn
import seaborn as sns
# %matplotlib inline
sns.heatmap(corr)
#Missing values
df= DataFrame([[5,4,3,2,7],[6,3,np.NaN,np.NaN,9],[np.NaN,3,4,2,np.NaN]])
df
x= df.dropna(axis=1) #dropping all the columns where null values is present
x
x=df.dropna(thresh=3)
x
##Fillna fills null with the values
df.fillna('5')
df=DataFrame(np.random.random_integers(25,size=25).reshape(5,5),index=['A','B','C','D','E'],
columns=['col1','col2','col3','col4','col5'])
df
df.drop('B')
# +
#selecting only two columns
df[['col1','col2']]
# -
df.ix['D'] #Selecting a index
df[df['col1']>10] ## selecting on a condition
#selecting values
df= Series([6,4,3,2,5],['A','B','C','D','E'])
df
df['D']
df[0:4]
#Sorting
df=Series([6,4,2,1],index=['P','R','S','Q'])
df
df.sort_index()
#Order by values
df.sort_values()
df.rank()
#Multiindexing
Sr = Series([5,4,3,7,9,6],index=[[1,1,2,2,3,3],['a','b',
'a','b','a','b']])
Sr
Sr.index
Sr[3]
#Get Values by Second Index
Sr[:,'b']
Sr.unstack()
| Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from czifile import CziFile
import numpy as np
pwd
datadir = '../data/20180108_htl_glc_sc11_mmzm.czi'
# The syntax that follows below was taken from this [tutorial] posted on *Scientific Programming with Python*.
#
# [tutorial]: (http://schryer.github.io/python_course_material/python/python_10.html)
with CziFile(datadir) as czi:
raw_im = czi.asarray()
type(raw_im),raw_im.shape
# The output object is a `numpy` array with 8 dimensions. Based on some notes in the [tutorial], we can guess what each dimension most likely contains.
#
# `(?, roi, channel, time, z, x, y, ?)`
# For this particular data, we collected a timelapse movie with 2 channels (GFP, brightfield). For this analysis, we only need the brightfield channel so we will index to select just the data that we want.
data = raw_im[:,:,1,:,:,:,:,:]
data.shape
# Our new array contains 156 timepoints, 1 channel, and xy dimensions of 1024x1024. We would like to reduce this array to the minimum number of dimensions (3), which we can do with the `np.squeeze` function.
data = np.squeeze(data)
data.shape
# Now we have the minimal number of dimensions in our array and we are ready to save the data in whatever format we prefer.
# # Test utility function in gbeflow
from imp import reload
import gbeflow
reload(gbeflow)
# The `gbeflow.CziImport` function reads in the file specified in the input and saves the squeezed array in the attribute `data`.
czi = gbeflow.CziImport(datadir)
czi.data.shape
# %load_ext watermark
# %watermark -v -p czifile
| notebooks/20181030-czifile_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## RSS
# You can find RSS feeds on many different sites. [Library of Congress](https://www.loc.gov/rss/) has a lot. Most blogs and news web sites have them, for example [Tech Crunch](https://techcrunch.com/rssfeeds/), [New York Times](http://www.nytimes.com/services/xml/rss/index.html), and [NPR](https://help.npr.org/customer/portal/articles/2094175-where-can-i-find-npr-rss-feeds-). The [DC Public Library](http://www.dclibrary.org/) even gives you an RSS feed of your [catalog searches](https://catalog.dclibrary.org/client/rss/hitlist/dcpl/qu=python). iTunes delivers podcasts by [aggregating RSS feeds](http://itunespartner.apple.com/en/podcasts/faq) from content creators.
#
# Today we are going to take a look at the [Netflix Top 100 DVDs](https://dvd.netflix.com/RSSFeeds). We will use the Python package [FeedParser](https://pypi.python.org/pypi/feedparser) to work with the RSS feed. FeedParser will allow us to deconstruct the data in the feed.
import feedparser
import pandas as pd
RSS_URL = "https://usa.newonnetflix.info/feed"#"http://dvd.netflix.com/Top100RSS"
feed = feedparser.parse(RSS_URL)
type(feed)
# "parse" is the primary function in FeedParser. The returned object is dictionary like and can be handled similarly to a dictionary. For example, we can look at the keys it contains and what type of items those keys are.
feed.keys()
type(feed.bozo)
type(feed.feed)
# We will look at some, but not all, of the data stored in the feed. For more information about the keys, see the [documentation](http://pythonhosted.org/feedparser/).
#
# We can use the version to check which type of feed we have.
feed.version
# Bozo is an interesing key to know about if you are going to parse a RSS feed in code. FeedParser sets the bozo bit when it detects a feed is not well-formed. (FeedParser will still parse the feed if it is not well-formed.) You can use the bozo bit to create error handling or just print a simple warning.
if feed.bozo == 0:
print("Well done, you have a well-formed feed!")
else:
print("Potential trouble ahead.")
# We can look at some of the feed elements through the feed attribute.
feed.feed.keys()
print(feed.feed.title)
print(feed.feed.link)
print(feed.feed.description)
# The [reference section](http://pythonhosted.org/feedparser/reference.html) of the feedparser documenation shows us all the inforamtion thatcan be in a feed. [Annotated Examples](http://pythonhosted.org/feedparser/annotated-examples.html) are also provided. But note the caution provided-
#
# "Caution: Even though many of these elements are required according to the specification, real-world feeds may be missing any element. If an element is not present in the feed, it will not be present in the parsed results. You should not rely on any particular element being present."
#
# For example, our feed is RSS 2.0. One of the elements available in this version is the published date.
feed.feed.published
# We can see from our error, our feed is not using 'published'.
#
# As with [standard python dictionaries](https://docs.python.org/3.5/library/stdtypes.html#dict), we can use the "get" method to see if a key exists. This is useful if we are writing code.
feed.feed.get('published', 'N/A')
# The data we are looking for are contained in the entries. Given the feed we are working with, how many entries do you think we have?
len(feed.entries)
# The items in entries are stored as a list.
type(feed.entries)
feed.entries[0].title
i = 0
for entry in feed.entries:
print(i, feed.entries[i].title)
i += 1
# Given that information, what is something we can do with this data? Why not make it a dataframe?
df = pd.DataFrame(feed.entries)
df.head()
# Challenge: write code to create a dataframe of the top 10 movies from the Netflix Top 100 DVDs and iTunes. Check to see if your feed is well formed. Compile the name of the feed as the souce, the published date, the movie ranking in the list, the movie title, a link to the movie, and the summary. If the published date does not exist in the feed, use the current date. Save your dataframe as a csv. Here is a link to one [possible solution](./rss_challenge.py).
| rss/rss_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2
#
# Before working on this assignment please read these instructions fully. In the submission area, you will notice that you can click the link to **Preview the Grading** for each step of the assignment. This is the criteria that will be used for peer grading. Please familiarize yourself with the criteria before beginning the assignment.
#
# An NOAA dataset has been stored in the file `data/C2A2_data/BinnedCsvs_d400/fb441e62df2d58994928907a91895ec62c2c42e6cd075c2700843b89.csv`. This is the dataset to use for this assignment. Note: The data for this assignment comes from a subset of The National Centers for Environmental Information (NCEI) [Daily Global Historical Climatology Network](https://www1.ncdc.noaa.gov/pub/data/ghcn/daily/readme.txt) (GHCN-Daily). The GHCN-Daily is comprised of daily climate records from thousands of land surface stations across the globe.
#
# Each row in the assignment datafile corresponds to a single observation.
#
# The following variables are provided to you:
#
# * **id** : station identification code
# * **date** : date in YYYY-MM-DD format (e.g. 2012-01-24 = January 24, 2012)
# * **element** : indicator of element type
# * TMAX : Maximum temperature (tenths of degrees C)
# * TMIN : Minimum temperature (tenths of degrees C)
# * **value** : data value for element (tenths of degrees C)
#
# For this assignment, you must:
#
# 1. Read the documentation and familiarize yourself with the dataset, then write some python code which returns a line graph of the record high and record low temperatures by day of the year over the period 2005-2014. The area between the record high and record low temperatures for each day should be shaded.
# 2. Overlay a scatter of the 2015 data for any points (highs and lows) for which the ten year record (2005-2014) record high or record low was broken in 2015.
# 3. Watch out for leap days (i.e. February 29th), it is reasonable to remove these points from the dataset for the purpose of this visualization.
# 4. Make the visual nice! Leverage principles from the first module in this course when developing your solution. Consider issues such as legends, labels, and chart junk.
#
# The data you have been given is near **Ann Arbor, Michigan, United States**, and the stations the data comes from are shown on the map below.
# +
import matplotlib.pyplot as plt
import mplleaflet
import pandas as pd
import numpy as np
def leaflet_plot_stations(binsize, hashid):
df = pd.read_csv('data/C2A2_data/BinSize_d{}.csv'.format(binsize))
station_locations_by_hash = df[df['hash'] == hashid]
lons = station_locations_by_hash['LONGITUDE'].tolist()
lats = station_locations_by_hash['LATITUDE'].tolist()
plt.figure(figsize=(8,8))
plt.scatter(lons, lats, c='r', alpha=0.7, s=200)
return mplleaflet.display()
leaflet_plot_stations(400,'fb441e62df2d58994928907a91895ec62c2c42e6cd075c2700843b89')
# +
df = pd.read_csv('data/C2A2_data/BinnedCsvs_d400/fb441e62df2d58994928907a91895ec62c2c42e6cd075c2700843b89.csv')
datesY = pd.to_datetime(df['Date']).values.astype('datetime64[Y]')
df_0514 = df.iloc[(datesY >= np.datetime64('2005')) & (datesY <= np.datetime64('2014'))].copy()
df_0514['MD'] = pd.to_datetime(df_0514['Date']).dt.strftime('%m-%d')
df_15 = df.iloc[(datesY >= np.datetime64('2015'))].copy()
df_15['MD'] = pd.to_datetime(df_15['Date']).dt.strftime('%m-%d')
# -
df_15_grp = df_15.groupby('MD').agg({'Data_Value':[np.max, np.min]})
df_0514_grp = df_0514.groupby('MD').agg({'Data_Value':[np.max, np.min]})
df_0514_grp = df_0514_grp.drop('02-29')
import matplotlib.ticker as ticker
# %matplotlib notebook
x_bound = np.repeat(np.array([0,31,28,31,30,31,30,31,31,30,31,30,31]),2)
x_bound = np.cumsum(x_bound[1:]/2)
# +
max_2015_over = df_15_grp['Data_Value']['amax'].values > df_0514_grp['Data_Value']['amax'].values
max_diff = df_15_grp['Data_Value']['amax'].values[max_2015_over] - df_0514_grp['Data_Value']['amax'].values[max_2015_over]
min_2015_under = df_15_grp['Data_Value']['amin'].values < df_0514_grp['Data_Value']['amin'].values
min_diff = df_0514_grp['Data_Value']['amin'].values[min_2015_under] - df_15_grp['Data_Value']['amin'].values[min_2015_under]
day_2015_plot = np.concatenate((np.arange(365)[max_2015_over],np.arange(365)[min_2015_under]))
data_2015_plot = np.concatenate((df_15_grp['Data_Value']['amax'].values[max_2015_over],df_15_grp['Data_Value']['amin'].values[min_2015_under]))
data_2015_plot = data_2015_plot/10.0
abs_diff = np.concatenate((max_diff, min_diff))
abs_diff[abs_diff<10] = 10
# +
plt.figure(figsize=(8,8))
plt.plot(df_0514_grp['Data_Value']['amax'].values/10, 'red', label='Max temps in the day of the years over 2005-2014')
plt.plot(df_0514_grp['Data_Value']['amin'].values/10, 'blue', label='Mmin temps in the day of the years over 2005-2014')
# fill the area between the linear data and exponential data
plt.gca().fill_between(range(365),
df_0514_grp['Data_Value']['amin'].values/10.0, df_0514_grp['Data_Value']['amax'].values/10.0,
facecolor='yellow',
alpha=0.25)
plt.scatter(day_2015_plot,data_2015_plot,60,'c', label='extreme temperature in 2015 broke records of the day over 2005-2014')
plt.ylabel('Temperature ($^\circ$C)')
plt.title('2015 extreme temperatures near Ann Arbor, Michigan, United States')
plt.axis([0,365,-50,50])
plt.xticks(x_bound[::2], ['JAN','FEB','MAR','APR','MAY','JUN','JUL','AUG','SEP','OCT','NOV','DEC'], horizontalalignment='left')
plt.legend(loc=4, frameon=False)
# -
plt.savefig('temperature2015.png')
| Applied Plotting, Charting & Data Representation in Python/Assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter: Recurrent Neural Networks
#
#
# # Topic: Predicting engine RUL usnig LSTM
# +
# import required packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
# settings for result reproducibility
np.random.seed(1234)
PYTHONHASHSEED = 0
# +
# read data
# training
train_df = pd.read_csv('PM_train.txt', sep=" ", header=None)
train_df.drop(train_df.columns[[26, 27]], axis=1, inplace=True) # last two columns are blank
train_df.columns = ['EngineID', 'cycle', 'OPsetting1', 'OPsetting2', 'OPsetting3', 's1', 's2', 's3',
's4', 's5', 's6', 's7', 's8', 's9', 's10', 's11', 's12', 's13', 's14',
's15', 's16', 's17', 's18', 's19', 's20', 's21']
# test
test_df = pd.read_csv('PM_test.txt', sep=" ", header=None)
test_df.drop(test_df.columns[[26, 27]], axis=1, inplace=True)
test_df.columns = ['EngineID', 'cycle', 'OPsetting1', 'OPsetting2', 'OPsetting3', 's1', 's2', 's3',
's4', 's5', 's6', 's7', 's8', 's9', 's10', 's11', 's12', 's13', 's14',
's15', 's16', 's17', 's18', 's19', 's20', 's21']
# actual RUL for each engine-id in the test data
truth_df = pd.read_csv('PM_truth.txt', sep=" ", header=None)
truth_df.drop(truth_df.columns[[1]], axis=1, inplace=True) # 2nd column is blank and thus, dropped
truth_df.columns = ['finalRUL'] # assigning column name as finalRUL
truth_df['EngineID'] = truth_df.index + 1 # adding new column EngineID
# +
#%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
## generate RUL data for training and test dataset
## %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
# +
# training dataset
maxCycle_df = pd.DataFrame(train_df.groupby('EngineID')['cycle'].max()).reset_index()
maxCycle_df.columns = ['EngineID', 'maxEngineCycle'] # column maxEngineCycle stores total cycles for an engine until failure
train_df = train_df.merge(maxCycle_df, on=['EngineID'], how='left')
train_df['engineRUL'] = train_df['maxEngineCycle'] - train_df['cycle'] # column engineRUL stores engine RUL at any given cycle
train_df.drop('maxEngineCycle', axis=1, inplace=True) # maxEngineCycle is not needed anymore
train_df.head()
# +
# compute maxEngineCycle for test data using data from test_df and truth_df
maxCycle_df = pd.DataFrame(test_df.groupby('EngineID')['cycle'].max()).reset_index()
maxCycle_df.columns = ['EngineID', 'maxEngineCycle']
truth_df['maxEngineCycle'] = maxCycle_df['maxEngineCycle'] + truth_df['finalRUL']
truth_df.drop('finalRUL', axis=1, inplace=True)
# generate engineRUL for test data
test_df = test_df.merge(truth_df, on=['EngineID'], how='left')
test_df['engineRUL'] = test_df['maxEngineCycle'] - test_df['cycle']
test_df.drop('maxEngineCycle', axis=1, inplace=True)
test_df.head()
# -
#%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
## clip RUL in training data at the threshold 150
## %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
RULthreshold = 150
train_df['engineRUL'] = np.where(train_df['engineRUL'] > RULthreshold, 150, train_df['engineRUL'])
# +
#%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
## scale training and test data
## %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
# training data: create temporary dataframe with columns to be scaled
all_cols = train_df.columns # get columns names
cols_to_scale = train_df.columns.difference(['EngineID','cycle','engineRUL']) # returns all column labels except these specified
train_df_with_cols_to_scale = train_df[cols_to_scale]
# scale and rejoin with columns that were not scaled
scaler = StandardScaler()
scaled_train_df_with_cols_to_scale = pd.DataFrame(scaler.fit_transform(train_df_with_cols_to_scale), columns=cols_to_scale) # transform returns a numpy array
train_df_scaled = train_df[['EngineID','cycle','engineRUL']].join(scaled_train_df_with_cols_to_scale) # join back non-scaled columns
train_df_scaled = train_df_scaled.reindex(columns = all_cols) # same columns order as before
# test data: repeat above steps
all_cols = test_df.columns
test_df_with_cols_to_scale = test_df[cols_to_scale]
scaled_test_df_with_cols_to_scale = pd.DataFrame(scaler.transform(test_df_with_cols_to_scale), columns=cols_to_scale) # transform returns a numpy array
test_df_scaled = test_df[['EngineID','cycle','engineRUL']].join(scaled_test_df_with_cols_to_scale) # join back non-scaled columns
test_df_scaled = test_df_scaled.reindex(columns = all_cols) # same columns order as before
# +
#%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
## re-format data into (samples, time steps, features) form
## %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
# -
# define utility function
def generate_LSTM_samples(engine_df, nSequenceSteps):
"""
This function generates list of LSTM samples (numpy arrays of size (nSequenceSteps, 24) each) for LSTM input
and list of output labels for LSTM
"""
engine_X_train_sequence = []
engine_y_train_sequence = []
engine_data = engine_df.values # converting to numpy
for sample in range(nSequenceSteps, engine_data.shape[0]):
engine_X_train_sequence.append(engine_data[sample-nSequenceSteps:sample,:-1]) # last column is output label
engine_y_train_sequence.append(engine_data[sample,-1])
return engine_X_train_sequence, engine_y_train_sequence
# +
nSequenceSteps = 50 # number of cycles in a sequence
X_train_sequence = []
y_train_sequence = []
# generate samples
for engineID in train_df_scaled['EngineID'].unique():
engine_df = train_df_scaled[train_df_scaled['EngineID'] == engineID]
engine_df = engine_df[['OPsetting1', 'OPsetting2', 'OPsetting3', 's1', 's2', 's3', 's4', 's5', 's6', 's7', 's8', 's9', 's10', 's11', 's12', 's13', 's14',
's15', 's16', 's17', 's18', 's19', 's20', 's21', 'engineRUL']]
engine_X_train_sequence, engine_y_train_sequence = generate_LSTM_samples(engine_df, nSequenceSteps)
X_train_sequence = X_train_sequence + engine_X_train_sequence # adding samples to the common list
y_train_sequence = y_train_sequence + engine_y_train_sequence
X_train_sequence, y_train_sequence = np.array(X_train_sequence), np.array(y_train_sequence) # convert list of (time steps, features) array into (samples, time steps, features) array
# +
#%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
## define and fit LSTM model
## %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, LSTM, Dropout
# custom metric
import tensorflow.keras.backend as K
def r2_custom(y_true, y_pred):
"""Coefficient of determination
"""
SS_res = K.sum(K.square(y_true - y_pred))
SS_tot = K.sum(K.square(y_true - K.mean(y_true)))
return (1 - SS_res/(SS_tot + K.epsilon()))
# define model
model = Sequential()
model.add(LSTM(units=100, return_sequences=True, input_shape=(nSequenceSteps, 24)))
model.add(Dropout(0.2))
model.add(LSTM(units=50))
model.add(Dropout(0.2))
model.add(Dense(1))
# model summary
model.summary()
# +
# compile model
model.compile(loss='mse', optimizer='Adam', metrics=r2_custom)
#%% fit model with early stopping
from tensorflow.keras.callbacks import EarlyStopping
es = EarlyStopping(monitor='val_loss', patience=10, restore_best_weights=True)
history = model.fit(X_train_sequence, y_train_sequence, epochs=200, batch_size=250, validation_split=0.3, callbacks=[es])
# +
# plot validation curve
plt.figure()
plt.title('Validation Curves: Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='val')
plt.legend()
plt.grid()
plt.show()
plt.figure()
plt.title('Validation Curves: R2')
plt.xlabel('Epoch')
plt.ylabel('R2')
plt.plot(history.history['r2_custom'], label='train')
plt.plot(history.history['val_r2_custom'], label='val')
plt.legend()
plt.grid()
plt.show()
# +
#%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
## evaluate model on test data
## %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
# +
# input/output test sequences (only the last sequence is used to predict failure)
X_test_sequence = []
y_test_sequence = []
for engineID in test_df_scaled['EngineID'].unique():
engine_df = test_df_scaled[test_df_scaled['EngineID'] == engineID]
if engine_df.shape[0] >= nSequenceSteps:
engine_df = engine_df[['OPsetting1', 'OPsetting2', 'OPsetting3', 's1', 's2', 's3', 's4', 's5', 's6', 's7', 's8', 's9', 's10', 's11', 's12', 's13', 's14',
's15', 's16', 's17', 's18', 's19', 's20', 's21', 'engineRUL']].values
X_test_sequence.append(engine_df[-nSequenceSteps:,:-1])
y_test_sequence.append(engine_df[-1,-1])
X_test_sequence, y_test_sequence = np.array(X_test_sequence), np.array(y_test_sequence)
# -
# evaluate performance
test_performance = model.evaluate(X_test_sequence, y_test_sequence)
print('R2_test: {}'.format(test_performance[1]))
# +
# plot
y_test_sequence_pred = model.predict(X_test_sequence)
plt.figure()
plt.plot(y_test_sequence_pred, color="blue", label='prediction')
plt.plot(y_test_sequence, color="green", label='actual')
plt.title('LSTM model prediction vs actual observation')
plt.ylabel('RUL')
plt.xlabel('Engine ID')
plt.legend(loc='upper left')
plt.show()
| Chapter_RNN/predictiveMaint_regression_RNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
import sqlite3
from sqlite3 import Error
print("imports done")
# -
def create_connection(db_file):
""" create a database connection to a SQLite database """
conn = None
try:
conn = sqlite3.connect(db_file)
print("connection success")
return conn
except Error as e:
print(e)
return conn
# +
def get_time():
conn=create_connection(r"../data1.db")
cur = conn.cursor()
cur.execute('select * from times')
b=0
while b!=None:
b=cur.fetchone()
if b!=None:
c=(b[1]+b[2]+b[3]+b[0])
print(c)
else:
break
conn.close()
get_time()
| Practice/Database Management/ip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright (c) Facebook, Inc. and its affiliates.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# # KubeFlow Pipelines : HPO with AX - Pytorch Cifar10 Image classification
#
# In this example, we train a Pytorch Lightning model to using image classification cifar10 dataset. A parent run will be created during the training process,which would dump the baseline model and relevant parameters,metrics and model along with its summary,subsequently followed by a set of nested child runs, which will dump the trial results. The best parameters would be dumped into the parent run once the experiments are completed.
#
# This notebook shows PyTorch CIFAR10 end-to-end classification example using Kubeflow Pipelines.
#
# An example notebook that demonstrates how to:
#
# * Get different tasks needed for the pipeline
# * Create a Kubeflow pipeline
# * Include Pytorch KFP components to preprocess, train, visualize and deploy the model in the pipeline
# * Submit a job for execution
# * Query(prediction and explain) the final deployed model
#
# ## import the necessary packages
# ! pip uninstall -y kfp
# ! pip install --no-cache-dir kfp ax-platform
# +
import kfp
import json
import os
from kfp.onprem import use_k8s_secret
from kfp import components
from kfp.components import load_component_from_file, load_component_from_url, func_to_container_op, InputPath
from kfp import dsl
from kfp import compiler
import numpy as np
import logging
from ax.service.ax_client import AxClient
import json
kfp.__version__
# -
# ## Enter your gateway and the auth token
# [Use this extension on chrome to get token]( https://chrome.google.com/webstore/detail/editthiscookie/fngmhnnpilhplaeedifhccceomclgfbg?hl=en)
#
# 
# ## Update values for the ingress gateway and auth session
INGRESS_GATEWAY='http://istio-ingressgateway.istio-system.svc.cluster.local'
AUTH="<auth-token>"
NAMESPACE="kubeflow-user-example-com"
COOKIE="authservice_session="+AUTH
EXPERIMENT="Default"
dist_volume = 'dist-vol'
volume_mount_path ="/model"
results_path = volume_mount_path+"/results.json"
# ## Set the Log bucket and Tensorboard Image
MINIO_ENDPOINT="http://minio-service.kubeflow:9000"
LOG_BUCKET="mlpipeline"
TENSORBOARD_IMAGE="public.ecr.aws/pytorch-samples/tboard:latest"
# ## Set the client and create the experiment
client = kfp.Client(host=INGRESS_GATEWAY+"/pipeline", cookies=COOKIE)
client.create_experiment(EXPERIMENT)
experiments = client.list_experiments(namespace=NAMESPACE)
my_experiment = experiments.experiments[0]
my_experiment
# ## Set the Inference parameters
DEPLOY_NAME="torchserve"
MODEL_NAME="cifar10"
ISVC_NAME=DEPLOY_NAME+"."+NAMESPACE+"."+"example.com"
INPUT_REQUEST="https://raw.githubusercontent.com/kubeflow/pipelines/master/samples/contrib/pytorch-samples/cifar10/input.json"
# ## Load the the components yaml files for setting up the components
# +
prepare_tensorboard_op = load_component_from_file("./common/tensorboard/component.yaml")
generate_trails_op = components.load_component_from_file(
"./common/ax_generate_trials/component.yaml"
)
complete_trails_op = components.load_component_from_file(
"./common/ax_complete_trials/component.yaml"
)
get_keys_op = components.load_component_from_file(
"../../../components/json/Get_keys/component.yaml"
)
get_element_op = components.load_component_from_file(
"../../../components/json/Get_element_by_key/component.yaml"
)
prep_op = components.load_component_from_file(
"./cifar10/yaml/pre_process/component.yaml"
)
# Uncomment hpo inputs in component yaml
train_op = components.load_component_from_file(
"./cifar10/yaml/train/component.yaml"
)
deploy_op = load_component_from_file("./common/deploy/component.yaml")
pred_op = load_component_from_file("./common/prediction/component.yaml")
minio_op = components.load_component_from_file(
"./common/minio/component.yaml"
)
kubernetes_create_pvc_op = load_component_from_file("../../../components/kubernetes/Create_PersistentVolumeClaim/component.yaml")
# +
from kubernetes.client.models import V1Volume, V1PersistentVolumeClaimVolumeSource
def create_dist_pipeline():
kubernetes_create_pvc_op(pvc_name=dist_volume, storage_size= "20Gi", namespace=NAMESPACE)
create_volume_run = client.create_run_from_pipeline_func(create_dist_pipeline, arguments={})
create_volume_run.wait_for_run_completion()
# + tags=[]
parameters = [
{"name": "lr", "type": "range", "bounds": [1e-4, 0.2], "log_scale": True},
{"name": "weight_decay", "type": "range", "bounds": [1e-4, 1e-2]},
{"name": "eps", "type": "range", "bounds": [1e-8, 1e-2]},
]
# -
# ## Define the pipeline
# + tags=[]
@dsl.pipeline(
name="AX Hpo", description="Estimating best parameters using AX"
)
def pytorch_ax_hpo( # pylint: disable=too-many-arguments
minio_endpoint=MINIO_ENDPOINT,
log_bucket=LOG_BUCKET,
log_dir=f"tensorboard/logs/{dsl.RUN_ID_PLACEHOLDER}",
mar_path=f"mar/{dsl.RUN_ID_PLACEHOLDER}/model-store",
config_prop_path=f"mar/{dsl.RUN_ID_PLACEHOLDER}/config",
model_uri=f"s3://mlpipeline/mar/{dsl.RUN_ID_PLACEHOLDER}",
best_params=f"hpo/{dsl.RUN_ID_PLACEHOLDER}",
tf_image=TENSORBOARD_IMAGE,
deploy=DEPLOY_NAME,
isvc_name=ISVC_NAME,
model=MODEL_NAME,
namespace=NAMESPACE,
confusion_matrix_log_dir=f"confusion_matrix/{dsl.RUN_ID_PLACEHOLDER}/",
checkpoint_dir="checkpoint_dir/cifar10",
input_req=INPUT_REQUEST,
cookie=COOKIE,
total_trials=2,
ingress_gateway=INGRESS_GATEWAY,
):
"""This method defines the pipeline tasks and operations"""
pod_template_spec = json.dumps({
"spec": {
"containers": [{
"env": [
{
"name": "AWS_ACCESS_KEY_ID",
"valueFrom": {
"secretKeyRef": {
"name": "mlpipeline-minio-artifact",
"key": "accesskey",
}
},
},
{
"name": "AWS_SECRET_ACCESS_KEY",
"valueFrom": {
"secretKeyRef": {
"name": "mlpipeline-minio-artifact",
"key": "secretkey",
}
},
},
{
"name": "AWS_REGION",
"value": "minio"
},
{
"name": "S3_ENDPOINT",
"value": f"{minio_endpoint}",
},
{
"name": "S3_USE_HTTPS",
"value": "0"
},
{
"name": "S3_VERIFY_SSL",
"value": "0"
},
]
}]
}
})
prepare_tb_task = prepare_tensorboard_op(
log_dir_uri=f"s3://{log_bucket}/{log_dir}",
image=tf_image,
pod_template_spec=pod_template_spec,
).set_display_name("Visualization")
prep_task = (
prep_op().after(prepare_tb_task).set_display_name("Preprocess & Transform")
)
gen_trials_task = generate_trails_op(total_trials, parameters, 'test-accuracy').after(prep_task).set_display_name("AX Generate Trials")
get_keys_task = get_keys_op(gen_trials_task.outputs["trial_parameters"]).after(gen_trials_task).set_display_name("Get Keys of Trials")
confusion_matrix_url = f"minio://{log_bucket}/{confusion_matrix_log_dir}"
script_args = f"model_name=resnet.pth," \
f"confusion_matrix_url={confusion_matrix_url}"
ptl_args = f"max_epochs=1, profiler=pytorch"
with dsl.ParallelFor(get_keys_task.outputs["keys"]) as item:
get_element_task = get_element_op(gen_trials_task.outputs["trial_parameters"], item).after(get_keys_task).set_display_name("Get Element from key")
train_task = (
train_op(
trial_id=item,
input_data=prep_task.outputs["output_data"],
cifar_script_args=script_args,
model_parameters=get_element_task.outputs["output"],
ptl_arguments=ptl_args,
results=results_path
).add_pvolumes({volume_mount_path: dsl.PipelineVolume(pvc=dist_volume)}).after(get_element_task).set_display_name("Training")
# For GPU uncomment below line and set GPU limit and node selector
# ).set_gpu_limit(1).add_node_selector_constraint('cloud.google.com/gke-accelerator','nvidia-tesla-p4')
)
complete_trials_task = complete_trails_op(gen_trials_task.outputs["client"], results_path).add_pvolumes({volume_mount_path: dsl.PipelineVolume(pvc=dist_volume)}).after(train_task).set_display_name("AX Complete Trials")
dsl.get_pipeline_conf().add_op_transformer(
use_k8s_secret(
secret_name="mlpipeline-minio-artifact",
k8s_secret_key_to_env={
"secretkey": "MINIO_SECRET_KEY",
"accesskey": "MINIO_ACCESS_KEY",
},
)
)
# -
# ## Compile the pipeline
compiler.Compiler().compile(pytorch_ax_hpo, 'pytorch.tar.gz', type_check=True)
# ## Execute the pipeline
run = client.run_pipeline(my_experiment.id, 'pytorch_ax_hpo', 'pytorch.tar.gz')
| samples/contrib/pytorch-samples/Pipeline-Cifar10-hpo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# # Airtable - Get data
# <img width="20%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/b2a2bo7r.png?w=800&h=400&fit=crop"/>
import naas_drivers
airtable = naas_drivers.airtable.connect('API_KEY', 'BASE_KEY', 'table_NAME')
data = airtable.get(view='All opportunities', maxRecords=20)
data
| Database/Airtable/Airtable_Get_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decision Analysis
# + [markdown] tags=[]
# Think Bayes, Second Edition
#
# Copyright 2020 <NAME>
#
# License: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# + tags=[]
# If we're running on Colab, install empiricaldist
# https://pypi.org/project/empiricaldist/
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
# !pip install empiricaldist
# + tags=[]
# Get utils.py
from os.path import basename, exists
def download(url):
filename = basename(url)
if not exists(filename):
from urllib.request import urlretrieve
local, _ = urlretrieve(url, filename)
print('Downloaded ' + local)
download('https://github.com/AllenDowney/ThinkBayes2/raw/master/soln/utils.py')
# + tags=[]
from utils import set_pyplot_params
set_pyplot_params()
# -
# This chapter presents a problem inspired by the game show *The Price is Right*.
# It is a silly example, but it demonstrates a useful process called Bayesian [decision analysis](https://en.wikipedia.org/wiki/Decision_analysis).
#
# As in previous examples, we'll use data and prior distribution to compute a posterior distribution; then we'll use the posterior distribution to choose an optimal strategy in a game that involves bidding.
#
# As part of the solution, we will use kernel density estimation (KDE) to estimate the prior distribution, and a normal distribution to compute the likelihood of the data.
#
# And at the end of the chapter, I pose a related problem you can solve as an exercise.
# ## The Price Is Right Problem
#
# On November 1, 2007, contestants named Letia and Nathaniel appeared on *The Price is Right*, an American television game show. They competed in a game called "The Showcase", where the objective is to guess the price of a collection of prizes. The contestant who comes closest to the actual price, without going over, wins the prizes.
#
# Nathaniel went first. His showcase included a dishwasher, a wine cabinet, a laptop computer, and a car. He bid \\$26,000.
#
# Letia's showcase included a pinball machine, a video arcade game, a pool table, and a cruise of the Bahamas. She bid \\$21,500.
#
# The actual price of Nathaniel's showcase was \\$25,347. His bid was too high, so he lost.
#
# The actual price of Letia's showcase was \\$21,578.
#
# She was only off by \\$78, so she won her showcase and, because her bid was off by less than 250, she also won Nathaniel's showcase.
# For a Bayesian thinker, this scenario suggests several questions:
#
# 1. Before seeing the prizes, what prior beliefs should the contestants have about the price of the showcase?
#
# 2. After seeing the prizes, how should the contestants update those beliefs?
#
# 3. Based on the posterior distribution, what should the contestants bid?
#
# The third question demonstrates a common use of Bayesian methods: decision analysis.
#
# This problem is inspired by [an example](https://nbviewer.jupyter.org/github/CamDavidsonPilon/Probabilistic-Programming-and-Bayesian-Methods-for-Hackers/blob/master/Chapter5_LossFunctions/Ch5_LossFunctions_PyMC3.ipynb) in <NAME>'s book, [*Probablistic Programming and Bayesian Methods for Hackers*](http://camdavidsonpilon.github.io/Probabilistic-Programming-and-Bayesian-Methods-for-Hackers).
# ## The Prior
#
# To choose a prior distribution of prices, we can take advantage of data from previous episodes. Fortunately, [fans of the show keep detailed records](https://web.archive.org/web/20121107204942/http://www.tpirsummaries.8m.com/).
#
# For this example, I downloaded files containing the price of each showcase from the 2011 and 2012 seasons and the bids offered by the contestants.
# + [markdown] tags=[]
# The following cells load the data files.
# + tags=[]
# Load the data files
download('https://raw.githubusercontent.com/AllenDowney/ThinkBayes2/master/data/showcases.2011.csv')
download('https://raw.githubusercontent.com/AllenDowney/ThinkBayes2/master/data/showcases.2012.csv')
# -
# The following function reads the data and cleans it up a little.
# +
import pandas as pd
def read_data(filename):
"""Read the showcase price data."""
df = pd.read_csv(filename, index_col=0, skiprows=[1])
return df.dropna().transpose()
# -
# I'll read both files and concatenate them.
# +
df2011 = read_data('showcases.2011.csv')
df2012 = read_data('showcases.2012.csv')
df = pd.concat([df2011, df2012], ignore_index=True)
# + tags=[]
print(df2011.shape, df2012.shape, df.shape)
# -
# Here's what the dataset looks like:
df.head(3)
# The first two columns, `Showcase 1` and `Showcase 2`, are the values of the showcases in dollars.
# The next two columns are the bids the contestants made.
# The last two columns are the differences between the actual values and the bids.
# ## Kernel Density Estimation
#
# This dataset contains the prices for 313 previous showcases, which we can think of as a sample from the population of possible prices.
#
# We can use this sample to estimate the prior distribution of showcase prices. One way to do that is kernel density estimation (KDE), which uses the sample to estimate a smooth distribution. If you are not familiar with KDE, you can [read about it here](https://mathisonian.github.io/kde).
#
# SciPy provides `gaussian_kde`, which takes a sample and returns an object that represents the estimated distribution.
#
# The following function takes `sample`, makes a KDE, evaluates it at a given sequence of quantities, `qs`, and returns the result as a normalized PMF.
# +
from scipy.stats import gaussian_kde
from empiricaldist import Pmf
def kde_from_sample(sample, qs):
"""Make a kernel density estimate from a sample."""
kde = gaussian_kde(sample)
ps = kde(qs)
pmf = Pmf(ps, qs)
pmf.normalize()
return pmf
# -
# We can use it to estimate the distribution of values for Showcase 1:
# +
import numpy as np
qs = np.linspace(0, 80000, 81)
prior1 = kde_from_sample(df['Showcase 1'], qs)
# -
# Here's what it looks like:
# + tags=[]
from utils import decorate
def decorate_value(title=''):
decorate(xlabel='Showcase value ($)',
ylabel='PMF',
title=title)
# + tags=[]
prior1.plot(label='Prior 1')
decorate_value('Prior distribution of showcase value')
# -
# **Exercise:** Use this function to make a `Pmf` that represents the prior distribution for Showcase 2, and plot it.
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Distribution of Error
#
# To update these priors, we have to answer these questions:
#
# * What data should we consider and how should we quantify it?
#
# * Can we compute a likelihood function; that is, for each hypothetical price, can we compute the conditional likelihood of the data?
#
# To answer these questions, I will model each contestant as a price-guessing instrument with known error characteristics.
# In this model, when the contestant sees the prizes, they guess the price of each prize and add up the prices.
# Let's call this total `guess`.
#
# Now the question we have to answer is, "If the actual price is `price`, what is the likelihood that the contestant's guess would be `guess`?"
#
# Equivalently, if we define `error = guess - price`, we can ask, "What is the likelihood that the contestant's guess is off by `error`?"
#
# To answer this question, I'll use the historical data again.
# For each showcase in the dataset, let's look at the difference between the contestant's bid and the actual price:
sample_diff1 = df['Bid 1'] - df['Showcase 1']
sample_diff2 = df['Bid 2'] - df['Showcase 2']
# To visualize the distribution of these differences, we can use KDE again.
qs = np.linspace(-40000, 20000, 61)
kde_diff1 = kde_from_sample(sample_diff1, qs)
kde_diff2 = kde_from_sample(sample_diff2, qs)
# Here's what these distributions look like:
# + tags=[]
kde_diff1.plot(label='Diff 1', color='C8')
kde_diff2.plot(label='Diff 2', color='C4')
decorate(xlabel='Difference in value ($)',
ylabel='PMF',
title='Difference between bid and actual value')
# -
# It looks like the bids are too low more often than too high, which makes sense. Remember that under the rules of the game, you lose if you overbid, so contestants probably underbid to some degree deliberately.
#
# For example, if they guess that the value of the showcase is \\$40,000, they might bid \\$36,000 to avoid going over.
# It looks like these distributions are well modeled by a normal distribution, so we can summarize them with their mean and standard deviation.
#
# For example, here is the mean and standard deviation of `Diff` for Player 1.
# +
mean_diff1 = sample_diff1.mean()
std_diff1 = sample_diff1.std()
print(mean_diff1, std_diff1)
# -
# Now we can use these differences to model the contestant's distribution of errors.
# This step is a little tricky because we don't actually know the contestant's guesses; we only know what they bid.
#
# So we have to make some assumptions:
#
# * I'll assume that contestants underbid because they are being strategic, and that on average their guesses are accurate. In other words, the mean of their errors is 0.
#
# * But I'll assume that the spread of the differences reflects the actual spread of their errors. So, I'll use the standard deviation of the differences as the standard deviation of their errors.
#
# Based on these assumptions, I'll make a normal distribution with parameters 0 and `std_diff1`.
#
# SciPy provides an object called `norm` that represents a normal distribution with the given mean and standard deviation.
# +
from scipy.stats import norm
error_dist1 = norm(0, std_diff1)
# -
# The result is an object that provides `pdf`, which evaluates the probability density function of the normal distribution.
#
# For example, here is the probability density of `error=-100`, based on the distribution of errors for Player 1.
error = -100
error_dist1.pdf(error)
# By itself, this number doesn't mean very much, because probability densities are not probabilities. But they are proportional to probabilities, so we can use them as likelihoods in a Bayesian update, as we'll see in the next section.
# ## Update
#
# Suppose you are Player 1. You see the prizes in your showcase and your guess for the total price is \\$23,000.
#
# From your guess I will subtract away each hypothetical price in the prior distribution; the result is your error under each hypothesis.
guess1 = 23000
error1 = guess1 - prior1.qs
# Now suppose we know, based on past performance, that your estimation error is well modeled by `error_dist1`.
# Under that assumption we can compute the likelihood of your error under each hypothesis.
likelihood1 = error_dist1.pdf(error1)
# The result is an array of likelihoods, which we can use to update the prior.
# + tags=[]
posterior1 = prior1 * likelihood1
posterior1.normalize()
# -
# Here's what the posterior distribution looks like:
# + tags=[]
prior1.plot(color='C5', label='Prior 1')
posterior1.plot(color='C4', label='Posterior 1')
decorate_value('Prior and posterior distribution of showcase value')
# -
# Because your initial guess is in the lower end of the range, the posterior distribution has shifted to the left. We can compute the posterior mean to see by how much.
prior1.mean(), posterior1.mean()
# Before you saw the prizes, you expected to see a showcase with a value close to \\$30,000.
# After making a guess of \\$23,000, you updated the prior distribution.
# Based on the combination of the prior and your guess, you now expect the actual price to be about \\$26,000.
# **Exercise:** Now suppose you are Player 2. When you see your showcase, you guess that the total price is \\$38,000.
#
# Use `diff2` to construct a normal distribution that represents the distribution of your estimation errors.
#
# Compute the likelihood of your guess for each actual price and use it to update `prior2`.
#
# Plot the posterior distribution and compute the posterior mean. Based on the prior and your guess, what do you expect the actual price of the showcase to be?
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Probability of Winning
#
# Now that we have a posterior distribution for each player, let's think about strategy.
#
# First, from the point of view of Player 1, let's compute the probability that Player 2 overbids. To keep it simple, I'll use only the performance of past players, ignoring the value of the showcase.
#
# The following function takes a sequence of past bids and returns the fraction that overbid.
def prob_overbid(sample_diff):
"""Compute the probability of an overbid."""
return np.mean(sample_diff > 0)
# Here's an estimate for the probability that Player 2 overbids.
prob_overbid(sample_diff2)
# Now suppose Player 1 underbids by \\$5000.
# What is the probability that Player 2 underbids by more?
#
# The following function uses past performance to estimate the probability that a player underbids by more than a given amount, `diff`:
def prob_worse_than(diff, sample_diff):
"""Probability opponent diff is worse than given diff."""
return np.mean(sample_diff < diff)
# Here's the probability that Player 2 underbids by more than \\$5000.
prob_worse_than(-5000, sample_diff2)
# And here's the probability they underbid by more than \\$10,000.
prob_worse_than(-10000, sample_diff2)
# We can combine these functions to compute the probability that Player 1 wins, given the difference between their bid and the actual price:
def compute_prob_win(diff, sample_diff):
"""Probability of winning for a given diff."""
# if you overbid you lose
if diff > 0:
return 0
# if the opponent overbids, you win
p1 = prob_overbid(sample_diff)
# or of their bid is worse than yours, you win
p2 = prob_worse_than(diff, sample_diff)
# p1 and p2 are mutually exclusive, so we can add them
return p1 + p2
# Here's the probability that you win, given that you underbid by \\$5000.
compute_prob_win(-5000, sample_diff2)
# Now let's look at the probability of winning for a range of possible differences.
xs = np.linspace(-30000, 5000, 121)
ys = [compute_prob_win(x, sample_diff2)
for x in xs]
# Here's what it looks like:
# + tags=[]
import matplotlib.pyplot as plt
plt.plot(xs, ys)
decorate(xlabel='Difference between bid and actual price ($)',
ylabel='Probability of winning',
title='Player 1')
# -
# If you underbid by \\$30,000, the chance of winning is about 30%, which is mostly the chance your opponent overbids.
#
# As your bids gets closer to the actual price, your chance of winning approaches 1.
#
# And, of course, if you overbid, you lose (even if your opponent also overbids).
# **Exercise:** Run the same analysis from the point of view of Player 2. Using the sample of differences from Player 1, compute:
#
# 1. The probability that Player 1 overbids.
#
# 2. The probability that Player 1 underbids by more than \\$5000.
#
# 3. The probability that Player 2 wins, given that they underbid by \\$5000.
#
# Then plot the probability that Player 2 wins for a range of possible differences between their bid and the actual price.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Decision Analysis
#
# In the previous section we computed the probability of winning, given that we have underbid by a particular amount.
#
# In reality the contestants don't know how much they have underbid by, because they don't know the actual price.
#
# But they do have a posterior distribution that represents their beliefs about the actual price, and they can use that to estimate their probability of winning with a given bid.
#
# The following function takes a possible bid, a posterior distribution of actual prices, and a sample of differences for the opponent.
#
# It loops through the hypothetical prices in the posterior distribution and, for each price,
#
# 1. Computes the difference between the bid and the hypothetical price,
#
# 2. Computes the probability that the player wins, given that difference, and
#
# 3. Adds up the weighted sum of the probabilities, where the weights are the probabilities in the posterior distribution.
def total_prob_win(bid, posterior, sample_diff):
"""Computes the total probability of winning with a given bid.
bid: your bid
posterior: Pmf of showcase value
sample_diff: sequence of differences for the opponent
returns: probability of winning
"""
total = 0
for price, prob in posterior.items():
diff = bid - price
total += prob * compute_prob_win(diff, sample_diff)
return total
# This loop implements the law of total probability:
#
# $$P(win) = \sum_{price} P(price) ~ P(win ~|~ price)$$
#
# Here's the probability that Player 1 wins, based on a bid of \\$25,000 and the posterior distribution `posterior1`.
total_prob_win(25000, posterior1, sample_diff2)
# Now we can loop through a series of possible bids and compute the probability of winning for each one.
# +
bids = posterior1.qs
probs = [total_prob_win(bid, posterior1, sample_diff2)
for bid in bids]
prob_win_series = pd.Series(probs, index=bids)
# -
# Here are the results.
# + tags=[]
prob_win_series.plot(label='Player 1', color='C1')
decorate(xlabel='Bid ($)',
ylabel='Probability of winning',
title='Optimal bid: probability of winning')
# -
# And here's the bid that maximizes Player 1's chance of winning.
prob_win_series.idxmax()
prob_win_series.max()
# Recall that your guess was \\$23,000.
# Using your guess to compute the posterior distribution, the posterior mean is about \\$26,000.
# But the bid that maximizes your chance of winning is \\$21,000.
# **Exercise:** Do the same analysis for Player 2.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Maximizing Expected Gain
#
# In the previous section we computed the bid that maximizes your chance of winning.
# And if that's your goal, the bid we computed is optimal.
#
# But winning isn't everything.
# Remember that if your bid is off by \\$250 or less, you win both showcases.
# So it might be a good idea to increase your bid a little: it increases the chance you overbid and lose, but it also increases the chance of winning both showcases.
#
# Let's see how that works out.
# The following function computes how much you will win, on average, given your bid, the actual price, and a sample of errors for your opponent.
def compute_gain(bid, price, sample_diff):
"""Compute expected gain given a bid and actual price."""
diff = bid - price
prob = compute_prob_win(diff, sample_diff)
# if you are within 250 dollars, you win both showcases
if -250 <= diff <= 0:
return 2 * price * prob
else:
return price * prob
# For example, if the actual price is \\$35000
# and you bid \\$30000,
# you will win about \\$23,600 worth of prizes on average, taking into account your probability of losing, winning one showcase, or winning both.
compute_gain(30000, 35000, sample_diff2)
# In reality we don't know the actual price, but we have a posterior distribution that represents what we know about it.
# By averaging over the prices and probabilities in the posterior distribution, we can compute the expected gain for a particular bid.
#
# In this context, "expected" means the average over the possible showcase values, weighted by their probabilities.
def expected_gain(bid, posterior, sample_diff):
"""Compute the expected gain of a given bid."""
total = 0
for price, prob in posterior.items():
total += prob * compute_gain(bid, price, sample_diff)
return total
# For the posterior we computed earlier, based on a guess of \\$23,000, the expected gain for a bid of \\$21,000 is about \\$16,900.
expected_gain(21000, posterior1, sample_diff2)
# But can we do any better?
#
# To find out, we can loop through a range of bids and find the one that maximizes expected gain.
# +
bids = posterior1.qs
gains = [expected_gain(bid, posterior1, sample_diff2) for bid in bids]
expected_gain_series = pd.Series(gains, index=bids)
# -
# Here are the results.
# + tags=[]
expected_gain_series.plot(label='Player 1', color='C2')
decorate(xlabel='Bid ($)',
ylabel='Expected gain ($)',
title='Optimal bid: expected gain')
# -
# Here is the optimal bid.
expected_gain_series.idxmax()
# With that bid, the expected gain is about \\$17,400.
expected_gain_series.max()
# Recall that your initial guess was \\$23,000.
# The bid that maximizes the chance of winning is \\$21,000.
# And the bid that maximizes your expected gain is \\$22,000.
# **Exercise:** Do the same analysis for Player 2.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Summary
#
# There's a lot going on this this chapter, so let's review the steps:
#
# 1. First we used KDE and data from past shows to estimate prior distributions for the values of the showcases.
#
# 2. Then we used bids from past shows to model the distribution of errors as a normal distribution.
#
# 3. We did a Bayesian update using the distribution of errors to compute the likelihood of the data.
#
# 4. We used the posterior distribution for the value of the showcase to compute the probability of winning for each possible bid, and identified the bid that maximizes the chance of winning.
#
# 5. Finally, we used probability of winning to compute the expected gain for each possible bid, and identified the bid that maximizes expected gain.
#
# Incidentally, this example demonstrates the hazard of using the word "optimal" without specifying what you are optimizing.
# The bid that maximizes the chance of winning is not generally the same as the bid that maximizes expected gain.
# ## Discussion
#
# When people discuss the pros and cons of Bayesian estimation, as contrasted with classical methods sometimes called "frequentist", they often claim that in many cases Bayesian methods and frequentist methods produce the same results.
#
# In my opinion, this claim is mistaken because Bayesian and frequentist method produce different *kinds* of results:
#
# * The result of frequentist methods is usually a single value that is considered to be the best estimate (by one of several criteria) or an interval that quantifies the precision of the estimate.
#
# * The result of Bayesian methods is a posterior distribution that represents all possible outcomes and their probabilities.
# Granted, you can use the posterior distribution to choose a "best" estimate or compute an interval.
# And in that case the result might be the same as the frequentist estimate.
#
# But doing so discards useful information and, in my opinion, eliminates the primary benefit of Bayesian methods: the posterior distribution is more useful than a single estimate, or even an interval.
# The example in this chapter demonstrates the point.
# Using the entire posterior distribution, we can compute the bid that maximizes the probability of winning, or the bid that maximizes expected gain, even if the rules for computing the gain are complicated (and nonlinear).
#
# With a single estimate or an interval, we can't do that, even if they are "optimal" in some sense.
# In general, frequentist estimation provides little guidance for decision-making.
#
# If you hear someone say that Bayesian and frequentist methods produce the same results, you can be confident that they don't understand Bayesian methods.
# ## Exercises
# **Exercise:** When I worked in Cambridge, Massachusetts, I usually took the subway to South Station and then a commuter train home to Needham. Because the subway was unpredictable, I left the office early enough that I could wait up to 15 minutes and still catch the commuter train.
#
# When I got to the subway stop, there were usually about 10 people waiting on the platform. If there were fewer than that, I figured I just missed a train, so I expected to wait a little longer than usual. And if there there more than that, I expected another train soon.
#
# But if there were a *lot* more than 10 passengers waiting, I inferred that something was wrong, and I expected a long wait. In that case, I might leave and take a taxi.
#
# We can use Bayesian decision analysis to quantify the analysis I did intuitively. Given the number of passengers on the platform, how long should we expect to wait? And when should we give up and take a taxi?
#
# My analysis of this problem is in `redline.ipynb`, which is in the repository for this book. [Click here to run this notebook on Colab](https://colab.research.google.com/github/AllenDowney/ThinkBayes2/blob/master/notebooks/redline.ipynb).
# **Exercise:** This exercise is inspired by a true story. In 2001 I created [Green Tea Press](https://greenteapress.com) to publish my books, starting with *Think Python*. I ordered 100 copies from a short run printer and made the book available for sale through a distributor.
#
# After the first week, the distributor reported that 12 copies were sold. Based that report, I thought I would run out of copies in about 8 weeks, so I got ready to order more. My printer offered me a discount if I ordered more than 1000 copies, so I went a little crazy and ordered 2000.
#
# A few days later, my mother called to tell me that her *copies* of the book had arrived. Surprised, I asked how many. She said ten.
#
# It turned out I had sold only two books to non-relatives. And it took a lot longer than I expected to sell 2000 copies.
# The details of this story are unique, but the general problem is something almost every retailer has to figure out. Based on past sales, how do you predict future sales? And based on those predictions, how do you decide how much to order and when?
#
# Often the cost of a bad decision is complicated. If you place a lot of small orders rather than one big one, your costs are likely to be higher. If you run out of inventory, you might lose customers. And if you order too much, you have to pay the various costs of holding inventory.
#
# So, let's solve a version of the problem I faced. It will take some work to set up the problem; the details are in the notebook for this chapter.
# + [markdown] tags=[]
# Suppose you start selling books online. During the first week you sell 10 copies (and let's assume that none of the customers are your mother). During the second week you sell 9 copies.
#
# Assuming that the arrival of orders is a Poisson process, we can think of the weekly orders as samples from a Poisson distribution with an unknown rate.
# We can use orders from past weeks to estimate the parameter of this distribution, generate a predictive distribution for future weeks, and compute the order size that maximized expected profit.
#
# * Suppose the cost of printing the book is \\$5 per copy,
#
# * But if you order 100 or more, it's \\$4.50 per copy.
#
# * For every book you sell, you get \\$10.
#
# * But if you run out of books before the end of 8 weeks, you lose \\$50 in future sales for every week you are out of stock.
#
# * If you have books left over at the end of 8 weeks, you lose \\$2 in inventory costs per extra book.
#
# For example, suppose you get orders for 10 books per week, every week. If you order 60 books,
#
# * The total cost is \\$300.
#
# * You sell all 60 books, so you make \\$600.
#
# * But the book is out of stock for two weeks, so you lose \\$100 in future sales.
#
# In total, your profit is \\$200.
#
# If you order 100 books,
#
# * The total cost is \\$450.
#
# * You sell 80 books, so you make \\$800.
#
# * But you have 20 books left over at the end, so you lose \\$40.
#
# In total, your profit is \\$310.
#
# Combining these costs with your predictive distribution, how many books should you order to maximize your expected profit?
# + [markdown] tags=[]
# To get you started, the following functions compute profits and costs according to the specification of the problem:
# + tags=[]
def print_cost(printed):
"""Compute print costs.
printed: integer number printed
"""
if printed < 100:
return printed * 5
else:
return printed * 4.5
# + tags=[]
def total_income(printed, orders):
"""Compute income.
printed: integer number printed
orders: sequence of integer number of books ordered
"""
sold = min(printed, np.sum(orders))
return sold * 10
# + tags=[]
def inventory_cost(printed, orders):
"""Compute inventory costs.
printed: integer number printed
orders: sequence of integer number of books ordered
"""
excess = printed - np.sum(orders)
if excess > 0:
return excess * 2
else:
return 0
# + tags=[]
def out_of_stock_cost(printed, orders):
"""Compute out of stock costs.
printed: integer number printed
orders: sequence of integer number of books ordered
"""
weeks = len(orders)
total_orders = np.cumsum(orders)
for i, total in enumerate(total_orders):
if total > printed:
return (weeks-i) * 50
return 0
# + tags=[]
def compute_profit(printed, orders):
"""Compute profit.
printed: integer number printed
orders: sequence of integer number of books ordered
"""
return (total_income(printed, orders) -
print_cost(printed)-
out_of_stock_cost(printed, orders) -
inventory_cost(printed, orders))
# + [markdown] tags=[]
# To test these functions, suppose we get exactly 10 orders per week for eight weeks:
# + tags=[]
always_10 = [10] * 8
always_10
# + [markdown] tags=[]
# If you print 60 books, your net profit is \\$200, as in the example.
# + tags=[]
compute_profit(60, always_10)
# + [markdown] tags=[]
# If you print 100 books, your net profit is \\$310.
# + tags=[]
compute_profit(100, always_10)
# + [markdown] tags=[]
# Of course, in the context of the problem you don't know how many books will be ordered in any given week. You don't even know the average rate of orders. However, given the data and some assumptions about the prior, you can compute the distribution of the rate of orders.
#
# You'll have a chance to do that, but to demonstrate the decision analysis part of the problem, I'll start with the arbitrary assumption that order rates come from a gamma distribution with mean 9.
#
# Here's a `Pmf` that represents this distribution.
# + tags=[]
from scipy.stats import gamma
alpha = 9
qs = np.linspace(0, 25, 101)
ps = gamma.pdf(qs, alpha)
pmf = Pmf(ps, qs)
pmf.normalize()
pmf.mean()
# + [markdown] tags=[]
# And here's what it looks like:
# + tags=[]
pmf.plot(color='C1')
decorate(xlabel=r'Book ordering rate ($\lambda$)',
ylabel='PMF')
# + [markdown] tags=[]
# Now, we *could* generate a predictive distribution for the number of books ordered in a given week, but in this example we have to deal with a complicated cost function. In particular, `out_of_stock_cost` depends on the sequence of orders.
#
# So, rather than generate a predictive distribution, I suggest we run simulations. I'll demonstrate the steps.
#
# First, from our hypothetical distribution of rates, we can draw a random sample of 1000 values.
# + tags=[]
rates = pmf.choice(1000)
np.mean(rates)
# + [markdown] tags=[]
# For each possible rate, we can generate a sequence of 8 orders.
# + tags=[]
np.random.seed(17)
order_array = np.random.poisson(rates, size=(8, 1000)).transpose()
order_array[:5, :]
# + [markdown] tags=[]
# Each row of this array is a hypothetical sequence of orders based on a different hypothetical order rate.
#
# Now, if you tell me how many books you printed, I can compute your expected profits, averaged over these 1000 possible sequences.
# + tags=[]
def compute_expected_profits(printed, order_array):
"""Compute profits averaged over a sample of orders.
printed: number printed
order_array: one row per sample, one column per week
"""
profits = [compute_profit(printed, orders)
for orders in order_array]
return np.mean(profits)
# + [markdown] tags=[]
# For example, here are the expected profits if you order 70, 80, or 90 books.
# + tags=[]
compute_expected_profits(70, order_array)
# + tags=[]
compute_expected_profits(80, order_array)
# + tags=[]
compute_expected_profits(90, order_array)
# + [markdown] tags=[]
# Now, let's sweep through a range of values and compute expected profits as a function of the number of books you print.
# + tags=[]
printed_array = np.arange(70, 110)
t = [compute_expected_profits(printed, order_array)
for printed in printed_array]
expected_profits = pd.Series(t, printed_array)
# + tags=[]
expected_profits.plot(label='')
decorate(xlabel='Number of books printed',
ylabel='Expected profit ($)')
# + [markdown] tags=[]
# Here is the optimal order and the expected profit.
# + tags=[]
expected_profits.idxmax(), expected_profits.max()
# + [markdown] tags=[]
# Now it's your turn. Choose a prior that you think is reasonable, update it with the data you are given, and then use the posterior distribution to do the analysis I just demonstrated.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
| notebooks/chap09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # IV.G - Timestamp Estimation
#
# Code for generating the consistency heatmaps for the timestamp estimation of a single image
# +
# # !pip install --upgrade scikit-image
# # !pip install pyproj
# -
# ### Imports & Definitions
# +
import numpy as np
import matplotlib.pyplot as plt
import os, sys
from itertools import product
import matplotlib.image as mpimg
import pyproj
from skimage import transform
sys.path.append("../../datasets")
from dataLoader import DataLoader, preprocess_time
from tf.keras.models import Model, load_model
from tf.keras.layers import Input
from tf.keras.losses import mean_squared_error
transLabels = ["dirty", "daylight", "night", "sunrisesunset", "dawndusk", "sunny", "clouds",
"fog", "storm", "snow", "warm", "cold", "busy", "beautiful", "flowers", "spring",
"summer", "autumn", "winter", "glowing", "colorful", "dull", "rugged", "midday",
"dark", "bright", "dry", "moist", "windy", "rain", "ice", "cluttered", "soothing",
"stressful", "exciting", "sentimental", "mysterious", "boring", "gloomy", "lush"]
batchSize = 1
pathToModel = "../IV.B_ablation_study/denseNet/gr_oh_loc_time_TA/weights.30-0.57407.hdf5"
gpuNumber = 4
## GPU selection
import tensorflow as tf
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_visible_devices(gpus[gpuNumber], 'GPU')
tf.config.experimental.set_memory_growth(gpus[gpuNumber], False)
#######################
## Custom MSE ##
#######################
# We will compute the MSE only for the consistent inputs
def transient_mse(y_true, y_pred):
return tf.sum(mean_squared_error(y_true[0::2,:], y_pred[0::2,:]), axis=-1)
#######################
## Deprocess time
#######################
def deprocess_time(time):
month, hour = time
month = (11.0 / 2.0) * (month + 1) + 1
hour = (23.0 / 2.0) * (hour + 1)
return (int(round(month)), int(round(hour)))
def deprocess_loc(loc):
_earth_radius = 6378137.0
x,y,z = loc
ecef = pyproj.Proj(proj='geocent', ellps='WGS84', datum='WGS84')
lla = pyproj.Proj(proj='latlong', ellps='WGS84', datum='WGS84')
lon, lat, alt = pyproj.transform(ecef, lla, x * _earth_radius, y*_earth_radius, z*_earth_radius, radians=False)
return (lat, lon, alt)
# -
# ### Load architecture and get pointers to specific layers
# As we will process the features from multiple timestamps, we avoid re-processing the features for the ground-level image, location, and satellite image
# +
baseModel = load_model(pathToModel, custom_objects={"transient_mse": transient_mse})
print(baseModel.summary())
groundBranchModel = Model(baseModel.get_layer("groundInput").input,
baseModel.get_layer("batch_normalization_2").output)
aerialBranchModel = Model(baseModel.get_layer("aerialInput").input,
baseModel.get_layer("batch_normalization_4").output)
locBranchModel = Model(baseModel.get_layer("locationInput").input,
baseModel.get_layer("batch_normalization_7").output)
timeBranchModel = Model(baseModel.get_layer("timeInput").input,
baseModel.get_layer("batch_normalization_10").output)
combinedFeaturesInput = Input(shape=(512,), name='concatenate_1_proxy')
combinedFeatures = baseModel.get_layer("consist_fc1")(combinedFeaturesInput)
combinedFeatures = baseModel.get_layer("batch_normalization_11")(combinedFeatures)
combinedFeatures = baseModel.get_layer("consist_fc2")(combinedFeatures)
combinedFeatures = baseModel.get_layer("batch_normalization_12")(combinedFeatures)
combinedFeatures = baseModel.get_layer("consist_fc3")(combinedFeatures)
combineModel = Model(combinedFeaturesInput, combinedFeatures)
# -
# --------------
# --------------
# --------------
# --------------
#
#
# ### Predicting the consistency probability for all hours and months
#
# The `skipCount` variable allows us to control which image would be selected (without needing to adapt much of the dataLoader)
# +
dl = DataLoader("test",
includeLocation = True,
includeSatellite = True,
outputTransientAttributes = True)
skipCount = 10
for batch, _ in dl.loadTestDataInBatches(batchSize, allTestSet=False):
if skipCount >= 1:
skipCount-=1
continue
grImg, aeImg, locInfo, timeInfo = [batch[i][0:1] for i in range(len(batch))]
dLoc = deprocess_loc(locInfo[0])
timeInfo = deprocess_time(timeInfo[0])
grFeatures = groundBranchModel.predict_on_batch(grImg)
aeFeatures = aerialBranchModel.predict_on_batch(aeImg)
locFeatures = locBranchModel.predict_on_batch(locInfo)
dLoc = deprocess_loc(locInfo[0])
timeList, predList = [], []
for month, hour in product(range(1,13), range(24)):
timeFeatures = timeBranchModel.predict_on_batch(preprocess_time((month, hour)).reshape(1,-1))
concatFV = np.hstack((grFeatures, aeFeatures, locFeatures, timeFeatures))
pred = consistModel.predict_on_batch(concatFV)
consistentProb = pred[0][0]
timeList += [(month, hour)]
predList += [consistentProb]
break
sortedTimeList = [(y,x) for y,x in sorted(zip(predList,timeList),
key=lambda pair: pair[0], reverse=True)]
sortedProbs = [l[0] for l in sortedTimeList]
sortedTimes = [l[1] for l in sortedTimeList]
# -
# Given the location (`dLoc`), we shifted the heatmap based on the timeZone to present it as local time rather than UCT.
#
# The timezone can be found manually, for example, with [this tool](https://timezonedb.com/).
# +
print(dLoc)
timeZone = +1 #Should change this considering the timeZone
# +
adjustedHeatMapMatrix = np.zeros((12, 24))
for i in range(len(sortedTimes)):
month, hour = sortedTimes[i]
adjustedHeatMapMatrix[month-1,((hour + timeZone)%24)] = sortedProbs[i]
fig=plt.figure(figsize=(8,8))
columns = 1
rows = 1
fig.add_subplot(rows, columns, 1)
fig.tight_layout()
plt.xticks(range(0,24), range(0,24), rotation=30, size=13)
plt.yticks(range(0,13),
["Jan.", "Feb.", "Mar.", "Apr.", "May", "Jun.", "Jul.", "Aug.", "Sept.", "Oct.", "Nov.", "Dec."],
size=13)
plt.scatter((timeInfo[1]+timeZone)%24, timeInfo[0]-1, lw=4, c='r', marker="*")
plt.imshow(adjustedHeatMapMatrix, cmap='viridis')
plt.show()
| paper_evaluation/IV.G_time_estimation/tifs_timeEstimation_singleImage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="color:#777777;background-color:#ffffff;font-size:12px;text-align:right;">
# prepared by <NAME> (QuSoft@Riga) | November 02, 2018
# </div>
# <table><tr><td><i> I have some macros here. If there is a problem with displaying mathematical formulas, please run me to load these macros.</i></td></td></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\inner}[2]{\langle #1,#2\rangle} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2> Matrices: Two Dimensional Lists </h2>
#
# A matrix is a list of vectors with the same dimensions.
#
# Here is an example matrix formed by 4 row vectors with dimension 5:
#
# $$
# M = \mymatrix{rrrrr}{8 & 0 & -1 & 0 & 2 \\ -2 & -3 & 1 & 1 & 4 \\ 0 & 0 & 1 & -7 & 1 \\ 1 & 4 & -2 & 5 & 9}.
# $$
#
# We can also say that $M$ is formed by 5 column vectors with dimension 4.
#
# $M$ is called an $ (4 \times 5) $-dimensional matrix. ($4 \times 5$: "four times five")
#
# We can represent $M$ as a two dimensional list in python.
# +
# we can break lines when defining our list
M = [
[8 , 0 , -1 , 0 , 2],
[-2 , -3 , 1 , 1 , 4],
[0 , 0 , 1 , -7 , 1],
[1 , 4 , -2 , 5 , 9]
]
# let's print matrix M
print(M)
# +
# let's print M in matrix form, row by row
for i in range(4): # there are 4 rows
print(M[i])
# -
# Remark that, by definition, the rows and columns of matrices are indexed starting from 1.
#
# The $ (i,j) $-th entry of $ M $ refers to the entry in $ i $-th row and $ j $-th column.
#
# (It is also denoted as $ M[i,j] $, $ M(i,j) $, or $ M_{ij} $.)
#
# On the other hand, in python, the indices start from zero.
#
# So, when we define a list for a matrix or vector in python, the value of an index is one less than the value of original index.
#
# Here are a few examples:
# +
M = [
[8 , 0 , -1 , 0 , 2],
[-2 , -3 , 1 , 1 , 4],
[0 , 0 , 1 , -7 , 1],
[1 , 4 , -2 , 5 , 9]
]
#let's print the element of M in the 1st row and the 1st column.
print(M[0][0])
#let's print the element of M in the 3rd row and the 4th column.
print(M[2][3])
#let's print the element of M in the 4th row and the 5th column.
print(M[3][4])
# -
# <h3> Multiplying a matrix with a number </h3>
#
# Let's multiply matrix $ M $ by $ -2 $.
#
# Each entry is multiplied by $ -2 $.
# +
# we use double nested for-loops
N =[] # the result matrix
for i in range(4): # for each row
N.append([]) # create an empty sub-list for each row in the result matrix
for j in range(5): # in row (i+1), for each column
N[i].append(M[i][j]*-2) # we add new elements into the i-th sub-list
# let's print M and N, and see the results
print("I am M:")
for i in range(4):
print(M[i])
print()
print("I am N:")
for i in range(4):
print(N[i])
# -
# After executing our program, we can write down the matrix $ N= -2 \cdot M $:
#
# $$
# N= -2 M = \mymatrix{rrrrr}{-16 & 0 & 2 & 0 & -4 \\ 4 & 6 & -2 & -2 & -8 \\ 0 & 0 & -2 & 14 & -2 \\ -2 & -8 & 4 & -10 & -18}.
# $$
# <h3> The summation of matrices</h3>
#
# If $ M $ and $ N $ are matrices with the same dimensions, then $ M+N $ is also a matrix with the same dimension.
#
# The summation of two matrices is similar to the summation of two vectors.
#
# If $ K = M +N $, then $ K[i,j] = M[i,j] + N[i,j] $ for every pair of $ (i,j) $.
#
# Let's find $ K $ in python.
# +
# create an empty list for the result matrix
K=[]
for i in range(len(M)): # len(M) return the number of rows in M
K.append([]) # we create a new row for K
for j in range(len(M[0])): # len(M[0]) returns the number of columns in M
K[i].append(M[i][j]+N[i][j]) # we add new elements into the i-th sublist/rows
# print each matrix in a single line
print("M=",M)
print("N=",N)
print("K=",K)
# -
# <b> Observation:</b>
#
# $ K = N +M $. We defined $ N $ as $ -2 M $.
#
# Thus, $ K = N+M = -2M + M = -M $.
#
# We can see that $ K = -M $ by looking the outcomes of our program.
# <h3> Task 1 </h3>
#
# Randomly create $ (3 \times 4) $-dimensional matrices $ A $ and $ B $.
#
# The entries can be from the list $ \{-5,\ldots,5\} $.
#
# Print the entries of both matrices.
#
# Find matrix $ C = 3A - 2B $, and print its entries. (<i>Note that $ 3A - 2B = 3A + (-2B) $</i>.)
#
# Verify the correctness your outcomes.
from random import randrange
#
# your solution is here
#
# <a href="..\bronze-solutions\B20_Python_Lists_Matrices_Solutions.ipynb#task1">click for our solution</a>
# <h3> Transpose of a matrix</h3>
#
# The transpose of a matrix is obtained by interchanging rows and columns.
#
# For example, the second row becomes the new second column, and third column becomes the new third row.
#
# The transpose of a matrix $ M $ is denoted $ M^T $.
#
# Here we give two examples.
#
# $$
# M = \mymatrix{rrrr}{-2 & 3 & 0 & 4\\ -1 & 1 & 5 & 9} ~~~~~ \Rightarrow ~~~~~ M^T=\mymatrix{rr}{-2 & -1 \\ 3 & 1 \\ 0 & 5 \\ 4 & 9} ~~~~~~~~ \mbox{ and } ~~~~~~~~
# N = \mymatrix{ccc}{1 & 2 & 3 \\ 4 & 5 & 6 \\ 7 & 8 & 9} ~~~~~ \Rightarrow ~~~~~ N^T = \mymatrix{ccc}{1 & 4 & 7 \\ 2 & 5 & 8 \\ 3 & 6 & 9}.
# $$
#
# Shortly, $ M[i,j] = M^T[j,i] $ and $ N[i,j] = N^T[j,i] $. (The indices are interchanged.)
# <h3> Task 2 </h3>
#
# Calculate $ M^T $ and $ N^T $ in python.
#
# Print all matrices and verify the correctness of your outcome.
M = [
[-2,3,0,4],
[-1,1,5,9]
]
N =[
[1,2,3],
[4,5,6],
[7,8,9]
]
#
# your solution is here
#
# <a href="..\bronze-solutions\B20_Python_Lists_Matrices_Solutions.ipynb#task2">click for our solution</a>
# <h3> Multiplication of a matrix with a vector </h3>
#
# We define a matrix $ M $ and a column vector $ v $:
#
# $$
# M = \mymatrix{rrr}{-1 & 0 & 1 \\ -2 & -3 & 4 \\ 1 & 5 & 6} ~~~~~~\mbox{and}~~~~~~ v = \myrvector{1 \\ -3 \\ 2}.
# $$
#
# The multiplication of $ M v $ is a new vector $ u $ shown as $ u = M v $:
# <ul>
# <li> The first entry of $u $ is the inner product of the first row of $ M $ and $ v $.</li>
# <li> The second entry of $ u $ is the inner product of the second row of $M$ and $ v $.</li>
# <li> The third entry of $ u $ is the inner product of the third row of $M$ and $v$. </li>
# </ul>
#
# Let's do the calculations in python.
# +
# matrix M
M = [
[-1,0,1],
[-2,-3,4],
[1,5,6]
]
# vector v
v = [1,-3,2]
# the result vector u
u = []
# for each row, we do an inner product
for i in range(3):
# inner product for one row is initiated
inner_result = 0 # this variable keeps the summation of the pairwise multiplications
for j in range(3): # the elements in the i-th row
inner_result = inner_result + M[i][j] * v[j]
# inner product for one row is completed
u.append(inner_result)
print("M is")
for i in range(len(M)):
print(M[i])
print()
print("v=",v)
print()
print("u=",u)
# -
# Let's check the calculations:
#
# $$
# \mbox{First row:}~~~~ \biginner{ \myrvector{-1 \\ 0 \\ 1} }{ \myrvector{1 \\ -3 \\ 2} } = (-1)\cdot 1 + 0 \cdot (-3) + 1 \cdot 2 = -1 + 0 + 2 = -1.
# $$
# $$
# \mbox{Second row:}~~~~ \biginner{ \myrvector{-2 \\ -3 \\ 4} }{ \myrvector{1 \\ -3 \\ 2} } = (-2)\cdot 1 + (-3) \cdot (-3) + 4 \cdot 2 = -2 + 9 + 8 = 15.
# $$
# $$
# \mbox{Third row:}~~~~ \biginner{ \myrvector{1 \\ 5 \\ 6} }{ \myrvector{1 \\ -3 \\ 2} } = 1\cdot 1 + 5 \cdot (-3) + 6 \cdot 2 = 1 - 15 + 12 = -2.
# $$
#
# Then,
# $$
# v' = \myrvector{-1 \\ 15 \\ -2 }.
# $$
# <b>Observations:</b>
# <ul>
# <li> The dimension of the row of $ M $ is the same as the dimension of $ v $. Otherwise, the inner product is not defined.</li>
# <li> The dimension of the result vector is the number of rows in $ M $, because we have an inner product for each row of $ M $</li>
# </ul>
# <h3> Task 3 </h3>
#
# Find $ u' = N u $ in python for the following matrix $ N $ and column vector $ u $:
#
# $$
# N = \mymatrix{rrr}{-1 & 1 & 2 \\ 0 & -2 & -3 \\ 3 & 2 & 5 \\ 0 & 2 & -2} ~~~~~~\mbox{and}~~~~~~ u = \myrvector{2 \\ -1 \\ 3}.
# $$
#
# your solution is here
#
# <a href="..\bronze-solutions\B20_Python_Lists_Matrices_Solutions.ipynb#task3">click for our solution</a>
# <h3> Multiplication of two matrices </h3>
#
# This is just the generalization of the procedure given above.
#
# Let's find matrix $ K = M \cdot N $ for given matrices
# $
# M = \mymatrix{rrr}{-1 & 0 & 1 \\ -2 & -1 & 2 \\ 1 & 2 & -2} ~~\mbox{and}~~
# N = \mymatrix{rrr}{0 & 2 & 1 \\ 3 & -1 & -2 \\ -1 & 1 & 0}.
# $
# Remark that the matrix $ N $ has three columns: $ v_1 = \myrvector{0 \\ 3 \\ -1} $, $ v_2 = \myrvector{2 \\ -1 \\ 1} $, and $ v_3 = \myrvector{1 \\ -2 \\ 0} $.
#
# We know how to calculate $ v_1' = M \cdot v_1 $.
#
# Similarly, we can calculate $ v_2' = M \cdot v_2 $ and $ v_3' = M \cdot v_3 $.
#
# You may have already guessed that these new column vectors ($v_1'$, $v_2'$, and $v_3'$) are the columns of $ K $.
#
# Technically, the inner product of the i-th row of $ M $ and $ j $-th row of $ N $ gives the $(i,j)$-th entry of $ K $.
# <h3> Task 4 </h3>
#
# Find matrix $ K $.
#
# This is a challanging task. You may use triple nested for-loops.
#
# You may also consider to write a function taking two lists and returning their inner product.
# +
# matrix M
M = [
[-1,0,1],
[-2,-1,2],
[1,2,-2]
]
# matrix N
M = [
[0,2,1],
[3,-1,-2],
[-1,1,0]
]
# matrix K
K = []
#
# your solution is here
#
# -
# <a href="..\bronze-solutions\B20_Python_Lists_Matrices_Solutions.ipynb#task4">click for our solution</a>
# <h3> Is $ A B = B A $? </h3>
#
# It is a well-known fact that the order of numbers does not matter in multiplication.
#
# For example, $ (-3) \cdot 4 = 4 \cdot (-3) $.
#
# Is it also true for matrices? For any given two matrices $ A $ and $ B $, is $ A \cdot B = B \cdot A $?
#
# There are some examples of $A$ and $B$ such that $ A \cdot B = B \cdot A $.
#
# But this is not true for most of the cases, and so this statement is false.
#
# We can falsify this statement by using our programming skill.
#
# We can also follow a probabilistic strategy.
#
# The idea is as follows: Randomly find two example matrices $ A $ and $ B $ such that $ AB \neq BA $.
#
# Remark that if $ AB = BA $, then $ AB - BA $ is a zero matrix.
# <h3> Task 5 </h3>
#
# Randomly define two $ (2 \times 2) $-dimensional matrices $A$ and $ B $.
#
# Then, find $ C= AB-BA $. If $ C $ is not a zero matrix, then we are done.
#
# <i>Remark: With some chances, we may find a pair of $ (A,B) $ such that $ AB = BA $.
#
# In this case, we should repeat our experiment. </i>
#
# your solution is here
#
# <a href="..\bronze-solutions\B20_Python_Lists_Matrices_Solutions.ipynb#task5">click for our solution</a>
| community/awards/teach_me_quantum_2018/bronze/bronze/B20_Lists_Matrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy
import cv2
im_grey = cv2.imread("smallgrey.png", 0)
im_grey
im_grey[0:2,2:4]
im_grey[:2,2:]
for i in im_grey:
print(i)
for i in im_grey.T:
print(i)
for i in im_grey.flat:
print(i)
ims = numpy.hstack((im_grey, im_grey))
ims
ims = numpy.hstack((im_grey, im_grey, im_grey))
ims
ims = numpy.vstack((im_grey, im_grey))
ims
lst = numpy.hsplit(ims, 5)
lst
lstv = numpy.vsplit(ims, 2)
lstv
| s07/lecture109.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import matplotlib.pyplot as plt
# %matplotlib notebook
def read_file(string):
with open(string) as f:
layerwise_magnitude = f.read().splitlines()
return layerwise_magnitude[:-1]
def parse_result(string):
first_split = string.split(",")
x, y = [(val.split(':'))[-1].strip() for val in first_split]
return [int(x), float(y)]
files = ['run_magnitude_all_layers_prune', 'run_magnitude_layerwise_prune', 'run_obd_all_layers_prune', 'run_obd_layerwise_prune', 'run_obd_all_layers_dropout']
files2=['run_obd_all_layers_prune','run_obd_all_layers_dropout']
fig = plt.figure(figsize=(12,10))
for f in files:
results = read_file("MNIST_Pruning_logs/"+f+"/results.txt")
x,y = zip(*[parse_result(line) for line in results])
plt.plot(x, y, linewidth='1.5')
plt.legend(files, loc='best')
plt.xlim(100, 30)
| Model Pruning/logs/.ipynb_checkpoints/ResultPlotScript-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/xBrymer/FreeCodeCamp-Projects/blob/master/Machine-Learning/cat_dog_classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Z0NKwmLOFnHz" colab_type="text"
# # Pre-requisites
# + id="W3TQ6NGeFRFA" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, Flatten, Dropout, MaxPooling2D, BatchNormalization
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="nA7B-QWbFrEN" colab_type="text"
# Downloading Data
# + id="g7wrmxdcFVjk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="dc6df50f-dd33-4879-99ec-ad0f9762413c"
URL = 'https://cdn.freecodecamp.org/project-data/cats-and-dogs/cats_and_dogs.zip'
path_to_zip = tf.keras.utils.get_file('cats_and_dogs.zip', origin=URL, extract=True)
PATH = os.path.join(os.path.dirname(path_to_zip), 'cats_and_dogs')
# + id="n_Kei-3nYl09" colab_type="code" colab={}
train_dir = os.path.join(PATH, 'train')
validation_dir = os.path.join(PATH, 'validation')
test_dir = os.path.join(PATH, 'test')
# + id="GbV4HRUpY6vf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="18a9137a-ff6c-4da3-812f-66c7376b0690"
print(os.listdir(test_dir))
# + [markdown] id="v2EmhcOYFw69" colab_type="text"
# Get number of files in each directory. The train and validation directories each have the subdirectories "dogs" and "cats".
# + id="Ntj4L7TaFuh7" colab_type="code" colab={}
total_train = sum([len(files) for r, d, files in os.walk(train_dir)])
total_val = sum([len(files) for r, d, files in os.walk(validation_dir)])
total_test = len(os.listdir(test_dir))
# + [markdown] id="Y49u4rAFXuRH" colab_type="text"
# # Preprocessing the Data
# + [markdown] id="HP-OLfGtGCL0" colab_type="text"
# Setting variables for preprocessing
# + id="5cXsexllGBVs" colab_type="code" colab={}
batch_size = 128
epochs = 15
IMG_HEIGHT = 150
IMG_WIDTH = 150
# + id="Gi7Gq7CaGGgH" colab_type="code" colab={}
train_image_generator = ImageDataGenerator(
rescale = (1/255),
)
validation_image_generator = ImageDataGenerator(
rescale = (1/255),
)
test_image_generator = ImageDataGenerator(
rescale = (1/255),
)
# + id="4bExK-X_JUCO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="8c9a8b10-cb55-4a9a-f8b8-dd554696511f"
train_data_gen = train_image_generator.flow_from_directory(
train_dir,
target_size = (IMG_HEIGHT, IMG_WIDTH),
batch_size = batch_size,
color_mode = 'rgb',
shuffle = True,
class_mode = 'binary'
)
val_data_gen = validation_image_generator.flow_from_directory(
validation_dir,
target_size = (IMG_HEIGHT, IMG_WIDTH),
batch_size = batch_size,
color_mode = 'rgb',
shuffle = True,
class_mode = 'binary'
)
test_data_gen = test_image_generator.flow_from_directory(
PATH,
target_size = (IMG_HEIGHT, IMG_WIDTH),
batch_size = batch_size,
color_mode = 'rgb',
shuffle = False,
classes = ['test'],
)
# + [markdown] id="COYPsUOCjGD7" colab_type="text"
# Visualizing the images
# + id="bkRM2kc2i81X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 846} outputId="c30659ca-d33d-43e0-ca02-da4399ded4ae"
def plotImages(images_arr, probabilities = False):
fig, axes = plt.subplots(len(images_arr), 1, figsize=(5,len(images_arr) * 3))
if probabilities is False:
for img, ax in zip( images_arr, axes):
ax.imshow(img)
ax.axis('off')
else:
for img, probability, ax in zip( images_arr, probabilities, axes):
ax.imshow(img)
ax.axis('off')
if probability > 0.5:
ax.set_title("%.2f" % (probability*100) + "% dog")
else:
ax.set_title("%.2f" % ((1-probability)*100) + "% cat")
plt.show()
sample_training_images, _ = next(train_data_gen)
plotImages(sample_training_images[:5])
# + id="Y8T5mJXujaf_" colab_type="code" colab={}
train_image_generator = ImageDataGenerator(
rescale = (1/255.0),
shear_range = 0.025,
horizontal_flip = True,
zoom_range = [1.5,3.0],
rotation_range = 20,
vertical_flip = True,
brightness_range = [0.5, 1.25]
)
# + id="vjfdWpB_jJ70" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 865} outputId="5e748a3b-b8ce-45a6-c7e2-a3565da52219"
train_data_gen = train_image_generator.flow_from_directory(batch_size=batch_size,
directory=train_dir,
target_size=(IMG_HEIGHT, IMG_WIDTH),
class_mode='binary')
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
# + [markdown] id="qnf7jrB0qMqw" colab_type="text"
# # Training the model
# + [markdown] id="wvsnpj2jr1t3" colab_type="text"
# Vanilla model
# + id="oYwVYtQbqHlj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 599} outputId="a5063e47-5409-4986-c330-686d4eb8bc9e"
model = Sequential()
model.add(Conv2D(32, kernel_size = 3, input_shape = (IMG_WIDTH, IMG_HEIGHT, 3), activation = 'relu'))
model.add(MaxPooling2D(2,2))
model.add(Dropout(0.10))
model.add(Conv2D(64, kernel_size = 3, activation = 'relu'))
model.add(MaxPooling2D(2,2))
model.add(Dropout(0.10))
model.add(Conv2D(128, kernel_size = 3, activation = 'relu'))
model.add(MaxPooling2D(2,2))
model.add(Dropout(0.10))
model.add(Flatten())
model.add(Dense(512, activation = 'relu'))
model.add(Dense(2, activation = 'softmax'))
optimizer = tf.keras.optimizers.Adam(lr = 1e-4, decay = 1e-9 )
model.compile(optimizer = optimizer, loss= 'sparse_categorical_crossentropy', metrics = ['accuracy'])
model.summary()
# + [markdown] id="2x3UedPpsK_o" colab_type="text"
# Training the model
# + id="X_XJrZgBrU2e" colab_type="code" colab={}
history = model.fit(train_data_gen, validation_data=val_data_gen, epochs = 50)
# + id="uauikQykClrD" colab_type="code" colab={}
# !cd '/content/drive/My Drive/External Datasets/Models'
model.save('dogs-vs-cats-fcc-v1.h5')
# + [markdown] id="pB2bCAnYIXZJ" colab_type="text"
# #Evaluating the model
# + id="PeLEvfPpT1OD" colab_type="code" colab={}
from tqdm.notebook import tqdm
# + [markdown] id="c0r8xDHuohLp" colab_type="text"
# Loading the model, if runtime crashes.
# + id="DabxJMtWIiVv" colab_type="code" colab={}
model = tf.keras.models.load_model('dogs-vs-cats-fcc.h5')
# + id="vSJhGukmT9oH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="1a69815d-887d-4bbf-bd71-fae8cd70d6da"
print(os.listdir(test_dir))
# + id="wRI9twfgTzXX" colab_type="code" colab={}
def process_test_images(limit):
test_images = []
for image in tqdm(sorted(os.listdir(test_dir)[:limit])):
if image == '.DS_Store' :
pass
else:
image = tf.keras.preprocessing.image.load_img(
f'{test_dir}/{image}',
color_mode="rgb",
interpolation="bilinear",
target_size=(IMG_HEIGHT, IMG_WIDTH)
)
image = tf.keras.preprocessing.image.img_to_array(image)
image = np.array([image])/255
image.reshape(IMG_HEIGHT, IMG_WIDTH, 3)
test_images.append(image)
return test_images
# + id="ne6YdddnurHo" colab_type="code" colab={}
probability_model = tf.keras.Sequential([
model, tf.keras.layers.Softmax()
])
# + id="WoZ4wb70mLXT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["9671c1945cdc485d80f14649557358a0", "57fbe81affd347b29d8f971d3b113076", "dda0c43276154683a2b615977e431e20", "832fff4a39eb40bd91e9d6c9173eae14", "5d6a78366a47423ca71dd939d81b3c92", "<KEY>", "8faa6baedd704f3b934893802ebebfe2", "22b5a73aaa2946a3ac3e80390fdfb541"]} outputId="6bd2a6f0-1a34-4eb5-ad8b-811c7525522e"
test_images = process_test_images(51)
# + id="PKFox73emlh5" colab_type="code" colab={}
def get_probabilites(images):
predictions = []
for image in images:
img_predict = np.argmax(probability_model(image))
if img_predict ==1:
predictions.append(1)
else:
predictions.append(0)
return predictions
# + id="Y9RMR2YL_jTZ" colab_type="code" colab={}
show_images = np.reshape(test_images, (-1,150,150,3))
# + id="KoKQi0v__vFH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="68bfc0a0-dfe0-4b94-80e3-01a036636ec6"
plt.imshow(show_images[2])
# + id="G-CKXh2GnBmD" colab_type="code" colab={}
probabilities = get_probabilites(test_images)
# + id="oNbcgHFUnFqk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="05795b38-1ba5-4a7e-bb7c-27ce400d3adc"
print(probabilities)
# + [markdown] id="DJcSGmTErjkU" colab_type="text"
# Testing the model
# + id="zNHbP0kErm3G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="d00bd1ab-73ce-4bd3-ec23-5157d702685e"
answers = [1, 0, 0, 1, 0, 0, 0, 0, 1, 1, 0,
1, 0, 1, 0, 1, 1, 0, 1, 1, 0, 0,
1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 1,
1, 0, 1, 1, 1, 1, 0, 1, 0, 1, 1,
0, 0, 0, 0, 0, 0]
correct = 0
for probability, answer in zip(probabilities, answers):
if round(probability) == answer:
correct +=1
percentage_identified = (correct / len(answers))
passed_challenge = percentage_identified > 0.63
print(f"Your model correctly identified {round(percentage_identified, 2)}% of the images of cats and dogs.")
if passed_challenge:
print("You passed the challenge!")
else:
print("You haven't passed yet. Your model should identify at least 63% of the images. Keep trying. You will get it!")
# + [markdown] id="-NYeXAqQrymH" colab_type="text"
#
| Machine-Learning/cat_dog_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="BGhKN00OU-IE"
# ! git clone https://github.com/sz128/slot_filling_and_intent_detection_of_SLU.git
# + id="VGYDhXVHBZBa"
# !wget http://nlp.stanford.edu/data/glove.6B.zip
# !unzip glove.6B.zip
# !ls -lat
# + id="OOX47KpNYGWP" colab={"base_uri": "https://localhost:8080/"} outputId="74d812b2-e6c8-4819-b71d-13fe9b8eafe3"
from torch import nn
import torch
'''
A simple bidirectional lstm for joint slot filling and intent classification
'''
class IntentSlot(nn.Module):
def __init__(self, embedding_size, vocab_size, hidden_size, intent_size, slot_size):
super(IntentSlot, self).__init__()
self.embedding = nn.Embedding(vocab_size, embedding_size, padding_idx=0)
self.lstm = nn.LSTM(input_size=embedding_size,
hidden_size=hidden_size,
num_layers=1,
bidirectional=True)
print(intent_size, slot_size)
self.classifier_slot = nn.Linear(2*hidden_size, slot_size)
self.classifier_intent = nn.Linear(2*hidden_size, intent_size)
def forward(self, x, mask):
x = self.embedding(x)
x, _ = self.lstm(x)
agg = torch.sum(x * mask.unsqueeze(-1), dim=1)
agg = agg / (1e-9 + torch.sum(mask, dim=1, keepdim=True))
slots = self.classifier_slot(x)
intent = self.classifier_intent(agg)
return slots, intent
if __name__ == '__main__':
model = IntentSlot(100, 10, 10, 4, 2)
x = torch.tensor([[1, 4], [5, 6]])
mask = torch.tensor([[1, 1], [1, 1]])
slots, intent = model(x, mask)
print(slots.size(), intent.size())
# + id="h5R_p9p6E8og"
import numpy as np
import random
from torch.nn.utils.rnn import pad_sequence
import torch
'''
create a tokenized dataset using the glove embeddings with a custom iterator
'''
class dataset:
def __init__(self, train_file, val_file, test_file, embedding_dim=100):
self.glove_vocab = self._build_glove_vocab(embedding_dim)
self.w_to_t, self.t_to_w, self.s_to_l, self.l_to_s, self.i_to_l, self.l_to_i \
= {'<PAD>': 0, '<UNK>': 1}, {0: '<PAD>', 1: '<UNK>'}, {'<PAD>': 0}, {0: '<PAD>'} \
, {}, {}
self._create_vocabulary(train_file, embedding_dim)
self._create_vocabulary(val_file, embedding_dim)
self._create_vocabulary(test_file, embedding_dim)
self.intent_size = len(self.i_to_l)
self.slot_size = len(self.s_to_l)
self.embedding_size = embedding_dim
self.vocab_size = len(self.t_to_w)
self.embedding = [np.zeros((1, embedding_dim)), np.random.randn(1, embedding_dim)]
for i in range(2, len(self.t_to_w)):
self.embedding.append(
self.glove_vocab[self.t_to_w[i]]
)
tokenized_train = self._process(train_file)
random.shuffle(tokenized_train)
tokenized_val = self._process(val_file)
tokenized_test = self._process(test_file)
self.splits = {'train': tokenized_train,
'val': tokenized_val,
'test': tokenized_test}
def iterate(self, mode, batch_size=64):
data = self.splits[mode]
idx = np.arange(len(data))
n_chunks = idx.shape[0] // batch_size + 1
for chunk_id, chunk in enumerate(np.array_split(idx, n_chunks)):
sentences = [torch.tensor(data[idx][0]) for idx in chunk]
slots = [torch.tensor(data[idx][1]) for idx in chunk]
intents = [torch.tensor(data[idx][2]).reshape(1) for idx in chunk]
padded_sentences = pad_sequence(sentences, batch_first=True, padding_value=0)
padded_slots = pad_sequence(slots, batch_first=True, padding_value=0)
mask = (padded_slots != 0).float()
intents = torch.cat(intents, 0)
yield padded_sentences, padded_slots, intents, mask
def _process(self, input_file):
with open(input_file, 'rt') as fi:
sentences = fi.read().strip().split('\n')
tokenized = []
for i, sentence in enumerate(sentences):
text, intent = sentence.split(' <=> ')
tokens, slots = [], []
for word_slot in text.split(' '):
word, slot = word_slot.split(":")
if word not in self.w_to_t:
tokens.append(1)
else:
tokens.append(self.w_to_t[word])
slots.append(self.s_to_l[slot])
tokenized.append((tokens, slots, self.i_to_l[intent]))
return tokenized
def _create_vocabulary(self, train_file, embedding_dim):
with open(train_file, 'rt') as fi:
sentences = fi.read().strip().split('\n')
for sentence in sentences:
text, intent = sentence.split(' <=> ')
for idx, word_slot in enumerate(text.split(' ')):
word, slot = word_slot.split(":")
if slot not in self.s_to_l:
x = self.s_to_l[slot] = len(self.s_to_l)
self.l_to_s[x] = slot
if intent not in self.i_to_l:
x = self.i_to_l[intent] = len(self.i_to_l)
self.l_to_i[x] = intent
if word not in self.w_to_t and word in self.glove_vocab:
x = self.w_to_t[word] = len(self.w_to_t)
self.t_to_w[x] = word
def _build_glove_vocab(self, dimension=100):
vocab = {}
with open('glove.6B.{}d.txt'.format(dimension),'rt') as fi:
full_content = fi.read().strip().split('\n')
for line in full_content:
splits = line.split()
word, embedding = splits[0], list(map(float, splits[1:]))
vocab[word] = np.array(embedding)
print('finished reading glove', len(vocab))
return vocab
# if __name__ == '__main__':
# directory = 'slot_filling_and_intent_detection_of_SLU/data/atis-2/'
# d = dataset(directory+'train', directory+'valid', directory+'test')
# for x,y1,y2 in d.iterate('train'):
# print(x.size(), y1.size(), y2.size())
# + id="c7dVSD9vnRMX" colab={"base_uri": "https://localhost:8080/"} outputId="52c934c9-cad4-4e86-88f6-31220f0db0e0"
import copy
def eval(model, mode='val'):
model.eval()
total_slot, total_intent, correct_slot, correct_intent = 0, 0, 0, 0
for x, y_s, y_i, mask in d.iterate(mode):
logits_slots, logits_intent = model(x, mask)
pred_slot = torch.argmax(logits_slots, dim=-1)
pred_intent = torch.argmax(logits_intent, dim=-1)
total_intent += y_i.size(0)
total_slot += torch.sum((y_s != 0).float()).item()
correct_slot += torch.sum(((pred_slot == y_s) & (y_s != 0)).float()).item()
correct_intent += torch.sum((pred_intent == y_i).float()).item()
return float(correct_slot) / total_slot, float(correct_intent) / total_intent
directory = 'slot_filling_and_intent_detection_of_SLU/data/atis-2/'
d = dataset(directory+'train', directory+'valid', directory+'test')
model = IntentSlot(embedding_size=d.embedding_size, vocab_size=d.vocab_size,
hidden_size = 512, intent_size=d.intent_size, slot_size=d.slot_size)
loss_slot = nn.CrossEntropyLoss(ignore_index=0)
loss_intent = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters())
eval_period, num_epochs, steps, best_acc = 20, 10, 0, 0
for epoch in range(num_epochs):
for x, y_s, y_i, mask in d.iterate('train'):
model.train()
optimizer.zero_grad()
logits_slots, logits_intent = model(x, mask)
l = loss_intent(logits_intent, y_i)
l = l + loss_slot(logits_slots.view(-1, d.slot_size), y_s.view(-1))
l.backward()
optimizer.step()
steps += 1
if steps % eval_period == 0:
slot_acc, intent_acc = eval(model, 'val')
print('val accuracy of slot filling is {0:.2f} and accuracy of intent \
classification is {1:.2f}'.format(slot_acc, intent_acc))
if slot_acc > best_acc:
best_model = copy.deepcopy(model)
best_acc = slot_acc
slot_acc, intent_acc = eval(best_model, 'test')
print('test accuracy of slot filling is {0:.2f} and accuracy of intent \
classification is {1:.2f}'.format(slot_acc, intent_acc))
# + [markdown] id="NxRzpZccUTZN"
# # New Section
| ATIS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Notebook for Creating Generator code for Keras and PyTorch
from __future__ import print_function, division
import os
import torch
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
#
# Import base tools
## Note, for mac osx compatability import something from shapely.geometry before importing fiona or geopandas
## https://github.com/Toblerity/Shapely/issues/553 * Import shapely before rasterio or fioana
from shapely import geometry
import rasterio
import random
from cw_tiler import main
from cw_tiler import utils
from cw_tiler import vector_utils
import numpy as np
import os
from tqdm import tqdm
import random
import cv2
import logging
# Setting Certificate Location for Ubuntu/Mac OS locations (Rasterio looks for certs in centos locations)
## TODO implement os check before setting
os.environ['CURL_CA_BUNDLE']='/etc/ssl/certs/ca-certificates.crt'
from cw_nets.tools import util as base_tools
argsdebug=True
logger = logging.getLogger(__name__)
if argsdebug:
logger.setLevel(logging.DEBUG)
else:
logger.setLevel(logging.INFO)
# Create the Handler for logging data to a file
logger_handler = logging.StreamHandler()
# Create a Formatter for formatting the log messages
logger_formatter = logging.Formatter('%(name)s - %(levelname)s - %(message)s')
# Add the Formatter to the Handler
logger_handler.setFormatter(logger_formatter)
# Add the Handler to the Logger
if argsdebug:
logger_handler.setLevel(logging.DEBUG)
else:
logger_handler.setLevel(logging.INFO)
logger.addHandler(logger_handler)
# -
class largeGeoTiff(Dataset):
"""Face Landmarks dataset."""
logger = logging.getLogger(__name__)
logger.setLevel(logging.DEBUG)
def __init__(self, raster_path,
stride_size_meters=150,
cell_size_meters = 200,
tile_size_pixels = 650,
transform=None,
quad_space=False,
sample=False,
testing=True
):
"""
Args:
rasterPath (string): Path to the rasterFile
stride_size_meters (float): sliding window stride size in meters
cell_size_meters (float): sliding window size in meters
tile_size_pixels (float): sliding window pixel dimensions
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.logger = logging.getLogger(__name__)
self.logger.setLevel(logging.DEBUG)
# Create the Handler for logging data to a file
logger_handler = logging.StreamHandler()
# Create a Formatter for formatting the log messages
logger_formatter = logging.Formatter('%(name)s - %(levelname)s - %(message)s')
# Add the Formatter to the Handler
logger_handler.setFormatter(logger_formatter)
# Add the Handler to the Logger
if argsdebug:
logger.setLevel(logging.DEBUG)
else:
logger.setLevel(logging.INFO)
self.logger.addHandler(logger_handler)
self.testing=testing
self.raster_path = raster_path
self.stride_size_meters = stride_size_meters
self.cell_size_meters = cell_size_meters
self.tile_size_pixels = tile_size_pixels
self.transform = transform
rasterBounds, dst_profile = base_tools.get_processing_details(self.raster_path, smallExample=sample)
self.src = rasterio.open(self.raster_path)
# Get Lat, Lon bounds of the Raster (src)
self.wgs_bounds = utils.get_wgs84_bounds(self.src)
# Use Lat, Lon location of Image to get UTM Zone/ UTM projection
self.utm_crs = utils.calculate_UTM_crs(self.wgs_bounds)
# Calculate Raster bounds in UTM coordinates
self.utm_bounds = utils.get_utm_bounds(self.src, self.utm_crs)
self.rasterBounds = rasterBounds
self.cells_list = base_tools.generate_cells_list_dict(rasterBounds,
self.cell_size_meters,
self.stride_size_meters,
self.tile_size_pixels,
quad_space=quad_space
)
self.cells_list = self.cells_list[0]
if self.testing:
with rasterio.open("test.tif", "w", **dst_profile) as dst:
self.cells_list = [window for ij, window in dst.block_windows()]
def __len__(self):
return len(self.cells_list)
def __getitem__(self, idx):
# Get Tile from bounding box
source_Raster=False
if source_Raster:
src_ras = self.raster_path
else:
src_ras = self.src
if self.testing:
sample = src_ras.read(window=self.cells_list[idx])
else:
cell_selection = self.cells_list[idx]
ll_x, ll_y, ur_x, ur_y = cell_selection
tile, mask, window, window_transform = main.tile_utm(src_ras,
ll_x, ll_y, ur_x, ur_y,
indexes=None,
tilesize=self.tile_size_pixels,
nodata=None,
alpha=None,
dst_crs=self.utm_crs)
#except:
# print(cell_selection)
sample = {'tile': tile.astype(np.float),
'mask': mask,
'window': window.toranges(),
'window_transform': window_transform}
if self.transform:
sample = self.transform(sample)
return sample
# +
from pylab import *
from skimage.morphology import watershed
import scipy.ndimage as ndimage
from PIL import Image, ImagePalette
from torch.nn import functional as F
from torchvision.transforms import ToTensor, Normalize, Compose
import torch
import tifffile as tiff
import cv2
import random
from pathlib import Path
img_transform = Compose([
ToTensor(),
Normalize(mean=[0.485, 0.456, 0.406, 0, 0, 0, 0, 0, 0, 0, 0],
std=[0.229, 0.224, 0.225, 1, 1, 1, 1, 1, 1, 1, 1])
])
def pad(img, pad_size=32):
"""
Load image from a given path and pad it on the sides, so that eash side is divisible by 32 (network requirement)
if pad = True:
returns image as numpy.array, tuple with padding in pixels as(x_min_pad, y_min_pad, x_max_pad, y_max_pad)
else:
returns image as numpy.array
"""
if pad_size == 0:
return img
height, width = img.shape[:2]
if height % pad_size == 0:
y_min_pad = 0
y_max_pad = 0
else:
y_pad = pad_size - height % pad_size
y_min_pad = int(y_pad / 2)
y_max_pad = y_pad - y_min_pad
if width % pad_size == 0:
x_min_pad = 0
x_max_pad = 0
else:
x_pad = pad_size - width % pad_size
x_min_pad = int(x_pad / 2)
x_max_pad = x_pad - x_min_pad
img = cv2.copyMakeBorder(img, y_min_pad, y_max_pad, x_min_pad, x_max_pad, cv2.BORDER_REFLECT_101)
return img, (x_min_pad, y_min_pad, x_max_pad, y_max_pad)
def minmax(img):
out = np.zeros_like(img).astype(np.float32)
if img.sum() == 0:
return out
for i in range(img.shape[2]):
c = img[:, :, i].min()
d = img[:, :, i].max()
t = (img[:, :, i] - c) / (d - c)
out[:, :, i] = t
return out.astype(np.float32)
def reform_tile(tile, rollaxis=True):
if rollaxis:
tile = np.rollaxis(tile, 0,3)
rgb = minmax(tile[:,:,(5,3,2)])
tf = tile.astype(np.float32)/ (2**11 - 1)
return np.concatenate([rgb, tf], axis=2) * (2**8 - 1)
def teranaus_transform(sample):
"""sample = {'tile': tile,
'mask': mask,
'window': window,
'window_transform': window_transform}
"""
img = reform_tile(sample['tile'])
img, pads = pad(img)
input_img = torch.unsqueeze(img_transform(img / 255), dim=0)
sample.update({'pad_img': img,
'pads': pads})
return sample
# +
# %%time
stride_size_meter = 150
cell_size_meter = 200
tile_size_pixels = 650
rasterPath = "/home/dlindenbaum/057341085010_01_assembley_MULPan_cog.tif"
#rasterPath = "/nfs/data/Datasets/CosmiQ_SpaceNet_Src/AOI_2_Vegas/srcData/rasterData/AOI_2_Vegas_MUL-PanSharpen_Cloud.tif"
#rasterPath = "s3://spacenet-dataset/AOI_2_Vegas/srcData/rasterData/AOI_2_Vegas_MUL-PanSharpen_Cloud.tif"
#rasterPath = "/home/dlindenbaum/cosmiqGit/cw-nets/cw_nets/cw_generator/RGBA.byte.tif"
spaceNetDatset = largeGeoTiff(rasterPath,
stride_size_meters=stride_size_meter,
cell_size_meters = cell_size_meter,
tile_size_pixels = tile_size_pixels,
transform=teranaus_transform,
sample=True
)
dataloader = DataLoader(spaceNetDatset, batch_size=10,
shuffle=False, num_workers=2)
from tqdm import tqdm
for idx, sample in tqdm(enumerate(dataloader)):
logger.info("Testing idx")
if idx == 10:
break
# -
len(sample)
| cw_nets/cw_generator/cw_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Autoratch/Algorithms/blob/master/verMay23-2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="jWlTS7WDk7cy" outputId="e10afde3-9cc6-4524-d3d1-7647fb0475c4"
from google.colab import drive
drive.mount('/content/drive')
# + id="S-E7wvs-ixAn"
import csv
import math
from datetime import datetime,timedelta
import matplotlib.pyplot as plt
import numpy as np
# default param val
platform = "MT"
forexpair = "EURUSD"
forextype = "M1"
yearmonth = "202204"
datafilepath = "/content/drive/My Drive/Colab Notebooks/forex_historical_data/DAT_"+platform+"_"+forexpair+"_"+forextype+"_"+yearmonth+".csv"
paramfilepath = "/content/drive/My Drive/Colab Notebooks/parameter.csv"
forexfilepath = "/content/drive/My Drive/Colab Notebooks/forex.csv"
bbVal = 20
sdVal = 2
rsiVal = 14
width = 5
rsiUpperLim = 70
rsiLowerLim = 30
martingale = 0
header = ["date","time","open","high","low","close"]
allData = [] # list of data
dayData = [] # list of each day containing list of data
weekData = [] # list of each week containing list of data
weekDataK = [] # list of each week containing list of data with width of K
bbDataK = [] # list of each week containing list of bb data with width of K
rsiDataK = [] # list of each week containing list of rsi data with width of K
dataIdx = [] # idx corresponding to itself
bbDataIdx = [] # idx corresponding to weekDataK
rsiDataIdx = [] # idx corresponding to weekDataK
avaDataIdx = [] # idx of available to indicate signals, combining bb and rsi; corresponding to weekDataK
dataAmount = [] # amount of data in each week
bbAmount = [] # amuount of bb in each week
rsiAmount = [] # amount of rsi in each week
signal = [] # signal for buy or sell; 1 for up, -1 for down
sigSuccess = 0 # no of success signal
sigFail = 0 # no of fail signal
sigAll = 0 # no of all signal
sigPercent = 0
totalDate = 0
totalMin = 0
totalWeek = 0
class eachData:
def __init__(self,date,time,open,high,low,close):
self.date = date
self.time = time
self.open = float(open)
self.high = float(high)
self.low = float(low)
self.close = float(close)
# def print(self):
# print(self.date+" "+self.time)
# print(self.open+" "+self.high+" "+self.low+" "+self.close)
class eachBB:
def __init__(self,middle,upper,lower):
self.middle = middle
self.upper = upper
self.lower = lower
def checkDiff(data1,data2): #check diff between data whether gap exist
format = "%Y.%m.%d"
datestr1 = datetime.strptime(data1.date,format)
datestr2 = datetime.strptime(data2.date,format)
expected = datestr1+timedelta(days=1)
if datestr2<=expected:
return 0 # no jump
else:
return 1 # gap exists
def plot(wk):
#plot data
data = weekDataK[wk]
x = dataIdx[wk]
y = []
for i in data:
y.append(i.open)
plt.plot(x,y) # data open
y = []
for i in data:
y.append(i.close)
plt.plot(x,y) # data close
#plot bb
bb = bbDataK[wk]
x = bbDataIdx[wk]
y = []
for i in x:
y.append(bb[i].middle)
plt.plot(x,y) # bb middle
y = []
for i in x:
y.append(bb[i].upper)
plt.plot(x,y) # bb upper
y = []
for i in x:
y.append(bb[i].lower)
plt.plot(x,y) # bb lower
plt.show()
#plot rsi
rsi = rsiDataK[wk]
x = rsiDataIdx[wk]
y = []
for i in x:
y.append(rsi[i])
plt.plot(x,y) # rsi val
y = []
for i in x:
y.append(rsiUpperLim)
plt.plot(x,y) # rsi upper
y = []
for i in x:
y.append(rsiLowerLim)
plt.plot(x,y) # rsi lower
plt.show()
# read parameter from csv file
file = open(paramfilepath)
csvreader = csv.reader(file)
for x in csvreader:
bbVal = int(x[0])
sdVal = int(x[1])
rsiVal = int(x[2])
width = int(x[3])
rsiUpperLim = int(x[4])
rsiLowerLim = int(x[5])
martingale = int(x[6])
file.close()
# read forex from csv file
file = open(forexfilepath)
csvreader = csv.reader(file)
for x in csvreader:
forexpair = x[0]
yearmonth = x[1]
file.close()
datafilepath = "/content/drive/My Drive/Colab Notebooks/forex_historical_data/DAT_"+platform+"_"+forexpair+"_"+forextype+"_"+yearmonth+".csv"
# read data from csv file
file = open(datafilepath)
csvreader = csv.reader(file)
for min in csvreader:
allData.append(eachData(min[0],min[1],min[2],min[3],min[4],min[5]))
file.close()
# store and organize into days and weeks data list
totalMin = len(allData)
tmp = ""
for min in allData:
if min.date!=tmp:
tmp = min.date
totalDate+=1
for i in range(totalDate):
dayData.append([])
tmp = ""
totalDate = 0
for min in allData:
if min.date!=tmp:
tmp = min.date
totalDate+=1
dayData[totalDate-1].append(min)
# for today in range(totalDate):
# print(dayData[today][0].date+" : "+dayData[today][0].time+"->"+dayData[today][len(dayData[today])-1].time)
first = 1
tmp = []
for i in range(len(allData)):
if first==1 or checkDiff(allData[i-1],allData[i])==0:
tmp.append(allData[i])
first = 0
else:
weekData.append(tmp)
weekDataK.append([])
bbDataK.append([])
rsiDataK.append([])
tmp = []
tmp.append(allData[i])
if tmp!=[]:
weekData.append(tmp)
weekDataK.append([])
rsiDataK.append([])
bbDataK.append([])
# print(len(weekData))
totalWeek = len(weekData)
# for i in range(len(weekData)):
# print(weekData[i][0].date+" "+weekData[i][0].time+" -> "+weekData[i][len(weekData[i])-1].date+" "+weekData[i][len(weekData[i])-1].time)
for i in range(len(weekData)):
for j in range(len(weekData[i])):
if j%width==width-1:
tmp = weekData[i][j]
tmp.open = weekData[i][j-width+1].open
weekDataK[i].append(tmp)
# for i in range(len(weekDataK)):
# for j in range(len(weekDataK[i])):
# tmp = weekDataK[0][j]
# print(tmp.date+" "+tmp.time)
for i in range(totalWeek):
dataAmount.append(len(weekDataK[i]))
bbAmount.append(0)
rsiAmount.append(0)
# calculate bollinger band
for i in range(totalWeek):
each = weekDataK[i]
for j in range(len(each)):
if j>=bbVal-1:
curSum = 0
for k in range(j-bbVal+1,j): # only 19 in front
curSum+=float(each[k].close)
curAvr = curSum/(bbVal-1)
curSD = 0
for k in range(j-bbVal+1,j):
curSD+=(curAvr-float(each[k].close))*(curAvr-float(each[k].close))
curSD/=(bbVal-1)
curSD = math.sqrt(curSD)
curBB = eachBB(curAvr,curAvr+sdVal*curSD,curAvr-sdVal*curSD)
# print(each[k].close,end=" ")
# print(curAvr,end=" ")
# print(curAvr+sdVal*curSD,end=" ")
# print(curAvr-sdVal*curSD)
bbDataK[i].append(curBB)
bbAmount[i]+=1
else:
bbDataK[i].append(eachBB(-1,-1,-1))
# calculate rsi value
for i in range(totalWeek):
each = weekDataK[i]
for j in range(len(each)):
if j>=rsiVal-1:
avrGain = 0
avrLoss = 0
for k in range(j-rsiVal+1,j+1): # all 14 including the current one
change = each[k].close-each[k].open
if change>0:
avrGain+=change
else:
avrLoss-=change
avrGain/=rsiVal
avrLoss/=rsiVal
if avrLoss!=0:
rs = avrGain/avrLoss
rsi = 100-(100/(1+rs))
else:
rsi = 100
rsiDataK[i].append(rsi)
rsiAmount[i]+=1
else:
rsiDataK[i].append(-1)
# create idx of each data bb rsi
for i in range(totalWeek):
data = weekDataK[i]
bb = bbDataK[i]
rsi = rsiDataK[i]
dataIdx.append(range(dataAmount[i]))
bbDataIdx.append(range(dataAmount[i]-bbAmount[i],dataAmount[i]))
rsiDataIdx.append(range(dataAmount[i]-rsiAmount[i],dataAmount[i]))
avaDataIdx.append(range(max(dataAmount[i]-bbAmount[i],dataAmount[i]-rsiAmount[i]),dataAmount[i]))
# plot(i)
# creating and evaluating signal
for i in range(totalWeek):
signal.append([])
for j in range(dataAmount[i]):
signal[i].append(0)
prevBbSig = -5
prevRsiSig = -5
curSideway = 0
curSuccess = 0
for j in avaDataIdx[i]:
if weekDataK[i][j].close>=bbDataK[i][j].upper:
bbSig = -1
elif weekDataK[i][j].close<=bbDataK[i][j].lower:
bbSig = 1
else:
bbSig = 0
if rsiDataK[i][j]>=rsiUpperLim:
rsiSig = -1
elif rsiDataK[i][j]<=rsiLowerLim:
rsiSig = 1
else:
rsiSig = 0
curSig = 0
if bbSig==rsiSig and bbSig!=0:
if bbSig!=prevBbSig or rsiSig!=prevRsiSig:
curSig = bbSig
curSideway = 0
curSuccess = 0
sigAll+=1
else:
curSideway+=1
if curSideway<=martingale and curSuccess==0:
curSig = bbSig
signal[i][j] = curSig
prevBbSig = bbSig
prevRsiSig = rsiSig
if curSig!=0 and j<dataAmount[i]-1:
curStatus = -1
if signal[i][j]==-1 and weekDataK[i][j].close>weekDataK[i][j+1].close:
curStatus = 1
if signal[i][j]==1 and weekDataK[i][j].close<weekDataK[i][j+1].close:
curStatus = 1
if weekDataK[i][j].close==weekDataK[i][j+1].close:
curStatus = 0
if curStatus==0:
curSideway-=1
elif curStatus==1:
curSuccess = 1
sigSuccess+=1
# print(cursig,end = " ")
# print()
sigPercent = sigSuccess/sigAll*100
file = open("/content/drive/My Drive/Colab Notebooks/result.csv","w")
bbP = str(bbVal)
rsiP = str(rsiVal)
sigP = str(sigPercent)
file.write(bbP+","+rsiP+","+sigP)
file.close()
#print(sigPercent)
# + colab={"base_uri": "https://localhost:8080/"} id="uvQ4Hb6WUEpN" outputId="8c4988c4-c8fc-4d9f-fe0d-19da6ffa0199"
bbLo = 10
bbHi = 30
sdVal = 2
rsiLo = 8
rsiHi = 20
width = 5
rsiUpperLim = 70
rsiLowerLim = 30
martingale = 1
def printToFile(bbVal,sdVal,rsiVal,width,rsiUpperLim,rsiLowerLim,martingale):
bbVal = str(bbVal)
sdVal = str(sdVal)
rsiVal = str(rsiVal)
width = str(width)
rsiUpperLim = str(rsiUpperLim)
rsiLowerLim = str(rsiLowerLim)
martingale = str(martingale)
filepath = "/content/drive/My Drive/Colab Notebooks/parameter.csv"
file = open(filepath,"w")
file.write(bbVal+","+sdVal+","+rsiVal+","+width+","+rsiUpperLim+","+rsiLowerLim+","+martingale)
file.close()
file = open("/content/drive/My Drive/Colab Notebooks/forex.csv","w")
file.write("EURJPY,202204")
file.close()
bestSigPercent = -1
for bbVal in range(bbLo,bbHi+1):
for rsiVal in range(rsiLo,rsiHi+1):
printToFile(bbVal,sdVal,rsiVal,width,rsiUpperLim,rsiLowerLim,martingale)
print("BB:",end = "")
print(bbVal,end = " ")
print("RSI:",end = "")
if rsiVal<10:
print(" ",end="")
print(rsiVal,end = "")
print("; percent accuracy : ",end = "")
exec(open("/content/drive/My Drive/Colab Notebooks/run.py").read())
file = open("/content/drive/My Drive/Colab Notebooks/result.csv")
csvreader = csv.reader(file)
for x in csvreader:
sigPercent = float(x[2])
print(sigPercent)
if sigPercent>bestSigPercent:
bestSigPercent = sigPercent
bestBB = bbVal
bestRSI = rsiVal
print()
print("best parameter config is")
print("BB:",end = "")
print(bestBB,end = " ")
print("RSI:",end = "")
if bestRSI<10:
print(" ",end="")
print(bestRSI,end = "")
print("; percent accuracy : ",end = "")
print(bestSigPercent)
| verMay23-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Vaincookie/FundamentalAnalysis/blob/master/Apex_Improved.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="yBeiMvfKGBUu"
import numpy as np
import pandas as pd
from tqdm import trange
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.data as data
train= pd.read_csv('/content/train.csv')
test= pd.read_csv('/content/test.csv')
test_ids= test['id'].to_numpy()
# + id="zm2XO2OWxtBG"
from apex import amp
# + id="SVnZitCrG2Ws"
def preprocess(df):
#some new
dfr= pd.get_dummies(df['R'], prefix= "R_")
df= pd.concat([df, dfr], axis= 1)
dfc= pd.get_dummies(df['C'], prefix= "C_")
df= pd.concat([df, dfc], axis= 1)
df= df.drop(['R', 'C'], axis= 1)
df['u_in_cumsum']= df['u_in'].groupby(df['breath_id']).cumsum()
df['time_step_cumsum']= df['time_step'].groupby(df['breath_id']).cumsum()
df['u_in_min']= df['u_in'].groupby(df['breath_id']).transform('min')
df['u_in_max']= df['u_in'].groupby(df['breath_id']).transform('max')
df['u_in_mean']= df['u_in'].groupby(df['breath_id']).transform('mean')
df['u_in_lag2']= df['u_in'].groupby(df['breath_id']).shift(2)
df['u_in_lag1']= df['u_in'].groupby(df['breath_id']).shift(1)
df['u_in_lag-1']= df['u_in'].groupby(df['breath_id']).shift(-1)
df['u_in_lag-2']= df['u_in'].groupby(df['breath_id']).shift(-2)
df= df.fillna(0)
df['u_in_diff1']= df['u_in']- df['u_in_lag1']
df['u_in_diff2']= df['u_in']- df['u_in_lag2']
df['u_in_diff3']= df['u_in_max']- df['u_in']
df['u_in_diff4']= df['u_in_mean']- df['u_in']
df1= df[df['u_out'] == 0]
df['mean_inspiratory_uin']= df1['u_in'].groupby(df['breath_id']).transform('mean')
df2= df[df['u_out'] == 1]
df['mean_expiratory_uin']= df2['u_in'].groupby(df['breath_id']).transform('mean')
df['u_in_diff5']= df['mean_inspiratory_uin']- df['u_in']
df['u_in_diff6']= df['mean_expiratory_uin']- df['u_in']
df= df.fillna(0)
df['delta_t']= df.groupby('breath_id')['time_step'].diff().fillna(0)
df['delta_uin']= df.groupby('breath_id')['u_in'].diff().fillna(0)
df['area']= df['u_in']*df['delta_t']
df['area']= df.groupby('breath_id')['area'].cumsum()
df['slope']= (df['delta_uin']/df['delta_t']).fillna(0)
return df
# + id="C0R1mjgoG407"
groups= train.breath_id.values.reshape(-1, 80)[:, 0]
groups.shape
train= preprocess(train)
targets= train['pressure'].to_numpy().reshape(-1, 80)
train.drop(['id','pressure', "breath_id"], axis= 1, inplace= True)
test= preprocess(test)
test.drop(['id', "breath_id"], axis= 1, inplace= True)
y_test= np.zeros(test.shape[0]).reshape(-1, 80)
from sklearn.preprocessing import RobustScaler
RS = RobustScaler()
train = RS.fit_transform(train)
test = RS.transform(test)
num_features= train.shape[-1]
train= train.reshape(-1, 80, num_features)
test= test.reshape(-1, 80, num_features)
# + id="fD9C9PU-G6xb"
class CustomDataset:
def __init__(self, data, target):
self.data= data
self.target= target
def __len__(self):
return len(self.data)
def __getitem__(self, idx):
current_sample= self.data[idx, :, :]
current_target= self.target[idx, :]
return torch.tensor(current_sample, dtype= torch.float), torch.tensor(current_target, dtype= torch.float)
# + id="Jsiq1QJ3G9wH"
class RNNModel(nn.Module):
def __init__(self, input_dim, output_dim):
super(RNNModel, self).__init__()
hidden_dim= [400, 300, 200, 100]
self.bilstm1= nn.LSTM(input_dim, hidden_dim[0], batch_first= True, bidirectional= True)
self.norm1= nn.LayerNorm(hidden_dim[0]*2)
self.bilstm2= nn.LSTM(hidden_dim[0]*2, hidden_dim[1], batch_first= True, bidirectional= True)
self.norm2= nn.LayerNorm(hidden_dim[1]*2)
self.bilstm3= nn.LSTM(hidden_dim[1]*2, hidden_dim[2], batch_first= True, bidirectional= True)
self.norm3= nn.LayerNorm(hidden_dim[2]*2)
self.bilstm4= nn.LSTM(hidden_dim[2]*2, hidden_dim[3], batch_first= True, bidirectional= True)
self.norm4= nn.LayerNorm(hidden_dim[3]*2)
self.d= nn.Dropout(p= 0.002)
self.fc1= nn.Linear(hidden_dim[3]*2, 100)
self.fc2= nn.Linear(100, output_dim)
# self.fc3= nn.Linear(32, output_dim)
def forward(self, X):
pred, _= self.bilstm1(X)
pred= self.norm1(pred)
pred, _= self.bilstm2(pred)
pred= self.norm2(pred)
pred, _= self.bilstm3(pred)
pred= self.norm3(pred)
pred, _= self.bilstm4(pred)
pred= self.norm4(pred)
pred= self.d(pred)
pred= self.fc1(pred)
pred= F.selu(pred)
pred= self.fc2(pred)
# pred= F.selu(pred)
# pred= self.fc3(pred)
pred= pred.squeeze(dim= 2)
return pred
# + id="B42ndDxqHABf"
def initialize_parameters(m):
if isinstance(m, nn.LSTM):
nn.init.orthogonal_(m.weight_ih_l0.data, gain= nn.init.calculate_gain('tanh'))
nn.init.orthogonal_(m.weight_hh_l0.data, gain= nn.init.calculate_gain('tanh'))
nn.init.orthogonal_(m.weight_ih_l0_reverse.data, gain= nn.init.calculate_gain('tanh'))
nn.init.orthogonal_(m.weight_hh_l0_reverse.data, gain= nn.init.calculate_gain('tanh'))
nn.init.constant_(m.bias_ih_l0.data, 0)
nn.init.constant_(m.bias_hh_l0.data, 0)
nn.init.constant_(m.bias_ih_l0_reverse.data, 0)
nn.init.constant_(m.bias_hh_l0_reverse.data, 0)
if isinstance(m, nn.Linear):
nn.init.xavier_normal_(m.weight.data)
nn.init.constant_(m.bias.data, 0)
# + id="p2wybfiRHBYz" colab={"base_uri": "https://localhost:8080/"} outputId="0bf2b364-0e7a-4ab9-c861-2c090847b89e"
device= "cuda" if torch.cuda.is_available() else 'cpu'
INPUT_DIM= num_features
OUTPUT_DIM= 1
BATCH_SIZE= 1024
model= RNNModel(input_dim= INPUT_DIM, output_dim= OUTPUT_DIM).to(device)
model.apply(initialize_parameters)
criterion= nn.L1Loss()
criterion.to(device)
optimizer= optim.Adam(model.parameters(), lr= 0.001)
model, optimizer = amp.initialize(model, optimizer, opt_level="O1") # 这里是“欧一”,不是“零一”
scheduler= optim.lr_scheduler.ReduceLROnPlateau(optimizer, factor= 0.5, patience= 10, verbose= True)
# + id="IVaZ-At0HDMr"
def train_model(dataloader, model, criterion, optimizer):
size= len(dataloader.dataset)
model.train()
batches= len(dataloader)
train_loss= 0
for batch_idx, (X, y) in enumerate(dataloader):
X, y= X.to(device), y.to(device)
scores= model(X)
loss= criterion(scores, y)
optimizer.zero_grad()
with amp.scale_loss(loss, optimizer) as scaled_loss:
scaled_loss.backward()
optimizer.step()
loss= loss.item()
train_loss += loss
train_loss_avg= train_loss/batches
print(f"avg. train loss: {train_loss_avg}")
return train_loss_avg
# + id="MFfEFzzFHEwD"
def val_model(dataloader, model, criterion):
size= len(dataloader.dataset)
batches= len(dataloader)
model.eval()
test_loss= 0
with torch.no_grad():
for X, y in (dataloader):
X, y= X.to(device), y.to(device)
scores= model(X)
test_loss += criterion(scores, y)
test_loss /= batches
print(f"avg test loss : {test_loss}")
return test_loss
# + id="3oO8dweLHF_H"
def predict_model(dataloader, model):
model.eval()
y_pred= np.array([])
with torch.no_grad():
for X , y in dataloader:
X, y= X.to(device), y.to(device)
preds= model(X)
preds= preds.flatten().cpu().numpy()
y_pred= np.concatenate((y_pred, preds))
return y_pred
# + colab={"base_uri": "https://localhost:8080/"} id="oVk82GdIHHnn" outputId="3ab69ff7-f29b-4bc3-c176-17344a2cc82f"
from sklearn.model_selection import GroupKFold
kfold= GroupKFold(n_splits= 5)
EPOCHS= 200
cv_scores= []
predictions= np.zeros(test_ids.shape[0])
for fold, (train_idx, val_idx) in enumerate(kfold.split(train, targets, groups= groups)):
X_train, X_val= train[train_idx], train[val_idx]
y_train, y_val= targets[train_idx], targets[val_idx]
train_dataset= CustomDataset(data= X_train, target= y_train)
val_dataset= CustomDataset(data= X_val, target= y_val)
train_loader= data.DataLoader(train_dataset, batch_size= BATCH_SIZE)
val_loader= data.DataLoader(val_dataset, batch_size= BATCH_SIZE)
best_valid_loss= float('inf')
avg_train_losses= []
avg_val_losses= []
for t in trange(EPOCHS):
print(f"Epoch: {t+1}")
train_loss= train_model(train_loader, model, criterion, optimizer)
val_loss= val_model(val_loader, model, criterion)
avg_train_losses.append(train_loss)
avg_val_losses.append(val_loss)
if (val_loss< best_valid_loss):
best_valid_loss= val_loss
ofilename = 'ventilator%d.pth' % fold
torch.save(model.state_dict(), ofilename)
scheduler.step(val_loss)
cv_scores.append(best_valid_loss)
test_dataset= CustomDataset(data= test, target= y_test)
test_loader= data.DataLoader(test_dataset, batch_size= BATCH_SIZE)
model.load_state_dict(torch.load('ventilator%d.pth' % fold, map_location=device))
predictions += (predict_model(test_loader, model))
break
# + id="SxHPfJcfHJTJ"
sub= pd.DataFrame({'id': test_ids, 'pressure': predictions})
sub.to_csv('submission.csv',index = False)
# + id="k8yo_vCCHM6Y"
import matplotlib.pyplot as plt
plt.plot(avg_train_losses)
plt.plot(avg_val_losses)
# + id="fQb_l1wiHOkm"
| Apex_Improved.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kyriell2/linear-algebra/blob/main/coding_act_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="AbVCi59Ku-yx"
# Coding act #3
# + colab={"base_uri": "https://localhost:8080/"} id="Dv73lmVMwjWj" outputId="2c106341-629f-4a7b-cb65-98617c45aedc"
#Python program to inverse and transpose a 3x3 matrix
import numpy as np
A = np.array([[1,-2,3],[4,25,17],[8,9,-10]])
print(A,"\n")
#transposing the matrix
B = np.transpose(A)
print(B)
# + colab={"base_uri": "https://localhost:8080/"} id="sU4VGZzp16kM" outputId="c84f4adb-8a64-4733-e6e5-3e0699416603"
#inversing the matrix
B = np.linalg.inv(A)
print(B)
# + colab={"base_uri": "https://localhost:8080/"} id="5Uu7hRFX2J84" outputId="bbb68f9c-28d5-49be-d2af-a4503a750b58"
#Python program to inverse and transpose a 4x4 matrix
C = np.array([[1,2,3,-4],[4,5,-6,7],[8,9,-10,11],[12,13,14,-15]])
print(C,"\n")
#tansposing the matrix
D = np.transpose(C)
print(D,"\n")
#inversing the matrix
invC = np.linalg.inv(C)
print(invC)
| coding_act_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def table_to_df():
import boto3
dynamodb = boto3.resource('dynamodb', region_name='us-east-1')
table = dynamodb.Table('url_text')
response = table.scan()
data = response['Items']
while 'LastEvaluatedKey' in response:
response = table.scan(ExclusiveStartKey=response['LastEvaluatedKey'])
data.extend(response['Items'])
title_list, text_raw_list, text_clean_list, test_urls, NER_list, time_list = [], [], [], [], [], []
for item in data:
NER_list.append(item['NER'])
title_list.append(item['title'])
text_raw_list.append(item['text_raw'])
text_clean_list.append(item['text_clean'])
time_list.append(item['timestamp'])
test_urls.append(item['url'])
summary = {'url': test_urls, 'title': title_list, 'text_raw': text_raw_list, 'text_clean':text_clean_list, 'NER': NER_list, 'timestamp': time_list}
summary_df = pd.DataFrame(summary)
return summary_df
summary_df = table_to_df()
summary_df.head()
| example/Demo_get_NoSQL_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from autoplotter import run_app
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/ersaurabhverma/autoplotter/master/Data/data.csv')
run_app(df, mode='inline')
run_app(df,mode='external')
| Notebook/Example.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.1
# language: julia
# name: julia-1.1
# ---
# # Unsupervised Learning using Bayesian Mixture Models
# The following tutorial illustrates the use *Turing* for clustering data using a Bayesian mixture model. The aim of this task is to infer a latent grouping (hidden structure) from unlabelled data.
#
# More specifically, we are interested in discovering the grouping illustrated in figure below. This example consists of 2-D data points, i.e. $\boldsymbol{x} = \{x_i\}_{i=1}^N \,, x_i \in \mathcal{R}^2$, which are distributed according to Gaussian distributions. For simplicity, we use isotropic Gaussian distributions but this assumption can easily be relaxed by introducing additional parameters.
# +
using Distributions, StatsPlots, Random
# Set a random seed.
Random.seed!(3)
# Construct 30 data points for each cluster.
N = 30
# Parameters for each cluster, we assume that each cluster is Gaussian distributed in the example.
μs = [-3.5, 0.0]
# Construct the data points.
x = mapreduce(c -> rand(MvNormal([μs[c], μs[c]], 1.), N), hcat, 1:2)
# Visualization.
scatter(x[1,:], x[2,:], legend = false, title = "Synthetic Dataset")
# -
# ## Gaussian Mixture Model in Turing
# To cluster the data points shown above, we use a model that consists of two mixture components (clusters) and assigns each datum to one of the components. The assignment thereof determines the distribution that the data point is generated from.
#
# In particular, in a Bayesian Gaussian mixture model with $1 \leq k \leq K$ components for 1-D data each data point $x_i$ with $1 \leq i \leq N$ is generated according to the following generative process.
# First we draw the parameters for each cluster, i.e. in our example we draw location of the distributions from a Normal:
# $$
# \mu_k \sim Normal() \, , \; \forall k \\
# $$
# and then draw mixing weight for the $K$ clusters from a Dirichlet distribution, i.e.
# $$
# w \sim Dirichlet(K, \alpha) \, . \\
# $$
# After having constructed all the necessary model parameters, we can generate an observation by first selecting one of the clusters and then drawing the datum accordingly, i.e.
# $$
# z_i \sim Categorical(w) \, , \; \forall i \\
# x_i \sim Normal(\mu_{z_i}, 1.) \, , \; \forall i
# $$
#
# For more details on Gaussian mixture models, we refer to <NAME>, *Pattern Recognition and Machine Learning*, Section 9.
# +
using Turing, MCMCChains
# Turn off the progress monitor.
Turing.turnprogress(false)
# -
@model GaussianMixtureModel(x) = begin
D, N = size(x)
# Draw the paramters for cluster 1.
μ1 ~ Normal()
# Draw the paramters for cluster 2.
μ2 ~ Normal()
μ = [μ1, μ2]
# Uncomment the following lines to draw the weights for the K clusters
# from a Dirichlet distribution.
# α = 1.0
# w ~ Dirichlet(2, α)
# Comment out this line if you instead want to draw the weights.
w = [0.5, 0.5]
# Draw assignments for each datum and generate it from a multivariate normal.
k = Vector{Int}(undef, N)
for i in 1:N
k[i] ~ Categorical(w)
x[:,i] ~ MvNormal([μ[k[i]], μ[k[i]]], 1.)
end
return k
end
# After having specified the model in Turing, we can construct the model function and run a MCMC simulation to obtain assignments of the data points.
# Set the automatic differentiation backend to forward differentiation.
# Note, this is temporary while the reverse differentiation functionality
# is being improved.
Turing.setadbackend(:forward_diff)
gmm_model = GaussianMixtureModel(x);
# To draw observations from the posterior distribution, we use a [particle Gibbs](https://www.stats.ox.ac.uk/~doucet/andrieu_doucet_holenstein_PMCMC.pdf) sampler to draw the discrete assignment parameters as well as a Hamiltonion Monte Carlo sampler for continous parameters.
#
# Note that we use a `Gibbs` sampler to combine both samplers for Bayesian inference in our model. We are also calling `mapreduce` to generate multiple chains, particularly so we test for convergence. The `chainscat` function simply adds multiple chains together.
gmm_sampler = Gibbs(100, PG(100, 1, :k), HMC(1, 0.05, 10, :μ1, :μ2))
tchain = mapreduce(c -> sample(gmm_model, gmm_sampler), chainscat, 1:3);
# ## Visualize the Density Region of the Mixture Model
# After sucessfully doing posterior inference, we can first visualize the trace and density of the parameters of interest.
#
# In particular, in this example we consider the sample values of the location parameter for the two clusters.
ids = findall(map(name -> occursin("μ", name), names(tchain)));
p=plot(tchain[:, ids, :], legend=true, labels = ["Mu 1" "Mu 2"], colordim=:parameter)
# You'll note here that it appears the location means are switching between chains. We will address this in future tutorials. For those who are keenly interested, see [this](https://mc-stan.org/users/documentation/case-studies/identifying_mixture_models.html) article on potential solutions.
#
# For the moment, we will just use the first chain to ensure the validity of our inference.
tchain = tchain[:, :, 1];
# As the samples for the location parameter for both clusters are unimodal, we can safely visualize the density region of our model using the average location.
# Helper function used for visualizing the density region.
function predict(x, y, w, μ)
# Use log-sum-exp trick for numeric stability.
return Turing.logaddexp(
log(w[1]) + logpdf(MvNormal([μ[1], μ[1]], 1.), [x, y]),
log(w[2]) + logpdf(MvNormal([μ[2], μ[2]], 1.), [x, y])
)
end
contour(range(-5, stop = 3), range(-6, stop = 2),
(x, y) -> predict(x, y, [0.5, 0.5], [mean(tchain[:μ1].value), mean(tchain[:μ2].value)])
)
scatter!(x[1,:], x[2,:], legend = false, title = "Synthetic Dataset")
# ## Infered Assignments
# Finally, we can inspect the assignments of the data points infered using Turing. As we can see, the dataset is partitioned into two distinct groups.
assignments = collect(skipmissing(mean(tchain[:k].value, dims=1).data))
scatter(x[1,:], x[2,:],
legend = false,
title = "Assignments on Synthetic Dataset",
zcolor = assignments)
| 1_GaussianMixtureModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
% matplotlib inline
import numpy as np
import itertools
import matplotlib.pyplot as plt
# ## Birthday Problem
#
# +
people= int(input())
def calc_Birthday_Probability(people):
# -
# ## Visualizing P and V
def visualizing_permutations():
return list(itertools.permutations(range(0,5)))
print(visualizing_permutations())
print("END OF PERMUTATIONS!")
def visualizing_variations():
return list(itertools.permutations(range(0,11),3))
print(visualizing_variations())
print("END OF VARIATIONS!")
word="emits"
def visualizing_anagrams(word):
return list(itertools.permutations(word))
print(visualizing_anagrams(word))
# ## Shuffling a Deck of Cards
def generate_cards():
suits = ["♥","♦", "♣", "♠"]
faces = list(range(2, 11)) + ["J", "Q", "K", "A"]
return list(map(lambda x: str(x[0]) + x[1],itertools.product(faces, suits)))
print(generate_cards())
def shuffle_cards(cards):
for i in range(51,-1,-1):
j=np.random.randint(0,52)
cards[i]=cards[j]
return cards
print(shuffle_cards(generate_cards()))
# ## Visualizing C
# +
def get_combinations(numbers, k):
return list(itertools.combinations(numbers,k))
numbers = np.random.uniform(-10, 10, 20)
print(numbers)
print(get_combinations(numbers, 4))
fruits = ["apple", "grapes", "banana", "kiwi", "orange",
"apricot", "cherry", "blueberry", "strawberry"]
# +
def print_salad_recipes(fruits):
for i in range(2,len(fruits)+1):
print(list(itertools.combinations(fruits,i)))
print_salad_recipes(fruits)
# -
# ## Random Uniform Numbers
# +
def generate_random_numbers(n):
return np.random.uniform(0, 5, n)
for n in [10, 1000, 10000, 100000]:
plt.figure()
plt.hist(generate_random_numbers(n))
plt.show()
# -
# ## Coin Toss
# +
def Cointossing():
value= np.random.random()
if value<0.5:
return 0
else:
return 1
print(Cointossing())
result=[]
for i in range(10000):
result.append(Cointossing())
plt.title("Fair Coin")
plt.hist(result, bins = 2)
plt.show()
def UnfairCointossing():
value= np.random.random()
if value<0.7:
return 0
else:
return 1
result=[]
for i in range(10000):
result.append(UnfairCointossing())
plt.title("Unfair Coin")
plt.hist(result, bins=2)
plt.show()
# -
# ## Playing Darts
def dartsThrow():
x = np.random.random_integers(0,10)
y = np.random.random_integers(0,10)
distance = np.sqrt(x**2+y**2)
if distance<2:
return 0
elif distance>=2 and distance<4:
return 0.2
elif distance>=4 and distance<6:
return 0.4
elif distance>=6 and distance<8:
return 0.6
elif distance>=8 and distance <10:
return 0.8
elif distance>=10:
return 1
result= []
for i in range(10000):
result.append(dartsThrow())
plt.hist(result, bins = 10)
plt.show()
# ## Binomial Distribution
# ## Poisson Distribution
| 03.Probability Concepts/Probability Concepts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Often, you have data and you need to determine which questions were answered which weren't.
# The computer calls a value that has no data either a null or sometimes not-a-number.
# When you are performing calculations with these missing values, you must be careful about how you handle these.
#
#
# # Masking
#
# # Finding Nulls
#
# # Any and All
# +
data = '''
household,dorm,phone_energy,laptop_energy
A,tuscany,,50
B,sauv,30,
C,tuscany,12,45
D,sauv,,
'''
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from io import StringIO
from tabulate import tabulate
df = pd.read_csv(StringIO(data))
df
# -
# note that you are getting a list of boolean values
# these boolean values can be used as an index to get rows of the data frame
df['phone_energy'].isnull()
# you can look at the entries where the laptop energy values are blank
df[df['laptop_energy'].isnull()]
# you can look at the entries where the laptop energy values are not blank
df[df['laptop_energy'].notnull()]
df['phone_energy'].value_counts()
# you can count how many nulls but you won't know where they are
df['phone_energy'].value_counts(dropna=False)
# lets say you want rows 1 and 3 from the data frame
df.iloc[[1,3]]
# you can get the same thing by passing in a list of False and True values
# where only values corresponding to row 1 and 3 are True
df[[False, True, False, True]]
# you can generate a list of True and False values using functions in Pandas
df['laptop_energy'].isnull()
df['laptop_energy'] > 48
| data-analysis/finding-null-values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import operator
import random
import numpy
from deap import base
from deap import benchmarks
from deap import creator
from deap import tools
creator.create("FitnessMax", base.Fitness, weights=(1.0,))
creator.create("Particle", list, fitness=creator.FitnessMax, speed=list,
smin=None, smax=None, best=None)
def generate(size, pmin, pmax, smin, smax):
part = creator.Particle(random.uniform(pmin, pmax) for _ in range(size))
part.speed = [random.uniform(smin, smax) for _ in range(size)]
part.smin = smin
part.smax = smax
return part
def updateParticle(part, best, phi1, phi2):
u1 = (random.uniform(0, phi1) for _ in range(len(part)))
u2 = (random.uniform(0, phi2) for _ in range(len(part)))
v_u1 = map(operator.mul, u1, map(operator.sub, part.best, part))
v_u2 = map(operator.mul, u2, map(operator.sub, best, part))
part.speed = list(map(operator.add, part.speed, map(operator.add, v_u1, v_u2)))
for i, speed in enumerate(part.speed):
if speed < part.smin:
part.speed[i] = part.smin
elif speed > part.smax:
part.speed[i] = part.smax
part[:] = list(map(operator.add, part, part.speed))
toolbox = base.Toolbox()
toolbox.register("particle", generate, size=2, pmin=-6, pmax=6, smin=-3, smax=3)
toolbox.register("population", tools.initRepeat, list, toolbox.particle)
toolbox.register("update", updateParticle, phi1=2.0, phi2=2.0)
toolbox.register("evaluate", benchmarks.h1)
def main():
pop = toolbox.population(n=5)
stats = tools.Statistics(lambda ind: ind.fitness.values)
stats.register("avg", numpy.mean)
stats.register("std", numpy.std)
stats.register("min", numpy.min)
stats.register("max", numpy.max)
logbook = tools.Logbook()
logbook.header = ["gen", "evals"] + stats.fields
GEN = 1000
best = None
for g in range(GEN):
for part in pop:
part.fitness.values = toolbox.evaluate(part)
if not part.best or part.best.fitness < part.fitness:
part.best = creator.Particle(part)
part.best.fitness.values = part.fitness.values
if not best or best.fitness < part.fitness:
best = creator.Particle(part)
best.fitness.values = part.fitness.values
for part in pop:
toolbox.update(part, best)
# Gather all the fitnesses in one list and print the stats
logbook.record(gen=g, evals=len(pop), **stats.compile(pop))
print(logbook.stream)
return pop, logbook, best
if __name__ == "__main__":
pop, logbook, best = main()
print pop
print logbook
print best
# -
| weighting/PSO_deap_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import zipfile
import tarfile
import os.path
import imageio
import io
import time
import re
import webdataset as wds
# # Reading the ZipFile
#
# We're taking the data out of the original zip file without unpacking.
src = zipfile.ZipFile("fat.zip")
files = [s for s in src.filelist if not s.is_dir()]
bydir = {}
for fname in files:
dir = os.path.dirname(fname.filename)
bydir.setdefault(dir, []).append(fname)
len(bydir.keys())
# # Creating the Tar Files
#
# The Falling Things dataset is a collection of videos of falling things. However, there are different ways in which it can be used to create training samples:
#
# - each frame is a training sample, presented in random order
# - applications: pose estimation, stereo, monocular depth
# - a small sequence of frames makes up a training sample
# - applications: optical flow, motion segmentation, frame interpolatino
# - each full sequence is a training sample (only 330 training samples)
# - applications: physical modeling, long time tracking
#
# For random access datasets, the details of how data is broken up into training samples usually is hidden in the input pipeline. When using large scale, sequential training, this happens in a separate step while we generate training data sets.
#
# Note also that videos in this dataset are not represented as video files but as sequences of frames, distinguished by filename. We are going to do the same thing in the WebDataset representation.
#
# We first generate a full sequence dataset with a special structure of one shard per video:
#
# - one `.tar` file per directory in the ZIP file
# - duplicate the camera and object settings for each sample (so that we can later shuffle)
# - one sample (=basename) per frame
#
# This is a valid WebDataset, but it also still preserves the video data in sequence within each shard, giving us different processing options.
# !rm -rf sequences shuffled
# !mkdir sequences shuffled
def addfile(dst, name, data):
assert isinstance(name, str), type(name)
info = tarfile.TarInfo(name=name)
info.size = len(data)
info.uname, info.gname = "bigdata", "bigdata"
info.mtime = time.time()
dst.addfile(info, fileobj=io.BytesIO(data))
count = 0
for dir in sorted(bydir.keys()):
tname = f"sequences/falling-things-{count:06d}.tar"
nfiles = 0
with tarfile.open(tname, "w|") as dst:
fnames = bydir[dir]
print(count, tname, dir)
camera_settings = [s for s in fnames if "_camera_settings" in s.filename][0]
camera_settings = src.open(camera_settings, "r").read()
object_settings = [s for s in fnames if "_object_settings" in s.filename][0]
object_settings = src.open(object_settings, "r").read()
last_base = "NONE"
for finfo in sorted(fnames, key=lambda x: x.filename):
fname = finfo.filename
if fname.startswith("_"):
continue
base = re.sub(r"\..*$", "", os.path.basename(fname))
if base != last_base:
#print("base:", base)
addfile(dst, dir+"/"+base+".camera.json", camera_settings)
addfile(dst, dir+"/"+base+".object.json", object_settings)
last_base = base
with src.open(fname, "r") as stream:
data = stream.read()
addfile(dst, fname, data)
nfiles += 1
#break
count += 1
# !ls -lth sequences/*.tar | head
# !tar tf sequences/falling-things-000000.tar | head
# # Frame-Level Training
#
# To generate frame level training, we can simply shuffle the sequence data. For this to work, it is important that we associated the sequence level information (camera, object) with each frame, as we did in the construction of the sequence dataset.
# We can now shuffle with:
#
# ```Bash
# $ tarp cat -m 5 -s 500 -o - by-dir/*.tar | tarp split - -o shuffled/temp-%06d.tar
# $ tarp cat -m 10 -s 1000 -o - shuffled/temp-*.tar | tarp split - -o shuffled/falling-things-shuffled-%06d.tar
# ```
#
# If your machine has more memory, you can adjust the `-m` and `-s` options.
# + language="bash"
# tarp cat -m 5 -s 500 -o - sequences/*.tar | tarp split - -o shuffled/temp-%06d.tar &&
# tarp cat -m 10 -s 1000 -o - shuffled/temp-*.tar | tarp split - -o shuffled/falling-things-shuffled-%06d.tar &&
# rm shuffled/temp-*.tar
# -
# # Loading the Frame-Level Data
ds = wds.Dataset("shuffled/falling-things-shuffled-000033.tar").decode()
for sample in ds:
break
sample.keys()
figsize(12, 8)
subplot(221); imshow(sample["left.jpg"])
subplot(222); imshow(sample["right.jpg"])
subplot(223); imshow(sample["left.seg.png"])
subplot(224); imshow(sample["right.seg.png"])
sample["camera.json"]
sample["object.json"]
sample["left.json"]
sample["right.json"]
| notebooks/convert-falling-things.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tuning the parameters of a PID controller
# In this notebook you can test your intuition for how to adjust the parameters of a PID controller.
#
# Start by watching [this excellent video](https://www.youtube.com/watch?v=XfAt6hNV8XM).
#
# ## Blockdiagram
# We consider the model used in the video: Velocity control of a car. The plant model describes how the velocity of the car responds to the position of the accelerator (the gas pedal). In addition to the signal from the accelerator, there are also unknown forces such as wind resistance and gravity when the car is going uphill or downhill. These forces are represented by a disturbance signal entering at the input to the system.
#
# <!---  -->
# <img src="cruise-control-pid-block.svg" alt="Block diagram of cruise control system" width="900">
#
# ## The PID controller
# The PID-controller is on so-called *parallel form*
# \begin{equation}
# F(s) = K_p + \frac{K_i}{s} + K_d s.
# \end{equation}
#
# ## The closed-loop system from the reference signal to the output
# The model is linear and hence the principle of superposition holds. This mean that we can look at the response to the reference signal and the response to the disturbance signal separately. Setting $$d=0,$$ we get a closed-loop response given by
# \begin{equation}
# Y(s) = \frac{\frac{1}{s(s+1)}F(s)}{1 + \frac{1}{s(s+1)}F(s)}R(s).
# \end{equation}
#
# ## The closed-loop system from disturbance to the output
# Setting $$r=0,$$ the reponse to the disturbance is given by
# \begin{equation}
# Y(s) = \frac{\frac{1}{s(s + 1)}}{1 + \frac{1}{s(s+1)}F(s)}D(s)
# \end{equation}
#
# ## The full closed-loop system
# We can find the response to a combination of input signals $r$ and $d$ by summation:
# \begin{equation}
# Y(s) = \frac{\frac{1}{s(s+1)}F(s)}{1 + \frac{1}{s(s+1)}F(s)}R(s) + \frac{\frac{1}{s(s + 1)}}{1 + \frac{1}{s(s+1)}F(s)}D(s)
# \end{equation}
#
#
# Uncomment and run the commands in this cell if a packages is missing
# !pip install slycot
# !pip install control
# %matplotlib widget
import ipywidgets as widgets
import matplotlib.pyplot as plt
import numpy as np
import control.matlab as cm
# ## Step response
# Below you can manipulate the $K_p$, $K_i$ and $K_d$ parameters of the PID-controller, and see a time-response of the system. At time $t_1=1$ there is a unit step in the reference signal, and at time $t_2=10$ yhere is a negative step in the disturbance signal. Note that since we scaled time using the time constant of the system, the time is not measured in seconds but in the length of time constant. So to get $t_2$ in seconds you will have to multiply with the time constant
# \begin{equation}
# t_2 = 5 T = 5 \frac{1}{\omega}
# \end{equation}
# where $\omega$ has unit $1/s$ or $Hz$.
# +
G1 = cm.tf([1.], [1, 1.])
Gint = cm.tf([1], [1, 0])
G = Gint*G1
print(G)
N = 600
t_end = 30
t = np.linspace(0, t_end, N)
# The reference signal
r = np.zeros(N)
r[int(N/t_end):] = 1.0
# The disturbance signal
d = np.zeros(N)
d[int(N/t_end)*10:] = -1.0
# set up plot
fig, ax = plt.subplots(figsize=(8, 3))
#ax.set_ylim([-.1, 4])
ax.grid(True)
def sim_PID(G, Kp, Ki, Kd, r,d,t):
"""
Returns the simulated response of the closed-loop system with a
PID controller.
"""
F = cm.tf([Kd, Kp, Ki], [1.0, 0])
Gr = cm.feedback(G*F,1)
Gd = cm.feedback(G,F)
yr = cm.lsim(Gr, r, t)
yd = cm.lsim(Gd, d, t)
return (yr, yd)
@widgets.interact(Kp=(0, 10, .2), Ki=(0, 8, .2), Kd=(0, 8, .2))
def update(Kp = 1.0, Ki=0, Kd=0):
"""Remove old lines from plot and plot new one"""
[l.remove() for l in ax.lines]
yr, yd = sim_PID(G, Kp, Ki, Kd, r,d,t)
ax.plot(yr[1], yr[0]+yd[0], color='C0')
#ax.plot(yd[1], yd[0], color='C1')
# -
# ### Exercise
# Try to find PID parameters that give
# 1. about 10% overshoot,
# 2. settling time of about 4,
# 3. negligable stationary error at $t=14$ (4 after onset of constant disturbance)
# ## Ramp response
# A negative unit ramp disturbance starts at time $t_1=0$.
# +
# The reference signal
dramp = np.linspace(0, t_end, N)
rr = np.zeros(N)
# set up plot
fig, ax = plt.subplots(figsize=(8, 3))
#ax.set_ylim([-.1, 4])
ax.grid(True)
@widgets.interact(Kp=(0, 10, .2), Ki=(0, 8, .2), Kd=(0, 8, .2))
def update(Kp = 1.0, Ki=1, Kd=0):
"""Remove old lines from plot and plot new one"""
[l.remove() for l in ax.lines]
yr, yd = sim_PID(G, Kp, Ki, Kd, rr,dramp,t)
ax.plot(yr[1], yr[0]+yd[0], color='C0')
#ax.plot(yd[1], yd[0], color='C1')
# -
# ### Exercise
# Why does the error in the ramp response keep growing although the controller contains an integrating term?
# ## Further reading
# - [Documentation for the Python Control Systems library](https://python-control.readthedocs.io/en/0.8.3/)
# - [PID control on Wikipedia](https://en.wikipedia.org/wiki/PID_controller)
#
| introduction/notebooks/Continuous-PID.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Cleaning FERC 1 Fuel data
# This notebook is meant to help get you understand the data cleaning process for FERC Form 1 fuel data for heat content and price per MMBTU data.
import sys
import os
sys.path.append(os.path.abspath(os.path.join('..','..')))
from pudl import pudl, ferc1, eia923, settings, constants
from pudl import models, models_ferc1, models_eia923
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
pudl_engine = pudl.connect_db()
# ## Define a function to clean data frames: What kind of errors are we fixing? Why is this kind of function a reasonable thing to do?
# Looking at some of the distributions of the data below, it becomes clear immediately that there are the data was originally entered in a variety of ways.
#
# Data has been entered in the different units, for example coal heat content values are entered in mmbtu per pound for some records while for most records it is entered in mmbtu per ton. Similarly we see data entered in different orders of magnitude, like gas heat content values entered in mmbtu per cubic foot rather than in mmbtu per thousand cubic foot.
#
# Since these data entry differences are separated by two or more orders of magnitude, we can often clearly delineate between populations. We can identify the primary distribution of values by looking at alternative sources, like the EIA, for the same data set and looking at physical properties of the fuels. For example, the average heat content of a ton of U.S. coal is roughly 20 mmbtu. We can identify the end points of the primary distrubtion in other data sources and apply that range of values to the FERC data to identify the primary distribtuion in the FERC data.
#
# Since data outside of the primary distribution is in most cases easily distinguishable and off by a an order of magnitude (e.g. 2000 for heat content per pound rather than per ton) we can with reasonable confidence use a small set of multipliers to bring outlying populations of data into the primary distribtuion, correcting for errors in how the data was entered.
#
# First let's pull in the applicable data from the PUDL FERC tables:
fuel_df = pd.read_sql('''SELECT * from fuel_ferc1''',pudl_engine)
coal = fuel_df[fuel_df.fuel=='coal']
gas = fuel_df[fuel_df.fuel=='gas']
oil = fuel_df[fuel_df.fuel=='oil']
# Then, we'll define a function that takes as arguments:
#
# - a data series to be cleaned
# - minimum and maximum values bounding the reasonable range of values for the series
# - multipliers (a list) to the minimum and maximum values are divided to define additional ranges of data, which are brought into the reasonable range by multiplying the values in the additional ranges by the multipliers (e.g. if a set of values is 1000 times too large to fit in the reasonable range, its multiplier will be .001).
#
# The function will return a cleaned series.
def fixit(tofix, min, max, mults):
fixed = tofix.copy()
fixed = fixed[fixed > 0]
for mult in mults:
fixed = fixed.apply(lambda x: x*mult if x > min/mult and x < max/mult else x)
fixed = fixed.apply(lambda x: np.nan if x < min or x > max else x)
return(fixed)
# We'll next define a function to show us graphs of what the data looks like before and after it is cleaned, and to show us how much data the cleaning process was not able to clean. This function will take 5 arguments:
#
# - a series of data to be cleaned
# - a series of data which has been cleaned by the cleaning function
# - the title (string) to apply to the before and after graphs
# - the x label (string) to apply to the before and after graphs
# - the y label (string) to apply to the before and after graphs
#
# The function will returned the before and after graphs and the percentage of how many values were not cleaned and were instead set to NaN.
#
# Our showfix function first displays unaltered populations on a log-log scale for two reasons. These populations are separated by orders of magnitude and it would be difficult to show them on the same graph otherwise. These populations are also often vastly different sizes: when one population is has thousands of occurences and the other tens, one is not able to see both without a logarthmic scale.
#
# The showfix function then displays the data once multipliers have been applied to the populations. This population necessarily lies in the same order of magnitude so a linear x axis scale makes sense for view. It's more intuitive for most to view the frequency of occurences on a linear scale as well so a linear y axis is used as well.
def showfix(tofix,fixed,title,xlabel,ylabel):
min_1 = tofix[tofix > 0].min()
max_1 = tofix.max()
fraction_lost = ( fixed.isnull().sum() / len(fixed) ) * 100
show_fraction = 'The percentage of values set to NaN is {} %'\
.format(round(fraction_lost,2))
font = {'family': 'serif',
'color': 'darkred',
'weight': 'normal',
'size': 14,
}
fig_1 = plt.figure(figsize=(16,12))
plt.subplot(2,1,1)
plt.title(title)
plt.xlabel(xlabel, fontsize=12)
plt.ylabel(ylabel, fontsize=12)
plt.grid()
ax = fig_1.axes[0]
ax.set_xscale('log')
ax.set_yscale('log')
plt.xlim(min_1,max_1)
plt.hist(tofix, bins=np.logspace(np.log10(min_1), np.log10(max_1), 100))
plt.subplot(2,1,2)
plt.xlabel(xlabel, fontsize=12)
plt.ylabel(ylabel, fontsize=12)
plt.grid()
plt.hist(fixed.dropna(),bins=100)
plt.text(0, .05, show_fraction, transform=fig_1.transFigure,fontdict=font)
plt.show()
# We'll define a simple graphing function to help us make the graph and a flexible graphing function to help us make slightly more complex graphs later on.
def simplegraph(series,xlabel,ylabel,title,range1,range2):
plt.figure(figsize=(10,7))
plt.xlabel(xlabel, fontsize=12)
plt.ylabel(ylabel, fontsize=12)
plt.title(title)
plt.hist(series,bins=100,range=(range1,range2))
plt.show()
def flexiblegraph(series,xlabel,ylabel,title,yscale,range1,range2,bins):
plt.figure(figsize=(10,7))
plt.xlabel(xlabel, fontsize=12)
plt.ylabel(ylabel, fontsize=12)
plt.title(title)
plt.yscale(yscale)
plt.hist(series,bins=bins,range=(range1,range2))
plt.show()
# ## FERC Cost per unit delivered
# ### FERC Coal cost per unit delivered
# Let's take a look at histograms of FERC Form 1 cost per unit delivered for coal. Nearly all of the fuel units are in tons.
coal.fuel_unit.value_counts()
# We'll start by dropping values below zero.
ferc_coal_cost_delivered = coal[coal.fuel_cost_per_unit_delivered > 0]
ferc_gas_cost_delivered = gas[gas.fuel_cost_per_unit_delivered > 0]
ferc_oil_cost_delivered = oil[oil.fuel_cost_per_unit_delivered > 0]
# The EIA reports that the "average annual sale prices of coal at mines producing each of the four major ranks of coal in 2015, in dollars per short ton (2,000 pounds)were: Bituminous—51.57, Subbituminous—14.63, Lignite—22.36, Anthracite—97.91..." (https://www.eia.gov/energyexplained/index.cfm?page=coal_prices).
#
# "In 2015, the average sales price of coal at the mine was 31.83 per ton, and the average delivered coal price to the electric power sector was 42.58 per ton, resulting in an average transportation cost of 10.75 per ton, or about 25 percent of the total delivered price."
#
# Looking at EIA data for average coal cost by state from 2008 through 2016, one sees a range from 14 dollars to 190 (for coal shipped to Maine in 2012!) dollars per ton (https://drive.google.com/open?id=0B5TMZz2CEFSbcjc5RmVSQUR4LXc). Let's use these bounds to identify the primary distribution for FERC coal cost per unit delivered.
plt.figure(figsize=(14,10))
plt.hist(ferc_coal_cost_delivered.fuel_cost_per_unit_delivered, bins=100,log=True,range=(0,200))
plt.title('FERC Coal cost per unit delivered')
plt.xlabel('Dollars per ton')
plt.ylabel('Number of occurences')
plt.show()
# There are only a handful of values near zero. It appears that these utilities entered the cost per mmbtu rather than the cost per ton. While some have listed mmbtu as the fuel unit, others have not. We could multiply these by the corrected average heat content per unit to get the price per ton but since it's a small population we will not worry about it for now.
simplegraph(ferc_coal_cost_delivered.fuel_cost_per_unit_delivered,'dollars per ton',\
'occurences','FERC coal cost per unit delivered',0,10)
ferc_coal_cost_delivered[ferc_coal_cost_delivered.fuel_cost_per_unit_delivered < 10]
# There are only 4 values above 200. There doesn't appear to be a common theme unifying these records.
ferc_coal_cost_delivered[ferc_coal_cost_delivered.fuel_cost_per_unit_delivered > 200]
# Weighting the cost per unit delivered shows us a more confined range of values. Still, values range out to nearly 200 but the distribution shows two sharper peaks near 30 dollars per ton. Since we don't have quantity delivered values, we weight the cost delivered data by quantity burned.
plt.figure(figsize=(14,10))
plt.hist(ferc_coal_cost_delivered.fuel_cost_per_unit_delivered,\
weights=ferc_coal_cost_delivered.fuel_qty_burned, bins=100,range=(0,200))
plt.title('Coal cost per unit delivered')
plt.xlabel('Dollars per ton')
plt.ylabel('Number of occurences')
plt.show()
# Let's use the cleaning functions to limit the data to the EIA state average values with a minimum of 14 and a maximum of 190.
ferc_delivered_coal = fixit(ferc_coal_cost_delivered.fuel_cost_per_unit_delivered, 14, 190, [1])
showfix(ferc_coal_cost_delivered.fuel_cost_per_unit_delivered,ferc_delivered_coal,'FERC Delivered coal',\
'Dollars per ton','number of occurences')
# ### FERC Gas cost per unit delivered
# Given that there's roughly 1 mmbtu per mcf of natural gas (on average 1.032 mmbtu according to EIA (https://www.eia.gov/tools/faqs/faq.php?id=45&t=8), the distribution for gas costs delivered should be very similar to that of the cost per mmbtu. The bounds for dollars per mmbtu from EIA data were from 1 to 35 dollars so the bounds for mcf delivered should be very similar, if only slightly higher. Let's start with the graphing the primary distribution.
plt.figure(figsize=(14,10))
plt.hist(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered, bins=100,range=(0,150))
plt.title('Gas cost per unit delivered')
plt.xlabel('Dollars per mcf')
plt.ylabel('Number of occurences')
plt.show()
# It looks like there are two tiny outlying populations - one that's 100 times too big and one that's 100 times too small. Some of the records in the population that's 100 times too big appear to be related to a dollars and cents issue - however it is only few records and the population appears overlaps with the primary distribution. Given the overlap, we won't include these values. Both populations are less than a percentage point.
ferc_delivered_gas = fixit(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered, 1, 35, [100])
showfix(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered,ferc_delivered_gas,'FERC Gas cost per unit delivered',\
'Dollars per mcf','occurences')
# Looking at the smaller of the two populations, it does not appear to mirror the main population and it's not clear what would make the population 100 times too small.
plt.figure(figsize=(14,10))
plt.hist(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered, bins=100,range=(0,.75))
plt.title('Gas cost per unit delivered')
plt.xlabel('Dollars per mcf')
plt.ylabel('Number of occurences')
plt.show()
ferc_gas_cost_delivered[(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered > 0) & (ferc_gas_cost_delivered.fuel_cost_per_unit_delivered > 1)]
# There are only a handful of values above 100. It appears that some but not all of these are off by a factor of 100 because the utility entered cents rather than dollars.
plt.figure(figsize=(14,10))
plt.hist(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered, bins=100,range=(100,350))
plt.title('Gas cost per unit delivered')
plt.xlabel('Dollars per mcf')
plt.ylabel('Number of occurences')
plt.show()
ferc_gas_cost_delivered[ferc_gas_cost_delivered.fuel_cost_per_unit_delivered > 100]
# Since we don't have quantity delivered values, we weight the graph above by quantity burned. This distribution also appears to fit nicely with the EIA 923 bounds of 1 and 35.
plt.figure(figsize=(14,10))
plt.hist(ferc_gas_cost_delivered.fuel_cost_per_unit_delivered,\
weights=ferc_gas_cost_delivered.fuel_qty_burned,bins=100,range=(0,40))
plt.title('Gas cost per unit delivered')
plt.xlabel('Dollars per mcf')
plt.ylabel('mcf')
plt.show()
# ### FERC Oil cost per unit delivered
# The West Texas Intermediate price of a barrel of oil has fluctuated between 30 and 145 dollars per barrel over the last decade. In our previous analysis of EIA cost per mmbtu data weighted by quantity delivered, we found that the primary distribution lay between 5 and 33 dollars per mmbtu.
#
# Given that the range of mmbtu per unit in the EIA 923 instructions is 3 to 6.9 mmbtu per barrel of various petroleum products, the widest range one could possibly expect to see cost per barrel delivered would be 15 to 228 dollars. But given that most petroleum-fired power in the U.S. relies on residual fuel oil and distillate fuel oil (https://www.eia.gov/energyexplained/index.cfm?page=electricity_in_the_united_states), a more reasonable cost range rely on an mmbtu per unit range of 5.7 (distillate low end) to 6.9 (residual high end). This would result in a cost per barrel delivered range of 28 to 228 dollars. Let's take a look at the FERC data in that range:
plt.figure(figsize=(14,10))
plt.hist(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered, bins=100,range=(0,250))
plt.title('Oil cost per unit delivered')
plt.xlabel('Dollars per barrel')
plt.ylabel('Number of occurences')
plt.show()
# The primary distribution above appears to begin around 20 and drop off considerably around 160.
simplegraph(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered,'Dollars per barrel','Number of occurences','FERC Oil cost per barrel',0,50)
simplegraph(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered,'Dollars per barrel',\
'Number of occurences','FERC Oil cost per barrel',100,250)
# When weighted by quantity burned, the distribution is much tighter, and as expected given the range in the price of oil above, begins around 30 and drops off considerably before 145.
plt.figure(figsize=(14,10))
plt.hist(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered,weights=ferc_oil_cost_delivered.fuel_qty_burned,\
bins=100,range=(0,200))
plt.title('Oil cost per unit delivered weighted by quantity delivered')
plt.xlabel('Dollars per barrel')
plt.ylabel('Barrels')
plt.show()
ferc_delivered_oil = fixit(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered, 28, 228, [42])
showfix(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered,ferc_delivered_oil,'FERC Oil cost per unit delivered',\
'Dollars per barrel','occurences')
# There is a small population of values near zero, which appears to have been entered in dollars per gallon rather than dollars per barrel. The distribution mirrors the primary distribution in shape and the mmbtu per unit values are also in mmbtu per gallon rather than per barrel.
simplegraph(ferc_oil_cost_delivered.fuel_cost_per_unit_delivered,'Dollars per barrel',\
'Number of occurences','FERC Oil cost per barrel',0,5)
ferc_oil_cost_delivered[ferc_oil_cost_delivered.fuel_cost_per_unit_delivered < 5]
# ## FERC Cost per unit burned
# ### FERC Coal cost per unit burned
# Let's take a look at histograms for FERC cost per unit burned for coal. We can rely on the same range of values used for the cost of fuel delivered. We expect that the values will be very similar to the results for cost per unit delivered.
ferc_coal_cost_burned = coal[coal.fuel_cost_per_unit_burned > 0]
ferc_gas_cost_burned = gas[gas.fuel_cost_per_unit_burned > 0]
ferc_oil_cost_burned = oil[oil.fuel_cost_per_unit_burned > 0]
ferc_coal_burned_fixed = fixit(ferc_coal_cost_burned.fuel_cost_per_unit_burned, 14, 190, [1])
showfix(ferc_coal_cost_burned.fuel_cost_per_unit_burned,ferc_coal_burned_fixed, \
'FERC Coal cost per unit burned','cost per ton','occurences')
# As expected, there is great overlap between the two distributions. Given the lag between fuel delivery and fuel consumption, it is not surprising that they are not precisely the same.
plt.figure(figsize=(14,10))
plt.hist(ferc_delivered_coal,bins=100, range=(0,200),alpha=.5,label='cost of coal delivered')
plt.hist(ferc_coal_burned_fixed,bins=100, range=(0,200),alpha=.5,label='cost of coal burned')
plt.xlabel('Dollars per ton')
plt.ylabel('Occurences')
plt.legend()
plt.show()
# ### FERC Gas cost per unit burned
# Let's take a look at histograms for FERC cost per unit burned for gas. Again, we can rely on the same range of values used for the cost of fuel delivered. We expect that the values will be very similar to the results for cost per unit delivered.
ferc_gas_burned_fixed = fixit(ferc_gas_cost_burned.fuel_cost_per_unit_burned, 1, 35, [1])
showfix(ferc_gas_cost_burned.fuel_cost_per_unit_burned,ferc_gas_burned_fixed, \
'FERC Coal cost per unit burned','cost per ton','occurences')
plt.figure(figsize=(14,10))
plt.hist(ferc_delivered_gas,bins=100, range=(0,50),alpha=.5,label='cost of gas delivered')
plt.hist(ferc_gas_burned_fixed,bins=100, range=(0,50),alpha=.5,label='cost of gas burned')
plt.xlabel('Dollars per mcf')
plt.ylabel('Occurences')
plt.legend()
plt.show()
# Once again, there's considerable overlap between the cost of fuel delivered and the cost of fuel burned.
# ### FERC Oil cost per unit burned
# Let's take a look at histograms for FERC cost per unit burned for oil. Again, we can rely on the same range of values used for the cost of fuel delivered. We expect that the values will be very similar to the results for cost per unit delivered.
simplegraph(ferc_oil_cost_burned.fuel_cost_per_unit_burned,'cost per barrel',\
'occurences','FERC Oil cost per unit burned',0,250)
# There is an outlying population near zero. Upon taking a closer look, these values appear to be off by a factor of 42, similar to the outlying population in the fuel delivered data. Once again, the mmbtu per unit values are also off by a factor of 42.
simplegraph(ferc_oil_cost_burned.fuel_cost_per_unit_burned,'cost per barrel',\
'occurences','FERC Oil cost per unit burned',0,5)
ferc_oil_cost_burned[ferc_oil_cost_burned.fuel_cost_per_unit_burned < 5]
ferc_oil_burned_fixed = fixit(ferc_oil_cost_burned.fuel_cost_per_unit_burned, 28, 228, [42])
showfix(ferc_oil_cost_burned.fuel_cost_per_unit_burned,ferc_oil_burned_fixed, \
'FERC Oil cost per unit burned','cost per barrel','occurences')
plt.figure(figsize=(14,10))
plt.hist(ferc_delivered_oil,bins=100, range=(20,200),alpha=.5,label='cost of oil delivered')
plt.hist(ferc_oil_burned_fixed,bins=100, range=(20,200),alpha=.5,label='cost of oil burned')
plt.xlabel('Dollars per barrel')
plt.ylabel('Occurences')
plt.legend()
plt.show()
# The number of occurences for oil burned and delivered has significant overlap but less so than the other fuel types.
| results/notebooks/ferc1/ferc1_fuel_cleaning/ferc1_fuel_cleaning_cost_delivered_burned.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from numpy import loadtxt
from numpy import unique
data = loadtxt('oil-spill.csv', delimiter=',')
# summarize the number of unique values in each column
for i in range(data.shape[1]):
print(i, len(unique(data[:, i])))
# -
from pandas import read_csv
df = read_csv('oil-spill.csv', header=None)
print(df.nunique())
from pandas import read_csv
# load the dataset
df = read_csv('oil-spill.csv', header=None)
print(df.shape)
# get number of unique values for each column
counts = df.nunique()
# record columns to delete
to_del = [i for i,v in enumerate(counts) if v == 1]
print(to_del)
# drop useless columns
df.drop(to_del, axis=1, inplace=True)
print(df.shape)
# +
# summarize the percentage of unique values for each column using numpy
from numpy import loadtxt
from numpy import unique
data = loadtxt('oil-spill.csv', delimiter=',')
# summarize the number of unique values in each column
for i in range(data.shape[1]):
num = len(unique(data[:, i]))
percentage = float(num) / data.shape[0] * 100
print('%d, %d, %.1f%%' % (i, num, percentage))
# -
# summarize the percentage of unique values for each column using numpy
from numpy import loadtxt
from numpy import unique
# load the dataset
data = loadtxt('oil-spill.csv', delimiter=',')
# summarize the number of unique values in each column
for i in range(data.shape[1]):
num = len(unique(data[:, i]))
percentage = float(num) / data.shape[0] * 100
if percentage < 1:
print('%d, %d, %.1f%%' % (i, num, percentage))
# delete columns where number of unique values is less than 1% of the rows/only implies for numerical values not categorical data
# deleting these columns before looking closely to them doesn't make sense we should be first sure that
# these columns doesn't contain critical information and if not then delete them
from pandas import read_csv
# load the dataset
df = read_csv('oil-spill.csv', header=None)
print(df.shape)
# get number of unique values for each column
counts = df.nunique()
# record columns to delete
to_del = [i for i,v in enumerate(counts) if (float(v)/df.shape[0]*100) < 1]
print(to_del)
# drop useless columns
df.drop(to_del, axis=1, inplace=True)
print(df.shape)
# example of applying the variance threshold for feature selection
from pandas import read_csv
from sklearn.feature_selection import VarianceThreshold
# load the dataset
df = read_csv('oil-spill.csv', header=None)
# split data into inputs and outputs
data = df.values
X = data[:, :-1]
y = data[:, -1]
print(X.shape, y.shape)
# define the transform
transform = VarianceThreshold() # we can pass the threshold into parantheses
# transform the input data
X_sel = transform.fit_transform(X)
print(X_sel.shape)
# +
from numpy import arange
from pandas import read_csv
from sklearn.feature_selection import VarianceThreshold
from matplotlib import pyplot
df = read_csv('oil-spill.csv', header=None)
# split data into inputs and outputs
data = df.values
X = data[:, :-1]
y = data[:, -1]
print(X.shape, y.shape)
# define thresholds to check / here we should check whether our data are numeric or categorical before deleting
thresholds = arange(0.0, 0.55, 0.05)
# apply transform with each threshold
results = list()
for t in thresholds:
# define the transform
transform = VarianceThreshold(threshold=t)
# transform the input data
X_sel = transform.fit_transform(X)
# determine the number of input features
n_features = X_sel.shape[1]
print('>Threshold=%.2f, Features=%d' % (t, n_features))
# store the result
results.append(n_features)
# plot the threshold vs the number of selected features
pyplot.plot(thresholds, results)
pyplot.show()
# -
| Data Cleansing Master/Sparse Column Identification and Removal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://data36.com/scatter-plot-pandas-matplotlib/
import numpy as np
import pandas as pd
# %matplotlib inline
pd.read_csv('zoo.csv', delimiter = ',')
pd.read_csv('pandas_tutorial_read.csv', delimiter=';')
# +
pd.read_csv('pandas_tutorial_read.csv', delimiter=';', names = ['my_datetime', 'event', 'country', 'user_id', 'source', 'topic'])
# -
article_read = pd.read_csv('pandas_tutorial_read.csv', delimiter=';', names = ['my_datetime', 'event', 'country', 'user_id', 'source', 'topic'])
article_read
article_read.tail()
pd.read_csv('zoo.csv', delimiter = ',', names = ['hi', 'hello', 'hey'])
article_read.head()
article_read.sample(20)
article_read[['country', 'user_id']].sample(10)
article_read[article_read.source == 'SEO']
article_read.topic == 'Asia'
article_read[article_read.topic == 'Asia']
article_read[['user_id','country','topic']][article_read.country == 'country_2'].head()
zoo = pd.read_csv('zoo.csv', delimiter = ',')
zoo.count()
zoo[['water_need','uniq_id']].sum()
zoo.water_need.median()
zoo.uniq_id.mean()
zoo.groupby('animal').mean()
zoo.groupby('water_need').mean()
article_read
article_read.groupby('source').count()
article_read[article_read.country == 'country_2'].groupby(['source', 'topic']).count()
zoo_eats = pd.DataFrame([['elephant','vegetables'], ['tiger','meat'], ['kangaroo','vegetables'], ['zebra','vegetables'], ['giraffe','vegetables']], columns=['animal', 'food'])
zoo.merge(zoo_eats)
zoo.merge(zoo_eats, how = 'outer')
zoo.merge(zoo_eats, how = 'left')
zoo.merge(zoo_eats, how = 'left', left_on = 'animal', right_on = 'animal')
zoo.sort_values('animal')
zoo.sort_values(by = ['water_need','animal'], ascending = False)
zoo.sort_values(by = ['water_need'], ascending = False).reset_index()
zoo.sort_values(by = ['water_need'], ascending = False).reset_index(drop = True)
zoo.merge(zoo_eats, how = 'left').fillna('unknown')
# !wget 172.16.31.10/dilan/pandas_tutorial_buy.csv
blog_buy = pd.read_csv('pandas_tutorial_buy.csv', delimiter=';', names = ['my_date_time', 'event', 'user_id', 'amount'])
blog_buy
ans = article_read.merge(blog_buy,how = 'left', left_on = 'user_id', right_on = 'user_id').fillna(0)
ans = ans.groupby('country').sum().amount.sort_values(ascending = False).head(3)
ans
mu = 168 #mean
sigma = 5 #stddev
sample = 250
np.random.seed(0)
height_f = np.random.normal(mu, sigma, sample).astype(int)
mu = 176 #mean
sigma = 6 #stddev
sample = 250
np.random.seed(1)
height_m = np.random.normal(mu, sigma, sample).astype(int)
gym = pd.DataFrame({'height_f': height_f, 'height_m': height_m})
gym.plot()
gym.groupby('height_m').count().plot.bar()
gym.groupby('height_f').count().plot.bar()
gym.hist()
gym.plot.hist(bins=20)
gym.plot.hist(bins=20, alpha=0.7)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(0)
mu = 170 #mean
sigma = 6 #stddev
sample = 100
height = np.random.normal(mu, sigma, sample)
weight = (height-100) * np.random.uniform(0.75, 1.25, 100)
gym = pd.DataFrame({'height': height, 'weight': weight})
gym
gym.plot.scatter(x = 'weight', y = 'height')
x = gym.weight
y = gym.height
plt.scatter(x,y)
| pandas_tutorial_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: turtle
# language: python
# name: turtle
# ---
# # Canvas Drawing Test
# +
from ipycanvas import Canvas
canvas = Canvas(width=200, height=200)
canvas.fill_rect(25, 25, 100, 100)
canvas.clear_rect(45, 45, 60, 60)
canvas.stroke_rect(50, 50, 50, 50)
canvas
# -
| docs/jupyter/canvas-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# * https://github.com/netty/netty
# * https://github.com/waylau/netty-4-user-guide-demos
# * [Netty 实现 Echo 服务器、客户端](https://github.com/waylau/netty-4-user-guide-demos/blob/master/netty4-demos/src/main/java/com/waylau/netty/demo/echo)
# * [基于 WebSocket 的聊天室](https://github.com/waylau/netty-4-user-guide-demos/tree/master/netty4-demos/src/main/java/com/waylau/netty/demo/websocketchat)
# * https://github.com/sanshengshui/netty-learning-example
# * [Netty 实现高性能 IoT 服务器 (Groza) 之精尽代码篇中](https://www.cnblogs.com/sanshengshui/p/9859030.html)
| Reference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import torch
import torch.nn as nn
import torchvision
import torchvision.datasets as datasets
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader, Subset
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
def encdec_accuracy(encoder, decoder, inf_gen, test_steps, device=DEVICE):
correct = 0
num_data = 0
#grab a batch from the test loader
for j in range(test_steps):
examples, labels = next(inf_gen)
embedding = encoder.forward(examples.to(device))
outputs = decoder.forward(embedding.to(device))
#for each output in the batch, check if the label is correct
for i, output in enumerate(outputs):
num_data += 1
max_i = np.argmax(output.detach().cpu().numpy())
if max_i == labels[i]:
correct += 1
acc = float(correct)/num_data
return acc
def inf_loader_generator(dataloader):
'''
Generates a function that infinitely samples a dataloader
'''
while True:
for x, y in dataloader:
yield x, y
# +
class mnist_Classifier(nn.Module):
def __init__(self):
super(mnist_Classifier, self).__init__()
self.conv1 = nn.Conv2d(1, 28, kernel_size=(5,5))
self.conv2 = nn.Conv2d(28, 32, kernel_size=(5,5))
#ok, i undersatnd this now. the output from conv2 is 32 channels. the remaining width and height after
# two 5x5 filters with NO PADDING and STRIDE 1 (defaults) is 20x20. So 32 channels * 20 h * 20 w
self.fc1 = nn.Linear(32*20*20, 16)
self.fc2 = nn.Linear(16, 10)
def forward(self, x):
x = self.conv1(x)
x = nn.ReLU()(x)
x = self.conv2(x)
x = nn.ReLU()(x)
# print(x.size()) --> torch.Size([16, 32, 20, 20])
x = x.view(-1, 32*20*20)
x = self.fc1(x)
x = nn.ReLU()(x)
x = self.fc2(x)
x = nn.ReLU()(x)
return torch.softmax(x,dim=1)
class EncoderCNN(nn.Module):
def __init__(self, input_size=1, hidden_size=28):
super().__init__()
self.hidden_size = hidden_size
self.conv = nn.Conv2d(input_size, hidden_size, kernel_size=(5,5))
self.output_shape = (hidden_size,24,24)
def forward(self, x):
x = self.conv(x)
x = nn.ReLU()(x)
return x
class DecoderCNN(nn.Module):
def __init__(self, hidden_size=28):
super().__init__()
self.hidden_size = hidden_size
self.conv = nn.Conv2d(hidden_size, 32, kernel_size=(5,5))
self.fc1 = nn.Linear(32*20*20, 16)
self.fc2 = nn.Linear(16, 10)
def forward(self, x):
x = self.conv(x)
x = nn.ReLU()(x)
x = x.view(-1, 32*20*20)
x = self.fc1(x)
x = nn.ReLU()(x)
x = self.fc2(x)
x = nn.ReLU()(x)
return torch.softmax(x,dim=1)
class EncoderCNN2(nn.Module):
def __init__(self, input_size=1, hidden_size=28):
super().__init__()
self.hidden_size = hidden_size
self.conv1 = nn.Conv2d(input_size, hidden_size, kernel_size=(5,5))
self.conv2 = nn.Conv2d(hidden_size, 32, kernel_size=(5,5))
self.output_shape = (hidden_size,24,24)
self.fc1 = nn.Linear(32*20*20, 16)
self.fc2 = nn.Linear(16, 10)
def forward(self, x):
x = self.conv1(x)
x = nn.ReLU()(x)
x = self.conv2(x)
x = nn.ReLU()(x)
x = x.view(-1, 32*20*20)
x = self.fc1(x)
x = nn.ReLU()(x)
x = self.fc2(x)
x = nn.ReLU()(x)
return x
class DecoderCNN2(nn.Module):
def __init__(self, hidden_size=28):
super().__init__()
self.hidden_size = hidden_size
def forward(self, x):
return torch.softmax(x,dim=1)
# -
def compute_immediate_sensitivity(model, inp, loss) -> list:
"""Core. Computes immediate sensitivity"""
# (1) first-order gradient (wrt parameters)
first_order_grads = torch.autograd.grad(
loss,
model.parameters(),
retain_graph=True,
create_graph=True,
# allow_unused=True
)
# (2) L2 norm of the gradient from (1)
grad_l2_norm = torch.norm(torch.cat([x.view(-1) for x in first_order_grads]), p=2)
# (3) Gradient (wrt inputs) of the L2 norm of the gradient from (2)
sensitivity_vec = torch.autograd.grad(grad_l2_norm, inp, retain_graph=True)[0]
# (4) L2 norm of (3) - "immediate sensitivity"
# sensitivity = [torch.norm(v, p=2).item() for v in sensitivity_vec]
sensitivity = torch.norm(
sensitivity_vec.view(sensitivity_vec.shape[0], -1), p=2, dim=1
)
return sensitivity
def multiple_data_generators(n, dataset="mnist", folds=50, batch_sizes=16):
if n > folds:
raise ValueError(f"n {n} > {folds} folds")
total = 60000
fold_size = total//folds
index = np.arange(total)
np.random.shuffle(index)
if np.array(batch_sizes).ndim == 0:
batch_sizes = [batch_sizes] * n
train_data_loaders = []
test_data_loaders = []
for i in range(n):
batch_size = batch_sizes[i]
if dataset == "mnist":
mnist_trainset = datasets.MNIST(root='./data', train=True, download=True, transform=torchvision.transforms.ToTensor())
mnist_testset = datasets.MNIST(root='./data', train=False, download=True, transform=torchvision.transforms.ToTensor())
elif dataset == "fashionmnist":
mnist_trainset = datasets.FashionMNIST(root='./data', train=True, download=True, transform=torchvision.transforms.ToTensor())
mnist_testset = datasets.FashionMNIST(root='./data', train=False, download=True, transform=torchvision.transforms.ToTensor())
else:
raise ValueError(f"dataset {dataset}")
mnist_trainset = Subset(mnist_trainset, index[i * fold_size:(i+1) * fold_size])
print(f"data slice [{index[i * fold_size]}, ..., {index[(i+1)* fold_size - 1]}]")
#print(i * fold_size, (i+1) * fold_size)
#print(len(mnist_trainset))
#print(len(mnist_testset))
max_trainsteps = len(mnist_trainset) // batch_size
max_teststeps = len(mnist_testset) // batch_size
#print(max_trainsteps)
#print(max_teststeps)
train_loader = DataLoader(mnist_trainset, batch_size=batch_size, shuffle=True, drop_last=True)
test_loader = DataLoader(mnist_testset, batch_size=batch_size, shuffle=True, drop_last=True)
inf_train = inf_loader_generator(train_loader)
inf_test = inf_loader_generator(test_loader)
iters_per_epoch_train = len(mnist_trainset) // batch_size
iters_per_epoch_test = len(mnist_testset) // batch_size
train_data_loaders.append(inf_train)
test_data_loaders.append(inf_test)
return train_data_loaders, test_data_loaders
class MultipleDecoder:
def __init__(self, n, hidden_size=28, device=DEVICE, alpha=20, epsilon=1e6, first_models=True, separate_encoders=False, separate_decoders=True, dataset="mnist", same_decoder_init=False, unbalanced=False):
"""
Creates n decoders for single encoder
"""
self.device = device
self.n = n
self.hidden_size = hidden_size
self.alpha = alpha
self.epsilon = epsilon
self.dataset = dataset
#if not first_models:
# print("Second Models")
# EncoderCNN = EncoderCNN2
# DecoderCNN = DecoderCNN2
# print(EncoderCNN)
# print(DecoderCNN)
#else:
# EncoderCNN = EncoderCNN
# DecoderCNN = DecoderCNN
self.separate_encoders = bool(separate_encoders)
self.separate_decoders = bool(separate_decoders)
if self.separate_encoders:
self.encoders = []
self.encoder_optimizers = []
for i in range(self.n):
self.encoders.append(EncoderCNN(1, self.hidden_size).to(self.device))
self.encoder_optimizers.append(optim.Adam(self.encoders[i].parameters(),lr=0.001))
else:
self.encoder = EncoderCNN(1, self.hidden_size).to(self.device)
self.encoder_optimizer = optim.Adam(self.encoder.parameters(),lr=0.001)
if not self.separate_decoders:
self.decoder = DecoderCNN(self.hidden_size).to(self.device)
self.decoder_criterion = nn.CrossEntropyLoss()
self.decoder_optimizer = optim.Adam(self.decoder.parameters(),lr=0.001)
else:
self.decoders = []
self.decoder_criterions = []
self.decoder_optimizers = []
if same_decoder_init:
decoder = DecoderCNN(self.hidden_size).to(self.device)
for i in range(self.n):
if i == 0:
self.decoders.append(decoder)
else:
import copy
self.decoders.append(copy.deepcopy(decoder).to(self.device))
self.decoder_criterions.append(nn.CrossEntropyLoss())
self.decoder_optimizers.append(optim.Adam(self.decoders[i].parameters(),lr=0.001))
else:
for i in range(self.n):
self.decoders.append(DecoderCNN(self.hidden_size).to(self.device))
self.decoder_criterions.append(nn.CrossEntropyLoss())
self.decoder_optimizers.append(optim.Adam(self.decoders[i].parameters(),lr=0.001))
if unbalanced:
self.train_data_generators, self.test_data_generators = multiple_data_generators_unbalanced(self.n, batch_sizes=16, dataset=dataset)
else:
self.train_data_generators, self.test_data_generators = multiple_data_generators(self.n, batch_sizes=16, dataset=dataset)
def train(self, epochs=20, train_steps_per_epoch=75, test_steps_per_epoch=50, final_test_steps=625, randomize_order=False, dp=True):
self._train_init(epochs=epochs, train_steps_per_epoch=train_steps_per_epoch, test_steps_per_epoch=test_steps_per_epoch, randomize_order=randomize_order)
import time
start = time.time()
elapsed_test_time = 0.0
while True:
if self.model_steps >= (self.train_steps_per_epoch*self.epochs):
print("Finished Training")
break
if self.model_steps % self.train_steps_per_epoch == 0:
print("Start of Epoch", (self.model_steps // self.train_steps_per_epoch))
self._train_batch(dp=dp)
if self.model_steps % self.train_steps_per_epoch == 0:
train_losses = [round(10000 * np.mean(x))/10000 for x in self.train_losses]
print(f"Average Train Loss:", train_losses)
#for i in range(self.n):
# print(f"Average Train Loss for {i}:", np.mean(self.train_losses[i]))
if test_steps_per_epoch:
start_test = time.time()
for i in range(self.n):
if self.separate_encoders:
encoder = self.encoders[i]
else:
encoder = self.encoder
if self.separate_decoders:
decoder = self.decoders[i]
else:
decoder = self.decoder
self.test_accs[i].append(encdec_accuracy(encoder, decoder, self.test_data_generators[i], self.test_steps_per_epoch, device=self.device))
test_accs = [x[-1] for x in self.test_accs]
print(f"Test Accuracy:", test_accs)
end_test = time.time()
elapsed_test_time += end_test - start_test
#for i in range(self.n):
# print(f"Test Accuracy {i}:", self.test_accs[i][-1])
end = time.time()
elapsed = end - start
self.training_time = elapsed - elapsed_test_time
print(f"Elapsed Test Time:", elapsed_test_time)
print(f"Elapsed Training Time:", self.training_time)
for i in range(self.n):
if self.separate_encoders:
encoder = self.encoders[i]
else:
encoder = self.encoder
if self.separate_decoders:
decoder = self.decoders[i]
else:
decoder = self.decoder
self.test_accs[i].append(encdec_accuracy(encoder, decoder, self.test_data_generators[i], final_test_steps, device=self.device))
test_accs = [x[-1] for x in self.test_accs]
print(f"Final Test Accuracy:", test_accs)
def _train_init(self, epochs=20, train_steps_per_epoch=375, test_steps_per_epoch=100, randomize_order=False):
self.epochs = epochs
self.model_steps = 0
if self.alpha is None or self.epsilon is None:
self.epsilon_iter = None
else:
self.epsilon_iter = self.epsilon / self.epochs
self.train_steps_per_epoch = train_steps_per_epoch
self.test_steps_per_epoch = test_steps_per_epoch
self.randomize_order = randomize_order
# plotting criteria
self.train_losses = [[] for x in range(self.n)]
self.test_accs = [[] for x in range(self.n)]
def _train_batch(self, dp=True):
index = list(range(self.n))
if self.randomize_order:
np.random.shuffle(index)
for i in index:
if dp:
self._train_decoder_batch_dp(i)
else:
self._train_decoder_batch_no_dp(i)
self.model_steps += 1
def _train_decoder_batch_no_dp(self, i):
"""
Train decoder i on the next batch without differential privacy
"""
if self.separate_encoders:
encoder = self.encoders[i]
encoder_optimizer = self.encoder_optimizers[i]
else:
encoder = self.encoder
encoder_optimizer = self.encoder_optimizer
if self.separate_decoders:
decoder = self.decoders[i]
decoder_optimizer = self.decoder_optimizers[i]
decoder_criterion = self.decoder_criterions[i]
else:
decoder = self.decoder
decoder_optimizer = self.decoder_optimizer
decoder_criterion = self.decoder_criterion
x_batch_train, y_batch_train = (x.to(self.device) for x in next(self.train_data_generators[i]))
encoder_optimizer.zero_grad()
decoder_optimizer.zero_grad()
# compute loss
embedding = encoder.forward(x_batch_train)
outputs = decoder.forward(embedding)
loss = decoder_criterion(outputs, y_batch_train)
loss.backward()
self.train_losses[i].append(loss.item())
# perform the a gradient step
encoder_optimizer.step()
decoder_optimizer.step()
def _train_decoder_batch_dp(self, i):
"""
Train decoder i on the next batch using differential privacy
"""
if self.separate_encoders:
encoder = self.encoders[i]
encoder_optimizer = self.encoder_optimizers[i]
else:
encoder = self.encoder
encoder_optimizer = self.encoder_optimizer
if self.separate_decoders:
decoder = self.decoders[i]
decoder_optimizer = self.decoder_optimizers[i]
decoder_criterion = self.decoder_criterions[i]
else:
decoder = self.decoder
decoder_optimizer = self.decoder_optimizer
decoder_criterion = self.decoder_criterion
x_batch_train, y_batch_train = (x.to(self.device) for x in next(self.train_data_generators[i]))
encoder_optimizer.zero_grad()
decoder_optimizer.zero_grad()
# Need grad on input for sensitivity; not sure if cloning is needed
x_batch_train = torch.autograd.Variable(torch.clone(x_batch_train).to(self.device), requires_grad=True)
# compute loss
embedding = encoder.forward(x_batch_train)
outputs = decoder.forward(embedding)
loss = decoder_criterion(outputs, y_batch_train)
batch_sensitivities = compute_immediate_sensitivity(
encoder, x_batch_train, loss
)
batch_sensitivity = torch.max(batch_sensitivities) / len(x_batch_train)
loss.backward()
# step 4. compute noise
# this is the scale of the Gaussian noise to be added to the batch gradient
sigma = torch.sqrt(
(batch_sensitivity ** 2 * self.alpha) / (2 * self.epsilon_iter)
)
# logging
#x = loss.item()
#if np.isnan(x):
# log.debug(
# f"Epoch (before adj): {epoch}; batch_idx: {batch_idx} loss: {loss}"
# )
# x = 0
# log.debug(
# f" epoch (adfter adj): {epoch}; batch_idx: {batch_idx} loss: {loss}"
# )
#train_loss += x
self.train_losses[i].append(loss.item())
# step 5. update gradients with computed sensitivities
with torch.no_grad():
for p in encoder.parameters():
p.grad += sigma * torch.randn(1).to(self.device)
# perform the a gradient step
encoder_optimizer.step()
decoder_optimizer.step()
def fine_tune(self, dp=False, **kwargs):
"""
Used to fine tune individual encoder / decoder pairs AFTER primary collective training
"""
import copy
self.separate_encoders = True
if self.separate_encoders:
self.encoders = []
self.encoder_optimizers = []
for i in range(self.n):
self.encoders.append(copy.deepcopy(self.encoder).to(self.device))
self.encoder_optimizers.append(optim.Adam(self.encoders[i].parameters(),lr=0.001))
self.training_time_dp = self.training_time
self.test_accs_dp = self.test_accs
self.train(dp=dp, **kwargs)
self.training_time += self.training_time_dp
# + jupyter={"outputs_hidden": true}
m = MultipleDecoder(10, epsilon=1e6)
m.train(dp=True, epochs=3)
m.fine_tune(epochs=1)
test_accs = [x[-1] for x in m.test_accs]
training_times = m.training_time
print(f"Average training time", np.mean(training_times))
print(f"Average test accuracy", np.mean(test_accs))
print(f"Median test accuracy", np.median(test_accs))
print(f"Standard deviation test accuracy", np.std(test_accs))
# + jupyter={"outputs_hidden": true}
m = MultipleDecoder(10, epsilon=1e6)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
test_accs = [x[-1] for x in m.test_accs]
training_times = m.training_time
print(f"Average training time", np.mean(training_times))
print(f"Average test accuracy", np.mean(test_accs))
print(f"Median test accuracy", np.median(test_accs))
print(f"Standard deviation test accuracy", np.std(test_accs))
# -
def multiple_data_generators_unbalanced(n, dataset="mnist", folds=50, batch_sizes=16):
if n != 10:
raise NotImplementedError("n must currently be 10")
if batch_sizes != 16:
raise NotImplementedError("batch_sizes must currently be 16")
if folds != 50:
raise NotImplementedError("folds must currently be 50")
if n > folds:
raise ValueError(f"n {n} > {folds} folds")
total = 60000
if np.array(batch_sizes).ndim == 0:
batch_sizes = [batch_sizes] * n
main_total = 660 # primary class
off_total = 60 # other classes
batch_size = 16
if dataset == "mnist":
trainset = datasets.MNIST(root='./data', train=True, download=True, transform=torchvision.transforms.ToTensor())
elif dataset == "fashionmnist":
trainset = datasets.FashionMNIST(root='./data', train=True, download=True, transform=torchvision.transforms.ToTensor())
else:
raise ValueError(f"dataset {dataset}")
dataloader = DataLoader(trainset, batch_size=16, shuffle=False, drop_last=True)
ys = []
for x, y in dataloader:
ys.append(y)
y = np.hstack(ys)
classes = []
n = 10 # number of classes
for i in range(n):
classes.append(np.where(y == i)[0])
np.random.shuffle(classes[i])
indexes = [[] for x in range(n)]
for i in range(n): # for each class
c = classes[i]
start = 0
for j in range(n): # for each organization
index = indexes[j]
if i == j: # main total
index.extend(c[start:start+main_total])
start += main_total
else: # off total
index.extend(c[start:start+off_total])
start += off_total
for i in range(n):
print(np.bincount(y[indexes[i]]))
train_data_loaders = []
test_data_loaders = []
for i in range(n):
batch_size = batch_sizes[i]
if dataset == "mnist":
mnist_trainset = datasets.MNIST(root='./data', train=True, download=True, transform=torchvision.transforms.ToTensor())
mnist_testset = datasets.MNIST(root='./data', train=False, download=True, transform=torchvision.transforms.ToTensor())
elif dataset == "fashionmnist":
mnist_trainset = datasets.FashionMNIST(root='./data', train=True, download=True, transform=torchvision.transforms.ToTensor())
mnist_testset = datasets.FashionMNIST(root='./data', train=False, download=True, transform=torchvision.transforms.ToTensor())
else:
raise ValueError(f"dataset {dataset}")
mnist_trainset = Subset(mnist_trainset, indexes[i])
#print(i * fold_size, (i+1) * fold_size)
print(len(mnist_trainset))
print(len(mnist_testset))
max_trainsteps = len(mnist_trainset) // batch_size
max_teststeps = len(mnist_testset) // batch_size
#print(max_trainsteps)
#print(max_teststeps)
train_loader = DataLoader(mnist_trainset, batch_size=batch_size, shuffle=True, drop_last=True)
test_loader = DataLoader(mnist_testset, batch_size=batch_size, shuffle=True, drop_last=True)
inf_train = inf_loader_generator(train_loader)
inf_test = inf_loader_generator(test_loader)
iters_per_epoch_train = len(mnist_trainset) // batch_size
iters_per_epoch_test = len(mnist_testset) // batch_size
train_data_loaders.append(inf_train)
test_data_loaders.append(inf_test)
return train_data_loaders, test_data_loaders
# +
test_accs = []
training_times = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=True)
m.train(dp=False, epochs=20)
test_accs.extend([x[-1] for x in m.test_accs])
training_times.append(m.training_time)
print(f"Average training time", np.mean(training_times))
print(f"Average test accuracy", np.mean(test_accs))
print(f"Median test accuracy", np.median(test_accs))
print(f"Standard deviation test accuracy", np.std(test_accs))
# +
test_accs_dp2 = []
training_times_dp2 = []
for i in range(10):
m = MultipleDecoder(2, separate_encoders=False)
m.train(dp=True, epochs=10)
m.fine_tune(epochs=10)
test_accs_dp2.extend([x[-1] for x in m.test_accs])
training_times_dp2.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp2))
print(f"Average test accuracy", np.mean(test_accs_dp2))
print(f"Median test accuracy", np.median(test_accs_dp2))
print(f"Standard deviation test accuracy", np.std(test_accs_dp2))
# +
test_accs_dp5 = []
training_times_dp5 = []
for i in range(4):
m = MultipleDecoder(5, separate_encoders=False)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
test_accs_dp5.extend([x[-1] for x in m.test_accs])
training_times_dp5.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp5))
print(f"Average test accuracy", np.mean(test_accs_dp5))
print(f"Median test accuracy", np.median(test_accs_dp5))
print(f"Standard deviation test accuracy", np.std(test_accs_dp5))
# -
test_accs_dp10 = []
training_times_dp10 = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=False)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
# +
fashion_test_accs = []
fashion_training_times = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=True, dataset="fashionmnist")
m.train(dp=False, epochs=20)
fashion_test_accs.extend([x[-1] for x in m.test_accs])
fashion_training_times.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times))
print(f"Average test accuracy", np.mean(fashion_test_accs))
print(f"Median test accuracy", np.median(fashion_test_accs))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs))
# -
fashion_test_accs_dp10 = []
fashion_training_times_dp10 = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=False, dataset="fashionmnist")
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
fashion_test_accs_dp10.extend([x[-1] for x in m.test_accs])
fashion_training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_dp10))
print(f"Average test accuracy", np.mean(fashion_test_accs_dp10))
print(f"Median test accuracy", np.median(fashion_test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_dp10))
fashion_test_accs_dp5 = []
fashion_training_times_dp5 = []
for i in range(4):
m = MultipleDecoder(5, separate_encoders=False, dataset="fashionmnist")
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
fashion_test_accs_dp5.extend([x[-1] for x in m.test_accs])
fashion_training_times_dp5.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_dp5))
print(f"Average test accuracy", np.mean(fashion_test_accs_dp5))
print(f"Median test accuracy", np.median(fashion_test_accs_dp5))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_dp5))
fashion_test_accs_dp2 = []
fashion_training_times_dp2 = []
for i in range(10):
m = MultipleDecoder(2, separate_encoders=False, dataset="fashionmnist")
m.train(dp=True, epochs=10)
m.fine_tune(epochs=10)
fashion_test_accs_dp2.extend([x[-1] for x in m.test_accs])
fashion_training_times_dp2.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_dp2))
print(f"Average test accuracy", np.mean(fashion_test_accs_dp2))
print(f"Median test accuracy", np.median(fashion_test_accs_dp2))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_dp2))
fashion_test_accs_dp10_same = []
fashion_training_times_dp10_same = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=False, dataset="fashionmnist", same_decoder_init=True)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
fashion_test_accs_dp10_same.extend([x[-1] for x in m.test_accs])
fashion_training_times_dp10_same.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_dp10_same))
print(f"Average test accuracy", np.mean(fashion_test_accs_dp10_same))
print(f"Median test accuracy", np.median(fashion_test_accs_dp10_same))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_dp10_same))
# + jupyter={"outputs_hidden": true}
fashion_test_accs_dp5_same = []
fashion_training_times_dp5_same = []
for i in range(4):
m = MultipleDecoder(5, separate_encoders=False, dataset="fashionmnist", same_decoder_init=True)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
fashion_test_accs_dp5_same.extend([x[-1] for x in m.test_accs])
fashion_training_times_dp5_same.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_dp5_same))
print(f"Average test accuracy", np.mean(fashion_test_accs_dp5_same))
print(f"Median test accuracy", np.median(fashion_test_accs_dp5_same))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_dp5_same))
# + jupyter={"outputs_hidden": true}
test_accs_unbalanced = []
training_times_unbalanced = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=True, dataset="mnist", unbalanced=True)
m.train(dp=False, epochs=20)
test_accs_unbalanced.extend([x[-1] for x in m.test_accs])
training_times_unbalanced.append(m.training_time)
print(f"Average training time", np.mean(training_times_unbalanced))
print(f"Average test accuracy", np.mean(test_accs_unbalanced))
print(f"Median test accuracy", np.median(test_accs_unbalanced))
print(f"Standard deviation test accuracy", np.std(test_accs_unbalanced))
# + jupyter={"outputs_hidden": true}
fashion_test_accs_unbalanced = []
fashion_training_times_unbalanced = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=True, dataset="fashionmnist", unbalanced=True)
m.train(dp=False, epochs=20)
fashion_test_accs_unbalanced.extend([x[-1] for x in m.test_accs])
fashion_training_times_unbalanced.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_unbalanced))
print(f"Average test accuracy", np.mean(fashion_test_accs_unbalanced))
print(f"Median test accuracy", np.median(fashion_test_accs_unbalanced))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_unbalanced))
# + jupyter={"outputs_hidden": true}
test_accs_dp_unbalanced = []
training_times_dp_unbalanced = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=False, dataset="mnist", unbalanced=True)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
test_accs_dp_unbalanced.extend([x[-1] for x in m.test_accs])
training_times_dp_unbalanced.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp_unbalanced))
print(f"Average test accuracy", np.mean(test_accs_dp_unbalanced))
print(f"Median test accuracy", np.median(test_accs_dp_unbalanced))
print(f"Standard deviation test accuracy", np.std(test_accs_dp_unbalanced))
# + jupyter={"outputs_hidden": true}
fashion_test_accs_dp_unbalanced = []
fashion_training_times_dp_unbalanced = []
for i in range(2):
m = MultipleDecoder(10, separate_encoders=False, dataset="fashionmnist", unbalanced=True)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
fashion_test_accs_dp_unbalanced.extend([x[-1] for x in m.test_accs])
fashion_training_times_dp_unbalanced.append(m.training_time)
print(f"Average training time", np.mean(fashion_training_times_dp_unbalanced))
print(f"Average test accuracy", np.mean(fashion_test_accs_dp_unbalanced))
print(f"Median test accuracy", np.median(fashion_test_accs_dp_unbalanced))
print(f"Standard deviation test accuracy", np.std(fashion_test_accs_dp_unbalanced))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False)
m.train(dp=True, epochs=20)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# -
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=True)
m.train(dp=True, epochs=15)
m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# + jupyter={"outputs_hidden": true}
test_accs_dp50_model2 = []
training_times_dp50_model2 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=True, first_models=False)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=5)
test_accs_dp50_model2.extend([x[-1] for x in m.test_accs])
training_times_dp50_model2.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50_model2))
print(f"Average test accuracy", np.mean(test_accs_dp50_model2))
print(f"Median test accuracy", np.median(test_accs_dp50_model2))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50_model2))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False, first_models=False)
m.train(dp=False, epochs=5)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False, first_models=False)
m.train(dp=False, epochs=5)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False, first_models=False)
m.train(dp=True, epochs=5)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e5)
m.train(dp=True, epochs=5)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e4)
m.train(dp=True, epochs=5)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# + jupyter={"outputs_hidden": true}
test_accs_dp50 = []
training_times_dp50 = []
for i in range(1):
m = MultipleDecoder(50, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e3)
m.train(dp=True, epochs=5)
#m.fine_tune(epochs=5)
test_accs_dp50.extend([x[-1] for x in m.test_accs])
training_times_dp50.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp50))
print(f"Average test accuracy", np.mean(test_accs_dp50))
print(f"Median test accuracy", np.median(test_accs_dp50))
print(f"Standard deviation test accuracy", np.std(test_accs_dp50))
# -
class EmptyDecoder:
def __init__(self, n, hidden_size=28, device=DEVICE, alpha=20, epsilon=1e6, first_models=True, separate_encoders=False, separate_decoders=False, dataset="mnist", same_decoder_init=False, unbalanced=False):
"""
Creates n decoders for single encoder
"""
self.device = device
self.n = n
self.hidden_size = hidden_size
self.alpha = alpha
self.epsilon = epsilon
self.dataset = dataset
if not first_models:
print("Second Models")
EncoderCNN = EncoderCNN2
DecoderCNN = DecoderCNN2
print(EncoderCNN)
print(DecoderCNN)
else:
EncoderCNN = EncoderCNN2
DecoderCNN = DecoderCNN
self.separate_encoders = bool(separate_encoders)
self.separate_decoders = bool(separate_decoders)
if self.separate_encoders:
self.encoders = []
self.encoder_optimizers = []
for i in range(self.n):
self.encoders.append(EncoderCNN(1, self.hidden_size).to(self.device))
self.encoder_optimizers.append(optim.Adam(self.encoders[i].parameters(),lr=0.001))
else:
self.encoder = EncoderCNN(1, self.hidden_size).to(self.device)
self.encoder_optimizer = optim.Adam(self.encoder.parameters(),lr=0.001)
self.decoder = DecoderCNN(self.hidden_size).to(self.device)
self.decoder_criterion = nn.CrossEntropyLoss()
if unbalanced:
self.train_data_generators, self.test_data_generators = multiple_data_generators_unbalanced(self.n, batch_sizes=16, dataset=dataset)
else:
self.train_data_generators, self.test_data_generators = multiple_data_generators(self.n, batch_sizes=16, dataset=dataset)
def train(self, epochs=20, train_steps_per_epoch=75, test_steps_per_epoch=50, final_test_steps=625, randomize_order=False, dp=True):
self._train_init(epochs=epochs, train_steps_per_epoch=train_steps_per_epoch, test_steps_per_epoch=test_steps_per_epoch, randomize_order=randomize_order)
import time
start = time.time()
elapsed_test_time = 0.0
while True:
if self.model_steps >= (self.train_steps_per_epoch*self.epochs):
print("Finished Training")
break
if self.model_steps % self.train_steps_per_epoch == 0:
print("Start of Epoch", (self.model_steps // self.train_steps_per_epoch))
self._train_batch(dp=dp)
if self.model_steps % self.train_steps_per_epoch == 0:
train_losses = [round(10000 * np.mean(x))/10000 for x in self.train_losses]
print(f"Average Train Loss:", train_losses)
#for i in range(self.n):
# print(f"Average Train Loss for {i}:", np.mean(self.train_losses[i]))
if test_steps_per_epoch:
start_test = time.time()
for i in range(self.n):
if self.separate_encoders:
encoder = self.encoders[i]
else:
encoder = self.encoder
decoder = self.decoder
self.test_accs[i].append(encdec_accuracy(encoder, decoder, self.test_data_generators[i], self.test_steps_per_epoch, device=self.device))
test_accs = [x[-1] for x in self.test_accs]
print(f"Test Accuracy:", test_accs)
end_test = time.time()
elapsed_test_time += end_test - start_test
#for i in range(self.n):
# print(f"Test Accuracy {i}:", self.test_accs[i][-1])
end = time.time()
elapsed = end - start
self.training_time = elapsed - elapsed_test_time
print(f"Elapsed Test Time:", elapsed_test_time)
print(f"Elapsed Training Time:", self.training_time)
for i in range(self.n):
if self.separate_encoders:
encoder = self.encoders[i]
else:
encoder = self.encoder
decoder = self.decoder
self.test_accs[i].append(encdec_accuracy(encoder, decoder, self.test_data_generators[i], final_test_steps, device=self.device))
test_accs = [x[-1] for x in self.test_accs]
print(f"Final Test Accuracy:", test_accs)
def _train_init(self, epochs=20, train_steps_per_epoch=375, test_steps_per_epoch=100, randomize_order=False):
self.epochs = epochs
self.model_steps = 0
if self.alpha is None or self.epsilon is None:
self.epsilon_iter = None
else:
self.epsilon_iter = self.epsilon / self.epochs
self.train_steps_per_epoch = train_steps_per_epoch
self.test_steps_per_epoch = test_steps_per_epoch
self.randomize_order = randomize_order
# plotting criteria
self.train_losses = [[] for x in range(self.n)]
self.test_accs = [[] for x in range(self.n)]
def _train_batch(self, dp=True):
index = list(range(self.n))
if self.randomize_order:
np.random.shuffle(index)
for i in index:
if dp:
self._train_decoder_batch_dp(i)
else:
self._train_decoder_batch_no_dp(i)
self.model_steps += 1
def _train_decoder_batch_no_dp(self, i):
"""
Train decoder i on the next batch without differential privacy
"""
if self.separate_encoders:
encoder = self.encoders[i]
encoder_optimizer = self.encoder_optimizers[i]
else:
encoder = self.encoder
encoder_optimizer = self.encoder_optimizer
decoder = self.decoder
decoder_criterion = self.decoder_criterion
x_batch_train, y_batch_train = (x.to(self.device) for x in next(self.train_data_generators[i]))
encoder_optimizer.zero_grad()
# compute loss
embedding = encoder.forward(x_batch_train)
outputs = decoder.forward(embedding)
loss = decoder_criterion(outputs, y_batch_train)
loss.backward()
self.train_losses[i].append(loss.item())
# perform the a gradient step
encoder_optimizer.step()
def _train_decoder_batch_dp(self, i):
"""
Train decoder i on the next batch using differential privacy
"""
if self.separate_encoders:
encoder = self.encoders[i]
encoder_optimizer = self.encoder_optimizers[i]
else:
encoder = self.encoder
encoder_optimizer = self.encoder_optimizer
decoder = self.decoder
decoder_criterion = self.decoder_criterion
x_batch_train, y_batch_train = (x.to(self.device) for x in next(self.train_data_generators[i]))
encoder_optimizer.zero_grad()
# Need grad on input for sensitivity; not sure if cloning is needed
x_batch_train = torch.autograd.Variable(torch.clone(x_batch_train).to(self.device), requires_grad=True)
# compute loss
embedding = encoder.forward(x_batch_train)
outputs = decoder.forward(embedding)
loss = decoder_criterion(outputs, y_batch_train)
batch_sensitivities = compute_immediate_sensitivity(
encoder, x_batch_train, loss
)
batch_sensitivity = torch.max(batch_sensitivities) / len(x_batch_train)
loss.backward()
# step 4. compute noise
# this is the scale of the Gaussian noise to be added to the batch gradient
sigma = torch.sqrt(
(batch_sensitivity ** 2 * self.alpha) / (2 * self.epsilon_iter)
)
# logging
#x = loss.item()
#if np.isnan(x):
# log.debug(
# f"Epoch (before adj): {epoch}; batch_idx: {batch_idx} loss: {loss}"
# )
# x = 0
# log.debug(
# f" epoch (adfter adj): {epoch}; batch_idx: {batch_idx} loss: {loss}"
# )
#train_loss += x
self.train_losses[i].append(loss.item())
# step 5. update gradients with computed sensitivities
with torch.no_grad():
for p in encoder.parameters():
p.grad += sigma * torch.randn(1).to(self.device)
# perform the a gradient step
encoder_optimizer.step()
def fine_tune(self, dp=False, **kwargs):
"""
Used to fine tune individual encoder / decoder pairs AFTER primary collective training
"""
import copy
self.separate_encoders = True
if self.separate_encoders:
self.encoders = []
self.encoder_optimizers = []
for i in range(self.n):
self.encoders.append(copy.deepcopy(self.encoder).to(self.device))
self.encoder_optimizers.append(optim.Adam(self.encoders[i].parameters(),lr=0.001))
self.training_time_dp = self.training_time
self.test_accs_dp = self.test_accs
self.train(dp=dp, **kwargs)
self.training_time += self.training_time_dp
test_accs_dp10 = []
training_times_dp10 = []
for i in range(1):
m = EmptyDecoder(10, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e6)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=5)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
test_accs_dp10 = []
training_times_dp10 = []
for i in range(1):
m = EmptyDecoder(10, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=2e5)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=5)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
test_accs_dp10 = []
training_times_dp10 = []
for i in range(1):
m = EmptyDecoder(10, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e5)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=2)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
test_accs_dp10 = []
training_times_dp10 = []
for i in range(1):
m = EmptyDecoder(10, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e4)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=5)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
test_accs_dp10 = []
training_times_dp10 = []
for i in range(1):
m = EmptyDecoder(10, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e3)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=5)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
test_accs_dp10 = []
training_times_dp10 = []
for i in range(1):
m = EmptyDecoder(10, separate_encoders=False, separate_decoders=False, first_models=False, epsilon=1e2)
m.train(dp=True, epochs=10)
#m.fine_tune(epochs=5)
test_accs_dp10.extend([x[-1] for x in m.test_accs])
training_times_dp10.append(m.training_time)
print(f"Average training time", np.mean(training_times_dp10))
print(f"Average test accuracy", np.mean(test_accs_dp10))
print(f"Median test accuracy", np.median(test_accs_dp10))
print(f"Standard deviation test accuracy", np.std(test_accs_dp10))
| experiments/asynch/SiteVisitAsynchronous.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Science Case #5: MAPS: an ALMA large program
#
# The [Molecules with ALMA at Planet-forming Scales (MAPS)](http://alma-maps.info) survey, also known as 2018.1.01055.L
#
#
# See also
# https://almascience.eso.org/alma-data/lp/MAPS
#
# Data in **2018.1.01055.L** is about 2TB
# +
import pandas as pd
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('display.width', None)
pd.set_option('display.max_colwidth',25)
from astroquery.admit import ADMIT
# -
# For now a private database, just for MAPS
a = ADMIT('../query/admit_maps.db')
# a.check()
p = a.query(nchan='>0')
s = p['target_name'].unique()
print(len(p),'line cubes', len(s),' unique sources: ',s)
p = a.query(nchan='>2',mom0flux='>0')
print(len(p))
f=p['formula'].unique()
print(len(f),'unique lines: ',f)
# Note that their website reports 19 lines
#
# 13CN **13CO** C17O **C18O** C2H c-C3H2 CH3CN CN **CO** CS **DCN** H13CN H13CO+ **H2CO** HC15N **HC3N** **HCN** **HCO+** N2D+
#
#
# Should list VLSR used....
#
# ### Show me where CO was found
p1 = a.query(formula="CO")
len(p1)
p1
# great, CO was in 10 observations. Looks like in all sources, but in MWC three times, and GM_Aur only once.
#
# Now, are there any that have another line in the cube where CO was found?
p2 = a.query(formula="CO",nlines='>2')
len(p2)
# That's a no, too bad.
#
# How about any isotopologues of CO: 13CO and C18O in the sources when CO was found, and how much emission?
#
s1 = p1['target_name'].unique()
print(s1)
ci=['obs_id','target_name','spw','s_resolution', 'mom0flux']
print('Sources:',s)
for si in s1:
pi = a.query(source_name_alma=si,formula="CO")
pj = a.query(source_name_alma=si,formula="13CO")
pk = a.query(source_name_alma=si,formula="C18O")
print("CO=======",si)
print(pi[ci])
print("13CO=======",si)
if len(pj)>0:
print(pj[ci])
print("C18O=======",si)
if len(pk)>0:
print(pk[ci])
print("=========")
| notebooks/Case5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Built-in libraries
from datetime import datetime, timedelta
import math
# NumPy, SciPy and Pandas
import pandas as pd
import numpy as np
# Scikit-Learn
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.preprocessing import PolynomialFeatures
from sklearn import linear_model
# -
"""
Function that calculates a load profile curve, for each building on a dataframe, based on the specified function.
Currently the following functions are supported:
- Average
- Median
And the currently resolution:
- Daily
The name parameter allow us to save the resulting csv with a more comprehensive title
"""
def doAggregation(datasetName, context, function, resolution='day'):
dataframe = pd.read_csv('../data/processed/{}_{}_dataset.csv'.format(datasetName, context), parse_dates=True,
infer_datetime_format=True, index_col=0)
df_load_curves = pd.DataFrame() # dataframe that will hold all load curves
# resample based on parameter
if (resolution == 'day'):
availableSamples = (dataframe.resample('1D').asfreq()).index # get list of timestamps group by day
delta = 23 # timedelta based on resample
else:
print("Please choose a valid resolution")
exit()
# iterate through all buildings (column)
for column in range(len(dataframe.columns)):
df_sampledReadings = pd.DataFrame() # dataframe to hold new samples for a column
currentColumn = pd.DataFrame(dataframe.iloc[:, column])
# iterate through each day
for timestamp in availableSamples:
# update time limits to the current date
start = timestamp
end = timestamp + timedelta(hours=delta)
# get meter data from only this resolution
df_reading = currentColumn[(currentColumn.index >= start) & (currentColumn.index <= end)]
# ignore index since they are unique timestamps
df_reading.reset_index(drop=True, inplace=True)
# append new sample as columns
df_sampledReadings = pd.concat([df_sampledReadings, df_reading], axis=1)
# make sure sure there are no columns with NaN values
df_sampledReadings.dropna(axis=1, how='all', inplace=True)
df_sampledReadings = df_sampledReadings.T # transpose it so it's easier to see and operate
# up to this point, the matrix above has the shape nxm where is the number of instances and m is the number of readings
# if any NaN prevailed
df_sampledReadings.fillna(value=0, inplace=True)
# calculate load curve based on function
if function == 'average':
load_curve = np.mean(df_sampledReadings, axis = 0)
elif function =='median':
load_curve = np.median(df_sampledReadings, axis = 0)
else:
print("Please choose a valid context")
exit()
# turn into one column dataframe for easier manipulation
load_curve = pd.DataFrame(load_curve)
# keep the instance name as column name
instance_name = []
instance_name.append(df_sampledReadings.index[0])
load_curve.columns = instance_name
# append current load curve to dataframe
df_load_curves = pd.concat([df_load_curves, load_curve], axis=1)
# end of for loop for one column
# replace NaN's with 0
df_load_curves = df_load_curves.replace(0.0, np.nan)
# drop rows with all nan values
df_load_curves = df_load_curves.dropna(axis=1, how='all')
# particular to the DGS dataset
if datasetName =='DGS':
# drop columns with more than 4 nan values (seems to be a sweet spot)
df_load_curves = df_load_curves.dropna(thresh=len(df_load_curves) - 1, axis=1)
df_load_curves.fillna(value=0, inplace=True)
df_load_curves = df_load_curves.T # rotate the final dataframe
# save the file and return the dataframe
df_load_curves.to_csv("../data/processed/{}_{}_{}_dataset.csv".format(datasetName, context, function))
return df_load_curves
df_BDG_weekday_average = doAggregation('BDG', 'weekday', 'average')
df_BDG_weekday_median = doAggregation('BDG', 'weekday', 'median')
df_BDG_weekend_average = doAggregation('BDG', 'weekend', 'average')
df_BDG_weekend_median = doAggregation('BDG', 'weekend', 'median')
df_BDG_fullweek_average = doAggregation('BDG', 'fullweek', 'average')
df_BDG_fullweek_median = doAggregation('BDG', 'fullweek', 'median')
df_DGS_weekday_average = doAggregation('DGS', 'weekday', 'average')
df_DGS_weekday_median = doAggregation('DGS', 'weekday', 'median')
df_DGS_weekend_average = doAggregation('DGS', 'weekend', 'average')
df_DGS_weekend_median = doAggregation('DGS', 'weekend', 'median')
df_DGS_fullweek_average = doAggregation('DGS', 'fullweek', 'average')
df_DGS_fullweek_median = doAggregation('DGS', 'fullweek', 'median')
| Preprocessing/.ipynb_checkpoints/load_cuve_generation-checkpoint.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # secret_santa
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/secret_santa.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/secret_santa.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Secret Santa problem in Google CP Solver.
From Ruby Quiz Secret Santa
http://www.rubyquiz.com/quiz2.html
'''
Honoring a long standing tradition started by my wife's dad, my friends
all play a Secret Santa game around Christmas time. We draw names and
spend a week sneaking that person gifts and clues to our identity. On the
last night of the game, we get together, have dinner, share stories, and,
most importantly, try to guess who our Secret Santa was. It's a crazily
fun way to enjoy each other's company during the holidays.
To choose Santas, we use to draw names out of a hat. This system was
tedious, prone to many 'Wait, I got myself...' problems. This year, we
made a change to the rules that further complicated picking and we knew
the hat draw would not stand up to the challenge. Naturally, to solve
this problem, I scripted the process. Since that turned out to be more
interesting than I had expected, I decided to share.
This weeks Ruby Quiz is to implement a Secret Santa selection script.
Your script will be fed a list of names on STDIN.
...
Your script should then choose a Secret Santa for every name in the list.
Obviously, a person cannot be their own Secret Santa. In addition, my friends
no longer allow people in the same family to be Santas for each other and your
script should take this into account.
'''
Comment: This model skips the file input and mail parts. We
assume that the friends are identified with a number from 1..n,
and the families is identified with a number 1..num_families.
Compare with the following model:
* MiniZinc: http://www.hakank.org/minizinc/secret_santa.mzn
This model gives 4089600 solutions and the following statistics:
- failures: 31264
- branches: 8241726
- WallTime: 23735 ms (note: without any printing of the solutions)
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
import sys
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver('Secret Santa problem')
#
# data
#
family = [1, 1, 1, 1, 2, 3, 3, 3, 3, 3, 4, 4]
num_families = max(family)
n = len(family)
#
# declare variables
#
x = [solver.IntVar(0, n - 1, 'x[%i]' % i) for i in range(n)]
#
# constraints
#
solver.Add(solver.AllDifferent(x))
# Can't be one own's Secret Santa
# Ensure that there are no fix-point in the array
for i in range(n):
solver.Add(x[i] != i)
# No Secret Santa to a person in the same family
for i in range(n):
solver.Add(family[i] != solver.Element(family, x[i]))
#
# solution and search
#
db = solver.Phase(x, solver.INT_VAR_SIMPLE, solver.INT_VALUE_SIMPLE)
solver.NewSearch(db)
num_solutions = 0
while solver.NextSolution():
num_solutions += 1
print('x:', [x[i].Value() for i in range(n)])
print()
print('num_solutions:', num_solutions)
print('failures:', solver.Failures())
print('branches:', solver.Branches())
print('WallTime:', solver.WallTime(), 'ms')
| examples/notebook/contrib/secret_santa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Question 1
#
import numpy as np
input_mat = np.matrix([[1,4,2,8],[3,4,5,1],[8,9,6,4],[3,8,2,6]])
con_mat = np.matrix([[1,0],[1,1]])
def conv(input_matrix,kernel,stride,pad):
#Generating a new matrix which is input matrix padded with zeros
input_height,input_width = input_matrix.shape
pad_matrix = np.zeros((input_height+2*pad,input_width+2*pad),dtype=int)
pad_matrix[pad : pad + input_height , pad : pad + input_width]= input_matrix
#Determinig the size of output matrix
output_height,output_width = tuple(map(lambda i,j : int(((i-j + 2*pad) //stride) + 1),input_matrix.shape,kernel.shape))
output_matrix = np.zeros((output_height , output_width),dtype = int)
#Kernel size. Kernel matrix would always be in square form.
k = kernel.shape[0]
#Perform dot product with the new pad_matrix and the kernel
i_s = 0
for i in range(output_height):
j_s = 0
for j in range(output_width):
output_matrix[i,j] = np.sum(np.multiply(pad_matrix[i_s:i_s+k, j_s:j_s+k],kernel))
j_s = j_s + stride
i_s = i_s + stride
print(output_matrix)
# ### a
## 1
conv(input_mat,con_mat,1,0)
## 2
conv(input_mat,con_mat,2,2)
# ### b
def conv_avg(input_matrix,kernel,stride,pad):
#Generating a new matrix which is input matrix padded with zeros
input_height,input_width = input_matrix.shape
pad_matrix = np.zeros((input_height+2*pad,input_width+2*pad))
pad_matrix[pad : pad + input_height , pad : pad + input_width]= input_matrix
#Determinig the size of output matrix
output_height,output_width = tuple(map(lambda i,j : int(((i-j + 2*pad) //stride) + 1),input_matrix.shape,kernel.shape))
output_matrix = np.zeros((output_height , output_width))
#Kernel size. Kernel matrix would always be in square form.
k = kernel.shape[0]
#Perform dot product with the new pad_matrix and the kernel
i_s = 0
for i in range(output_height):
j_s = 0
for j in range(output_width):
output_matrix[i,j] = np.mean(np.multiply(pad_matrix[i_s:i_s+k, j_s:j_s+k],kernel))
j_s = j_s + stride
i_s = i_s + stride
print(output_matrix)
kernel_unit = np.ones((3,3))
print(kernel_unit)
input_mat_avg = np.matrix([[1,4,2,7],[3,4,2,1],[3,9,6,4],[3,5,2,6]])
conv_avg(input_mat_avg,kernel_unit,1,0)
| Assignments/HW4_Q1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image processing
#
# ## Load an image to numpy
from typing import List
import PIL
from PIL import Image, ImagePalette
import numpy as np
print('Pillow Version:', PIL.__version__)
import pandas as pd
# Let's create a helper to manipulate the image with a palette
class ImageReaderHelper:
def __init__(self, name: str):
self.name = name
def load(self):
self.img = Image.open(f'{self.name}.png').convert('P')
self.palette = self.img.getpalette()
self.img_arr = np.asarray(self.img)
self.palette_arr = np.asarray(self.palette).reshape(256, 3)
print(self.palette_arr)
print(f"{self.name} shape: {self.img_arr.shape} dtype: {self.img_arr.dtype}")
print(f"{self.name} palette: {self.palette_arr.shape} dtype: {self.palette_arr.dtype}")
print('Unique colors used from palette in the image')
self.colors_used = np.unique(self.img_arr)
print(self.colors_used)
print('Unique RGB colors used')
self.min_color_idx = np.amin(self.colors_used)
self.max_color_idx = np.amax(self.colors_used)
print(f'Color index range: {self.min_color_idx} to {self.max_color_idx}')
def print_palette_info(self):
rgb_colors_used = self.palette_arr[self.min_color_idx:self.max_color_idx+1]
print(rgb_colors_used)
color_id, color_frequency = np.unique(self.img_arr, return_counts = True)
print('Color frequency')
print(color_frequency)
def get_pixels(self):
return self.img_arr
def get_colors(self):
return self.palette_arr
def get_shuffle_colors(self):
new_palette_arr = self.palette_arr[self.min_color_idx:self.max_color_idx+1].copy()
np.random.shuffle(new_palette_arr)
return new_palette_arr
veg2020 = ImageReaderHelper('vegetation2020')
veg2020.load()
# ## Understand the palette
#
# We can check the how many colors from the palette are been used in the entire image. For vegetation2020, it is a palette of 30 colors.
veg2020.print_palette_info()
# ## Change the palette
class ImageWriteHelper:
def __init__(self, name: str):
self.name = name
def set_colors(self,colors: List):
# Palette would be provided as [(R, G, B), ..]
self.colors = colors
def set_pixels(self,pixels: List):
self.pixels = np.asarray(pixels).astype(np.uint8)
def _create_image_palette(self):
# The list must be aligned by channel (All R values must be contiguous in the list before G and B values.)
colors = np.asarray(self.colors).astype(np.uint8).flatten()
r_colors = colors[0::3]
g_colors = colors[1::3]
b_colors = colors[2::3]
cols_palette = np.concatenate((r_colors, g_colors, b_colors), axis=None).tolist()
img_palette = ImagePalette.ImagePalette(mode='RGB', palette=cols_palette, size=len(cols_palette))
return img_palette
def _create_image(self):
new_img = Image.fromarray(self.pixels, 'P')
new_img.putpalette(self._create_image_palette())
return new_img
def save(self):
self._create_image().save(f"{self.name}.png")
img_rand_palette = ImageWriteHelper('img-exp-random-palette')
img_rand_palette.set_pixels(veg2020.get_pixels())
img_rand_palette.set_colors(veg2020.get_shuffle_colors())
img_rand_palette.save()
# ## Create a tri-colors grid image
col_white = (255, 255, 255)
col_black = (0, 0, 0)
col_grey = (85, 86, 87)
col_navy = (0,0,128)
# +
img_grid = ImageWriteHelper('img-exp-grid')
img_grid.set_colors([col_white, col_black, col_grey])
pix_grid = np.ones((1080, 1920))
select_color = lambda i, j : 1 if i % 30 <10 and j % 30 < 10 else (2 if i % 30 > 20 and j % 30 > 20 else 0)
for i in range(1080):
for j in range(1920):
pix_grid[i, j]= select_color(i, j)
print(pix_grid)
img_grid.set_pixels(pix_grid)
img_grid.save()
# -
# ## Convert an image with palette to monochrome
img_converted_mono = ImageWriteHelper('img-exp-converted-monochrome')
img_converted_mono.set_colors([col_white, col_black, col_grey])
img_converted_mono_shape = veg2020.get_pixels().shape
it_pix_converted_mono = np.nditer(veg2020.get_pixels(), flags=['multi_index'])
pix_converted_mono = np.array([1 if v >0 else 0 for v in it_pix_converted_mono]).reshape(img_converted_mono_shape)
img_converted_mono.set_pixels(pix_converted_mono)
img_converted_mono.save()
# ## Add two images
img_exp_add = ImageWriteHelper('img-exp-add')
img_exp_add.set_colors([col_white, col_black, col_grey, col_navy])
pixel_exp_add = pix_converted_mono + pix_grid
img_exp_add.set_pixels(pixel_exp_add)
img_exp_add.save()
| exp2021-02/exp2021-02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="v1CUZ0dkOo_F"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="qmkj-80IHxnd"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="_xnMOsbqHz61"
# # CycleGAN
# + [markdown] colab_type="text" id="Ds4o1h4WHz9U"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/generative/cyclegan"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/generative/cyclegan.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/generative/cyclegan.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/generative/cyclegan.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="ITZuApL56Mny"
# This notebook demonstrates unpaired image to image translation using conditional GAN's, as described in [Unpaired Image-to-Image Translation using Cycle-Consistent Adversarial Networks](https://arxiv.org/abs/1703.10593), also known as CycleGAN. The paper proposes a method that can capture the characteristics of one image domain and figure out how these characteristics could be translated into another image domain, all in the absence of any paired training examples.
#
# This notebook assumes you are familiar with Pix2Pix, which you can learn about in the [Pix2Pix tutorial](https://www.tensorflow.org/tutorials/generative/pix2pix). The code for CycleGAN is similar, the main difference is an additional loss function, and the use of unpaired training data.
#
# CycleGAN uses a cycle consistency loss to enable training without the need for paired data. In other words, it can translate from one domain to another without a one-to-one mapping between the source and target domain.
#
# This opens up the possibility to do a lot of interesting tasks like photo-enhancement, image colorization, style transfer, etc. All you need is the source and the target dataset (which is simply a directory of images).
#
# 
# 
# + [markdown] colab_type="text" id="e1_Y75QXJS6h"
# ## Set up the input pipeline
# + [markdown] colab_type="text" id="5fGHWOKPX4ta"
# Install the [tensorflow_examples](https://github.com/tensorflow/examples) package that enables importing of the generator and the discriminator.
# + colab={} colab_type="code" id="bJ1ROiQxJ-vY"
# !pip install git+https://github.com/tensorflow/examples.git
# + colab={} colab_type="code" id="lhSsUx9Nyb3t"
try:
# # %tensorflow_version only exists in Colab.
import tensorflow.compat.v2 as tf
except Exception:
pass
tf.enable_v2_behavior()
# + colab={} colab_type="code" id="YfIk2es3hJEd"
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow_datasets as tfds
from tensorflow_examples.models.pix2pix import pix2pix
import os
import time
import matplotlib.pyplot as plt
from IPython.display import clear_output
tfds.disable_progress_bar()
AUTOTUNE = tf.data.experimental.AUTOTUNE
# + [markdown] colab_type="text" id="iYn4MdZnKCey"
# ## Input Pipeline
#
# This tutorial trains a model to translate from images of horses, to images of zebras. You can find this dataset and similar ones [here](https://www.tensorflow.org/datasets/datasets#cycle_gan).
#
# As mentioned in the [paper](https://arxiv.org/abs/1703.10593), apply random jittering and mirroring to the training dataset. These are some of the image augmentation techniques that avoids overfitting.
#
# This is similar to what was done in [pix2pix](https://www.tensorflow.org/tutorials/generative/pix2pix#load_the_dataset)
#
# * In random jittering, the image is resized to `286 x 286` and then randomly cropped to `256 x 256`.
# * In random mirroring, the image is randomly flipped horizontally i.e left to right.
# + colab={} colab_type="code" id="iuGVPOo7Cce0"
dataset, metadata = tfds.load('cycle_gan/horse2zebra',
with_info=True, as_supervised=True)
train_horses, train_zebras = dataset['trainA'], dataset['trainB']
test_horses, test_zebras = dataset['testA'], dataset['testB']
# + colab={} colab_type="code" id="2CbTEt448b4R"
BUFFER_SIZE = 1000
BATCH_SIZE = 1
IMG_WIDTH = 256
IMG_HEIGHT = 256
# + colab={} colab_type="code" id="Yn3IwqhiIszt"
def random_crop(image):
cropped_image = tf.image.random_crop(
image, size=[IMG_HEIGHT, IMG_WIDTH, 3])
return cropped_image
# + colab={} colab_type="code" id="muhR2cgbLKWW"
# normalizing the images to [-1, 1]
def normalize(image):
image = tf.cast(image, tf.float32)
image = (image / 127.5) - 1
return image
# + colab={} colab_type="code" id="fVQOjcPVLrUc"
def random_jitter(image):
# resizing to 286 x 286 x 3
image = tf.image.resize(image, [286, 286],
method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
# randomly cropping to 256 x 256 x 3
image = random_crop(image)
# random mirroring
image = tf.image.random_flip_left_right(image)
return image
# + colab={} colab_type="code" id="tyaP4hLJ8b4W"
def preprocess_image_train(image, label):
image = random_jitter(image)
image = normalize(image)
return image
# + colab={} colab_type="code" id="VB3Z6D_zKSru"
def preprocess_image_test(image, label):
image = normalize(image)
return image
# + colab={} colab_type="code" id="RsajGXxd5JkZ"
train_horses = train_horses.map(
preprocess_image_train, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
train_zebras = train_zebras.map(
preprocess_image_train, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
test_horses = test_horses.map(
preprocess_image_test, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
test_zebras = test_zebras.map(
preprocess_image_test, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
# + colab={} colab_type="code" id="e3MhJ3zVLPan"
sample_horse = next(iter(train_horses))
sample_zebra = next(iter(train_zebras))
# + colab={} colab_type="code" id="4pOYjMk_KfIB"
plt.subplot(121)
plt.title('Horse')
plt.imshow(sample_horse[0] * 0.5 + 0.5)
plt.subplot(122)
plt.title('Horse with random jitter')
plt.imshow(random_jitter(sample_horse[0]) * 0.5 + 0.5)
# + colab={} colab_type="code" id="0KJyB9ENLb2y"
plt.subplot(121)
plt.title('Zebra')
plt.imshow(sample_zebra[0] * 0.5 + 0.5)
plt.subplot(122)
plt.title('Zebra with random jitter')
plt.imshow(random_jitter(sample_zebra[0]) * 0.5 + 0.5)
# + [markdown] colab_type="text" id="hvX8sKsfMaio"
# ## Import and reuse the Pix2Pix models
# + [markdown] colab_type="text" id="cGrL73uCd-_M"
# Import the generator and the discriminator used in [Pix2Pix](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/pix2pix/pix2pix.py) via the installed [tensorflow_examples](https://github.com/tensorflow/examples) package.
#
# The model architecture used in this tutorial is very similar to what was used in [pix2pix](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/pix2pix/pix2pix.py). Some of the differences are:
#
# * Cyclegan uses [instance normalization](https://arxiv.org/abs/1607.08022) instead of [batch normalization](https://arxiv.org/abs/1502.03167).
# * The [CycleGAN paper](https://arxiv.org/abs/1703.10593) uses a modified `resnet` based generator. This tutorial is using a modified `unet` generator for simplicity.
#
# There are 2 generators (G and F) and 2 discriminators (X and Y) being trained here.
#
# * Generator `G` learns to transform image `X` to image `Y`. $(G: X -> Y)$
# * Generator `F` learns to transform image `Y` to image `X`. $(F: Y -> X)$
# * Discriminator `D_X` learns to differentiate between image `X` and generated image `X` (`F(Y)`).
# * Discriminator `D_Y` learns to differentiate between image `Y` and generated image `Y` (`G(X)`).
#
# 
# + colab={} colab_type="code" id="8ju9Wyw87MRW"
OUTPUT_CHANNELS = 3
generator_g = pix2pix.unet_generator(OUTPUT_CHANNELS, norm_type='instancenorm')
generator_f = pix2pix.unet_generator(OUTPUT_CHANNELS, norm_type='instancenorm')
discriminator_x = pix2pix.discriminator(norm_type='instancenorm', target=False)
discriminator_y = pix2pix.discriminator(norm_type='instancenorm', target=False)
# + colab={} colab_type="code" id="wDaGZ3WpZUyw"
to_zebra = generator_g(sample_horse)
to_horse = generator_f(sample_zebra)
plt.figure(figsize=(8, 8))
contrast = 8
imgs = [sample_horse, to_zebra, sample_zebra, to_horse]
title = ['Horse', 'To Zebra', 'Zebra', 'To Horse']
for i in range(len(imgs)):
plt.subplot(2, 2, i+1)
plt.title(title[i])
if i % 2 == 0:
plt.imshow(imgs[i][0] * 0.5 + 0.5)
else:
plt.imshow(imgs[i][0] * 0.5 * contrast + 0.5)
plt.show()
# + colab={} colab_type="code" id="O5MhJmxyZiy9"
plt.figure(figsize=(8, 8))
plt.subplot(121)
plt.title('Is a real zebra?')
plt.imshow(discriminator_y(sample_zebra)[0, ..., -1], cmap='RdBu_r')
plt.subplot(122)
plt.title('Is a real horse?')
plt.imshow(discriminator_x(sample_horse)[0, ..., -1], cmap='RdBu_r')
plt.show()
# + [markdown] colab_type="text" id="0FMYgY_mPfTi"
# ## Loss functions
# + [markdown] colab_type="text" id="JRqt02lupRn8"
# In CycleGAN, there is no paired data to train on, hence there is no guarantee that the input `x` and the target `y` pair are meaningful during training. Thus in order to enforce that the network learns the correct mapping, the authors propose the cycle consistency loss.
#
# The discriminator loss and the generator loss are similar to the ones used in [pix2pix](https://www.tensorflow.org/tutorials/generative/pix2pix#define_the_loss_functions_and_the_optimizer).
# + colab={} colab_type="code" id="cyhxTuvJyIHV"
LAMBDA = 10
# + colab={} colab_type="code" id="Q1Xbz5OaLj5C"
loss_obj = tf.keras.losses.BinaryCrossentropy(from_logits=True)
# + colab={} colab_type="code" id="wkMNfBWlT-PV"
def discriminator_loss(real, generated):
real_loss = loss_obj(tf.ones_like(real), real)
generated_loss = loss_obj(tf.zeros_like(generated), generated)
total_disc_loss = real_loss + generated_loss
return total_disc_loss * 0.5
# + colab={} colab_type="code" id="90BIcCKcDMxz"
def generator_loss(generated):
return loss_obj(tf.ones_like(generated), generated)
# + [markdown] colab_type="text" id="5iIWQzVF7f9e"
# Cycle consistency means the result should be close to the original input. For example, if one translates a sentence from English to French, and then translates it back from French to English, then the resulting sentence should be the same as the original sentence.
#
# In cycle consistency loss,
#
# * Image $X$ is passed via generator $G$ that yields generated image $\hat{Y}$.
# * Generated image $\hat{Y}$ is passed via generator $F$ that yields cycled image $\hat{X}$.
# * Mean absolute error is calculated between $X$ and $\hat{X}$.
#
# $$forward\ cycle\ consistency\ loss: X -> G(X) -> F(G(X)) \sim \hat{X}$$
#
# $$backward\ cycle\ consistency\ loss: Y -> F(Y) -> G(F(Y)) \sim \hat{Y}$$
#
#
# 
# + colab={} colab_type="code" id="NMpVGj_sW6Vo"
def calc_cycle_loss(real_image, cycled_image):
loss1 = tf.reduce_mean(tf.abs(real_image - cycled_image))
return LAMBDA * loss1
# + [markdown] colab_type="text" id="U-tJL-fX0Mq7"
# As shown above, generator $G$ is responsible for translating image $X$ to image $Y$. Identity loss says that, if you fed image $Y$ to generator $G$, it should yield the real image $Y$ or something close to image $Y$.
#
# $$Identity\ loss = |G(Y) - Y| + |F(X) - X|$$
# + colab={} colab_type="code" id="05ywEH680Aud"
def identity_loss(real_image, same_image):
loss = tf.reduce_mean(tf.abs(real_image - same_image))
return LAMBDA * 0.5 * loss
# + [markdown] colab_type="text" id="G-vjRM7IffTT"
# Initialize the optimizers for all the generators and the discriminators.
# + colab={} colab_type="code" id="iWCn_PVdEJZ7"
generator_g_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
generator_f_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
discriminator_x_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
discriminator_y_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
# + [markdown] colab_type="text" id="aKUZnDiqQrAh"
# ## Checkpoints
# + colab={} colab_type="code" id="WJnftd5sQsv6"
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(generator_g=generator_g,
generator_f=generator_f,
discriminator_x=discriminator_x,
discriminator_y=discriminator_y,
generator_g_optimizer=generator_g_optimizer,
generator_f_optimizer=generator_f_optimizer,
discriminator_x_optimizer=discriminator_x_optimizer,
discriminator_y_optimizer=discriminator_y_optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
# if a checkpoint exists, restore the latest checkpoint.
if ckpt_manager.latest_checkpoint:
ckpt.restore(ckpt_manager.latest_checkpoint)
print ('Latest checkpoint restored!!')
# + [markdown] colab_type="text" id="Rw1fkAczTQYh"
# ## Training
#
# Note: This example model is trained for fewer epochs (40) than the paper (200) to keep training time reasonable for this tutorial. Predictions may be less accurate.
# + colab={} colab_type="code" id="NS2GWywBbAWo"
EPOCHS = 40
# + colab={} colab_type="code" id="RmdVsmvhPxyy"
def generate_images(model, test_input):
prediction = model(test_input)
plt.figure(figsize=(12, 12))
display_list = [test_input[0], prediction[0]]
title = ['Input Image', 'Predicted Image']
for i in range(2):
plt.subplot(1, 2, i+1)
plt.title(title[i])
# getting the pixel values between [0, 1] to plot it.
plt.imshow(display_list[i] * 0.5 + 0.5)
plt.axis('off')
plt.show()
# + [markdown] colab_type="text" id="kE47ERn5fyLC"
# Even though the training loop looks complicated, it consists of four basic steps:
#
# * Get the predictions.
# * Calculate the loss.
# * Calculate the gradients using backpropagation.
# * Apply the gradients to the optimizer.
# + colab={} colab_type="code" id="KBKUV2sKXDbY"
@tf.function
def train_step(real_x, real_y):
# persistent is set to True because the tape is used more than
# once to calculate the gradients.
with tf.GradientTape(persistent=True) as tape:
# Generator G translates X -> Y
# Generator F translates Y -> X.
fake_y = generator_g(real_x, training=True)
cycled_x = generator_f(fake_y, training=True)
fake_x = generator_f(real_y, training=True)
cycled_y = generator_g(fake_x, training=True)
# same_x and same_y are used for identity loss.
same_x = generator_f(real_x, training=True)
same_y = generator_g(real_y, training=True)
disc_real_x = discriminator_x(real_x, training=True)
disc_real_y = discriminator_y(real_y, training=True)
disc_fake_x = discriminator_x(fake_x, training=True)
disc_fake_y = discriminator_y(fake_y, training=True)
# calculate the loss
gen_g_loss = generator_loss(disc_fake_y)
gen_f_loss = generator_loss(disc_fake_x)
total_cycle_loss = calc_cycle_loss(real_x, cycled_x) + calc_cycle_loss(real_y, cycled_y)
# Total generator loss = adversarial loss + cycle loss
total_gen_g_loss = gen_g_loss + total_cycle_loss + identity_loss(real_y, same_y)
total_gen_f_loss = gen_f_loss + total_cycle_loss + identity_loss(real_x, same_x)
disc_x_loss = discriminator_loss(disc_real_x, disc_fake_x)
disc_y_loss = discriminator_loss(disc_real_y, disc_fake_y)
# Calculate the gradients for generator and discriminator
generator_g_gradients = tape.gradient(total_gen_g_loss,
generator_g.trainable_variables)
generator_f_gradients = tape.gradient(total_gen_f_loss,
generator_f.trainable_variables)
discriminator_x_gradients = tape.gradient(disc_x_loss,
discriminator_x.trainable_variables)
discriminator_y_gradients = tape.gradient(disc_y_loss,
discriminator_y.trainable_variables)
# Apply the gradients to the optimizer
generator_g_optimizer.apply_gradients(zip(generator_g_gradients,
generator_g.trainable_variables))
generator_f_optimizer.apply_gradients(zip(generator_f_gradients,
generator_f.trainable_variables))
discriminator_x_optimizer.apply_gradients(zip(discriminator_x_gradients,
discriminator_x.trainable_variables))
discriminator_y_optimizer.apply_gradients(zip(discriminator_y_gradients,
discriminator_y.trainable_variables))
# + colab={} colab_type="code" id="2M7LmLtGEMQJ"
for epoch in range(EPOCHS):
start = time.time()
n = 0
for image_x, image_y in tf.data.Dataset.zip((train_horses, train_zebras)):
train_step(image_x, image_y)
if n % 10 == 0:
print ('.', end='')
n+=1
clear_output(wait=True)
# Using a consistent image (sample_horse) so that the progress of the model
# is clearly visible.
generate_images(generator_g, sample_horse)
if (epoch + 1) % 5 == 0:
ckpt_save_path = ckpt_manager.save()
print ('Saving checkpoint for epoch {} at {}'.format(epoch+1,
ckpt_save_path))
print ('Time taken for epoch {} is {} sec\n'.format(epoch + 1,
time.time()-start))
# + [markdown] colab_type="text" id="1RGysMU_BZhx"
# ## Generate using test dataset
# + colab={} colab_type="code" id="KUgSnmy2nqSP"
# Run the trained model on the test dataset
for inp in test_horses.take(5):
generate_images(generator_g, inp)
# + [markdown] colab_type="text" id="ABGiHY6fE02b"
# ## Next steps
#
# This tutorial has shown how to implement CycleGAN starting from the generator and discriminator implemented in the [Pix2Pix](https://www.tensorflow.org/tutorials/generative/pix2pix) tutorial. As a next step, you could try using a different dataset from [TensorFlow Datasets](https://www.tensorflow.org/datasets/datasets#cycle_gan).
#
# You could also train for a larger number of epochs to improve the results, or you could implement the modified ResNet generator used in the [paper](https://arxiv.org/abs/1703.10593) instead of the U-Net generator used here.
#
#
#
# Try using a different dataset from . You can also implement the modified ResNet generator used in the [paper](https://arxiv.org/abs/1703.10593) instead of the U-Net generator that's used here.
| site/en/tutorials/generative/cyclegan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python - Sequenze (Stringa, Lista e Tupla)
#
# > ### Definizione di sequenza
# > ### Costruzione di una *sequenza*
# > ### Dimensione di una *sequenza*
# > ### Accesso agli elementi di una *sequenza*
# > ### Concatenazione di *sequenze*
# > ### Ripetizione di una *sequenza*
# > ### Scansione degli elementi di una *sequenza*
# > ### Aggiornamento di una lista
# > ### Assegnamento multiplo
# > ### List comprehension
# ## Definizione di *sequenza*
#
# Stringhe, liste e tuple sono ***sequenze***, cioé oggetti su cui si può iterare e i cui elementi sono indicizzati tramite posizione.
#
# **Stringa**: sequenza di caratteri
# - oggetto di tipo `str`
# - oggetto ___immutabile___
#
# ---
#
# **Lista**: sequenza di valori (oggetti) anche di tipo diverso
# - oggetto di tipo `list`
# - oggetto ***mutabile***
#
# **Tupla**: sequenza di valori (oggetti) anche di tipo diverso
# - oggetto di tipo `tuple`
# - oggetto ***immutabile***
# ## Costruzione di una *sequenza*
# ### Costruzione di una stringa
# - tramite letterale (sequenza di caratteri racchiusi tra singoli apici `'` o doppi apici `"`)
# - tramite funzione `str()`
# Letterale con doppi apici:
print("ciao")
# Letterale con singoli apici:
print('ciao')
# Singoli apici nel valore della stringa:
print('\'ciao\'')
print("'ciao'")
# Doppi apici nel valore della stringa:
print("\"ciao\"")
print('"ciao"')
# *Newline* `\n` nel valore della stringa:
print('ciao\nciao')
# Costruzione della stringa vuota:
""
''
# #### Esempi di costruzione tramite funzione `str()`
# Stringa vuota:
str()
# Costruzione con letterale stringa:
str("ciao")
# Costruzione con espressione aritmetica:
str(3+4)
# Costruzione con espressione di confronto:
str(34 > 0)
# **NOTA BENE**: il valore restituito dalla chiamata `str(34 >0)` è la stringa dei caratteri `T`, `r`, `u` ed `e` e non il valore `True` di tipo `bool`.
# ### Costruzione di una lista
#
# - tramite letterale `[value1, value2, ..., valueN]`
# - tramite funzione `list()`
# #### Esempi di costruzione tramite letterale
# Lista vuota:
[]
# Lista di un solo valore:
[4]
# Lista di tre valori dello stesso tipo:
[1,2,3]
# Lista di tre valori di tipo diverso:
[10, 7.8, "ab"]
# Lista di tre valori di tipo diverso di cui il terzo di tipo `list` (lista annidata):
[10, 7.8, [True, "ab"]]
# #### Esempi di costruzione tramite la funzione `list()`
# Lista vuota:
list()
# Costruzione con stringa:
list("abcde")
# Costruzione con lista:
list([1,2,3,4])
# ### Costruzione di una tupla
# - tramite letterale `(value1, value2, ..., valueN)`
#
# - tramite la funzione `tuple()`
# #### Esempi di costruzione tramite letterale
# Tupla vuota:
()
# Tupla di un solo valore:
(4,)
# Tupla di tre valori dello stesso tipo:
(1,2,3)
# Tupla di tre valori di tipo diverso:
(10, 7.8, "ab")
# Tupla di quattro valori di tipo diverso di cui il terzo di tipo `list` (lista annidata) e il quarto di tipo `tuple` (tupla annidata):
(10, 7.8, [1,2], (3,4))
# **NOTA BENE**: le parentesi tonde nel letterale di una tupla sono opzionali (non possono però essere omesse nel caso di letterale di una tupla vuota).
10, 7.8, [1,2], (3,4)
# #### Esempi di costruzione tramite la funzione `tuple()`
# Tupla vuota:
tuple()
# Costruzione con stringa:
tuple("abcde")
# Costruzione con lista:
tuple([1,2,3])
# Costruzione con tupla:
tuple((1,2,3))
# ## Dimensione di una *sequenza*
# La funzione `len()` restituisce la dimensione della *sequenza* passata come argomento.
#
# Dimensione di una stringa:
len("abcde")
# Dimensione di una lista:
len([1,2,3,4])
# Dimensione di una tupla:
len((1,2,3,4,5))
# ## Accesso agli elementi di una *sequenza*
# Le posizioni degli elementi all'interno di una *sequenza* sono indicate tramite:
#
# - indici positivi
# - 0: posizione del primo elemento
# - 1: posizione del secondo elemento
# - ...
# - dimensione della sequenza decrementato di 1: posizione dell'ultimo elemento
#
#
# - indici negativi
# - -1: posizione dell'ultimo elemento
# - -2: posizione del penultimo elemento
# - ...
# - negazione aritmetica della dimensione della sequenza: posizione del primo elemento
# ### Accesso a un solo elemento di una *sequenza*
# L'espressione:
#
# my_sequence[some_index]
#
# restituisce l'elemento della *sequenza* `my_sequence` in posizione `some_index`.
# **Esempi di accesso a un carattere di una stringa**
# Accesso al quinto carattere tramite indice positivo:
stringa = 'Hello world!'
stringa[4]
# **NOTA BENE**: `stringa[4]` restituisce un oggetto di tipo `str`.
# Accesso al quinto carattere tramite indice negativo:
stringa = 'Hello world!'
stringa[-8]
# **Esempi di accesso a un elemento di una lista**
# Accesso al quarto elemento tramite indice positivo:
lista = [1, 2, 3, [4, 5]]
lista[3]
# Accesso al primo elemento della lista annidata:
lista = [1, 2, 3, [4, 5]]
lista[3][0]
# Accesso al primo elemento tramite indice negativo:
lista = [1, 2, 3, [4, 5]]
lista[-4]
# **Esempi di accesso a un elemento di una tupla**
# Accesso al terzo elemento tramite indice positivo:
tupla = (1, 2, 3, 4)
tupla[2]
# Accesso all'ultimo elemento tramite indice negativo:
tupla = (1, 2, 3, 4)
tupla[-1]
# ### Accesso a elementi consecutivi di una *sequenza* (*slicing1*)
# L'epressione:
#
# my_sequence[start_index:end_index]
#
# restituisce gli elementi della *sequenza* `my_sequence` da quello in posizione `start_index` fino a quello in posizione `end_index-1`.
# **Esempi di accesso a caratteri consecutivi di una stringa (sottostringa)**
#
# Accesso alla sottostringa dal terzo al settimo carattere tramite indici positivi:
stringa = 'Hello world!'
stringa[2:7]
# Accesso alla sottostringa dal terzo al settimo carattere tramite indici negativi:
stringa = 'Hello world!'
stringa[-10:-5]
# Accesso al prefisso dei primi tre caratteri tramite indici positivi:
stringa = 'Hello world!'
stringa[0:3]
stringa = 'Hello world!'
stringa[:3]
# Accesso al suffisso che parte dal nono carattere tramite indici positivi:
stringa = 'Hello world!'
stringa[8:len(stringa)]
stringa = 'Hello world!'
stringa[8:]
# Ottenere una copia della stringa:
stringa = 'Hello world!'
stringa[0:len(stringa)]
stringa = 'Hello world!'
stringa[:]
# **Esempi di accesso a elementi consecutivi di una lista/tupla (sottolista/sottotupla)**
# Accesso alla sottolista dal terzo al quarto elemento tramite indici positivi:
lista = [1, 2, 3, 4, 5, 6]
lista[2:4]
# Accesso al suffisso di lista che parte dal terzo elemento tramite indici positivi:
lista = [1, 2, 3, 4, 5, 6]
lista[2:]
# Accesso al prefisso di tupla dei primi quattro elementi tramite indici positivi:
tupla = (1, 2, 3, 4, 5, 6)
tupla[:4]
# ### Accesso a elementi non consecutivi di una *sequenza* (*slicing2*)
# L'epressione:
#
# my_sequence[start_index:end_index:step]
#
# equivale a
#
# - considerare la sottosequenza di elementi consecutivi `my_sequence[start_index:end_index]`
#
# e
#
# - restituire di questa sottosequenza gli elementi a partire dal primo, saltando ogni volta `step-1` elementi
#
# **Esempi di accesso a caratteri non consecutivi di una stringa**
#
# Accesso ai caratteri a partire dal secondo, saltando ogni volta due caratteri e terminando non oltre il settimo:
stringa = 'xAxxBxxCxx'
stringa[1:8:3]
# Lo *slicing* con passo può essere utilizzato per invertire una sequenza:
stringa = 'Hello world!'
stringa[::-1]
lista = [1, 2, 3, 4, 5, 6, 7, 8, 9]
lista[::-1]
# ## Concatenazione di *sequenze*
#
# L'espressione:
#
# my_sequence1 + my_sequence2 + ... + my_sequenceN
#
# restituisce la concatenazione di N sequenze dello stesso tipo.
"ciao " + "mondo"
[1,2] + [3,4,5]
(1,2) + (3,4,5)
# ## Ripetizione di una *sequenza*
#
# L'espressione:
#
# my_sequence * times
#
# restituisce la ripetizione di `my_sequence` per `times` volte.
"ciao " * 4
[1,2] * 4
(1,2) * 4
# ## Scansione degli elementi una *sequenza*
# ### Operatore `in`
# L'espressione:
#
# my_element in my_sequence
#
# restituisce `True` se `my_element` è presente in `my_sequence`, altrimenti restituisce `False`
#
# Controllo della presenza del carattere `H`:
stringa = 'Hello world!'
"H" in stringa
# Controllo della presenza della sottostringa `world` nella stringa:
stringa = 'Hello world!'
"llo" in stringa
# Controllo della presenza della stringa `ca` nella lista:
lista = [1, 2, 'acaa', 10.5]
"ca" in lista
# Controllo della presenza della lista `[1,2]` nella lista:
lista = [1, 2, 'acaa', 10.5]
[1,2] in lista
# ### Scansione con operatore `in`
# **Sintassi di scansione di una *sequenza***:
#
# for element in my_sequence:
# do_something
#
# - `my_sequence`: *sequenza* da scandire da sinistra a destra
# - `element`: variabile di scansione
# - `do_something`: blocco di istruzioni da eseguire per ogni elemento considerato
#
# **Regola**:
#
# - le istruzioni in `do_something` devono essere indentate 4 volte rispetto alla riga di intestazione
#
# Scansione e stampa a video dei caratteri della stringa:
stringa = 'world'
for c in stringa:
print(c)
# Scansione e stampa a video degli elementi della lista:
lista = [3, 45.6, 'ciao']
for element in lista:
print(element)
# Scansione e stampa a video degli elementi della tupla:
tupla = 3, 45.6, 'ciao'
for element in tupla:
print(element)
# ## Aggiornamento di una lista
# ### Aggiornamento di un elemento di una lista
# L'istruzione:
#
# my_list[some_index] = new_value
#
# sostituisce l'elemento della lista `my_list` in posizione `some_index` con il nuovo valore `new_value`.
#
# Sostituzione del quarto elemento della lista con il valore 0:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[3] = 0
lista
# ### Aggiornamento di più elementi di una lista
# Le istruzioni:
#
# my_list[start_index:end_index] = new_list
# my_list[start_index:end_index:step] = new_list
#
# sostituiscono la sottolista di `my_list` ottenuta tramite *slicing* con la lista `new_list`.
# Sostituzione dei tre elementi consecutivi dal quarto al sesto della lista:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[3:6] = ['*', '*', '*']
lista
# Cancellazione dei tre elementi consecutivi dal quarto al sesto della lista:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[3:6] = []
lista
# Inserimento di tre elementi asterisco prima del secondo elemento:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[1:1] = ['*', '*', '*']
lista
# Aggiunta in coda di tre elementi asterisco:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[len(lista):len(lista)] = ['*', '*', '*']
lista
# Aggiunta in testa di tre elementi asterisco:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[:0] = ['*', '*', '*']
lista
# Aggiornamento al valore 0 degli elementi in posizione di indice pari:
lista = [1, 2, 3, 4, 5, 6, 7]
lista[::2] = [0, 0, 0, 0]
lista
# ### Cancellazione di un elemento di una lista con l'operatore `del`
# L'istruzione:
#
# del my_list[some_index]
#
# rimuove dalla lista `my_list` l'elemento in posizione di indice `some_index`.
#
# Cancellazione del terzo elemento della lista:
lista = [1, 2, 3, 4]
del lista[2]
lista
# ### Cancellazione di più elementi di una lista con l'operatore `del`
# Le istruzioni:
#
# del my_list[start_index:end_index]
# del my_list[start_index:end_index:step]
#
# rimuovono dalla lista `my_list` gli elementi prodotti dall'operazione di *slicing*.
#
# Cancellazione degli elementi dal terzo al quinto:
lista = [1, 2, 3, 4, 5, 6, 7, 8]
del lista[2:5]
lista
# Cancellazione degli elementi in posizione dispari:
lista = [0, 1, 2, 3, 4, 5, 6, 7]
del lista[1::2]
lista
# ## Assegnamento multiplo
# L'istruzione di assegnamento:
#
# (my_var1, my_var2, ..., my_varN) = my_sequence
#
# assegna alle N variabili specificate nella tupla a sinistra i valori della *sequenza* `my_sequence` specificata a destra.
#
# Le parentesi sono opzionali:
#
# my_var1, my_var2, ..., my_varN = my_sequence
#
# **NOTA BENE**: `my_sequence` deve avere dimensione pari a N.
#
# Assegnamento dei quattro caratteri della stringa "1234" a quattro variabili diverse:
v1, v2, v3, v4 = "1234"
v4
# Assegnamento dei quattro elementi della lista `[1,2,3,4]` a quattro variabili diverse:
v1, v2, v3, v4 = [1,2,3,4]
v4
# Assegnamento dei quattro elementi della tupla `(1,2,3,4)` a quattro variabili diverse:
v1, v2, v3, v4 = 1,2,3,4
# Scambiare il valore tra due variabili:
v1,v2 = v2,v1
v2
# Scandire una lista di tuple di due elementi stampando ogni volta i due elementi separatamente:
lista = [(1, 2), (3, 4), (5, 6)]
for (a,b) in lista:
print(str(a)+' '+str(b))
# ## List comprehension
# L'istruzione:
#
# [some_expression for element in my_sequence if condition]
#
# equivale a scrivere:
#
# for element in my_sequence:
# if condition:
# some_expression
#
# con la differenza che i valori restituiti da `some_expression` vengono restituiti tutti in un oggetto di tipo `list`.
#
# **NB**: la clausola if è opzionale.
#
# Data una lista, scrivere la list comprehension che produce la lista degli elementi di valore pari incrementati di 1:
lista = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
[element+1 for element in lista if element % 2 == 0]
# Data una lista, scrivere la list comprehension che produce la lista dei caratteri concatenati alla cifra 1:
stringa = 'ciao'
[e+"1" for e in stringa]
# La list comprehension nella sua sintassi più generale:
#
# [some_expression for e1 in my_seq1 for e2 in my_se2 ... for eN in my_seqN if condition]
#
# equivale a scrivere:
#
# for e1 in my_seq1:
# for e2 in my_seq2:
# ...
# for eN in my_seqN:
# if condition:
# some_expression
# Scrivere la list comprehension che produce la lista delle stringhe ottenute combinando tra di loro in tutti i modi possibili i caratteri della stringa, gli elementi (valori interi) della lista e gli elementi (caratteri) della tupla:
stringa = 'ciao'
lista = [1, 2, 3, 4]
tupla = ('a', 'b', 'c')
[x+str(y)+z for x in stringa for y in lista for z in tupla]
| laboratorio/lezione2-01ott21/lezione2-python-sequenze.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # Run this cell but don't modify it.
# # %load_ext autoreload
# # %autoreload 2
# import numpy as np
# import os
# import cv2
# import glob
# import matplotlib.pyplot as plt
# import pickle
# import matplotlib.image as mpimg
# from lane_lines import Lane_Lines
# +
# # Load our image
# binary_w= mpimg.imread('output_images/warped/test2.jpg')
# binary_warped = Lane_Lines(np.copy(binary_w[:,:,1]))
# # binary_warped = Lane_Lines(np.copy(binary_w))
# out_img = binary_warped.fit_polynomial()
# left_curverad, right_curverad = binary_warped.measure_curvature_real()
# print('Left Lane =', left_curverad, 'm and Right Lane =', right_curverad, 'm')
# print('Radius of road =',((left_curverad+right_curverad)/2),'m')
# plt.imshow(out_img)
# -
# +
# Run this cell but don't modify it.
# # %load_ext autoreload
# # %autoreload 2
import numpy as np
import os
import cv2
import glob
import matplotlib.pyplot as plt
import pickle
import matplotlib.image as mpimg
from lane_lines import Lane_Lines
# -
# Load our image
binary_w= mpimg.imread('output_images/warped/straight_lines1.jpg')
binary_warped = Lane_Lines(np.copy(binary_w[:,:,1]))
# binary_warped = Lane_Lines(np.copy(binary_w))
out_img = binary_warped.find_lane_pixels()
left_curverad, right_curverad, center_offset_m = binary_warped.measure_curvature_real()
print('Left Lane =' ,"%.2f"% left_curverad, 'm and Right Lane =', "%.2f"% right_curverad, 'm')
print('Radius of road =',"%.2f"% ((left_curverad+right_curverad)/2),'m')
print('Center Offset =',"%.2f"% center_offset_m,'m')
plt.imshow(out_img)
| Trials/Lane_Lines_Trial_Class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduccción
#
# Luego de un primer período con bajos contagios y poca propagación a lo largo de Argentina, desde hace dos meses el COVID-19 viene pegando fuerte en el país. El Ministerio de Salud dispone desde el 15 de Mayo de un dataset con cada caso registrado por localidad. Es una base de datos bastante amplia que hemos decidido analizar, considerando en esta primer aproximación algunos factores que nos parecieron interesantes, tales como:
# <ul>
# <li> Contagios por rango etario </li>
# <li> Mapa con densidad de muertes por departamento/comuna </li>
# <li> Predicciones en base a estudios previos </li>
# <li> Diferencias entre datasets: Min de Salud vs. COVID Stats AR </li>
# </ul>
#
# <h2> Problemática </h2>
# Sin bien la pandemia es una temática bastante cubierta por los medios, el análisis de datos es infrecuente y poco detallado. Con este proyecto intentaremos responder a algunas inquitudes particulares, sin que las mismas sean extensivas a una mirada general de la pandemia. Consideramos oportuno responder a algunas preguntas que con suerte, nos pueden ayudar a comprender mejor el lugar en el que estamos parados, cuánto falta hasta que la situación se estabilice o si por el contrario ya lo hizo; como así también algunos aspectos interesantes sobre la carga de datos.
#
# Esperamos con esto llevar tranquilidad e información de calidad a los ciudadanos y hacer foco en algunos problemas puntuales al sistema de salud.
# +
import pandas as pd
import numpy as np
# Uncomment if you want to download the csv file -- daily updated
# #!wget https://sisa.msal.gov.ar/datos/descargas/covid-19/files/Covid19Casos.csv
df = pd.read_csv("Covid19Casos.csv")
df.head()
# -
# # Limpieza de datos para análisis de fallecidos
#
# Sobre el dataset del Ministerio de salud, vamos a tomar algunos campos para analizar las tendencias sobre la curva de fallecidos diarios y acumulados.
# +
df['fallecido'] = df['fallecido'].replace("SI", 1)
df['fallecido'] = df['fallecido'].replace("NO", 0)
classFilter = (df.fallecido == 1) & (df.clasificacion_resumen == 'Confirmado')
dailyDeathsMS = df.loc[classFilter]
dailyDeathsMS['fecha'] = pd.to_datetime(dailyDeathsMS['fecha_fallecimiento'], format='%Y-%m-%d')
dailyDeathsMS = dailyDeathsMS[['fallecido', 'fecha']].groupby('fecha').sum()
dailyDeathsMS.reset_index(inplace=True)
# -
# Luego de limpiar los datos, nos quedamos con un dataframe que contiene únicamente el número de fallecidos y la fecha. Podemos hacer un primer análisis de la distribución de los decesos en función del tiempo y observar la tendencia.
# +
import matplotlib.pyplot as plt
dailyDeathsMS.plot(kind='scatter', x='fecha', y='fallecido', color='r')
plt.title('Fallecidos diarios')
plt.ylabel('fallecidos')
plt.xlabel('fecha')
plt.rcParams['figure.dpi'] = 200
# -
# Un estudio realizado por <NAME> y <NAME> de la Universidad de Kobe sobre curvas de ajuste para el desempeño del COVID-19 ha demostrado resultados satisfactorios. De esta manera, los autores han podido predecir el pico de casos en distintos lugares del mundo. Como la metodología de evaluación de contagiados cambia a medida que avanza la epidemia, nos hemos centrado en los fallecimientos. De esta manera podemos analizar las curvas bajo un mismo criterio. Para más información sobre la curva de ajuste a utilizar, pueden visitar el paper correspondiente en: https://www.medrxiv.org/content/10.1101/2020.07.02.20144899v2
#
# La curva de ajuste a utilizar en particular es la siguiente:
#
# <img src="./extras/formula.png" alt="Carga de Casos">
#
# Definimos en la siguiente función "log_normal" el comportamiento de la curva y fijamos 3 parámetros de ajuste: a, b y c. La particularidad de esta curva, tal como se puede observar en el gráfico inferior, es una pendiente de subida rápida y un descenso lento. Este comportamiento se ajusta considerablemente bien a las curvas de infección y decesos del COVID-19 y otras epidemias. Nuevamente, existe más información al respecto en el paper citado previamente.
# +
# Define fitting curve
def log_normal(t, a, b, c):
y = (a/t)*np.exp(-((np.log(t)-b)**2)/c**2)
return y
time = np.arange(0.1,10, 0.1)
y = [log_normal(t, 1, 1, 1) for t in time]
plt.title('Función log_normal - parámetros unitarios')
plt.ylabel('log_normal(x)')
plt.xlabel('x')
plt.rcParams['figure.dpi'] = 50
plt.plot(time, y)
# -
# convert dates to Date of Year format
dailyDeathsMS['DoY'] = dailyDeathsMS.fecha.apply(lambda x: x.strftime('%j'))
dailyDeathsMS.tail(14)
# Tal como se observa en la tabla anterior y la figura del gráfico de diseprsión, el número de fallecidos disminuye abruptamente para los últimos días. Este es un comportamiento que se repite en todas las actualizaciones del dataset del Ministerio de Salud y está relacionado con la demora en la carga de datos. El análisis de la demora requiere el desarrollo de un apartado particular y se encuentra fuera del análisis de este informe. De todas maneras, en la siguiente página se puede encontrar información al respecto: https://covidstats.com.ar/reportediario
#
# A modo de ejemplo, para el día de análisis de los datos, 23 de Septiembre, la carga de fallecimientos en función de la fecha de deceso es la siguiente:
#
# <img src="./extras/carga_casos.png" alt="Carga de Casos">
#
# Para la provincia de Buenos Aires, existe una demora de prácticamente un mes en la carga de datos, algo consistente, se repite diariamente este fenómeno. Lamentablemente esto impacta negativamente en la calidad de las predicciones del pico de la pandemia en Argentina, al menos de esta primer ola de contagios y fallecidos.
#
# Mas allá de este fenómeno de demora para la provincia de Buenos Aires, podemos observar que la carga de datos suele reunir las incidencias de los últimos 5 días. Es por esto que vamos a quitar estos días para hacer el análisis correspondiente.
# # Ajuste de curva Log Normal
# +
# Cast column values and set first day as #1
dailyDeathsMS['DoY'] = dailyDeathsMS.DoY.astype(int)
dailyDeathsMS['DoS'] = dailyDeathsMS.DoY.apply(lambda x: x - 66)
dailyDeathsMS.head()
# Normalize values to estimate fitting correctly and delete last days from analysis
x_data = dailyDeathsMS.DoS[:-5]
y_data = dailyDeathsMS.fallecido[:-5]
xdata =x_data/max(x_data)
ydata =y_data/max(y_data)
# +
# Find best parameters which optimizes the curve
from scipy.optimize import curve_fit
popt, pcov = curve_fit(log_normal, xdata, ydata)
print(popt)
# +
# Generate the fitting curve points
xfit = np.arange(0.005, 3, 0.005)
yfit = [log_normal(x, *popt) for x in xfit]
# Convert to numpy array to multiply it by a constant
xfit = np.asarray(xfit)
yfit = np.asarray(yfit)
# Re-scale arrays
xdata = xdata * max(x_data)
xfit = xfit * max(x_data)
ydata = ydata * max(y_data)
yfit = yfit * max(y_data)
plt.title('Ajuste con función log normal en base a Min. de Salud')
plt.ylabel('fallecidos')
plt.xlabel('Días desde inicio de epidemia')
plt.rcParams['figure.dpi'] = 50
p0 = plt.plot(xdata, ydata, 'r.')
p1 = plt.plot(xfit, yfit)
plt.legend((p0[0], p1[0]), ('Fallecidos diarios', 'Curva de ajuste'))
plt.show()
# Find the curve peak
maxNumCases = max(yfit)
dayMaxCases = xfit[np.where(yfit == maxNumCases)][0]
print("Max number of cases on DoY {} with {} deaths".format(dayMaxCases+66, maxNumCases))
# -
# <h2> Datos de COVID Stats AR </h2>
#
# A modo de referencia, es interesante analizar los resultados teniendo en cuenta los datos publicados en COVID Stats. Podemos exportar los datos a un csv accediendo directamente a https://covidstats.com.ar/exportar
covidStats = pd.read_csv("2020-09-25_Nacional.csv", skiprows=3)
covidStats.head()
# Limpiamos los datos para ajustarlos al formato de interés
covidStats['fecha'] = pd.to_datetime(covidStats['fecha'], format='%Y-%m-%d')
dailyDeathsCS = covidStats.loc[covidStats.fallecidos > 0]
dailyDeathsCS['DoY'] = [date.strftime('%j') for date in dailyDeathsCS.fecha]
dailyDeathsCS['DoY'] = dailyDeathsCS.DoY.astype(int)
dailyDeathsCS['DoS'] = [(i - 66) for i in dailyDeathsCS.DoY]
dailyDeathsCS = dailyDeathsCS[['fecha', 'fallecidos', 'DoY', 'DoS']]
dailyDeathsCS.tail(14)
# Aparentemente los datos son los mismos, veamos qué predicción obtenemos a partir del ajuste con la curva Log Normal
# +
x_data = dailyDeathsCS.DoS[:-5]
y_data = dailyDeathsCS.fallecidos[:-5]
xdata =x_data/max(x_data)
ydata =y_data/max(y_data)
popt, pcov = curve_fit(log_normal, xdata, ydata)
# Generate fitting curve
xfit = np.arange(0.005, 3, 0.005)
yfit = [log_normal(x, *popt) for x in xfit]
# Convert to numpy array to multiply it by a constant
xfit = np.asarray(xfit)
yfit = np.asarray(yfit)
# Re-scale arrays
xdata = xdata * max(x_data)
xfit = xfit * max(x_data)
ydata = ydata * max(y_data)
yfit = yfit * max(y_data)
plt.title('Ajuste con función log normal en base a COVID Stats AR')
plt.ylabel('Fallecidos')
plt.xlabel('Días desde inicio de epidemia')
plt.rcParams['figure.dpi'] = 50
p0 = plt.plot(xdata, ydata, 'r.')
p1 = plt.plot(xfit, yfit)
plt.legend((p0[0], p1[0]), ('Fallecidos diarios', 'Curva de ajuste'))
plt.show()
maxNumCases = max(yfit)
dayMaxCases = xfit[np.where(yfit == maxNumCases)][0]
print("Max number of cases on day {} with {} deaths".format(dayMaxCases+66, maxNumCases))
# -
# Evidentemente, los resultados son los mismos, esto es interesante ya que puede usarse el dataset de COVID Stats alternativamente para el modelaje de la curva, siendo esta base de datos mucho más liviana ya que la información es previamente procesada.
# <h2> Ajustes con la función logística </h2>
#
# Es común observar ajustes de infección y muertes realizados con esta curva, ya que su comportamiento muestra un punto de inflexión donde la curva cambia de velocidad. Este punto corresponde con el pico de contagios o muertes tal como suele observarse en la distribución de casos diarios.
#
# Para utilizar esta curva en particular es importante primero, tener una columna de los casos acumulados. Como las curvas de interés son analizadas en un marco temporal diario, acumularemos dichos resultados de dicha manera.
# Definimos entonces, una función para poder acumular los casos en una nueva columna "fall_tot"
# +
def acum_deaths(acumArray):
acum = []
count = 0
for f in acumArray:
count = f + count
acum.append(count)
return acum
dailyDeathsMS['fall_tot'] = acum_deaths(dailyDeathsMS.fallecido)
dailyDeathsCS['fall_tot'] = acum_deaths(dailyDeathsCS.fallecidos)
# -
# Definimos la función logística, también conocida como función sigmoid
# +
def sigmoid(x, Beta_1, Beta_2):
y = 1 / (1 + np.exp(-Beta_1*(x-Beta_2)))
return y
def lin_reg(x, a, b, c):
y = c / (1 + a * np.exp(-b*x))
return y
# +
x_data = dailyDeathsMS.DoS[:-5]
y_data = dailyDeathsMS.fall_tot[:-5]
yd_data = dailyDeathsMS.fallecido[:-5]
xdata =x_data/max(x_data)
ydata =y_data/max(y_data)
p0 = np.random.exponential(size=3)
bounds = (0, [10000, 50, 10])
popt, pcov = curve_fit(lin_reg, xdata, ydata, p0=p0, bounds=bounds)
print(*popt)
xfit = np.arange(0.005, 1.5, 0.005)
yfit = [lin_reg(x, *popt) for x in xfit]
# Convert to numpy array to multiply it by a constant
xfit = np.asarray(xfit)
yfit = np.asarray(yfit)
# Re-scale arrays
xdata = xdata * max(x_data)
xfit = xfit * max(x_data)
ydata = ydata * max(y_data)
yfit = yfit * max(y_data)
dydx_yfit = np.gradient(yfit)
#fig, axs = plt.subplots(1, 2, figsize=(20, 10))
plt.subplots(1, 2, figsize=(20, 10))
plt.subplot(121)
plt.plot(xdata, ydata, 'r.', label='total de fallecidos')
plt.plot(xfit, yfit, label='curva de ajuste')
plt.legend()
plt.ylabel('total de fallecidos')
plt.xlabel('Días desde inicio de epidemia')
plt.rcParams['figure.dpi'] = 50
plt.subplot(122)
plt.plot(xdata, yd_data, 'r.', label='fallecidos diarios')
plt.plot(xfit, dydx_yfit, label='curva de ajuste')
plt.legend()
plt.ylabel('fallecidos diarios')
plt.xlabel('Días desde inicio de epidemia')
plt.rcParams['figure.dpi'] = 50
plt.suptitle('Ajuste con modelo de regresión logística')
plt.show()
# -
maxNumCases = max(dydx_yfit)
dayMaxCases = xfit[np.where(dydx_yfit == maxNumCases)][0]
print("Max number of cases on day {} with {} deaths".format(dayMaxCases+66, maxNumCases))
# Existe un estudio de la Escuela de Ingeniería de Información de la Universidad de Geociencias de China que realiza predicciones basada en este modelo de función logística https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7328553/
#
# El comportamiento de los casos acumulados ajusta bien con la curva, como puede observarse, la derivada de la función logística -gráfico a la derecha- muestra el comportamiento de los casos diarios. Como vemos también ajusta muy bien a las actualizaciones diarias del número de fallecidos. En este caso, la predicción del modelo muestra que el pico de fallecidos se obtuvo el día 235 (22 de Agosto) con 181 decesos.
# # Mapa de Densidad de Fallecidos por Departamento
#
# Analizaremos en este apartado la distribución geográfica de fallecidos. En particular estamos interesados en el comportamiento de la pandemia en zonas densamente pobladas, donde estimamos que puede existir una mayor velocidad de contagio.
#
# Comenzaremos seleccionando los campos relevantes del dataset del Ministerio de Salud. Lamentablemente, no disponemos de datos con mayor granularidad que la clasificación de cada caso por departamento de cada provincia. Esto incluye a su vez, las comunas de CABA.
classFilter = (df.fallecido == 1) & (df.clasificacion_resumen == 'Confirmado')
deadMS = df.loc[classFilter]
deadMS = deadMS[['residencia_provincia_nombre', 'residencia_departamento_nombre', 'fallecido', 'residencia_provincia_id',
'residencia_departamento_id']]
# <h2> Data Cleaning </h2>
#
# Vamos a generar un campo con un código de identificación único para cada departamento, basado en la nomenclatura utilizada por el INDEC y el archivo con puntos geográficos que utilizaremos posteriormente.
# +
def make_in1(row):
idProv = row[3]
idDpto = row[4]
return(f'{idProv:02}{idDpto:03}')
deadMS['in1'] = [make_in1(row) for row in deadMS.values]
# -
# Algunas provincias no disponen del departamento de residencia del fallecido. El criterio utilizado es repartir los casos proporcionalmente en todas las demás comunas o departamentos. Si bien la decisión puede ser cuestionada, es una manera de no perder datos y distribuir los mismos de manera balanceada.
# +
provSinEsp = deadMS.loc[deadMS.residencia_departamento_nombre == 'SIN ESPECIFICAR']\
.groupby('residencia_provincia_nombre').sum().index
provSinEsp = list(provSinEsp)
if 'SIN ESPECIFICAR' in provSinEsp:
# Remove from provinces list
provSinEsp.remove('SIN ESPECIFICAR')
try:
# Delete all ocurrencies from dataset where not province specified
sinEsp = deadMS[deadMS['residencia_provincia_nombre'] == 'SIN ESPECIFICAR'].index
deadMS.drop(sinEsp, inplace=True)
except:
print('province sin especificar not found')
deadMSGrouped = deadMS.groupby(['residencia_provincia_nombre', 'residencia_departamento_nombre', 'in1'])\
.sum().sort_values('fallecido', ascending=False)
for provincia in provSinEsp:
totDeadProv = deadMSGrouped.loc[provincia, ].sum().fallecido
notSpecProv = deadMSGrouped.loc[provincia, 'SIN ESPECIFICAR', ].fallecido[0]
realDeadProv = totDeadProv - notSpecProv
for i, value in enumerate(deadMSGrouped.loc[provincia, ].values):
val = value[0]
index = deadMSGrouped.loc[provincia, ].index[i]
if index[0] != 'SIN ESPECIFICAR':
# Add proportionally the values in 'SIN ESPECIFICAR'
val = (val/realDeadProv) * (notSpecProv) + val
deadMSGrouped.loc[provincia, index[0], index[1]][0] = int(val)
deadMSGrouped.drop((provincia, 'SIN ESPECIFICAR'), inplace=True)
deadMSGrouped.reset_index(inplace=True)
# -
# Carguemos ahora los datos de población por cada departamento. Los datos se pueden encontrar en la página del Indec, luego se exporta una planilla de cálculos. En el repositorio hay un script para tomar los datos de dicha planilla y generar el csv correspondiente. https://redatam.indec.gob.ar/argbin/RpWebEngine.exe/PortalAction?&MODE=MAIN&BASE=CPV2010B&MAIN=WebServerMain.inl&_ga=2.177965477.1629507683.1526925251-993948438.1526925251
#
# Los parámetros utilizados en la generación del documento son: edades quinquenales, departamento/partido, y el total de toda la base.
# +
def get_population(code):
row = pobDeptos.loc[pobDeptos.in1 == code]
try:
return(row.iloc[0, 2])
except:
print(f'Population not found for {code}')
pobDeptos = pd.read_csv('poblacion_dpto.csv')
# Standarize codes to 5 digits with zero padding
pobDeptos['in1'] = [f'{code:05}' for code in pobDeptos.in1]
# Clean dataset and free some resources
deadMS = deadMSGrouped.drop(columns=['residencia_provincia_id','residencia_departamento_id'])
del(deadMSGrouped)
deadMS.rename(columns={'residencia_provincia_nombre': 'provincia',
'residencia_departamento_nombre': 'departamento',
'fallecido': 'fallecidos'},
inplace=True)
deadMS['poblacion'] = [get_population(in1) for in1 in deadMS.in1]
deadMS['deathRatio'] = [(fallecidos*10000/poblacion) for fallecidos, poblacion in \
zip(deadMS.fallecidos, deadMS.poblacion)]
deadMS.head()
# +
# Data cleaning of wrong inputs
deptos = ['06466', '38182']
for d in deptos:
print(deadMS.loc[deadMS.in1 == d].index)
# 272 258
deadMS.drop(index=[177, 240], inplace=True)
# -
# Finalmente, generemos un listado de los departamentos con mayor cantidad de muertos por 10 mil habitantes.
deadMS.sort_values('deathRatio', ascending=False).head(30)
# Graficamos los departamentos con la capa de densidad de muertos cada 10 mil habitantes. El archivo GeoJson puede ser descargado de https://ramsac.ign.gob.ar/api/v1/capas-sig/Geodesia+y+demarcaci%C3%B3n/L%C3%ADmites/departamento/json
#
# Sin embargo, la codificación no es totalmente compatible con la utilizada por INDEC. Algunos códigos, en particular los correspondientes a CABA no coinciden y han tenido que ser editados a mano. Se puede encontrar el GeoJson utilizado en la carpeta /extras
#
# Tener en cuenta que renderizar el mapa consume muchos recursos. El requisito es tener al menos <b> 1GB de RAM disponible.</b>
import folium
# +
argMap = folium.Map(
location=[-40, -61],
zoom_start=4 # Limited levels of zoom for free Mapbox tiles.
)
departments = 'departamento.json'
folium.Choropleth(
geo_data=departments,
name='choropleth',
data=deadMS,
columns=['in1', 'deathRatio'],
key_on='properties.in1',
fill_color='YlGn',
fill_opacity=0.7,
line_opacity=0.2,
nan_fill_color='white',
nan_fill_opacity=0.7,
legend_name='Fallecidos cada 10 mil habitantes'
).add_to(argMap)
#argMap # <-- uncomment if you want to display it in the notebook, otherwise it will be saved to disk
argMap.save('fallecidosArg.html')
# -
| augusto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import sys
import os
import argparse
import logging
import shutil
import re
from PIL import Image
from skimage import io
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import torch
import torch.nn.functional as F
import torchvision.models as models
import torch.optim as optim
from torchvision.datasets import VisionDataset
from torchvision import transforms
import torch.backends.cudnn as cudnn
from torch.utils.data import DataLoader
from sklearn.model_selection import train_test_split
from utils import *
from auto_tqdm import tqdm
import augmentations
# %matplotlib inline
DATA_DIR = '/data/amiratag/prostate'
MEM_DIR = '/home/amiratag/Projects/sslp'
DATA_DIR = "/export/medical_ai/kaggle_panda" # Stores data related to images
MEM_DIR = "/export/home/code/metamind/precision_oncology/simclr" #stores data related to model checkpoints
# -
def return_embeddings(slide, mask):
''' Function that transforms a slide into the tensor with 128 channels.'''
width, height = slide.shape[:2]
embeddings = np.zeros((width // im_shape[0], height // im_shape[1], embedding_size)).astype(np.float32)
batch, w_idxs, h_idxs = [], [], []
for w in range(embeddings.shape[0]):
for h in range(embeddings.shape[1]):
tile_mask = mask[w * im_shape[0]: (w+1) * im_shape[0], h * im_shape[1]: (h+1) * im_shape[1], 0]
if np.mean(np.equal(tile_mask, 0)) < 0.95:
tile = slide[w * im_shape[0]: (w+1) * im_shape[0], h * im_shape[1]: (h+1) * im_shape[1]] / 255.
batch.append(torch.tensor(tile.astype(np.float32).transpose(2,0,1)))
w_idxs.append(w)
h_idxs.append(h)
if len(batch) == batch_size or \
(w == embeddings.shape[0] - 1 and h == embeddings.shape[1] - 1):
if not len(batch):
continue
X = torch.stack(batch).to(device)
pred = model(X).detach().cpu().numpy()
embeddings[w_idxs, h_idxs] = pred
batch, w_idxs, h_idxs = [], [], []
return embeddings
# Read data
# +
dataset = pd.read_csv(os.path.join(DATA_DIR, 'train.csv'))
valid_idxs = np.array([i for i in range(len(dataset)) if dataset['data_provider'][i] == 'radboud' and
os.path.exists(os.path.join(DATA_DIR, 'train_images', dataset['image_id'][i])+'.tiff')])
train_idxs = valid_idxs[:-1000]
test_idxs = valid_idxs[-1000:]
train_slides = dataset['image_id'][train_idxs].values
train_isup_dic = dict(zip(train_slides, dataset['isup_grade'][train_idxs]))
test_slides = dataset['image_id'][test_idxs].values
test_isup_dic = dict(zip(test_slides, dataset['isup_grade'][test_idxs]))
# -
cudnn.benchmark = True
use_cuda = torch.cuda.is_available()
device = torch.device('cuda') if use_cuda else 'cpu'
# Load and run patch level model
model_dir = os.path.join(MEM_DIR, 'simclr/translationrotation')
im_shape = (224, 224)
embedding_size = 128
model = ResNetSimCLR('resnet34', embedding_size)
if use_cuda:
model = torch.nn.DataParallel(model).cuda()
checkpoints = [i for i in os.listdir(model_dir) if '.pt' in i]
checkpoints_epoch_num = [int(re.search('epoch(\d+)', c).group(1)) for c in checkpoints]
last_epoch = np.max(checkpoints_epoch_num)
checkpoint = torch.load(os.path.join(model_dir, 'checkpoint-epoch{}.pt'.format(last_epoch)))
print('Loading checkpoint from epoch #{}'.format(last_epoch))
state_dict = checkpoint.get('state_dict', checkpoint)
model.load_state_dict(state_dict)
model.eval()
# Computed and save training slides embeddings
batch_size = 320
overwrite = False
save_dir = 'simclr_translationrotation_predicted_masks'
if not os.path.exists(save_dir):
os.makedirs(save_dir)
for i, train_slide in tqdm(enumerate(train_slides)):
image_path = os.path.join(DATA_DIR, 'train_images', train_slide)+'.tiff'
mask_path = os.path.join(DATA_DIR, 'train_label_masks', train_slide)+'_mask.tiff'
save_path = os.path.join(save_dir, train_slide + '.npy')
if os.path.exists(save_path) and not overwrite:
continue
try:
slide = io.imread(image_path)
mask = io.imread(mask_path)
except:
continue
embeddings = return_embeddings(slide, mask)
np.save(save_path, embeddings)
# Computed and save test slides embeddings
save_dir = 'test_simclr_translationrotation_predicted_masks'
if not os.path.exists(save_dir):
os.makedirs(save_dir)
for i, test_slide in tqdm(enumerate(test_slides)):
image_path = os.path.join(DATA_DIR, 'train_images', test_slide)+'.tiff'
mask_path = os.path.join(DATA_DIR, 'train_label_masks', test_slide)+'_mask.tiff'
save_path = os.path.join(save_dir, test_slide + '.npy')
if os.path.exists(save_path) and not overwrite:
continue
try:
slide = io.imread(image_path)
mask = io.imread(mask_path)
except:
continue
embeddings = return_embeddings(slide, mask)
np.save(save_path, embeddings)
# Create slide-level dataset
image_names = os.listdir('simclr_translationrotation_predicted_masks')
X, y = [], []
for image_name in train_slides:
pm_path = os.path.join('simclr_translationrotation_predicted_masks', image_name+'.npy')
if os.path.exists(pm_path):
X.append(pm_path)
y.append(train_isup_dic[image_name])
X_test, y_test = [], []
for image_name in test_slides:
pm_path = os.path.join('test_simclr_translationrotation_predicted_masks', image_name+'.npy')
if os.path.exists(pm_path):
X_test.append(pm_path)
y_test.append(test_isup_dic[image_name])
X, y, X_test, y_test = np.array(X), np.array(y), np.array(X_test), np.array(y_test)
# Slide-level ConvNet
# +
class CustomDataset(VisionDataset):
def __init__(self, X, y):
super(CustomDataset, self).__init__('', transform=None,
target_transform=None)
self.X = X
self.y = y
assert len(self.X) == len(self.y)
def __len__(self):
return len(self.X)
def __getitem__(self, index):
img, target = self.X[index], self.y[index]
output = np.zeros((224, 224, 128)).astype(np.float32)
loaded_image = np.load(img)
shape = loaded_image.shape[:2]
r1 = np.random.choice(224 - shape[0])
r2 = np.random.choice(224 - shape[1])
output[r1:loaded_image.shape[0]+r1, r2:loaded_image.shape[1]+r2] = loaded_image
output = torch.Tensor(output.transpose((2,0,1)))
return output, target
def extra_repr(self):
return "Split: {}".format("Train" if self.train is True else "Test")
def train(device, model, train_loader, optimizer, epoch):
model.train()
train_metrics = []
for batch_idx, (data, target) in enumerate(train_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
logits = model(data)
loss = F.cross_entropy(logits, target)
loss.backward()
optimizer.step()
train_metrics.append(dict(
epoch=epoch,
loss=loss.item()))
print(
'Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader) * batch_size,
100. * batch_idx / len(train_loader), loss.item()))
return train_metrics
class ConvLayer(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size, stride=1):
super().__init__()
self.conv = torch.nn.Conv2d(in_channels, out_channels, kernel_size=kernel_size,
stride=stride)
self.bn = torch.nn.BatchNorm2d(out_channels)
self.relu = torch.nn.ReLU(inplace=True)
def forward(self, x):
h = self.conv(x)
h = self.bn(h)
h = self.relu(h)
return h
class MiniModel(nn.Module):
def __init__(self, num_classes):
super().__init__()
self.cl1_1 = ConvLayer(128, 128, 3, 1)
self.cl1_2 = ConvLayer(128, 128, 3, 1)
self.cl2_1 = ConvLayer(128, 128, 3, 1)
self.cl2_2 = ConvLayer(128, 128, 3, 1)
self.cl3_1 = ConvLayer(128, 128, 3, 1)
self.cl3_2 = ConvLayer(128, 128, 3, 1)
self.fc = torch.nn.Linear(128, num_classes)
def forward(self, x):
h = self.cl1_1(x)
h = self.cl1_2(h)
h = torch.nn.MaxPool2d(3, 3)(h)
h = self.cl2_1(h)
h = self.cl2_2(h)
h = torch.nn.MaxPool2d(3, 3)(h)
h = self.cl3_1(h)
h = self.cl3_2(h)
h = torch.nn.MaxPool2d(3, 3)(h)
h = torch.mean(h, axis=(-1, -2))
return self.fc(h)
# -
batch_size = 64
train_dataset = CustomDataset(X, y)
test_dataset = CustomDataset(X_test, y_test)
kwargs = {'num_workers': 12, 'pin_memory': False} if use_cuda else {}
kwargs = {}
train_loader = DataLoader(train_dataset, batch_size=batch_size, shuffle=True, drop_last=True, **kwargs)
test_loader = DataLoader(test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
model = MiniModel(num_classes=6)
if use_cuda:
model = torch.nn.DataParallel(model).cuda()
optimizer = optim.SGD(model.parameters(), lr=0.05,
momentum=0.9,
weight_decay=5e-4,
nesterov=True)
for epoch in range(0, 50):
if epoch % 5 == 0:
model.eval()
true, predicted = [], []
for batch_idx, (data, target) in enumerate(test_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
logits = model(data)
predicted.append(logits.argmax(-1).detach().cpu().numpy())
true.append(target.detach().cpu().numpy())
print(np.mean(np.concatenate(predicted) == np.concatenate(true)))
lr = 0.05 * 0.5 * (1 + np.cos((epoch - 1) / 50 * np.pi))
print('Setting learning rate to %g' % lr)
for param_group in optimizer.param_groups:
param_group['lr'] = lr
train(device, model, train_loader, optimizer, epoch)
model.eval()
true, predicted = [], []
for batch_idx, (data, target) in enumerate(test_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
logits = model(data)
predicted.append(logits.argmax(-1).detach().cpu().numpy())
true.append(target.detach().cpu().numpy())
print(np.mean(np.concatenate(predicted) == np.concatenate(true)))
from sklearn.metrics import confusion_matrix
fig = plt.figure(figsize=(8, 8))
cm = confusion_matrix(np.concatenate(true), np.concatenate(predicted))
normalized_cm = cm/np.sum(cm, -1, keepdims=True)
plt.imshow(normalized_cm, vmin=0., vmax=1)
plt.imshow(normalized_cm, vmin=0., vmax=1)
plt.colorbar()
plt.xticks(np.arange(6), ['ISUP {}'.format(i) for i in range(6)], fontsize=12)
plt.yticks(np.arange(6), ['ISUP {}'.format(i) for i in range(6)], fontsize=12)
plt.xlabel('Predicted', fontsize=15)
plt.ylabel('True', fontsize=15)
plt.title('Prediction Accuracy = {0:.1f}%'.format(100 * np.mean(np.concatenate(true) == np.concatenate(predicted))), fontsize=15)
for i in range(6):
for j in range(6):
plt.annotate('{0:.2f}%'.format(100 * normalized_cm[i, j]), (j-0.3, i+0.1), color='white', fontsize=12)
fig = plt.figure(figsize=(30, 30))
isups = np.array([train_isup_dic[train_slide] for train_slide in train_slides])
for i in range(6):
idxs = np.where(isups == i)[0]
for j in range(3):
plt.subplot(6, 3, 3 * i + j + 1)
slide = io.imread(os.path.join(DATA_DIR, 'train_images', train_slides[np.random.choice(idxs)]) + '.tiff')
plt.imshow(Image.fromarray(slide).resize((slide.shape[1] // 20, slide.shape[0]//20)))
plt.axis('off')
# Semi-supervised learning (assume 10000 tiles are labeled)
from sklearn.cluster import KMeans
predicted_masks = os.listdir(save_dir)
all_dims = []
for predicted in tqdm(predicted_masks):
all_dims.append(np.load(os.path.join(save_dir, predicted)).reshape((-1, 128)))
all_dims = np.concatenate(all_dims)
km = KMeans(50, verbose=3, max_iter=20, n_init=1)
km.fit(all_dims)
centers = km.cluster_centers_
profiles = np.zeros((len(predicted_masks), len(centers)))
labels = []
for i, predicted in tqdm(enumerate(predicted_masks)):
predicted_mask = np.load(os.path.join(save_dir, predicted)).reshape((-1, 128))
distances = np.linalg.norm(np.expand_dims(predicted_mask, 1) - np.expand_dims(centers, 0), axis=-1)
assignments = np.argmin(distances, -1)
idxs, counts = np.unique(assignments, return_counts=True)
profiles[i, idxs] = counts / np.sum(counts)
labels.append(isup_dic[predicted.replace('.npy', '')])
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
lr = LogisticRegression()
cross_val_score(lr, np.stack(profiles, 0), np.stack(labels, 0), cv=5)
representations = []
tiles = []
for i in tqdm(range(1000)):
image_path, mask_path = image_paths[i], mask_paths[i]
image_name = image_path.split('/')[-1].split('.')[0]
try:
slide = io.imread(image_path)
mask = io.imread(mask_path)
except:
continue
if os.path.exists(save_path) and not overwrite:
continue
size = slide.shape
while True:
w = np.random.choice(size[0] // im_shape[0])
h = np.random.choice(size[1] // im_shape[1])
msk = mask[w * im_shape[0]: (w+1) * im_shape[0], h * im_shape[1]: (h+1) * im_shape[1]][..., 0]
if np.mean(np.equal(msk, 0)) > 0.95:
continue
tile = slide[w * im_shape[0]: (w+1) * im_shape[0], h * im_shape[1]: (h+1) * im_shape[1]] / 255.
tiles.append(tiles)
X = torch.stack([torch.tensor(tile.astype(np.float32).transpose(2,0,1))])
if use_cuda:
X = X.to(device)
representations.append(model(X).detach().cpu().numpy())
break
# + active=""
# representations = np.concatenate(representations)
# tiles = np.concatenate(tiles)
# distances = np.linalg.norm(np.expand_dims(representations, 1) - np.expand_dims(centers, 0), axis=-1)
# assignments = np.argmin(distances, -1)
# -
fig = plt.figure(figsize=(12, 12))
i = 0
idxs = np.where(assignments == i)[0]
for n in range(16):
plt.subplot(4, 4, n+1)
plt.imshow(tiles[idxs[n]])
plt.axis('off')
i = 10
fig = plt.figure(figsize=(20, 20))
image_name = image_paths[i].split('/')[-1].split('.')[0]
msk = io.imread(mask_paths[i])
img = io.imread(image_paths[i])
size = msk.shape
plt.subplot(1, 3, 3)
plt.title('Predicted Map', fontsize=25)
predicted_mask = np.load(os.path.join('predicted_masks', image_name+'.npy'))
plt.imshow((predicted_mask * 60).astype(np.uint8), cmap='hot')
plt.axis('off')
plt.subplot(1, 3, 1)
plt.title('Slide', fontsize=25)
plt.imshow(np.array(Image.fromarray(img).resize((size[1]//10, size[0]//10))))
plt.axis('off')
plt.subplot(1, 3, 2)
msk = io.imread(mask_paths[i])
size = msk.shape
plt.title('True Map', fontsize=25)
plt.imshow(np.array(Image.fromarray(msk * 60).resize((size[1]//10, size[0]//10)))[..., 0], cmap='hot')
plt.axis('off')
| SSL/simclr/simclr_run-PANDAs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://github.com/lab-ml/python_autocomplete)
# [](https://colab.research.google.com/github/lab-ml/python_autocomplete/blob/master/notebooks/evaluate.ipynb)
#
# # Evaluate a model trained on predicting Python code
#
# This notebook evaluates a model trained on Python code.
#
# Here's a link to [training notebook](https://github.com/lab-ml/python_autocomplete/blob/master/notebooks/train.ipynb)
# [](https://colab.research.google.com/github/lab-ml/python_autocomplete/blob/master/notebooks/train.ipynb)
# ### Install dependencies
# !pip install labml labml_python_autocomplete
# Imports
# +
import string
import torch
from torch import nn
import numpy as np
from labml import experiment, logger, lab
from labml_helpers.module import Module
from labml.analytics import ModelProbe
from labml.logger import Text, Style, inspect
from labml.utils.pytorch import get_modules
from labml.utils.cache import cache
from labml_helpers.datasets.text import TextDataset
from python_autocomplete.train import Configs
from python_autocomplete.evaluate import Predictor
from python_autocomplete.evaluate.beam_search import NextWordPredictionComplete
# -
# We load the model from a training run. For this demo I'm loading from a run I trained at home.
#
# [](https://web.lab-ml.com/run?uuid=39b03a1e454011ebbaff2b26e3148b3d)
#
# If you have a locally trained model load it directly with:
#
# ```python
# run_uuid = 'RUN_UUID'
# checkpoint = None # Get latest checkpoint
# ```
#
# `load_bundle` will download an archive with a saved checkpoint (pretrained model).
# +
# run_uuid = 'a6cff3706ec411ebadd9bf753b33bae6'
# checkpoint = None
run_uuid, checkpoint = experiment.load_bundle(
lab.get_path() / 'saved_checkpoint.tar.gz',
url='https://github.com/lab-ml/python_autocomplete/releases/download/0.0.5/bundle.tar.gz')
# -
# We initialize `Configs` object defined in [`train.py`](https://github.com/lab-ml/python_autocomplete/blob/master/python_autocomplete/train.py).
conf = Configs()
# Create a new experiment in evaluation mode. In evaluation mode a new training run is not created.
experiment.evaluate()
# Load custom configurations/hyper-parameters used in the training run.
custom_conf = experiment.load_configs(run_uuid)
custom_conf
# Set the custom configurations
# +
# custom_conf['device.use_cuda'] = False
# -
experiment.configs(conf, custom_conf)
# Set models for saving and loading. This will load `conf.model` from the specified run.
experiment.add_pytorch_models({'model': conf.model})
# Specify which run to load from
experiment.load(run_uuid, checkpoint)
# Start the experiment
experiment.start()
# Initialize the `Predictor` defined in [`evaluate.py`](https://github.com/lab-ml/python_autocomplete/blob/master/python_autocomplete/evaluate.py).
#
# We load `stoi` and `itos` from cache, so that we don't have to read the dataset to generate them. `stoi` is the map for character to an integer index and `itos` is the map of integer to character map. These indexes are used in the model embeddings for each character.
p = Predictor(conf.model, conf.text.tokenizer,
state_updater=conf.state_updater,
is_token_by_token=conf.is_token_by_token)
# Set model to evaluation mode
_ = conf.model.eval()
# Setup probing to extract attentions
probe = ModelProbe(conf.model)
# A python prompt to test completion.
PROMPT = """from torch import nn
from labml_helpers.module import Module
from labml_nn.lstm import LSTM
class LSTM(Module):
def __init__(self, *,
n_tokens: int,
embedding_size: int,
hidden_size int,
n_layers int):
"""
# Get a token. `get_token` predicts character by character greedily (no beam search) until it find and end of token character (non alpha-numeric character).
stripped, prompt = p.rstrip(PROMPT)
rest = PROMPT[len(stripped):]
prediction_complete = NextWordPredictionComplete(rest, 5)
prompt = torch.tensor(prompt, dtype=torch.long).unsqueeze(-1)
# %%time
predictions = p.get_next_word(prompt, None, rest, [1.], prediction_complete, 5)
predictions.sort(key=lambda x: -x[0])
[(pred.prob, pred.text[len(rest):]) for pred in predictions]
# ## Lets analyze attentions
tokens = [p.tokenizer.itos[i[0]] for i in prompt]
inspect(tokens)
# Lets run the transformer XL model without cached memory to get the full attention matrix
inspect(p._get_predictions(prompt, None)[0])
# We capture the outputs after the [attention softmax](https://nn.labml.ai/transformers/mha.html#section-34)
inspect(probe.forward_output['*softmax*'])
attn = probe.forward_output['*softmax*'].get_list()
# Attentions have shape `[source, destination, batch, heads]`
inspect(attn[0].shape)
attn_maps = torch.stack([a.permute(2, 3, 0, 1)[0] for a in attn])
np.save('attentions.npy', attn_maps.detach().numpy())
| notebooks/evaluate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/angelabaltes/FindmyFigurine/blob/main/findmyfigurineapp.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pzGV21wmArkc" outputId="b4a3100e-789b-4b35-bbfd-266e766cdd2e" colab={"base_uri": "https://localhost:8080/", "height": 35}
# %cd /content/drive/My Drive/Insight/moeimouto-faces
# + id="HWkej4BuA1PX" outputId="406998f1-826a-4213-f6f2-2e446fbc87d5" colab={"base_uri": "https://localhost:8080/", "height": 121}
# !pip install -q streamlit
# !pip install pyngrok
# !pip install classify
# + id="5cy4dbERA1T-"
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.applications.inception_v3 import preprocess_input
import PIL
from pyngrok import ngrok
from tensorflow.keras.applications.inception_v3 import InceptionV3
# + id="AkRqpPJlA1WO" outputId="1eed4f90-c829-42f1-87c4-b299e8326d7e" colab={"base_uri": "https://localhost:8080/", "height": 35}
# %%writefile findmyfigurineapp.py
import streamlit as st
import tensorflow as tf
import numpy as np
from pyngrok import ngrok
import cv2
from PIL import Image, ImageOps
from keras.applications.inception_v3 import decode_predictions
model = tf.keras.models.load_model('my_model.hdf7')
st.title("""
FindmyFigurine
"""
)
st.write("This is an Image Classification Web App to Predict Anime Characters")
st.set_option('deprecation.showfileUploaderEncoding', False)
uploaded_file= st.file_uploader("Please upload an image file of an anime character", type=["jpg", "png"])
def import_and_predict(image_data, model):
size = (150,150)
image = ImageOps.fit(image_data, size, Image.ANTIALIAS)
image = np.asarray(image)
img = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
img_resize = (cv2.resize(img, dsize=(75, 75), interpolation=cv2.INTER_CUBIC))/255.
img_reshape = img_resize[np.newaxis,...]
prediction = model.predict(img_reshape)
return prediction
if uploaded_file is not None:
image = Image.open(uploaded_file)
st.image(image, caption='Uploaded Image.', use_column_width=True)
st.write("")
st.write("Classifying...")
prediction = import_and_predict(image, model)
if np.argmax(prediction) == 0:
st.write("**Aika Granzchesta!**")
st.write("Select a link to find a figurine!")
st.write("https://solarisjapan.com/search#q=Aika%20Granzchesta&page=0")
st.write("https://www.ebay.com/sch/i.html?_from=R40&_trksid=p2047675.m570.l1313&_nkw=aika+granzchesta&_sacat=0")
st.write("https://japanimedia-ex.com/aria-the-natural-aika-s-granzchesta-1-6-figure-solid-works-japan-anime-manga/")
if np.argmax(prediction) == 1:
st.write("**Aisaka Taiga!**")
st.write("Select a link to find a figurine!")
st.write("https://www.amazon.com/s?k=Aisaka+Taiga+figurine&ref=nb_sb_noss_2")
st.write("https://www.cosplayshow.com/TIGER-DRAGON-Aisaka-Taiga-Vinyl-PVC-Anime-Action-Figure-p144936.html")
st.write("ebay.com/sch/i.html?_from=R40&_trksid=p2334524.m570.l2632&_nkw=aisaka+taiga+figure&_sacat=13658&LH_TitleDesc=0&_osacat=0&_odkw=aika+granzchesta")
if np.argmax(prediction) == 2:
st.write("<NAME>!")
if np.argmax(prediction) == 3:
st.write("Akiy<NAME>!")
if np.argmax(prediction) == 4:
st.write("<NAME>!")
if np.argmax(prediction) == 5:
st.write("<NAME>!")
if np.argmax(prediction) == 6:
st.write("<NAME>!")
if np.argmax(prediction) == 7:
st.write("<NAME>!")
if np.argmax(prediction) == 8:
st.write("<NAME>!")
if np.argmax(prediction) == 9:
st.write("<NAME>!")
if np.argmax(prediction) == 10:
st.write("<NAME>!")
if np.argmax(prediction) == 11:
st.write("<NAME>!")
if np.argmax(prediction) == 12:
st.write("<NAME>!")
if np.argmax(prediction) == 13:
st.write("<NAME>!")
if np.argmax(prediction) == 14:
st.write("<NAME>!")
if np.argmax(prediction) == 15:
st.write("<NAME>!")
if np.argmax(prediction) == 16:
st.write("<NAME>!")
if np.argmax(prediction) == 17:
st.write("Belldandy!")
if np.argmax(prediction) == 18:
st.write("Black Rock Shooter!")
if np.argmax(prediction) == 19:
st.write("C.C!")
if np.argmax(prediction) == 20:
st.write("Canal Volphied!")
if np.argmax(prediction) == 21:
st.write("<NAME>!")
if np.argmax(prediction) == 22:
st.write("Chii!")
if np.argmax(prediction) == 23:
st.write("Cirno!")
if np.argmax(prediction) == 24:
st.write("<NAME>!")
if np.argmax(prediction) == 25:
st.write("<NAME>!")
if np.argmax(prediction) == 26:
st.write("<NAME>!")
if np.argmax(prediction) == 27:
st.write("Erio Mondial!")
if np.argmax(prediction) == 28:
st.write("Fate Testarossa!")
if np.argmax(prediction) == 29:
st.write("Feena Fam Earthlight!")
if np.argmax(prediction) == 30:
st.write("<NAME>!")
if np.argmax(prediction) == 31:
st.write("Fujibayashi Kyou!")
if np.argmax(prediction) == 32:
st.write("Fukuzawa Yumi!")
if np.argmax(prediction) == 33:
st.write("F<NAME>!")
if np.argmax(prediction) == 34:
st.write("Furukawa Nagisa!")
if np.argmax(prediction) == 35:
st.write("Fuyou Kaede!")
if np.argmax(prediction) == 36:
st.write("Golden Darkness!")
if np.argmax(prediction) == 37:
st.write("Hakurei Reimu!")
if np.argmax(prediction) == 38:
st.write("Hatsune Miku!")
if np.argmax(prediction) == 39:
st.write("Hayama Mizuki!")
if np.argmax(prediction) == 40:
st.write("Hayase Mitsuki!")
if np.argmax(prediction) == 41:
st.write("Hiiragi Kagami!")
if np.argmax(prediction) == 42:
st.write("Hiiragi Tsukasa!")
if np.argmax(prediction) == 43:
st.write("Hinamori Amu!")
if np.argmax(prediction) == 44:
st.write("Hirasawa Yui")
if np.argmax(prediction) == 45:
st.write("Horo!")
if np.argmax(prediction) == 46:
st.write("Houjou Reika!")
if np.argmax(prediction) == 47:
st.write("Houjou Satoko!")
if np.argmax(prediction) == 48:
st.write("Ibuki Fuuko!")
if np.argmax(prediction) == 49:
st.write("Ichinose Kotomi!")
if np.argmax(prediction) == 50:
st.write("Ikari Shinji!")
if np.argmax(prediction) == 51:
st.write("<NAME>!")
if np.argmax(prediction) == 52:
st.write("Ito Chika!")
if np.argmax(prediction) == 53:
st.write("Ito Nobue!")
if np.argmax(prediction) == 54:
st.write("Izayoi Sakuyae!")
if np.argmax(prediction) == 55:
st.write("Izumi Konata!")
if np.argmax(prediction) == 56:
st.write("K<NAME>!")
if np.argmax(prediction) == 57:
st.write("K<NAME>!")
if np.argmax(prediction) == 58:
st.write("Kagurazaka Asuna!")
if np.argmax(prediction) == 59:
st.write("<NAME>!")
if np.argmax(prediction) == 60:
st.write("K<NAME>!")
if np.argmax(prediction) == 61:
st.write("Kanu!")
if np.argmax(prediction) == 62:
st.write("K<NAME>!")
if np.argmax(prediction) == 63:
st.write("<NAME>!")
if np.argmax(prediction) == 64:
st.write("K<NAME>!")
if np.argmax(prediction) == 65:
st.write("Kawashima Ami!")
if np.argmax(prediction) == 66:
st.write("Kikuchi Makoto!")
if np.argmax(prediction) == 67:
st.write("Kinom<NAME>!")
if np.argmax(prediction) == 68:
st.write("Kirisame Marisa!")
if np.argmax(prediction) == 69:
st.write("Kisaragi Chihaya!")
if np.argmax(prediction) == 70:
st.write("Kobayakawa Yutaka!")
if np.argmax(prediction) == 71:
st.write("Kochiya Sanae!")
if np.argmax(prediction) == 72:
st.write("Koizumi Itsuki!")
if np.argmax(prediction) == 73:
st.write("Komaki Manaka!")
if np.argmax(prediction) == 74:
st.write("Konpaku Youmu!")
if np.argmax(prediction) == 75:
st.write("Kotegawa Yui!")
if np.argmax(prediction) == 76:
st.write("Kotobuki Tsumugi!")
if np.argmax(prediction) == 77:
st.write("Kousaka Tamaki!")
if np.argmax(prediction) == 78:
st.write("Kururugi Suzaku!")
if np.argmax(prediction) == 79:
st.write("Kusugawa Sasara!")
if np.argmax(prediction) == 80:
st.write("Kyon!")
if np.argmax(prediction) == 81:
st.write("**<NAME>!**")
if np.argmax(prediction) == 82:
st.write("Le<NAME>ouge!")
if np.argmax(prediction) == 83:
st.write("<NAME>!")
if np.argmax(prediction) == 84:
st.write("Lisianthus!")
if np.argmax(prediction) == 85:
st.write("Louise!")
if np.argmax(prediction) == 86:
st.write("<NAME>!")
if np.argmax(prediction) == 87:
st.write("Maria!")
if np.argmax(prediction) == 88:
st.write("<NAME>!")
if np.argmax(prediction) == 89:
st.write("Matsuoka Miu!")
if np.argmax(prediction) == 90:
st.write("<NAME>!")
if np.argmax(prediction) == 91:
st.write("<NAME>!")
if np.argmax(prediction) == 92:
st.write("Midori!")
if np.argmax(prediction) == 93:
st.write("<NAME>!")
if np.argmax(prediction) == 94:
st.write("<NAME>!")
if np.argmax(prediction) == 95:
st.write("Min<NAME>!")
if np.argmax(prediction) == 96:
st.write("<NAME>!")
if np.argmax(prediction) == 97:
st.write("<NAME>!")
if np.argmax(prediction) == 98:
st.write("<NAME>!")
if np.argmax(prediction) == 99:
st.write("<NAME>!")
if np.argmax(prediction) == 100:
st.write("N<NAME>!")
if np.argmax(prediction) == 101:
st.write("N<NAME>uki!")
if np.argmax(prediction) == 102:
st.write("Nagi!")
if np.argmax(prediction) == 103:
st.write("Nak<NAME>!")
if np.argmax(prediction) == 104:
st.write("Nanael!")
if np.argmax(prediction) == 105:
st.write("N<NAME>!")
if np.argmax(prediction) == 106:
st.write("Nerine!")
if np.argmax(prediction) == 107:
st.write("Nia!")
if np.argmax(prediction) == 108:
st.write("Njihara Ink!")
if np.argmax(prediction) == 109:
st.write("Nog<NAME>!")
if np.argmax(prediction) == 110:
st.write("<NAME>!")
if np.argmax(prediction) == 111:
st.write("Nunn<NAME>!")
if np.argmax(prediction) == 112:
st.write("Ogasawara Sachiko!")
if np.argmax(prediction) == 113:
st.write("Okazaki Tomoya!")
if np.argmax(prediction) == 114:
st.write("Pastel Inke!")
if np.argmax(prediction) == 115:
st.write("**Patchouli Knowledge!**")
st.write("Select a link to find a figurine!")
st.write("https://solarisjapan.com/search#q=Patchouli%20Knowledge&page=0")
st.write("https://otakumode.com/search?category=figures-dolls&keyword=Patchouli")
st.write("https://www.amazon.com/HTQING-Touhou-Project-Patchouli-Knowledge/dp/B08H5S86HR/ref=sr_1_48?dchild=1&keywords=Patchouli+knowledge+figurine&qid=1601487342&sr=8-48")
if np.argmax(prediction) == 116:
st.write("Primula!")
if np.argmax(prediction) == 117:
st.write("<NAME>!")
if np.argmax(prediction) == 118:
st.write("Reina!")
if np.argmax(prediction) == 119:
st.write("<NAME>!")
if np.argmax(prediction) == 120:
st.write("<NAME>!")
if np.argmax(prediction) == 121:
st.write("<NAME>!")
if np.argmax(prediction) == 122:
st.write("<NAME>!")
if np.argmax(prediction) == 123:
st.write("R<NAME>!")
if np.argmax(prediction) == 124:
st.write("Ryuuguu Rena!")
if np.argmax(prediction) == 125:
st.write("Saber!")
if np.argmax(prediction) == 126:
st.write("Saigyouji Yuyuko!")
if np.argmax(prediction) == 127:
st.write("<NAME>!")
if np.argmax(prediction) == 128:
st.write("Sakagami Tomoyo!")
if np.argmax(prediction) == 129:
st.write("Sakai Yuuji!")
if np.argmax(prediction) == 130:
st.write("San<NAME>!")
if np.argmax(prediction) == 131:
st.write("Saotome Alto!")
if np.argmax(prediction) == 132:
st.write("Sendou Erika!")
if np.argmax(prediction) == 133:
st.write("Seto San!")
if np.argmax(prediction) == 134:
st.write("<NAME>!")
if np.argmax(prediction) == 135:
st.write("Shameimaru Aya!")
if np.argmax(prediction) == 136:
st.write("Shana!")
if np.argmax(prediction) == 137:
st.write("<NAME>!")
if np.argmax(prediction) == 138:
st.write("Sh<NAME>!")
if np.argmax(prediction) == 139:
st.write("Shigure Asa!")
if np.argmax(prediction) == 140:
st.write("Shih<NAME>!")
if np.argmax(prediction) == 141:
st.write("Sh<NAME>!")
if np.argmax(prediction) == 142:
st.write("Shindou Kei!")
if np.argmax(prediction) == 143:
st.write("Shinku'!")
if np.argmax(prediction) == 144:
st.write("Shirakawa Kotori!")
if np.argmax(prediction) == 145:
st.write("<NAME>!")
if np.argmax(prediction) == 146:
st.write("Shirou Kamui!")
if np.argmax(prediction) == 147:
st.write("**Siesta!**")
st.write("Select a link to find a figurine!")
st.write("https://www.amazon.com/s?k=Zero+no+Tsukaima+siesta")
st.write("https://otakumode.com/search?category=figures-dolls&keyword=The%20Familiar%20of%20Zero")
st.write("https://solarisjapan.com/search#q=siesta&page=0")
if np.argmax(prediction) == 148:
st.write("Sonozaki Mion!")
if np.argmax(prediction) == 149:
st.write("Sonsaku Hakufu!")
if np.argmax(prediction) == 150:
st.write("Souryuu Asuka Langley!")
if np.argmax(prediction) == 151:
st.write("Subaru Nakajima'!")
if np.argmax(prediction) == 152:
st.write("Suigintou'!")
if np.argmax(prediction) == 153:
st.write("Suzumiya Akane!")
if np.argmax(prediction) == 154:
st.write("Suzumiya Haruhi!")
if np.argmax(prediction) == 155:
st.write("Suzumiya Haruka!")
if np.argmax(prediction) == 156:
st.write("Tainaka Ritsu'!")
if np.argmax(prediction) == 157:
st.write("Takamachi Nanoha!")
if np.argmax(prediction) == 158:
st.write("Takara Miyuki!")
if np.argmax(prediction) == 159:
st.write("Takatsuki Yayoi!")
if np.argmax(prediction) == 160:
st.write("Teana Lanster!")
if np.argmax(prediction) == 161:
st.write("Tohsaka Rin!")
if np.argmax(prediction) == 162:
st.write("Tsukimura Mayu!")
if np.argmax(prediction) == 163:
st.write("Tsuruya!")
if np.argmax(prediction) == 164:
st.write("Vita!")
if np.argmax(prediction) == 165:
st.write("Vivio'!")
if np.argmax(prediction) == 166:
st.write("Yagami Hayate!")
if np.argmax(prediction) == 167:
st.write("Yakumo Yukari!")
if np.argmax(prediction) == 168:
st.write("Yoko!")
if np.argmax(prediction) == 169:
st.write("Yoshida Kazumi!")
if np.argmax(prediction) == 170:
st.write("Yuno!")
if np.argmax(prediction) == 171:
st.write("Yuuki Mikan!")
if np.argmax(prediction) == 172:
st.write("Yuzuhara Konomi!")
# + id="bxtY69bsDCHZ"
# !streamlit run findmyfigurineapp.py &>/dev/null&
# + id="cvp_tayTDCPB"
#Check running processes
# !pgrep streamlit
# + id="zh9mqPKvYcyN"
# !ps -eaf | grep streamlit
# + id="NTAy4mnHDU_a"
# Setup a tunnel to the streamlit port 8501
public_url = ngrok.connect(port='8501')
public_url
# + id="2VVYC8iXDZSP"
#shutdown specific running process
# !kill 7963
# + id="8LmzLCyhIjSl"
#shutdown all ngrok processes
ngrok.kill()
| streamlit/findmyfigurineapp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
import numpy as np
import tensorflow as tf
# +
import json
with open('train-test.json') as fopen:
dataset = json.load(fopen)
with open('dictionary.json') as fopen:
dictionary = json.load(fopen)
# -
train_X = dataset['train_X']
train_Y = dataset['train_Y']
test_X = dataset['test_X']
test_Y = dataset['test_Y']
dictionary.keys()
# +
dictionary_from = dictionary['from']['dictionary']
rev_dictionary_from = dictionary['from']['rev_dictionary']
dictionary_to = dictionary['to']['dictionary']
rev_dictionary_to = dictionary['to']['rev_dictionary']
# -
GO = dictionary_from['GO']
PAD = dictionary_from['PAD']
EOS = dictionary_from['EOS']
UNK = dictionary_from['UNK']
# +
for i in range(len(train_X)):
train_X[i] += ' EOS'
train_X[0]
# +
for i in range(len(test_X)):
test_X[i] += ' EOS'
test_X[0]
# +
from tensorflow.python.util import nest
from tensorflow.python.layers.core import Dense
def gnmt_residual_fn(inputs, outputs):
def split_input(inp, out):
out_dim = out.get_shape().as_list()[-1]
inp_dim = inp.get_shape().as_list()[-1]
return tf.split(inp, [out_dim, inp_dim - out_dim], axis=-1)
actual_inputs, _ = nest.map_structure(split_input, inputs, outputs)
def assert_shape_match(inp, out):
inp.get_shape().assert_is_compatible_with(out.get_shape())
nest.assert_same_structure(actual_inputs, outputs)
nest.map_structure(assert_shape_match, actual_inputs, outputs)
return nest.map_structure(lambda inp, out: inp + out, actual_inputs, outputs)
class GNMTAttentionMultiCell(tf.nn.rnn_cell.MultiRNNCell):
def __init__(self, attention_cell, cells, use_new_attention=True):
cells = [attention_cell] + cells
self.use_new_attention = use_new_attention
super(GNMTAttentionMultiCell, self).__init__(
cells, state_is_tuple=True)
def __call__(self, inputs, state, scope=None):
"""Run the cell with bottom layer's attention copied to all upper layers."""
if not nest.is_sequence(state):
raise ValueError(
"Expected state to be a tuple of length %d, but received: %s"
% (len(self.state_size), state))
with tf.variable_scope(scope or "multi_rnn_cell"):
new_states = []
with tf.variable_scope("cell_0_attention"):
attention_cell = self._cells[0]
attention_state = state[0]
cur_inp, new_attention_state = attention_cell(
inputs, attention_state)
new_states.append(new_attention_state)
for i in range(1, len(self._cells)):
with tf.variable_scope("cell_%d" % i):
cell = self._cells[i]
cur_state = state[i]
if self.use_new_attention:
cur_inp = tf.concat(
[cur_inp, new_attention_state.attention], -1)
else:
cur_inp = tf.concat(
[cur_inp, attention_state.attention], -1)
cur_inp, new_state = cell(cur_inp, cur_state)
new_states.append(new_state)
return cur_inp, tuple(new_states)
class Translator:
def __init__(self, size_layer, num_layers, embedded_size,
from_dict_size, to_dict_size, learning_rate, beam_width = 5):
def cells(size,reuse=False):
return tf.nn.rnn_cell.GRUCell(size,reuse=reuse)
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.X_seq_len = tf.count_nonzero(self.X, 1, dtype=tf.int32)
self.Y_seq_len = tf.count_nonzero(self.Y, 1, dtype=tf.int32)
batch_size = tf.shape(self.X)[0]
encoder_embeddings = tf.Variable(tf.random_uniform([from_dict_size, embedded_size], -1, 1))
decoder_embeddings = tf.Variable(tf.random_uniform([to_dict_size, embedded_size], -1, 1))
encoder_embedded = tf.nn.embedding_lookup(encoder_embeddings, self.X)
main = tf.strided_slice(self.Y, [0, 0], [batch_size, -1], [1, 1])
decoder_input = tf.concat([tf.fill([batch_size, 1], GO), main], 1)
decoder_embedded = tf.nn.embedding_lookup(encoder_embeddings, decoder_input)
num_residual_layer = num_layers - 2
num_bi_layer = 1
num_ui_layer = num_layers - num_bi_layer
for n in range(num_bi_layer):
(out_fw, out_bw), (state_fw, state_bw) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(size_layer),
cell_bw = cells(size_layer),
inputs = encoder_embedded,
sequence_length = self.X_seq_len,
dtype = tf.float32,
scope = 'bidirectional_rnn_%d'%(n))
encoder_embedded = tf.concat((out_fw, out_bw), 2)
gru_cells = tf.nn.rnn_cell.MultiRNNCell([cells(size_layer) for _ in range(num_ui_layer)])
encoder_outputs, encoder_state = tf.nn.dynamic_rnn(
gru_cells,
encoder_embedded,
dtype=tf.float32,
sequence_length=self.X_seq_len)
encoder_state = (state_bw,) + (
(encoder_state,) if num_ui_layer == 1 else encoder_state)
decoder_cells = []
for n in range(num_layers):
cell = cells(size_layer)
if (n >= num_layers - num_residual_layer):
cell = tf.nn.rnn_cell.ResidualWrapper(cell, residual_fn = gnmt_residual_fn)
decoder_cells.append(cell)
attention_cell = decoder_cells.pop(0)
to_dense = tf.layers.Dense(to_dict_size)
with tf.variable_scope('decode'):
attention_mechanism = tf.contrib.seq2seq.BahdanauAttention(
num_units = size_layer,
memory = encoder_outputs,
memory_sequence_length = self.X_seq_len)
att_cell = tf.contrib.seq2seq.AttentionWrapper(
cell = attention_cell,
attention_mechanism = attention_mechanism,
attention_layer_size = None,
alignment_history = True,
output_attention = False)
gcell = GNMTAttentionMultiCell(att_cell, decoder_cells)
self.initial_state = tuple(
zs.clone(cell_state=es)
if isinstance(zs, tf.contrib.seq2seq.AttentionWrapperState) else es
for zs, es in zip(
gcell.zero_state(batch_size, dtype=tf.float32), encoder_state))
training_helper = tf.contrib.seq2seq.TrainingHelper(
decoder_embedded,
self.Y_seq_len,
time_major = False
)
training_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = gcell,
helper = training_helper,
initial_state = self.initial_state,
output_layer = to_dense)
training_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = training_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.Y_seq_len))
self.training_logits = training_decoder_output.rnn_output
with tf.variable_scope('decode', reuse=True):
encoder_out_tiled = tf.contrib.seq2seq.tile_batch(encoder_outputs, beam_width)
encoder_state_tiled = tf.contrib.seq2seq.tile_batch(encoder_state, beam_width)
X_seq_len_tiled = tf.contrib.seq2seq.tile_batch(self.X_seq_len, beam_width)
attention_mechanism = tf.contrib.seq2seq.BahdanauAttention(
num_units = size_layer,
memory = encoder_out_tiled,
memory_sequence_length = X_seq_len_tiled)
att_cell = tf.contrib.seq2seq.AttentionWrapper(
cell = attention_cell,
attention_mechanism = attention_mechanism,
attention_layer_size = None,
alignment_history = False,
output_attention = False)
gcell = GNMTAttentionMultiCell(att_cell, decoder_cells)
self.initial_state = tuple(
zs.clone(cell_state=es)
if isinstance(zs, tf.contrib.seq2seq.AttentionWrapperState) else es
for zs, es in zip(
gcell.zero_state(batch_size * beam_width, dtype=tf.float32), encoder_state_tiled))
predicting_decoder = tf.contrib.seq2seq.BeamSearchDecoder(
cell = gcell,
embedding = decoder_embeddings,
start_tokens = tf.tile(tf.constant([GO], dtype=tf.int32), [batch_size]),
end_token = EOS,
initial_state = self.initial_state,
beam_width = beam_width,
output_layer = to_dense,
length_penalty_weight = 0.0)
predicting_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = predicting_decoder,
impute_finished = False,
maximum_iterations = 2 * tf.reduce_max(self.X_seq_len))
self.predicting_ids = predicting_decoder_output.predicted_ids[:, :, 0]
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
self.cost = tf.contrib.seq2seq.sequence_loss(logits = self.training_logits,
targets = self.Y,
weights = masks)
self.optimizer = tf.train.AdamOptimizer(learning_rate).minimize(self.cost)
y_t = tf.argmax(self.training_logits,axis=2)
y_t = tf.cast(y_t, tf.int32)
self.prediction = tf.boolean_mask(y_t, masks)
mask_label = tf.boolean_mask(self.Y, masks)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# -
size_layer = 512
num_layers = 3
learning_rate = 1e-3
batch_size = 64
epoch = 20
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Translator(size_layer, num_layers, size_layer,
len(dictionary_from), len(dictionary_to), learning_rate)
sess.run(tf.global_variables_initializer())
# +
def str_idx(corpus, dic):
X = []
for i in corpus:
ints = []
for k in i.split():
ints.append(dic.get(k,UNK))
X.append(ints)
return X
def pad_sentence_batch(sentence_batch, pad_int):
padded_seqs = []
seq_lens = []
max_sentence_len = max([len(sentence) for sentence in sentence_batch])
for sentence in sentence_batch:
padded_seqs.append(sentence + [pad_int] * (max_sentence_len - len(sentence)))
seq_lens.append(len(sentence))
return padded_seqs, seq_lens
# -
train_X = str_idx(train_X, dictionary_from)
test_X = str_idx(test_X, dictionary_from)
train_Y = str_idx(train_Y, dictionary_to)
test_Y = str_idx(test_Y, dictionary_to)
sess.run(model.predicting_ids, feed_dict = {model.X: [train_X[0]]}).shape
# +
import tqdm
for e in range(epoch):
pbar = tqdm.tqdm(
range(0, len(train_X), batch_size), desc = 'minibatch loop')
train_loss, train_acc, test_loss, test_acc = [], [], [], []
for i in pbar:
index = min(i + batch_size, len(train_X))
maxlen = max([len(s) for s in train_X[i : index] + train_Y[i : index]])
batch_x, seq_x = pad_sentence_batch(train_X[i : index], PAD)
batch_y, seq_y = pad_sentence_batch(train_Y[i : index], PAD)
feed = {model.X: batch_x,
model.Y: batch_y}
accuracy, loss, _ = sess.run([model.accuracy,model.cost,model.optimizer],
feed_dict = feed)
train_loss.append(loss)
train_acc.append(accuracy)
pbar.set_postfix(cost = loss, accuracy = accuracy)
pbar = tqdm.tqdm(
range(0, len(test_X), batch_size), desc = 'minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_X))
batch_x, seq_x = pad_sentence_batch(test_X[i : index], PAD)
batch_y, seq_y = pad_sentence_batch(test_Y[i : index], PAD)
feed = {model.X: batch_x,
model.Y: batch_y,}
accuracy, loss = sess.run([model.accuracy,model.cost],
feed_dict = feed)
test_loss.append(loss)
test_acc.append(accuracy)
pbar.set_postfix(cost = loss, accuracy = accuracy)
print('epoch %d, training avg loss %f, training avg acc %f'%(e+1,
np.mean(train_loss),np.mean(train_acc)))
print('epoch %d, testing avg loss %f, testing avg acc %f'%(e+1,
np.mean(test_loss),np.mean(test_acc)))
# -
rev_dictionary_to = {int(k): v for k, v in rev_dictionary_to.items()}
# +
test_size = 20
batch_x, seq_x = pad_sentence_batch(test_X[: test_size], PAD)
batch_y, seq_y = pad_sentence_batch(test_Y[: test_size], PAD)
feed = {model.X: batch_x}
logits = sess.run(model.predicting_ids, feed_dict = feed)
logits.shape
# +
rejected = ['PAD', 'EOS', 'UNK', 'GO']
for i in range(test_size):
predict = [rev_dictionary_to[i] for i in logits[i] if rev_dictionary_to[i] not in rejected]
actual = [rev_dictionary_to[i] for i in batch_y[i] if rev_dictionary_to[i] not in rejected]
print(i, 'predict:', ' '.join(predict))
print(i, 'actual:', ' '.join(actual))
print()
# -
| mlmodels/model_dev/nlp_tfflow/neural-machine-translation/48.google-nmt.ipynb |
# # 📝 Exercise M6.01
#
# The aim of this notebook is to investigate if we can tune the hyperparameters
# of a bagging regressor and evaluate the gain obtained.
#
# We will load the California housing dataset and split it into a training and
# a testing set.
# +
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
data, target = fetch_california_housing(as_frame=True, return_X_y=True)
target *= 100 # rescale the target in k$
data_train, data_test, target_train, target_test = train_test_split(
data, target, random_state=0, test_size=0.5)
# -
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">If you want a deeper overview regarding this dataset, you can refer to the
# Appendix - Datasets description section at the end of this MOOC.</p>
# </div>
# Create a `BaggingRegressor` and provide a `DecisionTreeRegressor`
# to its parameter `base_estimator`. Train the regressor and evaluate its
# statistical performance on the testing set using the mean absolute error.
# +
# Write your code here.
from sklearn.ensemble import BaggingRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
bagged_trees = BaggingRegressor(
base_estimator=DecisionTreeRegressor())
_ = bagged_trees.fit(data_train, target_train)
y_pred = bagged_trees.predict(data_test)
print(f'MAE: {mean_absolute_error(target_test, y_pred):0.02f} k$')
# -
# Now, create a `RandomizedSearchCV` instance using the previous model and
# tune the important parameters of the bagging regressor. Find the best
# parameters and check if you are able to find a set of parameters that
# improve the default regressor still using the mean absolute error as a
# metric.
#
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Tip</p>
# <p class="last">You can list the bagging regressor's parameters using the <tt class="docutils literal">get_params</tt>
# method.</p>
# </div>
bagged_trees.get_params()
import sklearn
sorted(sklearn.metrics.SCORERS.keys())
# +
# Write your code here.
from sklearn.model_selection import RandomizedSearchCV
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import cross_validate
from scipy.stats import randint
param_grid = {
"max_depth": [3, 5, 8, None],
"min_samples_split": [2, 10, 30, 50],
"min_samples_leaf": [0.01, 0.05, 0.1, 1]}
param_grid = {
"n_estimators": randint(10, 30),
"max_samples": [0.5, 0.8, 1.0],
"max_features": [0.5, 0.8, 1.0],
"base_estimator__max_depth": randint(3, 10),
}
search = RandomizedSearchCV(
bagged_trees, param_grid, n_iter=20, scoring="neg_mean_absolute_error"
)
_ = search.fit(data_train, target_train)
# -
y_pred = search.predict(data_test)
from sklearn.metrics import mean_absolute_error
mean_absolute_error(target_test, y_pred)
# +
import pandas as pd
columns = [f"param_{name}" for name in param_grid.keys()]
columns += ["mean_test_score", "std_test_score", "rank_test_score"]
cv_results = pd.DataFrame(search.cv_results_)
cv_results = cv_results[columns].sort_values(by="rank_test_score")
cv_results["mean_test_score"] = -cv_results["mean_test_score"]
cv_results
# -
target_predicted = search.predict(data_test)
print(f"Mean absolute error after tuning of the bagging regressor:\n"
f"{mean_absolute_error(target_test, target_predicted):.2f} k$")
# We see that the bagging regressor provides a predictor in which fine tuning
# is not as important as in the case of fitting a single decision tree.
| notebooks/ensemble_ex_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bag of Words (Or: *What is unique about working with text*)
# **Q**: How is text different from the types of data you've seen so far?
# * It's a string, so can't do mathematical operations on it
# * It can be ambigious
# * It can be in a wide range of formats / files
# * Has a wide granularity level (file, paragraph, sentence, word, character)
# * It is unstructured
# **Q**: What does this mean for how you can work with text compared to types of data you've worked with so far?
# * Need to make it structured
# * Need to clean it well
# There are two main aspect of working with text that helps us feed it into computers / machine learning models:
# 1. Data/text preprocessing
# 2. Turning text into features
# ### 1. Data Preprocessing
# You already know that data cleaning is a large part of a data scientist's workflow. When working with unstructured data such as text, data cleaning plays an even bigger role.
#
# **Q**: Things we may want to do to clean our textual data:
# * Lemmatization
# * Stemming
# * Tokenization
# * Remove small words that don't contribute to the meaning
# * ...
# * Split the corpus into individual words / tokens
# * Remove punctuation
# * Deal with capitalization
# * Remove most common words:
# * Use a list of words to remove (stop words)
# * Remove the words that appear in more than X% of documents
# * Reduce words to their base parts:
# * *Stemming*: process of removing and replacing suffixes to get to the root of the words, *stem*.
# * based on heuristics (e.g. *-ational -> -ate*, *-tional -> -tion*)
# * does not always produce a word
# * feet->feet, wolves->wolv, cats->cat, talked->talk
# * *Lemmatization*: uses vocabulary and morphological analysis to return the base or dictionary form of a word, *lemma*.
# * does not always return a reduced form
# * feet->foot, wolves->wolf, cats->cat, talked->talked
corpus = ["we all love a yellow submarine", # Beatles
"yesterday, my submarine was in love", # Beatles
"we are love trouble with loyalty here", # Eminem
"loyalty to us is worth more than love is"] # Eminem
labels = ['Beatles'] * 2 + ['Eminem'] * 2
# ### 2. Turning text into features
# **Q**: Once you have your data cleaned and preprocessed like this, what can you imagine using as your features?
# * Amount of certain words
# * Length of song / words in song
# * Categorize words, then use word count
# #### Bag of words
# Each token/word will be a feature/column.
# * large sparse matrix
# * word order lost
# * counts not normalized
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer()
X = vectorizer.fit_transform(corpus)
X
pd.DataFrame(X.todense(), columns=vectorizer.get_feature_names(), index=labels)
# ##### Exercise: How can we remove the most common words?
# * Using a list of stop words
# * Removing the words that appear in more than X% of documents
# See `CountVectorizer` documentation for how to do each of these: https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html
# Remove most common words using these two methods. Use `.vocabulary_` and `.stop_words_` attributes to see which words have remained and which are removed (latter only in the case of the second method).
#
# What do you notice?
# * Using a list of stop words
vectorizer = CountVectorizer(stop_words='english')
X = vectorizer.fit_transform(corpus)
X_df = pd.DataFrame(X.todense(), columns=vectorizer.get_feature_names(), index=labels)
X_df
# * Removing the words that appear in more than X% of documents
vectorizer = CountVectorizer(max_df=0.75)
X = vectorizer.fit_transform(corpus)
X_df = pd.DataFrame(X.todense(), columns=vectorizer.get_feature_names(), index=labels)
X_df
vectorizer.stop_words_
X_df.columns
vectorizer.vocabulary_
vectorizer.get_feature_names()
# #### n-grams
# Instead of single tokens, we now also count token pairs (bigrams), triplets (trigrams), etc.
# * even larger sparser matrix
# * preserves local word order
# * counts not normalized
# * too many features:
# * remove high-frequency n-grams: can include stop words; not very informative
# * remove low-frequency n-grams: typos and rare n-grams; likely to overfit
vectorizer = CountVectorizer(ngram_range=(1, 2))
X = vectorizer.fit_transform(corpus)
X_df = pd.DataFrame(X.todense(), columns=vectorizer.get_feature_names(), index=labels)
X_df
# #### TF-IDF
# Stands for `term frequency - inverse document frequency` and aims to address the popularity/frequency of words in a corpus(not just inside of a single document).
# ##### TF = term frequency
#
# TF(t, d) - frequency of term (n-gram) _t_ in document _d_
# ##### IDF(t) = inverse document frequency (of term _t_ in the whole corpus)
#
# $ IDF(t) = \log \frac{1+N}{1+N_t}+1 $
#
#
# If term _t_ doesn't appear in many documents: IDF is "big".
#
# If term _t_ appears in many documents: IDF is close to 1 ("small") -> common terms are penalized.
# $ TFIDF(t, d) = TF(t,d)*IDF(t) $
#
# **Q**: What kind of terms will have high TF-IDF?
# * Those that appear a lot in small number of documents/songs.
# ##### Exercise: Implement TF-IDF vectorizer
# Look up how to implement TF-IDF vectorizer in `scikit-learn`. How does your features dataframe differ from the `CountVectorizer` one?
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer()
X = vectorizer.fit_transform(corpus)
X_df = pd.DataFrame(X.todense(), columns=vectorizer.get_feature_names(), index=labels)
X_df
# *Bonus question*: What can you say about the values in your new `X_df` (think about sums, normalizations, etc.)? Post your guesses in Slack!
# ##### Extra exercise
# Use your scraped lyrics and run them through a vectorizer of your choice. Then split the resulting feature vector `X` and your labels into training and test set and train a logistic regression model. Check how the choice of vectorizers and the parameters we mentioned affects the performance of your model.
#
# Once you have your final model, run the following lines of code to get the words that are strongest predictors for each of your bands and post them in Slack.
# `import operator`
# `model = LogisticRegression()`
# `print(operator.itemgetter(*np.argsort(model.coef_[0]))(vectorizer.get_feature_names())[-20:])`
# `print(operator.itemgetter(*np.argsort(model.coef_[0]))(vectorizer.get_feature_names())[:20])`
| week_04/Bag of Words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Containers
# ### Checking for containment.
# The `list` we saw is a container type: its purpose is to hold other objects. We can ask python whether or not a
# container contains a particular item:
'Dog' in ['Cat', 'Dog', 'Horse']
'Bird' in ['Cat', 'Dog', 'Horse']
2 in range(5)
99 in range(5)
# ### Mutability
# A list can be modified:
name = "<NAME>".split(" ")
print(name)
# +
name[0] = "Dr"
name[1:3] = ["Griffiths-"]
name.append("PhD")
print(" ".join(name))
# -
# ### Tuples
# A `tuple` is an immutable sequence. It is like a list, execpt it cannot be changed. It is defined with round brackets.
x = 0,
type(x)
my_tuple = ("Hello", "World")
my_tuple[0] = "Goodbye"
type(my_tuple)
# `str` is immutable too:
fish = "Hake"
fish[0] = 'R'
# But note that container reassignment is moving a label, **not** changing an element:
fish = "Rake" ## OK!
# *Supplementary material*: Try the [online memory visualiser](http://www.pythontutor.com/visualize.html#code=name+%3D++%22James+Philip+John+Hetherington%22.split%28%22+%22%29%0A%0Aname%5B0%5D+%3D+%22Dr%22%0Aname%5B1%3A3%5D+%3D+%5B%22Griffiths-%22%5D%0Aname.append%28%22PhD%22%29%0A%0Aname+%3D+%22Bilbo+Baggins%22&mode=display&origin=opt-frontend.js&cumulative=false&heapPrimitives=true&textReferences=false&py=2&rawInputLstJSON=%5B%5D&curInstr=0) for this one.
# ### Memory and containers
#
# The way memory works with containers can be important:
#
#
#
x = list(range(3))
x
y = x
y
z = x[0:3]
y[1] = "Gotcha!"
x
y
z
z[2] = "Really?"
x
y
z
# *Supplementary material*: This one works well at the [memory visualiser](http://www.pythontutor.com/visualize.html#code=x+%3D+%5B%22What's%22,+%22Going%22,+%22On%3F%22%5D%0Ay+%3D+x%0Az+%3D+x%5B0%3A3%5D%0A%0Ay%5B1%5D+%3D+%22Gotcha!%22%0Az%5B2%5D+%3D+%22Really%3F%22&mode=display&origin=opt-frontend.js&cumulative=false&heapPrimitives=true&textReferences=false&py=2&rawInputLstJSON=%5B%5D&curInstr=0).
# The explanation: While `y` is a second label on the *same object*, `z` is a separate object with the same data. Writing `x[:]` creates a new list containing all the elements of `x` (remember: `[:]` is equivalent to `[0:<last>]`). This is the case whenever we take a slice from a list, not just when taking all the elements with `[:]`.
#
# The difference between `y=x` and `z=x[:]` is important!
# Nested objects make it even more complicated:
x = [['a', 'b'] , 'c']
y = x
z = x[0:2]
x[0][1] = 'd'
z[1] = 'e'
x
y
z
# Try the [visualiser](http://www.pythontutor.com/visualize.html#code=x%3D%5B%5B'a','b'%5D,'c'%5D%0Ay%3Dx%0Az%3Dx%5B0%3A2%5D%0A%0Ax%5B0%5D%5B1%5D%3D'd'%0Az%5B1%5D%3D'e'&mode=display&origin=opt-frontend.js&cumulative=false&heapPrimitives=true&textReferences=false&py=2&rawInputLstJSON=%5B%5D&curInstr=0) again.
#
# *Supplementary material*: The copies that we make through slicing are called *shallow copies*: we don't copy all the objects they contain, only the references to them. This is why the nested list in `x[0]` is not copied, so `z[0]` still refers to it. It is possible to actually create copies of all the contents, however deeply nested they are - this is called a *deep copy*. Python provides methods for that in its standard library, in the `copy` module. You can read more about that, as well as about shallow and deep copies, in the [library reference](https://docs.python.org/3/library/copy.html).
# ### Identity vs Equality
#
#
# Having the same data is different from being the same actual object
# in memory:
[1, 2] == [1, 2]
[1, 2] is [1, 2]
# The == operator checks, element by element, that two containers have the same data.
# The `is` operator checks that they are actually the same object.
# But, and this point is really subtle, for immutables, the python language might save memory by reusing a single instantiated copy. This will always be safe.
"Hello" == "Hello"
"Hello" is "Hello"
# This can be useful in understanding problems like the one above:
x = range(3)
y = x
z = x[:]
x == y
x is y
x == z
x is z
| ch00python/025containers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calculating Charity Scores from relevant charity data on charitydata.ca (using dataframe from Data_Collection.ipyntb)
# +
#importing required libraries and setting the driver path
import pandas as pd
import numpy as np
import json
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.action_chains import ActionChains
from selenium.webdriver.common.desired_capabilities import DesiredCapabilities
from selenium.common.exceptions import NoSuchElementException
from selenium.common.exceptions import TimeoutException
driver_path = "C:/Users/16475/Documents/GitHub/CharityScore.ca/chromedriver.exe"
driver = webdriver.Chrome(executable_path=driver_path)
# +
# import the url data from the dataframe
df = pd.read_csv("expense data.csv")
# remove duplicate index column
df = pd.read_csv("expense data.csv").drop(['Unnamed: 0'],axis=1)
print(df.head(1))
# +
# find expense tables on charitydata.ca and add them to the dataframe as string representations of dictionaries
for i in range(1, 1):
driver.get(df.loc[i,"URL"])
# check if url page found, else skip
try:
url_check = driver.find_element_by_xpath("/html/body/main/div/section[2]/div/div[2]/p").text
except NoSuchElementException:
url_check = ""
if url_check == "The requested page could not be found.":
continue
# open expense table
try:
WebDriverWait(driver, 30).until(EC.presence_of_element_located((By.LINK_TEXT, 'Expenses'))).click()
expense_table = driver.find_element_by_xpath('//html/body/main/div/section[3]/div/div[2]/div[4]/div[2]/table[1]')
except TimeoutException:
try:
driver.get(df.loc[i,"URL"])
WebDriverWait(driver, 30).until(EC.presence_of_element_located((By.LINK_TEXT, 'Expenses'))).click()
expense_table = driver.find_element_by_xpath('//html/body/main/div/section[3]/div/div[2]/div[4]/div[2]/table[1]')
except:
continue
# create dictionary representation of expense table
expense_dict = {}
expense_rows = expense_table.find_elements(By.TAG_NAME, "tr")
header_row = expense_rows.pop(0)
headers = header_row.find_elements(By.TAG_NAME, "th")
header_key = headers.pop(0).text
headers = list(map(lambda x: int(x.text), headers))
expense_dict[header_key] = headers
for row in expense_rows:
row_key = row.find_element(By.TAG_NAME, "th").text
row_series = row.find_elements(By.TAG_NAME, "td")
num_list = []
for num in row_series:
if num.text[0] != "(":
num_list +=[int(num.text.replace(",",""))]
else:
num_list += [int(num.text[1:-1].replace(",",""))]
expense_dict[row_key] = num_list
df.loc[df.index[i], 'Expense_Table'] = str(expense_dict)
df.to_csv("expense data.csv")
# quit driver and export to csv
driver.quit()
df.to_csv("expense data.csv")
# +
# Adding columns for averages and percentages of expenditures per category
import pandas as pd
import numpy as np
import ast
import matplotlib.pyplot as plt
# import the url data from the dataframe
df = pd.read_csv("expense data.csv")
# remove duplicate index column
df = pd.read_csv("expense data.csv").drop(['Unnamed: 0'],axis=1)
# create list of common keys to access by index rather than name
ck = list(ast.literal_eval(df.loc[1,"Expense Table"]).keys())
# create data labels from common keys
# for each line in the dataframe, find the expense dictionary and create arrays for each bucket
for i in range(1,len(df)):
if df.loc[i, "Expense Table"] != "":
try:
expense_dict = ast.literal_eval(df.loc[i, "Expense Table"])
except ValueError:
expense_dict = None
if expense_dict != None:
reporting_years = expense_dict[ck[0]]
total_exp = expense_dict[ck[7]]
charitable_exp = expense_dict[ck[1]]
mgmt_exp = expense_dict[ck[2]]
fundraising_exp = expense_dict[ck[3]]
pol_exp = expense_dict[ck[4]]
other_exp = expense_dict[ck[5]]
gift_exp = expense_dict[ck[6]]
expense_matrix = np.matrix([reporting_years, total_exp, charitable_exp, mgmt_exp, fundraising_exp, pol_exp, gift_exp, other_exp])
# create a new columns with values from most recent year
df.loc[i, 'Reporting Year'] = expense_matrix[:,0][0].item()
df.loc[i, 'Total Expenditure'] = expense_matrix[:,0][1].item()
df.loc[i, 'Total Expenditure on Charitable Activities'] = expense_matrix[:,0][2].item()
df.loc[i, 'Total Expenditure on Management and Administration'] = expense_matrix[:,0][3].item()
df.loc[i, 'Total Expenditures on Fundraising'] = expense_matrix[:,0][4].item()
df.loc[i, 'Total Expenditures on Political Activities'] = expense_matrix[:,0][5].item()
df.loc[i, 'Total Other Expenditures '] = expense_matrix[:,0][6].item()
df.loc[i, 'Total Amount of Gifts Made to All Qualified Donees'] = expense_matrix[:,0][7].item()
df.to_csv("expense data calcs.csv")
print(expense_matrix[:,0])
| Score_Calculation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import numpy.ma as ma
import os
import string
import sys
import xarray as xr
import netCDF4 as nc4
import scipy.io as io
import glob
import matplotlib.pyplot as plt
import pickle
import itertools
from scipy.signal import butter, lfilter, filtfilt
import warnings
warnings.filterwarnings("ignore")
base = os.getcwd()+"/"
print(base)
# # Files
def get_model_names(filenames, model_list):
'''
Gets a list of available models (midHolocene & piControl) from the
curated_ESGF replica directory.
INPUTS:
- filenames: list, a file glob of the available sos & tos files
- model_list: list, an empty list to put the model names in
RETURNS:
- Nothing
'''
for path in filenames:
model_name = path.split('/')[4]
model_list.append(model_name)
# +
sos_filenames_mh = glob.glob('/data/CMIP/curated_ESGF_replica/*/midHolocene/sos*.nc')
sos_filenames_ctrl = glob.glob('/data/CMIP/curated_ESGF_replica/*/piControl/sos*.nc')
sos_filenames_hi = glob.glob('/data/CMIP/curated_ESGF_replica/*/historical/sos*.nc')
tos_filenames_mh = glob.glob('/data/CMIP/curated_ESGF_replica/*/midHolocene/tos*.nc')
tos_filenames_ctrl = glob.glob('/data/CMIP/curated_ESGF_replica/*/piControl/tos*.nc')
tos_filenames_hi = glob.glob('/data/CMIP/curated_ESGF_replica/*/historical/tos*.nc')
sos_models_mh=[]
sos_models_ctrl=[]
sos_models_hi=[]
tos_models_mh=[]
tos_models_ctrl=[]
tos_models_hi=[]
get_model_names(sos_filenames_mh, sos_models_mh)
get_model_names(sos_filenames_ctrl, sos_models_ctrl)
get_model_names(sos_filenames_hi, sos_models_hi)
get_model_names(tos_filenames_mh, tos_models_mh)
get_model_names(tos_filenames_ctrl, tos_models_ctrl)
get_model_names(tos_filenames_hi, tos_models_hi)
# -
def get_filenames(sos_models, tos_models, expt, var, array_list, model_namelist):
'''
Opens up the sos and tos files into x-array datasets.
INPUTS:
- sos_models, tos_models: list of model names
- expt: string, midHolocene or piControl
- var: string, sos or tos
- array_list: list to put x_array dataset in
- model_namelist: empty list, to put new model names in
RETURNS:
- Nothing
'''
# are both sos and tos present in piControl/midHolocene folders?
for model in sos_models:
if model in tos_models:
# {model}{expt}{var}
fn_format= "/data/CMIP/curated_ESGF_replica/{}/{}/{}*.nc"
# make a file-glob by putting the model into format
files = fn_format.format(model, expt, var)
print(files)
# open datasets & put them in a list
for fname in glob.iglob(files):
array_list.append(xr.open_dataset(fname))
model_namelist.append(model)
print('\n')
# +
#-------------------------------------------------------------------------------
new_sos_models_mh=[]
new_sos_models_ctrl=[]
new_sos_models_hi=[]
new_tos_models_mh=[]
new_tos_models_ctrl=[]
new_tos_models_hi=[]
sos_data_mh = []
sos_data_ctrl = []
sos_data_hi = []
tos_data_mh = []
tos_data_ctrl = []
tos_data_hi = []
get_filenames(sos_models_mh, tos_models_mh, 'midHolocene', 'sos', sos_data_mh,
new_sos_models_mh)
get_filenames(sos_models_mh, tos_models_mh, 'midHolocene', 'tos', tos_data_mh,
new_tos_models_mh)
get_filenames(sos_models_ctrl, tos_models_ctrl, 'piControl', 'sos', sos_data_ctrl,
new_sos_models_ctrl)
get_filenames(sos_models_ctrl, tos_models_ctrl, 'piControl', 'tos', tos_data_ctrl,
new_tos_models_ctrl)
get_filenames(sos_models_hi, tos_models_hi, 'historical', 'sos', sos_data_hi,
new_sos_models_hi)
get_filenames(sos_models_hi, tos_models_hi, 'historical', 'tos', tos_data_hi,
new_tos_models_hi)
# -
# # Coordinate system
#-------------------------------------------------------------------------------
def get_coord_names(fx):
'''
Discovers what the lat/lon variable in a dataset is called and returns this
as an array. Also converts -180-->180 deg longitudes to 0-->360 deg
longitudes.
INPUTS:
- fx: xarray DataSet e.g. sos_data_ctrl[0]
RETURNS:
- lat: array of latitudes
- lon: array of longitudes
'''
# work out what lat/lon var is called
if 'lat' in fx.variables:
lat = fx.variables['lat'].values
lon = fx.variables['lon'].values
if lon.max() < 300: # convert -180-->180 lons to 0-->360 lons
lon %= 360
elif 'nav_lat' in fx.variables:
lat = fx.variables['nav_lat'].values
lon = fx.variables['nav_lon'].values
if lon.max() < 300:
lon %= 360
elif 'latitude' in fx.variables:
lat = fx.variables['latitude'].values
lon = fx.variables['longitude'].values
if lon.max() < 300:
lon %= 360
else:
print("!!LAT/LON VAR NAME NOT RECOGNISED!!")
return lat, lon
#-------------------------------------------------------------------------------
def get_curvi_coords(fx, var, min_lat, max_lat, min_lon, max_lon, verbose):
'''
This code was developed by <NAME>, PhD candidate,
Geography UCL, 2017.
Returns variable over a specific lat-lon region by taking a subset
of the curvilinear coords i.e. for a variable X:
latitude.shape = (y.y)
longitude.shape = (x.x)
INPUTS:
- fx: xarray DataSet e.g. sos_data_ctrl[0]
- var: xarray DataArray e.g. sos_data_ctrl[0].sos
- min_lat: the minimum latitude (deg N)
- max_lat: the maximum latitude (deg N)
- min_lon: the minimum longitude (deg E)
- max_lon: the maximum longitude (deg E)
- verbose: if True, calculate the variable (e.g. sos) over the AOI,
if False, calculate the lat/lon variable over the AOI
(curvilinear coords)
RETURNS:
- var_ai: var in a specific lat-lon region
'''
print('***getting curvi coordinates***')
area = [min_lat, max_lat, min_lon, max_lon]
lat, lon = get_coord_names(fx)
# Specify area of interest as lat-lon degrees
# Produces boolean array
latt = np.logical_and(lat >= area[0], lat <= area[1])
lonn = np.logical_and(lon >= area[2], lon <= area[3])
# Select area of interest from combination of lat-lon arrays
# Produces boolean array
a_int = latt * lonn
# Indices for area of interest
# nonzero returns indices of elements that are non-zero (True)
(ii,jj) = a_int.nonzero()
if verbose:
# Var over AOI
# shape change: e.g. (8400, 384, 320) --> (8400, 185, 239)
var_ai = var[:, ii.min():ii.max(),jj.min():jj.max()] \
*a_int[ii.min():ii.max(),jj.min():jj.max()]
'''
# Show lat/lon field
# Boolean array, var*AOI, var over AOI only
vvv = [a_int, var[0,:,:]*a_int, var_ai[0,:,:]]
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(10, 2))
for vv,ax in zip(vvv,axes.flat):
im = ax.imshow(vv, origin=(20,10))
'''
else:
# Coords over AOI
# shape change: e.g. (384, 320) --> (185, 239)
var_ai = var[ii.min():ii.max(),jj.min():jj.max()] \
*a_int[ii.min():ii.max(),jj.min():jj.max()]
'''
# Show lat/lon field
# Boolean array, var*AOI, var over AOI only
vvv = [a_int, var[:,:]*a_int, var_ai[:,:]]
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(10, 2))
for vv,ax in zip(vvv,axes.flat):
im = ax.imshow(vv, origin=(20,10))
'''
print(var.shape, '-->', var_ai.shape)
print('***finished curvi coordinates***')
return var_ai
#-------------------------------------------------------------------------------
def get_pacific_coords(lat, lon, fs, ft, start_lat, end_lat, start_lon, end_lon):
'''
Determines whether lat/lon variable is rectilinear or curvilinear. If the
former, it selects a subset of the variable according to a start_lat,
end_lat, start_lon and end_lon. It then takes the subsequent lat/lon coord
as the lat/lon object. If the latter, it works out a subset of the variable
and lat/lon variable using the get_curvi_coords function. To gain a subset of
the lat/lon coordinates, it works out what the lat/lon variable is called
in the file.
INPUTS:
- lat: array of latitudes, e.g. fx.variables['lat'].values
- lon: array of longitudes, e.g. fx.variables['lon'].values
- start_lat: the minimum latitude (deg N)
- end_lat: the maximum latitude (deg N)
- start_lon: the minimum longitude (deg E)
- end_lon: the maximum longitude (deg E)
- fs, ft: xarray DataSet e.g. tos_data_ctrl[0]
RETURNS:
- IPsosVar, IPtosVar: var in a specific lat-lon region
- sos_latobj: latitude in a specific lat-lon region
- sos_lonobj: longitude in a specific lat-lon region
'''
# rectilinear
#try:
if len(lat.shape) == 1:
IPsosVar = fs.sos.sel(lat=slice(start_lat, end_lat),
lon=slice(start_lon, end_lon))
IPtosVar = ft.tos.sel(lat=slice(start_lat, end_lat),
lon=slice(start_lon, end_lon))
sos_latobj = IPsosVar.lat
sos_lonobj = IPsosVar.lon
# curvilinear
else:
IPsosVar = get_curvi_coords(fs, fs.sos, start_lat, end_lat,
start_lon, end_lon, verbose=True)
IPtosVar = get_curvi_coords(ft, ft.tos, start_lat, end_lat,
start_lon, end_lon, verbose=True)
if 'lat' in fs.variables:
sos_latobj = get_curvi_coords(fs, fs.lat, start_lat, end_lat,
start_lon, end_lon, verbose=False)
sos_lonobj = get_curvi_coords(ft, ft.lon, start_lat, end_lat,
start_lon, end_lon, verbose=False)
elif 'nav_lat' in fs.variables:
sos_latobj = get_curvi_coords(fs, fs.nav_lat, start_lat, end_lat,
start_lon, end_lon, verbose=False)
sos_lonobj = get_curvi_coords(ft, ft.nav_lon, start_lat, end_lat,
start_lon, end_lon, verbose=False)
elif 'latitude' in fs.variables:
sos_latobj = get_curvi_coords(fs, fs.latitude, start_lat, end_lat,
start_lon, end_lon, verbose=False)
sos_lonobj = get_curvi_coords(ft, ft.longitude, start_lat, end_lat,
start_lon, end_lon, verbose=False)
return IPsosVar, IPtosVar, sos_latobj, sos_lonobj
# # Compute corals
#-------------------------------------------------------------------------------
def coral_sensor_field(latArray, lonArray, sst, sss):
'''
This function implements the bivariate model of [1] to SST and SSS fields.
Adapted from: https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/pmip3_pcoral.py
INPUTS:
- latArray, lonArray, numpy 1D arrays
- sst (in K or degC), masked array
- sss (in psu*), masked array
RETURNS
- coral, the pseudocoral at the same locations as SST, SSS
- tosContri, the thermal contribution
- sosContri, the hydrological contribution
* assumes SSS in psu, so need to convert if this is not the case
[1] <NAME>. , <NAME> , <NAME> , <NAME> ,
and <NAME> (2011), Comparison of observed and simulated tropical
climate trends using a forward model of coral δ18O, Geophys. Res.
Lett., 38, L14706, doi:10.1029/2011GL048224.
'''
print('***doing coral_sensor_field***')
print('centering the fields')
# center the fields
nt, ny, nx = sss.shape
# Mean over time
sss_m = ma.mean(sss, axis=0)
sss_c = sss - np.tile(sss_m, (nt,1,1))
sst_m = ma.mean(sst, axis=0)
sst_c = sst - np.tile(sst_m, (nt,1,1))
print('assigning b-values')
# assign different b values based on location
a = -0.22
b1 = 0.3007062
b2 = 0.1552032
b3 = 0.2619054
b4 = 0.436509
b = np.empty((len(latArray),len(lonArray)))
for lat in range(len(latArray)):
for lon in range(len(lonArray)):
#Red sea
if lonArray[lon]>=32.83 and lonArray[lon]<=43.5 and \
latArray[lat]>=12.38 and latArray[lat]<=28.5:
b[lat][lon]=b1
#Indian ocean
elif lonArray[lon]<=120:
b[lat][lon]=b2
#Tropical Pacific
elif latArray[lat]>= -5 and latArray[lat]<=13:
b[lat][lon]=b3
#South Pacific
elif latArray[lat]< -5:
b[lat][lon]=b4
#Default: Tropical Pacific
else:
b[lat][lon]=b3
print('storing b-values')
# store coordinates of four b values seperately
b1_index = np.where(b == b1)
b2_index = np.where(b == b2)
b3_index = np.where(b == b3)
b4_index = np.where(b == b4)
# create a new array with the same shape as IPsos and compute coral
coral = np.empty_like(sss)
tosContri = np.empty_like(sst)
sosContri = np.empty_like(sss)
print('calculating contributions')
# hydrological contribution
for b_index, b in ((b1_index, b1), (b2_index, b2),
(b3_index, b3), (b4_index, b4)):
sosContri[:, b_index[0], b_index[1]] = b * sss_c[:, b_index[0],
b_index[1]]
# thermal contribution
tosContri = a * sst_c
# total contribution
coral = sosContri + tosContri
print('***finished coral_sensor_field***')
# export all three
return coral, tosContri, sosContri
#-------------------------------------------------------------------------------
def compute_corals(IPsosVar, IPtosVar, sos_latobj, sos_lonobj):
'''
This function implements the bivariate model of [1] to SST and SSS fields.
Adapted from: https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/pmip3_pcoral.py
INPUTS:
- IPsosVar, IPtosVar: var in a specific lat-lon region
- sos_latobj: latitude in a specific lat-lon region
- sos_lonobj: longitude in a specific lat-lon region
RETURNS
- tobj: the time variable
- sos_latobj: latitude in a specific lat-lon region
- sos_lonobj: longitude in a specific lat-lon region
- coral2: the pseudocoral at the same locations as SST, SSS
- tosContri: the thermal contribution
- sosContri: the hydrological contribution
* assumes SSS in psu, so need to convert if this is not the case
[1] <NAME>. , <NAME> , <NAME> , <NAME> ,
and <NAME> (2011), Comparison of observed and simulated tropical
climate trends using a forward model of coral δ18O, Geophys. Res.
Lett., 38, L14706, doi:10.1029/2011GL048224.
'''
print('***starting compute_corals***')
# define missing values
ma.set_fill_value(IPsosVar, 1e20)
ma.set_fill_value(IPtosVar, 1e20)
# load into arrays
IPsos = IPsosVar.values
IPtos = IPtosVar.values
# get the values for computations
sos_ma = ma.masked_equal(IPsos, 1e20)
sos_ma = ma.array(sos_ma, mask=np.isnan(sos_ma))
tos_ma = ma.masked_equal(IPtos, 1e20)
tos_ma = ma.array(tos_ma, mask=np.isnan(tos_ma))
# get the means map
sos_mean = ma.mean(sos_ma, axis=0)
tos_mean = ma.mean(tos_ma, axis=0)
# total means no seasonal cycle is removed from the computation
sos_mean_total = ma.mean(sos_ma)
if sos_mean_total <= 1: # MORE SOPHISTICATED EXCEPTION HANDLING HERE?
print ('times sos by 1000')
sos_ma = sos_ma * 1000
tobj = IPsosVar.time
timeArray = tobj.values
print('creating lat/lon arrays')
# detect whether variable is in curvilinear grid
# curvilinear
if len(sos_latobj.shape) == 2:
latArray = sos_latobj[:,0]
lonArray = sos_lonobj[0,:]
# rectangular
else:
latArray = sos_latobj
lonArray = sos_lonobj
# apply coral sensor model
coral, tosContri, sosContri = coral_sensor_field(latArray, lonArray,
tos_ma, sos_ma)
print('creating masked arrays')
coral2 = ma.masked_equal(coral, 1e20) # coral2.shape = 1200, 30, 108
tosContri = ma.masked_equal(tosContri, 1e20)
sosContri = ma.masked_equal(sosContri, 1e20)
print('***finished compute_corals***')
return tobj, sos_latobj, sos_lonobj, coral2, tosContri, sosContri
#-------------------------------------------------------------------------------
def coral_sensor_apply(ft, fs, expt, model):
'''
This function converts model output to pseudocoral, according to the
bivariate model of [1].
Adapted from: https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/pmip3_pcoral.py
INPUTS:
- ft: filename for SST field, with variable name tname [default = 'tos']
- fs: filename object field, with variable name sname [default = 'sos']
RETURNS
- eastern_vars, central_vars, western_vars: tuples of objects -->
e_tobj, e_sos_latobj, e_sos_lonobj, e_coral2, e_tosContri, e_sosContri
[1] <NAME>. , <NAME> , <NAME> , <NAME> ,
and <NAME> (2011), Comparison of observed and simulated tropical
climate trends using a forward model of coral δ18O, Geophys. Res. Lett.,
38, L14706, doi:10.1029/2011GL048224.
'''
print('***starting coral_sensor_apply***')
# get the start and end time steps
start_time_sos = fs.time[0]
end_time_sos = fs.time[-1]
start_time_tos = ft.time[0]
end_time_tos = ft.time[-1]
print('getting variables & lat/lon objects')
sos_latobj, sos_lonobj = get_coord_names(fs)
# EAST PACIFIC: -10, 0, 270, 280
e_IPsosVar, e_IPtosVar, e_sos_latobj, e_sos_lonobj = \
get_pacific_coords(sos_latobj, sos_lonobj, fs, ft, -10, 0, 270, 280)
# CENTRAL PACIFIC: -5, 5, 190, 240
c_IPsosVar, c_IPtosVar, c_sos_latobj, c_sos_lonobj = \
get_pacific_coords(sos_latobj, sos_lonobj, fs, ft, -5, 5, 190, 240)
# WEST PACIFIC: -20, 0, 120, 180
w_IPsosVar, w_IPtosVar, w_sos_latobj, w_sos_lonobj = \
get_pacific_coords(sos_latobj, sos_lonobj, fs, ft, -20, 0, 120, 180)
e_tobj, \
e_sos_latobj, \
e_sos_lonobj, \
e_coral2, \
e_tosContri, \
e_sosContri = compute_corals(e_IPsosVar,e_IPtosVar,e_sos_latobj,e_sos_lonobj)
c_tobj, \
c_sos_latobj, \
c_sos_lonobj, \
c_coral2, \
c_tosContri, \
c_sosContri = compute_corals(c_IPsosVar,c_IPtosVar,c_sos_latobj,c_sos_lonobj)
w_tobj, \
w_sos_latobj, \
w_sos_lonobj, \
w_coral2, \
w_tosContri, \
w_sosContri = compute_corals(w_IPsosVar,w_IPtosVar,w_sos_latobj,w_sos_lonobj)
eastern_vars = e_tobj, e_sos_latobj, e_sos_lonobj, e_coral2, e_tosContri, e_sosContri
central_vars = c_tobj, c_sos_latobj, c_sos_lonobj, c_coral2, c_tosContri, c_sosContri
western_vars = w_tobj, w_sos_latobj, w_sos_lonobj, w_coral2, w_tosContri, w_sosContri
###########################
# create a dictionary to store corals
corals = {}
corals['east'] = e_coral2
corals['central'] = c_coral2
corals['west'] = w_coral2
# save dictionary to a pickle file
pickle.dump(corals, open(base + 'corals/coral_{}_{}.p'.format(expt, model), "wb" ))
# save .mat
io.savemat(base + 'corals/coral_{}_{}.mat'.format(expt, model), corals)
print("saved!")
return eastern_vars, central_vars, western_vars
# # Bandpass & bootstrapping
def butter_bandpass(lowcut, highcut, fs, order=4):
'''
Adapted from https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/bandpass.py
'''
nyq = 0.5 * fs
low = lowcut / nyq
high = highcut / nyq
b, a = butter(order, [low, high], btype='bandpass')
return b, a
def butter_bandpass_filter(data, lowcut, highcut, fs, order=4):
'''
Adapted from https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/bandpass.py
'''
b, a = butter_bandpass(lowcut, highcut, fs, order=order)
y = filtfilt(b, a, data)
return y
# The bootstrap method is a statistical technique for estimating quantities about a population by averaging estimates from multiple small data samples. Samples are constructed by drawing observations from a large data sample one at a time and returning them to the data sample after they have been chosen. This allows a given observation to be included in a given small sample more than once. This approach to sampling is called sampling with replacement.
#
# <h6>Moving blocks sampling</h6>
# Break the series into roughly equal-length blocks of consecutive observations, to resample the block with replacement, and then paste the blocks together. There are $(n - b + 1)$ such blocks available, with consecutive samples of length b. This preserves dependency in the original samples to length b.
#
# For each boostrap sample, randomly select blocks and assemble into a length-n timeseries, then compute $\hat{\beta}^*$ for each such length-n series.
def block_bootstrap_ET(X, Lb, Nb):
'''
Implement Block Bootstrap as in:
http://nbviewer.ipython.org/github/welch/stats-notebooks/
blob/master/SubsamplingBootstrap.ipynb.
Adapted from https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/bootstrap.py.
INPUTS:
- X: the bootstrap sample, array
- Lb: needed to sample multiples of 12 years
- Nb: number of bootstrap samples
RETURNS:
- Xb: numpy array, resampled version, or "replicate" of data
'''
nt = len(X)
ns = int(np.ceil(nt/Lb))
Xb = np.zeros((Nb, nt))
for b in range(Nb):
for block_i, start in enumerate(np.random.randint(nt - Lb + 1, size=ns)):
try:
Xb[b, block_i*Lb:(block_i+1)*Lb] = X[start:start+Lb]
except ValueError:
# changing Lb to 12 as 24 would make (block_i+1)*Lb out of range for X
Xb[b, block_i*12:(block_i+1)*12] = X[start:start+12]
return Xb
def seasonal_cycle(Xb):
'''
Compute and isolate the seasonal cycle.
Adapted from https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/pcoral_bootstrap.py
INPUTS:
- Xb: numpy array, resampled version, or "replicate" of data
RETURNS:
- clim:
- anom:
'''
nb,nt = Xb.shape
ny = int(nt/12)
clim = np.empty((nb,12))
for i in range(12):
clim[:,i] = Xb[:,i::12].mean(axis=1)
print("clim", clim.shape)
anom = Xb - np.tile(clim,(1, ny))
return clim, anom
def computer(coral, Nb, Lb, windows):
'''
Compute variance & seasonal amplitude.
Adapted from https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/pcoral_bootstrap.py
INPUTS:
- coral:
- Nb: number of bootstrap samples
- Lb: needed to sample multiples of 12 years
- windows: sampling windows
RETURNS:
- variance, seasonal_amp:
'''
# filtering parameters
fs = 1
f_hi = 1/(12*2.0)
f_lo = fs/(12*7.0)
# compute spatial mean
spatial_mean = coral.mean(axis=(1,2))
print("spatial_mean", spatial_mean.shape)
print("coral", coral.shape)
# generate boostrap samples
Xb = block_bootstrap_ET(spatial_mean, Lb, Nb)
nw = len(windows) # number of windows
seasonal_amp = np.empty((nw, Nb))
variance = np.empty((nw, Nb))
index = 0 # loop over windows
for i in windows:
Xw = Xb[:, :i*12] # sample over window
clim, anom = seasonal_cycle(Xw) # isolate seasonal cycle
# compute seasonal amplitude
smax = np.nanmax(clim, axis=1)
smin = np.nanmin(clim, axis=1)
seasonal_amp[index, :] = smax - smin
# compute ENSO variance
anom2_7 = np.empty(anom.shape)
for b in range(Nb):
# apply bandpass filter
anom2_7[b, :] = butter_bandpass_filter(anom[b,:], f_lo, f_hi, fs)
# compute variance per se
variance[index,:] = np.var(anom2_7,axis=1)
index +=1 # update index
return (variance, seasonal_amp)
#-------------------------------------------------------------------------------
def create_stats(tos_data, sos_data, expt, model):
'''
Creates and stores statistics.
Adapted from https://github.com/CommonClimate/EmileGeay_NatGeo2015/blob/
master/code/python/pmip3_pcoral_bootstrap.py
INPUTS:
- tos_data
- sos_data
RETURNS:
- variance, seasonal_amp:
'''
# This script uses block bootstrap to randomize coral data [uses different
# sampling time length to generate distribution plot of seasonal cycle amplitude]?
eastern_vars, central_vars, western_vars = coral_sensor_apply(tos_data,
sos_data,
expt, model)
Nb = 1000 # number of bootstrap samples
Lb = 24 # needed to sample multiples of 12 years
windows = [50] # observation windows
nw = windows.__len__()
pcoral_boot_exp = {}; variance = {}; seasonal_amp = {}
# compute bootstrapped climate statistics on the three AOI
variance_e, seasonal_amp_e = computer(eastern_vars[3], Nb, Lb, windows)
variance_c, seasonal_amp_c = computer(central_vars[3], Nb, Lb, windows)
variance_w, seasonal_amp_w = computer(western_vars[3], Nb, Lb, windows)
# store variance results
variance = np.empty((3*nw, Nb))
variance[0:nw, :] = variance_e
variance[nw:2*nw, :] = variance_c
variance[2*nw:3*nw, :] = variance_w
# store seasonal amplitude results
seasonal_amp = np.empty((3*nw, Nb))
seasonal_amp[0:nw, :] = seasonal_amp_e
seasonal_amp[nw:2*nw, :] = seasonal_amp_c
seasonal_amp[2*nw:3*nw, :] = seasonal_amp_w
pcoral_boot_exp['var'] = variance
pcoral_boot_exp['seas'] = seasonal_amp
print(variance.shape, seasonal_amp.shape) # 0:6=east, 6:12=central, 12:18=west
print("Done!")
###########################
# save dictionary to a pickle file
pickle.dump(pcoral_boot_exp, open( base + 'bootstrapped_corals/pcoral_bootstrap_{}_{}.p'. \
format(expt, model), "wb" ))
# save .mat
io.savemat(base + 'bootstrapped_corals/pcoral_bootstrap_{}_{}.mat'. \
format(expt, model), pcoral_boot_exp)
print("saved!")
for i, model in enumerate(new_sos_models_mh):
outfile='corals/coral_mh_MODNAME.mat'
already_exists=os.path.isfile(outfile.replace('MODNAME',new_sos_models_mh[i]))
if not already_exists:
print("DOING MODEL: ", new_sos_models_mh[i])
create_stats(tos_data_mh[i], sos_data_mh[i], 'mh', new_sos_models_mh[i])
for i, model in enumerate(new_sos_models_ctrl):
outfile='corals/coral_ctrl_MODNAME.mat'
already_exists=os.path.isfile(outfile.replace('MODNAME',new_sos_models_ctrl[i]))
if not already_exists:
print("DOING MODEL: ", new_sos_models_ctrl[i])
create_stats(tos_data_ctrl[i], sos_data_ctrl[i], 'ctrl', new_sos_models_ctrl[i])
for i, model in enumerate(new_sos_models_hi):
outfile='corals/coral_hi_MODNAME.mat'
already_exists=os.path.isfile(outfile.replace('MODNAME',new_sos_models_hi[i]))
if not already_exists:
print("DOING MODEL: ", new_sos_models_hi[i])
create_stats(tos_data_hi[i], sos_data_hi[i], 'hi', new_sos_models_hi[i])
| pseudocorals/coral_Emile_Geay_all_files_15May.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import keras_ocr
# keras-ocr will automatically download pretrained
# weights for the detector and recognizer.
pipeline = keras_ocr.pipeline.Pipeline()
# Get a set of three example images
images = [
keras_ocr.tools.read(url) for url in [
'https://upload.wikimedia.org/wikipedia/commons/b/bd/Army_Reserves_Recruitment_Banner_MOD_45156284.jpg',
'https://upload.wikimedia.org/wikipedia/commons/e/e8/FseeG2QeLXo.jpg',
'https://upload.wikimedia.org/wikipedia/commons/b/b4/EUBanana-500x112.jpg'
]
]
# Each list of predictions in prediction_groups is a list of
# (word, box) tuples.
prediction_groups = pipeline.recognize(images)
# Plot the predictions
fig, axs = plt.subplots(nrows=len(images), figsize=(20, 20))
for ax, image, predictions in zip(axs, images, prediction_groups):
keras_ocr.tools.drawAnnotations(image=image, predictions=predictions, ax=ax)
# -
| OCR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Eodag basics
# ____________
#
# We present `eodag`'s basic features for **searching and downloading EO products**.
#
# To be able to follow this tutorial, you will need to install the additional Python package [ipyleaflet](https://ipyleaflet.readthedocs.io/en/latest/installation.html).
#
# ## Configuration
#
# Let's start by setting your personal credentials to access [PEPS service](https://peps.cnes.fr) by filling your username and password below:
import os
os.environ["EODAG__PEPS__AUTH__CREDENTIALS__USERNAME"] = "PLEASE_CHANGE_ME"
os.environ["EODAG__PEPS__AUTH__CREDENTIALS__PASSWORD"] = "<PASSWORD>"
# Let's check that `ipyleaflet` is available:
import ipyleaflet as ipyl
# Then we create a workspace directory `eodag_workspace_basics` where all our files and configuration will live:
workspace = 'eodag_workspace_basics'
if not os.path.isdir(workspace):
os.mkdir(workspace)
# Next, we create a configuration YAML file based on the following minimal content and save it in the workspace folder as `eodag_conf.yml`. Note that if we don't create such a configuration file `eodag` will load a default configuration from `~/.config/eodag/eodag.yml`.
#
# ```yaml
# peps:
# download:
# outputs_prefix: "absolute_path_to_eodag_workspace"
# extract: true
# ```
#
# This configuration file will indicate `eodag` to search for products made available by the PEPS service only, to download them in our workspace folder and to extract the downloaded products.
#
# The credentials could also have been defined in the configuration file. Since we have set them with environment variables earlier, we are good to go.
yaml_content = """
peps:
download:
outputs_prefix: "{}"
extract: true
""".format(os.path.abspath(workspace))
with open(os.path.join(workspace, 'eodag_conf.yml'), "w") as f_yml:
f_yml.write(yaml_content.strip())
# ## Search
#
# The first step before performing a **search** is to initialize the session by creating an `EODataAccessGateway` instance with the previous configuration file.
# +
# To have some basic feedback on what eodag is doing, we configure logging to output minimum information
from eodag.utils.logging import setup_logging
setup_logging(verbose=1)
from eodag.api.core import EODataAccessGateway
dag = EODataAccessGateway(os.path.join(workspace, 'eodag_conf.yml'))
# -
# Now let's search for Sentinel 2 L1C products over a period of 6 months in 2018 in the South of France (time in UTC), the search area being defined by its bounding box.
# +
from datetime import date
product_type = 'S2_MSI_L1C'
extent = {
'lonmin': 0,
'lonmax': 2,
'latmin': 43,
'latmax': 45
}
products, estimated_total_nbr_of_results = dag.search(
productType=product_type,
start='2018-06-01',
end='2018-12-01',
geom=extent
)
# -
estimated_total_nbr_of_results
len(products)
products[:3]
# `eodag` sent requests to the PEPS service and got as a result an estimated number of 767 (you might get a different value) products available matching the search criteria. The search also returned 20 products stored in the `products` variable that is a `SearchResult` instance, a list-like object. Each one of the products is an `EOProduct` instance that contains all the information required to download the product later.
#
# You can see that the number of retrieved products is lower than the estimated number of products available. This is the same behaviour as when you make a search on Google where you get 10 results per page. `eodag` works the same way, it is set by default to return the results from *page 1*, a single page being defined by a collection of *20 products*. This behaviour can be adapted with the parameters `page` and `items_per_page` of the `search` method.
# Now that we have found some products, we can easily store them in a GeoJSON file that we may use later to download them:
search_result_file = dag.serialize(products, filename=os.path.join(workspace, 'search_results.geojson'))
# The products can also be converted into a Python dictionnary structured as GeoJSON file. Doing so allows us to pass this dictionnary to an interactive plotting library such as `ipyleaflet` to check the extents of the products before downloading them:
# +
m = ipyl.Map(center=[43.6, 1.5], zoom=5)
layer = ipyl.GeoJSON(data=products.as_geojson_object(), hover_style={'fillColor': 'yellow'})
m.add_layer(layer)
m
# -
# ## Download
#
# Finally we could easily **download** all the returned products with `dag.download_all(products)`. Since products are heavy (a single product may already be over 700 Mo) we show how to download a single product with `dag.download(product)`. We also demonstrate the use of the `NotebookProgressCallback` to add a nice download progress bar:
# +
from eodag.utils import NotebookProgressCallback
# Download the second product found.
path = dag.download(products[1], progress_callback=NotebookProgressCallback())
path
# -
# When the download has completed you should find the product in your workspace folder. Note that above we set up the download with the Python object (`SearchResult`) returned by the search. It is also possible to recreate this search ouput thanks to the GeoJSON file we have saved previously with `dag.deserialize(path_to_geojson)`.
#
# This concludes the tutorial for the basic usage of `eodag`.
| examples/tuto_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming_Assingment7
# ### 1. Write a Python Program to find sum of array?
#
#Solution 1
lst = [1,2,3,4,5,6,7,8,9]
sum(lst)
#Solution 2
sum =0
lst = [1,2,3,4,5,6,7,8,9]
for i in range(0,len(lst)):
sum+=lst[i]
print("Sum is",sum)
# ### 2. Write a Python Program to find largest element in an array?
#
#solution 1
lst = [100,240,310,67,445,9]
max(lst)
print("lstgetst elemnt in the array {} is {}".format(lst,max(lst)))
# +
#Solution2
lst = [100,240,310,67,445,9]
big = 0
for i in range(0,len(lst)):
if big < lst[i]:
big = lst[i]
print("Largest element in the array {} is {} ".format(lst,big))
# -
# ### 3. Write a Python Program for array rotation?
#
# +
n = int(input("Enter the lenth of your list : "))
lst = []
for i in range(n):
lst.append(int(input()))
print("The List is ",lst)
rotation = input("Enter Rotation Right/Left : ")
nele = int(input("Enter the number of elements to rotate < : "))
if nele > n:
print("can not rotate as elements to rotate is larger then list lengh")
else:
rlst = []
if rotation.upper() == "RIGHT":
rlst[:] = lst[-nele:] + lst[:(n-nele)]
print("After right rotation : ",rlst)
elif rotation.upper() == "LEFT":
rlst[:] = lst[nele:n] + lst[:nele]
print("After Left rotation : ",rlst)
else:
print("Invalid Entry")
# -
# ### 4. Write a Python Program to Split the array and add the first part to the end?
#
# +
n = int(input("Enter the lenth of your list : "))
lst = []
for i in range(n):
lst.append(int(input()))
print("The List is ",lst)
nele = int(input("Enter the number of elements to split < : "))
if nele > n:
print("can not split as elements to split is larger then list lengh")
else:
print("The split list is :",lst[:nele])
rlst[:] = lst[nele:n] + lst[:nele]
print("The List after split and add :", rlst)
# -
# ### 5. Write a Python Program to check if given array is Monotonic?
# - An array is said to be monotonic in nature if it is either continuously increasing or continuously decreasing.
# +
n = int(input("Enter the lenth of your list : "))
lst = []
for i in range(n):
lst.append(int(input()))
print("The List is ",lst)
if all((lst[i] <= lst[i+1] for i in range(n-1)) or (lst[i] >= lst[i+1] for i in range(n-1))):
print("Monotonic")
else:
print("Not Monotonic")
| Programming_Assingment7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Tutorial 15. Feature engineering with audio files
#
# Created by <NAME> 2019 All content contained in this notebook is licensed under a [Creative Commons License 4.0 BY NC](https://creativecommons.org/licenses/by-nc/4.0/). The code is licensed under a [MIT license](https://opensource.org/licenses/MIT).
#
# This notebook is completely based on the following tutorial .
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats as stats
import matplotlib as mpl
import librosa
import librosa.display
import sklearn
import IPython.display as ipd
import TCD19_utils as TCD
TCD.set_plotting_style_2()
palette = sns.cubehelix_palette( rot = 0.3, as_cmap= True)
# %matplotlib inline
# This enables high res graphics inline
# %config InlineBackend.figure_format = 'svg'
# -
# ### Librosa is a great library to work with audio analysis in python
### Generating a sine wave with librosa
sr = 22050 # sample rate
T = 2.0 # seconds
t = np.linspace(0, T, int(T*sr), endpoint=False) # time variable
sine_wave = np.exp(0.2*np.sin(1.5*np.pi*300*t))# pure sine wave at 220 Hz
ipd.Audio(sine_wave, rate=sr) # load a NumPy array
plt.figure(figsize = (10, 4))
librosa.display.waveplot(sine_wave, sr=sr, color = 'salmon', alpha = 0.7);
audio_path = '../data/audio_sample.wav'
x , sr = librosa.load(audio_path)
import librosa.display
plt.figure(figsize=(8, 4))
librosa.display.waveplot(x, sr=sr, color = 'salmon', alpha = 0.7);
# ## Spectrogram
# A spectrogram is a visual representation of the spectrum of frequencies of sound or other signals as they vary with time.
spectrogram = np.abs(librosa.stft(x))
Xdb = librosa.amplitude_to_db(spectrogram)
# +
plt.figure(figsize=(8, 4))
librosa.display.specshow(Xdb, sr=sr, x_axis='time', y_axis='log', cmap = palette)
plt.colorbar(format='%+2.0f dB');
# -
# ## Zero crossing rate
# The zero crossing rate is the rate of sign-changes along a signal, i.e., the rate at which the signal changes from positive to negative or back. This feature has been used heavily in both speech recognition and music information retrieval. It usually has higher values for highly percussive sounds like those in metal and rock.
# +
n0 = 8500
n1 = 9100
plt.figure(figsize=(8, 4))
plt.plot(x[n0:n1], color = 'purple', alpha = 0.7)
plt.plot([-70, 700], [0, 0], lw = 4, color = 'gray', alpha = 0.4)
plt.xlim([-70, 700])
plt.grid()
# -
x.shape
zero_crossings = librosa.zero_crossings(x[n0:n1], pad=False)
print(sum(zero_crossings))
# ## spectral centroid
#
# It indicates where the ”centre of mass” for a sound is located and is calculated as the weighted mean of the frequencies present in the sound. Consider two songs, one from a blues genre and the other belonging to metal. Now as compared to the blues genre song which is the same throughout its length, the metal song has more frequencies towards the end. So spectral centroid for blues song will lie somewhere near the middle of its spectrum while that for a metal song would be towards its end.
#
#
# +
spectral_centroids = librosa.feature.spectral_centroid(x, sr=sr)[0]
spectral_centroids.shape
# Computing the time variable for visualization
frames = range(len(spectral_centroids))
t = librosa.frames_to_time(frames)
# Normalising the spectral centroid for visualisation
def normalize(x, axis=0):
return sklearn.preprocessing.minmax_scale(x, axis=axis)
#Plotting the Spectral Centroid along the waveform
librosa.display.waveplot(x, sr=sr, color = 'purple', alpha = 0.4)
plt.plot(t, normalize(spectral_centroids), color='lightgrey', lw = 3)
plt.ylim(-0.4, 0.4);
# -
# ## Spectral Rolloff
# It is a measure of the shape of the signal. It represents the frequency below which a specified percentage of the total spectral energy, e.g. 85%, lies.
spectral_rolloff = librosa.feature.spectral_rolloff(x+0.01, sr=sr)[0]
librosa.display.waveplot(x, sr=sr, color = 'purple', alpha = 0.4)
plt.plot(t, normalize(spectral_rolloff), color='lightgrey', lw = 4)
plt.ylim(-0.4, 0.4);
# ## Mel-Frequency Cepstral Coefficients
# The Mel frequency cepstral coefficients (MFCCs) of a signal are a small set of features (usually about 10–20) which concisely describe the overall shape of a spectral envelope. It models the characteristics of the human voice.
x.shape
mfccs = librosa.feature.mfcc(x, sr=sr, n_mfcc = 20)
print(mfccs.shape)
#Displaying the MFCCs:
plt.figure(figsize=(8, 4))
librosa.display.specshow(mfccs, sr=sr, x_axis='time', cmap = palette)
plt.colorbar();
# Here mfcc computed 20 MFCC s over 97 frames.
#
# We can also perform feature scaling such that each coefficient dimension has zero mean and unit variance:
# +
mfccs_scaled = sklearn.preprocessing.scale(mfccs, axis=1)
plt.figure(figsize=(8, 4))
librosa.display.specshow(mfccs_scaled, sr=sr, x_axis='time', cmap = palette)
plt.colorbar();
# -
# ## Chroma frequencies
# +
hop_length = 512
chromagram = librosa.feature.chroma_stft(x, sr=sr, hop_length=hop_length)
plt.figure(figsize=(8, 5))
librosa.display.specshow(chromagram, x_axis='time', y_axis='chroma',
hop_length=hop_length, cmap = palette)
plt.colorbar();
# -
| notebooks/t15-TCD_feature_engineering_audio_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: test
# language: python
# name: test
# ---
# + deletable=true editable=true
"""Required imports"""
import json
import urllib
from bs4 import BeautifulSoup
# -
# We have to generate the webpage dynamically since it's mostly JS.
from contextlib import closing
from selenium.webdriver import Chrome # pip install selenium
from selenium.webdriver.support.ui import WebDriverWait
# +
class FailedBookRetrievalError(Exception):
def __init__(self):
super(FailedBookRetrievalError, self).__init__()
class FailedAuthorRetrievalError(Exception):
def __init__(self):
super(FailedAuthorRetrievalError, self).__init__()
class Book(object):
def __init__(self, title, author, gr_id):
self.title = title.encode('ascii', 'ignore')
self.author = author
self.id = gr_id
self.href = 'https://www.goodreads.com/book/show/%s' % self.id
def __eq__(self, other):
return self.id == other.id
def __str__(self):
return "%s by %s" % (self.title, self.author.name)
def __hash__(self):
return hash(str(self.id))
def __repr__(self):
return "%s: %s" % (str(self), self.href)
class Author(object):
def __init__(self, name, gr_id):
self.name = name.encode('ascii', 'ignore')
self.id = gr_id
self.href = 'https://www.goodreads.com/author/show/%s' % self.id
def __eq__(self, other):
return self.id == other.id
def __str__(self):
return self.name
def __hash__(self):
return hash(str(self.id))
def __repr__(self):
return "%s: %s" % (str(self), self.href)
# +
def extract_author(anchor):
'''
Given an anchor .authorName extracts an Author object.
'''
try:
gr_id = anchor['href'].split('/')[-1]
name = anchor.find('span').contents[0]
except:
raise FailedAuthorRetrievalError(anchor)
return Author(name=name, gr_id=gr_id)
def extract_book(div):
'''
Given an .elementList div for a book, returns a Book object.
'''
try:
anchor_tag = div.find('a', {'class': 'bookTitle'})
title = anchor_tag.contents[0]
gr_id = anchor_tag['href'].split('/')[-1]
except:
raise FailedBookRetrievalError(anchor_tag)
try:
author_tag = div.find('a', {'class': 'authorName'})
author = extract_author(author_tag)
except:
author = None
return Book(title=title, author=author, gr_id=gr_id)
# -
def get_books(html):
'''
Extracts a set of Books from the html raw source if possible.
'''
soup = BeautifulSoup(html, 'html.parser')
result_list = soup.find('div', { 'class': 'leftContainer'})
try:
results = result_list.find_all('div', {'class': 'elementList'})
except:
print "Failed to find any results on page."
return set()
books = []
for res in results:
try:
books.append(extract_book(res))
except FailedBookRetrievalError:
print "Failed to extract book from %s" % res
pass
return set(books)
def get_raw_html(browser, shelf, page=1):
'''
Fetches the raw html for the page corresponding to `shelf`
and `page` number.
'''
service_url = 'https://www.goodreads.com/shelf/show/'
params = {
'page': page
}
url = service_url + shelf + '?' + urllib.urlencode(params)
browser.get(url)
return browser.page_source
def get_shelf(browser, term, n = 100):
'''
Given a browser and a shelf name, returns a list
of n `Book`s as determined by Good Reads.
'''
page = 1
books = set()
failed_sequence = 0
while len(books) < n:
html = get_raw_html(browser, term, page)
books_in_page = get_books(html)
if len(books_in_page) == 0: # no new books.
failed_sequence += 1
else:
failed_sequence = 0
if failed_sequence > 10:
break
books = books.union(books_in_page)
page += 1
return books
def login(browser, email, password):
'''
Login to GoodReads so we can access more data.
'''
url = 'https://www.goodreads.com/user/sign_in'
browser.get(url)
browser.find_element_by_id('user_email').send_keys(email)
browser.find_element_by_id('user_password').send_keys(password)
browser.find_element_by_name('next').click()
# + deletable=true editable=true
import pickle
def dump_book_set(s, filename):
with open(filename, 'w+') as handle:
pickle.dump(s, handle)
# -
def find_books(tags, n = 500):
'''
Given a list of tags, finds their intersection by looking
at the top n books in each tag and intersecting the resulting
sets.
Returns:
A set of Book objects. It may be helpful to run something like:
for book in book:
print book
'''
book_set = set()
with closing(Chrome('./chromedriver')) as browser:
login(browser, '<EMAIL>', 'luis3137')
for tag in tags:
shelf = get_shelf(browser, tag, n = n)
dump_book_set(shelf, "%s_n=%s_set.pk" % (tag, n))
book_set = book_set.intersection(shelf)
print "Finished collecting data for tag %s." % tag
return book_set
books = find_books(['adult'], n = 100000)
for book in books:
print book.href + " author: " + str(book.author)
# Load all of the results and intersect.
a, b, c, d = (pickle.load(open('strong-heroine_n=100000_set.pk')),
pickle.load(open('romance_n=100000_set.pk')),
pickle.load(open('female-lead_n=100000_set.pk')),
pickle.load(open('young-adult_n=100000_set.pk')))
for book in set.intersection(a,b,c,d):
print book.href + " author: " + str(book.author)
# + deletable=true editable=true
# Good reads data pulling.
def get_shelf(shelf_name):
service_url = 'https://www.goodreads.com/shelf/show/'
url = service_url + '?' + shelf_name
response = urllib.urlopen(url).read()
return response
# + deletable=true editable=true
get_shelf("fiction")
# + deletable=true editable=true
| Good Reads.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Set routines
import numpy as np
np.__version__
author = 'kyubyong. <EMAIL>'
# ## Making proper sets
# Q1. Get unique elements and reconstruction indices from x. And reconstruct x.
# +
x = np.array([1, 2, 6, 4, 2, 3, 2])
# -
# ## Boolean operations
# Q2. Create a boolean array of the same shape as x. If each element of x is present in y, the result will be True, otherwise False.
x = np.array([0, 1, 2, 5, 0])
y = np.array([0, 1])
# Q3. Find the unique intersection of x and y.
x = np.array([0, 1, 2, 5, 0])
y = np.array([0, 1, 4])
# Q4. Find the unique elements of x that are not present in y.
x = np.array([0, 1, 2, 5, 0])
y = np.array([0, 1, 4])
# Q5. Find the xor elements of x and y.
# +
x = np.array([0, 1, 2, 5, 0])
y = np.array([0, 1, 4])
out1 = np.setxor1d(x, y)
out2 = np.sort(np.concatenate((np.setdiff1d(x, y), np.setdiff1d(y, x))))
assert np.allclose(out1, out2)
# -
# Q6. Find the union of x and y.
x = np.array([0, 1, 2, 5, 0])
y = np.array([0, 1, 4])
out1 = np.union1d(x, y)
out2 = np.sort(np.unique(np.concatenate((x, y))))
assert np.allclose(out1, out2)
| Set_routines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The neuron can be thought of as an electrical circuit. It has ion fluxes through protein doorways that are called ion channels that are sensitive to voltage across the cell membrane. Ionic fluxes through ion channels in the cell membrane have dynamics that can be described by physics and mathematical models. Using concepts from electrical physics we can study how brief pulses of depolarization (the change of the relative voltage between the inside and outside of the cell) called action potentials are generated. Action potentials serve as physical substrates of information exchange between neurons as they form intricate communication and information processing networks.
#
# We begin studying the electrical properties of neurons with simplified models that serve as stepping stones to understanding the electro-chemical dynamics of information processing in the brain. In particular, we study how membrane capacitance and resistance allow neurons to integrate and filter synaptic currents over time.
# # The Cell Membrane
# The membrane of a cell is what defines its identity in relation to its surroundings. It is an extremely thin sheet (5-10 nm, 1000 x thinner than the human hair) of polarized phospholipids. A phospholipid molecule consists of a charged phosphite molecule and a long electrically neutral hydrocarbon chain. When placed in water, phospholipids self-organize into a bilayer, meaning that the charged hydrophilic phosphite molecules interface with water while the hydrophobic electrically neutral hydrocarbon chains hide themselves from the water medium. It's like a molecular sandwich with the phosphites forming the bun and the hydrocarbon chains forming the veggie patty.
#
# The cell membrane insulates the inside from the outside and its hydrophobic part prevents the movement of ions across it. However, life would be boring if the cell was merely a blob that could not exchange matter information with its environment. That's why cells evolved membrane spanning proteins called ion channels. They are like doors, some of them with key holes (meaning that they only open when a specific molecule binds to it), others opening in response to the change the difference in positive and negative ion concentration between the inside and outside of the cell (charge difference or voltage) and still others that are always open. These ion channels allow specific ions to pass through the membrane and thus enable intermembrane ion fluxes.
#
# In nerve cells, the concentration of ions is different in the inside and outside of the cell. The water in which the cell is embedded and is filled with acts as a polar solvent that dissolves small inorganic molecules. The water is a medium or a "wire" that conducts the various ionic flows that act as a substrate for biological information processing. Sodium, chloride and calcium ions are more abundant outside of the cell membrane and potassium ions are more abundant inside (a good way to remember that is that Na and Cl are basically dissociated salt molecules, therefore the neuron is essentially embedded in salt water exactly as it evolved from single cells in the primodial ocean, there are about $6
# \cdot 10^{19}$ salt molecules dissolved in a cubic centimeter of an extracellular solution-- by comparison the age of the Universe is on the order of $10^{17}$ seconds). Also, there is a difference in charge concetration between the inside and the outside of the membrane which gives rise to a voltage difference. This voltage difference is essential for neural signaling and one of the bases of action potentials.
#
# The so called power supplies of the membrane, i.e. the molecular elements that maintain a voltage difference between the inside and outside of the cell, are the membrane-spanning ion pumps. They use energy to export three sodium ions and import two potassium ions in one cycle of operation. They need ATP (energy) for that because sodium is more abundant outside of the cell and potassium is more abundant inside the cell, they are thus working against the concentration gradient.
#
#
# It is the movement of ions across the cell membrane that generates the electrical signals in the brain. Ion channels are the basis of neural signaling that forms the information content of all of our thoughts.
# # Capacitance
# A capacitor in an electric circuit is an element that doesn't let charged particles get through. Ions accumulate on the opposing sides of the capacitor. It is thus said to store charge. In the neuronal membrane, the capacitor is the phospholipid bilayer. It lets the neuronal membrane to act as an integrator of past inputs.
#
# The formula for capicance is:
#
# $C=\frac{q}{V}$
#
# where $q$ is charge and $V$ is voltage.
#
# The physical unit of capacitance is Farad.
#
# # Resistance
#
#
# The capacitor and resistor that are connected in parallel in an electric circuit act as temporal filters that smooth the synaptic input over time.
# # Voltage
# Voltage is the difference in electrical potential between two points (e.g. how much net positive and negative charge are at two points). The formula for voltage is:
#
# $V=I \cdot R$
#
# where $I$ is the current and $R$ is the resistance.
#
# The physical unit of voltage is Volt.
#
# Neural computation and signaling is influenced by the fluctuations of voltage differences across the cell membrane. The voltage across the cell membrane changes because of the injection of synaptic currents. This is how information flow from the neuronal network that a cell is embedded in drives these voltage fluctuations.
#
# Voltage changes are very fast, so they are an ideal substrate for propagating information in the brain, considering that the world changes very fast and neural systems have to be able to detect and react to these events.
# # Ionic current
# Currents flow through the cell membrane due to two driving forces: diffusion and electrostatic forces (drift of charged particles in an electric field). Because there is an uneven concentration of different ions inside and outside the cell membrane, these concentration gradients act as one of the driving forces for intracellular ionic currents and when ion channels for a particular ion open in the membrane, those particular ions will flow down their concentration gradient. This is because they "want" to achieve an equilibrium in their distribution, i.e. they will tend to redistribute themselves so that their distribution is even, meaning that they will flow in a direction where there is less of them. Diffusion depends on the temperature of the system. Diffusion is the cause of the redistribution of dye when you drop a few drops into a water container.
#
# During diffusion particles move and collide with each other and thus trace out random trajectories called a random walk. For example, a sodium ion in a physiological solution will collide with another molecule $10^{13}$ times per second. Diffusion is fast at short length scales. To diffuse across a cell body (10 micrometers) takes 50 milliseconds. Diffusion is slow at large length scales. To diffuse down a dendrite (1 millimeter) takes 10 minutes. To diffuse from the cell body down the motor neuron axon (1 m) it takes 10 years.
# # RC-Circuit -- The Passive Membrane
# In the passive membrane model, the resistance of the membrane, which corresponds to the ion channels embedded in it, is independent of the voltage across the membrane. In this model, the membrane is an RC-circuit composed of ion channels that are not sensitive to voltage-- the resistors R and the lipid bilayer which insulates the inside of the cell from its outside-- the capacitor C.
#
# When a pre-synaptic spike induces the transport of neurotransmitters across the synaptic junction and their binding to the post-synaptic ion channel receptors, these ion channels can open and let ions through. This flow of ions across the cell membrane is a current.
#
# The ionic current in this process thus splits into two components: the current that spreads itself on the capacitor and the current that goes through the resistor (through the ion channels). The current is thus "filtered" by the membrane.
#
# The impulse response function of this membrane, i.e. the RC-circuit, in terms of the voltage fluctuation induced by a brief pulse of current is an exponentially decaying function. Because the RC-circuit is linear, when we simulate multiple pulses of current going through the circuit, the resulting voltage response of the circuit is the convolution of the impulse response function with the timeseries of pulses. This means that the impulse responses add up linearly. For arbitrary time-dependent current input, we again convolve this current with the impulse response function.
#
# Let's now derive a differential equation to describe the response of the RC-circuit to a time-dependent current input.
#
# Membrane spanning ion pumps and the push-and-pull of diffusion and electrostatic forces on ions create an equilibrium potential that is maintained in the lack of current input. This membrane potential is called $V_{rest}$ and is like a battery in the RC-circuit. Okay, so we inject some current into the circuit. As mentioned before this current will split into a current that goes onto the capacitor ($I_{C}$) and a current that goes through the resistor ($I_{R}$). Therefore, because the total current is conserved:
#
# $I_{ext}=I_{R}+I_{C}$
#
# Let's take a look at the capacitive current (see also the section above).
#
# $C=\frac{q}{V}$
#
# $q=V \cdot C $
#
# Because current is just a flow of charge in time, we can construct the following differential equation by differentiating both terms with respect to time:
#
# $I_{C}=\frac{dq}{dt}=C \cdot \frac{dV}{dt}$
#
# What about the current on the resistor? The voltage across the resistor is described by the following equation:
#
#
# $V-V_{rest}=I_{R} \cdot R$
#
# $I_{R}=\frac{V-V_{rest}}{R}$
#
# Pluggin these two equations into the equation for the total current we have:
#
# $I_{ext}=\frac{V-V_{rest}}{R}+C \cdot \frac{dV}{dt}$
#
# Because we care about the evolution of voltage across the cell membrane, we can then re-arrange the terms:
#
# $C \cdot \frac{dV}{dt}=-\frac{V-V_{rest}}{R}+ I_{ext}$
#
# Another slight re-arrangement: we can multiply both sides by $R$. The term $R \cdot C$ is a constant that characterizes how fast the RC-circuit integrates inputs and it's called $\tau$ (it's the time-constant of the RC-circuit).
#
# $\tau \cdot \frac{dV}{dt}=-(V-V_{rest})+ R \cdot I_{ext}$
#
#
# # Response to current input
# We want to see how an RC-circuit responds to different current inputs.
#
# Consider the following experimental set up: a neuron is resting in a saline dish. We insert two electrodes through its membrane, one to measure the voltage difference between the intracellular and extracellular medium and one to inject currents into the neuron. We do this because we want to understand how the neuron transforms the synaptic currents from sensory input and other neurons into a characteristic response. We now set out to model this experimental set up.
| NEU_DYN_COUR/RC-Circuit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Feature Selection
#
# In the following cells, we will select a group of variables, the most predictive ones, to build our machine learning model.
#
# We will select variables using the **Lasso regression:** Lasso has the property of setting the coefficient of non-informative variables to zero.
#
# This has some interesting properties:
#
# 1. For production: Fewer variables mean smaller client input requirements (e.g. customers filling out a form on a website or mobile app), and hence less code for error handling. This reduces the chances of introducing bugs.
#
# 2. For model performance: Fewer variables mean simpler, more interpretable, better generalizing models
# +
# Import basic libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Feature selection packages
from sklearn.linear_model import Lasso
from sklearn.feature_selection import SelectFromModel
# Visualize all columns in the dataframe
pd.pandas.set_option('display.max_columns', None)
# +
# Load the data obtained in the feature engineering notebook
X_train = pd.read_csv('xtrain.csv')
X_test = pd.read_csv('xtest.csv')
X_train.head()
# +
# Define target variables
y_train = X_train['Survived']
y_test = X_test['Survived']
# Drop target variables from X's datasets
X_train = X_train.drop(['Survived'], axis=1)
X_test = X_test.drop(['Survived'], axis=1)
# -
X_train.head()
# ### Feature Selection
#
# Keep in mind we are in the research enviroment of the ML pipeline. Therefore, we applying Lasso Regression it is importan to set the seed.
# +
# We specify Lasso Regression model with a penalty coefficient
# (alpha). The bigger the alpha, the less features will be
# selected.
# selectFromModel object from sklearn, which will select
# automatically the features which coefficients are non-zero
selector = SelectFromModel(Lasso(alpha=0.005, random_state=0))
# Train the selector to choose features
selector.fit(X_train, y_train)
# -
# We can visualize which features are selected
selector.get_support()
# It is good that Lasso Regressor does not consider *Fare* variable, as the transformation of that variable was quite tricky and far from be Gaussian.
# +
# Make a list of selected features
selected_feat = X_train.columns[(selector.get_support())]
# Print the numbers of features
print('total features: {}'.format((X_train.shape[1])))
print('selected features: {}'.format(len(selected_feat)))
print('features with coefficients shrank to zero: {}'.format(
np.sum(selector.estimator_.coef_ == 0)))
# -
# Print selected variables
selected_feat
# Store the selected features
pd.Series(selected_feat).to_csv('selected_features.csv', index=False, header=False)
| Research_environment/ML pipeline research environment/ML_pipeline_research_feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
'''
Graph and Loss visualization using Tensorboard.
'''
from __future__ import print_function
import tensorflow as tf
# -
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
# Parameters
learning_rate = 0.01
training_epochs = 25
batch_size = 100
display_epoch = 1
logs_path = '/users/ektas/tensorboardlogs_mnist/'
# tf Graph Input
# mnist data image of shape 28*28=784
x = tf.placeholder(tf.float32, [None, 784], name='InputData')
# 0-9 digits recognition => 10 classes
y = tf.placeholder(tf.float32, [None, 10], name='LabelData')
# Set model weights
W = tf.Variable(tf.zeros([784, 10]), name='Weights')
b = tf.Variable(tf.zeros([10]), name='Bias')
# Construct model and encapsulating all ops into scopes, making
# Tensorboard's Graph visualization more convenient
with tf.name_scope('Model'):
# Model
pred = tf.nn.softmax(tf.matmul(x, W) + b) # Softmax
with tf.name_scope('Loss'):
# Minimize error using cross entropy
cost = tf.reduce_mean(-tf.reduce_sum(y*tf.log(pred), reduction_indices=1))
with tf.name_scope('SGD'):
# Gradient Descent
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
with tf.name_scope('Accuracy'):
# Accuracy
acc = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
acc = tf.reduce_mean(tf.cast(acc, tf.float32))
# Initialize the variables (i.e. assign their default value)
init = tf.global_variables_initializer()
# Create a summary to monitor cost tensor
tf.summary.scalar("loss", cost)
# Create a summary to monitor accuracy tensor
tf.summary.scalar("accuracy", acc)
# Merge all summaries into a single op
merged_summary_op = tf.summary.merge_all()
# Start training
with tf.Session() as sess:
# Run the initializer
sess.run(init)
# op to write logs to Tensorboard
summary_writer = tf.summary.FileWriter(logs_path, graph=tf.get_default_graph())
# Training cycle
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = int(mnist.train.num_examples/batch_size)
# Loop over all batches
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# Run optimization op (backprop), cost op (to get loss value)
# and summary nodes
_, c, summary = sess.run([optimizer, cost, merged_summary_op],
feed_dict={x: batch_xs, y: batch_ys})
# Write logs at every iteration
summary_writer.add_summary(summary, epoch * total_batch + i)
# Compute average loss
avg_cost += c / total_batch
# Display logs per epoch step
if (epoch+1) % display_epoch == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", "{:.9f}".format(avg_cost))
print("Optimization Finished!")
summary_writer.close()
# Test model
# Calculate accuracy
print("Accuracy:", acc.eval({x: mnist.test.images, y: mnist.test.labels}))
| Section 7/Code/tensorboard_MNIST_Section7.ipynb |
# + [markdown] colab_type="text" id="view-in-github"
# [View in Colaboratory](https://colab.research.google.com/github/DillipKS/MLCC_assignments/blob/master/intro_to_sparse_data_and_embeddings.ipynb)
# + [markdown] colab_type="text" id="JndnmDMp66FL"
# #### Copyright 2017 Google LLC.
# + colab={} colab_type="code" id="hMqWDc_m6rUC"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="PTaAdgy3LS8W"
# # Intro to Sparse Data and Embeddings
#
# **Learning Objectives:**
# * Convert movie-review string data to a sparse feature vector
# * Implement a sentiment-analysis linear model using a sparse feature vector
# * Implement a sentiment-analysis DNN model using an embedding that projects data into two dimensions
# * Visualize the embedding to see what the model has learned about the relationships between words
#
# In this exercise, we'll explore sparse data and work with embeddings using text data from movie reviews (from the [ACL 2011 IMDB dataset](http://ai.stanford.edu/~amaas/data/sentiment/)). This data has already been processed into `tf.Example` format.
# + [markdown] colab_type="text" id="2AKGtmwNosU8"
# ## Setup
#
# Let's import our dependencies and download the training and test data. [`tf.keras`](https://www.tensorflow.org/api_docs/python/tf/keras) includes a file download and caching tool that we can use to retrieve the data sets.
# + colab={"base_uri": "https://localhost:8080/", "height": 130} colab_type="code" id="jGWqDqFFL_NZ" outputId="1ae1631e-07e4-4450-9e2f-79a87f8bfa3c"
from __future__ import print_function
import collections
import io
import math
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from IPython import display
from sklearn import metrics
tf.logging.set_verbosity(tf.logging.ERROR)
train_url = 'https://dl.google.com/mlcc/mledu-datasets/sparse-data-embedding/train.tfrecord'
train_path = tf.keras.utils.get_file(train_url.split('/')[-1], train_url)
test_url = 'https://dl.google.com/mlcc/mledu-datasets/sparse-data-embedding/test.tfrecord'
test_path = tf.keras.utils.get_file(test_url.split('/')[-1], test_url)
# + [markdown] colab_type="text" id="6W7aZ9qspZVj"
# ## Building a Sentiment Analysis Model
# + [markdown] colab_type="text" id="jieA0k_NLS8a"
# Let's train a sentiment-analysis model on this data that predicts if a review is generally *favorable* (label of 1) or *unfavorable* (label of 0).
#
# To do so, we'll turn our string-value `terms` into feature vectors by using a *vocabulary*, a list of each term we expect to see in our data. For the purposes of this exercise, we've created a small vocabulary that focuses on a limited set of terms. Most of these terms were found to be strongly indicative of *favorable* or *unfavorable*, but some were just added because they're interesting.
#
# Each term in the vocabulary is mapped to a coordinate in our feature vector. To convert the string-value `terms` for an example into this vector format, we encode such that each coordinate gets a value of 0 if the vocabulary term does not appear in the example string, and a value of 1 if it does. Terms in an example that don't appear in the vocabulary are thrown away.
# + [markdown] colab_type="text" id="2HSfklfnLS8b"
# **NOTE:** *We could of course use a larger vocabulary, and there are special tools for creating these. In addition, instead of just dropping terms that are not in the vocabulary, we can introduce a small number of OOV (out-of-vocabulary) buckets to which you can hash the terms not in the vocabulary. We can also use a __feature hashing__ approach that hashes each term, instead of creating an explicit vocabulary. This works well in practice, but loses interpretability, which is useful for this exercise. See the tf.feature_column module for tools handling this.*
# + [markdown] colab_type="text" id="Uvoa2HyDtgqe"
# ## Building the Input Pipeline
# + [markdown] colab_type="text" id="O20vMEOurDol"
# First, let's configure the input pipeline to import our data into a TensorFlow model. We can use the following function to parse the training and test data (which is in [TFRecord](https://www.tensorflow.org/programmers_guide/datasets) format) and return a dict of the features and the corresponding labels.
# + colab={} colab_type="code" id="SxxNIEniPq2z"
def _parse_function(record):
"""Extracts features and labels.
Args:
record: File path to a TFRecord file
Returns:
A `tuple` `(labels, features)`:
features: A dict of tensors representing the features
labels: A tensor with the corresponding labels.
"""
features = {
"terms": tf.VarLenFeature(dtype=tf.string), # terms are strings of varying lengths
"labels": tf.FixedLenFeature(shape=[1], dtype=tf.float32) # labels are 0 or 1
}
parsed_features = tf.parse_single_example(record, features)
terms = parsed_features['terms'].values
labels = parsed_features['labels']
return {'terms':terms}, labels
# + [markdown] colab_type="text" id="SXhTeeYMrp-l"
# To confirm our function is working as expected, let's construct a `TFRecordDataset` for the training data, and map the data to features and labels using the function above.
# + colab={"base_uri": "https://localhost:8080/", "height": 36} colab_type="code" id="oF4YWXR0Omt0" outputId="05daded9-06af-4d1e-8968-5810af9864f5"
# Create the Dataset object.
ds = tf.data.TFRecordDataset(train_path)
# Map features and labels with the parse function.
ds = ds.map(_parse_function)
ds
# + [markdown] colab_type="text" id="bUoMvK-9tVXP"
# Run the following cell to retrieve the first example from the training data set.
# + colab={"base_uri": "https://localhost:8080/", "height": 564} colab_type="code" id="Z6QE2DWRUc4E" outputId="b646618c-bf9b-44f8-d7bf-e49f8d7e18fd"
n = ds.make_one_shot_iterator().get_next()
sess = tf.Session()
sess.run(n)
# + [markdown] colab_type="text" id="jBU39UeFty9S"
# Now, let's build a formal input function that we can pass to the `train()` method of a TensorFlow Estimator object.
# + colab={} colab_type="code" id="5_C5-ueNYIn_"
# Create an input_fn that parses the tf.Examples from the given files,
# and split them into features and targets.
def _input_fn(input_filenames, num_epochs=None, shuffle=True):
# Same code as above; create a dataset and map features and labels.
ds = tf.data.TFRecordDataset(input_filenames)
ds = ds.map(_parse_function)
if shuffle:
ds = ds.shuffle(10000)
# Our feature data is variable-length, so we pad and batch
# each field of the dataset structure to whatever size is necessary.
ds = ds.padded_batch(25, ds.output_shapes)
ds = ds.repeat(num_epochs)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# + [markdown] colab_type="text" id="Y170tVlrLS8c"
# ## Task 1: Use a Linear Model with Sparse Inputs and an Explicit Vocabulary
#
# For our first model, we'll build a [`LinearClassifier`](https://www.tensorflow.org/api_docs/python/tf/estimator/LinearClassifier) model using 50 informative terms; always start simple!
#
# The following code constructs the feature column for our terms. The [`categorical_column_with_vocabulary_list`](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_list) function creates a feature column with the string-to-feature-vector mapping.
# + colab={} colab_type="code" id="B5gdxuWsvPcx"
# 50 informative terms that compose our model vocabulary
informative_terms = ("bad", "great", "best", "worst", "fun", "beautiful",
"excellent", "poor", "boring", "awful", "terrible",
"definitely", "perfect", "liked", "worse", "waste",
"entertaining", "loved", "unfortunately", "amazing",
"enjoyed", "favorite", "horrible", "brilliant", "highly",
"simple", "annoying", "today", "hilarious", "enjoyable",
"dull", "fantastic", "poorly", "fails", "disappointing",
"disappointment", "not", "him", "her", "good", "time",
"?", ".", "!", "movie", "film", "action", "comedy",
"drama", "family")
terms_feature_column = tf.feature_column.categorical_column_with_vocabulary_list(key="terms", vocabulary_list=informative_terms)
# + [markdown] colab_type="text" id="eTiDwyorwd3P"
# Next, we'll construct the `LinearClassifier`, train it on the training set, and evaluate it on the evaluation set. After you read through the code, run it and see how you do.
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="HYKKpGLqLS8d" outputId="611b139c-5a3e-429c-9e68-1c21155be636"
my_optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
feature_columns = [ terms_feature_column ]
classifier = tf.estimator.LinearClassifier(
feature_columns=feature_columns,
optimizer=my_optimizer,
)
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] colab_type="text" id="J0ubn9gULS8g"
# ## Task 2: Use a Deep Neural Network (DNN) Model
#
# The above model is a linear model. It works quite well. But can we do better with a DNN model?
#
# Let's swap in a [`DNNClassifier`](https://www.tensorflow.org/api_docs/python/tf/estimator/DNNClassifier) for the `LinearClassifier`. Run the following cell, and see how you do.
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="jcgOPfEALS8h" outputId="00568e99-75ba-4b40-da25-3ab6f316b25f"
##################### Here's what we changed ##################################
classifier = tf.estimator.DNNClassifier( #
feature_columns=[tf.feature_column.indicator_column(terms_feature_column)], #
hidden_units=[20,20], #
optimizer=my_optimizer, #
) #
###############################################################################
try:
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
except ValueError as err:
print(err)
# + [markdown] colab_type="text" id="cZz68luxLS8j"
# ## Task 3: Use an Embedding with a DNN Model
#
# In this task, we'll implement our DNN model using an embedding column. An embedding column takes sparse data as input and returns a lower-dimensional dense vector as output.
# + [markdown] colab_type="text" id="AliRzhvJLS8k"
# **NOTE:** *An embedding_column is usually the computationally most efficient option to use for training a model on sparse data. In an [optional section](#scrollTo=XDMlGgRfKSVz) at the end of this exercise, we'll discuss in more depth the implementational differences between using an `embedding_column` and an `indicator_column`, and the tradeoffs of selecting one over the other.*
# + [markdown] colab_type="text" id="F-as3PtALS8l"
# In the following code, do the following:
#
# * Define the feature columns for the model using an `embedding_column` that projects the data into 2 dimensions (see the [TF docs](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column) for more details on the function signature for `embedding_column`).
# * Define a `DNNClassifier` with the following specifications:
# * Two hidden layers of 20 units each
# * Adagrad optimization with a learning rate of 0.1
# * A `gradient_clip_norm` of 5.0
# + [markdown] colab_type="text" id="UlPZ-Q9bLS8m"
# **NOTE:** *In practice, we might project to dimensions higher than 2, like 50 or 100. But for now, 2 dimensions is easy to visualize.*
# + [markdown] colab_type="text" id="mNCLhxsXyOIS"
# ### Hint
# + colab={} colab_type="code" id="L67xYD7hLS8m"
# Here's a example code snippet you might use to define the feature columns:
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
# + [markdown] colab_type="text" id="iv1UBsJxyV37"
# ### Complete the Code Below
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="5PG_yhNGLS8u" outputId="b1ddb549-1282-4df3-e3dc-edf4c59413f7"
########################## YOUR CODE HERE ######################################
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column,dimension=2)
# no. of dimensions kept at 2 for 2D visualisation
# increasing dimensions to 20 leads to better accuracy of >0.85
feature_columns = [ terms_embedding_column ]
classifier = tf.estimator.DNNClassifier( #
feature_columns=feature_columns,
hidden_units=[20,20], #
optimizer=my_optimizer, #
)
################################################################################
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] colab_type="text" id="eQS5KQzBybTY"
# ### Solution
#
# Click below for a solution.
# + colab={} colab_type="code" id="R5xOdYeQydi5"
########################## SOLUTION CODE ########################################
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
my_optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
classifier = tf.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[20,20],
optimizer=my_optimizer
)
#################################################################################
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] colab_type="text" id="aiHnnVtzLS8w"
# ## Task 4: Convince yourself there's actually an embedding in there
#
# The above model used an `embedding_column`, and it seemed to work, but this doesn't tell us much about what's going on internally. How can we check that the model is actually using an embedding inside?
#
# To start, let's look at the tensors in the model:
# + colab={"base_uri": "https://localhost:8080/", "height": 300} colab_type="code" id="h1jNgLdQLS8w" outputId="75169d02-5739-414f-f9a6-f02b69aff2e6"
classifier.get_variable_names()
# + [markdown] colab_type="text" id="Sl4-VctMLS8z"
# Okay, we can see that there is an embedding layer in there: `'dnn/input_from_feature_columns/input_layer/terms_embedding/...'`. (What's interesting here, by the way, is that this layer is trainable along with the rest of the model just as any hidden layer is.)
#
# Is the embedding layer the correct shape? Run the following code to find out.
# + [markdown] colab_type="text" id="JNFxyQUiLS80"
# **NOTE:** *Remember, in our case, the embedding is a matrix that allows us to project a 50-dimensional vector down to 2 dimensions.*
# + colab={"base_uri": "https://localhost:8080/", "height": 36} colab_type="code" id="1xMbpcEjLS80" outputId="53739219-a255-42a0-f3eb-a1722f18b665"
classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights').shape
# + [markdown] colab_type="text" id="MnLCIogjLS82"
# Spend some time manually checking the various layers and shapes to make sure everything is connected the way you would expect it would be.
# + [markdown] colab_type="text" id="rkKAaRWDLS83"
# ## Task 5: Examine the Embedding
#
# Let's now take a look at the actual embedding space, and see where the terms end up in it. Do the following:
# 1. Run the following code to see the embedding we trained in **Task 3**. Do things end up where you'd expect?
#
# 2. Re-train the model by rerunning the code in **Task 3**, and then run the embedding visualization below again. What stays the same? What changes?
#
# 3. Finally, re-train the model again using only 10 steps (which will yield a terrible model). Run the embedding visualization below again. What do you see now, and why?
# + colab={"base_uri": "https://localhost:8080/", "height": 863} colab_type="code" id="s4NNu7KqLS84" outputId="f8315e26-a593-4b18-8f60-af31d72511ee"
import numpy as np
import matplotlib.pyplot as plt
embedding_matrix = classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights')
for term_index in range(len(informative_terms)):
# Create a one-hot encoding for our term. It has 0s everywhere, except for
# a single 1 in the coordinate that corresponds to that term.
term_vector = np.zeros(len(informative_terms))
term_vector[term_index] = 1
# We'll now project that one-hot vector into the embedding space.
embedding_xy = np.matmul(term_vector, embedding_matrix)
plt.text(embedding_xy[0],
embedding_xy[1],
informative_terms[term_index])
# Do a little setup to make sure the plot displays nicely.
plt.rcParams["figure.figsize"] = (15, 15)
plt.xlim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.ylim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="Vu5kN1foVXsf" outputId="e80993ce-d0c0-4181-a17d-44922daf070d"
########################## YOUR CODE HERE ######################################
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
classifier = tf.estimator.DNNClassifier( #
feature_columns=feature_columns,
hidden_units=[20,20], #
optimizer=my_optimizer, #
)
################################################################################
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=10) #only 10 steps
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + colab={"base_uri": "https://localhost:8080/", "height": 863} colab_type="code" id="mpSETCfkVfep" outputId="ffbb82fb-c2f6-4959-a1d3-b17bed9265bd"
import numpy as np
import matplotlib.pyplot as plt
embedding_matrix = classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights')
for term_index in range(len(informative_terms)):
# Create a one-hot encoding for our term. It has 0s everywhere, except for
# a single 1 in the coordinate that corresponds to that term.
term_vector = np.zeros(len(informative_terms))
term_vector[term_index] = 1
# We'll now project that one-hot vector into the embedding space.
embedding_xy = np.matmul(term_vector, embedding_matrix)
plt.text(embedding_xy[0],
embedding_xy[1],
informative_terms[term_index])
# Do a little setup to make sure the plot displays nicely.
plt.rcParams["figure.figsize"] = (15, 15)
plt.xlim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.ylim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.show()
# + [markdown] colab_type="text" id="pUb3L7pqLS86"
# ## Task 6: Try to improve the model's performance
#
# See if you can refine the model to improve performance. A couple things you may want to try:
#
# * **Changing hyperparameters**, or **using a different optimizer** like Adam (you may only gain one or two accuracy percentage points following these strategies).
# * **Adding additional terms to `informative_terms`.** There's a full vocabulary file with all 30,716 terms for this data set that you can use at: https://dl.google.com/mlcc/mledu-datasets/sparse-data-embedding/terms.txt You can pick out additional terms from this vocabulary file, or use the whole thing via the `categorical_column_with_vocabulary_file` feature column.
# + colab={} colab_type="code" id="6-b3BqXvLS86"
# Download the vocabulary file.
terms_url = 'https://dl.google.com/mlcc/mledu-datasets/sparse-data-embedding/terms.txt'
terms_path = tf.keras.utils.get_file(terms_url.split('/')[-1], terms_url)
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="0jbJlwW5LS8-" outputId="1c100bd5-9944-43ca-b285-b9d097a37604"
# Create a feature column from "terms", using a full vocabulary file.
informative_terms = None
with io.open(terms_path, 'r', encoding='utf8') as f:
# Convert it to a set first to remove duplicates.
informative_terms = list(set(f.read().split()))
terms_feature_column = tf.feature_column.categorical_column_with_vocabulary_list(key="terms",
vocabulary_list=informative_terms)
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
my_optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
classifier = tf.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[10,10],
optimizer=my_optimizer
)
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + colab={"base_uri": "https://localhost:8080/", "height": 863} colab_type="code" id="wJE8n3XvaBHq" outputId="db523183-f17e-448a-da18-4b9d44567911"
import numpy as np
import matplotlib.pyplot as plt
embedding_matrix = classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights')
for term_index in range(len(informative_terms)):
# Create a one-hot encoding for our term. It has 0s everywhere, except for
# a single 1 in the coordinate that corresponds to that term.
term_vector = np.zeros(len(informative_terms))
term_vector[term_index] = 1
# We'll now project that one-hot vector into the embedding space.
embedding_xy = np.matmul(term_vector, embedding_matrix)
plt.text(embedding_xy[0],
embedding_xy[1],
informative_terms[term_index])
# Do a little setup to make sure the plot displays nicely.
plt.rcParams["figure.figsize"] = (15, 15)
plt.xlim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.ylim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 36} colab_type="code" id="TGo_qCztcHDw" outputId="26853961-935f-4c71-9bcf-6bf3d9445a46"
classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights').shape
# + [markdown] colab_type="text" id="ew3kwGM-LS9B"
# ## A Final Word
#
# We may have gotten a DNN solution with an embedding that was better than our original linear model, but the linear model was also pretty good and was quite a bit faster to train. Linear models train more quickly because they do not have nearly as many parameters to update or layers to backprop through.
#
# In some applications, the speed of linear models may be a game changer, or linear models may be perfectly sufficient from a quality standpoint. In other areas, the additional model complexity and capacity provided by DNNs might be more important. When defining your model architecture, remember to explore your problem sufficiently so that you know which space you're in.
# + [markdown] colab_type="text" id="9MquXy9zLS9B"
# ### *Optional Discussion:* Trade-offs between `embedding_column` and `indicator_column`
#
# Conceptually when training a `LinearClassifier` or a `DNNClassifier`, there is an adapter needed to use a sparse column. TF provides two options: `embedding_column` or `indicator_column`.
#
# When training a LinearClassifier (as in **Task 1**), an `embedding_column` is used under the hood. As seen in **Task 2**, when training a `DNNClassifier`, you must explicitly choose either `embedding_column` or `indicator_column`. This section discusses the distinction between the two, and the trade-offs of using one over the other, by looking at a simple example.
# + [markdown] colab_type="text" id="M_3XuZ_LLS9C"
# Suppose we have sparse data containing the values `"great"`, `"beautiful"`, `"excellent"`. Since the vocabulary size we're using here is $V = 50$, each unit (neuron) in the first layer will have 50 weights. We denote the number of terms in a sparse input using $s$. So for this example sparse data, $s = 3$. For an input layer with $V$ possible values, a hidden layer with $d$ units needs to do a vector-matrix multiply: $(1 \times V) * (V \times d)$. This has $O(V * d)$ computational cost. Note that this cost is proportional to the number of weights in that hidden layer and independent of $s$.
#
# If the inputs are one-hot encoded (a Boolean vector of length $V$ with a 1 for the terms present and a 0 for the rest) using an [`indicator_column`](https://www.tensorflow.org/api_docs/python/tf/feature_column/indicator_column), this means multiplying and adding a lot of zeros.
# + [markdown] colab_type="text" id="I7mR4Wa2LS9C"
# When we achieve the exact same results by using an [`embedding_column`](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column) of size $d$, we look up and add up just the embeddings corresponding to the three features present in our example input of "`great`", "`beautiful`", "`excellent`": $(1 \times d) + (1 \times d) + (1 \times d)$. Since the weights for the features that are absent are multiplied by zero in the vector-matrix multiply, they do not contribute to the result. Weights for the features that are present are multiplied by 1 in the vector-matrix multiply. Thus, adding the weights obtained via the embedding lookup will lead to the same result as in the vector-matrix-multiply.
#
# When using an embedding, computing the embedding lookup is an $O(s * d)$ computation, which is computationally much more efficient than the $O(V * d)$ cost for the `indicator_column` in sparse data for which $s$ is much smaller than $V$. (Remember, these embeddings are being learned. In any given training iteration it is the current weights that are being looked up.)
# + [markdown] colab_type="text" id="etZ9qf0kLS9D"
# As we saw in **Task 3**, by using an `embedding_column` in training the `DNNClassifier`, our model learns a low-dimensional representation for the features, where the dot product defines a similarity metric tailored to the desired task. In this example, terms that are used similarly in the context of movie reviews (e.g., `"great"` and `"excellent"`) will be closer to each other in the embedding space (i.e., have a large dot product), and terms that are dissimilar (e.g., `"great"` and `"bad"`) will be farther away from each other in the embedding space (i.e., have a small dot product).
# + colab={} colab_type="code" id="j47yabtfZ54X"
| intro_to_sparse_data_and_embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from PIL import Image, ImageDraw
import pandas as pd
import numpy as np
import zipfile
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import torch
import torchvision
from torch import nn
import torchvision.transforms as transforms
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
from sklearn.metrics import accuracy_score
# -
def read_zip(zipfile_str):
"""this function will taked a zipfile and return a list of image"""
imgzip = zipfile.ZipFile(zipfile_str)
inflist = imgzip.infolist()
image_list =[]
for f in inflist:
ifile = imgzip.open(f)
img = Image.open(ifile)
image_list.append(img)
return image_list
def diget_list(images):
"""
this function take an image and corp it to 9 smal images,
it return a list of images
"""
image = images.save('images.png')
image = images.resize((int(1500),int(1500)))
x = 0
y = 0
h = 500
w = 500
image_list = []
for i in range(9):
diget_image=image.crop((x,y,x+w,y+h))
if x+w >= image.width:
x = 0
y = y + h
else:
x=x+500
diget_image = diget_image.resize((int(28),int(28)))
image_list.append(diget_image)
return image_list
# +
images = read_zip('IMG_diget.zip') # get a listof the images
final_image_list =[]
for image in images: # a for loop to get the 90 image in one list
image_list = diget_list(image)
final_image_list = final_image_list + image_list
# creat a dataframe of the list
image_dataFrame = pd.DataFrame(data =final_image_list, columns =['images'])
# -
# creat a list of the label
g = 0
n = 0
label_list =[]
for i in range(100):
if g < 9:
label_list.append(n)
g = g+1
else:
n = n+1
g = 0
# add the label column
image_dataFrame['Label'] = label_list
# shuffle your dataframe
image_dataFrame = image_dataFrame.sample(frac=1, random_state=42).reset_index(drop=True)
for i in range(9):
# define subplot
plt.subplot(330 + 1 + i)
# plot raw pixel data
plt.imshow(image_dataFrame['images'][i], cmap=plt.get_cmap('gray'))
# show the figure
plt.show()
# #### transform image to Tensors of normalized range.
# +
X=[]
y=[]
for i in image_dataFrame['images']:
i = i.convert('L')
X.append(np.array(i))
for i in image_dataFrame['Label']:
y.append(i)
# -
type(X)
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size= 0.2)
# convert image aray to tourch.tensor
X_train = torch.Tensor(X_train)
X_test = torch.Tensor(X_test)
Y_train = torch.Tensor(Y_train)
Y_test = torch.Tensor(Y_test)
# reshaping the data to fit the model
X_train = X_train.reshape(72, 1, 28, 28)
X_test = X_test.reshape(18, 1, 28, 28)
print(f"X_train type is {type(X_train)}, y_train type is {type(Y_train)}")
print(f"X_test type is {type(X_test)}, y_test type is {type(Y_test)}")
# ### Define and intialize the neural network
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(1, 32, 3, 1),
)
self.layer2 = nn.Sequential(
nn.Conv2d(32, 64, 3, 1),
)
self.dropout1 = nn.Dropout2d(0.25)
self.dropout2 = nn.Dropout2d(0.5)
# First fully connected layer
self.fc1 = nn.Linear(9216, 128)
# Second fully connected layer
self.fc2 = nn.Linear(128, 10)
def forward(self, x):
# Pass data through conv1
x = self.layer1(x)
# Use the ReLU activation function
x = F.relu(x)
# Pass data through conv2
x = self.layer2(x)
x = F.relu(x)
# Run max pooling over x
x = F.max_pool2d(x, 2)
# Pass data through dropout1
x = self.dropout1(x)
x = self.dropout2(x)
# Flatten x with start_dim=1
x = torch.flatten(x, 1)
# Pass data through fc1
x = self.fc1(x)
x = F.relu(x)
x = self.dropout2(x)
x = self.fc2(x)
# Apply softmax to x
output = F.log_softmax(x, dim=1)
return output
model = Net()
print(model)
# #### Define a Loss function and optimizer
# +
# defining the loss function
criterion = nn.CrossEntropyLoss()
# checking if GPU is available
if torch.cuda.is_available():
model = model.cuda()
criterion = criterion.cuda()
# defining the optimizer
optimizer = optim.SGD(model.parameters(), lr=0.001, momentum=0.9)
# -
# #### Train the network
def train(epoch):
model.train()
tr_loss = 0
# getting the training set
x_train, y_train = Variable(X_train), Variable(Y_train)
# getting the validation set
x_val, y_val = Variable(X_test), Variable(Y_test)
# converting the data into GPU format
if torch.cuda.is_available():
x_train = x_train.cuda()
y_train = y_train.cuda()
x_val = x_val.cuda()
y_val = y_val.cuda()
# clearing the Gradients of the model parameters
optimizer.zero_grad()
# prediction for training and validation set
output_train = model(x_train)
output_val = model(x_val)
# computing the training and validation loss
y_train = y_train.type(torch.LongTensor).cuda()
y_val = y_val.type(torch.LongTensor).cuda()
loss_train = criterion(output_train, y_train)
loss_val = criterion(output_val, y_val)
train_losses.append(loss_train)
val_losses.append(loss_val)
# computing the updated weights of all the model parameters
loss_train.backward()
optimizer.step()
tr_loss = loss_train.item()
if epoch%2 == 0:
# printing the validation loss
print('Epoch : ',epoch+1, '\t', 'loss :', loss_val)
# defining the number of epochs
n_epochs = 25
# empty list to store training losses
train_losses = []
# empty list to store validation losses
val_losses = []
# training the model
for epoch in range(n_epochs):
train(epoch)
train_losses_values = []
val_losses_values = []
for i in train_losses:
train_losses_values.append(i.item())
for i in val_losses:
val_losses_values.append(i.item())
# plotting the training and validation loss
plt.plot(train_losses_values, label='Training loss')
plt.plot(val_losses_values, label='Validation loss')
plt.legend()
plt.show()
# #### check the accuracy of the model
# +
# prediction for training set
with torch.no_grad():
output = model(X_train.cuda())
softmax = torch.exp(output).cpu()
prob = list(softmax.numpy())
predictions = np.argmax(prob, axis=1)
# accuracy on training set
accuracy_score(Y_train, predictions)
# +
# prediction for validation set
with torch.no_grad():
output = model(X_test.cuda())
softmax = torch.exp(output).cpu()
prob = list(softmax.numpy())
predictions = np.argmax(prob, axis=1)
# accuracy on validation set
accuracy_score(Y_test, predictions)
# -
| mini project/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# %matplotlib inline
import numpy as np
from matplotlib import pyplot as plt
# ## Build a baseline model of stellar mass
# +
from halotools.sim_manager import CachedHaloCatalog
halocat = CachedHaloCatalog()
from halotools.empirical_models import Moster13SmHm
model = Moster13SmHm()
halocat.halo_table['stellar_mass'] = model.mc_stellar_mass(
prim_haloprop=halocat.halo_table['halo_mpeak'], redshift=0)
# -
# ## Define a simple model for $M_{\ast}-$dependence of ${\rm B/T}$ power law index
def powerlaw_index(log_mstar):
abscissa = [9, 10, 11.5]
ordinates = [3, 2, 1]
return np.interp(log_mstar, abscissa, ordinates)
# ## Calculate the spin-percentile
# +
from halotools.utils import sliding_conditional_percentile
x = halocat.halo_table['stellar_mass']
y = halocat.halo_table['halo_spin']
nwin = 201
halocat.halo_table['spin_percentile'] = sliding_conditional_percentile(x, y, nwin)
# -
# ## Use CAM to generate a Monte Carlo realization of ${\rm B/T}$
a = powerlaw_index(np.log10(halocat.halo_table['stellar_mass']))
u = halocat.halo_table['spin_percentile']
halocat.halo_table['bulge_to_total_ratio'] = 1 - powerlaw.isf(1 - u, a)
# ### Plot the results
# +
fig, ax = plt.subplots(1, 1)
mask1 = halocat.halo_table['stellar_mass'] < 10**9.5
mask2 = halocat.halo_table['stellar_mass'] > 10**10.5
__=ax.hist(halocat.halo_table['bulge_to_total_ratio'][mask1],
bins=100, alpha=0.8, normed=True, color='blue',
label=r'$\log M_{\ast} < 9.5$')
__=ax.hist(halocat.halo_table['bulge_to_total_ratio'][mask2],
bins=100, alpha=0.8, normed=True, color='red',
label=r'$\log M_{\ast} > 10.5$')
legend = ax.legend()
xlabel = ax.set_xlabel(r'${\rm B/T}$')
ylabel = ax.set_ylabel(r'${\rm PDF}$')
title = ax.set_title(r'${\rm Bulge}$-${\rm to}$-${\rm Total\ M_{\ast}\ Ratio}$')
figname = 'cam_example_bt_distributions.png'
fig.savefig(figname, bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight')
# +
xmin, xmax = 9, 11.25
fig, ax = plt.subplots(1, 1)
xscale = ax.set_xscale('log')
from scipy.stats import binned_statistic
log_mass_bins = np.linspace(xmin, xmax, 25)
mass_mids = 10**(0.5*(log_mass_bins[:-1] + log_mass_bins[1:]))
median_bt, __, __ = binned_statistic(
halocat.halo_table['stellar_mass'], halocat.halo_table['bulge_to_total_ratio'],
bins=10**log_mass_bins, statistic='median')
std_bt, __, __ = binned_statistic(
halocat.halo_table['stellar_mass'], halocat.halo_table['bulge_to_total_ratio'],
bins=10**log_mass_bins, statistic=np.std)
low_spin_mask = halocat.halo_table['spin_percentile'] < 0.5
median_bt_low_spin, __, __ = binned_statistic(
halocat.halo_table['stellar_mass'][low_spin_mask],
halocat.halo_table['bulge_to_total_ratio'][low_spin_mask],
bins=10**log_mass_bins, statistic='median')
std_bt_low_spin, __, __ = binned_statistic(
halocat.halo_table['stellar_mass'][low_spin_mask],
halocat.halo_table['bulge_to_total_ratio'][low_spin_mask],
bins=10**log_mass_bins, statistic=np.std)
high_spin_mask = halocat.halo_table['spin_percentile'] > 0.5
median_bt_high_spin, __, __ = binned_statistic(
halocat.halo_table['stellar_mass'][high_spin_mask],
halocat.halo_table['bulge_to_total_ratio'][high_spin_mask],
bins=10**log_mass_bins, statistic='median')
std_bt_high_spin, __, __ = binned_statistic(
halocat.halo_table['stellar_mass'][high_spin_mask],
halocat.halo_table['bulge_to_total_ratio'][high_spin_mask],
bins=10**log_mass_bins, statistic=np.std)
y1 = median_bt_low_spin - std_bt_low_spin
y2 = median_bt_low_spin + std_bt_low_spin
__=ax.fill_between(mass_mids, y1, y2, alpha=0.8, color='red',
label=r'${\rm low\ spin\ halos}$')
y1 = median_bt_high_spin - std_bt_high_spin
y2 = median_bt_high_spin + std_bt_high_spin
__=ax.fill_between(mass_mids, y1, y2, alpha=0.8, color='blue',
label=r'${\rm high\ spin\ halos}$')
ylim = ax.set_ylim(0, 1)
legend = ax.legend(loc='upper left')
xlabel = ax.set_xlabel(r'${\rm M_{\ast}/M_{\odot}}$')
ylabel = ax.set_ylabel(r'$\langle{\rm B/T}\rangle$')
figname = 'cam_example_bulge_disk_ratio.png'
fig.savefig(figname, bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight')
# -
| docs/notebooks/cam_modeling/cam_disk_bulge_ratios_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import psycopg2
import pandas as pd
import numpy as np
import os
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="ticks", color_codes=True)
import matplotlib
import warnings
import yellowbrick as yb
matplotlib.style.use('ggplot')
import numpy as np
import seaborn as sns
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.metrics import f1_score
from sklearn.pipeline import Pipeline
from sklearn.svm import LinearSVC, NuSVC, SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.linear_model import LogisticRegressionCV, LogisticRegression, SGDClassifier
from sklearn.ensemble import BaggingClassifier, ExtraTreesClassifier, AdaBoostClassifier, RandomForestClassifier, GradientBoostingClassifier
import matplotlib.pyplot as plt
df = pd.read_csv("LasVegas.csv", sep=';')
df
df.info()
#identify any null values
null_columns=df.columns[df.isnull().any()]
df[null_columns].isnull().sum()
df['Score'].value_counts()
df['Gym'].value_counts()
df['Spa'].value_counts()
sns.heatmap(df.corr())
sns.set(style="whitegrid")
sns.violinplot(x="Gym", y="Score", hue="Casino", data=df, figsize = (15,15))
corr_matrix = df.corr()
corr_matrix['Score'].sort_values(ascending=False)
sns.catplot(x='Score', y='Spa', kind="box", data=df);
sns.catplot(x='Score', y='Gym', kind="box", data=df);
sns.catplot(x='Score', y='Pool', kind="box", data=df);
LABEL_MAP = {
"YES": 1,
"NO": 0
}
# Convert categorical labels into incremental value
df['Pool'] = df['Pool'].map(LABEL_MAP).to_frame()
df['Gym'] = df['Gym'].map(LABEL_MAP).to_frame()
df['Tennis court'] = df['Tennis court'].map(LABEL_MAP).to_frame()
df['Spa'] = df['Spa'].map(LABEL_MAP).to_frame()
df['Free internet'] = df['Free internet'].map(LABEL_MAP).to_frame()
df['Casino'] = df['Casino'].map(LABEL_MAP).to_frame()
# +
df = df.drop(columns=['User continent', 'User country', 'Period of stay', 'Traveler type', 'Hotel name', 'Hotel stars',
'Review month', 'Review weekday'])
# +
from sklearn.ensemble import RandomForestClassifier
from yellowbrick.features import FeatureImportances
model = RandomForestClassifier(n_estimators=10)
viz = FeatureImportances(model, size=(1080, 1720))
viz.fit(X, y)
viz.show()
# -
corrmat = df.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(20,20))
#plot heat map
g=sns.heatmap(df[top_corr_features].corr(),annot=True,cmap="RdYlGn")
y = df['Score']
X = df.loc[:, df.columns.difference(['Score'])]
y
from imblearn.over_sampling import SMOTE
sm = SMOTE(random_state = 33)
X_sm, y_sm = sm.fit_sample(X, y.ravel())
pd.Series(y_sm).value_counts().plot.bar()
X_sm, y_sm = sm.fit_sample(X_sm, y_sm.ravel())
pd.Series(y_sm).value_counts().plot.bar()
X_sm, y_sm = sm.fit_sample(X_sm, y_sm.ravel())
pd.Series(y_sm).value_counts().plot.bar()
X_sm, y_sm = sm.fit_sample(X_sm, y_sm.ravel())
pd.Series(y_sm).value_counts().plot.bar()
# Create the train and test data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X_sm, y_sm, test_size=0.2)
def score_model(X, y, estimator, **kwargs):
y = LabelEncoder().fit_transform(y)
model = Pipeline([
('one_hot_encoder', OneHotEncoder(handle_unknown='ignore')),
('estimator', estimator)
])
model.fit(X_train, y_train, **kwargs)
expected = y_test
predicted = model.predict(X_test)
# Compute and return F1 (harmonic mean of precision and recall)
print("{}: {}".format(estimator.__class__.__name__, f1_score(expected, predicted, average='micro')))
# +
models = [
SVC(),
#NuSVC(),
LinearSVC(),
SGDClassifier(), KNeighborsClassifier(),
LogisticRegression(),
#aggingClassifier(),
ExtraTreesClassifier(n_estimators=100),
RandomForestClassifier(n_estimators=100),
GradientBoostingClassifier(n_estimators=100,learning_rate=.3)
#learning_rate=.5,max_depth=4, min_samples_leaf=75
]
for model in models:
score_model(X_train, y_train, model)
# -
| ADS-Fall2019A/Las Vegas Hotel TripAdvisor Reviews/Restaurant Suggestion (2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
cancer_data_31 = pd.read_csv('../final_code/data/31DayData.csv')
cancer_data_62 = pd.read_csv('../final_code/data/62DayData.csv')
diagnostics_data_HB = pd.read_csv('../final_code/data/diagnostics_by_board_september_2021.csv', sep=',', engine='python').sort_values('MonthEnding').reset_index(drop=True)
diagnostics_data_scot = pd.read_csv('../final_code/data/diagnostics_scotland_september_2021.csv', engine='python').sort_values('MonthEnding').reset_index(drop=True).rename(columns={'Country':'HBT'})
cancellation_data = pd.read_csv('../final_code/data/cancellations_by_board_november_2021.csv', engine='python').sort_values('Month').reset_index(drop=True)
AE_data = pd.read_csv('../final_code/data/monthly_ae_waitingtimes_202111.csv', engine='python').sort_values('Month').reset_index(drop=True)
# +
def plot_cancer_data(data, regions, cancers, years, rows):
y=0
year_index = [min(data.index[data['Quarter']==years[0]]),max(data.index[data['Quarter']==years[1]])]
yearData = data.iloc[year_index[0]:year_index[1]+1,:]
for HB in regions:
for CT in cancers:
typeData = yearData[yearData['CancerType'] == CT]
regionData = typeData[typeData['HB'] == HB]
aggregation_functions = {'Quarter':'first', 'HB': 'first', 'CancerType': 'first', rows: 'sum'}
finalData = regionData.groupby(regionData['Quarter'], as_index=False).aggregate(aggregation_functions)
y += finalData[rows]
x = finalData['Quarter']
plt.plot(x,y, label = rows)
def plot_diagnostics_data(data, regions, types, description, years, rows):
y=0
year_index = [min(data.index[data['MonthEnding']==years[0]]),max(data.index[data['MonthEnding']==years[1]])]
yearData = data.iloc[year_index[0]:year_index[1]+1,:]
for HBT in regions:
for T in types:
for D in description:
typeData = yearData[yearData['DiagnosticTestType'] == T]
describData = typeData[typeData['DiagnosticTestDescription'] == D]
regionData = describData[describData['HBT'] == HBT]
aggregation_functions = {'MonthEnding':'first', 'HBT': 'first', 'DiagnosticTestType': 'first', rows: 'sum'}
finalData = regionData.groupby(regionData['MonthEnding'], as_index=False).aggregate(aggregation_functions)
y += finalData[rows]
finalData.to_csv('data.csv')
x = list(map(str,finalData['MonthEnding']))
plt.plot(x,y, label = rows)
def plot_cancellation_data(data, regions, years, rows):
y=0
year_index = [min(data.index[data['Month']==years[0]]),max(data.index[data['Month']==years[1]])]
yearData = data.iloc[year_index[0]:year_index[1]+1,:]
for HBT in regions:
regionData = yearData[yearData['HBT'] == HBT]
aggregation_functions = {'Month':'first', 'HBT': 'first', rows: 'sum'}
finalData = regionData.groupby(regionData['Month'], as_index=False).aggregate(aggregation_functions)
y += finalData[rows]
finalData.to_csv('data.csv')
x = list(map(str,finalData['Month']))
plt.plot(x,y, label = rows)
# +
all_regions = ['S08000015', 'S08000016', 'S08000017', 'S08000019', 'S08000020', 'S08000022', 'S08000024', 'S08000025', 'S08000026', 'S08000028', 'S08000029', 'S08000030', 'S08000031', 'S08000032']
fig, axs = plt.subplots(1, 1, figsize=(20,5))
plot_cancer_data(cancer_data_31, all_regions, ['All Cancer Types'], ['2012Q1','2021Q3'], 'NumberOfEligibleReferrals31DayStandard')
plot_cancer_data(cancer_data_31, all_regions, ['All Cancer Types'], ['2012Q1','2021Q3'], 'NumberOfEligibleReferralsTreatedWithin31Days')
#axs.plot([32.5,32.5], [0,1600], color='r', linewidth=3)
every_nth = 4
for n, label in enumerate(axs.xaxis.get_ticklabels()):
if n % every_nth != 0:
label.set_visible(False)
plt.legend()
plt.show()
fig, axs = plt.subplots(1, 1, figsize=(20,5))
plot_cancer_data(cancer_data_62, all_regions, ['All Cancer Types'], ['2015Q1','2021Q3'], 'NumberOfEligibleReferrals62DayStandard')
plot_cancer_data(cancer_data_62, all_regions, ['All Cancer Types'], ['2015Q1','2021Q3'], 'NumberOfEligibleReferralsTreatedWithin62Days')
#axs.plot([32.5,32.5], [0,1600], color='r', linewidth=3)
every_nth = 4
for n, label in enumerate(axs.xaxis.get_ticklabels()):
if n % every_nth != 0:
label.set_visible(False)
plt.legend()
plt.show()
# +
report_endoscopy = [9476, 9108, 8225, 7875, 7347, 6954, 6129, 6398, 6290, 6990, 7887, 7009, 9340, 19648, 22757, 23088, 23358, 23787, 23383, 22789, 21098, 21460, 22662, 22075, 21878, 22445, 22248, 21899, 22791, 22919, 22947]
report_radiology = [4854, 7338, 8167, 8337, 10985, 10068, 9380, 10561, 8811, 10916, 10640, 6491, 11710, 45663, 43969, 40581, 33963, 27034, 24585, 25546, 21327, 23056, 25687, 21338, 18941, 18956, 17799, 21178, 27400, 29249, 30076]
finalData = pd.read_csv('data.csv')
fig, axs = plt.subplots(1, 1, figsize=(20,10))
plt.plot(list(map(str,finalData['MonthEnding'])),report_endoscopy, 'tab:gray',linewidth=4, label='report data endoscopy')
plt.plot(list(map(str,finalData['MonthEnding'])),report_radiology, 'tab:gray',linewidth=4, label='report data radiology')
plot_diagnostics_data(diagnostics_data_HB, all_regions, ['Imaging'], ['All Imaging'], [20190331,20210930], 'NumberWaitingOverSixWeeks')
#plot_diagnostics_data(diagnostics_data_HB, all_regions, ['Imaging'], ['All Imaging'], [20190331,20210930], 'NumberWaitingOverFourWeeks')
plot_diagnostics_data(diagnostics_data_scot, ['S92000003'], ['Endoscopy'], ['All Endoscopy'], [20190331,20210930], 'NumberWaitingOverSixWeeks')
plot_diagnostics_data(diagnostics_data_scot, ['S92000003'], ['Imaging'], ['All Imaging'], [20190331,20210930], 'NumberWaitingOverSixWeeks')
#axs.plot([32.5,32.5], [0,1600], color='r', linewidth=3)
every_nth = 2
for n, label in enumerate(axs.xaxis.get_ticklabels()):
if n % every_nth != 0:
label.set_visible(False)
plt.legend()
plt.show()
# +
fig, axs = plt.subplots(1, 1, figsize=(20,10))
plot_cancellation_data(cancellation_data, all_regions, [201805,202111], 'TotalCancelled')
plot_cancellation_data(cancellation_data, all_regions, [201805,202111], 'CancelledByPatientReason')
plot_cancellation_data(cancellation_data, all_regions, [201805,202111], 'ClinicalReason')
plot_cancellation_data(cancellation_data, all_regions, [201805,202111], 'NonClinicalCapacityReason')
#axs.plot([32.5,32.5], [0,1600], color='r', linewidth=3)
every_nth = 2
for n, label in enumerate(axs.xaxis.get_ticklabels()):
if n % every_nth != 0:
label.set_visible(False)
plt.legend()
plt.show()
# +
fig, axs = plt.subplots(1, 1, figsize=(20,10))
plot_cancellation_data(AE_data, all_regions, [201805,202111], 'AttendanceGreater8hrs')
plot_cancellation_data(AE_data, all_regions, [201805,202111], 'AttendanceGreater12hrs')
plot_cancellation_data(AE_data, all_regions, [201805,202111], 'NumberOfAttendancesAggregate')
plot_cancellation_data(AE_data, all_regions, [201805,202111], 'NumberMeetingTargetAggregate')
#axs.plot([32.5,32.5], [0,1600], color='r', linewidth=3)
every_nth = 2
for n, label in enumerate(axs.xaxis.get_ticklabels()):
if n % every_nth != 0:
label.set_visible(False)
plt.legend()
plt.show()
# -
| datasets_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
FN = '161103-run-plot'
# Plot validation accuracy vs. noise level for different experiments
# the results of the experiments are accumlated in:
FN1 = '160916-run-plot'
# each experiment is run by `run.bash` which runs the same experiment with different training size (`down_sample=.2,.5,1`) and in each time it runs (`run_all.bash`) with 5 different seeds.
# Each such run of `jacob-reed.py` includes a loop over different `noise_level`
# + magic_args="-s \"{FN1}\"" language="bash"
# echo $1
# # ./run.bash --FN=data/$1
# # ./run.bash --FN=data/$1 --model=simple --beta=0
# # ./run.bash --FN=data/$1 --model=complex --beta=0 --pretrain=2
# # ./run.bash --FN=data/$1 --model=reed_hard --beta=0.8
# # ./run.bash --FN=data/$1 --model=reed_soft --beta=0.95
# -
import fasteners
import time
import os
from collections import defaultdict
import cPickle as pickle
import warnings ; warnings.filterwarnings("ignore")
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['axes.facecolor'] = 'white'
with fasteners.InterProcessLock('/tmp/%s.lock_file'%FN1):
with open('data/%s.results.pkl'%FN1,'rb') as fp:
results = pickle.load(fp)
experiments = set([k[0] for k in results.keys()]) # find all unique experiments from experiment,noise keys
experiments = sorted(experiments)
# The different models for which we have results. Composed from the following:
# * B baseline, S simple, C complex, R reed soft, r reed hard
# * M for MLP, C for CNN
# * If the model is not baseline then beta is less than 1 and its value **after the decimal dot** appears (e.g. beta=0.85 will show as 85.) Note that if beta happens to be zero then nothing will appear
# * Pre-training using labels generated by: p=baseline, q=simple model. r=use simple weights as a start point for the bias of the channel matrix instead of rebuilding it from the labels predicted by the simple model
# * If model is complex then the value (usually 0) used to initialize channel matrix weights (unless it was 0.1)
print "Models:",', '.join(set(x.split('-')[0] for x in experiments))
# The noise used:
print "Noise:",', '.join(set(x.split('-')[1].split('_')[0] for x in experiments))
# different seeds used
# size of training data used. Multiply by 10 to get percentage, s at the end indicate that the number of labels in training set is stratified
print "Training size:",', '.join(set(x.split('-')[1].split('_')[1] for x in experiments))
# Count how many experiments (by different seeds) we did for every model/train-size combinations
e2n = defaultdict(int)
for e in reversed(experiments):
model_noise, seed, train_size = e.split('_')
e2n['_'.join([model_noise, train_size])] += 1
# e2n
#http://people.duke.edu/~ccc14/pcfb/analysis.html
def bootstrap(data, num_samples, statistic, alpha):
"""Returns bootstrap estimate of 100.0*(1-alpha) CI for statistic."""
n = len(data)
idx = np.random.randint(0, n, (num_samples, n))
samples = data[idx]
stat = np.sort(statistic(samples, 1))
return (stat[int((alpha/2.0)*num_samples)],
stat[int((1-alpha/2.0)*num_samples)])
# convert model code to label we will use in the graph's legend
namelookup = {'CMp0':'Complex baseline-labels',
'CMq0':'Complex', # simple-labels',
'CCq0':'Complex CNN', # simple-labels',
'CMr0':'Complex simple-weights',
'BM':'Baseline','BC':'Baseline CNN',
'SMp':'Simple','SMP':'Simple soft confusion',
'SCp':'Simple CNN','SCP':'Simple CNN soft confusion',
'rM8p':'Reed hard','RM95p':'Reed soft',
'rC8p':'Reed hard','RC95p':'Reed soft',
'rMp':'Reed hard beta=0','RMp':'Reed soft beta=0'}
# the order in which the graphs will appear in the legend (and colors)
model_order = {'SMp':3, 'SCp':3, 'CMp0':0, 'CMq0':1, 'CCq0':1, 'CMr0':2,
'rM8p':4, 'rC8p':4, 'rMp':5,
'RM95p':6, 'RC95p':6, 'RMp':7,
'BM':8,'BC':9, 'SMP':10}
import numpy as np
import matplotlib.pyplot as plt
def plot(idx, models=None, perm=None, down_samples=None,xlim=(0.3,0.5),ylim=(0.4,1),title='', stat=np.mean):
"""
models - list of str
take only experiments that starts with one of the str in models.
down_samples - list of float or str
take only results with specificed down sample
title - if None then dont produce any title. If string then build a title which includes in it the given str
"""
if down_samples is not None:
if not isinstance(down_samples,list):
down_samples = [down_samples]
down_samples = [down_sample if isinstance(down_sample, basestring)
else '%g'%(down_sample*10)
for down_sample in down_samples]
plt.subplot(idx)
e2xy = defaultdict(lambda : defaultdict(list))
for e in reversed(experiments):
if down_samples is not None:
for down_sample in down_samples:
if e.endswith('_'+down_sample) or e.endswith('_'+down_sample+'s'):
break
else:
continue
if models:
for model in models:
if e.startswith(model):
break
else:
continue
else:
model = e.split('-')[0]
down_sample = e.split('_')[-1]
if perm is not None:
pperm = e.split('-')[1].split('_')[0]
if pperm != perm:
continue
X = [k[1] for k in results.keys() if k[0] == e]
Y = []
for n in X:
d = results[(e,n)][1]
for k in ['baseline_acc', 'acc']:
if k in d:
Y.append(d[k])
break
else:
raise Exception('validation accuracy not found')
X, Y = zip(*sorted(zip(X,Y),key=lambda x: x[0]))
for x,y in zip(X,Y):
e2xy[(model,down_sample)][x] += [y]
Y = np.array(Y)
keys = sorted(e2xy.keys(),key=lambda md: (model_order[md[0].split('-')[0]], (' ' if len(md[1])-int(md[1].endswith('s'))==1 else '') + md[1]))
# determine if all down_sample are stratified (True) or all not stratified (False) or mixed (Nones)
all_stratified = None
for model,down_sample in keys:
if all_stratified is None:
all_stratified = down_sample.endswith('s')
elif all_stratified != down_sample.endswith('s'):
all_stratified = None
break
for c, (model,down_sample) in enumerate(keys):
xy = e2xy[(model,down_sample)]
X = []
Y = []
Ys = []
for x in sorted(xy.keys()):
X.append(x)
Y.append(stat(xy[x]))
Ys.append(xy[x])
error_bars = []
for y,ys in zip(Y,Ys):
ym, yp = bootstrap(np.array(ys), 10000, stat, 0.05)
error_bars.append((y-ym,yp-y))
X = np.array(X)
colors = ['b','g','r','c','m','k']
linestyles=['solid', 'dashed','dashdot','dotted']
color = colors[c % len(colors)]
linestyle = linestyles[(c//len(colors))%len(linestyles)]
plt.errorbar(X + 0.001*np.random.random(len(X)),
Y, yerr=np.array(error_bars).T, ecolor=color, elinewidth=2, alpha=0.2,
fmt='none')
# label of each graph is the model name in English and
# if we have more than one downsample option then also add it to the label
if model in namelookup:
label = namelookup[model]
else:
label = namelookup[model.split('-')[0]] + ' ' + model.split('-')[1]
if down_samples is None or len(down_samples) > 1:
if all_stratified is None:
label += ' '+(down_sample[:-1]+'0% stratified' if down_sample.endswith('s') else down_sample+'0%')
else:
label += ' '+(down_sample[:-1] if down_sample.endswith('s') else down_sample)+'0%'
plt.plot(X,Y, color=color, linestyle=linestyle, label=label)
plt.legend(loc='lower left')
plt.ylabel('test accuracy', fontsize=18)
plt.xlabel('noise fraction', fontsize=18);
if title is not None:
title = 'MNIST '+title
if all_stratified:
title += ' stratified'
plt.title(title)
# The range of X and Y is exactly like in other paper
if xlim is not None:
plt.xlim(*xlim)
if ylim is not None:
plt.ylim(*ylim)
plt.figure(figsize=(12,10))
plot(111, down_samples=1.0, models=['BM','SMp','CMq0','rM8p','RM95p'], title=None, perm='7904213568')
plt.figure(figsize=(12,10))
plot(111, down_samples=.5, models=['BM','SMp','CMq0','rM8p','RM95p'], title=None, perm='7904213568')
plt.figure(figsize=(12,10))
plot(111, down_samples=.2, models=['BM','SMp','CMq0','rM8p','RM95p'], title=None, perm='7904213568')
| Materials/161103-run-plot4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hhw
# language: python
# name: hhw
# ---
from __future__ import absolute_import, division, print_function
import tensorflow as tf
tf.enable_eager_execution()
with tf.device("/cpu"):
x = [[2.]]
m = tf.matmul(x, x)
print("hello, {}".format(m))
| play_with_api/basic_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chunribu/biotable/blob/main/src/omim_db_full_data_sqlite3_to_tsv.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DYq5a7-VzUSn"
# ## Install `omim` providing OMIM data (maybe need update manually)
# + colab={"base_uri": "https://localhost:8080/"} id="rWpgOROgxOrJ" outputId="d851c4ed-7a1e-4afb-a70e-f69651bdc834"
# !pip install --quiet omim
# !omim -h
# + [markdown] id="Mpde3p72yogG"
# ## Load OMIM DB data provided by omim package
# + id="563fcea9-d5f0-463d-a373-b4f3d344135c"
import pandas as pd
import sqlite3
import json
# + id="a58ae998-4edc-4bc4-9f66-790484c997a3"
# use the path to database file printed in help
con = sqlite3.connect("/usr/local/lib/python3.7/dist-packages/omim/data/omim.sqlite3")
omim = pd.read_sql('select * from omim', con)
omim.to_csv('omim_package_db.tsv.gz', sep='\t')
con.close()
# + tags=[] colab={"base_uri": "https://localhost:8080/", "height": 971} id="ccac594a-6d64-45ac-9720-7415b94eff84" outputId="1da4da99-e452-4e09-8672-6685e57d0682"
omim
# + [markdown] id="Iq4lwsJHzOy6"
# ## Merge geneMap and phenotypeMap informations together
# + colab={"base_uri": "https://localhost:8080/"} id="795456f9-d2c7-46d3-9bb9-8676024e2c6f" outputId="8d56096f-18f7-4005-c5bb-a9f21509aa24"
_map = omim.geneMap.copy()
print(_map.isna().value_counts()[False])
_map.update(omim.phenotypeMap)
_map.isna().value_counts()[False]
# + [markdown] id="uOXeOtIC0yBK"
# ## Parse `_map` in `json` module
# + id="ade356e4-1b3f-4125-88c6-06087f435db7"
_map_list = _map.apply(lambda x: json.loads(x) if x!=None else [{}])
# + [markdown] id="ZIRQ2BUh1Nzs"
# ## Make duplicate rows according to the length of each `_map_list`
# + id="1051f3a6-b474-4226-87c7-dde8e0c66433"
lens = _map_list.str.len()
idx = omim.index.repeat(lens)
# + colab={"base_uri": "https://localhost:8080/"} id="19e137d5-564c-47dc-a8c7-9dbd3d90e146" outputId="f994a1ca-0cf4-46fa-f464-a1ae281d80d5"
lens.sum()
# + colab={"base_uri": "https://localhost:8080/"} id="2b2190ee-e19e-41a3-bbdd-94ed9b617d23" outputId="cb575c42-a8d4-42cd-b705-78294ccc2e6e"
len(_map_list.sum())
# + id="9fc868bb-d731-4f20-b4bb-22016855141e"
df_map = pd.DataFrame(_map_list.sum())
# + id="c2f68bd3-fef4-4fcf-8d8c-38e99206ddda"
df_map = df_map.replace('', None)
# + [markdown] id="aBiWjmlf12P7"
# ## Merge the original and new-generated DataFrames
# + id="4ff19d99-1cee-4abc-a5ca-8e1acb052801"
omim_merged = omim.loc[idx].reset_index().merge(df_map, left_index=True, right_index=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 669} id="b7a88984-1435-4496-aa53-7e21d5ffa115" outputId="5d39ec21-71c1-4929-fb5c-60f9dc156943"
omim_merged.head()
# + [markdown] id="N9CjhMf52InE"
# ## Select fields to save locally
# + id="88a5193c-0da2-48b0-8288-8070ef4c08ea"
omim_merged = omim_merged[['mim_number','mim_type','prefix','title','references','entrez_gene_id','ensembl_gene_id','hgnc_gene_symbol','Phenotype','Phenotype MIM number','Inheritance','Location','Gene/Locus','Gene/Locus MIM number','generated']]
# + id="caac5fb5-162c-449b-8154-f1c18a4d777f"
omim_merged.to_csv('OMIM.flat.tsv.gz', sep='\t', index=False)
| src/omim_db_full_data_sqlite3_to_tsv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import matplotlib as mpl
import numpy as np
from matplotlib.offsetbox import TextArea, DrawingArea, OffsetImage, AnnotationBbox
# %config InlineBackend.figure_format = 'retina'
WRYH = mpl.font_manager.FontProperties(
fname='WeiRuanYaHei-1.ttf')
plt.rcParams['font.sans-serif'] = ['Songti SC']
# +
plt.figure(figsize=(9, 6), dpi=150)
# 修改为你的数据
good = [93, 97, 91, 82, 78]
bad = [7, 3, 9, 18, 22]
x = [i+1 for i in range(len(good))]
plt.xlim(1, len(good)+1)
plt.ylim(0, 100)
plt.plot(x, good, linestyle='-', marker='D',
markerfacecolor='#FFF', linewidth=1.5, markersize=5.5, color='#00C5D2')
plt.plot(x, bad, linestyle='-', marker='D',
markerfacecolor='#FFF', linewidth=1.5, markersize=5.5, color='#4E70F0')
plt.fill_between(x, good, bad, interpolate=True, color='#EDF6FF')
ax = plt.gca()
ax.spines['right'].set_color('None')
# ax.spines['left'].set_color('None')
ax.spines['top'].set_color('None')
# ax.spines['bottom'].set_color('None')
locs, labels = plt.xticks()
# 下面的x轴坐标需要修改
new_xticks = ['淘票票', '猫眼', '豆瓣', '烂番茄', 'IMDB', '电影平台']
plt.xticks(locs, new_xticks, rotation=345,
horizontalalignment='center') # 修改刻度
locs, labels = plt.yticks()
# 下面的y轴坐标需要修改
new_yticks = ['0%', '20%', '40%', '60%', '80%', '100%']
plt.yticks(locs, new_yticks,
horizontalalignment='right') # 修改刻度
ax.tick_params(bottom=False, top=False, left=True, right=False) # 隐藏刻度线
# 下面的文字与坐标需要修改
plt.grid(linewidth=0.5, alpha=0.9, axis='y', linestyle=(0, (10, 3)))
plt.text(0.86, 87.5, ' 93% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#00C5D2", ec='#333',), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(1.86, 101, ' 97% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#00C5D2", ec='#333',), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(2.86, 85.7, ' 91% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#00C5D2", ec='#333',), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(3.86, 76.7, ' 82% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#00C5D2", ec='#333',), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(4.86, 72.7, ' 78% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#00C5D2", ec='#333',), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(0.86, 11.5, ' 7% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#5172F0", ec='#333', linestyle='-'), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(2.1, -0.3, ' 3% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#5172F0", ec='#333', linestyle='-'), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(2.87, 12.5, ' 9% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#5172F0", ec='#333', linestyle='-'), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(3.87, 22, ' 18% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#5172F0", ec='#333', linestyle='-'), fontsize=8.1, family='DejaVu Serif', weight='heavy')
plt.text(4.87, 26, ' 22% ', color='#FFF', bbox=dict(boxstyle="round,pad = 0.5",
fc="#5172F0", ec='#333', linestyle='-'), fontsize=8.1, family='DejaVu Serif', weight='heavy')
ax.annotate(' 好评率 ', [5.1, 78.5],
(5.8, 78.5),
ha="right", va="center",
size=12,
color='w',
arrowprops=dict(arrowstyle= 'fancy,head_length=0.5,head_width=0.6,tail_width=0.8',
shrinkA=5,
shrinkB=50,
fc="#00C5D2", ec="k",
connectionstyle="arc3,rad=0.03"
),
bbox=dict(boxstyle="square", fc="#00C5D2",linestyle = '--' ,linewidth=0.6))
ax.annotate(' 差评率 ', [5.1, 21.5],
(5.8, 21.5),
ha="right", va="center",
size=12,
color='w',
arrowprops=dict(arrowstyle= 'fancy,head_length=0.5,head_width=0.6,tail_width=0.8',
shrinkA=5,
shrinkB=50,
fc="#5172F0", ec="k",
connectionstyle="arc3,rad=0.03"
),
bbox=dict(boxstyle="square", fc="#5172F0",linestyle = '--' ,linewidth=0.6))
plt.text(1, 115, '2009—2021年,电影《阿凡达》在国内外不同电影平台', size=20)
plt.text(4.6, 103, '评分几何?', size=26
,color= '#2259A0')
plt.text(3.95,95.7,'数据来源:各大电影平台官方页面|采集时间:4/6/2021',alpha = 0.5,size = 8, fontproperties=WRYH)
plt.show()
# -
| B-折线图/高级折线图/高级折线图.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Mh6dGF4NpnxX" colab_type="text"
# # Introduction to Tensorflow and Sonnet
#
# By the end of this colab you will have trained a neural net to approximate the NXOR function based on some data. In the process you will have learnt about
#
# * some useful tensorflow tensor operations
# * building a model with *Tensorflow* and *Sonnet*
# * visualizing the model you built
# * getting the data into your model
# * backpropagation as implemented by tensorflow
# * debugging tensorflow models
# * how to actually train the network.
#
# Recall: you can use the outline on the right hand side to navigate the colab easier.
# + id="fTqcFF-_gDRD" colab_type="code" outputId="e3458a38-a2ad-4339-850d-3e4e29e3d36e" colab={"base_uri": "https://localhost:8080/", "height": 109} executionInfo={"status": "ok", "timestamp": 1543882811820, "user_tz": -60, "elapsed": 2982, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
#@title Fetching (DM) sonnet from pip. Run this cell.
# !pip install dm-sonnet
# + id="HFzztlr1p1F9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 74} outputId="8f4e1879-8b45-4c76-dcba-4e2d37dddc74" executionInfo={"status": "ok", "timestamp": 1543882823296, "user_tz": -60, "elapsed": 10123, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
#@title Imports. Run this cell.
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import random
import seaborn as sns
import numpy as np
import tensorflow as tf
import sonnet as snt
from matplotlib import pyplot as plt
from google.colab import files
from scipy.stats import multivariate_normal
from IPython.display import clear_output, Image, display, HTML
sns.set_style('ticks')
# + id="naSbSB6q1K1e" colab_type="code" cellView="form" colab={}
#@title Utility functions. Run this cell.
def get_data(num_examples):
inputs = 2*np.random.random((num_examples, 2)) - 1
labels = np.prod(inputs, axis=1)
labels[labels <= 0] = -1
labels[labels > 0] = 1
return inputs, labels
def plot_nxor_data(inputs, labels, title):
MARKER_COLORS = np.array([
[1.0, 0.0, 0.0], # red for -1
[0.0, 1.0, 0.0], # green for +1
])
class_idx = (labels + 1 / 2.0).astype(np.int)
plt.figure()
plt.title(title)
plt.scatter(
x=inputs[:, 0], y=inputs[:, 1], c=MARKER_COLORS[class_idx], alpha=0.9)
plt.legend()
plt.show()
def strip_consts(graph_def, max_const_size=32):
"""Strip large constant values from graph_def."""
strip_def = tf.GraphDef()
for n0 in graph_def.node:
n = strip_def.node.add()
n.MergeFrom(n0)
if n.op == 'Const':
tensor = n.attr['value'].tensor
size = len(tensor.tensor_content)
if size > max_const_size:
tensor.tensor_content = "<stripped %d bytes>"%size
return strip_def
def show_graph(graph_def=None, max_const_size=32):
"""Visualize TensorFlow graph. Default to the default graph."""
if graph_def is None:
graph_def = tf.get_default_graph()
if hasattr(graph_def, 'as_graph_def'):
graph_def = graph_def.as_graph_def()
strip_def = strip_consts(graph_def, max_const_size=max_const_size)
code = """
<script src="//cdnjs.cloudflare.com/ajax/libs/polymer/0.3.3/platform.js"></script>
<script>
function load() {{
document.getElementById("{id}").pbtxt = {data};
}}
</script>
<link rel="import" href="https://tensorboard.appspot.com/tf-graph-basic.build.html" onload=load()>
<div style="height:460px">
<tf-graph-basic id="{id}"></tf-graph-basic>
</div>
""".format(data=repr(str(strip_def)), id='graph'+str(np.random.rand()))
iframe = """
<iframe seamless style="width:950px;height:480px;border:0" srcdoc="{}"></iframe>
""".format(code.replace('"', '"'))
display(HTML(iframe))
# + [markdown] id="UVPFcNlJftrQ" colab_type="text"
# ## The TensorFlow Paradigm
#
# This section is not necessarily a fully complete introduction to tensorflow. If you are not familiar with tensorflow or don't feel comfortable with some of the content consider using a third party tutorial or the tensorflow documentation.
#
# Instead this colab focuses on exploring the ideas underlying tensorflow and working with it, highlighting important concepts along the way.
#
# + [markdown] id="aPaS-ObaIuyE" colab_type="text"
# **There are two distinct phases when it comes to working with tensorflow:**
#
# 1. Constructing the computation graph, our model,
# 2. Running data through this graph.
#
# We soon see what this means.
#
# *Note:* that with TensorFlow *Eager mode* this is not the case anymore: there the two phases happen hand in hand. Here we work with *Graph mode*, however.
# + [markdown] id="OYynIRfG150V" colab_type="text"
# ### Building and displaying graphs
#
# Let's build a simple computation graph:
# + id="gpcEcp-8sKwE" colab_type="code" outputId="04591d41-a889-4293-8300-7ce460540595" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882829209, "user_tz": -60, "elapsed": 492, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
a = tf.constant([5, 3, 1])
b = tf.constant([-1, 2, 5])
c = a + b
c
# + [markdown] id="1Z7kU1QAsYRz" colab_type="text"
# Notice that `c` has no value associated. It is actually a (reference to a) node in the computation graph we just defined: tensorflow knows that to find the value of `c`, it needs to query the values of the nodes `a` and `b` and add them.
#
# + [markdown] id="pIhpA_m-vcjw" colab_type="text"
# **In tensorflow all computation is implemented as operations on tensors (or variables, etc), and this computation forms a graph.**
#
# * We add tensors and operations to a graph with our Python code and libraries.
# * The tensorflow API [docs](https://www.tensorflow.org/api_docs/python/) list all available operations.
# * In practice many -- if not most -- `numpy` operations have a tensorflow counterpart, though often not by that same name.
# + [markdown] id="iYB9fb3KzbdN" colab_type="text"
# We can visualize the graph we have built so far. `show_graph()` is a utility function we defined above<sup>1</sup>; it shows the tensorboard graph representation of the graph you pass to it, right here in colab.
#
# <small>1: The graph visualization code is from the [J<NAME> Blog](https://blog.jakuba.net/2017/05/30/tensorflow-visualization.html#Using-a-cloud-hosted-TensorBoard-instance-to-do-the-rendering).</small>
#
# + id="i7mfP36k1dbz" colab_type="code" outputId="de9fc373-0c59-4260-a9f4-022b9b567915" colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"status": "ok", "timestamp": 1543882833523, "user_tz": -60, "elapsed": 491, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
show_graph(tf.get_default_graph())
# + [markdown] id="RC17oYAE1dhx" colab_type="text"
# Note that in tensorflow you can have many graphs at the same time. By default, unless otherwise specified, we are building the so called "default graph" that we accessed with `tf.get_default_graph()`.
# + [markdown] id="kXEufNds1uM-" colab_type="text"
# ### Resetting the default graph
#
# Recall that colab cells run in arbitrary order, maintaining python state between them. Therefore, if you run a cell that adds some tensors or operations to the graph, you will add more and more copies of them to the graph. This is probably not what you want.
#
# **Try running the cell where we defined node `c` a few more times, then visualizing the graph.** You will see multiply copies the same nodes.
# + [markdown] id="Yfv-3PiDL4Tz" colab_type="text"
# To solve this issue, tensorflow has `tf.reset_default_graph()`, which clears everything from the default graph.
# + id="yn7tl-2I2eqz" colab_type="code" outputId="00a8c576-1a63-4dc2-b2e1-5afb832e376e" colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"status": "ok", "timestamp": 1543882840863, "user_tz": -60, "elapsed": 498, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
a = tf.constant(5, name='a')
b = tf.constant(-1, name='b')
c = tf.add(a, b, name='c')
show_graph(tf.get_default_graph())
# + [markdown] id="7Sg1A_YK1uSX" colab_type="text"
# Whenever in doubt about your current graph, you can just reset it and rebuild it.
#
# By the way, notice that in the previous code cell we labelled nodes in the graph using the `name` argument. This can often help us interpret the graph.
# + [markdown] id="i6GT1cHLp1Lk" colab_type="text"
# ### Running the graph
#
# Recall that `c` had no associated value -- we were merely informed that it is a tensor, it's shape, etc. **Tensors only have values when 'run' in a session**.
# + id="x9CnLzuW4DQW" colab_type="code" outputId="7e570c14-0df1-414f-ca0f-b5c79e380a84" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882847265, "user_tz": -60, "elapsed": 1009, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
a = tf.constant([5, 2], name='a')
b = tf.constant([-1, 0], name='b')
c = tf.add(a, b, name='c')
with tf.Session() as session:
print(session.run(c))
# + [markdown] id="N-DIUd_B0HBg" colab_type="text"
# What really happens is that when you pass a graph node (operation, tensor, etc) to `session.run()`, tensorflow figures out what is the minimal subset of the graph to run in order to satisfy your request, and runs only that. It's difficult to appreciate this in the context of the simple graphs we had so far, but we will see a good example shortly.
#
# + [markdown] id="KLSwi1NBxj11" colab_type="text"
# You can run any node from your graph, or a combination of them.
# + id="1gibFQ3_xkz1" colab_type="code" outputId="e2810d4c-8ab0-410c-cdf9-5f35aca54bbb" colab={"base_uri": "https://localhost:8080/", "height": 72} executionInfo={"status": "ok", "timestamp": 1543882850182, "user_tz": -60, "elapsed": 542, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
with tf.Session() as session:
print('a:', session.run(a)) # As noted above, in this case addition
# (required to find the value of c) is not even
# executed.
print('[b, c]:', session.run([b, c]))
print(session.run({'a': a, 'c': c}))
# + [markdown] id="UY1I6AnRzxKM" colab_type="text"
# The data flows through the graph just once, but tensorflow runs all requested operations and tensors (along with their dependencies), returning their calculated values. We can easily illustrate how this work with tensors that get a new random value each time you run them. **Try predicting the pattern before inspecting the printed results!**
# + id="o0x5QAcq1rc4" colab_type="code" outputId="98f20567-212b-4533-8e45-239a93b19bee" colab={"base_uri": "https://localhost:8080/", "height": 74} executionInfo={"status": "ok", "timestamp": 1543882852915, "user_tz": -60, "elapsed": 494, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
r = tf.random_normal(shape=(3,), mean=0.0, stddev=1.0)
x1 = r + 1 # Shifted +1
x2 = r - 1 # Shifted -1
with tf.Session() as session:
print('x1, x2 run separately:', session.run(x1), session.run(x2))
print('x1, x2 run together :', session.run([x1, x2]))
# + [markdown] id="HEP8J_ON3IhI" colab_type="text"
# Notice that
#
# * when x1 and x2 were run together, the difference between correpsonding entries is always 2,
# * while this is not the case when they were run separately.
#
# This is because when run together, `r` is sampled once, and both `x1` and `x2` use this same value.
# + [markdown] id="U_6zhb8OPQ68" colab_type="text"
# We now highlight what this means for neural network training implemented in tensorflow.
#
# ### A neural network example of tensorflow's computational model
#
# All computation required to train the network will be implemented as a tensorflow computation graph. In particular you will have tensor and operations like
#
# * `train`: take a training step on some data,
# * `loss`: calculate the loss on some data,
# * `outputs`: give you predictions on some data,
# * and so on.
#
# Given the computation model of tensorflow:
#
# * You will be able to `run(loss)` to calculate the loss, and **without triggering the training step computation**.
# * On the other hand, running `train` will calculate the `loss` since this is what it needs to optimize.
#
# If you `run([loss, train])`, tensorflow will take a training step and report the loss, **both based on the same data**.
#
#
# As a final note, the fact that only the **minimal required subset of nodes are run** is going to be crucial when using BatchNorm: the ops that update the statistics kept in BatchNorm are not dependencies of any other ops, therefore will not get run automatically. You will experiment with this in the ConvNets and Vision Lab.
# + [markdown] id="vSLzoomC4yY3" colab_type="text"
# ### Running a graph with state and inputs
#
# Our examples so far have been silly in the sense that they were straightforward computation on constants, not warranting a computation graph. We now showcase a situation where the value of a tensor is not defined until it is run; this is because the value is dependent on data fed to the graph at running time.
# + id="3GjCHShI468O" colab_type="code" outputId="838efa9c-1d94-4553-a72c-66105b01ceb4" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1543882858537, "user_tz": -60, "elapsed": 515, "user": {"displayName": "Luk\u00e1\u016<NAME>00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
a = tf.placeholder(dtype=tf.int32, shape=(), name='input')
b = tf.constant(-1, name='b')
c = tf.add(a, b, name='c')
with tf.Session() as session:
print(session.run(c, feed_dict={a: 3}))
print(session.run(c, feed_dict={a: 10}))
# + [markdown] id="XEeJPNHFgtlW" colab_type="text"
# We used a `tf.placeholder`. These are tensors that have no value or computation associated to them by default, instead they simply take data so this data can be computed on by the rest of the graph.
#
# Note that, at the same time, **any tensor may be fed with some data**.
# + [markdown] id="88G0QbH44DWX" colab_type="text"
# Another strength of the computation graph approach is that some nodes may be stateful. The most common stateful node is a *variable*. **A variable is a tensor that remembers its value between run calls**. This also means **it must be initialized**.
#
# In the following example `a` will be a variable. We also define an `inc` operation that increments the value of `a` by 1 each time this operation is run.
# + id="Dym9zktt6MkU" colab_type="code" outputId="be5df224-e4ea-4366-d1ed-4a1a8e2606ea" colab={"base_uri": "https://localhost:8080/", "height": 72} executionInfo={"status": "ok", "timestamp": 1543882861922, "user_tz": -60, "elapsed": 506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
a = tf.get_variable('counter', shape=(), dtype=tf.int64)
inc = tf.assign(a, a+1)
init_op = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init_op) # Sets an initial value for a.
print(session.run(a)) # By default, this is 0.
print(session.run(a))
session.run(inc)
session.run(inc)
print(session.run(a)) # We see the variable was incremented (twice).
# If you were to print the output of inc, you see that it actually
# returns the value of a post-increment. This is a convenience feature
# of tf.assign().
# + [markdown] id="6gcSLObKiB2-" colab_type="text"
# Statefulness is highly relevant to us since the weights of our machine learning models are stored as variables and are updated by some operations in `session.run` calls during training.
# + [markdown] id="SMndofivB1UA" colab_type="text"
# ### Quick Tour of Control dependencies and Race conditions
#
# *These topics do not often come up when training simple neural networks, but they are core concepts of tensorflow and you should be familiar with them.*
#
# With the introduction of stateful graph components we need to revisit the rule that tensorflow only executes the minimal set of operations required by a `run()` call. **Try predicting the output of the following cell.**
#
# + id="CcfLCBN6B1wg" colab_type="code" outputId="1b0e2b38-f704-4ad7-d56e-9a1ad5762bc1" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882864988, "user_tz": -60, "elapsed": 484, "user": {"displayName": "Luk\u00e1\u01<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
x = tf.get_variable("x", shape=(), initializer=tf.zeros_initializer())
assign_x = tf.assign(x, 10.0)
z = x + 1.0
init = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init)
print(session.run(z))
# + [markdown] id="2GG5vevHC2Ln" colab_type="text"
# From tensorflow's perspective,
#
# * There is a variable `x`, which starts with value 0,
# * `z` is always `x+1`,
# * with `assign_x` you can set the value of `x` to 10.
#
# So if you simply ask for the value of `z`, tensorflow evaluates the minimal subset of the graph it needs and reports that `z = 0 + 1`. This is reflected in the graph as well.
# + id="RWragQhLC2Sq" colab_type="code" outputId="ed172ee5-d4c8-4d36-b136-fb3fcd81d44c" colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"status": "ok", "timestamp": 1543882866972, "user_tz": -60, "elapsed": 526, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
show_graph()
# + [markdown] id="9j_eg2vUC2ZA" colab_type="text"
# If you want `x` incremented by 10 before using it to calculate `z`, you need to tell tensorflow. You can do so by specifying `assign_x` as a (control_)dependency of z.
# + id="5ZPI-FIqDdKD" colab_type="code" outputId="cc030202-2147-4a93-d2cc-87efa9af1aa4" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882870491, "user_tz": -60, "elapsed": 493, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
x = tf.get_variable("x", shape=(), initializer=tf.zeros_initializer())
assign_x = tf.assign(x, 10.0)
with tf.control_dependencies([assign_x]):
z = x + 1.0
init = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init)
print(session.run(z))
# + [markdown] id="2rvcRUhGDdQu" colab_type="text"
# To be precise, `tf.control_dependencies` ensures all operations / tensors passed to it are run before running the the operations defined inside its body.
# + [markdown] id="9jIrvWWzFMZN" colab_type="text"
# The other rule to keep in mind is that **Tensorflow is inherently parallel.** If there are computation subgraphs that do not depend on each other, they can -- and likely will be -- evaluated in parallel. We use the same generic example to illustrate this.
# + id="yyN-AnRfGABu" colab_type="code" outputId="0aeb5e1f-5954-46c3-83e5-07f8c3b9a636" colab={"base_uri": "https://localhost:8080/", "height": 201} executionInfo={"status": "ok", "timestamp": 1543882873965, "user_tz": -60, "elapsed": 490, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
x = tf.get_variable("x", shape=(), initializer=tf.zeros_initializer())
assign_x10 = tf.assign(x, 10.0)
assign_x5 = tf.assign(x, 5.0)
z = x + 1.0
init = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init)
for _ in range(10):
_, _, z_val = session.run([assign_x10, assign_x5, z])
print(z_val)
# + [markdown] id="68duXIjaFMis" colab_type="text"
# We can see that `z` can take various values: its value will depend on what order the different operations get run -- which we don't control. (If you do not see different values, re-run the cell until you do. If it doesn't help, this property has probably changed in some recent update of TensorFlow.)
# + [markdown] id="akMRszLOMFXi" colab_type="text"
# The lesson is that **if you care about the order of otherwise independent operations, you must be explicit about this**.
# + [markdown] id="woBAfFv7h5ua" colab_type="text"
# ### Exercise: "Interactive Tensorflow Summing Machine"
#
# Write a tensorflow graph which keeps a running sum of the integers passed to it through a `feed_dict`. To make sure it works feed the machine a few numbers, printing the cumulative sum after each step.
# + id="PhmThcmDh6FR" colab_type="code" colab={}
#@title Your Code
# + id="9jTyd1LrilBt" colab_type="code" cellView="form" outputId="6224b095-6a07-434c-d92d-f77d80639255" colab={"base_uri": "https://localhost:8080/", "height": 109} executionInfo={"status": "ok", "timestamp": 1543882882225, "user_tz": -60, "elapsed": 503, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
#@title Solution
tf.reset_default_graph()
cumulative_sum = tf.get_variable('sum', shape=(), dtype=tf.int64)
to_add = tf.placeholder(dtype=tf.int64, shape=(), name='input')
add = tf.assign(cumulative_sum, cumulative_sum + to_add)
init_op = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init_op) # Sets an initial value for a.
for i in range(1, 6):
print('cumulative sum={}; adding {}.'.format(session.run(cumulative_sum), i))
session.run(add, feed_dict={to_add: i})
# + [markdown] id="XvriNEgIAbms" colab_type="text"
# ## A word (section) on tensorflow tensor shapes
#
# Tensors in Tensorflow have **static and dynamic shape**.
#
# * Static shape information is known or can be deduced at graph construction time,
# * Dynamic shape information is only available when data is available.
#
# **Static shape may be and is often only partially defined**. For example, we may know that our model expect a batch of examples, each of shape `2 x 2`, but not how large these batches are. This will allow us to feed the computation graph with batches of any size. Once data is fed the tensors will have a known **dynamic shape**.
# + id="dceGjHt6Ab6y" colab_type="code" outputId="92250c9f-2cf2-4c2a-d64f-bcc38cce27d2" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882888041, "user_tz": -60, "elapsed": 516, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
inputs = tf.placeholder(dtype=tf.int32, shape=(None, 2, 2), name='input')
print('static shape:', inputs.shape)
# + [markdown] id="KOZU2rgKBQRl" colab_type="text"
# We pass `None` for axes that we do not know the static length of when specifying a shape. When a tensor or its shape is printed, this is denoted by a question mark, `?`, as seen above.
#
# + [markdown] id="rW8LjIbXhPKG" colab_type="text"
# **Bug-alert:** Be careful not to confuse passing `(None)` vs `(None,)` as a desired shape. The next cell illustrates the consequences:
# + id="FtF9B5mngHMt" colab_type="code" outputId="bb143e8f-87b3-41b8-86ba-97825c814a03" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1543882891488, "user_tz": -60, "elapsed": 595, "user": {"displayName": "Luk\u00e1\u016<NAME>\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
inputs_1 = tf.placeholder(dtype=tf.int32, shape=(None), name='input')
inputs_2 = tf.placeholder(dtype=tf.int32, shape=(None,), name='input')
print(inputs_1.shape) # Shape that we know nothing about, not even rank.
print(inputs_2.shape) # Tensorflow will assert that the tensor is of rank 1,
# albeit with unknwon length.
# + [markdown] id="2xvgx50cf3B2" colab_type="text"
# The static shape information is used to
#
# * verify operations make sense (think matrix multiplication),
# * infer the static shape of tensors defined through operations (so they can also be checked) .
#
# **Example**
#
# 1. We take `batch (?) x 2 x 2`-shaped tensors, flatten each example in the batch to be a vector of length `4`. Tensorflow will infer the shape of the flattened tensor automatically.
# 2. Then we multiply the now `? x 4`-shaped tensor with a vector. Tensorflow will only allow this to happen if the vector is of length 4, as otherwise the operation makes no sense.
#
# (In practice the `tf.matmul` operation we use does not accept vectors, so we will use a `4 x 1` matrix instead.)
# + id="s9ANUmVrB7qY" colab_type="code" outputId="599ab562-2ad5-4dd7-bd7a-611ae84d46ec" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1543882895442, "user_tz": -60, "elapsed": 591, "user": {"displayName": "Luk\u00e1\u016<NAME>\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
inputs = tf.placeholder(dtype=tf.int32, shape=(None, 2, 2), name='input')
flat_inputs = tf.contrib.layers.flatten(inputs)
print('flat_inputs static shape', flat_inputs.shape)
result = tf.matmul(flat_inputs, tf.constant([[0], [1], [2], [3]], name='ok'))
print('result static shape', result.shape)
# Uncomment and run to see
#
# ValueError: Dimensions must be equal, but are 4 and 3 for 'MatMul_4'
# (op: 'MatMul') with input shapes: [?,4], [3,1].
#
# tf.matmul(flat_inputs, tf.constant([[0], [1], [2]], name='shape_mismatch'))
# + [markdown] id="TlHbOikVBvwo" colab_type="text"
# It happens sometimes (e.g. for custom operations) that tensorflow is not be able to infer the static shape of the resulting tensor. f you know the expected shape, you can explicitly set it using `tensor.set_shape()`. This will allow tensorflow to infer and check later shapes.
# + [markdown] id="bolN-5BRF2Pf" colab_type="text"
# Finally, let us try working with the dynamic shape of a tensor.
# + id="x9U6SsqVBQXc" colab_type="code" outputId="a36d424d-328e-4c59-803a-ea530428b01c" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882899913, "user_tz": -60, "elapsed": 483, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
print('dynamic shape:', tf.shape(inputs))
# + [markdown] id="4sXdJmSLF1l-" colab_type="text"
# The **dynamic shape itself is a tensor** and may (only) be evaluated or computed with once the graph is run in a session.
# + id="3tFB0_e9GDqi" colab_type="code" outputId="21d9b346-157f-45cd-c4a1-bcf0923a5f23" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882902616, "user_tz": -60, "elapsed": 617, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
shape = tf.shape(inputs)
num_total_elements = tf.reduce_prod(shape)
with tf.Session() as session:
print(session.run([shape, num_total_elements], feed_dict={
inputs: np.array(np.random.random((3, 2, 2)))
}))
# + [markdown] id="R05v98YcrjeR" colab_type="text"
# ### Broadcasting
#
# Tensorflow automatically broadcasts operations, similarly to `numpy`. We covered broadcasting in detail in the `numpy` colab. Here we include three common examples.
# + id="yVAhM_F0qIjP" colab_type="code" outputId="2e0dec62-4b4c-42b1-cf6b-21f8ce476d7b" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1543882905044, "user_tz": -60, "elapsed": 482, "user": {"displayName": "Luk\u00e1\u016<NAME>00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
a = tf.constant([[1, 2, 3], [4, 5, 6]], name='a')
c = a - 1 # `1` is first turned into a constant,
# then broadcast across the full tensor
with tf.Session() as session:
print(session.run(c))
# + id="Wj449M26rj9Z" colab_type="code" outputId="b8f8fc72-a088-4c65-e758-4caf9054d483" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1543882906663, "user_tz": -60, "elapsed": 554, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
tf.reset_default_graph()
a = tf.constant([[1, 2, 3], [4, 5, 6]], name='a')
b = tf.constant([1000, 100, 10], name='b')
c = a + b
# a: 2 x 3
# b: 3
# --> b is copied over across the first axis to calculate c.
with tf.Session() as session:
print(session.run(c))
# + id="0eC_cWM1s6Ub" colab_type="code" colab={}
tf.reset_default_graph()
a = tf.constant([[1, 2, 3], [4, 5, 6]], name='a')
b = tf.constant([100, 10], name='b')
# a: 2 x 3
# b: 2
# --> a and b are not compatible;
# a + b # Raises an error.
# Uncomment lines below to see the error
#c = a + b
#with tf.Session() as session:
# print(session.run(c))
# + id="itipg8-pgG_w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="f161c598-ed76-4b4c-8384-677f17ed873d" executionInfo={"status": "ok", "timestamp": 1543882909639, "user_tz": -60, "elapsed": 542, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
# Instead, b can be defined as [[100], [10]] so that
# a: 2 x 3
# b: 2 x 1
# --> b is copied across the last axis.
tf.reset_default_graph()
a = tf.constant([[1, 2, 3], [4, 5, 6]], name='a')
b = tf.constant([[100], [10]], name='b')
c = a + b
with tf.Session() as session:
print(session.run(c))
# + [markdown] id="X_FfiXzOqb6R" colab_type="text"
# **As a general rule of thumb**
#
# * use broadcasting in the simple cases
# * prefer explicit broadcasting in complex situations.
#
# This will result in code that is **easier to read** and has **fewer bugs**.
# + [markdown] id="7-_uTExc6M7e" colab_type="text"
# ## Building a simple network with Sonnet
#
# Instead of building our neural networks in plain Tensorflow, we use the [sonnet](https://github.com/deepmind/sonnet) library.
#
# **Sonnet uses an object-oriented approach, similar to Torch/NN.**
#
# * This allows modules to be created, which define the forward pass of some computation.
# * Modules are ‘called’ with some input Tensors, which adds ops to the Graph and returns output Tensors.
#
# We call this a **configure-then-connect principle**, which allows for easy reuse of complex modules.
#
# + id="vaXAee8c_VwJ" colab_type="code" colab={}
tf.reset_default_graph() # You can always clear the current graph and
# add exactly what you need to it.
# + [markdown] id="_JX0ugUkl7dA" colab_type="text"
# Start by creating a Linear module (dense layer).
# + id="bXRJB-UN9QgM" colab_type="code" outputId="d7acf79a-c47c-4e2f-9478-954a0a223220" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882915201, "user_tz": -60, "elapsed": 509, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
linear = snt.Linear(output_size=5)
linear
# + [markdown] id="Nv4ROlyl9Qms" colab_type="text"
# Our input will be batches of 2-long vectors, and we will feed that data to the graph using `feed_dict`s.
# + id="mzMXfwp0_YeT" colab_type="code" colab={}
inputs_placeholder = tf.placeholder(tf.float32, shape=(None, 2), name='inputs')
# + [markdown] id="hQO7qLZM_Yrb" colab_type="text"
# As in tensorflow, we "call" the module on the tensor that we want it to compute on. This yields a tensor, the output of the calculation.
# + id="1_4Kmhdp_8jP" colab_type="code" colab={}
pre_activations = linear(inputs_placeholder)
# + [markdown] id="4wgBGo-f_8rI" colab_type="text"
# To complete our model, we apply a ReLU non-linearity and add a final linear layer with just 1 output.
# + id="8pKD3uf9_Y1u" colab_type="code" outputId="921d0735-fb44-4d9b-91b2-fcc7727f996c" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882922527, "user_tz": -60, "elapsed": 489, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
activations = tf.nn.relu(pre_activations)
outputs = snt.Linear(output_size=1)(activations)
outputs
# + [markdown] id="TH0GimMtsgqZ" colab_type="text"
# We drop the final singleton axis so that `outputs` becomes a vector.
# + id="Nn8wiKbAshd-" colab_type="code" outputId="00229253-34ab-45fc-84f4-1aacb7551523" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882925472, "user_tz": -60, "elapsed": 494, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
outputs = tf.squeeze(outputs, axis=-1)
outputs
# + [markdown] id="Xst87vC7kaV0" colab_type="text"
# Let's see the graph we built.
# + id="KJvV-Mggka2s" colab_type="code" outputId="3037e710-7b53-4487-aa41-31737862d778" colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"status": "ok", "timestamp": 1543882927897, "user_tz": -60, "elapsed": 502, "user": {"displayName": "Luk\u00e1\u016<NAME>\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
show_graph() # With no arguments show_graph() shows the default graph.
# + [markdown] id="9H3q1yfMkewp" colab_type="text"
# You can explore the exact set of tensorflow operations that were created the sonnet code by expanding colored boxes. **We can verify that each linear layer implements $WX+b$ for $X$ inputs and $W$ weights and $b$ bias with basic tensorflow operations**.
# + [markdown] id="0I76TX0ql3Q8" colab_type="text"
# Let's pass some data through our model. We will use the data generating function we wrote in the numpy colab. (It is redefined at the top of this colab).
# + id="Dugx7fxknxsV" colab_type="code" colab={}
init_op = tf.global_variables_initializer()
# + id="tCtNXgVAke4Q" colab_type="code" outputId="3d116021-0a6d-4181-df2e-3da38b621486" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1543882935743, "user_tz": -60, "elapsed": 743, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
inputs_np, unused_labels_np = get_data(num_examples=8)
with tf.Session() as session:
session.run(init_op) # Initializes the weights in the network.
outputs_np = session.run(outputs, feed_dict={
inputs_placeholder: inputs_np,
})
outputs_np
# + [markdown] id="ZfkWabTike_i" colab_type="text"
# You can rerun the above cell to see the output on new and new batches. The one thing that now remains is...
#
#
# + [markdown] id="LPlrr6NvqhjE" colab_type="text"
# ## Training a tensorflow model
#
# This is the same with or without sonnet.
#
# We will start by
#
# 1. Making the correct labels available to the graph,
# 2. Using these to define and calculate the loss on the output of the network.
# + id="5NjyuVM0nzVC" colab_type="code" colab={}
labels_placeholder = tf.placeholder(tf.float32, shape=(None,), name='labels')
# + [markdown] id="WM5liEzKTa0q" colab_type="text"
# Here we will simply regress onto the labels with the squared loss. (It would be better to calculate a cross entropy.)
# + id="dveVOLfC9GKf" colab_type="code" outputId="5f8fa5cf-4195-4194-dafc-67665d3d02a5" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882942175, "user_tz": -60, "elapsed": 495, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
with tf.name_scope('element_wise_loss'):
loss = tf.square(labels_placeholder - outputs)
loss
# + [markdown] id="9lTQIdr79e-_" colab_type="text"
# The loss tensor now calculates the loss per example. We want one scalar to optimize:
# + id="pT_esW_H9Veo" colab_type="code" outputId="a0da5ea9-8647-4b72-a3fc-e635663918dc" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543882945462, "user_tz": -60, "elapsed": 530, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
loss = tf.reduce_mean(loss, name='batch_mean_loss')
loss
# + [markdown] id="f6i4I7WInzlO" colab_type="text"
# We can verify on the graph that everything is as expected. The `name_scope` and `name` instructions make the graph easier to interpret.
# + id="AYrxqaptsnA-" colab_type="code" outputId="23021ca9-e49c-4f64-a660-b17c984b35ef" colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"status": "ok", "timestamp": 1543882948254, "user_tz": -60, "elapsed": 558, "user": {"displayName": "Luk\u00e1\u01<NAME>\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
show_graph()
# + [markdown] id="5Ad9LzFJsnHk" colab_type="text"
# We need to tell the computation graph that we want to minimize this loss.
# + id="mn5gwBw5s2kq" colab_type="code" colab={}
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
train_op = optimizer.minimize(loss)
# + [markdown] id="iLQ37jcduDwE" colab_type="text"
# **It is worth noting here the effect of this call on the graph.**
# + id="bqEc0SO-totM" colab_type="code" outputId="c2319beb-092d-4016-db3e-dac0c50e67ed" colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"status": "ok", "timestamp": 1543882963921, "user_tz": -60, "elapsed": 522, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
show_graph()
# + [markdown] id="BCcYLuZms1-B" colab_type="text"
# The minimization call added
#
# * gradient calculation operations
# * operations that update the weights based on these gradients.
#
# In fact, **we could have built the graph corresponding to `minimize()` manually** by
#
# * calculating the gradients of the loss with respect to the weights with the `tf.gradients(loss, [list of weights])` operation,
# * potentially scaling these gradients and adding them to the existing weights.
#
# + [markdown] id="JSOTFgNDnxQv" colab_type="text"
# By running the returned `train_op`, we take one gradient step, fitting the data just a bit better. Let's do this! But first some setup.
# + id="g1rq6u6JHuYV" colab_type="code" outputId="97ffe744-57c1-417c-a0a3-41655888bb0b" colab={"base_uri": "https://localhost:8080/", "height": 405} executionInfo={"status": "ok", "timestamp": 1543882970469, "user_tz": -60, "elapsed": 922, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
# Get some training data, and plot it. This is based on earlier exercises.
inputs_np, true_labels_np = get_data(num_examples=128)
plot_nxor_data(inputs_np, true_labels_np, title='Train data')
# Show some statistics that can help with debugging
print('Mean label on train data:', np.mean(true_labels_np))
# + id="-1rqXbD66xH4" colab_type="code" colab={}
init_op = tf.global_variables_initializer()
# + [markdown] id="kep9Tdjln0Ki" colab_type="text"
# **The final training script.**
#
# This cell contains all training and some reporting code. For now you can just run it, but for the next exercise you will have to understand it.
#
# *Note that sometimes we can get a bad weight initialization, but in a few runs you can easily get below 5% error.*
# + id="HxIP3cY4kfIA" colab_type="code" outputId="5345523c-8560-4f0e-8ec3-20edf65ab923" colab={"base_uri": "https://localhost:8080/", "height": 626} executionInfo={"status": "ok", "timestamp": 1543882990523, "user_tz": -60, "elapsed": 15650, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-<KEY>/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
RECORD_PERIOD = int(1e3)
training_steps = 10000 #@param {'type': 'integer'}
print('Losses:')
with tf.Session() as session:
session.run(init_op) # Initializes the weights in the network.
for i in range(training_steps):
_, loss_np = session.run(
[train_op, loss],
feed_dict={
inputs_placeholder: inputs_np,
labels_placeholder: true_labels_np,
})
if (i % RECORD_PERIOD) == 0:
print(' ', loss_np)
if loss_np < 0.01:
print()
print('Loss hit threshold after {} steps, stopping.'.format(i))
break
print()
# The model is ready to be evaluated. Fetch the predicted outputs.
predictions_np = session.run(outputs,
feed_dict={
inputs_placeholder: inputs_np,
})
# Actual label predictions given as {-1, +1}.
predictions_np[predictions_np <= 0] = -1
predictions_np[predictions_np > 0] = 1
# Prediction errors and plotting.
num_correct = np.count_nonzero(np.isclose(predictions_np, true_labels_np))
num_examples = true_labels_np.shape[0]
print('Prediction error:', (num_examples-num_correct)/num_examples)
plot_nxor_data(inputs_np, predictions_np, title='Predictions')
# + [markdown] id="BA0DfXiDWVym" colab_type="text"
# Notice that the prediction error calculation was inside the `with tf.Session()` context manager. This because **the graph state (including weights) is only maintained on a per session basis**. It is possible to save (and load) graphs, including their weights, with a [`tf.train.Saver`](https://www.tensorflow.org/api_docs/python/tf/train/Saver).
# + [markdown] id="SJLoWp90ueo4" colab_type="text"
# ## Exercise: Evaluate the trained model
#
# We have seen how to train the model -- that is -- we saw that the model can fit the training set well. But we are actually interested in generalizing to new examples from the same data distribution.
#
# 1. Define a training and a test dataset using our data generation function.
# 2. Fit the training data using the model we defined above.
# 3. Instead of reporting the prediction error only on the training set, also report it on the test set.
# 4. Plot the predictions on the test set using the pre-defined plotting function.
# + [markdown] id="Z-3LJCcXxSxO" colab_type="text"
# For simplicity, the full model building code is included in the cell below:
# + id="0-i6rjzYXGa8" colab_type="code" colab={}
tf.reset_default_graph()
# Inputs.
inputs_placeholder = tf.placeholder(tf.float32, shape=(None, 2), name='inputs')
labels_placeholder = tf.placeholder(tf.float32, shape=(None,), name='labels')
# All network and loss definition.
activations = tf.nn.relu(
snt.Linear(output_size=5)(inputs_placeholder))
outputs = tf.squeeze(
snt.Linear(output_size=1)(activations), axis=-1)
loss = tf.reduce_mean(
tf.squared_difference(labels_placeholder, outputs))
# Optimizer and initializer.
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.03)
train_op = optimizer.minimize(loss)
init_op = tf.global_variables_initializer()
# + id="uYdMlXp6uehe" colab_type="code" colab={}
#@title Your Code
# + id="nIDH0C6lWVku" colab_type="code" cellView="code" outputId="5f0bd437-0058-4460-bf16-c523749ff068" colab={"base_uri": "https://localhost:8080/", "height": 829} executionInfo={"status": "ok", "timestamp": 1543883031534, "user_tz": -60, "elapsed": 30995, "user": {"displayName": "Luk\u00e1\u016<NAME>\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
#@title Solution
# The solution is very similar to the previous training script, except care
# needs to be taken to have a separate train and test set.
train_inputs_np, train_labels_np = get_data(num_examples=256)
test_inputs_np, test_labels_np = get_data(num_examples=128)
TRAINING_STEPS = int(2e4)
RECORD_PERIOD = int(1e3)
def _get_predictions(inputs):
predictions_np = session.run(outputs,
feed_dict={
inputs_placeholder: inputs,
})
# Actual label predictions given as {-1, +1}.
predictions_np[predictions_np <= 0] = -1
predictions_np[predictions_np > 0] = 1
return predictions_np
def _get_error(predictions, true_labels):
num_correct = np.count_nonzero(np.isclose(predictions, true_labels))
num_examples = true_labels.shape[0]
return (num_examples-num_correct) / num_examples
print('Losses:')
with tf.Session() as session:
session.run(init_op) # Initializes the weights in the network.
for i in range(TRAINING_STEPS):
_, loss_np = session.run(
[train_op, loss],
feed_dict={
inputs_placeholder: train_inputs_np,
labels_placeholder: train_labels_np,
})
if (i % RECORD_PERIOD) == 0:
print(' ', loss_np)
if loss_np < 0.01:
print()
print('Loss hit threshold after {} steps, stopping.'.format(i))
break
print()
# The model is ready to be evaluated.
train_predictions = _get_predictions(train_inputs_np)
train_error = _get_error(train_predictions, train_labels_np)
test_predictions = _get_predictions(test_inputs_np)
test_error = _get_error(test_predictions, test_labels_np)
print('Train error:', train_error)
print('Test error:', test_error)
plot_nxor_data(test_inputs_np, test_predictions, title='Predictions')
# + [markdown] id="MAJnizTyrIan" colab_type="text"
# ## Datasets
#
# So far we used a `feed_dict`s to pass data to the computation graph. Another, often more efficient solution is to have nodes in the graph read, maninpulate, and make data available.
#
# Tensorflow has a dedicated `tf.data` module. Tensorflow's [Importing Data Guide](https://www.tensorflow.org/guide/datasets) guide is a great resource for learning about it. **Read this guide up to and including the "Reading input data > Consuming NumPy arrays"** section.
#
#
#
#
# + [markdown] id="3nrz4qxB_CwD" colab_type="text"
# ### Exercise: define a tensorflow dataset
#
# 1. Use the `get_data` function from before to generate a training dataset of 1000 examples and a test dataset of 500 examples.
# 2. Using `from_tensor_slices()`, define a training and a test `tf.data.Dataset`.
# 3. Ensure that the train data is (a) fully shuffled (b) can be iterated infinitely (c) is batched with a batch size of 64.
# 4. We do not shuffle the test data and we only want to iterate it once. We still batch it up so that the amount of data we compute on is limited.
#
# **Write a function called `get_tf_dataset()` that returns a (`train_dataset, test_dataset`)-tuple according to these instructions.** Print the returned datasets in order to verify they are correctly defined.
# + id="pNOBmK79HTpD" colab_type="code" colab={}
tf.reset_default_graph()
# + id="lksIuNH5_DjN" colab_type="code" colab={}
#@title Your Code
# + id="XxX8XhTKAS-9" colab_type="code" cellView="both" outputId="f23275ce-de9c-408c-b6d1-ff7a9430ce31" colab={"base_uri": "https://localhost:8080/", "height": 55} executionInfo={"status": "ok", "timestamp": 1543883052900, "user_tz": -60, "elapsed": 510, "user": {"displayName": "Luk\u00e1\u016<NAME>\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
#@title Solution
BATCH_SIZE = 64
train_data_np = get_data(1000)
test_data_np = get_data(500)
def get_tf_dataset():
train_dataset = tf.data.Dataset.from_tensor_slices(train_data_np)
train_dataset = train_dataset.shuffle(1000).repeat().batch(BATCH_SIZE)
test_dataset = tf.data.Dataset.from_tensor_slices(test_data_np)
test_dataset = test_dataset.batch(BATCH_SIZE)
return train_dataset, test_dataset
print(get_tf_dataset())
# + [markdown] id="3pxg1Tvj9l_X" colab_type="text"
# We need to access the data as tensors. We can do so by asking for an iterator over the dataset. We use the simplest iterator, which simply iterates over the dataset:
# + id="t_MqBL1AOLve" colab_type="code" colab={}
train_dataset, test_dataset = get_tf_dataset()
# + id="rMCtoQalD6go" colab_type="code" outputId="e97e52e2-ebfc-47f1-d93c-96b61f99a67b" colab={"base_uri": "https://localhost:8080/", "height": 111} executionInfo={"status": "ok", "timestamp": 1543883058986, "user_tz": -60, "elapsed": 554, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
train_data_iter = train_dataset.make_one_shot_iterator()
(train_inputs, train_labels) = train_data_iter.get_next()
train_inputs, train_labels
# + id="nO8DGVktHARQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 74} outputId="8cbc6b6a-ec35-4bb7-d80d-8feb2ffc568f" executionInfo={"status": "ok", "timestamp": 1543883065836, "user_tz": -60, "elapsed": 597, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
test_data = test_dataset.make_one_shot_iterator().get_next()
# + [markdown] id="PZ9iRdHjD6nE" colab_type="text"
# Now we can use `train_inputs` and `train_labels` like any other tensor. Each time we use them in a `session.run()` the tensor will hold a new batch.
# + id="STpmjTfOEduK" colab_type="code" outputId="254529b2-df0d-4043-8334-20ccc6df18ce" colab={"base_uri": "https://localhost:8080/", "height": 552} executionInfo={"status": "ok", "timestamp": 1543883072887, "user_tz": -60, "elapsed": 548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
def _print_some(np_array, descr):
print(descr + ':')
print(' shape: {}'.format(np_array.shape))
print(' first examples in batch: {}'.format(np_array[:4]))
with tf.Session() as session:
# Train data.
for _ in range(2):
train_inputs_np, train_labels_np = session.run([train_inputs, train_labels])
_print_some(train_inputs_np, 'train_inputs')
_print_some(train_labels_np, 'train_labels')
print()
# Test data.
test_inputs_np, test_labels_np = session.run(test_data)
_print_some(test_inputs_np, 'test_inputs')
_print_some(test_labels_np, 'test_labels')
# + [markdown] id="hZckeyEaTgO8" colab_type="text"
# We defined the test dataset to supply data for exacly one full iteration of the test dataset. We can fetch data until tensorflow lets us know there is no more data.
# + id="QMFducB2Tnbo" colab_type="code" outputId="91361058-e0bd-460a-8f2d-5c7eab0635f7" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1543883077854, "user_tz": -60, "elapsed": 530, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
with tf.Session() as session:
counter = 0
while True:
try:
test_inputs_np, test_labels_np = session.run(test_data)
counter += 1
except tf.errors.OutOfRangeError:
break
print('Counted {} batches of test examples.'.format(counter))
# + [markdown] id="M2nTc8Rxzytz" colab_type="text"
# The `make_one_shot_iterator()` function returns an iterator that, when exhausted, cannot be restarted.
# + [markdown] id="22XTfhplEd0s" colab_type="text"
# There are many utility functions in the `tf.data` both for reading in and manipulating data; chances are, whatever you would like to do it is already available there.
# + [markdown] id="8sUCWDkSrS2s" colab_type="text"
# ### Queues
#
# In earlier versions of tensorflow datasets had to be manipulated with so called [Queues](https://www.tensorflow.org/api_guides/python/threading_and_queues). They allowed data loading and preprocessing to be asynchronous, making the input pipeline faster. Their use for input pipelines is now deprecated, if you are interested in increasing the performance of your input pipeline read the [official guide on this topic](https://www.tensorflow.org/performance/datasets_performance).
#
# Queues are still used for pushing data between different threads, potentially on different machines, but we will not cover them in this lab.
#
# + [markdown] id="BtoJu2JQufKa" colab_type="text"
# ## The Power of Sonnet
#
# The Sonnet library has two key selling points:
#
# * Complex networks are easily reused.
# * Variable sharing is handled transparently by automatically reusing variables on subsequent calls to the same module.
#
# We will now see these features in action.
#
# + [markdown] id="1WJZFNUPJfCW" colab_type="text"
# We start by defining a sonnet module corresponding to the classifier we have been working with. The section on [defining your own submodules](https://deepmind.github.io/sonnet/#defining-your-own-modules) in the sonnet documentation is both helpful and precise. The key points are:
#
# * Inherit from snt.AbstractModule
# * Call superclass constructor
# * Implement the `_build()` method
#
# The `_build()` method is meant to construct all computation graph corresponding to this module. It takes as argument the inputs to the module, and returns the outputs.
#
#
# + id="YlUQsYkdq1Om" colab_type="code" colab={}
class MySimpleModule(snt.AbstractModule):
def __init__(self, num_hidden, nonlinearity=tf.nn.relu,
name="my_simple_module"):
super(MySimpleModule, self).__init__(name=name)
self._num_hidden = num_hidden
self._nonlinearity = nonlinearity
def _build(self, inputs):
# Inputs has shape batch_size x ?.
pre_activations = snt.Linear(output_size=self._num_hidden)(inputs)
activations = self._nonlinearity(pre_activations)
outputs = snt.Linear(output_size=1)(activations)
return tf.squeeze(outputs, axis=-1) # Shape: [batch_size].
# + [markdown] id="c02sphDjlqkB" colab_type="text"
# Aside: since this module is simply a sequence of other modules and tensorflow ops (e.g. the non-linearity), the module could have been made using the `snt.Sequential()` wrapper.
# + [markdown] id="NArC4JAtOl7I" colab_type="text"
# We can make a particular instance of the module we defined like so:
# + id="NECiDI2bOnRc" colab_type="code" colab={}
tf.reset_default_graph()
model = MySimpleModule(num_hidden=5)
# + id="xa8C-nTsmntL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="dc57c42a-0b27-4970-d0fe-4e9562786f7c" executionInfo={"status": "ok", "timestamp": 1543883098521, "user_tz": -60, "elapsed": 661, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
model
# + [markdown] id="IcYsy2PDMZRQ" colab_type="text"
# No graph has actually been created so far, since only the constructor of the class ran. Let's connect this module to the training data.
#
# *Note that while it is encouraged to only create graph in the `_build()` method, some sonnet modules may already do so in their constructor.*
# + id="bQGjSNnGObv2" colab_type="code" colab={}
train_dataset, test_dataset = get_tf_dataset()
train_inputs, train_labels = train_dataset.make_one_shot_iterator().get_next()
train_ouputs = model(train_inputs)
# + [markdown] id="_6TgD66tP3Zb" colab_type="text"
# The connection triggered the `_build()` function and we can see the graph corresponding to the model is built.
# + id="M5Fa4BhDP3fQ" colab_type="code" colab={}
# show_graph()
# + [markdown] id="X1y4JiTqJcvl" colab_type="text"
# The beauty of sonnet is that we can **connect the same `model` instance to the test data tensor and it will automatically share variables**.
# + id="0cqTPzOHRDpO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 74} outputId="d701b44d-626a-44d7-f809-f7ff0bf7fa4e" executionInfo={"status": "ok", "timestamp": 1543883189055, "user_tz": -60, "elapsed": 517, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
test_inputs, test_labels = test_dataset.make_one_shot_iterator().get_next()
test_outputs = model(test_inputs)
# + [markdown] id="Pr39gvUaX8O1" colab_type="text"
# Of course creating another instance will not share variables. Can you tell, based on the graph (not considering the names) which modules share weights?
# + id="oWSEVHdkYB8T" colab_type="code" colab={}
unshared_test_outputs = MySimpleModule(num_hidden=5, name='unshared_simple_module')(test_inputs)
# show_graph()
# + [markdown] id="lAdu7dhvRDxb" colab_type="text"
# The fact that `train_outputs` and `test_outputs` use shared variables means that training based on `train_outputs` will improve the quality of `test_ouputs` as well. We show this next.
#
# We base the training script here based on our previous one. Some modifications are required:
#
# * The references to the dataset must be updated. We do not use `feed_dicts`, but we must take care to run `test_outputs` or `train_outputs`.
# * In order to get the true (test) labels, we need to run the `test_labels` tensor.
# * We need to iterate over the full test dataset.
#
# Another change is that now each training step uses a different batch of data, while our earlier version used the full (smaller) dataset.
#
# + id="P4qhLEYPJc4r" colab_type="code" colab={}
# CHANGED HERE:
loss = tf.reduce_mean(tf.squared_difference(train_labels, train_ouputs))
# Optimizer and initializer.
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.03)
train_op = optimizer.minimize(loss)
init_op = tf.global_variables_initializer()
# + id="AjmiYrqOq1W1" colab_type="code" outputId="33776adf-9a0a-4c41-f1c2-ecea4a7bf661" colab={"base_uri": "https://localhost:8080/", "height": 441} executionInfo={"status": "ok", "timestamp": 1543883260476, "user_tz": -60, "elapsed": 29973, "user": {"displayName": "Luk\u00e1\u0161 Mart\u00e1k", "photoUrl": "https://lh5.googleusercontent.com/-Cf7M0y9YIEc/AAAAAAAAAAI/AAAAAAAAAFg/Mii_7w1relg/s64/photo.jpg", "userId": "04521090243879256607"}}
TRAINING_STEPS = int(2e4)
RECORD_PERIOD = int(1e3)
def _num_correct(predictions_np, true_labels_np):
# Actual label predictions given as {-1, +1}.
predictions_np[predictions_np <= 0] = -1
predictions_np[predictions_np > 0] = 1
# Count correct predictions.
return np.count_nonzero(np.isclose(predictions_np, true_labels_np))
print('Losses:')
with tf.Session() as session:
session.run(init_op) # Initializes the weights in the network.
for i in range(TRAINING_STEPS):
_, loss_np = session.run([train_op, loss]) # CHANGED HERE.
if (i % RECORD_PERIOD) == 0:
print(' ', loss_np)
print()
# The model is ready to be evaluated. Fetch the predicted outputs.
num_correct = 0
num_elements = 0
while True:
try:
# CHANGES HERE.
predictions_np, true_labels_np = session.run([test_outputs, test_labels])
num_elements += predictions_np.shape[0]
num_correct += _num_correct(predictions_np, true_labels_np)
except tf.errors.OutOfRangeError:
break
print('The prediction error on the test set:',
(num_elements - num_correct) / num_elements)
# + [markdown] id="PorU6KbWWxg0" colab_type="text"
# We will see another convenient feature of Sonnet when working with generative models in the VAE and GAN lab.
# + [markdown] id="0jXX5yAJrIil" colab_type="text"
# ## Debugging Tensorflow
#
# Debugging tensorflow code and models can be challenging when compared to debugging 1) simple python code or even 2) other machine learning code. This is due to the separate building and running phases* of tensorflow:
#
# * You cannot simply just stop the computation midway in a `run()` call and inspect what is going on. **
# * If an error is only revealed in a `session.run()` call, Tensorflow may often be unable to point you to the python code that generated the offending operation.
# * Race conditions may occur. These can be hard to detect because the race condition may only occur very very infrequently.
#
# In this section we list some practical advice to debugging tensorflow.
#
# <small>**Tensorflow's Eager mode removes this separation, making debugging simpler.</small><br />
# <small>****There is a [tensorflow debugger](https://www.tensorflow.org/programmers_guide/debugger) that tries to address this problem.*</small>
#
#
# + [markdown] id="_CiUuqj3rH5I" colab_type="text"
# * **Check your shapes**. It is possible that something is not of the shape you expect, but due to broadcasting the graph still computes something -- but not what you want.
# * **Check the graph with tensorboard**. Does it do what you wanted it to?
# * **Print and/or assert values of tensors**. While you cannot stop your graph mid-computation, you can print the values going through them. Unfortunately this [does not currently work](https://www.tensorflow.org/api_docs/python/tf/Print) in notebooks.
# + [markdown] id="NTvYunx7-J7u" colab_type="text"
# ## Not covered: Control Flow
#
# In tensorflow you can define logical operations such as conditionals, loops, etc. In fact, Tensorflow is Turing-complete. We do not cover them as these operations are not usually required for training neural nets, and it is better to avoid them unless really needed due their added compexity.
#
# + id="1pNx0PCN7o0M" colab_type="code" colab={}
| introductory/Intro_Tensorflow_and_Sonnet.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="18AF5Ab4p6VL"
# **Copyright 2018 Google LLC.**
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + [markdown] id="crfqaJOyp8bq"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="B_XlLLpcWjkA"
# # Training a Simple Neural Network, with tensorflow/datasets Data Loading
#
# [](https://colab.research.google.com/github/google/jax/blob/master/docs/notebooks/neural_network_with_tfds_data.ipynb)
#
# _Forked from_ `neural_network_and_data_loading.ipynb`
#
# 
#
# Let's combine everything we showed in the [quickstart notebook](https://colab.research.google.com/github/google/jax/blob/master/notebooks/quickstart.ipynb) to train a simple neural network. We will first specify and train a simple MLP on MNIST using JAX for the computation. We will use `tensorflow/datasets` data loading API to load images and labels (because it's pretty great, and the world doesn't need yet another data loading library :P).
#
# Of course, you can use JAX with any API that is compatible with NumPy to make specifying the model a bit more plug-and-play. Here, just for explanatory purposes, we won't use any neural network libraries or special APIs for builidng our model.
# + id="OksHydJDtbbI"
import jax.numpy as jnp
from jax import grad, jit, vmap
from jax import random
# + [markdown] id="MTVcKi-ZYB3R"
# ## Hyperparameters
# Let's get a few bookkeeping items out of the way.
# + id="-fmWA06xYE7d" outputId="520e5fd5-97c4-43eb-ef0e-b714d5287689"
# A helper function to randomly initialize weights and biases
# for a dense neural network layer
def random_layer_params(m, n, key, scale=1e-2):
w_key, b_key = random.split(key)
return scale * random.normal(w_key, (n, m)), scale * random.normal(b_key, (n,))
# Initialize all layers for a fully-connected neural network with sizes "sizes"
def init_network_params(sizes, key):
keys = random.split(key, len(sizes))
return [random_layer_params(m, n, k) for m, n, k in zip(sizes[:-1], sizes[1:], keys)]
layer_sizes = [784, 512, 512, 10]
param_scale = 0.1
step_size = 0.01
num_epochs = 10
batch_size = 128
n_targets = 10
params = init_network_params(layer_sizes, random.PRNGKey(0))
# + [markdown] id="BtoNk_yxWtIw"
# ## Auto-batching predictions
#
# Let us first define our prediction function. Note that we're defining this for a _single_ image example. We're going to use JAX's `vmap` function to automatically handle mini-batches, with no performance penalty.
# + id="7APc6tD7TiuZ"
from jax.scipy.special import logsumexp
def relu(x):
return jnp.maximum(0, x)
def predict(params, image):
# per-example predictions
activations = image
for w, b in params[:-1]:
outputs = jnp.dot(w, activations) + b
activations = relu(outputs)
final_w, final_b = params[-1]
logits = jnp.dot(final_w, activations) + final_b
return logits - logsumexp(logits)
# + [markdown] id="dRW_TvCTWgaP"
# Let's check that our prediction function only works on single images.
# + id="4sW2A5mnXHc5" outputId="ce9d86ed-a830-4832-e04d-10d1abb1fb8a"
# This works on single examples
random_flattened_image = random.normal(random.PRNGKey(1), (28 * 28,))
preds = predict(params, random_flattened_image)
print(preds.shape)
# + id="PpyQxuedXfhp" outputId="f43bbc9d-bc8f-4168-ee7b-79ee9d33f245"
# Doesn't work with a batch
random_flattened_images = random.normal(random.PRNGKey(1), (10, 28 * 28))
try:
preds = predict(params, random_flattened_images)
except TypeError:
print('Invalid shapes!')
# + id="oJOOncKMXbwK" outputId="fa380024-aaf8-4789-d3a2-f060134930e6"
# Let's upgrade it to handle batches using `vmap`
# Make a batched version of the `predict` function
batched_predict = vmap(predict, in_axes=(None, 0))
# `batched_predict` has the same call signature as `predict`
batched_preds = batched_predict(params, random_flattened_images)
print(batched_preds.shape)
# + [markdown] id="elsG6nX03BvW"
# At this point, we have all the ingredients we need to define our neural network and train it. We've built an auto-batched version of `predict`, which we should be able to use in a loss function. We should be able to use `grad` to take the derivative of the loss with respect to the neural network parameters. Last, we should be able to use `jit` to speed up everything.
# + [markdown] id="NwDuFqc9X7ER"
# ## Utility and loss functions
# + id="6lTI6I4lWdh5"
def one_hot(x, k, dtype=jnp.float32):
"""Create a one-hot encoding of x of size k."""
return jnp.array(x[:, None] == jnp.arange(k), dtype)
def accuracy(params, images, targets):
target_class = jnp.argmax(targets, axis=1)
predicted_class = jnp.argmax(batched_predict(params, images), axis=1)
return jnp.mean(predicted_class == target_class)
def loss(params, images, targets):
preds = batched_predict(params, images)
return -jnp.mean(preds * targets)
@jit
def update(params, x, y):
grads = grad(loss)(params, x, y)
return [(w - step_size * dw, b - step_size * db)
for (w, b), (dw, db) in zip(params, grads)]
# + [markdown] id="umJJGZCC2oKl"
# ## Data Loading with `tensorflow/datasets`
#
# JAX is laser-focused on program transformations and accelerator-backed NumPy, so we don't include data loading or munging in the JAX library. There are already a lot of great data loaders out there, so let's just use them instead of reinventing anything. We'll use the `tensorflow/datasets` data loader.
# + id="uWvo1EgZCvnK"
import tensorflow_datasets as tfds
data_dir = '/tmp/tfds'
# Fetch full datasets for evaluation
# tfds.load returns tf.Tensors (or tf.data.Datasets if batch_size != -1)
# You can convert them to NumPy arrays (or iterables of NumPy arrays) with tfds.dataset_as_numpy
mnist_data, info = tfds.load(name="mnist", batch_size=-1, data_dir=data_dir, with_info=True)
mnist_data = tfds.as_numpy(mnist_data)
train_data, test_data = mnist_data['train'], mnist_data['test']
num_labels = info.features['label'].num_classes
h, w, c = info.features['image'].shape
num_pixels = h * w * c
# Full train set
train_images, train_labels = train_data['image'], train_data['label']
train_images = jnp.reshape(train_images, (len(train_images), num_pixels))
train_labels = one_hot(train_labels, num_labels)
# Full test set
test_images, test_labels = test_data['image'], test_data['label']
test_images = jnp.reshape(test_images, (len(test_images), num_pixels))
test_labels = one_hot(test_labels, num_labels)
# + id="7VMSC03gCvnO" outputId="e565586e-d598-4fa1-dd6f-10ba39617f6a"
print('Train:', train_images.shape, train_labels.shape)
print('Test:', test_images.shape, test_labels.shape)
# + [markdown] id="xxPd6Qw3Z98v"
# ## Training Loop
# + id="X2DnZo3iYj18" outputId="bad334e0-127a-40fe-ec21-b0db77c73088"
import time
def get_train_batches():
# as_supervised=True gives us the (image, label) as a tuple instead of a dict
ds = tfds.load(name='mnist', split='train', as_supervised=True, data_dir=data_dir)
# You can build up an arbitrary tf.data input pipeline
ds = ds.batch(batch_size).prefetch(1)
# tfds.dataset_as_numpy converts the tf.data.Dataset into an iterable of NumPy arrays
return tfds.as_numpy(ds)
for epoch in range(num_epochs):
start_time = time.time()
for x, y in get_train_batches():
x = jnp.reshape(x, (len(x), num_pixels))
y = one_hot(y, num_labels)
params = update(params, x, y)
epoch_time = time.time() - start_time
train_acc = accuracy(params, train_images, train_labels)
test_acc = accuracy(params, test_images, test_labels)
print("Epoch {} in {:0.2f} sec".format(epoch, epoch_time))
print("Training set accuracy {}".format(train_acc))
print("Test set accuracy {}".format(test_acc))
# + [markdown] id="xC1CMcVNYwxm"
# We've now used the whole of the JAX API: `grad` for derivatives, `jit` for speedups and `vmap` for auto-vectorization.
# We used NumPy to specify all of our computation, and borrowed the great data loaders from `tensorflow/datasets`, and ran the whole thing on the GPU.
| docs/notebooks/neural_network_with_tfds_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Forecasting Traffic Volume (고속도로 교통량 예측)
# 이 소스는 https://github.com/chrisking/ForecastPOC.git 를 원본으로 하고 있습니다.
#
# - 미국 미네라폴리스 근처 고속도로의 차량 통행량을 시간별로 예측하는 문제를 풀어가는 과정을 아래 4개의 노트북으로 구성 함.
# - 이 사례에 대한 분석 보고서는 아래 PDF를 참고 하세요.
# - [Forecasting_Traffic_Volume_Model_Analysis](20200313_Forecasting_Traffic_Volume_Model_Analysis_Gonsoo.pdf)
#
# ## 실행 단계
# 1.0~4.0번까지의 노트북을 실행하시면 됩니다.
#
# **옵션으로서 1.0 을 통해서 데이터에 대한 이해를 높이면 좋습니다.**
#
# - 1.0 Validating_and_Importing_Target_Time_Series_Data
# - Target Data 를 만들고 data import 합니다.
# - 2.0.Creating_and_Evaluating_Predictors.ipynb
# - Predictor, Forecast를 생성하고 평가 합니다.
# - 3.0.Validating_and_Importing_Related_Time_Series_Data.ipynb
# - Target, Related 데이타 셋을 준비하고 Data Import 합니다.
# - 4.0.Creating_and_Evaluating_Related_Time_Predictors.ipynb
# - Predictor, Forecast를 생성하고 평가 합니다.
#
# 위의 노트북은 아래와 같은 기술적 부분을 가지고 있습니다.
#
# * Compare Target Data vs. Target + Related Data
# * Compare ARIMA, Prophet and DeepARP performance with Acutal value
#
#
| TrafficVolume/README.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PROJ_irox_oer] *
# language: python
# name: conda-env-PROJ_irox_oer-py
# ---
# # Import Modules
# +
import os
print(os.getcwd())
import sys
from pathlib import Path
import numpy as np
import pandas as pd
import plotly.graph_objs as go
import chart_studio.plotly as py
# #########################################################
from vasp.vasp_methods import parse_incar
# +
compenv = os.environ["COMPENV"]
vasp_dir = "."
# For testing purposes
if compenv == "wsl":
vasp_dir = os.path.join(
os.environ["PROJ_irox_oer"],
"__test__/anal_job_out")
# +
from vasp.parse_oszicar import parse_oszicar
out_dict = parse_oszicar(vasp_dir=vasp_dir)
ion_step_conv_dict = out_dict["ion_step_conv_dict"]
N_tot = out_dict["N_tot"]
# +
path_i = os.path.join(
vasp_dir,
"INCAR")
my_file = Path(path_i)
if my_file.is_file():
with open(path_i, "r") as f:
incar_lines = f.read().splitlines()
incar_dict = parse_incar(incar_lines)
nelm_i = incar_dict["NELM"]
# +
# y_plot_quant = "E"
# y_plot_quant = "dE"
y_plot_quant = "dE_abs"
# spacing = N_tot / 15
# spacing = N_tot / 10
spacing = int(N_tot / 5)
# -
data = []
x_axis_cum = 0
for i_cnt, ion_step_i in enumerate(list(ion_step_conv_dict.keys())):
# if i_cnt == 2:
# break
df_i = ion_step_conv_dict[ion_step_i]
num_N_i = df_i.N.max() - df_i.N.min()
print("")
print("num_N_i:", num_N_i)
extra_spacing = nelm_i - num_N_i - spacing
print("extra_spacing:", extra_spacing)
df_i["below_0"] = df_i.dE < 0.
df_i["dE_abs"] = np.abs(df_i["dE"])
# x_array = df_i.N + x_axis_cum + 50
x_array = df_i.N + x_axis_cum
# y_array = df_i.E
y_array = df_i[y_plot_quant]
color_array = df_i["below_0"]
color_array_2 = []
for i in color_array:
if i:
color_array_2.append("red")
else:
color_array_2.append("black")
num_N = df_i.N.max()
# x_axis_cum += num_N + 100
# x_axis_cum += num_N + spacing + extra_spacing
x_axis_cum += num_N + (nelm_i - num_N_i) + spacing
# #####################################################
trace_i = go.Scatter(
x=x_array,
y=y_array,
mode="markers",
opacity=0.8,
marker_color=color_array_2,
)
data.append(trace_i)
# #####################################################
trace_i = go.Scatter(
x=2 * [x_array[0] + nelm_i],
y=[1e-10, 1e10],
mode="lines",
line_color="grey",
# opacity=0.8,
# marker_color=color_array_2,
)
data.append(trace_i)
# +
max_list = []
min_list = []
for i_cnt, ion_step_i in enumerate(list(ion_step_conv_dict.keys())):
df_i = ion_step_conv_dict[ion_step_i]
max_dE = df_i.dE_abs.max()
min_dE = df_i.dE_abs.min()
# print("")
# print("max_dE:", max_dE)
# print("min_dE:", min_dE)
max_list.append(max_dE)
min_list.append(min_dE)
max_y = np.max(max_list)
min_y = np.min(min_list)
# +
num_N_i = df_i.N.max() - df_i.N.min()
print("num_N_i:", num_N_i)
extra_spacing = nelm_i - num_N_i - spacing
print("extra_spacing:", extra_spacing)
# +
# assert False
# +
fig = go.Figure(data=data)
fig.update_layout(
title=os.getcwd(),
xaxis=go.layout.XAxis(
title="dE",
),
yaxis=go.layout.YAxis(
title="N",
type="log",
# range=[min_y, max_y],
range=[-7, 6],
),
# xaxis_type="log",
# yaxis_type="log",
)
if compenv == "wsl":
fig.show()
# +
from plotting.my_plotly import my_plotly_plot
if compenv != "wsl":
write_png = True
else:
write_png = False
my_plotly_plot(
figure=fig,
plot_name="scf_convergence",
write_html=True,
write_png=write_png,
# png_scale=6.0,
# write_pdf=False,
# write_svg=False,
try_orca_write=True,
)
# -
# # Copy figure html file to Dropbox with rclone
# +
rclone_comm = "rclone copy out_plot/scf_convergence.html " + os.environ["rclone_dropbox"] + ":__temp__/"
import subprocess
result = subprocess.run(
rclone_comm.split(" "),
stdout=subprocess.PIPE)
# -
print(40 * "*")
print("*** Script finished running " + 12 * "*")
print(40 * "*")
# + active=""
#
#
#
# + jupyter={}
# root_dir = "."
# # path_i = "./job.out"
# path_i = os.path.join(root_dir, "OSZICAR")
# compenv = os.environ["COMPENV"]
# if compenv == "wsl":
# root_dir = os.path.join(
# os.environ["PROJ_irox_oer"],
# "__test__/anal_job_out")
# # path_i = "./OSZICAR"
# # path_i = "./OSZICAR.new"
# path_i = os.path.join(
# root_dir,
# # os.environ["PROJ_irox_oer"],
# # "__test__/anal_job_out/OSZICAR.new",
# "OSZICAR",
# )
# with open(path_i, "r") as f:
# oszicar_lines = f.read().splitlines()
# from vasp.vasp_methods import parse_incar
# from pathlib import Path
# path_i = os.path.join(
# root_dir,
# "INCAR")
# my_file = Path(path_i)
# if my_file.is_file():
# with open(path_i, "r") as f:
# incar_lines = f.read().splitlines()
# incar_dict = parse_incar(incar_lines)
# nsw_i = incar_dict["NSW"]
# nelm_i = incar_dict["NELM"]
# incar_parsed = True
# else:
# incar_parsed = False
# line_beginnings = ["DAV:", "RMM:", ]
# lines_groups = []
# group_lines_i = []
# for line_i in oszicar_lines:
# if line_i[0:4] in line_beginnings:
# group_lines_i.append(line_i)
# if "F= " in line_i:
# # print("IDJIFSD")
# lines_groups.append(group_lines_i)
# group_lines_i = []
# # This should add the final group_lines in the case that it hasn't finished yet
# if "F= " not in oszicar_lines[-1]:
# lines_groups.append(group_lines_i)
# N_tot = 0.
# ion_step_conv_dict = dict()
# for ion_step_i, lines_group_i in enumerate(lines_groups):
# data_dict_list = []
# for line_i in lines_group_i:
# data_dict_i = dict()
# line_list_i = [i for i in line_i.split(" ") if i != ""]
# N_i = line_list_i[1]
# data_dict_i["N"] = int(N_i)
# E_i = line_list_i[2]
# data_dict_i["E"] = float(E_i)
# dE_i = line_list_i[3]
# data_dict_i["dE"] = float(dE_i)
# d_eps_i = line_list_i[4]
# data_dict_i["d_eps"] = float(d_eps_i)
# ncg_i = line_list_i[5]
# data_dict_i["ncg"] = int(ncg_i)
# rms_i = line_list_i[6]
# data_dict_i["rms"] = float(rms_i)
# if len(line_list_i) > 7:
# rms_c_i = line_list_i[7]
# data_dict_i["rms_c"] = float(rms_c_i)
# # #################################################
# data_dict_list.append(data_dict_i)
# df_i = pd.DataFrame(data_dict_list)
# # print(N_tot)
# N_tot += df_i.N.max()
# ion_step_conv_dict[ion_step_i] = df_i
# print("N_tot", N_tot)
| dft_workflow/bin/anal_job_out.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # imshow demonstration
# - This notebook is part of biokit
# - https://github.com/biokit/biokit
# - https://pypi.python.org/pypi/biokit
#
# - imshow with pandas dataframe : imshow
# %pylab inline
from biokit.viz import Imshow
# # Imshow
# - we overwrite the pylab.imshow to change the default behaviour (no interpolation, hot cmap)
# - input can be a dataframe, in which case, X and Y ticks are filled automatically
data = numpy.random.randn(10,10)
imshow(data) # this is the default behaviour in matplotlib
im = Imshow(data)
im.plot(cmap='jet')
import pandas as pd
df = pd.DataFrame({'A':[1,2], 'B':[3,4]}, index=['X','Y'])
df
# by default xticks are rotated by 90 degrees since names could be long
# but in this example names are short, so we reset the rotation.
Imshow(df).plot(rotation_x=0)
# + jupyter={"outputs_hidden": true}
# -
| notebooks/viz/imshow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #README
#
# This is our readme file about Data Curation. We are the Masters of the Universe (and you can, too...)
#
# [IMAGE HERE: show in browser.]
# [4 tabs: directory of files, json, open refine, and notebook with commands.]
# [Then end in image.]
#
# When is a toy not a toy? (When it is a record...: SHOW record here.)
#
# * First, you have to have a file structure...
#
# ###Exploration
#
# * "Mattel" at the V&A?: 2 commands plus json
# * This stage shows how to use a structured set of results from the V&A Museum that we obtained by using the V&A API to query the word "Mattel." Doing this returned this record structure as a json file.
#
# [show file]
#
# * code example: using the API
#
# Here, we're reading the json file with shell command "cat," then piping it into several instances of the tool "jq."
#
# The first command lists all the values of the object key records; the second command grabs all the keys.
#
# * record structure
#
# The value of "records" is an array. From there, we grab each item of the array, and each item of the array is an object. We want all the keys of the "fields" object, as well as the values of the "title" and "object."
#
#
# ###Evaluation
#
# * Open (formerly Google) Refine
# (plug json into Open Refine)
#
# * What this helps you do: get data organized (cluster and edit [on the "Artist" field], plus facet, filter, etc...)
#
# ###Design
#
# * Which fields don't you need? (Museum-specific ones: "Last checked," etc...)
# * This data comes from the V&A: when you import other people's data, you get more than you need. (Usually.)
# * What do you -- or your users -- care about? (Who's your data audience?)
#
# ###Transformation
#
# * Obtained as json files
# * Linking (values of) entities to authority files (is Battle Cat in VIAF? Probably not, but you can look elsewhere...)
# * linking "Mattel"
# * "title" + "object": make a google query to make a large, merged image group
# * explaining how we did this is important: not just a Google search
#
# ###Documentation
#
# * Do this. Often. Redundantly. Again.
# * README.txt files
# * "data dictionaries": why you want them
#
# ###Sharing (yes, you want to...)
#
# * GenCon!
#
| dhcuration.README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/microprediction/timemachines/blob/main/CompareToNaive.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="3_J2q5zchh4k"
# !pip install timemachines
# !pip install --upgrade statsmodels
# + [markdown] id="KRKKP5HMlaM5"
# # Example of comparing against naive forecast
# For a given univariate time-series y, this runs checks against some time-series models in the timemachines library and reports any that beat the naive forecast. We proceed from models that can generate forecasts quickly to others that are more painful. First a quick plot...
# + id="RhgSyaQFqaEG"
import numpy as np
y = np.cumsum(np.random.randn(1000)) # <--- Swap this out for your time series
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="wryoJjfUhn1g" outputId="a281eb86-2b0d-4b5b-de59-d4f7dca0b4b8"
from timemachines.skatertools.visualization.priorplot import prior_plot
import matplotlib.pyplot as plt
from timemachines.skaters.simple.movingaverage import EMA_SKATERS
# Plot an example
f = EMA_SKATERS[0]
k = 1
prior_plot(f=f, k=k, y=y, n=450, n_plot=50)
plt.show()
# + id="2aIznKUXjBko"
threshold = 0.95 # What constitutes an interesting error, relative to naive
from pprint import pprint
from timemachines.skatertools.evaluation.evaluators import evaluate_mean_squared_error
from timemachines.skaters.simple.movingaverage import empirical_last_value
def report(y,k=1,models=EMA_SKATERS,n_burn=25, threshold=0.98):
"""
k - how many step to foreast ahead
threshold - fraction of naive forecast error that would constitute an interesting finding
n_burn - number of points to toss prior to error evaluation, to allow model to warm up
"""
lv_err = evaluate_mean_squared_error(f=empirical_last_value, y=y, k=k, a=None, t=None, e=None, r=None, n_burn=n_burn)
interesting = dict()
for f in models:
err = evaluate_mean_squared_error(f=f, y=y, k=1, a=None, t=None, e=None, r=None, n_burn=25)
if err<threshold*lv_err:
print(f.__name__+' error is '+str(err)+" versus "+str(lv_err)+' for naive forecast ')
interesting[f.__name__] = err
return interesting
# + [markdown] id="_TGJsPLLm3Bu"
# # Fast models....
# + colab={"base_uri": "https://localhost:8080/"} id="B-JdmJHfmQtF" outputId="c025934c-cba1-4e48-a1c1-e27649a48d12"
from timemachines.skaters.simple.thinking import THINKING_SKATERS
from timemachines.skaters.simple.hypocraticensemble import HYPOCRATIC_ENSEMBLE_SKATERS
FAST_MODELS = EMA_SKATERS + THINKING_SKATERS+HYPOCRATIC_ENSEMBLE_SKATERS
report(y=y,k=k,models=EMA_SKATERS + THINKING_SKATERS+HYPOCRATIC_ENSEMBLE_SKATERS)
# + [markdown] id="2sc5jAxom7Ii"
# # Slow models
# + id="8U568oIwmznt"
from timemachines.skaters.tsa.alltsaskaters import TSA_SKATERS # might also consider DLM
SLOW_MODELS = TSA_SKATERS
report(y=y,k=k,models=SLOW_MODELS)
# + [markdown] id="XgSxmaDlnFRh"
# # Interminable models
# This is here really to make the point that some models are intended for one-off forecasting on a historical data set. Don't hold your breath waiting for this to run.
# + id="vRn6MzAInMJ7"
from timemachines.skaters.prop.prophetskaterssingular import fbprophet_univariate
from timemachines.skaters.nproph.nprophetskaters import NPROPHET_UNIVARIATE_SKATERS
INTERMINABLE_MODELS = [fbprophet_univariate] + NPROPHET_UNIVARIATE_SKATERS
report(y=y,k=k,models=INTERMINABLE_MODELS)
| CompareToNaive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="k0IFBGCx8_im"
# # Recommendation Systems with TensorFlow
#
# + [markdown] colab_type="text" id="Le0Z54X68_iq"
# # Introduction
#
# In this lab, we will create a movie recommendation system based on the [MovieLens](https://movielens.org/) dataset available [here](http://grouplens.org/datasets/movielens/). The data consists of movies ratings (on a scale of 1 to 5).
# Specifically, we'll be using matrix factorization to learn user and movie embeddings. Concepts highlighted here are also available in the course on [Recommendation Systems](https://developers.google.com/machine-learning/recommendation/).
#
# ## Objectives
# 1. Explore the MovieLens Data
# 1. Train a matrix factorization model
# 1. Inspect the Embeddings
# 1. Perform Softmax model training
#
# -
# Ensure the right version of Tensorflow is installed.
# !pip freeze | grep tensorflow==2.6
# + cellView="both" colab={} colab_type="code" id="8vRUh2Mzo4s1"
from __future__ import print_function
import numpy as np
import pandas as pd
import collections
from mpl_toolkits.mplot3d import Axes3D
from IPython import display
from matplotlib import pyplot as plt
import sklearn
import sklearn.manifold
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
# Add some convenience functions to Pandas DataFrame.
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.3f}'.format
def mask(df, key, function):
"""Returns a filtered dataframe, by applying function to key"""
return df[function(df[key])]
def flatten_cols(df):
df.columns = [' '.join(col).strip() for col in df.columns.values]
return df
pd.DataFrame.mask = mask
pd.DataFrame.flatten_cols = flatten_cols
# +
#Let's install Altair for interactive visualizations
# !pip install git+git://github.com/altair-viz/altair.git
import altair as alt
alt.data_transformers.enable('default', max_rows=None)
#alt.renderers.enable('colab')
# + [markdown] colab_type="text" id="KhAVFaqCo4s5"
# We then download the MovieLens Data, and create DataFrames containing movies, users, and ratings.
# + cellView="both" colab={} colab_type="code" id="O3bcgduFo4s6"
# Download MovieLens data.
print("Downloading movielens data...")
from urllib.request import urlretrieve
import zipfile
urlretrieve("http://files.grouplens.org/datasets/movielens/ml-100k.zip", "movielens.zip")
zip_ref = zipfile.ZipFile('movielens.zip', "r")
zip_ref.extractall()
print("Done. Dataset contains:")
print(zip_ref.read('ml-100k/u.info'))
# +
# Load each data set (users, ratings, and movies).
users_cols = ['user_id', 'age', 'sex', 'occupation', 'zip_code']
users = pd.read_csv(
'ml-100k/u.user', sep='|', names=users_cols, encoding='latin-1')
ratings_cols = ['user_id', 'movie_id', 'rating', 'unix_timestamp']
ratings = pd.read_csv(
'ml-100k/u.data', sep='\t', names=ratings_cols, encoding='latin-1')
# The movies file contains a binary feature for each genre.
genre_cols = [
"genre_unknown", "Action", "Adventure", "Animation", "Children", "Comedy",
"Crime", "Documentary", "Drama", "Fantasy", "Film-Noir", "Horror",
"Musical", "Mystery", "Romance", "Sci-Fi", "Thriller", "War", "Western"
]
movies_cols = [
'movie_id', 'title', 'release_date', "video_release_date", "imdb_url"
] + genre_cols
movies = pd.read_csv(
'ml-100k/u.item', sep='|', names=movies_cols, encoding='latin-1')
# Since the ids start at 1, we shift them to start at 0. This will make handling of the
# indices easier later
users["user_id"] = users["user_id"].apply(lambda x: str(x-1))
movies["movie_id"] = movies["movie_id"].apply(lambda x: str(x-1))
movies["year"] = movies['release_date'].apply(lambda x: str(x).split('-')[-1])
ratings["movie_id"] = ratings["movie_id"].apply(lambda x: str(x-1))
ratings["user_id"] = ratings["user_id"].apply(lambda x: str(x-1))
ratings["rating"] = ratings["rating"].apply(lambda x: float(x))
# +
# Compute the number of movies to which a genre is assigned.
genre_occurences = movies[genre_cols].sum().to_dict()
# Since some movies can belong to more than one genre, we create different
# 'genre' columns as follows:
# - all_genres: all the active genres of the movie.
# - genre: randomly sampled from the active genres.
def mark_genres(movies, genres):
def get_random_genre(gs):
active = [genre for genre, g in zip(genres, gs) if g==1]
if len(active) == 0:
return 'Other'
return np.random.choice(active)
def get_all_genres(gs):
active = [genre for genre, g in zip(genres, gs) if g==1]
if len(active) == 0:
return 'Other'
return '-'.join(active)
movies['genre'] = [
get_random_genre(gs) for gs in zip(*[movies[genre] for genre in genres])]
movies['all_genres'] = [
get_all_genres(gs) for gs in zip(*[movies[genre] for genre in genres])]
mark_genres(movies, genre_cols)
# Create one merged DataFrame containing all the movielens data.
movielens = ratings.merge(movies, on='movie_id').merge(users, on='user_id')
# -
# Utility to split the data into training and test sets.
def split_dataframe(df, holdout_fraction=0.1):
"""Splits a DataFrame into training and test sets.
Args:
df: a dataframe.
holdout_fraction: fraction of dataframe rows to use in the test set.
Returns:
train: dataframe for training
test: dataframe for testing
"""
test = df.sample(frac=holdout_fraction, replace=False)
train = df[~df.index.isin(test.index)]
return train, test
# + [markdown] colab_type="text" id="bfhOILgfo4s8"
# # Exploring the Movielens Data
# Before we dive into model building, let's inspect our MovieLens dataset. It is usually helpful to understand the statistics of the dataset.
# + [markdown] colab_type="text" id="EoB8Drqwo4s-"
# ### Users
# We start by printing some basic statistics describing the numeric user features.
# + colab={} colab_type="code" id="k-IRqWato4s-"
users.describe()
# + [markdown] colab_type="text" id="h3nZ1ARho4tC"
# We can also print some basic statistics describing the categorical user features
# + colab={} colab_type="code" id="jFGxSmTGo4tD"
users.describe(include=[np.object])
# + [markdown] colab_type="text" id="CH2_gW3Oo4tF"
# We can also create histograms to further understand the distribution of the users. We use Altair to create an interactive chart.
# + cellView="both" colab={} colab_type="code" id="ffkqaK_zo4tF"
# The following functions are used to generate interactive Altair charts.
# We will display histograms of the data, sliced by a given attribute.
# Create filters to be used to slice the data.
occupation_filter = alt.selection_multi(fields=["occupation"])
occupation_chart = alt.Chart().mark_bar().encode(
x="count()",
y=alt.Y("occupation:N"),
color=alt.condition(
occupation_filter,
alt.Color("occupation:N", scale=alt.Scale(scheme='category20')),
alt.value("lightgray")),
).properties(width=300, height=300, selection=occupation_filter)
# A function that generates a histogram of filtered data.
def filtered_hist(field, label, filter):
"""Creates a layered chart of histograms.
The first layer (light gray) contains the histogram of the full data, and the
second contains the histogram of the filtered data.
Args:
field: the field for which to generate the histogram.
label: String label of the histogram.
filter: an alt.Selection object to be used to filter the data.
"""
base = alt.Chart().mark_bar().encode(
x=alt.X(field, bin=alt.Bin(maxbins=10), title=label),
y="count()",
).properties(
width=300,
)
return alt.layer(
base.transform_filter(filter),
base.encode(color=alt.value('lightgray'), opacity=alt.value(.7)),
).resolve_scale(y='independent')
# + [markdown] colab_type="text" id="D93Jn4Pao4tH"
# Next, we look at the distribution of ratings per user. Clicking on an occupation in the right chart will filter the data by that occupation. The corresponding histogram is shown in blue, and superimposed with the histogram for the whole data (in light gray). You can use SHIFT+click to select multiple subsets.
#
# What do you observe, and how might this affect the recommendations?
# + colab={} colab_type="code" id="x4ahcH2do4tI"
users_ratings = (
ratings
.groupby('user_id', as_index=False)
.agg({'rating': ['count', 'mean']})
.flatten_cols()
.merge(users, on='user_id')
)
# Create a chart for the count, and one for the mean.
alt.hconcat(
filtered_hist('rating count', '# ratings / user', occupation_filter),
filtered_hist('rating mean', 'mean user rating', occupation_filter),
occupation_chart,
data=users_ratings)
# + [markdown] colab_type="text" id="pmC6wykPo4tK"
# ### Movies
#
# It is also useful to look at information about the movies and their ratings.
# + colab={} colab_type="code" id="tCdPQRsso4tL"
movies_ratings = movies.merge(
ratings
.groupby('movie_id', as_index=False)
.agg({'rating': ['count', 'mean']})
.flatten_cols(),
on='movie_id')
genre_filter = alt.selection_multi(fields=['genre'])
genre_chart = alt.Chart().mark_bar().encode(
x="count()",
y=alt.Y('genre'),
color=alt.condition(
genre_filter,
alt.Color("genre:N"),
alt.value('lightgray'))
).properties(height=300, selection=genre_filter)
# + colab={} colab_type="code" id="NH9QQNr6o4tO"
(movies_ratings[['title', 'rating count', 'rating mean']]
.sort_values('rating count', ascending=False)
.head(10))
# + colab={} colab_type="code" id="xNb5Lz1oo4tP"
(movies_ratings[['title', 'rating count', 'rating mean']]
.mask('rating count', lambda x: x > 20)
.sort_values('rating mean', ascending=False)
.head(10))
# + [markdown] colab_type="text" id="kW_WRQyDo4tR"
# Finally, the last chart shows the distribution of the number of ratings and average rating.
# + colab={} colab_type="code" id="z8wxFfV8o4tR"
# Display the number of ratings and average rating per movie.
alt.hconcat(
filtered_hist('rating count', '# ratings / movie', genre_filter),
filtered_hist('rating mean', 'mean movie rating', genre_filter),
genre_chart,
data=movies_ratings)
# + [markdown] colab_type="text" id="WEhp-q31o4tS"
# # Preliminaries
#
# Our goal is to factorize the ratings matrix $A$ into the product of a user embedding matrix $U$ and movie embedding matrix $V$, such that $A \approx UV^\top$ with
# $U = \begin{bmatrix} u_{1} \\ \hline \vdots \\ \hline u_{N} \end{bmatrix}$ and
# $V = \begin{bmatrix} v_{1} \\ \hline \vdots \\ \hline v_{M} \end{bmatrix}$.
#
# Here
# - $N$ is the number of users,
# - $M$ is the number of movies,
# - $A_{ij}$ is the rating of the $j$th movies by the $i$th user,
# - each row $U_i$ is a $d$-dimensional vector (embedding) representing user $i$,
# - each rwo $V_j$ is a $d$-dimensional vector (embedding) representing movie $j$,
# - the prediction of the model for the $(i, j)$ pair is the dot product $\langle U_i, V_j \rangle$.
#
#
# + [markdown] colab_type="text" id="yMYZw_Yco4tT"
# ## Sparse Representation of the Rating Matrix
#
# The rating matrix could be very large and, in general, most of the entries are unobserved, since a given user will only rate a small subset of movies. For effcient representation, we will use a [tf.SparseTensor](https://www.tensorflow.org/api_docs/python/tf/SparseTensor). A `SparseTensor` uses three tensors to represent the matrix: `tf.SparseTensor(indices, values, dense_shape)` represents a tensor, where a value $A_{ij} = a$ is encoded by setting `indices[k] = [i, j]` and `values[k] = a`. The last tensor `dense_shape` is used to specify the shape of the full underlying matrix.
#
# #### Toy example
# Assume we have $2$ users and $4$ movies. Our toy ratings dataframe has three ratings,
#
# user\_id | movie\_id | rating
# --:|--:|--:
# 0 | 0 | 5.0
# 0 | 1 | 3.0
# 1 | 3 | 1.0
#
# The corresponding rating matrix is
#
# $$
# A =
# \begin{bmatrix}
# 5.0 & 3.0 & 0 & 0 \\
# 0 & 0 & 0 & 1.0
# \end{bmatrix}
# $$
#
# And the SparseTensor representation is,
# ```python
# SparseTensor(
# indices=[[0, 0], [0, 1], [1,3]],
# values=[5.0, 3.0, 1.0],
# dense_shape=[2, 4])
# ```
#
#
# + [markdown] colab_type="text" id="vBNijwXuo4tU"
# ### Exercise 1: Build a tf.SparseTensor representation of the Rating Matrix.
#
# In this exercise, we'll write a function that maps from our `ratings` DataFrame to a `tf.SparseTensor`.
#
# Hint: you can select the values of a given column of a Dataframe `df` using `df['column_name'].values`.
# + cellView="both" colab={} colab_type="code" id="lwgUGTSmo4tW"
#Solution
def build_rating_sparse_tensor(ratings_df):
"""
Args:
ratings_df: a pd.DataFrame with `user_id`, `movie_id` and `rating` columns.
Returns:
a tf.SparseTensor representing the ratings matrix.
"""
indices = ratings_df[['user_id', 'movie_id']].values
values = ratings_df['rating'].values
return tf.SparseTensor(
indices=indices,
values=values,
dense_shape=[users.shape[0], movies.shape[0]])
# + [markdown] colab_type="text" id="8k9CSOH5o4tX"
# ## Calculating the error
#
# The model approximates the ratings matrix $A$ by a low-rank product $UV^\top$. We need a way to measure the approximation error. We'll start by using the Mean Squared Error of observed entries only (we will revisit this later). It is defined as
#
# $$
# \begin{align*}
# \text{MSE}(A, UV^\top)
# &= \frac{1}{|\Omega|}\sum_{(i, j) \in\Omega}{( A_{ij} - (UV^\top)_{ij})^2} \\
# &= \frac{1}{|\Omega|}\sum_{(i, j) \in\Omega}{( A_{ij} - \langle U_i, V_j\rangle)^2}
# \end{align*}
# $$
# where $\Omega$ is the set of observed ratings, and $|\Omega|$ is the cardinality of $\Omega$.
#
#
# + [markdown] colab_type="text" id="KI1Ft-I7o4tX"
# ### Exercise 2: Mean Squared Error
#
# Write a TensorFlow function that takes a sparse rating matrix $A$ and the two embedding matrices $U, V$ and returns the mean squared error $\text{MSE}(A, UV^\top)$.
#
# Hints:
# * in this section, we only consider observed entries when calculating the loss.
# * a `SparseTensor` `sp_x` is a tuple of three Tensors: `sp_x.indices`, `sp_x.values` and `sp_x.dense_shape`.
# * you may find [`tf.gather_nd`](https://www.tensorflow.org/api_docs/python/tf/gather_nd) and [`tf.losses.mean_squared_error`](https://www.tensorflow.org/api_docs/python/tf/losses/mean_squared_error) helpful.
# + cellView="both" colab={} colab_type="code" id="p43zJDN7o4ta"
#Solution
def sparse_mean_square_error(sparse_ratings, user_embeddings, movie_embeddings):
"""
Args:
sparse_ratings: A SparseTensor rating matrix, of dense_shape [N, M]
user_embeddings: A dense Tensor U of shape [N, k] where k is the embedding
dimension, such that U_i is the embedding of user i.
movie_embeddings: A dense Tensor V of shape [M, k] where k is the embedding
dimension, such that V_j is the embedding of movie j.
Returns:
A scalar Tensor representing the MSE between the true ratings and the
model's predictions.
"""
predictions = tf.gather_nd(
tf.matmul(user_embeddings, movie_embeddings, transpose_b=True),
sparse_ratings.indices)
loss = tf.losses.mean_squared_error(sparse_ratings.values, predictions)
return loss
# + [markdown] colab_type="text" id="IrteGTFgo4td"
# Note: One approach is to compute the full prediction matrix $UV^\top$, then gather the entries corresponding to the observed pairs. The memory cost of this approach is $O(NM)$. For the MovieLens dataset, this is fine, as the dense $N \times M$ matrix is small enough to fit in memory ($N = 943$, $M = 1682$).
#
# Another approach (given in the alternate solution below) is to only gather the embeddings of the observed pairs, then compute their dot products. The memory cost is $O(|\Omega| d)$ where $d$ is the embedding dimension. In our case, $|\Omega| = 10^5$, and the embedding dimension is on the order of $10$, so the memory cost of both methods is comparable. But when the number of users or movies is much larger, the first approach becomes infeasible.
# + cellView="both" colab={} colab_type="code" id="6Z0m3dL88Uxd"
#Alternate Solution
def sparse_mean_square_error(sparse_ratings, user_embeddings, movie_embeddings):
"""
Args:
sparse_ratings: A SparseTensor rating matrix, of dense_shape [N, M]
user_embeddings: A dense Tensor U of shape [N, k] where k is the embedding
dimension, such that U_i is the embedding of user i.
movie_embeddings: A dense Tensor V of shape [M, k] where k is the embedding
dimension, such that V_j is the embedding of movie j.
Returns:
A scalar Tensor representing the MSE between the true ratings and the
model's predictions.
"""
predictions = tf.reduce_sum(
tf.gather(user_embeddings, sparse_ratings.indices[:, 0]) *
tf.gather(movie_embeddings, sparse_ratings.indices[:, 1]),
axis=1)
loss = tf.losses.mean_squared_error(sparse_ratings.values, predictions)
return loss
# + [markdown] colab_type="text" id="eSfW6SwIo4tk"
# # Training a Matrix Factorization model
#
# ## CFModel (Collaborative Filtering Model) helper class
# This is a simple class to train a matrix factorization model using stochastic gradient descent.
#
# The class constructor takes
# - the user embeddings U (a `tf.Variable`).
# - the movie embeddings V, (a `tf.Variable`).
# - a loss to optimize (a `tf.Tensor`).
# - an optional list of metrics dictionaries, each mapping a string (the name of the metric) to a tensor. These are evaluated and plotted during training (e.g. training error and test error).
#
# After training, one can access the trained embeddings using the `model.embeddings` dictionary.
#
# Example usage:
# ```
# U_var = ...
# V_var = ...
# loss = ...
# model = CFModel(U_var, V_var, loss)
# model.train(iterations=100, learning_rate=1.0)
# user_embeddings = model.embeddings['user_id']
# movie_embeddings = model.embeddings['movie_id']
# ```
#
# + cellView="both" colab={} colab_type="code" id="6NHoOwido4tk"
class CFModel(object):
"""Simple class that represents a collaborative filtering model"""
def __init__(self, embedding_vars, loss, metrics=None):
"""Initializes a CFModel.
Args:
embedding_vars: A dictionary of tf.Variables.
loss: A float Tensor. The loss to optimize.
metrics: optional list of dictionaries of Tensors. The metrics in each
dictionary will be plotted in a separate figure during training.
"""
self._embedding_vars = embedding_vars
self._loss = loss
self._metrics = metrics
self._embeddings = {k: None for k in embedding_vars}
self._session = None
@property
def embeddings(self):
"""The embeddings dictionary."""
return self._embeddings
def train(self, num_iterations=100, learning_rate=1.0, plot_results=True,
optimizer=tf.train.GradientDescentOptimizer):
"""Trains the model.
Args:
iterations: number of iterations to run.
learning_rate: optimizer learning rate.
plot_results: whether to plot the results at the end of training.
optimizer: the optimizer to use. Default to GradientDescentOptimizer.
Returns:
The metrics dictionary evaluated at the last iteration.
"""
with self._loss.graph.as_default():
opt = optimizer(learning_rate)
train_op = opt.minimize(self._loss)
local_init_op = tf.group(
tf.variables_initializer(opt.variables()),
tf.local_variables_initializer())
if self._session is None:
self._session = tf.Session()
with self._session.as_default():
self._session.run(tf.global_variables_initializer())
self._session.run(tf.tables_initializer())
#tf.train.start_queue_runners()
with self._session.as_default():
local_init_op.run()
iterations = []
metrics = self._metrics or ({},)
metrics_vals = [collections.defaultdict(list) for _ in self._metrics]
# Train and append results.
for i in range(num_iterations + 1):
_, results = self._session.run((train_op, metrics))
if (i % 10 == 0) or i == num_iterations:
print("\r iteration %d: " % i + ", ".join(
["%s=%f" % (k, v) for r in results for k, v in r.items()]),
end='')
iterations.append(i)
for metric_val, result in zip(metrics_vals, results):
for k, v in result.items():
metric_val[k].append(v)
for k, v in self._embedding_vars.items():
self._embeddings[k] = v.eval()
if plot_results:
# Plot the metrics.
num_subplots = len(metrics)+1
fig = plt.figure()
fig.set_size_inches(num_subplots*10, 8)
for i, metric_vals in enumerate(metrics_vals):
ax = fig.add_subplot(1, num_subplots, i+1)
for k, v in metric_vals.items():
ax.plot(iterations, v, label=k)
ax.set_xlim([1, num_iterations])
ax.legend()
return results
# + [markdown] colab_type="text" id="bKis0EyHo4tn"
# ### Exercise 3: Build a Matrix Factorization model and train it
#
# Using your `sparse_mean_square_error` function, write a function that builds a `CFModel` by creating the embedding variables and the train and test losses.
# + cellView="form" colab={} colab_type="code" id="M9RxIX_Oo4tp"
#Solution
def build_model(ratings, embedding_dim=3, init_stddev=1.):
"""
Args:
ratings: a DataFrame of the ratings
embedding_dim: the dimension of the embedding vectors.
init_stddev: float, the standard deviation of the random initial embeddings.
Returns:
model: a CFModel.
"""
# Split the ratings DataFrame into train and test.
train_ratings, test_ratings = split_dataframe(ratings)
# SparseTensor representation of the train and test datasets.
A_train = build_rating_sparse_tensor(train_ratings)
A_test = build_rating_sparse_tensor(test_ratings)
# Initialize the embeddings using a normal distribution.
U = tf.Variable(tf.random.normal(
[A_train.dense_shape[0], embedding_dim], stddev=init_stddev))
V = tf.Variable(tf.random.normal(
[A_train.dense_shape[1], embedding_dim], stddev=init_stddev))
train_loss = sparse_mean_square_error(A_train, U, V)
test_loss = sparse_mean_square_error(A_test, U, V)
metrics = {
'train_error': train_loss,
'test_error': test_loss
}
embeddings = {
"user_id": U,
"movie_id": V
}
return CFModel(embeddings, train_loss, [metrics])
# + [markdown] colab_type="text" id="mHMTVERDo4ts"
# Great, now it's time to train the model!
#
# Go ahead and run the next cell, trying different parameters (embedding dimension, learning rate, iterations). The training and test errors are plotted at the end of training. You can inspect these values to validate the hyper-parameters.
#
# Note: by calling `model.train` again, the model will continue training starting from the current values of the embeddings.
# + colab={} colab_type="code" id="_BlRIQJYo4tt"
# Build the CF model and train it.
model = build_model(ratings, embedding_dim=30, init_stddev=0.5)
# -
model.train(num_iterations=1000, learning_rate=10.)
# + [markdown] colab_type="text" id="-RWinHdno4tu"
# The movie and user embeddings are also displayed in the right figure. When the embedding dimension is greater than 3, the embeddings are projected on the first 3 dimensions. The next section will have a more detailed look at the embeddings.
# + [markdown] colab_type="text" id="IO1r1A5Xo4tu"
# # Inspecting the Embeddings
#
# In this section, we take a closer look at the learned embeddings, by
# - computing your recommendations
# - looking at the nearest neighbors of some movies,
# - looking at the norms of the movie embeddings,
# - visualizing the embedding in a projected embedding space.
# + [markdown] colab_type="text" id="8Z3q8xZVo4tw"
# ### Exercise 4: Write a function that computes the scores of the candidates
# We start by writing a function that, given a query embedding $u \in \mathbb R^d$ and item embeddings $V \in \mathbb R^{N \times d}$, computes the item scores.
#
# As discussed in the lecture, there are different similarity measures we can use, and these can yield different results. We will compare the following:
# - dot product: the score of item j is $\langle u, V_j \rangle$.
# - cosine: the score of item j is $\frac{\langle u, V_j \rangle}{\|u\|\|V_j\|}$.
#
# Hints:
# - you can use [`np.dot`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html) to compute the product of two np.Arrays.
# - you can use [`np.linalg.norm`](https://docs.scipy.org/doc/numpy-1.14.0/reference/generated/numpy.linalg.norm.html) to compute the norm of a np.Array.
# + cellView="form" colab={} colab_type="code" id="PqDjkdlFo4t0"
DOT = 'dot'
COSINE = 'cosine'
def compute_scores(query_embedding, item_embeddings, measure=DOT):
"""Computes the scores of the candidates given a query.
Args:
query_embedding: a vector of shape [k], representing the query embedding.
item_embeddings: a matrix of shape [N, k], such that row i is the embedding
of item i.
measure: a string specifying the similarity measure to be used. Can be
either DOT or COSINE.
Returns:
scores: a vector of shape [N], such that scores[i] is the score of item i.
"""
u = query_embedding
V = item_embeddings
if measure == COSINE:
V = V / np.linalg.norm(V, axis=1, keepdims=True)
u = u / np.linalg.norm(u)
scores = u.dot(V.T)
return scores
# + [markdown] colab_type="text" id="EDpSY1REo4t1"
# Equipped with this function, we can compute recommendations, where the query embedding can be either a user embedding or a movie embedding.
# + cellView="both" colab={} colab_type="code" id="6dSF9U1fo4t2"
def user_recommendations(model, measure=DOT, exclude_rated=False, k=6):
if USER_RATINGS:
scores = compute_scores(
model.embeddings["user_id"][943], model.embeddings["movie_id"], measure)
score_key = measure + ' score'
df = pd.DataFrame({
score_key: list(scores),
'movie_id': movies['movie_id'],
'titles': movies['title'],
'genres': movies['all_genres'],
})
if exclude_rated:
# remove movies that are already rated
rated_movies = ratings[ratings.user_id == "943"]["movie_id"].values
df = df[df.movie_id.apply(lambda movie_id: movie_id not in rated_movies)]
display.display(df.sort_values([score_key], ascending=False).head(k))
def movie_neighbors(model, title_substring, measure=DOT, k=6):
# Search for movie ids that match the given substring.
ids = movies[movies['title'].str.contains(title_substring)].index.values
titles = movies.iloc[ids]['title'].values
if len(titles) == 0:
raise ValueError("Found no movies with title %s" % title_substring)
print("Nearest neighbors of : %s." % titles[0])
if len(titles) > 1:
print("[Found more than one matching movie. Other candidates: {}]".format(
", ".join(titles[1:])))
movie_id = ids[0]
scores = compute_scores(
model.embeddings["movie_id"][movie_id], model.embeddings["movie_id"],
measure)
score_key = measure + ' score'
df = pd.DataFrame({
score_key: list(scores),
'titles': movies['title'],
'genres': movies['all_genres']
})
display.display(df.sort_values([score_key], ascending=False).head(k))
# + [markdown] colab_type="text" id="fFaiyCtgo4t6"
# ### Movie Nearest neighbors
#
# Let's look at the neareast neighbors for some of the movies.
# + colab={} colab_type="code" id="ejJHfA-so4t8"
movie_neighbors(model, "Aladdin", DOT)
movie_neighbors(model, "Aladdin", COSINE)
# + [markdown] colab_type="text" id="JD3X1ua8o4uA"
# It seems that the quality of learned embeddings may not be very good. Can you think of potential techniques that could be used to improve them? We can start by inspecting the embeddings.
# + [markdown] colab_type="text" id="SOxdxqpKo4uB"
# ## Movie Embedding Norm
#
# We can also observe that the recommendations with dot-product and cosine are different: with dot-product, the model tends to recommend popular movies. This can be explained by the fact that in matrix factorization models, the norm of the embedding is often correlated with popularity (popular movies have a larger norm), which makes it more likely to recommend more popular items. We can confirm this hypothesis by sorting the movies by their embedding norm, as done in the next cell.
# + cellView="form" colab={} colab_type="code" id="G_Kko-YxF6YE"
def movie_embedding_norm(models):
"""Visualizes the norm and number of ratings of the movie embeddings.
Args:
model: A MFModel object.
"""
if not isinstance(models, list):
models = [models]
df = pd.DataFrame({
'title': movies['title'],
'genre': movies['genre'],
'num_ratings': movies_ratings['rating count'],
})
charts = []
brush = alt.selection_interval()
for i, model in enumerate(models):
norm_key = 'norm'+str(i)
df[norm_key] = np.linalg.norm(model.embeddings["movie_id"], axis=1)
nearest = alt.selection(
type='single', encodings=['x', 'y'], on='mouseover', nearest=True,
empty='none')
base = alt.Chart().mark_circle().encode(
x='num_ratings',
y=norm_key,
color=alt.condition(brush, alt.value('#4c78a8'), alt.value('lightgray'))
).properties(
selection=nearest).add_selection(brush)
text = alt.Chart().mark_text(align='center', dx=5, dy=-5).encode(
x='num_ratings', y=norm_key,
text=alt.condition(nearest, 'title', alt.value('')))
charts.append(alt.layer(base, text))
return alt.hconcat(*charts, data=df)
def visualize_movie_embeddings(data, x, y):
nearest = alt.selection(
type='single', encodings=['x', 'y'], on='mouseover', nearest=True,
empty='none')
base = alt.Chart().mark_circle().encode(
x=x,
y=y,
color=alt.condition(genre_filter, "genre", alt.value("whitesmoke")),
).properties(
width=600,
height=600,
selection=nearest)
text = alt.Chart().mark_text(align='left', dx=5, dy=-5).encode(
x=x,
y=y,
text=alt.condition(nearest, 'title', alt.value('')))
return alt.hconcat(alt.layer(base, text), genre_chart, data=data)
def tsne_movie_embeddings(model):
"""Visualizes the movie embeddings, projected using t-SNE with Cosine measure.
Args:
model: A MFModel object.
"""
tsne = sklearn.manifold.TSNE(
n_components=2, perplexity=40, metric='cosine', early_exaggeration=10.0,
init='pca', verbose=True, n_iter=400)
print('Running t-SNE...')
V_proj = tsne.fit_transform(model.embeddings["movie_id"])
movies.loc[:,'x'] = V_proj[:, 0]
movies.loc[:,'y'] = V_proj[:, 1]
return visualize_movie_embeddings(movies, 'x', 'y')
# + colab={} colab_type="code" id="SXP_eW-Xo4uD"
movie_embedding_norm(model)
# + [markdown] colab_type="text" id="5o1vQGyIo4uE"
# Note: Depending on how the model is initialized, you may observe that some niche movies (ones with few ratings) have a high norm, leading to spurious recommendations. This can happen if the embedding of that movie happens to be initialized with a high norm. Then, because the movie has few ratings, it is infrequently updated, and can keep its high norm. This can be alleviated by using regularization.
#
# Try changing the value of the hyperparameter `init_stddev`. One quantity that can be helpful is that the expected norm of a $d$-dimensional vector with entries $\sim \mathcal N(0, \sigma^2)$ is approximatley $\sigma \sqrt d$.
#
# How does this affect the embedding norm distribution, and the ranking of the top-norm movies?
# + colab={} colab_type="code" id="2I0FcjkPAoO4"
model_lowinit = build_model(ratings, embedding_dim=30, init_stddev=0.05)
model_lowinit.train(num_iterations=1000, learning_rate=10.)
movie_neighbors(model_lowinit, "Aladdin", DOT)
movie_neighbors(model_lowinit, "Aladdin", COSINE)
movie_embedding_norm([model, model_lowinit])
# + [markdown] colab_type="text" id="fF6dMP1To4uH"
# ## Embedding visualization
# Since it is hard to visualize embeddings in a higher-dimensional space (when the embedding dimension $k > 3$), one approach is to project the embeddings to a lower dimensional space. T-SNE (T-distributed Stochastic Neighbor Embedding) is an algorithm that projects the embeddings while attempting to preserve their pariwise distances. It can be useful for visualization, but one should use it with care. For more information on using t-SNE, see [How to Use t-SNE Effectively](https://distill.pub/2016/misread-tsne/).
# + colab={} colab_type="code" id="W6bkyeDao4uH"
tsne_movie_embeddings(model_lowinit)
# + [markdown] colab_type="text" id="y9j2Bbf8o4uI"
# You can highlight the embeddings of a given genre by clicking on the genres panel (SHIFT+click to select multiple genres).
#
# We can observe that the embeddings do not seem to have any notable structure, and the embeddings of a given genre are located all over the embedding space. This confirms the poor quality of the learned embeddings. One of the main reasons is that we only trained the model on observed pairs, and without regularization.
# + [markdown] colab_type="text" id="K7NJT9gbo4ub"
# # Softmax model
#
# In this section, we will train a simple softmax model that predicts whether a given user has rated a movie.
#
# The model will take as input a feature vector $x$ representing the list of movies the user has rated. We start from the ratings DataFrame, which we group by user_id.
# + colab={} colab_type="code" id="JJcaNiWdo4ub"
rated_movies = (ratings[["user_id", "movie_id"]]
.groupby("user_id", as_index=False)
.aggregate(lambda x: list(x)))
rated_movies.head()
# + [markdown] colab_type="text" id="X14ABp0Jo4uc"
# We then create a function that generates an example batch, such that each example contains the following features:
# - movie_id: A tensor of strings of the movie ids that the user rated.
# - genre: A tensor of strings of the genres of those movies
# - year: A tensor of strings of the release year.
# + cellView="both" colab={} colab_type="code" id="N_IsDAEAo4ud"
#@title Batch generation code (run this cell)
years_dict = {
movie: year for movie, year in zip(movies["movie_id"], movies["year"])
}
genres_dict = {
movie: genres.split('-')
for movie, genres in zip(movies["movie_id"], movies["all_genres"])
}
def make_batch(ratings, batch_size):
"""Creates a batch of examples.
Args:
ratings: A DataFrame of ratings such that examples["movie_id"] is a list of
movies rated by a user.
batch_size: The batch size.
"""
def pad(x, fill):
return pd.DataFrame.from_dict(x).fillna(fill).values
movie = []
year = []
genre = []
label = []
for movie_ids in ratings["movie_id"].values:
movie.append(movie_ids)
genre.append([x for movie_id in movie_ids for x in genres_dict[movie_id]])
year.append([years_dict[movie_id] for movie_id in movie_ids])
label.append([int(movie_id) for movie_id in movie_ids])
features = {
"movie_id": pad(movie, ""),
"year": pad(year, ""),
"genre": pad(genre, ""),
"label": pad(label, -1)
}
batch = (
tf.data.Dataset.from_tensor_slices(features)
.shuffle(1000)
.repeat()
.batch(batch_size)
.make_one_shot_iterator()
.get_next())
return batch
def select_random(x):
"""Selectes a random elements from each row of x."""
def to_float(x):
return tf.cast(x, tf.float32)
def to_int(x):
return tf.cast(x, tf.int64)
batch_size = tf.shape(x)[0]
rn = tf.range(batch_size)
nnz = to_float(tf.count_nonzero(x >= 0, axis=1))
rnd = tf.random_uniform([batch_size])
ids = tf.stack([to_int(rn), to_int(nnz * rnd)], axis=1)
return to_int(tf.gather_nd(x, ids))
# + [markdown] colab_type="text" id="Yef5q1Qlo4ue"
# ### Loss function
# Recall that the softmax model maps the input features $x$ to a user embedding $\psi(x) \in \mathbb R^d$, where $d$ is the embedding dimension. This vector is then multiplied by a movie embedding matrix $V \in \mathbb R^{m \times d}$ (where $m$ is the number of movies), and the final output of the model is the softmax of the product
# $$
# \hat p(x) = \text{softmax}(\psi(x) V^\top).
# $$
# Given a target label $y$, if we denote by $p = 1_y$ a one-hot encoding of this target label, then the loss is the cross-entropy between $\hat p(x)$ and $p$.
# + [markdown] colab_type="text" id="DDnGAN_0o4ue"
# ### Exercise 5: Write a loss function for the softmax model.
#
# In this exercise, we will write a function that takes tensors representing the user embeddings $\psi(x)$, movie embeddings $V$, target label $y$, and return the cross-entropy loss.
#
# Hint: You can use the function [`tf.nn.sparse_softmax_cross_entropy_with_logits`](https://www.tensorflow.org/api_docs/python/tf/nn/sparse_softmax_cross_entropy_with_logits), which takes `logits` as input, where `logits` refers to the product $\psi(x) V^\top$.
# + cellView="form" colab={} colab_type="code" id="hQNuR42eo4uf"
#Solution
def softmax_loss(user_embeddings, movie_embeddings, labels):
"""Returns the cross-entropy loss of the softmax model.
Args:
user_embeddings: A tensor of shape [batch_size, embedding_dim].
movie_embeddings: A tensor of shape [num_movies, embedding_dim].
labels: A tensor of [batch_size], such that labels[i] is the target label
for example i.
Returns:
The mean cross-entropy loss.
"""
# Verify that the embddings have compatible dimensions
user_emb_dim = user_embeddings.shape[1].value
movie_emb_dim = movie_embeddings.shape[1].value
if user_emb_dim != movie_emb_dim:
raise ValueError(
"The user embedding dimension %d should match the movie embedding "
"dimension % d" % (user_emb_dim, movie_emb_dim))
logits = tf.matmul(user_embeddings, movie_embeddings, transpose_b=True)
loss = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=logits, labels=labels))
return loss
# + [markdown] colab_type="text" id="Y9SvjO4go4ug"
# ### Exercise 6: Build a softmax model, train it, and inspect its embeddings.
#
# We are now ready to build a softmax CFModel. Complete the `build_softmax_model` function in the next cell. The architecture of the model is defined in the function `create_user_embeddings` and illustrated in the figure below. The input embeddings (movie_id, genre and year) are concatenated to form the input layer, then we have hidden layers with dimensions specified by the `hidden_dims` argument. Finally, the last hidden layer is multiplied by the movie embeddings to obtain the logits layer. For the target label, we will use a randomly-sampled movie_id from the list of movies the user rated.
#
# 
#
# Complete the function below by creating the feature columns and embedding columns, then creating the loss tensors both for the train and test sets (using the `softmax_loss` function of the previous exercise).
#
# + cellView="form" colab={} colab_type="code" id="zAAr73xno4uj"
# Solution
def build_softmax_model(rated_movies, embedding_cols, hidden_dims):
"""Builds a Softmax model for MovieLens.
Args:
rated_movies: DataFrame of traing examples.
embedding_cols: A dictionary mapping feature names (string) to embedding
column objects. This will be used in tf.feature_column.input_layer() to
create the input layer.
hidden_dims: int list of the dimensions of the hidden layers.
Returns:
A CFModel object.
"""
def create_network(features):
"""Maps input features dictionary to user embeddings.
Args:
features: A dictionary of input string tensors.
Returns:
outputs: A tensor of shape [batch_size, embedding_dim].
"""
# Create a bag-of-words embedding for each sparse feature.
inputs = tf.feature_column.input_layer(features, embedding_cols)
# Hidden layers.
input_dim = inputs.shape[1].value
for i, output_dim in enumerate(hidden_dims):
w = tf.get_variable(
"hidden%d_w_" % i, shape=[input_dim, output_dim],
initializer=tf.truncated_normal_initializer(
stddev=1./np.sqrt(output_dim))) / 10.
outputs = tf.matmul(inputs, w)
input_dim = output_dim
inputs = outputs
return outputs
train_rated_movies, test_rated_movies = split_dataframe(rated_movies)
train_batch = make_batch(train_rated_movies, 200)
test_batch = make_batch(test_rated_movies, 100)
with tf.variable_scope("model", reuse=False):
# Train
train_user_embeddings = create_network(train_batch)
train_labels = select_random(train_batch["label"])
with tf.variable_scope("model", reuse=True):
# Test
test_user_embeddings = create_network(test_batch)
test_labels = select_random(test_batch["label"])
movie_embeddings = tf.get_variable(
"input_layer/movie_id_embedding/embedding_weights")
test_loss = softmax_loss(
test_user_embeddings, movie_embeddings, test_labels)
train_loss = softmax_loss(
train_user_embeddings, movie_embeddings, train_labels)
_, test_precision_at_10 = tf.metrics.precision_at_k(
labels=test_labels,
predictions=tf.matmul(test_user_embeddings, movie_embeddings, transpose_b=True),
k=10)
metrics = (
{"train_loss": train_loss, "test_loss": test_loss},
{"test_precision_at_10": test_precision_at_10}
)
embeddings = {"movie_id": movie_embeddings}
return CFModel(embeddings, train_loss, metrics)
# + [markdown] colab_type="text" id="VZB1J8l9o4ul"
# ### Train the Softmax model
#
# We are now ready to train the softmax model. You can set the following hyperparameters:
# - learning rate
# - number of iterations. Note: you can run `softmax_model.train()` again to continue training the model from its current state.
# - input embedding dimensions (the `input_dims` argument)
# - number of hidden layers and size of each layer (the `hidden_dims` argument)
#
# Note: since our input features are string-valued (movie_id, genre, and year), we need to map them to integer ids. This is done using [`tf.feature_column.categorical_column_with_vocabulary_list`](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_list), which takes a vocabulary list specifying all the values the feature can take. Then each id is mapped to an embedding vector using [`tf.feature_column.embedding_column`](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column).
#
# + colab={} colab_type="code" id="jJsXAS_go4ul"
# Create feature embedding columns
def make_embedding_col(key, embedding_dim):
categorical_col = tf.feature_column.categorical_column_with_vocabulary_list(
key=key, vocabulary_list=list(set(movies[key].values)), num_oov_buckets=0)
return tf.feature_column.embedding_column(
categorical_column=categorical_col, dimension=embedding_dim,
# default initializer: trancated normal with stddev=1/sqrt(dimension)
combiner='mean')
with tf.Graph().as_default():
softmax_model = build_softmax_model(
rated_movies,
embedding_cols=[
make_embedding_col("movie_id", 35),
make_embedding_col("genre", 3),
make_embedding_col("year", 2),
],
hidden_dims=[35])
softmax_model.train(
learning_rate=8., num_iterations=3000, optimizer=tf.train.AdagradOptimizer)
# + [markdown] colab_type="text" id="-sYjuHxJo4un"
# ### Inspect the embeddings
#
# We can inspect the movie embeddings as we did for the previous models. Note that in this case, the movie embeddings are used at the same time as input embeddings (for the bag of words representation of the user history), and as softmax weights.
# + colab={} colab_type="code" id="mvJazlnMo4uo"
movie_neighbors(softmax_model, "Aladdin", DOT)
movie_neighbors(softmax_model, "Aladdin", COSINE)
# + colab={} colab_type="code" id="ReIP69-9o4ur"
movie_embedding_norm(softmax_model)
# + colab={} colab_type="code" id="Trq0yYrXo4us"
tsne_movie_embeddings(softmax_model)
# + [markdown] colab_type="text" id="fZkZJ8yco4uu"
# ## Congratulations!
#
# You have completed this lab.
#
# If you would like to further explore these models, we encourage you to try different hyperparameters and observe how this affects the quality of the model and the structure of the embedding space. Here are some suggestions:
# - Change the embedding dimension.
# - In the softmax model: change the number of hidden layers, and the input features. For example, you can try a model with no hidden layers, and only the movie ids as inputs.
# - Using other similarity measures: In this notebook, we used dot product $d(u, V_j) = \langle u, V_j \rangle$ and cosine $d(u, V_j) = \frac{\langle u, V_j \rangle}{\|u\|\|V_j\|}$, and discussed how the norms of the embeddings affect the recommendations. You can also try other variants which apply a transformation to the norm, for example $d(u, V_j) = \frac{\langle u, V_j \rangle}{\|V_j\|^\alpha}$.
# -
# ## Challenge
#
# With everything you learned during the Advanced Machine Learning on Google Cloud, can you try and push the model to the AI Platform for predictions?
# Copyright 2021 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/10_recommend/cf_softmax_model/solution/cfmodel_softmax_model_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Map and Lambda Function
cube = lambda x: x*x*x
# complete the lambda function
def fibonacci(n):
if n == 0:
l = []
return l
# return a list of fibonacci numbers
if n==2:
l = [0,1]
return l
# Second Fibonacci number is 1
elif n==1:
l = [0]
return l
else:
fibonacci_numbers = [0, 1]
for i in range(2,n):
fibonacci_numbers.append(fibonacci_numbers[i-1]+fibonacci_numbers[i-2])
return fibonacci_numbers
if __name__ == '__main__':
n = int(input())
print(list(map(cube, fibonacci(n))))
| Problem1/Scripts/Python Functionals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from lagnn.core import *
# # Lagrangian Neural Networks
#
# > Re-implementation of [Lagrangian Neural Networks](https://arxiv.org/abs/2003.04630) by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>
# ## Getting started
#
# Spin up a conda virtual environment and install the required libraries:
#
# ```bash
# conda create -n lagnn python=3.9
# pip install .
# ```
#
#
| notebooks/index.ipynb |